
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
REVIEW article
Front. Oncol. , 02 April 2025
Sec. Molecular and Cellular Oncology
Volume 15 - 2025 | https://doi.org/10.3389/fonc.2025.1519103
This article is part of the Research Topic Role of nanocarriers and non-coding RNAs in solid malignancies View all 4 articles
 Reem A. Assal1
Reem A. Assal1 Hannah H. Rashwan2
Hannah H. Rashwan2 Zeina I. Zakaria3
Zeina I. Zakaria3 Jana H. Sweillam4
Jana H. Sweillam4 Yasmine M. Fouda5
Yasmine M. Fouda5 Abdelhamid M. Abdelhamid6
Abdelhamid M. Abdelhamid6 Rana A. Youness4*
Rana A. Youness4*Maternally expressed gene 3 (MEG3), a long non-coding RNA, plays a pivotal role in various biological processes, including tumorigenesis. Aberrant expression of MEG3 has been implicated in several cancers, including genitourinary malignancies. This comprehensive review explores the multifaceted functions of MEG3 in the context of genitourinary cancers through unravelling the molecular mechanisms underlying the influence of MEG3 on cellular proliferation, apoptosis, invasion, and metastasis. Additionally, we discuss the potential clinical implications of MEG3 as a biomarker and therapeutic target in genitourinary cancers. By unraveling the intricate role of MEG3 in these biological processes, this review aims to contribute to the development of novel strategies for the diagnosis and treatment of genitourinary malignancies.
Long non-coding RNAs (lncRNAs) are non-protein coding RNA molecules composed of more than 200 nucleotides (1, 2). However, recent discoveries have identified rare exceptions where lncRNAs encode small peptides (2–5). LncRNAs are key players in cancer development and have been validated as diagnostic and prognostic biomarkers in multiple cancers (1, 6–10). The general mechanisms by which lncRNAs act at the cellular level include signaling, scaffolding, decoying, and guiding (11–14). Although most lncRNAs exert their effects through a combination of mechanisms, the functions of lncRNAs can be broadly categorized into three main levels, namely the epigenetic, transcriptional, and post-transcriptional levels (15–18).
First, lncRNAs play a role in the epigenetic regulation of various genes (12, 19). This includes chromatin remodeling by affecting chromatin structure and regulating gene expression, DNA methylation with subsequent downregulation in target gene expression, and histone modification which could silence target genomic regions (20–22).
Second, lncRNAs could regulate gene expression on the nuclear or transcriptional level by interacting with transcription factors acting as scaffolds to prevent their binding to their target genes (22, 23). Such interactions alter the nuclear architecture through the formation of nuclear bodies such as paraspeckles and nuclear speckles which are involved in RNA metabolism and modification of splicing factors, respectively (24). Moreover, such interactions might affect pre-mRNA splicing by interacting with splicing factors, either by acting as decoys to sequester splicing factors or by inducing the phosphorylation of these factors (25–27).
Third, lncRNAs could alter the gene expression on the cytoplasmic or post transcriptional level by affecting the RNA stability and translation, binding with miRNAs to form a sponge RNA that competes with competing endogenous RNA (ceRNAs) (28–31). Such ceRNAs prevent the sponged miRNAs from performing their function or interacting with their respective target proteins (32–34).
Physiologically, the lncRNA activity is tightly regulated (35, 36). However, the dysregulation of lncRNAs in tumors worsens cancer progression and prognosis by interfering with all the reported hallmarks of cancer until now (25, 37, 38).
The lncRNA human maternally expressed gene 3 (MEG3), an imprinted gene located on chromosome 14q32.3 within the Delta-like 1-MEG3 locus (39, 40), is exclusively maternally expressed (41, 42). Furthermore, the absence of a Kozak consensus sequence in the undefined open reading frame classifies it as a non-coding RNA (43, 44). MEG3 transcription is orchestrated by RNA polymerase 2, resulting in the splicing of the gene into 10 exons that encompass five distinct structural motifs, namely M-I to M-V (41, 45, 46). In its mature state, the 1.6 kb MEG3 transcript exhibits polyadenylation at its 3’ end and exists in both the nucleus and cytoplasm (47–49) (Figure 1).
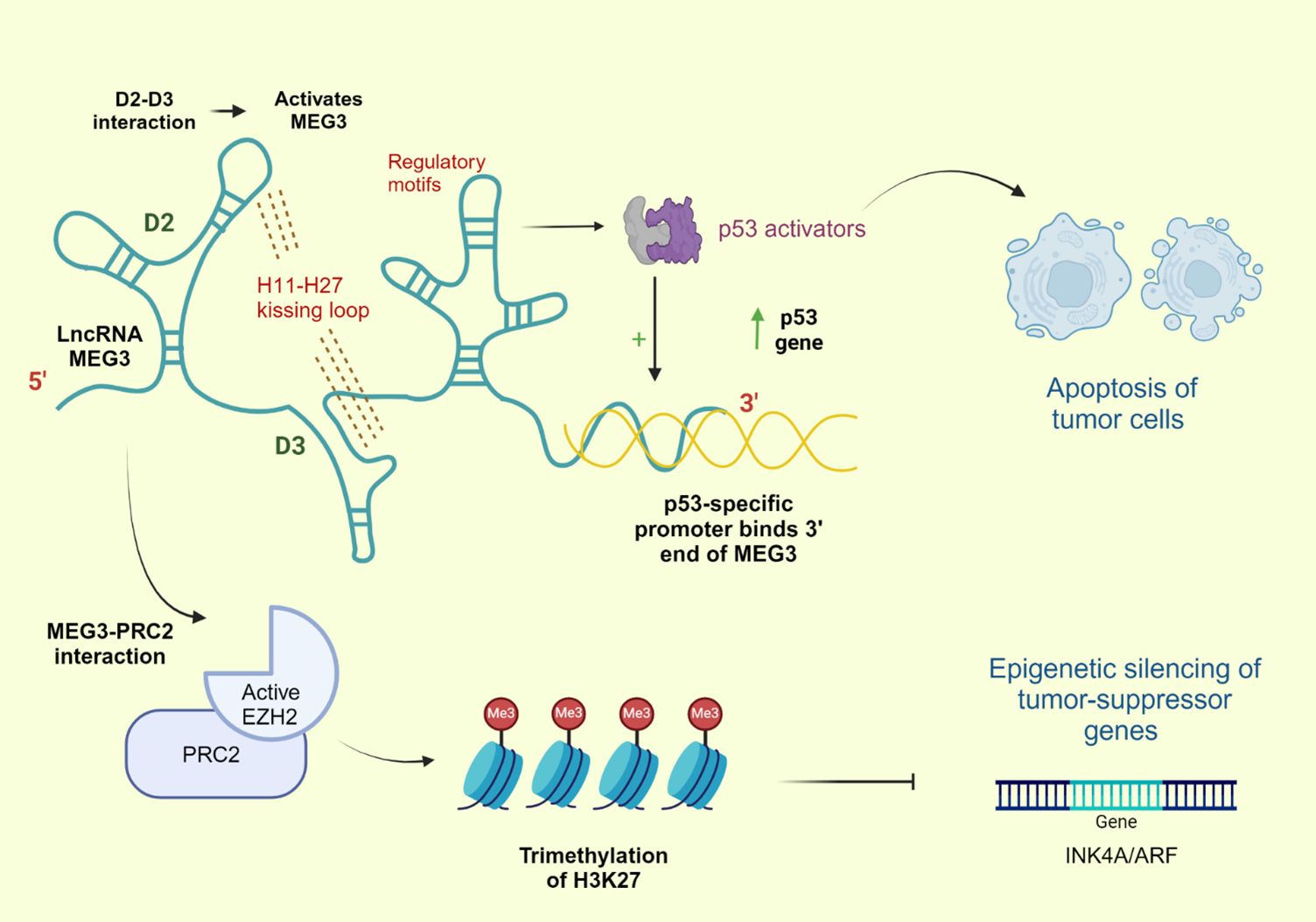
Figure 1. Structure and Function of MEG3. This figure illustrates the structure and function of long non-coding RNA (lncRNA) MEG3. The H11-H27 kissing loop activates MEG3 and the regulatory motifs play a role in activating p53. MEG3's interaction with p53 promotes apoptosis in tumor cells. Additionally, MEG3 interacts with PRC2 (via active EZH2), leading to the trimethylation of H3K27, which results in the epigenetic silencing of tumor-suppressor genes, such as INK4A/ARF.
The regulation of gene expression in the Delta-like 1-MEG3 region is governed by two differentially methylated regions (DMRs) consisting of multiple methylated CpG sites: the intergenic DMR, positioned approximately 13 kb upstream from the transcription start site of MEG3, and the post-fertilization-derived secondary DMR (MEG3-DMR), which overlaps with the promoter 1.5 kb upstream (40, 42). The alternative RNA splicing process of MEG3 results in the production of approximately 12 distinct transcript isoforms within exons 1–4 and 8–10. Within this set of isoforms, variation 1 emerges as the predominant transcript. The selective expression pattern of MEG3 isoforms is tissue- and cell type-specific. All twelve MEG3 isoforms are expressed in the liver of human fetuses, while five specific isoforms, namely MEG3, MEG3b, MEG3d, MEG3e, and MEG3g, are expressed in the tissues of the pituitary gland (42, 50).
Owing to its genomic location and complex molecular interplay, MEG3 plays a pivotal regulatory role in the processes of development and growth (40, 51). In addition, MEG3 plays a role in a wide variety of cellular processes, such as differentiation of osteogenic tissue and progression of bone-related conditions such as osteosarcoma, osteoarthritis, and osteoporosis (52, 53). MEG3 is highly expressed in various normal human tissues, particularly the brain and pituitary gland, especially in the gonadotrophin-producing cells (44, 54). Moreover, MEG3 has been implicated in several diseases, including ischemic neuronal death, atherosclerosis, and type II diabetes mellitus (49, 55–57). Its oncological relevance was first unveiled by Zhang et al., who revealed that MEG3 expression is deficient in pituitary adenomas, and observed that its ectopic expression resulted in the suppression of tumor cell proliferation (49, 54). Recent studies have intriguingly demonstrated a reduction in the MEG3 expression levels in various tumors, where it functions as a tumor suppressor through p53-dependent and p53-independent pathways (58, 59) (Figure 1).
MEG3, as a critical tumor suppressor gene, has the paramount potential to orchestrate many cancer hallmarks. Specifically, it can impede tumor cell proliferation, trigger apoptosis, inhibit invasion and metastasis, hinder angiogenesis, and suppress metabolic reprogramming in tumor cells (49). For instance, MEG3 demonstrated the ability to suppress the oncogenic activity of the proteins c-Myc and β-catenin in liver cancer (58). It also induces G0/G1 cell cycle arrest in prostate cancer cells (60) and G2/M arrest in cervical cancer cells (Figure 2) (61). Restoration of MEG3 expression suppresses tumor growth (54) and induces apoptosis in several human cancer cell lines, such as lung cancer A549 cells (62); SCC-15 and CAL27 tongue squamous cell carcinoma cell lines (63); and gastric cancer cell lines SGC7901, AGS, MGC803, MKN45, and BGC823 (59, 64).
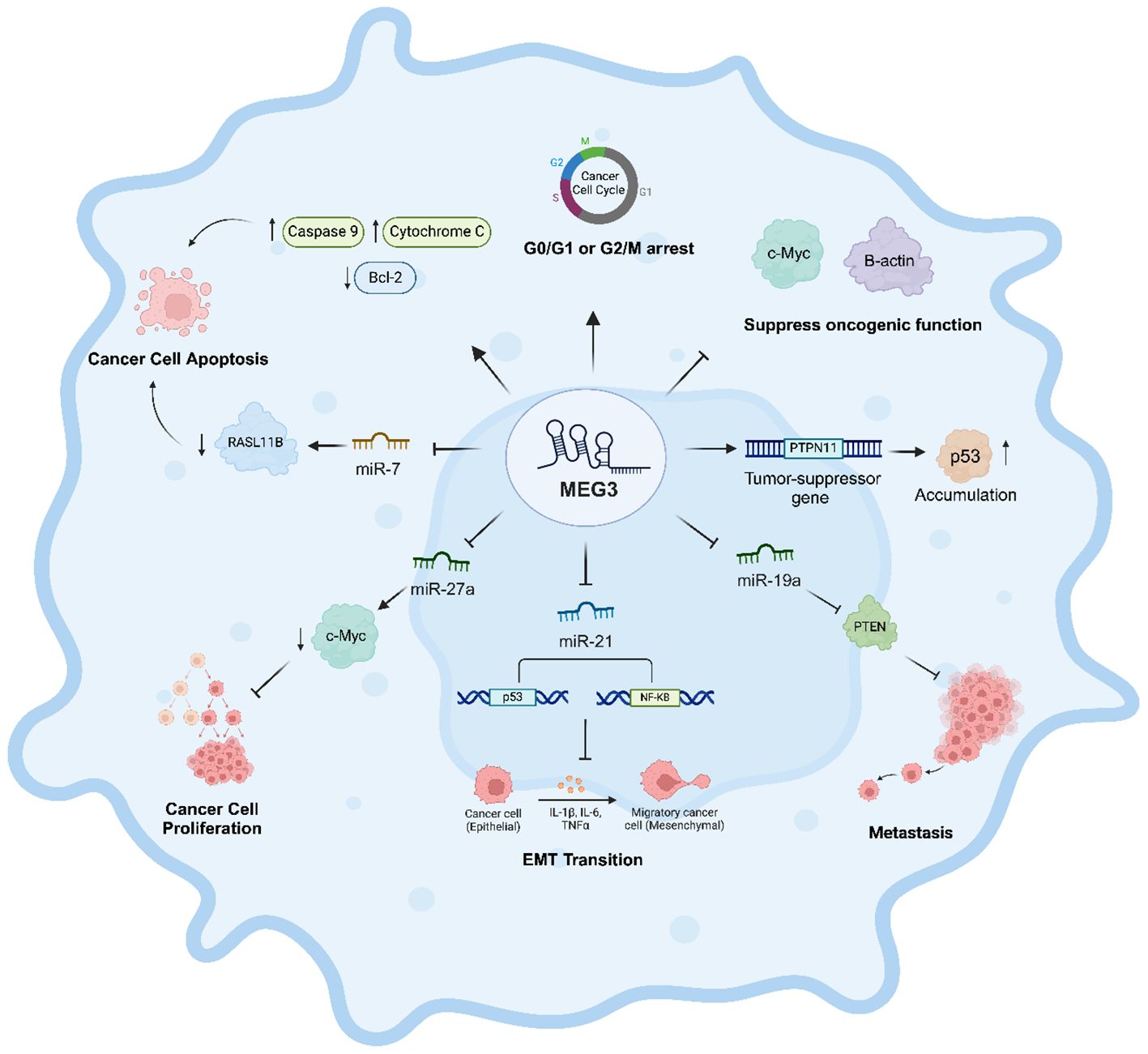
Figure 2. Regulatory role of MEG3 in cancer progression and suppression. This figure depicts the multifaceted tumor-suppressive role of MEG3 in cancer regulation. MEG3 induces cell cycle arrest at the G0/G1 or G2/M phases, restricting uncontrolled proliferation, and promotes apoptosis by upregulating Caspase 9 and Cytochrome C while downregulating the anti-apoptotic protein Bcl-2. It suppresses oncogenic activity by inhibiting c-Myc and B-actin and enhances tumor suppression by upregulating PTPN11, leading to increased p53 accumulation. MEG3 also inhibits metastasis by enhancing the expression of PTEN and modulates key microRNAs (miR-7, miR-27a, miR-19a, and miR-21) to balance oncogenic and tumor-suppressive pathways. Furthermore, it regulates epithelial-mesenchymal transition (EMT) by modulating p53 and NF-κB signaling, preventing the transition of epithelial cancer cells into migratory mesenchymal cells. Through these mechanisms, MEG3 acts as a crucial regulator of cancer progression and suppression.
MEG3 contains microRNA response elements, which enable MEG3 to function as a ceRNA and effectively sponge and sequester miRNAs that engage with diverse genes, proteins, such as p53, enhancer of zeste homologue 2, and nuclear factor-kappa B (Figure 2) (49). Through this mechanism, MEG3 can inhibit the epithelial-mesenchymal transition and metastatic potential of gastric cancer cells by sponging miR-21. This results in increased expression of the epithelial marker E-cadherin and reduced levels of mesenchymal markers, including N-cadherin, Snail, and β-catenin, along with migration-related proteins such as matrix metalloproteinase-2 (MMP-2), MMP-3, and MMP-9 (Figure 2) (42, 65). Furthermore, by sponging miR-19a, MEG3 modulates PTEN expression, thereby inhibiting the migratory and invasive capabilities of glioma cells (Figure 2) (49, 66).
Beyond its role as a ceRNA, MEG3 can influence gene activity through mechanisms such as translation, transcription, post-translational modifications, and epigenetic regulation. Several studies have linked MEG3 dysregulation to poor clinical outcomes and the development of drug resistance. For instance, MEG3 may ameliorate p53 levels by augmenting its transcriptional activity and post-translational modifications (49). It can also decrease the levels of murine double minute 2, an E3 ubiquitin ligase that facilitates p53 ubiquitination and proteasomal degradation, leading to the stabilization of p53 protein and activation of its downstream targets (49, 67). Additionally, MEG3 inhibition has been associated with nickel-induced hypermethylation through increased DNMT3b levels, while the suppression of PHLPP1 transcription has been attributed to decreased interaction between MEG3 and its repressive transcription partner c-Jun (53, 68). Moreover, upregulation of MEG3 expression increases the sensitization of tumor cells to radiation therapy and chemotherapy, thereby boosting the efficacy of current treatment approaches (50). Nevertheless, the functional properties of MEG3 and its involvement in both physiological and pathologic cellular processes are still under active investigation.
Upon examining the tumour-suppressive roles of lncRNAs in genitourinary cancers, MEG3 has emerged as a notable candidate. It is consistently downregulated in tumor tissues compared to normal ones and exhibits significant tumor suppressive roles across all genitourinary cancers. MEG3 primarily functions by promoting apoptosis and by inhibiting proliferation, angiogenesis and metastasis, marking it as a universal tumor suppressor in these cancers.
Among other tumour-suppressive lncRNAs, GAS5 stands out as most similar to MEG3. GAS5 has been identified as a robust tumor suppressor in several genitourinary cancers, including ovarian, RCC, bladder, prostate, and cervical cancers. In ovarian cancer, GAS5 disrupts mitochondrial membrane potential and influences apoptosis pathways, similarly to MEG3, through the promotion of BAX, BAK, and caspases (69, 70). Additionally, it regulates cell proliferation by inhibiting miR-21, thereby increasing SPRY2 expression, crucial for suppressing tumor growth and proliferation (71).
In RCC, GAS5 shares functions with MEG3 by inhibiting cell proliferation, migration, invasion, and inducing apoptosis and cell cycle arrest (Table 1) (72). In bladder cancer, while GAS5 suppresses cell proliferation by modulating cellular components like CCL1, it complements MEG3’s actions by also targeting major oncogenic pathways (73). In prostate cancer, GAS5 functions by inducing apoptosis and repressing androgen receptor (AR) activity, sequestering the androgen/AR complex and inhibiting the AR signaling pathway, which is vital for prostate cancer cell survival and growth (74, 75). Unlike MEG3, GAS5’s mechanism primarily focuses on disrupting hormone-driven growth signals. Furthermore, in cervical cancer, GAS5 acts as a growth and metastasis inhibitor by directly binding and downregulating miR-196a and miR-205, crucial for the proliferation, invasion, and apoptosis of cervical cancer cells (Table 1) (76, 77). GAS5’s broad impact on tumor suppression through miRNA regulation parallels MEG3’s function, though it targets different specific miRNAs, enhancing the collective miRNA regulatory spectrum in this cancer type.
The lncRNA MAGI2-AS3 exhibits tumour-suppressive functions in various cancers by sponging multiple miRNAs, including miR-15-5p, miR-374a-5p, miR-374b-5p, and miR-525-5p. In ovarian cancer, it inhibits MYC signaling, significantly reducing cell proliferation and migration, illustrating its capacity to suppress multiple oncogenic pathways (Table 1) (78, 79). This broad-spectrum targeting complements MEG3’s more focused approach against specific miRNAs and pathways like YBX1 and EGFR. In bladder cancer, MAGI2-AS3 operates through a ceRNA mechanism involving the MAGI2-AS3/miR-31–5p/TNS1 axis, with its downregulation linked to increased migration, proliferation, and invasion, emphasizing its role in metastasis and tumor staging (Table 1) (80).
Another lncRNA, CASC2, acts as a tumor suppressor in RCC by targeting miR-21, which leads to reduced proliferation and migration (81). Its effect on miR-21 provides a specific focal point that complements MEG3’s broader regulatory effects. In bladder cancer, CASC2 modulates the Wnt/β-catenin signaling pathway and promotes cell death, significantly impacting proliferation, migration, and invasion (Table 1) (82). These activities align with MEG3’s, presenting a coordinated defense against key signaling pathways that promote tumor growth. Moreover, PTENP1, functioning analogously to MEG3 in bladder cancer, enhances the expression of tumor suppressors like PDCD4 by suppressing miR-20a, thereby inhibiting tumor growth and metastasis (Table 1) (83).
In cervical cancer, PTENP1 targets key regulatory pathways by enhancing the expression of the tumor suppressor PTEN through the sequestration of miR-106b, inhibiting cell growth, motility, and EMT, crucial factors in cancer progression and metastasis (Table 1) (84, 85). Another lncRNA, TUSC8, demonstrated tumour-suppressive roles in cervical cancer where it focuses on the inhibition of invasion and migration through the miR-641/PTEN axis, similar to PTENP1, but with a specific miRNA target. By upregulating PTEN, TUSC8 curtails pathways essential for cancer cell invasiveness, supporting a role that complements both MEG3 and PTENP1 in modulating the tumor microenvironment (Table 1) (86).
In ovarian cancer, several lncRNAs like HAND2-AS1, HOXA11-AS, and HOTAIRM1 demonstrate tumor-suppressive roles similar to MEG3. HAND2-AS1 reduces metastatic potential by decreasing cell adhesion, migration, and viability, aligning with MEG3’s regulation of epithelial-mesenchymal transition (EMT) (87). HOXA11-AS, downregulated in epithelial ovarian cancer, likely modulates transcriptional pathways, complementing MEG3’s role in metastasis control (88). HOTAIRM1 suppresses proliferation and invasion by enhancing ARHGAP24 expression and sponging miR-106a-5p, mirroring MEG3’s anti-metastatic effects (89). In endometrial cancer, CASC2a and PCDH10 act as tumor suppressors with unique mechanisms. CASC2a, downregulated in the cancer, inhibits growth through epigenetic gene inactivation, contrasting with MEG3’s approach (90, 91).
PCDH10, re-expressed after methylation-induced downregulation, curbs growth and enhances apoptosis, paralleling MEG3’s effects but through methylation changes (92). LINC00261 also reduces proliferation, migration, and invasion by modulating the miRNA/FOXO1 axis, reflecting MEG3’s miRNA modulation strategy (93). TUSC7 and LINC00672 enhance chemotherapeutic responses in endometrial cancer, with TUSC7 increasing sensitivity to treatments like CDDP and Taxol, and LINC00672 promoting chemosensitivity through p53-mediated gene suppression. Both complement MEG3’s effects on cell cycle and apoptosis, offering synergistic benefits (Table 1) (94, 95).
In prostate cancer, lncRNAs such as H19 and PCAT29 play critical but distinct roles in modulating tumor behavior. H19, typically oncogenic in other cancers, serves as a tumor suppressor in metastatic prostate cancer. It specifically represses the effects of transforming growth factor beta 1 (TGFβ1), a key factor in metastasis, and modulates metastasis through the H19-miR-675 axis. This targeted regulation provides a pathway-specific suppression that complements MEG3’s broader influence on cellular apoptosis and invasion (Table 1) (74, 96). Additionally, PCAT29 is recognized as an androgen-regulated tumor suppressor whose expression is inversely correlated with the proliferation and migration of prostate cancer cells. This regulatory role on androgen signals offers a distinct but synergistic mechanism alongside MEG3’s modulation of epigenetic and miRNA pathways, enhancing the overall suppression of tumor growth and metastasis in prostate cancer (Table 1) (97).
In testicular germ cell tumors (TGCT), the lncRNA RFPL3S exhibits a significant reduction, which is associated with hypermethylation and low copy number variations. The downregulation of RFPL3S is linked to enhanced cell invasion and proliferation in TGCT, positioning it as a potential prognostic marker (Table 1) (98). This suggests that while MEG3 modulates miRNA interactions to affect cell survival pathways, RFPL3S’s influence primarily arises from epigenetic changes that affect its expression.
In RCC, the lncRNA SARCC targets VHL-mutant RCC cells, inhibiting proliferation by reducing the stability and expression of the androgen receptor (AR). This suppression of AR decreases HIF-2α and C-MYC expression, focusing on hormonal and hypoxia-related pathways (Table 1) (99). Unlike SARCC, MEG3 has a broader impact on signaling and apoptosis pathways, indicating a more extensive role in RCC tumor suppression. Additionally, NBAT1 functions as a tumor suppressor in RCC, inhibiting proliferation, migration, and invasion, aligning with MEG3’s effects and serving as a prognostic biomarker (Table 1) (100). In other findings, BX357664 acts as a tumor suppressor by modulating epithelial-mesenchymal transition (EMT) and affecting MMP2, MMP9, and TGF-β1/p38/HSP27 signaling. It inhibits proliferation, migration, invasion, and impacts the cell cycle, providing a complementary mechanism to MEG3’s modulation of the cell cycle and apoptosis (101).
In bladder cancer, BANCR increases apoptosis and reduces migration and proliferation when overexpressed, focusing on apoptosis rather than miRNA sponging. Its functional outcomes align with those of MEG3, making them cooperative in curbing tumor growth (102). MDC1-AS inhibits the malignant phenotype through the up-regulation of MDC1, decreasing proliferation, migration, and invasion, and supports MEG3’s effects on cell cycle and apoptosis (Table 1) (103). Furthermore, LINC00312 and LINC00641 target migration and invasion pathways by sponging miR-197-3p. Their approach mirrors MEG3’s strategy of miRNA modulation but targets different specific miRNAs, enhancing the suppression of invasive behavior in bladder cancer (104, 105).
Renal cell cancer, also known as renal cell carcinoma (RCC), is the most prevalent form of kidney cancer in adults, accounting for approximately 90% of cases. It includes clear cell, papillary, and chromophobe subtypes. Its incidence has consistently risen over the past few decades, and it poses a significant threat to global health (116). It has a mortality rate between 30 and 40% and is more prevalent in males than in females.
The aggressiveness of RCC varies considerably between individuals and is affected by several variables. More than 60% of patients with RCC are detected incidentally during routine ultrasound examination, and only 10% of patients exhibit characteristic clinical symptoms (117). Accordingly, a significant proportion of RCC cases are diagnosed at advanced stages, when the tumor has already spread beyond the kidney, resulting in inferior outcomes and limited treatment options. Certain histological RCC subtypes, including clear cell carcinoma and papillary carcinoma, are typically associated with increased aggressiveness and metastasis risk than other subtypes (118).
The role of MEG3 in RCC has been investigated in many studies, shedding light on its impact on cancer progression and potential therapeutic implications. MEG3 is downregulated in RCC tissues and cell lines. Its overexpression resulted in decreased proliferation, migration, and invasion of RCC cells via regulation of ST3Gal1 transcription and EGFR sialylation, thereby influencing the PI3K-AKT pathway (106). In addition, MEG3 was identified as an inducer of RCC cell apoptosis by activating the mitochondrial pathway, as indicated by decreased Bcl-2 expression and elevated levels of cleaved caspase-9 and cytochrome c protein (107) (Figure 3). Moreover, the combined expression of miR-124 and MEG3 emerged as an independent prognostic factor in RCC, where the overexpression of miR-124 or MEG3 inhibited cell proliferation, migration, and invasion, and restrained tumor growth. Additionally, MEG3 induced p53 protein accumulation and regulated the tumor-suppressive gene PTPN11 (119) (Table 2) (Figure 3). MEG3 regulates miR-7/RASL11B signaling in clear cell renal cell carcinoma, thereby promoting apoptosis; inhibiting cell proliferation, migration, and invasion; and inducing G0/G1 cell cycle arrest (120). In papillary renal cell carcinoma, MEG3 expression was downregulated in tumor tissues relative to that in adjacent normal tissues (Table 2) (133). These findings implicate MEG3 as a potential prognostic biomarker and molecular therapeutic target for RCC management.
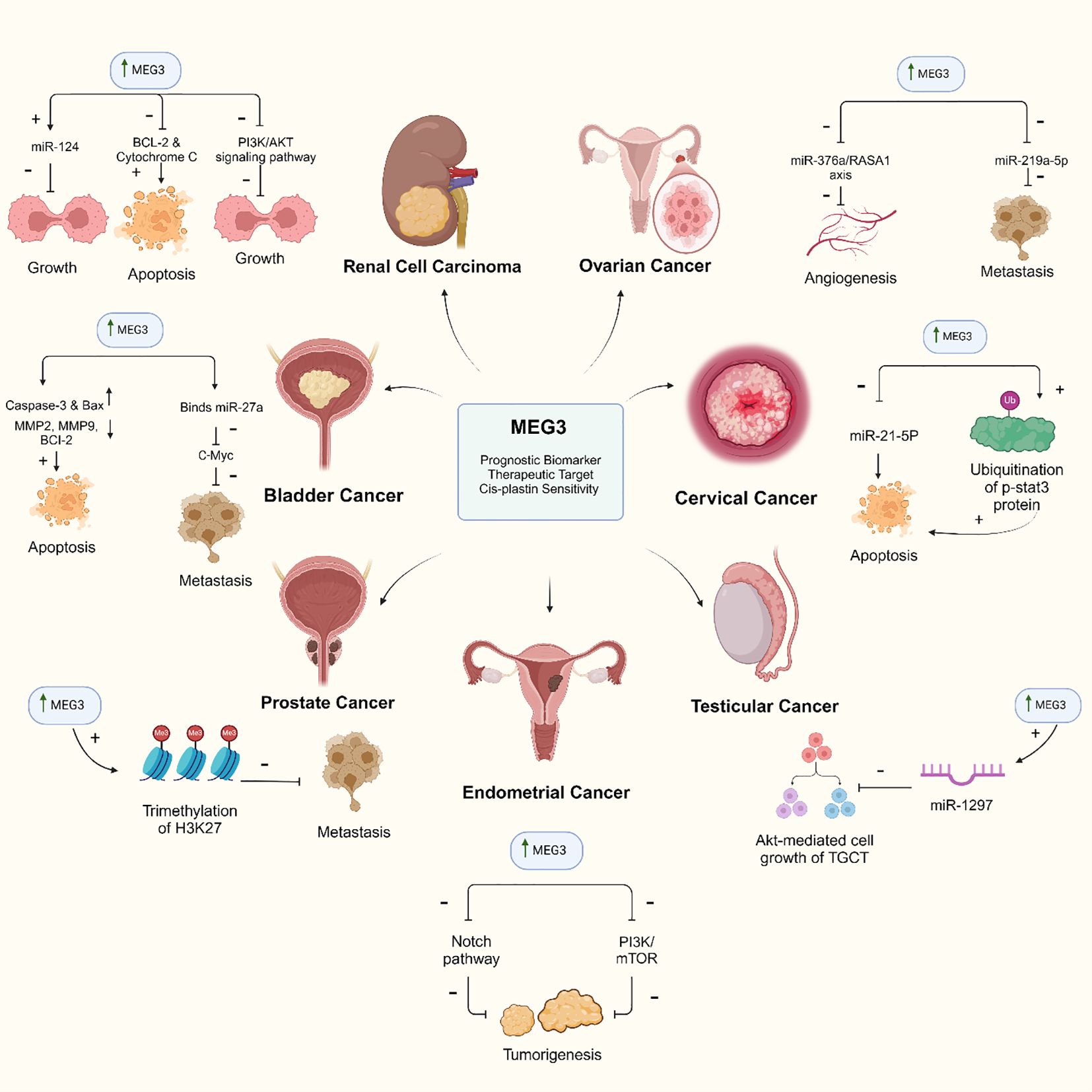
Figure 3. MEG3 lncRNA: a key regulator of tumor suppression across multiple cancer pathways. The diagram illustrates the role of MEG3 in various cancers as a prognostic biomarker, therapeutic target, and enhancer of cis-platin sensitivity. In renal cell carcinoma, MEG3 inhibits the PI3K/AKT signaling pathway, reducing tumor growth and promoting apoptosis. In ovarian cancer, it blocks angiogenesis and metastasis through specific miRNA pathways. MEG3 also induces apoptosis in bladder cancer and inhibits metastasis by downregulating oncogenes. In cervical cancer, MEG3 promotes apoptosis by inhibiting miR-21-5p and STAT3. It reduces cell growth in testicular cancer by regulating miR-1297. In endometrial cancer, MEG3 inhibits tumor growth by suppressing the Notch and PI3K/mTOR pathways, while in prostate cancer, it prevents metastasis through H3K27 trimethylation.
Furthermore, the expression of MEG3 has been studied in various types of renal cell carcinoma (RCC), including clear cell RCC (ccRCC) and papillary RCC (pRCC). In ccRCC, MEG3 expression is generally significantly reduced compared to normal kidney tissue. This downregulation of MEG3 has been associated with proliferation, migration and invasion in patients with ccRCC (120). The loss of MEG3 expression in ccRCC is thought to contribute to the pathogenesis of the disease through mechanisms such as up-regulating RASL11B to induce G0/G1 cell cycle arrest; and promoting cell apoptosis by suppressing miR-7 in ccRCC (120).
The expression patterns of MEG3 in pRCC are less well-defined but suggest a similar trend of downregulation compared to normal tissues (133). However, the extent and implications of MEG3 downregulation in pRCC are not as clearly established as in ccRCC. Studies suggest that the pathways and the impact of MEG3 downregulation in pRCC may differ slightly due to the histological and molecular differences between ccRCC and pRCC (118).Thus, while both ccRCC and pRCC show a trend of reduced MEG3 expression, the details, extent, and implications of this downregulation appear to vary, reflecting the distinct biological behaviors of these RCC subtypes. Further research is needed to fully elucidate the role of MEG3 in different RCC types and its potential as a therapeutic target or biomarker.
Bladder cancer is a common cancer with higher incidence rates in males than those in females. Trends in incidence vary by country, with stabilizing or declining rates among males and increasing rates among females. Mortality rates have been declining, especially in developed countries. The 5-year survival rate for muscle-invasive and metastatic bladder cancer is less than 5% (134).
The most common type of bladder cancer is transitional cell carcinoma. Other less common types include squamous cell carcinoma and adenocarcinoma. The extent of tumor invasion into the bladder wall and the presence of lymph node involvement or distal metastasis determines the tumor stage. The TNM staging system is commonly used to assess the stage of bladder cancer, with higher stages indicating a more aggressive disease. Molecular profiling advancements have expanded our understanding of the aggressiveness of bladder cancer. Mutations or alterations in genes such as TP53, RB1, and FGFR3 can indicate a more aggressive tumor behavior (135).
MEG3 plays a vital role in the suppression of tumors in several forms of cancer, including bladder cancer. In bladder cancer tissues, MEG3 is significantly reduced compared to healthy controls. The fact that MEG3 levels are negatively correlated with those of the autophagy marker LC3-II suggests that MEG3 may play a role in autophagy regulation. In human bladder cancer cell lines, activation of MEG3 inhibited autophagy, whereas inhibition of MEG3 induced autophagy. The downregulation of MEG3 inhibits apoptosis and induces cellular proliferation, but these effects are reversed when autophagy is inhibited (136).
MEG3 serves a crucial role in inhibiting the invasion and metastasis of bladder cancer cells. It functions as a ceRNA by negatively modulating c-Myc through competing with PHLPP2 mRNA for binding with miR-27a. The potent proto-oncogene c-Myc promotes proliferation, invasion, and apoptosis in numerous malignancies. Overexpression of MEG3 inhibited the transcription of c-Myc in a c-Jun-dependent manner, thereby preventing bladder cancer invasion and metastasis (108) (Figure 3).
Overexpression of MEG3 increases the sensitivity of bladder cancer cells to the chemotherapeutic agent, cisplatin. Mutual expression of MEG3 and p53 suggests that they may participate in a positive feedback cycle. In bladder cancer cells treated with cisplatin, MEG3 overexpression induces cell apoptosis, downregulates the anti-apoptotic protein Bcl2, and upregulates the pro-apoptotic proteins cleaved-caspase-3 and Bax. Moreover, overexpression of MEG3 inhibits cell invasion and downregulates MMP2 and MMP9 in bladder cancer cells (121) (Figure 3, Table 2).
Patients with bladder cancer have lower serum levels of MEG3 than healthy subjects and patients with benign disease. Moreover, a decrease in MEG3 expression is associated with a shortened relapse-free time (137) (Table 2). These findings demonstrate the therapeutic potential of MEG3 to improve the efficacy of chemotherapy and inhibit the progression of bladder cancer. They also indicate that MEG3 may serve as a diagnostic and prognostic marker for bladder cancer. Additional research is required to fully elucidate the underlying mechanisms and investigate the therapeutic potential of MEG3 for the management of bladder cancer.
Prostate cancer ranges from indolent, slowly growing tumors with a good prognosis to aggressive, metastasizing cancers with poor outcomes. Prostate cancer aggressiveness is determined by Gleason score, PSA levels, and tumor stage. Higher Gleason scores and PSA values indicate more aggressive illness. Genetic changes and biomarkers may also affect tumor behavior. The aggressiveness of prostate cancer requires early detection and personalized treatment. Surgery, radiation, hormone therapy, and targeted drugs can slow the progression of prostate cancer and improve survival. Early diagnosis and risk classification are key to adapting effective management approaches (138).
Various non-coding RNAs are implicated in the pathogenesis of prostate cancer. Among them, MEG3 was found to be down-regulated in prostate cancer and modulated the miR-9-5p/QKI-5 axis; thus, affecting cell proliferation, migration, invasion, and apoptosis rate (110). Additionally, MEG3 inhibits prostate cancer progression by facilitating H3K27 trimethylation of EN2 by binding to enhancer of zeste homologue 2 (111) (Figure 3). However, both MEG3 polymorphisms, rs11627993 C>T and rs7158663 A>G, did not appear to affect prostate cancer susceptibility (Table 2) (139). Collectively, these studies demonstrate the diverse regulatory functions of MEG3 in prostate cancer, providing potential insights into its impact on cancer progression and facilitating the identification of novel therapeutic targets for this disease.
While testicular cancers can arise from various cell types, the majority are germ cell tumors. These tumors predominantly occur in young men, although they can be diagnosed at any age (140).
Despite being a rare tumor, testicular germ cell tumor (TGCT) was reported as the most commonly occurring cancer in males aged 15–44 years in the US (141). It was predicted that the occurrence of TGCT will remain on the rise with the highest incidence in Hispanics compared to that in any other racial or ethnic group in the US (142). Histologically, the TGCTs can be classified into two groups, seminomas and non-seminomas (143). While seminomas are similar to PGCs, non-seminomas are divided into three classes, undifferentiated embryonal carcinoma, extra-embryonic (yolk sac choriocarcinoma) patterning, or differentiated as a teratoma (143). The European Association of Urology predicted the relapse of 15–30% of patients with TGCT after receiving the first line of chemotherapy and the patients will require further therapies (144). Moreover, primary TGCT can metastasize to several organs including the brain, heart, lung, liver, and lymph nodes (145). In normal spermatogenesis, primordial germ cells (PGCs) develop into gonocytes, followed by multiple developmental steps to eventually maturing into haploid spermatozoa (146, 147). However, if PGCs fail to mature into gonocytes either fetal or postnatally, intratubular germ cell neoplasia, also known as carcinoma in situ develops (148), which is similar to PGCs and their genomes remains undifferentiated and unmethylated (149). Next, intratubular germ cell neoplasia progresses to TGCTs through the loss of PTEN and p21 and the gain of murine double minute 2 expression (150, 151). Moreover, KRAS mutations, which are unique to TGCT, are not present in the intratubular germ cell neoplasia stage (152). Conventionally, the treatment of the TGCTs is a combination of radiotherapy along with chemotherapy, lymph node dissection, or radical orchiectomy.
The MEG3 expression levels are markedly reduced in TGCT, while the expression levels of miR-1297 were unchanged. Moreover, PTEN protein levels were lower in the tumor tissues compared to those in the paired adjacent non-tumor tissues; however, the PTEN transcript levels were unchanged. The authors hypothesized that there is post-transcriptional control of miR-1297 on the PTEN transcripts (Table 2) (12). Bioinformatic analyses revealed that miR-1297 can bind both MEG3 lncRNA and PTEN mRNA in TGCT cells. Overexpressing the MEG3 in TGCT cells led to the competitive binding of MEG3 lncRNA to miR-1297 rather than PTEN, allowing PTEN to suppress the Akt-mediated cell growth of the TGCT. Thus, overexpression of MEG3 could be a valuable tool for the treatment of TGCT (11) (Table 2, Figure 3).
Cervical cancer remains one of the most prevalent gynecologic malignancies worldwide. It ranks fourteenth among all malignancies and fourth among females worldwide (153). Each year, more than 500,000 females are diagnosed with cervical cancer, resulting in more than 300,000 fatalities worldwide (154). Globally, the age-adjusted incidence of cervical cancer was estimated to be 13.1 per 100,000 females, with rates ranging from less than 2 to 75 per 100,000 females, depending on the country. In eastern, western, central, and southern Africa, cervical cancer was the primary cause of cancer-related deaths among females (155).
Human papillomavirus (HPV) infection, particularly HPV types 16 and 18, is a significant etiologic factor for cervical cancer. Infections with high-risk HPV can result in persistent infection with subsequent genomic instability, thereby promoting the transformation of normal cervical cells into cancerous ones (156). Moreover, changes in particular genes and molecular pathways play a crucial role in the aggressiveness of cervical cancer. Mutations or dysregulations in TP53, p16INK4a, and PTEN can disrupt cell cycle control and promote uncontrolled cell proliferation, migration, and invasion (157) (158). In addition, the expression levels of specific biomarkers have been linked to the aggressiveness of cervical cancer as they facilitate extracellular matrix degradation and tissue remodeling. The altered expression of MMPs have been linked to an increase in tumor invasion and metastasis (159). Furthermore, studies have highlighted the function of lncRNAs in modulating the aggressiveness of cervical cancer. Certain lncRNAs have the potential to function as oncogenes or tumor suppressors, influencing cell proliferation, migration, and invasion (160).
MEG3 is a tumor suppressor that influences the growth, proliferation, and apoptosis of cervical cancer cells by modulating multiple molecular pathways and signaling cascades. MEG3 is consistently expressed at low levels in cervical cancer tissues and cell lines. However, increasing the MEG3 levels in cervical cancer cell lines can have significant effects. Elevated MEG3 levels inhibited cell proliferation, induced cell cycle arrest, and promoted apoptosis (122). In addition, MEG3 operates as a cancer suppressor by reducing the levels of miR-21-5p in cervical cancer cell lines (112) (Figure 3). Notably, inhibition of MEG3 in specific cell lines, such as HeLa and CaSki cells, significantly increased the expression of the oncomiR miR-21-5p (112). This highlights the potential function of MEG3 as a tumor suppressor that is capable of inhibiting the growth of cervical cancer (161). Downregulating MEG3 in cervical cancer promoted cellular proliferation, primarily via the regulation of miR-21 (112).
Moreover, MEG3 appears to have diagnostic and prognostic value in cervical cancer. It can be used as a diagnostic marker and prognostic indicator, especially for lymph node metastasis and FIGO staging. Low MEG3 levels, lymph node metastasis, and advanced FIGO stage (III and IV) have been identified as independent cervical cancer prognostic factors (113). A previous study revealed that the lncRNA-MEG3/miR-421/BTG1 pathway modulation by lidocaine inhibited the proliferation of cervical cancer cells and induced apoptosis (Table 2) (123). Additionally, MEG3 induced apoptosis in cervical carcinoma cells through endoplasmic reticulum stress and the miR-7-5p/STC1 axis (124). In cervical cancer tissues and cell lines, the expression of MEG3 and STC1 is diminished, while the expression levels of miR-7-5p are elevated. Overexpression of MEG3 or inhibition of miR-7-5p induced apoptosis in cervical carcinoma cells in response to endoplasmic reticulum stress (124). Furthermore, decreased MEG3 expression appears to be an important indicator of the transition from premalignant cervical lesions to invasive cancer (125). In addition, by modulating the miR-21/PTEN axis, MEG3 promoted cisplatin sensitivity in cervical cancer cells. Conversely, knocking down MEG3 promoted cervical cancer cell proliferation, migration, and inhibited apoptosis, particularly in the presence of cisplatin (126).
Besides being a tumor suppressor, MEG3 inhibits the proliferation of cervical cancer cells by promoting the ubiquitination-mediated degradation of P-STAT3 protein. MEG3 binds to P-STAT3 in cervical cancer cells, leading to its ubiquitination and subsequent degradation, resulting in apoptosis and inhibition of tumor cell proliferation (Table 2, Figure 3). A better understanding of the regulatory axis MEG3-STAT3 in cervical cancer may provide novel insights for potential treatment strategies (127). In conclusion, owing to its significant role in cervical cancer, MEG3 could potentially act as a diagnostic, prognostic, and therapeutic target.
Ovarian cancer is the seventh most prevalent cancer in females and the eighth leading cause of cancer-related mortality, with five-year survival rates below 45%. Although age-standardized rates are stable or declining in the majority of high-income nations, they are on the rise in many low- and middle-income nations (162). Ovarian cancer is a highly aggressive and frequently fatal form of gynecological cancer that originates in the ovaries. Lack of early symptoms contributes to its aggressiveness, resulting in late-stage diagnoses when the tumor has spread beyond the ovaries. The preponderance of ovarian cancers are epithelial ovarian carcinomas, and the most aggressive subtype is high-grade serous carcinoma. Due to the aggressive nature of ovarian cancer, prompt and accurate diagnosis, aggressive treatment strategies such as surgery and chemotherapy, and conducting research to identify effective targeted therapies are essential to improve patient outcomes (163).
Multiple studies have investigated the function of the lncRNA MEG3 in ovarian cancer. First, MEG3 was reported to regulate angiogenesis in ovarian cancer endothelial cells by sponging miRNA-376a and YBX1, with significantly lower expression in ovarian cancer endothelial cells than that in normal ovarian endothelial cells. Overexpressing MEG3 in ovarian cancer endothelial cells decreased tube formation through modulating the miR-376a/RASA1 axis (114) (Figure 3). Second, MEG3 inhibited the progression of ovarian cancer by sponging miR-30e-3p and regulating LAMA4 expression (115). In addition, MEG3 affected the epithelial-mesenchymal transition of ovarian cancer cells by sponging miR-219a-5p and regulating EGFR; low levels of MEG3 and miR-219a-5p were associated with a poor prognosis in patients with ovarian cancer (Figure 3). Knocking down MEG3 and miR-219a-5p in ovarian cancer cells increased the cellular viability, proliferation, invasion, and migration, and decreased apoptosis (Table 2) (129). Another study reported that MEG3 overexpression inhibited the cellular viability and invasion while promoting apoptosis in ovarian cancer through sponging miR-205-5p (130). A recent study in epithelial ovarian carcinoma (EOC) demonstrated that MEG-3 expression was significantly lower in EOC tissues compared to normal ovarian tissues. Receiver operating characteristic (ROC) analysis showed an AUC of 0.831, with high sensitivity (100%) and specificity (97.04%) in distinguishing malignant from normal tissues. Low MEG-3 expression correlated with advanced tumor stages, poor treatment response, and adverse prognostic factors, while high expression was associated with better survival outcomes (p < 0.001). These findings highlight MEG-3 as a promising biomarker for EOC diagnosis, prognosis, and personalized therapy (164). These studies collectively shed light on the diverse regulatory functions of MEG3 in ovarian cancer, indicating its potential role as a therapeutic target for this disease.
Endometrial cancer is the most prevalent gynecological cancer in high-income countries, and its incidence is increasing worldwide. The increasing prevalence of obesity is the most significant underlying cause of endometrial cancer (165). The incidence and mortality rates of endometrial cancer have increased by 21% and >1000%, respectively, over the past two decades (166). Non-endometrioid subtypes of cancer, such as serous and clear cell carcinomas, have a greater propensity for invasion and metastasis. These subtypes are frequently diagnosed at a later stage, which renders treatment as more challenging. The identification of specific molecular and genetic alterations associated with aggressive endometrial cancer has led to advancements in targeted therapies. Accordingly, biomarkers such as p53 mutations and DNA mismatch repair deficiencies may influence treatment decisions (167).
MEG3 was reported to be downregulated in highly invasive, sphere-forming, and TX-resistant endometrial cancer cell derivatives (168). In addition, MEG3 expression was substantially reduced in endometrial cancer tissues compared to that in adjacent normal tissues and normal endometrial cell lines, such as HEC-1A and KLE (132). Another study supported these findings by demonstrating that MEG3 expression was substantially lower in endometrial cancer samples compared to that in normal endometrial tissues (131). Notably, sustained overexpression of MEG3 in HEC-1B and Ishikawa cells induced apoptosis and decreased migration and invasion. In addition, MEG3 inhibited the tumorigenesis and progression of endometrial cancer by repressing the Notch signaling pathway (Table 2, Figure 3). Downregulation of MEG3 was associated with elevated levels of Notch1 and Hes1 proteins (132). In addition, MEG3 inhibited endometrial carcinoma cell proliferation, invasion, and metastasis, while inducing apoptosis and inhibiting the activation of the PI3K/m-TOR signaling pathway (131) (Figure 3). MEG3 could significantly suppress tumor growth as evidenced in tumor xenograft implantation in nude mice (131). Furthermore, MEG3 played a role in modulating PD-L1 expression in aggressive endometrial cancer cells; its expression was regulated by miR-216a (Table 2) (169). Together, these studies highlight the significant downregulation of MEG3 in endometrial cancer; its role in inhibiting cell proliferation, migration, and invasion; and its regulatory roles in modulating the Notch and PI3K pathways, thereby shedding light on potential therapeutic strategies for the treatment of endometrial cancer.
The unique nature of lncRNAs, including their high specificity, and sensitivity in detecting cancer, makes MEG3 a promising diagnostic biomarker (170, 171). MEG3 can be detected in easily accessible samples such as blood or serum, reducing the need for invasive procedures like tissue biopsies (171–174). Additionally, its stability in these fluids and ability to identify early-stage malignancies highlight its value in improving diagnostic accuracy and patient outcomes (164, 171, 173). These advantages suggest that MEG3 could revolutionize cancer diagnostics. However, integrating MEG3 diagnostics into clinical workflows presents challenges including cost-effectiveness and technical feasibility. In terms of cost-effectiveness, the development of MEG3-based assays, such as qRT-PCR or sequencing techniques, may incur higher expenses compared to conventional diagnostic methods like immunohistochemistry.
Moreover, technical feasibility also poses a hurdle for implementing MEG3 diagnostics on a broad scale, requiring standardized protocols to address pre-analytical variability, such as differences in sample collection, storage, and handling, which can impact RNA stability and assay reliability (164). Addressing these challenges through the development of robust, simplified protocols and user-friendly testing kits could significantly enhance the practicality of MEG3 diagnostics. Furthermore, multicenter clinical trials are necessary to validate MEG3’s diagnostic performance and establish its superiority over existing biomarkers. Overcoming these hurdles will pave the way for MEG3 diagnostics to become a practical tool in routine clinical settings.
Based on the aforementioned, MEG3’s tumor-suppressor role in genitourinary cancers positions it as a promising biomarker for early diagnosis and prognosis, while also opening exciting avenues for translating laboratory findings into clinical applications. For instance, a recent study investigating the role of MEG3 in a homogenous series of advanced HGSOC patients demonstrated that exogenous MEG3 expression in vitro and in vivo could reverse the malignant phenotype of HGSOC cells, inhibiting cell proliferation, migration, invasion, and spheroid formation in mice (128). Moreover, elevated MEG3 expression was independently associated with better progression-free and overall survival in patients. These findings are likely mediated through positive regulation of the PTEN network, as PTEN loss is associated with poor prognosis and impaired therapeutic response in HGSOC. These preclinical results reinforce MEG3’s role as a tumor suppressor and highlight its potential as a promising therapeutic target (128).
Another preclinical study demonstrated that upregulation of MEG3 in prostate cancer (PCa) cell lines induced apoptosis and G0/G1 phase arrest, to inhibit the expression of Bcl-2 and Cyclin D1. In vivo, MEG3 overexpression in PC3 cells resulted in significantly reduced tumor weight and volume in mice, reinforcing its tumor-suppressive properties. Overall, these results highlight the therapeutic potential of MEG3, opening up opportunities for its clinical application in prostate cancer treatment (60).
A new study demonstrates that MEG3 is downregulated in PCa tissues, with lower expression levels correlating with poorer prognosis. This study revealed that overexpression of MEG3 inhibited PCa cell proliferation and promoted apoptosis (175). Mechanistically, MEG3 acts as a ceRNA by binding miR-9-5p, thereby alleviating its repressive effect on NDRG1, a protein that regulates cell proliferation and apoptosis. Furthermore, preclinical studies have demonstrated that the upregulation of MEG3 significantly inhibits tumor growth in animal models. All of these findings support the therapeutic promise of MEG3 in PCa treatment (175).
Furthermore, another study MEG3 plays a critical role in bladder cancer by modulating miR-494 and the tumor suppressor gene PTEN. In vitro and in vivo assays have revealed that MEG3 inhibits the expression of miR-494 in bladder cancer cells. Thus, restoration of MEG3 levels holds therapeutic promise for the treatment of bladder cancer (109). Collectively, these findings have propelled MEG3 into the spotlight as a target for therapeutic interventions, encouraging the need for well-designed clinical trials to evaluate its efficacy and safety in patients with genitourinary cancers.
However, while MEG3’s tumor suppressive functions are well-documented in genitourinary cancers, its potential as a standalone therapy remains underexplored and requires further investigation. Current data suggest that MEG3 may be more effective as an adjuvant therapy rather than a primary treatment. For instance, in bladder and cervical cancers, MEG3 overexpression has been shown to improve the efficacy of chemotherapeutic agents like cisplatin by promoting apoptosis and mitigating cellular invasion and survival pathways (121, 126). This synergistic effect highlights MEG3’s potential to enhance chemosensitivity and improve outcomes in advanced or treatment-resistant cancers.
Despite these promising findings, significant challenges remain in MEG3-based therapies into clinical practice. A primary barrier is the effective delivery of MEG3 to target tissues. RNA-based therapeutics, including antisense oligonucleotides (ASOs) and small interfering RNAs (siRNAs), those aimed at modulating MEG3 expression, often encounter issues such as rapid degradation by nucleases, poor cellular uptake, and limited bioavailability at the tumor site (176, 177). These limitations hinder the precise delivery of MEG3-targeting agents to their intended sites of action. Delivery systems such as viral vectors, lipid nanoparticles, and exosome-based platforms offer potential solutions. However, each of these systems presents unique challenges regarding efficiency, safety, and specificity (176, 177). For instance, while lipid nanoparticles can enhance cellular uptake, they may also provoke immune responses that could compromise their therapeutic efficacy (177). Therefore, further research is essential to optimize these delivery mechanisms, ensuring that therapeutic agents, such as MEG3 effectively reach their intended targets within the tumor microenvironment.
Another major concern is the potential for off-target effects. As a ceRNA, MEG3 interacts with various miRNAs and downstream pathways, which could lead to adverse effects if misregulated, leading to undesirable unintended consequences, such as toxicity (177). To mitigate this, future studies should focus on engineering MEG3-based therapies to maximize specificity and minimize unintended interactions. High-throughput screening approaches could play a pivotal role in mitigating off-target effects, thus enhancing the safety of MEG3-based interventions (177).
While preclinical models may demonstrate safety, human responses can vary considerably, as evidenced by immune-related toxicities observed in some clinical trials of RNA-based therapeutics. These toxicities highlight the importance of rigorous validation and meticulous testing in preclinical models and patient cohorts to minimize adverse effects and improve therapeutic outcomes. Finally, while MEG3-centered therapeutic strategies offer significant promise in cancer treatment, addressing the limitations related to delivery mechanisms and off-target effects is essential. Continued research and development efforts are necessary to overcome these challenges, ultimately enhancing the therapeutic benefits of targeting MEG3 in cancer therapy.
This review provides a comprehensive overview of the emerging roles of the tumor suppressor MEG3 in the genitourinary cancers, positioning it as a promising therapeutic target. MEG3’s potential for improving diagnostics and therapeutic strategies offers exciting opportunities for advancing treatment options in these cancers. However, significant challenges remain in translating MEG3-based therapies into clinical practice, particularly in optimizing delivery mechanisms and minimizing off-target effects.
Future research should focus on refining MEG3-based strategies, exploring synergistic effects with conventional treatments like chemotherapy and immunotherapy, and addressing the logistical challenges associated with clinical implementation. By overcoming these barriers, MEG3 could significantly improve outcomes for patients with genitourinary cancers and pave the way for its integration into precision oncology.
RA: Data curation, Writing – original draft, Writing – review & editing. HR: Data curation, Methodology, Writing – original draft, Writing – review & editing. ZZ: Data curation, Methodology, Writing – original draft, Writing – review & editing. JS: Visualization, Writing – original draft, Writing – review & editing. YF: Data curation, Writing – original draft, Writing – review & editing. AA: Data curation, Methodology, Writing – original draft, Writing – review & editing. RY: Conceptualization, Data curation, Formal Analysis, Funding acquisition, Investigation, Methodology, Project administration, Resources, Supervision, Validation, Visualization, Writing – original draft, Writing – review & editing.
The author(s) declare that no financial support was received for the research and/or publication of this article.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
1. Selem NA, Youness RA, Gad MZ. What is beyond LncRNAs in breast cancer: A special focus on colon cancer-associated Transcript-1 (CCAT-1). Noncoding RNA Res. (2021) 6:174–86. doi: 10.1016/j.ncrna.2021.11.001
2. Youness RA, Gad MZ. Long non-coding RNAs: Functional regulatory players in breast cancer. Noncoding RNA Res. (2019) 4:36–44. doi: 10.1016/j.ncrna.2019.01.003
3. Wei C, Xu Y, Shen Q, Li R, Xiao X, Saw PE, et al. Role of long non-coding RNAs (lncRNAs) in cancer: from subcellular localization to nanoparticles (NPs)-mediated targeted regulation. Mol Therapy-Nucleic Acids. (2023). doi: 10.1016/j.omtn.2023.07.009
4. Zhang Y. LncRNA-encoded peptides in cancer. J Hematol Oncol. (2024) 17:66. doi: 10.1186/s13045-024-01591-0
5. Dawoud A, Zakaria Ihab Z, Rashwan Hisham H, Braoudaki M, Youness RA, et al. Circular RNAs: New layer of complexity evading breast cancer heterogeneity. Noncoding RNA Res. (2023) 8:60–74. doi: 10.1016/j.ncrna.2022.09.011
6. Lee B, Mazar J, Aftab MN, Qi F, Shelley J, Li J-L, et al. Long noncoding RNAs as putative biomarkers for prostate cancer detection. J Mol Diagnostics. (2014) 16:615–26. doi: 10.1016/j.jmoldx.2014.06.009
7. Beylerli O, Gareev I, Sufianov A, Ilyasova T, Guang Y. Long noncoding RNAs as promising biomarkers in cancer. Non-coding RNA Res. (2022) 7:66–70. doi: 10.1016/j.ncrna.2022.02.004
8. Mekky RY, Ragab MF, Manie T, Attia AA, Youness RA. MALAT-1: Immunomodulatory lncRNA hampering the innate and the adaptive immune arms in triple negative breast cancer. Trans Oncol. (2023) 31:101653. doi: 10.1016/j.tranon.2023.101653
9. Youness RA, Hafez HM, Khallaf E, Assal RA, Motaal Abdel A, Gad MZ. The long noncoding RNA sONE represses triple-negative breast cancer aggressiveness through inducing the expression of miR-34a, miR-15a, miR-16, and let-7a. J Cell Physiol. (2019) 234:20286–97. doi: 10.1002/jcp.v234.11
10. Youness R, Assal R, Hafez M, Motaal AA, Gad M. PO-347 sONE, a novel tumour suppressor lncRNA, with diminished expression level in young triple negative breast cancer (TNBC) patients with lymphnode metastasis and large tumour size. ESMO Open. (2018) 3:A364–5. doi: 10.1136/esmoopen-2018-EACR25.859
11. Qian Y, Shi L, Luo Z. Long non-coding RNAs in cancer: implications for diagnosis, prognosis, and therapy. Front Med. (2020) 7:612393. doi: 10.3389/fmed.2020.612393
12. Elmasri RA, Rashwan AA, Gaber SH, Rostom MM, Karousi P, Yasser MB, et al. Puzzling out the role of MIAT LncRNA in hepatocellular carcinoma. Non-coding RNA Res. (2024) 9:547–59. doi: 10.1016/j.ncrna.2024.01.006
13. Soliman R, Youness R, El-Shazly M, Handoussa H, Gad M. Regulatory interacting network between the immunomodulatory non-coding RNAs: miR-17-5p, MALAT1 and H19 lncRNAs in modulating the tumour microenvironment in TNBC. Ann Oncol. (2019) 30:xi57. doi: 10.1093/annonc/mdz452.032
14. Abdelhamid A, Zeinelabdeen Y, Manie T, Khallaf E, Assal R, Youness R. miR-17-5p/STAT3/H19: A novel regulatory axis tuning ULBP2 expression in young breast cancer patients. Pathology-Research Pract. (2024), 155638. doi: 10.1016/j.prp.2024.155638
15. Zhang X, Wang W, Zhu W, Dong J, Cheng Y, Yin Z, et al. Mechanisms and functions of long non-coding RNAs at multiple regulatory levels. Int J Mol Sci. (2019) 20:5573. doi: 10.3390/ijms20225573
16. Selem NA, Nafae H, Manie T, Youness RA, Gad MZ. Let-7a/cMyc/CCAT1/miR-17-5p circuit re-sensitizes atezolizumab resistance in triple negative breast cancer through modulating PD-L1. Pathol Res Pract. (2023) 248:154579. doi: 10.1016/j.prp.2023.154579
17. Youness RA, Assal RA, Motaal Abdel A, Gad MZ. A novel role of sONE/NOS3/NO signaling cascade in mediating hydrogen sulphide bilateral effects on triple negative breast cancer progression. Nitric Oxide. (2018) 80:12–23. doi: 10.1016/j.niox.2018.07.004
18. El-Khazragy N, Esmaiel MA, Mohamed MM, Hassan NS. Upregulation of long noncoding RNA Lnc-IRF2-3 and Lnc-ZNF667-AS1 is associated with poor survival in B-chronic lymphocytic leukemia. Int J Lab Hematol. (2020) 42:284–91.
19. Zeinelabdeen Y, Abaza T, Yasser MB, Elemam NM, Youness RA. MIAT LncRNA: A multifunctional key player in non-oncological pathological conditions. Noncoding RNA Res. (2024) 9:447–62. doi: 10.1016/j.ncrna.2024.01.011
20. Wang W, Min L, Qiu X, Wu X, Liu C, Ma J. Biological function of long non-coding RNA (LncRNA) Xist. Front Cell Dev Biol. (2021) 9:645647. doi: 10.3389/fcell.2021.645647
21. Morlando M, Fatica A. Alteration of epigenetic regulation by long noncoding RNAs in cancer. Int J Mol Sci. (2018) 19:570. doi: 10.3390/ijms19020570
22. Assal RA, Elemam NM, Mekky RY, Attia AA, Soliman AH, Gomaa AI, et al. A novel epigenetic strategy to concurrently block immune checkpoints PD-1/PD-L1 and CD155/TIGIT in hepatocellular carcinoma. Trans Oncol. (2024) 45:101961. doi: 10.1016/j.tranon.2024.101961
23. Dawoud A, Elmasri RA, Mohamed AH, Mahmoud A, Rostom MM, Youness RA. Involvement of circRNAs in regulating the “New generation of cancer hallmarks”: A special depiction on hepatocellular carcinoma. Crit Rev Oncol Hematol. (2024) 196:104312. doi: 10.1016/j.critrevonc.2024.104312
24. El-Aziz MKA, Dawoud A, Kiriacos CJ, Fahmy SA, Hamdy NM, Youness RA. Decoding hepatocarcinogenesis from a noncoding RNAs perspective. J Cell Physiol. (2023) 238:1982–2009. doi: 10.1002/jcp.v238.9
25. Ahmad M, Weiswald LB, Poulain L, Denoyelle C, Meryet-Figuiere M. Involvement of lncRNAs in cancer cells migration, invasion and metastasis: cytoskeleton and ECM crosstalk. J Exp Clin Cancer Res. (2023) 42:173. doi: 10.1186/s13046-023-02741-x
26. Sun Q, Hao Q, Prasanth KV. Nuclear long noncoding RNAs: key regulators of gene expression. Trends Genet. (2018) 34:142–57. doi: 10.1016/j.tig.2017.11.005
27. Sun W, Yang Y, Xu C, Guo J. Regulatory mechanisms of long noncoding RNAs on gene expression in cancers. Cancer Genet. (2017) 216:105–10. doi: 10.1016/j.cancergen.2017.06.003
28. Soliman AH, Youness RA, Sebak AA, Handoussa H. Phytochemical-derived tumor-associated macrophage remodeling strategy using Phoenix dactylifera L. boosted photodynamic therapy in melanoma via H19/iNOS/PD-L1 axis. Photodiagnosis Photodyn Ther. (2023) 44:103792. doi: 10.1016/j.pdpdt.2023.103792
29. Abdel-Latif M, Riad A, Soliman RA, Elkhouly AM, Nafae H, Gad MZ, et al. MALAT-1/p53/miR-155/miR-146a ceRNA circuit tuned by methoxylated quercitin glycoside alters immunogenic and oncogenic profiles of breast cancer. Mol Cell Biochem. (2022) 477:1281–93. doi: 10.1007/s11010-022-04378-4
30. Nafea H, Youness RA, Abou-Aisha K, Gad MZ. LncRNA HEIH/miR-939-5p interplay modulates triple-negative breast cancer progression through NOS2-induced nitric oxide production. J Cell Physiol. (2021) 236:5362–72. doi: 10.1002/jcp.v236.7
31. Abdallah RM, Elkhouly AM, Soliman RA, Mechawy El N, Sebaei El A, Motaal AA, et al. Hindering the Synchronization Between miR-486-5p and H19 lncRNA by Hesperetin Halts Breast Cancer Aggressiveness Through Tuning ICAM-1. Anticancer Agents Med Chem. (2022) 22:586–95. doi: 10.2174/1871520621666210419093652
32. Noh JH, Kim KM, McClusky WG, Abdelmohsen K, Gorospe M. Cytoplasmic functions of long noncoding RNAs. Wiley Interdiscip Reviews: RNA. (2018) 9:e1471. doi: 10.1002/wrna.2018.9.issue-3
33. Dykes IM, Emanueli C. Transcriptional and post-transcriptional gene regulation by long non-coding RNA. Genomics Proteomics Bioinf. (2017) 15:177–86. doi: 10.1016/j.gpb.2016.12.005
34. Shi X, Sun M, Wu Y, Yao Y, Liu H, Wu G, et al. Post-transcriptional regulation of long noncoding RNAs in cancer. Tumor Biol. (2015) 36:503–13. doi: 10.1007/s13277-015-3106-y
35. Abaza T, El-Aziz MKA, Daniel KA, Karousi P, Papatsirou M, Fahmy SA, et al. Emerging role of circular RNAs in hepatocellular carcinoma immunotherapy. Int J Mol Sci. (2023) 24. doi: 10.3390/ijms242216484
36. Youness RA, Hassan HA, Abaza T, Hady AA, Magdoub El HM, Ali M, et al. A Comprehensive Insight and In Silico Analysis of CircRNAs in Hepatocellular Carcinoma: A Step toward ncRNA-Based Precision Medicine. Cells. (2024) 13. doi: 10.3390/cells13151245
37. Bartonicek N, Maag JL, Dinger ME. Long noncoding RNAs in cancer: mechanisms of action and technological advancements. Mol Cancer. (2016) 15:1–10. doi: 10.1186/s12943-016-0530-6
38. Youness R, Attia A. 42P sONE and MEG3: Novel immunomodulatory tumor suppressor LncRNAs in HCV-induced hepatocellular carcinoma patients. ESMO Open. (2024) 9. doi: 10.1016/j.esmoop.2024.102377
39. Wylie AA, Murphy SK, Orton TC, Jirtle RL. Novel imprinted DLK1/GTL2 domain on human chromosome 14 contains motifs that mimic those implicated in IGF2/H19 regulation. Genome Res. (2000) 10:1711–8. doi: 10.1101/gr.161600
40. Al-Rugeebah A, Alanazi M, Parine NR. MEG3: an oncogenic long non-coding RNA in different cancers. Pathol Oncol Res. (2019) 25:859–74. doi: 10.1007/s12253-019-00614-3
41. Miyoshi N, Wagatsuma H, Wakana S, Shiroishi T, Nomura M, Aisaka K, et al. Identification of an imprinted gene, Meg3/Gtl2 and its human homologue MEG3, first mapped on mouse distal chromosome 12 and human chromosome 14q. Genes to Cells. (2000) 5:211–20. doi: 10.1046/j.1365-2443.2000.00320.x
42. Rashwan H, Taher A, Hassan H, Awaji A, Kiriacos C, Assal R, et al. Harnessing the supremacy of MEG3 LncRNA to defeat gastrointestinal Malignancies. Pathology-Research Pract. (2024), 155223. doi: 10.1016/j.prp.2024.155223
43. Zhang X, Gejman R, Mahta A, Zhong Y, Rice KA, Zhou Y, et al. Maternally expressed gene 3, an imprinted noncoding RNA gene, is associated with meningioma pathogenesis and progression. Cancer Res. (2010) 70:2350–8. doi: 10.1158/0008-5472.CAN-09-3885
44. Balik V, Srovnal J, Sulla I, Kalita O, Foltanova T, Vaverka M, et al. MEG3: a novel long noncoding potentially tumour-suppressing RNA in meningiomas. J neuro-oncol. (2013) 112:1–8. doi: 10.1007/s11060-012-1038-6
45. Zhou Y, Zhang X, Klibanski A. MEG3 noncoding RNA: a tumor suppressor. J Mol Endocrinol. (2012) 48:R45–53. doi: 10.1530/JME-12-0008
46. Sherpa C, Rausch JW, Le Grice SF. Structural characterization of maternally expressed gene 3 RNA reveals conserved motifs and potential sites of interaction with polycomb repressive complex 2. Nucleic Acids Res. (2018) 46:10432–47. doi: 10.1093/nar/gky722
47. Li R, Fang L, Pu Q, Bu H, Zhu P, Chen Z, et al. MEG3-4 is a miRNA decoy that regulates IL-1β abundance to initiate and then limit inflammation to prevent sepsis during lung infection. Sci Signaling. (2018) 11:eaao2387. doi: 10.1126/scisignal.aao2387
48. Azam S, Hou S, Zhu B, Wang W, Hao T, Bu X, et al. Nuclear retention element recruits U1 snRNP components to restrain spliced lncRNAs in the nucleus. RNA Biol. (2019) 16:1001–9. doi: 10.1080/15476286.2019.1620061
49. Zhang L, Zhao F, Li W, Song G, Kasim V, Wu S. The biological roles and molecular mechanisms of long non-coding RNA MEG3 in the hallmarks of cancer. Cancers. (2022) 14:6032. doi: 10.3390/cancers14246032
50. Xu J, Wang X, Zhu C, Wang K. A review of current evidence about lncRNA MEG3: A tumor suppressor in multiple cancers. Front Cell Dev Biol. (2022) 10:997633. doi: 10.3389/fcell.2022.997633
51. da Rocha ST, Edwards CA, Ito M, Ogata T, Ferguson-Smith AC. Genomic imprinting at the mammalian Dlk1-Dio3 domain. Trends Genet. (2008) 24:306–16. doi: 10.1016/j.tig.2008.03.011
52. Sun H, Peng G, Wu H, Liu M, Mao G, Ning X, et al. Long non−coding RNA MEG3 is involved in osteogenic differentiation and bone diseases. Biomed Rep. (2020) 13:15–21. doi: 10.3892/br.2020.1305
53. Hussain MS, Majami AA, Ali H, Kumar G, Almalki WH, Alzarea SI, et al. The complex role of MEG3: An emerging long non-coding RNA in breast cancer. Pathology-Research Pract. (2023), 154850. doi: 10.1016/j.prp.2023.154850
54. Zhang X, Zhou Y, Mehta KR, Danila DC, Scolavino S, Johnson SR, et al. A pituitary-derived MEG3 isoform functions as a growth suppressor in tumor cells. J Clin Endocrinol Metab. (2003) 88:5119–26. doi: 10.1210/jc.2003-030222
55. Zhang Y, Liu X, Bai X, Lin Y, Li Z, Fu J, et al. Melatonin prevents endothelial cell pyroptosis via regulation of long noncoding RNA MEG3/miR-223/NLRP3 axis. J pineal Res. (2018) 64:e12449. doi: 10.1111/jpi.2018.64.issue-2
56. Sathishkumar C, Prabu P, Mohan V, Balasubramanyam M. Linking a role of lncRNAs (long non-coding RNAs) with insulin resistance, accelerated senescence, and inflammation in patients with type 2 diabetes. Hum Genomics. (2018) 12:1–9. doi: 10.1186/s40246-018-0173-3
57. Yan H, Rao J, Yuan J, Gao L, Huang W, Zhao L, et al. Long non-coding RNA MEG3 functions as a competing endogenous RNA to regulate ischemic neuronal death by targeting miR-21/PDCD4 signaling pathway. Cell Death Dis. (2017) 8:3211. doi: 10.1038/s41419-017-0047-y
58. Zheng Q, Lin Z, Xu J, Lu Y, Meng Q, Wang C, et al. Long noncoding RNA MEG3 suppresses liver cancer cells growth through inhibiting β-catenin by activating PKM2 and inactivating PTEN. Cell Death Dis. (2018) 9:253. doi: 10.1038/s41419-018-0305-7
59. Zhang Z, Shi S, Li J, Costa M. Long non-coding RNA MEG3 in metal carcinogenesis. Toxics. (2023) 11:157. doi: 10.3390/toxics11020157
60. Luo G, Wang M, Wu X, Tao D, Xiao X, Wang L, et al. Long non-coding RNA MEG3 inhibits cell proliferation and induces apoptosis in prostate cancer. Cell Physiol Biochem. (2015) 37:2209–20. doi: 10.1159/000438577
61. Qin R, Chen Z, Ding Y, Hao J, Hu J, Guo F. Long non-coding RNA MEG3 inhibits the proliferation of cervical carcinoma cells through the induction of cell cycle arrest and apoptosis. Neoplasma. (2013) 60:486–92. doi: 10.4149/neo_2013_063
62. Kruer TL, Dougherty SM, Reynolds L, Long E, Silva T, Lockwood WW, et al. Expression of the lncRNA maternally expressed gene 3 (MEG3) contributes to the control of lung cancer cell proliferation by the Rb pathway. PloS One. (2016) 11:e0166363. doi: 10.1371/journal.pone.0166363
63. Jia LF, Wei SB, Gan YH, Guo Y, Gong K, Mitchelson K, et al. Expression, regulation and roles of miR-26a and MEG3 in tongue squamous cell carcinoma. Int J Cancer. (2014) 135:2282–93. doi: 10.1002/ijc.v135.10
64. Sun M, Xia R, Jin F, Xu T, Liu Z, De W, et al. Downregulated long noncoding RNA MEG3 is associated with poor prognosis and promotes cell proliferation in gastric cancer. Tumor Biol. (2014) 35:1065–73. doi: 10.1007/s13277-013-1142-z
65. Jiao J, Zhang S. Long non−coding RNA MEG−3 suppresses gastric carcinoma cell growth, invasion and migration via EMT regulation. Mol Med Rep. (2019) 20:2685–93. doi: 10.3892/mmr.2019.10515
66. Qin N, Tong GF, Sun LW, Xu XL. Long noncoding RNA MEG3 suppresses glioma cell proliferation, migration, and invasion by acting as a competing endogenous RNA of miR-19a. Oncol Res. (2017) 25:1471. doi: 10.3727/096504017X14886689179993
67. Huang Z-F, Tang Y-L, Shen Z-L, Yang K-Y, Gao K. UXT, a novel DNMT3b-binding protein, promotes breast cancer progression via negatively modulating lncRNA MEG3/p53 axis. Mol Therapy-Oncolytics. (2022) 24:497–506. doi: 10.1016/j.omto.2021.12.008
68. Shihabudeen Haider Ali MS, Cheng X, Moran M, Haemmig S, Naldrett MJ, Alvarez S, et al. LncRNA Meg3 protects endothelial function by regulating the DNA damage response. Nucleic Acids Res. (2019) 47:1505–22. doi: 10.1093/nar/gky1190
69. Gao J, Liu M, Zou Y, Mao M, Shen T, Zhang C, et al. Long non-coding RNA growth arrest-specific transcript 5 is involved in ovarian cancer cell apoptosis through the mitochondria-mediated apoptosis pathway. Oncol Rep. (2015) 34:3212–21. doi: 10.3892/or.2015.4318
70. Tripathi MK, Doxtater K, Keramatnia F, Zacheaus C, Yallapu MM, Jaggi M, et al. Role of lncRNAs in ovarian cancer: defining new biomarkers for therapeutic purposes. Drug Discovery Today. (2018) 23:1635–43. doi: 10.1016/j.drudis.2018.04.010
71. Ma N, Li S, Zhang Q, Wang H, Qin H, Wang S, et al. Long non-coding RNA GAS5 inhibits ovarian cancer cell proliferation via the control of microRNA-21 and SPRY2 expression. Exp Ther Med. (2018) 16:73–82. doi: 10.3892/etm.2018.6188
72. Qiao HP, Gao WS, Huo JX, Yang ZS. Long non-coding RNA GAS5 functions as a tumor suppressor in renal cell carcinoma. Asian Pac J Cancer Prev. (2013) 14:1077–82. doi: 10.7314/APJCP.2013.14.2.1077
73. Cao Q, Wang N, Qi J, Gu Z, Shen H. Long non−coding RNA−GAS5 acts as a tumor suppressor in bladder transitional cell carcinoma via regulation of chemokine (C−C motif) ligand 1 expression. Mol Med Rep. (2016) 13:27–34. doi: 10.3892/mmr.2015.4503
74. Misawa A, Takayama KI, Inoue S. Long non-coding RNAs and prostate cancer. Cancer Sci. (2017) 108:2107–14. doi: 10.1111/cas.2017.108.issue-11
75. Pickard MR, Mourtada-Maarabouni M, Williams GT. Long non-coding RNA GAS5 regulates apoptosis in prostate cancer cell lines. Biochim Biophys Acta. (2013) 1832:1613–23. doi: 10.1016/j.bbadis.2013.05.005
76. Yang W, Xu X, Hong L, Wang Q, Huang J, Jiang L, et al. Upregulation of lncRNA GAS5 inhibits the growth and metastasis of cervical cancer cells. J Cell Physiol. (2019) 234:23571–80. doi: 10.1002/jcp.v234.12
77. Yang W, Hong L, Xu X, Wang Q, Huang J, Jiang L. LncRNA GAS5 suppresses the tumorigenesis of cervical cancer by downregulating miR-196a and miR-205. Tumour Biol. (2017) 39:1010428317711315. doi: 10.1177/1010428317711315
78. Gokulnath P, Cristofaro T, Manipur I, Palma Di T, Soriano AA, Guarracino MR, et al. Long non-coding RNA MAGI2-AS3 is a new player with a tumor suppressive role in high grade serous ovarian carcinoma. Cancers (Basel). (2019) 11. doi: 10.3390/cancers11122008
79. Chang H, Zhang X, Li B, Meng X. MAGI2-AS3 suppresses MYC signaling to inhibit cell proliferation and migration in ovarian cancer through targeting miR-525-5p/MXD1 axis. Cancer Med. (2020) 9:6377–86. doi: 10.1002/cam4.v9.17
80. Tang C, Cai Y, Jiang H, Lv Z, Yang C, Xu H, et al. LncRNA MAGI2-AS3 inhibits bladder cancer progression by targeting the miR-31-5p/TNS1 axis. Aging (Albany NY). (2020) 12:25547–63. doi: 10.18632/aging.104162
81. Cao Y, Xu R, Xu X, Zhou Y, Cui L, He X. Downregulation of lncRNA CASC2 by microRNA-21 increases the proliferation and migration of renal cell carcinoma cells. Mol Med Rep. (2016) 14:1019–25. doi: 10.3892/mmr.2016.5337
82. Pei Z, Du X, Song Y, Fan L, Li F, Gao Y, et al. Down-regulation of lncRNA CASC2 promotes cell proliferation and metastasis of bladder cancer by activation of the Wnt/beta-catenin signaling pathway. Oncotarget. (2017) 8:18145–53. doi: 10.18632/oncotarget.15210
83. Zhong XL, Wang L, Yan X, Yang XK, Xiu H, Zhao M, et al. MiR-20a acted as a ceRNA of lncRNA PTENPL and promoted bladder cancer cell proliferation and migration by regulating PDCD4. Eur Rev Med Pharmacol Sci. (2020) 24:2955–64.
84. Wu C, Wang F, Tan L. Role and the molecular mechanism of lncRNA PTENP1 in regulating the proliferation and invasion of cervical cancer cells. Gene Ther. (2022) 29:464–75. doi: 10.1038/s41434-020-00189-8
85. Fan Y, Sheng W, Meng Y, Cao Y, Li R. LncRNA PTENP1 inhibits cervical cancer progression by suppressing miR-106b. Artif Cells Nanomed Biotechnol. (2020) 48:393–407. doi: 10.1080/21691401.2019.1709852
86. Zhu Y, Liu B, Zhang P, Zhang J, Wang L. LncRNA TUSC8 inhibits the invasion and migration of cervical cancer cells via miR-641/PTEN axis. Cell Biol Int. (2019) 43:781–8. doi: 10.1002/cbin.11152
87. Gokulnath P, Cristofaro T, Manipur I, Palma Di T, Soriano AA, Guarracino MR. Long non-coding RNA HAND2-AS1 acts as a tumor suppressor in high-grade serous ovarian carcinoma. Int J Mol Sci. (2020) 21. doi: 10.3390/ijms21114059
88. Richards EJ, Permuth-Wey J, Li Y, Chen YA, Coppola D, Reid BM, et al. A functional variant in HOXA11-AS, a novel long non-coding RNA, inhibits the oncogenic phenotype of epithelial ovarian cancer. Oncotarget. (2015) 6:34745–57. doi: 10.18632/oncotarget.v6i33
89. Chao H, Zhang M, Hou H, Zhang Z, Li N. HOTAIRM1 suppresses cell proliferation and invasion in ovarian cancer through facilitating ARHGAP24 expression by sponging miR-106a-5p. Life Sci. (2020) 243:117296. doi: 10.1016/j.lfs.2020.117296
90. Baldinu P, Cossu A, Manca A, Satta MP, Sini MC, Palomba G, et al. CASC2a gene is down-regulated in endometrial cancer. Anticancer Res. (2007) 27:235–43.
91. Baldinu P, Cossu A, Manca A, Satta MP, Sini MC, Rozzo C, et al. Identification of a novel candidate gene, CASC2, in a region of common allelic loss at chromosome 10q26 in human endometrial cancer. Hum Mutat. (2004) 23:318–26. doi: 10.1002/humu.20015
92. Zhao Y, Yang Y, Trovik J, Sun K, Zhou L, Jiang P, et al. A novel wnt regulatory axis in endometrioid endometrial cancer. Cancer Res. (2014) 74:5103–17. doi: 10.1158/0008-5472.CAN-14-0427
93. Fang Q, Sang L, Du S. Long noncoding RNA LINC00261 regulates endometrial carcinoma progression by modulating miRNA/FOXO1 expression. Cell Biochem Funct. (2018) 36:323–30. doi: 10.1002/cbf.v36.6
94. Shang C, Lang B, Ao CN, Meng L. Long non-coding RNA tumor suppressor candidate 7 advances chemotherapy sensitivity of endometrial carcinoma through targeted silencing of miR-23b. Tumour Biol. (2017) 39:1010428317707883. doi: 10.1177/1010428317707883
95. Li W, Li H, Zhang L, Hu M, Li F, Deng J, et al. Long non-coding RNA LINC00672 contributes to p53 protein-mediated gene suppression and promotes endometrial cancer chemosensitivity. J Biol Chem. (2017) 292:5801–13. doi: 10.1074/jbc.M116.758508
96. Zhu M, Chen Q, Liu X, Sun Q, Zhao X, Deng R, et al. lncRNA H19/miR-675 axis represses prostate cancer metastasis by targeting TGFBI. FEBS J. (2014) 281:3766–75. doi: 10.1111/febs.2014.281.issue-16
97. Malik R, Patel L, Prensner JR, Shi Y, Iyer MK, Subramaniyan S, et al. The lncRNA PCAT29 inhibits oncogenic phenotypes in prostate cancer. Mol Cancer Res. (2014) 12:1081–7. doi: 10.1158/1541-7786.MCR-14-0257
98. Guo J, Wang S, Jiang Z, Tang L, Liu Z, Cao J, et al. Long non-coding RNA RFPL3S functions as a biomarker of prognostic and immunotherapeutic prediction in testicular germ cell tumor. Front Immunol. (2022) 13:859730. doi: 10.3389/fimmu.2022.859730
99. Zhai W, Sun Y, Jiang M, Wang M, Gasiewicz TA, Zheng J, et al. Differential regulation of LncRNA-SARCC suppresses VHL-mutant RCC cell proliferation yet promotes VHL-normal RCC cell proliferation via modulating androgen receptor/HIF-2alpha/C-MYC axis under hypoxia. Oncogene. (2017) 36:4525. doi: 10.1038/onc.2016.514
100. Xue S, Li QW, Che JP, Guo Y, Yang FQ, Zheng JH. Decreased expression of long non-coding RNA NBAT-1 is associated with poor prognosis in patients with clear cell renal cell carcinoma. Int J Clin Exp Pathol. (2015) 8:3765–74.
101. Liu Y, Qian J, Li X, Chen W, Xu A, Zhao K, et al. Long noncoding RNA BX357664 regulates cell proliferation and epithelial-to-mesenchymal transition via inhibition of TGF-beta1/p38/HSP27 signaling in renal cell carcinoma. Oncotarget. (2016) 7:81410–22. doi: 10.18632/oncotarget.12937
102. He A, Liu Y, Chen Z, Li J, Chen M, Liu L, et al. Over-expression of long noncoding RNA BANCR inhibits Malignant phenotypes of human bladder cancer. J Exp Clin Cancer Res. (2016) 35:125. doi: 10.1186/s13046-016-0397-9
103. Xue Y, Ma G, Zhang Z, Hua Q, Chu H, Tong N, et al. A novel antisense long noncoding RNA regulates the expression of MDC1 in bladder cancer. Oncotarget. (2015) 6:484–93. doi: 10.18632/oncotarget.2861
104. Li Z, Hong S, Liu Z. LncRNA LINC00641 predicts prognosis and inhibits bladder cancer progression through miR-197-3p/KLF10/PTEN/PI3K/AKT cascade. Biochem Biophys Res Commun. (2018) 503:1825–9. doi: 10.1016/j.bbrc.2018.07.120
105. Wang YY, Wu ZY, Wang GC, Liu K, Niu XB, Gu S, et al. LINC00312 inhibits the migration and invasion of bladder cancer cells by targeting miR-197-3p. Tumour Biol. (2016) 37:14553–63. doi: 10.1007/s13277-016-5303-8
106. Gong A, Zhao X, Pan Y, Qi Y, Li S, Huang Y, et al. The lncRNA MEG3 mediates renal cell cancer progression by regulating ST3Gal1 transcription and EGFR sialylation. J Cell Sci. (2020) 133. doi: 10.1242/jcs.244020
107. Wang M, Huang T, Luo G, Huang C, Xiao XY, Wang L, et al. Long non-coding RNA MEG3 induces renal cell carcinoma cells apoptosis by activating the mitochondrial pathway. J Huazhong Univ Sci Technolog Med Sci. (2015) 35:541–5. doi: 10.1007/s11596-015-1467-5
108. Huang C, Liao X, Jin H, Xie F, Zheng F, Li J, et al. MEG3, as a competing endogenous RNA, binds with miR-27a to promote PHLPP2 protein translation and impairs bladder cancer invasion. Mol Ther Nucleic Acids. (2019) 16:51–62. doi: 10.1016/j.omtn.2019.01.014
109. Shan G, Tang T, Xia Y, Qian HJ. MEG3 interacted with miR-494 to repress bladder cancer progression through targeting PTEN. J Cell Physiol. (2020) 235:1120–8. doi: 10.1002/jcp.v235.2
110. Wu M, Huang Y, Chen T, Wang W, Yang S, Ye Z, et al. LncRNA MEG3 inhibits the progression of prostate cancer by modulating miR-9-5p/QKI-5 axis. J Cell Mol Med. (2019) 23:29–38. doi: 10.1111/jcmm.2019.23.issue-1
111. Zhou Y, Yang H, Xia W, Cui L, Xu R, Lu H, et al. LncRNA MEG3 inhibits the progression of prostate cancer by facilitating H3K27 trimethylation of EN2 through binding to EZH2. J Biochem. (2020) 167:295–301. doi: 10.1093/jb/mvz097
112. Zhang J, Yao T, Wang Y, Yu J, Liu Y, Lin Z. Long noncoding RNA MEG3 is downregulated in cervical cancer and affects cell proliferation and apoptosis by regulating miR-21. Cancer Biol Ther. (2016) 17:104–13. doi: 10.1080/15384047.2015.1108496
113. Wan S, Zhao H. Analysis of diagnostic and prognostic value of lncRNA MEG3 in cervical cancer. Oncol Lett. (2020) 20:183. doi: 10.3892/ol.2020.12044
114. Li Y, Zhang L, Zhao Y, Peng H, Zhang N, Bai W. MEG3 sponges miRNA-376a and YBX1 to regulate angiogenesis in ovarian cancer endothelial cells. Heliyon. (2023) 9:e13204. doi: 10.1016/j.heliyon.2023.e13204
115. Liu Y, Xu Y, Ding L, Yu L, Zhang B, Wei D. LncRNA MEG3 suppressed the progression of ovarian cancer via sponging miR-30e-3p and regulating LAMA4 expression. Cancer Cell Int. (2020) 20:181. doi: 10.1186/s12935-020-01259-y
116. Hsieh JJ, Purdue MP, Signoretti S, Swanton C, Albiges L, Schmidinger M, et al. Renal cell carcinoma. Nat Rev Dis Primers. (2017) 3:17009. doi: 10.1038/nrdp.2017.9
117. Petejova N, Martinek A. Renal cell carcinoma: Review of etiology, pathophysiology and risk factors. BioMed Pap Med Fac Univ Palacky Olomouc Czech Repub. (2016) 160:183–94. doi: 10.5507/bp.2015.050
118. Muglia VF, Prando A. Renal cell carcinoma: histological classification and correlation with imaging findings. Radiol Bras. (2015) 48:166–74. doi: 10.1590/0100-3984.2013.1927
119. Zhou H, Tang K, Liu H, Zeng J, Li H, Yan L, et al. Regulatory network of two tumor-suppressive noncoding RNAs interferes with the growth and metastasis of renal cell carcinoma. Mol Ther Nucleic Acids. (2019) 16:554–65. doi: 10.1016/j.omtn.2019.04.005
120. He H, Dai J, Zhuo R, Zhao J, Wang H, Sun F, et al. Study on the mechanism behind lncRNA MEG3 affecting clear cell renal cell carcinoma by regulating miR-7/RASL11B signaling. J Cell Physiol. (2018) 233:9503–15. doi: 10.1002/jcp.v233.12
121. Feng SQ, Zhang XY, Fan HT, Sun QJ, Zhang M. Up-regulation of LncRNA MEG3 inhibits cell migration and invasion and enhances cisplatin chemosensitivity in bladder cancer cells. Neoplasma. (2018) 65:925–32. doi: 10.4149/neo_2018_180125N55
122. Gomez N, Unzeta M, Tipton KF, Anderson MC, AM. Determination of monoamine oxidase concentrations in rat liver by inhibitor binding. Biochem Pharmacol. (1986) 35:4467–72. doi: 10.1016/0006-2952(86)90765-3
123. Zhu J, Han S. Lidocaine inhibits cervical cancer cell proliferation and induces cell apoptosis by modulating the lncRNA-MEG3/miR-421/BTG1 pathway. Am J Transl Res. (2019) 11:5404–16.
124. Pan X, Cao YM, Liu JH, Ding J, Xie XY, Cao PG. MEG3 induces cervical carcinoma cells’ Apoptosis through endoplasmic reticulum stress by miR-7-5p/STC1 axis. Cancer Biother Radiopharm. (2021) 36:501–10. doi: 10.1089/cbr.2019.3344
125. Dudea-Simon M, Mihu D, Pop LA, Ciortea R, Malutan AM, Diculescu D, et al. Alteration of gene and miRNA expression in cervical intraepithelial neoplasia and cervical cancer. Int J Mol Sci. (2022) 23. doi: 10.3390/ijms23116054
126. Du Y, Geng G, Zhao C, Gao T, Wei B. LncRNA MEG3 promotes cisplatin sensitivity of cervical cancer cells by regulating the miR-21/PTEN axis. BMC Cancer. (2022) 22:1145. doi: 10.1186/s12885-022-10188-0
127. Zhang J, Gao Y. Long non-coding RNA MEG3 inhibits cervical cancer cell growth by promoting degradation of P-STAT3 protein via ubiquitination. Cancer Cell Int. (2019) 19:175. doi: 10.1186/s12935-019-0893-z
128. Buttarelli M, Donato De M, Raspaglio G, Babini G, Ciucci A, Martinelli E, et al. Clinical value of lncRNA MEG3 in high-grade serous ovarian cancer. Cancers (Basel). (2020) 12. doi: 10.3390/cancers12040966
129. Wang L, Yu M, Zhao S. lncRNA MEG3 modified epithelial-mesenchymal transition of ovarian cancer cells by sponging miR-219a-5p and regulating EGFR. J Cell Biochem. (2019) 120:17709–22. doi: 10.1002/jcb.v120.10
130. Tao P, Yang B, Zhang H, Sun L, Wang Y, Zheng W. The overexpression of lncRNA MEG3 inhibits cell viability and invasion and promotes apoptosis in ovarian cancer by sponging miR-205-5p. Int J Clin Exp Pathol. (2020) 13:869–79.
131. Sun KX, Wu DD, Chen S, Zhao Y, Zong ZH. LncRNA MEG3 inhibit endometrial carcinoma tumorigenesis and progression through PI3K pathway. Apoptosis. (2017) 22:1543–52. doi: 10.1007/s10495-017-1426-7
132. Guo Q, Qian Z, Yan D, Li L, Huang L. LncRNA-MEG3 inhibits cell proliferation of endometrial carcinoma by repressing Notch signaling. BioMed Pharmacother. (2016) 82:589–94. doi: 10.1016/j.biopha.2016.02.049
133. Luo Q, Cui M, Deng Q, Liu J. Comprehensive analysis of differentially expressed profiles and reconstruction of a competing endogenous RNA network in papillary renal cell carcinoma. Mol Med Rep. (2019) 19:4685–96. doi: 10.3892/mmr.2019.10138
134. Antoni S, Ferlay J, Soerjomataram I, Znaor A, Jemal A, Bray F. Bladder cancer incidence and mortality: A global overview and recent trends. Eur Urol. (2017) 71:96–108. doi: 10.1016/j.eururo.2016.06.010
135. Lopez-Beltran A. Bladder cancer: clinical and pathological profile. Scand J Urol Nephrol Suppl. (2008) 218):95–109. doi: 10.1080/03008880802325226
136. Ying L, Huang Y, Chen H, Wang Y, Xia L, Chen Y, et al. Downregulated MEG3 activates autophagy and increases cell proliferation in bladder cancer. Mol Biosyst. (2013) 9:407–11. doi: 10.1039/c2mb25386k
137. Liu G, Zhao X, Zhou J, Cheng X, Ye Z, Ji Z, et al. Long non-coding RNA MEG3 suppresses the development of bladder urothelial carcinoma by regulating miR-96 and TPM1. Cancer Biol Ther. (2018) 19:1039–56. doi: 10.1080/15384047.2018.1480279
138. Madu CO, Lu Y. Novel diagnostic biomarkers for prostate cancer. J Cancer. (2010) 1:150–77. doi: 10.7150/jca.1.150
139. Xu B, Zhang M, Liu C, Wang C, You Z, Wang Y, et al. Association of long non-coding RNA MEG3 polymorphisms and risk of prostate cancer in Chinese Han population. Urol J. (2020) 18:176–80.
140. Rajpert-De-Meyts E, Aksglaede L, Bandak M, Toppari J, Jørgensen N. Testicular cancer: pathogenesis, diagnosis and management with focus on endocrine aspects. Endotext (2023).
141. Ghazarian AA, Trabert B, Devesa SS, McGlynn KA. Recent trends in the incidence of testicular germ cell tumors in the United States. Andrology. (2015) 3:13–8. doi: 10.1111/andr.2015.3.issue-1
142. Ghazarian AA, Kelly SP, Altekruse SF, Rosenberg PS, McGlynn KA. Future of testicular germ cell tumor incidence in the United States: Forecast through 2026. Cancer. (2017) 123:2320–8. doi: 10.1002/cncr.v123.12
143. Vasdev N, Moon A, Thorpe AC. Classification, epidemiology and therapies for testicular germ cell tumours. Int J Dev Biol. (2013) 57:133–9. doi: 10.1387/ijdb.130031nv
144. Albers P, Albrecht W, Algaba F, Bokemeyer C, Cohn-Cedermark G, Fizazi K, et al. Guidelines on testicular cancer: 2015 update. Eur Urol. (2015) 68:1054–68. doi: 10.1016/j.eururo.2015.07.044
145. Batool A, Karimi N, Wu X-N, Chen S-R, Liu Y-X. Testicular germ cell tumor: a comprehensive review. Cell Mol Life Sci. (2019) 76:1713–27. doi: 10.1007/s00018-019-03022-7
146. Chen SR, Liu YX. Regulation of spermatogonial stem cell self-renewal and spermatocyte meiosis by Sertoli cell signaling. Reproduction. (2015) 149:R159–67. doi: 10.1530/REP-14-0481
147. Chen SR, Liu YX. Disruption of genital ridge development causes aberrant primordial germ cell proliferation but does not affect their directional migration. BMC Biol. (2013) 11:22. doi: 10.1186/1741-7007-11-22
148. Skakkebaek NE. Possible carcinoma-in-situ of the testis. Lancet. (1972) 2:516–7. doi: 10.1016/S0140-6736(72)91909-5
149. Kristensen DG, Nielsen JE, Jørgensen A, Skakkebæk NE, Meyts Rajpert-De E, Almstrup K, et al. Evidence that active demethylation mechanisms maintain the genome of carcinoma in situ cells hypomethylated in the adult testis. Br J Cancer. (2014) 110:668–78. doi: 10.1038/bjc.2013.727
150. Di Vizio D, Cito L, Boccia A, Chieffi P, Insabato L, Pettinato G, et al. Loss of the tumor suppressor gene PTEN marks the transition from intratubular germ cell neoplasias (ITGCN) to invasive germ cell tumors. Oncogene. (2005) 24:1882–94. doi: 10.1038/sj.onc.1208368
151. Datta MW, Macri E, Signoretti S, Renshaw AA, Loda M. Transition from in situ to invasive testicular germ cell neoplasia is associated with the loss of p21 and gain of mdm-2 expression. Mod Pathol. (2001) 14:437–42. doi: 10.1038/modpathol.3880331
152. Taylor-Weiner A, Zack T, E, Guerriero JL, Bernard B, Reddy A, et al. Genomic evolution and chemoresistance in germ-cell tumours. Nature. (2016) 540:114–8. doi: 10.1038/nature20596
153. Fowler JR, Maani EV, Dunton CJ, Jack BW. Cervical Cancer. Treasure Island (FL: StatPearls (2023). ineligible companies. Disclosure: Elizabeth Maani declares no relevant financial relationships with ineligible companies. Disclosure: Charles Dunton declares no relevant financial relationships with ineligible companies. Disclosure: Brian Jack declares no relevant financial relationships with ineligible companies.
154. Cohen PA, Jhingran A, Oaknin A, Denny L. Cervical cancer. Lancet. (2019) 393:169–82. doi: 10.1016/S0140-6736(18)32470-X
155. Correction to lancet glob health 2020; 8: e191-203. Lancet Glob Health. (2022) 10:e41. doi: 10.1016/S2214-109X(21)00554-4
156. Burd EM. Human papillomavirus and cervical cancer. Clin Microbiol Rev. (2003) 16:1–17. doi: 10.1128/CMR.16.1.1-17.2003
157. McLaughlin-Drubin ME, Park D, Munger K. Tumor suppressor p16INK4A is necessary for survival of cervical carcinoma cell lines. Proc Natl Acad Sci U.S.A. (2013) 110:16175–80.
158. Harima Y, Sawada S, Nagata K, Sougawa M, Ostapenko V, Ohnishi T. Mutation of the PTEN gene in advanced cervical cancer correlated with tumor progression and poor outcome after radiotherapy. Int J Oncol. (2001) 18:493–7. doi: 10.3892/ijo.18.3.493
159. Sidorkiewicz I, Piskor B, Dabrowska E, Guzinska-Ustymowicz K, Pryczynicz A, Zbucka-Kretowska M, et al. Plasma levels and tissue expression of selected cytokines, metalloproteinases and tissue inhibitors in patients with cervical cancer. Anticancer Res. (2019) 39:6403–12. doi: 10.21873/anticanres.13854
160. Aalijahan H, Ghorbian S. Long non-coding RNAs and cervical cancer. Exp Mol Pathol. (2019) 106:7–16. doi: 10.1016/j.yexmp.2018.11.010
161. Tornesello ML, Faraonio R, Buonaguro L, Annunziata C, Starita N, Cerasuolo A, et al. The role of microRNAs, long non-coding RNAs, and circular RNAs in cervical cancer. Front Oncol. (2020) 10:150. doi: 10.3389/fonc.2020.00150
162. Webb PM, Jordan SJ. Epidemiology of epithelial ovarian cancer. Best Pract Res Clin Obstet Gynaecol. (2017) 41:3–14. doi: 10.1016/j.bpobgyn.2016.08.006
163. Momenimovahed Z, Tiznobaik A, Taheri S, Salehiniya H. Ovarian cancer in the world: epidemiology and risk factors. Int J Womens Health. (2019) 11:287–99. doi: 10.2147/IJWH.S197604
164. Almalki AA, Gharib AF, Al-Shehri SS, Alghamdi A, Alrehaili AA, Banjer HJ, et al. Exploring the potential of LNCRNA-MEG-3 as a diagnostic and prognostic marker in epithelial ovarian cancer. Eur J Gynaecological Oncol. (2024) 45.
165. Crosbie EJ, Kitson SJ, McAlpine JN, Mukhopadhyay A, Powell ME, Singh N. Endometrial cancer. Lancet. (2022) 399:1412–28. doi: 10.1016/S0140-6736(22)00323-3
166. Sorosky JI. Endometrial cancer. Obstet Gynecol. (2012) 120:383–97. doi: 10.1097/AOG.0b013e3182605bf1
167. Singh N, Hirschowitz L, Zaino R, Alvarado-Cabrero I, Duggan MA, Ali-Fehmi R, et al. Pathologic prognostic factors in endometrial carcinoma (Other than tumor type and grade). Int J Gynecol Pathol. (2019) 38 Suppl 1:S93–S113. doi: 10.1097/PGP.0000000000000524
168. Dong P, Xiong Y, Yue J, Xu D, Ihira K, Konno Y, et al. Long noncoding RNA NEAT1 drives aggressive endometrial cancer progression via miR-361-regulated networks involving STAT3 and tumor microenvironment-related genes. J Exp Clin Cancer Res. (2019) 38:295. doi: 10.1186/s13046-019-1306-9
169. Xu D, Dong P, Xiong Y, Chen R, Konno Y, Ihira K, et al. PD-L1 Is a Tumor Suppressor in Aggressive Endometrial Cancer Cells and Its Expression Is Regulated by miR-216a and lncRNA MEG3. Front Cell Dev Biol. (2020) 8:598205. doi: 10.3389/fcell.2020.598205
170. Wang Y-M, Trinh MP, Zheng Y, Guo K, Jimenez LA, Zhong W, et al. Analysis of circulating non-coding RNAs in a non-invasive and cost-effective manner. TrAC Trends Analytical Chem. (2019) 117:242–62. doi: 10.1016/j.trac.2019.07.001
171. Eldakhakhny B, Sutaih AM, Siddiqui MA, Aqeeli YM, Awan AZ, Alsayegh MY, et al. Exploring the role of noncoding RNAs in cancer diagnosis, prognosis, and precision medicine. Non-coding RNA Res. (2024). doi: 10.1016/j.ncrna.2024.06.015
172. Chen M-J, Wang X-G, Sun Z-X, Liu X-C. Diagnostic value of LncRNA-MEG3 as a serum biomarker in patients with hepatitis B complicated with liver fibrosis. Eur Rev Med Pharmacol Sci. (2019) 23.
173. Mishra S, Srivastava P, Pandey A, Agarwal A, Shukla S, Husain N, et al. Panel of serum long non-coding RNAs as potential non-invasive biomarkers for gallbladder carcinoma. Non-coding RNA Res. (2024) 9:583–93. doi: 10.1016/j.ncrna.2024.02.005
174. Russano M, Napolitano A, Ribelli G, Iuliani M, Simonetti S, Citarella F, et al. Liquid biopsy and tumor heterogeneity in metastatic solid tumors: the potentiality of blood samples. J Exp Clin Cancer Res. (2020) 39:1–13. doi: 10.1186/s13046-020-01601-2
175. Lian Z, Tian P, Ma S, Chang T, Liu R, Feng Q, et al. Long noncoding RNA MEG3 regulates cell proliferation and apoptosis by disrupting microRNA-9-5p-mediated inhibition of NDRG1 in prostate cancer. Aging (Albany NY). (2024) 16:1938. doi: 10.18632/aging.205472
176. Han S, Chen X, Huang L. The tumor therapeutic potential of long non-coding RNA delivery and targeting. Acta Pharm Sin B. (2023) 13:1371–82. doi: 10.1016/j.apsb.2022.12.005
Keywords: LncRNA - long noncoding RNA, MEG3 lncRNA, genitourinary cancers, renal cell carcinoma, bladder cancer, prostate cancer, testicular cancer, cervical cancer
Citation: Assal RA, Rashwan HH, Zakaria ZI, Sweillam JH, Fouda YM, Abdelhamid AM and Youness RA (2025) Deciphering the mysteries of MEG3 LncRNA and its implications in genitourinary cancers. Front. Oncol. 15:1519103. doi: 10.3389/fonc.2025.1519103
Received: 29 October 2024; Accepted: 28 February 2025;
Published: 02 April 2025.
Edited by:
Missaoui Nabiha, University of Sousse, TunisiaReviewed by:
Eswari Dodagatta-Marri, University of California, San Francisco, United StatesCopyright © 2025 Assal, Rashwan, Zakaria, Sweillam, Fouda, Abdelhamid and Youness. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Rana A. Youness, cmFuYS55b3VuZXNzMjFAZ21haWwuY29t
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.