
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Sustain. Food Syst. , 17 July 2020
Sec. Crop Biology and Sustainability
Volume 4 - 2020 | https://doi.org/10.3389/fsufs.2020.00097
 Karol L. Krey1*†
Karol L. Krey1*† Paul D. Nabity2
Paul D. Nabity2 Carmen K. Blubaugh3
Carmen K. Blubaugh3 Zhen Fu1,4
Zhen Fu1,4 James T. Van Leuven5
James T. Van Leuven5 John P. Reganold6
John P. Reganold6 Anna Berim7
Anna Berim7 David R. Gang7
David R. Gang7 Andrew S. Jensen8
Andrew S. Jensen8 William E. Snyder9
William E. Snyder9Plants deploy a variety of chemical and physical defenses to protect themselves against herbivores and pathogens. Organic farming seeks to enhance these responses by improving soil quality, ultimately altering bottom up regulation of plant defenses. While laboratory studies suggest this approach is effective, it remains unclear whether organic agriculture encourages more-active plant defenses under real-world conditions. Working on the farms of cooperating growers, we examined gene expression in the leaves of two potato (Solanum tuberosum) varieties, grown on organic vs. conventional farms. For one variety, Norkotah, we found significantly heightened initiation of genes associated with plant-defense pathways in plants grown in organic vs. conventional fields. Organic Norkotah fields exhibited lower levels of nitrate in soil and of nitrogen in plant foliage, alongside differences in communities of soil bacteria, suggesting possible links between soil management and observed differences in plant defenses. Additionally, numbers of predatory and phloem-feeding insects were higher in organic than conventional fields. A second potato variety, Alturas, which is generally grown using fewer inputs and in poorer-quality soils, exhibited lower overall herbivore and predator numbers, few differences in soil ecology, and no differences in gene-activity in organic and conventional farming systems. Altogether, our results suggest that organic farming has the potential to increase plants' resistance to herbivores, possibly facilitating reduced need for insecticide applications. These benefits appear to be mediated by plant variety and/or farming context.
Conventional farming methods often largely rely on chemical insecticides for the control of herbivorous insects (e.g., Soffe, 2002; Stockdale et al., 2002; Klonsky, 2012). This approach can inadvertently disrupt biological control and worsen pest outbreaks when broad-acting chemicals kill natural enemies (Penman and Chapman, 1988; Prischmann et al., 2005; Woods et al., 2012). Perhaps less appreciated, however, are the ways that conventional farming methods might disrupt “bottom-up” suppression of herbivores by plants. Plants utilize a broad range of chemical and physical defenses against their herbivores (Feeny, 1976; Price, 1991; Howe and Jander, 2008; Mithöfer and Boland, 2012), and induction of these defenses might be impacted by farming practices (e.g., Davis et al., 2001; Bonanomi et al., 2010). For example, synthetic chemical fertilizers provide sudden nutrient pulses that can be physiologically stressful for plants, including disruption of pathways related to plant defense against herbivores (Altieri and Nicholls, 2003; Bhardwaj et al., 2014). Likewise, several insecticides are known to alter plant physiology to the unintended benefit of plant-feeding insects (Karthikeyan et al., 2009; Ford et al., 2010; Szczepaniec et al., 2013). For example, neonicotinoid insecticides can suppress expression of important plant defense genes, alter levels of phytohormones involved in plant defense, and decrease plant resistance to herbivores such as spider mites (e.g., James and Price, 2002; Szczepaniec et al., 2013). The role of herbivores and their significance in causing large-scale changes in gene expression has been widely studied (Cheong et al., 2002; Delessert et al., 2004; Reymond et al., 2004; Smith et al., 2004; Voelckel and Baldwin, 2004; Zhu-Salzman et al., 2004; De Vos et al., 2005; Schmidt et al., 2005; Ralph et al., 2006; Thompson and Goggin, 2006; Broekgaarden et al., 2007). Therefore, conventional modern agriculture can impact both top-down herbivore suppression, by harming natural enemies, and bottom-up herbivore suppression, by disrupting plant defenses.
As an alternative approach, organic farming seeks to replace synthetic chemical insecticides with natural pest controls (Rigby and Cáceres, 2001; Pimentel et al., 2005). Indeed, reduced insecticide sprays in these systems likely contribute to more abundant and biodiverse communities of predatory insects (Crowder et al., 2010), spiders (Letourneau and Goldstein, 2001), and songbirds (Beecher et al., 2002; Bengtsson et al., 2005; Hole et al., 2005; Bouvier et al., 2011) on organic compared to conventional farms, which improves biological control of pest insects (Bengtsson et al., 2005; Birkhofer et al., 2008; Crowder et al., 2010). It is also possible that organic farming could improve bottom-up pest regulation (Altieri and Nicholls, 2003). For example, by eliminating synthetic chemical insecticides, organic methods would be expected to avoid any negative effects these chemicals have on plant defenses (discussed above). Beyond this, organic farming replaces synthetic chemical fertilizers with composted animal manures, cover crops, and other biological sources of fertility (Stockdale et al., 2002; Watson et al., 2002; Pimentel et al., 2005). These inputs tend to release nitrogen and other nutrients relatively slowly compared to synthetic fertilizers (Eigenbrode and Pimentel, 1988; Phelan et al., 1995, 1996; Hsu et al., 2009), and steady nutrient release likely exerts widespread impacts on plant physiology including defense (McGuiness, 1993; Phelan et al., 1995). These fertility-management approaches also increase organic matter in the soil, an explicit goal of organic farming (e.g., Jenny, 1980; Reganold et al., 1987; McGuiness, 1993; Altieri and Nicholls, 2003), which can increase biomass and biodiversity of soil-dwelling organisms (e.g., Gunapala and Scow, 1998; Swezey et al., 1998; Altieri, 1999; Mäder et al., 2002). Soil microbes are known to induce or prime plant defenses that provide indirect protection against herbivorous insects (Davis et al., 2001; Bonanomi et al., 2010). While studies in controlled settings have indeed suggested enhanced defenses for plants grown using organic vs. conventional soil amendments (e.g., Bloemberg and Lugtenberg, 2001; Lugtenberg et al., 2002; Bais et al., 2004; Morrissey et al., 2004; Goh et al., 2013), it remains unclear whether this would also be seen in the more-complex setting of real farms varying in a wide diversity of farming practices.
To examine the numerous physiological mechanisms by which organic farming may improve pest resistance, we used RNA sequencing (RNA-seq) and compared expression of defense genes in the foliage of commercially grown potato (Solanum tuberosum) crops collected from five organic and seven conventional fields. These farms were located across the potato-growing region of central and southern Washington (Supplementary Table 1), and we considered the two most common potato varieties grown in the region, Norkotah and Alturas. Our samples were timed to be at mid-growing season when plants in all fields were flowering and when pest damage, and thus risks to crop production, are greatest (O. Johnson, commercial potato grower, personal communication). To assess ecological factors beyond farming system that could contribute to defense-gene expression, we also intensively sampled arthropod communities and collected soils to document soil chemical and biological properties (including assessing soil microbial diversity using 16s rRNA sequencing), as we collected potato foliage. Our goal was not just to look for consistent defense-gene expression differences between the two farming systems, but also to initiate a search for specific factors differing between the two farming systems that could ultimately underlie correlative differences in gene expression patterns.
From fields of Norkotah and Alturas potato varieties of cooperating growers throughout central Washington (5 organic and 7 conventional, Supplementary Table 1), we collected damaged and undamaged potato foliage for RNA-seq analysis; intensively sampled communities of predatory and herbivorous arthropods; and sampled soils for chemical and microbial characterization. Below, we provide detailed methodologies for each of these components.
In Washington State, conventional potato fields typically manage pest insects using applications of broad-spectrum neonicotinoid and pyrethroid insecticides, while plant pathogens are controlled using pre-plant soil fumigation followed by seed treatment fungicides, late blight preventatives such as mancozeb, and various products aimed at pathogens that emerge in the crop during the growing season (Murray et al., 2019). In contrast, pest insects in organic potato fields are typically treated with neem extract and Bacillus thuringiensis, while plant pathogens are controlled using copper applications (Murray et al., 2019). Additional differences include herbicides for weed suppression and synthetic chemical fertilizers to enhance fertility in conventional fields, vs. frequent tillage and applications of manures and mulches in the organic system (Koss et al., 2005; Murray et al., 2019).
All potato plants, for both varieties and under each of the two farming systems, were sampled when plants were flowering, which occurs for several weeks during the mid-growing season. This sampling time coincides with particularly rapid foliar growth and tuber initiation, when pest damage can be particularly harmful to later yields (Stark et al., 2020). Within a 400-m2 area of each farm, we randomly chose 10 insect-damaged (with visible chewing or piercing sucking damage) leaves and 10 undamaged leaves from 20 individual plants that were located at mid-plant level (top leaves are newer and prone to excess defense; Zangerl and Bazzaz, 1992; Hartley and Jones, 1997). While wearing nitrile gloves, we used a sterile razor blade to quickly cut each leaf at the stem, inserting each leaf into a 50-ml conical Falcon tube and immediately placing it into a container of liquid nitrogen. Although all plants had some damage, the undamaged leaves chosen were visibly free of damage. The tubes were then stored in coolers with dry ice during transport back to the laboratory where they were immediately stored at −80°C.
Total RNA from each leaf collected as described above was extracted using the RNeasy Plant Mini kit (Qiagen Inc., Germany). RNA was then pooled with equal amounts of RNA concentration for a total of 5 unique samples per treatment combination (management and damage type) within a variety (see Supplementary Figure 1). RNase-free DNase I digestion was performed post extraction using the TURBO DNA-free Kit (Invitrogen Thermo Scientific, Lithuania). Initial integrity and quality of the total RNA was checked using a NanoDrop 1000 spectrophotometer and formaldehyde-agarose gel electrophoresis. Final RNA samples were assessed using a Fragment Analyzer (Advanced Analytical Technologies, Ankeny, IA) with the High Sensitivity RNA Analysis Kit and only RQNs ranging from 6 to 10 were used for RNA library preparation following standard protocols (TruSeq v2 Stranded mRNA Library Prep Kit; Illumina, San Diego, CA). The concentrations of RNA libraries were measured by StepOnePlus Real-Time PCR System (ThermoFisher Scientific, San Jose, CA) with the KAPA Library Quantification Kit (Kapabiosystems, Wilmington, MA). The libraries were diluted to 2 nM with RSB (10 mM Tris-HCl, pH 8.5) and denatured with 0.1 M NaOH. Eighteen pM libraries were clustered in a high-output flow cell using HiSeq Cluster Kit v4 on a cBot (Illumina) and sequenced using paired ends (100 bp) on a HiSeq 2500 using HiSeq SBS kit v4 (Illumina). The raw bcl files were converted to fastq files using software program bcl2fastq2.17.1.14.
The RNA-seq reads were adaptor trimmed and then aligned to the S. tuberosum genome using the aligner Tophat and Bowtie (Langmead et al., 2009; Trapnell et al., 2009; Bolger et al., 2014; Version12x; Phytozome Version 9, Joint Genome Institute) under default parameters, which allow up to two mismatches and report up to 40 alignments for reads mapping at multiple positions. Counts of gene abundance were made using Python Package HTSeq (Anders et al., 2015). Raw sequence data have been deposited in the Sequence Read Archive (http://www.ncbi.nlm.nih.gov/sra) under the BioProject numbers PRJNA417905 and PRJNA417851.
Our next objective was to estimate densities of predators and herbivores in each of the potato fields where foliage samples were collected. All predators were collected within the same area leaf tissue was collected, using a D-vac suction-sampling device and previously described methods (e.g., Koss et al., 2005). Briefly, we haphazardly identified 10 potato plants per field for sampling, walking in a zigzag pattern from the field edge toward the center of the field. We held the collecting cone over each plant, gently shaking the foliage for 20 s, and changed collecting bags between each group of 10 plants (Koss et al., 2005). We also visually examined an additional set of 10 plants, haphazardly chosen where we counted any arthropods present for 30 s. Arthropods in D-vac bags were immediately placed in a cooler on dry ice and transported back to the laboratory for further identification to the family level using the dichotomous keys in (Triplehorn et al., 2005).
At each farm site, five soil subsamples were collected randomly from each corner and middle within the same 400-m2 area in the field as noted above. Each of the five samples consisted of 20-cm deep soil that was taken with a soil corer. The soil replicates within each field were mixed and packed in sterile whirlpak bags, and each bag was sealed and transported to the laboratory at 4°C. A 100-g fraction of the mixed sample was stored at −20°C for molecular analysis, while ~200 g of the soil was immediately sent to SoilTest labs (Moses Lake, WA). There, soil samples were passed through a 2-mm sieve and analyzed for the following properties according to recommended soil testing methods by Gavlak et al. (2003): particle-size (% sand, silt, and clay); nitrate-nitrogen (N) and ammonium-N; extractable phosphorus, potassium, calcium, magnesium, sodium, sulfur, boron, zinc, manganese, copper, and iron; cation exchange capacity; total bases; % base saturation; exchangeable sodium percentage (ESP); pH; electrical conductivity; N mineralization; and percent organic matter (see Supplementary Table 2). Additionally, microbial biomass was estimated using a Solvita test (Haney et al., 2008) (Supplementary Table 2).
DNA was extracted from a 0.25 g sample using the PowerSoil DNA isolation Kit (MOBIO Laboratories, Inc., Carlsbad, CA, USA). Soil DNA extractions were sent to Oregon State University's Center for Genome Research and Biocomputing for 16S sequencing. This facility follows the Illumina 16S metagenomic sequencing library preparation protocol for MiSeq sequencing. The gene-specific sequences used in this protocol target the V3 and V4 region of bacterial 16S rRNA gene. The bacterial primer pairs and sequence analyses were done using previously described methods in Krey et al. (2019). Microbial sequencing data are available on GenBank under project number PRJNA553015.
Primary metabolite derivatization and gas chromatography were carried out using a slight modification of an established procedure (Lee and Fiehn, 2008). A defined amount of powdered freeze-dried potato leaves (ca. 20 mg) was suspended in 500 μL of extraction solvent containing methanol, 2-propanol, and water at a 5:2:2 ratio. After adding 1.5 μg of the internal standard ribitol, the material was extracted by shaking at room temperature for 10 min (Vortex) and sonication at room temperature for 10 min (Branson 5510 sonication bath). The extracts were then centrifuged for 10 min at 21,000 g, and the supernatants transferred into new vial. The extracts were dried in vacuum. Dry residues were suspended in 500 μL of 50% aqueous acetonitrile and re-extracted as above by sequential vortexing and sonication. The debris was again removed by centrifugation, and the supernatants were dried in vacuum. The dry residues were suspended in 10 μL O-methoxylamine hydrochloride (30 mg mL-1 in pyridine, both from Sigma) and incubated for 90 min at 30°C and 1000 rpm (Eppendorf Thermomixer).
Subsequently, samples were derivatized with 90 μL of MSTFA with 1% TMCS (Thermo-Pierce cat.-no.TS-48915) for 30 min at 37°C and 1000 rpm (Eppendorf Thermomixer). Gas chromatography-mass spectroscopy analysis was performed using a Pegasus 4D time-of-flight mass spectrometer (LECO) equipped with a Gerstel MPS2 autosampler and an Agilent 7890A oven. The derivatization products were separated on a 30 m, 0.25 mm i.d., 0.25 μm df Rxi-5Sil® column (Restek) with an IntegraGuard® pre-column using ultrapure He at a constant flow of 1 mL min-1 as carrier gas. The linear thermal gradient started with a 1-min hold at 50°C, followed by a ramp to 330°C at 20°C min-1. The final temperature was held for 5 min prior to returning to initial conditions. Mass spectra were collected at 17 spectra s-1. The injection port was held at 250°C, and 1 μL of the sample were injected at an appropriate split ratio.
Percent foliar nitrogen and carbon were quantified in leaf tissue using an Elemental Analyzer (ECS4010, Costech Analytical, Valencia, California, USA), following methods described by Krey et al. (2019).
We evaluated differences in plant gene expression, soil microbes, and insects across production systems (organic and conventional) and potato varieties (Norkotah and Alturas). Statistics were conducted in R (version 3.2.1; R Core Team, 2015) unless otherwise noted.
The EdgeR (Robinson et al., 2010) and limma (Ritchie et al., 2015) packages were used in R studio (v.0.99.903) to calculate differential gene expression on genes with > 2 read counts. In our study, we examined which genes are expressed at different levels when comparing leaves collected from plants grown under two management systems (organic vs. conventional), and when leaves had physical damage or appeared to be undamaged. To make these comparisons, we established a design matrix with management and insect as “treatment” (where field is the unit of replication) and made contrasts for pairwise comparisons separately for each cultivar (see Supplementary Table 3). We added field as a random effect using the duplicate Correlation function because leaf samples came from within a field, thus were not independent relative to the unit of replication. Counts with associated weighting were transformed to logCPM values using the voom function prior to tests of differential expression following standard practices (Chen et al., 2014; Smyth et al., 2020). Significance was defined using an FDR adjusted p-value cutoff at 10% (adj P < 0.1).
Using Blast2GO (v.4.1.3, Conesa et al., 2005) with Blast database set at “nr” and E-value at 1.0E-3, and the MapMan visualization tool kit (Thimm et al., 2004), we were able to assign gene ontology classes to our differentially expressed genes with BLAST matches to known sequences. Blast2GO also assigns biological functions based on BLAST sequence homologies and GO annotations (with respect to biological processes, molecular functions, and cellular components); each contrast that had differentially expressed genes (DEG) was assigned a biological function and categorized accordingly (Conesa et al., 2005). GO enrichment analysis was used to compute enrichment of GO terms found in MapMan in each of the contrast groups. The p-values were adjusted using the Benjamini and Hochberg FDR with a cutoff of < 0.1. Enrichment analysis was done using Fisher's Exact Test found in the R “stats” package.
Model assumptions were checked using histograms, Shapiro-Wilk tests, and residual plots, as necessary. To examine differences in insect densities across production regimes, we performed six generalized linear models assuming Poisson distributions: one for phloem feeders, thrips, and natural enemies within each of the two potato varieties examined. We considered the two potato varieties separately because of considerable differences in environments in which they are grown. To examine differences in the physical and nutritional profiles of soil between cropping systems and potato varieties, we used non-metric multi-dimensional scaling (NMDS) to collapse multi-dimensional soil data (Supplementary Table 2) onto two axes. We used the metaMDS function in the vegan package of R to create a dissimilarity matrix using Bray-Curtis distances, and used ordihull and ordiellipse functions to visualize distances between sites (Oksanen et al., 2007). Stress values were <0.2 (0.14), indicating that the ordination was an adequate representation of the data in reduced dimensions. To test for differences in soil management between production regimes and potato varieties, we performed analysis of similarity (anosim function in vegan), comparing ranked distances for each group to null distributions using 2000 permutations. Additionally, soil nutrient concentrations were compared using linear model, lm function in lmer4, with production regime (conventional and organic), variety (Norkotah and Alturas), and their interaction term as predictor variables.
Soil bacteria sequences were processed using previously described methods in Krey et al. (2019). Phyloseq v1.18.1 (McMurdie and Holmes, 2013) was used to visualize data. The differential abundance of bacteria between organic and conventional soils was assessed using Deseq2 (Love et al., 2014) within the phyloseq package (test = “Wald,” fitType = “local”) with FDR value < 0.1 and log2FoldChange> 4. Compositional diversity was assessed by applying the Shannon diversity index considering the number and abundance of species using the estimate_richness function in the “phyloseq” package. From each field, we calculated bacterial community evenness at the family level using Shannon (H)/ln(richness). Diversity and evenness indices were compared with ANOVA with production regime (conventional and organic) and variety (Norkotah and Alturas) as the predictor variables. Abundances of the top 10 bacteria families were ln(x+1) transformed and compared using ANOVA, with production regime (conventional and organic), variety (Norkotah and Alturas), and their interaction term as predictor variables.
Peak identification of foliar metabolites was conducted using Fiehn primary metabolite library (Kind et al., 2009). For identification, identity score cutoff of 700 was used. Peak alignment and spectrum comparisons were carried out using the Statistical Compare feature of the ChromaTOF® software (LECO). The internal standard ribitol and the initial tissue weight were used for normalization. Statistical analyses with selected metabolites were carried out using Metaboanalyst 3.0 (Xia and Wishart, 2016). Normalized data were uploaded and autoscaled for further uni- and multivariate analyses. Specific metabolites (see Supplementary Table 4) related to plant defense and health were selected and their concentrations were compared using a permutation multivariate analysis of variance using distance matrices (permutations = 999) using the adonis functions in the vegan package of R, with production regime (conventional and organic) and variety (Norkotah and Alturas) as predictor variables.
Percentages of carbon, nitrogen, and C:N ratio were measured in foliar tissue from 120 samples of herbivore-damaged and undamaged leaves from each sample site. Herbivore damage did not alter foliar nutrition (see Supplementary Table 5) and this enabled us to combine replicates (damaged and undamaged leaves) to increase our power to detect soil-mediated differences in plant quality across sites. We compared C, N, and C:N ratios with 3 separate linear mixed-effects models using the lme function in the nlme package of R, with production regime (conventional and organic), variety (Norkotah and Alturas), and their interaction term as predictor variables and field as a random effect.
Norkotah and Alturas varieties differed drastically in the number of differentially expressed genes in all treatments (DEG) (Supplementary Table 3). Examination of the transcriptome revealed 6,985 differentially expressed (DE) genes in Norkotah in damaged tissue (combining organic and conventional) relative to undamaged leaf tissue, while Alturas had 1,585 DE genes. For Norkotah, insect herbivory led to higher enrichment of defense-related genes in transcripts of plants from organic systems compared to conventional (stress-related genes, P = 0.03, secondary metabolite-related gene, P < 0.001; Figure 1, stress perception; Table 1; Supplementary Table 6), and had more upregulated genes related to plant stress (P = 0.01; Figure 1, stress perception; Table 1; Supplementary Table 6). In Norkotah, we saw a strong damage-induced response in the ethylene (ET) signaling pathway in organic systems (Figure 1, stress perception; Table 1), while Alturas had fewer differentially expressed genes overall, and most were downregulated across all treatments (Table 1; Supplementary Table 3). In Norkotah organic insect damaged leaves, genes associated with light harvesting and photosynthetic carbon assimilation strongly decreased (Figure 1, growth and development; Table 1; Supplementary Table 6), photosynthetic assimilation (PS) and nutrient transport related genes were more enriched in transcripts (PS, P < 0.001; Table 1; Supplementary Table 6, transport, P = 0.01; Table 1; Supplementary Table 6) and more PS genes were downregulated (P < 0.001; Table 1; Supplementary Table 6).
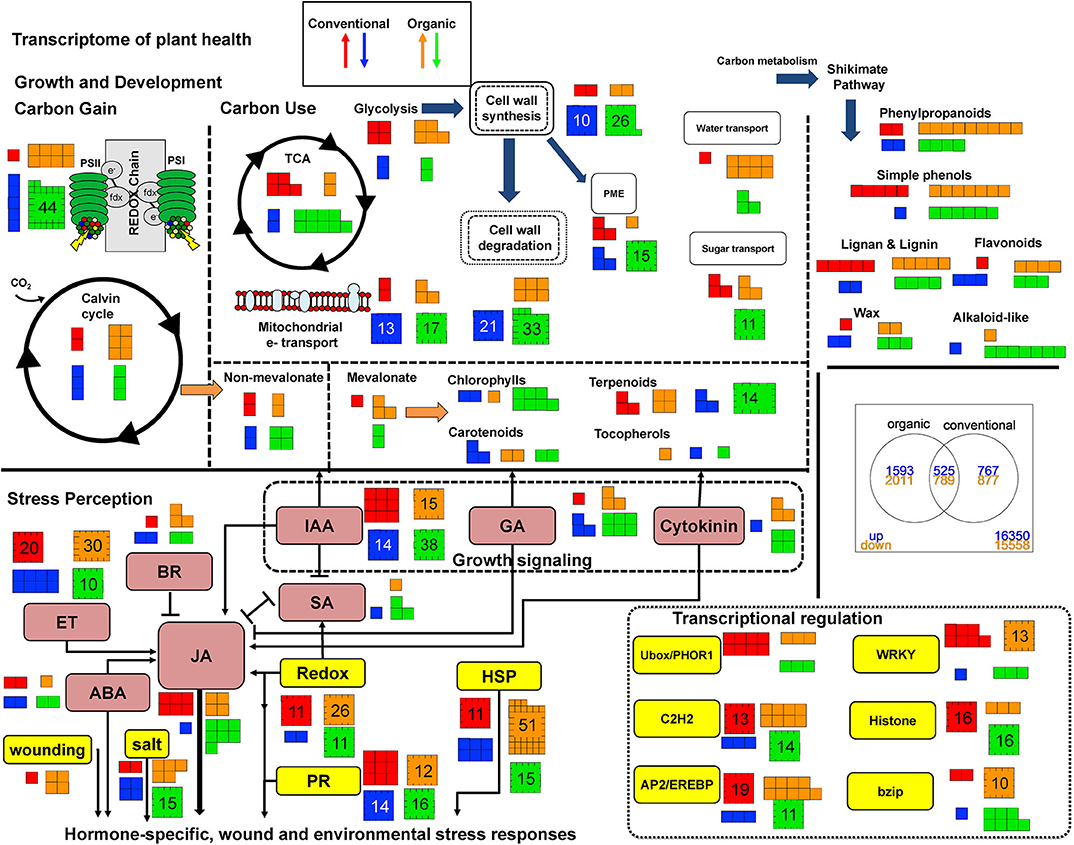
Figure 1. This graphical representation of the transcriptome for processes related to plant health and defense compares Norkotah potato plants grown under organic (orange and green arrows) and conventional (red and blue arrows) farming methods between insect undamaged and damaged leaves. Genes up or down are in reference to how plants respond to damage under different management regimes. Numbers indicate how many genes are up or down regulated. Text containing boxes are colored coded for metabolites specifically involved in hormone signaling/perception: pink; and non-hormonal processes mediating stress response/perception: yellow. Arrows link flow from primary to secondary metabolism (Top) and synergistic or antagonistic signaling during stress perception (Bottom).
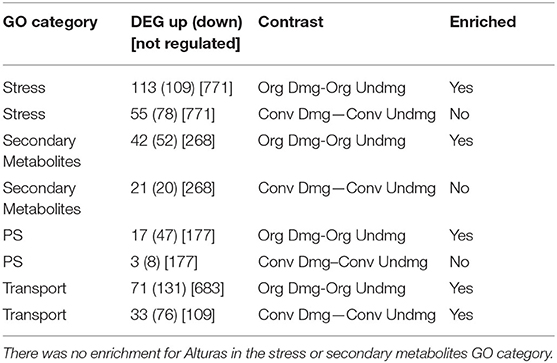
Table 1. Number of differentially expressed genes (DEG) in Norkotah with threshold of adjusted P-value < 0.1, categorized by Gene Ontology (GO) and compared via contrasts (see Supplemental Table 3).
Expression of genes associated with stress was enriched within the transcriptome for both varieties of damaged leaves (Norkotah, P = 0.005; Alturas, P = 0.001), while more stress-related genes were upregulated in Norkotah (P < 0.001; Table 1) and downregulated (P < 0.001) in Alturas. In Norkotah damaged leaves, there was an increase in the number of upregulated genes in salicylic acid (SA) and ET signaling pathways relative to undamaged leaves (Figure 1), while Alturas damaged leaves saw more downregulated genes (Supplementary Table 3, contrast 2 and 3). When we pooled samples from organic and conventional systems to improve our power to examine herbivore effects, damaged leaves led to more downregulated genes related to photosynthetic assimilation (PS) in Norkotah (P < 0.001; Table 1), while Alturas did not (P = 0.9445).
Organic Norkotah potatoes had higher numbers of phloem feeders compared to conventional plants (Coefficient = 0.606, SE = 0.191, Z = 3.16, P = 0.0016, Figure 2A), similar numbers of thrips (Coefficient = 0.051, SE = 0.073, Z = 0.694, P = 0.488, Figure 2B), and more natural enemies (Coefficient = 1.17, SE = 0.155, Z = 7.511, P = < 0.0001, Figure 2C). Organic Alturas potatoes had fewer thrips than conventional plants (Coefficient = 0.49, SE = 0.135, Z = −3.62, P = 0.0002, Figure 2D), similar numbers of phloem feeders (Coefficient = −0.265, SE = 0.222, Z = −1.193, P = 0.233, Figure 2E), and similar numbers of natural enemies (Coefficient = 0.361, SE = 0.228, Z = 1.583, P = 0.114, Figure 2F).
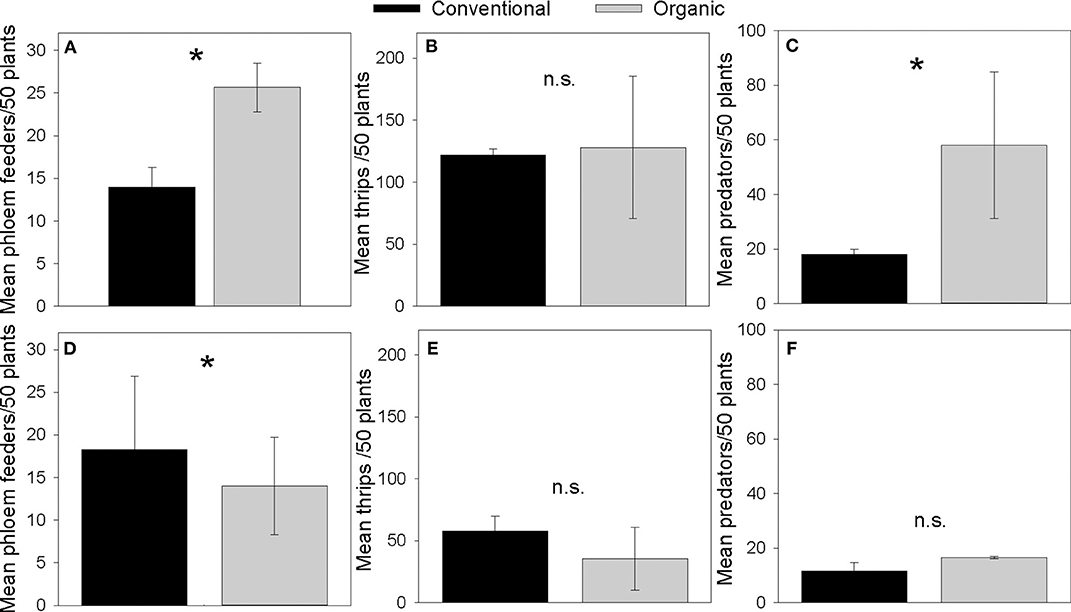
Figure 2. Mean counts of phloem feeding herbivores (A,D), thrips (B,E), and natural enemies (C,F), collected in vacuum samples from Norkotah (A–C) and Alturas (D–F) potato fields in 2015. *indicates P < 0.01.
Physical and chemical soil characteristics at organically managed sites where Norkotah potatoes were grown were significantly dissimilar from conventional sites and sites where Alturas potatoes grew (ANOSIM R = 0.2946, P = 0.049, Figure 3). Nearly 4x as many nitrates were available in conventional Norkotah fields compared to organic, whereas in Alturas nitrate levels were 4x lower in conventional fields compared to organic (management*variety interaction: F1,7 = 15.338, P = 0.005; Figure 4A). Phosphorus concentrations approximately doubled in organic and conventional Alturas fields and differed marginally between varieties [F(1, 7) = 4.162, P = 0.06; Figure 4B]. Microbial biomass was higher in Norkotah organic than Alturas [management*variety interaction: F(1, 7) = 6.458, P = 0.038; Figure 4E].
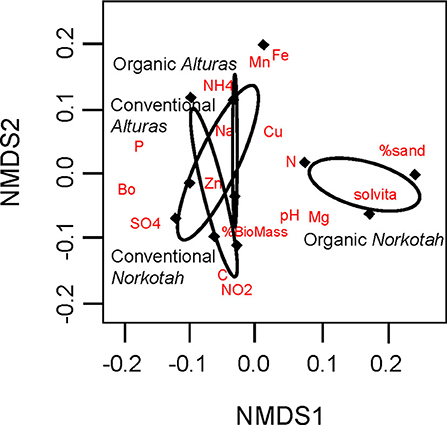
Figure 3. Physical and chemical profiles of soil displayed on two non-metric multidimensional scaled axes. Ellipses indicate clustering of our four treatment groups which consisted of two cropping system treatments (conventional and organic) and two potato varieties (Norkotah and Alturas). Red labels indicate the locations of soil characteristics that distinguish clusters.
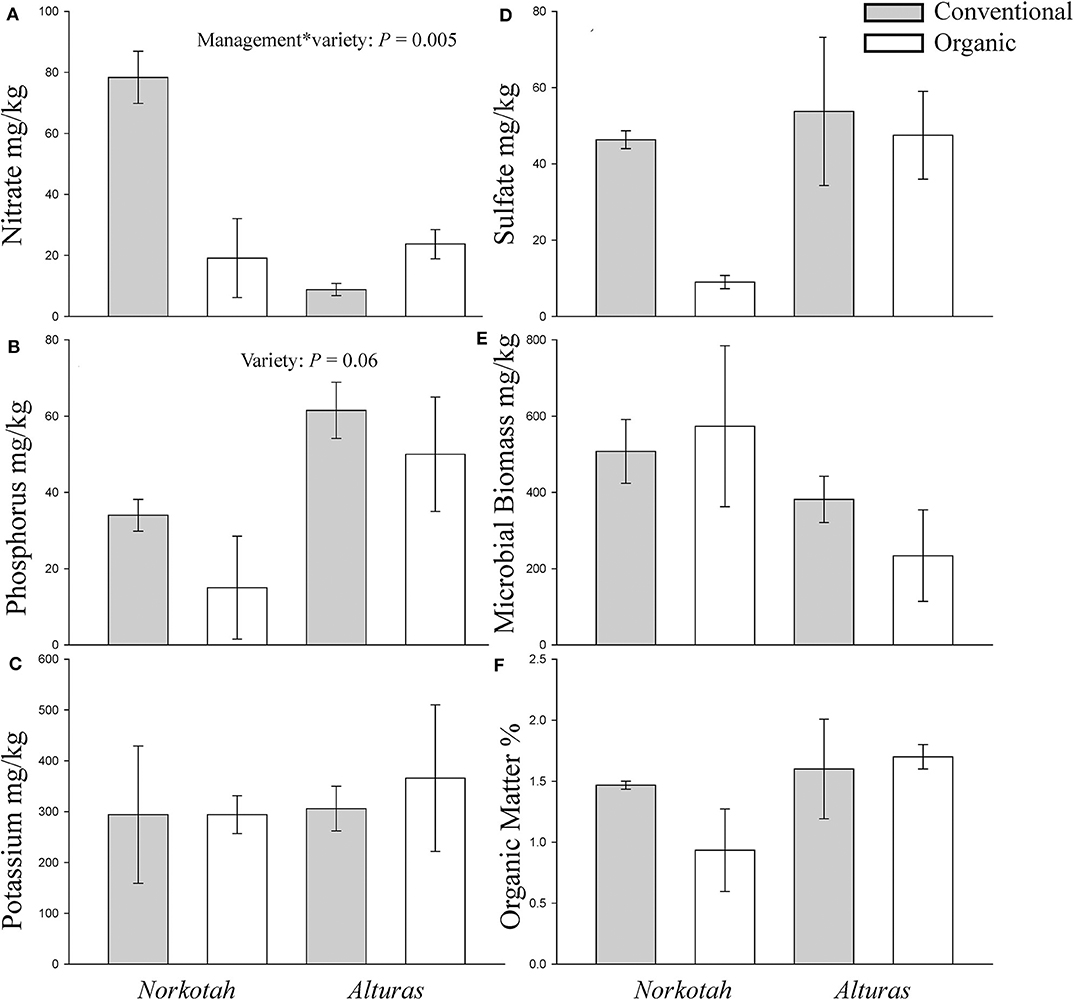
Figure 4. Concentrations of nutrients in soil collected from organic and conventional potato fields at mid-season. (A) Nitrate, (B) phosphorus, (C) potassium, (D) sulfate, (E) microbial biomass, and (F) organic matter. Gray bars indicate conventional soil and white bars organic.
There were no differences in soil bacterial diversity between management systems [F(1, 7) = 0.025, P = 0.879; Supplementary Table 7] or varieties [F(1, 7) = −0.05, P = 0.830; Supplementary Table 7], and community evenness similarly did not vary according to management system [F(1, 7) = 0.009, P = 0.926; Supplementary Table 7] or variety [F(1, 7) = 0.005, P = 0.944; Supplementary Table 7]. Ten bacterial taxa were differentially abundant within organic management when compared to conventional management (Padj<0.1; Supplementary Table 8). Bacteria from the family Bacilliaceae, Chitinophagaceae, and Planctomycetaceae were more abundant in organic Norkotah fields compared to conventional, but less abundant in organic Alturas fields compared to conventional (Bacilliaceae; management*variety interaction: F = 5.768, P = 0.0173, Chitinophagaceae; management*variety interaction: F = 3.921, P = 0.0478, Planctomycetaceae; management*variety interaction: F = 25.550, P < 0.001), while the other taxa were not significantly different by variety.
Percent carbon in leaf tissue did not differ among treatments (t = −0.321, P = 0.7491; Supplementary Table 5) or varieties (t = 0.436, P = 0.6637; Supplementary Table 5). Percent nitrogen was higher in Norkotah vs. Alturas (t = 2.667, P = 0.009), and the carbon:nitrogen ratio in conventional systems decreased marginally in the leaf tissue but only in Norkotah fields, whereas conventional management increased the carbon:nitrogen ratio in Alturas fields (t = 0.1.963, P = 0.052). The suite of primary metabolites (Supplementary Table 4) in leaf tissue did not differ based on variety, management, or herbivore damage (R2 = 0.126, df = 1, 11, P = 0.22).
Our central goal was to compare plant gene expression patterns in conventional vs. organic farming systems. To do this, we assessed herbivore-defense gene expression in plant tissue collected from open fields on working farms and quantified several potentially associated environmental variables (e.g., soil nutrients, microbial communities, herbivore pressure, etc.). The most punctuated effect of organic farming we observed was a strong increase in activity of several herbivore defense-associated genes in insect-damaged leaf tissue, whereas these genes were not differentially expressed in plants experiencing herbivory in conventional fields. Importantly, these responses were variety-specific: we saw this pattern of increased defense gene expression in Norkotah, which is typically marketed as a fresh product and is more susceptible to stress, but not in Alturas, which is primarily processed for dehydrated products and bred to be more tolerant to stresses (Novy et al., 2003; Stark et al., 2020).
Boorse (1977) defines unhealthy or disease states as occurring when normal function is impaired or limited. Thus, health is the absence of impairments or limitations. Given that normal function depends on a myriad of parameters, our definition of plant health for this paper uses a baseline of gene expression in conventionally managed potatoes. Any change in transcript abundance that represents an enhancement or limitation in processes related to metabolic homeostasis or perception of environmental stressors (abiotic or biotic) is considered a change in plant health. Given the number of interactions and pairwise comparisons that could be made, a logical baseline is an assessment of variation between undamaged conventionally grown plants and undamaged organically grown plants, hereafter referred to as conventional and organic plants, respectively. At this baseline, organic plants have higher abundance of growth transcripts and pathogen stress-related transcripts, and lower abundance of abiotic stress-related transcripts compared to conventional plants (Figure 1). More specifically, organic plants show greater expression in photosynthesis genes and cell wall metabolism (P < 0.01), and trends for higher development-related expression and reduced stress-related expression (P < 0.1). In contrast, Pacifico et al. (2017) found more stress related transcripts, with an emphasis on phenylpropanoid metabolism, in 1 year and enrichment in secondary metabolism (flavonoids) in a second year on potato crops. Energy metabolism, cell wall metabolism, signaling, and RNA regulation were consistently differentially expressed over both years compared to conventional plants, although the magnitudes and directions of the genes were variable (Pacifico et al., 2017). Furthermore, van Dijk et al. (2012) found organic fertilizer treatments in potato crops increased lipoxygenase-mediated defense gene expression but organic crop protection suppressed this type of gene expression. This directional expression would interact under field conditions where both fertilizer and crop protection strategies are applied simultaneously, and may explain why only two lox-related defense transcripts were found to be upregulated in organic plants compared to conventional plants in our data (Figure 1; Supplementary Table 6).
Some specific processes that showed enhanced transcript abundance included those involved in light harvesting, a reduced starch degradation gene (B-amlyase), and increasing mobilization of sugars for cell wall formation (Supplementary Table 6). Of interest is the upregulation of several genes related to pectin methyl esterases (PMEs), key enzymes that de-esterify pectins in support of reinforcing cell walls. PMEs also increase during pathogen recognition and depend on JA signaling mitigated by ethylene response factors (ERF) when necrotrophs attack, and both ERF and MYC signaling when biotrophs attack (Bethke et al., 2014). Constitutive JA activity in wildtype potato increases PME to reduce pectin methyl esterification and alter the pectin matrix in favor of increased resistance to some pathogens (Taurino et al., 2014).
GO analysis indicated stress responses trended toward enrichment (P < 0.1; Table 1), but several individual transcripts accumulated more in organic plants. Specifically, defense-related secondary metabolism (terpenoid, alkaloid) and hormone signaling (JA synthesis, ET synthesis/signaling) transcripts were greater, and abiotic stress response to temperature and wounding were reduced compared to conventional plants. Lox1 and HPL show greater expression in organic plants, indicating fatty acid biosynthesis (C-6 aldehydes and w-keto FA) or upstream defense metabolites may be elevated. This would suggest an active defense system if attacked or a higher constitutive defense level because of greater plant health, priming by soil microbes, or both. Biotic stress genes related to pathogen response showed elevated expression, as did miscellaneous p450s and UGTs, genes involved in altering the solubility of metabolites for secondary metabolism and detoxification. It should also be emphasized that, while up regulation of a suite of genes involved in defense response indicates a strong response, it does not necessarily indicate an effective response. We note that further studies are required to demonstrate that any differences in gene expression between farming systems are meaningfully increasing a plant's ability to suppress pest damage.
Overall, undamaged organic plants showed greater health as defined by transcript accumulation in favor of growth, development, and defense signaling. Given growth and defense tradeoffs are largely mediated by hormone interactions including antagonisms, a positive growth response alongside defense signaling likely means plants show a higher constitutive or primed state of health, rather than a defensive state caused by pest induction. Priming is most beneficial when attack is looming, which makes a mid-season organic potato field an optimal candidate to invest against late season attack where tuber filling is greatest. Thus, a future hypothesis to test is on the temporal nature of priming: does organic management always induce higher constitutive growth and defense processes, or is priming phenologically or temporally specific?
When damaged and undamaged plants (pooling across production systems), 6985 genes were differentially expressed. Of these, 1402 only appear significant when conventional and organic samples are combined, indicating an interaction between management strategy and damage. Indeed, the MDS plot illustrates the same interaction (Supplementary Figure 1). See Supplemental Table 4 for individual gene data (baseline O vs. C undam). Under damaged conditions, organic plants show more differentially expressed genes than conventional plants (3604 vs. 1644) but share 1,314 genes expressed in the same direction under damage (Supplementary Figures 2, 3). 30 genes were found to be expressed in opposite directions in management and the remaining genes were either up or downregulated in one production system but not significantly expressed in the other. The number of genes differentially expressed between conventional and organic production across all samples is low (2) because undamaged plants have gene expression patterns in opposite directions (Supplementary Figure 2). More genes are upregulated than downregulated in undamaged organic plants compared to conventional undamaged plants (Supplementary Figure 2). However, these differences in gene expression disappear when plants are attacked by chewing herbivores. Regardless of management strategy, growth-related transcripts were fewer and defense transcripts more abundant in Norkotah damaged plants. For example, genes related to photosynthesis, carbon mobilization, cell cycle, growth, and development showed more down than upregulated (Figure 1). Genes related to amino acid metabolism, secondary metabolism, redox regulation, and xenobiotic degradation had more up than downregulated (Figure 1). These patterns would suggest defense metabolism is active to defend against biotic stress caused during defoliation or other pest attack on damaged leaves while growth is suppressed.
When compared separately, 1960 more transcripts are differentially expressed in organic than in conventional plants experiencing biotic attack. Gene ontologies that are overrepresented include PS, lipid metabolism, cell wall metabolism, secondary metabolism, stress, and development, whereas conventional plants showed overrepresentation in photosynthesis, protein metabolism, and signaling genes (P < 0.05). The difference in response is intriguing because the gene expression enrichment profiles suggest organic plants are responding with a greater growth-defense tradeoff, which could be ascribed as a typical or healthy response. Conventional plants show enrichment in fewer pathways that include the expected suppression in photosynthesis under damage but show differential expression in signaling and protein turnover. Thus, the growth-defense tradeoff is active upon attack, but secondary metabolism and hormone mediated defense pathways are not enriched even if individual genes support this tradeoff.
Differences in defense gene expression between the two potato varieties in organic and conventional farming suggest that Alturas is generally less responsive to herbivore damage than Norkotah. This could be because Alturas is grown mostly for the dehydration industry (where tuber shape is not managed), and was bred for tolerance of low nitrogen rates, resistance to Verticillium dahliae (the cause of early dying), and other stresses (Novy et al., 2003). These traits may trade-off with herbivore resistance (e.g., Fineblum and Rausher, 1995; Rudgers et al., 2004). Also, due to their tolerance of suboptimal soil conditions, Alturas potatoes can be successfully grown in less desirable circumstances (e.g., fields with low soil fertility, marginal lands of rocks, tight potato rotations without green manures) than some other cultivars (Novy et al., 2003). For this reason, Alturas plants may be poorer quality hosts for herbivores, and therefore suffer less herbivory (e.g., Awmack and Leather, 2002). On the other hand, Norkotah plants are generally produced for fresh marketing in the Pacific Northwest and are relatively susceptible to herbivory, but production practices will vary substantially depending on whether the harvested crop will be stored vs. marketed direct from the field (Zvomuya et al., 2002; Stark et al., 2020). While Norkotah is resistant to common scab and tolerant of some viruses, a study showed green peach aphids had slower growth rates on Norkotah compared to Alturas plants (Draper et al., 2002; Davis et al., 2007; Stark et al., 2020). This could also partly explain why Norkotah had a higher defense response to herbivore attack than Alturas (Davis et al., 2007). Typically, better fields and higher inputs will be used to maintain the Norkotah plants as long as possible later in the season and may receive fewer inputs than other varieties (Zvomuya et al., 2002; Stark et al., 2020). Indeed, percent N in leaf tissue was significantly higher for Norkotah than Alturas, making it a higher quality food source for herbivores (Mattson, 1980; Awmack and Leather, 2002). Thus, chemical defense induction may be more important for Norkotah due to plant traits and relatively greater herbivore pressure (e.g., Wink, 1988).
We measured several environmental factors to identify those that might explain stronger upregulation of plant defense genes in organic systems compared to conventional. First, we quantified soil nutrients and sequenced soil microbial communities at each site where we sampled plants. In conventional fields, we saw a greater concentration of nitrates where Norkotah was grown compared to Alturas (Figure 4). Nitrate fertilization can improve host plant quality for herbivores and has been associated with increases in phloem-feeding insects (Strauss, 1987; Lou and Baldwin, 2004; Staley et al., 2010). Nitrogen augmentation can cause plants to reduce investments in chemical defenses (Glynn et al., 2003; Donaldson et al., 2006), particularly the quick-release N subsidies applied on conventional fields (Staley et al., 2010). Therefore, excess N availability in conventional fields where Norkotah grew (Figure 4) may have reduced defense gene expression as seen in the other treatments.
The use of organic soil amendments could also be associated with desirable soil properties, such as increased densities of beneficial soil microorganisms (Drinkwater et al., 1995; Doran, 2002). We saw an increase in soil microbial biomass in Norkotah organic compared to Alturas (Figure 4E). Several studies have shown that beneficial below-ground microorganisms may enhance plant stress tolerance and nutrient uptake (Bloemberg and Lugtenberg, 2001; Lugtenberg et al., 2002; Bais et al., 2004; Morrissey et al., 2004; Goh et al., 2013; Blundell et al., 2020). Many studies find increased microbial biomass (Gunapala and Scow, 1998; Swezey et al., 1998; Altieri, 1999; Mäder et al., 2002; Reganold et al., 2010) and diversity (Ge et al., 2008; Reilly et al., 2013; Lupatini et al., 2017) in response to organic soil management, but this is not consistent across every system (Liu et al., 2007) and can be site specific (Blundell et al., 2020). Alturas potatoes, which persist under nutrient poor conditions, may demonstrate another exception. It is important to also acknowledge that our fields were not paired side-by-side with matched soil-forming factors that have been shown to influence organic vs. conventional differences (Reganold et al., 1993, 2010; Reganold, 2013). When we compared the broad communities of bacteria in the soil, overall, we found few differences between organic and conventional fields (Supplementary Table 7).
Norkotah overall experienced higher herbivore pressure than Alturas and had a correspondingly stronger response in defense gene activity (Figures 1, 2). This pattern of herbivore pressure is one potential reason why induction of defense genes was variety-specific. Prior herbivore damage can prime induced defenses for stronger expression later in the season (Conrath et al., 2006; Frost et al., 2008) and may also be responsible for varying patterns of induction in our system. Insecticide use is another potential mechanism explaining differences in defense activity that we observed. In conventional potato crops, neonicotinoids are commonly applied, but are barred from use in organic fields (Koss et al., 2005). Previous work shows that neonicotinoids can attenuate defensive activity in plants (e.g., James and Price, 2002; Szczepaniec et al., 2013) and may have suppressed defense gene expression in our conventionally managed fields. Additionally, our study is rare in that it evaluates defense gene regulation in real-world farming systems characterized by numerous interacting environmental factors. It is important to acknowledge that this approach prohibited us from completely excluding herbivores on our “undamaged” plants, particularly phloem feeders, whose damage is often not visually detectable. Therefore, we cannot unequivocally distinguish changes in plant stress gene expression due to herbivory and soil characteristics. Still, bottom-up forces in organic production must be examined under field conditions if we are to understand their true importance in conferring herbivore resistance.
RNA sequencing is a powerful tool for transcriptional profiling to identify genes associated with plants, revealing networks of genes that function in a coordinated fashion to drive metabolic pathways or cellular processes. For example, farming systems have been shown to have an effect on gene expression, where organic management affected biological processes and pathways related to plant quality (Lu et al., 2005; Pacifico and Paris, 2016; Pacifico et al., 2017). However, almost all studies examining gene expression and farming systems are performed in controlled conditions and may be limited in understanding plant responses to multiple simultaneous stresses, as seen in real world conditions. When stresses occur in sequence or together, certain genes are activated that may not be induced independently (Rizhsky et al., 2002, 2004; Mittler, 2006). Future studies should take into account all the factors plants on commercial farms experience in the environment. Our study allowed us to show that specific patterns of gene expression may have useful applications in defining the differences between organically and conventionally grown plants. Linking gene expression to particular plant defensive traits is a key first step to exploiting these defenses to improve the control of herbivorous agricultural pests (Mitchell et al., 2016). For crop plants, this approach can suggest ways to use farming systems to enhance plant defenses and thus natural pest control.
Raw sequence data have been deposited in the Sequence Read Archive (http://www.ncbi.nlm.nih.gov/sra) under the BioProject numbers PRJNA417905 and PRJNA417851. Microbial sequencing data are available on GenBank under project number PRJNA553015. The data that support the findings of this study are openly available in FigShare doi: 10.6084/m9.figshare.11833311.
KK, PN, JR, AJ, and WS conceptualized the study. KK performed research. KK, PN, CB, ZF, JV, and AB analyzed and interpreted data. KK wrote the manuscript. All authors revised the manuscript.
This work was supported by Specialty Crop Research Initiative Grant no. 2015-51181-24292 from the USDA National Institute of Food and Agriculture, Northwest Potato Research Consortium, and National Institutes of Health award no. P20GM104420. This work was supported in part by the USDA National Institute of Food and Agriculture, Hatch project 227700.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
We thank the cooperating growers for their willing participation in this research. We thank Dr. W. Rodney Cooper and Dr. Michelle Heck for reviewing an earlier version of the manuscript and Samantha Beck for her assistance in field and laboratory help.
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fsufs.2020.00097/full#supplementary-material
Altieri, M. A. (1999). The ecological role of biodiversity in agroecosystems. Agric. Ecosyst. Environ. 74, 19–31. doi: 10.1016/S0167-8809(99)00028-6
Altieri, M. A., and Nicholls, C. I. (2003). Soil fertility management and insect pests: harmonizing soil and plant health in agroecosystems. Soil Tillage Res. 72, 203–211. doi: 10.1016/S0167-1987(03)00089-8
Anders, S., Pyl, P. T., and Huber, W. (2015). HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169. doi: 10.1093/bioinformatics/btu638
Awmack, C. S., and Leather, S. R. (2002). Host plant quality and fecundity in herbivorous insects. Ann. Rev. Entomol. 47, 817–844. doi: 10.1146/annurev.ento.47.091201.145300
Bais, H. P., Park, S.-W., Weir, T. L., Callaway, R. M., and Vivanco, J. M. (2004). How plants communicate using the underground information superhighway. Trends Plant Sci. 9, 26–32. doi: 10.1016/j.tplants.2003.11.008
Beecher, N. A., Johnson, R. J., Brandle, J. R., Case, R. M., and Young, L. J. (2002). Agroecology of birds in organic and nonorganic farmland. Conserv. Biol. 16, 1620–1631. doi: 10.1046/j.1523-1739.2002.01228.x
Bengtsson, J., Ahnström, J., and Weibull, A. C. (2005). The effects of organic agriculture on biodiversity and abundance: a meta-analysis. J. Appl. Ecol. 42, 261–269. doi: 10.1111/j.1365-2664.2005.01005.x
Bethke, G., Grundman, R. E., Sreekanta, S., Truman, W., Katagiri, F., and Glazebrook, J. (2014). Arabidopsis pectin methylesterases contribute to immunity against Pseudomonas syringae. Plant Physiol. 164, 1093–1107. doi: 10.1104/pp.113.227637
Bhardwaj, D., Ansari, M. W., Sahoo, R. K., and Tuteja, N. (2014). Biofertilizers function as key player in sustainable agriculture by improving soil fertility, plant tolerance and crop productivity. Microb. Cell Factor. 13:66. doi: 10.1186/1475-2859-13-66
Birkhofer, K., Bezemer, T. M., Bloem, J., Bonkowski, M., Christensen, S., Dubois, D., et al. (2008). Long-term organic farming fosters below and aboveground biota: Implications for soil quality, biological control and productivity. Soil Biol. Biochem. 40, 2297–2308. doi: 10.1016/j.soilbio.2008.05.007
Bloemberg, G. V., and Lugtenberg, B. J. J. (2001). Molecular basis of plant growth promotion and biocontrol by rhizobacteria. Curr. Opin. Plant Biol. 4, 343–350. doi: 10.1016/S1369-5266(00)00183-7
Blundell, R., Schmidt, J. E., Igwe, A., Cheung, A. L., Vannette, R. L., Gaudin, A. C., et al. (2020). Organic management promotes natural pest control through enhanced plant resistance to insects. Nat. Plants 6, 483–491. doi: 10.1038/s41477-020-0656-9
Bolger, A. M., Lohse, M., and Usadel, B. (2014). Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120. doi: 10.1093/bioinformatics/btu170
Bonanomi, G., Antignani, V., Capodilupo, M., and Scala, F. (2010). Identifying the characteristics of organic soil amendments that suppress soilborne plant diseases. Soil Biol. Biochem. 42, 136–144. doi: 10.1016/j.soilbio.2009.10.012
Bouvier, J.-C., Ricci, B., Agerberg, J., and Lavigne, C. (2011). Apple orchard pest control strategies affect bird communities in southeastern France. Environ. Toxicol. Chem. 30, 212–219. doi: 10.1002/etc.377
Broekgaarden, C., Poelman, E. H., Steenhuis, G., Voorrips, R. E., Dicke, M., and Vosman, B. (2007). Genotypic variation in genome-wide transcription profiles induced by insect feeding: Brassica oleracea – Pieris rapae interactions. BMC Genomics 8:239. doi: 10.1186/1471-2164-8-239
Chen, Y., McCarthy, D., Robinson, M., and Smyth, G. K. (2014). EdgeR: Differential Expression Analysis of Digital Gene Expression Data User's Guide. Bioconductor User's Guide. Available online at: http://www.bioconductor.org/packages/release/bioc/vignettes/edgeR/inst/doc/edgeRUsersGuide.~pdf (accessed May 16, 2020).
Cheong, Y. H., Chang, H.-S., Gupta, R., Wang, X., Zhu, T., and Luan, S. (2002). Transcriptional profiling reveals novel interactions between wounding, pathogen, abiotic stress, and hormonal responses in Arabidopsis. Plant Physiol. 129, 661–677. doi: 10.1104/pp.002857
Conesa, A., Götz, S., García-Gómez, J. M., Terol, J., Talón, M., and Robles, M. (2005). Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21, 3674–3676. doi: 10.1093/bioinformatics/bti610
Conrath, U., Beckers, G. J. M., Flors, V., García-Agustín, P., Jakab, G., Mauch, F., et al. (2006). Priming: getting ready for battle. Mol. Plant Microbe Interact. 19, 1062–1071. doi: 10.1094/MPMI-19-1062
Crowder, D. W., Northfield, T. D., Strand, M. R., and Snyder, W. E. (2010). Organic agriculture promotes evenness and natural pest control. Nature 466, 109–112. doi: 10.1038/nature09183
Davis, J. A., Radcliffe, E. B., and Ragsdale, D. W. (2007). Resistance to green peach aphid, Myzus persicae (Sulzer), and potato aphid, Macrosiphum euphorbiae (Thomas), in potato cultivars. Am. J. Potato Res. 84, 259–269. doi: 10.1007/BF02986276
Davis, J. R., Huisman, O. C., Everson, D. O., and Schneider, A. T. (2001). Verticillium wilt of potato: a model of key factors related to disease severity and tuber yield in Southeastern Idaho. Am. J. Potato Res. 78:291. doi: 10.1007/BF02875694
De Vos, M., van Oosten, V. R., van Poecke, R. M. P., van Pelt, J. A., Pozo, M. J., Mueller, M. J., et al. (2005). Signal signature and transcriptome changes of Arabidopsis during pathogen and insect attack. Mol. Plant Microbe Interact. 18, 923–937. doi: 10.1094/MPMI-18-0923
Delessert, C., Wilson, I., Straeten, D. V. D., Dennis, E., and Dolferus, R. (2004). Spatial and temporal analysis of the local response to wounding. Plant Mol. Biol. 55, 165–181. doi: 10.1007/s11103-004-0112-7
Donaldson, J. R., Kruger, E. L., and Lindroth, R. L. (2006). Competition- and resource-mediated tradeoffs between growth and defensive chemistry in trembling aspen (Populus tremuloides). New Phytol. 169, 561–570. doi: 10.1111/j.1469-8137.2005.01613.x
Doran, J. W. (2002). Soil health and global sustainability: translating science into practice. Agric. Ecosyst. Environ. 88, 119–127. doi: 10.1016/S0167-8809(01)00246-8
Draper, M. D., Pasche, J. S., and Gudmestad, N. C. (2002). Factors influencing PVY development and disease expression in three potato cultivars. Am. J. Potato Res. 79, 155–165. doi: 10.1007/BF02871931
Drinkwater, L. E., Letourneau, D. K., Workneh, F., van Bruggen, A. H. C., and Shennan, C. (1995). Fundamental differences between conventional and organic tomatoa in California. Ecol. Appl. 5, 1098–1112. doi: 10.2307/2269357
Eigenbrode, S. D., and Pimentel, D. (1988). Effects of manure and chemical fertilizers on insect pest populations on collards. Agric. Ecosyst. Environ. 20, 109–125. doi: 10.1016/0167-8809(88)90151-X
Feeny, P. (1976). “Plant apparency and chemical defense,” in Biochemical Interaction Between Plants and Insects, eds J. W. Wallace and R. L. Mansell (Boston, MA: Springer), 1–40. doi: 10.1007/978-1-4684-2646-5_1
Fineblum, W. L., and Rausher, M. D. (1995). Tradeoff between resistance and tolerance to herbivore damage in a morning glory. Nature 377:517. doi: 10.1038/377517a0
Ford, K. A., Casida, J. E., Chandran, D., Gulevich, A. G., Okrent, R. A., Durkin, K. A., et al. (2010). Neonicotinoid insecticides induce salicylate-associated plant defense responses. Proc. Natl. Acad. Sci. U.S.A. 107, 17527–17532. doi: 10.1073/pnas.1013020107
Frost, C. J., Mescher, M. C., Carlson, J. E., and Moraes, C. M. D. (2008). Plant defense priming against herbivores: getting ready for a different battle. Plant Physiol. 146, 818–824. doi: 10.1104/pp.107.113027
Gavlak, R., Horneck, D., and Miller, R. O. (2003). Soil, Plant and Water Reference Methods for the Western Region, 3rd Edn. Fort Collins, CO: WCC-103 Publication. Available online at: https://www.naptprogram.org/files/napt/western-states-method-manual-2005.pdf (accessed December 21, 2019).
Ge, Y., Zhang, J., Zhang, L., Yang, M., and He, J. (2008). Long-term fertilization regimes affect bacterial community structure and diversity of an agricultural soil in northern China. J. Soils Sediment. 8, 43–50. doi: 10.1065/jss2008.01.270
Glynn, C., Herms, D. A., Egawa, M., Hansen, R., and Mattson, W. J. (2003). Effects of nutrient availability on biomass allocation as well as constitutive and rapid induced herbivore resistance in poplar. Oikos 101, 385–397. doi: 10.1034/j.1600-0706.2003.12089.x
Goh, C.-H., Veliz Vallejos, D. F., Nicotra, A. B., and Mathesius, U. (2013). The impact of beneficial plant-associated microbes on plant phenotypic plasticity. J. Chem. Ecol. 39, 826–839. doi: 10.1007/s10886-013-0326-8
Gunapala, N., and Scow, K. M. (1998). Dynamics of soil microbial biomass and activity in conventional and organic farming systems. Soil Biol. Biochem. 30, 805–816. doi: 10.1016/S0038-0717(97)00162-4
Haney, R. L., Brinton, W. F., and Evans, E. (2008). Soil CO2 respiration: comparison of chemical titration, CO2 IRGA analysis and the solvita gel system. Renew. Agric. Food Syst. 23, 171–176. doi: 10.1017/S174217050800224X
Hartley, S. E., and Jones, C. G. (1997). “Plant chemistry and herbivory, or why the World is green,” in Plant Ecology, 2nd Edn, ed M. J. Crawley (Oxford, UK: Blackwell Publishing Ltd), 284–324.
Hole, D. G., Perkins, A. J., Wilson, J. D., Alexander, I. H., Grice, P. V., and Evans, A. D. (2005). Does organic farming benefit biodiversity? Biol. Conserv. 122, 113–130. doi: 10.1016/j.biocon.2004.07.018
Howe, G. A., and Jander, G. (2008). Plant immunity to insect herbivores. Ann. Rev. Plant Biol. 59, 41–66. doi: 10.1146/annurev.arplant.59.032607.092825
Hsu, Y.-T., Shen, T.-C., and Hwang, S.-Y. (2009). Soil fertility management and pest responses: a comparison of organic and synthetic fertilization. J. Econ. Entomol. 102, 160–169. doi: 10.1603/029.102.0123
James, D. G., and Price, T. S. (2002). Fecundity in twospotted spider mite (Acari: Tetranychidae) is increased by direct and systemic exposure to imidacloprid. J. Econ. Entomol. 95, 729–732. doi: 10.1603/0022-0493-95.4.729
Karthikeyan, G., Doraisamy, S., and Rabindran, R. (2009). Induction of systemic resistance in blackgram (Vigna mungo) against urdbean leaf crinkle virus by chemicals. Arch. Phytopathol. Plant Protect. 42, 1–15. doi: 10.1080/03235400600914314
Kind, T., Wohlgemuth, G., Lee, D. Y., Lu, Y., Palazoglu, M., Shahbaz, S., et al. (2009). FiehnLib: mass spectral and retention index libraries for metabolomics based on quadrupole and time-of-flight gas chromatography/mass spectrometry. Analy. Chem. 81, 10038–10048. doi: 10.1021/ac9019522
Klonsky, K. (2012). Comparison of production costs and resource use for organic and conventional production systems. Am. J. Agric. Econ. 94, 314–321. doi: 10.1093/ajae/aar102
Koss, A. M., Jensen, A. S., Schreiber, A., Pike, K. S., and Snyder, W. E. (2005). Comparison of predator and pest communities in Washington potato fields treated with broad-spectrum, selective, or organic insecticides. Environ. Entomol. 34, 87–95. doi: 10.1603/0046-225X-34.1.87
Krey, K. L., Blubaugh, C. K., Van Leuven, J. T., and Snyder, W. E. (2019). Organic soils control beetle survival while competitors limit aphid population growth. Environ. Entomol. 48:nvz100. doi: 10.1093/ee/nvz100
Langmead, B., Trapnell, C., Pop, M., and Salzberg, S. L. (2009). Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10:R25. doi: 10.1186/gb-2009-10-3-r25
Lee, D. Y., and Fiehn, O. (2008). High quality metabolomic data for Chlamydomonas reinhardtii. Plant Methods 4:7. doi: 10.1186/1746-4811-4-7
Letourneau, D. K., and Goldstein, B. (2001). Pest damage and arthropod community structure in organic vs. conventional tomato production in California. J. Appl. Ecol. 38, 557–570. doi: 10.1046/j.1365-2664.2001.00611.x
Liu, B., Tu, C., Hu, S., Gumpertz, M., and Ristaino, J. B. (2007). Effect of organic, sustainable, and conventional management strategies in grower fields on soil physical, chemical, and biological factors and the incidence of Southern blight. Appl. Soil Ecol. 37, 202–214. doi: 10.1016/j.apsoil.2007.06.007
Lou, Y., and Baldwin, I. T. (2004). Nitrogen supply influences herbivore-induced direct and indirect defenses and transcriptional responses in Nicotiana attenuata. Plant Physiol. 135, 496–506. doi: 10.1104/pp.104.040360
Love, M. I., Huber, W., and Anders, S. (2014). Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15:550. doi: 10.1186/s13059-014-0550-8
Lu, C., Hawkesford, M. J., Barraclough, P. B., Poulton, P. R., Wilson, I. D., Barker, G. L., et al. (2005). Markedly different gene expression in wheat grown with organic or inorganic fertilizer. Proc. Biol. Sci. 272, 1901–1908. doi: 10.1098/rspb.2005.3161
Lugtenberg, B. J. J., Chin-A-Woeng, T. F. C., and Bloemberg, G. V. (2002). Microbe–plant interactions: principles and mechanisms. Antonie Van Leeuwenhoek 81, 373–383. doi: 10.1023/A:1020596903142
Lupatini, M., Korthals, G. W., de Hollander, M., Janssens, T. K. S., and Kuramae, E. E. (2017). Soil microbiome is more heterogeneous in organic than in conventional farming system. Front. Microbiol. 7:2064. doi: 10.3389/fmicb.2016.02064
Mäder, P., Fliessbach, A., Dubois, D., Gunst, L., Fried, P., and Niggli, U. (2002). Soil fertility and biodiversity in organic farming. Science 296, 1694–1697. doi: 10.1126/science.1071148
Mattson, W. J. (1980). Herbivory in relation to plant nitrogen content. Ann. Rev. Ecol. Syst. 11, 119–161. doi: 10.1146/annurev.es.11.110180.001003
McGuiness, H. (1993). Living Soils: Sustainable Alternatives to Chemical Fertilizers or Developing Countries. New York, NY: Consumers Policy Institute.
McMurdie, P. J., and Holmes, S. (2013). phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8:e61217. doi: 10.1371/journal.pone.0061217
Mitchell, C., Brennan, R. M., Graham, J., and Karley, A. J. (2016). Plant defense against herbivorous pests: exploiting resistance and tolerance traits for sustainable crop protection. Front. Plant Sci. 7:1132. doi: 10.3389/fpls.2016.01132
Mithöfer, A., and Boland, W. (2012). Plant defense against herbivores: chemical aspects. Ann. Rev. Plant Biol. 63, 431–450. doi: 10.1146/annurev-arplant-042110-103854
Mittler, R. (2006). Abiotic stress, the field environment and stress combination. Trends Plant Sci. 11, 15–19. doi: 10.1016/j.tplants.2005.11.002
Morrissey, J. P., Dow, J. M., Mark, G. L., and O'Gara, F. (2004). Are microbes at the root of a solution to world food production? EMBO Rep. 5, 922–926. doi: 10.1038/sj.embor.7400263
Murray, K., Jepson, P., Sandlin, I., and Jensen, A. (2019). Integrated pest management Strategic Plan for Potatoes in Oregon, Washington and Idaho. NW Potato Research Consortium Oregon State University extension catalog. EM 9275. Avaliable online at: https://catalog.extension.oregonstate.edu/em9275 (accessed May 11, 2020).
Novy, R. G., Corsini, D. L., Love, S. L., Pavek, J. J., Mosley, A. R., James, S. R., et al. (2003). Alturas: a multi-purpose, Russet potato cultivar with high yield and tuber specific gravity. Am. J. Potato Res. 80:295. doi: 10.1007/BF02854312
Oksanen, J., Kindt, R., Legendre, P., O'Hara, B., Stevens, M. H. H., Oksanen, M. J., et al. (2007). The vegan package. Commun. Ecol. Package 10, 631–637. Available online at: www.sortie-nd.org/lme/R%20Packages/vegan.pdf
Pacifico, D., Onofri, C., Parisi, B., Ostano, P., and Mandolino, G. (2017). Influence of organic farming on the potato transcriptome. Sustainability 9:779. doi: 10.3390/su9050779
Pacifico, D., and Paris, R. (2016). Effect of organic potato farming on human and environmental health and benefits from new plant breeding techniques. is it only a matter of public acceptance? Sustainability 8:1054. doi: 10.3390/su8101054
Penman, D. R., and Chapman, R. B. (1988). Pesticide-induced mite outbreaks: pyrethroids and spider mites. Exp. Appl. Acarol. 4, 265–276. doi: 10.1007/BF01196190
Phelan, P. L., Mason, J. F., and Stinner, B. R. (1995). Soil-fertility management and host preference by European corn borer, Ostrinia nubilalis (Hübner), on Zea mays L.: a comparison of organic and conventional chemical farming. Agric. Ecosyst. Environ. 56, 1–8. doi: 10.1016/0167-8809(95)00640-0
Phelan, P. L., Norris, K. H., and Mason, J. F. (1996). Soil-management history and host preference by Ostrinia nubilalis: evidence for plant mineral balance mediating insect–plant interactions. Environ. Entomol. 25, 1329–1336. doi: 10.1093/ee/25.6.1329
Pimentel, D., Hepperly, P., Hanson, J., Douds, D., and Seidel, R. (2005). Environmental, energetic, and economic comparisons of organic and conventional farming systems. BioScience 55, 573–582. doi: 10.1641/0006-3568(2005)055[0573:EEAECO]2.0.CO;2
Price, P. W. (1991). The plant vigor hypothesis and herbivore attack. Oikos 62, 244–251. doi: 10.2307/3545270
Prischmann, D. A., James, D. G., and Snyder, W. E. (2005). Impact of management intensity on mites (Acari: Tetranychidae, Phytoseiidae) in Southcentral Washington wine grapes. Int. J. Acarol. 31, 277–288. doi: 10.1080/01647950508684432
R Core Team (2015). R: A Language and Environment for Statistical Computing. Vienna: R Foundation for Statistical Computing. Available online at: http://www.R-project.org/ (accessed May 6, 2017).
Ralph, S., Oddy, C., Cooper, D., Yueh, H., Jancsik, S., Kolosova, N., et al. (2006). Genomics of hybrid poplar (Populus trichocarpa× deltoides) interacting with forest tent caterpillars (Malacosoma disstria): normalized and full-length cDNA libraries, expressed sequence tags, and a cDNA microarray for the study of insect-induced defences in poplar. Mol. Ecol. 15, 1275–1297. doi: 10.1111/j.1365-294X.2006.02824.x
Reganold, J. P. (2013). Comparing organic and conventional farming systems: metrics and research approaches. Crop Manag. 12, 1–6. doi: 10.1094/CM-2013-0429-01-RS
Reganold, J. P., Andrews, P. K., Reeve, J. R., Carpenter-Boggs, L., Schadt, C. W., Alldredge, J. R., et al. (2010). Fruit and soil quality of organic and conventional strawberry agroecosystems. PLoS ONE 5:e12346. doi: 10.1371/annotation/1eefd0a4-77af-4f48-98c3-2c5696ca9e7a
Reganold, J. P., Elliott, L. F., and Unger, Y. L. (1987). Long-term effects of organic and conventional farming on soil erosion. Nature 330, 370–372. doi: 10.1038/330370a0
Reganold, J. P., Palmer, A. S., Lockhart, J. C., and Macgregor, A. N. (1993). Soil quality and financial performance of biodynamic and conventional farms in New Zealand. Science 260, 344–349. doi: 10.1126/science.260.5106.344
Reilly, K., Cullen, E., Lola-Luz, T., Stone, D., Valverde, J., Gaffney, M., et al. (2013). Effect of organic, conventional and mixed cultivation practices on soil microbial community structure and nematode abundance in a cultivated onion crop. J. Sci. Food Agric. 93, 3700–3709. doi: 10.1002/jsfa.6206
Reymond, P., Bodenhausen, N., Poecke, R. M. P. V., Krishnamurthy, V., Dicke, M., and Farmer, E. E. (2004). A conserved transcript pattern in response to a specialist and a generalist herbivore. Plant Cell Online 16, 3132–3147. doi: 10.1105/tpc.104.026120
Rigby, D., and Cáceres, D. (2001). Organic farming and the sustainability of agricultural systems. Agric. Syst. 68, 21–40. doi: 10.1016/S0308-521X(00)00060-3
Ritchie, M. E., Phipson, B., Wu, D., Hu, Y., Law, C. W., Shi, W., et al. (2015). limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43:e47. doi: 10.1093/nar/gkv007
Rizhsky, L., Liang, H., and Mittler, R. (2002). The combined effect of drought stress and heat shock on gene expression in tobacco. Plant Physiol. 130, 1143–1151. doi: 10.1104/pp.006858
Rizhsky, L., Liang, H., Shuman, J., Shulaev, V., Davletova, S., and Mittler, R. (2004). When defense pathways collide. the response of Arabidopsis to a combination of drought and heat stress. Plant Physiol. 134, 1683–1696. doi: 10.1104/pp.103.033431
Robinson, M. D., McCarthy, D. J., and Smyth, G. K. (2010). edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140. doi: 10.1093/bioinformatics/btp616
Rudgers, J. A., Strauss, S. Y., and Wendel, J. F. (2004). Trade-offs among anti-herbivore resistance traits: insights from Gossypieae (Malvaceae). Am. J. Botany 91, 871–880. doi: 10.3732/ajb.91.6.871
Schmidt, D. D., Voelckel, C., Hartl, M., Schmidt, S., and Baldwin, I. T. (2005). Specificity in ecological interactions. attack from the same lepidopteran herbivore results in species-specific transcriptional responses in two solanaceous host plants. Plant Physiol. 138, 1763–1773. doi: 10.1104/pp.105.061192
Smith, C. M., Rodriguez-Buey, M., Karlsson, J., and Campbell, M. M. (2004). The response of the poplar transcriptome to wounding and subsequent infection by a viral pathogen. New Phytol. 164, 123–136. doi: 10.1111/j.1469-8137.2004.01151.x
Smyth, G. K., Ritchie, M., Thorne, N., Wettenhall, J., Shi, W., and Hu, Y. (2020). Limma: Linear Models for Microarray and RNA-Seq Data User's Guide. Melbourne, VIC: Bioinformatics Division, The Walter and Eliza Hall Institute of Medical Research.
Soffe, R. J. (2002). Primrose McConnell's the Agricultural Notebook, 20th Edn. Oxford: UK Wiley-Blackwell Science.
Staley, J. T., Stewart-Jones, A., Pope, T. W., Wright, D. J., Leather, S. R., Hadley, P., et al. (2010). Varying responses of insect herbivores to altered plant chemistry under organic and conventional treatments. Proc. Biol. Sci. 277, 779–786. doi: 10.1098/rspb.2009.1631
Stark, J. C., Thornton, M., and Nolte, P. (eds.). (2020). Potato Production Systems. Cham: Springer Nature. doi: 10.1007/978-3-030-39157-7
Stockdale, E. A., Shepherd, M. A., Fortune, S., and Cuttle, S. P. (2002). Soil fertility in organic farming systems – fundamentally different? Soil Use Manage. 18, 301–308. doi: 10.1079/SUM2002143
Strauss, S. Y. (1987). Direct and indirect effects of host-plant fertilization on an insect community. Ecology 68, 1670–1678. doi: 10.2307/1939859
Swezey, S. L., Werner, M. R., Buchanan, M., and Allison, J. (1998). Comparison of conventional and organic apple production systems during three years of conversion to organic management in coastal California. Am. J. Alternat. Agric. 13, 162–180. doi: 10.1017/S0889189300007876
Szczepaniec, A., Raupp, M. J., Parker, R. D., Kerns, D., and Eubanks, M. D. (2013). Neonicotinoid insecticides alter induced defenses and increase susceptibility to spider mites in distantly related crop plants. PLoS ONE 8:e62620. doi: 10.1371/journal.pone.0062620
Taurino, M., Abelenda, J. A., Río-Alvarez, I., Navarro, C., Vicedo, B., Farmaki, T., et al. (2014). Jasmonate-dependent modifications of the pectin matrix during potato development function as a defense mechanism targeted by Dickeya dadantii virulence factors. Plant J. 77, 418–429. doi: 10.1111/tpj.12393
Thimm, O., Bläsing, O., Gibon, Y., Nagel, A., Meyer, S., Krüger, P., et al. (2004). mapman: a user-driven tool to display genomics data sets onto diagrams of metabolic pathways and other biological processes. Plant J. 37, 914–939. doi: 10.1111/j.1365-313X.2004.02016.x
Thompson, G. A., and Goggin, F. L. (2006). Transcriptomics and functional genomics of plant defence induction by phloem-feeding insects. J. Exp. Botany 57, 755–766. doi: 10.1093/jxb/erj135
Trapnell, C., Pachter, L., and Salzberg, S. L. (2009). TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111. doi: 10.1093/bioinformatics/btp120
Triplehorn, C. A. J., Borror, N. F., Triplehorn, D. J. C. A., and Johnson, N. F. (2005). Borror and DeLong's Introduction to the Study of Insects. Belmont, CA: Thomson Learning Inc.
van Dijk, J. P., Cankar, K., Hendriksen, P. J., Beenen, H. G., Zhu, M., Scheffer, S., et al. (2012). The identification and interpretation of differences in the transcriptomes of organically and conventionally grown potato tubers. J. Agric. Food Chem. 60, 2090–2101. doi: 10.1021/jf204696w
Voelckel, C., and Baldwin, I. T. (2004). Herbivore-induced plant vaccination. Part II. array-studies reveal the transience of herbivore-specific transcriptional imprints and a distinct imprint from stress combinations. Plant J. 38, 650–663. doi: 10.1111/j.1365-313X.2004.02077.x
Watson, C.A, Atkinson, D., Gosling, P., Jackson, L. R., and Rayns, F. W. (2002). Managing soil fertility in organic farming systems. Soil Use Manag. 18, 239–247. doi: 10.1079/SUM2002131
Wink, M. (1988). Plant breeding: importance of plant secondary metabolites for protection against pathogens and herbivores. Theor. Appl. Genet. 75, 225–233. doi: 10.1007/BF00303957
Woods, J. L., Dreves, A. J., Fisher, G. C., James, D. G., Wright, L. C., and Gent, D. H. (2012). Population density and phenology of Tetranychus urticae (Acari: Tetranychidae) in hop is linked to the timing of sulfur applications. Environ. Entomol. 41, 621–635. doi: 10.1603/EN11279
Xia, J., and Wishart, D. S. (2016). Using metaboanalyst 3.0 for comprehensive metabolomics data analysis. Curr. Protocols Bioinformatics 55, 14–10. doi: 10.1002/cpbi.11
Zangerl, A. R., and Bazzaz, F. A. (1992). “Theory and pattern in plant defense allocation,” in Plant Resistance to Herbivores and Pathogens, Ecology, Evolution and Genetics, eds R. S. Fritz and E. L. Simms (Chicago, IL: University of Chicago Press), 363–391.
Zhu-Salzman, K., Salzman, R. A., Ahn, J.-E., and Koiwa, H. (2004). Transcriptional regulation of sorghum defense determinants against a phloem-feeding aphid. Plant Physiol. 134, 420–431. doi: 10.1104/pp.103.028324
Keywords: plant defensive pathways, soil bacteria, plant stress, herbivores, farming systems, potato
Citation: Krey KL, Nabity PD, Blubaugh CK, Fu Z, Van Leuven JT, Reganold JP, Berim A, Gang DR, Jensen AS and Snyder WE (2020) Organic Farming Sharpens Plant Defenses in the Field. Front. Sustain. Food Syst. 4:97. doi: 10.3389/fsufs.2020.00097
Received: 24 March 2020; Accepted: 09 June 2020;
Published: 17 July 2020.
Edited by:
Giuseppina Rea, National Research Council (CNR), ItalyReviewed by:
Juan Manuel Alba, University of Amsterdam, NetherlandsCopyright © 2020 Krey, Nabity, Blubaugh, Fu, Van Leuven, Reganold, Berim, Gang, Jensen and Snyder. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Karol L. Krey, a2Fyb2wua3JleUB1c2RhLmdvdg==
†Present address: Karol L. Krey, USDA-ARS-Yakima Agricultural Research Laboratory, Wapato, WA, United States
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.