
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Sustain. Food Syst. , 15 October 2020
Sec. Sustainable Food Processing
Volume 4 - 2020 | https://doi.org/10.3389/fsufs.2020.550318
This article is part of the Research Topic Microbiological Risks in Food Processing View all 8 articles
Resistance to last-resort antibiotics is significant public health issue. Antibiotic use in animal husbandry may be a driver of resistance that can subsequently be disseminated via the food chain. This study sought to determine the level of polymyxin resistance in Gram-negative pathogens present in Australian poultry, particularly the presence of mobilizable mechanisms of polymyxin resistance. Cloacal swabs from 213 birds were taken in a point prevalence survey from six different farms at a Victorian chicken processing facility. Colistin resistant organisms were recovered by direct plating on CHROMagar COL-APSE media. Bacterial isolates were identified and analyzed by MALDI-TOF, biochemical and genotypic assays. The 213 specimens yielded 57 (26.8%) colistin-resistant Gram-negative organisms, all of which have been previously described as exhibiting intrinsic resistance to polymyxin antibiotics. The most frequent organism was identified as Hafnia paralvei (40/57; 70%). Other colistin-resistant organisms included Aeromonas hydrophila (16%), Myroides odoratus (7%), Alcaligenes faecalis (5%), and Pseudochrobactrum spp. (2%). No mobile colistin resistance (mcr) genes were detected, although the arnA gene was identified in two A. hydrophila isolates and may mediate colistin resistance in these isolates. Intrinsic polymyxin-resistant organisms are widely distributed in the food chain, with over a quarter of the birds tested yielding a polymyxin-resistant organism. However, strains containing mcr genes remain rare in Australian poultry.
Polymyxin B and E (colistin) are cationic antimicrobial peptides currently reserved as a last resort treatment for serious infections caused by multi-drug resistant Gram-negative bacteria (Poirel et al., 2017). Although colistin has been used in human and veterinary medicine for over 50 years, it has recently become increasingly important in the treatment of infections with carbapenem-resistant organisms where few treatment options now remain. Enterobacteriaceae such as Serratia spp. and Proteus spp., as well as Gram-negative cocci such as Neisseria are well-known to be intrinsically resistant to colistin. However, less has been documented about other intrinsically colistin-resistant organisms in the food chain. The increased reliance on this agent has forced us to reconsider how polymyxins are used, particularly in light of the emergence of mobilisable colistin resistance.
The first plasmid mediated mechanism of polymyxin resistance (MCR-1) was identified in China (Liu et al., 2016) and since this initial observation it has been identified globally (Arcilla et al., 2016; Wang et al., 2018). The mcr-1 gene encodes a phosphorylethanolamine transferase enzyme (PEtN) that modifies the lipopolysaccharide (LPS) in the outer membrane of Gram-negative bacteria and reduces the binding of polymyxins and other cationic peptides. Plasmid mediated colistin resistance genes (mcr-like) have been reported in Enterobacteriales from humans, food, food-producing animals and the environment (Wang et al., 2018). Often the discovery of these genes in food and food animals proceeds their identification in humans (Liu et al., 2016). Colistin has also been used widely in animal rearing in many countries, and therefore may provide a selective pressure for the emergence of polymyxin resistance. This has led to some countries (China and Brazil) to ban the use of colistin in agriculture. To the best of our knowledge, polymyxins are not routinely used in animal production systems in Australia, although they may be used therapeutically in cattle and sheep (Australian Pesticides Veterinary Medicines Authority, 2014). A position statement form the Australian Chicken Meat Federation (ACMF) states that polymyxins have never been approved for use in Australian poultry, and therefore should never have been used in Australian chicken flocks (Australian Chicken Meat Federation, 2018). Consequently, little has been documented in Australia about whether there in colistin resistance in poultry, or more broadly within the food chain.
The current study sought to determine if polymyxin resistant pathogens, are present in the Australian food chain. Poultry were selected as there are many reports of colistin-resistant organisms associated with poultry (Ling et al., 2017; Monte et al., 2017; Apostolakos and Piccirillo, 2018). Birds were screened for colistin-resistant organism using a recently described bacterial culture media; CHROMagar COL-APSE (Abdul Momin et al., 2017). CHROMagar COL-APSE is a selective medium for the isolation and differentiation of colistin-resistant Gram-negative bacteria. The utility of this media for prospective screening of veterinary specimens is also defined.
Poultry were sampled immediately post-slaughter on two separate visits to the abattoir in Victoria, Australia during December 2016. On each trip, birds from three different farms were sampled (Farms A–C and D–F, respectively). With the exception of farm A (n = 29 birds), a minimum of 30 birds were sampled with from each farm. Swabs (Copan, Brescia, Italy) were taken from poultry by insertion in the cloaca and placed in Aimes transport media with charcoal before being returned to the laboratory for processing. All birds sampled were indoor raised (not free range) meat chickens.
CHROMagar COL-APSE media (CHROMagar, Paris) was prepared from dehydrated powders, autoclaved, and quality controlled locally. Swabs were plated directly onto CHROMagar COL-APSE and incubated at 37°C for 24 h to screen for the carriage of polymyxin resistant Gram-negative bacteria present in the avian flora. The media was prepared with the addition of 50 mg/l 1 para-nitrophenyl glycerol (PNPG) to inhibit swarming of Proteus spp. Colistin resistant colonies were sub-cultured back on CHROMagar COL-APSE before long-term storage in brain-heart infusion broth (Oxoid, Basingstoke, UK) supplemented with 10% glycerol at −80°C.
All presumptive colistin-resistant bacteria were identified by matrix assisted laser desorption ionization-time of flight mass spectrometry (MALDI-TOF MS, Bruker, Coventry, UK). Single colonies were spotted in triplicate onto the target plate and overlaid with 1 μl α-cyano-4-hydroxycinnamic-acid matrix (10 mg/ml). Target plates were analyzed using the Bruker Microflex MALDI-TOF MS running Biotyper v2. The database used was Version 3.0.2.0. Results with a score value of <2.0 were discarded. A subset of isolates (including those which were not identified by the MALDI-TOF) had their identity confirmed by PCR amplification and DNA sequencing of their 16s rRNA genes (Weisburg et al., 1991). Biochemical testing including malonate assimilation and β-glucosidase activity were performed as described elsewhere (Huys et al., 2010).
Susceptibility to 11 antibiotic agents including, amoxicillin/clavulanic acid (30 μg), cephalexin (30 μg), cefpodoxime (10 μg), imipenem (10 μg), gentamicin (10 μg), streptomycin (10 μg), enrofloxacin (5 μg), nalidixic acid (30 μg), chloramphenicol (30 μg), doxycycline (30 μg), and cotrimoxazole (25 μg) were assessed in disc diffusion assays on Mueller-Hinton agar (Oxoid, Basingstoke, UK). The Clinical Laboratory Standards Institute (CLSI) breakpoints were used to define resistance where available (Clinical Laboratory Standards Institute, 2018). In the absence of guidelines for interpretation, the CLSI guidelines for Enterobacteriaceae were used. The minimum inhibitory concentrations (MICs) of colistin and polymyxin B were determined by agar dilution.
Genes encoding colistin resistance genes mcr-1, -2, -3,-4, and -5 were amplified using the polymerase chain reaction and primers and conditions described elsewhere (Liu et al., 2016; Xavier et al., 2016; Yin et al., 2017; Chen et al., 2018).
Selected isolates were further investigated by DNA sequencing using the Minion platform (Oxford Nanopore Technologies). The Rapid Sequencing Kit (SQK-RAD004) was used as per the manufacturer's instructions and the resulting reads analyzed by EPI2ME and the online AMR workflow.
Whole Genome Sequencing was performed on the Illumina HiSeq platform (Illumina, Inc., San Diego, CA) and the closest reference genome was identified using Kraken. The reads were mapped to the reference genome using Burrows-Wheeler aligner “mem” (BWA-mem) algorithm version 2. A de novo assembly of the reads was performed using SPAdes version 3.7.1, and the reads were again mapped back to the resultant contigs using BWA-mem. Annotation was performed by the NCBI Prokaryote Genome Annotation Pipeline (PGAP) version 4.6.
A total of 213 birds from six different farms were sampled at a chicken processing facility. From these specimens, 57 (26.8%) yielded colistin-resistant Gram-negative organisms (Table 1). The most commonly (40/57; 70.2%) recovered organism was identified as Hafnia alvei. Other organisms which were recovered on CHROMagar COL-APSE media included Aeromonas hydrophila (n = 9; 16.1%), Alcaligenes faecalis (n = 3; 5.3%), and Myroides odoratus (n = 4; 7.1%). A single isolate could not be identified by the Bruker MALDI-TOF system. Sequencing of the 16 rRNA gene identified the isolate as Pseudochrobactrum spp. The growth characteristics of each of these organisms on CHROMagar COL-APSE media is demonstrated in Figure 1.
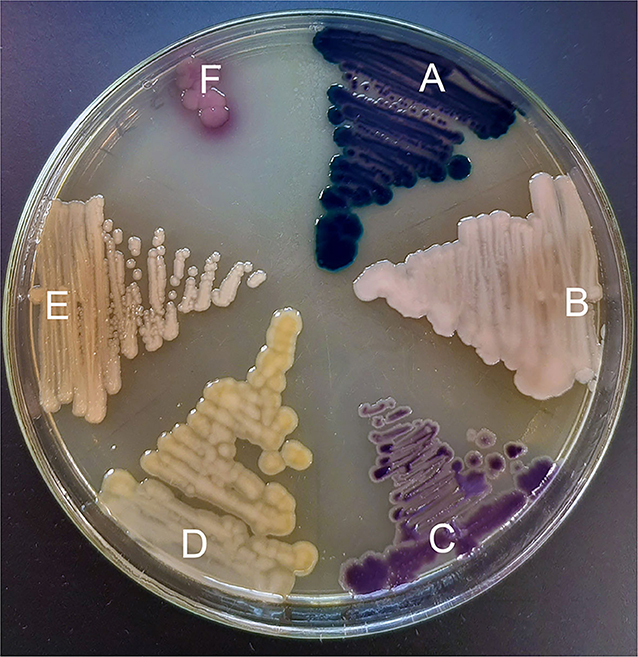
Figure 1. Colistin-resistant organisms grown on CHROMagar COL-APSE agar for 48 h; (A) Aeromonas hydrophila isolate C17; (B) Alcaligenes faecalis isolate F12; (C) Hafnia paralvei isolate E14; (D) Myroides odoratus isolate B27; (E) Pseudochrobactrum spp. isolate F17; (F) E. coli ATCC 25922.
The Hafnia isolates had an MIC90 for colistin and polymyxin B of 8 and 16 μg/ml, respectively (Table 2). Resistance to other antimicrobials was low among the Hafnia isolates, with the exception of AmpC mediated resistance to β-lactams (Table 3). As others have documented, the Bruker MALDI-TOF system cannot currently distinguish between the two Hafnia species: H. alvei and H. paralvei (Jayol et al., 2017). To determine which species these isolates belonged to, phenotypic testing using malonate assimilation and β-glucosidase activity was performed. These two tests have been shown to reliably discriminate between the two Hafnia species (Huys et al., 2010; Abbott et al., 2011). Based upon these biochemical tests all the Hafnia isolates were determined to be H. paralvei. To verify this a single isolate was submitted for whole genome sequencing which agreed with the phenotypic identification. This Whole Genome Shotgun project has been deposited at DDBJ/ENA/GenBank under the accession JAAIKS000000000. The version described in this paper is version JAAIKS010000000.
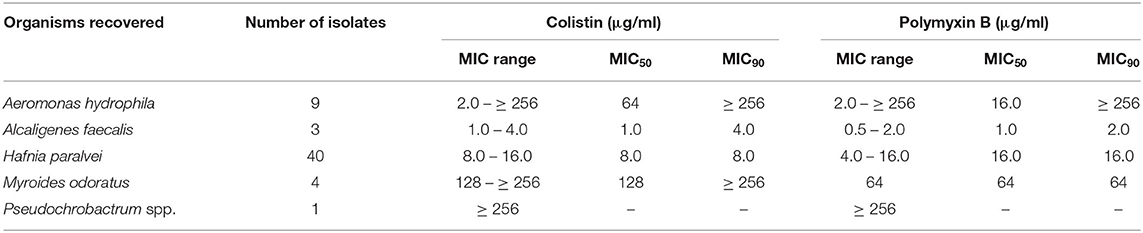
Table 2. Colistin and polymyxin B MICs of colistin-resistant organisms recovered from Victorian poultry farms.
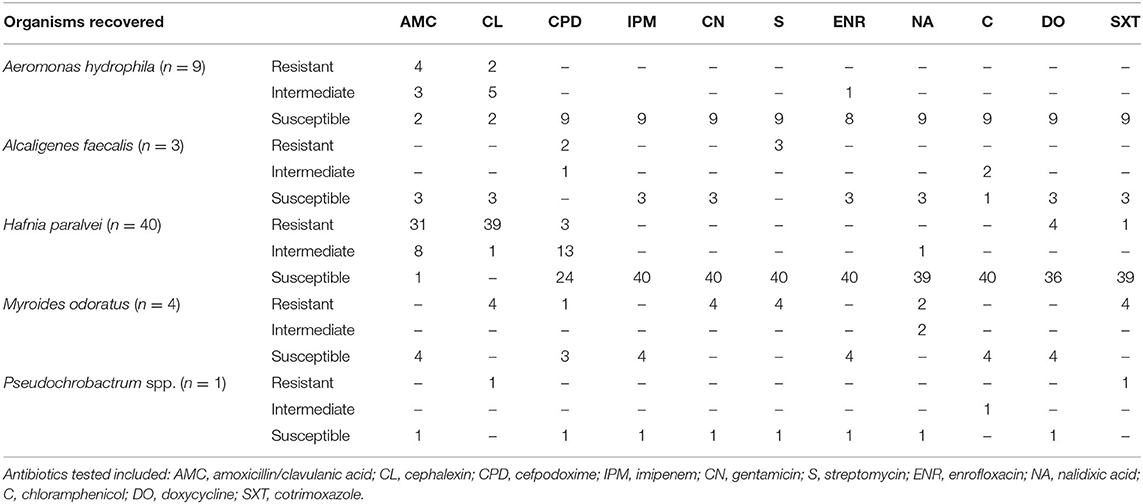
Table 3. Susceptibility to non-polymyxin antibiotics among polymyxin-resistant organisms recovered from Victorian poultry farms.
The level of polymyxin resistance varied among the other bacterial species: A. faecalis isolates had colistin and polymyxin B MICs of 1–4 μg/ml and 0.5–2 μg/ml, respectively. M. odoratus and Pseudochrobactrum spp. isolates had higher MICs of ≥128 μg/ml for colistin and ≥64 μg/ml for polymyxin B. Polymyxin MICs for A. hydrophila were the most variable ranging from 2 to ≥256 μg/ml for both colistin, and polymyxin B. Among these bacteria, Aeromonas species have previously been shown to harbor mcr-3 and mcr-5 like resistance genes (Ling et al., 2017; Ma et al., 2018). All nine isolates were screened by PCR for the presence of mcr-1, -2, -3,−4, and -5 genes and all were negative. Two isolates (A17 and C17, from farm A and C, respectively) were further investigated by MinION DNA sequencing. These isolates were chosen as they represent two different polymyxin resistance phenotypes (colistin/polymyxin B MICs of ≥256/≥256 μg/ml and 64/8 μg/ml, respectively). This revealed the presence of an arnA-like gene, which encodes a UDP-glucuronic acid (UDP-GlcUA) decarboxylase which may contribute toward their polymyxin resistance. This was the only gene identified in the AMR workflow that might contribute to polymyxin resistance.
While polymyxins are not routinely used in poultry production in Australia, several other classes of antimicrobial are used, primarily for the control of coccidiosis (Australian Pesticides Veterinary Medicines Authority, 2014). To assess if intrinsic colistin resistance is associated with resistance to other antibiotics, all isolates were tested against 12 unrelated antimicrobials (Table 3). The isolates recovered were generally susceptible to most other antibiotic classes (β-lactams, quinolones, aminoglycosides), with the notable exception of M. odoratus which was resistant to all the aminoglycosides tested as well as cephalexin, cotrimoxazole and nalidixic acid.
Polymyxin resistant organisms were recovered from Australian poultry samples, however all the organisms recovered have been previously described as intrinsically resistant to polymyxins. A notable exception is A. hydrophila, in which mcr genes have been previously detected, however no mcr-like genes were detected in the current study.
Colistin resistance, both acquired and intrinsic, is thought to occur by modification of the Gram-negative cell wall. Acquired colistin resistance encoded by mcr genes imparts resistance by catalyzing the addition of phosphoethanolamine (PEtN) to lipid A reducing the overall negative charge of the cell wall and therefore the binding affinity to polymyxins. LPS modification is also responsible for intrinsic polymyxin resistance in several bacterial species due to 4-amino-4-deoxy-L-arabinose (L-Ara4N) substitution (Olaitan et al., 2014). However, mechanisms of intrinsic colistin resistance remain unelucidated for many bacterial species.
CHROMagar COL-APSE media was successfully used for the detection and isolation of polymyxin-resistant organisms from veterinary specimens. Moreover, the media was suitable for direct inoculation from veterinary swabs. This media has been shown to be highly sensitive in the detection of organisms with acquired colistin resistance (Abdul Momin et al., 2017) and this current study further validates the utility of CHROMagar COL-APSE as a media for detection of organisms with intrinsic colistin-resistance.
CHROMagar COL-APSE performed well in the recovery of colistin-resistant organisms, including swabs with heavy fecal contamination. The ubiquity of Proteus, a bacterial genus intrinsically resistant to polymyxins, within the samples prompted the addition of the reagent PNPG. Proteus presents as a characteristic brown colony on CHROMagar COL-APSE media and its swarming motility quickly obscured other growth on the media. PNPG abolishes the swarming behavior of Proteus facilitating identification of other resistant organisms. Recent formulations of CHROMagar COL-APSE have found the addition of cefixime to be useful for the suppression of Proteus.
Of the polymyxin-resistant organisms recovered from Australian poultry Hafnia spp. were predominant. This genus is known to exhibit intrinsic resistance to polymyxins, and sequencing of 16S rRNA genes has revealed Hafnia to be phylogenetically related to other intrinsically colistin resistant members of the Enterobacteriales including Serratia spp. (Jayol et al., 2017). H. alvei was found as the most commonly isolated colistin-resistant organism from livestock in Switzerland, although it was not recovered from poultry, only from pigs and calves (Buess et al., 2017). Hafnia paralvei was only recently recognized as a distinct species (Huys et al., 2010). The H. paralvei isolates in the current study had median colistin and polymyxin B MICs of 8 and 16 μg/ml, comparable to what has been determined for the wild type population of this species elsewhere (Jayol et al., 2017). Little is known of the importance of H. paralvei as a veterinary pathogen. The biochemical characteristics of H. paralvei match those previously described as H. alvei genetic group 2: the same study, examining the ecology of Hafnia in Australia, found genetic group 1 was most commonly associated with birds (Okada and Gordon, 2003).
Aeromonas is a bacterial genera also considered intrinsically resistant to polymyxins. One Australian study found 44.5% of 193 Aeromonas isolates to have a colistin MIC ≥2 μg/ml by agar dilution testing (Aravena-Roman et al., 2012). More recently, Aeromonas spp. have been reported to carry acquired colistin resistance gene variants: mcr-3 has been detected in several Aeromonas species (Ling et al., 2017; Shen et al., 2018) and mcr-5 has been detected in A. hydrophila (Ma et al., 2018). Additionally, A. hydrophila appears to encompass a wide distribution of colistin MICs, with one collection of (78 isolates) ranging from <0.25 to >256 μg/ml by microbroth dilution testing after low-dose colistin induction (Fosse et al., 2003). The isolates described in our study fall within this MIC range, but were not shown to harbor any mcr-1, -2, -3,-4, or -5 like genes. DNA sequencing of two isolates identified the gene arnA, which may play a role in the polymyxin resistance observed in these isolates. ArnA is the first enzyme specific to the pathway which imparts polymyxin resistance by modification of lipid A of the outer membrane LPS with a 4-amino-4-deoxy-L-arabinose moiety (Breazeale et al., 2005). ArnA is bifunctional enzyme which catalyses firstly the decarboxylation of UDP-Glucuronic acid (UDP-GlcA) and subsequently formylates UDP-Ara4N (Gatzeva-Topalova et al., 2005).
Little has been documented about the MIC range and mechanisms of polymyxin resistance in the three other Gram-negative species identified: A. faecalis, M. odoratus, and Pseudochrobactrum. A study by Jacquier et al. found six of seven (86%) French isolates of A. faecalis had a colistin MIC ≥2 μg/ml (Jacquier et al., 2012), consistent with the susceptibility of the strains recovered here. M. odoratus (formerly Flavobacterium odoratum) is a Gram-negative non-fermentative opportunistic pathogen reportedly intrinsically resistant to many antibiotics, including polymyxins (Holmes et al., 1979). Colistin MICs for wild-type M. odoratus have previously been quantified at 32–256 μg/ml, although the mechanism of resistance has not been elucidated (Holmes et al., 1979; Gunzer et al., 2018). The genus Pseudochrobactrum was only described in 2006 (Kämpfer et al., 2006), and little is known of its clinical significance and associated antibiotic susceptibility.
The majority of the intrinsically polymyxin-resistant organisms identified in the current study have also well documented as meat spoilage organisms. Hafnia spp. in particular are important in spoilage of many different meat products including chicken meat (Säde et al., 2013; Höll et al., 2016), beef (Doulgeraki et al., 2011), and lamb (Kaur et al., 2017). Aeromonas spp. have also been associated with spoilage of poultry meat (Zhang et al., 2012). This implies that the polymyxin-resistant organisms survive poultry processing and will disseminate down the food chain, where traits could be passed on to human pathogens (Teale, 2002). Importantly, little is known about the mechanism of resistance in these organisms, and therefore the potential for the “escape” of resistance genes by horizontal gene transfer. There is a precedent for this: it has been hypothesized that the mcr-3 gene, for example, may have originated in Aeromonas species, and is now widespread in Enterobacteriaceae (Shen et al., 2018).
Colistin is not registered for use in any species of animal in Australia (Animal Health Australia, 2017; Australian Chicken Meat Federation, 2018). Acquired polymyxin-resistance remains rare in Australia with only a single report in humans (Ellem et al., 2017) and another in silver gulls (Mukerji et al., 2019). Despite this, the current study shows that intrinsically colistin resistant organisms are present in the food chain, particularly in food spoilage organisms. CHROMagar COL-APSE was a useful media for the recovery of these organisms. It remains to be determined if these pose a significant threat to human health. However, experience in other countries suggests that continued surveillance of the food chain is necessary to detect emerging resistances.
The datasets presented in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found here: PRJNA606458, SAMN14091501.
Ethical review and approval was not required for the animal study because animals were sampled post-slaughter.
All authors played an integral part of the project completion. Each author has read and approved the final version of the manuscript. Specifically, DB, SW, and DW conceived and designed the experiments. DB and SW performed the experiments. DB, SW, MA, and DW analyzed the data. DB, MA, and DW wrote the manuscript.
Reagents to prepare the CHROMagar COL-APSE culture media were provided free of charge for evaluation by CHROMagar.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The assistance of the staff of the Victorian poultry processing plant with obtaining the poultry cloacal swabs was gratefully acknowledged. Thanks also to CHROMagar for the generous gift of CHROMagar COL-APSE culture media.
Abbott, S. L., Moler, S., Green, N., Tran, R. K., Wainwright, K., and Janda, J. M. (2011). Clinical and laboratory diagnostic characteristics and cytotoxigenic potential of Hafnia alvei and Hafnia paralvei strains. J. Clin. Microbiol. 49, 3122–3126. doi: 10.1128/JCM.00866-11
Abdul Momin, M. H. F., Bean, D. C., Hendriksen, R. S., Haenni, M., Phee, L. M., and Wareham, D. W. (2017). CHROMagar COL-APSE: a selective bacterial culture medium for the isolation and differentiation of colistin-resistant gram-negative pathogens. J. Med. Microbiol. 66, 1554–1561. doi: 10.1099/jmm.0.000602
Animal Health Australia (2017). Animal Health in Australia 2016. Canberra, ACT: Animal Health Australia.
Apostolakos, I., and Piccirillo, A. (2018). A review on the current situation and challenges of colistin resistance in poultry production. Avian Pathol. 47, 546–558. doi: 10.1080/03079457.2018.1524573
Aravena-Roman, M., Inglis, T. J., Henderson, B., Riley, T. V., and Chang, B. J. (2012). Antimicrobial susceptibilities of Aeromonas strains isolated from clinical and environmental sources to 26 antimicrobial agents. Antimicrob. Agents Chemother. 56, 1110–1112. doi: 10.1128/AAC.05387-11
Arcilla, M. S., Van Hattem, J. M., Matamoros, S., Melles, D. C., Penders, J., de Jong, M. D., et al. (2016). Dissemination of the mcr-1 colistin resistance gene. Lancet Infect Dis. 16, 147–149. doi: 10.1016/S1473-3099(15)00541-1
Australian Chicken Meat Federation (2018). Position Statement: Responsible use of Antibiotics in the Australian Chicken Meat Industry. Sydney, NSW: Australian Chicken Meat Federation.
Australian Pesticides and Veterinary Medicines Authority (2014). Quantity of Antimicrobial Products Sold for Veterinary use in Australia: July 2005 to June 2010. Kingston, NY: Australian Pesticides and Veterinary Medicines Authority.
Breazeale, S. D., Ribeiro, A. A., Mcclerren, A. L., and Raetz, C. R. (2005). A formyltransferase required for polymyxin resistance in Escherichia coli and the modification of lipid A with 4-Amino-4-deoxy-L-arabinose. Identification and function of UDP-4-deoxy-4-formamido-L-arabinose. J. Biol. Chem. 280, 14154–14167. doi: 10.1074/jbc.M414265200
Buess, S., Nuesch-Inderbinen, M., Stephan, R., and Zurfluh, K. (2017). Assessment of animals as a reservoir for colistin resistance: no MCR-1/MCR-2-producing Enterobacteriaceae detected in Swiss livestock. J. Glob. Antimicrob. Resist. 8, 33–34. doi: 10.1016/j.jgar.2016.11.001
Chen, L., Zhang, J., Wang, J., Butaye, P., Kelly, P., Li, M., et al. (2018). Newly identified colistin resistance genes, mcr-4 and mcr-5, from upper and lower alimentary tract of pigs and poultry in China. PLoS ONE. 13:e0193957. doi: 10.1371/journal.pone.0193957
Clinical and Laboratory Standards Institute (2018). Performance Standards for Antimicrobial Susceptibility Testing; Twenty-Eighth Informational Supplement CLSI M100-S28. Wayne, PA: CLSI.
Doulgeraki, A. I., Paramithiotis, S., and Nychas, G.-J. E. (2011). Characterization of the Enterobacteriaceae community that developed during storage of minced beef under aerobic or modified atmosphere packaging conditions. Int. J. Food Microbiol. 145, 77–83. doi: 10.1016/j.ijfoodmicro.2010.11.030
Ellem, J. A., Ginn, A. N., Chen, S. C., Ferguson, J., Partridge, S. R., and Iredell, J. R. (2017). Locally acquired mcr-1 in Escherichia coli, Australia, 2011 and 2013. Emerg. Infect Dis. 23, 1160–1163. doi: 10.3201/eid2307.161638
Fosse, T., Giraud-Morin, C., and Madinier, I. (2003). Induced colistin resistance as an identifying marker for Aeromonas phenospecies groups. Lett. Appl. Microbiol. 36, 25–29. doi: 10.1046/j.1472-765X.2003.01257.x
Gatzeva-Topalova, P. Z., May, A. P., and Sousa, M. C. (2005). Structure and mechanism of ArnA: conformational change implies ordered dehydrogenase mechanism in key enzyme for polymyxin resistance. Structure 13, 929–942. doi: 10.1016/j.str.2005.03.018
Gunzer, F., Rudolph, W. W., Bunk, B., Schober, I., Peters, S., Müller, T., et al. (2018). Whole-genome sequencing of a large collection of Myroides odoratimimus and Myroides odoratus isolates and antimicrobial susceptibility studies. Emerg. Microbes Infect. 7:61. doi: 10.1038/s41426-018-0061-x
Höll, L., Behr, J., and Vogel, R. F. (2016). Identification and growth dynamics of meat spoilage microorganisms in modified atmosphere packaged poultry meat by MALDI-TOF MS. Food Microbiol. 60, 84–91. doi: 10.1016/j.fm.2016.07.003
Holmes, B., Snell, J. J., and Lapage, S. P. (1979). Flavobacterium odoratum: a species resistant to a wide range of antimicrobial agents. J. Clin. Pathol. 32, 73–77. doi: 10.1136/jcp.32.1.73
Huys, G., Cnockaert, M., Abbott, S. L., Janda, J. M., and Vandamme, P. (2010). Hafnia paralvei sp. nov., formerly known as Hafnia alvei hybridization group 2. Int. J. Syst. Evol. Microbiol. 60, 1725–1728. doi: 10.1099/ijs.0.018606-0
Jacquier, H., Le Monnier, A., Carbonnelle, E., Corvec, S., Illiaquer, M., Bille, E., et al. (2012). In vitro antimicrobial activity of “last-resort” antibiotics against unusual nonfermenting Gram-negative bacilli clinical isolates. Microb. Drug Resist. 18, 396–401. doi: 10.1089/mdr.2011.0195
Jayol, A., Saly, M., Nordmann, P., Menard, A., Poirel, L., and Dubois, V. (2017). Hafnia, an enterobacterial genus naturally resistant to colistin revealed by three susceptibility testing methods. J. Antimicrob. Chemother. 72, 2507–2511. doi: 10.1093/jac/dkx154
Kämpfer, P., Rosselló-Mora, R., Scholz, H. C., Welinder-Olsson, C., Falsen, E., and Busse, H.-J. (2006). Description of Pseudochrobactrum gen. nov., with the two species Pseudochrobactrum asaccharolyticum sp. nov. and Pseudochrobactrum saccharolyticum sp. nov. Int. J. Syst. Evol. Microbiol. 56, 1823–1829. doi: 10.1099/ijs.0.64256-0
Kaur, M., Shang, H., Tamplin, M., Ross, T., and Bowman, J. P. (2017). Culture-dependent and culture-independent assessment of spoilage community growth on VP lamb meat from packaging to past end of shelf-life. Food Microbiol. 68, 71–80. doi: 10.1016/j.fm.2017.06.015
Ling, Z., Yin, W., Li, H., Zhang, Q., Wang, X., Wang, Z., et al. (2017). Chromosome-mediated mcr-3 variants in Aeromonas veronii from chicken meat. Antimicrob. Agents Chemother. 61:e01272-17. doi: 10.1128/AAC.01272-17
Liu, Y. Y., Wang, Y., Walsh, T. R., Yi, L. X., Zhang, R., Spencer, J., et al. (2016). Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: a microbiological and molecular biological study. Lancet Infect Dis. 16, 161–168. doi: 10.1016/S1473-3099(15)00424-7
Ma, S., Sun, C., Hulth, A., Li, J., Nilsson, L. E., Zhou, Y., et al. (2018). Mobile colistin resistance gene mcr-5 in porcine Aeromonas hydrophila. J. Antimicrob. Chemother. 73, 1777–1780. doi: 10.1093/jac/dky110
Monte, D. F., Mem, A., Fernandes, M. R., Cerdeira, L., Esposito, F., Galvão, J. A., et al. (2017). Chicken meat as a reservoir of colistin-resistant Escherichia coli strains carrying mcr-1 genes in South America. Antimicrob. Agents Chemother. 61:e02718–16. doi: 10.1128/AAC.02718-16
Mukerji, S., Stegger, M., Truswell, A. V., Laird, T., Jordan, D., Abraham, R. J., et al. (2019). Resistance to critically important antimicrobials in Australian silver gulls (Chroicocephalus novaehollandiae) and evidence of anthropogenic origins. J. Antimicrob. Chemother. 74, 2566–2574. doi: 10.1093/jac/dkz242
Okada, S., and Gordon, D. M. (2003). Genetic and ecological structure of Hafnia alvei in Australia. Syst. Appl. Microbiol. 26, 585–594. doi: 10.1078/072320203770865891
Olaitan, A. O., Morand, S., and Rolain, J.-M. (2014). Mechanisms of polymyxin resistance: acquired and intrinsic resistance in bacteria. Front. Microbiol. 5:643. doi: 10.3389/fmicb.2014.00643
Poirel, L., Jayol, A., and Nordmann, P. (2017). Polymyxins: antibacterial activity, susceptibility testing, and resistance mechanisms encoded by plasmids or chromosomes. Clin. Microbiol. Rev. 30, 557–596. doi: 10.1128/CMR.00064-16
Säde, E., Murros, A., and Björkroth, J. (2013). Predominant enterobacteria on modified-atmosphere packaged meat and poultry. Food Microbiol. 34, 252–258. doi: 10.1016/j.fm.2012.10.007
Shen, Y., Xu, C., Sun, Q., Schwarz, S., Ou, Y., Yang, L., et al. (2018). Prevalence and genetic analysis of mcr-3-positive Aeromonas species from humans, retail meat, and environmental water samples. Antimicrob. Agents Chemother. 62:e00404-18. doi: 10.1128/AAC.00404-18
Teale, C. J. (2002). Antimicrobial resistance and the food chain. J. Appl. Microbiol. 92, 85S−89S. doi: 10.1046/j.1365-2672.92.5s1.20.x
Wang, R., van Dorp, L., Shaw, L. P., Bradley, P., Wang, Q., Wang, X., et al. (2018). The global distribution and spread of the mobilized colistin resistance gene mcr-1. Nat. Commun. 9:1179. doi: 10.1038/s41467-018-03205-z
Weisburg, W. G., Barns, S. M., Pelletier, D. A., and Lane, D. J. (1991). 16S ribosomal DNA amplification for phylogenetic study. J. Bact. 173, 697–703. doi: 10.1128/JB.173.2.697-703.1991
Xavier, B. B., Lammens, C., Ruhal, R., Kumar-Singh, S., Butaye, P., Goossens, H., et al. (2016). Identification of a novel plasmid-mediated colistin-resistance gene, mcr-2, in Escherichia coli, Belgium, June 2016. Euro Surveill. 21:30280. doi: 10.2807/1560-7917.ES.2016.21.27.30280
Yin, W., Li, H., Shen, Y., Liu, Z., Wang, S., Shen, Z., et al. (2017). Novel plasmid-mediated colistin resistance gene mcr-3 in Escherichia coli. mBio 8:e00543-17. doi: 10.1128/mBio.00543-17
Keywords: Aeromonas, polymyxin-resistance, colistin, poultry, Hafnia paralvei, antibiotics
Citation: Bean DC, Wigmore SM, Abdul Momin MHF and Wareham DW (2020) Polymyxin Resistant Bacteria in Australian Poultry. Front. Sustain. Food Syst. 4:550318. doi: 10.3389/fsufs.2020.550318
Received: 09 April 2020; Accepted: 31 August 2020;
Published: 15 October 2020.
Edited by:
Guadalupe Virginia Nevárez-Moorillón, Autonomous University of Chihuahua, MexicoReviewed by:
Raul Avila-Sosa, Meritorious Autonomous University of Puebla, MexicoCopyright © 2020 Bean, Wigmore, Abdul Momin and Wareham. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: David C. Bean, ZC5iZWFuQGZlZGVyYXRpb24uZWR1LmF1
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.