
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Sustain. Food Syst. , 27 March 2025
Sec. Agroecology and Ecosystem Services
Volume 9 - 2025 | https://doi.org/10.3389/fsufs.2025.1520713
This article is part of the Research Topic Innovative Approaches in Soil, Water, and Crop Management for Sustainable Agricultural Systems View all 7 articles
With the growing focus on agroecological interventions in crop production, more studies are required to understand the combined effects of multiple strategies on soil and plant health. In this study, we adopted a functional-trait approach to evaluate the resource acquisition strategies of crops in response to an organic amendment and a growth-promoting endophytic bacterial seed inoculant. Using wheat and soybean as model crops, we assessed crop performance traits [leaf chlorophyll (μmol m−2), base diameter (mm), biomass (g), nitrogen (N) usage index] and root traits [specific root length (SRL, m g−1), specific root tip density (SRTD, tips g−1), root diameter (root D, mm), root tissue density (root TD, g cm−3), root nitrogen (root N; %), root carbon-to-nitrogen ratios (root C/N)], along with the chemical and biological properties of rhizosphere soil at the end of the growing season. We employed three soil amendment treatments (organic, inorganic, and control) and two seed inoculant treatments (with and without seed inoculation). Soil amendments influenced the expression of wheat root traits, promoting acquisitive root traits—characterized by higher SRL, SRTD, and root N, along with lower root D—under organic amendments. There was no impact of soil amendments on soybean root trait expression. In contrast, seed inoculation positively affected the performance traits of base diameter and the nitrogen usage index of wheat. The significant interactive effects of soil amendments and seed inoculation were minimal. In the rhizosphere soil of wheat and soybeans, available phosphorus was significantly lower with organic amendments, while active carbon (POxC) was significantly higher with organic amendments for wheat. Well-established significant relationships between soil properties and root traits were also evident. Correlations between root traits and rhizosphere properties were apparent, specifically positive correlations between POxC and SRL, and negative correlations with the root C/N ratio. The composition of the rhizosphere soil fungal community was significantly explained by key root traits, including root D and root C/N ratio. Broadly, soil amendments had a stronger impact on crop root responses than seed inoculants, yet seed inoculants influenced direct measures of crop performance. Importantly, these two agroecological interventions did not significantly interact, indicating opportunities for better integration of agroecological strategies.
Declining soil health, coupled with the increasing prevalence of various abiotic and biotic stresses due to global climate change, poses significant challenges for the agricultural sector (Fiodor et al., 2021; Hamidov et al., 2018). Consequently, farmers increasingly require cost-effective integrative solutions to address both soil and plant health issues. Agroecological interventions that incorporate ecological principles into farming practices, such as agroforestry (Isaac et al., 2024), organic nutrient management (Chen et al., 2018), and the use of beneficial microorganisms (Batista and Singh, 2021), offer strategies that mitigate the adverse effects of synthetic fertilizer usage (Altieri, 2019; Isaac et al., 2018).
Organic amendments, unlike inorganic fertilizers, produce distinct nutrient release patterns and influence plant nutrient uptake. Inorganic fertilizers are characterized by large pulses of nutrients that are quickly available for uptake; however, indiscriminate application can lead to soil nutrient imbalances and losses (Drinkwater and Snapp, 2007). Through microbially mediated processes, organic inputs can maintain slow but steady nutrient supplies. Organic amendments, such as vermicompost, are also effective in promoting and sustaining long-term soil health (Chen et al., 2018; Lal, 2015). While inorganic fertilizers supply specific macronutrients, organic amendments provide diverse sources of carbon and organic forms of nutrients that stimulate soil microorganisms and induce changes in soil organic matter, structure, and water-holding capacity (Diacono and Montemurro, 2010; Malik et al., 2021). However, the improvements in soil health resulting from organic inputs do not always lead to increases in crop productivity (Schrama et al., 2018), and concerns regarding yield limitations due to slower nutrient turnover rates have, in part, hindered the widespread adoption of organic amendments (Röös et al., 2018).
Recent advances in microbial interventions indicate that plant growth-promoting bacteria (PGPB) and bioinoculants can reduce reliance on off-farm inputs, such as synthetic fertilizers and pesticides (Elnahal et al., 2022) while providing a promising solution to enhance plant productivity and health, thereby addressing yield concerns (Negi et al., 2021; Sessitsch and Mitter, 2015; Timmusk et al., 2017). Bacteria capable of colonizing host plant tissues without causing negative impacts are termed endophytic; ecologically, these bacteria have the potential to play a significant role as an agroecological intervention. Growth-promoting microorganisms inhabit areas within or around plant tissues and facilitate plant growth and responses to abiotic and biotic stresses through several mechanisms, including the production of 1-aminocyclopropane-1-carboxylate (ACC)-deaminase (Khan et al., 2016), various phytohormones (Sharma et al., 2023), volatile organic compounds (Almeida et al., 2023), siderophores (Loaces et al., 2011), and antibiotics (Narayanan and Glick, 2022). This contributes to the modulation of processes such as photosynthesis, pathogen resistance, and nutrient uptake.
One of the most recognized benefits of PGPB is its ability to mediate plant ethylene levels through the enzyme ACC-deaminase (Kang et al., 2019; Khan et al., 2016; Maxton et al., 2018). Ethylene, a stress-responsive hormone, typically elevates in plants undergoing various stressors, including injury, metal contamination, and extremes of soil salinity and water content (Orozco-Mosqueda et al., 2020). ACC-deaminase-producing bacteria help lower plant ethylene levels and thus promote growth by mitigating the plant stress response. Plantibacter flavus strains have been reported as endophytic inhabitants of wheat tissues (Robinson et al., 2016; Muvingi et al., 2023; Alikhani and Emami, 2020) and in the rhizosphere and rhizoplane of wheat (Braun-Kiewnick et al., 2024; Alikhani and Emami, 2020), indicating compatibility with wheat and the potential for optimizing inoculation procedures. However, the plant’s response to inoculating different PGPB can vary depending on host genotype and physiology (Elnahal et al., 2022; Mayer et al., 2019; Sessitsch and Mitter, 2015).
Beyond yield measurements, strong evidence indicates that functional traits—the morphological, chemical, and physiological components of plants that affect and respond to environmental factors (Bardgett et al., 2014; Reich, 2014; Violle et al., 2007; Martin and Isaac, 2015)—can be used as indicators of plant plasticity and trait trade-offs in relation to above- and belowground conditions (Gagliardi et al., 2015; Isaac and Borden, 2019; Freschet et al., 2021a; Yaffar et al., 2022), including those resulting from management regimes and interventions, such as organic amendments (Nimmo et al., 2023). Variations in plant functional traits are associated with agroecosystem functioning, including soil carbon dynamics (Buchanan et al., 2022), soil greenhouse gas fluxes (Buchanan et al., 2024), and yield (Martin and Isaac, 2021). Furthermore, plant traits may moderate positive soil outcomes from agroecological interventions (Rolhauser and Isaac, 2024). Morphological traits, such as high specific leaf area (SLA), and chemical traits, like increased leaf nitrogen content (leaf N), are generally classified as resource acquisitive traits, characterized by a fast-growing strategy and enhanced resource foraging capacity. In contrast, the opposite traits are defined by a slower-growing strategy and greater resource conservation capacity (Violle et al., 2007). While leaf functional trait trade-offs align with this universal spectrum of resource acquisition to resource conservation (Wright et al., 2004), root functional traits and resource acquisition present greater complexity, reacting to multiple soil nutrient and microbial characteristics and forming various root economic spectra (Bergmann et al., 2020). Root systems regulate metabolic construction costs, rhizosphere properties, and interactions with biota (Lynch et al., 2021). Morphological root traits can be classified as resource acquisitive, e.g., high root nitrogen content (root N) and high specific root tip densities (SRTD), or as resource conservative, e.g., high root C/N ratio and high root tissue density (root TD) (Isaac et al., 2021). They can also display do-it-yourself strategies, e.g., increased foraging through high specific root length (SRL) coupled with low root diameter (root D), or outsourcing strategies via fungal partnerships, e.g., enhanced fungal microbial symbiosis characterized by high root D paired with low SRL (Bergmann et al., 2020). Such root functional traits facilitate changes in rhizosphere nutrient availability and microbial community composition through the production and secretion of exudates such as amino acids, organic acids, sugars, and extracellular enzymes, such as phosphatases (Bargaz et al., 2017; Canarini et al., 2019; Han et al., 2022). Root acquisition strategies play a large role in inducing whole-plant fitness by modifying nutrient uptake and affecting key soil biological properties, including enzymatic activity and microbial communities (Fulthorpe et al., 2020; Spitzer et al., 2021).
In this study, we measured root functional trait responses, aboveground performance, and the chemical and biological properties of the rhizosphere in two important field crops, Triticum aestivum (wheat) and Glycine max (soybean), to examine the individual and combined effects of organic amendments and a growth-promoting endophyte bacterial seed inoculant. We hypothesized that the combination of the organic amendment and growth-promoting bacteria would influence root traits and shape resource acquisition strategies. Additionally, we hypothesized that root traits would affect rhizosphere properties, including fungal community composition, acid phosphatase activity (Apase), rhizosphere labile carbon, total N, root C/N ratio, and available phosphorus. Finally, we measured crop performance through aboveground biomass, base diameter, leaf chlorophyll content, and the Nitrogen Usage Index to assess the impact of seed inoculation and the combined treatment on aboveground plant performance traits. While research on the use of organic amendments and plant growth-promoting bacteria (PGPB) in sustainable agriculture is expanding, our understanding of the combined effects of multiple agroecological interventions and the mechanisms underlying interactions in these complex plant–soil-microbe matrices remains limited.
This study was conducted in experimental plots at the University of Toronto Scarborough Campus Farm (43.793, −79.186). The site had been left fallow for several years before being repurposed for educational research. The year prior to this study, the plots were used to grow soybeans and wheat. In May 2021, nine blocks measuring 5 m × 5 m were tilled. Three blocks received inorganic fertilizer, three were treated with vermicompost (worm castings containing approximately 32% carbon and approximately 2% nitrogen) as an organic amendment, and three were left unamended. Inorganic fertilizers and organic amendments were added at a rate of 75 kg N ha−1 and mixed into the top 2 inches of soil.
Spring wheat (variety AAC Brandon) and soybean were planted in 75 cm × 75 cm plots within the blocks. Wheat was sown in three rows per plot, spaced 20 cm apart, with a seeding density of 69 seeds per plot. Soybean was sown in three rows per plot, also spaced 20 cm apart, with a seeding density of 42 seeds per plot.
One hour prior to planting, half of the seeds for both species were inoculated with a novel growth-promoting bacterial endophyte (Lumactud et al., 2016; Lumactud et al., 2017; Lumactud and Fulthorpe, 2018; Mayer et al., 2019). The inoculum was prepared using the Plantibacter flavus strain 251, along with a liquid formulation containing 2 g xanthan gum, 50 ml glycerol, 1 L distilled water, and 4 g polyvinyl alcohol. This formulation was sterilized using an autoclave before the bacteria were introduced.
The bacterial cultures were grown for 48 hours at 30°C at 250 rpm in 30 ml of R2A liquid broth in 50 mL falcon tubes. Cell densities were enumerated using optical densities at 600 nm calibrated with plate counts. Seeds were coated with 4% of their weight in the bacterial formulation, supplemented with 1 × 1010 cells per seed. Control seeds were coated with the liquid formulation without bacteria at the same 4% seed weight ratio. All seed coatings were applied inside a biological safety hood in a sealed plastic bag 1 h before planting in the test plots.
Six plots of inoculated and uninoculated wheat and soybean were established in each amendment treatment, resulting in a total of 36 plots per species. The seed inoculation treatments (uninoculated/inoculated) were nested within each amendment treatment (inorganic fertilizer, organic amendment, no amendment). One soybean plot (inorganic fertilizer, inoculated) exhibited low germination and was removed from the dataset, reducing the total number of soybean plots from n = 36 to n = 35.
Soybean and wheat were sampled in late August 2021, at 70 days post-germination, with wheat at Feekes stage 11 (head ripening) and soybean at stage R6. We selected three plants per plot for analysis based on location (not along the edge of the plot), representativeness (average size within the plot), and health (not showing signs of disease or pests). For each of the three plants per plot, we measured base diameter (mm) using electronic calipers. We also assessed leaf chlorophyll content (μmol m−2) utilizing a portable chlorophyll meter (MC-100, Apogee Instruments Inc., Utah, USA) in accordance with the leaf chlorophyll measuring protocols established by Hayes et al. (2019), as this is a crucial leaf trait reflecting crop health. Aboveground biomass (g) was calculated from one whole plant sample per plot after drying at 60°C for 48 h. Nitrogen content was then scaled to the dry aboveground biomass, and a nitrogen usage index, as defined by Siddiqi and Glass (1981), was calculated as (aboveground biomass) * (aboveground biomass/nitrogen content of aboveground biomass).
One whole plant root per species per plot was analyzed for root traits following protocols adapted from Martin et al. (2019) and Isaac et al. (2017). Root samples were scanned using a flatbed scanner and analyzed with WinRHIZO imaging software (Reagent Instruments Inc., Quebec) to obtain average root D (mm), total root volume, total root length, and total number of root tips. The root samples were dried at 60°C for 48 h and weighed to determine dry root mass (g). From the root dry mass, total root TD (g cm−3) was calculated as the dry root mass divided by the total fresh root volume per diameter class (Freschet et al., 2021b). SRL (m g−1) was calculated as the total root length per dry mass of root, while SRTD (the number of tips g−1) was calculated from the number of tips per dry mass of root. Root chemical traits, including root N and root C (%), as well as root C/N, were determined from the dried root samples using a LECO elemental analyzer.
In the field, a root from each whole plant sample was gently shaken, and the soil that remained tightly adhered was collected and divided into three subsamples. An air-dried portion was used to analyze inorganic phosphorus (P). These air-dried samples were shaken with a Bray’s extracting solution and filtered into vials using P5 filter paper. The samples were analyzed for inorganic P (mg l−1) using a Lachat QuikChem 8,500 Series 2 Flow Injection Analyzer (Lachat Instruments, Loveland, CO, USA). An oven-dried subsample was used to analyze total nitrogen (N) and carbon (C). To quantify total soil N (%) and C (%) on a mass basis, a subsample of the soil was dried at 105°C for 72 h. The sample was then ground and analyzed in the same manner as the root samples using a LECO elemental analyzer. A subsample was placed in sterile vials and stored at −20°C (Munroe et al., 2015) for molecular analyses. Soil pH was determined with a pH meter on air-dried soil using a 1:2.5 soil-water ratio (FAO, 2021). Soil pH averaged [7.65 ± 0.30].
Permanganate oxidizable carbon (POxC-mg kg−1 soil), or active soil C, in rhizosphere soils was measured following the protocols outlined in Weil et al. (2003). Briefly, a mass of 2.5 g of air-dried rhizosphere soil was diluted with 18 ml of deionized water, and 2.0 ml of 0.2 M KMnO4 was added. The tubes were shaken for 2 min at 240 oscillations per minute. After the tubes were shaken, they were left to settle for 10 min in the dark. A volume of 0.5 ml of the supernatant was diluted with 49.5 ml of deionized water, and the absorbance was recorded using a UV–VIS spectrophotometer at 550 nm. POxC was calculated as the quantity of potassium permanganate reduced in each soil sample. Therefore, the lower the absorbance reading, the higher the POxC values. Final POxC values were calculated following Weil et al. (2003).
The soil enzyme Apase was measured according to Bargaz et al. (2012). A mass of 62.5 mg of fresh rhizosphere soil was mixed with 500 μl of 0.2 M sodium acetate buffer (pH < 6.5) and 125 μl of disodium p-nitrophenyl phosphate (p-NPP, 10 mM). The samples were incubated at 37°C for 1 h. After incubation, the reaction was stopped by adding 125 μl of CaCl2 (0.5 M), followed by 500 μl of NaOH (0.5 M). The samples were centrifuged at 20,000 g for 5 min, and the supernatant was analyzed using a UV–VIS spectrophotometer at 405 nm. Apase activity was expressed as the amount of p-nitrophenol (PNP) produced by the soil from the p-nitrophenyl phosphate substrate per gram of soil per unit of time (μM PNP/g soil/min).
Samples of rhizosphere soil were collected and immediately frozen at −20 °C until DNA extraction via DNAeasy PowerSoil Pro kits (Qiagen). DNA quality and concentrations were confirmed using the Qubit dsDNA HS Assay Kit (Thermo Fisher Scientific Inc.). Amplification and sequencing of fungal ITS fragments was performed by Metagenom Bio (Waterloo, Ontario, Canada) using primers BITS: 5′ ACCTGCGGARGGATCA and B58S3: 5′ – GAGATCCRTTGYTRAAAGTT (Bokulich and Mills, 2013). Demultiplexed sequences were processed using DADA2 v1.8 (Callahan et al., 2016) managed through QIIME 2 v.2019.7 (https://qiime2.org/, Caporaso et al., 2010). Both forward and reverse reads were truncated at decreasing quality (225 forward and 200 reverse), and paired reads were assembled after modeling and correcting for Illumina sequencing errors. Subsequently, chimeric amplified sequence variants (ASVs) were removed by reconstruction against more abundant parent ASVs. An ASV table was then constructed for downstream analysis. Taxonomy was assigned to representative sequences using a naive Bayesian classifier implemented in QIIME 2 with scikit-learn (v.0.21.3) trained against UNITEdb v.8, which is dynamically clustered for fungal ITS. Assignments were accepted above a 0.7 confidence threshold. Raw reads were deposited in the NCBI Short Read Archive (PRJNA1216962).
All statistical analyses were conducted using RStudio version 2021.09.1 + 372 (R Foundation for Statistical Computing, Vienna, Austria). Data were assessed for outliers, while residuals were assessed for normality, and all data underwent tests for homoscedasticity. Traits that were not normally distributed were log-transformed prior to analysis. To examine differences among soil amendment treatments, seed inoculation treatments, and their interactions regarding the measured plant traits, a two-way analysis of variance (ANOVA), followed by Tukey’s honestly significant difference (HSD) post hoc test, was performed. The impacts of the treatments on root trait variation were assessed by calculating coefficients of variation (CVs) as the ratio of the standard deviation to the mean for each root trait, grouped separately by soil amendment and seed inoculation treatments. Root functional traits were analyzed using principal component analysis (PCA) to identify multivariate patterns of root trait variation among the treatments. To determine whether the PCA scores significantly differed between the amendment treatments, the inoculation treatment, and their interaction, a multivariate permutational analysis of variance (PerMANOVA) was conducted using the “vegan” package.
To test for differences among soil amendment treatments on Apase, POxC, rhizosphere total N, total C, C/N ratio, and rhizosphere available P, a one-way analysis of variance (ANOVA) followed by Tukey’s honestly significant difference (HSD) post hoc test was employed. To assess the relationship between root traits and the measured rhizosphere properties, Pearson correlation coefficient matrices were computed. To examine the characteristics of the rhizosphere fungal community, fungal ASV abundance data were converted to percentages of total reads for each species pool, grouped by crop (analyzed separately). The most numerically abundant strains were ordered, retaining those within the top 30 ranked abundances, along with three of the most abundant AMF species (Fulthorpe et al., 2020). This resulted in 32 fungal genera being retained for both wheat and soybean. Nonmetric multidimensional scaling (NMDS) from the ‘vegan’ package was employed to determine whether fungal communities significantly differed between the amendment regimes, the inoculation treatment, and the interaction between the two, utilizing Bray–Curtis dissimilarity values transformed using a Wisconsin double standardization. The results of the NMDS were analyzed with a PerMANOVA via the adonis function. To evaluate whether fungal communities were significantly predicted by root functional traits, redundancy analyses (RDA) were conducted using Hellinger-transformed phylotype abundance data. The function “ordiR2step” with forward selection was used to reduce the number of root traits. The significance of each variable was assessed using a PerMANOVA implemented with the ‘anova.cca’ function in the ‘vegan’ R package (with 9,999 permutations employed).
There was a significant amendment effect on wheat root traits (Table 1). Wheat SRL (p = 0.007) was highest in the organic treatment (17.31 ± 9.84 m g1; Table 2), significantly higher than in the inorganic fertilizer treatment (9.75 ± 2.21 m g1). However, the no amendment wheat (11.73 ± 2.95 m g1) did not significantly differ from either the organic or inorganic fertilizer treatments. Wheat SRTD (p = 0.002) was also highest in the organic treatment (4857.4 ± 2057.6 # of tips g 1), which was significantly different from the inorganic fertilizer treatment (3005.3 ± 543.8 # of tips g1) and the no amendment treatment (3170.9 ± 685.2 # of tips g1). Wheat root D (p = 0.0002) was lowest in the organic treatment (0.46 ± 0.04 mm), which significantly differed from both the inorganic fertilizer treatment (0.57 ± 0.06 mm) and the no amendment treatment (0.54 ± 0.04 mm). Wheat root N content was also significantly affected by the amendment treatment (p = 0.049). Although the organic treatment (0.96 ± 0.28%) did not significantly differ from the inorganic fertilizer (0.82 ± 0.16%) or the no amendment treatment (1.03 ± 0.22%), the N content was significantly lower in the inorganic fertilizer treatment compared to the no amendment treatment. The inoculation treatment and the interaction between the amendment and the inoculation did not significantly affect wheat or soybean root traits. Additionally, there were no significant effects of either the amendment or the inoculation detected on soybean root traits.

Table 1. Results of a two-way analysis of variance for both root traits and performance traits of wheat and soybean for amendments, inoculation, and their interactions.
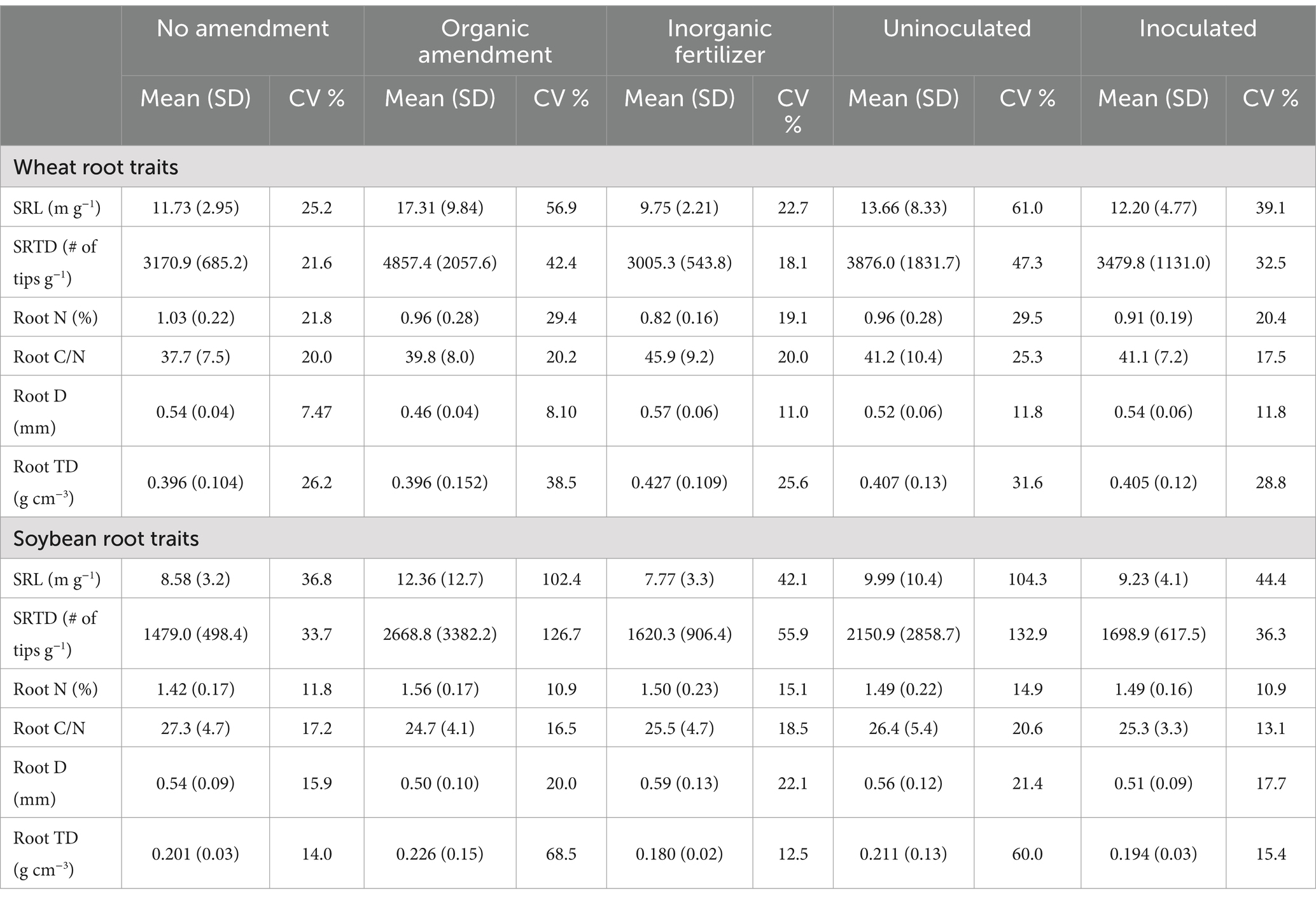
Table 2. Means, standard deviation (SD), and CV of wheat and soybean root functional traits [SRL; SRTD; root N; root C: N; root D; root TD] across three amendment treatments and two inoculation treatments.
The organic amendment resulted in higher CVs for wheat SRL, SRTD, TD, and root N, as well as for soybean SRL, SRTD, and TD, compared to root trait CVs under inorganic fertilizer (Table 2). This finding suggests low trait plasticity when crops are cultivated with inorganic fertilizers. Notably, across all root traits except for root D in both wheat and soybeans, the inoculation treatment resulted in lower trait variation than the uninoculated treatment. In the inoculated treatment, the CVs for the root traits ranged from 11.8 to 39.1% for wheat and 13.1 to 44.4% for soybean. Conversely, uninoculated plants exhibited CVs ranging from 11.8 to 61.0% for wheat and 14.9 to 104.3% for soybean.
The effects of the amendment treatment were observed on performance traits (Table 1). Wheat base diameter (p = 0.014) and nitrogen usage index (p = 0.050) demonstrated significant differences among inoculation treatments. Inoculation with the growth-promoting bacterial inoculant led to a notable increase of 28.5% in mean wheat base diameters and a 38.7% rise in the mean nitrogen usage index (Table 3). The mean leaf chlorophyll content was 45.9% higher in the inorganic fertilizer treatment compared to the no amendment treatment (p = 0.005). The mean base diameter was 32.1% larger in the inorganic fertilizer treatment than in the organic treatment (p = 0.014).
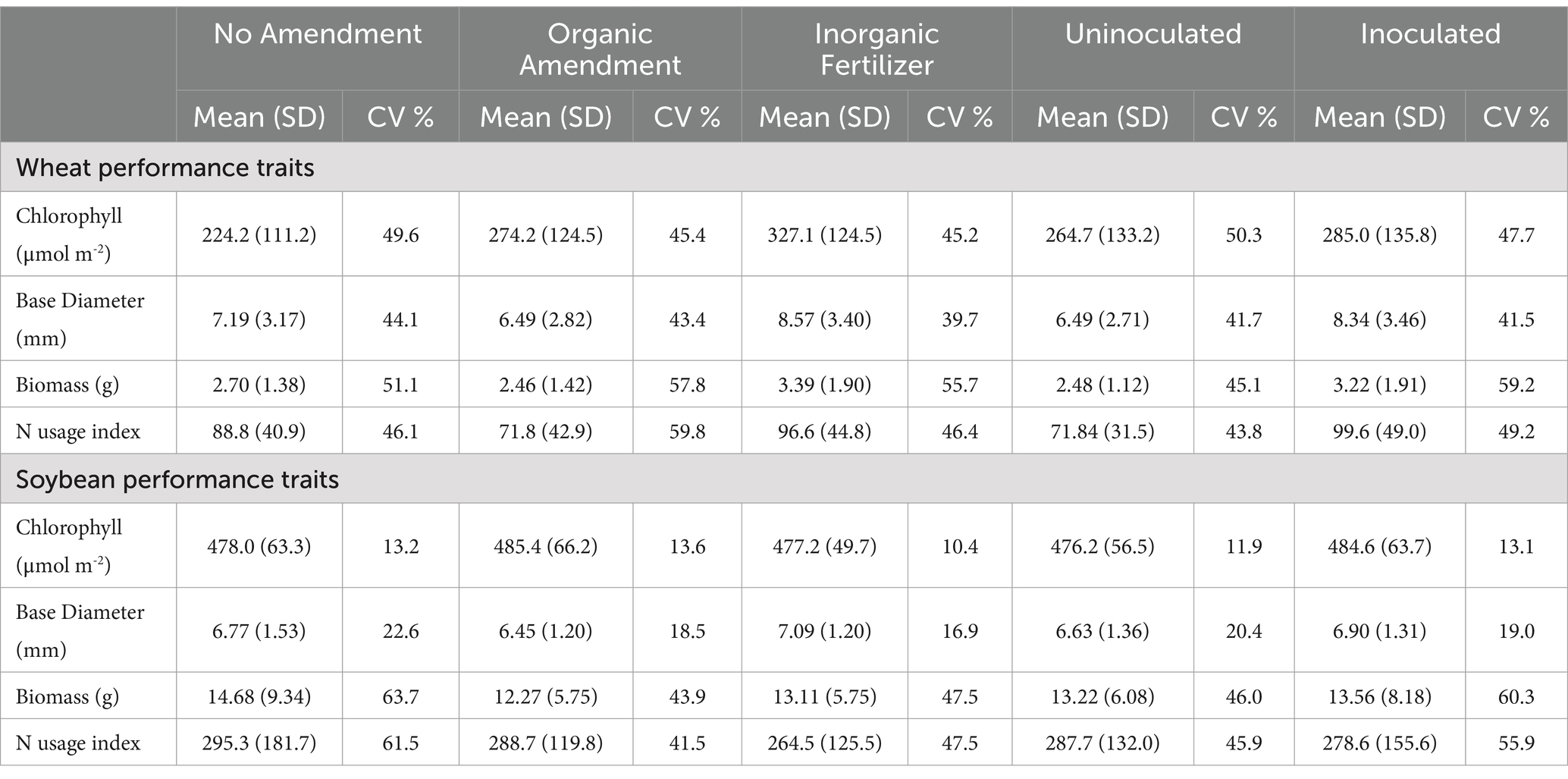
Table 3. Means, standard deviation (SD), and the CV of wheat and soybean performance traits across three amendment treatments and two inoculation treatments.
To measure multivariate root trait variation, PCAs were conducted for both crops. In wheat, the first two PCA axes accounted for a total of 75.7% of the variation in root functional traits (Figure 1). PCA axis 1 accounted for 44.8% of the variation in wheat root traits and was positively associated with conservative traits, root D, root C/N, and root TD, while being negatively associated with root acquisitive traits such as SRL, SRTD, and root N. PCA axis 2 accounted for an additional 30.9% of the variation in wheat root traits and was positively associated with root C/N, SRL, and SRTD, while negatively associated with root N and root TD.
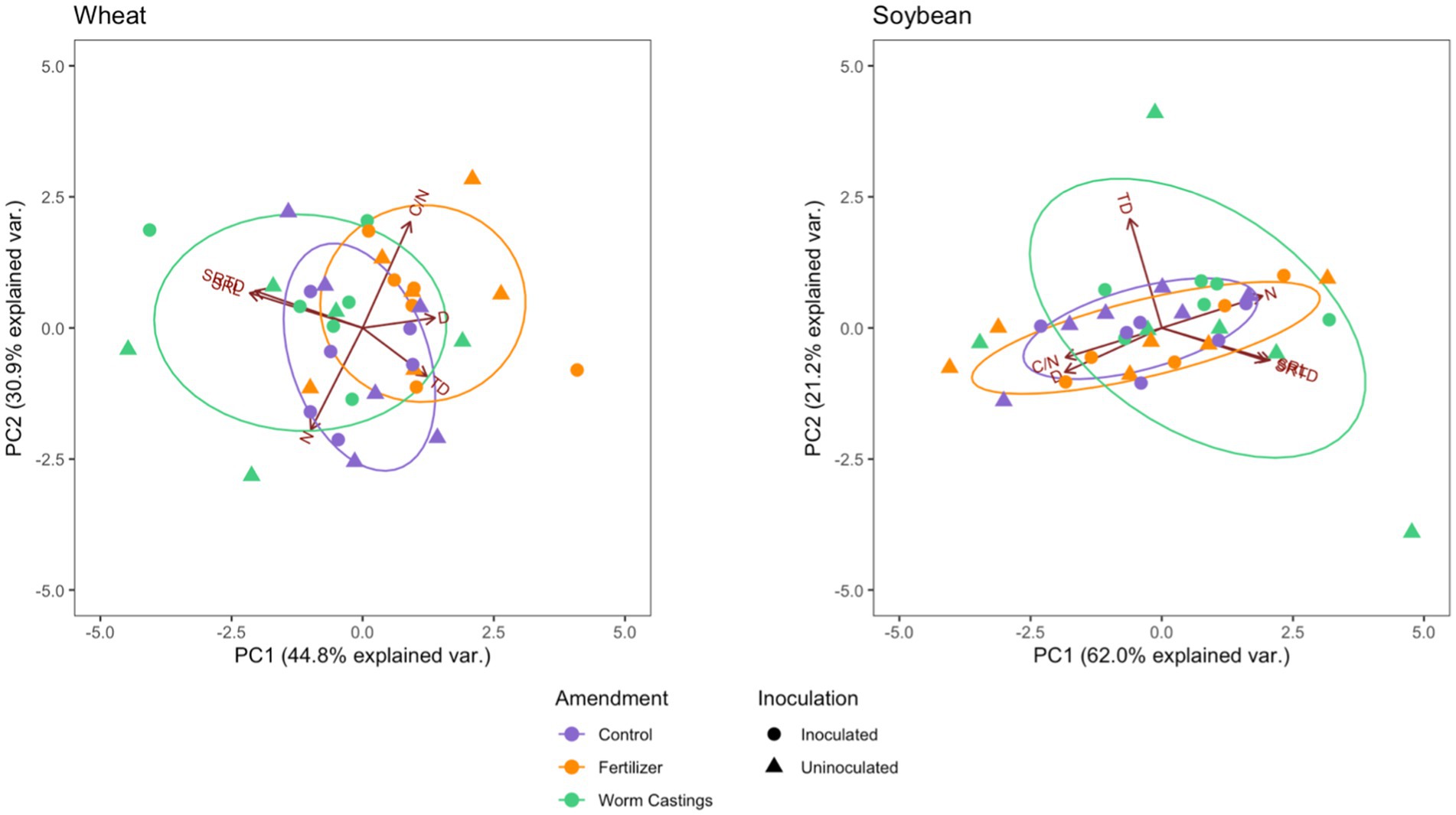
Figure 1. Principal component analysis (PCA) depicting the relationships among the six measured root functional traits: SRL, SRTD, root N, root C:N ratio, root D, and root TD in multivariate space for wheat and soybean. The shapes represent the two inoculation treatments, while the ellipses denote the three amendment treatments. Sample size n = 35.
For soybeans, the first two PCA axes explained a total of 83.2% of the variation in root functional traits (Figure 1). Similar to wheat, the first PCA axis accounted for the majority of the variation in soybean root traits (62.0%) and was positively associated with root acquisitive traits, root (N, SRL, and SRTD) while negatively associated with conservative traits, root D, root TD, and root C/N. The second PCA axis explained an additional 21.2% of the variation in soybean root traits and was negatively associated with root D, root C/N, SRL, and SRTD, while positively associated with root N and root TD.
PerMANOVA indicated that wheat multivariate traits varied significantly as a function of the amendment only, with a pairwise test indicating that the organic amendment and the inorganic fertilizer treatment significantly varied from each other (p = 0.021). Soybean multivariate traits did not vary significantly as a function of any of the treatments.
The mean and standard error for rhizosphere Apase, POxC, available P, total N, total C, and C/N ratio are shown in Table 4. Significant differences were found in the available P of wheat rhizosphere soil among the amendment treatments (p = 0.017), with notably higher values observed in the inorganic fertilizer treatment (24.8 ± 3.6 mg kg−1) compared to the organic treatment (5.9 ± 1.3 mg kg−1). Additionally, significant differences were noted in the POxC of wheat rhizosphere soil (p = 0.028), which was significantly lower in the inorganic fertilizer treatment (584.0 ± 20.5 mg C kg−1) compared to the organic treatment (761.3 ± 26.7 mg C kg−1). No significant differences were detected for wheat rhizosphere N, C, or C/N ratio, nor for Apase.
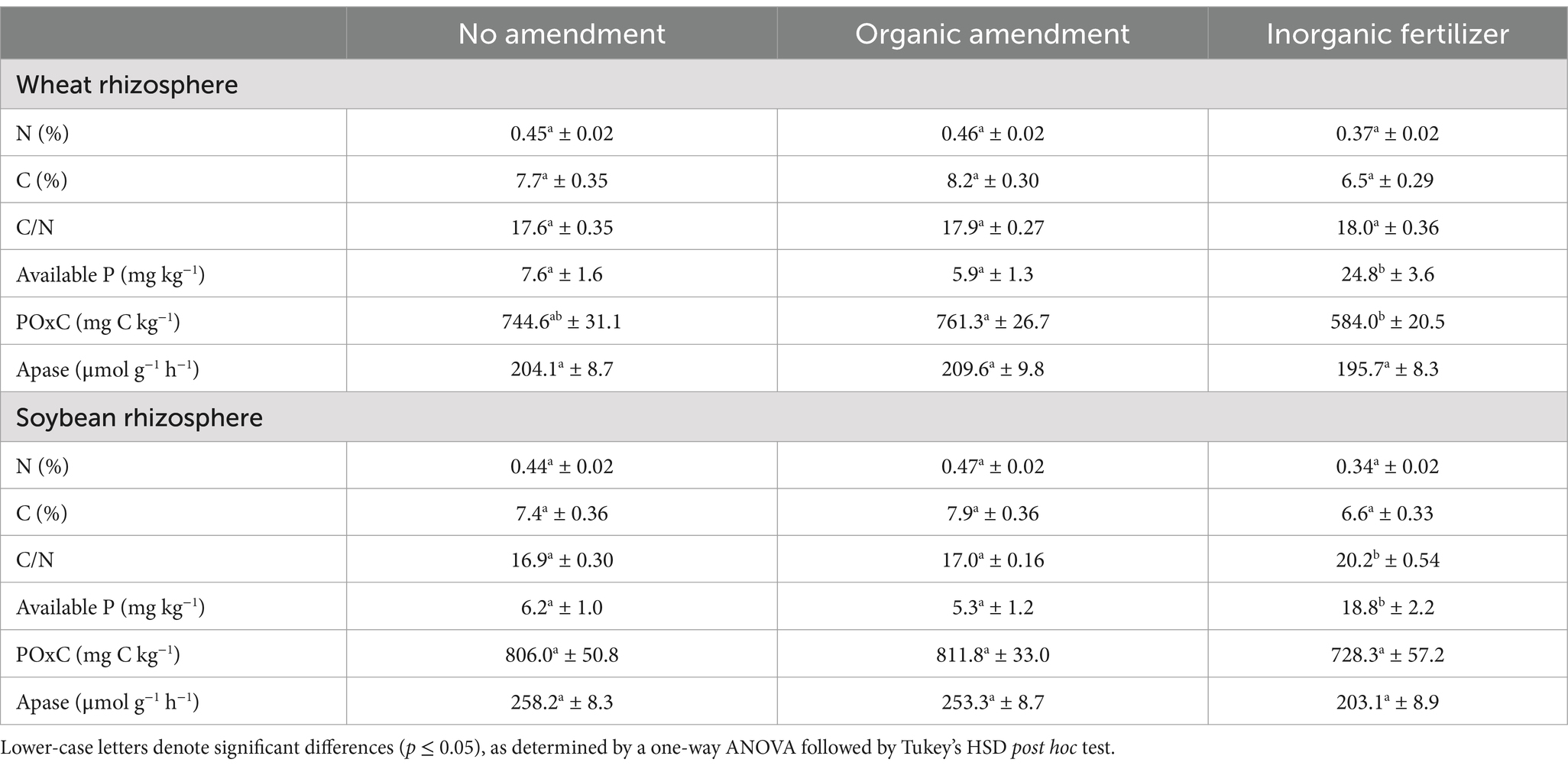
Table 4. Rhizosphere soil properties by amendment treatment with means and standard error (SE) for total N, total C, C/N ratio, available P, active C (POxC), and Apase activity.
Significant differences were also observed in the available P of the soybean rhizosphere in response to the amendment treatment (p = 0.007). Higher values were detected in the inorganic fertilizer treatment (18.8 ± 2.2 mg kg−1) compared to the organic treatment (5.3 ± 1.2 mg kg−1) and the no amendment treatment (6.2 ± 1.0 mg kg−1). The C/N ratio of the soybean rhizosphere was significantly influenced by the amendment treatment (p = 0.001), being higher in the inorganic fertilizer treatment (20.2 ± 0.54) relative to the organic treatment (17.0 ± 0.16) and the no amendment treatment (16.9 ± 0.30). Additionally, alkaline phosphatase (Apase) activity in the soybean rhizosphere was significantly affected by the amendment treatment (p = 0.039). Apase levels were lower in the inorganic fertilizer treatment (203.1 ± 8.9 μmol g−1 h−1) compared to the organic amendment (253.3 ± 8.7 μmol g−1 h−1). No significant differences were found for soybean rhizosphere N, C, or POxC.
The results of a redundancy analysis (RDA) indicated that wheat root functional traits explain fungal community composition, whereas soybean root functional traits do not (Table 5). Forward selection applied to the wheat RDA identified root D (p = 0.018) and root C/N ratio (p = 0.010) as the most significant traits for explaining fungal community composition, yielding a final model of R2 = 0.233 (p = 0.002). Rhizosphere fungal analysis revealed that the most abundant fungal genera across all measured wheat samples were (excluding unidentified genera) Mortierella (38.28%), Pyrenochaetopsis (4.62%), and Alternaria (2.72%). The most abundant genera across all soybean samples were (excluding unidentified genera) Mortierella (42.8%), Pyrenochaetopsis (4.90%), and Fusarium (1.92%). An NMDS analysis showed that fungal community composition did not significantly vary between the soil amendment treatments, the seed inoculation treatment, or the interaction between the soil amendment and seed inoculation treatments for both wheat and soybean.
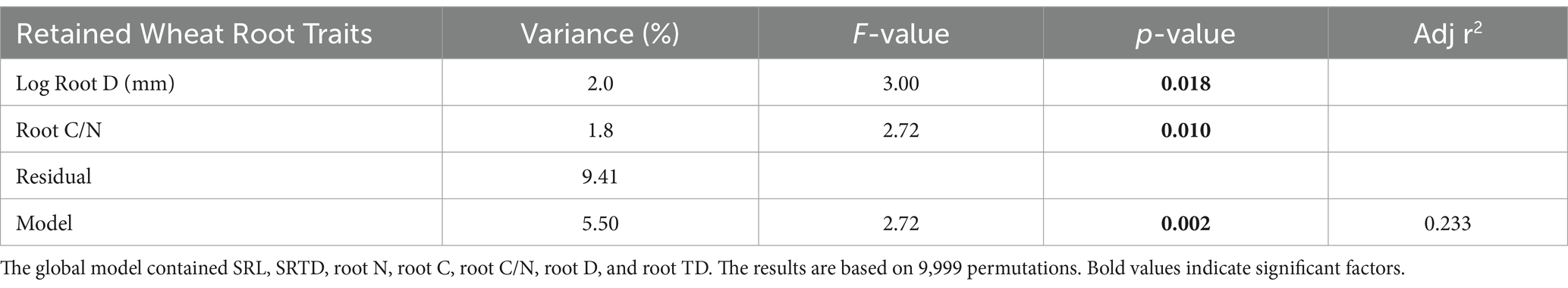
Table 5. Redundancy analysis for percent variance of fungal community composition explained by wheat root traits.
A Pearson correlation coefficient matrix was computed to assess the linear relationship between individual root traits and rhizosphere POxC, Apase, total N, total C, C/N ratio, and available P (Figure 2). In soybeans, SRL was found to be significantly positively correlated with POxC (r = 0.35, p = 0.041), total N (r = 0.40, p = 0.017), and total C (r = 0.35, p = 0.040), while being negatively correlated with C/N (r = −0.34, p = 0.046). Additionally, for soybeans, POxC was negatively correlated with root carbon (r = −0.34, p = 0.046) and the root C/N ratio (r = −0.35, p = 0.040).
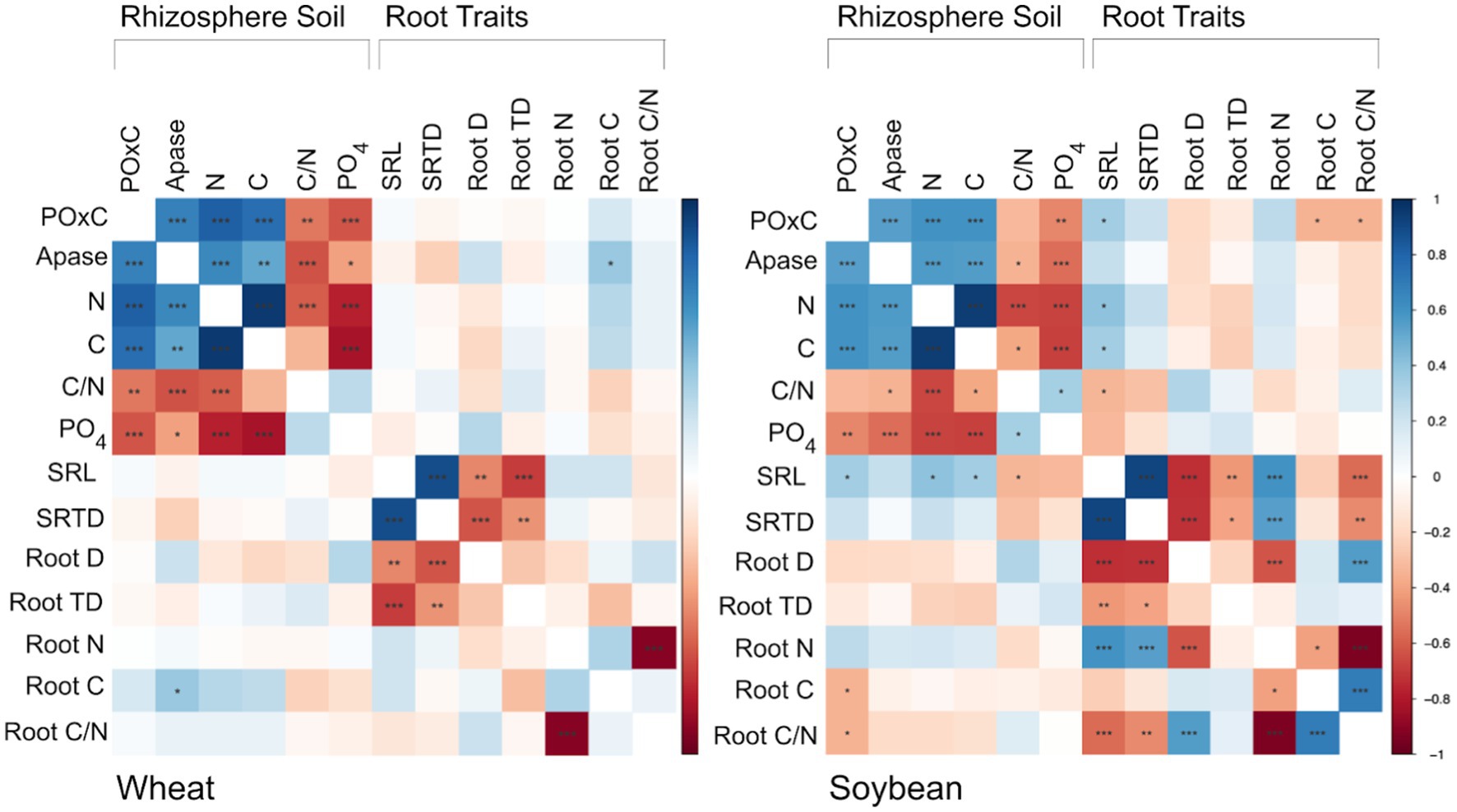
Figure 2. Pearson correlation matrix depicting relationships among the six measured root functional traits [SRL; SRTD; root N; root C/N; root D; root TD] and the six measured rhizosphere soil properties [total N, total C, C/N ratio, available P (PO4), active C (POxC), and Apase activity]. Significance is denoted by asterisks, where * p = 0.05, ** p = 0.01, *** p = 0.001. Wheat rhizosphere total N, C, C/N, and P and root SRL, SRTD, and root N and soybean rhizosphere POxC, total N, C, C/N and P and root SRL, SRTD, root TD, and root C/N were log-transformed prior to analysis.
Our results reveal that resource acquisition strategies are influenced by amendment regimes in wheat; organic amendments induce root trait expression toward the acquisitive end of the RES (Figure 1). We measured an increase in wheat root acquisitive traits, largely characterized by shifts in SRL and SRTD, as a result of adding the organic amendment compared to both the inorganic treatment and the no amendment treatment (Table 3). The substantial increase in SRL noted suggests that organic amendments induce changes in trait expression that allow for an increase in the volume of soil explored per unit investment of carbon. It has been proposed that thin, acquisitive root traits have a greater potential to adapt to fluctuating soil resources (Eissenstat et al., 2015). Given that organic amendments are characterized by heterogeneous nutrient cycling, resulting in nutrient hotspots, this may represent an advantageous resource strategy in agroecosystems (Borden et al., 2020). Similar shifts in root traits are also evident in long-term studies; Siddiqui et al. (2022) identified long-term root adaptations to organic versus conventional cropping systems. Organic-adapted populations were characterized by thin root Ds, longer roots with narrow root angles, larger surface areas, increased root mass density, and a greater number of metaxylem vessels, whereas conventionally adapted populations exhibited larger root volumes, thicker diameters, and fewer metaxylem vessels.
In contrast to wheat, soybean did not show any significant changes in root traits due to the treatments (Table 1). It is possible that soybean, as an N2-fixer, may have been more effective at managing organic forms of nitrogen and, therefore, exhibited fewer significant changes in root trait expression due to nutrient differences compared to wheat. Additionally, the dosage of fertilizer used in this study (75 kg N ha−1) may not have been sufficient to induce variation in soybean root functional traits. For example, McCoy et al. (2018) reported no difference in soybean root length and area when N fertilizer levels were applied at a rate of 45 kg ha−1 N; however, changes in root length and area were observed when N fertilizer was applied at a rate of 135 kg ha−1 N.
Despite minimal shifts in root traits in soybeans with treatments, variations in root traits, as measured through CVs, differed among treatments (Table 2). The root trait CVs increased in response to organic amendments for many individual root traits, specifically root acquisitive traits such as SRL and SRTD in both wheat and soybean. This indicates higher variability in a more heterogeneous environment. Siddiqui et al. (2022) found increased genetic variation in barley root system traits under organic conditions. Interestingly, while individual root traits were not significantly affected by inoculation, seed inoculation led to a general decrease in root trait variation in both soybean and wheat, as indicated by the CVs for each root trait (Table 2). The observed lower CV values suggest a tighter and more stable expression of root traits, which may indicate a fitness advantage in certain stable environments and may help in predicting trait outcomes (Dalal et al., 2017). Given the known positive effects of plant growth-promoting bacteria (PGPB) on enhancing plant stress responses (Glick, 2005; Grover et al., 2021), it is possible that the increased ability of inoculated plants to mediate stressful changes in the soil environment resulted in a more constrained or stable expression of root functional traits.
Although changes due to the inoculation treatment were not observed at the root level, significant effects were noted above ground, particularly for wheat. This indicates a decoupling of above- and belowground processes in response to the combined agroecological interventions (Table 1). Potential relationships between performance traits and root traits, as well as the effects of inoculation on root traits due to changes in overall plant fitness, may have been obscured by the plastic response of traits that respond differently to above- and belowground resources (Isaac et al., 2017; Weemstra et al., 2022). Several studies have also shown that endophyte bacteria can simultaneously upregulate and downregulate plant genes and metabolites (Timm et al., 2016; Weston et al., 2012). It is possible that overlapping signals from the inoculation masked certain belowground plant trait responses (Henning et al., 2016). Additionally, low or heterogeneous inoculation success may have occurred in the crops. Furthermore, the PGPB may have induced changes in earlier developmental stages that were not measured in this study, which focused on the later stages of phenological development.
Inoculation-induced changes have also been shown to be host-specific (Henning et al., 2016). This study noted host-specific changes. For wheat, significant positive effects were detected for the base diameter and the nitrogen usage index (Table 1). The positive effects on above-ground traits for wheat are intriguing, as larger base diameters may create stronger structural support, thereby promoting lodging resistance (Niu et al., 2022). Mayer et al. (2019) demonstrated that this particular strain positively affected the growth of Arabidopsis, basil, lettuce, and bok choy in laboratory settings. The observed increase in the N usage index is also favorable for nutrient management since optimizing nitrogen usage is an important strategy for maximizing yield and agronomic potential while reducing nitrogen inputs (Baligar et al., 2001). It is possible that the significant effect of inoculation on performance traits indicates a higher degree of trait integration; however, further research is needed to investigate and confirm this. A higher degree of integration may benefit crops in cases where a coordinated above-ground phenotype is required to combat a potentially stressful environment (Matesanz et al., 2021).
In soybeans, the relationships between root traits and rhizosphere properties are evident, with SRL emerging as a key trait (Figure 2). Soybean SRL is positively correlated with permanganate oxidizable carbon (POxC), total nitrogen (N), and total carbon (C), while it is negatively related to the carbon/nitrogen (C/N) ratio. Other studies have reported positive correlations between SRL and root exudation rates (Guyonnet et al., 2018; Meier et al., 2020). These positive relationships between rhizosphere properties and SRL may be explained by increases in exudation corresponding to higher SRL. Additionally, root C/N is negatively correlated with soybean rhizosphere POxC. Permanganate oxidizable carbon, or labile carbon, is measured by the amount of potassium permanganate (KMnO4) reduced in soil, indicating the active carbon pool in soils. Research suggests that POxC serves as a sensitive indicator of changes in soil organic carbon resulting from various soil management practices (Culman et al., 2012). A negative correlation between root carbon content and the C/N ratio may be related to increased exudation rates, leading to higher POxC content in the rhizosphere. In contrast, POxC is not associated with root traits in wheat. Variations in POxC among amendment regimes in wheat rhizosphere samples are likely due to changes in labile carbon resulting from the amendments or differences in microbial action, rather than changes induced by roots. Furthermore, we demonstrated that soil available P and Apase are negatively correlated in both soybean and wheat rhizospheres, highlighting the strong relationships between nutrient availability and the enzymatic activity required to release nutrients from organic sources—a crucial dynamic in organic agricultural systems a crucial dynamic in organic agricultural systems.
Fungal community redundancy analysis (RDA) indicated that, for wheat, individual root traits help explain fungal community composition (Table 5). Fungal communities, particularly arbuscular mycorrhizal fungi, are expected to be closely linked with SRL and root D due to a proposed trade-off between foraging and outsourcing strategies (Bergmann et al., 2020; Sweeney et al., 2021). Fulthorpe et al. (2020) also demonstrated that low root nitrogen, large root Ds, and high root C/N ratios were strong predictors of fungal endophyte community composition. In this analysis, RDA showed that wheat root D and root C/N were significant predictors of wheat rhizosphere fungal community composition, further suggesting an outsourcing strategy.
Root exudation and its effects on nutrient pools and fungal community characteristics are often complex and interconnected. Root functional traits are proposed to regulate the degree and extent of rhizosphere alteration, and the magnitude of the rhizosphere effect may depend on plant resource strategies, which can be categorized into acquisitive and conservative patterns (Han et al., 2020). Increased root exudation is negatively correlated with conservative root traits (Sun et al., 2021). Here, we demonstrate that the properties of the rhizosphere are influenced by root trait expression. It is reasonable to expect that treatments affecting root traits will produce corresponding effects on rhizosphere properties. Roots are recognized as key contributors to the alteration of the physical, chemical, and biological properties of the rhizosphere through root trait expression and exudation (Canarini et al., 2019). Rhizodeposits are diverse and may include carbohydrates, enzymes, hormones, proteins, organic acids, and amino acids, many of which are associated with nutrient acquisition (Hunter et al., 2014). Additionally, it is important to note that rhizodeposition can be complex and influenced by factors beyond root trait expression, including plant nutrient status, ontogeny, and environmental conditions (Dakora and Phillips, 2002).
In this study, we demonstrated that using an endophyte bacterial inoculant can lead to limited interactive effects among fertility treatments. This indicates a more significant impact of soil amendments on belowground traits when agroecological interventions are combined. We also show that the magnitude of above-ground benefits can remain relatively consistent across soil conditions within a specific crop. Microbial solutions can complement the integration of best management practices for soil health (e.g., organic amendments, crop rotation, reduced tillage) (Hartman et al., 2018; Norris and Congreves, 2018). However, it is essential to recognize that considerable work is required to identify suitable strains for specific hosts and effective solutions for scaling efficacy to field conditions. Rapid advancements in microbial products are anticipated, and research and industry partners should ensure they inform growers of realistic product expectations, development timelines, and potential limitations (Batista and Singh, 2021).
Our findings also indicate a need to optimize organic-adapted genotypes, as conventionally adapted genotypes may exhibit traits that do not maximize the agronomic potential in organic systems (see, Isaac et al., 2021; Warschefsky et al., 2014). Field crops like wheat and soybean have undergone significant aboveground selection pressure for traits that maximize yield. However, in low-input systems, these traits may not always be beneficial. For example, Li et al. (2021) examined the loss of functional traits for nutrient exploration and uptake due to selection under high nutrient availability. Efforts should be made to optimize genotypes for beneficial trait expression in agroecological cropping systems. Overall, soil amendments had a more substantial impact on crop root response than seed inoculants, although seed inoculants positively affected direct measures of crop performance. Importantly, these two agroecological interventions did not significantly interact, highlighting opportunities for improved integration of agroecological strategies.
The original contributions presented in the study are publicly available. Raw reads were deposited to NCBI short read archive (PRJNA1216962).
LR-T: Conceptualization, Data curation, Formal analysis, Investigation, Methodology, Visualization, Writing – original draft. RF: Conceptualization, Methodology, Funding acquisition, Project administration, Resources, Supervision, Writing – review & editing. MI: Conceptualization, Funding acquisition, Project administration, Resources, Supervision, Writing – review & editing, Data curation, Formal analysis.
The author(s) declare that financial support was received for the research and/or publication of this article. This work was supported by Natural Sciences and Engineering Research Council of Canada, Discovery Grant and the Canada Research Chairs program.
We thank Areem Siddiqi and Majura Peraparkaran for their field assistance. We acknowledge the NSERC Discovery Grant and the Canada Research Chairs program for their funding support.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The authors declare that no Gen AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Alikhani, H. A., and Emami, S. (2020). Investigating the Genetic Diversity of Wheat Growth Promoting Rhizosphere and Endophytic Bacteria. J. Microbiol. Biol, 9, 25–40. Available at: https://bjm.ui.ac.ir/article_24665.html
Almeida, O. A. C., Araujo, N. O. de., Mulato, A. T. N., Persinoti, G. F., Sfoorca, M. L., Calderan-Rodrigues, M. J., et al. (2023). Bacterial volatile organic compounds (VOCs) promote growth and induce metabolic changes in rice. Front. Plant Sci. 13:1056082. doi: 10.3389/fpls.2022.1056082
Altieri, M. A. (2019). Agroecology: The science of sustainable agriculture. 2nd Edn. Boca Raton, FL, USA: CRC Press.
Baligar, V. C., Fageria, N. K., and He, Z. L. (2001). Nutrient use efficiency in plants. Commun. Soil Sci. Plant Anal. 32, 921–950. doi: 10.1081/CSS-100104098
Bardgett, R. D., Mommer, L., and de Vries, F. T. (2014). Going underground: root traits as drivers of ecosystem processes. Trends Ecol. Evol. 29, 692–699. doi: 10.1016/j.tree.2014.10.006
Bargaz, A., Faghire, M., Abdi, N., Farissi, M., Sifi, B., Drevon, J.-J., et al. (2012). Low soil phosphorus availability increases acid phosphatases activities and affects P partitioning in nodules, seeds and rhizosphere of Phaseolus Vulgaris. Agriculture 2, 139–153. doi: 10.3390/agriculture2020139
Bargaz, A., Noyce, G. L., Fulthorpe, R., Carlsson, G., Furze, J. R., Jensen, E., et al. (2017). Species interactions enhance root allocation, microbial diversity and P Acquisition in Intercropped Wheat and Soybean under P deficiency. Appl. Soil Ecol. 120, 179–188. doi: 10.1016/j.apsoil.2017.08.011
Batista, B. D., and Singh, B. K. (2021). Realities and hopes in the application of microbial tools in agriculture. Microb. Biotechnol. 14, 1258–1268. doi: 10.1111/1751-7915.13866
Bergmann, J., Weigelt, A., van der Plas, F., Laughlin, D. C., Kuyper, T. W., Guerrero-Ramírez, N., et al. (2020). The fungal collaboration gradient dominates the root economics space in plants. Sci. Adv. 6:eaba3756. doi: 10.1126/sciadv.aba3756
Bokulich, N. A., and Mills, D. A. (2013). Improved selection of internal transcribed spacer-specific primers enables quantitative, ultra-high-throughput profiling of fungal communities. Appl. Environ. Microbiol. 79:12. doi: 10.1128/AEM.03870-12
Borden, K. A., Thomas, S. C., and Isaac, M. E. (2020). Variation in fine root traits reveals nutrient-specific acquisition strategies in agroforestry systems. Plant Soil 453, 139–151. doi: 10.1007/s11104-019-04003-2
Braun-Kiewnick, A., Giongo, A., Zamberlan, P. M., Pluta, P., Koch, H. J., Kage, H., et al. (2024). The Microbiome of Continuous Wheat Rotations: Minor Shifts in Bacterial and Archaeal Communities but a Source for Functionally Active Plant-Beneficial Bacteria. Phytobiomes J. doi: 10.1094/PBIOMES-05-24-0054-R
Buchanan, S. W., Mafa-Attoye, T., Dunfield, K., Thevathasan, N. V., and Isaac, M. E. (2022). The role of plant functional traits and diversity in soil carbon dynamics within riparian agroforests. J. Environ. Qual. 51, 33–43. doi: 10.1002/jeq2.20303
Buchanan, S.-W., Sauvadet, M., and Isaac, M. E. (2024). Decomposition of litter mixtures induces non-additive effects on soil priming across a riparian land use gradient. Soil Biol. Biochem. 190:109285. doi: 10.1016/j.soilbio.2023.109285
Callahan, B., McMurdie, P., Rosen, M., Han, A. W., Johnson, A. J. A., Holmes, S. P, et al. (2016). DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583. doi: 10.1038/nmeth.3869
Canarini, A., Kaiser, C., Merchant, A., Richter, A., and Wanek, W. (2019). Root exudation of primary metabolites: mechanisms and their roles in plant responses to environmental stimuli. Front. Plant Sci. 10:157. doi: 10.3389/fpls.2019.00157
Caporaso, J., Kuczynski, J., Stombaugh, J., Bittinger, K., Bushman, F. D., Costello, E. K., et al. (2010). QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336. doi: 10.1038/nmeth.f.303
Chen, Y., Camps-Arbestain, M., Shen, Q., Singh, B., and Cayuela, M. L. (2018). The long-term role of organic amendments in building soil nutrient fertility: a Meta-analysis and Review. Nutr. Cycl. Agroecosyst. 111, 103–125. doi: 10.1007/s10705-017-9903-5
Culman, S. W., Snapp, S. S., Freeman, M. A., Schipanski, M. E., Beniston, J., Lal, R., et al. (2012). Permanganate Oxidizable carbon reflects a processed soil fraction that is sensitive to management. Soil Sci. Soc. Am. J. 76, 494–504. doi: 10.2136/sssaj2011.0286
Dakora, F. D., and Phillips, D. A. (2002). Root exudates as mediators of mineral Acquisition in low-Nutrient Environments. Plant Soil 245, 35–47. doi: 10.1023/A:1020809400075
Dalal, A., Attia, Z., and Moshelion, M. (2017). To produce or to survive: how plastic is your crop stress physiology? Front. Plant Sci. 8:2067. doi: 10.3389/fpls.2017.02067
Diacono, M., and Montemurro, F. (2010). Long-term effects of organic amendments on soil fertility. A Review. Agron. Sustain. Dev. 30, 401–422. doi: 10.1051/agro/2009040
Drinkwater, L. E., and Snapp, S. S. (2007). “Nutrients in agroecosystems: rethinking the management paradigm” in Advances in agronomy, vol. 92, 163–186. doi: 10.1016/S0065-2113(04)92003-2
Eissenstat, D. M., Kucharski, J. M., Zadworny, M., Adams, T. S., and Koide, R. T. (2015). Linking root traits to nutrient foraging in arbuscular mycorrhizal trees in a temperate Forest. New Phytol. 208, 114–124. doi: 10.1111/nph.13451
Elnahal, A. S. M., El-Saadony, M. T., Saad, A. M., Desoky, E. M., El-Tahan, A. M., Rady, M. M., et al. (2022). The use of microbial inoculants for biological control, plant growth promotion, and sustainable agriculture: a Review. Eur. J. Plant Pathol. 162, 759–792. doi: 10.1007/s10658-021-02393-7
FAO (2021). Standard operating procedure for soil pH determination. Rome, Italy: Food and Agriculture Organization of the United Nations.
Fiodor, A., Singh, S., and Pranaw, K. (2021). The contrivance of plant growth promoting microbes to mitigate climate change impact in agriculture. Microorganisms 9:1841. doi: 10.3390/microorganisms9091841
Freschet, G. T., Pagès, L., Iversen, C. M., Comas, L. H., Rewald, B., Roumet, C., et al. (2021a). A starting guide to root ecology: strengthening ecological concepts and Standardising root classification, sampling, processing and trait measurements. New Phytol. 232, 973–1122. doi: 10.1111/nph.17572
Freschet, G. T., Roumet, C., Comas, L. H., Weemstra, M., Bengough, A. G., Rewald, B., et al. (2021b). Root traits as drivers of plant and ecosystem functioning: current understanding, pitfalls and future research needs. New Phytol. 232, 1123–1158. doi: 10.1111/nph.17072
Fulthorpe, R., Martin, A. R., and Isaac, M. E. (2020). Root endophytes of coffee (Coffea Arabica): variation across climatic gradients and relationships with functional traits. Phytobiomes J. 4, 27–39. doi: 10.1094/PBIOMES-04-19-0021-R
Gagliardi, S., Martin, A. R., Filho, E. d. M. V., Rapidel, B., and Isaac, M. E. (2015). Intraspecific leaf economic trait variation partially explains coffee performance across agroforestry management regimes. Agric. Ecosyst. Environ. 200, 151–160. doi: 10.1016/j.agee.2014.11.014
Glick, B. R. (2005). Modulation of plant ethylene levels by the bacterial enzyme ACC deaminase. FEMS Microbiol. Lett. 251, 1–7. doi: 10.1016/j.femsle.2005.07.030
Grover, M., Bodhankar, S., Sharma, A., Sharma, P., Singh, J., and Nain, L. (2021). PGPR mediated alterations in root traits: way toward sustainable crop production. Front. Sustain. Food Syst. 4:618230. doi: 10.3389/fsufs.2020.618230
Guyonnet, J. P., Cantarel, A. A. M., Simon, L., and Haichar, F. e. Z. (2018). Root exudation rate as functional trait involved in plant nutrient-use strategy classification. Ecol. Evol. 8, 8573–8581. doi: 10.1002/ece3.4383
Hamidov, A., Helming, K., Bellocchi, G., Bojar, W., Dalgaard, T., Ghaley, B. B., et al. (2018). Impacts of climate change adaptation options on soil functions: a Review of European case-studies. Land Degrad. Dev. 29, 2378–2389. doi: 10.1002/ldr.3006
Han, M., Chen, Y., Li, R., Yu, M., Fu, L., Li, S., et al. (2022). Root phosphatase activity aligns with the collaboration gradient of the root economics space. New Phytol. 234, 837–849. doi: 10.1111/nph.17906
Han, M., Sun, L., Gan, D., Fu, L., and Zhu, B. (2020). Root functional traits are key determinants of the rhizosphere effect on soil organic matter decomposition across 14 temperate hardwood species. Soil Biol. Biochem. 151:108019. doi: 10.1016/j.soilbio.2020.108019
Hartman, K., van der Heijden, M. G. A., Wittwer, R. A., Banerjee, S., Walser, J.-C., and Schlaeppi, K. (2018). Cropping practices manipulate abundance patterns of root and soil microbiome members paving the way to smart farming. Microbiome 6:14. doi: 10.1186/s40168-017-0389-9
Hayes, F. J., Buchanan, S. W., Coleman, B., Gordon, A. M., Reich, P. B., Thevathasan, N. V., et al. (2019). Intraspecific variation in soy across the leaf economics Spectrum. Ann. Bot. 123, 107–120. doi: 10.1093/aob/mcy147
Henning, J. A., Weston, D. J., Pelletier, D. A., Timm, C. M., Jawdy, S. S., and Classen, A. T. (2016). Root bacterial endophytes Alter Plant phenotype, but not physiology. PeerJ 4:e2606. doi: 10.7717/peerj.2606
Hunter, P. J., Teakle, G. R., and Bending, G. D. (2014). Root traits and microbial community interactions in relation to phosphorus availability and acquisition, with particular reference to Brassica. Front. Plant Sci. 5:27. doi: 10.3389/fpls.2014.00027
Isaac, M. E., and Borden, K. A. (2019). Nutrient acquisition strategies in agroforestry systems. Plant Soil 444, 1–19. doi: 10.1007/s11104-019-04232-5
Isaac, M. E., Martin, A. R., de Melo Virginio Filho, E., Rapidel, B., Roupsard, O., and van den Meersche, K. (2017). Intraspecific trait variation and coordination: root and leaf economics spectra in coffee across environmental gradients. Front. Plant Sci. 8:1196. doi: 10.3389/fpls.2017.01196
Isaac, M. E., Nimmo, V., Gaudin, A. C. M., Leptin, A., Schmidt, J. E., Kallenbach, C. M., et al. (2021). Crop domestication, root trait syndromes, and soil nutrient Acquisition in Organic Agroecosystems: a systematic Review. Front. Sustain. Food Syst. 5:716480. doi: 10.3389/fsufs.2021.716480
Isaac, M. E., Isakson, S. R., Dale, B., Levkoe, C. Z., Hargreaves, S. K., Mendez, V. E., et al. (2018). Agroecology in Canada: towards an integration of Agroecological practice, movement, and science. Sustain. For. 10:3299. doi: 10.3390/su10093299
Isaac, M. E., Sinclair, F., Laroche, G., Olivier, A., and Thapa, A. (2024). The ties that bind: how trees can enhance agroecological transitions. Agrofor. Syst. 98, 2369–2383. doi: 10.1007/s10457-024-01014-6
Kang, S.-M., Shahzad, R., Bilal, S., Khan, A. L., Park, Y.-G., Lee, K.-E., et al. (2019). Indole-3-acetic-acid and ACC deaminase producing Leclercia Adecarboxylata MO1 improves Solanum Lycopersicum L. growth and salinity stress tolerance by endogenous secondary metabolites regulation. BMC Microbiol. 19:80. doi: 10.1186/s12866-019-1450-6
Khan, A. L., Halo, B. A., Elyassi, A., Ali, S., Al-Hosni, K., Hussain, J., et al. (2016). Indole acetic acid and ACC deaminase from endophytic Bacteria improves the growth of Solanum Lycopersicum. Electron. J. Biotechnol. 21, 58–64. doi: 10.1016/j.ejbt.2016.02.001
Lal, R. (2015). Restoring soil quality to mitigate soil degradation. Sustain. For. 7, 5875–5895. doi: 10.3390/su7055875
Li, L., Li, Q., Davis, K. E., Patterson, C., Oo, S., Liu, W., et al. (2021). Response of root growth and development to nitrogen and potassium deficiency as well as microRNA-mediated mechanism in Peanut (Arachis Hypogaea L.). Front. Plant Sci. 12:695234. doi: 10.3389/fpls.2021.695234
Loaces, I., Ferrando, L., and Scavino, A. F. (2011). Dynamics, diversity and function of endophytic Siderophore-producing Bacteria in Rice. Microb. Ecol. 61, 606–618. doi: 10.1007/s00248-010-9780-9
Lumactud, R., and Fulthorpe, R. R. (2018). Endophytic Bacterial Community Structure and Function of Herbaceous Plants from Petroleum Hydrocarbon Contaminated and Non-Contaminated Sites. Front. Microbiol, 9, 1926. doi: 10.3389/fmicb.2018.01926
Lumactud, R., Fulthorpe, R., Sentchilo, V., and van der Meer, J. R. (2017). Draft Genome Sequence of Plantibacter flavus Strain 251 Isolated from a Plant Growing in a Chronically Hydrocarbon-Contaminated Site. Genome Announc, 5:e10–1128. doi: 10.1128/genomea.00276-17
Lumactud, R., Shen, S. Y., Lau, M., and Fulthorpe, R. (2016). Bacterial Endophytes Isolated from Plants in Natural Oil Seep Soils with Chronic Hydrocarbon Contamination. Front. Microbiol, 7:755. doi: 10.3389/fmicb.2016.00755
Lynch, J. P., Strock, C. F., Schneider, H. M., Sidhu, J. S., Ajmera, I., Galindo-Castaneda, T., et al. (2021). Root anatomy and soil resource capture. Plant Soil 466, 21–63. doi: 10.1007/s11104-021-05010-y
Malik, K. M., Khan, K. S., Rukh, S., Khan, A., Akbar, S., Billah, M., et al. (2021). Immobilization of cd, Pb and Zn through organic amendments in wastewater irrigated soils. Sustain. For. 13:2392. doi: 10.3390/su13042392
Martin, A. R., Hayes, F. J., Borden, K. A., Buchanan, S. W., Gordon, A. M., Isaac, M. E., et al. (2019). Integrating nitrogen fixing structures into above- and belowground functional trait spectra in soy (Glycine Max). Plant Soil 440, 53–69. doi: 10.1007/s11104-019-04058-1
Martin, A. R., and Isaac, M. E. (2015). REVIEW: plant functional traits in agroecosystems: a blueprint for research. J. Appl. Ecol. 52, 1425–1435. doi: 10.1111/1365-2664.12526
Martin, A. R., and Isaac, M. E. (2021). The leaf economics Spectrum’s morning coffee: plant size-dependent changes in leaf traits and reproductive onset in a perennial tree crop. Ann. Bot. 127, 483–493. doi: 10.1093/aob/mcaa199
Matesanz, S., Blanco-Sánchez, M., Ramos-Muñoz, M., de la Cruz, M., Benavides, R., and Escudero, A. (2021). Phenotypic integration does not constrain phenotypic plasticity: differential plasticity of traits is associated to their integration across environments. New Phytol. 231, 2359–2370. doi: 10.1111/nph.17536
Maxton, A., Singh, P., and Masih, S. A. (2018). ACC deaminase-producing Bacteria mediated drought and salt tolerance in Capsicum Annuum. J. Plant Nutr. 41, 574–583. doi: 10.1080/01904167.2017.1392574
Mayer, E., Dörr de Quadros, P., and Fulthorpe, R. (2019). Plantibacter Flavus, Curtobacterium Herbarum, Paenibacillus Taichungensis, and Rhizobium Selenitireducens endophytes provide host-specific growth promotion of Arabidopsis Thaliana, basil, lettuce, and bok choy plants. Appl. Environ. Microbiol. 85, e00383–e00319. doi: 10.1128/AEM.00383-19
McCoy, J. M., Kaur, G., Golden, B. R., Orlowski, J. M., Cook, D. R., Bond, J. A., et al. (2018). Nitrogen fertilization of soybean affects root growth and nodulation on two soil types in Mississippi. Commun. Soil Sci. Plant Anal. 49, 181–187. doi: 10.1080/00103624.2017.1421649
Meier, I. C., Tückmantel, T., Heitkötter, J., Müller, K., Preusser, S., Wrobel, T. J., et al. (2020). Root exudation of mature beech forests across a nutrient availability gradient: the role of root morphology and fungal activity. New Phytol. 226, 583–594. doi: 10.1111/nph.16389
Munroe, J. W., Soto, G., de Virginio Filho, E., Fulthorpe, R., and Isaac, M. E. (2015). Soil microbial and nutrient properties in the rhizosphere of coffee under agroforestry management. Appl. Soil Ecol. 93, 40–46. doi: 10.1016/j.apsoil.2015.04.003
Muvingi, M., Slovareva, O. Y., Yaremko, A. B., Zargar, M., Lyashko, M., Pakina, E., et al. (2023). Variations of the Bacterial Community in Wheat (Triticum aestivum), Oats (Avena sativa L.), Barley (Hordeum vulgare L.), and Triticale (Triticosecale) from Three Regions of the Republic of Crimea. Preprints, 202307.1559.v1. doi: 10.20944/preprints202307.1559.v1
Narayanan, Z., and Glick, B. R. (2022). Secondary metabolites produced by plant growth-promoting bacterial endophytes. Microorganisms 10:2008. doi: 10.3390/microorganisms10102008
Negi, Y. K., Sajwan, P., Uniyal, S., and Mishra, A. C. (2021). Enhancement in yield and nutritive qualities of strawberry fruits by the application of organic manures and biofertilizers. Sci. Hortic. 283:110038. doi: 10.1016/j.scienta.2021.110038
Nimmo, V., Violle, C., Entz, M., Rolhauser, A. G., and Isaac, M. E. (2023). Changes in crop trait plasticity with domestication history: management practices matter. Ecol. Evol. 13:e10690. doi: 10.1002/ece3.10690
Niu, Y., Chen, T., Zhao, C., Guo, C., and Zhou, M. (2022). Identification of QTL for stem traits in wheat (Triticum Aestivum L.). Front. Plant Sci. 13:962253. doi: 10.3389/fpls.2022.962253
Norris, C. E., and Congreves, K. A. (2018). Alternative management practices improve soil health indices in intensive vegetable cropping systems: a Review. Front. Environ. Sci. 6:50. doi: 10.3389/fenvs.2018.00050
Orozco-Mosqueda, M. d. C., Glick, B. R., and Santoyo, G. (2020). ACC deaminase in plant growth-promoting Bacteria (PGPB): an efficient mechanism to counter salt stress in crops. Microbiol. Res. 235:126439. doi: 10.1016/j.micres.2020.126439
Reich, P. B. (2014). The world-wide ‘fast–slow’ plant economics Spectrum: a traits manifesto. J. Ecol. 102, 275–301. doi: 10.1111/1365-2745.12211
Robinson, R. J., Fraaije, B. A., Clark, I. M., Jackson, R. W., Hirsch, P. R., Mauchline, T. H., et al. (2016). Wheat Seed Embryo Excision Enables the Creation of Axenic Seedlings and Kochs Postulates Testing of Putative Bacterial Endophytes. Sci. Rep. 6:25581. doi: 10.1038/srep25581
Rolhauser, A. G., and Isaac, M. E. (2024). Plant traits mediate agroecological outcomes in restored agricultural lands. Ecol. Indic. 161:111951. doi: 10.1016/j.ecolind.2024.111951
Röös, E., Mie, A., Wivstad, M., Salomon, A., Johansson, B., Gunnarsson, S., et al. (2018). Risks and opportunities of increasing yields in organic farming. A Review. Agron. Sustain. Dev. 38:14. doi: 10.1007/s13593-018-0489-3
Schrama, M., de Haan, J. J., Kroonen, M., Verstegen, H., and Van der Putte, W. H. (2018). Crop yield gap and stability in organic and conventional farming systems. Agric. Ecosyst. Environ. 256, 123–130. doi: 10.1016/j.agee.2017.12.023
Sessitsch, A., and Mitter, B. (2015). 21st century agriculture: integration of plant microbiomes for improved crop production and food security. Microb. Biotechnol. 8, 32–33. doi: 10.1111/1751-7915.12180
Sharma, A., Kumar, P., Pahal, V., Kumar, J., and Pandey, S. S. (2023). “Endophytic Phytohormone production and utilization of functional traits in plant growth promotion” in Plant Microbiome for Plant Productivity and Sustainable Agriculture. eds. S. Chhabra, et al. (Singapore: Springer Nature), 365–385.
Siddiqi, M., and Glass, A. (1981). Utilization index: A modified approach to the estimation and comparison of nutrient utilization efficiency in plants. J. Plant Nutr. 4. 289–302. doi: 10.1080/01904168109362919
Siddiqui, M. N., Schneider, M., Barbosa, M. B., Leon, J., and Ballvora, A. (2022). Natural selection under conventional and organic cropping systems affect root architecture in spring barley. Sci. Rep. 12:20095. doi: 10.1038/s41598-022-23298-3
Spitzer, C. M., Lindahl, B., Wardle, D. A., Sundqvist, M. K., Gundale, M. J., Fanin, N., et al. (2021). Root trait-microbial relationships across tundra plant species. New Phytol. 229, 1508–1520. doi: 10.1111/nph.16982
Sun, L., Ataka, M., Han, M., Han, Y., Gan, D., Xu, T., et al. (2021). Root exudation as a major competitive fine-root functional trait of 18 coexisting species in a subtropical Forest. New Phytol. 229, 259–271. doi: 10.1111/nph.16865
Sweeney, C. J., de Vries, F. T., van Dongen, B. E., and Bardgett, R. D. (2021). Root traits explain rhizosphere fungal community composition among temperate grassland plant species. New Phytol. 229, 1492–1507. doi: 10.1111/nph.16976
Timm, C. M., Pelletier, D. A., Jawdy, S. S., Gunter, L. E., Henning, J. A., Engle, N., et al. (2016). Two poplar-associated bacterial isolates induce additive favorable responses in a constructed plant-microbiome system. Front. Plant Sci. 7:497. doi: 10.3389/fpls.2016.00497
Timmusk, S., Behers, L., Muthoni, J., Muraya, A., and Aronsson, A. C. (2017). Perspectives and challenges of microbial application for crop improvement. Front. Plant Sci. 8:49. doi: 10.3389/fpls.2017.00049
Violle, C., Navas, M.-L., Vile, D., Kazakou, E., Fortunel, C., Hummel, I., et al. (2007). Let the concept of trait be functional! Oikos 116, 882–892. doi: 10.1111/j.0030-1299.2007.15559.x
Warschefsky, E., Penmetsa, R. V., Cook, D. R., and von Wettberg, E. J. B. (2014). Back to the wilds: tapping evolutionary adaptations for resilient crops through systematic hybridization with crop wild relatives. Am. J. Bot. 101, 1791–1800. doi: 10.3732/ajb.1400116
Weemstra, M., Roumet, C., Cruz-Maldonado, N., Anthelme, F., Stokes, A., and Freschet, G. T. (2022). Environmental variation drives the decoupling of leaf and root traits within species along an elevation gradient. Ann. Bot. 130, 419–430. doi: 10.1093/aob/mcac052
Weil, R. R., Islam, K. R., Stine, M. A., Gruver, J. B., and Samson-Liebig, S. E. (2003). Estimating active carbon for soil quality assessment: a simplified method for laboratory and field use. Am. J. Altern. Agric. 18, 3–17. doi: 10.1079/AJAA200228
Weston, D. J., Pelletier, D. A., Morrell-Falvey, J. L., Tschaplinski, T. J., Jawdy, S. S., Lu, T.-Y., et al. (2012). Pseudomonas fluorescens induces strain-dependent and strain-independent host plant responses in defense networks, primary metabolism, photosynthesis, and fitness. Mol. Plant-Microbe Interact. 25, 765–778. doi: 10.1094/MPMI-09-11-0253
Wright, I. J., Reich, P. B., Westoby, M., Ackerly, D. D., Baruch, Z., Bongers, F., et al. (2004). The worldwide leaf economics Spectrum. Nature 428, 821–827. doi: 10.1038/nature02403
Keywords: soil biogeochemistry, crop traits, root functional traits, organic agriculture, seed inoculants, wheat, soybean, Plantibacter flavus
Citation: Ritcey-Thorpe L, Fulthorpe RR and Isaac ME (2025) Effects of multiple agroecological interventions on crop nutrient acquisition and performance. Front. Sustain. Food Syst. 9:1520713. doi: 10.3389/fsufs.2025.1520713
Received: 31 October 2024; Accepted: 24 February 2025;
Published: 27 March 2025.
Edited by:
Anas Tallou, University of Bari Aldo Moro, ItalyReviewed by:
João Paulo Soto Veiga, Centro Universitário de Jaguariúna (UniFAJ), BrazilCopyright © 2025 Ritcey-Thorpe, Fulthorpe and Isaac. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Marney E. Isaac, bWFybmV5LmlzYWFjQHV0b3JvbnRvLmNh
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.