
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Environ. Sci. , 02 April 2025
Sec. Environmental Informatics and Remote Sensing
Volume 13 - 2025 | https://doi.org/10.3389/fenvs.2025.1577298
 Jamshid Ali1
Jamshid Ali1 Wang Haoran1
Wang Haoran1 Kaleem Mehmood2
Kaleem Mehmood2 Wakeel Hussain3
Wakeel Hussain3 Farhan Iftikhar4
Farhan Iftikhar4 Fahad Shahzad5
Fahad Shahzad5 Khadim Hussain6
Khadim Hussain6 Yin Qun1*
Yin Qun1* Jia Zhongkui1*
Jia Zhongkui1*Estimating above-ground biomass (AGB) is important for ecological assessment, carbon stock evaluation, and forest management. This research assesses the performance of the machine learning algorithms XGBoost, SVM, and RF using data from the Sentinel-2 and Landsat-9 satellites. The study assesses the influence of the significant spectral bands and vegetation indices on the accuracy of the AGB estimate. The results presented in the paper indicate that Sentinel-2 data were more effective than Landsat-9 data. This is mainly because it had higher spatial and spectral resolution, which enabled the model vegetation gradients and structural attributes more accurately. The XGBoost model performed the best with an R2 of 0.82 and RMSE of 0.73 Mg/ha with Sentinel-2 and R2 of 0.80 and RMSE of 0.71 Mg/ha with Landsat-9. In the current study, SVM also showed a substantial accuracy with an R2 of 0.79 and RMSE of 0.73 Mg/ha for Sentinel-2 and R2 of 0.76 and RMSE of 0.80 Mg/ha for Landsat-9. For Sentinel-2, the random forest achieved an R2 of 0.74 and an RMSE of 0.93 Mg/ha, and Landsat 9 yielded an R2 of 0.72 and an RMSE of 0.88 Mg/ha. Thus, using variable importance analysis, the results showed that vegetation indices and spectral bands have higher importance in predicting AGB. As expected from their application in biomass research, these predictors consistently emerged as highly significant across models and datasets. This study demonstrates the potential of integrating machine learning with remote sensing data to achieve accurate and efficient biomass assessment.
Forests are crucial for sustaining biodiversity, as they offer crucial habitats that sustain diverse biodiversity. They are essential for sustaining ecological balance and enhancing biodiversity (Ali et al., 2023; Stephenson and Damerell, 2022). Therefore, an estimate of the forestry biomass determines a given ecosystem’s ability to capture carbon and maintain a stable carbon stock (Jafri et al., 2022; Wang et al., 2022). Accurately assessing forest biomass is critical for analyzing the global carbon cycle and addressing numerous concerns, including climate change, Forest strength, and service regulation (Hu et al., 2022; Titus et al., 2021).
Conventional field measurements or remote sensing techniques often evaluate AGB in forests (Santoro et al., 2021). Satellite imagery is better than traditional forest inventories and surveys using LiDAR technology because it can cover larger areas at a lower cost and in less time (López Serrano et al., 2022). Integrating reference values within satellite data is a significant process in estimating aboveground biomass or forest inventories from airborne LiDAR more precisely (Campbell et al., 2021; Labrière et al., 2023).
The next step involves employing spatial prediction algorithms to generate precise geographic proportions of AGB (Das et al., 2024; Sun et al., 2023). Researchers have made significant progress in mapping forest AGB by combining modeling methods with better predictor applications from satellite data (Hojo et al., 2023). Past research has shown that a combination of diverse remote sensing techniques has successfully quantified and monitored biomass from forests at a regional level (Coops et al., 2023; Zhang and Shao, 2021). Consequently, current research indicates that diverse remote sensing methods, encompassing both passive and active sensors, can estimate AGB in a designated area (Ma et al., 2024). Researchers frequently utilize optical remote sensing imagery, distinguished by spatial, spectral, and temporal resolutions, to assess AGB at diverse scales (Sedano et al., 2021). Researchers primarily employ moderate and coarse-resolution data from the Moderate Resolution Imaging Spectroradiometer (MODIS) (Shahzad et al., 2025; Wongsai et al., 2020).
When looking at AGB on a small scale, medium-resolution data from the Sentinel-2 and Landsat satellites is needed, when using High-quality commercial satellite data from IKONOS, QuickBird, and WorldView-2, it is possible to get a pretty good idea of AGB at the forest stand-level (Fu et al., 2022; Lin et al., 2022). To estimate AGB at the regional level with average spatial resolution, microwave radar remote sensing data is required. These include synthetic aperture radar (SAR), interferometric SAR (InSAR), and polarimetric interferometric SAR (PolInSAR) data (Godinho Cassol et al., 2021; Ramachandran et al., 2023). The first and essential step in constructing accurate models for predicting AGB is indicating the correct algorithm (Araza et al., 2022; Li et al., 2021).
Previous research has frequently used the traditional statistical regression method for aboveground biomass (AGB) estimation despite its simplicity and ease of computation (Luo et al., 2024). This method employs a regression model that combines test data and remote-sensing features (Han et al., 2019; Hussain et al., 2024). It does not capture the correlation between forest AGB and RS data (Zhang et al., 2022). Standard methodologies for predicting and mapping AGB include interpolation techniques, non-parametric models, and kriging. Researchers have utilized geo statistics for AGB data to examine variations and develop sample designs for satellite and field-based forest monitoring (Li et al., 2020b; Su et al., 2020). It is challenging to map the continuous forest characteristics in large, steep areas. Important site factors like soil type, texture, nutrient status, solar flux density, moisture regime, and water holding capacity affect the key tree attributes within different stand types in terms of Diameter at breast height, height, and volume. When establishing the inventory, measurements of forest trees and AGB showed spatial dependency within small areas of stand types (Carmenta et al., 2020; Octavia et al., 2022).
However, this spatial autocorrelation varies depending on the community’s topographical conditions, residential zones, and the locations of commercial logging activities (Gibson, 2018; Shahzad et al., 2024). For AGB, it is clear that several studies have pointed out the integration of remote sensing technology with geostatistical and machine learning methods (Musekiwa et al., 2022; Prăvălie et al., 2023). This combination is especially advantageous for forecasting extensive regions characterized by diverse bioclimatic conditions and irregular terrain (Masereti Makori et al., 2024).
Remote sensing-based AGB estimation uses machine learning techniques, including decision trees, random forests, and support vector regression. These strategies improve the model’s ability to provide accurate biomass predictions, mainly where nonlinearity is a key reason. The literature published in the last decade reveals that decision tree-based algorithms like Random Forest (RF) and Gradient Boosting (GB) yield high accuracy in biomass estimation modeling (Cameron et al., 2022; Ghasemloo et al., 2022). Moreover, machine learning techniques encompass numerous adjustable hyperparameters significantly influencing the models. The adjustment of these settings has, at times, been overlooked. Previous studies have shown that the tuning procedure significantly influences the model performance, with the sensitivity of parameters varying between stochastic gradient boosting and random forests (Freeman et al., 2015; Li et al., 2020a; Prakash et al., 2022).
Research gaps persist in integrating RS data with machine learning models (XGBoost, SVM, and Random Forest) for biomass prediction despite applying RS data, particularly Sentinel-2 and Landsat-9 data, in biomass estimation. While earlier literature has established the superiority of high spatial resolution imagery such as Sentinel-2 over low spatial resolution imagery such as Landsat-9, there is still a lack of comparative research on how these datasets perform under different environmental conditions, especially in forests with structural homogeneity and a 50-year-old Larix principis-ruprechtii in Northern China. Some studies have established the significance of vegetation indices like NDVI, TNDVI, and NDI45 as biomass predictors. However, they have not comprehensively studied their contribution to machine learning algorithms such as XGBoost, SVM, and Random Forest. However, there is still a lack of comprehensive investigation into applying indices like GNDVI and NDI45 for biomass modeling, particularly in temperate forests.
The present study is expected to enhance the precision and generality of the AGB estimation in L. principis-rupprechtii Mayr plantations at the Saihanba Mechanical Forest Farm in Hebei Province, northern China. This is accomplished by integrating machine learning techniques with remote sensing data. This research aims to evaluate the performance of three popular machine learning algorithms, namely XGBoost, Support Vector Machine (SVM), and Random Forest (RF), to estimate AGB using Sentinel-2 and Landsat-9 data.
The study area included the Saihanba Forest Farm, which is located in Hebei Province, northern China, and ranges (41°22′– 42°58′N, 116°53′– 118°3′W). The research site is in the warm temperate continental monsoon climate zone. The altitude of the area is (1,010 ∼ 1,940 m). The mean annual temperature is (−1.2°C), and the average annual temperature range is from (−43.3°C–33.4°C). The annual rainfall is (452.2 mm), and the annual evaporation is 1,388 mm.
The typical soil types in the region include aeolian sandy soil, meadow soil, brown soil, and grey forest soil. The total operating area is 94,000 ha, of which the forest area is 73,333 ha, planted forest 57,333 ha, and natural forest 16,000 ha; the forest coverage rate is 80%, total forest volume is 5.025 million m3.
The most important vegetation zones include grassland, meadow, conifer and broad-leaved mixed forest, broad-leaved forest, and shrub forest; the forest density is 75.5%. The main trees are L. principis-rupprechtii, Picea asperata Mast., and Betula platyphylla Suk., and the main shrubs are Rhododendron micranthum Turcz., Syringa oblata Lindl. var. alba Rehder., and Sambucus williamsii Hance. The main herbaceous plants are Galium verum L. and Menyanthes trifoliata L. (Tao et al., 2023; Xu et al., 2022) (Figure 1).
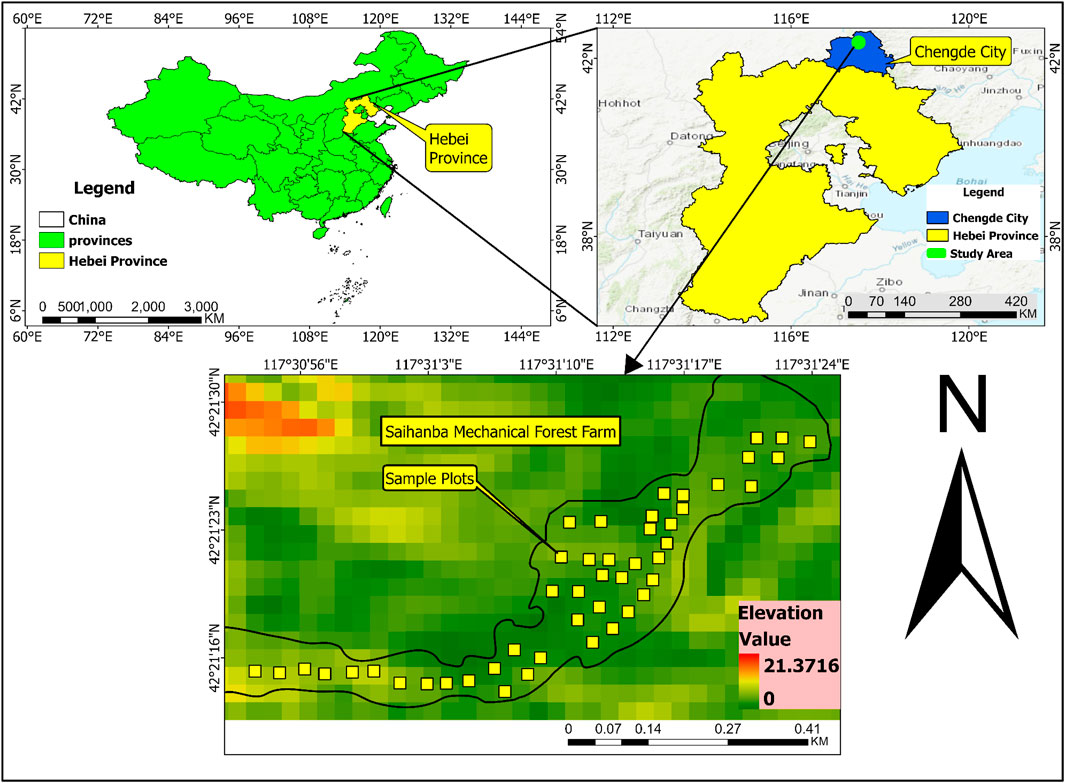
Figure 1. Location of the study sites in China, Chengde City, Hebei Province, Saihanba Mechanical Forest Farm.
We conducted a study for the forest inventory in August 2023. We meticulously chose the sampling spots, excluding non-forest regions. The study set up 45 sampling plots in total for the 50-year-old L. principis-rupprechtii Mayr plantation. We recorded the coordinates of each tree and plot, using Real-Time Kinematic (RTK). For analysis, we recorded each plot’s elevation, aspect, slope, height (in meters), stem density (trees per hectare), and DBH, measured 1.3 m above the ground using a calibrated diameter tape and caliper. Stem density was determined by counting the number of trees within each plot. Tree heights were measured using a Relascope (Almeida et al., 2021). Soil samples were obtained from the upper 20 cm layer using a soil auger to determine soil organic carbon (SOC) (Liu N. et al., 2021). These samples were put in plastic bags, allowed to air dry, and then taken to a laboratory for further tests.
To measure total biomass distribution, we use allometric equations of all tree elements, such as stem, branches, leaves, and roots. It allows for precise calculations of the distribution of above and belowground biomass (Zhao et al., 2016). To estimate the AGC and BGC amount, the obtained AGB and BGB values were multiplied by 0.5, assuming that the total amount of aboveground and belowground biomass had a 50% carbon content (Aye et al., 2022; Eshetu and Hailu, 2020), (Figure 2), (Supplementary Figure S3).
The European Space Agency’s Sentinel-2 provides medium-resolution multispectral imagery for Earth observation. Using the Google Earth Engine (GEE) platform, acquired and pre-processed Sentinel-2A and Landsat 9 images for the study area (https://earthengine.google.com/). The Sentinel-2A data were ortho-corrected bottom-atmospheric reflectance images, with Bands 2, 3, 4, and 8 selected for analysis, while Bands 1, 5, 6, and 9 were excluded due to their relevance to atmospheric correction and hydrological applications. Preprocessing included filtering out images with cloud cover exceeding 5%, performed using the Sen2Cor processor for Sentinel-2 Level-1C products. Cloud-covered pixels were identified, masked, and corrected accordingly. For Landsat 9, to do a complete analysis, carefully pick and extract (4) bands (Band 2, Band 3, Band 4, Band 5) that are thought to be important for lowering errors in estimating forest AGB and making comparisons useful. The images with less than 5% cloud cover were retained. We applied the CFMask algorithm, integrated into the Landsat Surface Reflectance product, to mask cloud pixels, replacing them with maximum value composites for data consistency. Both datasets were resampled to a common spatial resolution (10 m for Sentinel-2 and 30 m for Landsat 9) using bilinear interpolation and aligned to the WGS84 coordinate system. The final preprocessed data were split into training (70%) and validation (30%) samples. Comprehensive processing of Landsat 9 included orthorectification, georectification, and registration, ensuring high-quality data. (Li et al., 2024), (Table 1), (Figure 3).
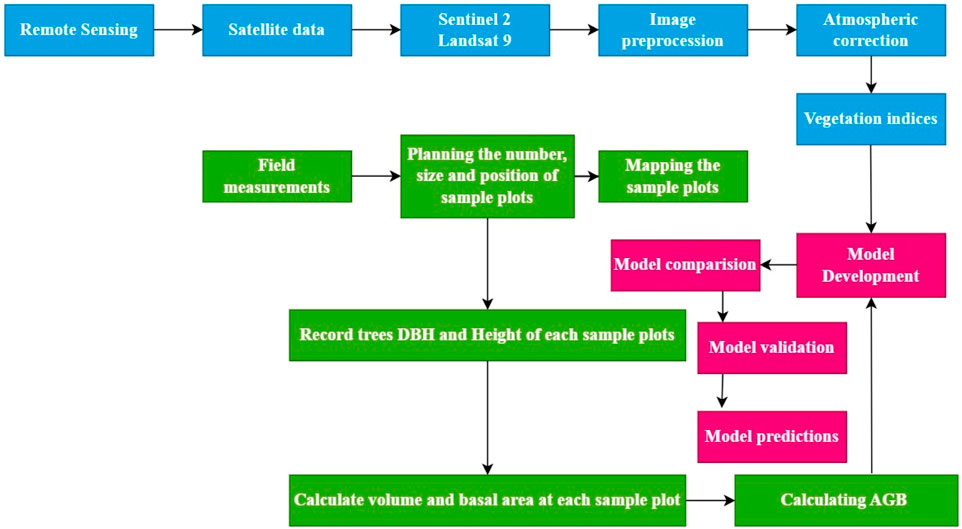
Figure 3. A flowchart for evaluating Vegetation Indices and modeling algorithms for mapping forest Above Ground Biomass using Sentinel 2 and Landsat 9 data.
Diverse methodological strategies are utilized to obtain field remote-sensing data (Aslam et al., 2024). In this study, the coordinates of the southwestern corner of each field plot were used to define the center point, serving as the geographic anchor for plot-level remote sensing data extraction. Satellite imagery from Sentinel-2 and Landsat 9 was resampled to match the spatial extent of the field plots as closely as possible. Despite these efforts, minor spatial mismatches remained between the pixel grid and plot boundaries due to geometric distortion, sensor resolution, and terrain variability.
To reduce the effect of such spatial discrepancies, circular buffer zones were applied: a 10-m radius for Sentinel-2 (10 m resolution) and a 30-m radius for Landsat 9 (30 m resolution) around each plot center. These buffer radii were selected to balance the minimization of geolocation error with the need to avoid spectral contamination from adjacent land covers with contrasting canopy structures. The average pixel value within each buffer was extracted to represent the spectral signal associated with each plot’s biophysical parameters (Turner et al., 2015). While the buffer-based averaging approach cannot eliminate all residual spatial discrepancies, it substantially reduces the influence of pixel-level misalignment by integrating spectral information across a spatial area representative of the plot. This methodological choice reflects a practical balance between spatial accuracy and ecological representativeness, commonly adopted in similar fields–remote sensing integration studies. We acknowledge that some degree of residual uncertainty may persist, but this was considered during model interpretation and is not expected to significantly bias the results at the scale of analysis.
The machine learning techniques were chosen because they can handle the complicated problems of Forest biomass estimation, where variables do not constantly interact in a straight line, there are many predictors, and there are many drivers. This makes their use, normalization, and insensitivity to outliers highly suitable. In the case of evaluating multisensory indices against field-measured AGB, we used Pearson’s product-moment correlation to perform a paired analysis (Chen et al., 2018).
We checked and analyzed the provided dataset to overcome the issue of multicollinearity. This approach included using the variance inflation factor (VIF) to determine whether any variables were redundant and, if so, to remove them (Mehmood et al., 2024a; Thompson et al., 2017). We systematically removed predictor variables with a coefficient magnitude of 0.8 or higher and high VIF values of 10 or more from the set in regression analysis (Kristensen et al., 2015; Pérez-Girón et al., 2020). The R Statistical Computation program performed the analytical operations (Table 2).
XGBoost, SVR, and RF should be used because these algorithms efficiently disentangle intricate and non-linear connections in natural systems such as the forest ecosystem (Zennaro et al., 2021). These algorithms can select several essential predictor factors independently and are well-equipped to handle datasets of high dimensionality. They are robust, can perform ensemble analysis, and employ state-of-the-art methods, aligning with our accurate AGB estimation objective. Detailed evaluation of several algorithms enhances the scientific credibility of the results and ensures the selection of the most appropriate approach for estimating AGB in temperate forests (Oehmcke et al., 2024; Pham et al., 2023).
XGBoost is the collective model for learning that includes gradient boosting and complicated regularization techniques to improve its predictiveness. It consistently generates new models to rectify errors in previous models while reducing a loss function given by the user through the second-order Taylor expansion. Researchers have found XGBoost highly effective for complex and multiple-variable predictive modeling due to its lack of missing values and overfitting problems (Mehmood et al., 2024b; Zhang and Jánošík, 2024).
The SVR regression method transforms the input data into the higher-dimensional space by the kernel function and attempts to minimize the prediction errors by shifting a hyperplane. This approach is beneficial in identifying non-linear relationships that might exist in the data. Different regression problems can employ SVR because it has low computational complexity and reasonable empirical risk (Hussain et al., 2025; Lee et al., 2020). Random Forest is a learning technique that integrates multiple decision trees through a process known as bagging. This technique grows each tree based on a bootstrap sample and employs the remaining data to compute the out-of-bag (OOB) error. To find the best partition, every node randomly chooses some explanatory variables. Environmental modeling and habitat suitability evaluations widely apply to random forests (RF) due to their applicability in classification and regression problems (Anees et al., 2024; Teng et al., 2023).
Some of the key parameters for hyperparameter tuning of the XGBoost model include features such as “nrounds”/“boosting iteration,” “max depth,” “min child weight,” “gamma,” and “subsample.” We used a grid search approach to enhance the model’s performance and identify the optimal parameters. The variable importance measures for XGBoost are “Gain” and “Frequency” (Asselman et al., 2023; Mehmood et al., 2025).
SVR tuning is done by choosing the kernel function and the “C” parameter that defines the width of the margin and permits misclassification. Thus, the authors have established that SVR’s flexibility allows for effectively handling complex decision limits (Mahmood et al., 2023; Shams et al., 2024). RF parameters include “ntree,” which measures the number of trees, and “mtry,” which indicates the number of features used randomly for splitting. Grid search optimized the values of these parameters. We measured the variable importance using the percent IncMSE and IncNodePurity (Bouslihim et al., 2024; Li Yudong et al., 2020) (Table 3).
The performance of the XGBoost, SVR, and RF algorithms for each variable group was then compared. The evaluations used the correlation coefficient r, RMAE, RRMSE, and mean error (MAE) to enable comparison based on the following Equations 1–5 (Bouslihim et al., 2024; Liu H. et al., 2021). The approach with the highest accuracy was assigned the predictive mapping of the AGB distribution for each variable group.
In Equations 1–5,
Table 4 presents a comprehensive statistical overview of a 50-year-old Larix principis-rupprechtii forest stand, providing insights into its structural attributes and biomass dynamics. Despite the homogeneity in species composition and age, the forest exhibits notable variability in several key parameters, reflecting the inherent complexity of natural systems. The mean DBH of 25.43 cm, accompanied by a standard deviation of 1.31 cm, suggests a moderate variation in tree size within the stand, typically ranging from 22.61 cm to 29.05 cm. Similarly, tree height, with a mean of 19.42 m and a relatively low standard deviation of 0.89 m, points to a uniform vertical structure, with tree heights distributed between 17.1 m and 21.64 m.
The stand density, averaging 677 trees per hectare, with an extensive standard deviation of 146 trees per hectare, highlights the variability in tree spacing and distribution, ranging from 450 to 1000 trees per hectare. The stand’s average AGB is 7.57 Mg/ha, with a standard deviation of 1.44 Mg/ha and values ranging from 5.07 to 10.29 Mg/ha. This variation in AGB reflects individual trees differing growth potential and carbon sequestration capacity within the stand (Table 4).
The correlation study of Sentinel-2 and Landsat-9 data provided key findings regarding the remote sensing predictors and AGB. For Sentinel-2, figure (A) with correlation coefficients varying from (0.32–0.58). Among the variables, vegetation indices emerged as the most significant predictors, particularly TNDVI and NDI45, showing correlations of (0.58). They emerged as the strongest predictors of AGB.
Other indices, such as NDVI (0.56) and GNDVI (0.52), also demonstrated strong correlations, reflecting their effectiveness in integrating vegetation density and photosynthetic activity—key factors influencing biomass accumulation. The NIR band (0.54) showed a strong correlation among individual spectral bands. Visible bands, such as Red (0.44) and Green (0.37), exhibited moderate correlations, with indices like SAVI (0.39) and MSAVI2 (0.42) showing secondary relevance. The WDVI (0.32) exhibited the weakest correlation.
In the case of Landsat-9 figure (B), the correlation coefficients varied between (0.30–0.56), with TNDVI (0.56), NDVI (0.53), and NDI45 (0.55) emerging as the top predictors of AGB. The band B5, (0.50) and band B4, (0.42) showed moderate correlations. However, soil background and atmospheric conditions often influence visible bands like B2 and B3, exhibiting weaker correlations (0.30) and (0.35). Some vegetation indices, such as PSSRa (0.43) and MSAVI2 (0.38), also showed moderate correlations, suggesting their complementary role in enhancing model performance.
The WDVI (0.30) displayed the weakest correlation, highlighting its limited predictive power for AGB in the context of Landsat-9 data. When comparing the two datasets, Sentinel-2 consistently outperformed Landsat-9 regarding correlation strength across all variables. Sentinel-2’s superior spatial and spectral resolution allows for precisely capturing vegetation characteristics like canopy structure, leaf area index, and photosynthetic activity. Stronger connections are seen between Sentinel-2-derived indices like TNDVI and NDVI, showing that it can accurately model AGB (Figure 4).
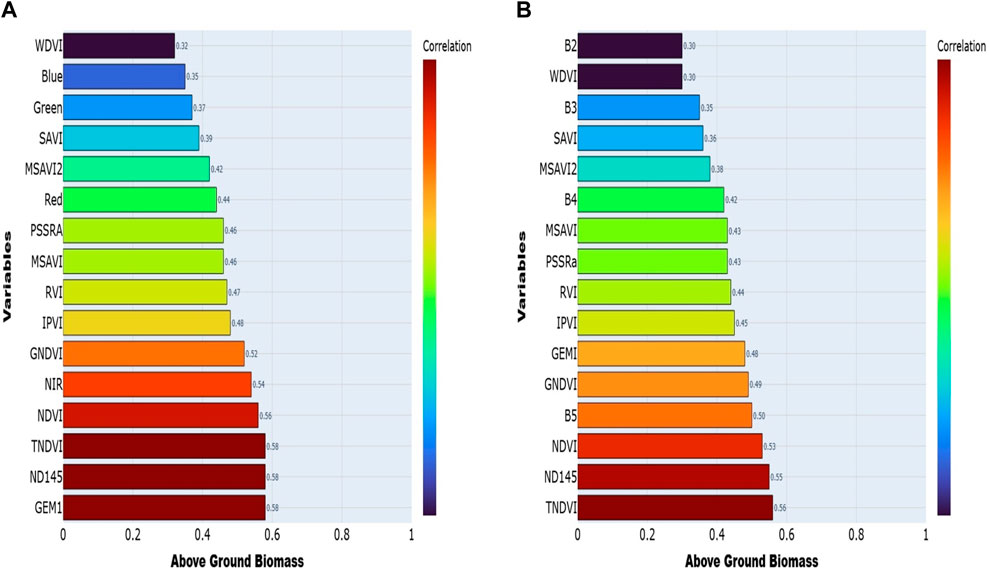
Figure 4. Demonstrates how different predictors relate to the field-measured biomass, Sentinel 2 (A) and Landsat 9 (B).
The variable importance analysis of predictors from Sentinel-2 and Landsat 9 images shows essential details about how they improve the performance of machine learning models such as XGBoost, SVM, and Random Forest (RF). The present work focuses on the effects of the spectral bands and vegetation indices on the model’s biomass and carbon stock assessment performance. Sentinel-2 shows that GNDVI and GEMI are the most influential predictors across all models, particularly within the XGBoost. GNDVI stands out due to its high sensitivity to plant properties, capturing subtle changes in vegetation vigor. It is a key variable for modeling above-ground biomass (AGB) and carbon stock because it can distinguish between important biophysical features like chlorophyll content and leaf area. Similarly, Landsat 9 highlights the ND145 index as the most important predictor, excelling particularly in XGBoost. ND145 can tell a lot about the health of plants; it can find changes in leaf area and chlorophyll content, which are important biophysical features for figuring out biomass and carbon stock. Both Sentinel-2 and Landsat 9 datasets reveal that indices such as SAVI, TNDVI, WDVI, PSSRa, and IPVI contribute moderately to the model’s performance. These indices provide valuable Supplementary Information on vegetation structure, density, and canopy properties, further refining biomass estimates.
In particular, WDVI and PSSRa in the Sentinel-2 and Landsat 9 datasets make notable contributions by capturing information related to vegetation moisture content and plant stress, which are important for biomass modeling under varying environmental conditions. To further clarify their relative importance, we quantified and compared the normalized variable importance scores of vegetation indices across the XGBoost, SVM, and RF models. This comparison revealed that although GNDVI consistently ranked highest in both datasets (e.g., 0.183 in Sentinel-2 and 0.064 in Landsat-9 using XGBoost), indices such as WDVI (Sentinel-2: 0.026 in XGBoost; Landsat-9: 0.045) and PSSRa (Sentinel-2: 0.030; Landsat-9: 0.045) demonstrated moderate yet model-consistent importance across all approaches. The systematic comparison also showed that WDVI and PSSRa ranked higher in RF and SVM models relative to XGBoost, indicating that their influence varies by algorithm but remains non-negligible. These findings are summarized in Supplementary Tables S1, S2, where the importance values of each VI across models are presented. This numerical evidence strengthens our interpretation of the ecological relevance of these indices in AGB estimation (Figure 5).
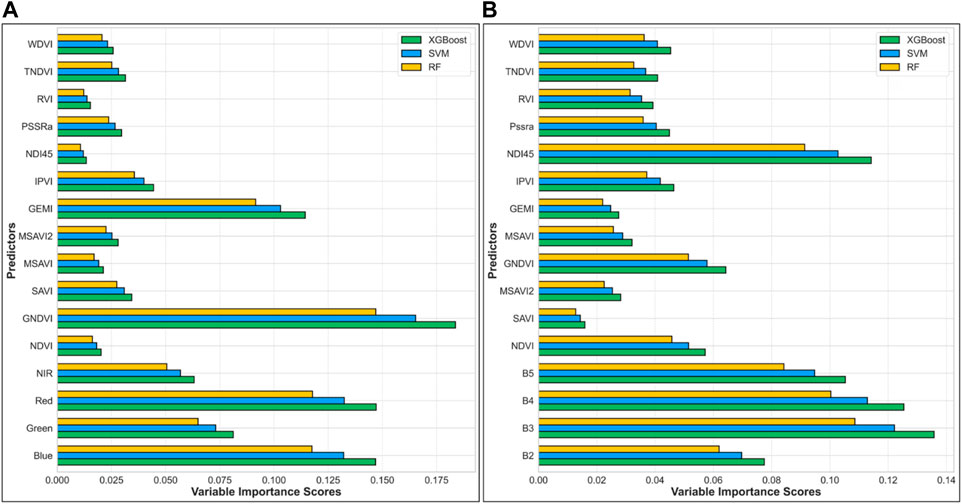
Figure 5. Shows the variable’s importance derived from XGboost, SVM, and RF from Sentinel 2 (A), Landsat 9 (B).
For this study, we used data from Sentinel-2 and Landsat-9 to compare how well three machine learning models—XGBoost, Support Vector Machine (SVM), and Random Forest (RF)—estimated AGB. Even though Landsat-9 has lower spatial and spectral resolution than Sentinel-2, both datasets helped estimate biomass, but Sentinel-2 consistently did better than Landsat-9.
XGBoost consistently delivered the best performance across both datasets. With Sentinel-2, it achieved a coefficient of determination R2 of 0.82, an RMSE of 0.73 Mg/ha, and an MAE of 0.60 Mg/ha, while for Landsat-9, it achieved an R2 of 0.80, an RMSE of 0.71 Mg/ha, and an MAE of 0.58 Mg/ha. Sentinel-2’s high spatial and spectral resolution made it easier for the model to pick up on small changes in canopy reflectance, vegetation structure, and biomass-related parameters, which led to lower error metrics. However, XGBoost maintained robust performance with Landsat-9, demonstrating adaptability across datasets with varying resolutions. The SVM model also exhibited strong performance, with Sentinel-2 achieving an R2 of 0.79, RMSE of 0.73 Mg/ha, and MAE of 0.63 Mg/ha, while Landsat-9 produced an R2 of 0.76, RMSE of 0.80 Mg/ha, and MAE of 0.66 Mg/ha.
SVM’s capacity to model non-linear relationships was evident, especially with appropriate kernel selection, making it a viable alternative for biomass estimation. In contrast, Random Forest (RF) showed the weakest performance, with Sentinel-2 yielding an R2 of 0.74, RMSE of 0.93 Mg/ha, and MAE of 0.76 Mg/ha, and Landsat-9 producing an R2 of 0.72, RMSE of 0.88 Mg/ha, and MAE of 0.74 Mg/ha. RF’s performance lagged behind the other models, particularly with Landsat-9, where its reduced ability to capture fine-scale variations in vegetation structure likely contributed to the lower accuracy. The coarser resolution of Landsat-9 likely hindered RF’s capacity to capture the variability needed for precise biomass estimation effectively. Overall, the results highlight the critical influence of satellite data resolution on model performance, with Sentinel-2 providing superior results due to its higher resolution. However, Landsat-9, despite its limitations, remains a valuable tool for global biomass estimation, particularly when paired with effective machine-learning models like XGBoost and SVM (Figure 6).
Comparing Sentinel-2 and Landsat-9-based AGB predictions made with XGBoost, SVM, and Random Forest models shows how spatial and spectral resolution affects the accuracy of biomass mapping. The Sentinel-2-based maps in Figure 7 (S1, S2, S3) had better spatial resolution. The AGB values ranged from (5.39–9.15 Mg/ha), showing apparent differences in biomass across the study area. The XGBoost model, in particular, excelled in delineating high and low-biomass zones, reflecting its ability to model complex spatial patterns with high precision.
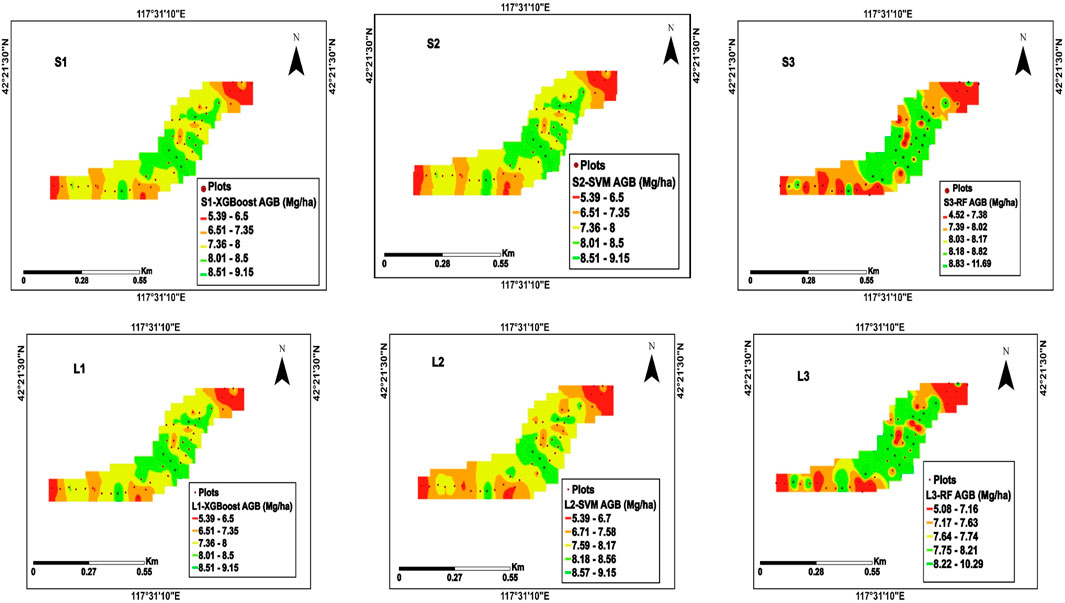
Figure 7. The study developed maps of AGB using the (XGBoost, SVM, RF) models, and data from the Sentinel 2 (S1, S2, S3) Landsat 9 (L1, L2, L3).
The SVM model had similar biomass ranges, but the changes between biomass classes were smoother because it uses a kernel-based approach that values global trends over local variability. The Random Forest model, in contrast, displayed more localized spatial noise, with a slightly wider range of AGB predictions from (4.71–10.15 Mg/ha).
When the same models were used on Landsat-9 data Figure 7 (L1, L2, L3), the lower spatial resolution, but the overall trends in biomass distribution were still well captured. XGBoost’s predictions for Landsat-9 resembled those of Sentinel-2, highlighting its robustness in handling data with lower resolution. The SVM model again showed smoother transitions, while the Random Forest model introduced more variability and noise, particularly with the Landsat-9 dataset. A comparison of the two data sets showed that Sentinel-2, which had a higher resolution, consistently made more accurate and detailed biomass maps. On the other hand, Landsat-9, which had a lower resolution, could still make accurate biomass predictions for larger-scale uses. Among the machine learning models, XGBoost consistently outperformed the others regarding spatial accuracy, its ability to capture non-linear relationships, and model complex interactions between input features. This study underscores the importance of selecting the appropriate remote sensing data and machine learning model for biomass estimation, with Sentinel-2 offering clear advantages for studies requiring fine-scale detail and XGBoost emerging as the most effective model for both datasets. The findings have significant implications for ecological monitoring, carbon accounting, and sustainable land management, highlighting the potential for combining high-resolution satellite data with advanced machine-learning techniques to improve AGB mapping (Figure 7).
This research assesses the capability of Sentinel-2 and Landsat-9 satellite data integrated with machine learning models for predicting AGB in a 50-year-old Larix principis-rupprechtii forest stand. It uses sound statistical and machine-learning techniques to demonstrate the usefulness of both Sentinel-2 and Landsat-9 satellite data in estimating forest biomass. The results are helpful for carbon stock assessment, forest evaluation, and land-use activities.
The results consistently demonstrate that Sentinel-2 outperforms Landsat-9 in AGB estimation, attributable to Sentinel-2’s higher spatial and spectral resolution. Indices from Sentinel-2, like TNDVI and ND145 (correlation = 0.58) and NDVI (correlation = 0.56), had stronger links with AGB. Landsat-9 counterparts (TNDVI = 0.56), (ND145 = 0.55), and NDVI = 0.53) did. This finding aligns with Castillo et al. (2017), who highlighted the advantages of higher-resolution imagery in capturing fine-scale vegetation gradients and structural attributes critical for biomass estimation.
Fassnacht et al. (2021) found through a correlation analysis between remote-sensing variables and field-measured AGB. These indices rely on the spectrum’s NIR and red edge regions, making them sensitive to canopy architecture, chlorophyll content, and vegetation vigor.
A study of the variable importance of Sentinel-2 and Landsat 9 images shows how important spectral bands and vegetation indices are in machine-learning models for estimating biomass and carbon stock. This study found that GNDVI was the best predictor for Sentinel-2 data in all models, especially in XGBoost. This is similar to Morales-Gallegos et al. (2023), Its sensitivity to subtle variations in vegetation properties, such as chlorophyll content and leaf area, positions it as an essential feature for biomass modeling. Similarly, NDI45 in Landsat 9 was the best predictor, especially in XGBoost. This aligns with recent research highlighting how indices like NDI45 and modified versions can capture important vegetation dynamics for biomass estimation (Pham et al., 2020). The study also revealed that other indices, such as WDVI and SAVI, had a relatively low correlation with biomass. Despite their application in remote sensing for vegetation and biomass estimation. The results are similar to those of Moghimi et al. (2024), who reported that these indices do not significantly contribute to biomass estimation.
Our findings demonstrate that WDVI and PSSRa in Landsat 9 play a crucial role. The results were consistent with the findings of Vidican et al. (2023), WDVI and PSSRa in Landsat 9 contributed to capturing moisture stress and vegetation health, which is crucial for biomass modeling, this study also found that spectral bands like Red and Blue in Sentinel-2 and Band 3 in Landsat 9 significantly affected biomass estimation.
These findings resonate with the work of Dong et al. (2020), who highlighted the importance of these bands for accurate biomass and carbon stock assessment. Overall, Sentinel-2 and Landsat 9 data complement each other well. Using them together improves model performance, making tracking vegetation and classifying land cover easier.
The results reveal that XGBoost consistently outperformed SVM and Random Forest across the Sentinel-2 and Landsat-9 datasets, achieving higher R2 values and lower error metrics. For Sentinel-2 data, XGBoost attained an (R2 = 0.82), SVM (R2 = 0.79), and Random Forest (R2 = 0.74). Similarly, with Landsat-9, XGBoost achieved an R2 of 0.80, outperforming SVM (R2 = 0.76) and Random Forest (R2 = 0.72). These results are similar to those of Liu H. et al. (2021), who reported that XGBoost’s gradient-boosting method is excellent at dealing with complicated, non-linear interactions in environmental datasets, especially when estimating biomass.
Similarly, Li et al. (2022) found that XGBoost outperformed traditional tree-based methods for forest biomass modeling, particularly when integrating multiple vegetation indices. New research from Miao et al. (2022) supports XGBoost’s ability to provide high accuracy, especially when combining data from different sources like Sentinel-2 and Landsat-9, which allows a more complex understanding of how plants change over time. SVM exhibited strong predictive capabilities, particularly in capturing non-linear relationships. However, its performance was slightly lower than XGBoost across both datasets, with Sentinel-2 results showing an RMSE of (0.73 Mg/ha) compared to XGBoost’s (0.69 Mg/ha). This finding is similar to Mehmood et al. (2012), but its kernel-based approach can sometimes make differences too smooth. also, this tendency highlights the need for careful kernel selection to ensure robust spatial transitions. Random Forest produced comparatively lower accuracy, particularly with Landsat-9 data, which recorded an RMSE of 0.88 Mg/ha. Thanh Noi and Kappas. (2017) reported that RF has been popular among ecological modeling techniques because of its stability and applicability for high-dimensional data. Yin et al. (2021), reported that its sensitivity to noise and potential overfitting in complex landscapes has been underscored in studies.
Future research should further integrate complementary datasets, such as LiDAR and hyperspectral imagery, to improve biomass prediction accuracy. Multitemporal analyses that account for seasonal and phenological variations could offer a more dynamic understanding of biomass changes. Also, creating ensemble methods that use the best parts of several machine learning models could avoid the problems that come with single algorithms and make predictions more accurate in complex and varied environments.
This study showed that remote sensing data from both Sentinel-2 and Landsat-9 can estimate the AGB of a L. principis-rupprechtii forest stand. It focused on the performance evaluation of the developed machine learning models, including XGBoost, SVM, and RF. The paper concludes that Sentinel-2, with higher spatial and spectral resolution, performs better in estimating biomass than Landsat-9, resulting in higher accuracy and detailed AGB prediction. It was discovered that to get a good picture of plants’ canopy structure and biomass’s most important chlorophyll content, use vegetation indices like TNDVI and NDI45 along with spectral bands like NIR.
The correlation analysis indicated that indices from Sentinel-2 had excellent correlations with AGB compared to the indices from Landsat-9. This demonstrates the significance of spatial and spectral resolution in remote sensing applications. Nevertheless, due to the coarser spatial resolution of Landsat-9, the results were helpful in larger-scale biomass mapping, especially when integrated with machine learning algorithms such as XGBoost and SVM, which showed good flexibility in handling data of different spatial resolutions. XGBoost was the best model in this study, with the highest accuracy in biomass predictions, followed by SVM, which was also very good at capturing non-linear patterns. The study that compared Sentinel-2 and Landsat-9 for AGB mapping demonstrates how valuable it could be to use remote sensing data in combination with modern machine learning techniques to obtain more accurate and less time-consuming biomass estimates. This approach has much potential for ecological assessment, carbon stock estimation, and sustainable forest management, particularly in areas that need accurate biomass information to address climate change and conserve biological diversity.
While Sentinel-2 provides superior accuracy for high-resolution biomass mapping, Landsat-9 remains a valuable tool for large-scale applications, especially when paired with effective machine-learning models. The findings from this study highlight the importance of selecting the appropriate remote sensing platform and machine learning technique to optimize biomass estimation, thereby contributing to the broader field of remote sensing-based environmental monitoring. Future research should focus on integrating multi-temporal satellite data and exploring more advanced machine learning algorithms to further enhance the accuracy and applicability of biomass mapping for global carbon accounting and sustainable land management initiatives.
The original contributions presented in the study are included in the article/Supplementary Material, further inquiries can be directed to the corresponding authors.
JA: Conceptualization, Data curation, Formal Analysis, Investigation, Methodology, Software, Validation, Visualization, Writing – original draft, Writing – review and editing. WHa: Resources, Writing – original draft, Writing – review and editing. KM: Software, Writing – original draft, Writing – review and editing. WHu: Formal Analysis, Software, Writing – original draft, Writing – review and editing. FI: Resources, Writing – original draft, Writing – review and editing. FS: Writing – original draft, Writing – review and editing. KH: Writing – original draft, Writing – review and editing. YQ: Funding acquisition, Project administration, Supervision, Writing – original draft, Writing – review and editing. JZ: Funding acquisition, Project administration, Supervision, Writing – original draft, Writing – review and editing.
The author(s) declare that financial support was received for the research and/or publication of this article. Effects of spatial variability and biological factors on trunk respiration of L. principis-rupprechtii and its internal mechanism: Grant Number: 31870387.
This work was made possible by the support, cooperation, and collaboration of the Saihanba Mechanical Forest Farm staff provided invaluable assistance and expertise throughout the project. The Hebei Forest Department members also played a crucial role in supporting our efforts. The (State Key Laboratory of Efficient Production of Forest Resources) contributed significantly with their advanced research and resources. Furthermore, the (Engineering Technology Research Center of Pinus tabuliformis of National Forestry and Grassland Administration) offered essential technological insights and guidance. Their collective efforts were instrumental in the successful completion of this work.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fenvs.2025.1577298/full#supplementary-material
Ali, J., Malik, S. U., Ashraf, M. I., Zhongkui, J., Husnain, Z., and Gulzar, S. (2023). Exploring the potential of carbon sequestration in sub-tropical pine forest ecosystem: a case study in district kurram, Pakistan. Sarhad J. Agric. 39. doi:10.17582/JOURNAL.SJA/2023/39.3.647.654
Almeida, A., Gonçalves, F., Silva, G., Mendonça, A., Gonzaga, M., Silva, J., et al. (2021). Individual tree detection and qualitative inventory of a eucalyptus sp. Stand using uav photogrammetry data. Remote Sens. (Basel) 13, 3655. doi:10.3390/rs13183655
Anees, S. A., Mehmood, K., Rehman, A., Rehman, N. U., Muhammad, S., Shahzad, F., et al. (2024). Unveiling fractional vegetation cover dynamics: a spatiotemporal analysis using MODIS NDVI and Machine Learning. Environ. Sustain Indic. 24, 100485. doi:10.1016/j.indic.2024.100485
Araza, A., de Bruin, S., Herold, M., Quegan, S., Labriere, N., Rodriguez-Veiga, P., et al. (2022). A comprehensive framework for assessing the accuracy and uncertainty of global above-ground biomass maps. Remote Sens. Environ. 272, 112917. doi:10.1016/j.rse.2022.112917
Aslam, R. W., Shu, H., Tariq, A., Naz, I., Ahmad, M. N., Quddoos, A., et al. (2024). Monitoring landuse change in Uchhali and Khabeki wetland lakes, Pakistan using remote sensing data. Gondwana Res. 129, 252–267. doi:10.1016/j.gr.2023.12.015
Asselman, A., Khaldi, M., and Aammou, S. (2023). Enhancing the prediction of student performance based on the machine learning XGBoost algorithm. Interact. Learn. Environ. 31, 3360–3379. doi:10.1080/10494820.2021.1928235
Aye, W. N., Tong, X., and Tun, A. W. (2022). Species diversity, biomass and carbon stock assessment of Kanhlyashay Natural Mangrove Forest. Forests 13, 1013. doi:10.3390/f13071013
Aziz, G., Minallah, N., Saeed, A., Frnda, J., and Khan, W. (2024). Remote sensing based forest cover classification using machine learning. Sci. Rep. 14, 69. doi:10.1038/s41598-023-50863-1
Bouslihim, Y., John, K., Miftah, A., Azmi, R., Aboutayeb, R., Bouasria, A., et al. (2024). The effect of covariates on Soil Organic Matter and pH variability: a digital soil mapping approach using random forest model. Ann GIS 30, 215–232. doi:10.1080/19475683.2024.2309868
Bulut, S. (2023). Machine learning prediction of above-ground biomass in pure Calabrian pine (Pinus brutia Ten.) stands of the Mediterranean region, Türkiye. Ecol. Inf. 74, 101951. doi:10.1016/j.ecoinf.2022.101951
Cameron, H. A., Panda, P., Barczyk, M., and Beverly, J. L. (2022). Estimating boreal forest ground cover vegetation composition from nadir photographs using deep convolutional neural networks. Ecol. Inf. 69, 101658. doi:10.1016/j.ecoinf.2022.101658
Campbell, M. J., Dennison, P. E., Kerr, K. L., Brewer, S. C., and Anderegg, W. R. L. (2021). Scaled biomass estimation in woodland ecosystems: testing the individual and combined capacities of satellite multispectral and lidar data. Remote Sens. Environ. 262, 112511. doi:10.1016/j.rse.2021.112511
Carmenta, R., Coomes, D. A., DeClerck, F. A. J., Hart, A. K., Harvey, C. A., Milder, J., et al. (2020). Characterizing and evaluating integrated landscape initiatives. One Earth 2, 174–187. doi:10.1016/j.oneear.2020.01.009
Castillo, J. A. A., Apan, A. A., Maraseni, T. N., and Salmo, S. G. (2017). Estimation and mapping of above-ground biomass of mangrove forests and their replacement land uses in the Philippines using Sentinel imagery. ISPRS J. Photogrammetry Remote Sens. 134, 70–85. doi:10.1016/j.isprsjprs.2017.10.016
Chen, L., Ren, C., Zhang, B., Wang, Z., and Xi, Y. (2018). Estimation of forest above-ground biomass by geographically weighted regression and machine learning with sentinel imagery. Forests 9, 582. doi:10.3390/f9100582
Coffie, G. H., and Cudjoe, S. K. F. (2023). Using extreme gradient boosting (XGBoost) machine learning to predict construction cost overruns. Int. J. Constr. Manag. 24, 1742–1750. doi:10.1080/15623599.2023.2289754
Coops, N. C., Tompalski, P., Goodbody, T. R. H., Achim, A., and Mulverhill, C. (2023). Framework for near real-Time forest inventory using multi source remote sensing data. Forestry 96, 1–19. doi:10.1093/forestry/cpac015
Das, B., Patnaik, S. K., Bordoloi, R., Paul, A., and Tripathi, O. P. (2024). Prediction of forest aboveground biomass using an integrated approach of space-based parameters, and forest inventory data. Geol. Ecol. Landscapes 8, 381–393. doi:10.1080/24749508.2022.2139484
Dong, T., Liu, J., Qian, B., He, L., Liu, J., Wang, R., et al. (2020). Estimating crop biomass using leaf area index derived from Landsat 8 and Sentinel-2 data. ISPRS J. Photogrammetry Remote Sens. 168, 236–250. doi:10.1016/j.isprsjprs.2020.08.003
Eshetu, E. Y., and Hailu, T. A. (2020). Carbon sequestration and elevational gradient: the case of Yegof mountain natural vegetation in North East, Ethiopia, implications for sustainable management. Cogent Food Agric. 6, 1733331. doi:10.1080/23311932.2020.1733331
Fassnacht, F. E., Poblete-Olivares, J., Rivero, L., Lopatin, J., Ceballos-Comisso, A., and Galleguillos, M. (2021). Using Sentinel-2 and canopy height models to derive a landscape-level biomass map covering multiple vegetation types. Int. J. Appl. Earth Observation Geoinformation 94, 102236. doi:10.1016/j.jag.2020.102236
Freeman, E. A., Moisen, G. G., Coulston, J. W., and Wilson, B. T. (2015). Random forests and stochastic gradient boosting for predicting tree canopy cover: comparing tuning processes and model performance. Can. J. For. Res. 46, 323–339. doi:10.1139/cjfr-2014-0562
Fu, B., Sun, J., Wang, Y., Yang, W., He, H., Liu, L., et al. (2022). Evaluation of LAI estimation of mangrove communities using DLR and ELR algorithms with UAV, hyperspectral, and SAR images. Front. Mar. Sci. 9. doi:10.3389/fmars.2022.944454
Ghasemloo, N., Matkan, A. A., Alimohammadi, A., Aghighi, H., and Mirbagheri, B. (2022). Estimating the agricultural Farm soil moisture using spectral indices of Landsat 8, and sentinel-1, and artificial neural networks. J. Geovisualization Spatial Analysis 6, 19. doi:10.1007/s41651-022-00110-4
Gibson, J. (2018). Forest loss and economic inequality in the Solomon Islands: using small-area estimation to link environmental change to welfare outcomes. Ecol. Econ. 148, 66–76. doi:10.1016/j.ecolecon.2018.02.012
Godinho Cassol, H. L., De Oliveira E Cruz De Aragão, L. E., Moraes, E. C., De Brito Carreiras, J. M., and Shimabukuro, Y. E. (2021). Quad-pol advanced land observing satellite/phased array L-band synthetic aperture radar-2 (ALOS/PALSAR-2) data for modelling secondary forest above-ground biomass in the central Brazilian amazon. Int. J. Remote Sens. 42, 4985–5009. doi:10.1080/01431161.2021.1903615
Han, L., Yang, G., Dai, H., Xu, B., Yang, H., Feng, H., et al. (2019). Modeling maize above-ground biomass based on machine learning approaches using UAV remote-sensing data. Plant Methods 15, 10. doi:10.1186/s13007-019-0394-z
Hojo, A., Avtar, R., Nakaji, T., Tadono, T., and Takagi, K. (2023). Modeling forest above-ground biomass using freely available satellite and multisource datasets. Ecol. Inf. 74, 101973. doi:10.1016/j.ecoinf.2023.101973
Hu, Y., Zhang, Q., Hu, S., Xiao, G., Chen, X., Wang, J., et al. (2022). Research progress and prospects of ecosystem carbon sequestration under climate change (1992–2022). Ecol. Indic. 145, 109656. doi:10.1016/j.ecolind.2022.109656
Hussain, K., Badshah, T., Mehmood, K., Shahzad, F., Anees, S. A., Khan, W. R., et al. (2025). Comparative analysis of sensors and classification algorithms for land cover classification in Islamabad, Pakistan. Earth Sci. Inf. 18, 212–222. doi:10.1007/s12145-025-01720-4
Hussain, K., Mehmood, K., Yujun, S., Badshah, T., Anees, S. A., Shahzad, F. N., et al. (2024). Analysing LULC transformations using remote sensing data: insights from a multilayer perceptron neural network approach. Ann. GIS, 1–28. Ann GIS 1–28. doi:10.1080/19475683.2024.2343399
Jafri, Y., Ahlström, J. M., Furusjö, E., Harvey, S., Pettersson, K., Svensson, E., et al. (2022). Double yields and negative emissions? Resource, climate and cost efficiencies in biofuels with carbon capture, storage and utilization. Front. Energy Res. 10. doi:10.3389/fenrg.2022.797529
Kristensen, T., Næsset, E., Ohlson, M., Bolstad, P. V., and Kolka, R. (2015). Mapping above- and below-ground carbon pools in boreal forests: the case for airborne lidar. PLoS One 10, e0138450. doi:10.1371/journal.pone.0138450
Labrière, N., Davies, S. J., Disney, M. I., Duncanson, L. I., Herold, M., Lewis, S. L., et al. (2023). Toward a forest biomass reference measurement system for remote sensing applications. Glob. Chang. Biol. 29, 827–840. doi:10.1111/gcb.16497
Lee, H., Wang, J., and Leblon, B. (2020). Using linear regression, random forests, and support vector machine with unmanned aerial vehicle multispectral images to predict canopy nitrogen weight in corn. Remote Sens. (Basel) 12, 2071. doi:10.3390/rs12132071
Li, X., Zhang, M., Long, J., and Lin, H. (2021). A novel method for estimating spatial distribution of forest above-ground biomass based on multispectral fusion data and ensemble learning algorithm. Remote Sens. (Basel) 13, 3910. doi:10.3390/rs13193910
Li, Y., Feng, Z., Chen, S., Zhao, Z., and Wang, F. (2020c). Application of the artificial neural network and support vector machines in forest fire prediction in the guangxi autonomous region, China. Discrete Dyn. Nat. Soc. 2020, 1–14. doi:10.1155/2020/5612650
Li, Y., Li, M., Li, C., and Liu, Z. (2020a). Forest aboveground biomass estimation using Landsat 8 and Sentinel-1A data with machine learning algorithms. Sci. Rep. 10, 9952. doi:10.1038/s41598-020-67024-3
Li, Y., Li, M., Liu, Z., and Li, C. (2020b). Combining kriging interpolation to improve the accuracy of forest aboveground biomass estimation using remote sensing data. IEEE Access 8, 128124–128139. doi:10.1109/ACCESS.2020.3008686
Li, Y., Li, M., and Wang, Y. (2022). Forest aboveground biomass estimation and response to climate change based on remote sensing data. Sustain. Switz. 14, 14222. doi:10.3390/su142114222
Li, Z., Yuan, Q., and Su, X. (2024). High-spatial-resolution surface soil moisture retrieval using the Deep Forest model in the cloud environment over the Tibetan Plateau. Geo-Spatial Inf. Sci., 1–20. doi:10.1080/10095020.2024.2307931
Lin, W., Lu, Y., Li, G., Jiang, X., and Lu, D. (2022). A comparative analysis of modeling approaches and canopy height-based data sources for mapping forest growing stock volume in a northern subtropical ecosystem of China. GIsci Remote Sens. 59, 568–589. doi:10.1080/15481603.2022.2044139
Liu, H., Jin, Y., Roche, L. M., O’Geen, A. T., and Dahlgren, R. A. (2021a). Understanding spatial variability of forage production in California grasslands: delineating climate, topography and soil controls. Environ. Res. Lett. 16, 014043. doi:10.1088/1748-9326/abc64d
Liu, N., Li, Y., Cong, P., Wang, J., Guo, W., Pang, H., et al. (2021b). Depth of straw incorporation significantly alters crop yield, soil organic carbon and total nitrogen in the North China Plain. Soil Tillage Res. 205, 104772. doi:10.1016/j.still.2020.104772
López Serrano, F. R., Rubio, E., García Morote, F. A., Andrés Abellán, M., Picazo Córdoba, M. I., García Saucedo, F., et al. (2022). Artificial intelligence-based software (AID-FOREST) for tree detection: a new framework for fast and accurate forest inventorying using LiDAR point clouds. Int. J. Appl. Earth Observation Geoinformation 113, 103014. doi:10.1016/j.jag.2022.103014
Luo, H., Qin, S., Li, J., Lu, C., Yue, C., and Ou, G. (2024). High-density forest AGB estimation in tropical forest integrated with PolInSAR multidimensional features and optimized machine learning algorithms. Ecol. Indic. 160, 111878. doi:10.1016/j.ecolind.2024.111878
Ma, T., Zhang, C., Ji, L., Zuo, Z., Beckline, M., Hu, Y., et al. (2024). Development of forest aboveground biomass estimation, its problems and future solutions: a review. Ecol. Indic. 159, 111653. doi:10.1016/j.ecolind.2024.111653
Mahmood, M. S., Elahi, A., Zaid, O., Alashker, Y., Șerbănoiu, A. A., Grădinaru, C. M., et al. (2023). Enhancing compressive strength prediction in self-compacting concrete using machine learning and deep learning techniques with incorporation of rice husk ash and marble powder. Case Stud. Constr. Mater. 19, e02557. doi:10.1016/j.cscm.2023.e02557
Masereti Makori, D., Abdel-Rahman, E. M., Odindi, J., Mutanga, O., Landmann, T., and Tonnang, H. E. Z. (2024). Multi-pronged abundance prediction of bee pests’ spatial proliferation in Kenya. Int. J. Appl. Earth Observation Geoinformation 128, 103738. doi:10.1016/j.jag.2024.103738
Mehmood, K., Anees, S. A., Muhammad, S., Hussain, K., Shahzad, F., Liu, Q., et al. (2024a). Analyzing vegetation health dynamics across seasons and regions through NDVI and climatic variables. Sci. Rep. 14, 11775. doi:10.1038/s41598-024-62464-7
Mehmood, K., Anees, S. A., Muhammad, S., Shahzad, F., Liu, Q., Khan, W. R., et al. (2025). Machine learning and spatio temporal analysis for assessing ecological impacts of the billion tree afforestation Project. Ecol. Evol. 15, e70736. doi:10.1002/ece3.70736
Mehmood, K., Anees, S. A., Rehman, A., Rehman, N. U., Muhammad, S., Shahzad, F., et al. (2024b). Assessment of climatic influences on net primary productivity along elevation gradients in temperate ecoregions. Trees, For. People 18, 100657. doi:10.1016/j.tfp.2024.100657
Mehmood, T., Liland, K. H., Snipen, L., and Sæbø, S. (2012). A review of variable selection methods in Partial Least Squares Regression. Chemom. Intelligent Laboratory Syst. 118, 62–69. doi:10.1016/j.chemolab.2012.07.010
Miao, J., Zhen, J., Wang, J., Zhao, D., Jiang, X., Shen, Z., et al. (2022). Mapping seasonal leaf nutrients of mangrove with sentinel-2 images and XGBoost method. Remote Sens. (Basel) 14, 3679. doi:10.3390/rs14153679
Moghimi, A., Tavakoli Darestani, A., Mostofi, N., Fathi, M., and Amani, M. (2024). Improving forest above-ground biomass estimation using genetic-based feature selection from Sentinel-1 and Sentinel-2 data (case study of the Noor forest area in Iran). Kuwait J. Sci. 51, 100159. doi:10.1016/j.kjs.2023.11.008
Morales-Gallegos, L. M., Martínez-Trinidad, T., Hernández-de la Rosa, P., Gómez-Guerrero, A., Alvarado-Rosales, D., and Saavedra-Romero, L. de L. (2023). Tree health condition in urban green areas assessed through crown indicators and vegetation indices. Forests 14, 1673. doi:10.3390/f14081673
Musekiwa, N. B., Angombe, S. T., Kambatuku, J., Mudereri, B. T., and Chitata, T. (2022). Can encroached rangelands enhance carbon sequestration in the African Savannah? Trees, For. People 7, 100192. doi:10.1016/j.tfp.2022.100192
Octavia, D., Suharti, S. M., Dharmawan, I. W. S., Nugroho, H. Y. S. H., Supriyanto, B., Rohadi, D., et al. (2022). Mainstreaming smart agroforestry for social forestry implementation to support sustainable development goals in Indonesia: a review. Sustain. Switz. 14, 9313. doi:10.3390/su14159313
Oehmcke, S., Li, L., Trepekli, K., Revenga, J. C., Nord-Larsen, T., Gieseke, F., et al. (2024). Deep point cloud regression for above-ground forest biomass estimation from airborne LiDAR. Remote Sens. Environ. 302, 113968. doi:10.1016/j.rse.2023.113968
Pérez-Girón, J. C., Álvarez-Álvarez, P., Díaz-Varela, E. R., and Mendes Lopes, D. M. (2020). Influence of climate variations on primary production indicators and on the resilience of forest ecosystems in a future scenario of climate change: application to sweet chestnut agroforestry systems in the Iberian Peninsula. Ecol. Indic. 113, 106199. doi:10.1016/j.ecolind.2020.106199
Pham, T. D., Ha, N. T., Saintilan, N., Skidmore, A., Phan, D. C., Le, N. N., et al. (2023). Advances in Earth observation and machine learning for quantifying blue carbon. Earth Sci. Rev. 243, 104501. doi:10.1016/j.earscirev.2023.104501
Pham, T. D., Le, N. N., Ha, N. T., Nguyen, L. V., Xia, J., Yokoya, N., et al. (2020). Estimating mangrove above-ground biomass using extreme gradient boosting decision trees algorithm with fused sentinel-2 and ALOS-2 PALSAR-2 data in can Gio biosphere reserve, Vietnam. Remote Sens. (Basel) 12, 777. doi:10.3390/rs12050777
Prakash, A. J., Behera, M. D., Ghosh, S. M., Das, A., and Mishra, D. R. (2022). A new synergistic approach for Sentinel-1 and PALSAR-2 in a machine learning framework to predict aboveground biomass of a dense mangrove forest. Ecol. Inf. 72, 101900. doi:10.1016/j.ecoinf.2022.101900
Prăvălie, R., Niculiță, M., Roșca, B., Marin, G., Dumitrașcu, M., Patriche, C., et al. (2023). Machine learning-based prediction and assessment of recent dynamics of forest net primary productivity in Romania. J. Environ. Manage 334, 117513. doi:10.1016/j.jenvman.2023.117513
Ramachandran, N., Saatchi, S., Tebaldini, S., d’Alessandro, M. M., and Dikshit, O. (2023). Mapping tropical forest aboveground biomass using airborne SAR tomography. Sci. Rep. 13, 6233. doi:10.1038/s41598-023-33311-y
Santoro, M., Cartus, O., Carvalhais, N., Rozendaal, D. M. A., Avitabile, V., Araza, A., et al. (2021). The global forest above-ground biomass pool for 2010 estimated from high-resolution satellite observations. Earth Syst. Sci. Data 13, 3927–3950. doi:10.5194/essd-13-3927-2021
Sedano, F., Lisboa, S. N., Sahajpal, R., Duncanson, L., Ribeiro, N., Sitoe, A., et al. (2021). The connection between forest degradation and urban energy demand in sub-Saharan Africa: a characterization based on high-resolution remote sensing data. Environ. Res. Lett. 16, 064020. doi:10.1088/1748-9326/abfc05
Shahzad, F., Mehmood, K., Anees, S. A., Adnan, M., Muhammad, S., Haidar, I., et al. (2025). Advancing forest fire prediction: a multi-layer stacking ensemble model approach. Earth Sci. Inf. 18, 270. doi:10.1007/s12145-025-01782-4
Shahzad, F., Mehmood, K., Hussain, K., Haidar, I., Anees, S. A., Muhammad, S., et al. (2024). Comparing machine learning algorithms to predict vegetation fire detections in Pakistan. Fire Ecol. 20, 57. doi:10.1186/s42408-024-00289-5
Shams, M. Y., Elshewey, A. M., El-kenawy, E. S. M., Ibrahim, A., Talaat, F. M., and Tarek, Z. (2024). Water quality prediction using machine learning models based on grid search method. Multimed. Tools Appl. 83, 35307–35334. doi:10.1007/s11042-023-16737-4
Stephenson, P. J., and Damerell, A. (2022). Bioeconomy and circular economy approaches need to enhance the focus on biodiversity to achieve sustainability. Sustain. Switz. 14, 10643. doi:10.3390/su141710643
Su, H., Shen, W., Wang, J., Ali, A., and Li, M. (2020). Machine learning and geostatistical approaches for estimating aboveground biomass in Chinese subtropical forests. For Ecosyst 7, 64. doi:10.1186/s40663-020-00276-7
Sun, L., Feng, Z., Shao, Y., Wang, L., Su, J., Ma, T., et al. (2023). The development of a set of novel low cost and data processing-free measuring instruments for tree diameter at breast height and tree position. Forests 14, 891. doi:10.3390/f14050891
Tao, C., Guo, T., Shen, M., and Tang, Y. (2023). Spatio-temporal dynamic of disturbances in planted and natural forests for the Saihanba region of China. Remote Sens. (Basel) 15, 4776. doi:10.3390/rs15194776
Teng, H., Chen, S., Hu, B., and Shi, Z. (2023). Future changes and driving factors of global peak vegetation growth based on CMIP6 simulations. Ecol. Inf. 75, 102031. doi:10.1016/j.ecoinf.2023.102031
Thanh Noi, P., and Kappas, M. (2017). Comparison of random forest, k-nearest neighbor, and support vector machine classifiers for land cover classification using sentinel-2 imagery. Sensors (Basel) 18, 18. doi:10.3390/s18010018
Thompson, C. G., Kim, R. S., Aloe, A. M., and Becker, B. J. (2017). Extracting the variance in flation factor and other multicollinearity diagnostics from typical regression results. Basic Appl. Soc. Psych. 39, 81–90. doi:10.1080/01973533.2016.1277529
Titus, B. D., Brown, K., Helmisaari, H. S., Vanguelova, E., Stupak, I., Evans, A., et al. (2021). Sustainable forest biomass: a review of current residue harvesting guidelines. Energy Sustain Soc. 11, 10. doi:10.1186/s13705-021-00281-w
Turner, W., Rondinini, C., Pettorelli, N., Mora, B., Leidner, A. K., Szantoi, Z., et al. (2015). Free and open-access satellite data are key to biodiversity conservation. Biol. Conserv. 182, 173–176. doi:10.1016/j.biocon.2014.11.048
Vidican, R., Mălinaș, A., Ranta, O., Moldovan, C., Marian, O., Ghețe, A., et al. (2023). Using remote sensing vegetation indices for the discrimination and monitoring of agricultural crops: a critical review. Agronomy. doi:10.3390/agronomy13123040
Wang, J., Shi, K., and Hu, M. (2022). Measurement of forest carbon sink efficiency and its influencing factors empirical evidence from China. Forests 13, 1909. doi:10.3390/f13111909
Wongsai, N., Wongsai, S., Lim, A., McNeil, D., and Huete, A. R. (2020). Impacts of spatial heterogeneity patterns on long-term trends of Moderate Resolution Imaging Spectroradiometer (MODIS) land surface temperature time series. J. Appl. Remote Sens. 14, 1. doi:10.1117/1.jrs.14.014513
Xu, A., Wang, D., Liu, Q., Zhang, D., Zhang, Z., and Huang, X. (2022). Incorporating stand density effects and regression techniques for stem taper modeling of a Larix principis-rupprechtii plantation. Front. Plant Sci. 13, 902325. doi:10.3389/fpls.2022.902325
Yin, J., Dong, J., Hamm, N. A. S., Li, Z., Wang, J., Xing, H., et al. (2021). Integrating remote sensing and geospatial big data for urban land use mapping: a review. Int. J. Appl. Earth Observation Geoinformation 103, 102514. doi:10.1016/j.jag.2021.102514
Zennaro, F., Furlan, E., Simeoni, C., Torresan, S., Aslan, S., Critto, A., et al. (2021). Exploring machine learning potential for climate change risk assessment. Earth Sci. Rev. 220, 103752. doi:10.1016/j.earscirev.2021.103752
Zhang, L., and Jánošík, D. (2024). Enhanced short-term load forecasting with hybrid machine learning models: CatBoost and XGBoost approaches. Expert Syst. Appl. 241, 122686. doi:10.1016/j.eswa.2023.122686
Zhang, Y., Ma, J., Liang, S., Li, X., and Liu, J. (2022). A stacking ensemble algorithm for improving the biases of forest aboveground biomass estimations from multiple remotely sensed datasets. GIsci Remote Sens. 59, 234–249. doi:10.1080/15481603.2021.2023842
Zhang, Y., and Shao, Z. (2021). Assessing of urban vegetation biomass in combination with LiDAR and high-resolution remote sensing images. Int. J. Remote Sens. 42, 964–985. doi:10.1080/01431161.2020.1820618
Keywords: above-ground biomass (AGB), Larix principis-rupprechtii, remote sensing, machine learning, Sentinel-2, and Landsat-9
Citation: Ali J, Haoran W, Mehmood K, Hussain W, Iftikhar F, Shahzad F, Hussain K, Qun Y and Zhongkui J (2025) Remote sensing and integration of machine learning algorithms for above-ground biomass estimation in Larix principis-rupprechtii Mayr plantations: a case study using Sentinel-2 and Landsat-9 data in northern China. Front. Environ. Sci. 13:1577298. doi: 10.3389/fenvs.2025.1577298
Received: 15 February 2025; Accepted: 24 March 2025;
Published: 02 April 2025.
Edited by:
Sawaid Abbas, University of the Punjab, PakistanReviewed by:
Faisal Mueen Qamer, International Centre for Integrated Mountain Development, NepalCopyright © 2025 Ali, Haoran, Mehmood, Hussain, Iftikhar, Shahzad, Hussain, Qun and Zhongkui. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Yin Qun, eWlucXVuMTAwMkBiamZ1LmVkdS5jbg==; Jia Zhongkui, amlhemtAMTYzLmNvbQ==
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.