
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Immunol. , 27 February 2025
Sec. Vaccines and Molecular Therapeutics
Volume 16 - 2025 | https://doi.org/10.3389/fimmu.2025.1534787
This article is part of the Research Topic Immunological Regulation to Enteroviruses and Respiratory Viruses: Infection and Vaccination Responses View all articles
 Ho Yu Ng1†
Ho Yu Ng1† Yunshi Liao2†
Yunshi Liao2† Ching Lung Cheung3,4
Ching Lung Cheung3,4 Ruiqi Zhang5
Ruiqi Zhang5 Kwok Hung Chan6
Kwok Hung Chan6 Wai-Kay Seto5
Wai-Kay Seto5 Wai K. Leung5
Wai K. Leung5 Ivan F. N. Hung5*
Ivan F. N. Hung5* Tommy T. Y. Lam2,4,7*
Tommy T. Y. Lam2,4,7* Ka Shing Cheung5*
Ka Shing Cheung5*Introduction: BNT162b2 immunogenicity wanes with time and we investigated association between gut microbiota and longer-term immunogenicity.
Methods: This cohort study prospectively recruited adult BNT162b2 two-dose recipients from three vaccination centers in Hong Kong. Blood samples were collected at baseline and day 180 after first dose, and tested for neutralizing antibodies (NAb) against receptor-binding domain (RBD) of wild type SARS-CoV-2 virus using chemiluminescence immunoassay. Shotgun DNA metagenomic sequencing was performed to characterize baseline stool microbiome. Baseline metabolites were measured by gas and liquid chromatography-tandem mass spectrometry (GC-MS/MS and LC-MS/MS). Primary outcome was persistent high NAb response (defined as top 25% of NAb level) at day 180. Putative bacterial species and metabolic pathways were identified using linear discriminant analysis [LDA] effect size analysis. Multivariable logistic regression adjusting for clinical factors was used to derive adjusted odds ratio (aOR) of outcome with bacterial species and metabolites.
Results: Of 242 subjects (median age: 50.2 years [IQR:42.5-55.6]; male:85 [35.1%]), 61 (25.2%) were high-responders while 33 (13.6%) were extreme-high responders (defined as NAb≥200AU/mL). None had COVID-19 at end of study. Ruminococcus bicirculans (log10LDA score=3.65), Parasutterella excrementihominis (score=2.82) and Streptococcus salivarius (score=2.31) were enriched in high-responders, while Bacteroides thetaiotaomicron was enriched in low-responders (score=-3.70). On multivariable analysis, bacterial species (R. bicirculans–aOR: 1.87, 95% CI: 1.02-3.51; P. excrementihominis–aOR: 2.2, 95% CI: 1.18-4.18; S. salivarius–aOR: 2.09, 95% CI: 1.13-3.94) but not clinical factors associated with high response. R. bicirculans positively correlated with most metabolic pathways enriched in high-responders, including superpathway of L-cysteine biosynthesis (score=2.25) and L-isoleucine biosynthesis I pathway (score=2.16) known to benefit immune system. Baseline serum butyrate (aOR:10.00, 95% CI:1.81-107.2) and isoleucine (aOR:1.17, 95% CI:1.04-1.35) significantly associated with extreme-high vaccine response.
Conclusion: Certain gut bacterial species, metabolic pathways and metabolites associate with longer-term COVID-19 vaccine immunogenicity.
Studies showed that there was a steady decline of antibody levels among COVID-19 vaccine recipients (1, 2), and vaccine effectiveness against infection decreased significantly from 92% at 15-30 days to 47% at 121-180 days (3). The durability of vaccine immunogenicity is in large contributed by the persistence of antibody levels, which in turn is affected by the amount of long-lived plasma cells that secrete the antibodies, and also by memory B cells which are responsible for secondary immune response during re-exposure to the pathogen (4, 5). Nevertheless, factors influencing the durability of vaccine immunogenicity are currently underinvestigated. While certain factors, such as age, comorbidities (such as obesity and diabetes mellitus) and prior infection were more established in affecting vaccine durability (6), the role of the gut microbiome in vaccine durability is relatively less understood and explored.
Gut microbiota modulates immune response toward various vaccines including influenza and oral rotavirus (7), possibly by secreting immunostimulatory short chain fatty acids (SCFAs) (8), secondary bile acids (9), lipopolysaccharides (LPS) (10), flagellin (11), and peptidoglycans (12). Notably, a randomized controlled trial (RCT) showed that antibiotic-induced gut microbiota dysbiosis attenuated the level of influenza vaccine-induced antibodies, increased inflammatory signaling and disturbed plasma metabolome (9). Moreover, it has been suggested that gut microbiota was linked to memory B cells differentiation, possibly through influencing the formation of germinal centers in the gut which was important for B cell differentiation into memory B cells (13).
Emerging evidence has reported the potential role of gut microbiota in COVID-19 vaccine immunogenicity (Supplementary Table 1) (14). For instance, two studies of mostly non-immunocompromised subjects reported that enrichment of certain bacterial species associated with higher antibody level, such as Eubacterium rectale and Roseburia faecis in one (15), and Collinsella aerofaciens, Fusicatenibacter saccharivorans in another (16). In another study conducted in infliximab-treated inflammatory bowel disease (IBD) patients, Bilophila was associated with above average immune response after second dose of vaccination with BNT162b2 or ChAdOx1 (17). However, these studies measured antibody response within a relatively short time frame from vaccination (15–18). Moreover, some had small sample sizes (17–19) and might even be focused on immunocompromised patient groups that did not reflect the healthy general population (17, 18). Some employed 16S rRNA sequencing which had lower resolution than shotgun metagenomic sequencing (17–19). Factors that might affect the microbiota, such as diet, were also often not adjusted (15–20). Therefore, the potential role of gut microbiota in persistence of longer-term immunogenicity toward COVID-19 vaccine in healthy adult subjects deserves further investigation.
We conducted this prospective cohort study to investigate association between gut microbiota composition and BNT162b2 immunogenicity in immunocompetent adults at 6 months after vaccination.
This was a prospective cohort study recruiting adult subjects receiving two doses of BNT162b2 vaccines containing mRNA encoding viral spike (S) protein of SARS-CoV-2 from three vaccination centers in Hong Kong (Sun Yat Sen Memorial Park Sports Centre, Ap Lei Chau HKU Vaccination Centre and Queen Mary Hospital) between May and August 2021. Exclusion criteria included age <18 years, IBD, immunocompromised status including post-transplantation and immunosuppressives/chemotherapy, other medical diseases (cancer, hematological, rheumatological and autoimmune diseases) and those with prior COVID-19 identified from history or the presence of antibodies to SARS-CoV-2 nucleocapsid (N) protein. This study was approved by the Institutional Review Board (IRB) of the University of Hong Kong and Hospital Authority Hong Kong West Cluster (UW 21-216). All participants provided written informed consent for participation in this study.
Basic demographics including age, sex, lifestyle factors (level of exercises, diet and smoking), overweight or obese (OWOB) (21–23), diabetes mellitus (DM) (21), hepatic steatosis (24, 25) as well as any use of antibiotics (26) and proton pump inhibitors (PPIs) within six months before vaccination were collected. Adequate level of exercises is defined as meeting WHO recommendation of at least 150 to 300 minutes of moderate-intensity aerobic physical activity per week, or at least 75 to 150 minutes of vigorous-intensity physical activity per week (27). Diet quality is assessed using the rapid Prime Diet Quality Score (rPDQS), which is a validated diet quality screener with a score ranging from 0 to 52, and a higher score reflecting better diet quality (28). We created tertile groups, and compared the highest tertile with the lowest/middle tertile similar to other studies (29).
Subjects received two doses of intramuscular BNT162b2 (0.3mL) 3 weeks apart as recommended by local health authority. Blood samples were collected (i) before vaccination (baseline) and (ii) at 180days after first dose. Baseline stool samples were collected in OMNIgene tube before vaccine administration and stored at -80°C until total genomic DNA was extracted from them using Qiagen QIAamp DNA stool Mini Kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions. Genomic DNA was then subjected to library preparation for shotgun metagenomic sequencing using Nextera DNA Library Prep Kit (Illumina, California, USA). In brief, genomic DNA was first fragmented and tagged with adapter sequences by engineered transposome. Index adapter sequences were then added to these tagged DNA using limited cycle PCR. After amplification, PCR amplicons were purified using AMPure XP beads (Beckman-Coulter). Quality of the DNA library was first assessed by a Qubit fluorometer (Thermo Fisher Scientific), then by a Bioanalyzer (Agilent Technologies). After library preparation, next-generation shotgun metagenomic sequencing was performed by the Illumina NovaSeq 6000 platform (Illumina, San Diego, US) running at paired-end 150 bp, resulting in 10 Gb raw data per sample.
We employed a cost-effective extreme case-control design in selecting the extreme high responders (defined as NAb > 200 AU/mL) and extreme low responders (defined as NAb < 50 AU/mL) (30), and performed targeted metabolomics on selected metabolites, which were SCFAs (acetate, propionate and butyrate) due to more evidence in association with vaccine immunogenicity from literature, as well as L-isoleucine which was implicated in our metabolic pathway analysis. Extreme case-control design utilizes an extreme-value sampling design approach to select individuals with extremely large or small values of the primary outcome for exposure data collection (30). It has been shown that such an approach with appropriate statistical analysis could control type I error well and achieve cost-effectiveness (30, 31). Furthermore, the main purpose of this design was to exclude participants with highly similar NAb levels in the intermediate range which would introduce noise into our analysis. Therefore, we selected an arbitrary cut-off of NAb ≥200 AU/mL and NAb<50AU/mL to only include participants who were in the extreme-high range and extreme-low range into analysis to further enhance the associations detected (32). Baseline serum SCFAs (acetate, propionate and butyrate) as well as L-isoleucine were measured using gas chromatography-tandem mass spectrometry (GC-MS/MS) and liquid chromatography-tandem mass spectrometry (LC-MS/MS), respectively. Detailed steps of GC-MS/MS and LC-MS/MS can be found in Supplementary File.
Vaccine immunogenicity was determined in terms of neutralizing antibody (NAbs) against SARS-CoV-2 receptor-binding domain (RBD). NAb level is a surrogate marker of vaccine effectiveness (33) that predicts protection from symptomatic COVID-19 (34, 35). Although the gold standard to measure NAb level is live virus microneutralization assay (vMN), this test must be conducted under biosafety level-3 containment and therefore is not widely applicable to daily clinical practice, in particular during period of infection outbreak. Our previous study showed that a surrogate NAb assay, performed using the new version of the iFlash-2019-nCoV NAb kit (chemiluminescent microparticle immunoassay; Shenzhen YHLO Biotech Co, Ltd., Shenzhen, China), had good diagnostic performance (sensitivity: 98%, specificity: 95%, positive predictive value: 98% and negative predictive value: 94%) and agreement of 94% relative to vMN assay (36).
In the current study, testing for NAb was performed using iFlash-2019-nCoV NAb kit. Briefly, serum samples and a reagent pack with 2019-nCoV RBD antigen (30KD)-coated paramagnetic microparticles and acridinium ester-labeled ACE2 conjugate were placed in sample loading area and reagent loading area respectively (36). The iFlash system then automatically performed all functions and measured signals elicited from chemiluminescent reactions. NAb seropositivity was defined as ≥15 AU/mL.
Primary outcome of interest was persistent high NAb response at day 180. We defined top 25% of NAb (i.e. above 75 percentile) as high NAb response similar to the study by Tang et al. (16) NAb seropositivity was not chosen as primary outcome because 95.9% of the cohort remained seropositive at day 180.
Raw NGS reads were processed by fastp v0.20.1 (37) to quality and adapter trimming to remove sequencing adapters and bases with poor quality. Trimmed reads were subjected to host sequence removal by Bowtie2 (38) to map reads against human reference genome GRCh38.p13. Composition of microbial communities at species level and functional profile in each sample were inferred from the cleaned reads using MetaPhlAn (v3.0) (39) and HUMAnN (v3.0) (40) respectively. Estimation of species coverage and relative abundance was determined. Low-abundance taxa were not excluded from analysis as it has been shown that low-abundance bacteria could contribute substantially to host phenotypes (41) and there is no consensus in the analytical approach as to which levels filtering methods should be applied to remove low-abundance taxa, which would influence the results of downstream analyses (42). Instead, we incorporated negative control sample into sequencing to make sure the inferred microbiome taxa were not derived from contaminants introduced during sampling, lab work, or sequencing.
All statistical analyses were conducted with R version 4.2.2 (R Foundation for Statistical Computing, Vienna, Austria) statistical software. Data was displayed as median (interquartile range [IQR]) for continuous variables, and as number of patients (percentage) for categorical variables. The Mann-Whitney U test and the Chi-square test or Fisher exact test was used for two continuous variables and categorical variables respectively.
Alpha-diversity in terms of observed species, Shannon and Simpson index was computed using vegan package in R Studio, and compared using Wilcoxon signed-rank test. Beta-diversity including Bray-Curtis compositional dissimilarity was compared using non-metric multidimensional scaling (NMDS). Permutational multivariate analysis of variance (PERMANOVA) was used to compare microbial communities of different samples. Putative gut bacterial species and metabolic pathways with an absolute value of linear discriminant analysis (LDA) score ≥ 2 were identified using LEfSe (linear discriminant analysis effect size). The median was used to define a high relative abundance of a particular bacterial species.
A univariate logistic and linear regression model was used to estimate odds ratio (OR) and beta coefficients, respectively, of high-response with the various aforementioned clinical factors and with a high relative abundance of putative gut bacterial species. A multivariable logistic regression model was used to estimate adjusted OR (aOR) of high response with clinical factors and putative bacterial species of p<0.15 on univariate logistic regression analysis as in previous study (43). The performance of the variables with p<0.15 in either univariate logistic regression or univariate linear regression in predicting vaccine response was assessed by random forest machine learning model to derive the area under receiver operating characteristic curve (AUC). Details of the machine learning model can be found in Supplementary Methods.
To explore the relationship between DM and P. excrementihominis and vaccine immunogenicity, we stratified the subjects based on the baseline relative abundance of P. excrementihominis and DM status, and assigned four groups (High abundance-Non-DM, High abundance-DM, Low abundance-Non-DM, Low abundance-DM) to perform trend test (ptrend). Similarly, to explore the relationship between OWOB and P. excrementihominis and vaccine immunogenicity, we stratified the subjects based on the baseline relative abundance of P. excrementihominis and OWOB status, and assigned four groups (High abundance-Normal weight, High abundance-OWOB, Low abundance-Normal weight, Low abundance-OWOB) to perform trend test (ptrend).
Spearman’s correlation tests were used to analyze correlation among continuous variables. False discovery rate (FDR) was used to correct for multiple comparisons in multiple hypothesis testing, including during LefSe analysis (44).
For the analysis of metabolites, multivariable logistic regression was used to evaluate the association of metabolites with extreme high response. To achieve an alpha value of 0.05 and a power of at least 80% to detect an association if the metabolite has an odds ratio of at least 1.5, the sample size will be 70 for the extreme case-control design.
A two-sided p-value ≤0.05 was considered as statistically significant, while a p-value ≤ 0.1 was considered borderline significant.
We recruited 242 eligible adults who had received two doses of BNT162b2. The median age was 50.2 years (IQR:42.5-55.6; range 18-75), and 85 (35.1%) were male. 232 (95.9%) remained seropositive (≥15 AU/mL) at day 180 (median NAb level:55.2 AU/mL; IQR:30.8-123.6). 61 (25.2%) were classified as persistent high-responders (median NAb level:205.4 AU/mL; IQR:164.6-291.3) while 181 (74.8%) were low-responders (median NAb level:40.9 AU/mL; IQR:25.8-66.4). There were 119 (51.1%) subjects who had adequate level of exercises as per WHO recommendations, and 97 (41.6%) had rPDQS at top tertile. Additionally, there were 134 (55.4%) subjects who were OWOB, and 16 (6.6%) had DM. Baseline characteristics were comparable between the high- and low-responders (all p>0.05) (Table 1). At day 180, none had SARS-CoV-2 after they received two doses of BNT162b2.
On univariate logistic regression, age ≥ 55 years old and DM were negatively associated with persistent high vaccine response with borderline statistical significance (Table 2). On univariate linear regression, OWOB and DM were negatively associated with persistent high vaccine response with borderline statistical significance (Supplementary Table 2). Other factors including lipid profile, lifestyle factors (level of exercises, diet and smoking), and antibiotic use were not significantly associated with persistent high vaccine response on both univariate logistic and univariate linear regression (Table 2, Supplementary Table 2).
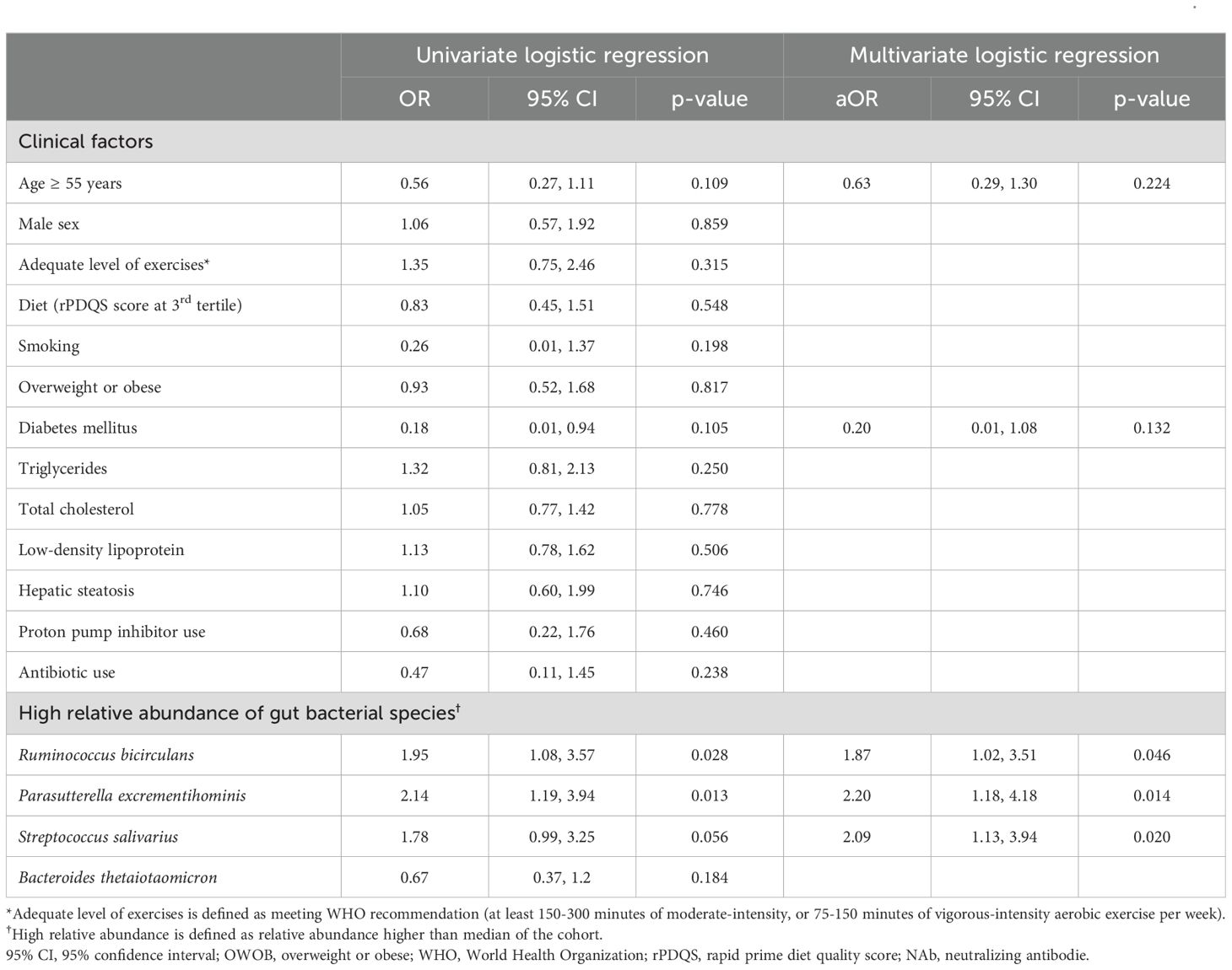
Table 2. Univariate and multivariate logistic regression between high-vaccine response and a combination of clinical factors and bacterial species.
There was no significant difference in alpha diversity (richness, Shannon and Simpson all p>0.05; Supplementary Figure 1) and beta diversity (PERMANOVA analysis, p=0.270; Supplementary Figure 2) between high- and low-responders.
Using LEfSe analysis, we found that five bacterial species (namely Ruminococcus bicirculans, Bacteroides intestinalis, Parasutterella excrementihominis, Streptococcus salivarius, and Prevotella sp CAG 891) and four (namely Prevotella sp CAG 279, Parabacteroides sp CAG 409, Bacteroides fragilis, and Bacteroides thetaiotaomicron) were enriched in high- and low-responders in baseline gut microbiome respectively (Figure 1). Among them, R. bicirculans, P. excrementihominis, S. salivarius and B. thetaiotaomicron were not zero-inflated (i.e. their median relative abundance was not equal to zero). Specifically, R. bicirculans (log10LDA score=3.65; 0.9% vs 0.1%; p=0.031), P. excrementihominis (log10LDA score=2.82; 0.08% vs 0.02%; p=0.040) and S. salivarius (log10LDA score=2.31; 0.06% vs 0.04%; p=0.031) were enriched in persistent high-responders, while B. thetaiotaomicron was enriched in low-responders (log10LDA score=-3.70; 2.1% vs 1.1%; p=0.040) (Figures 1A, B).
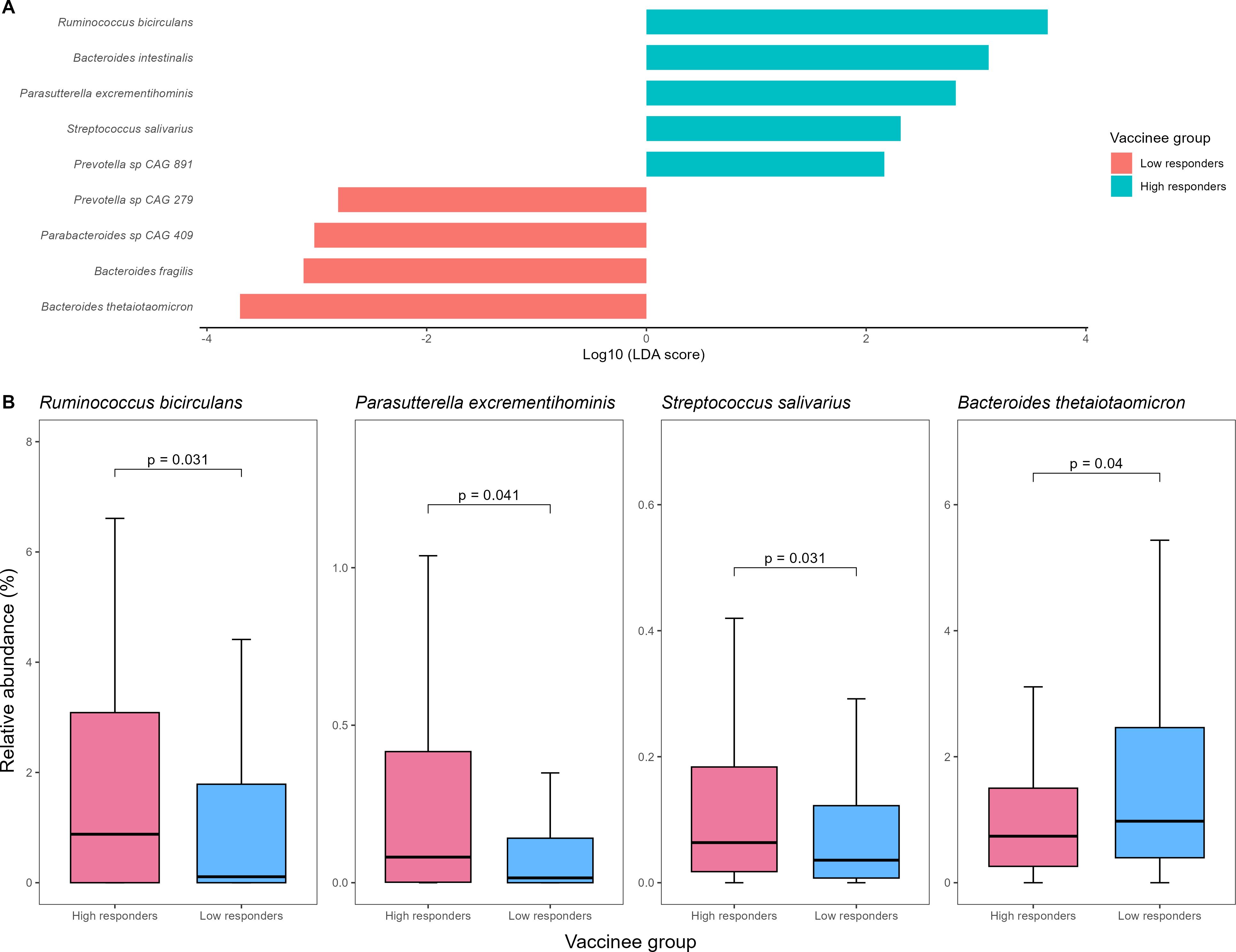
Figure 1. (A) Baseline gut bacterial species enriched in high- vs low-vaccine responders detected by LEfSe (B) Comparison of relative abundances of putative baseline gut bacterial species identified on LEfSe analysis between high- and low-vaccine responders. LEfSe, linear discriminant analysis effect size.
On univariate logistic regression, a higher abundance of R. bicirculans (OR:1.95; 95% CI:1.08-3.57), P. excrementihominis (OR:2.14; 95% CI:1.19-3.94) and S. salivarius (OR:1.78; 95% CI:0.99-3.25) was associated with high-response (all p<0.10) (Table 2).
We then incorporated clinical (age ≥ 55 years and DM) and the three bacterial species factors with p<0.15 on univariate logistic regression into multivariate logistic regression. Only bacterial species (R. bicirculans–aOR:1.87, 95% CI:1.02-3.51; P. excrementihominis–aOR:2.2, 95% CI:1.18-4.18; S. salivarius–aOR:2.09, 95% CI:1.13-3.94) but not clinical factors (age ≥55 years–aOR:0.63, 95% CI:0.29-1.30; DM–0.20, 95% CI:0.01-1.08) were associated with persistent high-response (Table 2). Sensitivity analysis incorporating OWOB also yielded similar results (Supplementary Table 3). The clinical factors (age ≥ 55 years, DM and OWOB) and the and the three bacterial species factors with p<0.15 on either univariate logistic regression or univariate linear regression collectively predict vaccine response with AUC of 0.712 by random forest machine learning model (Supplementary Figure 3).
As DM and OWOB were potential factors affecting vaccine immunogenicity on univariate analysis, we further investigated the relative abundances of bacterial markers in patient cohorts, as well as the role of these metabolic factors as an effect modifier of bacteria-immune response relationship. Supplementary Figure 4 shows that there was no significant difference in relative abundances of bacterial markers between non-DM and DM subjects as well as non-OWOB subjects and OWOB subjects, except for R. bicirculans which was significantly more abundant in non-DM subjects than DM subjects (0.27% vs 0%;p=0.044). Non-DM subjects with a high relative abundance of P. excrementihominis had significantly higher median NAb level than non-DM subjects with low abundance of P. excrementihominis (81.7 [IQR:37.8-165.0] vs 45.1 AU/mL [IQR:27.6-91.6];p=0.003) (Supplementary Figure 5A). When stratified by baseline relative abundance of P. excrementihominis and DM status, and taking high abundance-non DM as reference, there was a decreasing trend of NAb level at day 180 (p-trend=0.022, Supplementary Table 4). Similarly, non-OWOB subjects with high relative abundance of this species compared with non-OWOB subjects with low abundance of this species (101.0 [IQR:46.5-165.0] vs 42.7 AU/mL [IQR:26.8-91.6];p=0.009) (Supplementary Figure 6A). When stratified by baseline relative abundance of P. excrementihominis and OWOB status, and taking high abundance-normal weight as reference, there was a significant decreasing trend of NAb level at day 180 (p-trend=0.009, Supplementary Table 5).
We then considered having high relative abundance of P. excrementihominis, non-OWOB and non-DM as favorable factors, and found that subjects who had all three favorable factors had a significantly higher median NAb level (106.0 AU/mL [IQR:48.3-167.0]) than those with just two (55.1 [IQR:28.9-122.0];p=0.015), one (46.0 AU/mL [IQR:28.6-81.9];p=0.007) or no protective factors (20.6 AU/mL [IQR:19.7-35.0];p=0.014) (Supplementary Figure 7).
On the other hand, there were no similar observations among other three putative bacterial markers with either DM or OWOB on NAb level (Supplementary Figures 5B–D and Supplementary Figures 6B–D). While non-DM subjects with high relative abundance of S. salivarius had significantly higher median NAb level than DM subjects with low relative abundance of this species (60.0 [IQR:30.8-160.0] vs 25.9 AU/mL [IQR:19.7-35.2];p=0.026), there was no similar observations in non-OWOB subjects with high relative abundance of this species compared with OWOB subjects with low relative abundance of this species.
Additionally, we conducted subgroup analysis based on sex, and found that in male subjects, high abundance of P. excrementihominis were associated with high vaccine response at day 180 (aOR=4.59, 95% CI: 1.51-15.70, p=0.010) (Supplementary Table 6). On the other hand, in female subjects, high abundance of R. bicirculans (aOR=2.36, 95% CI: 1.09-5.35, p=0.034) and S. salivarius (aOR=2.29, 95% CI: 1.07-5.03, p=0.035) were associated with high vaccine response at day 180 (Supplementary Table 6).
We then investigated the metabolic pathways in baseline gut microbiome and found that six and five pathways were enriched in high- and low-responders respectively (Supplementary Figure 8). They were classified into “Biosynthesis” and “Degradation/Utilization/Assimilation” categories according to MetaCyc database (Supplementary Table 7). In persistent high-responders, enriched pathways included those related to energy production as well as amino acid biosynthesis, such as superpathway of L-cysteine biosynthesis (mammalian) (log10LDA score=2.25;p=0.005) and superpathway of L-isoleucine biosynthesis I (log10LDA score=2.16;p=0.020).
We performed Spearman’s correlation analysis between baseline gut bacterial species and metabolic pathways (Figure 2). Most of the metabolic pathways enriched in high-responders were positively associated with R. bicirculans. Notably, superpathway of L-cysteine biosynthesis (mammalian) and superpathway of L-isoleucine biosynthesis I showed positive correlation with R. bicirculans (Spearman’s r=0.45;p<0.001 and r=0.41;p<0.001 respectively) and S. salivarius (r=0.15;p=0.069 and r=0.21;p=0.007 respectively).
Thirty-three and 37 subjects were classified as extreme-high responders and extreme-low responders, respectively. Among them, 29 (41.4%) were male, and the median age was 50.2 years old (IQR:40.1-53.8). For SCFAs, butyrate was significantly associated with extreme high vaccine response (aOR:10.00, 95% CI:1.81-107.2; p=0.025), while propionate was of borderline significance (aOR:1.33, 95% CI: 0.97-1.92; p=0.091) and acetate did not show significant association (aOR:1.01, 95% CI:0.99-1.02; p=0.368). Baseline serum isoleucine was also significantly associated with extreme high vaccine response (aOR:1.17, 95% CI:1.04-1.35; p=0.016).
To our knowledge, our study was the first to show the association of baseline gut microbiota composition with persistently high immunogenicity toward BNT162b2 at 180 days post-vaccination. We identified four potential baseline microbial markers, namely R. bicirculans, P. excrementihominis, S. salivarius and B. thetaiotaomicron, as well as metabolic pathway markers that might predict longer-term vaccine immunogenicity. In particular, R. bicirculans positively correlated with most of the metabolic pathways enriched in persistent high-responders, highlighting it as a potential key bacterial species that might be beneficial to the immune system.
Gut microbiota is involved in modulating immune response toward vaccination via production of immunomodulatory metabolites. Some of these metabolites served as vaccine adjuvants through activating PRRs such as toll-like receptors (TLRs) or NOD-like receptors. TLR5-mediated sensing of flagellin enhances the presence of short-lived plasma cells and immune response to influenza vaccine (11). TLR-4 mediated sensing of bacterial LPS promotes type 1 T helper cells (Th1) and antibody production (10), while NOD2-mediated recognition of bacterial peptidoglycan contributes to the mucosal adjuvant activity of cholera toxin (12). Other key metabolites include short-chain fatty acids (SCFAs) and secondary bile acids. The former increases antibody production through promoting energy production in B cells (8), while the latter was negatively correlated with inflammatory signatures following influenza vaccination (9).
Additionally, the gut microbiota was implicated in promoting memory B cell formation, which is important for antibody durability. In mouse models, correction of gut microbiota dysbiosis were able to promote germinal center formation, which was required for memory B cell differentiation (45). Moreover, bacterial LPS could activate TLR4, which has been shown to enhance the persistence of germinal centers and early differentiation into long-lived memory cells in rhesus macaques (46). In humans, it has been shown that early colonization of microbiota such as Bifidobacteria in infants was associated with future increased frequency of CD27+ memory B cells (47, 48). Conversely, babies born to mothers with IBD were found to be depleted in Bifidobacteria, which was associated with fewer class-switched memory B cells in later timepoints (49). These suggest that the gut microbiota may have a role to play in promoting durable vaccine immunogenicity, which was largely contributed by memory B cell function.
On multivariable analysis, high abundance of R. bicirculans, P. excrementihominis and S. salivarius remained predictive of persistently high vaccine immunogenicity, while the clinical factors (age ≥ 55 years old and DM) were not. This suggests that high abundance of R. bicirculans, P. excrementihominis and S. salivarius in stool samples is a more important factor than routine clinical parameters in predicting vaccine immunogenicity. Consistently, the significant bacterial species identified from LEfSe analysis in our current study are capable of producing these immunomodulatory metabolites (Figure 3). Our metabolomics analysis also showed that baseline butyrate and propionate levels were associated with extreme high vaccine response at day 180 with statistical significance and borderline significance, respectively.
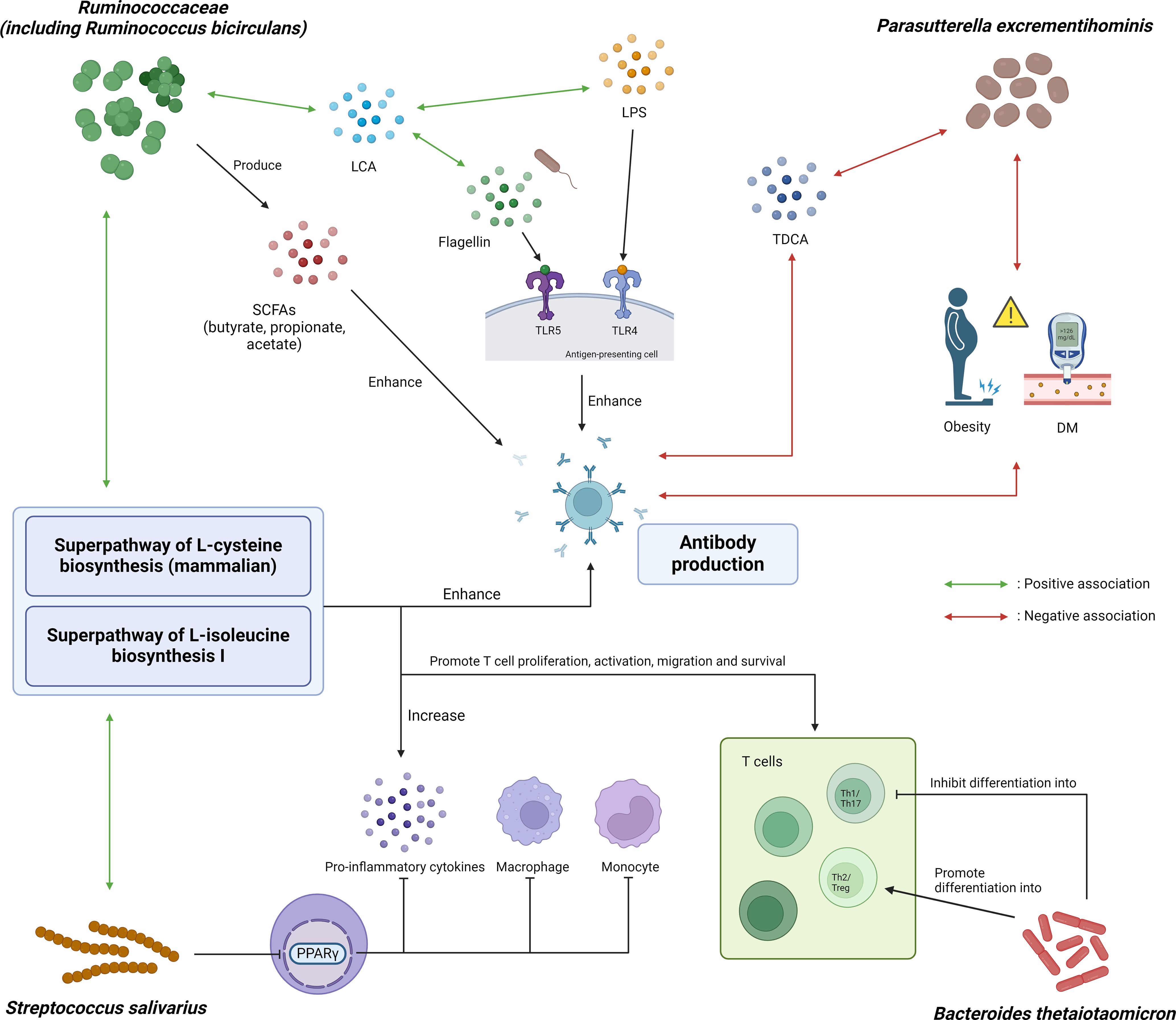
Figure 3. Graphical illustration of potential mechanistic links between microbial and metabolic markers with COVID-19 vaccine immunogenicity. Ruminococcaceae, including Ruminococcus bicirculans, can produce SCFAs such as butyrate, propionate and acetate, which in turn can enhance in vivo polyclonal antibody production. It is also positively associated with fecal lithocholic acid (LCA), which was shown to have positive association with flagellin and lipopolysaccharide (LPS) content in the stool after influenza vaccination. Parasutterella excrementihominis was negatively associated with obesity, diabetes mellitus and taurodeoxycholic acid (TDCA), all of which were associated with poorer response to vaccines. Streptococcus salivarius can inhibit monocytes, macrophages and production of pro-inflammatory cytokines through inhibiting PPARγ. Bacteroides thetaiotaomicron inhibits differentiation into Th1 cells which was important for protection against SARS-CoV-2, thereby attenuating antibody response to COVID-19 vaccine. Additionally, superpathway of L-cysteine biosynthesis (mammalian) and superpathway of L-isoleucine biosynthesis I were positively correlated with R. bicirculans and S. salivarius. Cysteine and isoleucine boost the immune system by promoting T cell proliferation, activation and survival, as well as enhancing the production of pro-inflammatory cytokines as well as antibody production. SCFA, short-chain fatty acid; LCA, lithocholic acid; LPS, lipopolysaccharide; TDCA, taurodeoxycholic acid; DM, diabetes mellitus; PPARγ, peroxisome proliferator-activated receptor gamma; TLR, toll-like receptor; Th1/2/17, T helper 1/2/17 cells; Treg, regulatory T cells.
Ruminococcacecae produces SCFA including butyrate, propionate and acetate (50, 51). SCFAs can boost antibody production by stimulating expression of genes necessary for plasma cell differentiation, and increasing energy production through glycolysis, oxidative phosphorylation and fatty acid synthesis (8, 52). In particular, butyrate can also boost antibody production through upregulating follicular helper T cells which promote the activation and differentiation of plasma cells (52). Apart from producing acetate (53, 54), R. bicirculans is positively associated with the levels of fecal lithocholic acid (LCA) (55), which was positively associated with flagellin and LPS content in stool after influenza vaccination (9). P. excrementihominis was associated with a reduction in taurodeoxycholic acid (TDCA) in a mice study (56), which in turn was associated with poorer response to COVID-19 vaccination (including BNT162b2 vaccine) in infliximab-treated IBD patients (17). S. salivarius was shown to be able to inhibit peroxisome proliferator-activated receptor gamma (PPARγ) activation (57), which negatively regulates monocytes and macrophages and inhibits the production of pro-inflammatory cytokines (58).
On the other hand, B. thetaiotaomicron, which was enriched in low responders, may promote the preferential differentiation of anti-inflammatory Treg/Th2 cells while suppressing the relative differentiation of pro-inflammatory Th1/Th17 cells (59). Moreover, the polysaccharide A (PSA) of B. thetaiotaomicron promotes the function of Treg cells through interaction with toll-like receptor 2 (TLR2) (60). Th1 profile was favored following BNT162b2 vaccination and was important for protection against SARS-CoV-2 infection (61). Therefore, suppression of Th1 cells differentiation might attenuate the antibody response to BNT162b2. On the other hand, Treg cells suppresses B cell functions, including NAb production and immune memory (62). Conversely it has been shown in mouse models that depletion of Treg cells using anti-CD25 monoclonal antibody could induce more durable immunogenicity to BCG and hepatitis B vaccinations (63). Therefore, the preferential differentiation of Treg cells due to B. thetaiotaomicron might also have dampened the antibody response to BNT162b2.
Additionally, we found that superpathway of L-cysteine biosynthesis (mammalian) and superpathway of L-isoleucine biosynthesis I pathway were enriched in persistent high responders, and were positively correlated with R. bicirculans and S. salivarius (Figure 3). The metabolic activities of Ruminococcus bicirculans can produce H2S and acetyl-CoA (64), which provide materials and an energy source for L-cysteine biosynthesis through the superpathway of L-cysteine biosynthesis (fungi). Acetyl-CoA also contributes to the synthesis of L-isoleucine as a material, along with oxaloacetate, through the superpathway of L-isoleucine biosynthesis I. Extracellular cysteine was important for T cell proliferation, activation and survival (65). Cysteine transporters were strongly upregulated during T cell activation, and DNA synthesis in T cells was dependent on cysteine (66, 67). Extracellular cysteine could also affect the level of intracellular glutathione, as well as the activity of NFκB pathway which was involved in the secretion of inflammatory cytokines (68). Deficiency of extracellular cysteine was associated with immunodeficiency-related conditions, including acquired immune deficiency syndrome (AIDS) and a variety of cancers (65, 69). Isoleucine is a branched chain amino acid (BCAA) which was important for immune cell proliferation and function as well as production of pro-inflammatory cytokines (70, 71), and was greatly incorporated into lymphocytes (72). Isoleucine could induce the expression of β-defensins (70, 73), which was involved in activating IFN-γ (74). IFN-γ was in turn positively correlated with SARS-CoV-2 neutralizing antibody titers (75). In another study, isoleucine supplementation resulted in increased production of immunoglobulins, RV-specific antibodies and cytokines in the intestines and serum of RV-infected piglets (76), supporting its potential beneficial role in boosting immunity against viral infections. Our current study also showed that the baseline isoleucine was significantly associated with extreme high vaccine response at day 180.
We also investigated the interaction between significant bacterial species and metabolic factors with NAb level at six months. Interestingly, we found that different combinations of DM status, OWOB status and baseline abundance of P. excrementihominis were associated with different levels of vaccine response (Figure 3). Those subjects who were non-DM and non-OWOB with a higher relative abundance of P. excrementihominis had significantly higher NAb level than those with only one or even without these protective factors. Obesity (21–23) and DM (21) are associated with lower vaccine immunogenicity to COVID-19 vaccines (21). Parasutterella was shown to be negatively associated with obesity in both mice models (77) and in Chinese adults (78). Parasutterella was also found to be enriched in healthy pregnant women compared to those with gestational DM (79), and several RCTs found that these interventions were able to reverse gut dysbiosis in DM patients by increasing abundance of Parasutterella (80–83). This might explain the interactions among these three protective factors, and the synergistic effect they exert on vaccine response.
It should be noted that other studies have identified different bacterial markers that might also predict COVID-vaccine immunogenicity. The discrepancy in results might be explained by inherent variations in gut microbiota composition across different populations due to factors including diet, lifestyle and socioeconomic status (7), and also by heterogenous study designs in terms of sample size and population, vaccine type, sequencing method, as well as the timepoint of measuring vaccine immunogenicity. Existing studies that investigated the association of gut microbiota with COVID-19 vaccination are summarized in Supplementary Table 1. Of note, most of them measured antibody response within a relatively short time frame from vaccination, ranging from one to three months (15–18). Several studies also only had small sample size (17–19), while some focused in special patient populations, such as one conducted in infliximab-treated IBD patients (17), and another in people living with HIV (18). In addition, factors that might affect the microbiota, such as diet, were often not statistically accounted for as in our study (15–20). Moreover, some studies employed 16S rRNA gene sequencing instead of shotgun metagenomic sequencing, which had lower resolution and could not profile gut microbiota down to species and strain level (17–19). As a result, these studies found various different bacterial markers that were associated with high vaccine response. Among immunocompetent individuals, these included Eubacterium rectale and Roseburia faecis in BNT162b2 vaccinees (15), Bifidobacterium adolescentis in CoronaVac (inactivated virus vaccine) vaccinees (15), as well as Collinsella aerofaciens and Fusicatenibacter saccharivorans in BBIBP-CorV (inactivated virus vaccine) vaccinees (16). Different bacterial markers were observed in immunosuppressed individuals. In one study on IBD patients receiving infliximab for >12 weeks, Bilophila was found to be enriched in high-responders while Streptococcus was enriched in low responders after two doses of BNT162b2 or ChAdOx1 vaccination (17). Another study on people living with HIV found that Flavonifractor, Lachnospira and Oscillibacter were enriched in high-responders after two doses of BNT162b2, while Butyricimonas and Paraprevotella were enriched in low-responders (18).
Several limitations of our study should be noted. First, our study findings, including the metabolic pathways, were correlative, and further studies on animal models, such as in germ-free mice or in microbiome-modulated animal models (such as through supplementation with a bacteria strain), were required to prove causality behind the association between the gut microbiota, metabolites and vaccine immunogenicity observed in our study. Second, our study findings may not be generalizable to other populations as variation of gut microbiota composition exist across different populations and geographical regions due to various factors, including diet, lifestyle and socioeconomic status (7). Furthermore, the use of extreme case-control design may also have limited the generalizability of our metabolomic findings to other populations. Third, longer follow-up on the durability of immunogenicity after two vaccine doses was not possible as the majority of participants received booster dose after six months.
Nevertheless, we are planning to further investigate whether gut microbiota can predict vaccine immunogenicity and durability after booster doses. The markers identified in this study might potentially apply to immunogenicity after booster doses as well and explain difference in the speed of waning of antibody levels. Moreover, the findings of our study might be translated to clinical applications by developing probiotics and amino acid supplements, which would be validated in animal models and in clinical RCTs on human subjects, to boost vaccine immunogenicity in the general population. In fact, similar studies were done in the past. For instance, one RCT investigated the effect of probiotic supplementation with Limosilactobacillus reuteri DSM 17938 for 6 months on serum antibody levels after COVID-19 vaccination, and found that probiotic supplementation exhibited significantly higher serum antibody levels than placebo arm at >28 days after vaccination (84). Thus, we were hopeful that our study could help to lay the foundation for further animal studies or even RCTs to validate and investigate the potential gut bacterial species that were implicated in enhancing durable vaccine immunogenicity for potential therapeutic use in the future.
Certain gut bacterial species could be associated with persistently high vaccine immunogenicity at six months after two doses of BNT162b2. These results may potentially facilitate the development of gut microbiota interventions to improve long-term durability of vaccine immune response.
All sequencing data generated in this study are deposited in the Sequence Read Archive under BioProject PRJNA983517.
The studies involving humans were approved by the Institutional Review Board (IRB) of the University of Hong Kong and Hospital Authority Hong Kong West Cluster (UW 21-216). The studies were conducted in accordance with the local legislation and institutional requirements. The participants provided their written informed consent to participate in this study.
HN: Formal analysis, Visualization, Writing – original draft, Writing – review & editing. YL: Formal analysis, Writing – original draft, Writing – review & editing. CC: Formal analysis, Writing – original draft, Writing – review & editing. RZ: Investigation, Writing – review & editing. KC: Investigation, Writing – review & editing. WS: Supervision, Writing – review & editing. WL: Supervision, Writing – review & editing. IH: Supervision, Writing – review & editing. TL: Formal analysis, Writing – original draft, Writing – review & editing. KC: Conceptualization, Formal analysis, Funding acquisition, Methodology, Supervision, Writing – original draft, Writing – review & editing.
The author(s) declare that financial support was received for the research, authorship, and/or publication of this article. Health and Medical Research Fund, Food and Health Bureau, The Government of the Hong Kong Special Administrative Region (Reference no: COVID1903010, Project 16). InnoHK funding from the Innovation and Technology Commission (D24H, C2i).
Figure 3 was created with Biorender.com.
Authors CC and TL were employed by Laboratory of Data Discovery for Health Limited. Author TL was employed by Centre for Immunology & Infection Limited.
The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fimmu.2025.1534787/full#supplementary-material
1. Naaber P, Tserel L, Kangro K, Sepp E, Jurjenson V, Adamson A, et al. Dynamics of antibody response to BNT162b2 vaccine after six months: a longitudinal prospective study. Lancet Reg Health Eur. (2021) 10:100208. doi: 10.1016/j.lanepe.2021.100208
2. Skowronski DM, De Serres G. Safety and efficacy of the BNT162b2 mRNA covid-19 vaccine. N Engl J Med. (2021) 384:1576–7. doi: 10.1056/NEJMc2036242
3. Nordström P, Ballin M, Nordström A. Risk of infection, hospitalisation, and death up to 9 months after a second dose of COVID-19 vaccine: a retrospective, total population cohort study in Sweden. Lancet. (2022) 399(10327):814–23. doi: 10.1016/S0140-6736(22)00089-7
4. Yoshida T, Mei H, Dorner T, Hiepe F, Radbruch A, Fillatreau S, et al. Memory B and memory plasma cells. Immunol Rev. (2010) 237:117–39. doi: 10.1111/j.1600-065X.2010.00938.x
5. Nguyen DC, Joyner CJ, Sanz I, Lee FE. Factors affecting early antibody secreting cell maturation into long-lived plasma cells. Front Immunol. (2019) 10:2138. doi: 10.3389/fimmu.2019.02138
6. Berber E, Ross TM. Factors predicting COVID-19 vaccine effectiveness and longevity of humoral immune responses. Vaccines (Basel). (2024) 12:1284. doi: 10.3390/vaccines12111284
7. Lynn DJ, Benson SC, Lynn MA, Pulendran B. Modulation of immune responses to vaccination by the microbiota: implications and potential mechanisms. Nat Rev Immunol. (2022) 22:33–46. doi: 10.1038/s41577-021-00554-7
8. Kim M, Qie Y, Park J, Kim CH. Gut microbial metabolites fuel host antibody responses. Cell Host Microbe. (2016) 20:202–14. doi: 10.1016/j.chom.2016.07.001
9. Hagan T, Cortese M, Rouphael N, Boudreau C, Linde C, Maddur MS, et al. Antibiotics-driven gut microbiome perturbation alters immunity to vaccines in humans. Cell. (2019) 178:1313–1328 e13. doi: 10.1016/j.cell.2019.08.010
10. Georg P, Sander LE. Innate sensors that regulate vaccine responses. Curr Opin Immunol. (2019) 59:31–41. doi: 10.1016/j.coi.2019.02.006
11. Oh JZ, Ravindran R, Chassaing B, Carvalho FA, Maddur MS, Bower M, et al. TLR5-mediated sensing of gut microbiota is necessary for antibody responses to seasonal influenza vaccination. Immunity. (2014) 41:478–92. doi: 10.1016/j.immuni.2014.08.009
12. Kim D, Kim YG, Seo SU, Kim DJ, Kamada N, Prescott D, et al. Nod2-mediated recognition of the microbiota is critical for mucosal adjuvant activity of cholera toxin. Nat Med. (2016) 22:524–30. doi: 10.1038/nm.4075
13. Yu B, Wang L, Chu Y. Gut microbiota shape B cell in health and disease settings. J Leukoc Biol. (2021) 110:271–81. doi: 10.1002/JLB.1MR0321-660R
14. Ng HY, Leung WK, Cheung KS. Association between gut microbiota and SARS-coV-2 infection and vaccine immunogenicity. Microorganisms. (2023) 11:452. doi: 10.3390/microorganisms11020452
15. Ng SC, Peng Y, Zhang L, Mok CK, Zhao S, Li A, et al. Gut microbiota composition is associated with SARS-CoV-2 vaccine immunogenicity and adverse events. Gut. (2022) 71:1106–16. doi: 10.1136/gutjnl-2021-326563
16. Tang B, Tang L, He W, Jiang X, Hu C, Li Y, et al. Correlation of gut microbiota and metabolic functions with the antibody response to the BBIBP-CorV vaccine. Cell Rep Med. (2022) 3:100752. doi: 10.1016/j.xcrm.2022.100752
17. Alexander JL, Mullish BH, Danckert NP, Liu Z, Olbei ML, Saifuddin A, et al. The gut microbiota and metabolome are associated with diminished COVID-19 vaccine-induced antibody responses in immunosuppressed inflammatory bowel disease patients. EBioMedicine. (2023) 88:104430. doi: 10.1016/j.ebiom.2022.104430
18. Ray S, Narayanan A, Vesterbacka J, Blennow O, Chen P, Gao Y, et al. Impact of the gut microbiome on immunological responses to COVID-19 vaccination in healthy controls and people living with HIV. NPJ Biofilms Microbiomes. (2023) 9:104. doi: 10.1038/s41522-023-00461-w
19. Seong H, Yoon JG, Nham E, Choi YJ, Noh JY, Cheong HJ, et al. The gut microbiota modifies antibody durability and booster responses after SARS-CoV-2 vaccination. J Transl Med. (2024) 22:827. doi: 10.1186/s12967-024-05637-2
20. Peng Y, Zhang L, Mok CKP, Ching JYL, Zhao S, Wong MKL, et al. Baseline gut microbiota and metabolome predict durable immunogenicity to SARS-CoV-2 vaccines. Signal Transduct Target Ther. (2023) 8:373. doi: 10.1038/s41392-023-01629-8
21. Boroumand AB, Forouhi M, Karimi F, Moghadam AS, Naeini LG, Kokabian P, et al. Immunogenicity of COVID-19 vaccines in patients with diabetes mellitus: A systematic review. Front Immunol. (2022) 13:940357. doi: 10.3389/fimmu.2022.940357
22. Faizo AA, Qashqari FS, El-Kafrawy SA, Barasheed O, Almashjary MN, Alfelali M, et al. A potential association between obesity and reduced effectiveness of COVID-19 vaccine-induced neutralizing humoral immunity. J Med Virol. (2023) 95:e28130. doi: 10.1002/jmv.28130
23. Piernas C, Patone M, Astbury NM, Gao M, Sheikh A, Khunti K, et al. Associations of BMI with COVID-19 vaccine uptake, vaccine effectiveness, and risk of severe COVID-19 outcomes after vaccination in England: a population-based cohort study. Lancet Diabetes Endocrinol. (2022) 10:571–80. doi: 10.1016/S2213-8587(22)00158-9
24. Cheung KS, Lam LK, Hui RWH, Mao X, Zhang RR, Chan KH, et al. Effect of moderate-to-severe hepatic steatosis on neutralising antibody response among BNT162b2 and CoronaVac recipients. Clin Mol Hepatol. (2022) 28:553–64. doi: 10.3350/cmh.2022.0082
25. Cheung KS, Lam LK, Mao X, Tan JT, Ooi PH, Zhang R, et al. Effect of Moderate to Severe Hepatic Steatosis on Vaccine Immunogenicity against Wild-Type and Mutant Virus and COVID-19 Infection among BNT162b2 Recipients. Vaccines (Basel). (2023) 11:497. doi: 10.3390/vaccines11030497
26. Cheung KS, Lam LK, Zhang R, Ooi PH, Tan JT, To WP, et al. Association between Recent Usage of Antibiotics and Immunogenicity within Six Months after COVID-19 Vaccination. Vaccines (Basel). (2022) 10:1122. doi: 10.3390/vaccines10071122
27. Bull FC, Al-Ansari SS, Biddle S, Borodulin K, Buman MP, Cardon G, et al. World Health Organization 2020 guidelines on physical activity and sedentary behaviour. Br J Sports Med. (2020) 54:1451–62. doi: 10.1136/bjsports-2020-102955
28. Kronsteiner-Gicevic S, Tello M, Lincoln LE, Kondo JK, Naidoo U, Fung TT, et al. Validation of the rapid prime diet quality score screener (rPDQS), A brief dietary assessment tool with simple traffic light scoring. J Acad Nutr Diet. (2023) 123:1541–1554.e7. doi: 10.1016/j.jand.2023.05.023
29. Yang J, Wang M, Tobias DK, Rich-Edwards JW, Darling AM, Abioye AI, et al. Dietary diversity and diet quality with gestational weight gain and adverse birth outcomes, results from a prospective pregnancy cohort study in urban Tanzania. Matern Child Nutr. (2022) 18:e13300. doi: 10.1111/mcn.13300
30. Zhang H, Bi W, Cui Y, Chen H, Chen J, Zhao Y, et al. Extreme-value sampling design is cost-beneficial only with a valid statistical approach for exposure–secondary outcome association analyses. Stat Methods Med Res. (2020) 29:466–80. doi: 10.1177/0962280219839093
31. Li GH, Cheung CL, Xiao SM, Lau KS, Gao Y, Bow CH, et al. Identification of QTL genes for BMD variation using both linkage and gene-based association approaches. Hum Genet. (2011) 130:539–46. doi: 10.1007/s00439-011-0972-2
32. Carr DF, O’Meara H, Jorgensen AL, Campbell J, Hobbs M, McCann G, et al. SLCO1B1 genetic variant associated with statin-induced myopathy: a proof-of-concept study using the clinical practice research datalink. Clin Pharmacol Ther. (2013) 94:695–701. doi: 10.1038/clpt.2013.161
33. Earle KA, Ambrosino DM, Fiore-Gartland A, Goldblatt D, Gilbert PB, Siber GR, et al. Evidence for antibody as a protective correlate for COVID-19 vaccines. Vaccine. (2021) 39:4423–8. doi: 10.1016/j.vaccine.2021.05.063
34. Khoury DS, Cromer D, Reynaldi A, Schlub TE, Wheatley AK, Juno JA, et al. Neutralizing antibody levels are highly predictive of immune protection from symptomatic SARS-CoV-2 infection. Nat Med. (2021) 27:1205–11. doi: 10.1038/s41591-021-01377-8
35. Bergwerk M, Gonen T, Lustig Y, Amit S, Lipsitch M, Cohen C, et al. Covid-19 breakthrough infections in vaccinated health care workers. N Engl J Med. (2021) 385:1474–84. doi: 10.1056/NEJMoa2109072
36. Chan KH, Leung KY, Zhang RR, Liu D, Fan Y, Chen H, et al. Performance of a surrogate SARS-coV-2-neutralizing antibody assay in natural infection and vaccination samples. Diagnostics (Basel). (2021) 11:1757. doi: 10.3390/diagnostics11101757
37. Chen S, Zhou Y, Chen Y, Gu J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics. (2018) 34:i884–90. doi: 10.1093/bioinformatics/bty560
38. Langmead B, Salzberg SL. Fast gapped-read alignment with Bowtie 2. Nat Methods. (2012) 9:357–9. doi: 10.1038/nmeth.1923
39. Truong DT, Franzosa EA, Tickle TL, Scholz M, Weingart G, Pasolli E, et al. MetaPhlAn2 for enhanced metagenomic taxonomic profiling. Nat Methods. (2015) 12:902–3. doi: 10.1038/nmeth.3589
40. Beghini F, McIver LJ, Blanco-Miguez A, Dubois L, Asnicar F, Maharjan S, et al. Integrating taxonomic, functional, and strain-level profiling of diverse microbial communities with bioBakery 3. Elife. (2021) 10:e65088. doi: 10.7554/eLife.65088
41. Han G, Vaishnava S. Microbial underdogs: exploring the significance of low-abundance commensals in host-microbe interactions. Exp Mol Med. (2023) 55:2498–507. doi: 10.1038/s12276-023-01120-y
42. Nikodemova M, Holzhausen EA, Deblois CL, Barnet JH, Peppard PE, Suen G, et al. The effect of low-abundance OTU filtering methods on the reliability and variability of microbial composition assessed by 16S rRNA amplicon sequencing. Front Cell Infect Microbiol. (2023) 13:1165295. doi: 10.3389/fcimb.2023.1165295
43. Wang X, Eijkemans MJ, Wallinga J, Biesbroek G, Trzcinski K, Sanders EA, et al. Multivariate approach for studying interactions between environmental variables and microbial communities. PloS One. (2012) 7:e50267. doi: 10.1371/journal.pone.0050267
44. Benjamini Y, Hochberg Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J R Stat Society. Ser B (Methodological). (1995) 57:289–300. doi: 10.1111/j.2517-6161.1995.tb02031.x
45. Stebegg M, Silva-Cayetano A, Innocentin S, Jenkins TP, Cantacessi C, Gilbert C, et al. Heterochronic faecal transplantation boosts gut germinal centres in aged mice. Nat Commun. (2019) 10:2443. doi: 10.1038/s41467-019-10430-7
46. Kasturi SP, Skountzou I, Albrecht RA, Koutsonanos D, Hua T, Nakaya H, et al. Programming the magnitude and persistence of antibody responses with innate immunity. Nature. (2011) 470:543–7. doi: 10.1038/nature09737
47. Lundell AC, Bjornsson V, Ljung A, Ceder M, Johansen S, Lindhagen G, et al. Infant B cell memory differentiation and early gut bacterial colonization. J Immunol. (2012) 188:4315–22. doi: 10.4049/jimmunol.1103223
48. Rudin A, Lundell AC. Infant B cell memory and gut bacterial colonization. Gut Microbes. (2012) 3:474–5. doi: 10.4161/gmic.21419
49. Torres J, Hu J, Seki A, Eisele C, Nair N, Huang R, et al. Infants born to mothers with IBD present with altered gut microbiome that transfers abnormalities of the adaptive immune system to germ-free mice. Gut. (2020) 69:42–51. doi: 10.1136/gutjnl-2018-317855
50. Louis P, Flint HJ. Formation of propionate and butyrate by the human colonic microbiota. Environ Microbiol. (2017) 19:29–41. doi: 10.1111/emi.2017.19.issue-1
51. Xie J, Li LF, Dai TY, Qi X, Wang Y, Zheng TZ, et al. Short-chain fatty acids produced by ruminococcaceae mediate α-linolenic acid promote intestinal stem cells proliferation. Mol Nutr Food Res. (2022) 66:e2100408. doi: 10.1002/mnfr.202100408
52. Sun J, Chen S, Zang D, Sun H, Sun Y, Chen J. Butyrate as a promising therapeutic target in cancer: From pathogenesis to clinic (Review). Int J Oncol. (2024) 64:44. doi: 10.3892/ijo.2024.5632
53. Wegmann U, Louis P, Goesmann A, Henrissat B, Duncan SH, Flint HJ. Complete genome of a new Firmicutes species belonging to the dominant human colonic microbiota (‘Ruminococcus bicirculans’) reveals two chromosomes and a selective capacity to utilize plant glucans. Environ Microbiol. (2014) 16:2879–90. doi: 10.1111/emi.2014.16.issue-9
54. Ishizaka S, Kikuchi E, Tsujii T. Effects of acetate on human immune system. Immunopharmacol Immunotoxicol. (1993) 15:151–62. doi: 10.3109/08923979309025991
55. Yu J, Zhang H, Chen L, Ruan Y, Chen Y, Liu Q. Disease-associated gut microbiota reduces the profile of secondary bile acids in pediatric nonalcoholic fatty liver disease. Front Cell Infect Microbiol. (2021) 11:698852. doi: 10.3389/fcimb.2021.698852
56. Ju T, Kong JY, Stothard P, Willing BP. Defining the role of Parasutterella, a previously uncharacterized member of the core gut microbiota. ISME J. (2019) 13:1520–34. doi: 10.1038/s41396-019-0364-5
57. Couvigny B, de Wouters T, Kaci G, Jacouton E, Delorme C, Dore J, et al. Commensal streptococcus salivarius modulates PPARgamma transcriptional activity in human intestinal epithelial cells. PloS One. (2015) 10:e0125371. doi: 10.1371/journal.pone.0125371
58. Heming M, Gran S, Jauch SL, Fischer-Riepe L, Russo A, Klotz L, et al. Peroxisome proliferator-activated receptor-gamma modulates the response of macrophages to lipopolysaccharide and glucocorticoids. Front Immunol. (2018) 9:893. doi: 10.3389/fimmu.2018.00893
59. Li K, Hao Z, Du J, Gao Y, Yang S, Zhou Y. Bacteroides thetaiotaomicron relieves colon inflammation by activating aryl hydrocarbon receptor and modulating CD4(+)T cell homeostasis. Int Immunopharmacol. (2021) 90:107183. doi: 10.1016/j.intimp.2020.107183
60. Tomkovich S, Jobin C. Microbiota and host immune responses: a love-hate relationship. Immunology. (2016) 147:1–10. doi: 10.1111/imm.2016.147.issue-1
61. Sahin U, Muik A, Derhovanessian E, Vogler I, Kranz LM, Vormehr M, et al. COVID-19 vaccine BNT162b1 elicits human antibody and T(H)1 T cell responses. Nature. (2020) 586:594–9. doi: 10.1038/s41586-020-2814-7
62. Gupta S, Su H, Agrawal S. CD8 treg cells inhibit B-cell proliferation and immunoglobulin production. Int Arch Allergy Immunol. (2020) 181:947–55. doi: 10.1159/000509607
63. Moore AC, Gallimore A, Draper SJ, Watkins KR, Gilbert SC, Hill AV. Anti-CD25 antibody enhancement of vaccine-induced immunogenicity: increased durable cellular immunity with reduced immunodominance. J Immunol. (2005) 175:7264–73. doi: 10.4049/jimmunol.175.11.7264
64. La Reau AJ, Suen G. The Ruminococci: key symbionts of the gut ecosystem. J Microbiol. (2018) 56:199–208. doi: 10.1007/s12275-018-8024-4
65. Sikalidis AK. Amino acids and immune response: a role for cysteine, glutamine, phenylalanine, tryptophan and arginine in T-cell function and cancer? Pathol Oncol Res. (2015) 21:9–17. doi: 10.1007/s12253-014-9860-0
66. Levring TB, Hansen AK, Nielsen BL, Kongsbak M, von Essen MR, Woetmann A, et al. Activated human CD4+ T cells express transporters for both cysteine and cystine. Sci Rep. (2012) 2:266. doi: 10.1038/srep00266
67. Levring TB, Kongsbak M, Rode AK, Woetmann A, Odum N, Bonefeld CM, et al. Human CD4+ T cells require exogenous cystine for glutathione and DNA synthesis. Oncotarget. (2015) 6:21853–64. doi: 10.18632/oncotarget.v6i26
68. Mihm S, Galter D, Droge W. Modulation of transcription factor NF kappa B activity by intracellular glutathione levels and by variations of the extracellular cysteine supply. FASEB J. (1995) 9:246–52. doi: 10.1096/fasebj.9.2.7781927
69. Droge W, Eck HP, Gmunder H, Mihm S. Modulation of lymphocyte functions and immune responses by cysteine and cysteine derivatives. Am J Med. (1991) 91:140S–4S. doi: 10.1016/0002-9343(91)90297-B
70. Nie C, He T, Zhang W, Zhang G, Ma X. Branched chain amino acids: beyond nutrition metabolism. Int J Mol Sci. (2018) 19:954. doi: 10.3390/ijms19040954
71. Fan P, Li L, Rezaei A, Eslamfam S, Che D, Ma X. Metabolites of dietary protein and peptides by intestinal microbes and their impacts on gut. Curr Protein Pept Sci. (2015) 16:646–54. doi: 10.2174/1389203716666150630133657
72. Calder PC. Branched-chain amino acids and immunity. J Nutr. (2006) 136:288S–93S. doi: 10.1093/jn/136.1.288S
73. Gu C, Mao X, Chen D, Yu B, Yang Q. Isoleucine plays an important role for maintaining immune function. Curr Protein Pept Sci. (2019) 20:644–51. doi: 10.2174/1389203720666190305163135
74. Mekonnen D, Mengist HM, Jin T. SARS-CoV-2 subunit vaccine adjuvants and their signaling pathways. Expert Rev Vaccines. (2022) 21:69–81. doi: 10.1080/14760584.2021.1991794
75. Kurteva E, Vasilev G, Tumangelova-Yuzeir K, Ivanova I, Ivanova-Todorova E, Velikova T, et al. Interferon-gamma release assays outcomes in healthy subjects following BNT162b2 mRNA COVID-19 vaccination. Rheumatol Int. (2022) 42:449–56. doi: 10.1007/s00296-022-05091-7
76. Mao X, Gu C, Ren M, Chen D, Yu B, He J, et al. l-isoleucine administration alleviates rotavirus infection and immune response in the weaned piglet model. Front Immunol. (2018) 9:1654. doi: 10.3389/fimmu.2018.01654
77. Zhang C, Zhang M, Pang X, Zhao Y, Wang L, Zhao L. Structural resilience of the gut microbiota in adult mice under high-fat dietary perturbations. ISME J. (2012) 6:1848–57. doi: 10.1038/ismej.2012.27
78. Zeng Q, Li D, He Y, Li Y, Yang Z, Zhao X, et al. Discrepant gut microbiota markers for the classification of obesity-related metabolic abnormalities. Sci Rep. (2019) 9:13424. doi: 10.1038/s41598-019-49462-w
79. Ma S, You Y, Huang L, Long S, Zhang J, Guo C, et al. Alterations in gut microbiota of gestational diabetes patients during the first trimester of pregnancy. Front Cell Infect Microbiol. (2020) 10:58. doi: 10.3389/fcimb.2020.00058
80. Ding Z, Zhao Y, Liu J, Ge W, Xu X, Wang S, et al. Dietary succinoglycan riclin improves glycemia control in mice with type 2 diabetes. J Agric Food Chem. (2022) 70:1819–29. doi: 10.1021/acs.jafc.1c06881
81. Xiao S, Liu C, Chen M, Zou J, Zhang Z, Cui X, et al. Scutellariae radix and coptidis rhizoma ameliorate glycolipid metabolism of type 2 diabetic rats by modulating gut microbiota and its metabolites. Appl Microbiol Biotechnol. (2020) 104:303–17. doi: 10.1007/s00253-019-10174-w
82. Li J, Jia S, Yuan C, Yu B, Zhang Z, Zhao M, et al. Jerusalem artichoke inulin supplementation ameliorates hepatic lipid metabolism in type 2 diabetes mellitus mice by modulating the gut microbiota and fecal metabolome. Food Funct. (2022) 13:11503–17. doi: 10.1039/D2FO02051C
83. Zhou R, He D, Zhang H, Xie J, Zhang S, Tian X, et al. Ginsenoside Rb1 protects against diabetes-associated metabolic disorders in Kkay mice by reshaping gut microbiota and fecal metabolic profiles. J Ethnopharmacol. (2023) 303:115997. doi: 10.1016/j.jep.2022.115997
84. Forsgard RA, Rode J, Lobenius-Palmer K, Kamm A, Patil S, Tacken MGJ, et al. Limosilactobacillus reuteri DSM 17938 supplementation and SARS-CoV-2 specific antibody response in healthy adults: a randomized, triple-blinded, placebo-controlled trial. Gut Microbes. (2023) 15:2229938. doi: 10.1080/19490976.2023.2229938
Keywords: gut microbiota, vaccine, COVID-19 vaccine, vaccine immunogenicity, BNT162b2 (Pfizer-BioNTech)
Citation: Ng HY, Liao Y, Cheung CL, Zhang R, Chan KH, Seto W-K, Leung WK, Hung IFN, Lam TTY and Cheung KS (2025) Gut microbiota is associated with persistence of longer-term BNT162b2 vaccine immunogenicity. Front. Immunol. 16:1534787. doi: 10.3389/fimmu.2025.1534787
Received: 26 November 2024; Accepted: 07 February 2025;
Published: 27 February 2025.
Edited by:
Huiwen Zheng, Chinese Academy of Medical Sciences and Peking Union Medical College, ChinaCopyright © 2025 Ng, Liao, Cheung, Zhang, Chan, Seto, Leung, Hung, Lam and Cheung. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Ka Shing Cheung, Y2tzNjM0QGhrdS5oaw==; Ivan F. N. Hung, aXZhbmZuQGdtYWlsLmNvbQ==; Tommy T. Y. Lam, dHR5bGFtQGhrdS5oaw==
†These authors share first authorship
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.