
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. For. Glob. Change , 17 March 2025
Sec. Forest Ecophysiology
Volume 8 - 2025 | https://doi.org/10.3389/ffgc.2025.1557824
 Tian-You He1†
Tian-You He1† Peng-Kai Zhu1,2*†
Peng-Kai Zhu1,2*† Cai-Ling Song1
Cai-Ling Song1 Min Wu1
Min Wu1 Jian-Nan Lin1
Jian-Nan Lin1 Yan-Ping Xu1
Yan-Ping Xu1 Jia-Lin Zhang1
Jia-Lin Zhang1 Ying-Hui Zhang3
Ying-Hui Zhang3 Jun-Dong Rong1
Jun-Dong Rong1 Kai Liu1
Kai Liu1 Li-Guang Chen1
Li-Guang Chen1 Yu-Shan Zheng1
Yu-Shan Zheng1 Ling-Yan Chen1*
Ling-Yan Chen1*In the context of climate change, assessing the adaptive potential of species and populations is crucial for developing effective conservation strategies. Changes in plant gene expression play a significant role in the adaptation process to climate change. This study aims to explore the adaptive responses of the conifer species Chamaecyparis hodginsii (the name has been revised from Fokienia hodginsii) to climate change and analyze the molecular-level reactions of these long-lived trees to climatic shifts. It seeks to understand their phenotypic responses to climate change, identify key environmental factors driving adaptive gene expression, and provide information for transplantation conservation strategies based on genetic adaptability. By conducting mixed-tissue RNA sequencing on samples from multiple provenances and employing redundancy analysis (RDA), weighted gene co-expression network analysis (WGCNA), and partial least squares path modeling (PLS-PM), the study assesses the impact of climatic variables on gene expression and phenotype. It identifies key gene groups associated with environmental responses and elucidates the complex relationships between environmental factors, functional gene groups, and phenotypic traits. The findings reveal that C. hodginsii adapts to environmental stresses by regulating specific gene activities related to morphological trait adjustments. Moreover, environmental factors such as the impact on tree architecture emphasize the importance of Precipitation Seasonality, Isothermality, and Precipitation of Driest Quarter for adapting to climate stresses. This research not only unveils the complex adaptive responses of C. hodginsii to climate change but also provides critical insights for the management and conservation of long-lived tree species facing climate change threats.
Climate change is one of the major threats to biodiversity, with studies showing it has led to localized extinctions of animal and plant species (Walther et al., 2002). Assessing the current adaptive state of species and populations is essential for effective conservation strategies (Eizaguirre and Baltazar-Soares, 2014; Razgour et al., 2019). For long-lived species, such as forest trees, rates of mutation and migration may be insufficient to match the pace of environmental change, thus limiting adaptive potential (Holliday et al., 2017; Bisbing et al., 2021; Dauphin et al., 2021). Consequently, shifts in gene expression are critical for plant adaptation to changing environments and can, to some extent, reflect the adaptive status of species. Rapid environmental shifts can induce changes in gene expression programs, thereby regulating plant metabolism to adapt to new conditions and increasing phenotypic variation (López-Maury et al., 2008; Berger et al., 2011). Changes in gene expression not only directly reflect the species’ adaption to current environments but are also essential for assessing the viability of natural species under climate change and for developing effective ecological and forest management strategies. During the process of adapting to antibiotic stress, plants undergo extensive changes in gene expression (Chen and Soltis, 2020; Kan et al., 2023), providing a basis for natural selection on short evolutionary timescales (Xu et al., 2016; Innes et al., 2023).
Differences in gene expression patterns can explain the varying abilities of different populations to adapt to environmental changes (Poncet et al., 2010; Xu et al., 2016). Traditional transplant or garden experiments are the gold standard for assessing plant adaptability (Weeks et al., 2011). Translocating organisms from their native habitats to new environments also serves as an effective method to conserve genetic diversity (Anderson et al., 2011; Fournier-Level et al., 2011; Sang et al., 2022), especially for threatened and rare species (Hufford and Mazer, 2003). Notably, seedlings, being in the early stages of growth, may be more sensitive to environmental changes (Hummel et al., 2004; Huang and Wang, 2016). RNA-seq has been successfully applied to adaptive studies in conifer species (Ma et al., 2019; Yang et al., 2022). Additionally, Weighted Gene Co-expression Network Analysis (WGCNA) can identify gene sets closely related to climate adaptation, revealing how plants regulate gene expression to cope with rapidly changing climates, thereby deepening the current understanding of species adaptation mechanisms.
Chamaecyparis hodginsii is classified as Vulnerable (VU) on the International Union for Conservation of Nature (IUCN) Red List (Thomas and Yang, 2013) and is listed as a Category II protected plant in China1. This Tertiary relict species has survived since the glacial periods (Yin et al., 2018; Rong et al., 2024). In some regions where it currently exists, it remains a dominant species; however, historical human activity and climate change have led to population decline and habitat fragmentation, which are the primary threats to its persistence (Yin Q.-Y. et al., 2021). Additionally, fossil records indicate that this genus was once widely distributed across the Northern Hemisphere, including locations such as Saskatchewan, Canada (McIver and Basinger, 1990), Jilin in northeastern China (Ying and Fu, 2002), and Zhejiang in eastern Chin (He et al., 2012). However, climate shifts have led to its disappearance in these areas, and C. hodginsii is now limited to southern China, northern Vietnam, and northern Laos (Yin et al., 2018). It is highly likely that C. hodginsii will face further population declines and heightened survival challenges under the current context of climate change. The recently released C. hodginsii genome (Rong et al., 2024) offers an excellent opportunity to evaluate its environmental plasticity at the genetic level.
Trees act as effective carbon sinks and are crucial in fighting climate change (Isabel et al., 2020; Zhu et al., 2023). Their long generation times and frequent extreme climate events make them vulnerable to climate change (Nepstad et al., 2007; Dauphin et al., 2021). This study focuses on C. hodginsii, a near-threatened species. We aim to understand its adaptability to climate change using mixed-tissue RNA sequencing. Our goals include revealing tree phenotypes’ response to climate change at the gene expression level, identifying key environmental factors affecting adaptability, and analyzing gene expression regulation. This work seeks to provide insights and strategies for protecting long-lived tree species against climate change.
All plant samples in this study were sourced from the Bailai State-owned Forest Farm in Hutou Town, Anxi County, Quanzhou City, Fujian Province, China (118°3′ E, 25°49’ N, 590–600 m above sea level). The plant experiments and field studies, including the collection of plant materials, complied with relevant institutional, national, and international guidelines and regulations. The germination and sprouting experiments of cypress seeds were conducted in the greenhouse of Fujian Agriculture and Forestry University, with seeds disinfected before sowing. At the end of 2015, seed germination began under controlled environmental conditions, maintaining a temperature of 22–25°C and a relative humidity of 70–80%. Successfully germinated seeds were then transferred to seedling trays filled with standard nutrient soil to promote seedling growth. Seedlings received at least 8 h of natural light, supplemented by grow lights to ensure adequate lighting, and were watered and fertilized according to standard care procedures. In April 2016, when the seedlings reached about 15 cm in height, they were transplanted to outdoor plots with similar soil texture, drainage conditions, and light exposure. The experiment employed a completely randomized block design (CRD) with 8 plots, each containing representative seedlings from 14 different provenances (Supplementary Table S1 and Supplementary Figure S1). Five seedlings from each provenance were randomly assigned to each plot to ensure even distribution across the experimental area, with a planting distance of 2 m to minimize competition. Appropriate measures were taken to alleviate transplant shock, and the growth and adaptability of the seedlings were regularly monitored post-transplantation. In April 2020, 4 years after transplantation, key growth parameters of the surviving seedlings were measured, including Height, Diameter at Breast Height (DBH), Crown Width (CW), and the Average Angle of Dips, the latter obtained by calculating the average angle of the three lowest secondary branches of each plant.
In April 2020, we randomly selected seven individuals from each seed provenance and collected their stem-differentiating xylem, leaves, branches, and roots. To ensure robustness and reliability of the data, samples were collected in two separate replicates for each seed provenance. These tissues were mixed by category and disinfected uniformly with alcohol for 30 s to 1 min, then rinsed in deionized water for 2 min to remove any alcohol residue and dead microbes. One gram of mixed tissue sample from each type was immediately weighed, flash-frozen in liquid nitrogen, and stored at −80°C for later use. Total RNA was extracted using the RNAprep Pure Plant Kit (Tiangen, Beijing, China) and the quality of RNA was assessed by 2% agarose gel electrophoresis. The concentration of RNA was determined by the NanoPhotometer® spectrophotometer (IMPLEN, California, United States) and the Qubit® 2.0 Fluorometer in conjunction with the Qubit® RNA Assay Kit (Life Technologies, California, United States). RNA integrity was evaluated using the RNA Nano 6,000 Assay Kit on the Agilent® 2,100 Bioanalyzer (Agilent Technologies, California, United States), with an RNA integrity number (RIN) greater than seven considered satisfactory. Subsequently, libraries were constructed using the NEB-Next® Ultra™ RNA Library Prep Kit for Illumina® (NEB, United States) and sequenced on the Illumina® NovaSeq 6,000 platform to produce 150 bp paired-end reads.
We performed initial processing of the raw RNA-Seq data (NGDC: CRX961097–CRX961124) using fastp v0.23.2 (Chen et al., 2018) with parameters set to ensure quality (−qualified_quality_phred 15 --unqualified_percent_limit 50 --n_base_limit 15), generating a total of 315.16 Gb of clean data. Subsequently, we obtained the C. hodginsii reference genome and structural annotations (Rong et al., 2024) from https://doi.org/10.6084/m9.figshare.26064412.v1. The clean data were aligned to the genome using the Two-pass alignment mode of STAR v2.7.8a (Dobin et al., 2013) with parameters of --alignIntronMin 20 --alignIntronMax 50,000 --sjdbOverhang 149 --outFilterMismatchNmax 2 --outSJfilterReads Unique --outSAMmultNmax 1 --outSAMmapqUnique 60 to enhance the accuracy and reliability of the alignment, resulting in an average depth of 48.63x per base. After alignment, gene expression levels for each gene were quantified using featureCounts v2.0.1 (Liao et al., 2014) with default parameters.
First, we obtained information on 19 environmental factors from various seed provenance locations (Supplementary Table S1), sourced from the WorldClim database (Fick and Hijmans, 2017). Spearman correlation analysis was employed to assess the interrelationships among these environmental factors. High correlations (correlation coefficient |r| > 0.65) were observed among many environmental factors, likely due to the close proximity of certain provenance locations. Such high correlations can lead to multicollinearity among predictors in redundancy analysis (RDA), reducing model accuracy. Therefore, following the method of Sang et al., we selectively retained the most significant variables from highly correlated groups (Sang et al., 2022). We conducted gradient forest model analysis using the R package gradientForest v0.1–37 (Ellis et al., 2012), setting the tree count to 500, to evaluate the impact of each environmental factor on gene expression levels and determine their importance. Based on the results of the gradient forest analysis and the correlations between environmental factors, we selected factors that were both significant and had low intercorrelations to reduce potential collinearity issues. The R package stats v4.3.3 is used to perform Principal Components Analysis (PCA) on various samples to initially characterize the potential associations between provenances and expression data. These selected environmental factors were then used as explanatory variables in subsequent Redundancy Analysis (RDA) to delve deeper into their relationship with gene expression data. RDA, known for its low false-positive rate (Capblancq and Forester, 2021), was conducted using the R package vegan v2.6–4 (Oksanen et al., 2017) to further reveal how environmental factors influence gene expression patterns. Significant environment-associated expressed genes were defined by their loadings on the tails of the distribution along one or more RDA axes.
To gain a deeper understanding of the modular characteristics of gene expression, this study employed WGCNA. We utilized the Python package PyWGCNA v2.0.1 (Rezaie et al., 2023) with default parameter settings. This analysis clusters genes into modules based on similar expression patterns, aiding in exploring the correlations among these modules, environmental factors, and morphological traits of each seed provenance. It helps identify groups of genes that may be influenced by specific environmental factors. Within each module significantly related to environmental and phenotypic factors, we further filtered to select genes with a KME value greater than 0.6 as key genes. These key genes are considered to play central roles within the modules and may be sensitive to environmental changes. Functional annotation and enrichment analysis identified genes with significant functions and strong correlations to environmental factors. Finally, we selected these genes for constructing a Partial Least Squares Path Model (PLS-PM) using the R package plspm v0.5.1 (Tenenhaus et al., 2005), setting the maximum iteration count to 300. We began by hypothesizing potential connections among three variable groups based on the prior assumption that environmental changes, mediated by gene expression, influence phenotypic traits. Using PLS-PM, we then assessed the strength, direction, and reliability of relationships between each factor within the environmental and gene expression datasets. This was followed by analyzing connections between gene expression and phenotypic changes, ultimately linking the three variable groups through the combined results. The PLS-PM model reveals the direct and indirect effects among these variables, offering profound insights into how gene expression is regulated by environmental factors at the seed provenance location, impacting the phenotype.
Through correlation analysis, we identified temperature and precipitation metrics that exhibit strong positive correlations (Figure 1A). To address these collinearity issues and ensure the accuracy of subsequent analyses, we employed a gradient forest model to assess the importance of various environmental factors on gene expression (Figure 1B). The results indicated that the precipitation of the driest quarter and the precipitation of the driest month were most critical to the predictive models, suggesting the significance of drought conditions in influencing gene expression. Furthermore, to avoid collinearity issues, we selected the most important representative factor from each group, ultimately identifying five relatively independent and significant environmental factors: precipitation of the driest quarter, mean temperature of the driest quarter, annual temperature range, isothermality, and precipitation seasonality. These factors are crucial for explaining variations in gene expression.
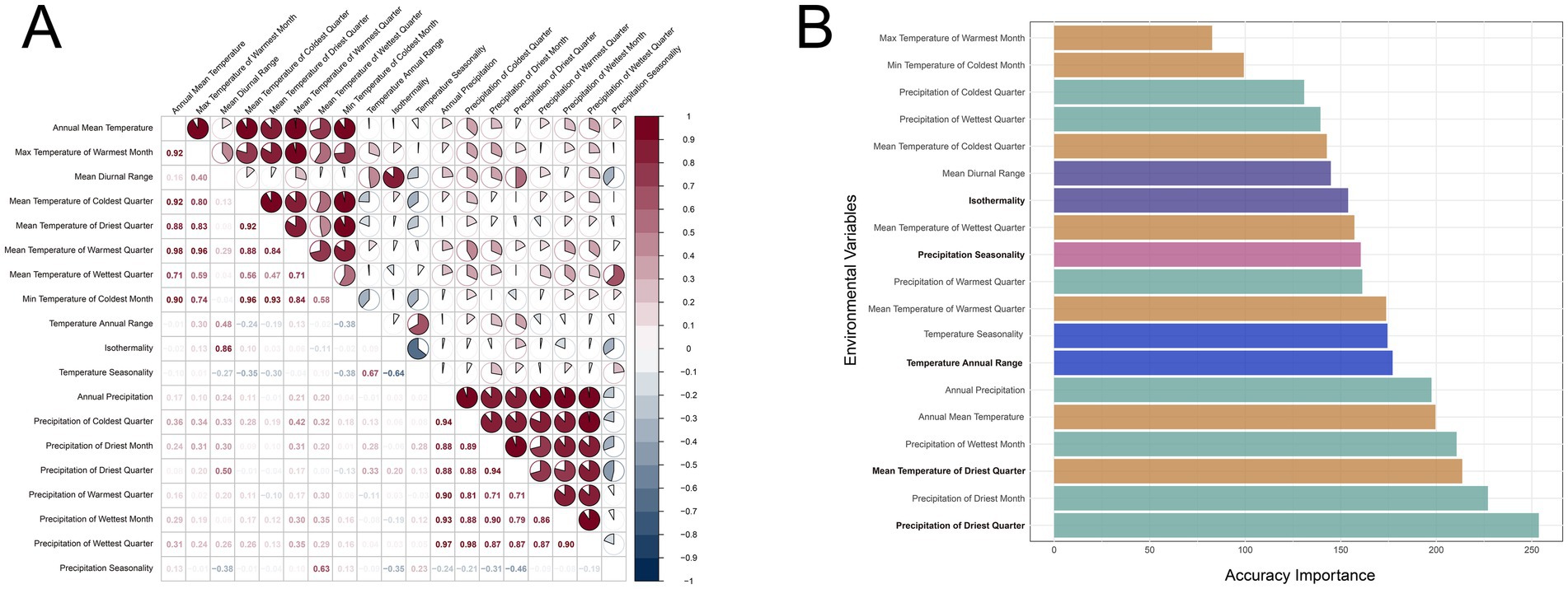
Figure 1. Correlation matrix (A) and importance barplot (B) of the 19 environmental factors. The depth of color and the fill level of the shapes are both used to represent the strength of the correlation, with dark colors and solid fills indicating strong correlations, and white and hollow shapes indicating no correlation. Environmental factor importance was calculated using the random forest algorithm. Within the highly correlated groups (correlation coefficient |r| > 0.65), the most important factors are highlighted in bold and the bars for the same factor group are colored the same.
To explore the expression heterogeneity of different provenances of C. hodginsii, we first characterized the distribution of each sample’s expression levels using PCA. The results showed a potential correlation between expression levels and the geographic distance of the provenances. For instance, CQJJ is geographically distant from other provenances (Supplementary Figure S1), and its samples also exhibit a more distinct distribution compared to others (Supplementary Figure S2). The RDA results further confirmed this connection, revealing a strong relationship between gene expression and environmental factors of the provenances, with the first axis explaining 38.6% of the environmental variation, while the second and third axes accounted for an additional 21.7 and 16.5% of variation, respectively, highlighting the profound impact of environmental factors on gene expression (Figures 2A,B and Supplementary Table S2). Notably, the significant orientation of Precipitation Seasonality on the first axis indicated it as a key environmental factor driving major variations in gene expression. The similar direction of arrows for Isothermality and Precipitation of Driest Quarter suggested some similarity in their effects on gene expression. In contrast, the unique direction of Mean Temperature of Driest Quarter pointed to a different pattern of influence compared to other factors. Furthermore, the directions of Temperature Annual Range and Precipitation of Driest Quarter nearly overlapped on the second and third axes (Figure 2B), suggesting similarities in their effects on gene expression patterns in these specific dimensions. However, this does not imply a lack of differences between these two factors; rather, such differences may not be evident on these two axes.
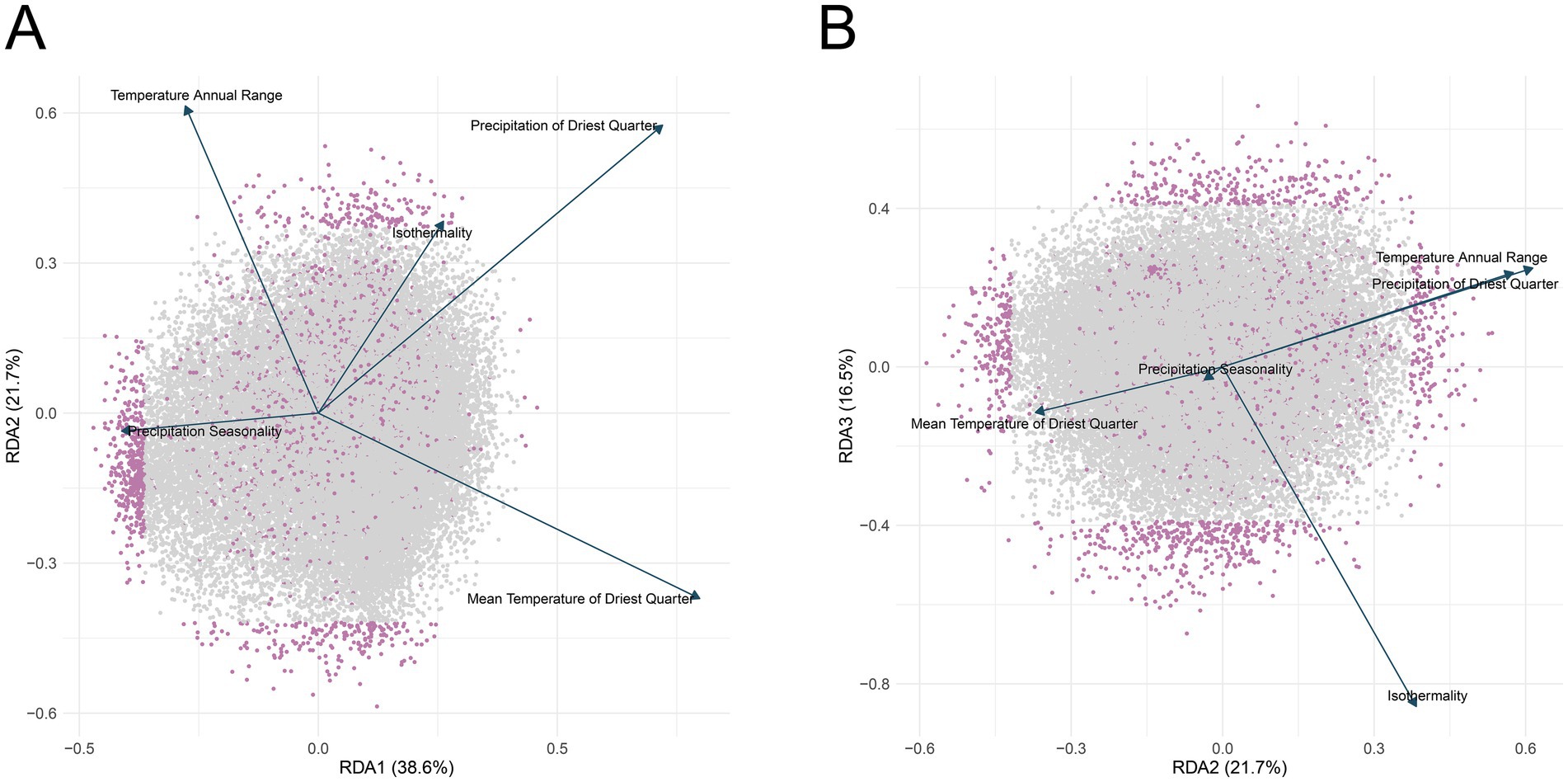
Figure 2. Ordination plots of redundancy analysis (RDA) of gene expression under different environmental conditions, showing RDA1 vs. RDA2 (A) and RDA2 vs. RDA3 (B). In both plots, individual genes are depicted as small points, with those more affected by environmental factors highlighted in purple. The vectors represent the direction and magnitude of the environmental factors’ influence on gene expression, with the length of the vector corresponding to the strength of the influence.
Further analysis revealed that 2,941 genes had significant loadings on at least one RDA axis, indicating high sensitivity to specific environmental factors and significant expression variation. Through 999 permutation tests, our model was statistically significant (p < 0.05), confirming the substantial impact of Precipitation of Driest Quarter and Mean Temperature of Driest Quarter on gene expression (Supplementary Table S3). Overall, environmental factors accounted for 25.06% of the total variance, with the remaining variance likely due to other unconsidered variables. These findings highlight the complexity of plant gene regulatory networks and suggest that plants may respond to diverse environmental conditions through transcriptional regulatory mechanisms.
Through WGCNA, we categorized the genes into 14 modules, each containing a cluster of genes with similar expression patterns (Supplementary Figure S3). The analysis indicated significant correlations between specific modules and environmental factors or phenotypic traits, suggesting that certain genes may respond to environmental stresses or phenotypic changes. For instance, the Rosybrown module showed a positive correlation with Precipitation of Driest Quarter (r = 0.65, p < 0.05), while the Brown module was negatively correlated with DBH (r = −0.63, p < 0.05) (Figure 3). To identify genes most sensitive to environmental changes, we selected those with module membership (KME) values greater than 0.6 as key genes, highly correlated with the representative gene expression patterns of their respective modules. Integrating these findings with RDA results, we further pinpointed key genes within the Dimgrey, Firebrick, Rosybrown, and Snow modules. These modules demonstrated strong correlations with environmental factors or phenotypic traits, reinforcing the credibility of the RDA findings. Notably, the Dimgrey module contributed the majority of key genes, with fewer key genes from the other three modules (Supplementary Figure S4), suggesting that while multiple modules are associated with environmental or phenotypic characteristics, the Dimgrey module may play a leading role in responding to environmental changes. Functional enrichment analysis of these key genes revealed their involvement in the synthesis of cyclic and aromatic compounds and various enzymatic activities (Figure 4). These functions are crucial for cellular metabolic activities, the transmission of genetic information, and genome integrity, highlighting the pivotal role of these genes in plant adaptive responses and phenotypic shaping.
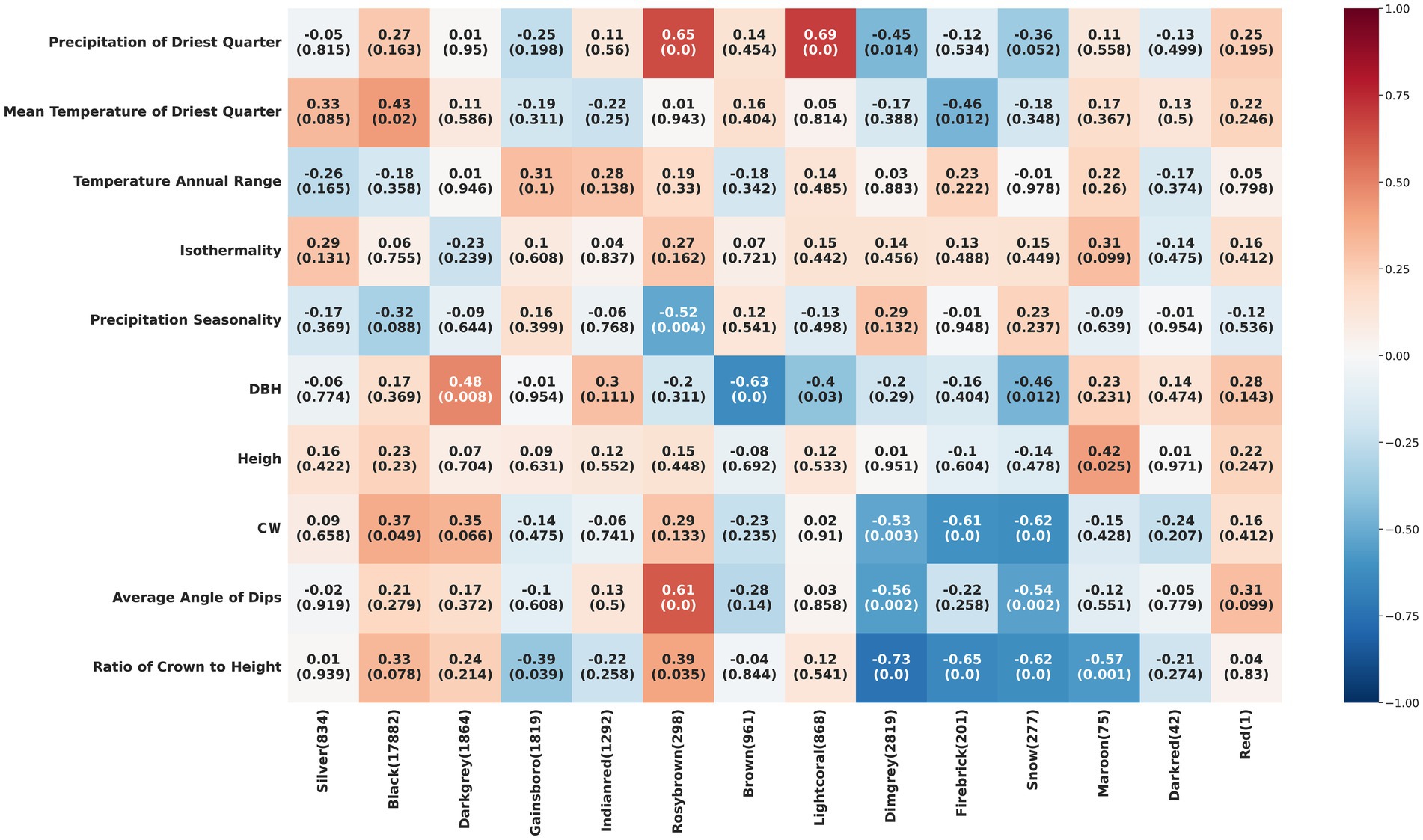
Figure 3. Heatmap showing the results of weighted gene co-expression network analysis (WGCNA) associating gene modules with environmental and phenotypic variables. Each cell in the heatmap represents the correlation coefficient and its significance in parentheses between a gene module and an environmental or phenotypic trait. The color scale on the right indicates the strength of the correlation.
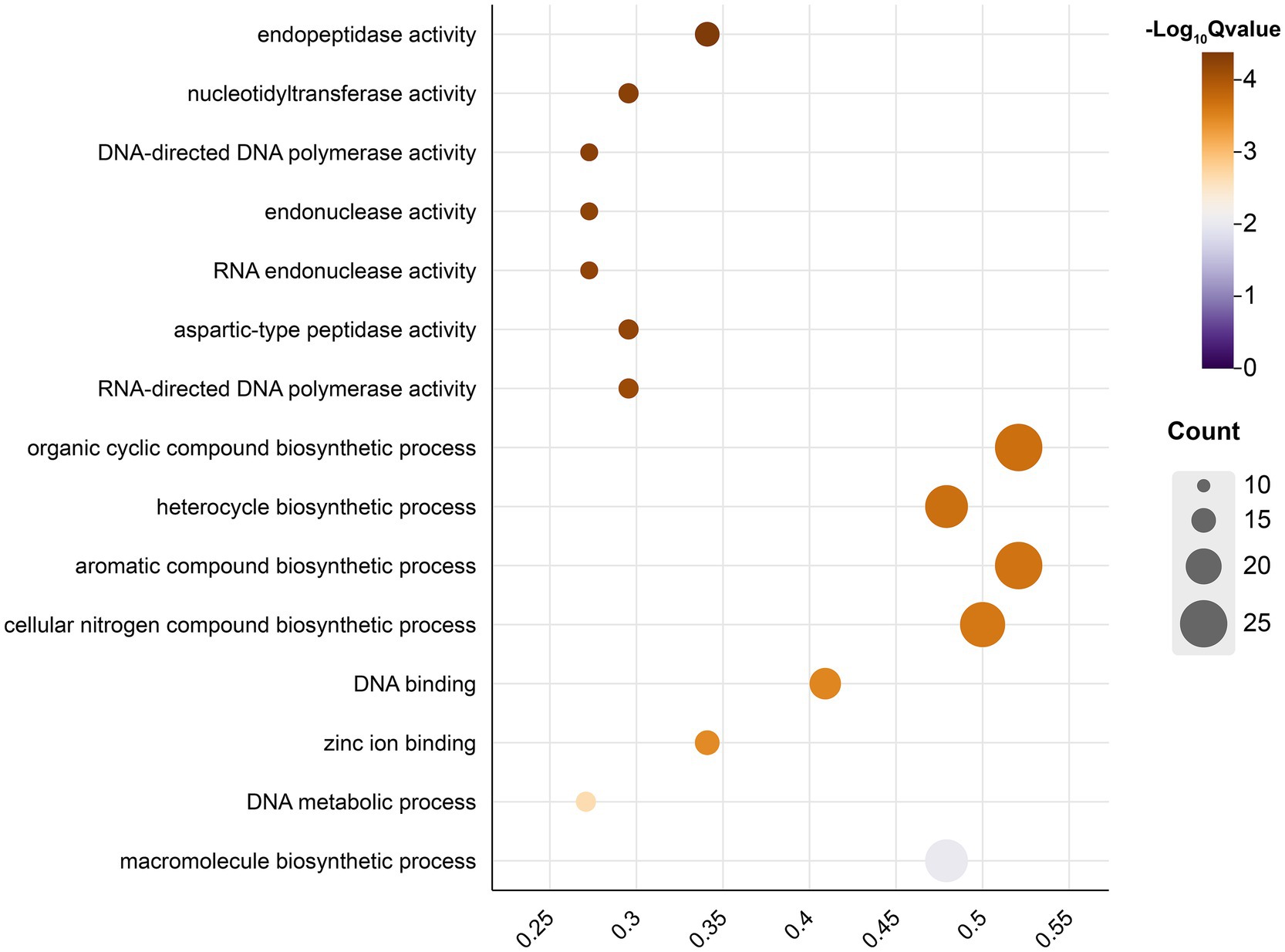
Figure 4. Gene ontology (GO) functional enrichment analysis of key genes identified with KME > 0.6 by WGCNA and RDA. The bubble plot represents various biological processes and molecular functions, with the size of each bubble reflecting the number of genes associated with that term and the color representing the -log10Qvalue of the enrichment significance.
To further understand how environmental factors influence plant phenotypic traits through key genes, we employed PLS-PM. This analysis revealed a network of relationships among environmental variables, functional gene groups within the key genes, and phenotypic traits (p < 0.05). The environmental variables included only Precipitation Seasonality, Isothermality, and Precipitation of Driest Quarter. These factors significantly impacted two groups of genes: one involved in the biosynthesis of cyclic and aromatic compounds, and the other related to peptidase and nucleotidyltransferase activities (Figure 5 and Supplementary Table S4). These relationships were further linked to two phenotypic traits of trees: CW and Average Angle of Dips. Precipitation Seasonality and Isothermality directly influenced the genes associated with cyclic and aromatic compound biosynthesis, subsequently affecting CW. In contrast, the mean temperatures of the three environmental variables influenced the gene groups related to peptidase and nucleotidyltransferase activities, which then affected the Average Angle of Dips. Additionally, our model explained 76.15% of the variance in CW and 68.24% of the variation in Average Angle of Dips (Supplementary Table S5), indicating that we could substantially explain the variations in phenotypic traits. This model helps visualize the complexity of gene responses to environmental cues and their potential impact on phenotypes, emphasizing the significance of specific environmental factors in the adaptive responses of the species.
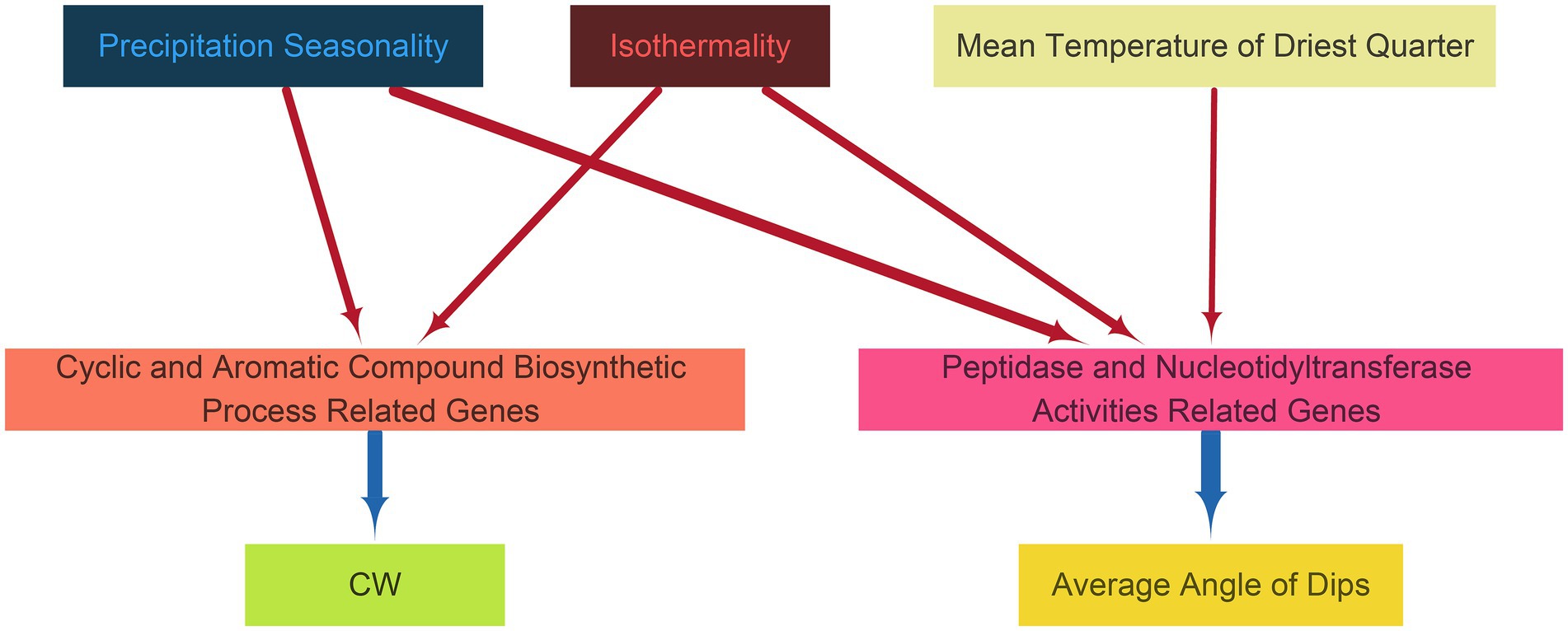
Figure 5. Partial least squares path modeling (PLS-PM) plot showing the relationships between specific functional gene categories and environmental and phenotypic data. These relationships are represented by the color and thickness of the arrows, with red indicating positive effects and blue indicating negative effects, and the thicker the arrow, the stronger the relationship.
The capacity of plants to adapt to environmental stress, particularly under rapidly changing climate conditions, is a central topic in ecological and evolutionary biology research. Previous studies have shown that plants can achieve local adaptation through genetic variation even over short geographical and temporal scales (Medrano et al., 2020; Innes et al., 2023), suggesting that plants can cope with diverse environmental challenges by selecting specific life-history traits (Poncet et al., 2010). However, long-lived and highly specialized species may face issues of adaptive lag, especially pronounced against the backdrop of rapid climate change (Holliday et al., 2017; Bisbing et al., 2021; Sang et al., 2022). In response to this challenge, using the translocation of C. hodginsii as an example, explored the potential adaptability of conifer trees under environmental change pressures. Climate change has significantly increased the intensity of droughts in humid regions, posing major challenges to plant growth and development (Ruffault et al., 2013; Grossiord et al., 2020). Temperature and moisture are key factors affecting plant growth and development, with changes in environmental temperature directly impacting plant physiological processes (Porporato et al., 2001; Ding et al., 2020).
The strong El Niño event of 2015–2016, part of the El Niño Southern Oscillation (ENSO) phenomenon (Qian, 2022), coincided with the transplantation period of our C. hodginsii seedlings. This severe drought may lead to a significant increase in tree mortality rates (Nepstad et al., 2007), presenting a unique challenge to plant growth and development. Under such harsh conditions, the early growth stages of plants are likely to be profoundly affected. Notably, transcriptomic analyses were conducted on these four-year-old transplanted C. hodginsii. The results showed a range of transcriptional changes closely related to the climate of their seed provenance locations (Figures 2, 3), especially genes involved in the biosynthesis of cyclic and aromatic compounds (Figure 4). This suggests that C. hodginsii might cope with early environmental stress by regulating the synthesis of aromatic compounds. These compounds not only play roles in signaling between plants, enhancing stress tolerance (Ahammed et al., 2015, 2020), but may also prime surrounding plant tissues for stronger responses to subsequent stresses (Erb et al., 2015). This mechanism is particularly important for coniferous trees known for their aromatic wood and contributes to their adaptation to rapidly changing environments.
Gymnosperms often have large genome sizes, largely attributed to historical transposon activity causing genomic redundancy (Zhu et al., 2023). In this study, we found a positive correlation between the climate of seed provenance locations and genes involved in peptidase and nucleotidyltransferase activities, enzymes associated with transposon activity (Hickman and Dyda, 2015, 2016). This suggests that there might be differences in transposon activity levels among C. hodginsii provenances. Additionally, the activity changes in these genes seem to be related to a decrease in the Average Angle of Dips (Figures 4, 5). Transposon activity plays a unique role in species diversification and gene function variation, even in species with narrower gene pools (Domínguez et al., 2020). Specifically, genic transposable element (TE) insertions can lead to the production of multiple transcriptional isoforms, thereby increasing the diversity of gene expression (Aktaş et al., 2017; Vandecraen et al., 2017). As a tertiary relict gymnosperm (Yin Q.-Y. et al., 2021), transposon activity in the evolutionary history of C. hodginsii has provided significant momentum (Dubin et al., 2018; Lisch, 2012). In the context of dramatic global climate changes, active transposons may offer ongoing dynamism for the environmental adaptability of C. hodginsii, facilitating phenotypic variation.
Precipitation and sunlight significantly influence tree architecture, where smaller branch angles and shorter branch lengths help reduce the risk of photodamage and cavitation in the water transport system (Lines et al., 2012; Ceulemans et al., 1990; Haworth and McPherson, 1995). Our findings indicate a negative correlation between changes in Precipitation Seasonality and both the CW and Average Angle of Dips of C. hodginsii (Figure 5). This suggests that C. hodginsii from regions with more seasonal precipitation may be more sensitive to extreme drought conditions.
Additionally, transposon activity may mediate the regulation of gene expression associated with plant hormone signaling and branch development, thereby indirectly influencing branching patterns (Lisch, 2009). For instance, a class of aromatic plant hormones, such as strigolactones, can regulate plant architecture (Gomez-Roldan et al., 2008; Crawford et al., 2010). Therefore, we speculate that transposon activity may regulate the branching pattern of C. hodginsii by influencing the expression of genes in hormone signaling pathways that regulate plant architecture, thereby helping it better adapt to its growing environment.
While our study provides insights into the potential interactions between environmental factors, transposon activity, and plant hormones, the way these elements collectively influence the adaptive development of relict plants like C. hodginsii requires further investigation. In particular, understanding how C. hodginsii utilizes these mechanisms to adapt to changing environments remains an open question. Future research should delve into the specific mechanisms of these interactions through genetic and physiological experiments to deepen our understanding of plant adaptive evolution. Moreover, considering that transcriptional regulatory networks and epigenetic mechanisms may be associated with transposon activity (Wan et al., 2022), exploring their roles in rapid environmental adaptation will be crucial. This necessitates not only broader genomic data but also specific studies on the role of epigenetic modifications in adaptive development.
The datasets presented in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found: https://ngdc.cncb.ac.cn/gsub/submit/gsa/subCRA023801, PRJCA023136.
T-YH: Data curation, Investigation, Writing – review & editing. P-KZ: Conceptualization, Data curation, Formal analysis, Writing – original draft. C-LS: Investigation, Writing – review & editing. MW: Investigation, Writing – review & editing. J-NL: Investigation, Writing – review & editing. Y-PX: Investigation, Writing – review & editing. J-LZ: Investigation, Writing – review & editing. Y-HZ: Funding acquisition, Writing – review & editing. J-DR: Data curation, Writing – review & editing. KL: Data curation, Writing – review & editing. L-GC: Funding acquisition, Writing – review & editing. Y-SZ: Resources, Writing – review & editing. L-YC: Supervision, Writing – review & editing.
The author(s) declare that financial support was received for the research and/or publication of this article. This work was supported by National Key Research and Development Program of China [2021YFD2200501]; Scientific Research Project of Fujian Province [2023 J01478]; Fujian Forestry Nursery Technology Tackle Key Issues Project Phase Seven [LZKG202207]; and Forestry Peak Discipline Construction Project from Fujian Agriculture and Forestry University [72202200205]. The funding agency was not involved in the design of the study, collection, analysis, interpretation of data and writing the manuscript.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Gen AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
The Supplementary material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/ffgc.2025.1557824/full#supplementary-material
Ahammed, G., Li, X., Liu, A., and Chen, S. (2020). Brassinosteroids in plant tolerance to abiotic stress. J. Plant Growth Regul. 39, 1451–1464. doi: 10.1007/s00344-020-10098-0
Ahammed, G., Xia, X., Li, X., Shi, K., Yu, J., and Zhou, Y. (2015). Role of brassinosteroid in plant adaptation to abiotic stresses and its interplay with other hormones. Curr. Protein Pept. Sci. 16, 462–473. doi: 10.2174/1389203716666150330141427
Aktaş, T., Avşar Ilık, İ., Maticzka, D., Bhardwaj, V., Pessoa Rodrigues, C., Mittler, G., et al. (2017). DHX9 suppresses RNA processing defects originating from the Alu invasion of the human genome. Nature 544, 115–119. doi: 10.1038/nature21715
Anderson, J. T., Willis, J. H., and Mitchell-Olds, T. (2011). Evolutionary genetics of plant adaptation. Trends Genet. 27, 258–266. doi: 10.1016/j.tig.2011.04.001
Berger, D., Bauerfeind, S. S., Blanckenhorn, W. U., and Schäfer, M. A. (2011). High temperatures reveal cryptic genetic variation in a polymorphic female sperm storage organ. Evolution 65, 2830–2842. doi: 10.1111/j.1558-5646.2011.01392.x
Bisbing, S. M., Urza, A. K., Buma, B. J., Cooper, D. J., Matocq, M., and Angert, A. L. (2021). Can long-lived species keep pace with climate change? Evidence of local persistence potential in a widespread conifer. Divers. Distrib. 27, 296–312. doi: 10.1111/ddi.13191
Capblancq, T., and Forester, B. R. (2021). Redundancy analysis: a Swiss Army knife for landscape genomics. Methods Ecol. Evol. 12, 2298–2309. doi: 10.1111/2041-210X.13722
Ceulemans, R., Stettler, R. F., Hinckley, T. M., Isebrands, J. G., and Heilman, P. E. (1990). Crown architecture of Populus clones as determined by branch orientation and branch characteristics. Tree Physiol. 7, 157–167. doi: 10.1093/treephys/7.1-2-3-4.157
Chen, Z.-H., and Soltis, D. E. (2020). Evolution of environmental stress responses in plants. Plant Cell Environ. 43, 2827–2831. doi: 10.1111/pce.13922
Chen, S., Zhou, Y., Chen, Y., and Gu, J. (2018). Fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890. doi: 10.1093/bioinformatics/bty560
Crawford, S. E., Shinohara, N., Sieberer, T., Williamson, L., George, G., Hepworth, J., et al. (2010). Strigolactones enhance competition between shoot branches by dampening auxin transport. Development 137, 2905–2913. doi: 10.1242/dev.051987
Dauphin, B., Rellstab, C., Schmid, M., Zoller, S., Karger, D. N., Brodbeck, S., et al. (2021). Genomic vulnerability to rapid climate warming in a tree species with a long generation time. Glob. Chang. Biol. 27, 1181–1195. doi: 10.1111/gcb.15469
Ding, Y., Shi, Y., and Yang, S. (2020). Molecular regulation of plant responses to environmental temperatures. Mol. Plant 13, 544–564. doi: 10.1016/j.molp.2020.02.004
Dobin, A., Davis, C. A., Schlesinger, F., Drenkow, J., Zaleski, C., Jha, S., et al. (2013). STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21. doi: 10.1093/bioinformatics/bts635
Domínguez, M., Dugas, E., Benchouaia, M., Leduque, B., Jiménez-Gómez, J. M., Colot, V., et al. (2020). The impact of transposable elements on tomato diversity. Nat. Commun. 11:4058. doi: 10.1038/s41467-020-17874-2
Dubin, M., Scheid, O. M., and Becker, C. (2018). Transposons: a blessing curse. Curr. Opin. Plant Biol. 42, 23–29. doi: 10.1016/j.pbi.2018.01.003
Eizaguirre, C., and Baltazar-Soares, M. (2014). Evolutionary conservation—evaluating the adaptive potential of species. Evol. Appl. 7, 963–967. doi: 10.1111/eva.12227
Ellis, N., Smith, S. J., and Pitcher, C. R. (2012). Gradient forests: calculating importance gradients on physical predictors. Ecology 93, 156–168. doi: 10.1890/11-0252.1
Erb, M., Veyrat, N., Robert, C. A. M., Xu, H., Frey, M., Ton, J., et al. (2015). Indole is an essential herbivore-induced volatile priming signal in maize. Nat. Commun. 6:6273. doi: 10.1038/ncomms7273
Fick, S. E., and Hijmans, R. J. (2017). WorldClim 2: new 1-km spatial resolution climate surfaces for global land areas. Int. J. Climatol. 37, 4302–4315. doi: 10.1002/joc.5086
Fournier-Level, A., Korte, A., Cooper, M. D., Nordborg, M., Schmitt, J., and Wilczek, A. M. (2011). A map of local adaptation in Arabidopsis thaliana. Science 334, 86–89. doi: 10.1126/science.1209271
Gomez-Roldan, V., Fermas, S., Brewer, P. B., Puech-Pagès, V., Dun, E. A., Pillot, J.-P., et al. (2008). Strigolactone inhibition of shoot branching. Nature 455, 189–194. doi: 10.1038/nature07271
Grossiord, C., Buckley, T. N., Cernusak, L. A., Novick, K. A., Poulter, B., Siegwolf, R. T. W., et al. (2020). Plant responses to rising vapor pressure deficit. New Phytol. 226, 1550–1566. doi: 10.1111/nph.16485
Haworth, K., and McPherson, G. R. (1995). Effects ofQuercus emoryitrees on precipitation distribution and microclimate in a semi-arid savanna. J. Arid Environ. 31, 153–170. doi: 10.1006/jare.1995.0057
He, W., Sun, B., and Liu, Y.-S. (2012). Fokienia shengxianensis sp. nov. (Cupressaceae) from the late Miocene of eastern China and its paleoecological implications. Rev. Palaeobot. Palynol. 176-177, 24–34. doi: 10.1016/j.revpalbo.2012.03.013
Hickman, A., and Dyda, F. (2015). Mechanisms of DNA transposition. Microbiol. Spectr. 3:MDNA3-0034–2014. doi: 10.1128/microbiolspec.MDNA3-0034-2014
Hickman, A., and Dyda, F. (2016). DNA transposition at work. Chem. Rev. 116, 12758–12784. doi: 10.1021/acs.chemrev.6b00003
Holliday, J. A., Aitken, S. N., Cooke, J. E. K., Fady, B., González-Martínez, S. C., Heuertz, M., et al. (2017). Advances in ecological genomics in forest trees and applications to genetic resources conservation and breeding. Mol. Ecol. 26, 706–717. doi: 10.1111/mec.13963
Huang, Q., and Wang, Y. (2016). Overexpression of TaNAC2D displays opposite responses to abiotic stresses between seedling and mature stage of transgenic Arabidopsis. Front. Plant Sci. 7:1754. doi: 10.3389/fpls.2016.01754
Hufford, K., and Mazer, S. (2003). Plant ecotypes: genetic differentiation in the age of ecological restoration. Trends Ecol. Evol. 18, 147–155. doi: 10.1016/S0169-5347(03)00002-8
Hummel, I., El-Amrani, A., Gouesbet, G., Hennion, F., and Couée, I. (2004). Involvement of polyamines in the interacting effects of low temperature and mineral supply on Pringlea antiscorbutica (Kerguelen cabbage) seedlings. J. Exp. Bot. 55, 1125–1134. doi: 10.1093/JXB/ERH126
Innes, P. A., Goebl, A. M., Smith, C. C. R., Rosenberger, K., and Kane, N. C. (2023). Gene expression and alternative splicing contribute to adaptive divergence of ecotypes. Heredity 132, 120–132. doi: 10.1038/s41437-023-00665-y
Isabel, N., Holliday, J. A., and Aitken, S. N. (2020). Forest genomics: advancing climate adaptation, forest health, productivity, and conservation. Evol. Appl. 13, 3–10. doi: 10.1111/eva.12902
Kan, Y., Mu, X.-R., Gao, J., Lin, H.-X., and Lin, Y. (2023). The molecular basis of heat stress responses in plants. Mol. Plant 16, 1612–1634. doi: 10.1016/j.molp.2023.09.013
Liao, Y., Smyth, G. K., and Shi, W. (2014). featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930. doi: 10.1093/bioinformatics/btt656
Lines, E. R., Zavala, M. A., Purves, D. W., and Coomes, D. A. (2012). Predictable changes in aboveground allometry of trees along gradients of temperature, aridity and competition. Glob. Ecol. Biogeogr. 21, 1017–1028. doi: 10.1111/j.1466-8238.2011.00746.x
Lisch, D. (2009). Epigenetic regulation of transposable elements in plants. Annu. Rev. Plant Biol. 60, 43–66. doi: 10.1146/annurev.arplant.59.032607.092744
Lisch, D. (2012). How important are transposons for plant evolution? Nat. Rev. Genet. 14, 49–61. doi: 10.1038/nrg3374
López-Maury, L., Marguerat, S., and Bähler, J. (2008). Tuning gene expression to changing environments: from rapid responses to evolutionary adaptation. Nat. Rev. Genet. 9, 583–593. doi: 10.1038/nrg2398
Ma, Y., Wang, J., Hu, Q., Li, J., Sun, Y., Zhang, L., et al. (2019). Ancient introgression drives adaptation to cooler and drier mountain habitats in a cypress species complex. Commun. Biol. 2:213. doi: 10.1038/s42003-019-0445-z
McIver, E. E., and Basinger, J. F. (1990). Fossil seed cones of Fokienia (Cupressaceae) from the Paleocene Ravenscrag formation of Saskatchewan, Canada. Can. J. Botany 68, 1609–1618. doi: 10.1139/b90-207
Medrano, M., Alonso, C., Bazaga, P., López, E., and Herrera, C. (2020). Comparative genetic and epigenetic diversity in pairs of sympatric, closely related plants with contrasting distribution ranges in south-eastern Iberian mountains. AoB Plants 12:plaa013. doi: 10.1093/aobpla/plaa013
Nepstad, D., Tohver, I., Ray, D. G., Moutinho, P., and Cardinot, G. (2007). Mortality of large trees and lianas following experimental drought in an Amazon forest. Ecology 88, 2259–2269. doi: 10.1890/06-1046.1
Oksanen, J., Gavin, L., Simpson, F. G. B., Kindt, R., Legendre, P., Minchin, P. R., et al. (2017). Vegan: Community ecology package. Available online at: https://cran.r-project.org/web/packages/vegan/index.html (Accessed February 20, 2024).
Poncet, B., Herrmann, D., Gugerli, F., Taberlet, P., Holderegger, R., Gielly, L., et al. (2010). Tracking genes of ecological relevance using a genome scan in two independent regional population samples of Arabis alpina. Mol. Ecol. 19, 2896–2907. doi: 10.1111/j.1365-294X.2010.04696.x
Porporato, A., Laio, F., Ridolfi, L., and Rodríguez-Iturbe, I. (2001). Plants in water-controlled ecosystems: active role in hydrologic processes and response to water stress: III. Vegetation water stress. Adv. Water Resour. 24, 725–744. doi: 10.1016/S0309-1708(01)00006-9
Qian, Y. (2022). “China: China and the 2015–2016 El Niño” in El Niño ready nations and disaster risk reduction. ed. M. H. Glantz (Cham: Springer International Publishing), 29–42.
Razgour, O., Forester, B., Taggart, J. B., Bekaert, M., Juste, J., Ibáñez, C., et al. (2019). Considering adaptive genetic variation in climate change vulnerability assessment reduces species range loss projections. Proc. Natl. Acad. Sci. 116, 10418–10423. doi: 10.1073/pnas.1820663116
Rezaie, N., Reese, F., and Mortazavi, A. (2023). PyWGCNA: a Python package for weighted gene co-expression network analysis. Bioinformatics 39:btad415. doi: 10.1093/bioinformatics/btad415
Rong, J., Zheng, Y., Zhang, Z., Zhang, J., Gu, Y., Hua, T., et al. (2024). De novo whole-genome assembly of the 10-Gigabase Fokienia hodginsii genome to reveal differential epigenetic events between callus and xylem. Adv. Sci. 11:2644. doi: 10.1002/advs.202402644
Ruffault, J., Martin-StPaul, N. K., Rambal, S., and Mouillot, F. (2013). Differential regional responses in drought length, intensity and timing to recent climate changes in a Mediterranean forested ecosystem. Clim. Chang. 117, 103–117. doi: 10.1007/s10584-012-0559-5
Sang, Y., Long, Z., Dan, X., Feng, J., Shi, T., Jia, C., et al. (2022). Genomic insights into local adaptation and future climate-induced vulnerability of a keystone forest tree in East Asia. Nat. Commun. 13:6541. doi: 10.1038/s41467-022-34206-8
Tenenhaus, M., Vinzi, V. E., Chatelin, Y.-M., and Lauro, C. (2005). PLS path modeling. Comput. Statist. Data Anal. 48, 159–205. doi: 10.1016/j.csda.2004.03.005
Thomas, P., and Yang, Y. (2013). Fokienia hodginsii. The IUCN Red List of Threatened Species 2013: e.T32351A2815809. (Accessed March 09 2025).
Vandecraen, J., Chandler, M., Aertsen, A., and Houdt, R. V. (2017). The impact of insertion sequences on bacterial genome plasticity and adaptability. Crit. Rev. Microbiol. 43, 709–730. doi: 10.1080/1040841X.2017.1303661
Walther, G.-R., Post, E., Convey, P., Menzel, A., Parmesan, C., Beebee, T. J. C., et al. (2002). Ecological responses to recent climate change. Nature 416, 389–395. doi: 10.1038/416389a
Wan, T., Gong, Y., Liu, Z., Zhou, Y., Dai, C., and Wang, Q. (2022). Evolution of complex genome architecture in gymnosperms. GigaScience 11:giac078. doi: 10.1093/gigascience/giac078
Weeks, A., Sgrò, C., Young, A., Frankham, R., Mitchell, N., Miller, K. A., et al. (2011). Assessing the benefits and risks of translocations in changing environments: a genetic perspective. Evol. Appl. 4, 709–725. doi: 10.1111/j.1752-4571.2011.00192.x
Xu, Q., Zhu, C., Fan, Y., Song, Z., Xing, S., Liu, W., et al. (2016). Population transcriptomics uncovers the regulation of gene expression variation in adaptation to changing environment. Sci. Rep. 6:25536. doi: 10.1038/srep25536
Yang, H., Li, J., Milne, R. I., Tao, W., Wang, Y., Miao, J., et al. (2022). Genomic insights into the genotype-environment mismatch and conservation units of a Qinghai-Tibet plateau endemic cypress under climate change. Evol. Appl. 15, 919–933. doi: 10.1111/eva.13377
Yin, Q., Chen, S., Guo, W., Huang, Y., Huang, Y., Zhou, R., et al. (2018). Pronounced genetic differentiation in Fokienia hodginsii revealed by simple sequence repeat markers. Ecol. Evol. 8, 10938–10951. doi: 10.1002/ece3.4560
Yin, Q.-Y., Fan, Q., Li, P., Truong, D., Zhao, W.-Y., Zhou, R.-C., et al. (2021). Neogene and quaternary climate changes shaped the lineage differentiation and demographic history of Fokienia hodginsii (Cupressaceae s.l.), a tertiary relict in East Asia. J. Syst. Evol. 59, 1081–1099. doi: 10.1111/jse.12582
Ying, G. S., and Fu, Z. G. (2002). Oligocene Sanhe flora in Longjing county of Jilin, Northeast China. Acta Palaeontol. Sin 41, 193–210. doi: 10.3969/j.issn.0001-6616.2002.02.004
Keywords: coniferous tree, environmental changes, gene expression, Chamaecyparis hodginsii, adaptability, seed experiments
Citation: He T-Y, Zhu P-K, Song C-L, Wu M, Lin J-N, Xu Y-P, Zhang J-L, Zhang Y-H, Rong J-D, Liu K, Chen L-G, Zheng Y-S and Chen L-Y (2025) Gene expression mediates environmentally induced phenotypic plasticity in Chamaecyparis hodginsii seedlings. Front. For. Glob. Change. 8:1557824. doi: 10.3389/ffgc.2025.1557824
Received: 09 January 2025; Accepted: 03 March 2025;
Published: 17 March 2025.
Edited by:
Pedro Giovâni Da Silva, Federal University of Minas Gerais, BrazilReviewed by:
Min Liu, Guangzhou University, ChinaCopyright © 2025 He, Zhu, Song, Wu, Lin, Xu, Zhang, Zhang, Rong, Liu, Chen, Zheng and Chen. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Ling-Yan Chen, ZmFmdWNseUBmYWZ1LmVkdS5jbg==; Peng-Kai Zhu, cGt6aHUyMjJAZ21haWwuY29t
†These authors have contributed equally to this work
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.