
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
REVIEW article
Front. Oncol. , 01 April 2025
Sec. Molecular and Cellular Oncology
Volume 15 - 2025 | https://doi.org/10.3389/fonc.2025.1552773
This article is part of the Research Topic Long Noncoding RNAs in Cancer View all articles
 WeiChi Liu
WeiChi Liu WeiFa Wang*
WeiFa Wang*Gastric cancer (GC) is an exceedingly aggressive disease and ranks as the third leading cause of cancer-related deaths, which poses a huge health burden globally. Chemotherapy is commonly employed during the middle to advanced stages of cancer, although it faces frequent treatment failures attributed to drug resistance. Thus, it is imperative for researchers to identify potential targets for overcoming therapeutic resistance, thereby facilitating the development of novel anti-cancer agents for GC patients with advanced stages. Long noncoding RNAs (lncRNAs) are a diverse group of transcripts with limited protein-coding capacity, which have been recognized for functional molecules for regulating cancer progression including cell proliferation, metastasis, and drug resistance in GC. In this review, we examine the intricate molecular networks on the role of lncRNAs in drug resistance of GC. LncRNAs conferred cancer cell resistance to anti-cancer drug through various molecular mechanisms, therefore functioning as promising therapeutic targets for GC patients. Additionally, we discuss current advancements of strategies targeting lncRNAs in cancer therapy, which may pave the way for lncRNA-mediated precision medicine for this malignant disease.
Gastric cancer (GC) remains the second most prevalent types of gastrointestinal cancers and ranks the fifth of cancer-related deaths globally in 2022, which significantly contributes to estimated 6.8% of the overall cancer mortality rate (1). Risk factors of gastric cancer include infection with Helicobacter pylori, advancing age, unhealthy lifestyle and diets (2). Nowadays, the disease diagnosis is made histologically following an endoscopic biopsy, and staging is performed using imaging examinations such as laparoscopy, CT scans, and endoscopic ultrasound (3). It exhibits significant molecular and phenotypic heterogeneity, which poses challenges for disease diagnosis and treatment (2). Currently, the primary treatment for early-stage GC is implicated in the surgical resection (4). But due to concealed symptoms in the tumorigenesis, most patients are diagnosed at advanced stages with distant metastasis, who are no longer a suitable candidate for surgical treatment (5). Therefore, chemotherapy, radiotherapy, molecular targeted therapy, immunotherapy or a combination of these methods function as feasible options for advanced GC patients with distant metastasis (5). Although tumors enter remission rapidly under standard therapy, they develop resistance, ultimately leading to treatment failure and disease relapse (6). Therefore, it is essential for researchers to identify potential targets for overcoming therapeutic resistance and pave the way for the developing novel anti-cancer agents in advanced GC.
Long noncoding RNAs (lncRNAs) are a diverse group of transcripts with limited protein-coding ability that exceed 200 nucleotides in length (7). Initially, lncRNAs were considered as “noise” produced during the RNA transcription process due to the lack of protein-coding capability (8). However, with ongoing research, lncRNAs have been found to possess limited protein-coding potential and play significant roles in various pathophysiological processes through complex molecular mechanisms (9–11). Dysregulated lncRNAs function as oncogenes or tumor suppressors that regulate cell proliferation, invasion, migration, metabolic reprogramming, and therapeutic resistance across multiple types of human cancers (12–15). LncRNAs are essential for gene regulation, participating in the regulation of genetic activation and silencing, epigenetic modifications and post-translational regulation, thus impacting tumor progression (15, 16). In the context of GC, abnormal lncRNA expression is closely associated with tumor progression, and therapeutic resistance, thus targeting lncRNAs hold great potential for GC treatment (17–19). However, the detailed molecular mechanisms await further investigations.
In this review, we elaborate the underlying mechanisms involved in the role of lncRNAs in GC progression. Specifically, we depict the intricate mechanistic network of lncRNAs in drug re istance, which indicting that lncRNAs are promising therapeutic targets for GC therapy. Furthermore, we discuss the emerging therapeutic strategies that targeting lncRNAs in cancer therapy, aiming to offer a thorough and structured resource for researchers focusing on lncRNA-based therapeutic therapies.
Since the initial identification of lncRNAs through high-throughput sequencing technologies, their extensive potential to modulate the gene expression, transcription, and protein translation has become increasingly evident (20). LncRNAs are synthesized by the activity of RNA polymerase II and display features commonly attributed to protein-coding messenger RNAs (mRNAs), including the presence of a poly-A+ tail and a 5' cap (21). LncRNAs can be classified as intronic, intergenic, sense, antisense, or bidirectional RNAs based on their genomic location, function, and structure (22). This diverse range of genomic locations plays a crucial role in the functional variety and regulatory functions of lncRNAs, which could be present in both the cytoplasm and the nucleus, enabling them to exert their influence and fulfill their functions effectively (23). LncRNAs can also be classified according to their mechanistic modes as signals, decoys, guides, and scaffolds (24). As signals, these RNAs interact with transcription factors in a spatiotemporal manner to regulate gene expression (25). In their decoy role, lncRNAs bind to transcription factors and other proteins, effectively sequestering them from chromatin or directing them into nuclear subdomains (26). Additionally, lncRNAs function as guides by associating with RNA-binding proteins (RBPs) to facilitate their binding to target genes (26). Finally, lncRNAs serve as flexible scaffolds that can accommodate various macromolecules, enable complex formation, and carry out a range of biological functions (27). LncRNAs regulate post-transcriptional gene expression levels in the cytoplasm by stabilizing mRNAs, promoting or inhibiting the translation of target mRNAs through acting as precursors to mRNAs or functioning as competing endogenous RNAs (ceRNAs) by sponging specific target microRNAs (miRNAs) and facilitating mRNA decay (28). In addition, lncRNAs are involved in alternative splicing, the formation of subcellular compartments, and the epigenetic regulation of specific genes (29). Moreover, lncRNAs are found to participate in regulating numerous signaling pathways and targeting multiple downstream target genes (7, 30) (Figure 1).
Numerous anticancer agents have been identified to trigger apoptosis and engage apoptosis-related signaling pathways (31). Nevertheless, the dysregulation of apoptotic processes frequently contributes to the development of drug resistance and the failure of treatment (31). The regulation of the mitochondrial apoptosis pathway relies on a delicate balance between pro-apoptotic and anti-apoptotic proteins. Disruption of this balance can lead to drug resistance in GC (32, 33). LncRNA plasmacytoma variant translocation 1 (PVT1) induced the expression of anti-apoptotic protein Bcl-2, thus inhibiting cell apoptosis and thereby enhancing the resistance of GC cells to 5-fluorouracil (5-FU) (34). Fang et al. discovered that lncRNA UCA1 sponged miR-27b to promote adriamycin and cisplatin resistance by increase Bcl-2 expression and decreased expression of caspase-3 (35). Small nucleolar RNA host gene (SNHG) encoded a snoRNA and generated a lncRNA, which can regulate gene expression (36). For example, SNHG5 downregulated Bax and upregulated Bcl-2, thereby inhibiting apoptosis and facilitating cisplatin resistance (36).
Caspase family plays an essential role in cell apoptosis and drug resistance. For example, apoptotic protease-activating factor 1 (APAF1)-binding lncRNA (ABL) significantly interacted with the insulin-like growth factor 2 mRNA-binding protein 1 (IGF2BP1), which facilitated the recognition of m6A modifications on ABL, promoting its stability. Additionally, ABL bound to APAF1, consequently blocking apoptosome formation and decreasing the expression caspases-9 and -3. These changes led to multidrug resistance (MDR) in GC (33). Furthermore, RP11-874J12.4 functioned as a molecular sponge for miR-397, which further enhanced the expression of signal sequence receptor subunit 2 (SSR2) that upregulated Bcl-2 expression and downregulated expression of cleaved caspase-3, caspase-9, and Bax, leading to resistance to chemotherapeutic drugs in GC (37). LncRNA FAM84B antisense RNA (FAM84B-AS) augmented cisplatin resistance by suppressing apoptosis via enhancing the expression of BCL-2 and BCL-xL and downregulating the expression of caspase-3, -7, and -9 (38) (Figure 2).
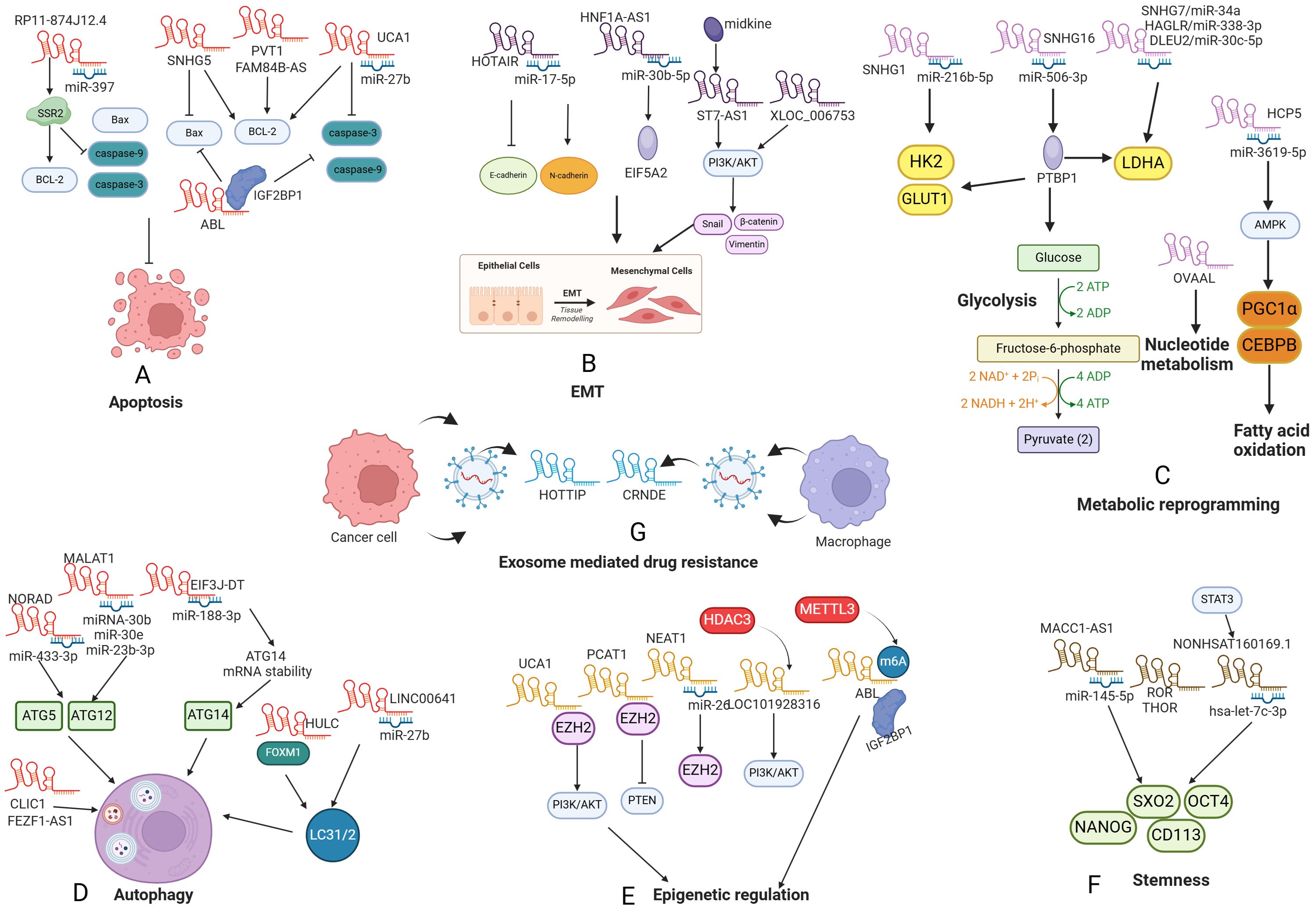
Figure 2. The role of lncRNAs in the drug resistance of gastric cancer. (A) LncRNAs promote drug resistance by regulating cell apoptosis; (B) LncRNAs promote drug resistance by inducing EMT; (C) LncRNAs promote drug resistance by metabolic reprogramming; (D) LncRNAs promote drug resistance by promoting cell autophagy; (E) LncRNAs promote drug resistance by epigenetic regulation; (F) LncRNAs promote drug resistance by promoting cancer stemness; (G) Exosomal lncRNAs promote drug resistance.
Epithelial-mesenchymal transition (EMT) refers to a biological process whereby epithelial cells undergo a phenotypic transformation to acquire mesenchymal characteristics (39, 40). In the context of cancer, EMT is linked to critical metastatic processes such as tumor metastasis and therapeutic resistance (41, 42). Emerging evidence has revealed that lncRNAs regulate GC drug resistance through EMT pathway (43, 44). LncRNAs play a crucial role in mediating EMT-associated drug resistance by modulating EMT markers and transcription factors. LncRNA HOTAIR directly sponged miR-17-5p, leading to downregulation of E-cadherin and upregulation of N-cadherin and Vimentin, thus facilitating both EMT and resistance to multiple drugs including: cisplatin, doxorubicin and 5-FU (45). LncRNA HNF1A-AS1 was reported to be highly expressed in GC tissues, which facilitated cell resistance to 5-FU by promoting miR-30b-5p/EIF5A2 axis-mediated EMT (43). Yang et al. revealed that cancer-associated fibroblast (CAF)-derived midkine (MK) significantly upregulated lncRNA ST7-AS1 in the context of GC, which involved in the cisplatin resistance. Knockdown of ST7-AS1 reversed cisplatin resistance via inhibiting phosphoinositide 3-kinase (PI3K)/AKT pathway and suppressing EMT (44). In line with this, LncRNA XLOC_006753 was found to be upregulated in MDR GC cell lines. The knockdown of XLOC_006753 resulted in decreased levels of Snail, β-catenin, and Vimentin, which ultimately reversed the EMT and enhanced the sensitivity of gastric cancer cells to cisplatin and 5-FU by inactivating PI3K/AKT pathway (46).
Cancer cells undergo metabolic reprogramming to support their survival and cancer progression. This involves heightened glycolytic dependency characterized by increased glucose uptake and fermentation to lactate, satisfying their anabolic requirements for proliferation (47). This phenomenon, known as the Warburg effect, has been reported to induce drug resistance in cancers (48). In the context of GC, emerging evidence has revealed that dysregulated lncRNAs might contribute to drug resistance via regulating glycolysis. Research performed by Xu and colleagues has demonstrated that lncRNA SNHG1 sponged miR-216b-5p and enhanced expression levels of hexokinase 2 (HK2), which functions as a critical enzyme in glycolysis. These changes conferred GC cell resistance to paclitaxel, which provided novel targets for treating chemoresistant patients (49). Furthermore, lncRNA SNHG16 and polypyrimidine Tract Binding Protein 1 (PTBP1) were upregulated in GC cell lines. Mechanistic studies revealed that SNHG16 severed as a ceRNA for miR-506-3p to target PTBP1, which resulting in enhanced mRNA stability of multiple glycolysis enzymes including Glucose Transporter Type 1 (GLUT1), HK2, and lactate dehydrogenase A (LDHA), thus promoted 5-FU resistance, which could be reversed by silencing SNHG16 in vivo (50). Bioinformatics analysis revealed that miR-34a has a potential binding site on SNHG7, and this negative relationship was validated in cisplatin-resistant GC tissues. Moreover, LDHA levels could be enhanced by lncRNA SNHG7/miR-34a axis and knockdown of SNHG7 could sensitized cisplatin-resistant GC cells by inhibiting LDHA, suggesting the SNHG7-miR-34a/LDHA-glycolysis axis contributes to cisplatin resistance (51). LncRNA HAGLR was highly expressed in GC cell lines and tissues. It sponged miR-338-3p to target LDHA-mediated glycolysis, thus facilitated 5-FU resistance in the context of GC (52). DLEU2, was another oncogenic lncRNA that targeted LDHA-mediated glycolysis to promote paclitaxel resistance by acting as ceRNA for miR-30c-5p (53). LncRNA HCP5 enhanced fatty acid oxidation via sequestering miR-3619-5p and modulating AMPK/PGC1α/CEBPB pathway, thereby facilitating chemoresistance in GC (54). Recently, Tian et al. reported that lncRNA OVAAL promoted resistance to 5-FU by enhancing pyrimidine biosynthesis, thereby counteracting the thymidylate synthase dysfunction caused by 5-FU (55), indicating that targeting OVAAL-mediated nucleotide metabolic reprogramming presents a promising strategy to overcome 5-FU resistance in GC.
Autophagy is a conserved cellular degradation process that encapsulates portions of the cytosol and damaged organelles within double-membrane vesicles called autophagosomes (56). Numerous studies have demonstrated that increased levels of autophagy not only improve tumor survival but also enhance drug resistance across various tumor types (56). IL-6 activated autophagy via the IL-6/JAK2 pathway and contributed to chemotherapy resistance (57). In GC, chloride channel 1 (CLIC1) was found to promote the activation of autophagy, thus reducing cellular sensitivity to cisplatin (58). Furthermore, dysregulated lncRNAs were found to induce MDR via targeting autophagy. For example, lncRNA EIF3J-DT was significantly overexpressed in drug-resistant GC cells. EIF3J-DT induced ATG14 expression through directly stabilizing ATG14 mRNA. Additionally, EIF3J-DT prevented the degradation of ATG14 mRNA by sponging miR-188-3p, thus promoting chemotherapeutic resistance (18). Hu et al. demonstrated that lncRNA metastasis-associated lung adenocarcinoma transcript-1 (MALAT1) functioned as a molecular sponge for miR-23b-3p, diminishing its inhibitory effects on the expression of ATG12, which led to the development of autophagy-mediated -MDR in cancer cells to cisplatin and vincristine (59). Researchers have demonstrated that MALAT1 sponged miRNA-30b and miR-30e to target ATG5, thus contributing to cisplatin resistance (60, 61). Furthermore, FEZF1-AS1 modulated MDR of GC cells by regulating ATG5 (62). LncRNA NORAD was an oxidative stress-induced lncRNA that facilitated oxaliplatin resistance by regulating miR-433-3p-ATG5/ATG12 pathway (63), which uncovered a complex interaction between cellular stress and lncRNAs in autophagy-mediated drug resistance.
Additionally, research by Xin et al. has shown that lncRNA HULC interacted with forkhead box M1 (FOXM1), stabilizing this protein, which enhances the B-light chain 3 (LC3)-II/LC3-I ratio and contributes to cisplatin resistance through autophagy. Consistent with these findings, the silencing of HULC has been demonstrated to inhibit autophagy and improve the sensitivity of GC cells to chemotherapy (64). LINC00641 was found to sponge miR-582-5p, activating autophagy flux by evaluating expression of microtubule-associated protein 1A/LC3 I/II and p62, thus conferring cancer cells resistance to oxaliplatin. And silencing LINC00641 could increase the sensitivity of GC cells to oxaliplatin (65).
Epigenetic modifications refer to influence gene expression without changing the underlying DNA sequences, which have been extensively explored in tumor progression and drug resistance (66). Enhancer of zeste homolog 2 (EZH2) is responsible for the trimethylation of lysine 27 on histone H3 (H3K27me3), serving an essential role in a Polycomb Repressive Complex 2 (PRC2)-dependent manner, which represses expression of multiple tumor suppressive genes through the H3K27me3 mediated by EZH2 (67). EZH2 was involved in drug resistance across various types of human cancers (68). In the context of GC, lncRNAs induced drug resistance by targeting EZH2. For example, Dai et al. revealed that lncRNA UCA1 was upregulated in GC cells and tissues. Furthermore, UCA1 could recruit EZH2 and promoted PI3K/AKT signaling pathway, which led to cisplatin resistance (69). LncRNA prostate cancer-associated transcript 1 (PCAT1) was upregulated in cisplatin-resistant GC tissues, exerting tumor-promoting effects by interacting with EZH2 and thus epigenetically silencing phosphatase and tensin homolog deleted on chromosome ten (PTEN), leading to enhanced H3K27 trimethylation (70). These changes resulted in GC cell resistance to cisplatin (70). LncRNA NEAT1 sequestered miR-26, which diminished the inhibitory effects on EZH2, which led to oxaliplatin resistance (71). Histone deacetylases (HDACs) plays essential roles in maintaining balance between epigenetic modifications, thus exerting greatly impacts on chemotherapeutic resistance (72). HDAC3 was upregulated in GC cisplatin-resistant cells, which further promoted transcription of lncRNA LOC101928316, leading to the activation of PI3K/Akt/mTOR signaling pathway and enhanced cisplatin resistance (73). m6A methylation is the most prevalent modification found in both mRNA and non-coding RNAs, exerting significant effects on RNA stability, splicing, localization, and translation (74). In MDR-GC cell lines, METTL3 has been shown to elevate the m6A levels of ABL, which interacted with IGF2BP1, a member of the m6A reader family, thus preventing APAF1 from forming apoptotic bodies. This mechanism inhibited apoptosis and contributes to drug resistance (33). These results revealed that lncRNA-mediated epigenetic regulations provide feasible therapeutic targets for overcoming drug resistance in GC.
Cancer stemness is considered a crucial element in tumor development. Cancer stem cells (CSCs) represent a distinct population of cancer cells characterized by their capacity for self-renewal and differentiation, contributing to tumor development and resistance to therapy (75). The stemness markers of CSCs primarily encompass cluster of differentiation 24 (CD24), CD133, SOX2(SRY-box transcription factor 2), SOX9 and c-Myc (76–78). Emerging evidence has revealed that lncRNAs regulating drug resistance via targeting stemness-related markers or pathways. For example, Wang and colleagues reported lncRNA ROR exhibited increased expression in the gastric cancer stem cells (GCSCs), which upregulated multiple stemness-related transcription factors including NANOG, SOX2, CD133 and OCT4. Furthermore, ROR promoted cell resistance to adriamycin and vincristine (79, 80). Zhao et al. revealed that lncRNA NONHSAT160169.1 was upregulated in lapatinib resistant GC cells. Further investigation into the mechanisms by which NONHSAT160169.1 conferred lapatinib resistance revealed that it was induced by the signal transducer and activator of transcription 3 (STAT3) pathway. NONHSAT160169.1 sponged hsa-let-7c-3p and thereby negating its suppressive impacts on the expression of SRY-box transcription factor 2 (SOX2), which suggested a NONHSAT160169.1/hsa-let-7c-3p/SOX2 axis in understanding lapatinib resistance in GC (81). LncRNA THOR mediated the mRNA stability of SOX9, thus enhancing stemness and cisplatin resistance in GC (82). LncRNA MACC1 antisense RNA 1 (MACC1-AS1) acted as a competitive antagonist to miR-145-5p, leading to the upregulation of diacylglycerol cholinephosphotransferase (CPT1) and acetyl-CoA synthetase (ACS), which enhanced expression of stemness markers such as LIN28, SOX2 and OCT4, which contributed to the 5-FU and oxaliplatin resistance (83).
Exosome plays a vital role in enabling cells to adapt to various internal and external changes involved in processes including injury response and tissue homeostasis. Exosome transfer biological mediators including lncRNA, miRNA, and proteins from donor cells to receipt cells, representing a specific and tightly modulatory communication, which significantly impacts on cancer progression (84). LncRNA CRNDE exhibited highly expression in tumor-associated macrophages (TAMs) in the context of GC. Further mechanistic studies have demonstrated that M2-polarized TAMs secreted exosomes, which transferred CRNDE from TAMs to tumor cells. This significantly enhanced the ubiquitination of PTEN through neural precursor cell expressed developmentally downregulated protein 4-1 (NEDD4-1)-mediated signaling pathways, leading to cisplatin resistance of GC cells (85). Exosomes derived from GC cells packaged lncRNA HOTTIP, which mediated high-mobility group A1 (HMGA1)/miR-218 axis to facilitate cisplatin resistance (86) (Table 1).
The disruption and function of lncRNAs are progressively being incorporated into cancer treatment strategies, offering benefits in both laboratory and clinical settings (87). The application of RNA-based therapies has emerged as a promising strategy for treating cancers (87). Certain endogenous lncRNAs can modulate the expression of genes associated with tumorigenesis, and their dysregulation may contribute to disease onset, underscoring the potential of these ncRNAs as targets for drug development (87, 88). In GC, lncRNAs can function as either oncogenes or tumor suppressors, leading to the abnormal inhibition or degradation of their target mRNAs. Consequently, they are considered critical therapeutic targets for cancer treatment. In this context, the application of lncRNA-based therapies presents advantages. These emerging strategies included the use of small interfering RNAs (siRNAs), antisense oligonucleotides (ASOs), and clustered regularly interspaced short palindromic repeats (CRISPR)-CRISPR-associated protein (Cas) 9 gene editing (89–91) (Figure 3).
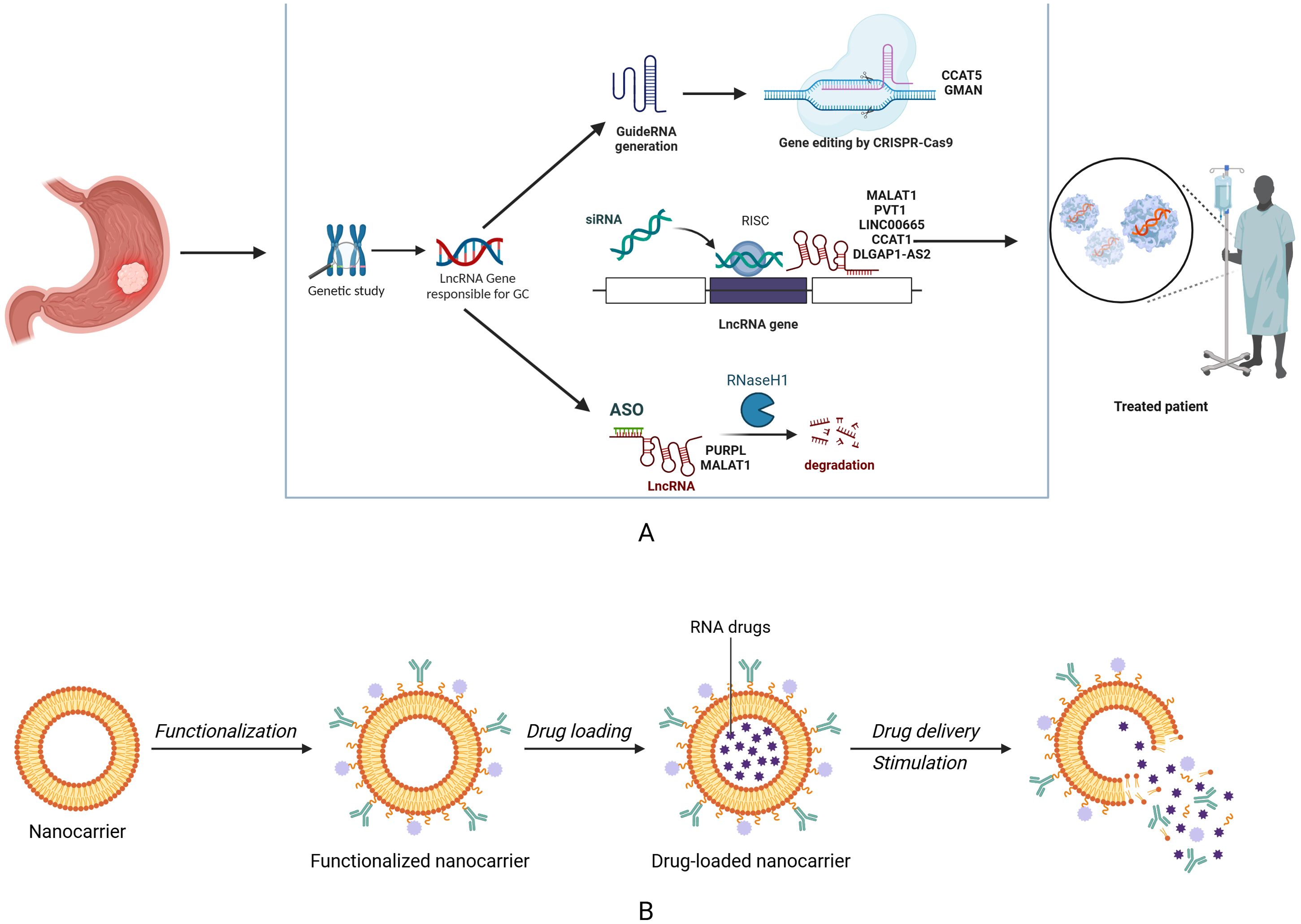
Figure 3. Therapeutic strategies for targeting lncRNAs in cancer therapy. (A) CRISPR/Cas 9 technique, antisense oligonucleotides (ASOs), and small interfering RNAs (siRNAs) are capable of degrading lncRNAs, thus providing novel insights into gastric cancer treatment. (B) Nanomedicine functions as essential nanocarrier for specifically delivering RNA-based drugs in gastric cancer treatment.
SiRNAs are a type of therapeutic agent derived from nucleic acids, functioning as drugs target cytoplasmic RNAs or to induce transcriptional silencing through histone modification and remodeling chromatin within the nucleus by binding to promoter regions (92). This potential has generated considerable excitement in the field of gene therapy. For example, MALAT1 was implicated in GC progression and drug resistance, functioning as a potential target for GC treatment. Experimental assays demonstrated that siRNA-mediated silencing of MALAT1 effectively inhibited the migration and invasion of GC cells (93). Exosomal MALAT1 derived from TAMs was found to enhance chemotherapy resistance. MALAT1 could be transferred from TAMs to cancer cells, while the dual silencing of MALAT1 in tumor cells and TAMs by transferring siRNAs via exosomes significantly enhanced chemosensitivity in the setting of GC (94). Bahar et al. demonstrated that knockdown of PVT1 by specific siRNA could reverse paclitaxel resistance and suppress tumor growth, which suggested a promising approach for GC treatment (95).
Oligonucleotide technology has been advanced to selectively modify the expression of critical genes involved in terminal illnesses, including human cancers (96). ASOs are chemically modified single-stranded nucleic acid sequences, which could bind complementarily to RNA sequences, thus modulating the functions of mRNA (96). ASOs facilitate the degradation of lncRNAs through the action of RNase H, thereby enabling the silencing and regulation of lncRNAs (97). These characteristics render ASOs a valuable asset in modern medicine, offering high target specificity for treating diseases (98). LncRNA p53-upregulated-regulator-of-p53-levels (PURPL) contributed to chemotherapy resistance, while ASO-based knockdown significantly reduced the expression of PURPL, leading to induced cell apoptosis and sensitized cancer cell to chemotherapeutic agents (99). Furthermore, LNA gapmeR, a specially designed ASO, has found extensive application in preclinical studies. This LNA gapmeR ASO directly induces the degradation of MALAT1, demonstrating remarkable antitumor activity and eliciting cytotoxic effects in a mouse models (100). ASOs that targeted the lncRNA MALAT1 in triple-negative breast cancer has been demonstrated to alleviate malignant characteristics by reshaping tumor microenvironment (101). And ASOs-based silencing of MALAT1 led to reduced cancer metastasis and reversed therapeutic resistance in various types of human cancers (102–104). Given the oncogenic role of MALAT1 in the progression and drug resistance in GC, the suppression of MALAT1 by ASOs hold immense therapeutic potential for GC treatment, which await further investigation both in the preclinical and clinical settings in the context of GC (105–107).
Recent advancements in the CRISPR-Cas9 technique have shown promising results in the precise modification of genes, potentially offering new avenues for cancer treatment (108). Studies employing CRISPR-Cas9 technology have shown encouraging, safe, and effective cancer therapies, marking a significant step forward in the field of precision oncology (109). Emerging preclinical studies have utilized CRISPR- Cas 9 system to target oncogenic lncRNAs, thus inhibiting tumor progression. The knockdowns of RP11-93B14.5, PANDAR, and NEAT1 lncRNAs have demonstrated great effects on the inhibition of cell proliferation, growth, metastasis and therapeutic responses in GC (110–112). In the context of GC, Liu et al. revealed that CRISPR/Cas9-knocknout system-mediated silences of lncRNA CCAT5 significantly impeded tumor growth in vivo (113). Moreover, a strategy utilizing CRISPR/Cas9 system to target the lncRNA GMAN led to a significant decrease in the number of metastases generated from GC cells in vivo (114). These findings highlight the significant potential of CRISPR-Cas9 technique for translating lncRNA research into clinical applications, which may pave the way for GC treatment.
Synthetic nanocarriers have long been employed for drug delivery purposes. They have the capacity to encapsulate both hydrophobic and hydrophilic drugs, and their formulations can be modified to enhance stability (115). Emerging evidence has revealed that nanocarriers serve as feasible platforms that can deliver lncRNAs to tumors, potentially enhancing therapeutic results for cancer patients (115). Wang and colleagues reported that lncRNA ABL promoted MDR and tumor growth in GC, and encapsulated liposomal siRNA targeting ABL could markedly increase the sensitivity of GC cells to chemotherapeutic agents (33). The nanocarrier chitosan-gelatin-EGCG (CGE) has been reported to delivery siRNA that targeted TMEM44‐AS1, which significantly abrogated 5-FU resistance in GC (116). The therapeutic delivery of lncRNA LINC00589 via polyethyleneimine-modified mesoporous silica nanoparticles demonstrated significant efficacy in attenuating peritoneal metastasis in GC experimental models (117), offers a promising strategy for enhancing therapeutic efficacy against various malignancies. The capability to modify polymeric nanoparticles facilitates accurate customization of lncRNA delivery systems. However, the intricate nature of these structures presents challenges for maintaining consistent production. Accordingly, progress in polymerization techniques is crucial for enhancing the therapeutic effectiveness of lncRNA-based cancer therapies.
Existing evidence has revealed that lncRNAs play an essential role in GC drug resistance. Bahar et al. reported that siPVT1 significantly enhanced chemosensitivity to paclitaxel in GC cells (95). Moreover, using siRNA silencing CCAT1 could reversed drug resistance in cisplatin-resistant GC cells by affecting the PI3K/AKT pathway (118). Cells subjected to LINC00665-shRNA treatment exhibited a marked decrease in trastuzumab responsiveness relative to other experimental cohorts (119). The combined application of siRNA-mediated DLGAP1-AS2 knockdown and oxaliplatin significantly enhanced the chemosensitivity of GC cells, reducing the effective dose of oxaliplatin, which markedly induced apoptosis while simultaneously suppressing cell proliferation, and metastatic potential compared to monotherapy (120). Additionally, several studies have revealed that targeting lncRNA NEAT1 (121), NONHSAT160169.1 (81), FUAT1 (122), HOTAIR (123) also reversed drug resistance in GC, providing promising therapeutic strategies in cancer therapy.
Nucleic acid-based therapies have received considerable attention as treatments for a range of diseases, including cancer; nevertheless, caution is necessary in their application. The implementation of nucleic acid-based therapies in vivo studies encounters several challenges. Inefficient delivery methods and the low bioavailability of siRNA observed in animal models limit their therapeutic efficacy (124). In addition, ASOs can become trapped within endosomes, which significantly diminishes their bioavailability (125). As a result, it is vital to establish that oligonucleotides have minimal or absent off-target effects (126). Moreover, when utilizing CRISPR/Cas9 technology, it is important to carefully evaluate potential off-target effects, as they could result in unintended consequences (127). Enhancing the stability of these therapies, prolonging their pharmacokinetics, and improving overall drug stability are therefore critical considerations (128). However, given the multiple roles of lncRNAs in various biological activities, thus modulating the expression of specific lncRNAs may trigger a cascade of reactions, potentially resulting in additional side effects during drug application, which await further investigations in clinical settings.
Although nanoparticle-enhanced delivery of lncRNAs holds significant promise in the field of oncology, there are still considerable challenges related to the scalable and cost-effective manufacturing of these nanoscale systems (129). The complexities involved in achieving desired characteristics such as biocompatibility, stability, and targeted specificity necessitate a meticulous design process (130). The intricate regulatory networks involving lncRNAs require additional research to clarify their specific roles and interactions with other molecular pathways. Furthermore, while specific inhibitors targeting lncRNAs have shown promising anti-cancer effects in preclinical studies, their safety and effectiveness in human clinical trials are still uncertain (131). This highlights the need for enhanced targeting strategies and delivery systems to ensure accurate and effective modulation of lncRNAs. In addition, lncRNA-based therapeutic approaches are faced by tolerance issues, particularly immunogenicity, arising from pathogen-associated molecular pattern receptors that recognize RNA structures, which can trigger immune responses (132, 133). Further research should focus on addressing these challenges, which may contribute to the further exploration of cancer treatment strategies.
LncRNAs represent a class of functional RNA transcripts present in human genomes. Mounting evidence has highlighted the diverse functions and mechanisms through which many lncRNAs operate, increasingly associating them with the onset and progression of cancers. Nonetheless, the exploration of many lncRNAs remains limited. In the context of GC, lncRNAs exert regulatory functions on chromatin remodeling, histone modification, sponging miRNAs, stabilizing mRNA and regulating translation of various proteins, thus significantly impacting cancer progression and drug resistance. Moreover, lncRNAs serve as essential biomarkers for early detection and monitoring of GC. LINC01133 was reported to be decreased in the serum of GC patients. Furthermore, expression of LINC01133 was associated with tumor markers such as CEA and CA19-9 along with various clinicopathological parameters, including tumor size, the tumor-node-metastasis (TNM) stage, and distant metastasis (134), indicating that LINC01133 was an independent prognostic factor for the disease. Research by Jiang et al. has revealed that lncRNA FAM87A served as an independent biomarker for the overall survival of GC patients (135). Further research performed by Liang et al. indicated that the expression levels of lncRNA XIST and ZFPM2-AS1 were significantly elevated in the plasma of GC patients. Additionally, the area under the receiver operating characteristic curve (AUC) was calculated to be 0.62 for ZFPM2-AS1 and 0.68 for XIST, which suggested that these lncRNAs could be used as candidate plasma biomarkers for GC patients (136). Overall, these findings indicate that lncRNAs are associated with various clinical characteristics of GC and could sever as essential biomarkers for early detection and diagnosis of GC. However, a common limitation in these studies is the relatively small sample size. To identify a lncRNA as a potential new biomarker, it is essential to have a sufficiently large study population that can reliably support the candidature of lncRNAs for biomarker development.
Moreover, lncRNAs play an essential role in regulate drug resistance, which includes multiple mechanisms including regulating cell apoptosis, EMT, metabolic reprogramming, autophagy, stemness, and mediating epigenetic regulation. Therefore, targeting lncRNAs may be feasible strategies for overcoming drug resistance and paving the way for GC treatment. Notably, strategies such as siRNAs, ASOs, and CRISPR/Cas 9 technique significantly exhibit remarkable tumor regression effects in the preclinical setting. But there is lacking of investigations on the anti-cancer effects in the clinical settings. Furthermore, concerns have been raised regarding the therapeutic potential of targeting individual lncRNAs, as well as the effectiveness of current targeting strategies, which underscores the necessity for improved targeting strategies and delivery systems to facilitate precise and effective modulation of lncRNAs. Future research should focus on these challenges and promote the translation of current findings into clinical applications, which may improve the prognosis of chemotherapy-resistant GC patients.
WL: Conceptualization, Data curation, Formal Analysis, Investigation, Methodology, Project administration, Resources, Software, Validation, Visualization, Writing – original draft, Writing – review & editing. WW: Data curation, Formal Analysis, Investigation, Resources, Supervision, Validation, Visualization, Writing – review & editing.
The author(s) declare that no financial support was received for the research and/or publication of this article.
We thank the BioRender drawing software (https://www.biorender.com/).
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
1. Bray F, Laversanne M, Sung H, Ferlay J, Siegel RL, Soerjomataram I, et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. (2024) 74(3):229–63. doi: 10.3322/caac.21834
2. Thrift AP, Wenker TN, El-Serag HB. Global burden of gastric cancer: epidemiological trends, risk factors, screening and prevention. Nat Rev Clin Oncol. (2023) 20(5):338–49. doi: 10.1038/s41571-023-00747-0
3. Yang WJ, Zhao HP, Yu Y, Wang JH, Guo L, Liu JY, et al. Updates on global epidemiology, risk and prognostic factors of gastric cancer. World J Gastroenterol. (2023) 29(16):2452–68. doi: 10.3748/wjg.v29.i16.2452
4. Yeni M, Kalayci T. Splenectomy during gastric cancer surgery versus splenectomy during extra-gastric abdominal cancer surgery. J Coll Physicians Surg Pak. (2022) 32(4):514–8. doi: 10.29271/jcpsp.2022.04.514
5. Sexton RE, Al Hallak MN, Diab M, Azmi AS. Gastric cancer: a comprehensive review of current and future treatment strategies. Cancer Metastasis Rev. (2020) 39(4):1179–203. doi: 10.1007/s10555-020-09925-3
6. Vasan N, Baselga J, Hyman DM. A view on drug resistance in cancer. Nature. (2019) 575(7782):299–309. doi: 10.1038/s41586-019-1730-1
7. Huang P, Wen F, Li Q. Current concepts of the crosstalk between lncRNA and E2F1: shedding light on the cancer therapy. Front Pharmacol. (2024) 15:1432490. doi: 10.3389/fphar.2024.1432490
8. Nojima T, Proudfoot NJ. Mechanisms of lncRNA biogenesis as revealed by nascent transcriptomics. Nat Rev Mol Cell Biol. (2022) 23(6):389–406. doi: 10.1038/s41580-021-00447-6
9. Yu J, Wang W, Yang J, Zhang Y, Gong X, Luo H, et al. LncRNA PSR regulates vascular remodeling through encoding a novel protein arteridin. Circ Res. (2022) 131(9):768–87. doi: 10.1161/circresaha.122.321080
10. Liang Q, Zhang W, Wu H, Liu B. LncRNA-disease association identification using graph auto-encoder and learning to rank. Brief Bioinform. (2023) 24(1). doi: 10.1093/bib/bbac539
11. Yan J, Wang R, Tan J. Recent advances in predicting lncRNA-disease associations based on computational methods. Drug Discovery Today. (2023) 28(2):103432. doi: 10.1016/j.drudis.2022.103432
12. Hsu CY, Jamal A, Kamal MA, Ahmad F, Bokov DO, Mustafa YF, et al. Pathological roles of lncRNA HOTAIR in liver cancer: An updated review. Gene. (2024) 2024:149180. doi: 10.1016/j.gene.2024.149180
13. Hu J, Li D, Liu C. LncRNAs chaperoning dynamic protein condensates in cancer cells. Mol Cell. (2024) 84(24):4695–7. doi: 10.1016/j.molcel.2024.11.028
14. Jiang J, Lin F, Wu W, Zhang Z, Zhang C, Qin D, et al. Exosomal long non-coding RNAs in lung cancer: A review. Med (Baltimore). (2024) 103(51):e38492. doi: 10.1097/md.0000000000038492
15. Yuan J, Lu J, Zhu J, Chen F, Zeng Z, Yan J, et al. LncRNA FIRRE drives gastric cancer progression via ZFP64-mediated TUBB3 promoter activation. Cancer Lett. (2024) 2024:217398. doi: 10.1016/j.canlet.2024.217398
16. Shiraishi T, Matsumoto A. From non-coding to coding: The importance of long non-coding RNA translation in de novo gene birth. Biochim Biophys Acta Gen Subj. (2024) 2024:130747. doi: 10.1016/j.bbagen.2024.130747
17. Fang L, Huang H, Lv J, Chen Z, Lu C, Jiang T, et al. m5C-methylated lncRNA NR_033928 promotes gastric cancer proliferation by stabilizing GLS mRNA to promote glutamine metabolism reprogramming. Cell Death Dis. (2023) 14(8):520. doi: 10.1038/s41419-023-06049-8
18. Luo Y, Zheng S, Wu Q, Wu J, Zhou R, Wang C, et al. (lncRNA) EIF3J-DT induces chemoresistance of gastric cancer via autophagy activation. Autophagy. (2021) 17(12):4083–101. doi: 10.1080/15548627.2021.1901204
19. Wang Y, Shen K, Cheng Q, Zhou X, Liu K, Xiao J, et al. The long noncoding RNA ELFN1-AS1 promotes gastric cancer growth and metastasis by interacting with TAOK1 to inhibit the hippo signaling pathway. Cell Death Discovery. (2024) 10(1):465. doi: 10.1038/s41420-024-02235-5
20. Qian X, Zhao J, Yeung PY, Zhang QC, Kwok CK. Revealing lncRNA structures and interactions by sequencing-based approaches. Trends Biochem Sci. (2019) 44(1):33–52. doi: 10.1016/j.tibs.2018.09.012
21. Núñez-Martínez HN, Recillas-Targa F. Emerging functions of lncRNA loci beyond the transcript itself. Int J Mol Sci. (2022) 23(11). doi: 10.3390/ijms23116258
22. Bridges MC, Daulagala AC, Kourtidis A. LNCcation: lncRNA localization and function. J Cell Biol. (2021) 220(2). doi: 10.1083/jcb.202009045
23. Peng WX, Koirala P, Mo YY. LncRNA-mediated regulation of cell signaling in cancer. Oncogene. (2017) 36(41):5661–7. doi: 10.1038/onc.2017.184
24. McCabe EM, Rasmussen TP. lncRNA involvement in cancer stem cell function and epithelial-mesenchymal transitions. Semin Cancer Biol. (2021) 75:38–48. doi: 10.1016/j.semcancer.2020.12.012
25. Kopp F, Mendell JT. Functional classification and experimental dissection of long noncoding RNAs. Cell. (2018) 172(3):393–407. doi: 10.1016/j.cell.2018.01.011
26. Yao ZT, Yang YM, Sun MM, He Y, Liao L, Chen KS, et al. New insights into the interplay between long non-coding RNAs and RNA-binding proteins in cancer. Cancer Commun (Lond). (2022) 42(2):117–40. doi: 10.1002/cac2.12254
27. Taiana E, Ronchetti D, Todoerti K, Nobili L, Tassone P, Amodio N, et al. LncRNA NEAT1 in paraspeckles: A structural scaffold for cellular DNA damage response systems? Noncoding RNA. (2020) 6(3). doi: 10.3390/ncrna6030026
28. Xu J, Xu J, Liu X, Jiang J. The role of lncRNA-mediated ceRNA regulatory networks in pancreatic cancer. Cell Death Discovery. (2022) 8(1):287. doi: 10.1038/s41420-022-01061-x
29. Liu Y, Liu X, Lin C, Jia X, Zhu H, Song J, et al. Noncoding RNAs regulate alternative splicing in cancer. J Exp Clin Cancer Res. (2021) 40(1):11. doi: 10.1186/s13046-020-01798-2
30. Mazarei M, Shahabi Rabori V, Ghasemi N, Salehi M, Rayatpisheh N, Jahangiri N, et al. LncRNA MALAT1 signaling pathway and clinical applications in overcome on cancers metastasis. Clin Exp Med. (2023) 23(8):4457–72. doi: 10.1007/s10238-023-01179-x
31. Carneiro BA, El-Deiry WS. Targeting apoptosis in cancer therapy. Nat Rev Clin Oncol. (2020) 17(7):395–417. doi: 10.1038/s41571-020-0341-y
32. Kong F, Wu K, Pang L, Huang Y, Li L, Xu J, et al. Inhibition of apoptosis-regulatory protein siva-1 reverses multidrug resistance in gastric cancer by targeting PCBP1. Oncol Res. (2022) 30(6):277–88. doi: 10.32604/or.2022.027301
33. Wang Q, Chen C, Xu X, Shu C, Cao C, Wang Z, et al. APAF1-binding long noncoding RNA promotes tumor growth and multidrug resistance in gastric cancer by blocking apoptosome assembly. Adv Sci (Weinh). (2022) 9(28):e2201889. doi: 10.1002/advs.202201889
34. Du P, Hu C, Qin Y, Zhao J, Patel R, Fu Y, et al. LncRNA PVT1 mediates antiapoptosis and 5-fluorouracil resistance via increasing Bcl2 expression in gastric cancer. J Oncol. (2019) 2019:9325407. doi: 10.1155/2019/9325407
35. Fang Q, Chen X, Zhi X. Long non-coding RNA (LncRNA) urothelial carcinoma associated 1 (UCA1) increases multi-drug resistance of gastric cancer via downregulating miR-27b. Med Sci Monit. (2016) 22:3506–13. doi: 10.12659/msm.900688
36. Li M, Zhang YY, Shang J, Xu YD. LncRNA SNHG5 promotes cisplatin resistance in gastric cancer via inhibiting cell apoptosis. Eur Rev Med Pharmacol Sci. (2019) 23(10):4185–91. doi: 10.26355/eurrev_201905_17921
37. Liu Y, Cao J, Pu YS, Ma Y, Wu M, Wang JH. RP11-874J12.4, a novel lncRNA, confers chemoresistance in human gastric cancer cells by sponging miR-3972 and upregulating SSR2 expression. Am J Transl Res. (2021) 13(6):5892–910.
38. Zhang Y, Li Q, Yu S, Zhu C, Zhang Z, Cao H, et al. Long non-coding RNA FAM84B-AS promotes resistance of gastric cancer to platinum drugs through inhibition of FAM84B expression. Biochem Biophys Res Commun. (2019) 509(3):753–62. doi: 10.1016/j.bbrc.2018.12.177
39. Akhmetkaliyev A, Alibrahim N, Shafiee D, Tulchinsky E. EMT/MET plasticity in cancer and go-or-Grow decisions in quiescence: the two sides of the same coin? Mol Cancer. (2023) 22(1):90. doi: 10.1186/s12943-023-01793-z
40. Huang P, Zhu S, Liang X, Zhang Q, Liu C, Song L. Revisiting lung cancer metastasis: Insight from the functions of long non-coding RNAs. Technol Cancer Res Treat. (2021) 20:15330338211038488. doi: 10.1177/15330338211038488
41. Ebrahimi N, Manavi MS, Faghihkhorasani F, Fakhr SS, Baei FJ, Khorasani FF, et al. Harnessing function of EMT in cancer drug resistance: a metastasis regulator determines chemotherapy response. Cancer Metastasis Rev. (2024) 43(1):457–79. doi: 10.1007/s10555-023-10162-7
42. Xue W, Yang L, Chen C, Ashrafizadeh M, Tian Y, Sun R. Wnt/β-catenin-driven EMT regulation in human cancers. Cell Mol Life Sci. (2024) 81(1):79. doi: 10.1007/s00018-023-05099-7
43. Jiang L, Zhang Y, Su P, Ma Z, Ye X, Kang W, et al. Long non-coding RNA HNF1A-AS1 induces 5-FU resistance of gastric cancer through miR-30b-5p/EIF5A2 pathway. Transl Oncol. (2022) 18:101351. doi: 10.1016/j.tranon.2022.101351
44. Yang KD, Wang Y, Zhang F, Li QL, Luo BH, Feng DY, et al. CAF-derived midkine promotes EMT and cisplatin resistance by upregulating lncRNA ST7-AS1 in gastric cancer. Mol Cell Biochem. (2022) 477(11):2493–505. doi: 10.1007/s11010-022-04436-x
45. Jia J, Zhan D, Li J, Li Z, Li H, Qian J. The contrary functions of lncRNA HOTAIR/miR-17-5p/PTEN axis and shenqifuzheng injection on chemosensitivity of gastric cancer cells. J Cell Mol Med. (2019) 23(1):656–69. doi: 10.1111/jcmm.13970
46. Zeng L, Liao Q, Zou Z, Wen Y, Wang J, Liu C, et al. Long non-coding RNA XLOC_006753 promotes the development of multidrug resistance in gastric cancer cells through the PI3K/AKT/mTOR signaling pathway. Cell Physiol Biochem. (2018) 51(3):1221–36. doi: 10.1159/000495499
47. Huang P, Zhu S, Liang X, Zhang Q, Luo X, Liu C, et al. Regulatory mechanisms of LncRNAs in cancer glycolysis: Facts and perspectives. Cancer Manag Res. (2021) 13:5317–36. doi: 10.2147/cmar.S314502
48. Icard P, Shulman S, Farhat D, Steyaert JM, Alifano M, Lincet H. How the warburg effect supports aggressiveness and drug resistance of cancer cells? Drug Resist Update. (2018) 38:1–11. doi: 10.1016/j.drup.2018.03.001
49. Xu J, Xu Y, Ye G, Qiu J. LncRNA-SNHG1 promotes paclitaxel resistance of gastric cancer cells through modulating the miR-216b-5p-hexokianse 2 axis. J Chemother. (2023) 35(6):527–38. doi: 10.1080/1120009x.2022.2157618
50. Ding Y, Gao S, Zheng J, Chen X. Blocking lncRNA-SNHG16 sensitizes gastric cancer cells to 5-fu through targeting the miR-506-3p-PTBP1-mediated glucose metabolism. Cancer Metab. (2022) 10(1):20. doi: 10.1186/s40170-022-00293-w
51. Pei LJ, Sun PJ, Ma K, Guo YY, Wang LY, Liu FD. LncRNA-SNHG7 interferes with miR-34a to de-sensitize gastric cancer cells to cisplatin. Cancer biomark. (2021) 30(1):127–37. doi: 10.3233/cbm-201621
52. Hu J, Huang L, Ding Q, Lv J, Chen Z. Long noncoding RNA HAGLR sponges miR-338-3p to promote 5-fu resistance in gastric cancer through targeting the LDHA-glycolysis pathway. Cell Biol Int. (2022) 46(2):173–84. doi: 10.1002/cbin.11714
53. Xiang W, Zhang B, Li H. LncRNA DLEU2 contributes to taxol resistance of gastric cancer cells through regulating the miR-30c-5p-LDHA axis. J Chemother. (2024) 36(1):49–60. doi: 10.1080/1120009x.2023.2203606
54. Wu H, Liu B, Chen Z, Li G, Zhang Z. MSC-induced lncRNA HCP5 drove fatty acid oxidation through miR-3619-5p/AMPK/PGC1α/CEBPB axis to promote stemness and chemo-resistance of gastric cancer. Cell Death Dis. (2020) 11(4):233. doi: 10.1038/s41419-020-2426-z
55. Tan JN, Zhou SN, Zhang W, Yang B, Zhong GY, Huang J, et al. Long noncoding RNA OVAAL enhances nucleotide synthesis through pyruvate carboxylase to promote 5-fluorouracil resistance in gastric cancer. Cancer Sci. (2022) 113(9):3055–70. doi: 10.1111/cas.15453
56. Chang H, Zou Z. Targeting autophagy to overcome drug resistance: further developments. J Hematol Oncol. (2020) 13(1):159. doi: 10.1186/s13045-020-01000-2
57. Hu F, Song D, Yan Y, Huang C, Shen C, Lan J, et al. IL-6 regulates autophagy and chemotherapy resistance by promoting BECN1 phosphorylation. Nat Commun. (2021) 12(1):3651. doi: 10.1038/s41467-021-23923-1
58. Nong ZL, Zhao K, Wang Y, Yu Z, Wang CJ, Chen JQ. CLIC1-mediated autophagy confers resistance to DDP in gastric cancer. Anticancer Drugs. (2024) 35(1):1–11. doi: 10.1097/cad.0000000000001518
59. YiRen H, YingCong Y, Sunwu Y, Keqin L, Xiaochun T, Senrui C, et al. Long noncoding RNA MALAT1 regulates autophagy associated chemoresistance via miR-23b-3p sequestration in gastric cancer. Mol Cancer. (2017) 16(1):174. doi: 10.1186/s12943-017-0743-3
60. Xi Z, Si J, Nan J. LncRNA MALAT1 potentiates autophagy-associated cisplatin resistance by regulating the microRNA-30b/autophagy-related gene 5 axis in gastric cancer. Int J Oncol. (2019) 54(1):239–48. doi: 10.3892/ijo.2018.4609
61. Zhang YF, Li CS, Zhou Y, Lu XH. Propofol facilitates cisplatin sensitivity via lncRNA MALAT1/miR-30e/ATG5 axis through suppressing autophagy in gastric cancer. Life Sci. (2020) 244:117280. doi: 10.1016/j.lfs.2020.117280
62. Gui Z, Zhao Z, Sun Q, Shao G, Huang J, Zhao W, et al. LncRNA FEZF1-AS1 promotes multi-drug resistance of gastric cancer cells via upregulating ATG5. Front Cell Dev Biol. (2021) 9:749129. doi: 10.3389/fcell.2021.749129
63. Wang J, Sun Y, Zhang X, Cai H, Zhang C, Qu H, et al. Oxidative stress activates NORAD expression by H3K27ac and promotes oxaliplatin resistance in gastric cancer by enhancing autophagy flux via targeting the miR-433-3p. Cell Death Dis. (2021) 12(1):90. doi: 10.1038/s41419-020-03368-y
64. Xin L, Zhou Q, Yuan YW, Zhou LQ, Liu L, Li SH, et al. METase/lncRNA HULC/FoxM1 reduced cisplatin resistance in gastric cancer by suppressing autophagy. J Cancer Res Clin Oncol. (2019) 145(10):2507–17. doi: 10.1007/s00432-019-03015-w
65. Hu Y, Su Y, Lei X, Zhao H, Wang L, Xu T, et al. LINC00641/miR-582-5p mediate oxaliplatin resistance by activating autophagy in gastric adenocarcinoma. Sci Rep. (2020) 10(1):14981. doi: 10.1038/s41598-020-70913-2
66. Wang N, Ma T, Yu B. Targeting epigenetic regulators to overcome drug resistance in cancers. Signal Transduct Target Ther. (2023) 8(1):69. doi: 10.1038/s41392-023-01341-7
67. Kaur P, Shankar E, Gupta S. EZH2-mediated development of therapeutic resistance in cancer. Cancer Lett. (2024) 586:216706. doi: 10.1016/j.canlet.2024.216706
68. Duan R, Du W, Guo W. EZH2: a novel target for cancer treatment. J Hematol Oncol. (2020) 13(1):104. doi: 10.1186/s13045-020-00937-8
69. Dai Q, Zhang T, Pan J, Li C. LncRNA UCA1 promotes cisplatin resistance in gastric cancer via recruiting EZH2 and activating PI3K/AKT pathway. J Cancer. (2020) 11(13):3882–92. doi: 10.7150/jca.43446
70. Li H, Ma X, Yang D, Suo Z, Dai R, Liu C. PCAT-1 contributes to cisplatin resistance in gastric cancer through epigenetically silencing PTEN via recruiting EZH2. J Cell Biochem. (2020) 121(2):1353–61. doi: 10.1002/jcb.29370
71. Li Y, Peng C, Fang C, Huang K. Upregulation of nuclear-enriched abundant transcript 1 confers oxaliplatin resistance to gastric cancer. Cell Biol Int. (2020) 44(2):446–55. doi: 10.1002/cbin.11245
72. Parveen R, Harihar D, Chatterji BP. Recent histone deacetylase inhibitors in cancer therapy. Cancer. (2023) 129(21):3372–80. doi: 10.1002/cncr.34974
73. Ren H, Tang L. HDAC3-mediated lncRNA-LOC101928316 contributes to cisplatin resistance in gastric cancer via activating the PI3K-Akt-mTOR pathway. Neoplasma. (2021) 68(5):1043–51. doi: 10.4149/neo_2021_210317N356
74. Zhao Y, Li J, Dian M, Bie Y, Peng Z, Zhou Y, et al. Role of N6-methyladenosine methylation in nasopharyngeal carcinoma: current insights and future prospective. Cell Death Discovery. (2024) 10(1):490. doi: 10.1038/s41420-024-02266-y
75. Loh JJ, Ma S. Hallmarks of cancer stemness. Cell Stem Cell. (2024) 31(5):617–39. doi: 10.1016/j.stem.2024.04.004
76. Biserova K, Jakovlevs A, Uljanovs R, Strumfa I. Cancer stem cells: Significance in origin, pathogenesis and treatment of glioblastoma. Cells. (2021) 10(3). doi: 10.3390/cells10030621
77. Walcher L, Kistenmacher AK, Suo H, Kitte R, Dluczek S, Strauß A, et al. Cancer stem cells-origins and biomarkers: Perspectives for targeted personalized therapies. Front Immunol. (2020) 11:1280. doi: 10.3389/fimmu.2020.01280
78. Huang P, Wen F, Li Y, Li Q. The tale of SOX2: Focusing on lncRNA regulation in cancer progression and therapy. Life Sci. (2024) 344:122576. doi: 10.1016/j.lfs.2024.122576
79. Wang S, Chen W, Yu H, Song Z, Li Q, Shen X, et al. lncRNA ROR promotes gastric cancer drug resistance. Cancer Control. (2020) 27(1):1073274820904694. doi: 10.1177/1073274820904694
80. Wang S, Liu F, Deng J, Cai X, Han J, Liu Q. Long noncoding RNA ROR regulates proliferation, invasion, and stemness of gastric cancer stem cell. Cell Reprogram. (2016) 18(5):319–26. doi: 10.1089/cell.2016.0001
81. Zhao X, Xu Z, Meng B, Ren T, Wang X, Hou R, et al. Long noncoding RNA NONHSAT160169.1 promotes resistance via hsa-let-7c-3p/SOX2 axis in gastric cancer. Sci Rep. (2023) 13(1):20858. doi: 10.1038/s41598-023-47961-5
82. Song H, Xu Y, Shi L, Xu T, Fan R, Cao M, et al. LncRNA THOR increases the stemness of gastric cancer cells via enhancing SOX9 mRNA stability. BioMed Pharmacother. (2018) 108:338–46. doi: 10.1016/j.biopha.2018.09.057
83. He W, Liang B, Wang C, Li S, Zhao Y, Huang Q, et al. MSC-regulated lncRNA MACC1-AS1 promotes stemness and chemoresistance through fatty acid oxidation in gastric cancer. Oncogene. (2019) 38(23):4637–54. doi: 10.1038/s41388-019-0747-0
84. Mashouri L, Yousefi H, Aref AR, Ahadi AM, Molaei F, Alahari SK. Exosomes: composition, biogenesis, and mechanisms in cancer metastasis and drug resistance. Mol Cancer. (2019) 18(1):75. doi: 10.1186/s12943-019-0991-5
85. Xin L, Zhou LQ, Liu C, Zeng F, Yuan YW, Zhou Q, et al. Transfer of LncRNA CRNDE in TAM-derived exosomes is linked with cisplatin resistance in gastric cancer. EMBO Rep. (2021) 22(12):e52124. doi: 10.15252/embr.202052124
86. Wang J, Lv B, Su Y, Wang X, Bu J, Yao L. Exosome-mediated transfer of lncRNA HOTTIP promotes cisplatin resistance in gastric cancer cells by regulating HMGA1/miR-218 axis. Onco Targets Ther. (2019) 12:11325–38. doi: 10.2147/ott.S231846
87. Shermane Lim YW, Xiang X, Garg M, Le MT, Li-Ann Wong A, Wang L, et al. The double-edged sword of H19 lncRNA: Insights into cancer therapy. Cancer Lett. (2021) 500:253–62. doi: 10.1016/j.canlet.2020.11.006
88. Chen B, Dragomir MP, Yang C, Li Q, Horst D, Calin GA. Targeting non-coding RNAs to overcome cancer therapy resistance. Signal Transduct Target Ther. (2022) 7(1):121. doi: 10.1038/s41392-022-00975-3
89. Rupaimoole R, Slack FJ. MicroRNA therapeutics: towards a new era for the management of cancer and other diseases. Nat Rev Drug Discovery. (2017) 16(3):203–22. doi: 10.1038/nrd.2016.246
90. Coan M, Haefliger S, Ounzain S, Johnson R. Targeting and engineering long non-coding RNAs for cancer therapy. Nat Rev Genet. (2024) 25(8):578–95. doi: 10.1038/s41576-024-00693-2
91. Warner KD, Hajdin CE, Weeks KM. Principles for targeting RNA with drug-like small molecules. Nat Rev Drug Discovery. (2018) 17(8):547–58. doi: 10.1038/nrd.2018.93
92. Pallathadka H, Jabir M, Rasool KH, Malathi H, Sharma N, Pramanik A, et al. siRNA-based therapy for overcoming drug resistance in human solid tumours; molecular and immunological approaches. Hum Immunol. (2024) 86(1):111221. doi: 10.1016/j.humimm.2024.111221
93. Chen D, Liu L, Wang K, Yu H, Wang Y, Liu J, et al. The role of MALAT-1 in the invasion and metastasis of gastric cancer. Scand J Gastroenterol. (2017) 52(6-7):790–6. doi: 10.1080/00365521.2017.1280531
94. Wang Y, Zhang J, Shi H, Wang M, Yu D, Fu M, et al. M2 tumor-associated macrophages-derived exosomal MALAT1 promotes glycolysis and gastric cancer progression. Adv Sci (Weinh). (2024) 11(24):e2309298. doi: 10.1002/advs.202309298
95. Naseri B, Farsad-Akhtar N, Mardi A, Baghbani E, Bornedeli S, Asadi M, et al. lncRNA PVT1 silencing inhibits gastric cancer cells' progression via enhancing chemosensitivity to paclitaxel. Gene. (2025) 932:148900. doi: 10.1016/j.gene.2024.148900
96. Ramasamy T, Ruttala HB, Munusamy S, Chakraborty N, Kim JO. Nano drug delivery systems for antisense oligonucleotides (ASO) therapeutics. J Control Release. (2022) 352:861–78. doi: 10.1016/j.jconrel.2022.10.050
97. Aguilar R, Mardones C, Moreno AA, Cepeda-Plaza M. A guide to RNA structure analysis and RNA-targeting methods. FEBS J. (2024). doi: 10.1111/febs.17368
98. Ma WK, Voss DM, Scharner J, Costa ASH, Lin KT, Jeon HY, et al. ASO-based PKM splice-switching therapy inhibits hepatocellular carcinoma growth. Cancer Res. (2022) 82(5):900–15. doi: 10.1158/0008-5472.Can-20-0948
99. Berhane T, Holm A, Karstensen KT, Petri A, Ilieva MS, Krarup H, et al. Knockdown of the long noncoding RNA PURPL induces apoptosis and sensitizes liver cancer cells to doxorubicin. Sci Rep. (2022) 12(1):19502. doi: 10.1038/s41598-022-23802-9
100. Amodio N, Stamato MA, Juli G, Morelli E, Fulciniti M, Manzoni M, et al. Drugging the lncRNA MALAT1 via LNA gapmeR ASO inhibits gene expression of proteasome subunits and triggers anti-multiple myeloma activity. Leukemia. (2018) 32(9):1948–57. doi: 10.1038/s41375-018-0067-3
101. Adewunmi O, Shen Y, Zhang XH, Rosen JM. Targeted inhibition of lncRNA Malat1 alters the tumor immune microenvironment in preclinical syngeneic mouse models of triple-negative breast cancer. Cancer Immunol Res. (2023) 11(11):1462–79. doi: 10.1158/2326-6066.Cir-23-0045
102. Gong N, Teng X, Li J, Liang XJ. Antisense oligonucleotide-conjugated nanostructure-targeting lncRNA MALAT1 inhibits cancer metastasis. ACS Appl Mater Interfaces. (2019) 11(1):37–42. doi: 10.1021/acsami.8b18288
103. Li GZ, Meng GX, Pan GQ, Zhang X, Yan LJ, Li RZ, et al. MALAT1/ mir-1-3p mediated BRF2 expression promotes HCC progression via inhibiting the LKB1/AMPK signaling pathway. Cancer Cell Int. (2023) 23(1):188. doi: 10.1186/s12935-023-03034-1
104. Wang L, Fu H, Song L, Wu Z, Yu J, Guo Q, et al. Overcoming AZD9291 resistance and metastasis of NSCLC via ferroptosis and multitarget interference by nanocatalytic sensitizer plus AHP-DRI-12. Small. (2023) 19(4):e2204133. doi: 10.1002/smll.202204133
105. Wang Z, Wang X, Zhang T, Su L, Liu B, Zhu Z, et al. LncRNA MALAT1 promotes gastric cancer progression via inhibiting autophagic flux and inducing fibroblast activation. Cell Death Dis. (2021) 12(4):368. doi: 10.1038/s41419-021-03645-4
106. Zhang Z, Li M, Zhang Z. lncRNA MALAT1 modulates oxaliplatin resistance of gastric cancer via sponging miR-22-3p. Onco Targets Ther. (2020) 13:1343–54. doi: 10.2147/ott.S196619
107. Zhu K, Ren Q, Zhao Y. lncRNA MALAT1 overexpression promotes proliferation, migration and invasion of gastric cancer by activating the PI3K/AKT pathway. Oncol Lett. (2019) 17(6):5335–42. doi: 10.3892/ol.2019.10253
108. Wang SW, Gao C, Zheng YM, Yi L, Lu JC, Huang XY, et al. Current applications and future perspective of CRISPR/Cas9 gene editing in cancer. Mol Cancer. (2022) 21(1):57. doi: 10.1186/s12943-022-01518-8
109. Fatima H, Singh D, Muhammad H, Acharya S, Aziz MA. Improving the use of CRISPR/Cas9 gene editing machinery as a cancer therapeutic tool with the help of nanomedicine. 3 Biotech. (2025) 15(1):17. doi: 10.1007/s13205-024-04186-1
110. Li Q, Zhu Z, Zhang H, et al. LncRNA RP11-93B14.5 promotes gastric cancer cell growth through PI3K/AKT signaling pathway. Mol Biotechnol. (2024) 66:2332–40. doi: 10.1007/s12033-023-00844-6
111. Liu J, Ben Q, Lu E, et al. Long noncoding RNA PANDAR blocks CDKN1A gene transcription by competitive interaction with p53 protein in gastric cancer. Cell Death Dis. (2018) 9:168. doi: 10.1038/s41419-017-0246-6
112. Haghighi N, Doosti A, Kiani J. Evaluation of CRISPR/cas9 system effects on knocking out NEAT1 gene in AGS gastric cancer cell line with therapeutic perspective. J Gastrointest Cancer. (2022) 53:623–31. doi: 10.1007/s12029-021-00669-z
113. Liu C, Shen A, Song J, Cheng L, Zhang M, Wang Y, et al. LncRNA-CCAT5-mediated crosstalk between wnt/β-catenin and STAT3 signaling suggests novel therapeutic approaches for metastatic gastric cancer with high wnt activity. Cancer Commun (Lond). (2024) 44(1):76–100. doi: 10.1002/cac2.12507
114. Zhuo W, Liu Y, Li S, Guo D, Sun Q, Jin J, et al. Long noncoding RNA GMAN, up-regulated in gastric cancer tissues, is associated with metastasis in patients and promotes translation of ephrin A1 by competitively binding GMAN-AS. Gastroenterology. (2019) 156(3):676–691.e611. doi: 10.1053/j.gastro.2018.10.054
115. Davodabadi F, Mirinejad S, Malik S, Dhasmana A, Ulucan-Karnak F, Sargazi S, et al. Nanotherapeutic approaches for delivery of long non-coding RNAs: an updated review with emphasis on cancer. Nanoscale. (2024) 16(8):3881–914. doi: 10.1039/d3nr05656b
116. Zhou M, Dong J, Huang J, Ye W, Zheng Z, Huang K, et al. Chitosan-Gelatin-EGCG nanoparticle-meditated LncRNA TMEM44-AS1 silencing to activate the P53 signaling pathway for the synergistic reversal of 5-FU resistance in gastric cancer. Adv Sci (Weinh). (2022) 9(22):e2105077. doi: 10.1002/advs.202105077
117. Wang S, Wo L, Zhang Z, et al. Delivery of LINC00589 via mesoporous silica nanoparticles inhibits peritoneal metastasis in gastric cancer. Cancer Lett. (2022) 549:215916. doi: 10.1016/j.canlet.2022.215916
118. Wu Q, Zhu C, Zhao T, Liu T, Da M. Downregulation of lncRNA CCAT1 enhances chemosensitivity in cisplatin-resistant gastric cancer cells. Drug Dev Res. (2025) 86:e70048. doi: 10.1002/ddr.70048
119. Wang B, Liu W, Song B, et al. Targeting LINC00665/miR-199b-5p/SERPINE1 axis to inhibit trastuzumab resistance and tumorigenesis of gastric cancer via PI3K/AKt pathway. Noncoding RNA Res. (2025) 10:153–62. doi: 10.1016/j.ncrna.2024.07.004
120. Azadi SS, Safaralizadeh R, Amini M, et al. Investigating the effect of LncRNA DLGAP1-AS2 suppression on chemosensitivity of gastric cancer to chemotherapy. Naunyn Schmiedebergs Arch Pharmacol. (2024) 397:7891–903. doi: 10.1007/s00210-024-03130-7
121. Wang J, Zhang J, Liu H, et al. N6-methyladenosine reader hnRNPA2B1 recognizes and stabilizes NEAT1 to confer chemoresistance in gastric cancer. Cancer Commun (Lond). (2024) 44:469–90. doi: 10.1002/cac2.12534
122. Liu M, Li H, Li X, et al. A novel lncRNA FUAT1/TNS4 axis confers chemoresistance by suppressing reactive oxygen species-mediated apoptosis in gastric cancer. Antioxid Redox Signal. (2024) 41:24–41. doi: 10.1089/ars.2023.0298
123. Luo Y, Lu X, Ma W, et al. Dampening HOTAIR sensitizes the gastric cancer cells to oxaliplatin through miR-195-5p and ABCG2 pathway. J Cell Mol Med. (2023) 27:3591–600. doi: 10.1111/jcmm.17925
124. Alshaer W, Zureigat H, Al Karaki A, Al-Kadash A, Gharaibeh L, Hatmal MM, et al. siRNA: Mechanism of action, challenges, and therapeutic approaches. Eur J Pharmacol. (2021) 905:174178. doi: 10.1016/j.ejphar.2021.174178
125. Dhuri K, Bechtold C, Quijano E, Pham H, Gupta A, Vikram A, et al. Antisense oligonucleotides: An emerging area in drug discovery and development. J Clin Med. (2020) 9(6). doi: 10.3390/jcm9062004
126. Hofman CR, Corey DR. Targeting RNA with synthetic oligonucleotides: Clinical success invites new challenges. Cell Chem Biol. (2024) 31(1):125–38. doi: 10.1016/j.chembiol.2023.09.005
127. Gupta D, Bhattacharjee O, Mandal D, Sen MK, Dey D, Dasgupta A, et al. CRISPR-Cas9 system: A new-fangled dawn in gene editing. Life Sci. (2019) 232:116636. doi: 10.1016/j.lfs.2019.116636
128. Sparmann A, Vogel J. RNA-based medicine: from molecular mechanisms to therapy. EMBO J. (2023) 42(21):e114760. doi: 10.15252/embj.2023114760
129. Patni H, Chaudhary R, Kumar A. Unleashing nanotechnology to redefine tumor-associated macrophage dynamics and non-coding RNA crosstalk in breast cancer. Nanoscale. (2024) 16(39):18274–94. doi: 10.1039/d4nr02795g
130. Hu X, Lv X, Zhang L, Li SS, Jin X. Noncoding RNA lipotherapeutics: A promising breakthrough in pulmonary hypertension treatment. Curr Pharm Biotechnol. (2025) 26(1):9–16. doi: 10.2174/0113892010302590240321073509
131. Muskan M, Abeysinghe P, Cecchin R, Branscome H, Morris KV, Kashanchi F. Therapeutic potential of RNA-enriched extracellular vesicles: The next generation in RNA delivery via biogenic nanoparticles. Mol Ther. (2024) 32(9):2939–49. doi: 10.1016/j.ymthe.2024.02.025
132. Khaleel AQ, Jasim SA, Menon SV, Kaur M, Sivaprasad GV, Rab SO, et al. siRNA-based knockdown of lncRNAs: A new modality to target tumor progression. Pathol Res Pract. (2024) 266:155746. doi: 10.1016/j.prp.2024.155746
133. Kumar A, Yap KC, BharathwajChetty B, Lyu J, Hegde M, Abbas M, et al. Regulating the regulators: long non-coding RNAs as autophagic controllers in chronic disease management. J BioMed Sci. (2024) 31(1):105. doi: 10.1186/s12929-024-01092-9
134. Sui X, Zhang Q, Hao M, Chen Y. Serum LINC01133 combined with CEA and CA19-9 contributes to the diagnosis and survival prognosis of gastric cancer. Med (Baltimore). (2024) 103(46):e40564. doi: 10.1097/md.0000000000040564
135. Jiang X, Wu X, Lu M, Fan W, Song J, Song F. Long non-coding RNA FAM87A is associated with overall survival and promotes cell migration and invasion in gastric cancer. Front Oncol. (2024) 14:1448502. doi: 10.3389/fonc.2024.1448502
Keywords: lncRNA, gastric cancer, drug resistance, therapy, precision medicine
Citation: Liu W and Wang W (2025) LncRNA in gastric cancer drug resistance: deciphering the therapeutic strategies. Front. Oncol. 15:1552773. doi: 10.3389/fonc.2025.1552773
Received: 29 December 2024; Accepted: 17 March 2025;
Published: 01 April 2025.
Edited by:
Khushbukhat Khan, Trials360, PakistanReviewed by:
Eswari Dodagatta-Marri, University of California, San Francisco, United StatesCopyright © 2025 Liu and Wang. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: WeiFa Wang, Mzk1Mjg5MTFAcXEuY29t
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.