
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
REVIEW article
Front. Oncol. , 03 April 2025
Sec. Molecular and Cellular Oncology
Volume 15 - 2025 | https://doi.org/10.3389/fonc.2025.1524763
 Shuzhen Mao1†
Shuzhen Mao1† Junyan Li2†
Junyan Li2† Jiahui Huang3
Jiahui Huang3 Lili Lv2
Lili Lv2 Qilian Zhang4
Qilian Zhang4 Qing Cheng3
Qing Cheng3 Xiaojing Liu3
Xiaojing Liu3 Zhiwei Bi3
Zhiwei Bi3 Jing Yao3*
Jing Yao3*Cancer is a complex and highly lethal disease marked by unchecked cell proliferation, aggressive behavior, and a strong tendency to metastasize. Despite significant advancements in cancer diagnosis and treatment, challenges such as early detection difficulties, drug resistance, and adverse effects of radiotherapy or chemotherapy continue to threaten patient survival. MicroRNAs (miRNAs) have emerged as critical regulators in cancer biology, with miR-506 being extensively studied and recognized for its tumor-suppressive effects across multiple cancer types. This review examines the regulatory mechanisms of miR-506 in common cancers, focusing on its role in the competing endogenous RNA (ceRNA) network and its effects on cancer cell proliferation, apoptosis, and migration. We also discuss the potential of miR-506 as a therapeutic target and its role in overcoming drug resistance in cancer treatment. Overall, these insights underscore the therapeutic potential of miR-506 and its promise in developing novel cancer therapies.
Cancer is a terrible disease, and cancer cells usually proliferate in an uncontrolled manner. Cancer cells demonstrate invasiveness and metastasis during proliferation, leading to serious consequences (1, 2). As the second leading cause of death globally, cancer treatment has always posed an enormous challenge. Current approaches to cancer management include surgery, chemotherapy, radiation therapy, targeted therapies, and their combinations of these. However, despite advancements in traditional treatment approaches and the emergence of new treatments, the cancer-related mortality rate continues to increase (3, 4). The main challenges currently faced in cancer treatment include difficulties in detecting and diagnosing early-stage cancer, susceptibility to metastasis, increased drug resistance, and individual variations during treatment. Therefore, actively exploring new approaches to the diagnosis and management of cancer is of great importance.
MicroRNAs (miRNAs) are a class of endogenous, highly conserved, non-coding single-stranded RNA molecules, typically consisting of 18 to 25 nucleotides in length (5, 6). They primarily function to regulate the stability of mRNA for protein-coding genes negatively. In most cases, miRNAs promote the degradation of target mRNA or reduce its proper translation initiation by binding to complementary sequences within the 3′ untranslated region (UTR) (4). The discovery of the first miRNA, lin-4, in 1993 unveiled a novel mechanism of gene regulation (7). An increasing body of research has demonstrated that dysregulation of miRNAs is closely associated with the onset and progression of various human diseases (8, 9), especially in cancer (10). miRNAs have now been widely recognized as tumor suppressors or oncogenes in numerous human malignancies, playing pivotal roles in cancer development and progression by regulating multiple signaling pathways and cellular functions (10, 11).
MicroRNA-506 (miR-506) is located on the q27.3 locus of the X chromosome (chrXq27.3) (12). This locus houses a key miRNA cluster containing 30 mature miRNAs, which are further categorized into two groups according to their genomic location: miR-506-514 and miR-888-892, with miR-506 being the most well-known member of this cluster (13). According to the source, miR-506 is classified as 5p or 3p, with miR-506-3p holding a prominent place in miR-506 (14). Recent preclinical studies have highlighted the tumor-suppressive potential of miR-506 across multiple cancer types, positioning it as a promising candidate for clinical translation but occasionally displaying oncogenic properties depending on the cancer type and molecular targets. miR-506 inhibits hepatocellular carcinoma (HCC) cell proliferation and tumorigenicity by targeting Rho-associated protein kinase 1 (ROCK1), which is upregulated in HCC tissues and inversely correlated with miR-506 levels, supporting miR-506 mimics as a therapeutic strategy (15). miR-506 suppresses Non-Small Cell Lung Cancer (NSCLC) progression by regulating Tubby-like protein 3 (TULP3), inducing mitochondrial apoptosis, and in vivo studies confirm its ability to reduce tumor growth, highlighting its role in NSCLC therapy (16). miR-506-3p targets MTMR6, inhibiting cell proliferation and promoting apoptosis. Its downregulation in ovarian tumors underscores its potential as a therapeutic target (17). Conversely, in colorectal cancer (CRC), miR-506 inhibits NR4A1, an orphan nuclear receptor driving oncogenic signaling (18). In breast cancer (BC), miR-506 modulates the specificity protein (SP)1/SP3 to demethylate the tumor-suppressive lncRNA MEG3, thereby inhibiting metastasis (19). These distinct targets underscore the need for cancer-specific mechanistic studies. While miR-506 is often recognized for its tumor-suppressive functions, limited evidence suggests it can also exhibit pro-cancer effects in specific tumor types. In melanoma, miR-506 has been identified as part of the miRNA-506-514 cluster, critical in initiating melanocyte transformation and promoting tumor growth. The overexpression of miR-506 in melanoma models has been linked to enhanced tumorigenicity, suggesting its potential as an oncogenic driver in this context (14). miRNA-506 expression is elevated in the plasma of esophageal squamous cell carcinoma (ESCC) patients compared to healthy individuals. It correlates with disease severity indicators, which exhibit diagnostic significance for ESCC and are associated with patient prognosis, suggesting its potential as a molecular marker for both diagnosis and outcome prediction in ESCC (20). Compared to miR-506-3p, miR-506-5p has been investigated in only a limited number of cancer types, including gastric cancer (GC) and glioma (21–24).
Despite the increasing research on miR-506 due to its recognized importance in cancer regulation, several unmet needs remain in its therapeutic application. The expression pattern and function of miR-506 are complex and, at times, contradictory, highlighting its distinct role across different cancer types. Growing evidence suggests that miR-506 primarily functions as a tumor suppressor. However, miR-506 exhibits oncogenic properties in specific contexts, depending on the cancer type and cellular environment. For instance, miR-506 promotes tumorigenesis in uveal melanoma (UM) (25) and contributes to chemoresistance in CC (26). Furthermore, tumor heterogeneity and microenvironmental factors, such as metabolic reprogramming, immune modulation, and its interactions with other non-coding RNAs, such as those involved in competing endogenous RNA (ceRNA) network, further modulate miR-506’s role in cancer progression (17). This functional paradox complicates the development of miR-506-based therapies, as their therapeutic efficacy varies across cancer types and microenvironments. miR-506 simultaneously targets multiple mRNAs, exerting broad regulatory effects across multiple genes (27). This characteristic makes miR-506 a promising candidate for treating multifactorial diseases such as cancer. However, its extensive regulatory activity also increases the risk of off-target effects, potentially disrupting normal cellular function (28). Due to its inherent instability and poor tissue specificity, effective delivery remains a significant challenge. Naked miR-506 molecules undergo rapid degradation in circulation, reducing their bioavailability and limiting their therapeutic potential (29). Exosome- and nanoparticle-based miRNA delivery systems have shown potential in enhancing miRNA stability and targeting specificity (30, 31). Despite these advancements, optimized delivery strategies are still needed to ensure safe and effective tumor-specific uptake while minimizing toxicity. The clinical validation of miR-506-based therapies remains limited, necessitating further large-scale, multicenter clinical studies to assess their safety, efficacy, and potential integration into standard cancer treatments (32).
This review distinguishes itself from previous studies by providing a comprehensive and up-to-date perspective on the role miR-506 in cancer therapy. Unlike previous reviews that primarily focused on its tumor-suppressive function, this review systematically examines the oncogenic and tumor-suppressive roles of miR-506, offering insights into the underlying mechanisms contributing to this paradox. Furthermore, this review highlights the influence of the ceRNA network on miR-506 function, an aspect that has not been extensively explored in previous studies. This review provides a detailed analysis of the molecular mechanisms through which miR-506 modulates drug resistance. Additionally, it discusses the challenges associated with the development of miRNA-based therapies. It aims to elucidate the potential of the miR-506 family as a therapeutic target in cancer, thereby advancing the development of more effective and safer cancer treatment strategies.
The ceRNA network is a relatively novel concept in molecular biology that describes how different RNA species—primarily messenger RNAs (mRNAs), long non-coding RNAs (lncRNAs), and circular RNAs (circRNAs)—interact with miRNAs in a manner that influences gene expression (33). ceRNAs are endogenous transcripts that harbor shared miRNA response elements (MREs) and regulate one another by competing for binding to shared miRNAs, thereby reducing the availability of miRNAs for their target mRNAs (33, 34). Since miRNAs typically function as post-transcriptional repressors by targeting mRNAs for degradation or translational inhibition, ceRNA competition can modulate mRNA stability and translation, ultimately affecting key cellular processes such as proliferation, differentiation, and apoptosis (35).
The interplay between ceRNAs is orchestrated by shared miRNAs, forming a finely balanced regulatory network (33). Dysregulation of ceRNA networks in cancer is often driven by genetic mutations, epigenetic alterations, or aberrant RNA expression, disrupting critical oncogenes and tumor suppressors (36). Increasing evidence suggests that ceRNAs play a crucial role in tumorigenesis by modulating the expression of oncogenes and tumor suppressors through miRNA-mediated interactions (37). Functionally, ceRNAs in cancer can be categorized into the following roles (34) (1): regulation of cancer cell proliferation, migration, and invasion. For instance, TINCR functions as a ceRNA by sequestering miR-544a from its target gene FBXW7, thereby suppressing lung cancer cell proliferation and invasion (38). Conversely, circ-TLK1 and lncRNA HOSD-AS1 enhance HCC progression by sponging miR-138-5p and miR-130a-3p, respectively, leading to upregulation of SOX4, a key oncogene that promotes proliferation, invasion, and metastasis (39) (2). modulation of epithelial-mesenchymal transition (EMT). lncRNA CCAT1 functions as a ceRNA to promote EMT in gliomas and retinoblastomas, thereby facilitating tumor progression (40, 41). In contrast, lncRNA CASC2 acts as a tumor suppressor by inhibiting HCC cell migration, invasion, and EMT progression through miRNA-mediated regulation (42) (3). promotion or suppression of tumor initiation. RSU1P2, for example, promotes cervical cancer development by competitively binding to the shared microRNA let-7a, thereby enhancing oncogene expression (43). In contrast, circRNA Cir-ITCH exerts tumor-suppressive effects by functioning as a ceRNA to inhibit bladder cancer progression (44). (4) regulation of chemotherapeutic sensitivity and resistance. lncRNA CCAT1 enhances HCC cell proliferation and reduces oxaliplatin-induced apoptosis, thereby contributing to chemoresistance (45). Conversely, lncRNA CASC2 increases the sensitivity of prostate cancer cells to docetaxel, highlighting its potential as a therapeutic target for overcoming drug resistance (46).
Emerging research has revealed that miR-506 not only directly targets mRNAs to regulate cancer progression but also participates in ceRNA networks by interacting with circRNAs and lncRNAs. These interactions create a multifaceted regulatory framework that amplifies miR-506's role in cancer biology (47). Below, we discuss the involvement of circRNAs and lncRNAs in miR-506-mediated ceRNA networks and their implications in cancer.
circRNAs are non-coding RNA molecules distinguished by their strong stability and have recently gained recognition as critical components of gene expression networks (48, 49). circRNAs possess multiple conserved miRNA binding sites and can function as efficient miRNA sponges that specifically adsorb miRNAs, thereby modulating their inhibitory effects on target genes (50). A growing body of evidence supports the notion that circRNAs, functioning as ceRNAs, dominate the regulation of the circRNA-miRNA-mRNA signaling axis (51). For instance, in NSCLC, circRNAFOXO3 (52) and circRNA100565 (53) can inhibit miR-506 by exerting a sponge effect, thereby up-regulating high mobility group box B3 (HMGB3) and high mobility group box A2 (HMGA2) and promoting disease progression. Additionally, has-circRNA0016788 can enhance HCC progression by modulating miR-506-3p and increasing Poly ADP-ribose Polymerase family 14 (PARP14) expression (54). Snail family transcriptional repressor 2 (Snail2) and yes-associated protein 1 (YAP1) are known oncogenes that are up-regulated in cancer (55, 56), circRNAPCNX serves as a sponge for miR-506 and inhibits the anti-cancer activity of miR-506 via the circRNAPCNX-miR-506-Snail2/YAP axis in HCC (57). Another study shows that circRNASYPL1 can down-regulate miR-506 and up-regulate zeste homolog 2 (EZH2) enhancer, facilitating cancer cell metastasis in HCC and exacerbating disease progression (58). As mentioned earlier, studies have shown that miR-506 can also act synergistically with other miRNAs as a target for ceRNA. For example, in CRC, circRNAPACRGL stimulates the growth, invasion, and migration of cancer cells through the miR-142-3p/miR-506-3p-TGF-β1 axis while also influencing neutrophil differentiation from N1 to N2, thereby affecting apoptosis (59).
lncRNAs, similar to circRNAs, have been recognized for their capacity to modulate miR-506 function in cancer through a ceRNA mechanism. lncRNAs, a class of non-coding transcripts exceeding 200 nucleotides in length, account for approximately 60% of the total human transcripts (60). The ongoing advancement of biotechnology in recent years has underscored that lncRNAs play a significant role in regulating cancer-related processes, including tumor cell proliferation, apoptosis, autophagy, EMT, and drug resistance (61). For instance, LINC01433, by targeting miR-506-3p, has been shown to enhance the proliferative, migratory, and invasive properties of nasopharyngeal carcinoma cells (62). In gliomas, lincRNA00963 reduces the expression level of miR-506, thereby affecting the downstream target protein branched-chain Amino Acid Transaminase 1 (BCAT1) and weakening the inhibitory effect of miR-506 on tumors (63). Huang et al. demonstrated through clinical samples and cell experiments that lncRNANEAT1 can exacerbate cancer progression by acting as a sponge for miR-506-3p (64). In summary, within the ceRNA network, circRNAs and lncRNAs typically serve as effective “sponges” for miRNAs, sequestering them from target mRNAs to indirectly regulate gene expression and influence tumor progression, including EMT and drug resistance (65). The relationships among circRNAs, lncRNAs, and miR-506 in different types of cancer are as follows (Tables 1, 2); Figure 1 shows the ceRNA network of lncRNA or circRNA with miR-506 and corresponding targets in cancer, as well as the direct target of miR-506.
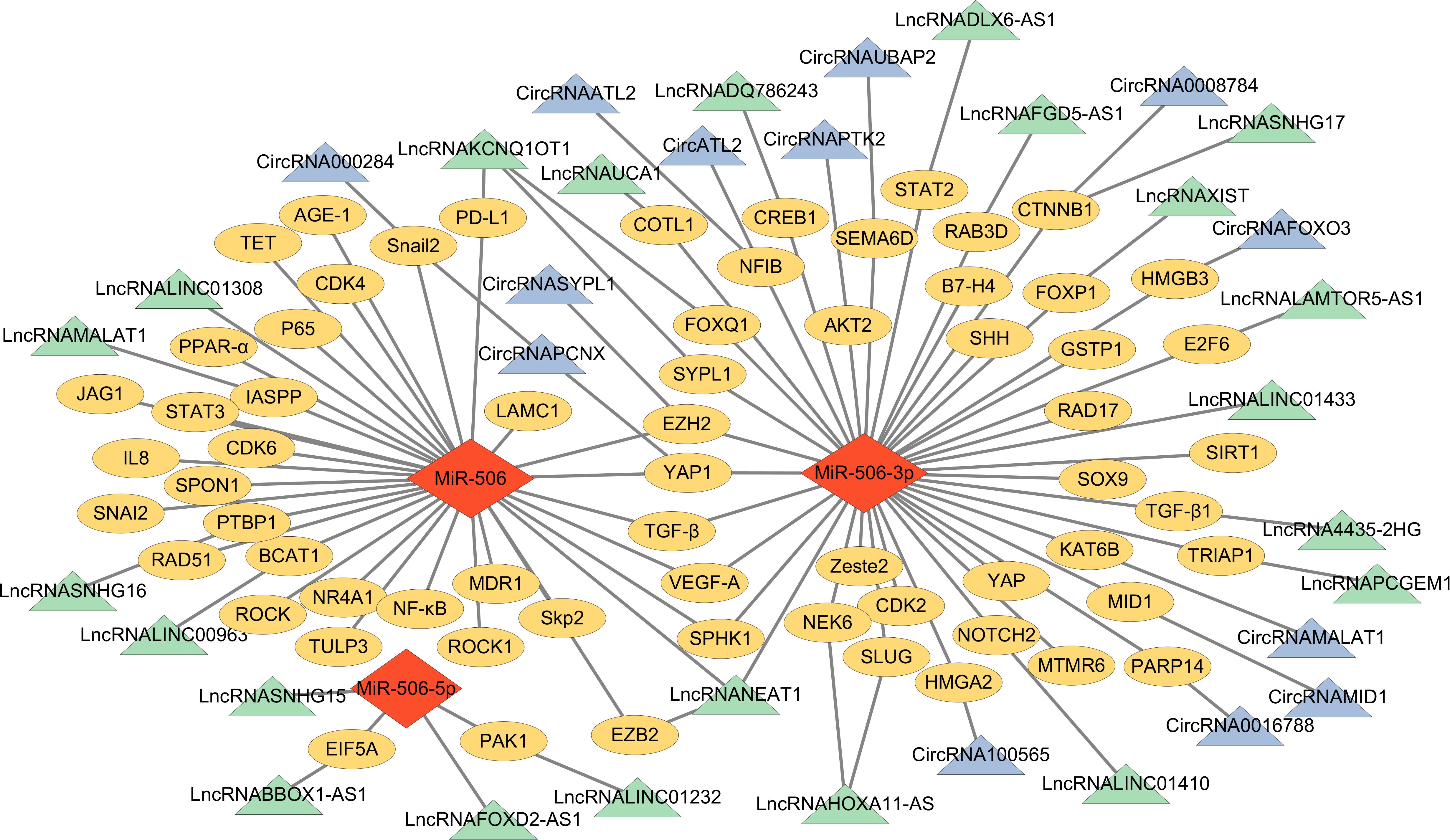
Figure 1. The ceRNA network of miR-506 family members and their target genes. ( ) = miR-506 family members; (
) = miR-506 family members; ( ) = circRNAs; (
) = circRNAs; ( ) = lncRNAs; (
) = lncRNAs; ( ) = target genes.
) = target genes.
Lung cancer remains the predominant cause of cancer-related mortality globally, with NSCLC accounting for over 85% of cases, and patients with NSCLC are usually diagnosed at advanced stages (66, 67). Decreased levels of miR-506-3p result in enhanced expression of coactosin-like protein 1 (COTL1), thereby promoting the progression of NSCLC (68). Yin et al. demonstrated that miR-506 can promote cancer cell apoptosis, reduce their activity, and achieve anti-cancer effects by participating in oxidative stress and targeting Nuclear factor kappa b (NF-κB) and p65 together (69). The tumor microenvironment affects the disease progression, including the changes in peripheral blood vessels. Vascular endothelial growth factor A (VEGF-A) is a critical cytokine in angiogenesis and is regulated by signal transducer and activator of transcription 3 (STAT3); alginic acid can lower miR-506 levels to inhibit angiogenesis, which is consistent with the results of xenograft experiments (70). Research also indicates that miR-506 can work synergistically with other miRNAs to impede cancer progression. Specifically, combination therapy of miR-143 and miR-506 downregulates the levels of cyclin-dependent kinases (CDK) 1, CDK4, and CDK6 and induces apoptosis in cancer cells, but it does not affect normal lung fibroblasts. This combination therapy can usually inhibit the transition of the cancer cell cycle, equivalent to the effect of clinical cell cycle inhibitors (71); Hossian et al.'s research also confirms this viewpoint (72). In conclusion, these findings demonstrate that miR-506 may function as a tumor suppressor in lung cancer.
Liver cancer (LC) ranks as the fourth most common cause of mortality attributed to cancer, with its incidence showing an upward trend (73). Typically, LC can be broadly categorized into two types: primary liver cancer and metastatic liver cancer. The former encompasses various types, including HCC, intrahepatic cholangiocarcinoma, and mixed liver cancer, with HCC being the most prevalent (74). Research highlights that miR-506-3p can inhibit the growth of LC cells and serve as a biomarker for the diagnosis and prognosis of LC (75). The Hippo signaling pathway is significantly linked to the onset of LC, with YAP critically involved in modulating the response genes within this pathway (76, 77). Researchers found that miR-506-3p can specifically target the 3' UTR of YAP, leading to the downregulation of YAP along with its associated target genes, including cellular myelocytomatosis (c-Myc) and connective tissue growth factor (CTGF), thereby inhibiting proliferation in LC cells (78). MiR-506 has been shown to trigger apoptosis in HCC cells. Deng et al. demonstrated through experiments that miR-506 restrains HCC cell proliferation in vitro and tumor growth in vivo. Rho-associated protein kinase 1 (ROCK1), a target that miR-506 may directly target in HCC cells, has been shown to have an inverse relationship with miR-506-3p expression levels in HCC tissue. Furthermore, miR-506 is capable of causing cell cycle arrest and promoting apoptosis in cancer cells (15). miR-506 also affects the microenvironment around HCC cells; rapid cancer growth requires an adequate blood supply, and tumor vascular growth is essential for cancer development (79). HCC is generally considered a hypervascular tumor because it exhibits arterial enhancement compared with normal liver tissue (74). Sphingosine kinase 1 (SPHK1) facilitates cell survival, proliferation, transformation, and angiogenesis, while miR-506 targets SPHK1 to reduce its expression at the cellular, mRNA, and protein levels, inhibiting liver cancer angiogenesis (80). Previous studies show that miR-506 can directly target specific proteins to inhibit the growth of HCC cells, such as interleukin-8 (IL-8) (81) and STAT3 (82). Moreover, prior research has demonstrated that the functions of Ras homolog A (RhoA) in cancer cell migration, apoptosis, and proliferation are critical (83). The ROCK family contributes to cancer advancement by modulating cell growth and migration processes (84). miR-506 can modulate LC cell proliferation and apoptosis by influencing the RhoA/ROCK signaling cascade (85). Additional research demonstrates that down-regulating the expression of sirtuin1 (SIRT1) may prevent cancer cells from invasively growing and metastasizing with the combined treatment of miR-124-3p and miR-506-3p, thereby delaying the development of LC (86). Summarizing the evidence, there is extensive linkage between LC and miR-506, suggesting t that miR-506 may serve as a promising therapeutic target for managing LC.
Osteosarcoma (OS) is a highly malignant bone tumor predominantly affecting adolescents, with approximately 75% of patients aged 15 to 25 years (87). Characterized by a tendency for early metastasis and poor prognosis, OS poses a significant threat to adolescent health, and searching for new and effective therapeutic targets is an urgent priority. A growing body of research indicates that miR-506 functions as an inhibitor of OS initiation and progression through the modulation of various target genes, and elevated levels of its expression generally correlate with improved patient survival rates (55, 88, 89). miR-506 typically exerts inhibitory effects on OS. Yu et al. demonstrated through clinical sample studies and cell experiments that an increase in miR-506 levels can reduce the expression of Snail2 (55). This transcriptional repressor can promote EMT and, consequently, accelerate the metastasis of cancer cells (90). Additionally, astrocyte elevated gene-1 (AGE-1) has been confirmed as a downstream effector of miR-506; this microRNA can suppress AGE-1 expression via the wingless-type MMTV integration site family/beta-catenin (Wnt/β-catenin) signaling pathway to exert its inhibitory effect on OS cells, which was validated in an OS mouse model (89). Further studies have also pointed out that miR-506 impedes the aggressive behavior of OS by targeting Ras-related protein Rab-3D (RAB3D) (91), S-phase kinase-associated protein 2 (Skp2) (92), or SPHK1 (88). Similarly, the NF-κB signaling pathway is significant in various diseases, including cancer; IL-1β can reduce the expression of miR-506 through this pathway, leading to increased Jagged 1 (JAG1) expression and exacerbating the progression of OS (93). However, most clinical samples related to OS research have not been conducted in specific experiments based on different tumor stages. Moreover, many studies on miR-506 are still at the experimental stage and lack extensive clinical validation. These limitations challenge the clinical application of miR-506 in diagnosis and treatment.
Globally, CRC is categorized as the third most frequently occurring malignant neoplasm and accounts for the second-highest number of cancer-related fatalities (94). Researchers have identified that several genes targeted by miR-506 impact the biological behavior of CRC, including EZH2, which is implicated in carcinogenesis as an oncogene (95). Research by Zhang et al. confirmed that miR-506 is significantly downregulated in CRC tissues and cells through both in vivo and in vitro studies. By directly targeting the 3' UTR of EZH2, miR-506 reduces its level and subsequently exerts an anti-tumor response mediated by the Wnt/β-catenin signaling pathway (96). Ten-eleven translocation (TET) is highly expressed in CRC; miRNA-506 targets TETs and regulates the levels of TET1, TET2, and TET3, ultimately inhibiting the proliferation and invasion of CRC cells in vivo and in vitro (97). Furthermore, overexpression of miR-506 has been shown to inhibit cancer cell viability, invasion, and colony formation ability of CRC cells; moreover, miR-506 also targets laminin subunit gamma-1 (LAMC1), which is an important extracellular matrix (ECM) regulator to slow down cell migration and then reduce the metastasis of cancer cells (98). Other studies have similarly suggested a suppressive role of miR-506-3p in CRC, such as its targeting of cystatin-p1 (CSTP1) (30) and EZH2 (99). Combination therapy is also an effective treatment; miR-124 and miR-506 jointly target DNA (cytosine-5)-methyltransferase 3B (DNMT3B) and dynamin-1 (DNM1) and reduce global DNA methylation to reduce the effect of CRC (100). The sensitivity of radiotherapy and chemotherapy also reflects the inhibitory effect of miR-506 on CRC. Studies have shown that, based on the novel network-based method, radiosensitive patients exhibit significantly higher expression levels of hsa-miR-506-3p than radioresistant patients, offering a new foundation for future CRC treatment (101). Early cancer diagnosis, timely surgical intervention, and effective disease control are crucial. It has been found that miR-506 is helpful for early diagnosis of CRC (102). Although miR-506 shows significant anti-cancer effects, numerous genetic factors and signaling cascades contribute to CRC onset and progression, so the single-target action of miR-506 may not fully control cancer progression. Further research is essential to uncover the precise mechanisms and specific gene targets through which miR-506 influences CRC.
The incidence of OC is notably high among women. A significant proportion of patients are diagnosed at an advanced stage of the disease, leading to high mortality (103). In the past few years, several miR-506 target genes affecting the biological behavior of OC have been identified. Sun et al. observed a relationship between miR-506 expression and increased levels of E-cadherin (E-cad), along with the downregulation of vimentin (VIM), N-cadherin (N-cad), and snail homolog 2 (SNAI2). miR-506 can bind to the specific binding site of VIM, affecting the migration and drug resistance of epithelial ovarian cancer (EOC). These findings were confirmed by studies on clinical samples and animal experiments (104). Another study shows that miR-506-3p can target myotubularin-related protein 6 (MTMR6) to prevent cancer cells from proliferating in OC tissues, arresting the cells in the G0/G1 phase and thereby inducing apoptosis (17). A similar study indicates that miR-506 can target EZH2, reducing its expression and inhibiting OC (95). Other studies have also confirmed this conclusion (105). The CDK4/6-forkhead box m1 (FOXM1) pathway is a crucial signal network frequently activated in high-grade serous OC cases (106). Liu et al. demonstrated that miR-506 might suppress this pathway through in vivo and in vitro studies and clinical sample investigations, induce cancer cell apoptosis, and inhibit tumor progression in OC (107). Sirtuin1 (SIRT1), a deacetylation-regulating intracellular protein, activates tumor suppressor protein p53 to control tumor growth (108). In OC, miR-506-3p has been observed to interact with the 3'UTR of SIRT1 and reduce its expression, while studies confirm that elevated expression of SIRT1 promotes the proliferation of OC cells and inhibits apoptosis (109). Through integrated analyses, Yang et al. observed that miR-506 may be regulated in serous OC. Further, they confirmed at the cellular level that miR-506 can target SNAI2, a transcriptional repressor of E-cadherin, blocking EMT induced by transforming growth factor β (TGF-β). miR-506 enhances the expression of E-cadherin, reduces the expression of SNAI2 and VIM, and inhibits cancer cell migration and invasion (110). Research has also reported that miR-506 can directly target RAD51, reduce the expression of RAD51, regulate PARP-induced DNA damage repair, and improve chemosensitivity to cisplatin (111). Ultimately, the beneficial effects miR-506 on OC are extensive, encompassing inhibition of cancer cell migration, promotion of apoptosis, and enhancement of chemosensitivity, providing a solid foundation for future OC treatments.
he impact of miR-506 on various cancer types has become increasingly recognized in the scientific community. The mechanisms of action and associated target genes of the miR-506 family in common cancers are illustrated in Figure 2. In particular, miR-506-3p has been shown to suppress the proliferation of papillary thyroid cancer (PTC) cells by targeting YAP1 (56). YAP1 is a known promoter of tumor progression (112) and acts as a carcinogen (113). YAP1 is the direct target gene of miR-506-3p, which targets YAP1 3’UTRs to down-regulate YAP1, allowing exposure of the CDK2/Cyclin E1 complex. Consequently, miR-506-3p is believed to inhibit the proliferation of PTC by suppressing YAP1 and modulating the YAP1-CDK2/Cyclin protein complex, which is involved in cell cycle control (56). YAP1 is an important target gene for many malignant cancers, such as HCC (78), papillary thyroid cancer (56), GC (114), and esophageal squamous cell carcinoma (ESCC) (115). In ESCC, YAP activates its downstream target SOX9 through TEA domain transcription factor 1 (TEAD1)-mediated binding transcription, targeting YAP’s miRNA, including miR-506-3p. This results in SOX9 inducing miR-506-3p, inhibiting YAP expression post-transcriptionally, creating a negative feedback mechanism (115). Other studies show that miR-506 functions as an oncosuppressive microRNA in nasopharyngeal carcinoma, exerting its tumor-suppressing activity primarily by suppressing Forkhead box protein Q1 (FOXQ1) expression (116). Similarly, in cervical cancer tissues, a reduction in miR-506 levels has been observed (117), which is negatively correlated with FOXQ1 expression. High expression of FOXQ1 enhances the migration and metastatic abilities of cervical cancer cells, promotes epithelial-mesenchymal transition (EMT), and decreases chemosensitivity. Based on these findings, there may be implications for developing treatment strategies for cervical cancer (118). Gli3 is a hedgehog pathway transcription factor. Wen et al. found that in cervical cancer, miR-506 can target Gil3, arrest cell cycle progression at the G1/S phase, curb proliferation, and induce apoptosis (119). Studies have shown that in prostate cancer cell lines (DU145 and PC-3), miR-506-3p expression is decreased, and its high expression inhibits tumor progression in vitro. At the same time, increased levels of N-acetylgalactosaminyltransferase 4 (GALNT4) negate the inhibitory effect of miR-506-3p on tumors (120). In addition, miR-506 targets ROCK1 and downregulates its expression, inhibiting neuroblastoma tumor growth (121). The association between BC survival and EMT has been widely confirmed in numerous studies, where miR-506 is shown to attenuate BC progression by regulating the EMT process (122, 123).
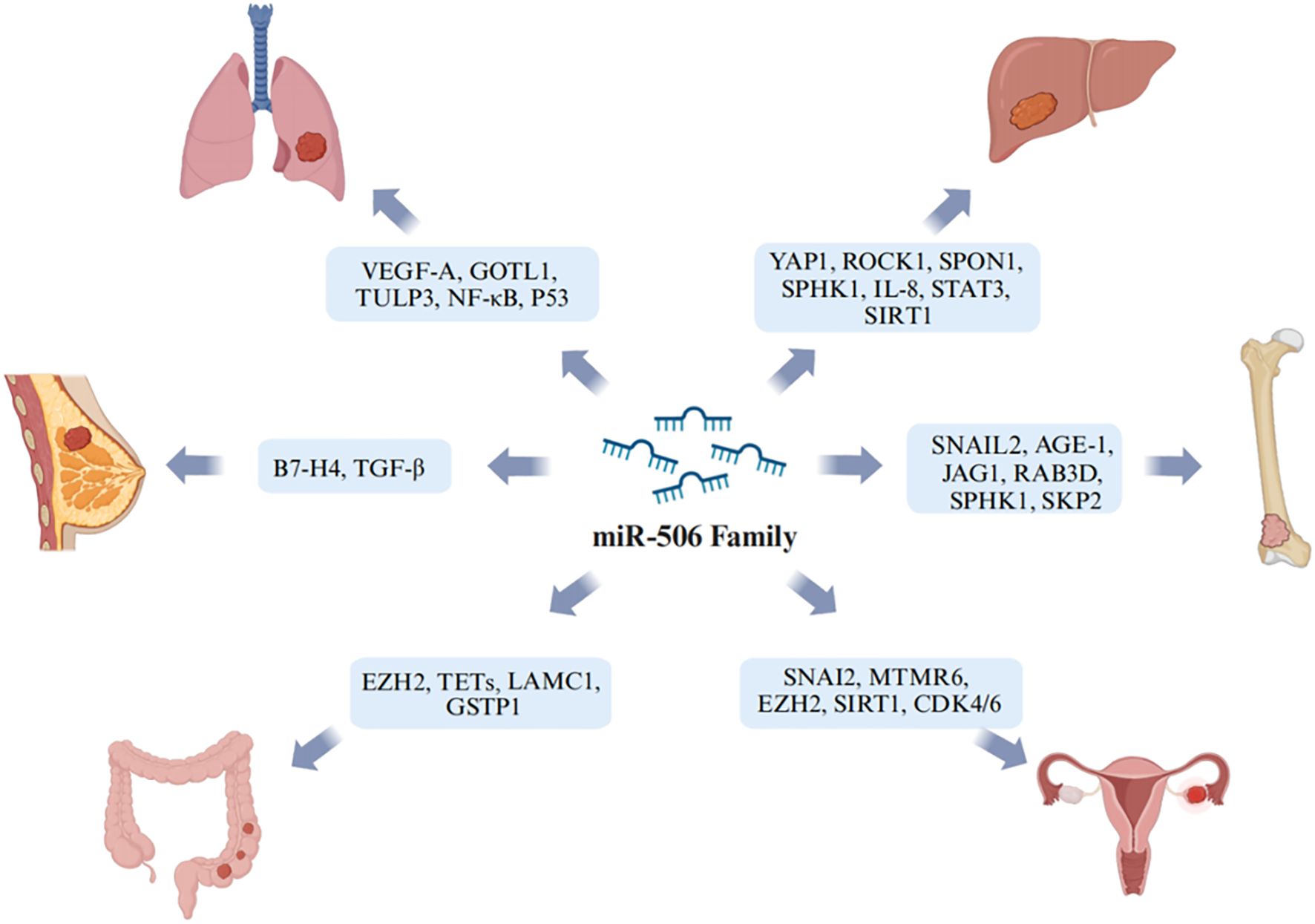
Figure 2. Schematic representation of the potential biological mechanisms through which miR-506 regulates various types of cancer. Image created with BioRender.com, with permission.
In cancer, the miR-506 family usually plays an inhibitory role; however, some research indicates that miR-506 may facilitate the progression of certain cancers. The miR-506-514 cluster is a collection of microRNAs on chromosome chrXq27.3, with miR-506-3p as its most prominent member (13). UM is the most prevalent primary intraocular malignancy in adults (124). It originates from melanocytes in the uveal tract, specifically in the choroid, iris, or ciliary body (125). Research suggests that the miR-506-514 cluster functions as a tumor suppressor in UM. Falzone et al. conducted a clinical sample study and divided the patients into T3-T4 vs. T1-T2, categorizing them as high-grade and low-grade groups. Compared with low-grade or surviving UM patients, the expression of mRNAs within the miRNA-506-514 cluster was significantly reduced in high-grade or deceased individuals (126). Nevertheless, findings from other relevant research studies indicate otherwise. Streicher et al. observed overexpression of the miR-506-514 cluster in melanoma patient tissues and cell lines, including SKMEL-2, SKMEL-5, A375, MALME-3M, and RPMI-7951. Inhibition of this cluster was shown to induce apoptosis in melanoma cells. Moreover, the sub-cluster effect was more evident than the whole cluster's (14). In addition, the role of miR-506 seems to be different between primary UM and metastatic melanoma. Specifically, miR-506 levels were elevated in patients with metastatic UM compared to those with primary UM, suggesting its potential as an early biomarker in disease progression (127). Melanoma is a biologically distinct malignancy that exhibits a strong response to immunotherapy, unlike most other solid tumors (128). Additionally, the anterior segment of the eye, particularly the iris, is more exposed to ultraviolet (UV) radiation, leading to DNA damage, distinguishing UM from other cancers (129). Tong et al. (26) reported that miR-506 is associated with chemoresistance in CRC. In the SW1161 CRC cell line, miR-506 induces resistance to hydroxy camptothecin by suppressing the expression of peroxisome proliferator-activated receptor alpha (PPARα), thereby reducing the drug’s efficacy. miR-506 can function as both a tumor suppressor and an oncogene in different cancer types, with several key factors contributing to this dual role. miR-506 exhibits tissue-specific expression patterns, which may determine whether it acts as a tumor suppressor or an oncogene. miR-506 is regulated by ceRNA networks, including lncRNAs and circRNAs, which act as "sponges" to modulate its availability (47). Its functional accessibility may vary significantly across different cancer types. Additionally, the tumor microenvironment, including immune infiltration, hypoxia, and stromal interactions, further regulates miR-506 activity, altering its downstream effects (28). Genetic variation is another major factor contributing to this paradox, encompassing structural genetic alterations such as chromosomal deletions/amplifications and mutations (130), defects in miRNA biogenesis mechanisms, and epigenetic changes, such as altered DNA methylation (131), which vary across different cancer types. Furthermore, no universally accepted protocol for miRNA sample collection, processing, detection, quantification, or therapeutic application leads to inconsistencies in research findings across different laboratories. As mentioned above, the role of the miR-506 family across various cancer types should be considered more comprehensively. Including more clinical samples and conducting more detailed cell and animal experiments would make experimental results more reliable. More specific studies should be performed according to different cancer types. Although the roles miR-506 in cancer have been partially identified, its exact across various cancer types remains incompletely understood.
Chemotherapy is a primary therapeutic method for the treatment of cancer, typically used as a monotherapy or in combination with surgery or radiation therapy to treat cancer patients (132). Chemotherapy mainly includes classic cytotoxic and molecular-targeted drugs (133), but drug resistance constantly challenges their efficacy. miRNAs play a significant role not only in cancer progression but also in modulating resistance to anticancer drugs. Since 2013, Blower et al. have elucidated the role of miRNAs in modulating chemotherapeutic responsiveness and resistance (134). Subsequent investigations have revealed that miRNAs can counteract chemoresistance by modulating genes implicated in drug resistance, optimizing drug targets, inhibiting cell migration, inducing cell apoptosis, and regulating the tumor microenvironment (135–137). As a microRNA with intricate regulatory functions, miR-506 is a key modulator in counteracting resistance to anticancer therapies. miR-506 is generally downregulated in drug-resistant tumors, whereas its upregulation has been shown to enhance tumor cell sensitivity to chemotherapeutic agents (138). The ability of miR-506 to sensitize drug-resistant cancer cells to chemotherapy is multifaceted, involving the regulation of cell cycle progression, apoptosis, drug transport, DNA repair, and EMT (139). Notably, the interaction between ceRNAs and miR-506 is pivotal in this process.
EMT is a critical biological process in which epithelial cells undergo phenotypic transition, characterized by the loss of cell-cell adhesion and the acquisition of mesenchymal traits, resulting in increased motility and invasiveness of cancer cells (140). In addition to its role in tumor progression, EMT is also implicated in chemoresistance (136, 141). In OC, miR-506 suppresses SNAI2-mediated EMT, thereby restricting the acquisition of stem-like properties in cancer cells and enhancing their response to chemotherapy and radiotherapy (111). Other studies have shown that the Wnt/β-catenin pathway, crucial in EMT and associated with cisplatin resistance, is targeted by miR-506 to suppress EMT and improve OC's susceptibility to PARP inhibitors and cisplatin (142). Overexpression of miR-506-3p in erlotinib-resistant (ER) cells has been demonstrated to downregulate sonic hedgehog (SHH) signaling, thereby reversing EMT-mediated erlotinib resistance (143). However, ceRNAs function as competitive endogenous RNAs that sequester miR-506, attenuating its EMT-inhibitory effects in cancer cells. For instance, FOXD2-AS1 promotes glioma cell proliferation and EMT by sequestering miR-506-5p, thereby modulating the expression of EMT-related genes such as CDH1, CDH2, and VIM (21). Similarly, HOXA11-AS facilitates HCC progression and EMT by regulating the miR-506-3p/Slug axis (144). Additionally, NEAT1 drives chemoresistance to gemcitabine in pancreatic cancer (PC) cells by regulating ZEB2 expression through the miR-506-3p/ZEB2/EMT axis (137). Chemoresistant cancer cells often evade apoptosis, a critical mechanism for treatment efficacy. Silencing of circRNAATL2 diminishes the ceRNA effect on miR-506-3p, curbing paclitaxel (PTX)-resistant OC cell drug resistance and up-regulated miR-506 can also induce cancer cell apoptosis by targeting NFIB, thus delaying disease progression (145). In CRC, overexpression of miR-506 in HCT116-OxR cells inhibits the expression of multidrug resistance protein 1 (MDR1)/P-glycoprotein 1 (P-gp) by down-regulating the Wnt/β-catenin pathway, promoting apoptosis in drug-resistant cells and enhancing sensitivity to oxaliplatin (146). Moreover, miR-506 can increase the sensitivity of drug-resistant cells to drugs by interfering with DNA damage repair. Homologous recombination (HR) is a critical pathway for the repair of DNA double-strand breaks (DSBs), and its dysfunction has been closely associated with increased sensitivity to DNA-damaging agents. RAD51 and RAD17 are two key components of the HR machinery. Previous studies have demonstrated that miR-506 directly targets RAD51, thereby sensitizing cancer cells to DNA damage and significantly enhancing the sensitivity of serous ovarian cancer (SOC) cells to cisplatin and PARP inhibitors (111). Recent evidence suggests that the miR-506-3p/RAD17 axis impairs DNA damage sensing in epithelial ovarian cancer (EOC) cells, leading to G2/M cell cycle arrest. This disruption may force severely damaged mitotic cells to enter mitosis, ultimately triggering mitotic catastrophe, thereby restoring miR-506-3p-induced platinum chemosensitivity (147). Moreover, in certain cancers, miR-506 enhances the sensitivity of cancer cells to anticancer agents by modulating metabolic pathways. miR-506 can restore chemosensitivity to 5-fluorouracil (5-FU) in drug-resistant cancer cells by downregulating polypyrimidine tract-binding protein 1 (PTBP1) expression and suppressing glycolytic activity. However, lncRNA SNHG16 acts as a molecular sponge for miR-506, thereby attenuating its inhibitory effects on PTBP1 and glycolysis (148). Notably, combining miR-506 with other miRNAs may serve as a promising strategy to overcome drug resistance. For example, miR-124-3P and miR-506-3p together can target EZH2, enhancing the sensitivity of sorafenib-resistant thyroid cancer cells to drugs (138).
Radiotherapy remains a cornerstone of cancer treatment and, together with surgery and chemotherapy, constitutes one of the three fundamental pillars of oncologic therapy. Increasing evidence underscores the critical role of microRNA dysregulation in modulating tumor radiosensitivity (101). Studies have demonstrated that DGCR5 knockdown enhances the radiosensitivity of human laryngeal squamous cell carcinoma by upregulating miR-506 and suppressing Wnt signaling (149). In OC, miR-506 suppresses SNAI2-mediated EMT, thereby restricting the acquisition of stem-like properties in cancer cells and enhancing their response to chemotherapy and radiotherapy (111). Fei et al. observed that plasma levels of hsa-miR-506-3p and hsa-miR-140-5p were significantly higher in radiosensitive CRC patients than in radioresistant patients. Subsequent experiments confirmed that overexpression of hsa-miR-506-3p and hsa-miR-140-5p significantly reduced cell proliferation, survival, and clonogenic potential in CRC cells following radiation exposure (101). Overall, research on mir-506 in improving tumor radioresistance remains limited, and further studies are needed to explore its role. Nevertheless, it is undeniable that mir-506 holds great potential as a promising radiosensitizer to enhance the efficacy of radiotherapy.
Although miR-506 holds promise in reversing drug resistance in certain cancers, its efficacy may vary depending on the cancer type and individual patient. Additionally, tumor cells may develop mechanisms to counteract miR-506-mediated reversal of resistance. Further research is needed to gain a deeper understanding of these processes. Table 3 presents an overview of the interplay between miR-506 and chemoresistance in cancer treatment.
Although the miR-506 family exhibits oncogenic or ambiguous roles in a few cancer types, miR-506 is among the most significantly downregulated miRNAs in various malignancies (68, 78, 99, 119). Both in vitro and in vivo studies have consistently demonstrated its potent tumor-suppressive effects, highlighting its potential as a promising candidate for clinical translation. We reviewed the literature to analyze the challenges and strategies in translational medicine for miRNA-506. miR-506 has shown the ability to enhance chemotherapy sensitivity, supporting its use with conventional therapies. Preclinical models of NSCLC have suggested that combining miR-506 with cisplatin or immune checkpoint inhibitors may improve therapeutic efficacy (16). Furthermore, liquid biopsy techniques detecting miR-506 in blood or exosomes could facilitate patient stratification for targeted clinical trials. Phase I/II trial design similar to NCT04285476, which analyzed miRNA profiles in thyroid carcinoma, could be employed to validate miR-506 as a predictive biomarker (150). Although no dedicated clinical trials for miR-506 have been completed, implementing other miRNAs in clinical settings offers valuable insights and potential precedents for future applications. For instance, MRG-106 (Cobomarsen) is currently in Phase I and II clinical trials for treating lymphoma and leukemia (151). MesomiR-1 has been tested in a Phase I clinical trial for mesothelioma patients (152). Miravirsen (miR-122 inhibitor) successfully reduced HCV replication in Phase II trials, highlighting TS-miRNAs' potential in oncology. By integrating innovative drug delivery methods, biomarker stratification, and lessons learned from existing miRNA therapeutics, miR-506 could transition from preclinical research to clinical application. Nevertheless, current studies are limited to some common cancers; even for target genes in these cancers, research on target proteins remains insufficient. Despite evidence of its involvement in regulating gene expression and cell function, a detailed understanding of the precise molecular mechanisms by which miR-506 operates remains limited. This hampers the development of targeted therapeutic strategies involving miR-506, underscoring the need for further research. However, a significant gap remains in understanding the role of miR-506 in drug resistance. Some studies have clarified that the gene can counteract drug resistance through particular target proteins. Still, the precise pathway of influence is inaccurate, which is what future research should focus on. Since EMT is an essential cancer cell phenotype contributing to drug resistance, EMT inhibitors like miR-506 could be employed as crucial components of chemotherapy or targeted treatment medications, enhancing the clinical outcomes of existing cancer therapies. Even though studies on the connection between miR-506, EMT, and cancer cell drug resistance have increased recently, much hasn't been done to date on using miR-506 as a target to reduce drug resistance. Thus, future studies should focus more on this issue. Moreover, several studies use animal or cell models without clinical samples, or the number of clinical samples is not large enough. Some studies neglect to conduct in-depth disease stage and grade analyses, leading to outcome discrepancies. These issues need to be considered in further studies in this area. Notably, the biological function and expression pattern miR-506 exhibit variability across distinct disease states, adding complexity to its potential therapeutic applications. The inconsistency effects observed in specific diseases may be attributed to various factors, including individual differences, disease subtypes, and tissue specificity.
Effective delivery remains a critical barrier due to miR-506’s instability and poor tissue specificity. Lipid nanoparticles successfully delivered miR-34a in clinical trials (NCT01829971), providing a template for miR-506 encapsulation (153). The gelatin nanosphere (GN) delivery system enables the sustained and controlled release of exogenous miR-506, effectively targeting PENK and inactivating the ERK/FOS signaling pathway, thereby suppressing the growth and metastasis of triple-negative breast cancer (TNBC) (31). The delivery of miR-506-3p encapsulated in DOPC nanoliposomes effectively inhibited tumor growth and significantly enhanced the therapeutic effects of olaparib and cisplatin in orthotopic ovarian cancer mouse models (111). Additionally, exosome-mediated delivery of miR-506-3p enhances tumor-targeting efficiency, reducing proliferation and increasing apoptosis in CRC cells through the downregulation of GSTP1 (30). As shown in preclinical breast cancer models, pH-sensitive polymeric nanoparticles could release miR-506 mimics selectively in acidic TME (154). Applying in situ-forming hydrogels in intratumoral drug delivery, including their advantages in reducing systemic toxicity, provides insights into developing clinical application pathways for miRNA-506 (155).
Future research should focus on several key areas to establish miR-506 as a viable therapeutic agent. First, large-scale clinical trials are needed to determine its safety and efficacy. Next, delivery methods must be optimized to ensure tumor-specific targeting. Additionally, biomarker-based screening strategies should be developed for patient selection. Finally, exploring combination therapy approaches will be essential to maximize its clinical utility.
Evidence has demonstrated that miRNAs play dual roles in various human malignancies, functioning as tumor suppressors (TS-miRNAs) or oncogenic miRNAs (oncomiRs). These small regulatory RNAs are involved in cancer initiation and progression by modulating multiple signaling pathways and cellular functions (156). Due to this multifunctionality, miRNAs have emerged as promising therapeutic targets with immense clinical potential. Two principal miRNA-based therapeutic strategies have been developed for cancer treatment: inhibition of overexpressed oncomiRs and restoration of downregulated TS-miRNAs (29). Specifically, three major approaches can be employed to silence oncomiRs, including miRNA sponges, antagomiRs/antimiRs, and CRISPR/Cas9 gene-editing technology (28). They sequester or destroy endogenous intracellular miRNAs, thereby preventing their binding availability to target mRNA. For restoring the function of tumor suppressor miRNAs that are downregulated or lost in cancer cells, miRNA replacement therapy using synthetic miRNA mimics is a viable strategy (157).
Despite the growing enthusiasm for miRNA-based therapeutics, several key challenges must be addressed before miRNA therapies can become routine clinical treatments. First, miRNA therapy faces the pharmacokinetic challenges of poor stability, short half-life, and uneven distribution in vivo. Naked miRNAs are highly susceptible to nuclease degradation, making it difficult to maintain therapeutic concentrations. Additionally, miRNAs are rapidly cleared from circulation and may accumulate in non-target tissues such as the liver and kidneys, reducing therapeutic efficacy (158). Improvement strategies include chemical modifications (2' -O-methylation, nucleic acid modification) and carrier encapsulation (lipid nanoparticles, polymer nanoparticles) to improve stability and targeting (28). The second is the challenge of off-target effects and targeted delivery. While offering therapeutic potential, miRNAs' extensive gene regulatory network also increases the risk of off-target effects that can disrupt normal cellular processes and induce unexpected biological effects. It has been shown that both single - and double-stranded oligonucleotides activate innate immune system responses (159) and can also be neurotoxic (160). Efficient and targeted delivery remains a significant hurdle. Viral vectors (e.g., AAV, lentivirus) offer high efficiency but raise concerns regarding immunogenicity and insertional mutagenesis. Non-viral delivery systems (e.g., lipid nanoparticles, exosomes) provide safer alternatives, but delivery efficiency, potential toxicity, and tissue specificity still need to be further optimized (28). Localized delivery strategies may offer an alternative approach. For example, Inoue et al. (161) demonstrated that a topical formulation containing miR-634 effectively suppressed tumor growth in two skin cancer models without systemic toxicity—however, only a limited number of cancers, such as primary and localized tumors. The third is the complexity of individualized therapy. miRNA expression varies significantly across cancer types, patient genetics, and tumor microenvironments, resulting in inter-patient variability in therapeutic response. Accurate identification of patients' miRNA expression profiles and the development of personalized miRNA therapy are key to future growth. In addition, combination treatment strategies (such as miRNA combined with targeted drugs or immunotherapy) may improve efficacy but still need to be validated in many clinical studies. Finally, there is no universally accepted protocol for miRNA detection, quantification, or therapeutic use, and an incomplete mass production and regulatory framework limits their clinical translation.
The current findings can further explore the molecular mechanisms by which miR-506 affects cancer cell growth. The infinite proliferation of cancer cells is driven by dysregulation of the cell cycle, and miR-506 plays a critical role in suppressing this process by targeting multiple cell cycle-related genes. Previous studies have demonstrated that miR-506 exerts its tumor-suppressive effects by directly targeting the 3′-UTR of CDK4/6, key regulators of the G1/S transition, thereby inducing G1 phase arrest and inhibiting cancer cell proliferation (72, 107). Furthermore, the combined application of miR-506 and miR-143 in the treatment of lung cancer LC and PC synergistically downregulates CDK1, CDK4, and CDK6, blocking both the G1/S and G2/M transitions and inducing robust apoptotic activity (71). One of the key mechanisms by which cancer cells evade immune surveillance is the suppression of apoptosis. miR-506 plays a crucial role in this process by regulating key apoptosis-related proteins. Apoptosis is tightly controlled by various genes and cytokines, with the Bcl-2 family serving as a central regulator. This family comprises two functionally opposing groups: anti-apoptotic proteins (e.g., Bcl-2, Bcl-xL) and pro-apoptotic proteins (e.g., Bax, Bak). These proteins govern mitochondrial membrane permeability and the subsequent release of cytochrome, ultimately determining whether a cell undergoes apoptosis (162). miR-506 influences the apoptotic process of cancer cells by regulating the expression of Bcl-2 family proteins in various cancer types. For instance, in NSCLC cell lines (16) and osteosarcoma cells (163), the overexpression of miR-506 significantly increases the sensitivity of cancer cells to apoptosis, which is closely associated with the upregulation of Bax and the downregulation of Bcl-2 at both the mRNA and protein levels. Additionally, studies have demonstrated that miR-506 can regulate NF-κB p65, specifically elevating reactive oxygen species (ROS) levels in tumor cells and activating the p53 pathway, thereby selectively inducing apoptosis in lung cancer cells (164). Moreover, miR-506 influences cancer cell fate by regulating other apoptosis-related genes. For instance, in Jurkat cells, a T-cell acute lymphoblastic leukemia (T-ALL) cell line, overexpression of miR-506 leads to decreased expression of pro-apoptotic genes such as p53 and p21 while increasing the expression of the anti-apoptotic gene Bcl-2 (165). This suggests that miR-506 may exhibit oncogenic properties in specific cancer types.
Cancer cells rely on DNA repair mechanisms to maintain genomic stability and survival. HR is a critical pathway for the repair of DNA double-strand breaks and is also associated with tumor chemoresistance. Studies have shown that miR-506 specifically targets the 3'-UTR of RAD51, thereby suppressing RAD51 gene expression, impairing the DNA damage response pathway, and enhancing chemosensitivity both in vitro and in vivo, ultimately reducing cancer cell growth (111, 166, 167). Furthermore, miR-506-3p has been identified to target RAD17, a key component of the HR pathway, thereby diminishing cancer cells' ability to sense DNA damage in OC. This leads to disrupting the G2/M cell cycle checkpoint, delaying G2/M cell cycle arrest, and potentially allowing severely DNA-damaged cells to enter mitosis, ultimately resulting in mitotic catastrophe (147). Aberrant human epidermal growth factor receptor-2 (HER2) signaling is implicated in various solid tumors, including BC, GC, biliary tract cancer, CRC, OC, and PC, where HER2 overexpression or amplification promotes tumor growth, invasion, and is associated with poor prognosis (168). Consequently, HER2 has been recognized as a key therapeutic target across multiple cancer types (169, 170). To date, no direct studies have demonstrated that miR-506 regulates HER2 expression or function. However, other miRNAs have been shown to modulate HER2. For instance, miR-101-5p (171), miR-489 (172), and miR-1226-3p (173) have been investigated for their roles in HER2-positive breast cancer. Given the significance of HER2 in tumor progression and treatment resistance, further research is warranted to explore the potential regulatory interactions between miR-506 and HER2, addressing this gap in the field.
Recent studies have highlighted the role of miR-506 in modulating tumor immune evasion and checkpoint regulation, suggesting its potential therapeutic significance. B7-H4, an immune checkpoint protein that negatively regulates T-cell activation, is overexpressed in BC and is associated with a poor immune response. It has been demonstrated that miR-506-3p downregulates the disease-associated rs10754339 "G" allele in B7-H4, thereby suppressing tumor progression (122). In addition, studies have shown that miR-506 promotes tumor immune escape by affecting programmed cell death-1 receptor (PD-1)/programmed cell death-ligand 1 (PD-L1) (174–176). As a potential regulator of immune checkpoint molecules, miR-506 has emerged as a promising candidate for combination therapies with immune checkpoint inhibitors (ICIs). Additionally, miR-506 reprograms tumor-associated macrophages (TAMs) from an immunosuppressive M2 phenotype to a pro-inflammatory M1 phenotype. This shift promotes cytotoxic T lymphocyte (CTL) infiltration, thereby reducing immune evasion in pancreatic ductal adenocarcinoma (PDAC) and strengthening immune-mediated tumor suppression (174).
Metabolic reprogramming, including increased glucose metabolism, fatty acid synthesis, and glutamine metabolism, is a major driver of cancer drug resistance. Among these, the enhancement of glycolysis is particularly critical (177). Tumor cells preferentially rely on glycolysis for energy production, even in the presence of sufficient oxygen, a phenomenon known as the “Warburg effect” (178). Studies have shown that enhanced glycolysis is closely associated with chemoresistance (179). For instance, PTBP1 promotes glycolysis by regulating the expression of pyruvate kinase M2 (PKM2), a key glycolytic enzyme, while miR-506-3p directly targets PTBP1, downregulating its expression and thereby reducing glycolysis, which restores 5-FU sensitivity in GC cells (148). Additionally, circHIPK3 binds to miR-506, suppressing its expression and subsequently upregulating pyruvate dehydrogenase kinase 2 (PDK2), leading to further activation of glycolysis and enhancing the proliferation and metastasis of HCC cells (180). Enolase 1 (ENO1), a key enzyme in glucose metabolism, is frequently overexpressed in various cancers and plays a crucial role in tumor progression (181). Beyond its role in glycolysis, ENO1 also interacts with choline kinase alpha (CHKα), a key enzyme in choline metabolism, stabilizing it and thereby promoting choline phospholipid metabolism and tumor cell proliferation (182). Dysregulated choline metabolism has been proposed as a novel cancer hallmark, with high CHKα expression being strongly associated with tumor progression and poor patient prognosis (182, 183). Furthermore, key glycolytic enzymes and substrates, including hexokinase 2 (HK2) (184), phosphoglycerate mutase 1 (PGAM1) (185), glucose transporters (GLUTs) (186), and lactate dehydrogenase (LDH) (187), are upregulated in various drug-resistant cancer cells, further supporting the critical role of glycolysis in chemoresistance. To date, limited studies have explored how miR-506 modulates metabolic pathways to overcome drug resistance, highlighting a promising avenue for future research
Continued research on miR-506 could provide essential insights into its complex mechanisms in different cancer types, providing significant clinical implications. miR-506 is down-regulated in a variety of cancers, including HCC (57), NSCLC (52), CC (59), and BC (123), and its expression is associated with tumor progression, drug resistance, and prognosis. Therefore, miR-506 can be a biomarker for cancer diagnosis and prognosis assessment (12). For example, in ovarian cancer, patients with low expression of miR-506 had poorer chemotherapy response (104), suggesting that miR-506 could be used to predict chemotherapy sensitivity and guide personalized treatment strategies. miR-506 regulates chemoresistance by modulating EMT, cell cycle progression, apoptosis, DNA damage repair pathways (e.g., RAD17), and key oncogenic signaling pathways (e.g., SHH, WNT/β-catenin) (139). Upregulating miR-506 expression through miRNA mimics or targeted therapeutic strategies may help reverse drug resistance and enhance chemotherapy efficacy, particularly for platinum-based compounds, doxorubicin, and EGFR-TKIs, ultimately improving patient outcomes (111, 142). Given its ability to suppress cancer cell proliferation and promote apoptosis, miR-506 represents a promising therapeutic target for miRNA-based gene therapy. Furthermore, lipid nanoparticles and viral vectors offer potential strategies for targeted miR-506 delivery to tumor tissues, presenting a novel and promising anticancer approach (12, 30, 139). miRNA has unique versatility and unique advantages in cancer treatment. Among them, miR-506 has become a key regulatory molecule in cancer biology and shows good therapeutic potential. Although its clinical application is still early, we are confident in its vast application potential. Future research should promote the advancement of miR therapy, including miR-506, by conducting comprehensive molecular studies, optimizing targeted drug delivery strategies, and launching large-scale randomized clinical trials.
JY: Conceptualization, Funding acquisition, Resources, Supervision, Writing – review & editing. SM: Conceptualization, Writing – original draft, Writing – review & editing. JL: Data curation, Writing – review & editing. JH: Data curation, Writing – review & editing. LL: Writing – review & editing. QZ: Writing – review & editing, Validation. QC: Writing – review & editing. XL: Writing – review & editing, Validation. ZB: Writing – review & editing.
The author(s) declare that financial support was received for the research and/or publication of this article. This work was supported by the Research Fund for Shandong Provincial Natural Science Foundation (No. ZR2024QH068) and the Research Fund for Lin He's Academician Workstation for New Medicine and Clinical Translation at Jining Medical University (No. JYHL2021FMS13).
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that Generative AI was used in the creation of this manuscript. The authors verify and take full responsibility for the use of generative AI in the preparation of this manuscript. Generative AI was used exclusively for language polishing and enhancement of the manuscript to ensure clarity, coherence, and adherence to academic standards of English language expression. No content generation, data analysis, or creative writing was performed by the AI. The author(s) have reviewed and edited the AI-assisted sections to maintain the integrity and accuracy of the research findings and discussions presented in the manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
1. Roy PS, Saikia BJ. Cancer and cure: A critical analysis. Indian J Cancer. (2016) 53:441–2. doi: 10.4103/0019-509x.200658
2. Nowell PC. The clonal evolution of tumor cell populations. Science. (1976) 194:23–8. doi: 10.1126/science.959840
3. Murray CJL, Lopez AD, Naghavi M, Wang H. Global, regional, and national life expectancy, all-cause mortality, and cause-specific mortality for 249 causes of death, 1980-2015: A systematic analysis for the global burden of disease study 2015. . Lancet. (2016) 388:1459–544. doi: 10.1016/s0140-6736(16)31012-1
4. Catela Ivkovic T, Voss G, Cornella H, Ceder Y. Micrornas as cancer therapeutics: A step closer to clinical application. Cancer Lett. (2017) 407:113–22. doi: 10.1016/j.canlet.2017.04.007
5. Yao Q, Chen Y, Zhou X. The roles of micrornas in epigenetic regulation. Curr Opin Chem Biol. (2019) 51:11–7. doi: 10.1016/j.cbpa.2019.01.024
6. Melo SA, Kalluri R. Molecular pathways: micrornas as cancer therapeutics. Clin Cancer Res. (2012) 18:4234–9. doi: 10.1158/1078-0432.CCR-11-2010
7. Almeida MI, Reis RM, Calin GA. Microrna history: discovery, recent applications, and next frontiers. Mutat Res. (2011) 717:1–8. doi: 10.1016/j.mrfmmm.2011.03.009
8. Dornan D, Settleman J. Cancer: mirna addiction - depending on life's little things. Curr Biol. (2010) 20:R812–3. doi: 10.1016/j.cub.2010.08.040
9. Daugaard I, Hansen TB. Biogenesis and function of ago-associated rnas. Trends Genet. (2017) 33:208–19. doi: 10.1016/j.tig.2017.01.003
10. Rupaimoole R, Slack FJ. Microrna therapeutics: towards a new era for the management of cancer and other diseases. Nat Rev Drug Discovery. (2017) 16:203–22. doi: 10.1038/nrd.2016.246
11. Croce CM. Causes and consequences of microrna dysregulation in cancer. Nat Rev Genet. (2009) 10:704–14. doi: 10.1038/nrg2634
12. Li J, Ju J, Ni B, Wang H. The emerging role of mir-506 in cancer. Oncotarget. (2016) 7:62778–88. doi: 10.18632/oncotarget.11294
13. Yoshida K, Yokoi A, Yamamoto Y, Kajiyama H. Chrxq27.3 mirna cluster functions in cancer development. J Exp Clin Cancer Res. (2021) 40:112. doi: 10.1186/s13046-021-01910-0
14. Streicher KL, Zhu W, Lehmann KP, Georgantas RW, Morehouse CA, Brohawn P, et al. A novel oncogenic role for the mirna-506-514 cluster in initiating melanocyte transformation and promoting melanoma growth. Oncogene. (2012) 31:1558–70. doi: 10.1038/onc.2011.345
15. Deng Q, Xie L, Li H. Mir-506 suppresses cell proliferation and tumor growth by targeting rho-associated protein kinase 1 in hepatocellular carcinoma. Biochem Biophys Res Commun. (2015) 467:921–7. doi: 10.1016/j.bbrc.2015.10.043
16. Li ZH, Zhou JH, Chen SN, Pan L, Feng Y, Luo MQ, et al. Microrna-506 has a suppressive effect on the tumorigenesis of nonsmall-cell lung cancer by regulating tubby-like protein 3. Bioengineered. (2021) 12:10176–86. doi: 10.1080/21655979.2021.2001216
17. Wang Y, Lei X, Gao C, Xue Y, Li X, Wang H, et al. Mir-506-3p suppresses the proliferation of ovarian cancer cells by negatively regulating the expression of mtmr6. J Biosci. (2019) 44(6):126. doi: 10.1007/s12038-019-9952-9
18. Huang M, Xie X, Song X, Gu S, Chang X, Su T, et al. Mir-506 suppresses colorectal cancer development by inhibiting orphan nuclear receptor nr4a1 expression. J Cancer. (2019) 10:3560–70. doi: 10.7150/jca.28272
19. Wang XX, Guo GC, Qian XK, Dou DW, Zhang Z, Xu XD, et al. Mir-506 attenuates methylation of lncrna meg3 to inhibit migration and invasion of breast cancer cell lines via targeting sp1 and sp3. Cancer Cell Int. (2018) 18:171. doi: 10.1186/s12935-018-0642-8
20. Li SP, Su HX, Zhao D, Guan QL. Plasma mirna-506 as a prognostic biomarker for esophageal squamous cell carcinoma. Med Sci Monit. (2016) 22:2195–201. doi: 10.12659/msm.899377
21. Zhao J, Zeng XB, Zhang HY, Xiang JW, Liu YS. Long non-coding rna foxd2-as1 promotes cell proliferation, metastasis and emt in glioma by sponging mir-506-5p. Open Med (Wars). (2020) 15:921–31. doi: 10.1515/med-2020-0175
22. Hu L, Cao H, Zheng L, Li R. Bbox1-as1 activates hedgehog signaling pathway to facilitate the proliferation and stemness of esophageal squamous cell carcinoma cells via mir-506-5p/eif5a/ptch1 axis. Curr Mol Pharmacol. (2023) 16:894–904. doi: 10.2174/1874467216666230130132927
23. Chen G, Liao J, Xu Y, Chen Y, Li J, Bu G, et al. Linc01232 promotes metastasis and emt by regulating mir-506-5p/pak1 axis in gastric cancer. Cancer Manag Res. (2022) 14:1729–40. doi: 10.2147/cmar.S352081
24. Chen Z, Zhong T, Li T, Zhong J, Tang Y, Liu Z, et al. Lncrna snhg15 modulates gastric cancer tumorigenesis by impairing mir-506-5p expression. Bioscience Rep. (2021) 41(7):BSR20204177. doi: 10.1042/bsr20204177
25. Cheng RF, Wang J, Zhang JY, Sun L, Zhao YR, Qiu ZQ, et al. Microrna-506 is up-regulated in the development of pancreatic ductal adenocarcinoma and is associated with attenuated disease progression. Chin J Cancer. (2016) 35:64. doi: 10.1186/s40880-016-0128-9
26. Tong JL, Zhang CP, Nie F, Xu XT, Zhu MM, Xiao SD, et al. Microrna 506 regulates expression of ppar alpha in hydroxycamptothecin-resistant human colon cancer cells. FEBS Lett. (2011) 585:3560–8. doi: 10.1016/j.febslet.2011.10.021
27. Shu J, Silva B, Gao T, Xu Z, Cui J. Dynamic and modularized microrna regulation and its implication in human cancers. Sci Rep. (2017) 7:13356. doi: 10.1038/s41598-017-13470-5
28. Reda El Sayed S, Cristante J, Guyon L, Denis J, Chabre O, Cherradi N. Microrna therapeutics in cancer: current advances and challenges. Cancers (Basel). (2021) 13(11):2680. doi: 10.3390/cancers13112680
29. Romano G, Acunzo M, Nana-Sinkam P. Micrornas as novel therapeutics in cancer. Cancers (Basel). (2021) 13(7):1526. doi: 10.3390/cancers13071526
30. Xuan B, Wang Y. Exosome-transmitted mir-506-3p inhibits colorectal cancer cell Malignancy via regulating gstp1. Appl Biochem Biotechnol. (2023) 195:2015–27. doi: 10.1007/s12010-022-04268-x
31. Liu XL, Liu WJ, Chen Q, Liu J, Yang CQ, Zhang G, et al. Mir-506-loaded gelatin nanospheres target penk and inactivate the erk/fos signaling pathway to suppress triple-negative breast cancer aggressiveness. Mol Carcinog. (2021) 60:538–55. doi: 10.1002/mc.23310
32. Borchardt H, Kogel A, Kalwa H, Weirauch U, Aigner A. Therapeutic mir-506-3p replacement in pancreatic carcinoma leads to multiple effects including autophagy, apoptosis, senescence, and mitochondrial alterations in vitro and in vivo. Biomedicines. (2022) 10(7):1692. doi: 10.3390/biomedicines10071692
33. Salmena L, Poliseno L, Tay Y, Kats L, Pandolfi PP. A cerna hypothesis: the rosetta stone of a hidden rna language? Cell. (2011) 146:353–8. doi: 10.1016/j.cell.2011.07.014
34. Qi X, Lin Y, Chen J, Shen B. Decoding competing endogenous rna networks for cancer biomarker discovery. Brief Bioinform. (2020) 21:441–57. doi: 10.1093/bib/bbz006
35. Tang Z, Li X, Zheng Y, Liu J, Liu C, Li X. The role of competing endogenous rna network in the development of hepatocellular carcinoma: potential therapeutic targets. Front Cell Dev Biol. (2024) 12:1341999. doi: 10.3389/fcell.2024.1341999
36. Xu D, Wang L, Pang S, Cao M, Wang W, Yu X, et al. The functional characterization of epigenetically related lncrnas involved in dysregulated cerna-cerna networks across eight cancer types. Front Cell Dev Biol. (2021) 9:649755. doi: 10.3389/fcell.2021.649755
37. Abdollahzadeh R, Daraei A, Mansoori Y, Sepahvand M, Amoli MM, Tavakkoly-Bazzaz J. Competing endogenous rna (Cerna) cross talk and language in cerna regulatory networks: A new look at hallmarks of breast cancer. J Cell Physiol. (2019) 234:10080–100. doi: 10.1002/jcp.27941
38. Liu X, Ma J, Xu F, Li L. Tincr suppresses proliferation and invasion through regulating mir-544a/fbxw7 axis in lung cancer. BioMed Pharmacother. (2018) 99:9–17. doi: 10.1016/j.biopha.2018.01.049
39. Lu Y, Liu Y, Zhang K, Jiang L. Circular rna tlk1 exerts oncogenic functions in hepatocellular carcinoma by acting as a cerna of mir-138-5p. J Oncol. (2022) 2022:2415836. doi: 10.1155/2022/2415836
40. Meng X, Zhang Y, Hu Y, Zhong J, Jiang C, Zhang H. Lncrna ccat1 sponges mir-218-5p to promote emt, cellular migration and invasion of retinoblastoma by targeting mtf2. Cell Signal. (2021) 86:110088. doi: 10.1016/j.cellsig.2021.110088
41. Cui B, Li B, Liu Q, Cui Y. Lncrna ccat1 promotes glioma tumorigenesis by sponging mir-181b. J Cell Biochem. (2017) 118:4548–57. doi: 10.1002/jcb.26116
42. Wang Y, Liu Z, Yao B, Li Q, Wang L, Wang C, et al. Long non-coding rna casc2 suppresses epithelial-mesenchymal transition of hepatocellular carcinoma cells through casc2/mir-367/fbxw7 axis. Mol Cancer. (2017) 16:123. doi: 10.1186/s12943-017-0702-z
43. Liu Q, Guo X, Que S, Yang X, Fan H, Liu M, et al. Lncrna rsu1p2 contributes to tumorigenesis by acting as a cerna against let-7a in cervical cancer cells. Oncotarget. (2017) 8:43768–81. doi: 10.18632/oncotarget.10844
44. Yang C, Yuan W, Yang X, Li P, Wang J, Han J, et al. Circular rna circ-itch inhibits bladder cancer progression by sponging mir-17/mir-224 and regulating P21, pten expression. Mol Cancer. (2018) 17:19. doi: 10.1186/s12943-018-0771-7
45. Xia C, Sun Y, Li Y, Ma J, Shi J. Lncrna ccat1 enhances chemoresistance in hepatocellular carcinoma by targeting qki-5. Sci Rep. (2022) 12:7826. doi: 10.1038/s41598-022-11644-4
46. Gao W, Lin S, Cheng C, Zhu A, Hu Y, Shi Z, et al. Long non-coding rna casc2 regulates sprouty2 via functioning as a competing endogenous rna for mir-183 to modulate the sensitivity of prostate cancer cells to docetaxel. Arch Biochem Biophys. (2019) 665:69–78. doi: 10.1016/j.abb.2018.01.013
47. Tay Y, Rinn J, Pandolfi PP. The multilayered complexity of cerna crosstalk and competition. Nature. (2014) 505:344–52. doi: 10.1038/nature12986
48. Hentze MW, Preiss T. Circular rnas: splicing's enigma variations. EMBO J. (2013) 32:923–5. doi: 10.1038/emboj.2013.53
49. Chen LL, Yang L. Regulation of circrna biogenesis. RNA Biol. (2015) 12:381–8. doi: 10.1080/15476286.2015.1020271
50. Hansen TB, Jensen TI, Clausen BH, Bramsen JB, Finsen B, Damgaard CK, et al. Natural rna circles function as efficient microrna sponges. Nature. (2013) 495:384–8. doi: 10.1038/nature11993
51. Qi X, Zhang DH, Wu N, Xiao JH, Wang X, Ma W. Cerna in cancer: possible functions and clinical implications. J Med Genet. (2015) 52:710–8. doi: 10.1136/jmedgenet-2015-103334
52. Yang M, Zheng E, Ni J, Xu X, Jiang X, Zhao G. Circular rna circfoxo3 facilitate non-small cell lung cancer progression through upregulating hmgb3 via sponging mir-545-3p/mir-506-3p. Tissue Cell. (2022) 75:101702. doi: 10.1016/j.tice.2021.101702
53. Li L, Wei H, Zhang H, Xu F, Che G. Circ_100565 promotes proliferation, migration and invasion in non-small cell lung cancer through upregulating hmga2 via sponging mir-506-3p. Cancer Cell Int. (2020) 20:160. doi: 10.1186/s12935-020-01241-8
54. Chen M, Hu G, Zhou X, Peng Z, Wen W. Hsa_Circ_0016788 regulates hepatocellular carcinoma progression via mir-506-3p/poly-adenosine diphosphate-ribose polymerase. J Gastroenterol Hepatol. (2021) 36:3457–68. doi: 10.1111/jgh.15635
55. Yu Z, Zhang Y, Gao N, Wang X. Overexpression of mir-506 inhibits growth of osteosarcoma through snail2. Am J Transl Res. (2015) 7:2716–23.
56. Chen L, Wang X, Ji C, Hu J, Fang L. Mir-506-3p suppresses papillary thyroid cancer cells tumorigenesis by targeting yap1. Pathol Res Pract. (2020) 216:153231. doi: 10.1016/j.prp.2020.153231
57. Sun P, Fan X, Hu X, Fu X, Wei Q, Zang Y. Circpcnx and pecanex promote hepatocellular carcinoma cell viability by inhibiting mir-506. Cancer Manag Res. (2019) 11:10957–67. doi: 10.2147/CMAR.S232940
58. Lei D, Wang T. Circsypl1 promotes the proliferation and metastasis of hepatocellular carcinoma via the upregulation of ezh2 expression by competing with hsa-mir-506-3p. J Oncol. (2022) 2022:2659563. doi: 10.1155/2022/2659563
59. Shang A, Gu C, Wang W, Wang X, Sun J, Zeng B, et al. Exosomal circpacrgl promotes progression of colorectal cancer via the mir-142-3p/mir-506-3p- tgf-beta1 axis. Mol Cancer. (2020) 19:117. doi: 10.1186/s12943-020-01235-0
60. Kopp F, Mendell JT. Functional classification and experimental dissection of long noncoding rnas. Cell. (2018) 172(3):393–407. doi: 10.1016/j.cell.2018.01.011
61. Lin YH. Crosstalk of lncrna and cellular metabolism and their regulatory mechanism in cancer. Int J Mol Sci. (2020) 21:2947. doi: 10.3390/ijms21082947
62. Zhou M, Dong Z, Hu S, Xiao M. Linc01433 targets mir-506-3p to promote the biological progress of nasopharyngeal carcinoma cells. Eur Arch Otorhinolaryngol. (2021) 278:3363–74. doi: 10.1007/s00405-021-06607-w
63. Ye F, Xu R, Ge Y, Zheng Y, Liu X, Deng P, et al. Linc00963 confers oncogenic properties in glioma by regulating the mir-506/bcat1 axis. Cancer Manage Res. (2020) 12:2339–51. doi: 10.2147/cmar.S246332
64. Huang B, Liu C, Wu Q, Zhang J, Min Q, Sheng T, et al. Long non-coding rna neat1 facilitates pancreatic cancer progression through negative modulation of mir-506-3p. Biochem Biophys Res Commun. (2017) 482:828–34. doi: 10.1016/j.bbrc.2016.11.120
65. Salmena L, Poliseno L, Tay Y, Kats L, Pandolfi PP. A cerna hypothesis: the rosetta stone of a hidden rna language? Cell. (2011) 3):146. doi: 10.1016/j.cell.2011.07.014
66. Santos FN, de Castria TB, Cruz MR, Riera R. Chemotherapy for advanced non-small cell lung cancer in the elderly population. Cochrane Database Syst Rev. (2015) 2015:CD010463. doi: 10.1002/14651858.CD010463.pub2
67. Chen Z, Fillmore CM, Hammerman PS, Kim CF, Wong KK. Non-small-cell lung cancers: A heterogeneous set of diseases. Nat Rev Cancer. (2014) 14:535–46. doi: 10.1038/nrc3775
68. Guo S, Yang P, Jiang X, Li X, Wang Y, Zhang X, et al. Genetic and epigenetic silencing of mircorna-506-3p enhances cotl1 oncogene expression to foster non-small lung cancer progression. Oncotarget. (2017) 8:644–57. doi: 10.18632/oncotarget.13501
69. Yin M, Ren X, Zhang X, Luo Y, Wang G, Huang K, et al. Selective killing of lung cancer cells by mirna-506 molecule through inhibiting nf-kappab P65 to evoke reactive oxygen species generation and P53 activation. Oncogene. (2015) 34:691–703. doi: 10.1038/onc.2013.597
70. Wang K, Wang B, Wang Z, Yang R. Alginic acid inhibits non-small cell lung cancer-induced angiogenesis via activating mir-506 expression. J Natural Medicines. (2021) 75:553–64. doi: 10.1007/s11418-021-01493-2
71. Hossian A, Mackenzie GG, Mattheolabakis G. Combination of mir−143 and mir−506 reduces lung and pancreatic cancer cell growth through the downregulation of cyclin−Dependent kinases. Oncol Rep. (2021) 45(4):2. doi: 10.3892/or.2021.7953
72. Hossian A, Sajib MS, Tullar PE, Mikelis CM, Mattheolabakis G. Multipronged activity of combinatorial mir-143 and mir-506 inhibits lung cancer cell cycle progression and angiogenesis in vitro. Sci Rep. (2018) 8:10495. doi: 10.1038/s41598-018-28872-2
73. Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. (2015) 65:87–108. doi: 10.3322/caac.21262
74. Zheng SG, Xu HX, Liu LN. Management of hepatocellular carcinoma: the role of contrast-enhanced ultrasound. World J Radiol. (2014) 6:7–14. doi: 10.4329/wjr.v6.i1.7
75. Xu H, Luo M, Xiang H, Liao W, Huang H, Wu Y, et al. Mir-506-3p can inhibit cell proliferation and is a diagnostic and prognostic marker of liver cancer. Am J Trans Res. (2021) 13:11531–9.
76. Xiao W, Wang J, Ou C, Zhang Y, Ma L, Weng W, et al. Mutual interaction between yap and C-myc is critical for carcinogenesis in liver cancer. Biochem Biophys Res Commun. (2013) 439:167–72. doi: 10.1016/j.bbrc.2013.08.071
77. Camargo FD, Gokhale S, Johnnidis JB, Fu D, Bell GW, Jaenisch R, et al. Yap1 increases organ size and expands undifferentiated progenitor cells. Curr Biol. (2007) 17:2054–60. doi: 10.1016/j.cub.2007.10.039
78. Wang Y, Cui M, Sun BD, Liu FB, Zhang XD, Ye LH. Mir-506 suppresses proliferation of hepatoma cells through targeting yap mrna 3'utr. Acta Pharmacol Sin. (2014) 35:1207–14. doi: 10.1038/aps.2014.59
79. Zhang X, Tang J, Zhi X, Xie K, Wang W, Li Z, et al. Mir-874 functions as a tumor suppressor by inhibiting angiogenesis through stat3/vegf-a pathway in gastric cancer. Oncotarget. (2015) 6:1605–17. doi: 10.18632/oncotarget.2748
80. Lu Z, Zhang W, Gao S, Jiang Q, Xiao Z, Ye L, et al. Mir-506 suppresses liver cancer angiogenesis through targeting sphingosine kinase 1 (Sphk1) mrna. Biochem Biophys Res Commun. (2015) 468:8–13. doi: 10.1016/j.bbrc.2015.11.008
81. Wang Z, Si M, Yang N, Zhang H, Fu Y, Yan K, et al. Microrna-506 suppresses invasiveness and metastasis of human hepatocellular carcinoma cells by targeting il8. Am J Cancer Res. (2018) 8:1586–94.
82. Su Z, Ye X, Shang L. Mir-506 promotes natural killer cell cytotoxicity against human hepatocellular carcinoma cells by targeting stat3. Yonsei Med J. (2019) 60:22–9. doi: 10.3349/ymj.2019.60.1.22
83. Ruihua H, Mengyi Z, Chong Z, Meng Q, Xin M, Qiulin T, et al. Rhoa regulates resistance to irinotecan by regulating membrane transporter and apoptosis signaling in colorectal cancer. Oncotarget. (2016) 7:87136–46. doi: 10.18632/oncotarget.13548
84. Jiang R, Zhang C, Liu G, Gu R, Wu H. Microrna-101 inhibits proliferation, migration and invasion in osteosarcoma cells by targeting rock1. Am J Cancer Res. (2017) 7:88–97.
85. Zhang L, Zhou H, Wei G. Mir-506 regulates cell proliferation and apoptosis by affecting rhoa/rock signaling pathway in hepatocellular carcinoma cells. Int J Clin Exp Pathol. (2019) 12:1163–73.
86. Xiang H, Luo M, Hou P, Xiao Z, Huang Z, Feng Q, et al. Mir-124-3p combined with mir-506-3p delay hepatic carcinogenesis via modulating sirtuin 1. Biomarkers. (2021) 26:196–206. doi: 10.1080/1354750X.2020.1854856
87. Jiashi W, Chuang Q, Zhenjun Z, Guangbin W, Bin L, Ming H. Microrna-506-3p inhibits osteosarcoma cell proliferation and metastasis by suppressing rab3d expression. Aging. (2018) 10:1294–305. doi: 10.18632/aging.101468
88. Wang D, Bao F, Teng Y, Li Q, Li J. Microrna-506-3p initiates mesenchymal-to-epithelial transition and suppresses autophagy in osteosarcoma cells by directly targeting sphk1. Biosci Biotechnol Biochem. (2019) 83:836–44. doi: 10.1080/09168451.2019.1569496
89. Yao J, Qin L, Miao S, Wang X, Wu X. Overexpression of mir-506 suppresses proliferation and promotes apoptosis of osteosarcoma cells by targeting astrocyte elevated gene-1. Oncol Lett. (2016) 12:1840–8. doi: 10.3892/ol.2016.4827
90. Guo J, Xia N, Yang L, Zhou S, Zhang Q, Qiao Y, et al. Gsk-3beta and vitamin D receptor are involved in beta-catenin and snail signaling in high glucose-induced epithelial-mesenchymal transition of mouse podocytes. Cell Physiol Biochem. (2014) 33:1087–96. doi: 10.1159/000358678
91. Jiashi W, Chuang Q, Zhenjun Z, Guangbin W, Bin L, Ming H. Microrna-506-3p inhibits osteosarcoma cell proliferation and metastasis by suppressing rab3d expression. Aging (Albany NY). (2018) 10:1294–305. doi: 10.18632/aging.101468
92. Ding L, Sun R, Yan Q, Wang C, Han X, Cui Y, et al. Mir-506 exerts antineoplastic effects on osteosarcoma cells via inhibition of the skp2 oncoprotein. Aging (Albany NY). (2021) 13:6724–39. doi: 10.18632/aging.202530
93. Hu M, Yuan X, Liu Y, Tang S, Miao J, Zhou Q, et al. Il-1beta-induced nf-kappab activation down-regulates mir-506 expression to promotes osteosarcoma cell growth through jag1. BioMed Pharmacother. (2017) 95:1147–55. doi: 10.1016/j.biopha.2017.08.120
94. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2015. CA: A Cancer J Clin. (2015) 65(1):5–29. doi: 10.3322/caac.21254
95. Stefansson OA, Esteller M. Ezh2-mediated epigenetic repression of DNA repair in promoting breast tumor initiating cells. Breast Cancer Res. (2011) 13:309–. doi: 10.1186/bcr2871
96. Zhang Y, Lin C, Liao G, Liu S, Ding J, Tang F, et al. Microrna-506 suppresses tumor proliferation and metastasis in colon cancer by directly targeting the oncogene ezh2. Oncotarget. (2015) 6:32586–601. doi: 10.18632/oncotarget.5309
97. Wu M, Zhang Y, Tang A, Tian L. Mir-506 inhibits cell proliferation and invasion by targeting tet family in colorectal cancer. Iran J Basic Med Sci. (2016) 19:316–22.
98. Zu C, Liu T, Zhang G. Microrna-506 inhibits Malignancy of colorectal carcinoma cells by targeting lamc1. Ann Clin Lab Sci. (2016) 46:666–74.
99. Ai L, Luo X, Yan X, Jiang S. Microrna-506-3p inhibits colorectal cancer cell proliferation through targeting enhancer of zeste homologue 2. Bioengineered. (2021) 12:4044–53. doi: 10.1080/21655979.2021.1951930
100. Chen Z, Liu S, Tian L, Wu M, Ai F, Tang W, et al. Mir-124 and mir-506 inhibit colorectal cancer progression by targeting dnmt3b and dnmt1. Oncotarget. (2015) 6:38139–50. doi: 10.18632/oncotarget.5709
101. Liao F, Chen X, Peng P, Dong W. Rwr-algorithm-based dissection of microrna-506-3p and microrna-140-5p as radiosensitive biomarkers in colorectal cancer. Aging (Albany NY). (2020) 12:20512–22. doi: 10.18632/aging.103907
102. Krawczyk P, Powrózek T, Olesiński T, Dmitruk A, Dziwota J, Kowalski D, et al. Evaluation of mir-506 and mir-4316 expression in early and non-invasive diagnosis of colorectal cancer. Int J Colorectal Dis. (2017) 32:1057–60. doi: 10.1007/s00384-017-2814-8
103. Siegel RL, Miller KD, Fuchs HE, Jemal A. Cancer statistics, 2021. CA Cancer J Clin. (2021) 71:7–33. doi: 10.3322/caac.21654
104. Sun Y, Hu L, Zheng H, Bagnoli M, Guo Y, Rupaimoole R, et al. Mir-506 inhibits multiple targets in the epithelial-to-mesenchymal transition network and is associated with good prognosis in epithelial ovarian cancer. J Pathol. (2015) 235:25–36. doi: 10.1002/path.4443
105. Sun Y, Meng C, Liu G. Microrna-506-3p inhibits ovarian cancer metastasis by down-regulating the expression of ezh2. J Cancer. (2022) 13:943–50. doi: 10.7150/jca.66959
106. Integrated genomic analyses of ovarian carcinoma. Nature. (2011) 474:609–15. doi: 10.1038/nature10166
107. Liu G, Sun Y, Ji P, Li X, Cogdell D, Yang D, et al. Mir-506 suppresses proliferation and induces senescence by directly targeting the cdk4/6-foxm1 axis in ovarian cancer. J Pathol. (2014) 233:308–18. doi: 10.1002/path.4348
108. Yi J, Luo J. Sirt1 and P53, effect on cancer, senescence and beyond. Biochim Biophys Acta. (2010) 1804:1684–9. doi: 10.1016/j.bbapap.2010.05.002
109. Xia XY, Yu YJ, Ye F, Peng GY, Li YJ, Zhou XM. Microrna-506-3p inhibits proliferation and promotes apoptosis in ovarian cancer cell via targeting sirt1/akt/foxo3a signaling pathway. Neoplasma. (2020) 67:344–53. doi: 10.4149/neo_2020_190517N441
110. Yang D, Sun Y, Hu L, Zheng H, Ji P, Pecot CV, et al. Integrated analyses identify a master microrna regulatory network for the mesenchymal subtype in serous ovarian cancer. Cancer Cell. (2013) 23:186–99. doi: 10.1016/j.ccr.2012.12.020
111. Liu G, Yang D, Rupaimoole R, Pecot CV, Sun Y, Mangala LS, et al. Augmentation of response to chemotherapy by microrna-506 through regulation of rad51 in serous ovarian cancers. J Natl Cancer Inst. (2015) 107(7):djv108. doi: 10.1093/jnci/djv108
112. Lee SE, Lee JU, Lee MH, Ryu MJ, Kim SJ, Kim YK, et al. Raf kinase inhibitor-independent constitutive activation of yes-associated protein 1 promotes tumor progression in thyroid cancer. Oncogenesis. (2013) 2:e55. doi: 10.1038/oncsis.2013.12
113. Lorenzetto E, Brenca M, Boeri M, Verri C, Modena P. Yap1 acts as oncogenic target of 11q22 amplification in multiple cancer subtypes. Oncotarget. (2014) 5(9):2608–21. doi: 10.18632/oncotarget.1844
114. Deng J, Lei W, Xiang X, Zhang L, Yu F, Chen J, et al. Microrna-506 inhibits gastric cancer proliferation and invasion by directly targeting yap1. Tumour Biol. (2015) 36:6823–31. doi: 10.1007/s13277-015-3364-8
115. Wang L, Zhang Z, Yu X, Huang X, Liu Z, Chai Y, et al. Unbalanced yap-sox9 circuit drives stemness and Malignant progression in esophageal squamous cell carcinoma. Oncogene. (2019) 38:2042–55. doi: 10.1038/s41388-018-0476-9
116. Zhang Z, Ma J, Luan G, Kang L, Su Y, He Y, et al. Mir-506 suppresses tumor proliferation and invasion by targeting foxq1 in nasopharyngeal carcinoma. PloS One. (2015) 10:e0122851. doi: 10.1371/journal.pone.0122851
117. Gong M, Chen C, Zhao H, Sun M, Song M. Mir-506 suppresses cervical cancer cell proliferation both in vitro and in vivo. Neoplasma. (2018) 65:331–8. doi: 10.4149/neo_2018_170112N25
118. Zhang M, Xu Q, Yan S, Li Z, Yan W, Jia X. Suppression of forkhead box Q1 by microrna-506 represses the proliferation and epithelial-mesenchymal transition of cervical cancer cells. Oncol Rep. (2016) 35:3106–14. doi: 10.3892/or.2016.4651
119. Wen SY, Lin Y, Yu YQ, Cao SJ, Zhang R, Yang XM, et al. Mir-506 acts as a tumor suppressor by directly targeting the hedgehog pathway transcription factor gli3 in human cervical cancer. Oncogene. (2015) 34:717–25. doi: 10.1038/onc.2014.9
120. Hu CY, You P, Zhang J, Zhang H, Jiang N. Mir-506-3p acts as a novel tumor suppressor in prostate cancer through targeting galnt4. Eur Rev Med Pharmacol Sci. (2019) 23:5133–8. doi: 10.26355/eurrev_201906_18177
121. Li D, Cao Y, Li J, Xu J, Liu Q, Sun X. Mir-506 suppresses neuroblastoma metastasis by targeting rock1. Oncol Lett. (2017) 13:417–22. doi: 10.3892/ol.2016.5442
122. El Din GS, Youness RA, Assal RA, Gad MZ. Mirna-506-3p directly regulates rs10754339 (a/G) in the immune checkpoint protein B7-H4 in breast cancer. Microrna. (2020) 9:346–53. doi: 10.2174/2211536609666201209152949
123. Arora H, Qureshi R, Park WY. Mir-506 regulates epithelial mesenchymal transition in breast cancer cell lines. PloS One. (2013) 8:e64273. doi: 10.1371/journal.pone.0064273
124. Chang AE, Karnell LH, Menck HR. The national cancer data base report on cutaneous and noncutaneous melanoma: A summary of 84,836 cases from the past decade. The american college of surgeons commission on cancer and the american cancer society. Cancer. (1998) 83:1664–78. doi: 10.1002/(sici)1097-0142(19981015)83:8<1664::aid-cncr23>3.0.co;2-g
125. McLaughlin CC, Wu XC, Jemal A, Martin HJ, Roche LM, Chen VW. Incidence of noncutaneous melanomas in the U. S. Cancer. (2005) 103:1000–7. doi: 10.1002/cncr.20866
126. Falzone L, Romano GL, Salemi R, Bucolo C, Tomasello B, Lupo G, et al. Prognostic significance of deregulated micrornas in uveal melanomas. Mol Med Rep. (2019) 19:2599–610. doi: 10.3892/mmr.2019.9949
127. Wróblewska JP, Lach MS, Ustaszewski A, Kulcenty K, Ibbs M, Jagiełło I, et al. The potential role of selected mirna in uveal melanoma primary tumors as early biomarkers of disease progression. Genes (Basel). (2020) 11(3):271. doi: 10.3390/genes11030271
128. Smit KN, Jager MJ, de Klein A, Kili E. Uveal melanoma: towards a molecular understanding. Prog Retin Eye Res. (2020) 75:100800. doi: 10.1016/j.preteyeres.2019.100800
129. Jager MJ, Shields CL, Cebulla CM, Abdel-Rahman MH, Grossniklaus HE, Stern MH, et al. Uveal melanoma. Nat Rev Dis Primers. (2020) 6:24. doi: 10.1038/s41572-020-0158-0
130. Bertucci A, Kim KH, Kang J, Zuidema JM, Lee SH, Kwon EJ, et al. Tumor-targeting, microrna-silencing porous silicon nanoparticles for ovarian cancer therapy. ACS Appl Mater Interfaces. (2019) 11:23926–37. doi: 10.1021/acsami.9b07980
131. Kafshdooz L, Pourfathi H, Akbarzadeh A, Kafshdooz T, Razban Z, Sheervalilou R, et al. The role of micrornas and nanoparticles in ovarian cancer: A review. Artif Cells Nanomed Biotechnol. (2018) 46:241–7. doi: 10.1080/21691401.2018.1454931
132. Luqmani YA. Mechanisms of drug resistance in cancer chemotherapy. Med Princ Pract. (2005) 14 Suppl 1:35–48. doi: 10.1159/000086183
133. Saijo N. History of cancer chemotherapy; from cytotoxic drugs to molecular targeted therapy. Nihon Rinsho. (2012) 70 Suppl 8:18–23.
134. Blower PE, Verducci JS, Lin S, Zhou J, Chung JH, Dai Z, et al. Microrna expression profiles for the nci-60 cancer cell panel. Mol Cancer Ther. (2007) 6:1483–91. doi: 10.1158/1535-7163.Mct-07-0009
135. Gomes BC, Rueff J, Rodrigues AS. Micrornas and cancer drug resistance. Methods Mol Biol. (2016) 1395:137–62. doi: 10.1007/978-1-4939-3347-1_9
136. Phi LTH, Sari IN, Yang YG, Lee SH, Jun N, Kim KS, et al. Cancer stem cells (Cscs) in drug resistance and their therapeutic implications in cancer treatment. Stem Cells Int. (2018) 2018:5416923. doi: 10.1155/2018/5416923
137. Fu X, Deng X, Xiao W, Huang B, Yi X, Zou Y. Downregulation of neat1 sensitizes gemcitabine-resistant pancreatic cancer cells to gemcitabine through modulation of the mir-506-3p/zeb2/emt axis. Am J Cancer Res. (2021) 11:3841–56.
138. Wang Z, Dai J, Yan J, Zhang Y, Yin Z. Targeting ezh2 as a novel therapeutic strategy for sorafenib-resistant thyroid carcinoma. J Cell Mol Med. (2019) 23:4770–8. doi: 10.1111/jcmm.14365
139. Nasrpour Navaei Z, Taghehchian N, Zangouei AS, Abbaszadegan MR, Moghbeli M. Microrna-506 as a tumor suppressor in anaplastic thyroid carcinoma by regulation of wnt and notch signaling pathways. Iran J Basic Med Sci. (2023) 26:594–602. doi: 10.22038/ijbms.2023.69174.15069
140. Kim DH, Xing T, Yang Z, Dudek R, Chen YH. Epithelial mesenchymal transition in embryonic development, tissue repair and cancer: A comprehensive overview. J Clin Med. (2018) 7:1. doi: 10.3390/jcm7010001
141. Hou Y. Mir-506 inhibits cell proliferation, invasion, migration and epithelial-to-mesenchymal transition through targeting rwdd4 in human bladder cancer. Oncol Lett. (2019) 17:73–8. doi: 10.3892/ol.2018.9594
142. Sun Y, Wu J, Dong X, Zhang J, Meng C, Liu G. Microrna-506-3p increases the response to parp inhibitors and cisplatin by targeting ezh2/β-catenin in serous ovarian cancers. Transl Oncol. (2021) 14:100987. doi: 10.1016/j.tranon.2020.100987
143. Haque I, Kawsar HI, Motes H, Sharma M, Banerjee S, Banerjee SK, et al. . doi: 10.3390/ijms21239307
144. Liu Y, Yan W, Zhou D, Jin G, Cheng X. Long non−Coding rna hoxa11−as accelerates cell proliferation and epithelial−Mesenchymal transition in hepatocellular carcinoma by modulating the mir−506−3p/slug axis. Int J Mol Med. (2020) 46:1805–15. doi: 10.3892/ijmm.2020.4715
145. Ying H, Zhao R, Yu Q, Zhang K, Deng Q. Circatl2 enhances paclitaxel resistance of ovarian cancer via impacting mir-506-3p/nfib axis. Drug Dev Res. (2022) 83:512–24. doi: 10.1002/ddr.21882
146. Zhou H, Lin C, Zhang Y, Zhang X, Zhang C, Zhang P, et al. Mir-506 enhances the sensitivity of human colorectal cancer cells to oxaliplatin by suppressing mdr1/P-gp expression. Cell Prolif. (2017) 50(3):e12341. doi: 10.1111/cpr.12341
147. Bagnoli M, Nicoletti R, Valitutti M, Rizzo A, Napoli A, Montalvao De Azevedo R, et al. Impairment of rad17 functions by mir-506-3p as a novel synthetic lethal approach targeting DNA repair pathways in ovarian cancer. Front Oncol. (2022) 12:923508. doi: 10.3389/fonc.2022.923508
148. Ding Y, Gao S, Zheng J, Chen X. Blocking lncrna-snhg16 sensitizes gastric cancer cells to 5-fu through targeting the mir-506-3p-ptbp1-mediated glucose metabolism. Cancer Metab. (2022) 10:20. doi: 10.1186/s40170-022-00293-w
149. Tang T, Shan G. Dgcr5 promotes cancer stem cell-like properties of radioresistant laryngeal carcinoma cells by sponging mir-506 via wnt pathway. J Cell Physiol. (2019) 234:18423–31. doi: 10.1002/jcp.28478
150. Kim T, Croce CM. Microrna: trends in clinical trials of cancer diagnosis and therapy strategies. Exp Mol Med. (2023) 55:1314–21. doi: 10.1038/s12276-023-01050-9
151. Witten L, Slack FJ. Mir-155 as a novel clinical target for hematological Malignancies. Carcinogenesis. (2020) 41:2–7. doi: 10.1093/carcin/bgz183
152. van Zandwijk N, Pavlakis N, Kao SC, Linton A, Boyer MJ, Clarke S, et al. Safety and activity of microrna-loaded minicells in patients with recurrent Malignant pleural mesothelioma: A first-in-man, phase 1, open-label, dose-escalation study. Lancet Oncol. (2017) 18:1386–96. doi: 10.1016/s1470-2045(17)30621-6
153. Hong DS, Kang YK, Borad M, Sachdev J, Ejadi S, Lim HY, et al. Phase 1 study of mrx34, a liposomal mir-34a mimic, in patients with advanced solid tumours. Br J Cancer. (2020) 122:1630–7. doi: 10.1038/s41416-020-0802-1
154. Qin J, Zhu Y, Zheng D, Zhao Q. Ph-sensitive polymeric nanocarriers for antitumor biotherapeutic molecules targeting delivery. Bio-Design Manufacturing. (2021) 4:612–26. doi: 10.1007/s42242-020-00105-4
155. Shin GR, Kim HE, Kim JH, Choi S, Kim MS. Advances in injectabl e in situ-forming hydrogels for intratumoral treatment. Pharmaceutics. (2021) 13(11):1953. doi: 10.3390/pharmaceutics13111953
156. Otmani K, Lewalle P. Tumor suppressor mirna in cancer cells and the tumor microenvironment: mechanism of deregulation and clinical implications. Front Oncol. (2021) 11:708765. doi: 10.3389/fonc.2021.708765
157. Mollaei H, Safaralizadeh R, Rostami Z. Microrna replacement therapy in cancer. J Cell Physiol. (2019) 234:12369–84. doi: 10.1002/jcp.28058
158. Wang H, Jiang Y, Peng H, Chen Y, Zhu P, Huang Y. Recent progress in microrna delivery for cancer therapy by non-viral synthetic vectors. Adv Drug Delivery Rev. (2015) 81:142–60. doi: 10.1016/j.addr.2014.10.031
159. Kabilova TO, Meschaninova MI, Venyaminova AG, Nikolin VP, Zenkova MA, Vlassov VV, et al. Short double-stranded rna with immunostimulatory activity: sequence dependence. Nucleic Acid Ther. (2012) 22:196–204. doi: 10.1089/nat.2011.0328
160. Lehmann SM, Krüger C, Park B, Derkow K, Rosenberger K, Baumgart J, et al. An unconventional role for mirna: let-7 activates toll-like receptor 7 and causes neurodegeneration. Nat Neurosci. (2012) 15:827–35. doi: 10.1038/nn.3113
161. Inoue J, Fujiwara K, Hamamoto H, Kobayashi K, Inazawa J. Improving the efficacy of egfr inhibitors by topical treatment of cutaneous squamous cell carcinoma with mir-634 ointment. Mol Ther Oncolytics. (2020) 19:294–307. doi: 10.1016/j.omto.2020.10.009
162. Kaloni D, Diepstraten ST, Strasser A, Kelly GL. Bcl-2 protein family: attractive targets for cancer therapy. Apoptosis. (2023) 28:20–38. doi: 10.1007/s10495-022-01780-7
163. Wang X, Ding R, Fu Z, Yang M, Li D, Zhou Y, et al. Overexpression of mir-506-3p reversed doxorubicin resistance in drug-resistant osteosarcoma cells. Front Pharmacol. (2024) 15:1303732. doi: 10.3389/fphar.2024.1303732
164. Yin M, Ren X, Zhang X, Luo Y, Wang G, Huang K, et al. Selective killing of lung cancer cells by mirna-506 molecule through inhibiting nf-κb P65 to evoke reactive oxygen species generation and P53 activation. Oncogene. (2015) 34:691–703. doi: 10.1038/onc.2013.597
165. Rostami Yasuj S, Obeidi N, Khamisipou G, Gharehdaghi Z, Zangeneh Z. Overexpression of mir-506 in jurkat (Acute T cell leukemia) cell line. Iran J Pathol. (2020) 15:282–91. doi: 10.30699/ijp.2020.119627.2298
166. Deng M, Sun J, Xie S, Zhen H, Wang Y, Zhong A, et al. Inhibition of mcm2 enhances the sensitivity of ovarian cancer cell to carboplatin. Mol Med Rep. (2019) 20:2258–66. doi: 10.3892/mmr.2019.10477
167. Liu G, Xue F, Zhang W. Mir-506: A regulator of chemo-sensitivity through suppression of the rad51-homologous recombination axis. Chin J Cancer. (2015) 34:485–7. doi: 10.1186/s40880-015-0049-z
168. Oh DY, Bang YJ. Her2-targeted therapies - a role beyond breast cancer. Nat Rev Clin Oncol. (2020) 17:33–48. doi: 10.1038/s41571-019-0268-3
169. Liu X, Song Y, Cheng P, Liang B, Xing D. Targeting her2 in solid tumors: unveiling the structure and novel epitopes. Cancer Treat Rev. (2024) 130:102826. doi: 10.1016/j.ctrv.2024.102826
170. Cheng X. A comprehensive review of her2 in cancer biology and therapeutics. Genes (Basel). (2024) 15(7):903. doi: 10.3390/genes15070903
171. Normann LS, Haugen MH, Aure MR, Kristensen VN, Mælandsmo GM, Sahlberg KK. Mir-101-5p acts as a tumor suppressor in her2-positive breast cancer cells and improves targeted therapy. Breast Cancer (Dove Med Press). (2022) 14:25–39. doi: 10.2147/bctt.S338404
172. Patel Y, Soni M, Awgulewitsch A, Kern MJ, Liu S, Shah N, et al. Overexpression of mir-489 derails mammary hierarchy structure and inhibits her2/neu-induced tumorigenesis. Oncogene. (2019) 38:445–53. doi: 10.1038/s41388-018-0439-1
173. Mohamadzade Z, Ms B, Ghaemi Z, Hoseinpour P. Cell specific tumor suppressor effect of hsa-mir-1226-3p through downregulation of her2, pik3r2, and akt1 genes. Int J Biochem Cell Biol. (2021) 134:105965. doi: 10.1016/j.biocel.2021.105965
174. Yang T, Han Y, Chen J, Liang X, Sun L. Mir-506 promotes antitumor immune response in pancreatic cancer by reprogramming tumor-associated macrophages toward an M1 phenotype. Biomedicines. (2023) 11(11):2874. doi: 10.3390/biomedicines11112874
175. Assal RA, Elemam NM, Mekky RY, Attia AA, Soliman AH, Gomaa AI, et al. A novel epigenetic strategy to concurrently block immune checkpoints pd-1/pd-L1 and cd155/tigit in hepatocellular carcinoma. Transl Oncol. (2024) 45:101961. doi: 10.1016/j.tranon.2024.101961
176. Zhong S, Golpon H, Zardo P, Borlak J. Mirnas in lung cancer. A systematic review identifies predictive and prognostic mirna candidates for precision medicine in lung cancer. Transl Res. (2021) 230:164–96. doi: 10.1016/j.trsl.2020.11.012
177. Peng J, Cui Y, Xu S, Wu X, Huang Y, Zhou W, et al. Altered glycolysis results in drug-resistant in clinical tumor therapy. Oncol Lett. (2021) 21:369. doi: 10.3892/ol.2021.12630
178. Koppenol WH, Bounds PL, Dang CV. Otto warburg's contributions to current concepts of cancer metabolism. Nat Rev Cancer. (2011) 11:325–37. doi: 10.1038/nrc3038
179. Ma L, Zong X. Metabolic symbiosis in chemoresistance: refocusing the role of aerobic glycolysis. Front Oncol. (2020) 10:5. doi: 10.3389/fonc.2020.00005
180. Yu Q, Chen W, Li Y, He J, Wang Y, Yang S, et al. The novel circular rna hipk3 accelerates the proliferation and invasion of hepatocellular carcinoma cells by sponging the micro rna-124 or micro rna-506/pyruvate dehydrogenase kinase 2 axis. Bioengineered. (2022) 13:4717–29. doi: 10.1080/21655979.2022.2031398
181. Zhou J, Zhang S, Chen Z, He Z, Xu Y, Li Z. Circrna-eno1 promoted glycolysis and tumor progression in lung adenocarcinoma through upregulating its host gene eno1. Cell Death Dis. (2019) 10:885. doi: 10.1038/s41419-019-2127-7
182. Ma Q, Jiang H, Ma L, Zhao G, Xu Q, Guo D, et al. The moonlighting function of glycolytic enzyme enolase-1 promotes choline phospholipid metabolism and tumor cell proliferation. Proc Natl Acad Sci U.S.A. (2023) 120::e2209435120. doi: 10.1073/pnas.2209435120
183. Chen X, Qiu H, Wang C, Yuan Y, Tickner J, Xu J, et al. Molecular structure and differential function of choline kinases chkα and chkβ in musculoskeletal system and cancer. Cytokine Growth Factor Rev. (2017) 33:65–72. doi: 10.1016/j.cytogfr.2016.10.002
184. Liu X, Miao W, Huang M, Li L, Dai X, Wang Y. Elevated hexokinase ii expression confers acquired resistance to 4-hydroxytamoxifen in breast cancer cells. Mol Cell Proteomics. (2019) 18:2273–84. doi: 10.1074/mcp.RA119.001576
185. Huang K, Liang Q, Zhou Y, Jiang LL, Gu WM, Luo MY, et al. A novel allosteric inhibitor of phosphoglycerate mutase 1 suppresses growth and metastasis of non-small-cell lung cancer. Cell Metab. (2019) 30:1107–19.e8. doi: 10.1016/j.cmet.2019.09.014
186. Hamadneh L, Abuarqoub R, Alhusban A, Bahader M. Upregulation of pi3k/akt/pten pathway is correlated with glucose and glutamine metabolic dysfunction during tamoxifen resistance development in mcf-7 cells. Sci Rep. (2020) 10:21933. doi: 10.1038/s41598-020-78833-x
187. Zhou M, Zhao Y, Ding Y, Liu H, Liu Z, Fodstad O, et al. Warburg effect in chemosensitivity: targeting lactate dehydrogenase-a re-sensitizes taxol-resistant cancer cells to taxol. Mol Cancer. (2010) 9:33. doi: 10.1186/1476-4598-9-33
188. Chen G, Wang K, Li G, Wang L, Xiao Y, Chen B. Long noncoding rna lamtor5-as1 interference affects microrna-506-3p/E2f6-mediated behavior of non-small cell lung cancer cells. Oncol Res. (2022) 28:945–59. doi: 10.3727/096504021X16328213967104
189. Zhang N, Li X, Zhang Y, Cheng S, Du J, Liu J. Long noncoding rna kcnq1ot1 promotes cell motility in hepatocellular carcinoma by regulating mir-506-3p/foxq1. J Biol regulators homeostatic Agents. (2020) 34:943–52. doi: 10.23812/20-188-a-55
190. Li S, Hu X, Yu S, Yi P, Chen R, Huang Z, et al. Hepatic stellate cell-released cxcl1 aggravates hcc Malignant behaviors through the mir4435-2hg/mir-506-3p/tgfb1 axis. Cancer Sci. (2022) 114(2):504–20. doi: 10.1111/cas.15605
191. Li C, Lin X, Zhang C, Wan L, Yin J, Wang B. Long non-coding rna fgd5-as1 enhances osteosarcoma cell proliferation and migration by targeting mir-506-3p/rab3d axis. Hum Cell. (2021) 34:1255–65. doi: 10.1007/s13577-021-00536-w
192. Xia X, Li Z, Li Y, Ye F, Zhou X. Lncrna xist promotes carboplatin resistance of ovarian cancer through activating autophagy via targeting mir-506-3p/foxp1 axis. J Gynecol Oncol. (2022) 33:e81. doi: 10.3802/jgo.2022.33.e81
193. Lei R, Xue M, Zhang L, Lin Z. Long noncoding rna malat1-regulated microrna 506 modulates ovarian cancer growth by targeting iaspp. Onco Targets Ther. (2017) 10:35–46. doi: 10.2147/OTT.S112686
194. Yan H, Silva MA, Li H, Zhu L, Li P, Li X, et al. Long noncoding rna dq786243 interacts with mir-506 and promotes progression of ovarian cancer through targeting camp responsive element binding protein 1. J Cell Biochem. (2018) 119:9764–80. doi: 10.1002/jcb.27295
195. Yong W, Yu D, Jun Z, Yachen D, Weiwei W, Midie X, et al. Long noncoding rna neat1, regulated by lin28b, promotes cell proliferation and migration through sponging mir-506 in high-grade serous ovarian cancer. Cell Death Dis. (2018) 9:861. doi: 10.1038/s41419-018-0908-z
196. Zhang YY, Li P, Zhu MZ, Guo Y, Yang J. Linc01308 accelerates the Malignant progression of ovarian cancer by binding to mirna-506. Eur Rev Med Pharmacol Sci. (2019) 23:3253–60. doi: 10.26355/eurrev_201904_17685
197. Liu H, He X, Li T, Qu Y, Xu L, Hou Y, et al. Pcgem1 promotes proliferation, migration and invasion in prostate cancer by sponging mir-506 to upregulate triap1. BMC Urol. (2022) 22:14. doi: 10.1186/s12894-022-00969-x
198. Hu Y, Sun H, Hu J, Zhang X. Lncrna dlx6-as1 promotes the progression of neuroblastoma by activating stat2 via targeting mir-506-3p. Cancer Manag Res. (2020) 12:7451–63. doi: 10.2147/CMAR.S252521
199. Zhao X, Shen F, Yang B. Lncrna linc01410 induced by myc accelerates glioma progression via sponging mir-506-3p and modulating notch2 expression to motivate notch signaling pathway. Cell Mol Neurobiol. (2022) 42:1513–21. doi: 10.1007/s10571-021-01042-1
200. Li H, Li T, Huang D, Zhang P. Long noncoding rna snhg17 induced by yy1 facilitates the glioma progression through targeting mir-506-3p/ctnnb1 axis to activate wnt/beta-catenin signaling pathway. Cancer Cell Int. (2020) 20:29. doi: 10.1186/s12935-019-1088-3
201. Ding Y, Duan H, Lin J, Zhang X. Yy1 accelerates oral squamous cell carcinoma progression through long non-coding rna kcnq1ot1/microrna-506-3p/sypl1 axis. J Ovarian Res. (2022) 15:77. doi: 10.1186/s13048-022-01000-5
202. Han N, Zuo L, Chen H, Zhang C, He P, Yan H. Long non-coding rna homeobox A11 antisense rna (Hoxa11-as) promotes retinoblastoma progression via sponging mir-506-3p. Onco Targets Ther. (2019) 12:3509–17. doi: 10.2147/OTT.S195404
203. Gong SX, Yang FS, Qiu DD. Circptk2 accelerates tumorigenesis of colorectal cancer by upregulating akt2 expression via mir-506-3p. Kaohsiung J Med Sci. (2022) 38:1060–9. doi: 10.1002/kjm2.12589
204. Yang FS, Gong SX, Qiu DD. Circ-malat1 accelerates cell proliferation and epithelial mesenchymal transformation of colorectal cancer through regulating mir-506-3p/kat6b axis. Kaohsiung J Med Sci. (2023) 39:862–72. doi: 10.1002/kjm2.12698
205. Gao F, Xu Q, Tang Z, Zhang N, Huang Y, Li Z, et al. Exosomes derived from myeloid-derived suppressor cells facilitate castration-resistant prostate cancer progression via S100a9/circmid1/mir-506-3p/mid1. J Transl Med. (2022) 20:346. doi: 10.1186/s12967-022-03494-5
206. Ma HB, Yao YN, Yu JJ, Chen XX, Li HF. Extensive profiling of circular rnas and the potential regulatory role of circrna-000284 in cell proliferation and invasion of cervical cancer via sponging mir-506. Am J Transl Res. (2018) 10:592–604.
207. Gong X, Li W, Dong L, Qu F. Circubap2 promotes sema6d expression to enhance the cisplatin resistance in osteosarcoma through sponging mir-506-3p by activating wnt/beta-catenin signaling pathway. J Mol Histol. (2020) 51:329–40. doi: 10.1007/s10735-020-09883-8
208. Zhao Z, Han X, Nie C, Lin S, Wang J, Fang H. Circ_0008784 activates wnt/β-catenin pathway to affect the proliferation and apoptosis of triple-negative breast cancer cells. Pathol - Res Pract. (2023) 241:154185. doi: 10.1016/j.prp.2022.154185
209. Zhu J, Tao L, Jin L. Microrna−506−3p reverses gefitinib resistance in non−Small cell lung cancer by targeting yes−Associated protein 1. Mol Med Rep. (2019) 19:1331–9. doi: 10.3892/mmr.2018.9710
210. Rencüzoğullari Ö, Arisan ED. Palbociclib suppresses the cancer stem cell properties and cell proliferation through increased levels of mir-506 or mir-150 in panc-1 and miapaca-2 cells. Turk J Biol. (2022) 46:342–60. doi: 10.55730/1300-0152.2622
211. Zhang K, Sun X, Sun W, Wang M, Han F. Exosomal microrna-506 inhibits biological activity of lung adenocarcinoma cells and increases sensitivity to cisplatin-based hyperthermia. Cell Signal. (2022) 100:110469. doi: 10.1016/j.cellsig.2022.110469
212. Chen C, Liu Y, Wang H, Zhang X, Shi Y, Chen J. Foxo1-mir-506 axis promotes chemosensitivity to temozolomide and suppresses invasiveness in glioblastoma through a feedback loop of foxo1/mir-506/ets1/foxo1. J Zhejiang Univ Sci B. (2023) 24:698–710. doi: 10.1631/jzus.B2200503
Keywords: cancer, microRNA-506, circular RNAs, long non-coding RNAs, drug resistance
Citation: Mao S, Li J, Huang J, Lv L, Zhang Q, Cheng Q, Liu X, Bi Z and Yao J (2025) Therapeutic potential of microRNA-506 in cancer treatment: mechanisms and therapeutic implications. Front. Oncol. 15:1524763. doi: 10.3389/fonc.2025.1524763
Received: 08 November 2024; Accepted: 17 March 2025;
Published: 03 April 2025.
Edited by:
Mohammad Hasanain, University of Miami Health System, United StatesReviewed by:
Tsung-Chieh Shih, University of Texas MD Anderson Cancer Center, United StatesCopyright © 2025 Mao, Li, Huang, Lv, Zhang, Cheng, Liu, Bi and Yao. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Jing Yao, eWFvc3l4el8xMTFAMTYzLmNvbQ==
†These authors have contributed equally to this work
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.