
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Vet. Sci. , 19 March 2025
Sec. Veterinary Epidemiology and Economics
Volume 12 - 2025 | https://doi.org/10.3389/fvets.2025.1538725
This article is part of the Research Topic Pathogen Transmission at the Domestic-Wildlife Interface: A Growing Challenge that Requires Integrated Solutions - Volume II View all 5 articles
 Zhen-Qiu Gao1†
Zhen-Qiu Gao1† Si-Yuan Qin2†
Si-Yuan Qin2† Lin-Hong Xie2†
Lin-Hong Xie2† Guang-Rong Bao3
Guang-Rong Bao3 Xingzhou Wang4
Xingzhou Wang4 Ya Qin3
Ya Qin3 Xuetao Han4
Xuetao Han4 Xiaoming Yu4
Xiaoming Yu4 Cong-Cong Lei2
Cong-Cong Lei2 Xiao-Tian Zhang2
Xiao-Tian Zhang2 Ming-Yuan Yu2
Ming-Yuan Yu2 He-Ting Sun2*
He-Ting Sun2* Shuo Liu3*
Shuo Liu3*Introduction: Blastocystis sp. and Enterocytozoon bieneusi are common zoonotic pathogens threatening human and animal health. These parasites are widely distributed in birds, and substantial research on their prevalence has been conducted. However, no studies on Blastocystis sp. and E. bieneusi in raptors exist.
Methods: The present study collected 335 fecal samples from raptors in Changdao, China. The prevalence and genotypes of Blastocystis sp. and E. bieneusi were determined through amplification of SSU rRNA and ITS gene. Phylogenetic analysis was performed using MEGA 11 with the neighbor-joining method (Kimura 2-parameter model, 1000 replicate).
Results: The overall infection rates of Blastocystis sp. and E. bieneusi in raptors were 1.19% (4/335) and 1.79% (6/335), respecttively. Among them, the highest infection rate of Blastocystis sp. was observed in Accipiter nisus (3.85%, 1/26), while Buteo japonicus showed the highest infection rate of E. bieneusi (33.33%, 1/3), followed by Asio otus (7.69%, 1/13). This study identified two Blastocystis sp. subtypes: ST3 and ST10 in raptors for the first time. Regarding E. bieneusi in raptors, we identified four genotypes: CHN-F1, HND-III, BEB6, and HLJD-I. Among these, BEB6 and CHN-F1 are notable for their zoonotic potential and the risk of waterborne outbreaks.
Discussion: These findings suggest that raptors may be potential transmitters of Blastocystis sp. and E. bieneusi to humans and other animals, as well as sources of water contamination. This study fills a gap in the research on Blastocystis sp. and E. bieneusi in raptors and is important for public health safety.
Blastocystis sp. is a zoonotic intestinal protozoan of the family Blastocystidae, which can be transmitted through foodborne, waterborne, person-to-person, and zoonotic routes (1). Approximately one billion people worldwide are infected with this parasite, with infection rates among diarrhea patients ranging from 0.8 to 100% (2). However, a large number of Blastocystis sp. have been detected in both symptomatic and asymptomatic individuals (3). Therefore, their role in the host's gut (whether as a symbiont or a parasite) remains unclear, and their pathogenicity has yet to be conclusively determined. Blastocystis sp. exhibits significant genetic diversity, with 44 subtypes reported, of which 38 STs have been confirmed as valid subtypes. Studies show that 14 subtypes, including ST1–ST10, ST12, ST14, ST16, and ST23, are recognized as zoonotic subtypes (4). Approximately 90% of human infections are caused by ST1–ST4, with ST3 being the dominant subtype (5). Other subtypes are primarily found in other animals (6).
Microsporidia are a highly diverse group of parasites, both in terms of host range and genetic variation (7). Over 200 genera and more than 1,700 species are known globally, capable of infecting up to 240 species of vertebrates and invertebrates (8). Among human infections caused by microsporidia, Enterocytozoon bieneusi is the most common pathogenic species, accounting for approximately 90% of all microsporidia infections in humans (9). The pathogenicity of E. bieneusi is closely linked to the host's immune status (10). In individuals with normal immune function, the infection is typically asymptomatic or presents with mild symptoms. However, in immunocompromised individuals, such as those with HIV, the infection can lead to diarrhea, abdominal pain, and, in severe cases, even death (11). Based on analysis of the Internal Transcribed Spacer (ITS) nucleotide sequence, 820 genotypes of E. bieneusi have been identified and further categorized into 15 phylogenetic groups (12). Group 1 includes most zoonotic genotypes, which exhibit strong cross-species transmission capabilities and high zoonotic potential (12). Group 2 primarily infects ruminants, although some genotypes (such as BEB4, BEB6, I, and J) have been reported to infect humans. Groups 3–15 exhibit stronger host specificity, and their zoonotic potential is either limited or unclear (8).
As global ecosystems change, the habitats of wildlife are shrinking, especially under the intensified influence of human activities. As a result, the interaction space between domesticated and wild animals has become increasingly complex. Raptors, as apex predators in the food chain, have close ecological interactions with various poultry, livestock, and other wildlife, making them potential carriers for parasite transmission. Studies have shown that the prevalence of Blastocystis sp. in global bird populations is 29% (2). In addition, a meta-analysis indicates that the infection rate of microsporidia in birds worldwide is 14.6%, with E. bieneusi accounting for 77.42% of microsporidian infections (13). These findings suggest that both pathogens are widely distributed among birds. If raptors were infected with these two parasites, it would not only threaten their health but also, through ecological interactions, may impact other species and even human health. However, there have been no reports of raptors being infected with Blastocystis sp. and E. bieneusi. This knowledge gap hinders our understanding of the role of raptors in the transmission of these parasites and their potential impacts on human and animal health.
This study is the first to investigate the prevalence and genetic diversity of Blastocystis sp. and E. bieneusi in raptors. The findings will provide data to support ecological restoration and public health safety and offer valuable insights into the transmission risks of infectious diseases in raptors and potential preventive measures.
From September to October 2024, a total of 335 samples were collected form raptors in Shandong Changdao National Nature Reserve, including Caprimulgus jotaka (n = 2), Asio flammeus (n = 3), Accipiter gentilis (n = 3), Pernis ptilorhynchus (n = 2), Otus sunia (n = 242), Otus lettia (n = 1), Falco tinnunculus (n = 2), Buteo japonicus (n = 3), Accipiter nisus (n = 26), Accipiter gularis (n = 28), Accipiter virgatus (n = 4), Falco subbuteo (n = 1), Ninox scutulata (n = 2), Falco peregrinus (n = 3), Asio otus (n = 13). We captured raptors using nets on Daheishan Island, Changdao, and released them after collecting cloacal swab samples. During sampling, we recorded the species and sampling time. All samples were transported to the laboratory on dry ice and stored at −80°C. All sampling procedures strictly adhered to the guidelines of the Ethics Committee of Qingdao Agricultural University.
After adding 500 μL of physiological saline to each cloacal swab, the mixture was vortexed at maximum speed until the feces were completely detached from the swab. According to the manufacturer's instructions, DNA was extracted using the E.Z.N.A. Stool DNA Extraction Kit (Omega Biotek Inc, Norcross, GA, USA). The extracted DNA samples were stored at −20°C until PCR analysis.
Nested PCR was performed to amplify the Internal Transcribed Spacer (ITS) region to detect the presence of E. bieneusi in the samples. In the first round, external primers NEBF1 (5′-GGTCATAGGGATGAAGAG-3′) and NEBR1 (5′-TTCGAGTTCTTTCGCGCTC-3′) were used, with the following reaction program: 94 for 5 min for pre-denaturation; 94°C for 45 s, 55°C for 45 s, 72°C for 1 min, for 35 cycles; and a final extension at 72°C for 10 min. In the second round, internal primers NEBF2 (5′-GCTCTGAATATCTATGGCT-3′) and NEBR2 (5′-ATCGCCGACGGATCCAAGTG-3′) were used, with an extension time of 40 s and other reaction conditions the same as the first round (14). Additionally, forward primer RD5 (5′-ATCTGGTTGATCCTGCCAGT-3′) and reverse primer BhRDr (5′-GAGCTTTTTAACTGCAACAACG-3′) were used to amplify the small subunit ribosomal RNA (SSU rRNA) gene to detect the presence of Blastocystis sp. in the samples, under the following conditions: 94°C for 5 min for pre-denaturation; 94°C for 45 s, 57°C for 45 s, 72°C for 1 min, for 35 cycles; and a final extension at 72°C for 10 min (15). All PCR reactions included positive and negative controls. PCR products were analyzed by electrophoresis on a 1.5% agarose gel to assess the amplification results.
All PCR-positive products were subjected to bidirectional sequencing by Qingdao Weilai Biotechnology Co., Ltd. Representative sequences were obtained by clustering at 100% similarity using cd-hit. These representative sequences were then compared with reference sequences from GenBank. A phylogenetic tree was constructed in MEGA 11 software using the neighbor-joining method (NJ) combined with the Kimura 2-parameter model, and the reliability of the results was assessed through 1,000 bootstrap replicates.
Data analysis was performed using SPSS software (IBM Corp., Armonk, NY, USA), calculating the odds ratio (OR) and 95% confidence intervals (95% CI) of the infection rate of parasites. A p-value of < 0.05 was considered statistically significant.
In this study, four Blastocystis sp.-positive samples were detected from 335 raptor fecal swab samples, with an overall prevalence was 1.19% (4/335, 95% CI 0.33–3.03). No significant differences in prevalence were found between different raptor species (χ2 = 4.45, df = 14, P = 0.9921). Among the species, Accipiter nisus had the highest prevalence (3.85%, 1/26), followed by Otus sunia (1.24%, 3/242). No Blastocystis sp. infection was detected in other species (Table 1, Figure 1).
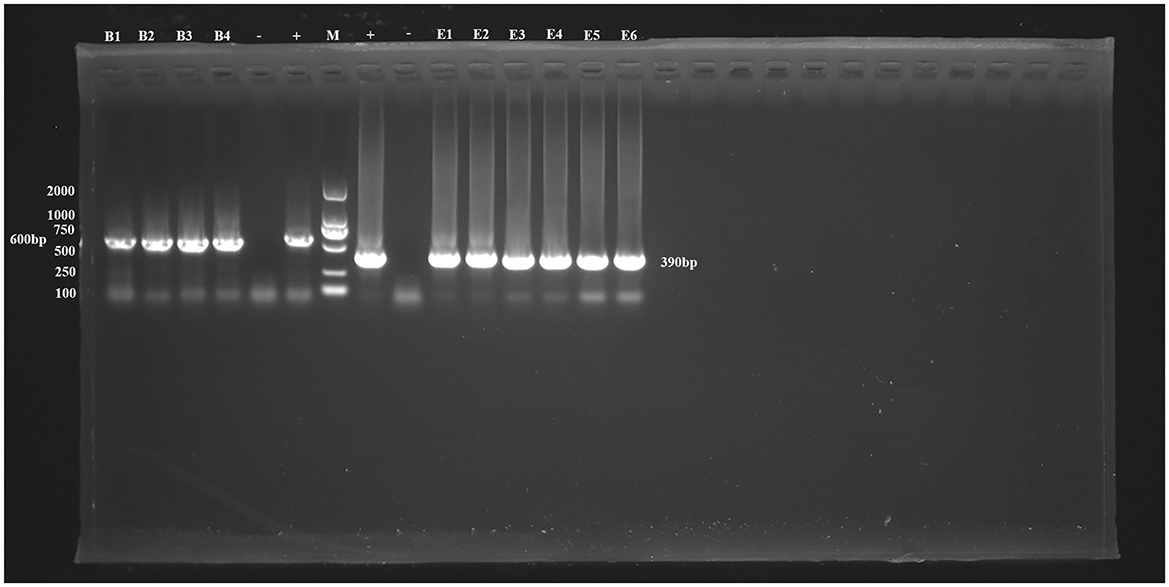
Figure 1. PCR amplification results for the SSU rRNA gene of Blastocystis sp. and the ITS gene of E. bieneusi. B1–B4 represent Blastocystis sp.-positive samples, E1–E6 represent E. bieneusi-positive samples. –, Negative control; +, Positive control; M, Marker.
Additionally, six positive samples of E. bieneusi were detected in this study, resulting in an overall infection rate of 1.79% (6/335, 95% CI 0.66–3.86). The highest infection rate was found in Buteo japonicus (33.33%, 1/3), followed by Asio otus (7.69%, 1/13), Accipiter gularis (3.57%, 1/28), and Otus sunia (1.24%, 3/242). No E. bieneusi infection was found in other species. The differences in infection rates between species were not statistically significant (χ2 = 9.45, df = 14, P = 0.8011; Table 2, Figure 1).
Through sequence analysis of Blastocystis sp.-positive samples, this study identified two Blastocystis sp. subtypes, ST10 and ST3, with ST10 being the dominant subtype (75%, 3/4), found in both Accipiter nisus and Otus sunia. ST3 was detected only in Otus sunia (Table 1).
Additionally, sequence analysis of six E. bieneusi-positive samples identified four genotypes: HLJD-I, BEB6, HND-III, and CHN-F1. BEB6 (33.33%, 2/6) was found in both Asio otus and Otus sunia, with HLJD-I (66.66%, 2/3) being the dominant genotype in Otus sunia. HND-III and CHN-F1 were detected in Accipiter gularis and Buteo japonicus, respectively (Table 2).
This study obtained two representative sequences of Blastocystis sp. through clustering. Phylogenetic analysis showed that PQ643313 clustered with the reference sequence MT798805 in the same clade, both belonging to the ST10 subtype. PQ643314 showed 100% similarity with the human-derived reference sequence MK782518 and clustered in the ST3 subtype (Figure 2).
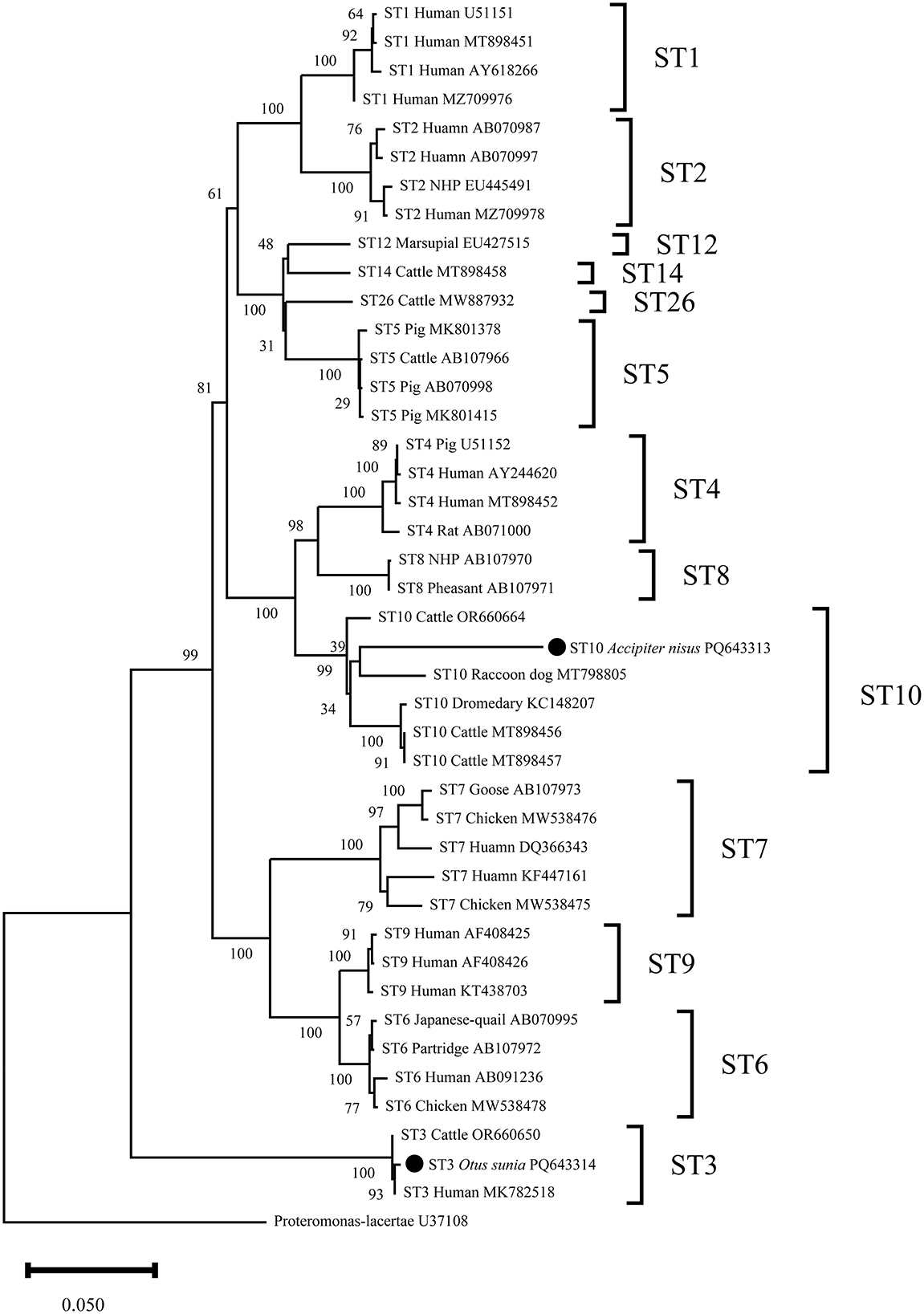
Figure 2. Phylogenetic relationship between the Blastocystis sp. sequences in this study and the reference sequences downloaded from GenBank was assessed using the neighbor-joining (NJ) method, with genetic distances calculated based on the Kimura 2-parameter model. The nucleotide sequences identified in this study are marked with black circles before the subtype names. The numbers on the branches represent the bootstrap percentage values from 1,000 repetitions, with values >50% shown in the tree.
The phylogenetic analysis of six representative E. bieneusi sequences revealed that PQ643841 shared 99% similarity with the reference sequence MW756993 and clustered in the first group. The other representative sequences clustered in the second group, with PQ643838 and PQ643843 located in the same branch. These two sequences, along with the reference sequence KX383618, formed a sister group and all belong to the HLJD-I genotype. PQ6463839 showed 99% similarity with the reference sequence KX383636 and formed a separate branch. PQ643840 and PQ643842 are of the BEB6 genotype and clustered with the reference sequence MK139947 in the same clade (Figure 3).
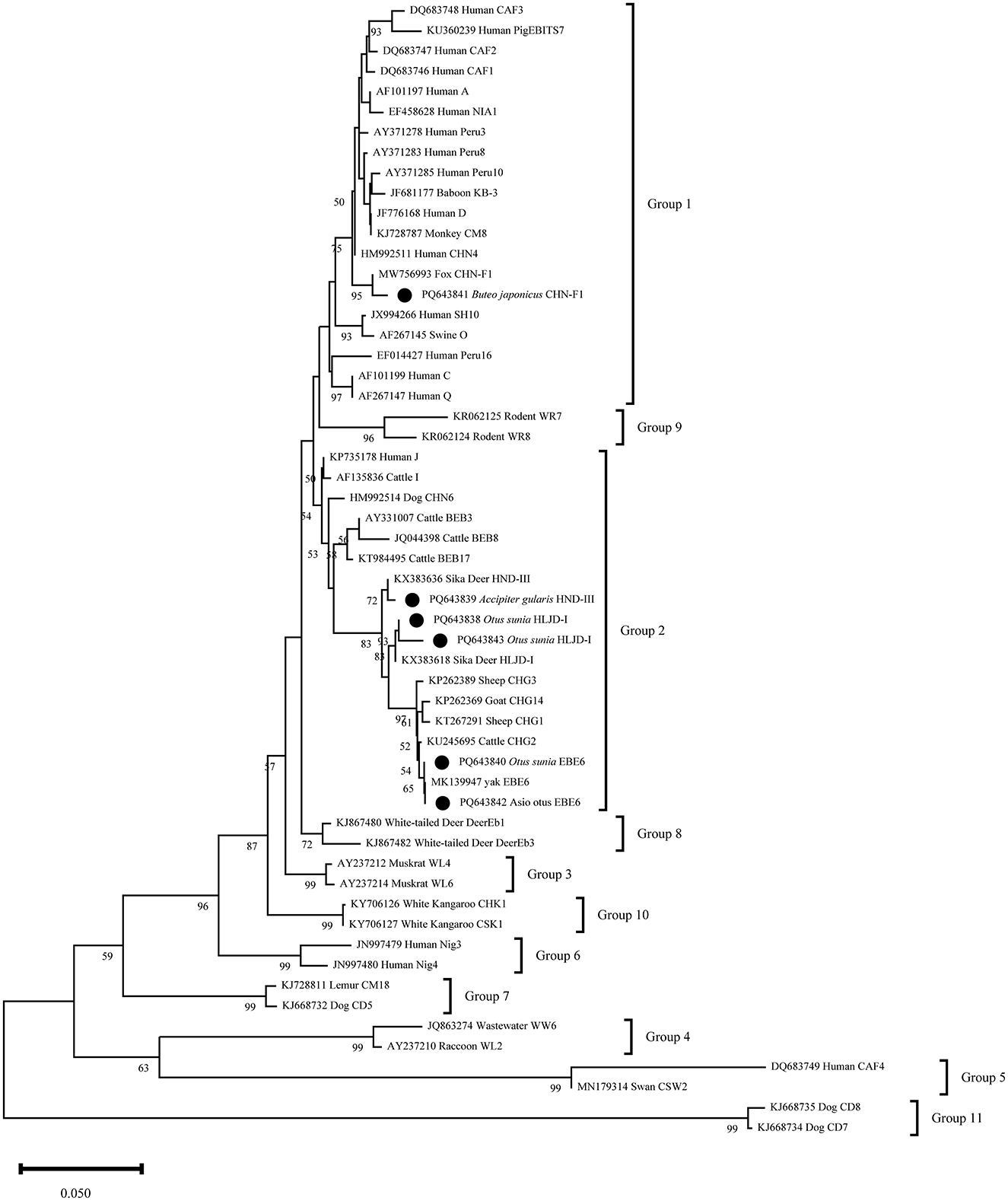
Figure 3. Phylogenetic relationship between the E. bieneusi sequences in this study and the reference sequences downloaded from GenBank was assessed using the neighbor-joining (NJ) method, with genetic distances calculated based on the Kimura 2-parameter model. The nucleotide sequences identified in this study are marked with black circles before the subtype names. The numbers on the branches represent the bootstrap percentage values from 1,000 repetitions, with values >50% shown in the tree.
A meta-analysis has shown that the global prevalence of Blastocystis sp. in birds is 29% (2). The prevalence is 35% in Chinese peafowls, 17.8% in Malaysian quails, and 42.9% in Iranian pigeons (16–18). These data indicate that Blastocystis sp. is widely distributed among birds globally. In this study, the overall prevalence of Blastocystis sp. in raptors was 1.19%, which is lower than the 4.92% prevalence reported in migratory birds in Northeast China (19). This difference may be attributed to sampling methods, geographic location, and other factors, and the specific reasons warrant further investigation. Additionally, the highest prevalence was observed in Accipiter nisus (3.85%, 1/26), followed by Otus sunia (1.24%, 3/242), with no Blastocystis sp. infection detected in other raptor species. This variation could be linked to the host's immune status. As a common parasite in birds, E. bieneusi had an overall infection rate of 1.79% (6/335) in this study, which is lower than the infection rates in Central European pigeons (13.29%), Iraqi pet birds (32.32%), and Chinese swans [7.49%; (20–22)]. The low prevalence in raptors may indicate a relatively low exposure or low suitability of raptors as hosts of E. bieneusi compared to other birds. Furthermore, the infection rates of E. bieneusi in different raptor species ranged from 0 to 33.33%, with the highest infection rate observed in Buteo japonicus (33.33%). This difference may be related to the the sample size as well as the differences in susceptibility among different species to parasites.
The known Blastocystis sp. subtypes that can infect humans range from ST1 to ST10, with ST6 and ST7 being widespread in various bird species such as poultry, peafowls, and quails, and are considered the dominant subtypes in birds (16, 17, 23). However, the results of this study differ from other bird studies. We did not detect ST6 or ST7 in raptors, but instead unexpectedly identified ST3 and ST10. Epidemiological data show that ST3 is the primary subtype responsible for human infections and is widely found in dogs, rodents, and cattle, indicating that this subtype has strong host adaptability and significant zoonotic potential (24–26). ST10 is widely distributed in animals such as cattle, sheep, and camels, and is considered one of the most specific subtypes in ruminants (24, 27, 28). The infection of ST3 and ST10 in raptors is likely due to interactions with domesticated or wild animals and there is also a risk of cross-species transmission to humans, particularly among birdwatchers. Moreover, the increasing encroachment of human activities into wildlife habitats may facilitate such cross-species transmission, highlighting the need for greater attention to zoonotic risks in wildlife monitoring. An important observation is that Noradilah et al. isolated ST10 from river water in Malaysia, further suggesting that water sources could serve as a significant route for the transmission of Blastocystis sp. (29). This further emphasizes the environmental context in which Blastocystis sp. transmission may occur, where water contamination from wildlife could contribute to broader ecological and public health risks.
This study identified four genotypes of E. bieneusi from raptors: CHN-F1, HLJD-I, BEB6, and HND-III. CHN-F1 is classified into Group 1, initially discovered by Zhao et al. in foxes in China and widely present in farmed fur animals in China (30, 31). Subsequently, Holubová et al. reported this genotype in pigeons in Central Europe (18). Although no human infections have been reported so far, CHN-F1 has the potential to cause zoonotic diseases, and its cross-species transmission ability should not be overlooked. HLJD-I, BEB6, and HND-III are classified into Group 2, which was initially considered to be specific to ruminants (8). However, subsequent studies have shown that genotypes such as BEB4, BEB6, I, and J have been found in humans and other animals, indicating that this group poses a public health risk (32). HLJD-I and HND-III were first discovered in sika deer in Heilongjiang and Henan, China, respectively, and this study is the first to detect these two genotypes in birds, suggesting that these genotypes are no longer restricted to their original hosts and may have cross-species transmission potential (33). To our knowledge, BEB6 was first discovered and named in cattle, but later studies confirmed that this genotype is more common in sheep and has also been sporadically found in non-human primates, raccoon dogs, and other animals, suggesting that it has low host specificity and high adaptability (22, 31, 34–36). Notably, as reported by Ye et al., this genotype has already been detected in a wastewater treatment facility in Zhengzhou, China (37), suggesting that raptors may contaminate surface water sources through their feces, thereby posing a potential risk for waterborne outbreaks. This finding further highlights the potential role of raptors in environmental pollution, and future research should focus on the ecological risks of waterborne transmission of zoonotic pathogens.
However, this study has some limitations. For example, the sample size for certain raptor species is relatively small, which may affect the accuracy of infection rate estimates, especially for rare species. Additionally, the study was conducted in a single geographic location, which may not fully reflect the distribution patterns of these parasites in other areas. Another limitation is the lack of consideration of seasonal variation in parasite prevalence, which is an important factor in understanding the dynamics of infection. Future research should focus on expanding the sample size, especially for rare species, and incorporating multiple geographic regions to capture a broader range of ecological and environmental gradients. Additionally, implementing long-term monitoring to assess the seasonal and temporal trends of parasite prevalence will greatly enhance our understanding of the epidemiological patterns of raptor parasitic infections.
In conclusion, this study is the first to reveal the prevalence of Blastocystis sp. and E. bieneusi in raptors, with infection rates of 1.19% and 1.79%, respectively. We identified two zoonotic Blastocystis sp. subtypes (ST3 and ST10), which were detected in birds for the first time. The study also identified four E. bieneusi genotypes (CHN-F1, HND-III, BEB6, and HLJD-I), among which BEB6 and CHN-F1, as zoonotic genotypes, were found in raptors for the first time. Both BEB6 and ST10 have the potential to cause waterborne outbreaks. These results suggest that raptors may be potential cross-species transmitters, creating new transmission pathways between humans and animals, and highlighting the potential ecological interactions and zoonotic risks between wild and domestic animals, particularly the threat of water contamination by raptors. Therefore, future research should focus on the interaction patterns between raptors, domestic animals, and other wildlife, and explore additional control strategies to reduce the risks of cross-species disease transmission and water contamination.
The representative sequences of Blastocystis sp. obtained in this study have been submitted to GenBank, with sequence numbers PQ643313 and PQ643314. The representative sequences of E. beneusi have been submitted to GenBank, with sequence numbers: PQ643838-PQ643843.
The animal study was approved by Research Ethics Committee for the Care and Use of Laboratory Animals in Qingdao Agricultural University, China. The study was conducted in accordance with the local legislation and institutional requirements.
Z-QG: Conceptualization, Methodology, Supervision, Writing – review & editing. S-YQ: Conceptualization, Methodology, Resources, Writing – review & editing. L-HX: Conceptualization, Methodology, Resources, Writing – review & editing. G-RB: Methodology, Resources, Writing – review & editing. XW: Writing – review & editing. YQ: Resources, Writing – review & editing. XH: Resources, Writing – review & editing. XY: Resources, Writing – review & editing. C-CL: Resources, Writing – review & editing. X-TZ: Resources, Writing – review & editing. M-YY: Resources, Writing – review & editing. H-TS: Conceptualization, Funding acquisition, Writing – review & editing. SL: Methodology, Software, Writing – original draft.
The author(s) declare that financial support was received for the research and/or publication of this article. This work was supported by the National Key Research and Development Program of China (2021YFC2300903).
We sincerely thank the Qingdao Birding Association for their invaluable support and assistance during the sampling process.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Gen AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
1. Asghari A, Hassanipour S, Hatam G. Comparative molecular prevalence and subtypes distribution of Blastocystis sp. a potentially zoonotic infection isolated from symptomatic and asymptomatic patients in Iran: a systematic review and meta-analysis. Acta Parasitologica. (2021) 66:745–59. doi: 10.1007/s11686-021-00360-0
2. Barati M, KarimiPourSaryazdi A, Rahmanian V, Bahadory S, Abdoli A, Rezanezhad H, et al. Global prevalence and subtype distribution of blastocystis sp. in rodents, birds, and water supplies: a systematic review and meta-analysis. Prev Vet Med. (2022) 208:105770. doi: 10.1016/j.prevetmed.2022.105770
3. Zhao W, Ren G, Wang L, Xie L, Wang J, Mao J, et al. Molecular prevalence and subtype distribution of Blastocystis spp. among children who have diarrheia or are asymptomatic in Wenzhou, Zhejiang Province, China. Parasite. (2024) 31:12. doi: 10.1051/parasite/2024012
4. Naguib D, Gantois N. Molecular epidemiology and genetic diversity of the enteric protozoan parasite Blastocystis sp. in the Northern Egypt population. Pathogens. (2023) 12:1359. doi: 10.3390/pathogens12111359
5. Alfellani MA, Taner-Mulla D, Jacob AS, Imeede CA, Yoshikawa H, Stensvold CR, et al. Genetic diversity of blastocystis in livestock and zoo animals. Protist. (2013) 164:497–509. doi: 10.1016/j.protis.2013.05.003
6. Su C, Mei X, Feng X, Zhang F, Wang P, He B, et al. Prevalence and molecular subtyping of Blastocystis sp. in rabbits in Henan, central China. Folia Parasitol. (2022) 69:2022.027. doi: 10.14411/fp.2022.027
7. Taghipour A, Bahadory S, Khazaei S, Zaki L, Ghaderinezhad S, Sherafati J, et al. Global molecular epidemiology of microsporidia in pigs and wild boars with emphasis on Enterocytozoon bieneusi: a systematic review and meta-analysis. Vet Med Sci. (2022) 8:1126–36. doi: 10.1002/vms3.751
8. Li W, Feng Y, Santin M. Host specificity of Enterocytozoon bieneusi and public health implications. Trends Parasitol. (2019) 35:436–51. doi: 10.1016/j.pt.2019.04.004
9. Wang Y, Li XM, Yang X, Wang XY, Wei YJ, Cai Y, et al. Global prevalence and risk factors of Enterocytozoon bieneusi infection in humans: a systematic review and meta-analysis. Parasite. (2024) 31:9. doi: 10.1051/parasite/2024007
10. Han B, Pan G, Weiss LM. Microsporidiosis in humans. Clin Microbiol Rev. (2021) 34:e0001020. doi: 10.1128/CMR.00010-20
11. Leśniańska K, Perec-Matysiak A. Wildlife as an environmental reservoir of Enterocytozoon bieneusi (Microsporidia)–analyses of data based on molecular methods. Ann Parasitol. (2017) 63:265–81. doi: 10.17420/ap6304.113
12. Zhao W, Zhou H, Yang L, Ma T, Zhou J, Liu H, et al. Prevalence, genetic diversity and implications for public health of Enterocytozoon bieneusi in various rodents from Hainan Province, China. Parasit Vectors. (2020) 13:438. doi: 10.1186/s13071-020-04314-9
13. Taghipour A, Ghodsian S, Jabbari M, Rajabpour V, Bahadory S, Malih N, et al. The global epidemiology of microsporidia infection in birds: a systematic review and meta-analysis. Int J Environ Health Res. (2024) 34:2180–96. doi: 10.1080/09603123.2023.2219988
14. Buckholt MA, Lee JH, Tzipori S. Prevalence of Enterocytozoon bieneusi in swine: an 18-month survey at a slaughterhouse in Massachusetts. Appl Environ Microbiol. (2002) 68:2595–9. doi: 10.1128/AEM.68.5.2595-2599.2002
15. Scicluna SM, Tawari B, Clark CG. DNA barcoding of blastocystis. Protist. (2006) 157:77–85. doi: 10.1016/j.protis.2005.12.001
16. Liu X, Zhang S, Dong H, Wang R, Zhang L, Ge Y. Prevalence and molecular characteristics of Blastocystis sp. from peafowl (Pavo cristatus) in China. J Parasitol. (2021) 107:790–3. doi: 10.1645/19-72
17. Rauff-Adedotun AA, Douglas SJ, Rajamanikam A, Amira Suriaty Y, Farah Haziqah MT. Natural infection of blastocystis ST6 among commercial quails (Coturnix) in Penang, Malaysia. Trop Biomed. (2022) 39:73–7. doi: 10.47665/tb.39.1.012
18. Rostami M, Fasihi-Harandi M, Shafiei R, Aspatwar A, Derakhshan FK, Raeghi S. Genetic diversity analysis of Blastocystis subtypes and their distribution among the domestic animals and pigeons in Northwest of Iran. Infect Genet Evol. (2020) 86:104591. doi: 10.1016/j.meegid.2020.104591
19. Wang HJ, Chen D, Ma ZH, Liu CF, Li WD, Hao Y, et al. Molecular detection, subtyping of Blastocystis sp. in migratory birds from nature reserves in Northeastern China. Acta Trop. (2024) 258:107355. doi: 10.1016/j.actatropica.2024.107355
20. Holubová N, Zikmundová V, Kicia M, Zajaczkowska Ż, Rajský M, Konečný R, et al. Genetic diversity of Cryptosporidium spp., Encephalitozoon spp. and Enterocytozoon bieneusi in feral and captive pigeons in Central Europe. Parasitol Res. (2024) 123:158. doi: 10.1007/s00436-024-08169-2
21. Abdullah DA, Alobaidii WA, Alkateb YNM, Ali FF, Ola-Fadunsin SD, Gimba FI. Molecular detection and prevalence of human-pathologic Enterocytozoon bieneusi among pet birds in Mosul, Iraq. Comp Immunol Microbiol Infect Dis. (2023) 95:101964. doi: 10.1016/j.cimid.2023.101964
22. Wang Y, Zhang K, Zhang Y, Wang K, Gazizova A, Wang L, et al. First detection of Enterocytozoon bieneusi in whooper swans (Cygnus) in China. Parasit Vectors. (2020) 13:5. doi: 10.1186/s13071-020-3884-y
23. Salehi R, Rostami A. Genetic characterization of Blastocystis from poultry, livestock animals and humans in the southwest region of Iran-zoonotic implications. Transbound Emerg Dis. (2022) 69:1178–85. doi: 10.1111/tbed.14078
24. Duan J, Qin H, Sun M, Fu Y, Lang J, Zhang A, et al. Occurrence and genotypic identification of Blastocystis sp. Enterocytozoon bieneusi, and Giardia duodenalis in dairy cattle in Heilongjiang Province, China. Parasitol Int. (2024) 100:102871. doi: 10.1016/j.parint.2024.102871
25. Liao S, Lin X, Sun Y, Qi N, Lv M, Wu C, et al. Occurrence and genotypes of Cryptosporidium spp, Giardia duodenalis, and Blastocystis sp in household, shelter, breeding, and pet market dogs in Guangzhou, Southern China. Scientific Reports. (2020) 10:17736. doi: 10.1038/s41598-020-74299-z
26. Mohammadpour I, Bozorg-Ghalati F, Gazzonis AL, Manfredi MT, Motazedian MH, Mohammadpour N. First molecular subtyping and phylogeny of Blastocystis sp. isolated from domestic and synanthropic animals (dogs, cats and brown rats) in Southern Iran. Parasit Vectors. (2020) 13:365. doi: 10.1186/s13071-020-04225-9
27. Shams M, Bahrami AM, Mousivand A, Shamsi L, Asghari A. First molecular characterization of Blastocystis subtypes from domestic animals (sheep and cattle) and their animal-keepers in Ilam, Western Iran: a zoonotic concern. J Eukaryot Microbiol. (2024) 71:e13019. doi: 10.1111/jeu.13019
28. Asghari A, Yousefi A, Badali R, Mohammadi MR, Shamsi L, Maleki F, et al. First molecular subtyping and zoonotic significance of Blastocystis sp. in dromedary (C. dromedarius) and Bactrian (C. bactrianus) camels in Iran: a molecular epidemiology and review of available literature. Vet Med Sci. (2024) 10:e1442. doi: 10.1002/vms3.1442
29. Attah AO, Sanggari A, Li LI, Nik Him NAII, Ismail AH, Meor Termizi FH. Blastocystis occurrence in water sources worldwide from 2005 to 2022: a review. Parasitology Researsh. (2023) 122:1–10. doi: 10.1007/s00436-022-07731-0
30. Zhao W, Zhang W, Yang Z, Liu A, Zhang L, Yang F, et al. Genotyping of Enterocytozoon bieneusi in farmed blue foxes (Alopex Lagopus) and raccoon dogs (Nyctereutes Procyonoides) in China. PLoS ONE. (2015) 10:e0142611. doi: 10.1371/journal.pone.0142611
31. Chen M, Wang H, Li X, Guo Y, Lu Y, Zheng L, et al. Molecular epidemiology of Enterocytozoon bieneusi from foxes and raccoon dogs in the Henan and Hebei provinces in China. BMC Vet Res. (2024) 20:53. doi: 10.1186/s12917-024-03883-6
32. Nourrisson C, Lavergne RA, Moniot M. Enterocytozoon bieneusi, a human pathogen. Emerg Microbes Infect. (2024) 13:2406276. doi: 10.1080/22221751.2024.2406276
33. Zhao W, Zhang W, Wang R, Liu W, Liu A, Yang D, et al. Enterocytozoon bieneusi in sika deer (Cervus Nippon) and red deer (Cervus Elaphus): deer specificity and zoonotic potential of its genotypes. Parasitol Res. (2014) 113:4243–50. doi: 10.1007/s00436-014-4100-9
34. Du SZ, Zhao GH, Shao JF, Fang YQ, Tian GR, Zhang LX, et al. Cryptosporidium spp, Giardia intestinalis, and Enterocytozoon bieneusi in captive non-human primates in Qinling Mountains. Korean J Parasitol. (2015) 53:395–402. doi: 10.3347/kjp.2015.53.4.395
35. Apaydin MH, Yetişmiş G. Molecular prevalence and phylogenetic characterization of Enterocytozoon bieneusi in sheep in the van region. Turkiye Parazitoloji Dergisi. (2023) 47:64–70. doi: 10.4274/tpd.galenos.2022.76476
36. Karim MR, Rume FI, Li D, Li J, Zhang L. First molecular characterization of Enterocytozoon bieneusi in children and calves in Bangladesh. Transbound Emerg Dis. (2022) 69:1999–2007. doi: 10.1111/tbed.14187
Keywords: Enterocytozoon bieneusi, Blastocystis sp., raptor, prevalence, China
Citation: Gao Z-Q, Qin S-Y, Xie L-H, Bao G-R, Wang X, Qin Y, Han X, Yu X, Lei C-C, Zhang X-T, Yu M-Y, Sun H-T and Liu S (2025) First report of Blastocystis sp. and Enterocytozoon bieneusi in raptors. Front. Vet. Sci. 12:1538725. doi: 10.3389/fvets.2025.1538725
Received: 03 December 2024; Accepted: 28 February 2025;
Published: 19 March 2025.
Edited by:
Saúl Jiménez-Ruiz, University of Cordoba, SpainReviewed by:
Houqiang Luo, Wenzhou Vocational College of Science and Technology, ChinaCopyright © 2025 Gao, Qin, Xie, Bao, Wang, Qin, Han, Yu, Lei, Zhang, Yu, Sun and Liu. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: He-Ting Sun, eGlhb2ZlbmdzaHRAMTYzLmNvbQ==; Shuo Liu, THMxNjY0MTFAMTYzLmNvbQ==
†These authors have contributed equally to this work
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.