
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
REVIEW article
Front. Immunol. , 19 March 2025
Sec. Molecular Innate Immunity
Volume 16 - 2025 | https://doi.org/10.3389/fimmu.2025.1563655
This article is part of the Research Topic Unraveling circRNAs and miRNAs: Key Regulators in Immune-Related Diseases View all 6 articles
Breast cancer is among the most prevalent malignant tumors worldwide, with triple-negative breast cancer (TNBC) being the most aggressive subtype and lacking effective treatment options. Circular RNAs (circRNAs) are noncoding RNAs that play crucial roles in the development of tumors, including breast cancer. This article examines the progress of research on circRNAs in breast cancer, focusing on four main areas: 1) breast cancer epidemiology, classification, and treatment; 2) the structure, discovery process, characteristics, formation, and functions of circRNAs; 3) the expression, mechanisms, clinical relevance, and recent advances in the study of circRNAs in breast cancer cells and the immune microenvironment, particularly in TNBC; and 4) the challenges and future prospects of the use of circRNAs in BC research.
Breast cancer remains a leading cause of cancer-related morbidity and mortality among women worldwide (1). According to the latest statistics, it accounts for a significant proportion of new cancer cases and deaths, underscoring the urgent need for effective prevention and treatment strategies (2). Among the various subtypes of breast cancer, triple-negative breast cancer (TNBC) is particularly notorious due to its aggressive nature and poor prognosis (3). TNBC is characterized by the absence of estrogen receptors, progesterone receptors, and human epidermal growth factor receptor 2 (HER2), which limits the options for targeted therapies that are effective in other breast cancer subtypes (4). Consequently, patients with TNBC often face a higher risk of recurrence and metastasis, making it imperative to explore novel therapeutic approaches and biomarkers (5).
Recent advances in molecular biology have shed light on the complex regulatory networks that govern cancer development and progression (6). One of the most intriguing discoveries in this context is the role of non-coding RNAs, particularly circular RNAs (circRNAs) (7). Unlike linear RNAs, circRNAs are characterized by their covalently closed circular structure, which confers them with increased stability and resistance to degradation (8). This unique feature enables circRNAs to function as molecular sponges for microRNAs (miRNAs), thereby modulating gene expression and influencing various cellular processes, including proliferation, apoptosis, and invasion (9).
The dysregulation of circRNAs has been implicated in numerous cancers, including breast cancer. Emerging evidence suggests that circRNAs can play multifaceted roles in tumor biology, acting as oncogenes or tumor suppressors depending on the context (10). For instance, certain circRNAs have been shown to promote tumor growth and metastasis by sequestering miRNAs that would otherwise inhibit oncogenic pathways. Conversely, other circRNAs may exert tumor-suppressive effects by promoting the expression of tumor suppressor genes or by interfering with pro-tumorigenic signaling pathways (11).
In the context of TNBC, circRNAs have garnered significant attention due to their potential to contribute to drug resistance, a major challenge in the treatment of this aggressive subtype (6). Studies have demonstrated that circRNAs can modulate the response of TNBC cells to chemotherapy and targeted therapies by regulating key signaling pathways, such as the PI3K/AKT/mTOR pathway (12). Furthermore, circRNAs may influence the tumor microenvironment (TME) by interacting with various immune cells, thereby shaping the immune landscape of the tumor and affecting its response to immunotherapy (13).
Despite the promising insights into the role of circRNAs in breast cancer, several challenges remain. The mechanisms by which circRNAs exert their effects on tumor biology are still not fully understood, and the functional diversity of circRNAs necessitates a comprehensive investigation into their specific roles in different breast cancer subtypes. Additionally, the potential of circRNAs as biomarkers for diagnosis, prognosis, and therapeutic response remains to be explored in clinical settings.
This review aims to provide a comprehensive overview of the current understanding of circRNAs in breast cancer, with a particular focus on TNBC. We will discuss the biogenesis, cellular localization, and degradation mechanisms of circRNAs, as well as their interactions with miRNAs and other molecular players in the tumor microenvironment. Furthermore, we will highlight the emerging evidence linking circRNAs to drug resistance and their potential as therapeutic targets. By synthesizing the existing literature, we hope to elucidate the multifaceted roles of circRNAs in breast cancer and pave the way for future research aimed at harnessing their potential for clinical application. Ultimately, understanding the intricate interplay between circRNAs and breast cancer biology could lead to innovative strategies for improving patient outcomes and advancing the field of cancer therapeutics.
Breast cancer arises from the carcinogenesis of breast epithelial tissue and has become one of the most prevalent malignant tumors worldwide, with an incidence rate second only to that of lung cancer (1) (Table 1). In 2022, breast cancer became the fifth leading cause of cancer-related deaths globally, accounting for 665,684 fatalities—a significantly greater number of deaths than other gynecological malignancies, such as cervical and ovarian cancers. Additionally, approximately 2.31 million women worldwide were newly diagnosed with breast cancer. The global incidence of breast cancer continues to increase at an annual rate of approximately 3.1% (2) (Table 2). In addition, the growth rate of breast cancer in Chinese women is much greater than that in the rest of the world (14). Furthermore, the age of breast cancer onset is decreasing. Combined with factors such as societal development, population growth, and an aging population, the economic burden of cancer on women worldwide has been steadily increasing (15).
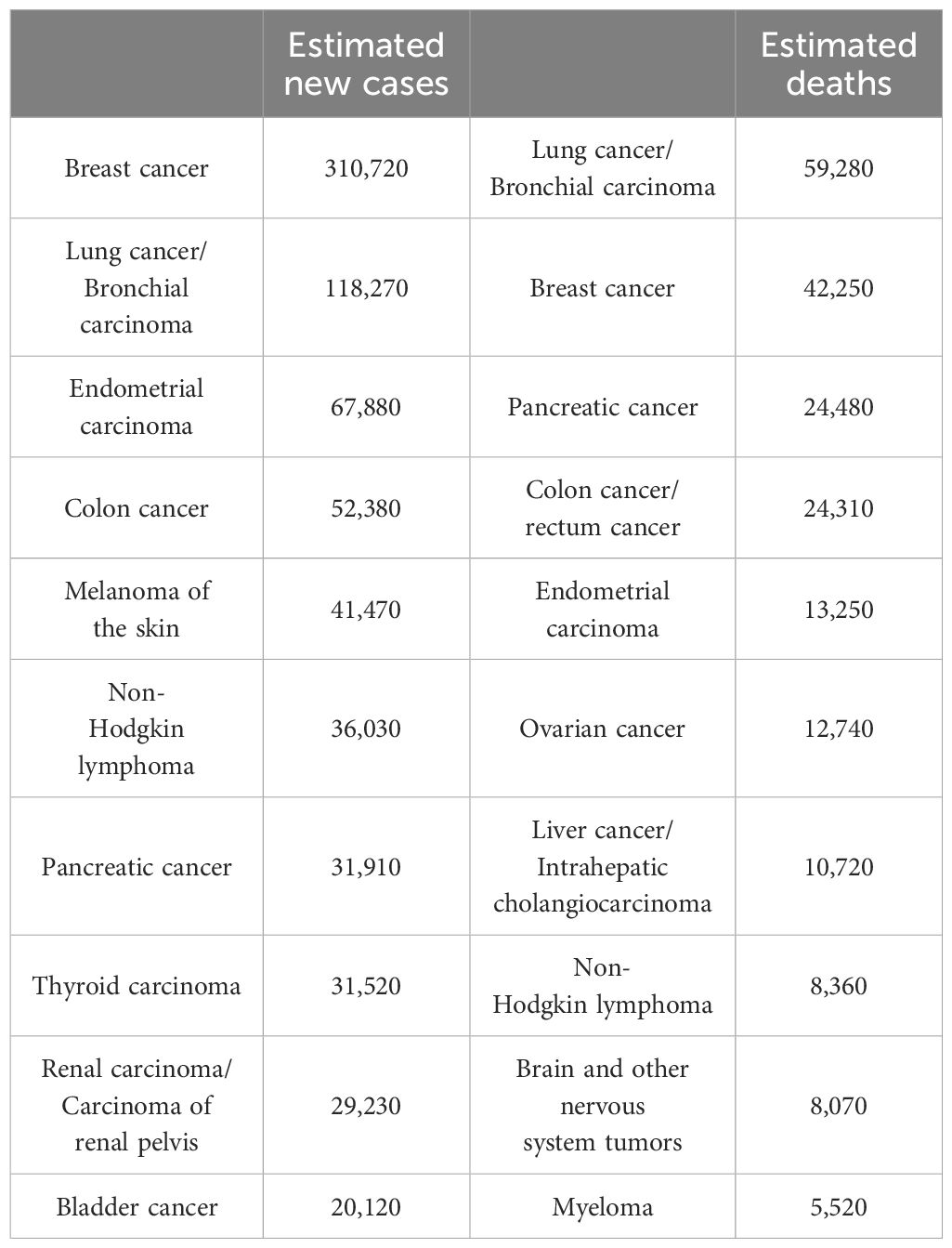
Table 1. The top ten cancers with estimated incidence rates and mortality rates in the female population, United States, 2024.
There are three main subtypes of breast cancer: ER/PR-overexpressing breast cancer (accounting for approximately 60% of cases), HER2-overexpressing breast cancer (approximately 20%), and triple-negative breast cancer (TNBC) (approximately 20%) (3). These subtypes differ in terms of gene expression, progression rates, metastasis potential, treatment approaches, and prognoses (4). The immune microenvironment of breast cancer is a complex network of immune cells, signaling molecules, and tumor components that play critical roles in tumor progression and immune evasion (16). The microenvironment includes various immune cells, such as T cells, macrophages, dendritic cells, and regulatory T cells, each of which contributes to either promoting or inhibiting the immune response. Tumor cells often employ immune checkpoint pathways, such as PD-L1, and secrete immunosuppressive factors such as TGF-β to evade immune detection (17). The characteristics of the immune microenvironment are closely linked to tumor aggressiveness, prognosis, and response to treatment, making the microenvironment a crucial focus for developing targeted immunotherapies (18). Breast cancer mortality rates have decreased due to early screening, detection, diagnosis, and improved treatment methods. However, TNBC is still associated with a poor prognosis and limited treatment options (19). Drug resistance, along with the potential for recurrence and metastasis, further complicates treatment (5). Recent evidence suggests that circRNAs may indirectly influence cancer progression through regulating gene expression (7). In particular, multiple circRNAs linked to breast cancer progression have been identified, which may serve as biomarkers and therapeutic targets for triple-negative breast cancer (6).
CircRNAs are noncoding, closed loop RNA molecules without 3’ and 5’ end caps that connect different splicing domains through covalent bonds. They have a highly stable structure and are resistant to degradation by nucleases (8). CircRNAs can be categorized into three types: exonic circRNAs (ecircRNAs), which contain only exon sequences; circular intronic RNAs (ciRNAs), which contain only intron sequences; and exon−intron circRNAs (EIciRNAs), which contain both exon and intron sequences (9). While most circRNAs are located in the cytoplasm, a few consisting solely of intron sequences are found in the nucleus (10).
CircRNAs were first discovered in 1976 in RNA viruses that cause diseases in higher plants (20). Subsequent research revealed additional circRNAs, with Hsu et al. identifying circRNA structures in the cytoplasm of HeLa cells in 1979 (21) and their presence in the hepatitis delta virus in 1986 (22). The stable structure of circRNAs was confirmed in 1988 (23), and endogenous circRNAs were identified in human cells in 1991 (24). Later, in 1995, Chen et al. demonstrated that engineered circRNAs can be translated in vitro, with the formation of circRNAs requiring reverse repeated recyclization (25, 26). In 1998, Perriman et al. confirmed that engineered circRNAs can be translated in vivo (27). In 2006, Suzuki et al. first enriched circRNAs using RNase R (28). Salzman et al. identified circRNAs using the gene expression program of human cells in 2012 (29). In 2013, Memczak et al. conducted functional analyses on circRNAs, providing insights into their potential roles in gene regulation (30). In 2016, researchers discovered that abnormally fused circRNAs can promote tumor growth and development (31), and in 2017, it was reported that some endogenous circRNAs can be translated, whereas others have no biological function in vivo (32, 33). In 2018, Huang et al. identified the regulatory mechanism of circRNA localization and posttranscriptional enucleation (34). Owing to the development of bioinformatics, over 183,000 circRNAs have been discovered in the human body (35) (Figure 1).
CircRNAs are widely distributed throughout organisms and are specifically expressed and regulated in various tissues and cells (36). Furthermore, they are detected in bodily fluids such as saliva (37), plasma (38), and exocrine bodies (39).
The specificity of circRNAs is prominently reflected in their dynamic and context-dependent expression patterns. CircRNAs exhibit strong tissue and cell type specificity, with distinct expression profiles across various biological contexts. Additionally, their spatiotemporal regulation allows them to play roles in specific developmental stages or physiological states, while aberrant expression is often linked to disease progression, including cancer, cardiovascular disorders, and neurodegenerative diseases (40, 41). These features make circRNAs highly specific biomarkers for diagnosis and potential therapeutic targets. A study in rats revealed that, during different lactation stages, many circRNAs are specifically expressed (42).
CircRNAs possess highly stable, closed-loop structures with connected 3’ and 5’ ends, rendering them resistant to degradation by nucleases, and their lack of 5’ cap and 3’ tail structures prevents degradation reactions such as decapping and deadenylation (43).
In terms of conservativeness, circRNAs are more conserved than their linear counterparts (44). Studies have shown that 4,522 of 15,849 circRNAs in mice have homologous sequences in humans (45), highlighting the potential translational value of circRNAs in future research.
CircRNAs are formed through one of three main models. The first model of circRNA formation is lasso-driven cyclization, in which partially folded pre-mRNAs cause exon hopping, connecting the downstream exon’s 3-SD (splicing donor) with the upstream exon’s 5’SA (splicing acceptor) to form an RNA lasso with both exons and introns. The introns are removed, resulting in ecircRNAs. The second model is intron-pairing-driven cyclization, which involves pairing and complementary binding between introns on the side of pre-mRNAs, forming a lasso that can either remove introns to create ecircRNAs or retain introns to form EIciRNAs. Finally, the third model involves RNA-binding protein-dependent cyclization, in which RNA-binding proteins (RBPs) bind to introns during transcription, leading to the formation of circRNAs (11, 12, 46–49). EcircRNAs are the most common type of circRNA (50) (Figure 2).
MicroRNAs (miRNAs) are small ncRNA molecules, approximately 19-24 nucleotides in length (51). By binding to genes through complementary base pairing, miRNAs can inhibit mRNA translation into proteins or cause mRNA degradation (52). CircRNAs possess various miRNA response elements (MREs) that complementarily bind to miRNAs, enabling interactions such as the sponge effect, where circRNAs regulate miRNA activity (53). Hansen et al. discovered the first sponge structure of a miRNA reaction element containing 74 miR-7 (54). The most well-known circRNA that acts as a miRNA sponge is cirRS-7 (55). By inhibiting miR-7, ciRS-7 can upregulate the expression of related genes (56). CcircRNA-Sry has been found to regulate tumor progression by sponging miR-138 (13). Studies have shown that circRNAs, as miRNA sponges, play crucial roles in regulating tumor progression (57, 58).
During the progression of breast cancer, various circRNAs are activated, including upregulated circRNAs such as hsa_circ_100876, which promote metastasis by inhibiting miR-361-3p (59), and circCER, which promotes metastasis by inhibiting miR-136 (60). Additionally, the downregulation of circRNAs such as hsa_circ_000911 via the inhibition of miR-449a negatively regulates the invasion of BC (61). Moreover, circGFRA1 promotes fat phagocytosis in TNBC through miR-34a (62).
CircRNAs have the ability to directly or indirectly regulate host gene expression. For example, circRNAs containing a starting codon, such as circ_CNOT2, can regulate linear genes by forcing them to use another starting codon for translation (63). Moreover, circRNAs can promote mRNA transcription (64) and are indirectly regulated by increasing the activity of RNA polymerase II (65). Li et al. reported that EIcRNAs can interact with nuclear ribonucleoprotein (U1snRNP) to promote host gene transcription (66). Additionally, SOX8 is involved in the maintenance of stem-like capacities in TNBC cells (67).
RBPs are crucial for posttranscriptional regulation, including mRNA splicing, stabilization, localization, modification, and translation (68). CircRNAs can interact with RBPs, altering the function of related proteins (69). For example, circCCNB1 can interact with cyclin-dependent kinase 1 and cyclin B1, affecting p53 gene mutation in breast cancer (70). Circ-PABPN1 can bind to the RBP HuR, hindering PABPN1 translation (71). Moreover, some circRNAs can act as scaffold molecules for protein−protein interactions. For example, circAmotl1 can bind to protein kinase B (AKT1) and inositol 3-phosphate-dependent protein kinase 1 (PDK1) (72).
CircRNAs reshape the tumor microenvironment (TME) by regulating epithelial–stromal transition, tumor angiogenesis, immune cell function, and the inflammatory response. Immune cells are the most abundant TME components and play a critical role in breast cancer cell progression (73). CircRNAs have been shown to promote cellular responses to external stimuli by binding to specific proteins. This is particularly evident in the rapid generation of immune responses following viral infections (74). Furthermore, some exogenous circRNAs can recognize the receptor RIG-1, consequently stimulating immune signal transduction in mammalian cells (75). CircRNAs in the TME can increase the expression of immune checkpoint molecules, including PD-L1, PD-1 and CD73, on the surface of tumor cells through miRNA sponge action and help tumor cells escape the toxic killing of immune cells (76).
Although circRNAs lack 3’ and 5’ end caps, experiments have revealed that artificial circRNAs containing open reading frames for green fluorescent protein can be translated into these proteins in E. coli (77). Some circRNAs containing open reading frames can participate in protein translation (11). CircRNAs are primarily involved in protein translation via the N6-methyladenosine-dependent pathway and the IRES-dependent pathway (78). They drive initial transcription and protein translation through base-modified N6 methyl adenosine (m6A) sites (79). CircRNAs with IRES sites are instrumental in protein translation after binding to ribosomes (80). Proteins translated by circRNAs potentially possess antitumor properties by blocking tumor metastasis or metabolism (81).
CircRNAs significantly impact the progression of BC by influencing various cellular processes, such as proliferation, invasion, apoptosis, and drug resistance (82). Wang et al. demonstrated that estrogen-induced circPGR plays a crucial role in ER-positive BC through circRNA sequencing (83). High-throughput sequencing has revealed a vast number of differentially expressed circRNAs in BC tissue compared with healthy tissue (84). Through gene chip analysis, 19 upregulated and 22 downregulated circRNAs were identified in breast cancer tissues compared with normal tissues (85). Specific circRNAs, such as circKIF4A, which is correlated with the survival rate of patients with TNBC (86), are upregulated, and circMYO9B is correlated with prognosis (87). Conversely, some circRNAs, such as circTADA2As, are downregulated, which is associated with the survival rate of patients with TNBC (88), and circ-NOL10, which is downregulated in breast cancer tissue (89) (Figure 3).
Breast cancer stem cells have been identified as major factors in the onset of breast cancer (90). These cells are also responsible for the frequent metastasis and recurrence of the disease because of their ability to promote tumor growth and their resistance to conventional treatments (91). Accordingly, research has identified breast cancer stem cells as potential targets for BC treatment (92, 93). Yan et al. hypothesized that circRNAs could act as miRNA sponges, influencing the regulation of breast cancer stem cell proliferation and renewal (94). Lin et al. discovered that TV-CircRGPD6 targets circRGPD6, which results in increased expression of p-H2AX, a DNA damage marker, and reduced expression of CD44, a stem cell marker (95). Additionally, circ_002178 overexpression promotes increased ALDH1 activity and increases the expression of stem cell markers in SUM149PT cells (96).
One of the circRNAs shown to play a role in BC is circRNF10-DHX15. This circRNA inhibits the proliferation of cancer cells by sponging the DHX15-NF-κB p65 positive feedback loop (97). Another circRNA, circZFAND6, which is a sponge of miR-647, promotes cell migration and proliferation in breast cancer (98). CircRNAs can also regulate the cell cycle and proliferation of BC through cancer-related signaling pathways. For example, circRNA-069718 promotes cell proliferation and invasion in TNBC via the Wnt activation/β-catenin pathway (99), whereas circ-ITCH inactivates the Wnt/β-catenin signal, inhibiting cell proliferation in TNBC (100).
EMT is a process in which epithelial cells transform into a mesenchymal phenotype through a specific program. As a result, a reduction in adhesion between tumor cells promotes tumor invasion and metastasis (101). EMT is known to cause tumorigenic changes in the tumor microenvironment (102). Several circRNAs have been shown to regulate EMT in BC. For example, circROBO1 promotes breast cancer carcinogenesis and liver metastasis via the circROFO1/KLF5/FUS feedback loop. Furthermore, circROBO1 inhibits the transcription of BECN1 and prevents the selective autophagy of afadin (103). Leng et al. reported that CIRC_0000043 regulates the miR-136/Smad3 axis to promote the EMT, invasion and metastasis of breast cancer (104). Several studies have examined the role of specific circRNAs in breast cancer metastasis. Song et al. reported that circHMCU targets EMT and G phase cell cycle checkpoints, thereby promoting the progression of BC (105). Additionally, the upregulation of circ_IRAK3 (hsa_circular RNA_0005505) in metastatic breast cancer is related to the recurrence of distant organs (106). Furthermore, Mao et al. reported that circRNA_000554 binds to miR-182 and suppresses epithelial stromal transformation in breast cancer (107). Conversely, Pan et al. demonstrated that the CIRC-TFF1 gene regulates the miR-326/TFF1 axis, thereby inhibiting the progression of BC (108).
Breast cancer cells thrive in a hypoxic tumor microenvironment, and one circRNA, HIF1α-associated circDENND4C, is known to promote their proliferation by combining with miR-200b and miR-200c, promoting glycolysis, invasion, and metastasis (109, 110). Li et al. reported that hsa_circ_0067842 is expressed in breast cancer, which promotes immune escape by stabilizing CMTM6 via HuR, subsequently inhibiting PD-L1 ubiquitination and degradation ultimately leading to increased PD-L1 expression and enhanced tumor immune evasion (111).
The most upregulated circRNA among the candidate genes in TNBC cells, circ-EPSTI1, can inhibit apoptosis, whereas knockout of circ-EPSTI1 can induce apoptosis (112). In addition, circ-FOXO3 is downregulated in BC cells but upregulated during apoptosis (113).
Many circRNAs have been shown to regulate the tumor immune microenvironment and cause immune escape of tumor cells (114). Several circRNAs have been proven to be related to the tumor immune microenvironment in breast cancer. Circ_0001142 is highly expressed in breast cancer cells, and the circ_0001142/miR-361-3p/PIK3CB pathway is involved in the polarization and autophagy of macrophages (115). In addition, circRNAs can regulate T-cell activity and immune checkpoint molecules. The circular RNA circWWC3 can increase IL-4 expression and secretion in breast cancer cells, which further enhances the expression of PD-L1 and facilitates breast cancer immune evasion (116). Ectopic circ_002172 expression could inhibit cytotoxic T lymphocyte infiltration in breast cancer and promote immune escape (117). CircRNAs have also been found to be associated with macrophages in breast cancer. Circ-Ccnb1 interacts with wild-type p53 and allows Bclaf1 to bind Bcl2, resulting in the induction of cell death (70). The circular RNA CDR1as/ciRS-7 plays an essential role in the immune microenvironment of breast cancer, including M2 macrophages. CDR1as regulates the TGF-β signaling pathway and ECM-receptor interaction to contribute to the immune microenvironment of breast cancer (118). These findings identify several circRNAs as diagnostic biomarkers and potential targets for breast cancer therapy.
CircRNAs are promising biomarkers for liquid biopsy because of their wide distribution in the body, stable structure, high sensitivity, and strong specificity (119, 120). There is a link between circUSP42 downregulation and the late clinical stage and lymph node metastasis of TNBC (121). The plasma concentration of hsa_circ_0001785 decreases significantly after surgical removal of breast cancer masses, suggesting its potential as a biomarker for evaluating prognosis (85). This may be attributed to the decrease in the amount of tumor-derived nucleic acid released after surgery (122). The overexpression of circWWC3 activates the RAS signaling pathway, which is associated with poor prognosis in BC patients (123).
Owing to their demonstrated association with cancer progression, multiple circRNAs have been identified as potential therapeutic targets for BC (124). CircRAD18 promotes the progression of TNBC by functioning as a miR-208a/3164 sponge to regulate IGF1 and FGF2 expression. Therefore, circRAD18 is a therapeutic target for TNBC (125). Through its involvement in the miR-675/NEDD4L axis, circKDM4B impedes breast cancer progression, particularly in terms of angiogenesis and tumor metastasis. Given these findings, circKDM4B has significant potential as a therapeutic target (126).
Chemotherapy reduces cancer metastasis and recurrence and ensures successful surgery. However, chemotherapy resistance arising from long-term use is the primary reason for breast cancer treatment failure (127). Common chemotherapy medications include adriamycin, cyclophosphamide, doxorubicin, paclitaxel, and 5-fluorourazalidine (128). Through the miR-361-5p/TLR4 pathway, circGFRA1 reduces TNBC cell sensitivity to paclitaxel (PTX) (129), whereas circ_0006528 promotes the progression of paclitaxel-resistant breast cancer cells. Nevertheless, it is possible to inhibit the growth of drug-resistant cells by silencing circ_0006528 (130). CircKDM4C binds to miR-548p and reduces breast cancer progression, thus slowing drug resistance to doxorubicin (131). Furthermore, circUBE2D2 disruption inhibits the invasion of TNBC and decreases doxorubicin resistance (132). Finally, studies have suggested that the downstream molecule Rafl of the hsa_circ_0006528/miRNA-7-5p axis influences BC cell resistance to doxorubicin (133).
CircRNAs affect the progression of TNBC, primarily as miRNA sponges. For example, Chen et al. discovered that estrogen receptor β2 (ERβ2) is responsible for mediating hsa_cir_0000732, acting as a sponge of miR-1184 and promoting the migration, growth and invasion of TNBC (134). Gong et al. reported that circUBR5 maintains the malignant growth of TNBC through miR-1179 absorption and UBR5 upregulation both in vitro and in vivo (135). According to Shao et al., circ_0004676 regulates miR-377-3p/E2F6/PNO1, playing a carcinogenic role in TNBC (136). In addition, hsa_cir_102229 regulates the expression of PFTK1 through combining with miR-152-3p, hence impacting TNBC cell function (137). In a study by Li et al., circ_0041732 promoted TNBC invasion and metastasis and regulated tumor characteristics via miR-149-5p/FGF5 (138). Finally, circFAM64A targets the 3’UTR of Cdc10-dependent transcript 1 (CDT1), and elevated CDT1 expression often correlates with a dismal prognosis (139). The circular RNA CDR1as/ciRS-7 has a comprehensive role in interacting with various immune cells, including T cells, NK cells and macrophages, and leads to TME reshaping and breast cancer progression (118).
Differential expression of circRNAs in TNBC and adjacent tissues has been previously observed. Magalhães et al. reported that 16 kinds of circRNAs were differentially expressed between TNBC patients and control individuals. Some circRNAs interact with RBPs involved in cancer and gene regulation pathways, including PTBP1, ELAVL1, EIF4A3, and AGO1/2 (140).
CircRNAs also impact cell cycle and proliferation of TNBC cells. For example, IGF2BP2 and HuR use circEIF3H as a scaffold to promote TNBC proliferation and metastasis, as reported by Song et al. (141). Additionally, Barznegar et al. reported that the upregulation of circ-ELP3 promotes TNBC development (142). The authors showed that circ_0076611 interacts with proliferation-related transcripts, thereby amplifying the progression of TNBC (143). In addition, Ruan et al. reported that circMETTL3 acts as a miR-34c-3p sponge to inhibit TNBC invasion and metastasis (144).
In TNBC, CircRNAs play a significant role in epithelial mesenchymal transformation, invasion, and metastasis. For example, overexpression of circBACH2 induces TNBC cell proliferation and epithelial mesenchymal transformation (145). CircCD44 promotes TNBC proliferation and invasion via the miR-502–5p/KRAS and IGF2BP2/Myc pathways (146). Zan et al. demonstrated that by targeting the miR-28-5p/LDHA pathway, circ-CSNK1G1 promotes TNBC cell invasion, metastasis, and glycolysis (147). CircPRKCI, as a miR-545-3p sponge, regulates the phosphorylation of WBP2 and AKT to increase TNBC migration (148). Via the miR-136–5p/PDK4 pathway, circERBB2 facilitates the Warburg effect and increases the growth of TNBC (149). Furthermore, the downregulation of circPTK2 promotes TNBC proliferation and invasion (150). Additionally, the knockdown of some circRNAs, such as circ_0062558, affects the miR-876-3p/SLC1A5 axis (151), circ_000520 affects the miR-1296/ZFX axis (152), and circDHDDS affects the miR-362-3p/DDX5 axis (153), induces the apoptosis of TNBC cells.
As a potential prognostic biomarker for TNBC, circNR3C2, combined with miR-513a-3p, inhibits HRD1-mediated tumor growth (154). Xing et al. reported that circ-PDCD11 overexpression is an independent risk factor associated with poor prognosis in TNBC patients (155). Similarly, circ-TRIO regulates miR-432-5p/CCD58 and can be a new prognostic marker for TNBC, as reported by Wang et al. (156). Moreover, Li et al. demonstrated that both the Wnt/beta-catenin pathway and MYH9 stabilization promote the progression of TNBC and that circ-EIF6-encoded EIF6-224a is involved in this process (157). These findings suggest that targeting circ-EIF6/EIF6-224 aa could be an approach for both prognostic and therapeutic purposes in TNBC treatment. CircPSMA1 can activate the miR-637/Akt1/β-catenin (cyclin D1) axis, promoting TNBC tumorigenesis and metastasis. These findings highlight the potential of circPSMA1 as a biomarker and target for TNBC immunotherapy (158). Chen et al. reported that circHIF1A regulates NFIB expression and translocation to promote TNBC progression. Additionally, circHIF1A is upregulated in the plasma, making it a potential target for TNBC diagnosis and treatment (159). The circRNA hsa_cir_0006220 has been found to inhibit tumor growth in TNBC by regulating miR-197-5p/CDH19, making it another promising therapeutic target (160). Furthermore, CircRAD54L2 modulates the miR-888 family/PDK1 axis to promote TNBC proliferation, invasion, and metastasis, making it another potential therapeutic target in TNBC (161).
CircRNAs can also affect the chemoresistance of TNBC. For example, cirCWAC induces chemoresistance by targeting miR-142, activating the PI3K/AKT pathway, and upregulating WWP1 (162). The circRNA CREIT reduces the stability of PKR and thus improves the drug resistance of TNBC to doxorubicin (163); moreover, CircINTS4 promotes the chemoresistance of TNBC through competitive binding with the miR-129-5p/POM121 axis (164). CircUBAP2, as a sponge of miR-300, upregulates ASF1B and triggers PI3K/AKT/mTOR (PAM) signaling to increase TNBC resistance to cisplatin (165) (Table 3).
1. Current studies have focused primarily on the role of circRNAs as miRNA molecular sponges in breast cancer, with less attention given to their interactions with other potential molecules, such as proteins or other RNA types. The biological functions of circRNAs and their role in regulating the breast tumor microenvironment remain incompletely understood. Furthermore, the clinical relevance of circRNAs in breast cancer, particularly with respect to chemotherapy resistance, has yet to be fully elucidated.
2. Some circRNAs undergo nuclear-to-cytoplasmic localization conversion, but the mechanisms underlying this process and their nuclear export remain poorly understood. Additionally, while circRNAs are characterized by stable closed-ring structures, their biogenesis and degradation mechanisms require further investigation.
3. Owing to their stability, abundance, and tissue specificity, circRNAs hold great potential in cancer diagnosis, prognosis, and treatment. They can serve as noninvasive biomarkers and therapeutic targets by regulating key pathways, such as miRNA sponging and drug resistance. Synthetic circRNAs or delivery systems could restore tumor-suppressing functions or counteract oncogenic signals. Additionally, their role in immune modulation offers prospects for enhancing immunotherapies. With further clinical validation, circRNAs could revolutionize precision medicine and address challenges such as drug resistance in cancer treatment. However, applying these findings to clinical practice presents a significant challenge.
Currently, research on circRNAs in breast cancer is an emerging field, and future directions can focus on the following areas:
1. Clarify circRNA biogenesis, cellular localization, and degradation mechanisms.
2. Investigate additional “sponge” effects of circRNAs on miRNAs and elucidate the mechanisms behind other biological functions of circRNAs.
3. Identify the mechanisms by which circRNAs interact with multiple immune cell types, reshape the breast TME, regulate breast cancer cells, and provide potential therapeutic targets.
4. Examine and identify the functions of circRNAs in BC, particularly their role in contributing to drug resistance in breast cancer treatments beyond chemotherapy (e.g., radiotherapy and immunotherapy).
5. Further screen and verify circRNA biomarker candidates and therapeutic targets are needed, with an emphasis on early clinical application.
6. Develop a standardized classification method for circRNA sources, distribution, functions, and naming conventions.
PC: Writing – original draft. JZ: Data curation, Writing – original draft. SW: Writing – original draft, Writing – review & editing. XZ: Data curation, Writing – original draft. WZ: Writing – original draft. ZG: Funding acquisition, Supervision, Writing – review & editing. HT: Conceptualization, Supervision, Writing – review & editing.
The author(s) declare that financial support was received for the research and/or publication of this article. This research was funded by the Guangdong Basic and Applied Basic Research Foundation (2021B1515120053, Ziyun Guan).
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
1. Siegel RL, Giaquinto AN, Jemal A. Cancer statistics, 2024. CA: Cancer J Clin. (2024) 74:12–49. doi: 10.3322/caac.21820
2. Bray F, Laversanne M, Sung H, Ferlay J, Siegel RL, Soerjomataram I, et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: Cancer J Clin. (2024) 74:229–63. doi: 10.3322/caac.21834
3. McAnena PF, McGuire A, Ramli A, Curran C, Malone C, McLaughlin R, et al. Breast cancer subtype discordance: impact on post-recurrence survival and potential treatment options. BMC cancer. (2018) 18:203. doi: 10.1186/s12885-018-4101-7
4. Neophytou C, Boutsikos P, Papageorgis P. Molecular mechanisms and emerging therapeutic targets of triple-negative breast cancer metastasis. Front Oncol. (2018) 8:31. doi: 10.3389/fonc.2018.00031
5. Garrido-Castro AC, Lin NU, Polyak K. Insights into molecular classifications of triple-negative breast cancer: improving patient selection for treatment. Cancer discovery. (2019) 9:176–98. doi: 10.1158/2159-8290.CD-18-1177
6. Zhang HD, Jiang LH, Sun DW, Hou JC, Ji ZL. CircRNA: a novel type of biomarker for cancer. Breast Cancer (Tokyo Japan). (2018) 25:1–7. doi: 10.1007/s12282-017-0793-9
7. Lyu L, Zhang S, Deng Y, Wang M, Deng X, Yang S, et al. Regulatory mechanisms, functions, and clinical significance of CircRNAs in triple-negative breast cancer. J Hematol Oncol. (2021) 14:41. doi: 10.1186/s13045-021-01052-y
8. Zhou WY, Cai ZR, Liu J, Wang DS, Ju HQ, Xu RH. Circular RNA: metabolism, functions and interactions with proteins. Mol cancer. (2020) 19:172. doi: 10.1186/s12943-020-01286-3
9. Qi Y, Han W, Chen D, Zhao J, Bai L, Huang F, et al. Engineering circular RNA regulators to specifically promote circular RNA production. Theranostics. (2021) 11:7322–36. doi: 10.7150/thno.56990
10. Cheng D, Wang J, Dong Z, Li X. Cancer-related circular RNA: diverse biological functions. Cancer Cell Int. (2021) 21:11. doi: 10.1186/s12935-020-01703-z
11. Kristensen LS, Andersen MS, Stagsted LVW, Ebbesen KK, Hansen TB, Kjems J. The biogenesis, biology and characterization of circular RNAs. Nat Rev Genet. (2019) 20:675–91. doi: 10.1038/s41576-019-0158-7
12. Tran AM, Chalbatani GM, Berland L, Cruz De Los Santos M, Raj P, Jalali SA, et al. A new world of biomarkers and therapeutics for female reproductive system and breast cancers: circular RNAs. Front Cell Dev Biol. (2020) 8:50. doi: 10.3389/fcell.2020.00050
13. Meng X, Li X, Zhang P, Wang J, Zhou Y, Chen M. Circular RNA: an emerging key player in RNA world. Briefings Bioinf. (2017) 18:547–57. doi: 10.1093/bib/bbw045
14. Chen W, Zheng R, Baade PD, Zhang S, Zeng H, Bray F, et al. Cancer statistics in China, 2015. CA: Cancer J Clin. (2016) 66:115–32. doi: 10.3322/caac.21338
15. Li X, Xin P, Wang C, Wang Z, Wang Q, Kuang H. Mechanisms of traditional Chinese medicine in the treatment of mammary gland hyperplasia. Am J Chin Med. (2017) 45:443–58. doi: 10.1142/S0192415X17500276
16. Harris MA, Savas P, Virassamy B, O’Malley MMR, Kay J, Mueller SN, et al. Towards targeting the breast cancer immune microenvironment. Nat Rev Cancer. (2024) 24:554–77. doi: 10.1038/s41568-024-00714-6
17. Onkar SS, Carleton NM, Lucas PC, Bruno TC, Lee AV, Vignali DAA, et al. The great immune escape: understanding the divergent immune response in breast cancer subtypes. Cancer discovery. (2023) 13:23–40. doi: 10.1158/2159-8290.CD-22-0475
18. Kundu M, Butti R, Panda VK, Malhotra D, Das S, Mitra T, et al. Modulation of the tumor microenvironment and mechanism of immunotherapy-based drug resistance in breast cancer. Mol cancer. (2024) 23:92. doi: 10.1186/s12943-024-01990-4
19. Zhang J, Guan H, Song C, Liu L, Tang H, Wang J. Locoregional treatments of metastatic internal mammary node following neoadjuvant chemotherapy. Chin Med J. (2025) 138:358–60. doi: 10.1097/CM9.0000000000003397
20. Sanger HL, Klotz G, Riesner D, Gross HJ, Kleinschmidt AK. Viroids are single-stranded covalently closed circular RNA molecules existing as highly base-paired rod-like structures. Proc Natl Acad Sci United States America. (1976) 73:3852–6. doi: 10.1073/pnas.73.11.3852
21. Hsu MT, Coca-Prados M. Electron microscopic evidence for the circular form of RNA in the cytoplasm of eukaryotic cells. Nature. (1979) 280:339–40. doi: 10.1038/280339a0
22. Kos A, Dijkema R, Arnberg AC, van der Meide PH, Schellekens H. The hepatitis delta (delta) virus possesses a circular RNA. Nature. (1986) 323:558–60. doi: 10.1038/323558a0
23. Harland R, Misher L. Stability of RNA in developing Xenopus embryos and identification of a destabilizing sequence in TFIIIA messenger RNA. Dev (Cambridge England). (1988) 102:837–52. doi: 10.1242/dev.102.4.837
24. Nigro JM, Cho KR, Fearon ER, Kern SE, Ruppert JM, Oliner JD, et al. Scrambled exons. Cell. (1991) 64:607–13. doi: 10.1016/0092-8674(91)90244-S
25. Chen CY, Sarnow P. Initiation of protein synthesis by the eukaryotic translational apparatus on circular RNAs. Sci (New York NY). (1995) 268:415–7. doi: 10.1126/science.7536344
26. Dubin RA, Kazmi MA, Ostrer H. Inverted repeats are necessary for circularization of the mouse testis Sry transcript. Gene. (1995) 167:245–8. doi: 10.1016/0378-1119(95)00639-7
27. Perriman R, Ares M Jr. Circular mRNA can direct translation of extremely long repeating-sequence proteins in vivo. RNA (New York NY). (1998) 4:1047–54. doi: 10.1017/S135583829898061X
28. Suzuki H, Zuo Y, Wang J, Zhang MQ, Malhotra A, Mayeda A. Characterization of RNase R-digested cellular RNA source that consists of lariat and circular RNAs from pre-mRNA splicing. Nucleic Acids Res. (2006) 34:e63. doi: 10.1093/nar/gkl151
29. Salzman J, Gawad C, Wang PL, Lacayo N, Brown PO. Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PloS One. (2012) 7:e30733. doi: 10.1371/journal.pone.0030733
30. Memczak S, Jens M, Elefsinioti A, Torti F, Krueger J, Rybak A, et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature. (2013) 495:333–8. doi: 10.1038/nature11928
31. Guarnerio J, Bezzi M, Jeong JC, Paffenholz SV, Berry K, Naldini MM, et al. Oncogenic role of fusion-circRNAs derived from cancer-associated chromosomal translocations. Cell. (2016) 165:289–302. doi: 10.1016/j.cell.2016.03.020
32. Pamudurti NR, Bartok O, Jens M, Ashwal-Fluss R, Stottmeister C, Ruhe L, et al. Translation of circRNAs. Mol Cell. (2017) 66:9–21.e7. doi: 10.1016/j.molcel.2017.02.021
33. Piwecka M, Glažar P, Hernandez-Miranda LR, Memczak S, Sa W, Rybak-Wolf A, et al. Loss of a mammalian circular RNA locus causes miRNA deregulation and affects brain function. Sci (New York NY). (2017) 357(6357):eaam8526. doi: 10.1126/science.aam8526
34. Huang C, Liang D, Tatomer DC, Wilusz JE. A length-dependent evolutionarily conserved pathway controls nuclear export of circular RNAs. Genes Dev. (2018) 32:639–44. doi: 10.1101/gad.314856.118
35. Chen LL. The expanding regulatory mechanisms and cellular functions of circular RNAs. Nat Rev Mol Cell Biol. (2020) 21:475–90. doi: 10.1038/s41580-020-0243-y
36. Xie J, Ye F, Deng X, Tang Y, Liang JY, Huang X, et al. Circular RNA: A promising new star of vaccine. J Trans Internal Med. (2023) 11:372–81. doi: 10.2478/jtim-2023-0122
37. Zhao SY, Wang J, Ouyang SB, Huang ZK, Liao L. Salivary circular RNAs hsa_Circ_0001874 and hsa_Circ_0001971 as novel biomarkers for the diagnosis of oral squamous cell carcinoma. Cell Physiol biochem: Int J Exp Cell physiol biochem Pharmacol. (2018) 47:2511–21. doi: 10.1159/000491624
38. Zhang H, Zhu L, Bai M, Liu Y, Zhan Y, Deng T, et al. Exosomal circRNA derived from gastric tumor promotes white adipose browning by targeting the miR-133/PRDM16 pathway. Int J cancer. (2019) 144:2501–15. doi: 10.1002/ijc.v144.10
39. Maass PG, Glažar P, Memczak S, Dittmar G, Hollfinger I, Schreyer L, et al. A map of human circular RNAs in clinically relevant tissues. Journal of molecular medicine (Berlin, Germany). J Mol Med (Berl). (2017) 95:1179–89. doi: 10.1007/s00109-017-1582-9
40. Qiao X, Ding Y, Altawil A, Yin Y, Wang Q, Wang W, et al. Roles of noncoding RNAs in chronic obstructive pulmonary disease. J Trans Internal Med. (2023) 11:106–10. doi: 10.2478/jtim-2023-0084
41. Ma B, Wang S, Wu W, Shan P, Chen Y, Meng J, et al. Mechanisms of circRNA/lncRNA-miRNA interactions and applications in disease and drug research. Biomed pharmacother = Biomed pharmacotherapie. (2023) 162:114672. doi: 10.1016/j.biopha.2023.114672
42. Shi Y, Guo Y, Zhou J, Cui G, Cheng JC, Wu Y, et al. A spatiotemporal gene expression and cell atlases of the developing rat ovary. Cell proliferation. (2023) 56:e13516. doi: 10.1111/cpr.v56.12
43. Zhang Z, Yang T, Xiao J. Circular RNAs: promising biomarkers for human diseases. EBioMedicine. (2018) 34:267–74. doi: 10.1016/j.ebiom.2018.07.036
44. Du WW, Fang L, Yang W, Wu N, Awan FM, Yang Z, et al. Induction of tumor apoptosis through a circular RNA enhancing Foxo3 activity. Cell Death differentiation. (2017) 24:357–70. doi: 10.1038/cdd.2016.133
45. Rybak-Wolf A, Stottmeister C, Glažar P, Jens M, Pino N, Giusti S, et al. Circular RNAs in the mammalian brain are highly abundant, conserved, and dynamically expressed. Mol Cell. (2015) 58:870–85. doi: 10.1016/j.molcel.2015.03.027
46. Jeck WR, Sorrentino JA, Wang K, Slevin MK, Burd CE, Liu J, et al. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA (New York NY). (2013) 19:141–57. doi: 10.1261/rna.035667.112
47. Barrett SP, Wang PL, Salzman J. Circular RNA biogenesis can proceed through an exon-containing lariat precursor. eLife. (2015) 4:e07540. doi: 10.7554/eLife.07540
48. Chen J, Gu J, Tang M, Liao Z, Tang R, Zhou L, et al. Regulation of cancer progression by circRNA and functional proteins. J Cell Physiol. (2022) 237:373–88. doi: 10.1002/jcp.v237.1
49. Gao Y, Li J, Ma M, Fu W, Ma L, Sui Y, et al. Prognostic prediction of m6A and ferroptosis-associated lncRNAs in liver hepatocellular carcinoma. J Trans Internal Med. (2024) 12:526–9. doi: 10.1515/jtim-2024-0023
50. Zaiou M. Circular RNAs as potential biomarkers and therapeutic targets for metabolic diseases. Adv Exp Med Biol. (2019) 1134:177–91. doi: 10.1007/978-3-030-12668-1_10
51. Misir S, Aliyazicioglu Y, Demir S, Turan I, Hepokur C. Effect of turkish propolis on miRNA expression, cell cycle, and apoptosis in human breast cancer (MCF-7) cells. Nutr cancer. (2020) 72:133–45. doi: 10.1080/01635581.2019.1616100
53. Wang X, Fang L. Advances in circular RNAs and their roles in breast Cancer. J Exp Clin Cancer research: CR. (2018) 37:206. doi: 10.1186/s13046-018-0870-8
54. Hansen TB, Wiklund ED, Bramsen JB, Villadsen SB, Statham AL, Clark SJ, et al. miRNA-dependent gene silencing involving Ago2-mediated cleavage of a circular antisense RNA. EMBO J. (2011) 30:4414–22. doi: 10.1038/emboj.2011.359
55. Kleaveland B, Shi CY, Stefano J, Bartel DP. A network of noncoding regulatory RNAs acts in the mammalian brain. Cell. (2018) 174:350–62.e17. doi: 10.1016/j.cell.2018.05.022
56. van Rossum D, Verheijen BM, Pasterkamp RJ. Circular RNAs: novel regulators of neuronal development. Front Mol Neurosci. (2016) 9:74. doi: 10.3389/fnmol.2016.00074
57. Zou Y, Ye F, Kong Y, Hu X, Deng X, Xie J, et al. The single-cell landscape of intratumoral heterogeneity and the immunosuppressive microenvironment in liver and brain metastases of breast cancer. Advanced Sci (Weinheim Baden-Wurttemberg Germany). (2023) 10:e2203699. doi: 10.1002/advs.202203699
58. Zhao C, Qian S, Tai Y, Guo Y, Tang C, Huang Z, et al. Proangiogenic role of circRNA-007371 in liver fibrosis. Cell proliferation. (2023) 56:e13432. doi: 10.1111/cpr.13432
59. Yang CY, Zhang FX, He JN, Wang SQ. CircRNA_100876 promote proliferation and metastasis of breast cancer cells through adsorbing microRNA-361-3p in a sponge form. Eur Rev Med Pharmacol Sci. (2019) 23:6962–70. doi: 10.26355/eurrev_201908_18736
60. Qu Y, Dou P, Hu M, Xu J, Xia W, Sun H. circRNA−CER mediates Malignant progression of breast cancer through targeting the miR−136/MMP13 axis. Mol Med Rep. (2019) 19:3314–20. doi: 10.3892/mmr.2019.9965
61. Wang H, Xiao Y, Wu L, Ma D. Comprehensive circular RNA profiling reveals the regulatory role of the circRNA-000911/miR-449a pathway in breast carcinogenesis. Int J Oncol. (2018) 52:743–54. doi: 10.3892/ijo.2018.4265
62. He R, Liu P, Xie X, Zhou Y, Liao Q, Xiong W, et al. circGFRA1 and GFRA1 act as ceRNAs in triple negative breast cancer by regulating miR-34a. J Exp Clin Cancer research: CR. (2017) 36:145. doi: 10.1186/s13046-017-0614-1
63. Smid M, Wilting SM, Uhr K, Rodríguez-González FG, de Weerd V, Prager-Van-der-Smissen WJC, et al. The circular RNome of primary breast cancer. Genome Res. (2019) 29:356–66. doi: 10.1101/gr.238121.118
64. Prats AC, David F, Diallo LH, Roussel E, Tatin F, Garmy-Susini B, et al. Circular RNA, the key for translation. Int J Mol Sci. (2020) 21 (22):8591. doi: 10.3390/ijms21228591
65. Zhao X, Cai Y, Xu J. Circular RNAs: biogenesis, mechanism, and function in human cancers. Int J Mol Sci. (2019) 20(16):3926. doi: 10.3390/ijms20163926
66. Li Z, Huang C, Bao C, Chen L, Lin M, Wang X, et al. Exon-intron circular RNAs regulate transcription in the nucleus. Nat Struct Mol Biol. (2015) 22:256–64. doi: 10.1038/nsmb.2959
67. Tang H, Chen B, Liu P, Xie X, He R, Zhang L, et al. SOX8 acts as a prognostic factor and mediator to regulate the progression of triple-negative breast cancer. Carcinogenesis. (2019) 40:1278–87. doi: 10.1093/carcin/bgz034
68. Okholm TLH, Sathe S, Park SS, Kamstrup AB, Rasmussen AM, Shankar A, et al. Transcriptome-wide profiles of circular RNA and RNA-binding protein interactions reveal effects on circular RNA biogenesis and cancer pathway expression. Genome Med. (2020) 12:112. doi: 10.1186/s13073-020-00812-8
69. Yang F, Liu Y, Xiao J, Li B, Chen Y, Hu A, et al. Circ-CTNNB1 drives aerobic glycolysis and osteosarcoma progression via m6A modification through interacting with RBM15. Cell proliferation. (2023) 56:e13344. doi: 10.1111/cpr.13344
70. Fang L, Du WW, Lyu J, Dong J, Zhang C, Yang W, et al. Enhanced breast cancer progression by mutant p53 is inhibited by the circular RNA circ-Ccnb1. Cell Death differentiation. (2018) 25:2195–208. doi: 10.1038/s41418-018-0115-6
71. Li XX, Xiao L, Chung HK, Ma XX, Liu X, Song JL, et al. Interaction between huR and circPABPN1 modulates autophagy in the intestinal epithelium by altering ATG16L1 translation. Mol Cell Biol. (2020) 40(6):e00492–19. doi: 10.1128/MCB.00492-19
72. Zeng Y, Du WW, Wu Y, Yang Z, Awan FM, Li X, et al. A circular RNA binds to and activates AKT phosphorylation and nuclear localization reducing apoptosis and enhancing cardiac repair. Theranostics. (2017) 7:3842–55. doi: 10.7150/thno.19764
73. Xu Z, Chen Y, Ma L, Chen Y, Liu J, Guo Y, et al. Role of exosomal non-coding RNAs from tumor cells and tumor-associated macrophages in the tumor microenvironment. Mol therapy: J Am Soc Gene Ther. (2022) 30:3133–54. doi: 10.1016/j.ymthe.2022.01.046
74. Liu CX, Li X, Nan F, Jiang S, Gao X, Guo SK, et al. Structure and degradation of circular RNAs regulate PKR activation in innate immunity. Cell. (2019) 177:865–80.e21. doi: 10.1016/j.cell.2019.03.046
75. Chen YG, Kim MV, Chen X, Batista PJ, Aoyama S, Wilusz JE, et al. Sensing self and foreign circular RNAs by intron identity. Mol Cell. (2017) 67:228–38.e5. doi: 10.1016/j.molcel.2017.05.022
76. Liu Z, Wang T, She Y, Wu K, Gu S, Li L, et al. N(6)-methyladenosine-modified circIGF2BP3 inhibits CD8(+) T-cell responses to facilitate tumor immune evasion by promoting the deubiquitination of PD-L1 in non-small cell lung cancer. Mol cancer. (2021) 20:105. doi: 10.1186/s12943-021-01398-4
77. Kong S, Tao M, Shen X, Ju S. Translatable circRNAs and lncRNAs: Driving mechanisms and functions of their translation products. Cancer letters. (2020) 483:59–65. doi: 10.1016/j.canlet.2020.04.006
78. Zhang L, Hou C, Chen C, Guo Y, Yuan W, Yin D, et al. The role of N(6)-methyladenosine (m(6)A) modification in the regulation of circRNAs. Mol cancer. (2020) 19:105. doi: 10.1186/s12943-020-01224-3
79. Yang Y, Fan X, Mao M, Song X, Wu P, Zhang Y, et al. Extensive translation of circular RNAs driven by N(6)-methyladenosine. Cell Res. (2017) 27:626–41. doi: 10.1038/cr.2017.31
80. Lei M, Zheng G, Ning Q, Zheng J, Dong D. Translation and functional roles of circular RNAs in human cancer. Mol cancer. (2020) 19:30. doi: 10.1186/s12943-020-1135-7
81. Wu P, Mo Y, Peng M, Tang T, Zhong Y, Deng X, et al. Emerging role of tumor-related functional peptides encoded by lncRNA and circRNA. Mol cancer. (2020) 19:22. doi: 10.1186/s12943-020-1147-3
82. Tang X, Ren H, Guo M, Qian J, Yang Y, Gu C. Review on circular RNAs and new insights into their roles in cancer. Comput Struct Biotechnol J. (2021) 19:910–28. doi: 10.1016/j.csbj.2021.01.018
83. Wang L, Yi J, Lu LY, Zhang YY, Wang L, Hu GS, et al. Estrogen-induced circRNA, circPGR, functions as a ceRNA to promote estrogen receptor-positive breast cancer cell growth by regulating cell cycle-related genes. Theranostics. (2021) 11:1732–52. doi: 10.7150/thno.45302
84. Wang J, Zhang Q, Zhou S, Xu H, Wang D, Feng J, et al. Circular RNA expression in exosomes derived from breast cancer cells and patients. Epigenomics. (2019) 11:411–21. doi: 10.2217/epi-2018-0111
85. Yin WB, Yan MG, Fang X, Guo JJ, Xiong W, Zhang RP. Circulating circular RNA hsa_circ_0001785 acts as a diagnostic biomarker for breast cancer detection. Clinica chimica acta; Int J Clin Chem. (2018) 487:363–8. doi: 10.1016/j.cca.2017.10.011
86. Tang H, Huang X, Wang J, Yang L, Kong Y, Gao G, et al. circKIF4A acts as a prognostic factor and mediator to regulate the progression of triple-negative breast cancer. Mol cancer. (2019) 18:23. doi: 10.1186/s12943-019-0946-x
87. Wang N, Gu Y, Li L, Wang F, Lv P, Xiong Y, et al. Circular RNA circMYO9B facilitates breast cancer cell proliferation and invasiveness via upregulating FOXP4 expression by sponging miR-4316. Arch Biochem biophysics. (2018) 653:63–70. doi: 10.1016/j.abb.2018.04.017
88. Xu JZ, Shao CC, Wang XJ, Zhao X, Chen JQ, Ouyang YX, et al. circTADA2As suppress breast cancer progression and metastasis via targeting miR-203a-3p/SOCS3 axis. Cell Death disease. (2019) 10:175. doi: 10.1038/s41419-019-1382-y
89. Cai Y, Zhao X, Chen D, Zhang F, Chen Q, Shao CC, et al. circ-NOL10 regulated by MTDH/CASC3 inhibits breast cancer progression and metastasis via multiple miRNAs and PDCD4. Mol Ther Nucleic Acids. (2021) 26:773–86. doi: 10.1016/j.omtn.2021.09.013
90. Cully M. Cancer: Fibroblast subtype provides niche for cancer stem cells. Nat Rev Drug discovery. (2018) 17:163. doi: 10.1038/nrd.2018.23
91. Gelsomino L, Panza S, Giordano C, Barone I, Gu G, Spina E, et al. Mutations in the estrogen receptor alpha hormone binding domain promote stem cell phenotype through notch activation in breast cancer cell lines. Cancer letters. (2018) 428:12–20. doi: 10.1016/j.canlet.2018.04.023
92. Zhang R, Tu J, Liu S. Novel molecular regulators of breast cancer stem cell plasticity and heterogeneity. Semin Cancer Biol. (2022) 82:11–25. doi: 10.1016/j.semcancer.2021.03.008
93. Wang H, Xu H, Chen W, Cheng M, Zou L, Yang Q, et al. Rab13 sustains breast cancer stem cells by supporting tumor-stroma cross-talk. Cancer Res. (2022) 82:2124–40. doi: 10.1158/0008-5472.CAN-21-4097
94. Yan N, Xu H, Zhang J, Xu L, Zhang Y, Zhang L, et al. Circular RNA profile indicates circular RNA VRK1 is negatively related with breast cancer stem cells. Oncotarget. (2017) 8:95704–18. doi: 10.18632/oncotarget.21183
95. Lin X, Chen W, Wei F, Xie X. TV-circRGPD6 Nanoparticle Suppresses Breast Cancer Stem Cell-Mediated Metastasis via the miR-26b/YAF2 Axis. Mol therapy: J Am Soc Gene Ther. (2021) 29:244–62. doi: 10.1016/j.ymthe.2020.09.005
96. Li W, Yang X, Shi C, Zhou Z. Hsa_circ_002178 promotes the growth and migration of breast cancer cells and maintains cancer stem-like cell properties through regulating miR-1258/KDM7A axis. Cell transplantation. (2020) 29:963689720960174. doi: 10.1177/0963689720960174
97. Zheng W, Wang X, Yu Y, Ji C, Fang L. CircRNF10-DHX15 interaction suppressed breast cancer progression by antagonizing DHX15-NF-κB p65 positive feedback loop. Cell Mol Biol letters. (2023) 28:34. doi: 10.1186/s11658-023-00448-7
98. Huang X, Tan W, Liu Z, Fu X, Li Z, Lai S, et al. EIF4A3-induced circZFAND6 promotes breast cancer proliferation and metastasis through the miR-647/FASN axis. Life Sci. (2023) 324:121745. doi: 10.1016/j.lfs.2023.121745
99. Zhang J, Xu HD, Xing XJ, Liang ZT, Xia ZH, Zhao Y. CircRNA_069718 promotes cell proliferation and invasion in triple-negative breast cancer by activating Wnt/β-catenin pathway. Eur Rev Med Pharmacol Sci. (2019) 23:5315–22. doi: 10.26355/eurrev_201906_18198
100. Wang ST, Liu LB, Li XM, Wang YF, Xie PJ, Li Q, et al. Circ-ITCH regulates triple-negative breast cancer progression through the Wnt/β-catenin pathway. Neoplasma. (2019) 66:232–9. doi: 10.4149/neo_2018_180710N460
101. Brabletz T, Kalluri R, Nieto MA, Weinberg RA. EMT in cancer. Nat Rev Cancer. (2018) 18:128–34. doi: 10.1038/nrc.2017.118
102. Singh S, Chakrabarti R. Consequences of EMT-driven changes in the immune microenvironment of breast cancer and therapeutic response of cancer cells. J Clin Med. (2019) 8(5):642. doi: 10.3390/jcm8050642
103. Wang Z, Yang L, Wu P, Li X, Tang Y, Ou X, et al. The circROBO1/KLF5/FUS feedback loop regulates the liver metastasis of breast cancer by inhibiting the selective autophagy of afadin. Mol cancer. (2022) 21:29. doi: 10.1186/s12943-022-01498-9
104. Leng X, Huang G, Ding J, Ma F. Circ_0000043 promotes the proliferation, migration, invasiveness, and epithelial-mesenchymal transition in breast cancer cells via the miR-136-Smad3 axis. Biochem Cell Biol = Biochimie biologie cellulaire. (2021) 99:277–85. doi: 10.1139/bcb-2020-0219
105. Song X, Liang Y, Sang Y, Li Y, Zhang H, Chen B, et al. circHMCU promotes proliferation and metastasis of breast cancer by sponging the let-7 family. Mol Ther Nucleic Acids. (2020) 20:518–33. doi: 10.1016/j.omtn.2020.03.014
106. Wu J, Jiang Z, Chen C, Hu Q, Fu Z, Chen J, et al. CircIRAK3 sponges miR-3607 to facilitate breast cancer metastasis. Cancer letters. (2018) 430:179–92. doi: 10.1016/j.canlet.2018.05.033
107. Mao Y, Lv M, Cao W, Liu X, Cui J, Wang Y, et al. Circular RNA 000554 represses epithelial-mesenchymal transition in breast cancer by regulating microRNA-182/ZFP36 axis. FASEB J. (2020) 34:11405–20. doi: 10.1096/fj.201903047R
108. Pan G, Mao A, Liu J, Lu J, Ding J, Liu W. Circular RNA hsa_circ_0061825 (circ-TFF1) contributes to breast cancer progression through targeting miR-326/TFF1 signalling. Cell proliferation. (2020) 53:e12720. doi: 10.1111/cpr.12720
109. Liang G, Liu Z, Tan L, Su AN, Jiang WG, Gong C. HIF1α-associated circDENND4C promotes proliferation of breast cancer cells in hypoxic environment. Anticancer Res. (2017) 37:4337–43. doi: 10.21873/anticanres.11827
110. Ren S, Liu J, Feng Y, Li Z, He L, Li L, et al. Knockdown of circDENND4C inhibits glycolysis, migration and invasion by up-regulating miR-200b/c in breast cancer under hypoxia. J Exp Clin Cancer research: CR. (2019) 38:388. doi: 10.1186/s13046-019-1398-2
111. Li J, Dong X, Kong X, Wang Y, Li Y, Tong Y, et al. Circular RNA hsa_circ_0067842 facilitates tumor metastasis and immune escape in breast cancer through HuR/CMTM6/PD-L1 axis. Biol direct. (2023) 18:48. doi: 10.1186/s13062-023-00397-3
112. Chen B, Wei W, Huang X, Xie X, Kong Y, Dai D, et al. circEPSTI1 as a prognostic marker and mediator of triple-negative breast cancer progression. Theranostics. (2018) 8:4003–15. doi: 10.7150/thno.24106
113. Lu WY. Roles of the circular RNA circ-Foxo3 in breast cancer progression. Cell Cycle (Georgetown Tex). (2017) 16:589–90. doi: 10.1080/15384101.2017.1278935
114. Guan L, Hao Q, Shi F, Gao B, Wang M, Zhou X, et al. Regulation of the tumor immune microenvironment by cancer-derived circular RNAs. Cell Death disease. (2023) 14:132. doi: 10.1038/s41419-023-05647-w
115. Lu C, Shi W, Hu W, Zhao Y, Zhao X, Dong F, et al. Endoplasmic reticulum stress promotes breast cancer cells to release exosomes circ_0001142 and induces M2 polarization of macrophages to regulate tumor progression. Pharmacol Res. (2022) 177:106098. doi: 10.1016/j.phrs.2022.106098
116. Zheng Y, Ren S, Zhang Y, Liu S, Meng L, Liu F, et al. Circular RNA circWWC3 augments breast cancer progression through promoting M2 macrophage polarization and tumor immune escape via regulating the expression and secretion of IL-4. Cancer Cell Int. (2022) 22:264. doi: 10.1186/s12935-022-02686-9
117. Li P, Ren X, Zheng Y, Sun J, Ye G. Tumor promoting effect of circ_002172 associates with induced immune escape in breast cancer via the miR-296-5p/CXCL12 axis. Int immunopharmacol. (2022) 106:108530. doi: 10.1016/j.intimp.2022.108530
118. Zou Y, Zheng S, Deng X, Yang A, Xie X, Tang H, et al. The role of circular RNA CDR1as/ciRS-7 in regulating tumor microenvironment: A pan-cancer analysis. Biomolecules. (2019) 9(9):429. doi: 10.3390/biom9090429
119. Mutebi M, Anderson BO, Duggan C, Adebamowo C, Agarwal G, Ali Z, et al. Breast cancer treatment: A phased approach to implementation. Cancer. (2020) 126 Suppl 10:2365–78. doi: 10.1002/cncr.v126.s10
120. Everaert C, Helsmoortel H, Decock A, Hulstaert E, Van Paemel R, Verniers K, et al. Performance assessment of total RNA sequencing of human biofluids and extracellular vesicles. Sci Rep. (2019) 9:17574. doi: 10.1038/s41598-019-53892-x
121. Yu J, Shen W, Xu J, Gong B, Gao B, Zhu J. circUSP42 is downregulated in triple-negative breast cancer and associated with poor prognosis. Technol Cancer Res Treat. (2020) 19:1533033820950827. doi: 10.1177/1533033820950827
122. Zhao T, Zheng Y, Hao D, Jin X, Luo Q, Guo Y, et al. Blood circRNAs as biomarkers for the diagnosis of community-acquired pneumonia. J Cell Biochem. (2019) 120:16483–94. doi: 10.1002/jcb.v120.10
123. Meng L, Liu S, Liu F, Sang M, Ju Y, Fan X, et al. ZEB1-Mediated Transcriptional Upregulation of circWWC3 Promotes Breast Cancer Progression through Activating Ras Signaling Pathway. Mol Ther Nucleic Acids. (2020) 22:124–37. doi: 10.1016/j.omtn.2020.08.015
124. Lei B, Tian Z, Fan W, Ni B. Circular RNA: a novel biomarker and therapeutic target for human cancers. Int J Med Sci. (2019) 16:292–301. doi: 10.7150/ijms.28047
125. Zou Y, Zheng S, Xiao W, Xie X, Yang A, Gao G, et al. circRAD18 sponges miR-208a/3164 to promote triple-negative breast cancer progression through regulating IGF1 and FGF2 expression. Carcinogenesis. (2019) 40:1469–79. doi: 10.1093/carcin/bgz071
126. Guo XY, Liu TT, Zhu WJ, Liu HT, Zhang GH, Song L, et al. CircKDM4B suppresses breast cancer progression via the miR-675/NEDD4L axis. Oncogene. (2022) 41:1895–906. doi: 10.1038/s41388-022-02232-x
127. Wang X, Xu Y, Wang X, Jiang C, Han S, Dong K, et al. LincRNA-p21 suppresses development of human prostate cancer through inhibition of PKM2. Cell proliferation. (2017) 50(6):e12395. doi: 10.1111/cpr.2017.50.issue-6
128. Fraguas-Sánchez AI, Martín-Sabroso C, Fernández-Carballido A, Torres-Suárez AI. Current status of nanomedicine in the chemotherapy of breast cancer. Cancer chemother Pharmacol. (2019) 84:689–706. doi: 10.1007/s00280-019-03910-6
129. Zheng SR, Huang QD, Zheng ZH, Zhang ZT, Guo GL. circGFRA1 affects the sensitivity of triple-negative breast cancer cells to paclitaxel via the miR-361-5p/TLR4 pathway. J Biochem. (2021) 169:601–11. doi: 10.1093/jb/mvaa148
130. Liu G, Zhang Z, Song Q, Guo Y, Bao P, Shui H. Circ_0006528 contributes to paclitaxel resistance of breast cancer cells by regulating miR-1299/CDK8 axis. OncoTargets Ther. (2020) 13:9497–511. doi: 10.2147/OTT.S252886
131. Liang Y, Song X, Li Y, Su P, Han D, Ma T, et al. circKDM4C suppresses tumor progression and attenuates doxorubicin resistance by regulating miR-548p/PBLD axis in breast cancer. Oncogene. (2019) 38:6850–66. doi: 10.1038/s41388-019-0926-z
132. Dou D, Ren X, Han M, Xu X, Ge X, Gu Y, et al. CircUBE2D2 (hsa_circ_0005728) promotes cell proliferation, metastasis and chemoresistance in triple-negative breast cancer by regulating miR-512-3p/CDCA3 axis. Cancer Cell Int. (2020) 20:454. doi: 10.1186/s12935-020-01547-7
133. Gao D, Zhang X, Liu B, Meng D, Fang K, Guo Z, et al. Screening circular RNA related to chemotherapeutic resistance in breast cancer. Epigenomics. (2017) 9:1175–88. doi: 10.2217/epi-2017-0055
134. Chen D, Wang M, Zhang H, Zhou S, Luo C. Estrogen receptor β2 (ERβ2)-mediated upregulation of hsa_circ_0000732 promotes tumor progression via sponging microRNA-1184 in triple-negative breast cancer (TNBC). Inflammation Res. (2022) 71:255–66. doi: 10.1007/s00011-021-01536-0
135. Wang ZA, Shen MM. Comparative lineage tracing reveals cellular preferences for prostate cancer initiation. Mol Cell Oncol. (2015) 2:e985548. doi: 10.4161/23723556.2014.985548
136. Shao G, Fan X, Zhang P, Liu X, Huang L, Ji S. Circ_0004676 exacerbates triple-negative breast cancer progression through regulation of the miR-377-3p/E2F6/PNO1 axis. Cell Biol toxicol. (2023) 39:2183–205. doi: 10.1007/s10565-022-09704-6
137. Du C, Zhang J, Zhang L, Zhang Y, Wang Y, Li J. Hsa_circRNA_102229 facilitates the progression of triple-negative breast cancer via regulating the miR-152-3p/PFTK1 pathway. J Gene Med. (2021) 23:e3365. doi: 10.1002/jgm.v23.9
138. Li H, Yin H, Yan Y. Circ_0041732 regulates tumor properties of triple-negative breast cancer cells by the miR-149-5p/FGF5 pathway. Int J Biol markers. (2022) 37:178–90. doi: 10.1177/03936155221086599
139. Maimaiti Y, Zhang N, Zhang Y, Zhou J, Song H, Wang S. CircFAM64A enhances cellular processes in triple-negative breast cancer by targeting the miR-149-5p/CDT1 axis. Environ toxicol. (2022) 37:1081–92. doi: 10.1002/tox.23466
140. Magalhães L, Ribeiro-Dos-Santos AM, Cruz RL, Nakamura KDM, Brianese R, Burbano R, et al. Triple-negative breast cancer circRNAome reveals hsa_circ_0072309 as a potential risk biomarker. Cancers. (2022) 14(13):3280. doi: 10.3390/cancers14133280
141. Song X, Chen B, Liang Y, Li Y, Zhang H, Han D, et al. CircEIF3H-IGF2BP2-HuR scaffold complex promotes TNBC progression via stabilizing HSPD1/RBM8A/G3BP1 mRNA. Cell Death discovery. (2022) 8:261. doi: 10.1038/s41420-022-01055-9
142. Barznegar M, Rahimi K, Mahdavi P, Menbari MN, Darvishi N, Vahabzadeh Z, et al. Relation between the circular and linear form of the Elongator Acetyltransferase Complex Subunit 3 in the progression of triple-negative breast cancer. Cell Biochem Funct. (2022) 40:550–8. doi: 10.1002/cbf.v40.6
143. Turco C, Esposito G, Iaiza A, Goeman F, Benedetti A, Gallo E, et al. MALAT1-dependent hsa_circ_0076611 regulates translation rate in triple-negative breast cancer. Commun Biol. (2022) 5:598. doi: 10.1038/s42003-022-03539-x
144. Ruan HG, Gu WC, Xia W, Gong Y, Zhou XL, Chen WY, et al. METTL3 is suppressed by circular RNA circMETTL3/miR-34c-3p signaling and limits the tumor growth and metastasis in triple negative breast cancer. Front Oncol. (2021) 11:778132. doi: 10.3389/fonc.2021.778132
145. Wang X, Xue B, Zhang Y, Guo G, Duan X, Dou D. Up-regulated circBACH2 contributes to cell proliferation, invasion, and migration of triple-negative breast cancer. Cell Death disease. (2021) 12:412. doi: 10.1038/s41419-021-03684-x
146. Li J, Gao X, Zhang Z, Lai Y, Lin X, Lin B, et al. CircCD44 plays oncogenic roles in triple-negative breast cancer by modulating the miR-502-5p/KRAS and IGF2BP2/Myc axes. Mol cancer. (2021) 20:138. doi: 10.1186/s12943-021-01444-1
147. Zan X, Li W, Wang G, Yuan J, Ai Y, Huang J, et al. Circ-CSNK1G1 promotes cell proliferation, migration, invasion and glycolysis metabolism during triple-negative breast cancer progression by modulating the miR-28-5p/LDHA pathway. Reprod Biol endocrinol: RB&E. (2022) 20:138. doi: 10.1186/s12958-022-00998-z
148. Wang X, Song H, Fang L, Wu T. EIF4A3-mediated circPRKCI expression promotes triple-negative breast cancer progression by regulating WBP2 and PI3K/AKT signaling pathway. Cell Death discovery. (2022) 8:92. doi: 10.1038/s41420-022-00892-y
149. Huang Y, Zheng S, Lin Y, Ke L. Circular RNA circ-ERBB2 elevates the warburg effect and facilitates triple-negative breast cancer growth by the microRNA 136-5p/pyruvate dehydrogenase kinase 4 axis. Mol Cell Biol. (2021) 41:e0060920. doi: 10.1128/MCB.00609-20
150. Wang M, Chen D, Zhang H, Luo C. Circular RNA circPTK2 modulates migration and invasion via miR-136/NFIB signaling on triple-negative breast cancer cells in vitro. Inflammation Res. (2022) 71:409–21. doi: 10.1007/s00011-022-01548-4
151. Yuan M, Zhang J, He Y, Yi G, Rong L, Zheng L, et al. Circ_0062558 promotes growth, migration, and glutamine metabolism in triple-negative breast cancer by targeting the miR-876-3p/SLC1A5 axis. Arch gynecology obstetrics. (2022) 306:1643–55. doi: 10.1007/s00404-022-06481-9
152. Zhou Y, Ma G, Peng S, Tuo M, Li Y, Qin X, et al. Circ_0000520 contributes to triple-negative breast cancer progression through mediating the miR-1296/ZFX axis. Thorac cancer. (2021) 12:2427–38. doi: 10.1111/1759-7714.14085
153. Cui S, Zhang Y, Xing L, Li R, Piao Y, Liu H. Circular RNA dehydrodolichyl diphosphate synthase facilitated triple-negative breast cancer progression via miR-362-3p/DDX5 axis. Environ toxicol. (2022) 37:1483–94. doi: 10.1002/tox.23500
154. Fan Y, Wang J, Jin W, Sun Y, Xu Y, Wang Y, et al. CircNR3C2 promotes HRD1-mediated tumor-suppressive effect via sponging miR-513a-3p in triple-negative breast cancer. Mol cancer. (2021) 20:25. doi: 10.1186/s12943-021-01321-x
155. Xing Z, Wang R, Wang X, Liu J, Zhang M, Feng K, et al. CircRNA circ-PDCD11 promotes triple-negative breast cancer progression via enhancing aerobic glycolysis. Cell Death discovery. (2021) 7:218. doi: 10.1038/s41420-021-00604-y
156. Wang Z, Li Y, Yang J, Liang Y, Wang X, Zhang N, et al. Circ-TRIO promotes TNBC progression by regulating the miR-432-5p/CCDC58 axis. Cell Death disease. (2022) 13:776. doi: 10.1038/s41419-022-05216-7
157. Li Y, Wang Z, Su P, Liang Y, Li Z, Zhang H, et al. circ-EIF6 encodes EIF6-224aa to promote TNBC progression via stabilizing MYH9 and activating the Wnt/beta-catenin pathway. Mol Ther. (2022) 30:415–30. doi: 10.1016/j.ymthe.2021.08.026
158. Yang SJ, Wang DD, Zhong SL, Chen WQ, Wang FL, Zhang J, et al. Tumor-derived exosomal circPSMA1 facilitates the tumorigenesis, metastasis, and migration in triple-negative breast cancer (TNBC) through miR-637/Akt1/β-catenin (cyclin D1) axis. Cell Death disease. (2021) 12:420. doi: 10.1038/s41419-021-03680-1
159. Chen T, Wang X, Li C, Zhang H, Liu Y, Han D, et al. CircHIF1A regulated by FUS accelerates triple-negative breast cancer progression by modulating NFIB expression and translocation. Oncogene. (2021) 40:2756–71. doi: 10.1038/s41388-021-01739-z
160. Shi Y, Han T, Liu C. CircRNA hsa_circ_0006220 acts as a tumor suppressor gene by regulating miR-197-5p/CDH19 in triple-negative breast cancer. Ann Trans Med. (2021) 9:1236. doi: 10.21037/atm-21-2934
161. He Q, Hao Q, Wu Y, Vadgama JV, Jiang Y. CircRAD54L2 promotes triple-negative breast cancer progression by regulating the miR-888 family/PDK1 axis. Life Sci. (2023) 312:121128. doi: 10.1016/j.lfs.2022.121128
162. Wang L, Zhou Y, Jiang L, Lu L, Dai T, Li A, et al. CircWAC induces chemotherapeutic resistance in triple-negative breast cancer by targeting miR-142, upregulating WWP1 and activating the PI3K/AKT pathway. Mol cancer. (2021) 20:43. doi: 10.1186/s12943-021-01332-8
163. Wang X, Chen T, Li C, Li W, Zhou X, Li Y, et al. CircRNA-CREIT inhibits stress granule assembly and overcomes doxorubicin resistance in TNBC by destabilizing PKR. J Hematol Oncol. (2022) 15:122. doi: 10.1186/s13045-022-01345-w
164. Tang Q, Zhou F, Yang C, Dai J, Li J, He Y. CircINTS4 facilitates chemoresistance of TNBC by competitively binding miR-129-5p/POM121 axis. J Oncol. (2022) 2022:2630864. doi: 10.1155/2022/2630864
Keywords: breast cancer, triple-negative breast cancer, CircRNAs, biomarkers, immune microenvironment
Citation: Chen P, Zhang J, Wu S, Zhang X, Zhou W, Guan Z and Tang H (2025) CircRNAs: a novel potential strategy to treat breast cancer. Front. Immunol. 16:1563655. doi: 10.3389/fimmu.2025.1563655
Received: 20 January 2025; Accepted: 20 February 2025;
Published: 19 March 2025.
Edited by:
Mohamad S. Hakim, Qassim University, Saudi ArabiaReviewed by:
Xing-xing Fan, Macau University of Science and Technology, Macao SAR, ChinaCopyright © 2025 Chen, Zhang, Wu, Zhang, Zhou, Guan and Tang. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Hailin Tang, dGFuZ2hsQHN5c3VjYy5vcmcuY24=; Ziyun Guan, bHlndWFuenlAc2N1dC5lZHUuY24=
†These authors have contributed equally to this work
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.