
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Sustain. Food Syst. , 19 March 2025
Sec. Crop Biology and Sustainability
Volume 9 - 2025 | https://doi.org/10.3389/fsufs.2025.1550106
This article is part of the Research Topic Innovative Solutions For Next-Generation Fertilizers View all 13 articles
Strawberry (Fragaria × ananassa Duchesne ex Rozier) is an important small fruit grown worldwide. Organic amendments can alter microbial communities and increase crop productivity. While research on organic amendments in strawberry cultivation has primarily focused on various regions in the U.S., especially the West Coast, there has been little to no investigation into their potential benefits in North Carolina (NC). A three-year trial was conducted from 2019 to 2022 at the Horticultural Crops Research Station in Castle Hayne, NC, U. S. A. The main objective of the study was to examine the effects of reduced rates of dried molasses (5.60 t/ha) and mustard meal (2.24 t/ha), a half-rate combination of both, a positive control (Pic Clor 60), and a negative control (no fumigated and no amended) on soil bacterial communities and strawberry yield. Our results from 16S microbiome amplicon sequencing showed significant variations in the composition of the soil bacterial community over time between the organic amendment treatments and the controls. The alpha diversity indices (Shannon index) of the soil bacterial microbiome were generally higher in plots with organic amendments than those treated with Pic Clor 60. Taxonomic classification revealed that the two phyla, Proteobacteria and Actinobacteriota, were prominent in the organic amendment treatments. The total marketable yield rankings for the three organic amendment treatments (dried molasses, mustard meal, and the half-rate combination of both) were comparable to those achieved through fumigation. The results indicated that bacterial structure and yield improved in the organic amendment plots, while microbial diversity decreased in the fumigation plots, and yields were lower in the untreated plots. This study will inform the selection of organic amendments to enhance microbial diversity and promote sustainability in strawberry farming in NC.
Strawberry (Fragaria × ananassa Duchesne ex Rozier) is a highly nutritious and vital crop (Hannum, 2004). The continuous cultivation of strawberries in fields for an extended period has decreased the diversity of bacterial communities and soil health parameters, while the presence of pathogenic fungi has increased (Li and Liu, 2019; Yang et al., 2023). Organic amendments provide essential nutrients, enhance the quality of soil, and impact microbial communities, disease management, and crop productivity in various regions (Oldfield et al., 2018; Strauss and Kluepfel, 2015). An organic amendment refers to any material that increases the organic matter content of soil and improves its chemical, physical, and biological properties (Goss et al., 2013; Rosskopf et al., 2020).
Soil management practices such as cover crops, reduced tillage, mulches, and organic amendments can improve sustainable soil quality, health, and crop productivity (Abawi and Widmer, 2000; Louws et al., 2000; Pankhurst, 1997; Rosskopf et al., 2020). Cover crops like ryegrass, pearl millet, oats, white clover, cowpea, and hairy vetch can be used as green manure crops before planting. They contribute to improved soil health and crop yield, erosion prevention, reduction of plant-parasitic nematode populations, and the addition of organic matter or nitrogen (e.g., legumes) to subsequent cash crops (Fageria et al., 2005). Two common organic amendments, dried molasses, and mustard meal are frequently used in combination with anaerobic soil disinfestation (ASD) to reduce soil-borne diseases and enhance microbial communities in strawberry and tomato production systems across the West Coast, Southeastern, and Midwestern regions (Mazzola et al., 2015; Testen and Miller, 2018; Vincent et al., 2022; Wang and Mazzola, 2019). Dried molasses is a byproduct of the sugar-making process obtained from crushed sugar cane and contains high sugar content which positively impacts soil microbiomes, improves soil structure and moisture retention; functions as a biopesticide, and promotes plant growth and high yields (Expósito et al., 2022). On the other hand, various Brassicaceae seed meals (SMs) like Sinapis alba L., Brassica juncea L., and B. carinata have been utilized as soil amendments in agricultural production systems (Mazzola and Brown, 2010; Mazzola et al., 2015). Among these, a commercial Brassica pellet called BioFence (Triumph Italia SPA) was incorporated into the soil to suppress soil-borne pathogens significantly, attributed to a chemically-derived mechanism by releasing biologically active glucosinolate hydrolysis products such as isothiocyanates (Matthiessen and Kirkegaard, 2006). Organic amendments must be effective, affordable, readily available, and biodegradable for soil microbiomes (Daugovish et al., 2021, 2023). It is crucial to evaluate how different organic amendments affect beneficial microbiomes and crop productivity in a new area (Mazzola et al., 2018; Larkin, 2015; Liu et al., 2021; Shu et al., 2022). This is necessary because soil, weather conditions, soil-borne pathogens, microbial communities, and cropping systems vary across regions (Strauss and Kluepfel, 2015).
California’s climate and soil composition are unique, making it possible for strawberries to grow throughout the year. Several studies conducted on the West Coast indicate that organic amendments as carbon sources impact soil microbial communities (Deng et al., 2019; Hewavitharana et al., 2014, 2019; Poret-Peterson et al., 2019, 2020). For example, rice bran used resulted in microbial communities with a distinct composition compared to those obtained from molasses (Mazzola et al., 2018). The main strawberry growing season in North Carolina typically occurs from October to early June. However, it may vary depending on geographic locations, soil types, strawberry genotypes, growing seasons, and weather conditions (Bandara and Janaranjana, 2021; Poling, 1993; Samtani et al., 2019; Rana and Gu, 2020). Strawberry growers use raised beds with black plastic mulch and buried drip irrigation lines. Plugs (rooted strawberry tips) are transplanted in the beds in mid-October and managed throughout the winter. Harvesting takes place from early April to early June, and marketable yields can range from 16.8 t/ha to 33.63 t/ha, with net returns from $4,856 t/ha to $6,475/ha (Rysin et al., 2015; Sydorovych et al., 2006). Intensive cropping systems have led to low soil carbon inputs and reduced soil fertility. Additionally, the lack of crop rotation and high dependence on fumigants have caused soil deterioration. Although various management practices such as organic amendments, cover crops, crop rotations, and biofumigation were evaluated in North Carolina (NC), U. S. A. (Louws, 2009; Welker et al., 2008), strawberry growers encounter unique challenges, like distinct pest problems, limited access to high-yield varieties, and the impact of climate change on disease outbreaks (Bandara and Janaranjana, 2021). Importantly, the findings from studies on organic amendments conducted on the West Coast may not be directly applicable to the East Coast, such as NC. Consequently, there is a knowledge gap in conducting a comprehensive assessment of the impacts of organic amendments on soil microbiome composition and structure, and their effect on strawberry production in NC. Depending on organic amendments, they can either increase or decrease soil microbial diversity compared to chemical fertilizers (Ahn et al., 2012; Bebber and Richards, 2020; Bonanomi et al., 2020; Feng et al., 2018; Shu et al., 2022). Soil microbiomes are a vital part of soil biodiversity and contribute to various ecosystem processes such as the decomposition of organic matter, nutrient cycling, and enhancement of plant productivity (Bernard et al., 2014; Bender et al., 2016; Bonanomi et al., 2020; Saleem et al., 2019; Shu et al., 2022; Xia et al., 2020). Overusing mineral fertilizers in the past depleted microbial diversity in agricultural soils (Zhou et al., 2020). On the other hand, organic amendments, including livestock manure, crop residue, cereal brans, molasses, mustard meal, and green manure, not only improve soil microbial activities but also alter microbial composition and diversity (Gravuer et al., 2019; Li et al., 2021; Liu et al., 2021; Rosskopf et al., 2020; Xia et al., 2020; Zhou et al., 2020).
We hypothesize that organic amendments could positively shift bacterial community assembly and improve strawberry yield, as this could be a critical and unexplored aspect in North Carolina. This study aimed to fill the gaps in our knowledge about the effects of two organic amendments: dried molasses and mustard meal on bacterial abundance, composition, and diversity in the soil, and on crop yield in NC. To achieve this, we conducted three-year field trials, extracted DNA from the soil, sequenced the V3-V4 region of the 16S ribosomal RNA (rRNA), and monitored the yield over time.
Field experiments were conducted from 2019 to 2022 at the Horticultural Crops Research Station in Castle Hayne, NC, U. S. A. The experimental site is situated at a latitude of 34° 21′ 20” N and a longitude of 77°53′ 59” W, and its elevation is nine meters above sea level. The climate of this area is mild, with warm summers and cool winters. The average daily temperature varies between 32°C in the summer and 15°C in the winter. The rainfall is evenly distributed throughout the year, with an average annual precipitation of 1,118 mm (Robinson, 2005). The baseline soil conditions, recorded during land preparation in 2019, consisted of a clay sandy loam with a pH of 5.9. The soil had an organic carbon content of 1.16%, a cation exchange capacity of 4.5 meq/100 cm2, and the following nutrient levels: phosphorus at 90 mg/kg, potash at 14 mg/kg, magnesium at 8 mg/kg, and calcium at 63 mg/kg. Soil depth varies depending on soil types, compactness, and crop root growth. For example, strawberry is a shallow-rooted crop, and our experimental site has clay sandy loam soil. Thus, we collected soils at a 12 to 15 cm depth to assess soil nutrients and microbiomes.
In mid-September of the years 2019, 2020, and 2021, three raised beds were created for each treatment plot, each 9.2 m long, 0.61 m wide, and 0.2 m high, with 1.5 m from the center of one bed to the other. Beds were pre-formed, and the organic amendment was added to the top of each bed and mixed with the soil at 15 cm soil depth as recommended previously (Murphy, 2020) using a rototiller to ensure even distribution. The beds were pulled and covered with a 0.025 mm black VaporSafe® totally impermeable film (TIF) made of polyethylene plastic mulch (Raven Industries, Inc., Sioux Falls, SD, U. S. A.). The experiment was arranged using a randomized block design, with each block or replication being the same size as the number of treatments (i.e., factor). Additionally, each treatment was assigned to one experimental unit within each block. The same design with four blocks was used each year. Each block consisted of 10 experimental units (plots) in 2020 and 2021, and 5 in 2022 and treatments were applied annually to the same plot area. Each treatment was applied to one experimental unit within the block were typically considered as random effects. For microbiome analysis, three replications per treatment were utilized to minimize sequencing costs, while four replications per treatment were used for yield data analysis.
Ten experimental treatments using black or clear plastic mulch were evaluated in 2020 and 2021 (Table 1). Only five effective treatments for higher yield in 2020 and 2021 were further selected and evaluated in 2022.
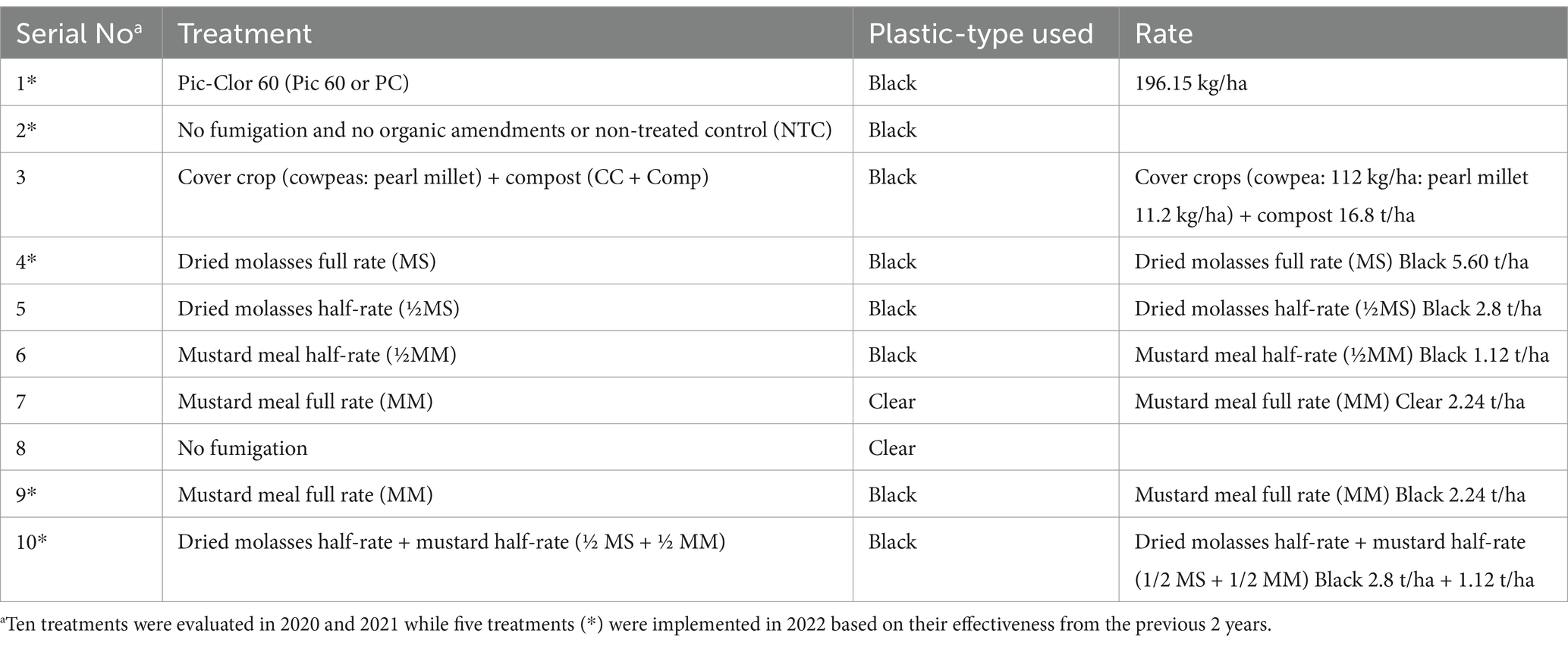
Table 1. Ten treatments were used in field experiments between 2019 and 2021, and five promising treatments/controls were selected and evaluated in 2022.
The following materials were used in these experiments. During the summer, a mixture of cowpea and pearl millet cover crops were seeded to promote optimal growth. Before seeding in June, a standard compost application rate of 16.8 metric tons per hectare was applied, as reported by a local waste management consortium (Nuova Amit, 2007) and previous experience (McWhirt et al., 2014). After this, the vegetation was flail-mowed early September to evenly distribute the cut residue across the cover crop plots. The residue from the cover crops was then incorporated into the soil to a depth of 20 to 30 cm using a PTO-driven rototiller, as recommended by McWhirt et al. (2014).
To reduce production costs, we purchased dried molasses (5.60 t/ha) from Agricultural Carbon Source in Plant City, FL, U. S. A. and mustard meal (MM) in the form of BioFence™ (2.24 t/ha) from Brassica carinata at Volunteer Ag Products in Greenfield, TN, U. S. A. A half rate of MS and MM (½ MS + ½ MM) was also prepared. Two control treatments were used: pre-plant chemical fumigation using Pic-Clor 60 (60% chloropicrin and 40% 1, 3-dichloro propene; 196.15 kg/ha) from TriEst Ag Group, Inc., (Greenville, NC, U. S. A.) as a positive control and no fumigation and no amendment as a negative control. Fumigant beds were prepared by injecting Pic-Clor 60 with two knives per bed evenly spaced (12 to 15 cm soil depth). Two drip lines were placed about 2.5 cm beneath the soil surface and spaced approximately 0.3 m apart. The beds were covered with 0.025 mm black VaporSafe® TIF plastic mulch (Raven Industries, Inc., Sioux Falls, SD, U. S. A.). As reported previously (Shennan et al., 2014), all beds treated with organic amendments were saturated to a depth of 20 to 30 cm. This saturation was achieved by applying water at a pressure of 83 kPa for up to 6 hours or until beds were saturated, which facilitated the rapid breakdown of the organic amendments.
The application of readily biodegradable organic amendments is followed by irrigating the soil to its field capacity and maintaining anaerobic conditions for a specific duration known as the incubation period, which typically lasts from 3 to 6 weeks (Meshram et al., 2024; Shennan et al., 2014). This process influences the composition of the soil microbial community. The standard wait time after fumigation is three weeks before planting to prevent crop damage due to chemical residues. Therefore, 3 weeks after adding the organic amendments, the beds were aerated by puncturing holes in the plastic. The beds were planted with five-week-old seedlings, known as plugs, from the commercial strawberry variety ‘Chandler’ (Poling, 1993). This variety is commonly grown in North Carolina, USA (Samtani et al., 2019), and the seedlings were sourced from Aarons Creek Farms Inc. in Junction, VA, or Fresh-Pik Produce Inc. in Wilson, NC, U. S. A. between October 17 and 18 each year. Sixty plants were planted on each bed with offset twin rows at ~0.30 m between each plant. Subsequently, plants were watered and fertilized according to standard production practices (Hoffmann et al., 2021). After the June harvest, the strawberry beds were broken up; the same treatment was applied to each plot and treated the same way every year.
Soil samples for microbiome analysis were collected from the harvesting area of each plot, covering approximately 1.9 m2 from the entrance of the central bed. To prevent plot interference across the years, all data were secured from the central beds of the three-bed plot. In 2019, soil samples (20 cores/plot) were collected from each treatment after applying organic amendments and Pic-Clor 60 and just one day before planting. For the second collection, soil samples were collected between the first and second week of April each year. A handheld soil probe (1.75 cm internal diameter) was used to collect 20 cores from the rhizosphere soil in the harvesting area at a depth of 15 cm. Approximately 500 grams of soil per replicate per treatment were collected and mixed well in a 10-liter bucket to create a bulk sample. In this study, the term “microbiome” pertains to the bacterial communities associated with this soil sample.
The yield data were collected from the inner 20-plant zone of the inner bed. The total marketable yield was recorded weekly in 2020 but semi-weekly in 2021 and 2022. Marketable yields were calculated in kilograms per hectare (kg/ha). The data was analyzed using the Statistical Analysis Software (SAS) Version 9.4 (SAS Institute, 2016) through analysis of variance (ANOVA). Two-way ANOVA was used to compare the effects of 10 treatments (Table 1) on strawberry yield. Mean values were then compared using Tukey’s HSD test at p ≤ 0.05 level. Spearman’s rank correlation coefficient analysis was performed to determine the strength and direction (negative or positive) and to assess the r and p-values for the slope of the relationship between the 2020 and 2021 yield data. Based on these results, five treatments: the full rate of dried molasses (MS), the full rate of mustard meal (MM), half the rate of dried molasses, and half the rate of mustard meal (½ MS + ½ MM), fumigation treatment with Pic-Clor 60 and a control treatment without fumigation were selected and evaluated in 2022.
DNA was extracted from 0.25 g of rhizosphere soil using the DNeasy PowerSoil® Pro kit (QIAGEN Sciences, Germantown, MD, U.S.A). The DNA was quantified using a Nanodrop 2000 spectrophotometer (Thermo Fisher Scientific, Waltham, MA, U.S.A). The final DNA concentration was adjusted to 20 ng/μl for each sample and then shipped to Novogene Corporation, Sacramento, CA, U.S.A.
The bacterial 16S ribosomal RNA (16S rRNA) gene amplicons were generated using the forward primer 515F and the reverse primer 806R, which target the V4 variable region (Caporaso et al., 2011). This approach enhances community clustering with the specified read length (Janssen, 2006; Klindworth et al., 2013; Liu et al., 2007). Standardized quantities of PCR products from each DNA sample were pooled and ligated with adapters following the manufacturer’s guidelines (Illumina, San Diego, CA, U.S.A.). Sequencing was performed on the Illumina MiSeq platform using the MiSeq reagent kit v2 for 250 bp paired-end runs (Novogene Corporation, Sacramento, CA, U.S.A.).
The sequencing data was processed using a Novogene proprietary pipeline.1 The Quantitative Insights Into Microbial Ecology (QIIME) software Version 1.7.0 (Caporaso et al., 2010) was used to filter the raw sequence and obtain a high-quality clean sequence (Bokulich et al., 2013). Aligned clean sequences with SILVA Version 128 (Quast et al., 2013) using a confidence threshold of 80%. Sequences of barcodes, primers, and short sequences containing ambiguous base calls and homopolymer runs exceeding 6 bp were removed. Sequences that were unassigned or classified as Eukaryota, Archaea, mitochondria, and chloroplasts were excluded. The aligned sequences were examined for chimeric sequences using UCHIME (Edgar et al., 2011), and any chimeric sequences were removed. The processed sequences were then clustered into operational taxonomic units (OTUs) at a 97% similarity level using UPARSE software (Edgar, 2013) and BLASTn from NCBI.2
Sequence data were analyzed using phyloseq Version 1.20.0 (McMurdie and Holmes, 2013) and an R software Version 4. 2.0 (R Core Team, 2022) and normalized using cumulative sum scaling in metagenomeSeq Version 1.18.0 (Paulson et al., 2014). The 16S rRNA training set from the Ribosomal Database Project Version 1.20 (Cole et al., 2014) was used to assign taxonomy to each OTU. Representative sequences were identified at the genus and species level, and the OTUs were utilized to compare bacterial communities in the soil across different treatments (Supplementary Table S1). Rarefaction curves were created based on the observed OTUs (Caporaso et al., 2010). The number of sequences was normalized across all samples to 9,999 reads using the random subsample method. The Shannon’s diversity index (H′) was used to assess bacterial community α-diversity (diversity within treatment) using mother Version 1. 34.1 (Schloss et al., 2009) using the minimum library size as the default with 1,000-bootstrap resampling. These indices were computed for treatments and years, and means were compared using Tukey’s Honest Significant Difference (HSD) test (p ≤ 0.001) to determine significant differences between treatments.
Venn diagrams (Heberle et al., 2015) were generated to compare OTU data of three organic amendment treatments (MS, MM, and ½ MS + ½ MM) with Pic-Clor 60 and NTC. The relative abundance of taxonomic ranks, such as phylum, genus, and species, were analyzed to identify the most abundant OTUs (greater than 0.3%) in each treatment. The two-part testing using a nonparametric Wilcoxon rank-based approach was used to compare the bacterial communities across different treatments and years (Wagner et al., 2011). Histograms were created to compare the common bacterial phyla with different organic amendment treatments and sampling years. They were selected to compare the abundance across treatments based on cumulative subsampling (CSS). Evolutionary trees were generated to compare the common bacterial genera using MUSCLE Version 3.8.31 (Edgar, 2004). Differentially enriched genera were visualized using ternary plots to identify the predominant bacterial species (Bulgarelli et al., 2015). This approach allowed for a comparison of individual organic amendments with Pic-Clor 60 and the negative control (NTC). The density of circles observed in the plots represents the relative abundance and distribution patterns of different bacterial species.
The β-diversity of the bacterial community was analyzed to generate Bray–Curtis dissimilarity using PAST software Version 4.03 (Hammer et al., 2001). After normalization, we examined the relative abundances of OTUs across each taxonomic rank in every sample to assess community dissimilarity coefficients between pairwise treatments using the R package metagenomeSeq Version 1.18.0 (Paulson et al., 2014). Heatmaps were employed to visualize the data, and a value of 0.00 indicated that the two treatments were identical. Tukey’s HSD test assessed mean differences using a significance level (p ≤ 0.05). To determine if bacterial community composition clustered according to treatments, non-metric multidimensional scaling (NMDS) was conducted using the Bray–Curtis dissimilarity index with PAST software Version 4.03 (Hammer et al., 2001). Additionally, a stress value greater than 0.3 indicates a poor representation, while a stress value below 0.01 signifies a uniform or excellent representation of the bacterial community composition. Normalized data were used to assess variations in bacterial taxa across treatments. The detected community of taxa in each sample was compared to those in all other samples using analysis of similarity (ANOSIM) (Hammer et al., 2001). In ANOSIM data analysis, a probability value (PANOSIM) of ≤0.01 and an average correlation coefficient (RANOSIM) of 0.98 indicates a strong relationship or significant similarities between the two treatments. In contrast, a high p value, such as PANOSIM = 0.5, along with a low RANOSIM (ranging from 0.2 to 0.4), suggests a weak correlation or notable dissimilarities between the two treatments.
Refraction curves typically reached a plateau, indicating that most of the community diversity had been sequenced and was sufficient for analysis (data not shown). Regardless of treatments, in 2020, amplified soil DNA showed the highest total raw and effective sequence (Supplementary Figures 1A–D).
The distribution of bacterial communities varied across treatments and years. The bacterial shared total OTUs in soil treated with MS, MM, and ½ MS + ½ MM differed significantly (p ≤ 0.0001) from Pic-Clor 60 and NTC each year (Supplementary Figures S2A–L). Overall, the total number of shared OTUs was significantly higher by 35 to 45% in the soil following all organic amendment treatments. The lowest total number of shared OTUs across all treatments occurred in 2019, with counts ranging from 3,078 to 3,265 (Supplementary Figures S2A–F). In contrast, all treatments achieved the highest number of shared OTUs in 2021, with counts ranging from 8,194 to 8,258 (Supplementary Figures S2G–L). The total shared OTUs for MS showed greater significance in 2019 and 2022, but not in the intervening years.
The CSS-normalized OTU data analysis of bacterial relative abundance indicated that eight phyla were prevalent across the treatments (Figures 1A–D). These predominant phyla included Acidobacteria, Actinobacteria, Bacteroidota, Chloroflexi, Firmicutes, Patescibacteria, Planctomycetes, and Proteobacteria, were common to all year soils sampled. The impact of organic amendments on the relative abundance of bacterial communities varied based on the treatments applied and the years examined. For example, significant differences were observed each year in the abundance of the bacterial phyla Actinobacteria and Proteobacteria between the organic amendment treatments (MS, MM, and ½ MS + ½ MM) and the positive control (Pic-Clor 60) (Figures 1A–D). Generally, the dominant bacterial phyla across all treatments were Firmicutes, Proteobacteria, and Actinobacteria, among others. Proteobacteria was more abundant in the plots treated with Pic-Clor 60 in 2022 than in other sampling years. Firmicutes were less abundant in the control treatments in 2019 and 2020 than those that received organic amendments. Proteobacteria showed a slight increase in abundance across all years following organic amendment treatments. Chloroflexota (specifically Chloroflexi) was more abundant in the MM and MS + MM treatments than in the others in 2021 (Figure 1C).
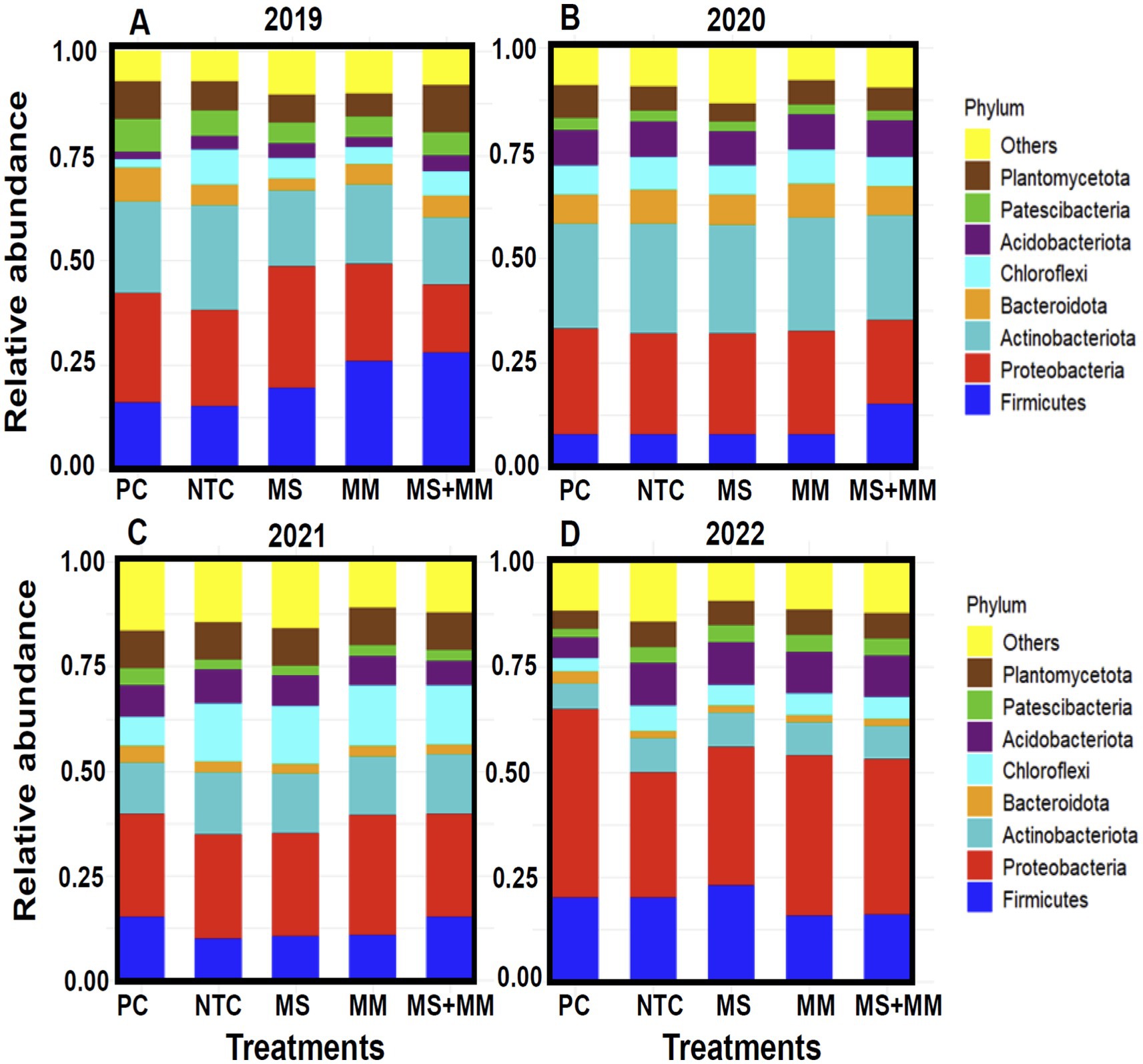
Figure 1. Stacked bar charts display the relative abundance of predominant bacteria at the phylum level (Y-axis) in soil samples collected over 4 years: 2019 (A), 2020 (B), 2021 (C), and 2022 (D). Sequence data from three replications in each treatment for 2019 to 2022 were used to determine the relative abundance of the top eight bacteria at the phylum level. Unknownphyla are categorized as “others.” Each bacterial phylum in each treatment isrepresented by the same color for each year, corresponding to three organic amendments (X-axis): dried molasses (MS), mustard meal (MM), and a combination of half-rate of both (½ MS + ½ MM). The control treatments included a non-amended control (NTC) and chemical soil fumigation with Pic-Clor 60 (PC).
Phylogenetic evolutionary trees were analyzed to determine the relationships among key bacterial genera under various treatments. For example, the genus Bacillus was found to be predominantly present in all three organic amendments across the years (Figures 2A–D). Similarly, regardless of the organic amendment treatments, Sphingomonas was observed in 2020, KD4-96 in 2021, and Burkholderia in 2022. The assembled reads were further analyzed using a ternary plot to confirm the effects of organic amendments on bacterial functional enrichment categories at the species level, which helped identify the bacterial species that support microbial establishment in the strawberry rhizosphere. The average abundance of the dominant bacterial species associated with the OTUs found in soil varied with different organic amendments and across the sampling years (Figures 3A–L). Ternary plots indicated that in 2019, the most frequently detected species included Chitophagaceae bacterium and Rhodanobacter. In 2020, Microcoleus spp., Bacillus aryabhattai, Terrabacter, and Clostridium saccharoperbutylaceto were predominant. In 2021, C. saccharoperbutylaceto and Bacillus funiculus were the most common, while in 2022; the frequently detected species included Bradyrhizobium alkalii, Komagataeibacter saccharivora, and Bacillus azotoformans (Figures 3A–L). Overall, the presence of these potentially beneficial bacteria demonstrated resilience across all the experiments.
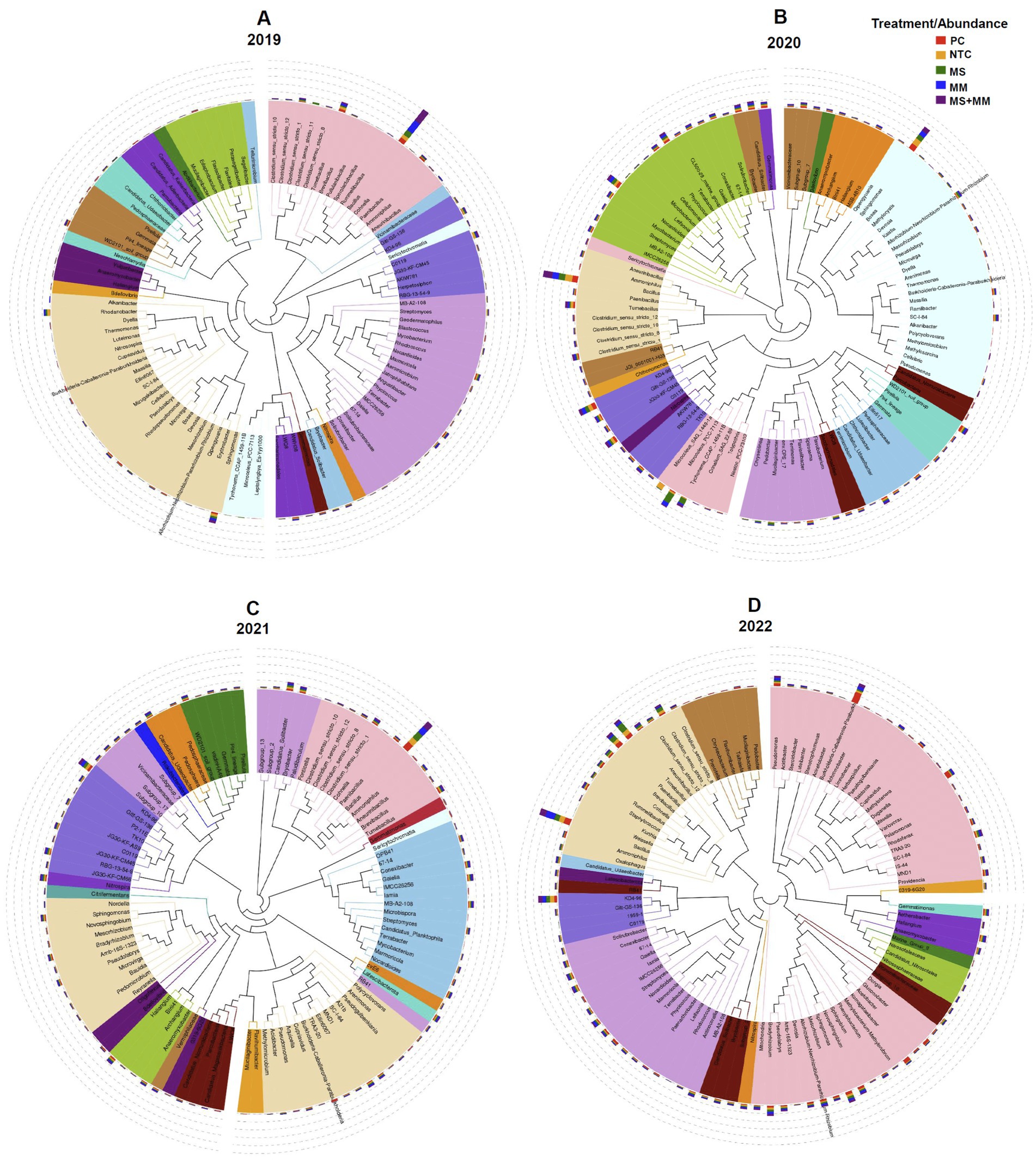
Figure 2. The evolutionary phylogenetic trees of the most abundant bacterial genera in soil samples collected over 4 years: 2019 (A), 2020 (B), 2021 (C), and 2022 (D) are presented in response to three organic amendments: dried molasses (MS), mustard meal (MM), and a combination of half-rate of both (½ MS + ½ MM). The controls included non-amended control (NTC) and chemical soil fumigation with Pic-Clor 60 (PC). The sequence data from three replications in each treatment were utilized for bacterial community analysis to construct the trees. In the innermost circle, different branches with colors represent bacterial phyla. The outermost circle with a stacked bar graph displays the relative abundance of the bacterial genus in each treatment. The same color for each year represents each treatment.
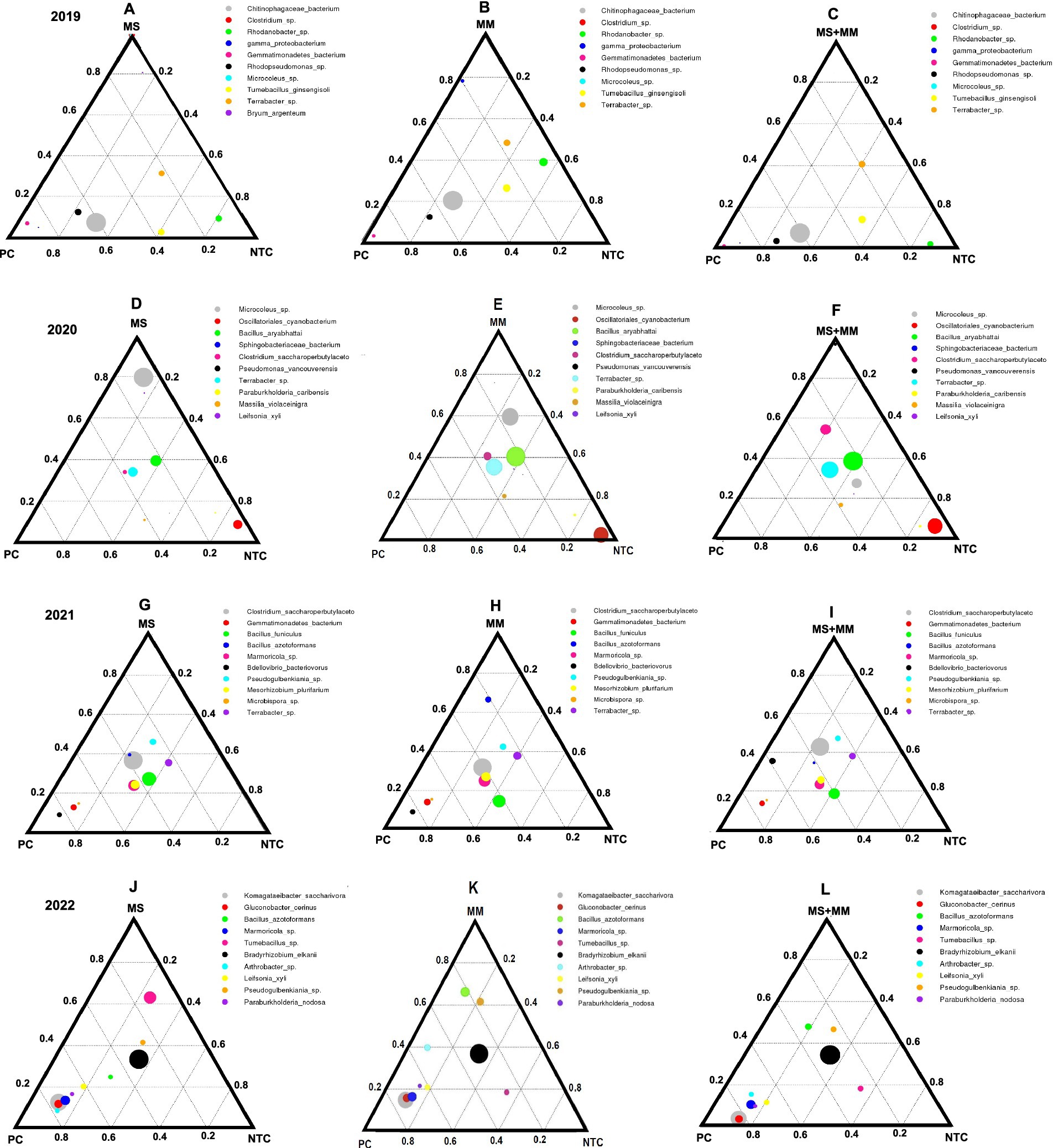
Figure 3. Ternary plots illustrate the average abundance of the dominant bacterial species found in soil samples collected over 4 years: 2019 (A), 2020 (B), 2021 (C), and 2022 (D). The sequence data from three replications in each treatment from 2019 to 2022 were used for the bacterial community analysis. Each ternary plot is represented as a triangle with sides scaled from zero to one, and the data can be expressed in terms of either proportions or absolute values. Each side of the triangle corresponds to one of the three measured components. The axes are arranged as follows: the X-axis is at the bottom of the plot, the Y-axis is on the right, and the Z-axis is on the left. A point is plotted on the triangle so that a line drawn perpendicular to each side from the point intersects at the respective component values. The impacts of three organic amendments were examined: dried molasses (MS), mustard meal (MM), and a combined half-rate of both (½ MS + ½ MM), along with a non-amended control (NTC) and a chemical soil fumigation treatment using Pic-Clor 60 (PC). The proportions of the three components in the plots are unconstrained, varying from zero to one. The size of each circle and color in the plots indicates the abundance of specific bacterial species in the soil.
The α-diversity for each sample using the Shannon diversity index (H′) was used to assess the diversity of bacterial communities in the soil. The results indicated that α-diversity varied significantly among treatments over the years (Figures 4A–D). Notably, the α-diversity in soil treated with Pic-Clor 60 was significantly lower (p ≤ 0.001) than in the other treatments. The lowest H′ ranging from 8.2 to 8.5, was recorded in 2019 with the application of Pic-Clor 60. Furthermore, the reduction in H′ was more pronounced across all treatments in 2019 compared to other years. In all organic treatments, H′ in 2021 was higher, falling between 11 and 11.5, compared to the range of 10 to 10.5 observed in 2019, 2020, and 2022 (Figures 4A–D). Additionally, the α-diversity of bacterial communities showed no significant differences in soil treated with MS, MM, and NTC in 2021, with H′ values ranging from 9.5 to 11.5.
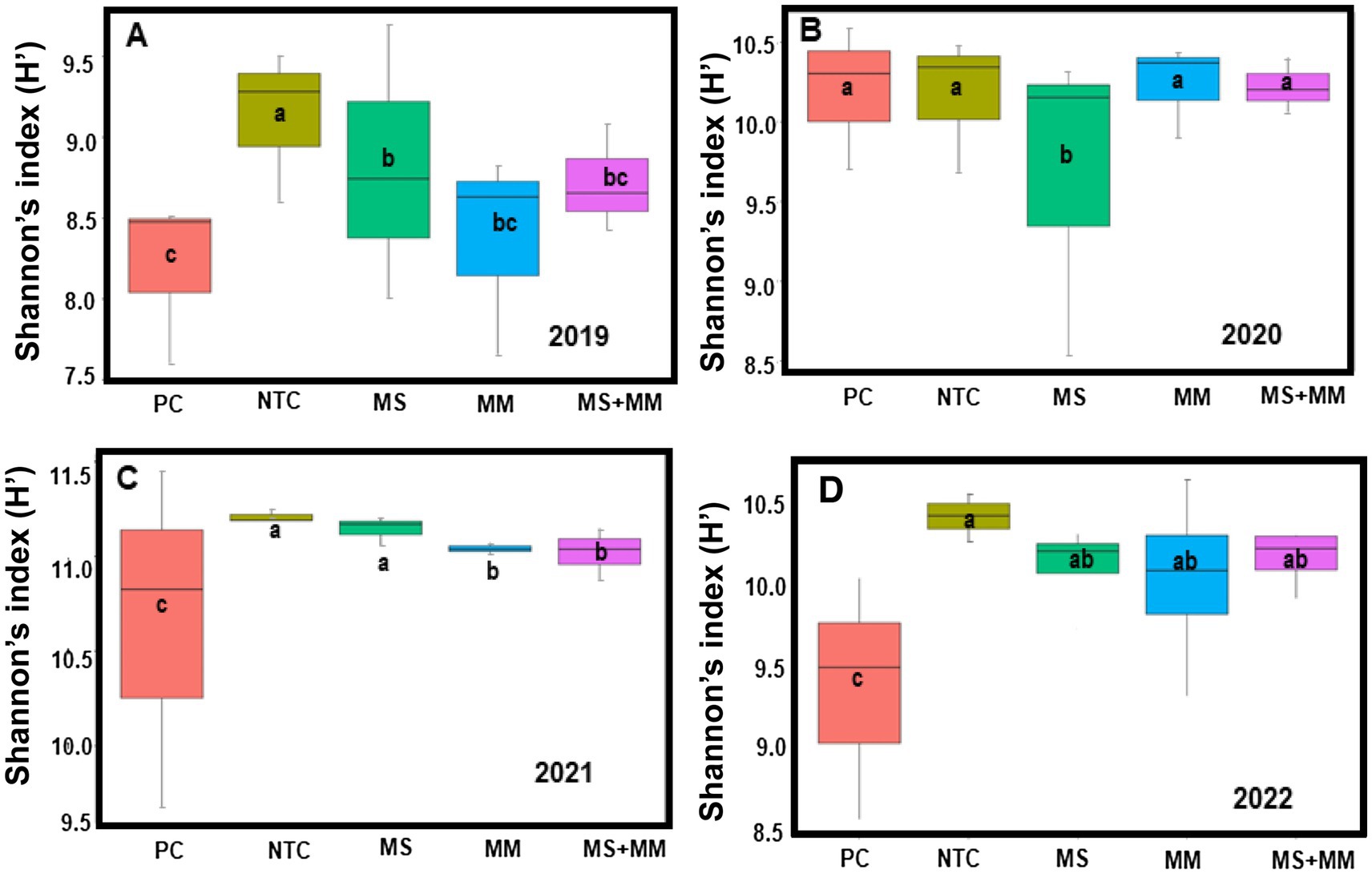
Figure 4. Comparison of the α-diversity of bacterial communities found in soil samples collected over 4 years: 2019 (A), 2020 (B), 2021 (C), and 2022 (D) in response to three organic amendments: dried molasses (MS), mustard meal (MM), and a combination half-rate of both (½ MS + ½ MM). The controls included non-amended control (NTC) and chemical soil fumigation with Pic-Clor 60 (PC). The sequence data from three replications in each treatment from 2019 to 2022 were used for bacterial community analysis in the soil. The boxplots display Shannon’s index (H′) on the Y-axis and treatments on the X-axis. The same color for each year represents each treatment. Mean values followed by the same letter are not significantly different based on Tukey’s honest significant difference test (p ≤ 0.05).
β-diversity analysis was performed to assess the differences in bacterial diversity among the treatments (Figures 5A–D). The results indicated that Pic-Clor 60 significantly influenced bacterial β-diversity in the soil (Tukey’s HSD test, p ≤ 0.001), with dissimilarity coefficients ranging from 0.329 in 2019 to 0.133 in 2020. Most samples from the respective treatments displayed low coefficient values and did not differ significantly from each other or negative control (NTC), indicating a high degree of similarity among them. For example, the dissimilarity coefficients for all organic amendment treatments (MS, MM, and ½ MS + ½ MM) and NTC were consistently below 0.1 and showed no significant differences across years (Figures 5A–D).
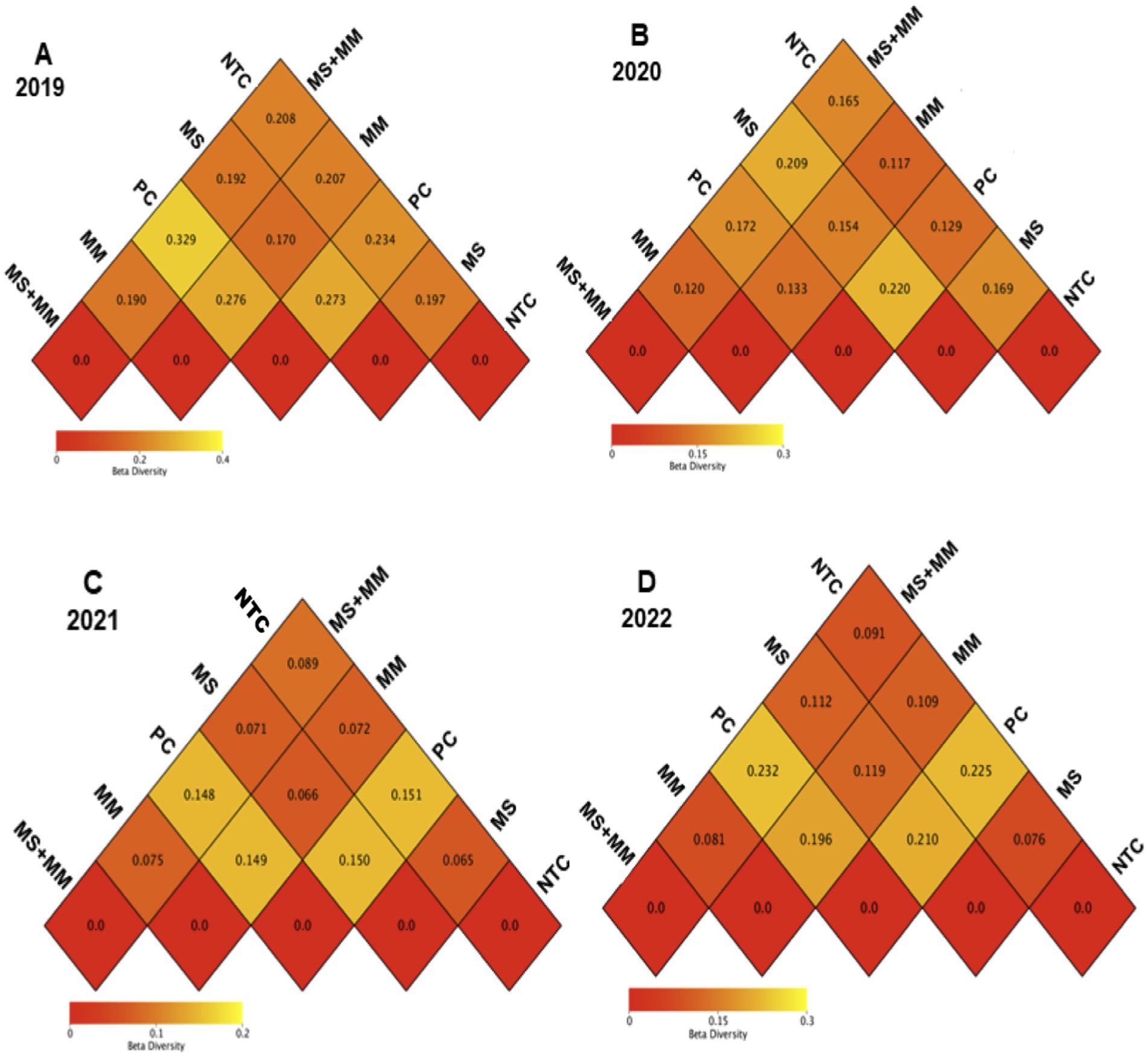
Figure 5. Comparison of the β-diversity of bacterial communities found in soil samples collected over 4 years: 2019 (A), 2020 (B), 2021 (C), and 2022 (D) in response to three organic amendments: dried molasses (MS), mustard meal (MM), and a combination half-rate of both (½ MS + ½ MM). The controls included non-amended control (NTC) and chemical soil fumigation with Pic-Clor 60 (PC). The sequence data from three replications in each treatment from 2019 to 2022 were used for bacterial community analysis in soil. Heatmaps were created to compare the dissimilarity coefficients and compared according to Tukey’s honest significant difference test (p ≤ 0.001). The dissimilarity coefficient score shows red for a decrease and green for an increase in bacterial β-diversity in treatment-paired comparisons.
To assess the variation in bacterial communities across different treatments and sampling years, we visualized the OTU data using Non-metric Multidimensional Scaling (NMDS) plots (Figures 6A–D). In both 2019 and 2022, the soil bacterial communities in plots treated with Pic-Clor 60 displayed distinct clustering and significantly different bacterial composition (p < 0.0001), indicating the highest degree of dissimilarity compared to other treatments (Figures 6A–D). In contrast, the bacterial communities in plots treated with organic amendments (e.g., MM, MS, or a combination of ½ MS and ½ MM) did not show significant differences in composition when compared to those treated with Pic-Clor 60 and the negative control (NTC) during 2020 and 2021. During these years, clustering by treatment was less distinct (Figures 6A–D). The stress values remained low throughout the years (i.e., < 0.3), ranging from 0.124 to 0.183. This indicates a strong representation of the bacterial community composition and a shift towards more stress-tolerant species. Overall, communities in soil amended with organic treatments were significantly different in composition from those in soil treated with Pic-Clor 60 in 2022 (PANOSIM ≤ 0.07 and RANOSIM ≥ 0.42). However, the bacterial communities observed in 2019 (PANOSIM ≤ 0.24 and RANOSIM ≥ 0.49) and 2020 (PANOSIM ≤ 0.73 and RANOSIM ≥ −0.13), and in 2021 (PANOSIM ≤ 0.78 and RANOSIM ≥ −0.14) suggested some degree of overlap between the bacterial compositions across treatments (Supplementary Table S2).
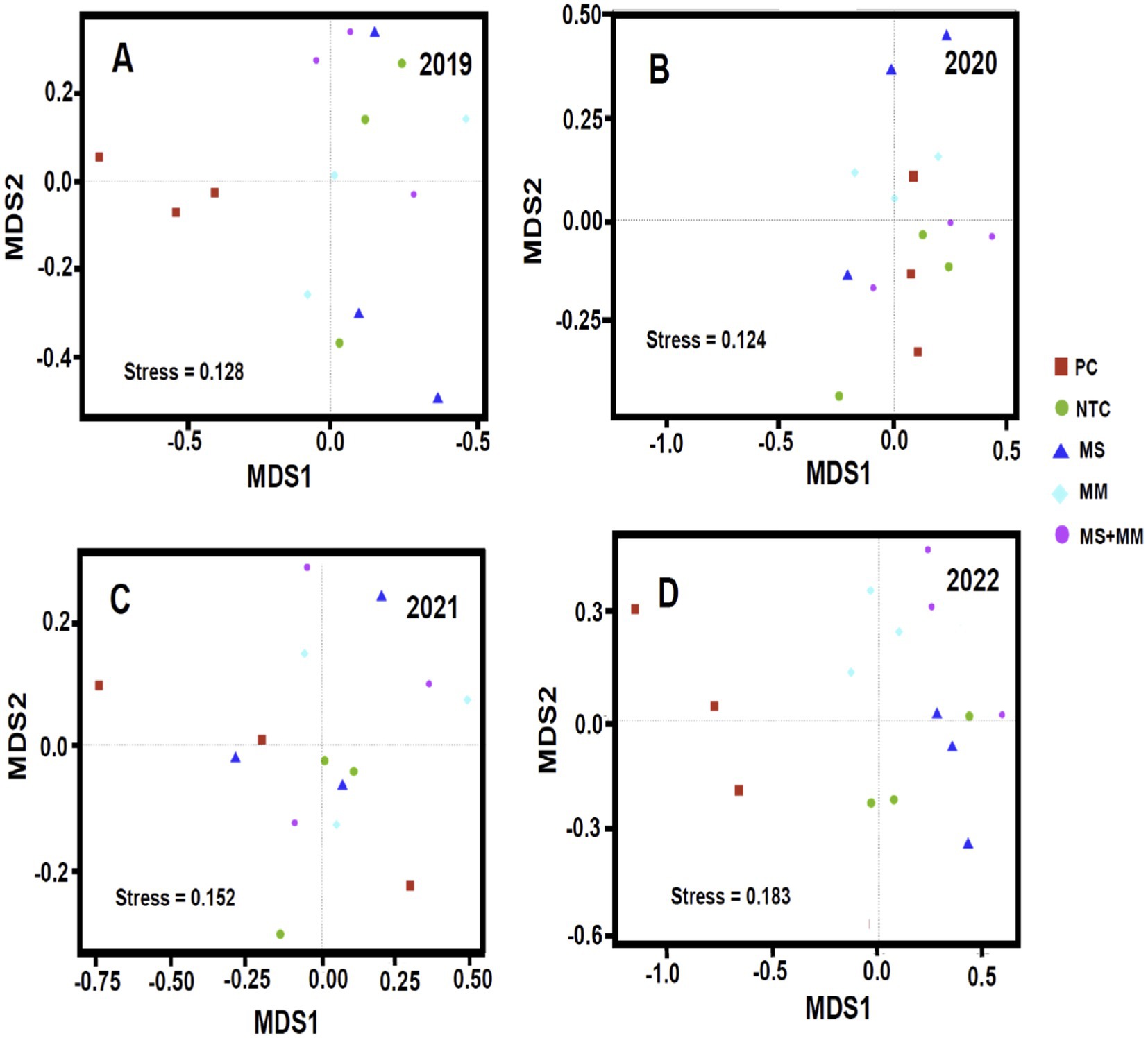
Figure 6. Ordination was conducted using nonmetric multidimensional scaling (NMDS) of operational taxonomic units, accompanied by the analysis of similarity (ANOSIM) (Hammer et al., 2001). This analysis was performed on soil samples collected over four years: 2019 (A), 2020 (B), 2021 (C), and 2022 (D). The study investigated the impact of three organic amendments: dried molasses (MS), mustard meal (MM), and a combination of half-rates of both (½ MS + ½ MM), along with a non-amended control (NTC) and chemical soil fumigation using Pic-Clor 60 (PC). Each treatment is represented by a symbol and organized by the same color for each year. The sequence data from three replications in each treatment from 2019 to 2022 were combined and used for bacterial community analysis. In ANOSIM data analysis, a probability value (PANOSIM) of ≤0.01 and an average correlation coefficient (RANOSIM) of 0.98 indicate a strong relationship or significant similarities between the two treatments. In contrast, a high p value, such as PANOSIM = 0.5, and a low RANOSIM (ranging from 0.2 to 0.4) suggests a weak correlation or dissimilarities between the two treatments.
Across 3 years, the total marketable yield fluctuated annually, with the lowest yield observed in 2020. Generally, cumulative marketable yields ranged from 4,775 kg/ha to 29,815 kg/ha (Figures 7A–C). Plots covered with clear plastic (NTC-CLR) in 2020 and 2021 displayed low marketable yields. Similarly, plots using compost/cover crop (CC + Comp) and plots covered with clear plastic (CLR), whether amended with MM full rate (MM-CLR) or not amended (NTC-CLR), also reported the lowest yields in both years. In 2021, all non-CLR treatments were similar (Figure 7B). Spearman’s correlation coefficient was used to examine organic amendments’ positive or negative influence on yield and the relationship between yields in 2020 and 2021. The coefficient value of 0.98 was highly significant (p ≤ 0.0001) (Supplementary Figure S3), indicating that the treatments that resulted in the highest yield in 2020 also led to the highest yield in 2021. The MS at full rate, the MM at half rate (½ MM), and the ½ MS + ½ MM combination produced higher yields than the NTC and showed similar yields as the Pic-Clor 60 in 2020 (Figure 7A). In 2022, the ½ MS + ½ MM combination and MS full rate amendments generated higher yields compared to both the NTC and Pic-Clor 60, while the MM full rate produced marketable berries equivalent to the Pic-Clor 60 treatment and higher than the NTC (Figure 7C).
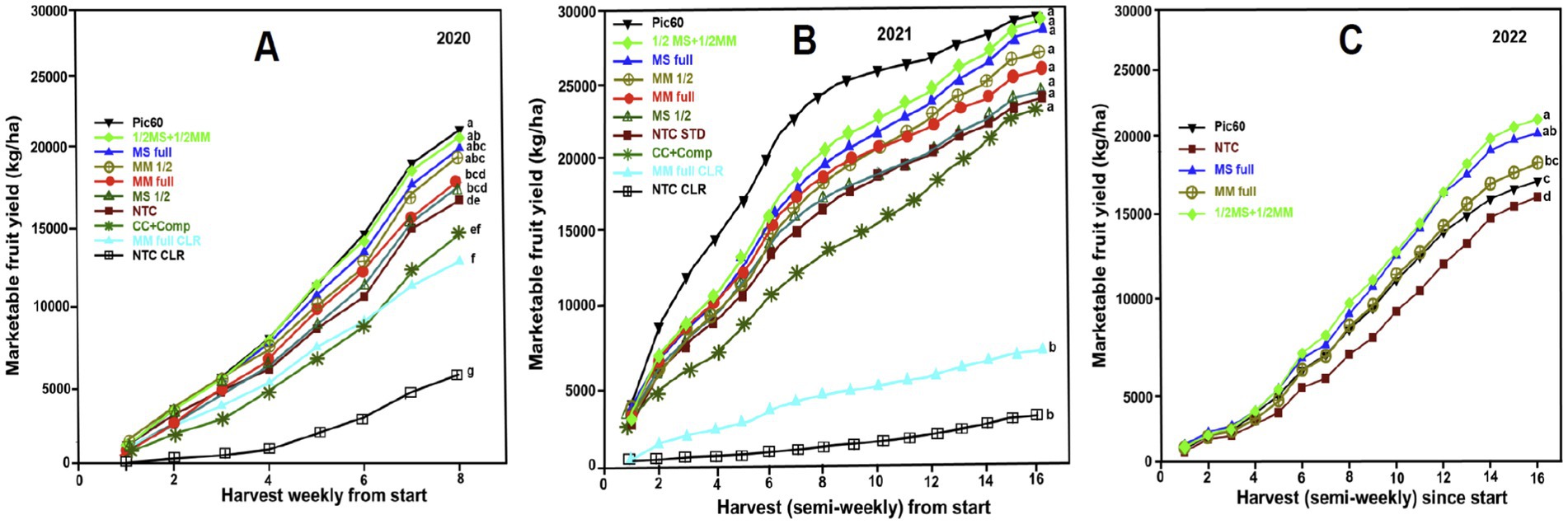
Figure 7. Impacts of organic amendments on strawberry yield in North Carolina, U. S. A. In 2020 and 2021, 10 treatments were evaluated, and each treatment was represented by a line graph and organized in the same color for each year. Acronyms for treatments are defined in materials and methods (Table 1). (A) The cumulative yield (Y-axis) was harvested over eight weekly harvests (X-axis) in 2020. As denoted by the same letter, the cumulative marketable yield progress was not significantly different based on repeated measures analysis and the Fisher Protected LSD (p ≤ 0.05). (B) The cumulative marketable yield progress curves at semi-weekly harvests in 2021 revealed that each curve when followed by the same letter, exhibited no significant difference based on repeated measures analysis and the Fisher Protected LSD (p ≤ 0.05). An independence analysis indicated a significant correlation between the 10 treatment ranking of total yield in 2020 and the yield in 2021 (Supplementary Figure S3). (C) Three promising organic amendments treatments: dried molasses (MS), mustard meal (MM), and a combination half-rate of both (½ MS + ½ MM), and the controls included non-amended control (NTC) and chemical soil fumigation with Pic-Clor 60 (PC) were selected and further evaluated in 2022. The cumulative marketable yield progress curves at semi-weekly harvests showed each amendment increased yield compared to the NTC, and yields were greater or similar to the fumigation treatment denoted by the same letter based on repeated measures analysis and the Fisher Protected LSD (p ≤ 0.05).
The experiments examine the association of bacterial communities with strawberry soil in North Carolina, USA. The results indicate that organic amendments can alter the bacterial microbiome in strawberry fields and increase crop yield. This effect selectively enriches certain bacteria populations known to be beneficial in plant production systems, aligning with previous studies that examined various organic amendments and agricultural byproducts used with anaerobic soil disinfestation (ASD) in different states across the United States (Adhikari and Louws, 2022; Adhikari et al., 2023; Bernard et al., 2014; Rosskopf et al., 2020; Strauss and Kluepfel, 2015; Testen et al., 2021). Therefore, investigating the interactions of the factors that shape the bacterial microbiome is crucial.
This study investigates the effects of two common organic amendments, molasses, and mustard meal on bacterial community composition, diversity, and strawberry yield. The results suggest that these treatments increased yields compared to the non-treated control (NTC) in two out of 3 years. They performed similarly to, or even better than, the fumigation treatment in all three years. This supports our hypothesis that organic amendments can positively impact microbial communities and enhance crop yields. A previous study demonstrated that organic amendments positively influenced bacterial diversity, which can enhance nutrient turnover and provide additional nutrients for plant growth (Fan et al., 2021). These amendments are essential for maintaining soil microbial communities and offer various benefits, including improved organic matter decomposition, enhanced soil structure, and health, increased crop yields, and suppression of soilborne pathogens (Bonilla et al., 2012; Liu et al., 2021; Luo et al., 2018; Thangarajan et al., 2013; Shu et al., 2022). Specifically, the application of mustard meal at rates of 4.4 to 6.6 tons per hectare and molasses at 20 to 25 tons per hectare has been shown to positively alter microbial communities and impact crop yields (Mazzola et al., 2015, 2018; Poret-Peterson et al., 2019; Testen and Miller, 2018).
Application different rates of the mustard meal (2.2, 4.4, and 6.6 t/ha) were compared on specific rootstock genotypes to manage apple replant disease in the Pacific region (Wang and Mazzola, 2019). Another organic amendment, dried molasses, has been applied at a rate of 20 to 22 t/ha to mitigate soil-borne pathogens in tomatoes in Florida and Ohio (Testen and Miller, 2018; Vincent et al., 2022). Liquid or dried MS has also been used at a rate of 20 to 22 t/ha in tomatoes in Florida and Ohio (Testen and Miller, 2018; Vincent et al., 2022). Although using higher rates of MS and MM has been shown to affect soil microbial communities positively (Mazzola et al., 2015; Testen and Miller, 2018; Vincent et al., 2022; Wang and Mazzola, 2019), excessive application of organic amendments can lead to increased production costs. Therefore, we determined whether dried molasses (5.60 t/ha) and mustard meal (2.24 t/ha) and a combination of half the rate of both (1:1 ratio), would still be effective in treating the bacterial communities over three years during strawberry production. These rates were lower than those used in previous studies in other states (Mazzola et al., 2015; Testen and Miller, 2018; Vincent et al., 2022). Specific bacterial and fungal phyla were found to be more abundant in MM-treated soil compared to the control (Wang and Mazzola, 2019).
Some dominant taxa, e.g., Actinobacteria and Proteobacteria, were identified in this study in response to organic amendments. The application of MS and composted poultry litter to the soil can significantly enhance the microbial communities, including actinomycetes, arbuscular mycorrhizal fungi, Gram-positive and Gram-negative bacteria, and protozoa (Welbaum et al., 2004). Additionally, MS contains free sugars that support microbial communities (Bonanomi et al., 2020) and members of the Bacteroidetes and Proteobacteria quickly flourish in nutrient-rich environments (Fierer et al., 2007). Intriguingly, the members of Firmicutes phylum exhibited low abundance in the soil initially, but it became more prevalent following all organic amendment treatments in the subsequent years. Previous investigations also suggest that members of Firmicutes were enriched in soils treated with organic amendments (Poret-Peterson et al., 2019; Strauss and Kluepfel, 2015). In contrast, soil treated with rice bran did not show the presence of the Firmicutes phylum (Mazzola et al., 2018). Field crops like rice (Edwards et al., 2015), wheat (Ai et al., 2015), and smooth cordgrass (Hong et al., 2015) have indicated an increase in the relative abundance of Proteobacteria. Evidence from several studies suggested that management practices (Edwards et al., 2015; Wipf et al., 2021), soil depths (Schlatter et al., 2018, 2020), host genotypes (Hassani et al., 2023; Peiffer et al., 2013), geographic sites (Edwards et al., 2015, 2018), and microenvironments, and soil physiochemical properties near the rhizoplane of plants play critical roles in plant-microbe associations (Singh et al., 2004; Trivedi et al., 2020). The differences in bacterial community composition observed across years in this study could be attributed to micro-environmental conditions in the soil, plant growth, and interactions between plants and microbiomes (Mowlick et al., 2013, 2014; Robinson, 2005; Stremińska et al., 2014; Stromberger et al., 2005).
Cumulative marketable yields varied considerably between years, but yields in 2020 and 2022 were lower than in 2021. Due to the COVID-19 pandemic during harvest in 2020, movement restrictions led to a lack of timely crop and pest management practices, resulting in irreversible losses at different crop stages and affecting the quantity and quality of the harvest. Variation in yield is frequently a function of fall and winter conditions that impact plant growth and yield potential (Poling, 1993). Throughout the study, only one strawberry cultivar., ‘Chandler’ was used. When plating soil suspensions on semi-selective media from various treatments, we observed several novel strains of beneficial Gram-positive and Gram-negative bacteria associated with strawberry plants. For example, one such strain is Bacillus aryabhattai, which can dissolve phosphate, synthesize siderophores, produce plant growth hormones, and enhance salt tolerance (Bhattacharyya et al., 2017; Liu et al., 2020; Park et al., 2017; Shahid et al., 2022). Although these organic amendments positively affected bacterial composition and strawberry yield, it is essential to note that the experiments were conducted in just one location. Conducting additional studies in different places in the future can help mitigate potential biases and offer a more comprehensive understanding of how microbial diversity responds to organic amendments.
In conclusion, low MS and MM rates, each at half the usual amount, can enhance beneficial bacterial microbiomes and marketable yield in field-grown strawberries. This approach can also lower production costs compared to the high rates used in other studies. These findings have practical implications for improving agricultural practices and increasing strawberry yield. Next-generation sequencing technologies and culturomics can provide further insights into microbiome functions and plant-microbiome interactions. Ultimately, these findings could lead to biologically based solutions for improving soil and plant health and increasing crop yield in conventional and organic strawberry production systems.
The original contributions presented in the study are included in the article/Supplementary material, further inquiries can be directed to the corresponding author.
TA: Conceptualization, Data curation, Formal analysis, Investigation, Methodology, Project administration, Resources, Supervision, Validation, Funding acquisition, Writing – original draft, Writing – review & editing. AP: Methodology, Writing – review & editing. FL: Funding acquisition, Writing – review & editing.
The author(s) declare that financial support was received for the research and/or publication of this article. The North Carolina Strawberry Association, the Southern Region Small Fruit Consortium, and USDA-NIFA-SCRI Grant # 2017-51181-26832 funded this study.
The authors thank John Garner and Jonathan Francki and other staff at Horticultural Crops Research Station, Castle Hayne, NC; David Baltzegar and other staff at Genomic Sciences Laboratory (GSL), North Carolina State University, Raleigh, NC; and Mohd Zubair and Cassidy Hardaway Novogene Corporation, Inc., Sacramento, CA, U. S. A. for their technical support.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declared that they were an editorial board member of Frontiers, at the time of submission. This had no impact on the peer review process and the final decision.
The author(s) declare that no Gen AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
The Supplementary material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fsufs.2025.1550106/full#supplementary-material
Abawi, G. S., and Widmer, T. L. (2000). Impact of soil health management practices on soilborne pathogens, nematodes and root diseases of vegetable crops. Appl. Soil Ecol. 15, 37–47. doi: 10.1016/S0929-1393(00)00070-6
Adhikari, T. B., and Louws, F. J. (2022). Improving soil health for strawberry production in the southeast: providing tools for growers. Project #: 2021-R-09. Submitted to the Southern Region Small Fruit Consortium (SRSFC), Raleigh, North Carolina, USA.
Adhikari, T. B., Philbrick, A., and Louws, F. J. (2023). Deciphering the active strawberry microbiome associated with roots and rhizosphere soil. The paper was presented at the 12th International Society of Plant Pathology at the Satellite event Phytobiomes Research for Plant Health, held from August 19-25, 2023, in Lyon, France.
Ahn, J.-H., Song, J., Kim, B.-Y., Kim, M.-S., Joa, J.-H., and Weon, H.-Y. (2012). Characterization of the bacterial and archaeal communities in rice field soils subjected to long-term fertilization practices. J. Microbiol. 50, 754–765. doi: 10.1007/s12275-012-2409-6
Ai, C., Liang, G., Sun, J., Wang, X., He, P., Zhou, W., et al. (2015). Reduced dependence of rhizosphere microbiome on plant-derived carbon in 32-year long-term inorganic and organic fertilized soils. Soil Biol. Biochem. 80, 70–78. doi: 10.1016/j.soilbio.2014.09.028
Bandara, B., and Janaranjana, S. (2021). Understanding the economic sustainability of strawberry farming in North Carolina. Int. J. Food Agric. Eco. 9, 191–202. doi: 10.2139/ssrn.4319525
Bebber, D. P., and Richards, V. R. (2020). A meta-analysis of the effect of organic and mineral fertilizers on soil microbial diversity. Ecology. 175:104450. doi: 10.1101/2020.10.04.325373
Bender, S. F., Wagg, C., and van der Heijden, M. G. A. (2016). An underground revolution: biodiversity and soil ecological engineering for agricultural sustainability. Trends Ecol. Evol. 31, 440–452. doi: 10.1016/j.tree.2016.02.016
Bernard, E., Larkin, R. P., Tavantzis, S., Erich, M. S., Alyokhin, A., and Gross, S. (2014). Rapeseed rotation, compost, and biocontrol amendments reduce soilborne diseases and increase tuber yield inorganic and conventional potato production systems. Plant Soil 374, 611–627. doi: 10.1007/s11104-013-1909-4
Bhattacharyya, C., Bakshi, U., Mallick, I., Mukherji, S., Bera, B., and Ghosh, A. (2017). Genome-guided insights into the plant growth promotion capabilities of the physiologically versatile Bacillus aryabhattai strain AB211. Front. Microbiol. 8:41. doi: 10.3389/fmicb.2017.00411
Bokulich, N. A., Subramanian, S., Faith, J. J., Gevers, D., Gordon, J. I., Knight, R., et al. (2013). Quality filtering vastly improves diversity estimates from Illumina amplicon sequencing. Nat. Methods 10, 57–59. doi: 10.1038/nmeth.2276
Bonanomi, G., De Filippis, F., Zotti, M., Idbella, M., Cesarano, G., Al-Rowaily, S., et al. (2020). Repeated applications of organic amendments promote beneficial microbiota, improve soil fertility, and increase crop yield. Appl. Soil Ecol. 156:103714. doi: 10.1016/j.apsoil.2020.103714
Bonilla, N., Gutiérrez-Barranquero, J. A., De Vicente, A., and Cazorla, F. M. (2012). Enhancing soil quality and plant health through suppressive organic amendments. Diversity 4, 475–491. doi: 10.3390/d4040475
Bulgarelli, D., Garrido-Oter, R., Münch, P. C., Weiman, A., Dröge, J., Pan, Y., et al. (2015). Structure and function of the bacterial root microbiota in wild and domesticated barley. Cell Host Microbe 17, 392–403. doi: 10.1016/j.chom.2015.01.011
Caporaso, J. G., Kuczynski, J., Stombaugh, J., Bittinger, K., Bushman, F. D., Costello, E. K., et al. (2010). QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336. doi: 10.1038/nmeth.f.303
Caporaso, J. G., Lauber, C. L., Walters, W. A., Berg-Lyons, D., Lozupone, C. A., Turnbaugh, P. J., et al. (2011). Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc. Natl. Acad. Sci. USA 108, 4516–4522. doi: 10.1073/pnas.1000080107
Cole, J. R., Wang, Q., Fish, J. A., Chai, B., McGarrell, D. M., Sun, Y., et al. (2014). Ribosomal database project: data and tools for high throughput rRNA analysis. Nucleic Acids Res. 42, D633–D642. doi: 10.1093/nar/gkt1244
Daugovish, O., Muramoto, J., Shennan, C., and Zavatta, M. (2021). Plant-derived carbon sources for anaerobic soil disinfestation in Southern California. Glob. J. Agric. Innov. Res. Dev. 8, 169–175. doi: 10.15377/2409-9813.2021.08.13
Daugovish, O., Valdes-Berriz, M., Muramoto, J., Shennan, C., Zavatta, M., and Henry, P. (2023). Carbon sources for anaerobic soil disinfestation in Southern California strawberry. Agronomy 13:1635. doi: 10.3390/agronomy13061635
Deng, S., Wipf, H. M. L., Pierroz, G., Raab, T. K., Khanna, R., and Coleman-Derr, D. (2019). A plant growth-promoting microbial soil amendment dynamically alters the strawberry root bacterial microbiome. Sci. Rep. 9:17677. doi: 10.1038/s41598-019-53623-2
Edgar, R. C. (2004). MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797. doi: 10.1093/nar/gkh340
Edgar, R. (2013). UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat. Methods 10, 996–998. doi: 10.1038/nmeth.2604
Edgar, R. C., Haas, B. J., Clemente, J. C., Quince, C., and Knight, R. (2011). UCHIME improves the sensitivity and speed of chimera detection. Bioinformatics 27, 2194–2200. doi: 10.1093/bioinformatics/btr381
Edwards, J., Johnson, C., Santos-Medellín, C., Lurie, E., Podishetty, N. K., Bhatnagar, S., et al. (2015). Structure, variation, and assembly of the root-associated microbiomes of rice. Proc. Natl. Acad. Sci. USA 112, 911–920. doi: 10.1073/pnas.1414592112
Edwards, J. A., Santos-Medellín, C. M., Liechty, Z. S., Nguyen, B., Lurie, E., Eason, S., et al. (2018). Compositional shifts in root-associated bacterial and archaeal microbiota track the plant life cycle in field-grown rice. PLoS Biol. 16, 1–28. doi: 10.1371/journal.pbio.2003862
Expósito, A., García, S., Giné, A., Escudero, N., Herranz, S., Pocurull, M., et al. (2022). Effect of molasses application alone or combined with Trichoderma asperellum T-34 on Meloidogyne spp. management and soil microbial activity in organic production systems. Agronomy 12:1508. doi: 10.3390/agronomy12071508
Fageria, N. K., Baligar, V. C., and Bailey, B. A. (2005). Role of cover crops in improving soil and row crop productivity. Comm. Soil Sci. Plant Anal. 36, 2733–2757. doi: 10.1080/00103620500303939
Fan, K., Delgado-Baquerizo, M., Guo, X., Wang, D., Zhu, Y., and Chu, H. (2021). Biodiversity of keystone phylotypes determines crop production in a 4-decade fertilization experiment. ISME J. 15, 550–561. doi: 10.1038/s41396-020-00796-8
Feng, M., Adams, J. M., Fan, K., Shi, Y., Sun, R., Wang, D., et al. (2018). Long-term fertilization influences community assembly processes of soil diazotrophs. Soil Biol. Biochem. 126, 151–158. doi: 10.1016/j.soilbio.2018.08.021
Fierer, N., Bradford, M. A., and Jackson, R. B. (2007). Toward an ecological classification of soil bacteria. Ecology 88, 1354–1364. doi: 10.1890/05-1839
Goss, M. J., Tubeileh, A., and Goorahoo, D. (2013). A review of the use of organic amendments and the risk to human health. Adv. Agron. 120, 275–379. doi: 10.1016/B978-0-12-407686-0.00005-1
Gravuer, K., Gennet, S., and Throop, H. L. (2019). Organic amendment additions to rangelands: a meta-analysis of multiple ecosystem outcomes. Glob. Chang. Biol. 25, 1152–1170. doi: 10.1111/gcb.14535
Hammer, O., Harper, D. A., and Ryan, P. D. (2001). PAST: paleontological statistics software package for education and data analysis. Palaeontol. Electron. 4:9. Available at: http://palaeo-electronica.org/2001_1/past/issue1_01.htm
Hannum, S. M. (2004). Potential impact of strawberries on- human health: a review of the science. Crit. Rev. Food Sci. Nutr. 44, 1–17. doi: 10.1080/10408690490263756
Hassani, M.-A., Gonzalez, O., Hunter, S. S., Holmes, G., Hewavitharana, S., Ivors, K., et al. (2023). Microbiome network connectivity and composition linked to disease resistance in strawberry plants. Phytobiomes 7, 298–311. doi: 10.1094/PBIOMES-10-22-0069-R
Heberle, H., Meirelles, G. V., da Silva, F. R., Telles, G. P., and Minghim, R. (2015). Interacti Venn: a web-based tool for the analysis of sets through Venn diagrams. BMC Bioinform. 16, 1–7. doi: 10.1186/s12859-015-0611-3
Hewavitharana, S. S., Klarer, E., Reed, A. J., Leisson, R., Poirier, B., Honaas, L., et al. (2019). Temporal dynamics of the soil metabolome and microbiome during simulated anaerobic soil disinfestation. Front. Microbiol. 10:2365. doi: 10.3389/fmicb.2019.02365
Hewavitharana, S. S., Ruddell, D., and Mazzola, M. (2014). Carbon source-dependent antifungal and nematicidal volatiles derived during anaerobic soil disinfestation. Eur. J. Plant Pathol. 140, 39–52. doi: 10.1007/s10658-014-0442-5
Hoffmann, M., Volk, E., and Peres, N. (2021). Strawberry plasticulture producers guide. Gen. Tech. Rep. SRS-260r. Asheville, NC: U.S. Department of Agriculture Forest Service, Southern Research Station. 53 p. Nulty, Steven; Gavazzi, Michael, eds. Hurricane preparation and recovery in the Southeastern United States.
Hong, Y., Liao, D., Hu, A., Wang, H., Chen, J., Khan, S., et al. (2015). Diversity of endophytic and rhizoplane bacterial communities associated with exotic Spartina alterniflora and native mangrove using Illumina amplicon sequencing. Can. J. Microbiol. 61, 723–733. doi: 10.1139/cjm-2015-0079
Janssen, P. H. (2006). Identifying the dominant soil bacterial taxa in libraries of 16S rRNA and 16S rRNA genes. Appl. Environ. Microbiol. 72, 1719–1728. doi: 10.1128/AEM.72.3.1719-1728.2006
Klindworth, A., Pruesse, E., Schweer, T., Peplies, J., Quast, C., Horn, M., et al. (2013). Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 41:e1. doi: 10.1093/nar/gks808
Larkin, R. P. (2015). Soil health paradigms and implications for disease management. Annu. Rev. Phytopathol. 53, 199–221. doi: 10.1146/annurev-phyto-080614-120357
Li, P., Li, Y., Xu, L., Zhang, H., Shen, X., Xu, H., et al. (2021). Crop yield-soil quality balance in double cropping in China’s upland by organic amendments: a meta-analysis. Geoderma 403:115197. doi: 10.1016/j.geoderma.2021.115197
Li, W., and Liu, Q. (2019). Changes in fungal community and diversity in strawberry rhizosphere soil after 12 years in the greenhouse. J. Integ. Agric. 18, 677–687. doi: 10.1016/S2095-3119(18)62003-9
Liu, H., Brettell, L. E., Qiu, Z., and Singh, B. K. (2020). Microbiome-mediated stress resistance in plants. Trends Plant Sci. 25, 733–743. doi: 10.1016/j.tplants.2020.03.014
Liu, S., Khan, M. H., Yuan, Z., Hussain, S., Cao, H., and Liu, Y. (2021). Response of soil microbiome structure and network profiles to four soil amendments in mono-cropping strawberry greenhouse. PLoS One 16:e0245180. doi: 10.1371/journal.pone.0245180
Liu, Z., Lozupone, C., Hamady, M., Bushman, F. D., and Knight, R. (2007). Short pyrosequencing reads suffice for accurate microbial community analysis. Nucleic Acids Res. 35:e12. doi: 10.1093/nar/gkm541
Louws, F. J. (2009). “IPM for soilborne disease management for vegetable and strawberry crops in SE” in Plant pathology in the 21st century: Contributions to the 9th international congress. eds. U. Gisi, I. Chet, and M. L. Gullino (New York: Springer), 217–227.
Louws, F. J., Grabowski, M. A., Fernandez, G. E., and Vollmer, J. B. (2000). Compost as an alternative to methyl bromide in plasticulture strawberry production. Proc. Intl. Res. Conf. Methyl Bromide Alternatives and Emissions Reduction 33/1–33/2.
Luo, G., Li, L., Friman, V.-P., Guo, J., Guo, S., Shen, Q., et al. (2018). Organic amendments increase crop yields by improving microbe-mediated soil functioning of agroecosystems: a meta-analysis. Soil Biol. Biochem. 124, 105–115. doi: 10.1016/j.soilbio.2018.06.002
Matthiessen, J. N., and Kirkegaard, J. A. (2006). Biofumigation and enhanced biodegradation: opportunity and challenge in soilborne pest and disease management. Crit. Rev. Plant Sci. 25, 235–265. doi: 10.1080/07352680600611543
Mazzola, M., and Brown, J. (2010). Efficacy of brassicaceous seed meal formulations for the control of apple replant disease in organic and conventional orchard production systems. Plant Dis. 94, 835–842. doi: 10.1094/PDIS-94-7-0835
Mazzola, M., Hewavitharana, S. S., and Strauss, S. L. (2015). Brassica seed meal soil amendments transform the rhizosphere microbiome and improve apple production through resistance to pathogen reinfestation. Phytopathology 105, 460–469. doi: 10.1094/PHYTO-09-14-0247-R
Mazzola, M., Muramoto, J., and Shennan, C. (2018). In California field trials, anaerobic disinfestation induced changes to the soil microbiome, disease incidence, and strawberry fruit yields. Appl. Soil Ecol. 127, 74–86. doi: 10.1016/j.apsoil.2018.03.009
McMurdie, P. J., and Holmes, S. (2013). Phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS One 8:e61217. doi: 10.1371/journal.pone.0061217
McWhirt, A., Fernandez, G., and Schroeder-Moreno, M. (2014). Sustainable practices for Plasticulture strawberry production in the southeast. NC Cooperative Extension Publication. Available online at: http://content.ces.ncsu.edu/sustainable-practices-for-plasticulturestrawberry-production-in-the-southeast/ (Accessed September 20, 2024).
Meshram, S., Philbrick, A. N., and Adhikari, T. B. (2024). Anaerobic soil disinfestation: a biologically-based solution for sustainable crop production. Front. Hortic. 3:1436782. doi: 10.3389/fhort.2024.1436782
Mowlick, S., Inoue, T., Takehara, T., Tonouchi, A., Kaku, N., Ueki, K., et al. (2014). Usefulness of Japanese-radish residue in biological soil disinfestation to suppress spinach wilt disease accompanying with proliferation of soil bacteria in the Firmicutes. Crop Prot. 61, 64–73. doi: 10.1016/j.cropro.2014.03.010
Mowlick, S., Yasukawa, H., Inoue, T., Takehara, T., Kaku, N., Ueki, K., et al. (2013). Suppression of spinach wilt disease by biological soil disinfestation incorporated with Brassica juncea plants in association with changes in soil bacterial communities. Crop protect. 54, 185-193.
Murphy, S. (2020). FS1328 Soil for raised beds. Rutgers Cooperative Extension Fact Sheet FS1328. (Accessed February 16, 2025).
Nuova Amit (2007). Company material published in the yearbook 2007 by CIC - consortium of Italian composters. Available online at: http://compost.it/schede_soci/nuovaamit.pdf.
Oldfield, E. E., Wood, S. A., and Bradford, M. A. (2018). Direct effects of soil organic matter on productivity mirror those observed with organic amendments. Plant Soil 423, 363–373. doi: 10.1007/s11104-017-3513-5
Pankhurst, C. E. (1997). “Biodiversity of soil organisms as an indicator of soil health” in Biological indicators of soil health. eds. C. E. Pankhurst, B. M. Doube, and V. V. S. R. Gupta (Wallingford: CAB International), 297–324.
Park, Y. G., Mun, B. G., Kang, S. M., Hussain, A., Shahzad, R., Seo, C. W., et al. (2017). Bacillus aryabhattai SRB02 tolerates oxidative and nitrosative stress and promotes the growth of soybean by modulating the production of phytohormones. PLoS One 12:e0173203. doi: 10.1371/journal.pone.0173203
Paulson, J. N., Talukder, H., Pop, M., and Bravo, H. C. (2014). metagenomeSeq: Statistical analysis for sparse high-throughput sequencing. Bioconductor. Available online at http://cbcb.umd.edu/software/metagenomeSeq (Accessed November 20, 2023).
Peiffer, J. A., Spor, A., Koren, O., Jin, Z., Tringe, S. G., Dangl, J. L., et al. (2013). Diversity and heritability of the maize rhizosphere microbiome under field conditions. Proc. Natl. Acad. Sci. USA 110, 6548–6553. doi: 10.1073/pnas.1302837110
Poling, E. B. (1993). Strawberry plasticulture in North Carolina: II. Preplant, planting, and postplant considerations for growing Chandler strawberry on black plastic mulch. Hort Technol. 3, 383–393. doi: 10.21273/HORTTECH.3.4.383
Poret-Peterson, A. T., Albu, S., McClean, A. E., and Kluepfel, D. A. (2019). Shifts in soil bacterial communities as a function of carbon source used during anaerobic soil disinfestation. Front. Environ. Sci. 6:60. doi: 10.3389/fenvs.2018.00160
Poret-Peterson, A. T., Sayed, N., Glyzewski, N., Forbes, H., González-Orta, E. T., and Kluepfel, D. A. (2020). Temporal responses of microbial communities to anaerobic soil disinfestation. Microb. Ecol. 80, 191–201. doi: 10.1007/s00248-019-01477-6
Quast, C., Pruesse, E., Yilmaz, P., Gerken, J., Schweer, T., Yarza, P., et al. (2013). The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596. doi: 10.1093/nar/gks1219
R Core Team (2022). A language and environment for statistical computing. Vienna, Austria: R Foundation for Statistical Computing.
Rana, T. S., and Gu, S. (2020). Growth and yield of organic day-neutral strawberries in low tunnels inside high tunnels in North Carolina. Hort Sci. 55, 336–343. doi: 10.21273/HORTSCI14491-19
Robinson, P. J. (2005). North Carolina weather and climate. North Carolina: University of North Carolina Press, Chapel Hill, 256.
Rosskopf, E., Di Gioia, F., Hong, J. C., Pisani, C., and Kokalis-Burelle, N. (2020). Organic amendments for pathogen and nematode control. Annu. Rev. Phytopathol. 58, 277–311. doi: 10.1146/annurev-phyto-080516-035608
Rysin, O., McWhirt, A., Fernandez, G., Louws, F. J., and Schroeder-Moreno, M. (2015). Economic viability and environmental impact assessment of three different strawberry production systems in the southeastern United States. Hort Technol. 25, 585–594. doi: 10.21273/HORTTECH.25.4.585
Saleem, M., Hu, J., and Jousset, A. (2019). More than the sum of its parts: microbiome biodiversity as a driver of plant growth and soil health. Annu. Rev. Ecol. Evol. Syst. 50, 145–168. doi: 10.1146/annurev-ecolsys-110617-062605
Samtani, J. B., Rom, C. R., Friedrich, H., Fennimore, S. A., Finn, C. E., and Petran, A. (2019). The status and future of the strawberry industry in the United States. Hort Technol. 29, 11–24. doi: 10.21273/HORTTECH04135-18
SAS Institute (2016). Statistical analysis software (SAS) User’s guide version 9.4. Cary, NC, USA: SAS Institute, Inc.
Schlatter, D. C., Kahl, K., Carlson, B., Huggins, D. R., and Paulitz, T. (2018). Fungal community composition and diversity vary with soil depth and landscape position in a no-till wheat-based cropping system. FEMS Microbiol. Ecol. 94:94. doi: 10.1093/femsec/fiy098
Schlatter, D. C., Kahl, K., Carlson, B. R., Huggins, D. R., and Paulitz, T. C. (2020). Soil acidification modifies soil depth-microbiome relationships in a no-till wheat cropping system. Soil Biol. Biochem. 149:107939. doi: 10.1016/j.soilbio.2020.107939
Schloss, P. D., Westcott, S. L., Ryabin, T., Hall, J. R., Hartmann, M., Hollister, E. B., et al. (2009). Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 75, 7537–7541. doi: 10.1128/AEM.01541-09
Shahid, M., Zeyad, M. T., Syed, A., Singh, U. B., Mohamed, A., Bahkali, A. H., et al. (2022). Stress-tolerant endophytic isolate Priestia aryabhattai BPR-9 modulates physio-biochemical mechanisms in wheat (Triticum aestivum L.) for enhanced salt tolerance. Int. J. Environ. Res. Public Health 19:10883. doi: 10.3390/ijerph191710883
Shennan, C., Muramoto, J., Lamers, J., Mazzola, M., Rosskopf, E. N., Kokalis-Burelle, N., et al. (2014). Anaerobic soil disinfestation for soilborne disease control in strawberry and vegetable systems: current knowledge and future directions. Acta Hortic. 1044, 165–175. doi: 10.17660/ActaHortic.2014.1044.20
Shu, X., He, J., Zhou, Z., Xia, L., Hu, Y., Zhang, Y., et al. (2022). Organic amendments enhance soil microbial diversity, microbial functionality and crop yields: a meta-analysis. Sci. Total Environ. 829:154627. doi: 10.1016/j.scitotenv.2022.154627
Singh, B. K., Millard, P., Whiteley, A. S., and Murrell, J. C. (2004). Unraveling rhizosphere–microbial interactions: opportunities and limitations. Trends Microbiol. 12, 386–393. doi: 10.1016/j.tim.2004.06.008
Strauss, S. L., and Kluepfel, D. A. (2015). Anaerobic soil disinfestation: a chemical-independent approach to pre-plant control of plant pathogens. J. Integr. Agric. 14, 2309–2318. doi: 10.1016/S2095-3119(15)61118-2
Stremińska, M. A., Runia, W. T., Termorshuizen, A. J., Feil, H., and van der Wurff, A. W. G. (2014). Anaerobic soil disinfestation in microcosms of two sandy soils. Commun. Agric. Appl. Biol. Sci. 79, 15–19
Stromberger, M. E., Klose, S., Ajwa, H., Trout, T., and Fennimore, S. (2005). Microbial populations and enzyme activities in soils fumigated with methyl bromide alternatives. Soil Sci. Soc. Am. 69, 1987–1999. doi: 10.2136/sssaj2005.0076
Sydorovych, O., Safley, C. D., Ferguson, L. M., Poling, E. B., Fernandez, G. E., Brannen, P. M., et al. (2006). Economic evaluation of the methyl bromide alternatives for the production of strawberries in the southeastern US. Hort Technol. 16, 118–128. doi: 10.21273/HORTTECH.16.1.0118
Testen, A. L., and Miller, S. A. (2018). Carbon source and soil origin shape soil microbiomes and tomato soilborne pathogen populations during anaerobic soil disinfestation. Phytobiomes 2, 138–150. doi: 10.1094/PBIOMES-02-18-0007-R
Testen, A. L., Rotondo, F., Mills, M. P., Horvat, M. M., and Miller, S. A. (2021). Evaluation of agricultural byproducts and cover crops as anaerobic soil disinfestation carbon sources for managing a soilborne disease complex in high tunnel tomatoes. Front. Sustain. Food Syst. 5:645197. doi: 10.3389/fsufs.2021.645197
Thangarajan, R., Bolan, N. S., Tian, G., Naidu, R., and Kunhikrishnan, A. (2013). Role of organic amendment application on greenhouse gas emission from soil. Sci. Total Environ. 465, 72–96. doi: 10.1016/j.scitotenv.2013.01.031
Trivedi, P., Leach, J. E., Tringe, S. G., Sa, T., and Singh, B. K. (2020). Plant–microbiome interactions: from community assembly to plant health. Nat. Rev. Microbiol. 18, 607–621. doi: 10.1038/s41579-020-0412-1
Vincent, I. R., Paudel, B. R., Guo, H., Rosskopf, E. N., Di Gioia, F., Hong, J. C., et al. (2022). Spatial and temporal changes of soil microbial communities in field tomato production as affected by anaerobic soil disinfestation. Front. Sustain. Food Syst. 6:838635. doi: 10.3389/fsufs.2022.838635
Wagner, B. D., Robertson, C. E., and Harris, J. K. (2011). Application of two-part of statistics for comparison of sequence variant counts. PLoS One 6:e20296. doi: 10.1371/journal.pone.0020296
Wang, L., and Mazzola, M. (2019). Field evaluation of reduced rate Brassicaceae seed meal amendment and rootstock genotype on the microbiome and control of apple replant disease. Phytopathology 109, 1378–1391. doi: 10.1094/PHYTO-02-19-0045-R
Welbaum, G. E., Sturz, A. V., Dong, Z., and Nowak, J. (2004). Managing soil microorganisms to improve productivity of agro-ecosystems. Crit. Rev. Plant Sci. 23, 175–193. doi: 10.1080/07352680490433295
Welker, R. M., Louws, F. J., Driver, J. G. Smith, and Schiemann, C. A. (2008). On-farm research and extension activities to implement MB alternatives: an area-wide initiative update. Proc. of the Int. Res. Conf. on Methyl Bromide Alternatives and Emissions Reduction. 9/1–9/3. Available online at: http://mbao.org/2008/Proceedings/009WelkerRMBAOAREAWIDEFJL.pdf (Accessed July 12, 2023).
Wipf, H. M. L., Xu, L., Gao, C., Spinner, H. B., Taylor, J., Lemaux, P., et al. (2021). Agricultural soil management practices differentially shape the bacterial and fungal microbiomes of Sorghum bicolor. Appl. Environ. Microbiol. 87, e02345–e02320. doi: 10.1128/AEM.02345-20
Xia, Q., Rufty, T., and Shi, W. (2020). Soil microbial diversity and composition: links to soil texture and associated properties. Soil Biol. Biochem. 149:107953. doi: 10.1016/j.soilbio.2020.107953
Yang, H., Zhang, X., Qiu, X., Chen, J., Wang, Y., Zhang, G., et al. (2023). Fusarium wilt invasion results in a strong impact on strawberry microbiomes. Plants (Basel). 12:4153. doi: 10.3390/plants12244153
Keywords: Fragaria × ananassa, microbiome, soil, 16S rRNA, organic amendments, strawberry
Citation: Adhikari TB, Philbrick AN and Louws FJ (2025) Elucidating the impact of organic amendments on soil bacterial communities and strawberry yield in North Carolina. Front. Sustain. Food Syst. 9:1550106. doi: 10.3389/fsufs.2025.1550106
Received: 22 December 2024; Accepted: 25 February 2025;
Published: 19 March 2025.
Edited by:
Marouane Baslam, Niigata University, JapanReviewed by:
Raj Majumdar, Agricultural Research Service (USDA), United StatesCopyright © 2025 Adhikari, Philbrick and Louws. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Tika B. Adhikari, dGJhZGhpa2FAbmNzdS5lZHU=
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.