
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
REVIEW article
Front. Immunol. , 03 April 2025
Sec. Cytokines and Soluble Mediators in Immunity
Volume 16 - 2025 | https://doi.org/10.3389/fimmu.2025.1549839
This article is part of the Research Topic Maternal-Fetal interface: new insight in placenta research Volume II View all 10 articles
Epigenetic regulation of placental development and pregnancy-related disease processes has recently been a hot research topic. Implantation and subsequent placental development depend on carefully orchestrated interactions between fetal and maternal tissues, involving a delicate balance of immune factors. Epigenetic regulation, which refers to altering gene expression and function without changing the DNA sequence, is an essential regulatory process in cell biology. Several epigenetic modifications are known, such as DNA methylation, histone modifications, non-coding RNA regulation, and RNA methylation. Recently, there has been increasing evidence that epigenetic modifications are critical for the immune microenvironment at the maternal-fetal interface. In this review, we highlight recent advances in the role of epigenetics in the immune microenvironment at the maternal-fetal interface and in epigenetic regulation and placenta-associated pregnancy complications.
Pregnancy is a physiological state in which the maternal immune system smoothly tolerates hemizygous fetal tissues, and several immune mechanisms at the maternal-fetal interface act synergistically to protect the fetus from rejection (1). The percentage and function of immune cells in utero changes dynamically at different stages of pregnancy (2). During normal pregnancy, dynamic interactions between trophoblast cells and decidual immune cells are required to provide a suitable immune microenvironment for successful embryo implantation and normal fetal development (3). The most important immune cells in the human decidua include uterine natural killer (uNK) cells, different T-cell subpopulations, dendritic cells, and macrophages (4). In addition, humoral factors such as sex hormones and cytokines secreted by several immune and non-immune cells play an important role in immune tolerance and pregnancy maintenance (5). An in-depth understanding of the regulatory mechanisms of the immune microenvironment in fetal and maternal tissues will help to understand better the pathophysiological characteristics of pregnancy-related complications such as infertility, miscarriage, and pre-eclampsia (PE), and at the same time, provide a foundation for improving adverse perinatal outcomes (6).
Epigenetics is defined as protein expression changes that occur without alterations in the DNA sequence. It can be stably transmitted during cell proliferation and organismal development and is closely linked to various pathophysiological processes (7). Epigenetics is a cutting-edge discipline that has gradually developed while studying many life phenomena that do not correspond to traditional genetics. The regulatory modes of action of this process include but are not limited to, DNA methylation, histone modification, non-coding RNA (ncRNA) modification, and RNA methylation. These modes of regulation act individually or interact with each other (8, 9).
The pivotal role of epigenetic regulation in developing the maternal-fetal immune microenvironment has garnered significant scientific interest, with many recent studies published. This paper aims to provide a review of the epigenetic mechanisms of the immune microenvironment at the maternal-fetal interface and an overview of recent advances in epigenetic regulation and placenta-associated pregnancy complications.
“Epigenetics” describes altering gene expression and function without altering the DNA sequence, resulting in heritable phenotypes. The concept of epigenetic inheritance was first proposed by Waddington in the journal Endeavour in 1942 (10). The primary focus of genetic research is the inheritance or transmission of genotypes, whereas the process by which genotypes produce phenotypes is the domain of epigenetics. The process of epigenetic inheritance encompasses a range of mechanisms, including DNA methylation, histone modification, the regulation of ncRNA RNAs, and RNA methylation (Figure 1). These mechanisms influence the functions and properties of genes, primarily by regulating their transcription or translation processes (11).
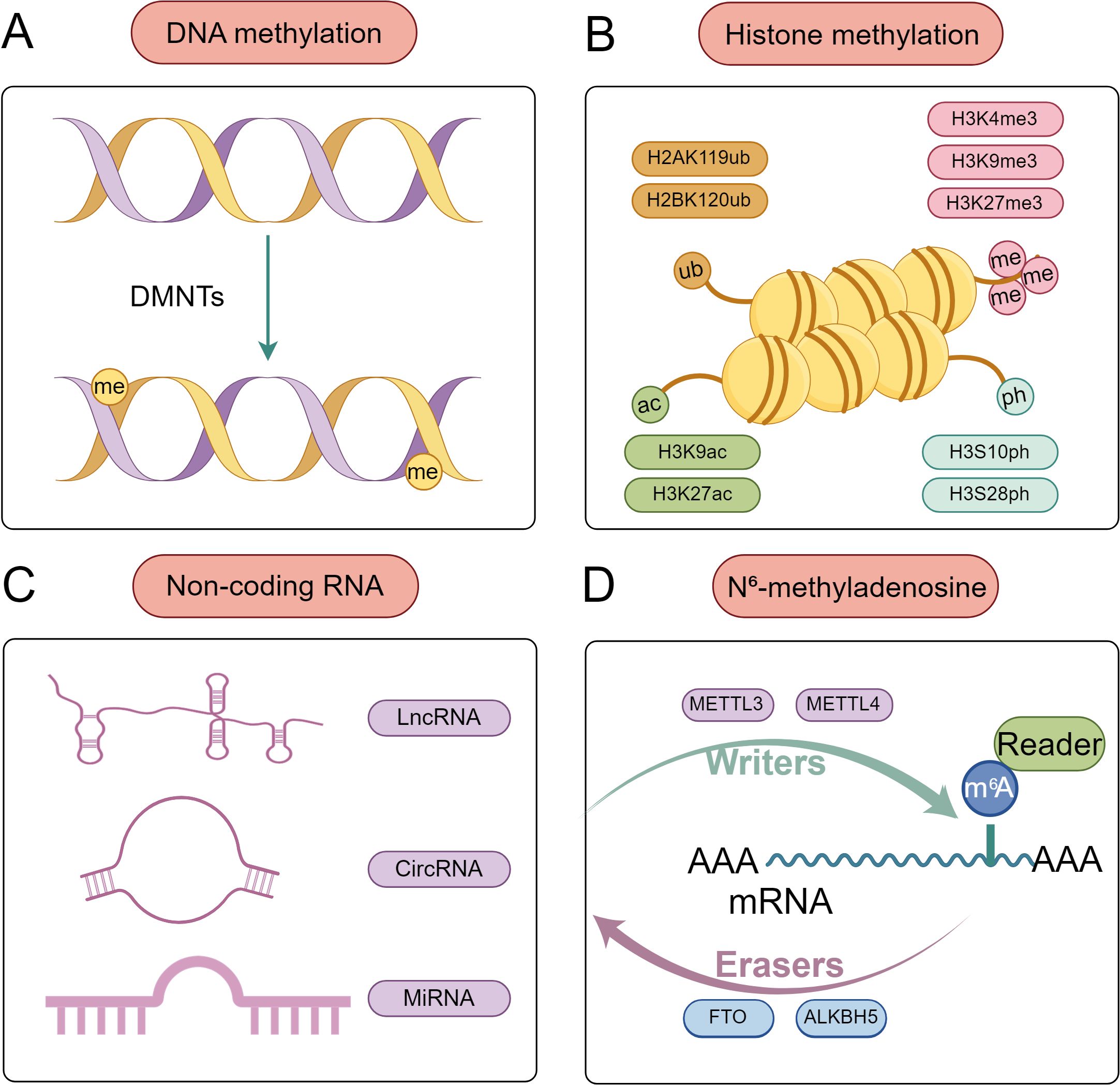
Figure 1. Common epigenetic modifications at the maternal-fetal interface. (A) DNA methylation. DNA methyltransferases (DNMTs) catalyze the covalent addition of a methyl group (-CH3) to the fifth carbon atom of cytosine (C) in DNA molecules, forming 5-methylcytosine (5mC). (B) Histone modifications. These include acetylation, phosphorylation, methylation, and ubiquitination, which play critical regulatory roles at the maternal-fetal interface by modulating chromatin structure and gene transcription activity. (C) Non-coding RNA. MiRNA mainly inhibits gene expression by binding to the 3 ‘untranslated region of mRNA; lncRNA is ncRNA with a length exceeding 200 nucleotides; CircRNAs are highly conserved circular ncRNAs that can act as miRNA sponges, bind to RNA binding proteins, or encode proteins. (D) N6-methyladenosine is the most common form of RNA methylation, which includes three regulatory systems: Writer, Eraser, and Readers. The Writer is responsible for catalyzing the addition of methyl groups to RNA, thereby promoting RNA methylation and ensuring its stability, such as METTL3 and METTL4. Eraser is responsible for mediating the demethylation modification process of RNA, such as FTO and ALKBH5. Readers can recognize and bind to the m6A site in mRNA.
DNA methylation is prevalent in the human genome, and the normal methylation state of DNA is essential for the typical structure and function of the organism’s cells. DNA methylation, as a significant modality in the regulation of epigenetic mechanisms, refers to the process by which the 5th carbon atom on the CpG dinucleotide cytosine of a DNA sequence receives a methyl group from the methyl donor S-adenosyl methionine (SAM) using covalent bonding under the action of the enzyme DNA methyltransferase (DNMT). Methyl donor S-adenosyl methionine (SAM) is used to obtain a methyl group, and the product of this reaction is called 5-methylcytosine (5-mC) (12). DNA methylation is associated with gene silencing, where methylation of CpG sequences in the promoter or non-promoter regions can inhibit promoter binding to transcription factors or be recognized and bound by methyl CpG binding domain (MBD), thereby directly or indirectly inhibiting gene expression (13). DNA methylation occurs mainly at CpG islands in eukaryotes and is mainly regulated by DNMTs and methyl-binding proteins. Five DNMT proteins have been identified in mammals, but only DNMT1, DNMT3A, and DNMT3B have methyltransferase activity (14). DNMT1 can maintain the methylation status of hemimethylated DNA, while DNMT3A and DNMT3B preferentially act on unmethylated and hemimethylated DNA, and the TET protein (ten-eleven translocation, TET) can convert 5-methylcytosine to 5-hydroxymethylcytosine in order to induce DNA demethylation (15). At the maternal-fetal interface, DNA methylation exhibits significant differences in immune cells, which are crucial for maintaining pregnancy immune tolerance and placental development (16–18). However, abnormal DNA methylation regulation may affect embryonic development ability and interfere with the immune microenvironment of embryo attachment and maternal-fetal interface, leading to adverse pregnancy outcomes. Studies have shown that the DNA methylation pattern in human placenta has a reliable and significant correlation with preeclampsia (PE) and gestational diabetes (GDM) (19). Especially in PE patients, the DNA methylation status of non-imprinting genes in the placenta undergoes significant changes, with genes involved in cell adhesion, proliferation, invasion, and other functions showing particularly prominent methylation abnormalities (20).
Histone modification refers to acetylation, phosphorylation, methylation, ubiquitination, and other modifications at histone amino acid sites (21). These modifications affect the compactness and accessibility of chromatin in different ways, affecting gene expression and, consequently, all aspects of biological physiology and developmental processes. At the same time, histone modification involves many modification sites and enzymes. Any abnormal link can lead to the occurrence of diabetes in pregnancy, recurrent abortion, PE, and other diseases.
Acetylation of histones involves introducing acetyl groups to lysine residues in histone tails. This process is carried out by histone acetyltransferase (HAT), which adds the acetyl group to catalyze histone acetylation, and histone deacetylase (HDAC), which removes the acetyl group for deacetylation (22). Under normal conditions, histone acetylation is associated with transcription activation, whereas histone deacetylation is associated with gene silencing (23, 24). The level of acetylation is determined by the balance between HAT and HDAC activity and plays an essential role in chromatin remodeling and transcriptional regulation (25). Specifically, histone deacetylases 8 and 9 (HDAC8, HDAC9) are involved in regulating M1/M2 polarization of macrophages, while HDAC1 and HDAC2 can promote differentiation and fusion of placental trophoblast cells by regulating the development of trophoblast cells (26, 27).
Histone methylation is an important form of histone modification that involves adding one, two, or three methyl groups to certain amino acids in histones, known as mono-, bi-, and trimethylation, respectively. Although methylation can occur at many sites in histones, it mainly occurs at lysine (K) and arginine (R) residues in the tail (28–30). Here, we focus on histone lysine methylation because of its importance and range of vital functions. Histone lysine methylation mainly occurs on histones H3 and H4, and the six loci of H3K4, H3K9, H3K27, H3K36, H3K79, and H4K20 have been studied more (31, 32). Usually, H3K4, H3K36, and H3K39 are involved in transcriptional activation, while H3K9, H3K27, and H4K20 are involved in transcriptional repression or silencing, and thus the biological effects will vary according to different methylation sites and methylation levels (33). At the maternal-fetal interface, the process of sensitization of trophoblast cells is closely related to the enhancement of various histone modifications, including H3K4 trimethylation (H3K4me3), H3K9 acetylation (H3K9ac), and H3K27 acetylation (H3K27ac). The activation of natural killer cells (NK cells) is closely related to H3K4 monomethylation (H3K4me1) and H3K27 acetylation (H3K27ac) (33).
Histone phosphorylation is achieved by adding phosphate groups to amino acid residues in histone tails (34). Several protein kinases catalyze this histone modification, are closely related to the cell cycle, and can affect the transcriptional activation of DNA by mediating the recruitment of DNA damage repair proteins (35). Almost all histones can be phosphorylated at specific sites, and together with other post-translational modifications, they regulate a variety of biological processes (36).
The human genome is highly active at the transcriptional level, but only about 1.9% of the gene sequence is transcribed into proteins, while the rest is transcribed into ncRNAs (37). ncRNAs mainly consist of microRNAs (miRNAs), long-stranded non-coding RNAs (LncRNAs), and cyclic RNAs (circRNAs) (38). Among them, miRNAs are between 19 and 23 nucleotide units in length and mainly play a role in silencing gene expression by antisense inhibition of the 3’untranslated regions (3’UTR) of mRNAs (39); LncRNAs are a type of ncRNAs with a length of >200 nucleotide units (40); circRNAs are a type of highly conserved ncRNAs that form a closed-loop structure with covalent bonds, and their mechanism of action mainly includes acting as miRNA sponges, binding to RNA-binding proteins to regulate transcription, and encoding proteins (41). Supplementary Table 1 summarizes the association of Maternal-fetal interface immune cells with ncRNAs.
RNA methylation is the most common type of RNA modification, including N6 methyladenosine (m6A), 5-methylcytosine (m5C), N7 methylguanine (m7G), and N1 methyladenosine (m1A) (42). Among them, m6A is the most abundant RNA modification, mainly located in the 3 ‘untranslated region (3’ UTR) near the mRNA protein coding sequence and stops codons, regulating gene expression and biological functions by controlling RNA metabolism, alternative splicing, degradation, and translation processes (43, 44). The m6A methylation modification regulatory system mainly includes three categories: Writer, Eraser, and Readers. The writer mainly promotes RNA methylation and maintains its stability, with the most common molecules being METTL3 and METTL4. Eraser is responsible for mediating the demethylation modification process of RNA, such as molecular FTO and ALKBH5. Readers can recognize and bind to the m6A site in mRNA, shortening the half-life of mRNA. As an important post-transcriptional regulatory mechanism, RNA methylation can significantly affect the stability, intracellular localization, transport process, splicing mode, and translation efficiency of RNA (45). Research has shown that this epigenetic modification exhibits abnormal expression patterns in various reproductive system diseases, including pathological states such as PE, spontaneous abortion, endometriosis, and premature ovarian failure. The root cause of many pregnancy complications lies in poor implantation. Zhan Hong Zheng et al. (46) found that mice with Mettl3 deficiency were completely infertile due to implantation and decidualization failure, revealing that the m6A modification in the 5 ‘- UTR of Pgr mRNA mediated by METTL3 is closely related to the signal transduction of normal progesterone. In addition, the elevated level of RNA demethylase ALKBH5 in placental villus tissue of patients with recurrent spontaneous abortion (RSA) may be due to its ability to shorten the half-life of CYR61 mRNA in an m6A dependent manner and reduce the level of total m6A modification, affecting the invasiveness of the trophoblast and leading to miscarriage (47).
The maternal-fetal interface consists mainly of decidua from the mother and trophoblast cells from the embryo. The interactions between decidual immune cells and trophoblast cells form a vast network of cellular connections. The decidual immune system is constituted by different subpopulations of maternal immune cells, including decidual natural killer (dNK) cells, macrophages, and T cells, among others. Modern reproductive immunology suggests that imbalances in the immune microenvironment at the maternal-fetal interface may be involved in the development of pregnancy-related diseases. Increasing evidence suggests that epigenetic modifications are critical to the immune microenvironment at the maternal-fetal interface, and studies on the correlation between the dynamic function and compositional changes of vital immune cells (NK cells, macrophages, and T cells) and trophoblasts at the maternal-fetal interface and epigenetics are described in detail below (Figure 2).
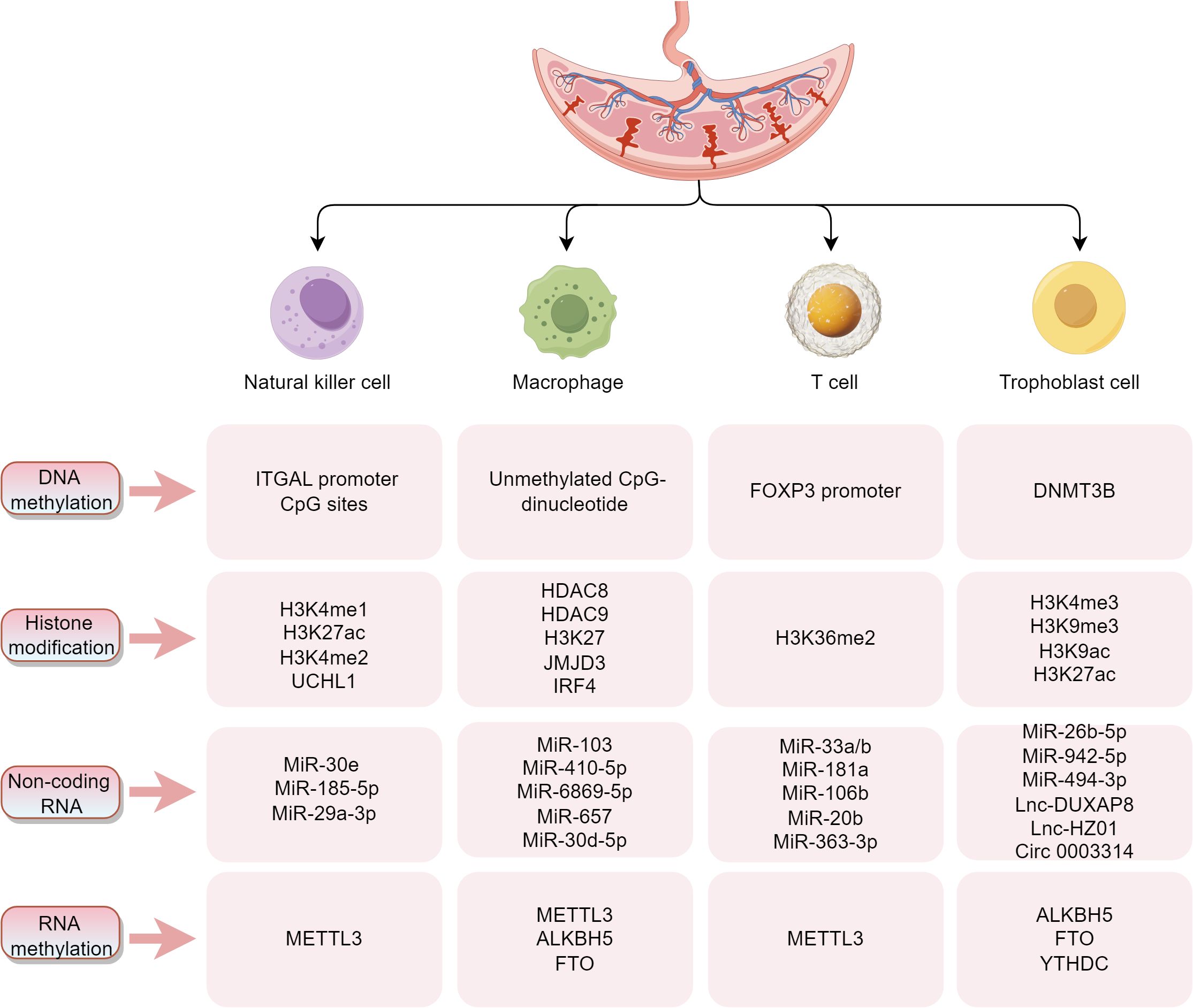
Figure 2. Summarizes the key molecules and modification sites associated with epigenetic regulation in four critical cell types (NK cells, macrophages, T cells, and trophoblasts) at the maternal-fetal interface. These epigenetic modifications, encompassing DNA methylation, histone modifications, non-coding RNAs (ncRNAs), and RNA methylation, play pivotal roles in regulating immune tolerance at the maternal-fetal interface and supporting placental development.
NK cells are intrinsic immune cells in the human body and are found in peripheral blood and endometrium (48). Among them, peripheral blood (pb) NK cells have two main subpopulations (CD56bright CD16dim and CD56dim CD16bright), and up to 90% of pNK cells are predominantly CD56dim CD16bright, and the primary function of pNK cells is highly cytotoxic (49). Compared to pbNK cells, dNK cells have a different phenotype; they resemble the CD56brightCD16dimpbNK subpopulation but have their unique characteristics, are mainly responsible for the production of cytokines and have poor cytotoxicity (50). Vento-Tormo et al. (51) have identified three significant subpopulations of dNK cells. In early pregnancy, NK cells are present in large numbers at the maternal-fetal interface, accounting for 70% to 80% of the decidual lymphocytes, whereas in the human decidua, 10% to 15% of the cells are decidual lymphocytes (52). Dnk cells are multifunctional, secreting various growth factors, angiogenic factors, chemokines, and cytokines. These not only synergistically promote the metamorphosis process of the endometrium and the remodeling of the spiral arteries of the uterus, thus laying a solid foundation for the growth of placental blood vessels and the construction of the placenta, but also exhibit a unique immune-regulatory mechanism. NK cells guide the migration of trophoblasts toward the fetal membranes and spiral arteries while preventing trophoblasts from being destroyed, thus regulating trophoblast invasion into the uterine wall and ensuring that the whole process is safe and smooth (52, 53).
Decidual NK cells promote angiogenesis and trophoblast invasion and are closely associated with placental development (54). Epigenetic mechanisms such as DNA demethylation and histone acetylation have been shown to dynamically regulate gene transcriptional activity in immune cells, including CD8 (+) T cells and NK cells (18). However, research on the DNA methylation regulatory network in NK cells at the maternal-fetal interface is still limited. It is worth noting that the DNA methylation status is closely related to the activation level of NK cells: activated NK cells exhibit low methylation characteristics of CpG sites, while primitive NK cells maintain relatively high methylation levels, indicating that the methylation status of NK cells has significant plasticity and reproductivity (55). Hu et al. (17) found that DNA methylation in chorionic extravillous trophoblast (EVT) cells is regulated by dNK and the soluble molecules it secretes, which in turn affects EVT differentiation, adhesion, and migration. This discovery extends the function of dNK from classical immune regulation to epigenetic-mediated regulation of trophoblast development. In addition, Ana Sofia Cerdeira et al. (56) successfully induced dNK-like cell generation through pharmacological demethylation intervention, providing evidence for the core role of epigenetic mechanisms in dNK development. In summary, DNA methylation is not only a key epigenetic switch for the dynamic regulation of NK cell function but also affects the immune microenvironment and placental development at the maternal-fetal interface through dNK-mediated methylation remodeling. Future research needs to reveal the specific regulatory network of DNA methylation in dNK cells and its impact on pregnancy outcomes.
Histone modifications are essential to promote normal NK cell development, and the histone H2A deubiquitinase MYSM1 regulates NK cells by controlling transcription factors, the lack of which severely impairs NK cell development (57). Giuseppe Sciumè et al. (58) found that activation of NK cells induced enhancer regions to H3K4me1 and H3K27ac histone modifications. Wiedemann et al. (59) used a multi-omics approach to analyze the complex interactions between cytokine signaling pathways in NK cells and found that IFN - α can enhance the epigenetic modification of the H3K4me3 promoter site. Ma et al. (60) demonstrated that SPA-VSMC has the potential to transform into CD56 dNK by identifying a small portion of dNK modified with H3K4me2 in the myosin heavy chain 11 (MYH11) promoter region through immunofluorescence-DNA in situ hybridization-neighbor-joining and chromatin immunoprecipitation experiments, which is the first demonstration that SPA-VSMC has the potential to transform into CD56 dNK. Zhang et al. (61) showed that ubiquitin C-terminal hydrolase L1 (UCHL1) can regulate decidualization through the JAK2/STAT3 signaling pathway. When UCHL1 is deficient, it can cause damage to decidualization during pregnancy in mice, leading to miscarriage, and is associated with a decrease in the number of decidual natural killer cells (dNKs). Through the above research, we can gain a deeper understanding of the epigenetic regulatory mechanisms of NK cells, as well as the pathogenesis of related diseases such as miscarriage.
In recent years, multiple studies have revealed the important role of NK cells in pregnancy and their epigenetic regulatory mechanisms. Gamliel et al. (62) discovered a unique phenotype and transcriptional features of NKG2C high NK cell population in the decidua of women with multiple pregnancies. These cells showed increased accessibility at the Ifng and Vegfa loci and exhibited more vigorous IFN - γ and VEGF-A secretion. It provides important clues for understanding how microenvironmental changes during pregnancy induce epigenetic reprogramming of NK cells. In terms of miRNA regulation of NK cells, research (63) found through comparative analysis that 36 miRNAs were expressed explicitly in decidual NK cells. In comparison, two miRNAs were only expressed in peripheral blood NK cells, revealing molecular differences in NK cells from different sources. Another study (64) through miRNA profiling analysis of decidual NK cells from patients with unexplained recurrent spontaneous abortion (URSA), 50 differentially expressed miRNAs were identified, of which 49 were upregulated, and 1 was downregulated, indicating a close association between abnormal miRNA expression and URSA. Specifically, researchers have found through in vitro experiments that upregulated miR-30e can reduce the cytotoxicity of peripheral blood and decidual NK cells by targeting PRF1, inhibit Th1 tolerance phenotype, and induce Th2 immune dominance, thereby promoting the formation of a micro immune tolerance environment at the maternal-fetal interface (65). In addition, miR-185-5p is involved in the occurrence of RSA by interfering with VEGF expression and angiogenesis in decidual NK cells (66). Fang et al. (67) found that a decrease in miR-29a-3p levels in villous-derived exosomes of patients with URSA can increase IFN - γ levels in decidual NK cells. In vivo, experiments in mice revealed that vEXOs carrying miR-29a-3p can reduce embryonic resorption in RPL mice. However, despite significant progress in these studies, the epigenetic regulatory mechanisms of decidual NK cells have not been fully elucidated. Further in-depth research is needed regarding the proportion, functional status, and relationship with epigenetic regulatory mechanisms of decidual NK cells.
In METTL3 deficient NK cells, the absence of m6A modification leads to a decrease in SHP-2 protein expression, inhibiting the activation of AKT and MAPK signaling pathways, and significantly reducing NK cell responsiveness to IL-15. In addition, the absence of METTL3 disrupts the homeostasis of NK cells. It inhibits their infiltration and function in the tumor microenvironment, ultimately accelerating the development of mouse tumors and shortening their survival period (68). Although the function of RNA methylation (especially m6A modification) in immune cells is gradually being revealed, research on its role in natural killer cells (NK cells) at the maternal-fetal interface is still relatively limited. Mother-fetal interface NK cells (NK cells) are crucial in early pregnancy embryo implantation, placental formation, and maternal-fetal immune tolerance. However, their functional regulatory mechanisms still need further exploration.
Decidual macrophages are the second largest immune cell population (10%-20%) in the uterine decidua in early human pregnancy, second only to the percentage of dNK cells (60%-80%) (69). Notably, macrophages can be categorized into classically activated (M1) and alternatively activated (M2) phenotypes based on their function and the types of cytokines they produce (70). M1 macrophages are pro-inflammatory and are characterized by the production of higher levels of IL-12 and IL-23, as well as lower levels of IL-10, thus killing intracellular microbes and inducing Th1 immunity. In contrast, M2 macrophages produce higher levels of IL-10 and lower levels of IL-12 and IL-23 and perform functions such as anti-inflammatory and tissue remodeling, as well as the removal of apoptotic cells and debris (71). At the maternal-fetal interface, the number and ratio of M1/M2 macrophages change during different gestation periods. Specifically, M1-type macrophages predominate during the precomputation period, followed by a transition to a mixed population of M1 and M2 types; after the establishment of the placenta, M2-type macrophages dominate (72). Evidence suggests that an imbalance of M1/M2 macrophages leads to a pro-inflammatory microenvironment in the endometrium, which is not conducive to inducing fetal tolerance and further leads to pregnancy-related disorders such as recurrent miscarriages (73, 74). M1 and M2 macrophages play different roles at different gestational stages; thus, the maintenance of a normal pregnancy requires that the M1/M2 macrophage is in a state of equilibrium.
A study (16) conducted genome-wide methylation analysis on maternal monocytes, fetal monocytes, decidual macrophages, and fetal placental macrophages (Hofbauer cells) using Illumina Infinium Human Methylation 27 BeadChip technology and found significant differences in DNA methylation among these cell populations. Notably, genes related to immune response are highly methylated in fetal cells, while maternal cells exhibit different methylation patterns. Meanwhile, excessive pro-inflammatory reactions at the maternal-fetal interface may jeopardize pregnancy maintenance. At the same time, DNA methylation may play a protective role in this process by regulating the anti-inflammatory function of macrophages. In premature birth studies, genome-wide placental DNA methylation analysis combined with methylated DNA immunoprecipitation sequencing (MeDIP-seq) technology revealed the gene functional characteristics of differentially methylated regions (DMRs), with a significant enrichment of Fc - γ receptor-mediated macrophage phagocytosis related pathways (75). This discovery suggests that DNA methylation may be involved in the mechanism of premature birth by regulating the phagocytic function of macrophages. In addition, unmethylated CpG dinucleotides derived from microorganisms can activate Toll-like receptor 9 (TLR-9), and these DNA activators exert phagocytic and clearance functions through the production of intrauterine immune cells (such as macrophages), thereby affecting the immune microenvironment at the maternal-fetal interface (76). These studies collectively reveal the important role of DNA methylation in regulating macrophage function at the maternal-fetal interface and maintaining pregnancy, providing a new perspective for a deeper understanding of the epigenetic mechanisms of pregnancy-related diseases.
It was found that the expression of HDAC8 was reduced in decidual macrophages from patients with RSA, and the knockdown of HDAC8 inhibited M2 macrophage activation and promoted apoptosis of differentiated THP-1 (dTHP-1) macrophages through ERK pathway (26). Meng et al. (71) reported that nuclear factor-κ B ligand (RANKL) receptor activator, secreted by human embryonic trophoblasts and maternal decidual stromal cells, polarizes decidual macrophages to an M2 phenotype, which is mediated by activation of Akt/signal transducer and activator of transcription factor 6 (STAT6) signaling, and correlates with the up-regulation of the histone H3 lysine 27 demethylases, Jmjd3 and IRF4, in decidual macrophages. Liu et al. (77) found that Hdac9/HDAC9 deficiency promotes macrophage polarization toward M2 macrophages, while Hdac9/HDAC9 ablation significantly enhanced phagocytosis of fluorescent microspheres in M2 Raw 264.7 cells, but reduced the capacity of THP-1-derived M1 macrophages. These studies collectively reveal the key role of histone modifying enzymes in macrophage polarization and functional regulation.
NcRNA can regulate macrophage function and M1/M2 polarization, closely related to the occurrence and development of various pregnancy-related diseases. Some in vitro experimental studies have shown that specific miRNAs affect macrophage polarization by regulating key signaling pathways. For example, miR-103 and miR-410-5p inhibit the STAT1-mediated signaling pathway, preventing polarization of M1 macrophages and preventing RSA (78, 79). MiR-6869-5p induces M2 polarization in gestational diabetes by regulating PTPRO (80). On the contrary, miR-657 promotes gestational glucose by targeting FAM46C (81). In addition, miRNA-30d-5p derived from placental exosomes induces M2 polarization by inhibiting HDAC9 expression (82). The low-level miR-455-3p in decidual stromal cells inhibits the invasion of trophoblast cells by promoting macrophage polarization (83). MiR-146a-5p has been shown to increase embryo resorption rate and promote M2 polarization of macrophages in URSA (84). The article also briefly introduces the relationship between some miRNAs and macrophages (85).
LncRNAs also participate in the regulatory network of macrophage polarization. Wu et al. (86) found that the expression of AOC4P was significantly up-regulated in trophoblast cells of patients with RSA. This molecule inhibits the degradation of EZH2 by regulating TRAF6, thus inhibiting glycolysis in trophoblast cells and actively participating in the polarization process of M2 macrophages. Xiujun Li et al. (87) confirmed through in vitro experiments that lncRNA MALAT1 not only regulates the proliferation and angiogenesis of mesenchymal stem cells (MSCs) but also promotes M2 polarization by regulating the expression of indoleamine 2,3-dioxygenase (IDO). LINC00240 could promote macrophage polarization toward the M2 type by regulating the miR-155/Nrf2 axis (88). The knockdown of LINC00221 negatively regulated the expression of miR-542-3p in trophoblasts to reduce macrophage migration and invasion (89). These studies provide a new perspective for elucidating the regulatory role of ncRNA in macrophage polarization. However, most current research is limited to in vitro experiments. The mechanism of action of lncRNA is highly complex, and its function may vary significantly depending on cell type and pathological state. Therefore, in the future, more in vivo experiments are needed to verify its biological functions and further explore its regulatory mechanisms and therapeutic potential in diseases by combining clinical samples.
METTL3 participates in macrophage regulation through different mechanisms as a “writer” in RNA methylation. Low expression of METTL3 can inhibit the degradation of NOD1 and RIPK2 mRNA mediated by YTHDF1 and YTHDF2, thereby upregulating the NOD1 pathway and subsequently promoting LPS-induced macrophage inflammatory response (90). Another study showed that METTL3 could promote oxLDL-induced macrophage inflammatory response by activating STAT1 signaling (91). Zhao et al. (92) co-cultured ALKBH5 overexpressing cell lines with THP-1 and found that the N6 methyladenosine regulatory factor ALKBH5 damaged macrophage recruitment and M2 differentiation by reducing VEGF secretion in stromal cells, leading to miscarriage. Meanwhile, inhibiting VEGF signaling can help discover changes in macrophage polarization in different pregnancy complications (93). In addition, Cox analysis found that M2 macrophages were positively correlated with the m6A regulatory factor FTO and negatively correlated with CBLL1 (94).
In early pregnancy, 30% to 45% of the decidual T cells are CD4+ T cells at the maternal-fetal interface, and 45% to 75% are CD8+ T cells (95). Under normal conditions, a dynamic balance is maintained between CD4+ and CD8+ T lymphocytes (96). CD4+ T cells are mainly helper T cells (Th) and regulatory T cells (Treg) (97). CD4+ helper T cells (Th) are classified into Th1, Th2, and Th17 cells according to the type of cytokines they secrete (98). Th1 cytokines promote macrophage activation and cytotoxicity and mainly mediate cellular immune responses (99). Th2 cytokines mediate humoral immune responses, promote eosinophil and mast cell differentiation, inhibit immune inflammation, reduce excessive damage, and have immunotoxic effects (100). After conception, maternal recognition of the fetus and maintenance of pregnancy is achieved through the immune balance between Th17/Treg, Th1/Th2 cells, and the dominant Th2-type cells (101). The large amount of Th2-type cytokines at the maternal-fetal interface inhibits the production of Th1-type cytokines through a negative feedback effect, suppresses TDTH and CTL cell activity, and thus inhibits rejection (102). Some subpopulations of maternal-fetal interface T cells help EVT invade the endometrium and promote embryo implantation and placenta formation, whereas other subpopulations are closely associated with pregnancy complications (103). The classical pattern of immune regulation in human pregnancy is a shift in maternal immune response from an inflammatory Th1 cytokine pattern to a Th2 pattern, with maternal-fetal interface decidual T cells playing a crucial role in regulating the placental microenvironment and recognizing fetal antigens (104).
Research has shown that the methylation levels of Th1/Th2 pathway genes exhibit dynamic changes at different stages of pregnancy. During normal pregnancy, the methylation levels of Th2 pathway genes change more significantly in the early stages of pregnancy. In contrast, the methylation levels of Th1 pathway CpG genes change more frequently in the late stages of pregnancy, suggesting that the temporal regulation of Th1/Th2 gene methylation is closely related to the maintenance of pregnancy (105). Abnormal DNA methylation can participate in the occurrence of pregnancy-related diseases through multiple pathways. It can disrupt the immune balance at the maternal-fetal interface by regulating the proliferation, differentiation, and cellular activity of Th and Treg cells, thereby promoting pathological processes such as recurrent RSA (106). Comparative analysis reveals that RSA patients exhibit abnormal FOXP3 promoter hypermethylation in decidual tissues, resulting in suppressed FOXP3 expression and subsequent disruption of maternal-fetal immune tolerance, potentially representing a key molecular pathway in RSA (107).
Histone methyltransferase Nsd2 was found to upregulate CXCR4 expression through H3K36me2 modification, promoting Treg cell recruitment into the decidua to ensure maternal-fetal immune tolerance (108).
Schjenken et al. (109) found that miR-155 is required to expand regulatory T cells to mediate robust pregnancy tolerance in mice. Notably, the expression of miR-33a/b and miR-181a was significantly down-regulated in patients with RSA and led to a reduction in the number of Treg cells by negatively regulating the expression level of the vital molecule sphingosine-1-phosphate receptor-1 (S1PR1) (110). In patients with PE, miR-106b inhibits Treg differentiation by suppressing TNF - β expression, while miR-20b and miR-363-3p upregulate TH17 cell transcription factor ROR γ t/STAT3 and enhance TH17 activity, respectively, leading to Treg/TH17 imbalance (111). It is worth noting that environmental factors can also interfere with ncRNA networks. For example, Jamie L. McCall et al. (112) confirmed that prenatal cadmium exposure activates CD4+T cells in offspring by upregulating lncRNA Snhg7, suggesting that exogenous toxic substances may participate in the pathological process of pregnancy through epigenetic mechanisms.
METTL3 deficient CD4 (+) T cells disrupt the homeostasis and differentiation of immature T cells (113). By hybridizing Mettl3f/f mice with Foxp3Cre YFP mice and specifically knocking out Mettl3 in Tregs, the results showed that the offspring mice’s peripheral lymph nodes and spleen significantly increased and developed severe autoimmune diseases and infertility. Mechanism studies have shown that the absence of Mettl3/m6A leads to an increase in Socs mRNA levels, inhibiting the IL-2-STAT5 signaling pathway, which is crucial for the function and stability of Treg cells (114). In addition, METTL3-dependent m6A methylation plays a critical role in regulating follicular helper T cell (TFH) lineage differentiation. Research has found that the absence of METTL3 in CD4+T cells impairs the function of TFH cells, leading to a significant reduction in germinal center (GC) response in METTL3 deficient mice after acute viral infection (115).
EVT are one of the main coordinators of immunity at the maternal-fetal interface (116), actively creating an environment for tolerogenic phenotypes by expressing a unique set of major histocompatibility complex (MHC) molecules (117).
DNA methylation in trophoblast cells continues to change dynamically after implantation and throughout pregnancy, with the differentiation of cytotrophoblasts into syngeneic trophoblasts and the acquisition of an invasive phenotype in EVT, both of which involve extensive DNA methylation changes (118). The study (119) identified ZBTB24 as an epigenetic regulator that modulates E-cadherin expression and, thus, cell viability, differentiation, and migration in trophoblasts by altering DNA methylation in the promoter region. In addition, dNK can regulate trophoblast function by altering gene expression through DNA methylation (120). In summary, these findings reveal the core regulatory role of DNA methylation in nourishing cells and pregnancy maintenance at the maternal-fetal interface.
During the sensitization process of BeWo trophoblast cells, histone markers associated with active transcription (H3K4me3, H3K9ac, and H3K27ac) significantly increased, while the levels of inhibitory histone modifications (H3K9me3 and H3K27me3) remained unchanged (121). This process is consistent with increased mRNA levels of EP300 and P300/CBP-related factors (PCAF). The binding of RNA polymerase II to specific gene promoters is enhanced during the process of trophoblast differentiation, which is closely related to active histone markers. Meanwhile, the downregulation of pro-inflammatory transcription factors after differentiation of multinucleated syncytiotrophoblasts is associated with reduced enrichment of H3K27Ac and H3K9Ac promoters, as well as enhanced binding of H3K9me3 to histone deacetylase 1 (122). Histone deacetylases 1 and 2 regulate the development of extrachorionic trophoblast cells and promote the differentiation and fusion of human placental trophoblast cells (27). In addition, glycolytic metabolism is involved in regulating the function of trophoblast cells. Yu et al. (123) found that glycolysis exhibited high activity in human trophoblast stem cells and cytotrophoblast cells, whereas it was significantly reduced in syncytiotrophoblast cells. However, when supplemented with the glycolytic derivative acetyl coenzyme A, the fusion function of the syncytiotrophoblast was restored. Of particular note, acetylation of H3K9, H3K18, H3K27, and H4K16 was particularly sensitive to glycolysis during syncytiotrophoblast fusion with the metabolism of acetyl-coenzyme A. These modifications regulated the promoters of relevant genes involved in syncytiotrophoblast fusion and metabolism of syncytiotrophoblast cells. In terms of pathological mechanisms, an in vitro experiment found that knocking down ACLY disrupts histone acetylation and IL-10 secretion in nourishing cells, thereby inhibiting M2 polarization of macrophages. This process is involved in the pathogenesis of RSA (124). Sumi et al. (125) found that LPS treatment increased SIRT1 expression in mouse placental tissue and human trophoblast cells and inhibited NLRP3 inflammasome activation in trophoblast cells by reducing oxidative stress, thereby reducing lipopolysaccharide-induced IL-1 β levels. These studies collectively reveal the important role of histone modifications in the differentiation, metabolic regulation, and pathological processes of trophoblast cells.
MiRNAs are equally important in regulating trophoblast function, and their abnormal expression is closely related to pregnancy-related diseases. Liang et al. (126) systematically summarized miRNA regulation on trophoblast function and its related signaling pathways, providing a theoretical basis for understanding the mechanisms of pregnancy diseases. Meanwhile, lncRNA also participates in trophoblast dysfunction through various mechanisms. The long noncoding RNA DUXAP8 is significantly upregulated in the placental tissue of PE and can inhibit biological functions such as proliferation, migration, and invasion of trophoblast cells. The research team constructed a pregnant rat PE model to further validate its molecular mechanism by mediating DUXAP8 overexpression through adenovirus vectors. The experimental results showed that DUXAP8 activates the AKT/mTOR signaling pathway, inhibits endoplasmic reticulum autophagy, and promotes the occurrence and development of PE (127). In vitro experiments, Inc-HZ01 with m6A RNA methylation modification forms a positive feedback loop with MXD1, which plays a vital role in regulating abortive and BPDE-exposed trophoblast cell function regulation. This loop increases Inc-HZ01 levels by enhancing the RNA stability of MXD1, which inhibits trophoblast proliferation and induces miscarriage (128). Circ_0003314 inhibits trophoblast function and induces cell apoptosis through the miR-26b-5p/IL1RAP signaling axis (129). In the PE study, circ_0015382 impaired trophoblast function through the miR-942-5p/NDRG1 axis, and its expression level was significantly elevated in the placental tissue of PE patients (130). In addition, Ou et al. (131) found through Pearson correlation analysis of placental tissue from 25 PE patients that the expression level of circ_0111277 was significantly higher than that of normal pregnancy placenta. Mechanism studies have shown that circ_0111277 participates in the occurrence of PE by regulating the miR-494-3p/HTRA1/Notch-1 signaling pathway, inhibiting the invasion and migration ability of trophoblast cells. These research results indicate that ncRNA participates in the precise regulation of trophoblast function through a complex regulatory network, providing potential molecular targets for diagnosing and treating pregnancy-related diseases. However, the exact molecular pathways of these regulatory mechanisms still need to be elucidated through more in-depth in vivo experimental studies, including the construction of conditional gene knockout animal models.
In vitro experiments showed that knocking down m6A demethylase ALKBH5 can promote the invasion of HTR-8/SVneo in trophoblast cells by regulating the stability of CYR61 mRNA, while overexpression of ALKBH5 has the opposite effect (132). Meanwhile, Chuanmei Qin et al. (133) found that the expression of ALKBH5 in the villus tissue of RSA patients was elevated compared with the control group and confirmed through in vitro experiments that ALKBH5-overexpression-inhibited RSL3-induced cell death in the trophoblast by promoting the expression of iron death-related gene FTL. In addition, the expression of m6A demethylase FTO protein increased in URSA trophoblast cells, inhibiting m6A modification of MEG3 and weakening the stabilizing effect of YTHDC1 protein on MEG3. MEG3 inhibits the expression of the TGF - β 1 gene by binding to the EZH2 protein and TGF - β 1 gene promoter, thereby affecting the invasion and proliferation of trophoblast cells (134). These findings reveal the key role of RNA methylation in regulating the function of trophoblast cells and its potential mechanisms in miscarriage.
As mentioned above, epigenetic regulation plays a moderately important role in the immune microenvironment at the maternal-fetal interface, and their absence or dysfunction may lead to complications during pregnancy (135). It has been suggested that normal and abnormal pregnancies lead to changes in cell populations, which may be facilitated by epigenetic modifications induced by different pregnancy-associated processes (136). The placenta, a uniquely evolved organ that develops externally to the embryo, undergoes rapid growth and fulfills diverse roles throughout pregnancy, primarily ensuring a stable and protective milieu for fetal development. Notably, in recent years, shifts in lifestyle have coincided with a steady increase in the incidence of placental-related pregnancy disorders, exerting profound consequences on both maternal and neonatal health (137). Table 1 summarizes the association between epigenetic regulation and placental related pregnancy complications.
Abnormal changes in DNA methylation are closely related to maternal health and fetal development. There are extensive DNA methylation changes in early-onset PE, among which TGFBR1 induces disease occurrence through a DNMT3A downregulation mediated DNA methylation-independent pathway (138). Meanwhile, the low methylation status of placental growth factor and Fms-related tyrosine kinase-1 further affects placental function (139). In terms of fetal development, changes in CpG methylation associated with maternal PE, pre-pregnancy weight loss, and type 2 diabetes risk are significantly associated with low birth weight, suggesting that placental DNA methylation may serve as a bridge between maternal metabolic status and the risk of chronic disease in offspring (140). In addition, epigenetic changes are also associated with other pregnancy-related diseases. Researchers recruited three decidua samples from patients with RSA and normal controls. They systematically identified key genes regulated by DNA methylation in the decidua and blood of RSA patients through genome-wide bisulfite sequencing (GWBS) and transcriptome sequencing. They found that 23 genes exhibited significant methylation and expression differences between RSA patients and healthy controls, with hypomethylated differential methylation regions (DMR) and upregulated differential gene expression (DGE) co-enriched in the Rap1, GnRH, and Estrogen signaling pathways. Additionally, 32 genes from these three pathways showed significant differences in DMR between RSA patients and the control group. Hi-MethylSeq analysis further revealed that SGK1 in RSA patients’ blood and decidua samples exhibited a high methylation state. At the same time, SGK3 and CREB5 also showed significant changes in methylation levels in the decidua (141). T lymphocyte CD3 methylation may mediate psychiatric symptoms such as depression and anxiety during pregnancy (142). The above research reveals the extensive role of epigenetic modifications in pregnancy-related diseases, providing a new perspective for the mechanism research of related diseases.
HDAC 2 expression and activity are down-regulated in monocytes/macrophages of patients with gestational diabetes, suggesting the role of histone deacetylation in metabolic disorders (143). The difference in placental chromatin activity is associated with fetal growth restriction, and H3K27 acetylation may be involved in regulating placental function (144). Men1, a member of the H3K4 methyltransferase complex, is crucial for endometrial decidual transformation. Research has shown that abnormal differentiation of uterine stromal cells in mice with Men1 deficiency can lead to embryo resorption and pregnancy failure (145). Intriguingly, novel histone acetylation modifications, including H3K18la, have been identified as critical regulators of embryo implantation, with reduced levels potentially contributing to pregnancy loss. Optimal lactate concentrations facilitate endometrial cell proliferation and apoptosis regulation by enhancing histone acetylation, creating a favorable environment for successful embryo implantation (146, 147). Qianying Yang et al. (147) detected relatively high levels of lactate in the fetal membranes and endometrial tissues of pregnant sheep and found that lactate can induce H3K18 acetylation and regulate the balance of redox homeostasis and apoptosis in the endometrium to ensure successful implantation RSA is closely related to the abnormal proliferation and differentiation of early trophoblast cells. Histone acetyltransferase KAT8 activates downstream gene CDX2 through H4K16ac, regulating trophoblast cell proliferation. Its absence increases the risk of embryo implantation failure. Clinical analysis shows that decreased expression of KAT8, CDX2, and H4K16ac is associated with RSA (148). Meanwhile, H3K4-specific demethylase KDM5C affects the function of trophoblast cells by regulating H3K4me3 demethylation at the promoter of TGF β 2 and RAGE genes. Overexpression of KDM5C can lead to decreased proliferation and invasion ability of trophoblast cells. In vivo experiments further confirmed that overexpression of KDM5C leads to a significant increase in mouse embryo absorption rate (149). In addition, the expression of H3K4me3 and H3K9ac in the placenta of patients with PE is reduced, and these modifications are regulated by substances such as Gal-2 and PPAR γ, suggesting the potential role of epigenetic modifications in gestational hypertension (150, 151). In conclusion, histone modification plays an important role in embryo implantation, placental development, and pregnancy maintenance by regulating gene expression and cell function. Its abnormal expression may lead to complications such as pregnancy diabetes, RSA, and PE.
Abnormal expression of ncRNA can affect pregnancy outcomes by regulating the function of trophoblast cells. In RSA, miR-153-3p is highly expressed in extracellular vesicles derived from decidual macrophages, inhibiting the proliferation and migration of trophoblast cells through the IDO/STAT3 pathway (152). Meanwhile, miR-185-5p expression was elevated in the villi of RPL patients, and further in vitro experiments showed that miR-185-5p leads to pregnancy failure by reducing VEGF expression in dNK cells and decreasing angiogenesis (66). In PE, multiple in vitro experiments have found that miRNA participates in disease development by regulating trophoblast cell proliferation, migration, and apoptosis. For example, miR-515-5P inhibits the proliferation, migration, and invasion of HTR-8/SVneo in trophoblast cells by reducing HDAC2 levels (153). MiR-23a inhibits HDAC2 and activates NF-κB, hindering the migration of trophoblast cells and promoting apoptosis (154). MiR-199a-5p inhibits trophoblast cell invasion by suppressing VEGFA expression (155). In addition, Fu et al. (156) have outlined the role of ncRNAs in pregnancy-related complications, which will not be described in detail here. The results indicate that ncRNA significantly contributes to the pathogenesis of RSA and PE by modulating crucial molecular pathways.
N6-methyladenosine (m6A) RNA modification is a pivotal regulatory mechanism in pregnancy-related disorders. In PE, in vitro studies has found the methylation level of m6A RNA in the placental trophoblast is significantly increased, and the methyltransferase METTL3 participates in the pathogenesis by regulating the expression of hnRNPC1/C2 (157). METTL14 upregulates FOXO3a expression through m6A dependent mechanism, inhibiting proliferation and invasion of trophoblast cells (158). At the same time, the m6A-modified circRNA in PE placenta generally increases, while the m6A modification of circPAPP2 is enhanced, but its expression is reduced. Its stability is regulated by IGF2BP3 (159). In gestational diabetes (GDM), the researchers found that the overall mRNA m6A methylation level in the fetal liver of the mouse was significantly increased by constructing a GDM mouse model, and in vitro experiments revealed that RBM15 mediated m6A modification affected the insulin sensitivity of offspring by regulating the expression of CLDN4, leading to metabolic syndrome (160). Compared with normal early pregnant women, the m6A methylation level in placental villus tissue of RM patients was significantly reduced, while ALKBH5 expression was not explicitly regulated. In vitro experiments have shown that knocking down ALKBH5 promotes trophoblast invasion, while overexpression inhibits invasion. It may be that ALKBH5 governs the stability of CYR61 mRNA through a dependent mechanism, thereby affecting the function of the trophoblast (132). In addition, In vitro studies have found that reduced m6A modification mediated by METTL3 can increase ZBTB4 expression, impair the invasive ability of the trophoblast layer, and lead to adverse pregnancy outcomes (161). Suqi Wu et al. (162) further elaborated on the role of m6A modification in maternal-fetal crosstalk and its potential mechanisms in pregnancy-related diseases. These findings suggest that m6A modification plays an important role in pregnancy-related diseases by regulating different target genes. However, the complex regulatory network of m6A modification and its specific functions under different pathological conditions still needs further exploration. Future research should combine multi-omics analysis and in vivo experiments to deeply reveal the dynamic regulatory mechanism of m6A modification at the maternal-fetal interface.
Successful pregnancy depends on the dynamic balance of multiple immune factors in the maternal-fetal interface immune microenvironment (3). The production of maternal autoimmune antibodies and the imbalance of immune regulation at the maternal-fetal interface are important factors affecting embryo implantation and leading to RSA. In recent years, epigenetic research has revealed the key roles of DNA methylation, histone modification, ncRNA and RNA methylation in immune regulation at the maternal-fetal interface. Especially DNA methylation, as the most promising and sensitive biological marker in epigenetic modifications, its abnormalities are not only closely related to the occurrence of various immune-related diseases but can also affect the immune tolerance at the maternal-fetal interface, leading to the maternal immune system producing a defensive rejection response to the embryo (13, 54). Research on DNA methylation in oncology has shown that it occurs earlier than gene mutations and can directly obtain relevant genetic material from tissues, providing the possibility for early prevention and treatment of diseases. The pathogenesis of immune-type RSA is similar to that of tumors, characterized by immune cell dysfunction and immune response suppression. With the deepening of epigenetic research, DNA methylation is expected to become a key breakthrough point for targeted diagnosis and precise treatment of immune type RSA, promoting the rapid development of reproductive medicine.
The epigenetic study of immune regulation at the maternal-fetal interface provides a new perspective for analyzing pregnancy’s physiological and pathological mechanisms. By revealing the role of epigenetic modifications in the maternal-fetal interface immune microenvironment, researchers can gain a deeper understanding of the pathogenesis of pregnancy-related diseases and identify effective targets for immunotherapy. For example, dynamic changes in DNA methylation and histone modification may become key nodes in regulating maternal-fetal interface immune tolerance. In addition, the role of ncRN (such as miRNA and lncRNA) in regulating immune cell function at the maternal-fetal interface is gradually revealed. These findings lay a theoretical foundation for developing intervention strategies for pregnancy-related diseases based on epigenetics.
The fundamental causes of many placental diseases are still unclear. However, it is now widely believed that immune dysregulation at the placental interface can lead to many diseases (135, 163). Meanwhile, compared to genetic changes, epigenetic abnormalities are more straightforward to reverse. Drugs targeting specific epigenetic mechanisms involved in gene expression regulation, and even some nutrients, may become emerging tools for disease prevention or treatment. In this review, some evidence suggests that epigenetic modifications of inhibitory compounds or regulation of noncoding RNA expression through genetic tools may alleviate placental pregnancy-related diseases caused by immune dysregulation at the placental interface. These will provide new ideas for precise prevention and targeted treatment of diseases related to placental pregnancy.
Although significant progress has been made in the study of epigenetics in the maternal-fetal interface immune microenvironment, some limitations remain. Existing research has focused chiefly on single epigenetic modifications (such as DNA methylation or specific noncoding RNA). In contrast, research on the interactions and synergistic regulatory mechanisms between multiple epigenetic modifications is still insufficient. In addition, most studies rely on in vitro cell experiments or animal models, which make it difficult to fully simulate the complex immune microenvironment of the human maternal-fetal interface. However, a great deal of further research is still needed. Examples include (1) exploring how dysregulation of immunoregulation at the placental interface leads to epigenetic alterations in specific genes; (2) identifying human susceptibility genes in epigenetic alterations induced by dysregulation of immunoregulation at the placental interface; (3) demonstrating whether epigenetic alterations can be used as a biomarker for the early detection of, and therapeutic targeting for, pregnancy-associated diseases induced by dysregulation of immunoregulation at the placental interface; (4) developing novel inhibitors targeting epigenetic modifications;and (5) In-depth study of the interaction network and synergistic regulatory mechanism among various epigenetic modifications.
XQM: Visualization, Writing – original draft, Writing – review & editing. XC: Visualization, Writing – original draft, Writing – review & editing. XFM: Conceptualization, Supervision, Writing – review & editing. MC: Supervision, Writing – review & editing. YZ: Conceptualization, Supervision, Validation, Writing – review & editing.
The author(s) declare that financial support was received for the research and/or publication of this article. This work was supported by the Key Research and Development Program of Hubei Province (Grant No. 2022BCA010), Hubei Provincial Key Laboratory Open Project - Seed Fund Project (Grant No. 2024KFZ001).
Figures were created using Figdraw (www.figdraw.com).
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fimmu.2025.1549839/full#supplementary-material
1. PrabhuDas M, Bonney E, Caron K, Dey S, Erlebacher A, Fazleabas A, et al. Immune mechanisms at the maternal-fetal interface: perspectives and challenges. Nat Immunol. (2015) 16:328–34. doi: 10.1038/ni.3131
2. Ghaebi M, Nouri M, Ghasemzadeh A, Farzadi L, Jadidi-Niaragh F, Ahmadi M, et al. Immune regulatory network in successful pregnancy and reproductive failures. Biomed pharmacother = Biomed pharmacotherapie. (2017) 88:61–73. doi: 10.1016/j.biopha.2017.01.016
3. Mor G, Aldo P, Alvero AB. The unique immunological and microbial aspects of pregnancy. Nat Rev Immunol. (2017) 17:469–82. doi: 10.1038/nri.2017.64
4. Li D, Zheng L, Zhao D, Xu Y, Wang Y. The role of immune cells in recurrent spontaneous abortion. Reprod Sci (Thousand Oaks Calif.). (2021) 28:3303–15. doi: 10.1007/s43032-021-00599-y
5. Robinson DP, Klein SL. Pregnancy and pregnancy-associated hormones alter immune responses and disease pathogenesis. Hormones Behav. (2012) 62:263–71. doi: 10.1016/j.yhbeh.2012.02.023
6. Li M, Sun F, Qian J, Chen L, Li D, Wang S, et al. Tim-3/CTLA-4 pathways regulate decidual immune cells-extravillous trophoblasts interaction by IL-4 and IL-10. FASEB J. (2021) 35:e21754. doi: 10.1096/fj.202100142R
7. Feinberg AP. Epigenetics at the epicenter of modern medicine. JAMA. (2008) 299:1345–50. doi: 10.1001/jama.299.11.1345
8. Cavalli G, Heard E. Advances in epigenetics link genetics to the environment and disease. Nature. (2019) 571:489–99. doi: 10.1038/s41586-019-1411-0
9. Harvey ZH, Chen Y, Jarosz DF. Protein-based inheritance: epigenetics beyond the chromosome. Mol Cell. (2018) 69:195–202. doi: 10.1016/j.molcel.2017.10.030
10. Nicoglou A, Merlin F. Epigenetics: A way to bridge the gap between biological fields. Stud history philosophy Biol Biomed Sci. (2017) 66:73–82. doi: 10.1016/j.shpsc.2017.10.002
11. Inbar-Feigenberg M, Choufani S, Butcher DT, Roifman M, Weksberg R. Basic concepts of epigenetics. Fertility sterility. (2013) 99:607–15. doi: 10.1016/j.fertnstert.2013.01.117
12. Ginder GD, Williams DC Jr. Readers of DNA methylation, the MBD family as potential therapeutic targets. Pharmacol Ther. (2018) 184:98–111. doi: 10.1016/j.pharmthera.2017.11.002
13. Greenberg MVC, Bourc’his D. The diverse roles of DNA methylation in mammalian development and disease. Nat Rev Mol Cell Biol. (2019) 20:590–607. doi: 10.1038/s41580-019-0159-6
14. Chen Z, Zhang Y. Role of mammalian DNA methyltransferases in development. Annu Rev Biochem. (2020) 89:135–58. doi: 10.1146/annurev-biochem-103019-102815
15. Schübeler D. Function and information content of DNA methylation. Nature. (2015) 517:321–6. doi: 10.1038/nature14192
16. Kim SY, Romero R, Tarca AL, Bhatti G, Kim CJ, Lee J, et al. Methylome of fetal and maternal monocytes and macrophages at the feto-maternal interface. Am J Reprod Immunol (New York N.Y.: 1989). (2012) 68:8–27. doi: 10.1111/j.1600-0897.2012.01108.x
17. Hu Y, Yuen R, Eastabrook G, Dutz J, Tan R, von Dadelszen P, et al. PP065. dNK and dNK-CM mediated alterations of DNA methylation in extravillous cytotrophoblasts (EVTS). Pregnancy hypertension. (2012) 2:277. doi: 10.1016/j.preghy.2012.04.176
18. Fernández-Sánchez A, Baragaño Raneros A, Carvajal Palao R, Sanz AB, Ortiz A, Ortega F, et al. DNA demethylation and histone H3K9 acetylation determine the active transcription of the NKG2D gene in human CD8+ T and NK cells. Epigenetics. (2013) 8:66–78. doi: 10.4161/epi.23115
19. Liu L, Zhang X, Rong C, Rui C, Ji H, Qian YJ, et al. Distinct DNA methylomes of human placentas between pre-eclampsia and gestational diabetes mellitus. Cell Physiol Biochem. (2014) 34:1877–89. doi: 10.1159/000366386
20. Deshpande SS, Balasinor NH. Placental defects: an epigenetic perspective. Reprod Sci (Thousand Oaks Calif.). (2018) 25:1143–60. doi: 10.1177/1933719118766265
21. Alaskhar Alhamwe B, Khalaila R, Wolf J, von Bülow V, Harb H, Alhamdan F, et al. Histone modifications and their role in epigenetics of atopy and allergic diseases. Allergy asthma Clin Immunol. (2018) 14:39. doi: 10.1186/s13223-018-0259-4
22. Ding P, Ma Z, Liu D, Pan M, Li H, Feng Y, et al. Lysine acetylation/deacetylation modification of immune-related molecules in cancer immunotherapy. Front Immunol. (2022) 13:865975. doi: 10.3389/fimmu.2022.865975
23. Jenuwein T, Allis CD. Translating the histone code. Sci (New York N.Y.). (2001) 293:1074–80. doi: 10.1126/science.1063127
24. Filippakopoulos P, Knapp S. Targeting bromodomains: epigenetic readers of lysine acetylation. Nat Rev Drug Discovery. (2014) 13:337–56. doi: 10.1038/nrd4286
25. Hyndman KA, Knepper MA. Dynamic regulation of lysine acetylation: the balance between acetyltransferase and deacetylase activities. Am J Physiol Renal Physiol. (2017) 313:F842–f846. doi: 10.1152/ajprenal.00313.2017
26. Yao Y, Hao F, Tang LC, Xu XH, Jin L. Downregulation of HDAC8 expression decreases CD163 levels and promotes the apoptosis of macrophages by activating the ERK signaling pathway in recurrent spontaneous miscarriage. Mol Hum Reprod. (2020) 26:521–31. doi: 10.1093/molehr/gaaa035
27. Jaju Bhattad G, Jeyarajah MJ, McGill MG, Dumeaux V, Okae H, Arima T, et al. Histone deacetylase 1 and 2 drive differentiation and fusion of progenitor cells in human placental trophoblasts. Cell Death Dis. (2020) 11:311. doi: 10.1038/s41419-020-2500-6
28. Martin C, Zhang Y. The diverse functions of histone lysine methylation. Nat Rev Mol Cell Biol. (2005) 6:838–49. doi: 10.1038/nrm1761
29. Bannister AJ, Kouzarides T. Reversing histone methylation. Nature. (2005) 436:1103–6. doi: 10.1038/nature04048
30. Bassett SA, Barnett MP. The role of dietary histone deacetylases (HDACs) inhibitors in health and disease. Nutrients. (2014) 6:4273–301. doi: 10.3390/nu6104273
31. Jenuwein T. The epigenetic magic of histone lysine methylation. FEBS J. (2006) 273:3121–35. doi: 10.1111/j.1742-4658.2006.05343.x
32. Berger SL. The complex language of chromatin regulation during transcription. Nature. (2007) 447:407–12. doi: 10.1038/nature05915
33. Li Y, Ge K, Li T, Cai R, Chen Y. The engagement of histone lysine methyltransferases with nucleosomes: structural basis, regulatory mechanisms, and therapeutic implications. Protein Cell. (2023) 14:165–79. doi: 10.1093/procel/pwac032
34. Rossetto D, Avvakumov N, Côté J. Histone phosphorylation: a chromatin modification involved in diverse nuclear events. Epigenetics. (2012) 7:1098–108. doi: 10.4161/epi.21975
35. Fillingham J, Keogh MC, Krogan NJ. GammaH2AX and its role in DNA double-strand break repair. Biochem Cell Biol = Biochimie biologie cellulaire. (2006) 84:568–77. doi: 10.1139/o06-072
36. Banerjee T, Chakravarti D. A peek into the complex realm of histone phosphorylation. Mol Cell Biol. (2011) 31:4858–73. doi: 10.1128/MCB.05631-11
37. Dahariya S, Paddibhatla I, Kumar S, Raghuwanshi S, Pallepati A, Gutti RK, et al. Classification, biogenesis and functions in blood cells. Mol Immunol. (2019) 112:82–92. doi: 10.1016/j.molimm.2019.04.011
38. Abraham JM, Meltzer SJ. Long noncoding RNAs in the pathogenesis of Barrett’s esophagus and esophageal carcinoma. Gastroenterology. (2017) 153:27–34. doi: 10.1053/j.gastro.2017.04.046
39. Pinho JD, Silva GEB, Teixeira-Júnior AAL, Rocha TMS, Batista LL, de Sousa AM, et al. Non-coding RNA in penile cancer. Front Oncol. (2022) 12:812008. doi: 10.3389/fonc.2022.812008
40. Ran R, Liao HY, Wang ZQ, Gong CY, Zhou KS, Zhang HH. Mechanisms and functions of long noncoding RNAs in intervertebral disc degeneration. Pathology Res Pract. (2022) 235:153959. doi: 10.1016/j.prp.2022.153959
41. Huang Z, Shan R, Wen W, Li J, Zeng X, Wan R. The emerging roles of circ-ABCB10 in cancer. Front Cell Dev Biol. (2022) 10:782938. doi: 10.3389/fcell.2022.782938
42. Hopfinger MC, Kirkpatrick CC, Znosko BM. Predictions and analyses of RNA nearest neighbor parameters for modified nucleotides. Nucleic Acids Res. (2020) 48:8901–13. doi: 10.1093/nar/gkaa654
43. Jiang X, Liu B, Nie Z, Duan L, Xiong Q, Jin Z, et al. The role of m6A modification in the biological functions and diseases. Signal transduction targeted Ther. (2021) 6:74. doi: 10.1038/s41392-020-00450-x
44. Harper JE, Miceli SM, Roberts RJ, Manley JL. Sequence specificity of the human mRNA N6-adenosine methylase in vitro. Nucleic Acids Res. (1990) 18:5735–41. doi: 10.1093/nar/18.19.5735
45. Oerum S, Meynier V, Catala M, Tisné C. A comprehensive review of m6A/m6Am RNA methyltransferase structures. Nucleic Acids Res. (2021) 49:7239–55. doi: 10.1093/nar/gkab378
46. Zheng ZH, Zhang GL, Jiang RF, Hong YQ, Zhang QY, He JP, et al. METTL3 is essential for normal progesterone signaling during embryo implantation via m(6)A-mediated translation control of progesterone receptor. Proc Natl Acad Sci United States America. (2023) 120:e2214684120. doi: 10.1073/pnas.2214684120
47. Liu C, Ma Y, Shang Y, Huo R, Li W. Post-translational regulation of the maternal-to-zygotic transition. Cell Mol Life sciences: CMLS. (2018) 75:1707–22. doi: 10.1007/s00018-018-2750-y
48. Yovel G, Shakhar K, Ben-Eliyahu S. The effects of sex, menstrual cycle, and oral contraceptives on the number and activity of natural killer cells. Gynecologic Oncol. (2001) 81:254–62. doi: 10.1006/gyno.2001.6153
49. Moffett-King A. Natural killer cells and pregnancy. Nat Rev Immunol. (2002) 2:656–63. doi: 10.1038/nri886
50. Gaynor LM, Colucci F. Uterine natural killer cells: functional distinctions and influence on pregnancy in humans and mice. Front Immunol. (2017) 8:467. doi: 10.3389/fimmu.2017.00467
51. Vento-Tormo R, Efremova M, Botting RA, Turco MY, Vento-Tormo M, Meyer KB, et al. Single-cell reconstruction of the early maternal-fetal interface in humans. Nature. (2018) 563:347–53. doi: 10.1038/s41586-018-0698-6
52. Fu B, Zhou Y, Ni X, Tong X, Xu X, Dong Z, et al. Natural killer cells promote fetal development through the secretion of growth-promoting factors. Immunity. (2017) 47:1100–13.e6. doi: 10.1016/j.immuni.2017.11.018
53. Colucci F, Kieckbusch J. Maternal uterine natural killer cells nurture fetal growth: in medio stat virtus. Trends Mol Med. (2015) 21:60–7. doi: 10.1016/j.molmed.2014.12.009
54. Zhang Q, Xiao Z, Lee CL, Duan YG, Fan X, Yeung WSB, et al. The regulatory roles of chemerin-chemokine-like receptor 1 axis in placental development and vascular remodeling during early pregnancy. Front Cell Dev Biol. (2022) 10:883636. doi: 10.3389/fcell.2022.883636
55. Wiencke JK, Butler R, Hsuang G, Eliot M, Kim S, Sepulveda MA, et al. The DNA methylation profile of activated human natural killer cells. Epigenetics. (2016) 11:363–80. doi: 10.1080/15592294.2016.1163454
56. Cerdeira AS, Rajakumar A, Royle CM, Lo A, Husain Z, Thadhani RI, et al. Conversion of peripheral blood NK cells to a decidual NK-like phenotype by a cocktail of defined factors. J Immunol (Baltimore Md.: 1950). (2013) 190:3939–48. doi: 10.4049/jimmunol.1202582
57. Nandakumar V, Chou Y, Zang L, Huang XF, Chen SY. Epigenetic control of natural killer cell maturation by histone H2A deubiquitinase, MYSM1. Proc Natl Acad Sci United States America. (2013) 110:E3927–36. doi: 10.1073/pnas.1308888110
58. Sciumè G, Mikami Y, Jankovic D, Nagashima H, Villarino AV, Morrison T, et al. Rapid enhancer remodeling and transcription factor repurposing enable high magnitude gene induction upon acute activation of NK cells. Immunity. (2020) 53:745–58.e4. doi: 10.1016/j.immuni.2020.09.008
59. Wiedemann GM, Santosa EK, Grassmann S, Sheppard S, Le Luduec JB, Adams NM, et al. Deconvoluting global cytokine signaling networks in natural killer cells. Nat Immunol. (2021) 22:627–38. doi: 10.1038/s41590-021-00909-1
60. Ma Y, Yu X, Zhang L, Liu J, Shao X, Li YX, et al. Uterine decidual niche modulates the progressive dedifferentiation of spiral artery vascular smooth muscle cells during human pregnancy†. Biol Reprod. (2021) 104:624–37. doi: 10.1093/biolre/ioaa208
61. Zhang J, Xue M, Huang J, He S, Zhu L, Zhao X, et al. Deficiency of UCHL1 results in insufficient decidualization accompanied by impaired dNK modulation and eventually miscarriage. J Trans Med. (2024) 22:478. doi: 10.1186/s12967-024-05253-0
62. Gamliel M, Goldman-Wohl D, Isaacson B, Gur C, Stein N, Yamin R, et al. Trained memory of human uterine NK cells enhances their function in subsequent pregnancies. Immunity. (2018) 48:951–62.e5. doi: 10.1016/j.immuni.2018.03.030
63. Carlino C, Rippo MR, Lazzarini R, Monsurrò V, Morrone S, Angelini S, et al. Differential microRNA expression between decidual and peripheral blood natural killer cells in early pregnancy. Hum Reprod (Oxford England). (2018) 33:2184–95. doi: 10.1093/humrep/dey323
64. Li D, Li J, Jia B, Wang Y, Zhang J, Liu G. Genome-wide identification of microRNAs in decidual natural killer cells from patients with unexplained recurrent spontaneous abortion. Am J Reprod Immunol (New York N.Y.: 1989). (2018) 80:e13052. doi: 10.1111/aji.2018.80.issue-5
65. Huang Q, Ding J, Gong M, Wei M, Zhao Q, Yang J. Effect of miR-30e regulating NK cell activities on immune tolerance of maternal-fetal interface by targeting PRF1. Biomed pharmacother = Biomed pharmacotherapie. (2019) 109:1478–87. doi: 10.1016/j.biopha.2018.09.172
66. Xiong Y, Fang Z, Dong J, Chen S, Mao J, Zhang W, et al. Maternal circulating exosomal miR-185-5p levels as a predictive biomarker in patients with recurrent pregnancy loss. J assisted Reprod Genet. (2023) 40:553–66. doi: 10.1007/s10815-023-02733-y
67. Fang Z, Mao J, Huang J, Sun H, Lu X, Lei H, et al. Increased levels of villus-derived exosomal miR-29a-3p in normal pregnancy than uRPL patients suppresses decidual NK cell production of interferon-γ and exerts a therapeutic effect in abortion-prone mice. Cell communication signaling: CCS. (2024) 22:230. doi: 10.1186/s12964-024-01610-0
68. Song H, Song J, Cheng M, Zheng M, Wang T, Tian S, et al. METTL3-mediated m(6)A RNA methylation promotes the anti-tumour immunity of natural killer cells. Nat Commun. (2021) 12:5522. doi: 10.1038/s41467-021-25803-0
69. Sheng YR, Hu WT, Wei CY, Tang LL, Liu YK, Liu YY, et al. IL-33/ST2 axis affects the polarization and efferocytosis of decidual macrophages in early pregnancy. Am J Reprod Immunol (New York N.Y.: 1989). (2018) 79:e12836. doi: 10.1111/aji.2018.79.issue-6
70. Murray PJ. Macrophage polarization. Annu Rev Physiol. (2017) 79:541–66. doi: 10.1146/annurev-physiol-022516-034339
71. Meng YH, Zhou WJ, Jin LP, Liu LB, Chang KK, Mei J, et al. RANKL-mediated harmonious dialogue between fetus and mother guarantees smooth gestation by inducing decidual M2 macrophage polarization. Cell Death Dis. (2017) 8:e3105. doi: 10.1038/cddis.2017.505
72. Mor G, Cardenas I, Abrahams V, Guller S. Inflammation and pregnancy: the role of the immune system at the implantation site. Ann New York Acad Sci. (2011) 1221:80–7. doi: 10.1111/j.1749-6632.2010.05938.x
73. Tsao FY, Wu MY, Chang YL, Wu CT, Ho HN. M1 macrophages decrease in the deciduae from normal pregnancies but not from spontaneous abortions or unexplained recurrent spontaneous abortions. J Formosan Med Assoc = Taiwan yi zhi. (2018) 117:204–11. doi: 10.1016/j.jfma.2017.03.011
74. Schonkeren D, van der Hoorn ML, Khedoe P, Swings G, van Beelen E, Claas F, et al. Differential distribution and phenotype of decidual macrophages in preeclamptic versus control pregnancies. Am J Pathol. (2011) 178:709–17. doi: 10.1016/j.ajpath.2010.10.011
75. Schuster J, Uzun A, Stablia J, Schorl C, Mori M, Padbury JF. Effect of prematurity on genome wide methylation in the placenta. BMC Med Genet. (2019) 20:116. doi: 10.1186/s12881-019-0835-6
76. Walsh SW, Chumble AA, Washington SL, Archer KJ, Sahingur SE, Strauss JF 3rd. Increased expression of toll-like receptors 2 and 9 is associated with reduced DNA methylation in spontaneous preterm labor. J Reprod Immunol. (2017) 121:35–41. doi: 10.1016/j.jri.2017.05.003
77. Liu Y, Du M, Lin HY. Histone deacetylase 9 deficiency exaggerates uterine M2 macrophage polarization. J Cell Mol Med. (2021) 25:7690–708. doi: 10.1111/jcmm.v25.16
78. Zhu X, Liu H, Zhang Z, Wei R, Zhou X, Wang Z, et al. MiR-103 protects from recurrent spontaneous abortion via inhibiting STAT1 mediated M1 macrophage polarization. Int J Biol Sci. (2020) 16:2248–64. doi: 10.7150/ijbs.46144
79. Yang J, Li L, Wang L, Chen R, Yang X, Wu J, et al. Trophoblast-derived miR-410-5p induces M2 macrophage polarization and mediates immunotolerance at the fetal-maternal interface by targeting the STAT1 signaling pathway. J Trans Med. (2024) 22:19. doi: 10.1186/s12967-023-04831-y
80. Wang P, Ma Z, Wang Z, Wang X, Zhao G, Wang Z. MiR-6869-5p induces M2 polarization by regulating PTPRO in gestational diabetes mellitus. Mediators Inflammation. (2021) 2021:6696636. doi: 10.1155/2021/6696636
81. Wang P, Wang Z, Liu G, Jin C, Zhang Q, Man S, et al. miR-657 promotes macrophage polarization toward M1 by targeting FAM46C in gestational diabetes mellitus. Mediators Inflammation. (2019) 2019:4851214. doi: 10.1155/2019/4851214
82. Bai K, Li J, Lin L, Zhang Q, Zhong J, Liu X, et al. Placenta exosomal miRNA-30d-5p facilitates decidual macrophage polarization by targeting HDAC9. J leukocyte Biol. (2023) 113:434–44. doi: 10.1093/jleuko/qiad022
83. Ma C, Zhao M, Wang X, Zhong W, Zhang Y, Xu L. Downregulation of miR-455-3p in decidual cells promotes macrophage polarization and suppresses trophoblasts invasion. Exp Cell Res. (2023) 425:113510. doi: 10.1016/j.yexcr.2023.113510
84. Ye HX, Liao GN, Dong YJ, Li L, Wang XM, Shu J, et al. miR-146a-5p enhances embryo survival in unexplained recurrent spontaneous abortion by promoting M2 polarization of decidual macrophages. Int Immunopharmacol. (2022) 110:108930. doi: 10.1016/j.intimp.2022.108930
85. Yin T, Li X, Li Y, Zang X, Liu L, Du M. Macrophage plasticity and function in cancer and pregnancy. Front Immunol. (2023) 14:1333549. doi: 10.3389/fimmu.2023.1333549
86. Wu S, Han L, Zhou M, Li X, Luo L, Wang Z, et al. LncRNA AOC4P recruits TRAF6 to regulate EZH2 ubiquitination and participates in trophoblast glycolysis and M2 macrophage polarization which is associated with recurrent spontaneous abortion. Int Immunopharmacol. (2023) 125:111201. doi: 10.1016/j.intimp.2023.111201
87. Li X, Song Y, Liu F, Liu D, Miao H, Ren J, et al. Long non-coding RNA MALAT1 promotes proliferation, angiogenesis, and immunosuppressive properties of mesenchymal stem cells by inducing VEGF and IDO. J Cell Biochem. (2017) 118:2780–91. doi: 10.1002/jcb.v118.9
88. Wu HY, Liu K, Zhang JL. LINC00240/miR-155 axis regulates function of trophoblasts and M2 macrophage polarization via modulating oxidative stress-induced pyroptosis in preeclampsia. Mol Med (Cambridge Mass.). (2022) 28:119. doi: 10.1186/s10020-022-00531-3
89. Zhao G, Wang Y, Wang Y, Cui X, Liu L, Zhi Y, et al. Downregulation of LINC00221 promotes angiopoiesis in HUVEC and inhibits recruitment of macrophages by augmenting miR-542-3p in trophoblast cells. J assisted Reprod Genet. (2022) 39:2381–93. doi: 10.1007/s10815-022-02593-y
90. Cai Y, Yu R, Kong Y, Feng Z, Xu Q. METTL3 regulates LPS-induced inflammatory response via the NOD1 signaling pathway. Cell signalling. (2022) 93:110283. doi: 10.1016/j.cellsig.2022.110283
91. Li Z, Xu Q, Huangfu N, Chen X, Zhu J. Mettl3 promotes oxLDL-mediated inflammation through activating STAT1 signaling. J Clin Lab Anal. (2022) 36:e24019. doi: 10.1002/jcla.24019
92. Zhao Y, Sun J, Jin L. The N6-methyladenosine regulator ALKBH5 mediated stromal cell-macrophage interaction via VEGF signaling to promote recurrent spontaneous abortion: A bioinformatic and in vitro study. Int J Mol Sci. (2022) 23:15819. doi: 10.3390/ijms232415819
93. Wheeler KC, Jena MK, Pradhan BS, Nayak N, Das S, Hsu CD, et al. VEGF may contribute to macrophage recruitment and M2 polarization in the decidua. PloS One. (2018) 13:e0191040. doi: 10.1371/journal.pone.0191040
94. Zheng W, Fu Z, Tan X, Liang X, Cao L. Bioinformatic analysis of m6A regulator-mediated RNA methylation modification patterns and immune microenvironment characterization in endometriosis. Biochem Genet. (2025) 63:433–64. doi: 10.1007/s10528-024-10725-5
95. Qin XY, Shen HH, Zhou WJ, Mei J, Lu H, Tan XF, et al. Insight of autophagy in spontaneous miscarriage. Int J Biol Sci. (2022) 18:1150–70. doi: 10.7150/ijbs.68335
96. Wei J, Zheng W, Chapman NM, Geiger TL, Chi H. T cell metabolism in homeostasis and cancer immunity. Curr Opin Biotechnol. (2021) 68:240–50. doi: 10.1016/j.copbio.2021.02.003
97. Wei Y, Ding J, Li J, Cai S, Liu S, Hong L, et al. Metabolic reprogramming of immune cells at the maternal-fetal interface and the development of techniques for immunometabolism. Front Immunol. (2021) 12:717014. doi: 10.3389/fimmu.2021.717014
98. Saravia J, Chapman NM, Chi H. Helper T cell differentiation. Cell Mol Immunol. (2019) 16:634–43. doi: 10.1038/s41423-019-0220-6
99. Nonaka K, Saio M, Umemura N, Kikuchi A, Takahashi T, Osada S, et al. Th1 polarization in the tumor microenvironment upregulates the myeloid-derived suppressor-like function of macrophages. Cell Immunol. (2021) 369:104437. doi: 10.1016/j.cellimm.2021.104437
100. Liu G, Liu F. Bach2: A key regulator in Th2-related immune cells and Th2 immune response. J Immunol Res. (2022) 2022:2814510. doi: 10.1155/2022/2814510
101. Gao X, Zhong Y, Liu Y, Ding R, Chen J. The role and function of regulatory T cells in toxoplasma Gondii-induced adverse pregnancy outcomes. J Immunol Res. (2021) 2021:8782672. doi: 10.1155/2021/8782672
102. Lombardelli L, Logiodice F, Kullolli O, Haller H, Agostinis C, Bulla R, et al. At embryo implantation site IL-35 secreted by trophoblast, polarizing T cells towards IL-35+ IL-10+ IL-4+ Th2-type cells, could favour fetal allograft tolerance and pregnancy success. Int J Mol Sci. (2022) 23:4926. doi: 10.3390/ijms23094926
103. Schwede S, Alfer J, von Rango U. Differences in regulatory T-cell and dendritic cell pattern in decidual tissue of placenta accreta/increta cases. Placenta. (2014) 35:378–85. doi: 10.1016/j.placenta.2014.03.004
104. Wang W, Sung N, Gilman-Sachs A, Kwak-Kim J. T helper (Th) cell profiles in pregnancy and recurrent pregnancy losses: Th1/Th2/Th9/Th17/Th22/Tfh cells. Front Immunol. (2020) 11:2025. doi: 10.3389/fimmu.2020.02025
105. Iqbal S, Lockett GA, Holloway JW, Arshad SH, Zhang H, Kaushal A, et al. Changes in DNA methylation from age 18 to pregnancy in type 1, 2, and 17 T helper and regulatory T-cells pathway genes. Int J Mol Sci. (2018) 19:477. doi: 10.3390/ijms19020477
106. Zhou Q, Xiong Y, Qu B, Bao A, Zhang Y. DNA methylation and recurrent pregnancy loss: A mysterious compass? Front Immunol. (2021) 12:738962. doi: 10.3389/fimmu.2021.738962
107. Hou W, Li Z, Li Y, Fang L, Li J, Huang J, et al. Correlation between protein expression of FOXP3 and level of FOXP3 promoter methylation in recurrent spontaneous abortion. J obstetrics gynaecology Res. (2016) 42:1439–44. doi: 10.1111/jog.2016.42.issue-11
108. Zhang L, Long X, Yin Y, Wang J, Zhu H, Chen J, et al. Histone methyltransferase Nsd2 ensures maternal-fetal immune tolerance by promoting regulatory T-cell recruitment. Cell Mol Immunol. (2022) 19:634–43. doi: 10.1038/s41423-022-00849-2
109. Schjenken JE, Moldenhauer LM, Zhang B, Care AS, Groome HM, Chan HY, et al. MicroRNA miR-155 is required for expansion of regulatory T cells to mediate robust pregnancy tolerance in mice. Mucosal Immunol. (2020) 13:609–25. doi: 10.1038/s41385-020-0255-0
110. Geng X, Mao G, Zhao D, Xiang Y, Wang M, Yu G, et al. Downregulation of miR-33a/b and miR-181a contributes to recurrent pregnancy loss by upregulating S1PR1 and repressing regulatory T cell differentiation. Placenta. (2022) 121:137–44. doi: 10.1016/j.placenta.2022.03.011
111. Zolfaghari MA, Arefnezhad R, Parhizkar F, Hejazi MS, Motavalli Khiavi F, Mahmoodpoor A, et al. T lymphocytes and preeclampsia: The potential role of T-cell subsets and related MicroRNAs in the pathogenesis of preeclampsia. Am J Reprod Immunol (New York N.Y.: 1989). (2021) 86:e13475. doi: 10.1111/aji.13475
112. McCall JL, Varney ME, Rice E, Dziadowicz SA, Hall C, Blethen KE, et al. Prenatal cadmium exposure alters proliferation in mouse CD4(+) T cells via lncRNA Snhg7. Front Immunol. (2021) 12:720635. doi: 10.3389/fimmu.2021.720635
113. Li HB, Tong J, Zhu S, Batista PJ, Duffy EE, Zhao J, et al. m(6)A mRNA methylation controls T cell homeostasis by targeting the IL-7/STAT5/SOCS pathways. Nature. (2017) 548:338–42. doi: 10.1038/nature23450
114. Tong J, Cao G, Zhang T, Sefik E, Amezcua Vesely MC, Broughton JP, et al. m(6)A mRNA methylation sustains Treg suppressive functions. Cell Res. (2018) 28:253–6. doi: 10.1038/cr.2018.7
115. Yao Y, Yang Y, Guo W, Xu L, You M, Zhang YC, et al. METTL3-dependent m(6)A modification programs T follicular helper cell differentiation. Nat Commun. (2021) 12:1333. doi: 10.1038/s41467-021-21594-6
116. Gregori S, Amodio G, Quattrone F, Panina-Bordignon P. HLA-G orchestrates the early interaction of human trophoblasts with the maternal niche. Front Immunol. (2015) 6:128. doi: 10.3389/fimmu.2015.00128
117. Moffett A, Loke C. Immunology of placentation in eutherian mammals. Nat Rev Immunol. (2006) 6:584–94. doi: 10.1038/nri1897
118. Vlahos A, Mansell T, Saffery R, Novakovic B. Human placental methylome in the interplay of adverse placental health, environmental exposure, and pregnancy outcome. PloS Genet. (2019) 15:e1008236. doi: 10.1371/journal.pgen.1008236
119. Ruan H, Dai Z, Yan J, Long X, Chen Y, Yang Y, et al. ZBTB24 (Zinc Finger and BTB Domain Containing 24) prevents recurrent spontaneous abortion by promoting trophoblast proliferation, differentiation and migration. Bioengineered. (2022) 13:2777–90. doi: 10.1080/21655979.2021.2019655
120. Hu Y, Blair JD, Yuen RK, Robinson WP, von Dadelszen P. Genome-wide DNA methylation identifies trophoblast invasion-related genes: Claudin-4 and Fucosyltransferase IV control mobility via altering matrix metalloproteinase activity. Mol Hum Reprod. (2015) 21:452–65. doi: 10.1093/molehr/gav007
121. Shankar K, Kang P, Zhong Y, Borengasser SJ, Wingfield C, Saben J, et al. Transcriptomic and epigenomic landscapes during cell fusion in BeWo trophoblast cells. Placenta. (2015) 36:1342–51. doi: 10.1016/j.placenta.2015.10.010
122. Kwak YT, Muralimanoharan S, Gogate AA, Mendelson CR. Human trophoblast differentiation is associated with profound gene regulatory and epigenetic changes. Endocrinology. (2019) 160:2189–203. doi: 10.1210/en.2019-00144
123. Yu X, Wu H, Su J, Liu X, Liang K, Li Q, et al. Acetyl-CoA metabolism maintains histone acetylation for syncytialization of human placental trophoblast stem cells. Cell Stem Cell. (2024) 31:1280–97.e7. doi: 10.1016/j.stem.2024.07.003
124. Chen X, Song QL, Li ZH, Ji R, Wang JY, Ge C, et al. Deletion of ACLY disrupts histone acetylation and IL-10 secretion in trophoblasts, which inhibits M2 polarization of macrophages: A possible role in recurrent spontaneous abortion. Oxid Med Cell Longevity. (2022) 2022:5216786. doi: 10.1155/2022/5216786
125. Park S, Shin J, Bae J, Han D, Park SR, Shin J, et al. SIRT1 alleviates LPS-induced IL-1β Production by suppressing NLRP3 inflammasome activation and ROS production in trophoblasts. Cells. (2020) 9:728. doi: 10.3390/cells9030728
126. Liang L, Chen Y, Wu C, Cao Z, Xia L, Meng J, et al. MicroRNAs: key regulators of the trophoblast function in pregnancy disorders. J assisted Reprod Genet. (2023) 40:3–17. doi: 10.1007/s10815-022-02677-9
127. Wei XH, Liao LY, Yin YX, Xu Q, Xie SS, Liu M, et al. Overexpression of long noncoding RNA DUXAP8 inhibits ER-phagy through activating AKT/mTOR signaling and contributes to preeclampsia. Cell Mol Life sciences: CMLS. (2024) 81:336. doi: 10.1007/s00018-024-05385-y
128. Xu Z, Tian P, Guo J, Mi C, Liang T, Xie J, et al. Lnc-HZ01 with m6A RNA methylation inhibits human trophoblast cell proliferation and induces miscarriage by up-regulating BPDE-activated lnc-HZ01/MXD1 positive feedback loop. Sci total Environ. (2021) 776:145950. doi: 10.1016/j.scitotenv.2021.145950
129. Li X, Li L, Ruan L. Circ_0003314 combines with the miR-26b-5p/IL1RAP axis to inhibit HTR-8/SVneo cell proliferation, migration, invasion and tube formation and promote apoptosis. Appl Biochem Biotechnol. (2024) 197:443–58. doi: 10.1007/s12010-024-05011-4
130. Wu H, Zhao L. Circ_0015382 regulates the miR-942-5p/NDRG1 axis to suppress trophoblast cell functions. Reprod Sci (Thousand Oaks Calif.). (2024). doi: 10.1007/s43032-024-01685-7
131. Ou Y, Zhu L, Wei X, Bai S, Chen M, Chen H, et al. Circular RNA circ_0111277 attenuates human trophoblast cell invasion and migration by regulating miR-494/HTRA1/Notch-1 signal pathway in pre-eclampsia. Cell Death Dis. (2020) 11:479. doi: 10.1038/s41419-020-2679-6
132. Li XC, Jin F, Wang BY, Yin XJ, Hong W, Tian FJ. The m6A demethylase ALKBH5 controls trophoblast invasion at the maternal-fetal interface by regulating the stability of CYR61 mRNA. Theranostics. (2019) 9:3853–65. doi: 10.7150/thno.31868
133. Qin C, Wu J, Wei X, Liu X, Lin Y. ALKBH5 modulation of ferroptosis in recurrent miscarriage: implications in cytotrophoblast dysfunction. PeerJ. (2024) 12:e18227. doi: 10.7717/peerj.18227
134. Zhang J, Liu X, Gao Y. FTO protein regulates the TGF-β signalling pathway through RNA N6-methyladenosine modification to induce unexplained recurrent spontaneous abortion. FEBS J. (2024) 291:1545–59. doi: 10.1111/febs.v291.7
135. Brown LY, Bonney EA, Raj RS, Nielsen B, Brown S. Generalized disturbance of DNA methylation in the uterine decidua in the CBA/J x DBA/2 mouse model of pregnancy failure. Biol Reprod. (2013) 89:120. doi: 10.1095/biolreprod.113.113142
136. Huang X, Wang L, Zhao S, Liu H, Chen S, Wu L, et al. Pregnancy induces an immunological memory characterized by maternal immune alterations through specific genes methylation. Front Immunol. (2021) 12:686676. doi: 10.3389/fimmu.2021.686676
137. Lee ED, Mistry HD. Placental related disorders of pregnancy. Int J Mol Sci. (2022) 23:3519. doi: 10.3390/books978-3-0365-5899-8
138. Jia Y, Xie H, Zhang J, Ying H. Induction of TGF-β receptor I expression in a DNA methylation-independent manner mediated by DNMT3A downregulation is involved in early-onset severe preeclampsia. FASEB J. (2020) 34:13224–38. doi: 10.1096/fj.202000253RR
139. D. KM, Kaur L, Randhir KN, Mehendale SS, Sundrani DP, Chandak GR, et al. Placental growth factor and Fms related tyrosine kinase-1 are hypomethylated in preeclampsia placentae. Epigenomics. (2021) 13:257–69. doi: 10.2217/epi-2020-0318
140. Tekola-Ayele F, Zeng X, Ouidir M, Workalemahu T, Zhang C, Delahaye F, et al. DNA methylation loci in placenta associated with birthweight and expression of genes relevant for early development and adult diseases. Clin Epigenet. (2020) 12:78. doi: 10.1186/s13148-020-00873-x
141. Zhou L, Pu Y, Zhou Y, Wang B, Chen Y, Bai Y, et al. Genome wide methylation analysis to uncover genes related to recurrent pregnancy loss. Genes Genomics. (2021) 43:361–9. doi: 10.1007/s13258-020-01020-9
142. Robakis TK, Lee S, Werner E, Liu G, Miller M, Wylie D, et al. DNA methylation patterns in T lymphocytes are generally stable in human pregnancies but CD3 methylation is associated with perinatal psychiatric symptoms. Brain behavior Immun - Health. (2020) 3:100044. doi: 10.1016/j.bbih.2020.100044
143. Qu X, Yu H, Jia B, Yu X, Cui Q, Liu Z, et al. Association of downregulated HDAC 2 with the impaired mitochondrial function and cytokine secretion in the monocytes/macrophages from gestational diabetes mellitus patients. Cell Biol Int. (2016) 40:642–51. doi: 10.1002/cbin.10598
144. Paauw ND, Lely AT, Joles JA, Franx A, Nikkels PG, Mokry M, et al. H3K27 acetylation and gene expression analysis reveals differences in placental chromatin activity in fetal growth restriction. Clin Epigenet. (2018) 10:85. doi: 10.1186/s13148-018-0508-x
145. Liu M, Deng W, Tang L, Liu M, Bao H, Guo C, et al. Menin directs regionalized decidual transformation through epigenetically setting PTX3 to balance FGF and BMP signaling. Nat Commun. (2022) 13:1006. doi: 10.1038/s41467-022-28657-2
146. Zhang D, Tang Z, Huang H, Zhou G, Cui C, Weng Y, et al. Metabolic regulation of gene expression by histone lactylation. Nature. (2019) 574:575–80. doi: 10.1038/s41586-019-1678-1
147. Yang Q, Liu J, Wang Y, Zhao W, Wang W, Cui J, et al. A proteomic atlas of ligand-receptor interactions at the ovine maternal-fetal interface reveals the role of histone lactylation in uterine remodeling. J Biol Chem. (2022) 298:101456. doi: 10.1016/j.jbc.2021.101456
148. Bi S, Huang L, Chen Y, Hu Z, Li S, Wang Y, et al. KAT8-mediated H4K16ac is essential for sustaining trophoblast self-renewal and proliferation via regulating CDX2. Nat Commun. (2024) 15:5602. doi: 10.1038/s41467-024-49930-6
149. Xiao M, Zheng Y, Wang MX, Sun YH, Chen J, Zhu KY, et al. Elevated histone demethylase KDM5C increases recurrent miscarriage risk by preventing trophoblast proliferation and invasion. Cell Death Discovery. (2022) 8:495. doi: 10.1038/s41420-022-01284-y
150. Hahn L, Meister S, Mannewitz M, Beyer S, Corradini S, Hasbargen U, et al. Gal-2 increases H3K4me(3) and H3K9ac in trophoblasts and preeclampsia. Biomolecules. (2022) 12:707. doi: 10.3390/biom12050707
151. Meister S, Hahn L, Beyer S, Paul C, Mitter S, Kuhn C, et al. Regulation of epigenetic modifications in the placenta during preeclampsia: PPARγ Influences H3K4me3 and H3K9ac in extravillous trophoblast cells. Int J Mol Sci. (2021) 22:12469. doi: 10.3390/ijms222212469
152. Ying X, Jin X, Zhu Y, Liang M, Chang X, Zheng L. Exosomes released from decidual macrophages deliver miR-153-3p, which inhibits trophoblastic biological behavior in unexplained recurrent spontaneous abortion. Int Immunopharmacol. (2020) 88:106981. doi: 10.1016/j.intimp.2020.106981
153. Zhang K, Zhang H, Gao S, Sun C, Wang B. Effect and mechanism of microRNA-515-5p in proliferation and apoptosis of trophoblast cells in preeclampsia via manipulating histone deacetylase 2. Mol Reprod Dev. (2023) 90:59–66. doi: 10.1002/mrd.23649
154. Fan Y, Dong Z, Zhou G, Fu J, Zhan L, Gao M, et al. Elevated miR-23a impairs trophoblast migration and invasiveness through HDAC2 inhibition and NF-κB activation. Life Sci. (2020) 261:118358. doi: 10.1016/j.lfs.2020.118358
155. Mei Z, Huang B, Zhang Y, Qian X, Mo Y, Deng N. Histone deacetylase 6 negatively regulated microRNA-199a-5p induces the occurrence of preeclampsia by targeting VEGFA in vitro. Biomed pharmacother = Biomed pharmacotherapie. (2019) 114:108805. doi: 10.1016/j.biopha.2019.108805
156. Fu X, Li Y, Zhang Z, Wang B, Wei R, Chu C, et al. Emerging role of miRNAs, lncRNAs, and circRNAs in pregnancy-associated diseases. Chin Med J. (2023) 136:1300–10. doi: 10.1097/CM9.0000000000002595
157. Gu Y, Chu X, Morgan JA, Lewis DF, Wang Y. Upregulation of METTL3 expression and m6A RNA methylation in placental trophoblasts in preeclampsia. Placenta. (2021) 103:43–9. doi: 10.1016/j.placenta.2020.10.016
158. Fan W, Zhou W, Yan Q, Peng Y, Wang H, Kong C, et al. Upregulation of METTL14 contributes to trophoblast dysfunction by elevating FOXO3a expression in an m(6)A-dependent manner. Placenta. (2022) 124:18–27. doi: 10.1016/j.placenta.2022.05.008
159. Zhang Y, Yang H, Long Y, Zhang Y, Chen R, Shi J, et al. circRNA N6-methyladenosine methylation in preeclampsia and the potential role of N6-methyladenosine-modified circPAPPA2 in trophoblast invasion. Sci Rep. (2021) 11:24357. doi: 10.1038/s41598-021-03662-5
160. Fang J, Wu X, He J, Zhang H, Chen X, Zhang H, et al. RBM15 suppresses hepatic insulin sensitivity of offspring of gestational diabetes mellitus mice via m6A-mediated regulation of CLDN4. Mol Med (Cambridge Mass.). (2023) 29:23. doi: 10.1186/s10020-023-00615-8
161. Huang N, Gao Y, Zhang M, Guo L, Qin L, Liao S, et al. METTL3-mediated m(6)A RNA methylation of ZBTB4 interferes with trophoblast invasion and maybe involved in RSA. Front Cell Dev Biol. (2022) 10:894810. doi: 10.3389/fcell.2022.894810
162. Wu S, Liu K, Zhou B, Wu S. N6-methyladenosine modifications in maternal-fetal crosstalk and gestational diseases. Front Cell Dev Biol. (2023) 11:1164706. doi: 10.3389/fcell.2023.1164706
Keywords: epigenetics, DNA methylation, histone modification, non-coding RNA, RNA methylation, placental interface, immune microenvironment
Citation: Ma X, Chen X, Mu X, Cao M and Zhang Y (2025) Epigenetics of maternal-fetal interface immune microenvironment and placental related pregnancy complications. Front. Immunol. 16:1549839. doi: 10.3389/fimmu.2025.1549839
Received: 22 December 2024; Accepted: 17 March 2025;
Published: 03 April 2025.
Edited by:
Alicia E. Damiano, University of Buenos Aires, ArgentinaReviewed by:
Hong Liu, Huazhong University of Science and Technology, ChinaCopyright © 2025 Ma, Chen, Mu, Cao and Zhang. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Yan Zhang, cm0wMDIxMjVAd2h1LmVkdS5jbg==
†These authors have contributed equally to this work
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.