
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
MINI REVIEW article
Front. Immunol. , 21 March 2025
Sec. Cancer Immunity and Immunotherapy
Volume 16 - 2025 | https://doi.org/10.3389/fimmu.2025.1573686
This article is part of the Research Topic The Role of Metabolic Reprogramming in Tumor Therapy View all 7 articles
Metabolic reprogramming is a hallmark of ovarian cancer, enabling tumor progression, immune evasion and drug resistance. The tumor microenvironment (TME) further shapes metabolic adaptations, enabling cancer cells to withstand hypoxia and nutrient deprivation. While organoid models provide a physiologically relevant platform for studying these processes, they still lack immune and vascular components, limiting their ability to fully recapitulate tumor metabolism and drug responses. In this study, we investigated the key metabolic mechanisms involved in ovarian cancer progression, focusing on glycolysis, lipid metabolism and amino acid metabolism. We integrated metabolomic analyses and drug sensitivity assays to explore metabolic-TME interactions using patient-derived, adult stem cell-derived and iPSC-derived organ tissues. Among these, we found that glycolysis, lipid metabolism and amino acid metabolism play a central role in tumor progression and chemotherapy resistance. We identified methylglyoxal (MGO)-mediated BRCA2 dysfunction as a driver of immune escape, a role for sphingolipid signaling in tumor proliferation and a role for kynurenine metabolism in CD8+ T cell suppression. In addition, PI3K/AKT/mTOR and Wnt/β-catenin pathways promote chemoresistance through metabolic adaptation. By elucidating the link between metabolic reprogramming and immune evasion, this study identifies key metabolic vulnerabilities and potential drug targets in ovarian cancer. Our findings support the development of metabolically targeted therapies and increase the utility of organoid-based precision medicine models.
Ovarian cancer is a deadly gynecologic disease, causing over 152,000 deaths annually (1). While most patients initially respond to platinum-based therapies, resistance mechanisms, including paclitaxel resistance, hinder treatment progress (2, 3). This highlights the urgent need to study ovarian cancer metabolism and drug resistance to achieve precision therapy (3). However, ovarian cancer’s high heterogeneity complicates research (4). Organoid technology preserves the tumor microenvironment (TME), mimics tumor heterogeneity (5–7). It allows the study of developmental and pathogenic pathways that traditional animal models struggle to replicate and supports real-time imaging at lower costs and higher throughput than animal models, with a broader range of cell types than 2D cultures (8, 9). Additionally, it simulates drug penetration and metabolism, aiding in personalized therapy (10).
In ovarian cancer, dysregulated signaling pathways and metabolic reprogramming empower cancer cells to proliferate and persist within the adverse tumor microenvironment (TME) (11). Various signaling pathways control cell proliferation, survival, and stemness, while metabolic reprogramming closely links to cisplatin resistance (12–14). ovarian cancer organoids provide a realistic model to study these pathways and processes, enhancing the understanding of drug resistance mechanisms (15). This paper examines different drug-related metabolic and signaling pathways in ovarian cancer organoids, summarizes drug resistance, and discusses limitations in personalized treatment to inform clinical applications.
Organoids are tissue analogs with a certain spatial structure formed by in vitro three-dimensional culture using adult stem cells, pluripotent stem cells, or patient-derived cells (16, 17).They have stable biological characteristics: tumor-derived organoids accurately replicate their structural, phenotypic, and genetic features, have similar tumor heterogeneity, in TME effectively mimic immune cells and stromal components, can be cultured in vitro long-term culture (17), and the ability to preserve metabolic adaptations in tumors, dynamically regulate glycolysis, lipid metabolism and amino acid metabolism, to study metabolic reprogramming in a way that more closely approximates the in vivo microenvironment, and to generalize the formation of two events occurring in organoid bodies, namely, the cellular adhesion of categorical aggregates and the spatially specific cellular hematopoietic stereotypes (18). (Table 1).
Induced pluripotent stem cells (iPSCs) derived from patients with early-onset ovarian cancer carrying BRCA1 gene mutations possess the capability to differentiate into fallopian tube epithelial (FTE)-like structures (19). Pathogenic BRCA1 mutations promote the transformation of normal FTECs into high-grade serous ovarian cancer cells (20). Additionally, these induced pluripotent stem cells (iPSCs) exhibit embryonic stem cell-like morphology and display pluripotent characteristics, including RNA expression of key pluripotency factors and the production of pluripotency marker proteins (21).
Ovarian cancer-like organs derived from ASCs retain the intratumoral heterogeneity and subclonal evolutionary capacity of the primary tumor (21). Different tumor subtypes of these organoids have differential responses to standard platinum-based chemotherapy, including subtypes that become chemoresistant in recurrent ovarian cancer, e.g. PDO can be used in drug screening assays to determine and capture the response of different tumor subtypes to platinum-based chemotherapy (22, 23).
Ovarian cancer patient-derived organoids (OC PDOs), established from ovarian cancer patient specimens, preserve the genomic features of the original tumor, including mutations, copy number variations, and other genomic alterations (24).OC PDOs also replicate the heterogeneity of the original tumor and exhibit drug responses that align with the clinical outcomes observed in patients (25).
The fallopian tube is notably recognized as the primary tissue of origin for ovarian cancer, particularly high-grade serous ovarian carcinoma (HGSOC), with fallopian tube epithelial cells (FTEC) considered the cells of origin for HGSOC. These cells are enriched with stem cells dependent on Wnt signaling and have the ability to form organoids stably and efficiently (26). Patient-derived HGSOC organoids are morphologically and molecularly consistent with their parent tumors, exhibiting features of nuclear pleomorphism, prominent nucleoli and dense chromatin formed by TP53 mutations (27).
HGSOC frequently harbors BRCA1 or BRCA2 mutations and homologous recombination (HR) defects, leading to synthetic lethality in HR-deficient cells and sensitivity to PARPi (e.g., olaparib) (28). However, most HGSOC-like organoids possess functional HR repair, making them resistant to PARPi (27).
Both HGSOC and their organoids exhibit genomic instability, TP53 mutations, and distinct mRNA and miRNA expression profiles (29, 30). While many differentiated HGSOC-like organoids lack the patient’s immune microenvironment and vascular system, some newer versions show histological and molecular diversity, including CD34+ endothelial cells that retain critical immune and vascular structures (31). This enhancement better replicates tumor complexity, providing a more effective model for studying HGSOC pathomechanisms. (Figure 1A)
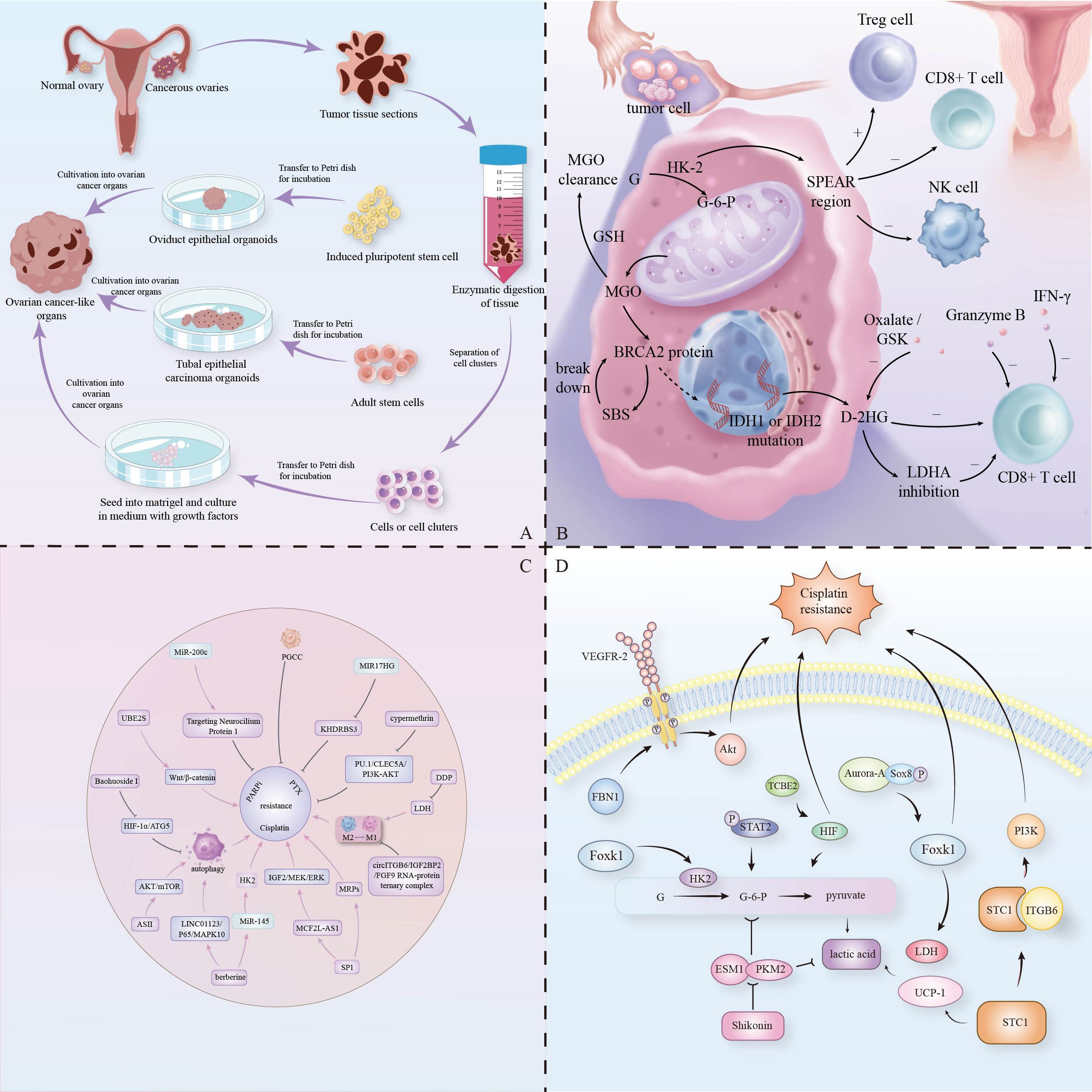
Figure 1. (A) Organoid models of ovarian cancer from three different sources. (B) Schematic diagram of the glycolytic metabolic pathway in ovarian cancer cells, with the dotted line indicating the normal pathway in the absence of SBS action. (C) Chemotherapeutic agents and their targets. Mechanisms of action of chemotherapeutic agents and networks of resistance-related gene and protein interactions. (D) Mechanisms of metabolic regulation by signaling pathways.
Metabolic reprogramming, a hallmark of ovarian cancer, allows cells to adjust energy metabolism for rapid proliferation and survival in diverse conditions (32). Ovarian cancer cells typically achieve metabolic pattern remodeling by enhancing glycolytic activity and inducing the Warburg effect (33). Additionally, these cells can regulate fatty acid and amino acid metabolism and inhibit oxidative phosphorylation to meet their growth requirements.
Methylglyoxal (MGO) as a highly reactive dicarbonyl metabolite. It is an important endogenous dicarbonyl metabolic component in the human body and is present in many different tissues and organs (34).
As obtained in studies of non-malignant breast cells or patient-derived organ tissues, MGO induces cancer-associated mutant single base substitutions (SBS), triggers BRCA2 proteolysis, and transiently disables the tumor-suppressor function of BRCA2 in DNA repair and replication, leading to functional haploinsufficiency (35, 36). According to Knudson’s “two strikes” theory, tumor development usually requires two mutations: the first mutation may be inherited, and the second occurs in somatic cells (37). And defects in functional haploinsufficiency mean that even without complete loss of complete mutations in both genes, tumors may still result. This approach bypasses Knudson’s “double whammy” requirement for tumor initiation in the presence of partially preserved BRCA2 function (38).
Smith-Beckerman DM et al. reported that glyoxalase 1 (Glo1) is upregulated in FH-OSE cells of women with ovarian cancer carrying a BRCA1 mutation. Glo1 plays a key role in clearing methylglyoxal (MGO), catalyzing its conjugation with GSH to form less toxic products (39). This suggests that MGO accumulation in ovarian cancer may serve a protective role when BRCA2 is impaired, potentially impacting BRCA2 function and influencing ovarian cancer progression through Glo1 regulation. However, the role of Glo1 in MGO metabolism suggests that MGO may affect BRCA2 function, but whether it directly induces BRCA2 degradation requires further experimental verification.
D-2HG is a metabolite produced primarily by isocitrate dehydrogenase (IDH) mutant tumor cells (40). In TME, D-2HG significantly weakened the defense function of nearby CD8+ T cells and acutely impaired their antitumor activity, including decreasing granzyme B expression and inhibiting IFN-γ release, which further weakened their immune response (41, 42). In addition, D-2HG inhibits the proliferation and cytokine (e.g., IL-2, IFN-γ, TNF-α) production of immune cells by inhibiting LDH activity, disrupting glycolytic fluxes and NAD+/NADH balance, thereby reducing their ability to kill cancer cells (43, 44). D-2HG also modifies the glucose metabolism of CD8+ T cells, further inhibiting their proliferation and cytotoxicity (42, 45, 46).
D-2HG accumulation is usually caused by LDH1 or LDH2 mutations. In EOC cells, low LDH1 expression leads to D-2HG accumulation, which inhibits CD8+ T cell proliferation and cytotoxicity and promotes immune escape (47). It has been shown that the use of LDH inhibitors (e.g., oxalate and GSK) significantly reduces D-2HG production in samples from glioma patients with LDH1 mutations, thereby inhibiting immune escape and delaying tumor progression. It was also demonstrated that LDH2 enhances metabolic remodeling and promotes OC growth by enhancing the activity of the TCA cycle (48). These results suggest that in ovarian cancer-like organs, LDH inhibitors may restore T cell activity and enhance tumor immune response by reducing D-2HG concentration (42).
Hexokinase-2 (HK2), which catalyzes the first step of glycolysis, is highly expressed in ovarian cancer cells (49). It was shown that HK2 supports rapid proliferation and survival of tumor cells by promoting glycolysis and enhancing cellular energy production (50). In the presence of sufficient oxygen, tumor cells increase glucose uptake and tend to ferment glucose into lactic acid, a phenomenon known as the “Warburg effect” (51). HK2 induces it, resulting in the formation of a more acidic environment around the cancer cells (50). This acidic microenvironment suppresses immune cell function through a combination of direct and indirect mechanisms. Lactate accumulates in the tumor microenvironment and binds to GAPDH, thereby directly inhibiting immune cell function and proliferation (52). In addition, the acidic environment not only inhibits the production of cytokines associated with anti-tumor immune responses, but also enhances the migration of tumor cells by decreasing the migration and infiltration ability of the immune cells while enhancing their migration (53–56). Together, these factors make it more difficult for immune cells to effectively reach and function at the tumor site. In addition, the acidic environment can promote the proliferation and activation of regulatory T cells (Tregs), thus indirectly inhibiting the function of CD8+ T cells and NK cells (57, 58). The acidic environment reduces the viability and killing ability of anti-tumor immune cells (e.g., CD8+ T cells and NK cells) and promotes the proliferation of regulatory T cells (Tregs) (59, 60). Therefore, it can be hypothesized that the HK2-induced Warburg effect can regulate the CD8+ and NK cells through acidifying the microenvironment in ovarian carcinoma carcinoids (51).
In ovarian cancer cells, there exists a negative regulatory relationship between AMPK phosphorylation and HIF-1α protein (61, 62). The activation of AMPK inhibits glycolysis and further suppresses the function of HIF-1α by promoting its degradation (63). However, the overexpression of HIF-1α can conversely inhibit the activation of AMPK and prevent the metabolic shift from glycolysis to OXPHOS (64). Therefore, the proliferation of ovarian cancer cells can be inhibited by silencing the activation of AMPK and suppressing the HIF-1α protein through transient receptor potential melastatin 7 (TRPM7), thereby adjusting the metabolic network and shifting the preferred glycolysis to oxidative phosphorylation (OXPHOS) (65, 66).
Sphingolipids are mainly composed of sphingomyelin and fatty acids, which are widely present in cell membranes (67). synthesizing sphingolipids from scratch is an important pathway for tumor immune evasion. Sphingolipids, as specific sphingolipids, can promote cell growth and survival in cancer cells (68).
The hypoxic tumor microenvironment induces the expression of the secreted protein ESM1 through the transcription of HIF-1α (57, 69), which in turn promotes the SUMOylation of PKM2 and the subsequent formation of PKM2 dimers (70). This process promotes the Warburg effect and the nuclear translocation of PKM2, which ultimately leads to STAT3 phosphorylation (71, 72). Enhancement of fatty acid synthesis in ovarian cancer (70). Because it was found that shikonin inhibits the molecular interaction between ESM1 and PKM2, thus preventing the formation of PKM2 dimers and inhibiting glycolysis and fatty acid synthesis in ovarian cancer (70).
Amino acid metabolism regulates immune escape by modulating immune cell function (73, 74). For example, the Kynurenine pathway in tryptophan metabolism directly promotes immune escape by activating Treg cells and inhibiting CD8+ T cell activity (45, 46, 75). Arginine promotes T cell and NK cell function, resulting in enhanced proliferation, differentiation and effector function, thereby increasing the anti-tumor activity of immune cells (76, 77).
Therefore, it can be inferred that in the study of ovarian cancer-like organs, the survival of tumor cells can be effectively interfered with by regulating the amino acid metabolic pathway. For example, interfering with tyrosine catabolism in epithelial ovarian cancer (EOC) as the main disease model can inhibit DNA damage caused by genotoxic chemotherapeutic drugs (78). Depletion of fumarate acetoacetate hydrolase (FAH), a key enzyme in tyrosine metabolism, reduces the sensitivity of EOC to chemotherapy (78). In addition, tyrosine metabolism also plays an important role in tumor immune escape. Tyrosine can be converted to tryptophan, which in turn affects the activity of the Kynurenine pathway and promotes tumor cell immune escape (75, 79). Therefore, intervening in tyrosine metabolism may not only enhance the anti-tumor response of immune cells, but also improve the efficacy of chemotherapeutic agents and enhance the therapeutic effect. (Figure 1B)
The tumor microenvironment (TME) consists of a wide range of non-cancer cells, non-cellular components and molecules released by them (80, 81), and is characterized by hypoxia, low pH and high redox state (82). However, cancer cells can survive in this hypoxic, nutrient-depleted environment (83). In particular, ovarian cancer cells often adapt to this microenvironment by enhancing glycolysis, and this metabolic reprogramming also enables tumor cells to rapidly produce energy and metabolic intermediates that support tumor cell proliferation and survival (11).
Multiple signaling pathways regulate glycolysis, impacting tumor proliferation, migration, survival, and resistance to chemotherapy. For instance, serine/threonine kinase Aurora-A binds to the transcription factor SOX8, phosphorylating its Ser327 site, which increases FOXK1 expression and regulates genes linked to cellular senescence and glycolysis, such as LDHA and HK2. This promotes glucose metabolism and induces cisplatin resistance. Similarly (84), the TCEB2/HIF1A axis enhances cisplatin resistance by promoting glycolysis and angiogenesis (85). Furthermore, increased FBN1 expression in cisplatin-resistant ovarian cancer models indicates that the Fibrillin-1/VEGFR2/STAT2 axis plays a regulatory role in glycolysis and angiogenesis, thereby contributing to cisplatin resistance (86).
Fatty metabolism also undergoes metabolic reprogramming (11). For example, STC1 in the FOXC2/ITGB6 signaling axis can promote lipid metabolism by up-regulating lipid-related genes such as mitochondrial brown lipolytic coupling protein 1 (UCP1), TOM20 and perilipin1 (87). ESM1 promotes fatty acid synthesis and angiogenesis in ovarian cancer through the PKM2-dependent Warburg effect within a hypoxic TME, while shikonin effectively disrupts the interaction between ESM1 and PKM2, subsequently inhibiting glycolysis, fatty acid synthesis, and angiogenesis in ovarian cancer (70). (Figure 1D)
Berberine, an alkaloid with antitumor properties, inhibits the Warburg effect in ovarian cancer cells by enhancing TET3-mediated demethylation via the TET3/miR-145/HK2 signaling pathway (88). Moreover, berberine modulates autophagy and glycolysis in ovarian cancer through the LINC01123/P65/MAPK10 signaling axis (89).
Cryptotanshinone (CT) can inhibit cancer growth by inhibiting glycolysis and oxidative phosphorylation (90). Isovalerolactone, has anticancer activity. It has been shown to inhibit glycolysis and suppress cisplatin resistance in ovarian cancer cells (91). LDH can repolarize macrophages from a tumor-promoting M2 phenotype to a tumor-suppressive M1 phenotype by facilitating this phenotypic shift (92). Conversely, the platinum complex (DDP) downregulates the expression of LDHA and LDHB, thereby inhibiting glycolysis and glucose oxidation (93).
FOXO1 acts as a cell-specific coregulated transcription factor (94). For instance, within the miRNA-374a/FOXO1 signaling axis, propofol induces cell cycle arrest and decreases ovarian cancer cell viability by downregulating miR-374a, thereby alleviating its inhibition of FOXO1 (95). Additionally, FOXO1 influences ovarian cancer cell proliferation, migration, and invasion through apoptosis regulation in the EZH2/FOXO1 pathway (96). As an antitumor flavonoid, baicalein promotes apoptosis through the inhibition of the EZH2/FOXO1 signaling pathway (97).
The PI3K/AKT/mTOR pathway is a highly active cell signaling cascade in advanced ovarian cancer (12, 98). Xanthopterin suppresses ovarian cancer proliferation and cisplatin resistance by targeting the PU.1/CLEC5A/PI3K-AKT pathway (99). Ropivacaine, on the other hand, inhibits stemness and accelerates iron death of ovarian cancer cells by inactivating the PI3K/AKT signaling pathway (100).
The transcription factor SP1 enhances cisplatin resistance through multiple mechanisms. Beyond SP1-induced MCF2L-AS1, which activates the IGF2/MEK/ERK pathway (101), SP1 also transcribes multidrug resistance-associated proteins (MRPs) and key enzymes involved in arachidonic acid (AA) metabolism (e.g., 12-lipoxygenase), thereby promoting chemoresistance and metastasis in ovarian cancer (102). Moreover, circITGB6 promotes cisplatin resistance in ovarian cancer by forming the circITGB6/IGF2BP2/FGF9 RNA-protein ternary complex, which facilitates the polarization of tumor-associated macrophages (TAMs) towards the M2 phenotype (103, 104).
We also identified ex vivo and in vivo factors such as miR-219-5p, Baohuoside I, and Astragaloside II (ASII) that could modulate cisplatin resistance by regulating autophagy in ovarian cancer cells. ASII can induce autophagy by inhibiting the AKT/mTOR signaling pathway (105), while Boswellia serrata I suppresses autophagy through downregulation of the HIF-1α/ATG5 signaling axis (106). Moreover, miR-219-5p inhibits Wnt/β-catenin signaling and autophagy in ovarian cancer cells by targeting HMGA2 (105) and reduces cisplatin resistance by suppressing Wnt/β-catenin signaling and autophagy through HMGA2 targeting in these cells (107).
PARPi resistance involves various heterogeneous mechanisms, making it crucial to understand its molecular basis in patients to address chemoresistance and refine personalized treatments (108). Utilizing organoid technology, researchers are creating patient-derived organoids to predict responses to PARP inhibitors (PARPi) and explore strategies to overcome drug resistance in ovarian cancer (109). Notably, the Wnt/β-catenin pathway contributes to resistance in both cisplatin and PARPi (107). For instance, the ubiquitin-conjugating enzyme E2S (UBE2S) promotes PARPi resistance in ovarian cancer by activating Wnt/β-catenin signaling (110). Additionally, targeting polyploid giant cancer cells (PGCC) in ovarian cancer organoids could enhance therapeutic efficacy and reduce PARPi resistance (111). (Figure 1C)
This review systematically explores the application of ovarian cancer organoid models in the study of metabolic reprogramming, immune escape and drug resistance mechanisms, introducing and integrating organoids into the metabolism-immunity-drug resistance interaction framework. Metabolic reprogramming serves as a crucial indicator of tumor progression and therapeutic resistance (112). Organoid-based drug screening is more accurate and physiologically relevant than traditional techniques and is more likely to identify new strategies for treating metabolic disorders in ovarian cancer.
There are still limitations to this approach. Current cancer organoid culture techniques lack control and reproducibility, requiring the development of stable, reproducible platforms (9). Additionally, organoid models often lack vascular networks and immune systems, failing to fully replicate in vivo organ structure and function (113). Organoid techniques are also more time-intensive than 2D cultures and often require specialized media and growth factors (114). Despite advances, organoids lack vascular and immune components, limiting their ability to mimic tumor microenvironments and drug responses, necessitating further improvements. In the future, we can integrate immune and matrix co-culture, microfluidic dynamic culture system, single cell metabolomics and clinical metabolism data to optimize model stability and physiological relevance, so that it can better simulate the in vivo metabolic environment and help precision medicine research.
LZ: Conceptualization, Visualization, Writing – original draft, Writing – review & editing. JZ: Visualization, Writing – original draft. CS: Conceptualization, Writing – original draft. JW: Investigation, Writing – original draft. LJ: Validation, Writing – original draft. HC: Funding acquisition, Resources, Writing – original draft, Writing – review & editing. QW: Funding acquisition, Resources, Writing – original draft, Writing – review & editing.
The author(s) declare that financial support was received for the research and/or publication of this article. This study was supported by grants from the Sichuan Province Science and Technology Department of foreign (border) high-end talent introduction project (No: 2023ZHYZ0009) and Luzhou Science and Technology Department Applied Basic Research program (No:2022-WYC-196).
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
1. Zhang R, Siu MKY, Ngan HYS, Chan KKL. Molecular biomarkers for the early detection of ovarian cancer. Int J Mol Sci. (2022) 23:12041. doi: 10.3390/ijms231912041
2. Wang L, Wang X, Zhu X, Zhong L, Jiang Q, Wang Y, et al. Drug resistance in ovarian cancer: from mechanism to clinical trial. Mol Cancer. (2024) 23:66. doi: 10.1186/s12943-024-01967-3
3. Chan WS, Mo X, Ip PPC, Tse KY. Patient-derived organoid culture in epithelial ovarian cancers-Techniques, applications, and future perspectives. Cancer Med. (2023) 12:19714–31. doi: 10.1002/cam4.v12.19
4. Li S, Lei N, Chen M, Guo R, Han L, Qiu L, et al. Exploration of organoids in ovarian cancer: From basic research to clinical translation. Transl Oncol. (2024) 50:102130. doi: 10.1016/j.tranon.2024.102130
5. Haque MR, Rempert TH, Al-Hilal TA, Wang C, Bhushan A, Bishehsari F. Organ-chip models: opportunities for precision medicine in pancreatic cancer. Cancers (Basel). (2021) 13:4487. doi: 10.3390/cancers13174487
6. Liu J, Huang X, Huang L, Huang J, Liang D, Liao L, et al. Organoid: next-generation modeling of cancer research and drug development. Front Oncol. (2021) 11:826613. doi: 10.3389/fonc.2021.826613
7. Zhou S, Lu J, Liu S, Shao J, Liu Z, Li J, et al. Role of the tumor microenvironment in Malignant melanoma organoids during the development and metastasis of tumors. Front Cell Dev Biol. (2023) 11:1166916. doi: 10.3389/fcell.2023.1166916
8. Romero-Guevara R, Ioannides A, Xinaris C. Kidney organoids as disease models: strengths, weaknesses and perspectives. Front Physiol. (2020) 11:563981. doi: 10.3389/fphys.2020.563981
9. LeSavage BL, Suhar RA, Broguiere N, Lutolf MP, Heilshorn SC. Next-generation cancer organoids. Nat Mater. (2022) 21:143–59. doi: 10.1038/s41563-021-01057-5
10. Li J, Zhou M, Xie J, Chen J, Yang M, Ye C, et al. Organoid modeling meets cancers of female reproductive tract. Cell Death Discovery. (2024) 10:410. doi: 10.1038/s41420-024-02186-x
11. Yan J, Xu F, Zhou D, Zhang S, Zhang B, Meng Q, et al. Metabolic reprogramming of three major nutrients in platinum-resistant ovarian cancer. Front Oncol. (2023) 13:1231460. doi: 10.3389/fonc.2023.1231460
12. Ediriweera MK, Tennekoon KH, Samarakoon SR. Role of the PI3K/AKT/mTOR signaling pathway in ovarian cancer: Biological and therapeutic significance. Semin Cancer Biol. (2019) 59:147–60. doi: 10.1016/j.semcancer.2019.05.012
13. Wang L, Zhao X, Fu J, Xu W, Yuan J. The role of tumour metabolism in cisplatin resistance. Front Mol Biosci. (2021) 8:691795. doi: 10.3389/fmolb.2021.691795
14. Nguyen VHL, Hough R, Bernaudo S, Peng C. Wnt/β-catenin signaling in ovarian cancer: Insights into its hyperactivation and function in tumorigenesis. J Ovarian Res. (2019) 12:122. doi: 10.1186/s13048-019-0596-z
15. Zhang M, Yin R, Li K. Advances and challenges in the origin and evolution of ovarian cancer organoids. Front Oncol. (2024) 14:1429141. doi: 10.3389/fonc.2024.1429141
16. Chen D, Su X, Chen H, Chen S, Zhao Y, Wei W. Human organoids as a promising platform for fighting COVID-19. Int J Biol Sci. (2022) 18:901–10. doi: 10.7150/ijbs.64993
17. Ren X, Chen W, Yang Q, Li X, Xu L. Patient-derived cancer organoids for drug screening: Basic technology and clinical application. J Gastroenterol Hepatol. (2022) 37:1446–54. doi: 10.1111/jgh.15930
18. Jin RU, Mills JC. Tumor organoids to study gastroesophageal cancer: a primer. J Mol Cell Biol. (2020) 12:593–606. doi: 10.1093/jmcb/mjaa035
19. Van Lent J, Baggiolini A. Harmony in chaos: understanding cancer through the lenses of developmental biology. Mol Oncol. (2024) 18:793–6. doi: 10.1002/1878-0261.13594
20. Dai Y, Xu J, Gong X, Wei J, Gao Y, Chai R, et al. Human fallopian tube-derived organoids with TP53 and RAD51D mutations recapitulate an early stage high-grade serous ovarian cancer phenotype in vitro. Int J Mol Sci. (2024) 25:886. doi: 10.3390/ijms25020886
21. Yucer N, Ahdoot R, Workman MJ, Laperle AH, Recouvreux MS, Kurowski K, et al. Human iPSC-derived fallopian tube organoids with BRCA1 mutation recapitulate early-stage carcinogenesis. Cell Rep. (2021) 37:110146. doi: 10.1016/j.celrep.2021.110146
22. Kopper O, de Witte CJ, Lõhmussaar K, Valle-Inclan JE, Hami N, Kester L, et al. An organoid platform for ovarian cancer captures intra- and interpatient heterogeneity. Nat Med. (2019) 25:838–49. doi: 10.1038/s41591-019-0422-6
23. Tao M, Wu X. The role of patient-derived ovarian cancer organoids in the study of PARP inhibitors sensitivity and resistance: from genomic analysis to functional testing. J Exp Clin Cancer Res. (2021) 40:338. doi: 10.1186/s13046-021-02139-7
24. Jiang X, Oyang L, Peng Q, Liu Q, Xu X, Wu N, et al. Organoids: opportunities and challenges of cancer therapy. Front Cell Dev Biol. (2023) 11:1232528. doi: 10.3389/fcell.2023.1232528
25. Spagnol G, Sensi F, De Tommasi O, Marchetti M, Bonaldo G, Xhindoli L, et al. Patient derived organoids (PDOs), extracellular matrix (ECM), tumor microenvironment (TME) and drug screening: state of the art and clinical implications of ovarian cancer organoids in the era of precision medicine. Cancers (Basel). (2023) 15:2059. doi: 10.3390/cancers15072059
26. Chang YH, Chu TY, Ding DC. Human fallopian tube epithelial cells exhibit stemness features, self-renewal capacity, and Wnt-related organoid formation. J BioMed Sci. (2020) 27:32. doi: 10.1186/s12929-019-0602-1
27. Hill SJ, Decker B, Roberts EA, Horowitz NS, Muto MG, Worley MJ Jr., et al. Prediction of DNA repair inhibitor response in short-term patient-derived ovarian cancer organoids. Cancer Discovery. (2018) 8:1404–21. doi: 10.1158/2159-8290.CD-18-0474
28. Lord CJ, Ashworth A. PARP inhibitors: Synthetic lethality in the clinic. Science. (2017) 355:1152–8. doi: 10.1126/science.aam7344
29. Bell D, Berchuck A, Birrer M, Imielinski M, Chien J, Cramer DW, et al. Integrated genomic analyses of ovarian carcinoma. Nature. (2011) 474:609–15. doi: 10.1038/nature10166
30. Vang R, Levine DA, Soslow RA, Zaloudek C, Shih Ie M, Kurman RJ. Molecular alterations of TP53 are a defining feature of ovarian high-grade serous carcinoma: A rereview of cases lacking TP53 mutations in the cancer genome atlas ovarian study. Int J Gynecol Pathol. (2016) 35:48–55. doi: 10.1097/PGP.0000000000000207
31. Zhao Y, Wang C, Deng W, Li L, Liu J, Shi Y, et al. Patient-derived ovarian cancer organoid carries immune microenvironment and blood vessel keeping high response to cisplatin. MedComm. (2024) 5:e697. doi: 10.1002/mco2.697
32. Liu Z, Fan M, Hou J, Pan S, Xu Y, Zhang H, et al. Serine hydroxymethyltransferase 2 knockdown induces apoptosis in ccRCC by causing lysosomal membrane permeabilization via metabolic reprogramming. Cell Death Dis. (2023) 14:144. doi: 10.1038/s41419-023-05677-4
33. Fernández LP, Gómez de Cedrón M, Ramírez de Molina A. Alterations of lipid metabolism in cancer: implications in prognosis and treatment. Front Oncol. (2020) 10:577420. doi: 10.3389/fonc.2020.577420
34. Schalkwijk CG, Stehouwer CDA. Methylglyoxal, a highly reactive dicarbonyl compound, in diabetes, its vascular complications, and other age-related diseases. Physiol Rev. (2020) 100:407–61. doi: 10.1152/physrev.00001.2019
35. Chen S, Parmigiani G. Meta-analysis of BRCA1 and BRCA2 penetrance. J Clin Oncol. (2007) 25:1329–33. doi: 10.1200/JCO.2006.09.1066
36. King MC, Marks JH, Mandell JB. Breast and ovarian cancer risks due to inherited mutations in BRCA1 and BRCA2. Science. (2003) 302:643–6. doi: 10.1126/science.1088759
37. Luke B, Brown MB, Nichols HB, Schymura MJ, Browne ML, Fisher SC, et al. Assessment of birth defects and cancer risk in children conceived via in vitro fertilization in the US. JAMA Netw Open. (2020) 3:e2022927. doi: 10.1001/jamanetworkopen.2020.22927
38. Kong LR, Gupta K, Wu AJ, Perera D, Ivanyi-Nagy R, Ahmed SM, et al. A glycolytic metabolite bypasses “two-hit” tumor suppression by BRCA2. Cell. (2024) 187:2269–2287.e16. doi: 10.1016/j.cell.2024.03.006
39. Smith-Beckerman DM, Fung KW, Williams KE, Auersperg N, Godwin AK, Burlingame AL. Proteome changes in ovarian epithelial cells derived from women with BRCA1 mutations and family histories of cancer. Mol Cell Proteomics. (2005) 4:156–68. doi: 10.1074/mcp.M400157-MCP200
40. Savino AM, Stuani L. Targeting glycolysis to rescue 2-hydroxyglutarate immunosuppressive effects in dendritic cells and acute myeloid leukemia. Haematologica. (2024) 109:2388–90. doi: 10.3324/haematol.2023.284893
41. Borodovsky A, Barbon CM, Wang Y, Ye M, Prickett L, Chandra D, et al. Small molecule AZD4635 inhibitor of A(2A)R signaling rescues immune cell function including CD103(+) dendritic cells enhancing anti-tumor immunity. J Immunother Cancer. (2020) 8:e000417. doi: 10.1136/jitc-2019-000417
42. Notarangelo G, Spinelli JB, Perez EM, Baker GJ, Kurmi K, Elia I, et al. Oncometabolite d-2HG alters T cell metabolism to impair CD8(+) T cell function. Science. (2022) 377:1519–29. doi: 10.1126/science.abj5104
43. Ackermann T, Zuidhof HR, Müller C, Kortman G, Rutten MGS, Broekhuis MJC, et al. C/EBPβ-LIP mediated activation of the malate-aspartate shuttle sensitizes cells to glycolysis inhibition. Mol Metab. (2023) 72:101726. doi: 10.1016/j.molmet.2023.101726
44. Manohar SM. At the crossroads of TNF α Signaling and cancer. Curr Mol Pharmacol. (2024) 17:e060923220758. doi: 10.2174/1874467217666230908111754
45. Peng S, Lin A, Jiang A, Zhang C, Zhang J, Cheng Q, et al. CTLs heterogeneity and plasticity: implications for cancer immunotherapy. Mol Cancer. (2024) 23:58. doi: 10.1186/s12943-024-01972-6
46. Gao Z, Bai Y, Lin A, Jiang A, Zhou C, Cheng Q, et al. Gamma delta T-cell-based immune checkpoint therapy: attractive candidate for antitumor treatment. Mol Cancer. (2023) 22:31. doi: 10.1186/s12943-023-01722-0
47. Zhou J, Xu Y, Wang L, Cong Y, Huang K, Pan X, et al. LncRNA IDH1-AS1 sponges miR-518c-5p to suppress proliferation of epithelial ovarian cancer cell by targeting RMB47. J BioMed Res. (2023) 38:51–65. doi: 10.7555/JBR.37.20230097
48. Chen J, Hong JH, Huang Y, Liu S, Yin J, Deng P, et al. EZH2 mediated metabolic rewiring promotes tumor growth independently of histone methyltransferase activity in ovarian cancer. Mol Cancer. (2023) 22:85. doi: 10.1186/s12943-023-01786-y
49. Xu L, Li Y, Zhou L, Dorfman RG, Liu L, Cai R, et al. SIRT3 elicited an anti-Warburg effect through HIF1α/PDK1/PDHA1 to inhibit cholangiocarcinoma tumorigenesis. Cancer Med. (2019) 8:2380–91. doi: 10.1002/cam4.2019.8.issue-5
50. Lin TY, Gu SY, Lin YH, Shih JH, Lin JH, Chou TY, et al. Paclitaxel-resistance facilitates glycolytic metabolism via Hexokinase-2-regulated ABC and SLC transporter genes in ovarian clear cell carcinoma. BioMed Pharmacother. (2024) 180:117452. doi: 10.1016/j.biopha.2024.117452
51. Wang Y, Patti GJ. The Warburg effect: a signature of mitochondrial overload. Trends Cell Biol. (2023) 33:1014–20. doi: 10.1016/j.tcb.2023.03.013
52. Zhou J, Gu J, Qian Q, Zhang Y, Huang T, Li X, et al. Lactate supports Treg function and immune balance via MGAT1 effects on N-glycosylation in the mitochondria. J Clin Invest. (2024) 134:e175897. doi: 10.1172/JCI175897
53. Ciepła J, Smolarczyk R. Tumor hypoxia unveiled: insights into microenvironment, detection tools and emerging therapies. Clin Exp Med. (2024) 24:235. doi: 10.1007/s10238-024-01501-1
54. Zhou J, Yang R, Chen Y, Chen D. Efficacy tumor therapeutic applications of stimuli-responsive block copolymer-based nano-assemblies. Heliyon. (2024) 10:e28166. doi: 10.1016/j.heliyon.2024.e28166
55. Pastornická A, Rybárová S, Drahošová S, Mihalik J, Kreheľová A, Pavliuk-Karachevtseva A, et al. Influence of paclitaxel and doxorubicin therapy of ßIII-tubulin, carbonic anhydrase IX, and survivin in chemically induced breast cancer in female rat. Int J Mol Sci. (2021) 22:6363. doi: 10.3390/ijms22126363
56. Qiong L, Shuyao X, Shan X, Qian F, Jiaying T, Yao X, et al. Recent advances in the glycolytic processes linked to tumor metastasis. Curr Mol Pharmacol. (2024) 17:e18761429308361. doi: 10.2174/0118761429308361240823061634
57. Glaviano A, Lau HS, Carter LM, Lee EHC, Lam HY, Okina E, et al. Harnessing the tumor microenvironment: targeted cancer therapies through modulation of epithelial-mesenchymal transition. J Hematol Oncol. (2025) 18:6. doi: 10.1186/s13045-024-01634-6
58. Liu S, Sun Q, Ren X. Novel strategies for cancer immunotherapy: counter-immunoediting therapy. J Hematol Oncol. (2023) 16:38. doi: 10.1186/s13045-023-01430-8
59. Feng Q, Bennett Z, Grichuk A, Pantoja R, Huang T, Faubert B, et al. Severely polarized extracellular acidity around tumor cells. Nat BioMed Eng. (2024) 8:787–99. doi: 10.1038/s41551-024-01178-7
60. Steenbrugge J, Pauwelyn G, Demeyere K, Devriendt N, de Rooster H, Sanders NN, et al. Xenogeneic equine stem cells activate anti-tumor adaptive immunity in a 4T1-based intraductal mouse model for triple-negative breast cancer: proof-of-principle. Front Immunol. (2023) 14:1252374. doi: 10.3389/fimmu.2023.1252374
61. Li W, Ma W, Zhong H, Liu W, Sun Q. Metformin inhibits proliferation of human keratinocytes through a mechanism associated with activation of the MAPK signaling pathway. Exp Ther Med. (2014) 7:389–92. doi: 10.3892/etm.2013.1416
62. Wang Z, Yan Y, Lou Y, Huang X, Liu L, Weng Z, et al. Diallyl trisulfide alleviates chemotherapy sensitivity of ovarian cancer via the AMPK/SIRT1/PGC1α pathway. Cancer Sci. (2023) 114:357–69. doi: 10.1111/cas.v114.2
63. Yin CL, Ma YJ. The regulatory mechanism of hypoxia-inducible factor 1 and its clinical significance. Curr Mol Pharmacol. (2024) 17:e18761429266116. doi: 10.2174/0118761429266116231123160809
64. Chen Y, Liu L, Xia L, Wu N, Wang Y, Li H, et al. TRPM7 silencing modulates glucose metabolic reprogramming to inhibit the growth of ovarian cancer by enhancing AMPK activation to promote HIF-1α degradation. J Exp Clin Cancer Res. (2022) 41:44. doi: 10.1186/s13046-022-02252-1
65. Zheng Z, Ke L, Ye S, Shi P, Yao H. Pharmacological mechanisms of cryptotanshinone: recent advances in cardiovascular, cancer, and neurological disease applications. Drug Des Devel Ther. (2024) 18:6031–60. doi: 10.2147/DDDT.S494555
66. Mostafavi S, Hassan ZM. The anti-neoplastic effects of metformin modulate the acquired phenotype of fibroblast cells in the breast cancer-normal fibroblast co-culture system. Oncol Res. (2024) 32:477–87. doi: 10.32604/or.2023.043926
67. Tian L, Ogretmen B, Chung BY, Yu XZ. Sphingolipid metabolism in T cell responses after allogeneic hematopoietic cell transplantation. Front Immunol. (2022) 13:904823. doi: 10.3389/fimmu.2022.904823
68. Kim SB, Limbu KR, Oh YS, Kim SL, Park SK, Baek DJ, et al. Novel dimer derivatives of PF-543 as potential antitumor agents for the treatment of non-small cell lung cancer. Pharmaceutics. (2022) 14:2035. doi: 10.3390/pharmaceutics14102035
69. Han Y, Pan Q, Guo Z, Du Y, Zhang Y, Liu Y, et al. BMP9-induced vascular normalisation improves the efficacy of immunotherapy against hepatitis B virus-associated hepatocellular carcinoma. Clin Transl Med. (2023) 13:e1247. doi: 10.1002/ctm2.v13.5
70. Zhang J, Ouyang F, Gao A, Zeng T, Li M, Li H, et al. ESM1 enhances fatty acid synthesis and vascular mimicry in ovarian cancer by utilizing the PKM2-dependent warburg effect within the hypoxic tumor microenvironment. Mol Cancer. (2024) 23:94. doi: 10.1186/s12943-024-02009-8
71. Barrero CA, Datta PK, Sen S, Deshmane S, Amini S, Khalili K, et al. HIV-1 Vpr modulates macrophage metabolic pathways: a SILAC-based quantitative analysis. PloS One. (2013) 8:e68376. doi: 10.1371/journal.pone.0068376
72. Kilpatrick D, Walker SJ, Bell AJ. Importance of the great vessels in the genesis of the electrocardiogram. Circ Res. (1990) 66:1081–7. doi: 10.1161/01.RES.66.4.1081
73. Zang S, Lv LX, Liu CF, Zhang P, Li C, Wang JX. Metabolomic investigation of ultraviolet ray-inactivated white spot syndrome virus-induced trained immunity in marsupenaeus japonicus. Front Immunol. (2022) 13:885782. doi: 10.3389/fimmu.2022.885782
74. Zhang W, Zhou M, Chen C, Wu S, Wang L, Xia B, et al. Identification of CD98 as a novel biomarker for HIV-1 permissiveness and latent infection. mBio. (2022) 13:e0249622. doi: 10.1128/mbio.02496-22
75. Zhang Q, Li W. Correlation between amino acid metabolism and self-renewal of cancer stem cells: Perspectives in cancer therapy. World J Stem Cells. (2022) 14:267–86. doi: 10.4252/wjsc.v14.i4.267
76. Abdesheikhi J, Sedghy F, Mahmoodi M, Fallah H, Ranjkesh M. Metabolic regulation of T cell activity: implications for metabolic-based T-cell therapies for cancer. Iran BioMed J. (2023) 27:1–14. doi: 10.52547/ibj.3811
77. Sung JY, Cheong JH. New immunometabolic strategy based on cell type-specific metabolic reprogramming in the tumor immune microenvironment. Cells. (2022) 11(5):768. doi: 10.3390/cells11050768
78. Li J, Zheng C, Mai Q, Huang X, Pan W, Lu J, et al. Tyrosine catabolism enhances genotoxic chemotherapy by suppressing translesion DNA synthesis in epithelial ovarian cancer. Cell Metab. (2023) 35:2044–2059.e8. doi: 10.1016/j.cmet.2023.10.002
79. Xia D, Lai X, Wu K, Zhou P, Li L, Guo Z, et al. Metabolomics study of fasudil on cisplatin-induced kidney injury. Biosci Rep. (2019) 39:BSR20192940. doi: 10.1042/BSR20192940
80. Naser R, Fakhoury I, El-Fouani A, Abi-Habib R, El-Sibai M. Role of the tumor microenvironment in cancer hallmarks and targeted therapy (Review). Int J Oncol. (2023) 62:23. doi: 10.3892/ijo.2022.5471
81. Xiao Y, Yu D. Tumor microenvironment as a therapeutic target in cancer. Pharmacol Ther. (2021) 221:107753. doi: 10.1016/j.pharmthera.2020.107753
82. Zhang Z, Ding C, Sun T, Wang L, Chen C. Tumor therapy strategies based on microenvironment-specific responsive nanomaterials. Adv Healthc Mater. (2023) 12:e2300153. doi: 10.1002/adhm.202300153
83. Corn KC, Windham MA, Rafat M. Lipids in the tumor microenvironment: From cancer progression to treatment. Prog Lipid Res. (2020) 80:101055. doi: 10.1016/j.plipres.2020.101055
84. Sun H, Wang H, Wang X, Aoki Y, Wang X, Yang Y, et al. Aurora-A/SOX8/FOXK1 signaling axis promotes chemoresistance via suppression of cell senescence and induction of glucose metabolism in ovarian cancer organoids and cells. Theranostics. (2020) 10:6928–45. doi: 10.7150/thno.43811
85. Deng Z, Li B, Wang W, Xia W, Zhang L, Chen L, et al. TCEB2/HIF1A signaling axis promotes chemoresistance in ovarian cancer cells by enhancing glycolysis and angiogenesis. Eur J Med Res. (2024) 29:456. doi: 10.1186/s40001-024-02050-9
86. Wang Z, Chen W, Zuo L, Xu M, Wu Y, Huang J, et al. The Fibrillin-1/VEGFR2/STAT2 signaling axis promotes chemoresistance via modulating glycolysis and angiogenesis in ovarian cancer organoids and cells. Cancer Commun (Lond). (2022) 42:245–65. doi: 10.1002/cac2.12274
87. Lin F, Li X, Wang X, Sun H, Wang Z, Wang X. Stanniocalcin 1 promotes metastasis, lipid metabolism and cisplatin chemoresistance via the FOXC2/ITGB6 signaling axis in ovarian cancer. J Exp Clin Cancer Res. (2022) 41:129. doi: 10.1186/s13046-022-02315-3
88. Li J, Zou Y, Pei M, Zhang Y, Jiang Y. Berberine inhibits the Warburg effect through TET3/miR-145/HK2 pathways in ovarian cancer cells. J Cancer. (2021) 12:207–16. doi: 10.7150/jca.48896
89. Yan X, Yuan C, Wang Z, Xu Z, Wu Z, Wang M, et al. Berberine modulates ovarian cancer autophagy and glycolysis through the LINC01123/P65/MAPK10 signaling axis. Phytomedicine. (2024) 135:156121. doi: 10.1016/j.phymed.2024.156121
90. Wang T, Zhang M, Khan M, Li J, Wu X, Ma T, et al. Cryptotanshinone suppresses ovarian cancer via simultaneous inhibition of glycolysis and oxidative phosphorylation. BioMed Pharmacother. (2024) 170:115956. doi: 10.1016/j.biopha.2023.115956
91. Chun J. Isoalantolactone suppresses glycolysis and resensitizes cisplatin-based chemotherapy in cisplatin-resistant ovarian cancer cells. Int J Mol Sci. (2023) 24:12397. doi: 10.3390/ijms241512397
92. De Palma M, Biziato D, Petrova TV. Microenvironmental regulation of tumor angiogenesis. Nat Rev Cancer. (2017) 17:457–74. doi: 10.1038/nrc.2017.51
93. Jin S, Yin E, Feng C, Sun Y, Yang T, Yuan H, et al. Regulating tumor glycometabolism and the immune microenvironment by inhibiting lactate dehydrogenase with platinum(iv) complexes. Chem Sci. (2023) 14:8327–37. doi: 10.1039/D3SC01874A
94. Adiguzel D, Celik-Ozenci C. FoxO1 is a cell-specific core transcription factor for endometrial remodeling and homeostasis during menstrual cycle and early pregnancy. Hum Reprod Update. (2021) 27:570–83. doi: 10.1093/humupd/dmaa060
95. Sun Y, Peng YB, Ye LL, Ma LX, Zou MY, Cheng ZG. Propofol inhibits proliferation and cisplatin resistance in ovarian cancer cells through regulating the microRNA−374a/forkhead box O1 signaling axis. Mol Med Rep. (2020) 21:1471–80. doi: 10.3892/mmr.2020.10943
96. Zheng M, Cao MX, Luo XJ, Li L, Wang K, Wang SS, et al. EZH2 promotes invasion and tumor glycolysis by regulating STAT3 and FoxO1 signaling in human OSCC cells. J Cell Mol Med. (2019) 23:6942–54. doi: 10.1111/jcmm.v23.10
97. Lang X, Chen Z, Yang X, Yan Q, Xu M, Liu W, et al. Scutellarein induces apoptosis and inhibits proliferation, migration, and invasion in ovarian cancer via inhibition of EZH2/FOXO1 signaling. J Biochem Mol Toxicol. (2021) 35:e22870. doi: 10.1002/jbt.v35.10
98. Xiao Y, Yu Y, Jiang P, Li Y, Wang C, Zhang R. The PI3K/mTOR dual inhibitor GSK458 potently impedes ovarian cancer tumorigenesis and metastasis. Cell Oncol (Dordr). (2020) 43:669–80. doi: 10.1007/s13402-020-00514-8
99. Guo A, Lin J, Zhong P, Chen J, Wang L, Lin X, et al. Phellopterin attenuates ovarian cancer proliferation and chemoresistance by inhibiting the PU.1/CLEC5A/PI3K-AKT feedback loop. Toxicol Appl Pharmacol. (2023) 477:116691. doi: 10.1016/j.taap.2023.116691
100. Bélanger PM, Lalande M, Doré F, Labrecque G. Time-dependent variations in the organ extraction ratios of acetaminophen in rat. J Pharmacokinet Biopharm. (1987) 15:133–43. doi: 10.1007/BF01062340
101. Zhu Y, Yang L, Wang J, Li Y, Chen Y. SP1-induced lncRNA MCF2L-AS1 promotes cisplatin resistance in ovarian cancer by regulating IGF2BP1/IGF2/MEK/ERK axis. J Gynecol Oncol. (2022) 33:e75. doi: 10.3802/jgo.2022.33.e75
102. Zhang Q, Yan G, Lei J, Chen Y, Wang T, Gong J, et al. The SP1-12LOX axis promotes chemoresistance and metastasis of ovarian cancer. Mol Med. (2020) 26:39. doi: 10.1186/s10020-020-00174-2
103. Li H, Luo F, Jiang X, Zhang W, Xiang T, Pan Q, et al. CircITGB6 promotes ovarian cancer cisplatin resistance by resetting tumor-associated macrophage polarization toward the M2 phenotype. J Immunother Cancer. (2022) 10:e004029. doi: 10.1136/jitc-2021-004029
104. Nie S, Wan Y, Wang H, Liu J, Yang J, Sun R, et al. CXCL2-mediated ATR/CHK1 signaling pathway and platinum resistance in epithelial ovarian cancer. J Ovarian Res. (2021) 14:115. doi: 10.1186/s13048-021-00864-3
105. Zhang L, Liu Y, Lei X, Liu X, Sun H, Liu S. Astragaloside II enhanced sensitivity of ovarian cancer cells to cisplatin via triggering apoptosis and autophagy. Cell Biol Int. (2023) 47:1600–13. doi: 10.1002/cbin.12055
106. Zhou Y, Liu T, Wu Q, Wang H, Sun Y. Baohuoside I inhibits resistance to cisplatin in ovarian cancer cells by suppressing autophagy via downregulating HIF-1α/ATG5 axis. Mol Carcinog. (2023) 62:1474–86. doi: 10.1002/mc.v62.10
107. Song Z, Liao C, Yao L, Xu X, Shen X, Tian S, et al. miR-219-5p attenuates cisplatin resistance of ovarian cancer by inactivating Wnt/β-catenin signaling and autophagy via targeting HMGA2. Cancer Gene Ther. (2023) 30:596–607. doi: 10.1038/s41417-022-00574-y
108. Chiappa M, Guffanti F, Bertoni F, Colombo I, Damia G. Overcoming PARPi resistance: Preclinical and clinical evidence in ovarian cancer. Drug Resist Update. (2021) 55:100744. doi: 10.1016/j.drup.2021.100744
109. Tao M, Sun F, Wang J, Wang Y, Zhu H, Chen M, et al. Developing patient-derived organoids to predict PARP inhibitor response and explore resistance overcoming strategies in ovarian cancer. Pharmacol Res. (2022) 179:106232. doi: 10.1016/j.phrs.2022.106232
110. Hu W, Li M, Chen Y, Gu X. UBE2S promotes the progression and Olaparib resistance of ovarian cancer through Wnt/β-catenin signaling pathway. J Ovarian Res. (2021) 14:121. doi: 10.1186/s13048-021-00877-y
111. Zhang X, Yao J, Li X, Niu N, Liu Y, Hajek RA, et al. Targeting polyploid giant cancer cells potentiates a therapeutic response and overcomes resistance to PARP inhibitors in ovarian cancer. Sci Adv. (2023) 9:eadf7195. doi: 10.1126/sciadv.adf7195
112. Mukha A, Kahya U, Linge A, Chen O, Löck S, Lukiyanchuk V, et al. GLS-driven glutamine catabolism contributes to prostate cancer radiosensitivity by regulating the redox state, stemness and ATG5-mediated autophagy. Theranostics. (2021) 11:7844–68. doi: 10.7150/thno.58655
113. Wang Y, Wang P, Qin J. Human organoids and organs-on-chips for addressing COVID-19 challenges. Adv Sci (Weinh). (2022) 9:e2105187. doi: 10.1002/advs.202105187
114. Xia T, Du WL, Chen XY, Zhang YN. Organoid models of the tumor microenvironment and their applications. J Cell Mol Med. (2021) 25:5829–41. doi: 10.1111/jcmm.v25.13
115. Yan HHN, Chan AS, Lai FP, Leung SY. Organoid cultures for cancer modeling. Cell Stem Cell. (2023) 30:917–37. doi: 10.1016/j.stem.2023.05.012
116. Smith BA, Balanis NG, Nanjundiah A, Sheu KM, Tsai BL, Zhang Q, et al. A human adult stem cell signature marks aggressive variants across epithelial cancers. Cell Rep. (2018) 24:3353–3366.e5. doi: 10.1016/j.celrep.2018.08.062
Keywords: ovarian cancer, organoid, drug resistance, metabolic reprogramming, molecular mechanisms, immune escape, personalized therapy
Citation: Zhang L, Zhao J, Su C, Wu J, Jiang L, Chi H and Wang Q (2025) Organoid models of ovarian cancer: resolving immune mechanisms of metabolic reprogramming and drug resistance. Front. Immunol. 16:1573686. doi: 10.3389/fimmu.2025.1573686
Received: 09 February 2025; Accepted: 04 March 2025;
Published: 21 March 2025.
Edited by:
Haixia Zhu, Nantong Tumor Hospital, ChinaReviewed by:
Jian Zhang, Southern Medical University, ChinaCopyright © 2025 Zhang, Zhao, Su, Wu, Jiang, Chi and Wang. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Hao Chi, Y2hpaGFvQHN3bXUuZWR1LmNu; Qin Wang, d2FuZ3FAc3dtdS5lZHUuY24=
†These authors have contributed equally to this work
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.