
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Nutr. , 10 March 2025
Sec. Nutrigenomics
Volume 12 - 2025 | https://doi.org/10.3389/fnut.2025.1538505
This article is part of the Research Topic Genome-based Nutrition Strategies for Preventing Diet-related Chronic Diseases: Where Genes, Diet, and Food Culture Meet, Volume II View all articles
 Sarah A. Blomquist1
Sarah A. Blomquist1 Jil H. Albrecht1
Jil H. Albrecht1 Brian Hallmark2,3
Brian Hallmark2,3 Yann C. Klimentidis3,4
Yann C. Klimentidis3,4 Luis A. Garcia5,6
Luis A. Garcia5,6 Lawrence J. Mandarino5,6
Lawrence J. Mandarino5,6 Dawn K. Coletta5,6,7
Dawn K. Coletta5,6,7 Floyd H. Chilton1,3,8,9*
Floyd H. Chilton1,3,8,9*Background: Latinos, the largest racial/ethnic minority group in the United States, have high rates of cardiometabolic diseases, hypothesized due in part to genetic variation in the fatty acid desaturase (FADS) cluster that is associated with reduced omega-3 (n-3) highly unsaturated fatty acid (HUFA) biosynthesis. This study examined how variations in FADS and other HUFA pathway-related genes (ELOVL5 and ELOVL2) impact cardiometabolic disease risk factors in Latinos of Mexican Ancestry (LMA).
Results: This study analyzed 493 self-identified LMA from the Arizona Insulin Resistance registry (AIR) and found a marked enrichment in FADS alleles linked the ancestral haplotype (AH) compared to European Americans. LMA individuals with two AH alleles produced markedly lower levels of n-6 and n-3 HUFAs. However, this was more pronounced with the n-3 HUFAs, eicosapentaenoic acid (EPA) and docosahexaenoic acid (DHA), where the n-6 arachidonic acid (ARA) to EPA and DHA ratios were 30:1 and 5:1, respectively, and circulating EPA levels were reduced to <5 ng/mL. Importantly, genetic variations in both FADS and ELOVL2/5 regions also were strongly associated with several cardiometabolic disease (CMD) markers, with the presence of two FADS AH alleles corresponding to a 45, 33, and 41% increase in fasting insulin, triglyceride levels and HOMA-IR, respectively.
Conclusion: This study reveals the potential impact of genetically influenced HUFA regulation and n-3 HUFA deficiency on cardiometabolic disease risk within LMA. These insights provide a strong rationale for future studies and clinical trials that focus on n-3 HUFA supplementation to mitigate CMD disparities in LMA populations.
The nutritional content of food in developed nations has changed dramatically over the past 75 years (1), mirroring the rise in non-communicable diseases, which account for approximately 75% of deaths in developed countries (2–4). Cardiometabolic diseases (CMD) remain the leading cause of death globally, with disparities particularly evident among various racial/ethnic minority groups in the United States (USA). Latino populations, especially Mexican Americans (MxAm), have higher rates of obesity, diabetes, hyperlipidemia, and cardiovascular diseases compared Non-Hispanic White (5–7) populations.
It is now estimated that over 70% of daily energy consumed in a modern Western diet (MWD) is from sources that were absent from our ancestors’ hunter-gatherer diet (8). A notable example of this shift is the change in dietary polyunsaturated fatty acids (PUFA) consumption, which can be traced back to recommendations from health agencies to reduce saturated fat intake in favor of PUFAs, predominantly derived from vegetable oils such as soybean, corn, and canola (9, 10). These recommendations led to a ~ 4-fold increase in the dietary consumption of the omega-6 (n-6) PUFA linoleic acid (LA) without significantly altering the consumption of dietary omega-3 (n-3) PUFAs, thereby altering the ratio of dietary n-6 to n-3 PUFAs to greater than 10:1 (1).
The dietary n-6 and n-3, 18-carbon PUFAs LA and alpha-linolenic acid [ALA, omega-3 (n-3)] are converted by the liver and other tissues to long-chain (>20 carbon), highly unsaturated fatty acids (HUFAs) utilizing a sequential set of desaturases and elongases (11, 12). Figure 1 highlights genes and enzymes key to HUFA biosynthesis and metabolism. In two parallel and competing pathways, 18-carbon n-6 or n-3 PUFAs are converted into HUFAs. In the n-6 arm of the pathway (left), ARA is synthesized from LA, utilizing 2 desaturation and 1 elongation enzymatic steps (13, 14). In the n-3 arm of the pathway (right), EPA and DHA are synthesized from dietary ALA. This second desaturation step (Δ-5 desaturase or FADS1) is rate-limiting, and genetic variation throughout the FADS gene cluster impacts the pathway efficiency via altered FADS1 transcription (15–17).
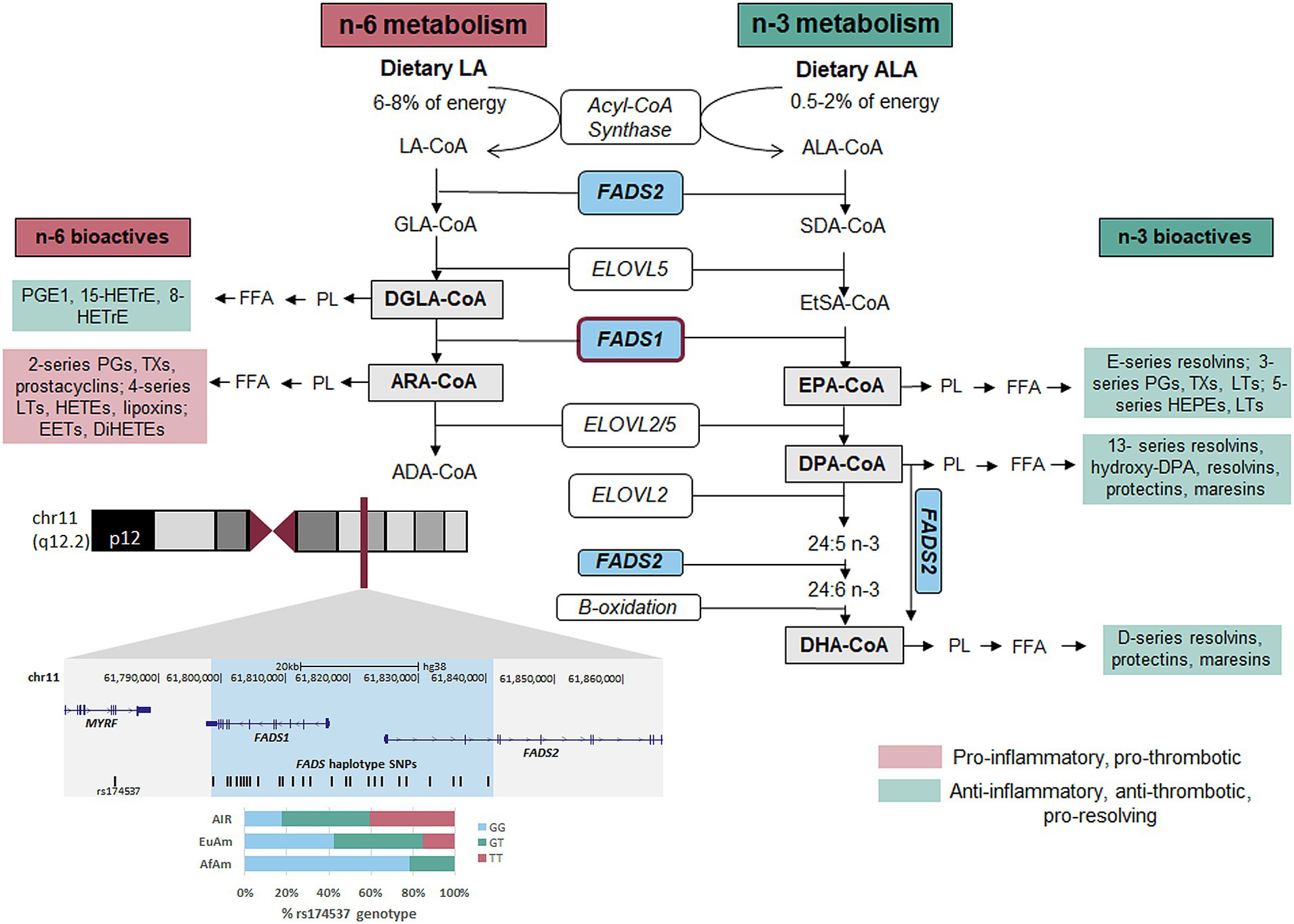
Figure 1. n-6 and n-3 HUFA biosynthetic pathway. FADS1/2, fatty acid desaturase 1 or 2; ELOVL2/5, fatty acid elongase 2 or 5; ALA, alpha-linolenic acid; SDA, stearidonic acid; EtSA, eicosatetranenoic acid; EPA, eicosapentaenoic acid; DPA, docosapentaenoic acid; DHA, docosahexaenoic acid; LA, linoleic acid; GLA, gamma-linolenic acid; DGLA, dihomogammalinoleic acid; ARA, arachidonic acid; ADA, adrenic acid; PL, phospholipid; FFA, free fatty acid; HETrE, hydroxyeicosatrienoic acid; PG, prostaglandin; PGE1, prostaglandin E1; TX, thromboxane; LT, leukotriene; EET, epoxyeicosatetraenoic acid; DiHETE, dihydroxyeicosatetraenoic acid; HETE, hydroxyeicosatetraenoic acid; HEPE, hydroxyeicosapentaenoic acid; Co-A, Coenzyme A.
Once ARA is formed and placed into membrane phospholipids, it can be mobilized (by phospholipases) and metabolized [by cyclooxygenase (COX), lipoxygenase (LOX) and P450 pathways] into oxylipins including eicosanoids such as prostaglandins, thromboxanes and leukotrienes (Figure 1) (18–20). These then act locally to promote inflammation and thrombosis in CVD and other diseases (20). By contrast, DGLA, the n-6 HUFA proximal to FADS1, is metabolized by COX 1/2 to 1-series PGs, particularly PGE1, and by 15-LOX to 15-HETrE (21). These two metabolites of DGLA suppress inflammation, promote vasodilation, lower blood pressure, inhibit smooth muscle cell proliferation, and exert anti-neoplastic activities (22–25). Similarly, EPA and DHA are metabolized to anti-inflammatory, anti-thrombotic and “pro-resolving” oxylipins including resolvins, protectins, and maresins (19, 26, 27). With few exceptions, DGLA, EPA, DHA and their oxylipin products exert effects that differ from or oppose ARA-derived oxylipins (Figure 1) regarding inflammation, hyperlipidaemia and disease modification. Consequently, DGLA conversion to ARA and the balance of ARA to DGLA, EPA and DHA have the potential to influence the prevalence and severity of numerous human diseases.
Numerous studies have established a connection between genetic variations in the FADS cluster and the efficiency of the HUFA biosynthetic pathway, and thus HUFA levels (28, 29). Early research indicated that high levels of ARA, as well as the ARA/LA ratio, are independent predictors for coronary artery disease (30). Furthermore, the role of FADS variants in complex lipid and inflammatory phenotypes has been extensively documented (31). Notably, the FADS gene region is recognized as the locus associated with the largest multimorbidity cluster in the entire human genome (32).
Moreover, research from various laboratories has revealed significant ancestry-based discrepancies in the frequency of FADS variants affecting HUFAs and their corresponding oxylipin metabolites, such as eicosanoids (33–36). Evolutionary pressures have shaped the allele frequencies of FADS variants across different populations, reflecting adaptations to diverse diets and environments. For example, the “derived” FADS haplotype experienced ancient positive selection in Africa, leading to its high frequency in present-day African populations (37, 38). For example, ∼80% of African Americans (AfAm), as compared to ∼45% of European Americans (EuAm), carry two copies of derived alleles associated with increased FADS1 activity, resulting in a marked increase in HUFAs, especially ARA and ARA-derived metabolites (34, 35). The introduction of agriculture is also thought to have influenced FADS allele frequencies. However, strong selection on the derived allele in European populations occurred much later than the Neolithic transition, possibly as late as the Bronze Age (39). The findings of Fumagalli et al. (40) on genetic adaptations in the Greenlandic Inuit population provided an important rationale as to why a high proportion of Indigenous American ancestry populations carry two copies of “ancestral” alleles associated with reduced FADS1 activity, resulting in a reduction in HUFAs, especially EPA production (36, 37, 41, 42). Their research demonstrated that the FADS gene cluster shows the strongest signal of genetic adaptation, reflecting evolutionary responses to a traditional marine-based diet and extreme Arctic conditions (40). Specifically, the ancestral FADS alleles were positively selected in the Inuit and remain prevalent among populations with high proportions of Indigenous American ancestry such as MxAm.
An unresolved question is if the variation in FADS in Hispanic populations that induces very low levels of n-3 HUFAs and imbalances in n-6 to n-3 HUFA ratios influences the risk of cardiometabolic diseases often observed in these populations. To address this important question, we investigated the relationship between variation the in genes (FADS1,2,3 and ELOVL2,5) that code for HUFA biosynthetic enzymes on a robust set of cardiometabolic markers and traits in adult Latino participants from the Arizona Insulin Resistance registry (AIR) (43).
The Arizona Insulin Resistance registry (AIR) cohort comprises 667 self-reported Latinos of Mexican Ancestry (LMA) residing in Arizona, ranging in age from 8 to 83 years. A detailed description of these participants can be found in a prior publication by Shaibi et al. (44). In the subset of n = 497 adult participants, 35% were male, the mean age was 36.4 and the mean BMI was 29.8. No socioeconomic status or lifestyle factors were available for reporting. Of these, 353 participants were spread across 92 families, while the remainder were single, unrelated individuals. The distribution of relative pairs within the entire cohort is documented by DeMenna et al. (43). In the current study, we focused solely on adult participants aged 18 and above (n = 497). In this subset, 81% were overweight or obese, and 45% had hyperglycemia (44). Comprehensive data, including metabolic, anthropometric, demographic, and medical history, were collected for all participants (44).
A total of 493 participants were successfully genotyped for 20 SNPs (three ELOVL5, six ELOVL2, and eleven FADS1/2/3) using the Agena Bioscience® MALDI-TOF™ instrument. The 20 SNPs were chosen based on previous literature that revealed significant associations between various SNPs and HUFA levels (17, 35, 39, 40, 45–50), given that the SNPs could be successfully multiplexed. Primers for multiplex PCR and extension PCR were designed with Agena’s Online Assay Design Suite (v2.0). The assays were validated against the human genome database GRCH37/hg19 (dbSNP138). Oligos were synthesized by Integrated DNA Technologies (Coralville, Iowa), and hydrated to stock concentrations (250 μM for PCR primers and 500 μM for extension primers).
All reagents used were part of the iPlex Gold Genotyping kit (Agena Biosciences, San Diego, CA), and used according to their recommended protocols. High-quality genomic DNA was obtained from whole blood using the PAXgene Blood DNA procedure per the manufacturer’s instructions (Qiagen, CA, United States) (43). Samples were plated at 20 ng per well on a 96-well plate. A 5 μL multiplex polymerase chain reaction (PCR) was performed on each sample. Following PCR, extra nucleotides were neutralized with a shrimp alkaline phosphatase (SAP) reaction. A third reaction utilized iPLEX Gold chemistry, terminator nucleotides and a probe (extension primer) that anneals directly adjacent to the SNP site, to carry out single-base extension onto the SNP site. The reactions were cleaned using a salt-sequestering resin, and a small volume (nanoliters) of each was spotted onto the matrix of a specialized silicon chip. This chip was then analyzed using Agena’s Matrix Assisted Laser Desorption/Ionization – Time of Flight (MALDI-TOF) mass-spectrometry instrument and MassARRAY Typer software v4.1.83. Samples were genotyped using a 20-SNP custom array designed for the Agena MassARRAY MALDI-TOF instrument. The work was performed by the University of Arizona Genetics Core.
Collection of samples and clinical phenotypes are described in more detail in previous publications (43, 44). Briefly, blood samples were collected after a 12-h fast to assess biomarkers including glucose, insulin, triglycerides, total cholesterol, high-density lipoprotein cholesterol, low-density lipoprotein cholesterol, and very-low-density lipoprotein cholesterol. Participants also underwent a 2-h oral glucose tolerance test. Diabetes and metabolic phenotypes, including indices of insulin action and secretion, were also collected. The following clinical indices were evaluated: body mass index (kg/m2), fat mass (%), waist circumference (cm), hip circumference (cm), systolic blood pressure (mmHg), diastolic blood pressure (mmHg), fasting plasma insulin (μIU/mL), fasting plasma glucose (mg/dL), hemoglobin A1c (%), 2 h oral glucose tolerance test (mg/dL), homeostasis model assessment for insulin resistance (HOMA-IR), Matsuda index, disposition index, prediabetes status, diabetes status, alanine aminotransferase (IU/L), aspartate aminotransferase (IU/L), and adiponectin (μg/mL). Participants were excluded from the study if they had any of the following: overt diabetes, untreated metabolic disease, HIV/AIDS, active cancer (or in remission for less than 3 years), acute illness, or currently pregnant.
Serum samples were collected from participants and stored at −80°C until the fatty acid (FA) analyses. Fatty acid methyl esters (FAMEs) were prepared (51) after alkaline hydrolysis of complex lipids in duplicate samples (100 μL) in the presence of the C:17 internal standard triheptadecanoin (Nuchek Prep). This standard was included for FA quantification, as previously described (52, 53). A standard panel of 37 FAs (Supelco, which accounts for 99% of FAs in the sample) was quantified by gas chromatography with flame ionization detection (GC-FID) using an Agilent 9000 Intuvo gas chromatographer with a DB-FastFAME column (20 m, 0.18 mm ID, 0.2 μm film) for a split injection with a split ratio of 15:1. The instrument response factor was calculated based on external standard sets for quality assurance purposes, and a mixture of known FAMEs was run with each sample set to monitor instrument performance (54). Individual FAs were expressed as ng/ml and were calculated using the internal standard response factor and by taking the average of duplicate samples.
All analyses were conducted using R Studio, version 4.2.0 or later. We examined associations between SNPs in FADS, ELOVL2, and ELOVL5 with fatty acids (FAs) and other clinical indices using regression analyses. The major alleles found in the sampled population were chosen as the alleles whose effect was to be tested, regardless of their presence or absence in the human genome reference. Genotypes were coded based on the number of major alleles (0, 1, or 2 copies). For FADS SNPs, most major alleles were ancestral (i.e., the same as the most recent common ancestors of humans). The only exception was rs174538. After analyzing the 1,000 Genomes haplotype structure, it was found that the derived allele of rs174538 consistently aligns with the ancestral background. Consequently, the coding for this SNP was inverted for our analysis to again count the number of major alleles.
Linear mixed models (55) (using the lmekin package in R) with genotype coded both additively and as a factor model were performed, accounting for relatedness between individuals using self-reported pedigree information. All regression analyses included covariate adjustments for age, sex, type 2 diabetes status and body mass index (BMI). Models were also assessed after removal of BMI. Outcome variables were transformed to approximate normality using logarithmic transformations. Variables were checked for outlier data points > 3 standard deviations from the mean, but ultimately nothing was removed from the data set. Missing values were handled using listwise deletion. Summary statistic tables for all permutations of regression analysis are included in Supplementary File 1, and list β-values (estimate, or effect size per increasing number of major alleles), and p-values (significance). Due to logarithmic transformations, estimates required transformation in order to be expressed as percentages (56). Forest Plots were generated using a mixed regression model with 95% confidence intervals and are based on the standard p-value threshold. The false discovery rate (FDR) was controlled using the Benjamini-Hochberg procedure (57), and was performed two ways. The first set was computed from the list of all p-values within each individual SNP, where a threshold of 0.1 was used to examine significant associations [referred to as “pval (fdr)” in raw datasheets]. A second global FDR (q-values, referred to as “global_fdr” in raw datasheets) was computed using a p-value from all tests performed for this analysis with a 0.2 threshold for a cutoff. To account for multiple hypothesis testing and minimize false positives, we applied a false discovery rate (FDR) threshold to balance discovery and reproducibility. An FDR cutoff of 0.2, a widely used standard in genomics research (57), was initially applied to identify significant associations. To further assess the robustness of our findings, we also applied a more stringent FDR threshold of 0.1, which reduces the number of significant results by increasing conservatism. By comparing results at both thresholds, we identified the most robust associations that remained significant under the stricter criterion.
Before examining allele frequencies in our study, it is important to point out that allele frequencies can vary significantly across different ethnic groups. An allele that is the major allele in one population may be the minor allele in another population. In the current study, the major alleles in Indigenous American ancestry populations (1000 Genomes MxAm and AIR) are markedly different than in AfAm and EuAm populations. As the focus of this manuscript is Indigenous American ancestry populations, the major alleles in these populations will be designated as the alleles of interest for all populations.
As discussed above and detailed in Table 1, the allele frequencies of SNPs in FADS and ELOVL2/5 varied among different populations of different ancestries. For the FADS SNPs, the allele frequencies (based on 1,000 Genomes samples) of the alleles of interest range from 0.19 to 0.39 in EuAm populations. In AfAm populations, they range from 0.11 to 0.89, with notably smaller linkage disequilibrium (LD) blocks in the FADS cluster of this group, resulting in rs174594, rs174602, and rs174455 not in strong LD with other FADS SNPs within this population. In contrast, in 1000 Genomes MxAm populations, the frequencies were higher and ranged from 0.61 to 0.73, while in AIR, they ranged from 0.54 to 0.70.
We also observed significant variations in genotypic frequencies across populations. As illustrated in Supplementary Figure 2, the TT genotype at rs174537 occurs in less than 1% of AfAm populations, 15% of EuAm, 48% of MxAm, and 41% of AIR participants. In contrast, the GG genotype was found in 79% AfAm, 42% EuAm, 9% MxAm and 18% AIR populations.
For the three ELOVL5 SNPs examined, the allele frequencies in MxAm and AIR were similar to those in EuAm, ranging between 0.71 and 0.82. In contrast, in AfAm populations, they ranged from 0.35 to 0.44. Among the six ELOVL2 SNPs, EuAm populations exhibited a frequency range from 0.44 to 0.75. AfAm populations had a broader range from 0.32 to 0.91. In MxAm and AIR populations, the frequencies lie from 0.64 to 0.81 and 0.61 to 0.75, respectively.
Figures 2A–D display Forest Plots illustrating the associations between FADS and ELOVL2/5 SNP alleles and circulating n-6 and n-3 HUFA levels in the AIR cohort. All FADS alleles showed significant associations with the n-6 HUFAs, ARA and DGLA. For ARA levels, there was a significant decrease ranging from 8.0 to 20.4% per FADS major allele (the allele most common in the AIR cohort). In contrast, these same FADS alleles were associated with increases in DGLA levels, ranging from 9.5 to 14.3% per major allele (PMA). All but two FADS SNPs were associated with EPA levels, where each major allele decreased EPA levels between 11.2 and 24.7%. Similarly, all but three FADS SNPs were associated with DHA levels, with decreases ranging from 6.0 to 9.3% per allele. In contrast to FADS SNPs, only two ELOVL2/5 SNPs had allelic associations with ARA, where each allele decreased levels by ~6% PMA. No ELOVL2/5 alleles impacted DGLA, EPA, or DHA levels.
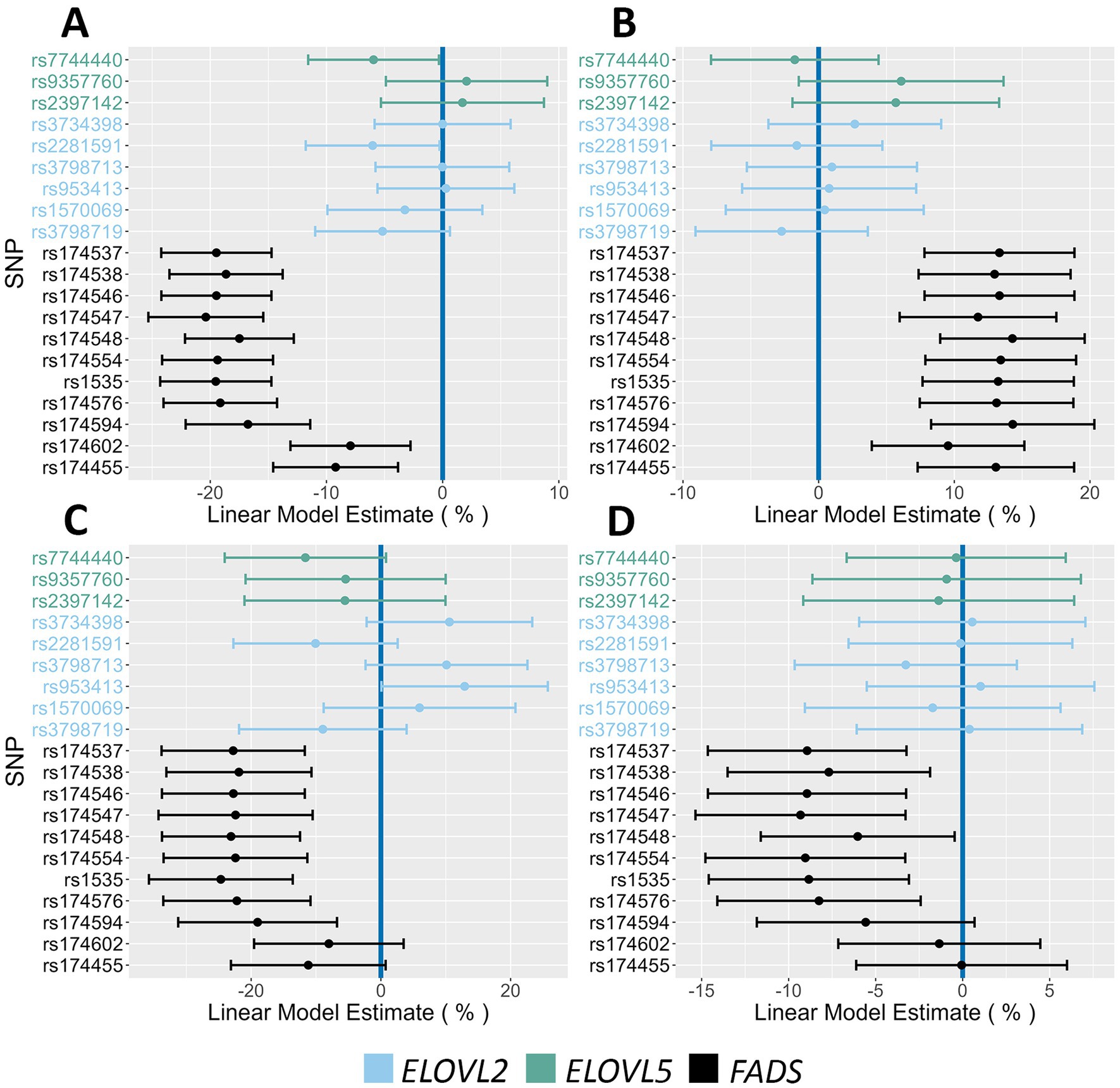
Figure 2. Impact of FADS and ELOVL alleles on n-6 and n-3 HUFA plasma levels. This figure contains Forest Plots that illustrate the impact of FADS and ELOVL reference alleles on n-6 and n-3 HUFA levels. (A,B) Allelic effects on ARA and DGLA, respectively. (C,D) Allelic effects on EPA and DHA, respectively. Green, blue and black lines represent ELOVL5, ELOVL2 and FADS alleles, respectively. The 95% confidence intervals are indicated by the horizontal lines.
Figures 3A–D display boxplots of HUFA levels as a function of rs174537 genotypes. Rs174537 was chosen as a representative FADS SNP based on its strong associations with ARA levels in previous GWAS studies, its wide use in association studies, and the fact that it sits in LD with two proposed FADS1 functional SNPs (58, 59). Table 2 illustrates HUFA ratios, estimates (β) and significance values when comparing TT and GG genotypes at rs174537. Figures 2, 3 and Table 2 demonstrate a 33% reduction in ARA concentrations when comparing the GG to TT genotypes. This trend was reversed for DGLA, the HUFA proximal to the FADS1 enzymatic step, with a 32% increase in the TT compared to the GG genotype. EPA and DHA concentrations were decreased by 38 and 15% when comparing GG to TT genotypes, respectively.
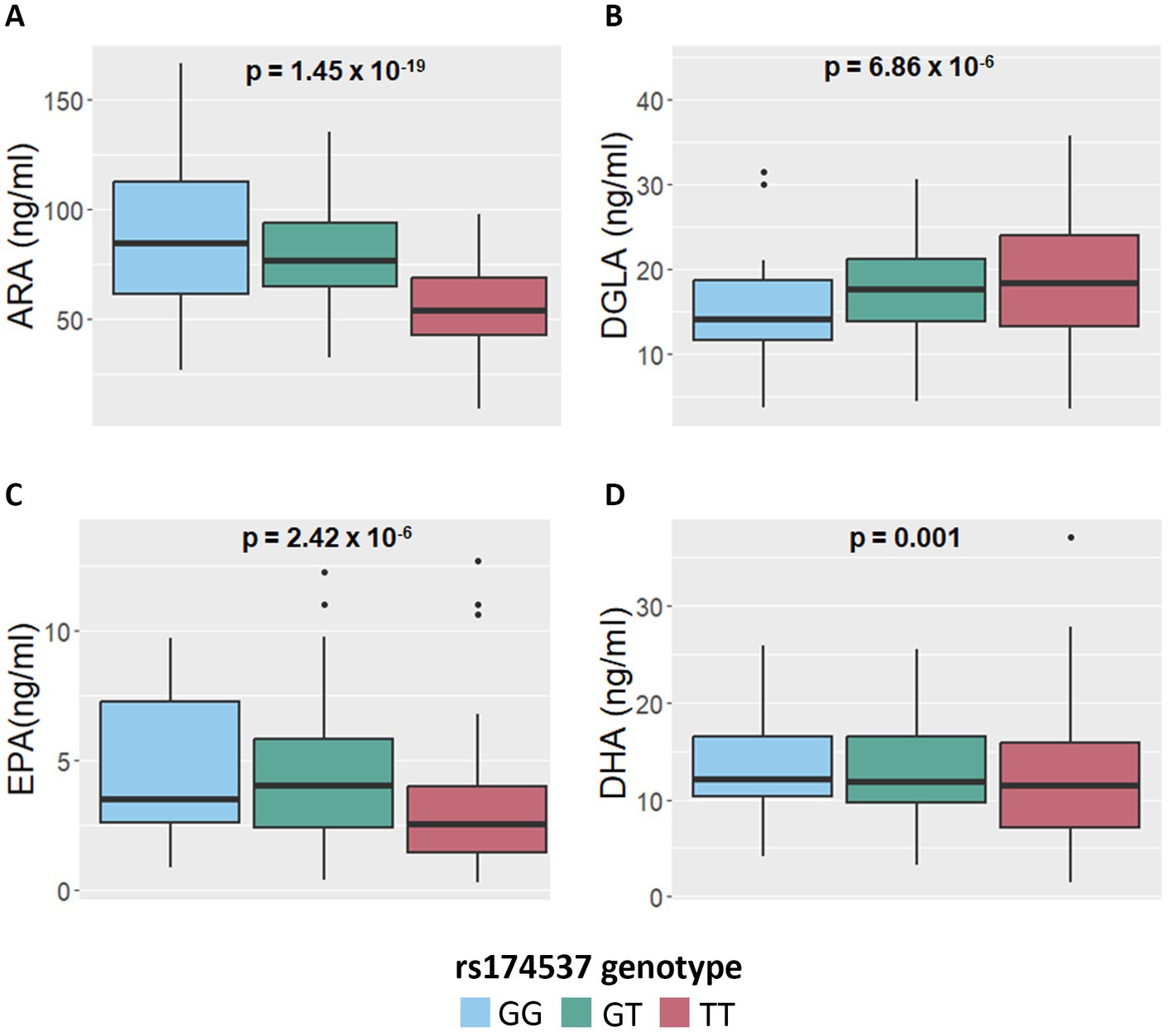
Figure 3. Impact of FADS and ELOVL genotypes on n-6 and n-3 HUFA plasma concentrations. (A,B) Concentrations of ARA and DGLA, respectively, as a function of the FADS SNP, rs174537, genotypes. (C,D) Concentrations of EPA and DHA, respectively, as a function of rs174537 genotypes. p-values remained significant after FDR adjustment and were derived from an additive linear mixed model.
Determining HUFA ratios throughout the biosynthesis pathway, and particularly those flanking the FADS1 biosynthetic step, has been utilized to assess the metabolic flux through the pathway (15, 16, 46). The opposing effects of n-6/n-3 HUFA on inflammatory modulation and their associated circulating levels also illustrate the balance between anti-inflammatory and pro-resolving substrates. Table 2 shows there is a 30–40% decrease in levels of both ARA and EPA individually, in individuals with the TT genotype at rs174537 (homozygous for the AIR major allele) as opposed to the GG genotype (minor in AIR but common in other populations). This is notable as these HUFAs sit at the same point (distal to the FADS1 biochemical step) in the n-6 and n-3 HUFA biosynthetic pathways.
Table 2 also reveals ARA/DGLA ratios are 3.1 and 6.1 for the TT and GG genotypes, respectively (p = 1.6 × 10−76, β = −2.99), suggesting that the metabolic flux through the FADS1 portion of the pathway (Figure 1) is markedly decreased in individuals with the TT genotype. When comparing the ratios of the n-6 HUFA, ARA, to n-3 HUFAs, ARA/EPA ratios are not statistically different when comparing the TT and GG genotypes (p = 0.9, β = −0.63). In contrast, ARA/DHA ratios are statistically lower for the TT genotype (4.98 and 6.4 in the TT and GG genotypes, respectively, p = 8.3×10−12, β = −1.31). Within the n-3 HUFA biosynthetic pathway, DHA/EPA ratios were not significantly different when comparing TT and GG genotypes.
Supplementary Figure 3 illustrates allelic effects on HUFA ratios among all assayed SNPs. FADS major alleles (AIR population) were associated with decreased ratios of ARA/DGLA and ARA/DHA, where each major allele reduced the ARA/DGLA ratio by 0.75–1.5 and the ARA/DHA ratio by 0.4–0.75. FADS alleles at rs174548 and rs1535 also increase DHA/EPA by ~1.1. Notably, several ELOVL2/5 alleles also impacted HUFA ratios. For example, ARA/EPA decreased by 5.7–6.4 PMA at rs3734398 and rs953413, and for ARA/DHA, rs7744440, rs2281591 and rs3798719 show a ratio decrease ranging from 0.31 to 0.33 PMA. For DHA/EPA, rs3734398, rs3798713 and rs953413 show a ratio decrease ranging from 1.2 to 1.4 PMA.
Figure 4 displays Forest Plots that examine the associations with phenotypes linked to insulin regulation. The FADS SNP rs174455 was associated with both HOMA-IR (14.6% increase PMA) and fasting insulin levels (12.8% increase PMA). Three ELOVL2/5 SNPs (rs7744440, rs2281591, rs3798719) were associated with 2 h glucose (4.3–4.5% increase PMA), fasting insulin (14.3–19.8% increase PMA), and HOMA-IR (13.3–17.2% increase PMA). Forest Plots in Figure 5 illustrate associations between the AIR cohort major alleles and fasting lipids. There were strong and consistent effects between FADS major alleles and increased levels of triglyceride (6.7–14.2% PMA) and VLDL (6.5–13.3% PMA). The associations of FADS and ELOVL2/5 alleles with anthropometric characteristics by sex were also examined. Notably, in females, FADS major alleles were associated with decreased hip circumference (ranging from 0.7 to 1.2% PMA) and weight (4.1–4.2% PMA) (Figure 6). Neither FADS nor ELOVL2/5 alleles had any effect on anthropomorphic traits in men (data not shown).
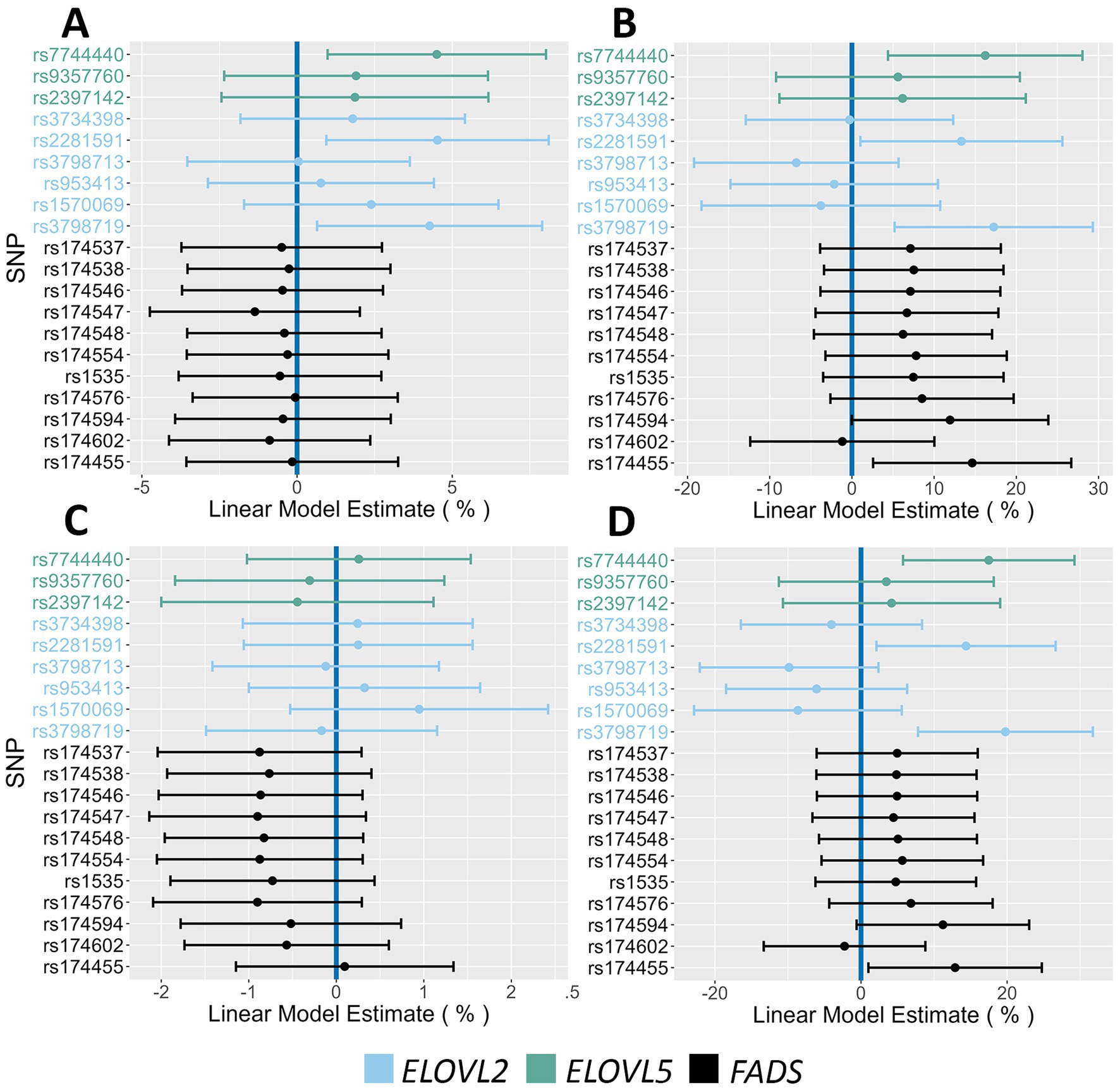
Figure 4. Impact of FADS and ELOVL genotypes on insulin action and glucose phenotypes. (A,B) Associations between 2 h glucose and HOMA-IR, respectively, and all assayed SNPs. (C,D) Associations between fasting glucose and fasting insulin, respectively, and all assayed SNPs. The 95% confidence intervals are indicated by the horizontal lines.
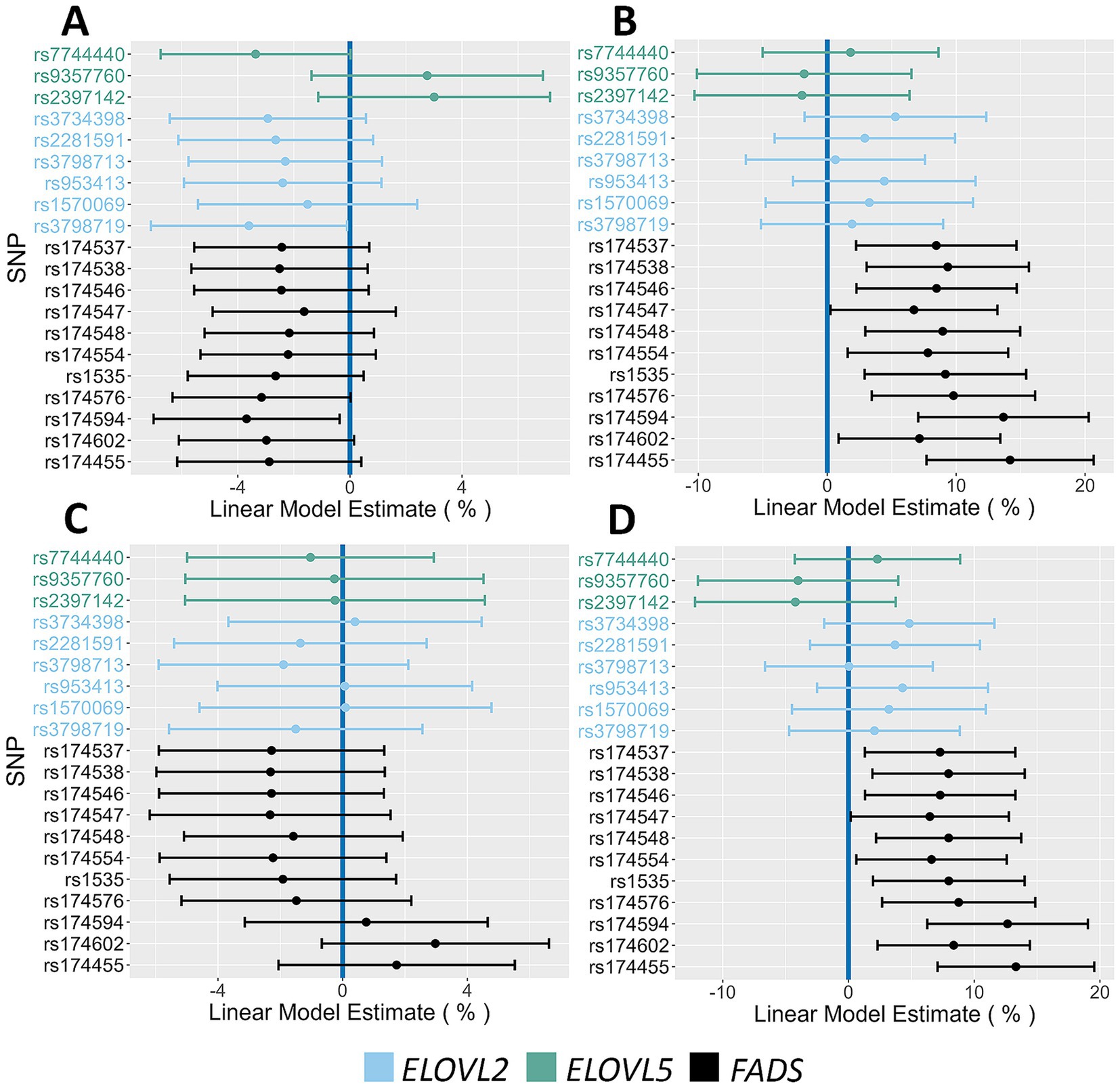
Figure 5. Impact of FADS and ELOVL genotypes on fasting lipid phenotypes. (A,B) Associations between HDL and Triglycerides, respectively, and all assayed SNPs. (C,D) Associations between LDL and VLDL, respectively, and all assayed SNPs. The 95% confidence intervals are indicated by the horizontal lines.
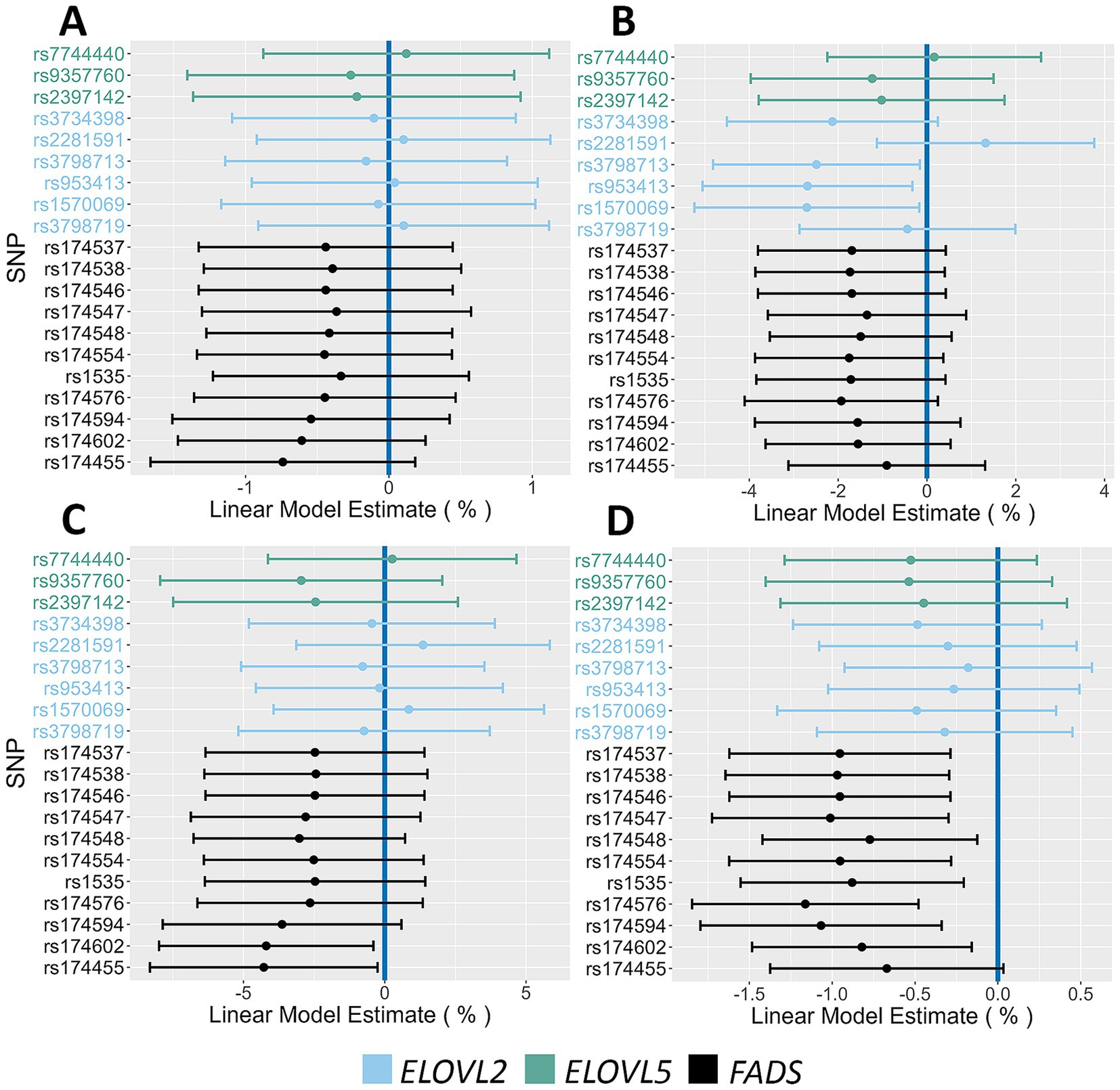
Figure 6. Impact of FADS and ELOVL genotypes on anthropometric phenotypes in females. (A,B) Associations between height and fat mass, respectively, and all assayed SNPs. (C,D) Associations between weight and hip circumference, respectively, and all assayed SNPs. The 95% confidence intervals are indicated by the horizontal lines.
The FADS variant that showed the most consistent and strongest associations with several cardiometabolic biomarkers when comparing homozygous major alleles to homozygous minor alleles was rs174455 (Table 3). For instance, HOMA-IR values were significantly higher in the homozygous major allele genotype (GG) group compared to the homozygous minor allele genotype (AA), with a 45% increase. Similarly, fasting insulin levels were elevated by an average of 43% in individuals with the GG compared to the AA genotype. Triglyceride levels were also significantly higher in the GG group, with a 33% increase, as were VLDL levels, which rose by nearly 30%. There were no significant differences observed between GG and AA genotypes for males regarding fat mass, hip circumference, height, or weight. In females, weight was significantly different between the genotypes, with those carrying the GG genotype weighing ~10% less than their AA counterparts.
Diabetes and hyperlipidemia are significant health concerns in Mexican and MxAm populations, who have higher prevalence rates compared to Non-Hispanic White (NHW) populations (60, 61). For example, MxAm are 1.7 times more likely to be diagnosed with diabetes than NHW, and over 50% of Hispanic adults (including MxAm) are predicted to develop type 2 diabetes in their lifetime (62). MxAm with diabetes also tend to have worse glycemic control and higher rates of diabetes-related complications, such as kidney disease. Additionally, MxAm are more likely to have higher triglyceride levels and lower HDL cholesterol levels compared to NHW (61, 62). Non-alcoholic fatty liver disease (NAFLD) affects about 25% of the general population, but the prevalence is higher in Hispanic populations, with MxAm having the highest rate at 42.8% (63).
Socioeconomic factors, including lower income, decreased access to healthcare, and language barriers, clearly contribute to the higher prevalence of these conditions in MxAm populations (61, 64). Genetic variants also play a role. For instance, the SLC16A11 risk variants, more common in people of Mexican ancestry, are associated with increased diabetes risk (64–66). Variations in the PNPLA3 gene, which predisposes people to store extra fat in liver cells, are found more often in those of Hispanic origin and are also associated with greater severity of NAFLD (67, 68).
In the current study of Latino adults of MxAm descent from the Southwest US, we observed high frequencies of alleles linked to the ancestral FADS haplotype, comparable to those observed in MxAm populations from the US West Coast. In contrast, these alleles are much less frequent in African and European ancestry populations and are also associated with a reduced capacity to synthesize n-6 and n-3 HUFAs, largely due to their impact on the rate-limiting FADS1 enzymatic step (15). This observation was confirmed in our study and mirrored findings in other Latino populations (36, 41, 42), where a 33 and 38% reduction in circulating levels of ARA and EPA, respectively, was seen when individuals with the TT genotype at rs174537 were compared to those with the GG genotype.
Within the TT genotype at rs174537, mean ARA levels (56 ng/mL) were 18-fold and 5-fold higher than mean EPA (3 ng/mL) and mean DHA (12 ng/mL) levels, respectively. Moreover, individual ARA/EPA ratios display a high level of variability (coefficient of variation ~109%) with a strong positive skew, leading to a mean ARA/EPA ratio of ~30 to 1 for the TT group. Although this ratio does not differ significantly between genotypes, the extremely low concentrations of EPA in TT individuals raise an important question as to whether there is an EPA deficiency with the potential to impact cardiometabolic disease. This question is especially relevant given findings from recent notable n-3 HUFA supplementation randomized controlled trials such as ASCEND, VITAL, and REDUCE-IT (69–71). Notably, only REDUCE-IT provided higher doses of EPA to patients at high risk of CVD and demonstrated a reduction in composite CVD morbidity and mortality (including CVD mortality, non-fatal MI, non-fatal stroke, cardiovascular revascularization or unstable angina) in the treatment group (69).
While the current study corroborated the frequencies of the ancestral haplotype and its impact on n-6 and n-3 HUFA levels as has been observed in other African, European and Indigenous American ancestry cohorts, the major objective here was to determine whether FADS/ELOVL variants impact important cardiometabolic traits, including triglycerides, HDL, VLDL, fat mass, and insulin resistance markers. Previously, our laboratory examined the impact of the ancestral FADS haplotype on clinical phenotypes in six Hispanic subgroups in the Multi-Ethic Study of Atherosclerosis (MESA) (42). Results from this study suggested that FADS variation was associated with circulating levels of triglycerides, HDL-C, E-selectin, s-ICAM, waist-hip ratio, height, and weight. However, only associations for triglycerides and height remained significant when the regression model included the principal components of ancestry. That study underscored the need to investigate the clinical impact of FADS variation in a more homogeneous cohort.
The key findings of the current study demonstrate that the ancestral haplotype is strongly associated with several cardiometabolic endpoints. For example, at rs174455, the homozygous ancestral FADS genotype (compared to the homozygous derived genotype) was linked to a ~40% increase in fasting insulin and HOMA-IR, a ~30% increase in triglycerides and VLDL, and modest decreases in HDL and female anthropometric measures. Interestingly, ELOVL2/5 variants were also associated with increased glucose, insulin, and HOMA-IR, indicating variation in the HUFA biosynthetic pathway outside of the FADS cluster may also affect cardiometabolic endpoints. The underlying molecular mechanisms that link FADS, circulating HUFAs, and cardiometabolic endpoints are not yet clearly understood. However, n-3 HUFA supplementation has been shown to impact insulin-stimulated glucose disposal, potentially improving insulin sensitivity (72, 73). Additionally, the ratio of EPA to ARA has been shown to be a promising indicator of glycemic control (74).
Interestingly, alleles associated with the ancestral haplotype are often strongly linked to significantly higher triglyceride levels (36, 38, 75), whereas n-3 HUFA supplementation is well-documented to reduce circulating triglyceride levels (42, 76, 77). Mechanistically, n-3 HUFAs have been shown to: (1) inhibit hepatic synthesis and secretion of VLDL particles, the primary carriers of triglycerides in the bloodstream; (2) enhance the activity of lipoprotein lipase, an enzyme that hydrolyzes triglycerides in VLDL and chylomicrons, accelerating their clearance from circulation; (3) inhibit acyl-CoA:1,2-diacylglycerol acyltransferase, a key enzyme in triglyceride biosynthesis; and (4) increase peroxisomal β-oxidation, thereby promoting fatty acid metabolism (75, 76). These findings support our hypothesis that n-3 HUFA deficiency, particularly of EPA, that results from FADS gene-by-diet PUFA interactions in MxAm individuals of Indigenous American ancestry, may contribute to insulin resistance, elevated triglyceride levels, and an increased risk of diabetes and NAFLD. Notably, our study suggests that these metabolic conditions, when linked to n-3 HUFA deficiency, may be specifically mitigated through targeted n-3 HUFA supplementation.
A limitation of our study design is the lack of dietary data in our cohort, which prevents a complete assessment of gene-by-dietary PUFA interactions. However, we recently published data from a similar cohort of MxAm from the same region in Arizona that provide relevant context (77). In that cohort, approximately 35% of total caloric intake came from fat, with an average total fat intake of 59.6 g/day. PUFAs contributed 5–10% of energy, primarily from the n-6 PUFA, LA, with a mean intake of 12 ± 6.5 g/day (6.9 ± 2.2% of total energy). The mean n-6 to n-3 PUFA ratio in this population was 8.8:1, closely resembling the dietary patterns of other populations consuming a modern Western diet. Another limitation of our study is the lack of data on lifestyle factors and supplementation (e.g., PUFA supplements), which could influence the interpretation of our findings.
While the study’s focus on self-reported Latinos of Mexican origin from the AIR registry provided a targeted dataset, it may limit the generalizability of our findings to other Latino populations due to potential underrepresentation of broader genetic diversity. However, our analysis of the MESA cohort corroborates these findings, revealing strong negative associations between Indigenous American genetic ancestry and HUFA levels. Furthermore, the FADS rs174537 SNP accounted for a substantial portion of the ancestry-related effects on n-3 HUFAs, including EPA, and showed strong associations with various metabolic, inflammatory, and anthropometric traits, particularly circulating triglycerides.
Taken together, the current study suggests that gene-by-diet interactions associated with the quantities and ratios of dietary PUFAs, combined with the FADS ancestral haplotype, may play an important role in risk, complications and cardiometabolic disease disparities observed in MxAm populations. This study highlights the complex interplay between genetic variants, particularly in the FADS genes, and cardiometabolic traits in MxAm populations. The findings also underscore the importance of considering both genetic predispositions and environmental factors, such as diet, in understanding the health disparities that disproportionately affect these populations. Importantly, the gene-by-diet interactions involving PUFAs, HUFAs, and their downstream metabolic effects point to potential avenues for personalized nutrition and therapeutic strategies to mitigate cardiometabolic risk in this population.
Future research should further explore how dietary interventions, particularly those aimed at optimizing PUFA intake or supplementing with n-3 HUFAs, could modify the effects of FADS variants. This is particularly relevant when interactions between the FADS gene and dietary PUFAs result in an increased risk of n-3 HUFA deficiency, which would be expected in the 40–50% of MxAm homozygous for the ancestral haplotype. Given the high prevalence of hyperlipidemia in this population and the biological mechanisms linking n-3 HUFAs to elevated lipids, individuals in this group may especially benefit from n-3 HUFA supplementation. Supplementation with doses ranging from 2 to 4 grams per day has been shown to be most effective in lowering triglyceride levels, and the beneficial effects are typically observed within 6 weeks and can continue to improve over a period of up to 6 months in most studies (78). Additionally, expanding studies to other Latino populations and controlling for other socioeconomic and lifestyle factors will be critical in developing targeted prevention and treatment programs. Ultimately, addressing these gene-by-diet interactions could help reduce the burden of diabetes, cardiovascular disease, and related conditions in MxAm, Mexican and other Latino communities.
The datasets presented in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found in the article/Supplementary material.
The studies involving humans were approved by the Institutional Review Board (IRB) of Arizona State University, and each participant provided written informed consent before taking part in the study. AIR is registered as an exempt protocol under the UA IRB #1703255156. The studies were conducted in accordance with the local legislation and institutional requirements. The participants provided their written informed consent to participate in this study.
SB: Data curation, Formal analysis, Investigation, Software, Validation, Visualization, Writing – original draft, Writing – review & editing. JA: Investigation, Writing – original draft. BH: Formal analysis, Methodology, Validation, Writing – review & editing. YK: Methodology, Writing – review & editing. LG: Investigation, Writing – original draft. LM: Resources, Writing – review & editing. DC: Methodology, Resources, Writing – review & editing, Supervision. FC: Conceptualization, Funding acquisition, Methodology, Project administration, Resources, Supervision, Writing – review & editing.
The author(s) declare that financial support was received for the research, authorship, and/or publication of this article. This study was funded by NIH (National Center for Complementary and Integrative Health) (R01 AT008621 to FHC) and U.S. Department of Agriculture (ARZT-1361680-H23-157 to FHC).
FC hilton is a cofounder of Resonance Pharma, Inc. (Ann Arbor, MI, USA). This company develops diagnostics and therapeutics for phospholipase enzymes. This relationship is managed by the Office for Responsible Outside Interests at the University of Arizona.
The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The authors declare that no Gen AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
The Supplementary material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fnut.2025.1538505/full#supplementary-material
1. Blasbalg, TL, Hibbeln, JR, Ramsden, CE, Majchrzak, SF, and Rawlings, RR. Changes in consumption of omega-3 and omega-6 fatty acids in the United States during the 20th century. Am J Clin Nutr. (2011) 93:950–62. doi: 10.3945/ajcn.110.006643
2. Hambleton, IR, Caixeta, R, Jeyaseelan, SM, Luciani, S, and Hennis, AJM. The rising burden of non-communicable diseases in the Americas and the impact of population aging: a secondary analysis of available data. Lancet Reg Health Am. (2023) 21:21. doi: 10.1016/j.lana.2023.100483
3. Richards, NC, Gouda, HN, Durham, J, Rampatige, R, Rodney, A, and Whittaker, M. Disability, noncommunicable disease and health information. Bull World Health Organ. (2016) 94:230–2. doi: 10.2471/blt.15.156869
4. World Health Organization. Noncommunicable diseases: Mortality (2023). Available online at: https://www.who.int/data/gho/data/themes/topics/topic-details/GHO/ncd-mortality (Accessed September 26, 2024).
5. Koyama, AK, McKeever Bullard, K, Xu, F, Onufrak, S, Jackson, SL, Saelee, R, et al. Prevalence of cardiometabolic diseases among racial and ethnic subgroups in adults - behavioral risk factor surveillance system, United States, 2013-2021. MMWR Morb Mortal Wkly Rep. (2024) 73:51–6. doi: 10.15585/mmwr.mm7303a1
6. Tsao, CW, Aday, AW, Almarzooq, ZI, Alonso, A, Beaton, AZ, Bittencourt, MS, et al. Heart disease and stroke statistics—2022 update: a report from the American Heart Association. Circulation. (2022) 145:e153–639. doi: 10.1161/CIR.0000000000001052
7. Lopez-Neyman, SM, Davis, K, Zohoori, N, Broughton, KS, Moore, CE, and Miketinas, D. Racial disparities and prevalence of cardiovascular disease risk factors, cardiometabolic risk factors, and cardiovascular health metrics among us adults: NHANES 2011-2018. Sci Rep. (2022) 12:19475. doi: 10.1038/s41598-022-21878-x
8. Cordain, L, Eaton, SB, Sebastian, A, Mann, N, Lindeberg, S, Watkins, BA, et al. Origins and evolution of the Western diet: health implications for the 21st century. Am J Clin Nutr. (2005) 81:341–54. doi: 10.1093/ajcn.81.2.341
9. Miller, M, Stone, NJ, Ballantyne, C, Bittner, V, Criqui, MH, Ginsberg, HN, et al. Triglycerides and cardiovascular disease: a scientific statement from the American Heart Association. Circulation. (2011) 123:2292–333. doi: 10.1161/CIR.0b013e3182160726
10. Dietary fat and its relation to heart attacks and strokes. Report by the Central Committee for Medical and Community Program of the American Heart Association. JAMA. (1961) 175:389–91. doi: 10.1001/jama.1961.63040050001011
11. Brenna, JT, Salem, N Jr, Sinclair, AJ, and Cunnane, SC. Alpha-Linolenic acid supplementation and conversion to n-3 long-chain polyunsaturated fatty acids in humans. Prostaglandins Leukot Essent Fatty Acids. (2009) 80:85–91. doi: 10.1016/j.plefa.2009.01.004
12. Zhang, JY, Kothapalli, KS, and Brenna, JT. Desaturase and elongase-limiting endogenous long-chain polyunsaturated fatty acid biosynthesis. Curr Opin Clin Nutr Metab Care. (2016) 19:103–10. doi: 10.1097/mco.0000000000000254
13. Park, WJ, Kothapalli, KS, Lawrence, P, Tyburczy, C, and Brenna, JT. An alternate pathway to long-chain polyunsaturates: the FADS2 gene product Delta8-desaturates 20:2n-6 and 20:3n-3. J Lipid Res. (2009) 50:1195–202. doi: 10.1194/jlr.M800630-JLR200
14. Sprecher, H, and Chen, Q. Polyunsaturated fatty acid biosynthesis: a microsomal-peroxisomal process. Prostaglandins Leukot Essent Fatty Acids. (1999) 60:317–21. doi: 10.1016/s0952-3278(99)80006-4
15. Reynolds, LM, Dutta, R, Seeds, MC, Lake, KN, Hallmark, B, Mathias, RA, et al. FADS genetic and metabolomic analyses identify the ∆5 desaturase (FADS1) step as a critical control point in the formation of biologically important lipids. Sci Rep. (2020) 10:15873. doi: 10.1038/s41598-020-71948-1
16. Kothapalli, KSD, Park, HG, and Brenna, JT. Polyunsaturated fatty acid biosynthesis pathway and genetics. implications for interindividual variability in prothrombotic, inflammatory conditions such as COVID-19. Prostaglandins Leukot Essent Fat Acids. (2020) 162:102183. doi: 10.1016/j.plefa.2020.102183
17. Gieger, C, Geistlinger, L, Altmaier, E, Hrabé de Angelis, M, Kronenberg, F, Meitinger, T, et al. Genetics meets metabolomics: a genome-wide association study of metabolite profiles in human serum. PLoS Genet. (2008) 4:e1000282. doi: 10.1371/journal.pgen.1000282
18. Haeggström, JZ, and Funk, CD. Lipoxygenase and leukotriene pathways: biochemistry, biology, and roles in disease. Chem Rev. (2011) 111:5866–98. doi: 10.1021/cr200246d
19. Serhan, CN, and Chiang, N. Resolution phase lipid mediators of inflammation: agonists of resolution. Curr Opin Pharmacol. (2013) 13:632–40. doi: 10.1016/j.coph.2013.05.012
20. Smith, WL. The eicosanoids and their biochemical mechanisms of action. Biochem J. (1989) 259:315–24. doi: 10.1042/bj2590315
21. Borgeat, P, Hamberg, M, and Samuelsson, B. Transformation of arachidonic acid and homo-gamma-linolenic acid by rabbit polymorphonuclear leukocytes. Monohydroxy acids from novel lipoxygenases. J Biol Chem. (1976) 251:7816–20. doi: 10.1016/S0021-9258(19)57008-9
22. Sergeant, S, Rahbar, E, and Chilton, FH. Gamma-linolenic acid, dihommo-gamma linolenic, eicosanoids and inflammatory processes. Eur J Pharmacol. (2016) 785:77–86. doi: 10.1016/j.ejphar.2016.04.020
23. Fan, YY, Ramos, KS, and Chapkin, RS. Dietary gamma-linolenic acid modulates macrophage-vascular smooth muscle cell interactions. Evidence for a macrophage-derived soluble factor that downregulates DNA synthesis in smooth muscle cells. Arterioscler Thromb Vasc Biol. (1995) 15:1397–403. doi: 10.1161/01.atv.15.9.1397
24. Zurier, RB. Role of prostaglandins E in inflammation and immune responses. Adv Prostaglandin Thromboxane Leukot Res. (1991) 21b:947–53.
25. Tabolacci, C, Lentini, A, Provenzano, B, Gismondi, A, Rossi, S, and Beninati, S. Similar antineoplastic effects of nimesulide, a selective Cox-2 inhibitor, and prostaglandin E1 on B16-F10 murine melanoma cells. Melanoma Res. (2010) 20:273–9. doi: 10.1097/CMR.0b013e328339d8ac
26. Serhan, CN, Arita, M, Hong, S, and Gotlinger, K. Resolvins, docosatrienes, and neuroprotectins, novel omega-3-derived mediators, and their endogenous aspirin-triggered epimers. Lipids. (2004) 39:1125–32. doi: 10.1007/s11745-004-1339-7
27. Ariel, A, and Serhan, CN. Resolvins and protectins in the termination program of acute inflammation. Trends Immunol. (2007) 28:176–83. doi: 10.1016/j.it.2007.02.007
28. Schaeffer, L, Gohlke, H, Müller, M, Heid, IM, Palmer, LJ, Kompauer, I, et al. Common genetic variants of the FADS1 FADS2 gene cluster and their reconstructed haplotypes are associated with the fatty acid composition in phospholipids. Hum Mol Genet. (2006) 15:1745–56. doi: 10.1093/hmg/ddl117
29. Chilton, FH, Dutta, R, Reynolds, LM, Sergeant, S, Mathias, RA, and Seeds, MC. FADS genotypes and desaturase activity estimated by the ratio of arachidonic acid to linoleic acid are associated with inflammation and coronary artery disease. Nutrients. (2017) 9:1165. doi: 10.3390/nu9111165
30. Martinelli, N, Girelli, D, Malerba, G, Guarini, P, Illig, T, Trabetti, E, et al. FADS genotypes and desaturase activity estimated by the ratio of arachidonic acid to linoleic acid are associated with inflammation and coronary artery disease. Am J Clin Nutr. (2008) 88:941–9. doi: 10.1093/ajcn/88.4.941
31. Koletzko, B, Reischl, E, Tanjung, C, Gonzalez-Casanova, I, Ramakrishnan, U, Meldrum, S, et al. FADS1 and FADS2 polymorphisms modulate fatty acid metabolism and dietary impact on health. Annu Rev Nutr. (2019) 39:21–44. doi: 10.1146/annurev-nutr-082018-124250
32. Fadason, T, Schierding, W, Lumley, T, and O’Sullivan, JM. Chromatin interactions and expression quantitative trait loci reveal genetic drivers of multimorbidities. Nat Commun. (2018) 9:5198. doi: 10.1038/s41467-018-07692-y
33. Hester, AG, Murphy, RC, Uhlson, CJ, Ivester, P, Lee, TC, Sergeant, S, et al. Relationship between a common variant in the fatty acid desaturase (FADS) cluster and eicosanoid generation in humans. J Biol Chem. (2014) 289:22482–9. doi: 10.1074/jbc.M114.579557
34. Mathias, RA, Sergeant, S, Ruczinski, I, Torgerson, DG, Hugenschmidt, CE, Kubala, M, et al. The impact of FADS genetic variants on ω6 polyunsaturated fatty acid metabolism in African Americans. BMC Genet. (2011) 12:50. doi: 10.1186/1471-2156-12-50
35. Sergeant, S, Hugenschmidt, CE, Rudock, ME, Ziegler, JT, Ivester, P, Ainsworth, HC, et al. Differences in arachidonic acid levels and fatty acid desaturase (FADS) gene variants in African Americans and European Americans with diabetes or the metabolic syndrome. Br J Nutr. (2012) 107:547–55. doi: 10.1017/s0007114511003230
36. Sergeant, S, Keith, BA, Seeds, MC, Legins, JA, Young, CB, Vitolins, MZ, et al. Impact of FADS gene variation and dietary fatty acid exposure on biochemical and anthropomorphic phenotypes in a Hispanic/Latino cohort. Front Nutr. (2023) 10:1111624. doi: 10.3389/fnut.2023.1111624
37. Harris, DN, Ruczinski, I, Yanek, LR, Becker, LC, Becker, DM, Guio, H, et al. Evolution of hominin polyunsaturated fatty acid metabolism: from Africa to the New World. Genome Biol Evol. (2019) 11:1417–30. doi: 10.1093/gbe/evz071
38. Mathias, RA, Fu, W, Akey, JM, Ainsworth, HC, Torgerson, DG, Ruczinski, I, et al. Adaptive evolution of the FADS gene cluster within Africa. PLoS One. (2012) 7:e44926. doi: 10.1371/journal.pone.0044926
39. Mathieson, S, and Mathieson, I. FADS1 and the timing of human adaptation to agriculture. Mol Biol Evol. (2018) 35:2957–70. doi: 10.1093/molbev/msy180
40. Fumagalli, M, Moltke, I, Grarup, N, Racimo, F, Bjerregaard, P, Jorgensen, ME, et al. Greenlandic Inuit show genetic signatures of diet and climate adaptation. Science. (2015) 349:1343–7. doi: 10.1126/science.aab2319
41. Downie, CG, Highland, HM, Alotaibi, M, Welch, BM, Howard, AG, Cheng, S, et al. Genome-wide association study reveals shared and distinct genetic architecture underlying fatty acid and bioactive oxylipin metabolites in the Hispanic Community Health Study/Study of Latinos (HCHS/SOL). medRxiv. (2024). doi: 10.1101/2024.05.21.24307719
42. Yang, C, Hallmark, B, Chai, JC, O'Connor, TD, Reynolds, LM, Wood, AC, et al. Impact of Amerind ancestry and FADS genetic variation on omega-3 deficiency and cardiometabolic traits in Hispanic populations. Commun Biol. (2021) 4:918. doi: 10.1038/s42003-021-02431-4
43. DeMenna, J, Puppala, S, Chittoor, G, Schneider, J, Kim, JY, Shaibi, GQ, et al. Association of common genetic variants with diabetes and metabolic syndrome related traits in the Arizona Insulin Resistance registry: a focus on Mexican American families in the Southwest. Hum Hered. (2014) 78:47–58. doi: 10.1159/000363411
44. Shaibi, GQ, Coletta, DK, Vital, V, and Mandarino, LJ. The design and conduct of a community-based registry and biorepository: a focus on cardiometabolic health in Latinos. Clin Transl Sci. (2013) 6:429–34. doi: 10.1111/cts.12114
45. Tanaka, T, Shen, J, Abecasis, GR, Kisialiou, A, Ordovas, JM, Guralnik, JM, et al. Genome-wide association study of plasma polyunsaturated fatty acids in the Inchianti study. PLoS Genet. (2009) 5:e1000338. doi: 10.1371/journal.pgen.1000338
46. Mathias, RA, Vergara, C, Gao, L, Rafaels, N, Hand, T, Campbell, M, et al. FADS genetic variants and omega-6 polyunsaturated fatty acid metabolism in a homogeneous island population. J Lipid Res. (2010) 51:2766–74. doi: 10.1194/jlr.M008359
47. Lemaitre, RN, Tanaka, T, Tang, W, Manichaikul, A, Foy, M, Kabagambe, EK, et al. Genetic loci associated with plasma phospholipid n-3 fatty acids: a meta-analysis of genome-wide association studies from the CHARGE Consortium. PLoS Genet. (2011) 7:e1002193. doi: 10.1371/journal.pgen.1002193
48. Buckley, MT, Racimo, F, Allentoft, ME, Jensen, MK, Jonsson, A, Huang, H, et al. Selection in Europeans on fatty acid desaturases associated with dietary changes. Mol Biol Evol. (2017) 34:1307–18. doi: 10.1093/molbev/msx103
49. Li, X, Gan, ZW, Ding, Z, Wu, YX, Chen, XY, Tian, HM, et al. Genetic variants in the ELOVL5 but not ELOVL2 gene associated with polyunsaturated fatty acids in Han Chinese breast milk. Biomed Environ Sci. (2017) 30:64–7. doi: 10.3967/bes2017.008
50. Wu, Y, Wang, Y, Tian, H, Lu, T, Yu, M, Xu, W, et al. DHA intake interacts with ELOVL2 and ELOVL5 genetic variants to influence polyunsaturated fatty acids in human milk. J Lipid Res. (2019) 60:1043–9. doi: 10.1194/jlr.M090951
51. Metcalfe, LD, Schmitz, AA, and Pelka, JR. Rapid preparation of fatty acid esters from lipids for gas chromatographic analysis. Anal Chem. (1966) 38:514–5. doi: 10.1021/ac60235a044
52. Sergeant, S, Ruczinski, I, Ivester, P, Lee, TC, Morgan, TM, Nicklas, BJ, et al. Impact of methods used to express levels of circulating fatty acids on the degree and direction of associations with blood lipids in humans. Br J Nutr. (2016) 115:251–61. doi: 10.1017/S0007114515004341
53. Weaver, KL, Ivester, P, Seeds, M, Case, LD, Arm, JP, and Chilton, FH. Effect of dietary fatty acids on inflammatory gene expression in healthy humans. J Biol Chem. (2009) 284:15400–7. doi: 10.1074/jbc.M109.004861
54. Brenna, JT, Plourde, M, Stark, KD, Jones, PJ, and Lin, YH. Best practices for the design, laboratory analysis, and reporting of trials involving fatty acids. Am J Clin Nutr. (2018) 108:211–27. doi: 10.1093/ajcn/nqy089
55. McArdle, PF, O'Connell, JR, Pollin, TI, Baumgarten, M, Shuldiner, AR, Peyser, PA, et al. Accounting for relatedness in family based genetic association studies. Hum Hered. (2007) 64:234–42. doi: 10.1159/000103861
56. Interpreting log transformations in a linear model. Available online at: https://library.virginia.edu/data/articles/interpreting-log-transformations-in-a-linear-model (Accessed October 3, 2024).
57. Benjamini, Y, Drai, D, Elmer, G, Kafkafi, N, and Golani, I. Controlling the false discovery rate in behavior genetics research. Behav Brain Res. (2001) 125:279–84. doi: 10.1016/s0166-4328(01)00297-2
58. Hawe, JS, Wilson, R, Schmid, KT, Zhou, L, Lakshmanan, LN, Lehne, BC, et al. Genetic variation influencing DNA methylation provides insights into molecular mechanisms regulating genomic function. Nat Genet. (2022) 54:18–29. doi: 10.1038/s41588-021-00969-x
59. Hermant, X, Delay, C, Flaig, A, Luque-Bedregal, J, Briand, G, Bout, MA, et al. Identification of a functional FADS1 3’UTR variant associated with erythrocyte n-6 polyunsaturated fatty acids levels. J Clin Lipidol. (2018) 12:1280–9. doi: 10.1016/j.jacl.2018.07.012
60. Statistics About Diabetes: American Diabetes Association. Available online at: https://diabetes.org/about-diabetes/statistics/about-diabetes (Accessed September 11, 2024).
61. Aguayo-Mazzucato, C, Diaque, P, Hernandez, S, Rosas, S, Kostic, A, and Caballero, AE. Understanding the growing epidemic of type 2 diabetes in the Hispanic population living in the United States. Diabetes Metab Res Rev. (2019) 35:e3097. doi: 10.1002/dmrr.3097
62. Vidal, TM, Williams, CA, Ramoutar, UD, and Haffizulla, F. Type 2 diabetes mellitus in Latinx populations in the United States: a culturally relevant literature review. Cureus. (2022) 14:e23173. doi: 10.7759/cureus.23173
63. Saab, S, Manne, V, Nieto, J, Schwimmer, JB, and Chalasani, NP. Nonalcoholic fatty liver disease in Latinos. Clin Gastroenterol Hepatol. (2016) 14:5–12. doi: 10.1016/j.cgh.2015.05.001
64. Fernandez, ML. Lifestyle factors and genetic variants associated to health disparities in the Hispanic population. Nutrients. (2021) 13:2189. doi: 10.3390/nu13072189
65. Mercader, JM, and Florez, JC. The genetic basis of type 2 diabetes in Hispanics and Latin Americans: challenges and opportunities. Front Public Health. (2017) 5:5. doi: 10.3389/fpubh.2017.00329
66. Williams, AL, Jacobs, SBR, Moreno-Macías, H, Huerta-Chagoya, A, Churchhouse, C, Márquez-Luna, C, et al. Sequence variants in SLC16A11 are a common risk factor for type 2 diabetes in Mexico. Nature. (2014) 506:97–101. doi: 10.1038/nature12828
67. Trépo, E, Romeo, S, Zucman-Rossi, J, and Nahon, P. PNPLA3 gene in liver diseases. J Hepatol. (2016) 65:399–412. doi: 10.1016/j.jhep.2016.03.011
68. Romeo, S, Kozlitina, J, Xing, C, Pertsemlidis, A, Cox, D, Pennacchio, LA, et al. Genetic variation in PNPLA3 confers susceptibility to nonalcoholic fatty liver disease. Nat Genet. (2008) 40:1461–5. doi: 10.1038/ng.257
69. Bhatt, DL, Steg, PG, Miller, M, Brinton, EA, Jacobson, TA, Ketchum, SB, et al. Cardiovascular risk reduction with icosapent ethyl for hypertriglyceridemia. N Engl J Med. (2019) 380:11–22. doi: 10.1056/NEJMoa1812792
70. Bowman, L, Mafham, M, Wallendszus, K, Stevens, W, Buck, G, Barton, J, et al. Effects of n-3 fatty acid supplements in diabetes mellitus. N Engl J Med. (2018) 379:1540–50. doi: 10.1056/NEJMoa1804989
71. Manson, JE, Cook, NR, Lee, IM, Christen, W, Bassuk, SS, Mora, S, et al. Marine n-3 fatty acids and prevention of cardiovascular disease and cancer. N Engl J Med. (2019) 380:23–32. doi: 10.1056/NEJMoa1811403
72. Egalini, F, Guardamagna, O, Gaggero, G, Varaldo, E, Giannone, B, Beccuti, G, et al. The effects of omega 3 and omega 6 fatty acids on glucose metabolism: an updated review. Nutrients. (2023) 15:15 (12). doi: 10.3390/nu15122672
73. Sinha, S, Haque, M, Lugova, H, and Kumar, S. The effect of omega-3 fatty acids on insulin resistance. Life. (2023) 13:1322. doi: 10.3390/life13061322
74. Poreba, M, Rostoff, P, Siniarski, A, Mostowik, M, Golebiowska-Wiatrak, R, Nessler, J, et al. Relationship between polyunsaturated fatty acid composition in serum phospholipids, systemic low-grade inflammation, and glycemic control in patients with type 2 diabetes and atherosclerotic cardiovascular disease. Cardiovasc Diabetol. (2018) 17:29. doi: 10.1186/s12933-018-0672-5
75. Bays, HE, Tighe, AP, Sadovsky, R, and Davidson, MH. Prescription omega-3 fatty acids and their lipid effects: physiologic mechanisms of action and clinical implications. Expert Rev Cardiovasc Ther. (2008) 6:391–409. doi: 10.1586/14779072.6.3.391
76. Shearer, GC, Savinova, OV, and Harris, WS. Fish oil -- how does it reduce plasma triglycerides? Biochim Biophys Acta. (2012) 1821:843–51. doi: 10.1016/j.bbalip.2011.10.011
77. Lopez-Pentecost, M, Hallmark, B, Thomson, CA, Chilton, F, and Garcia, DO. Association between dietary fatty acid intake and liver steatosis and fibrosis in a sample of Mexican-origin Hispanic adults with overweight or obesity. Int J Environ Res Public Health. (2023) 20:3103. doi: 10.3390/ijerph20043103
Keywords: nutrigenomics, health disparities, omega-6, omega-3, lipids, nutrition, precision nutrition
Citation: Blomquist SA, Albrecht JH, Hallmark B, Klimentidis YC, Garcia LA, Mandarino LJ, Coletta DK and Chilton FH (2025) The influence of FADS genetic variation and omega-3 fatty acid deficiency on cardiometabolic disease risk in a Mexican American population. Front. Nutr. 12:1538505. doi: 10.3389/fnut.2025.1538505
Received: 02 December 2024; Accepted: 18 February 2025;
Published: 10 March 2025.
Edited by:
Sonia Roman, University of Guadalajara, MexicoReviewed by:
Ramona Suharoschi, University of Agricultural Sciences and Veterinary Medicine of Cluj-Napoca, RomaniaCopyright © 2025 Blomquist, Albrecht, Hallmark, Klimentidis, Garcia, Mandarino, Coletta and Chilton. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Floyd H. Chilton, ZmNoaWx0b25AYXJpem9uYS5lZHU=
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.