- 1Department of Pharmacy, the Second Affiliated Hospital of Shandong First Medical University, Tai’an, China
- 2Department of Stomatology, the Second Affiliated Hospital of Shandong First Medical University, Tai’an, China
Objective: To evaluate the characteristics of the circulating microRNA expression profiles in patients with osteoporosis.
Methods: A systematic literature search was performed using the Web of Science, PubMed, Embase, Cochrane Library, China National Knowledge Infrastructure (CNKI), VIP, and WANFANG databases from inception until 1 March 2024. The search strategy employed keywords, encompassing “osteoporosis”, “bone loss”, or “osteopenia” and “miRNA” or “microRNA”. The Newcastle-Ottawa Scale (NOS) quality assessment scale was used to evaluate the methodological quality. Heterogeneity tests and statistical analyses of all data were performed by Stata 16.0. The differences in microRNA levels between groups were illustrated by the weighted mean difference (WMD) and 95% confidence interval (95% CI).
Results: A total of 27 studies were included and analyzed in the meta-analysis, with 2,263 participants. The results showed that miR-21-5p (WMD 0.88, 95% CI: 0.22 to 1.55), miR-125b-5p (WMD 6.63, 95% CI: 0.19 to 13.08), miR-483-5p(WMD 6.43, 95% CI: 3.26 to 9.61), miR-133a (WMD 1.43, 95% CI: 1.39 to 1.47), miR-422a (WMD 1, 95% CI: 0.28 to 1.72), and miR-214-3p (WMD 2.03, 95% CI: 0.14 to 3.92) were significantly upregulated, and miR-497-5p (WMD -0.57, 95% CI: -0.98 to -0.17) was significantly downregulated.
Conclusion: miR-21-5p, miR-125b-5p, miR-483-5p, miR-133a, miR-497-5p, miR-422a, and miR-214-3p might serve as potential diagnostic biomarkers for osteoporosis. In the future, integrating these miRNAs to build a diagnostic model might be a promising diagnosis strategy for osteoporosis.
Systematic review registration: https://www.crd.york.ac.uk/PROSPERO/, identifier CRD42023481209.
1 Introduction
Osteoporosis (OP), a progressive skeletal disorder characterized by reduced bone mineral density, compromised bone strength, and increased risk of fracture, has garnered widespread attention because it has a significant impact on health-related quality of life (1). Nearly 200 million people worldwide are diagnosed with osteoporosis each year, and nearly 9 million osteoporotic fractures occur each year (2). Hip and vertebrae fractures are associated with particularly high morbidity and mortality, posing a high socioeconomic burden on overstretched health systems (3). Therefore, fracture prevention through early diagnosis is the primary goal of osteoporosis management. However, the onset of osteoporosis is usually asymptomatic, which highlights the need for a strategy that allows a quick and effective diagnosis of osteoporosis to mitigate further disease progress and subsequent fractures (4).
Currently, bone mineral density (BMD), measured using dual-energy x-ray absorptiometry (DXA), is employed to diagnose osteoporosis (5). Higher BMD indicates denser, stronger bones and lower BMD indicates less dense, weaker bones. Notably, not only bone mass but also bone quality is a factor in osteoporosis. Unfortunately, DXA cannot accurately assess bone quality (6). In addition, spinal deformities, previous compression fractures, and aortic atherosclerosis can lead to an increase in effective x-ray uptake, resulting in a pseudo-elevation of the T-Score (7, 8). Overall, DXA scans are not accurate enough to diagnose osteoporosis. Therefore, an effective diagnostic indicator to identify osteoporosis is urgently required.
MicroRNAs (miRNAs), which are non-encoded, endogenous, and single-stranded RNAs of ~22 nucleotides, are important regulators of numerous biological processes through the posttranscriptional regulation of gene expression (9, 10). A large number of in vitro and in vivo studies suggest that miRNAs are involved in cell differentiation, proliferation, autophagy, and apoptosis in the bone microenvironment. Differential expression of miRNAs in different stages of the development of osteoporosis have shown that they are closely related to the occurrence and development of osteoporosis (11).
Recently, numerous research studies have focused on circulating miRNAs as a potential biomarker for the early detection of osteoporosis (12). Some studies indicated up or downregulation of diverse miRNAs in osteoporotic postmenopausal women compared with healthy postmenopausal women (13). Due to different technological platforms and small sample sizes among various studies, conflicting results regarding the direction of regulation have been found for some miRNAs. Furthermore, the characteristic expression of circulating miRNAs in osteoporosis has not been accurately evaluated. This meta-analysis was performed to further clarify the characteristics of the circulating miRNA expression profiles in osteoporosis, and explore the potential value of circulating miRNAs for the diagnosis of osteoporosis.
2 Methods
This systematic review and meta-analysis was conducted in line with the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) statement. The protocol for this meta-analysis was prospectively registered in the PROSPERO database (CRD42023481209).
2.1 Data sources and retrieval strategy
A systematic literature search was performed using the Web of Science, PubMed, Embase, Cochrane Library, CNKI, VIP, and WANFANG databases. The literature search included studies published in English or Chinese from inception until 1 March 2024. The search strategy employed the following keywords: “osteoporosis”, “bone loss”, or “osteopenia” and “miRNA” or “microRNA”. For the identification of additional relevant studies, the reference lists of related reviews and included articles were screened manually.
2.2 Selection criteria
The inclusion criteria were defined as (1): the study should have the miRNA expression profiles of patients with osteoporosis (2); osteoporosis patients were included in the experimental group, and healthy individuals were included in the control group (3); the relative miRNA expression must be profiled by RT-qPCR (4); the sample size was reported (5); the mean and standard deviation of the miRNA expression profiles could be obtained; and (6) all the patients with osteoporosis were diagnosed by DXA. The exclusion criteria were as follows (1): studies on animals (2); simple descriptive literature without a control group (3); review literature, case report, abstract, and letter; and (4) literature for which the relevant data could not be obtained.
2.3 Data extraction
The following study data was extracted: first author, year of publication, country, age, sex, source of samples, miRNA expression profile, assay type, and sample size. Two investigators independently extracted the data following the pre-defined inclusion and exclusion criteria. If different sample sources were provided in the same study, data extraction and analysis were carried out respectively. Any discrepancies were resolved through discussions and, if necessary, a third senior investigator was consulted until a consensus was reached. The corresponding authors of the original articles were contacted to obtain relevant data that were not present in the full text and supplementary information. If data were only shown by graphs, GetData Graph Digitizer software (version 2.26) was used to extract numerical values.
2.4 Quality assessment
Two independent investigators used the Newcastle-Ottawa Scale (NOS) to assess the quality of the included studies. The scale involved three aspects with a total of 9 points: the selection of cohorts (0–4 points), the comparability of cohorts (0–2 points), and the assessment of the outcome (0–3 points). Quality was classified as high with an NOS score of ≥6 points and low with an NOS score of <6 points (14). Disagreements were resolved until a consensus was reached by mutual discussion with a third senior investigator.
2.5 Statistical analysis
Stata software (version 16.0, Stata Corp LP, College Station, TX, USA) was used to analyze the data. The weighted mean difference (WMD) and 95% confidence interval (95% CI) were calculated to indicate the effect size of the differences in miRNA expression levels between groups. The heterogeneity of the included studies was usually tested by Cochrane’s Q test and I2 statistic. For Cochrane’s Q test, P < 0.1 indicated significant heterogeneity between the studies. The fixed effect model was selected if I2 was < 50% and the random effect model was selected if I2 was ≥ 50%. Subgroup and regression analyses were conducted to explore the possible sources of heterogeneity. A sensitivity analysis was performed to analyze the stability and reliability of the effect size using the “remove one study” method. Finally, Egger’s and Begg’s tests, as well as the visual inspection of funnel plots, were used to assess any potential publication bias.
3 Results
3.1 Search results and study characteristics
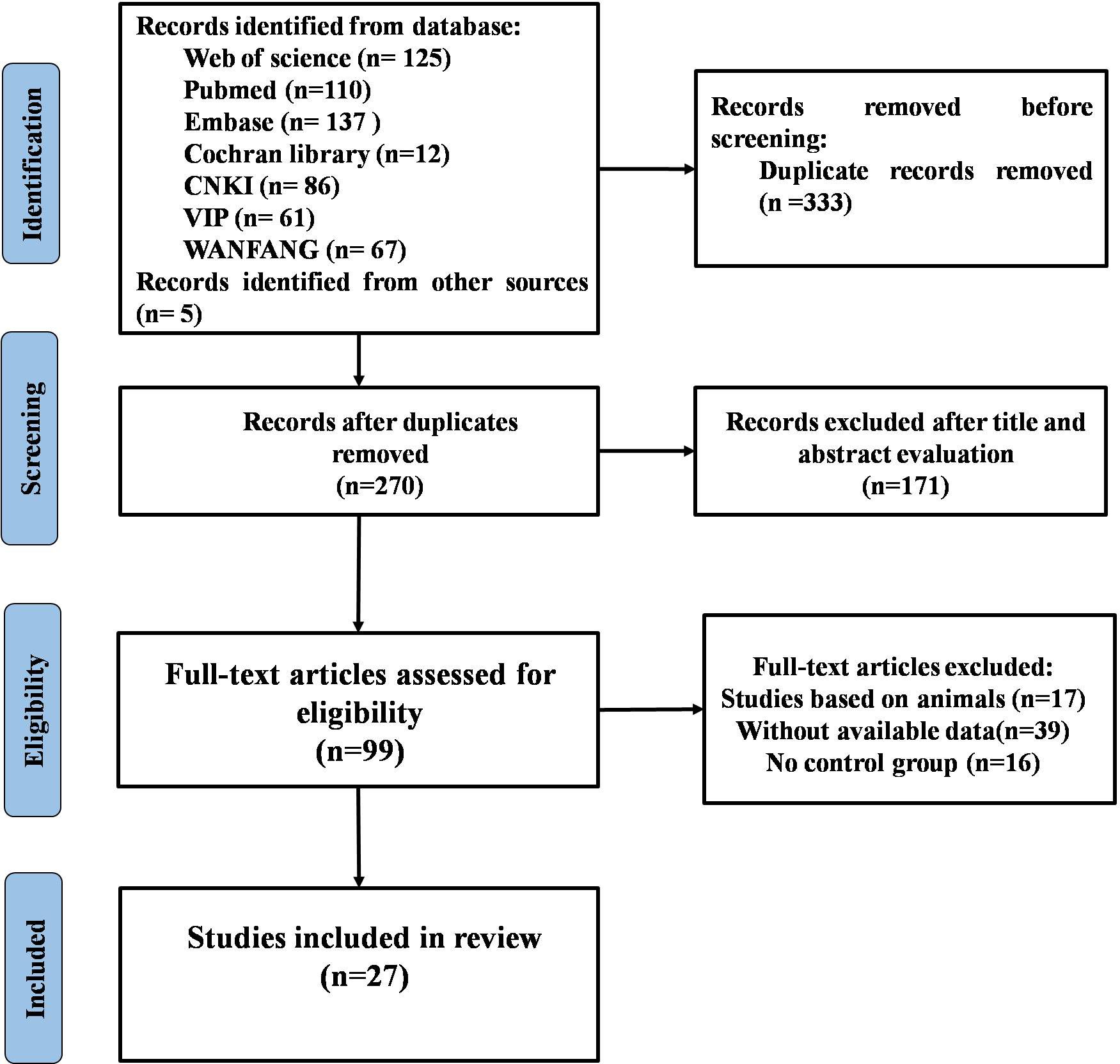
The selection process of the studies is presented in Figure 1. Our search returned a total of 603 studies. After the elimination of duplicates, 270 articles were reviewed. Of these, 171 were excluded by browsing the title and abstract, leaving 99 full-text studies to be reviewed. Finally, 27 studies that met the inclusion criteria were included and analyzed in the meta-analysis, with 2,263 participants. Details of the included studies are shown in Supplementary Table 1. All the studies were published between 2014 and 2023, with 19 conducted in Asia, 4 in Europe, 3 in North America, and 1 in Africa. The average age of all participants ranged from 39.1 to 80 years old, and most of the participants were female.
3.2 Quality evaluation results
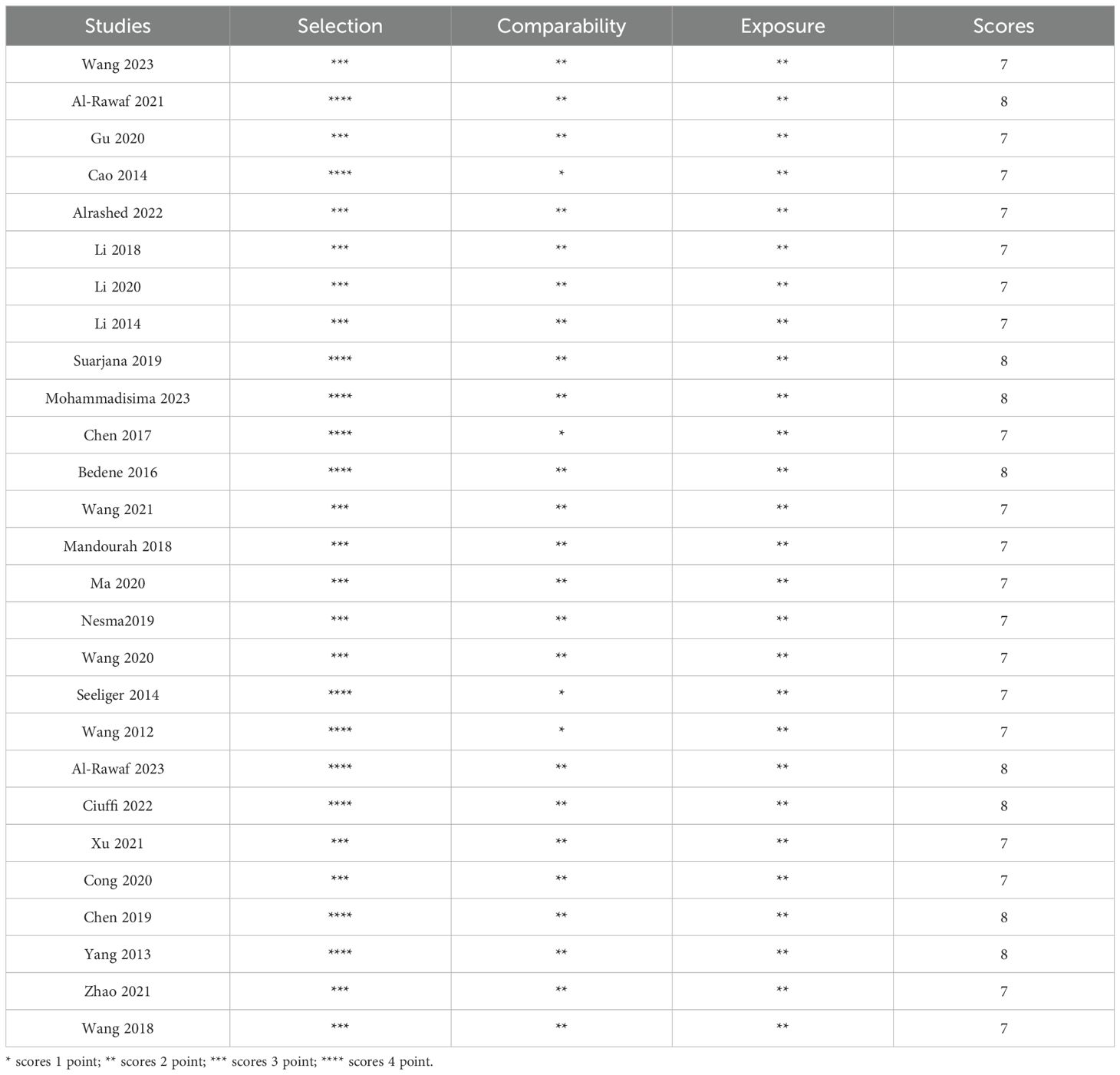
The quality of the included studies was evaluated by applying the NOS, as shown in Table 1. The NOS scores ranged from 7 to 8, and, as such, all the included studies were determined to be of high quality.
3.3 Results of the meta-analysis of the expression of each circulating miRNA in patients with osteoporosis
Six miRNAs were evidently upregulated and had been screened out as follows: miR-21-5p (WMD 0.88, 95% CI: 0.22 to 1.55, p = 0.009, I2 = 98.6%, Figure 2A), miR-125b-5p (WMD 6.63, 95% CI: 0.19 to 13.08, p = 0.044, I2 = 99.8%, Figure 2B), miR-483-5p (WMD 6.43, 95% CI: 3.26 to 9.61, p < 0.001, I2 = 92.1%, Figure 2C), miR-133a (WMD 1.43, 95% CI: 1.39 to 1.47, p < 0.001, I2 = 0%, Figure 2D), miR-422a (WMD 1, 95% CI: 0.28 to 1.72, p = 0.007, I2 =87.8%, Figure 2G), and miR-214-3p (WMD 2.03, 95% CI: 0.14 to 3.92, p = 0.036, I2 =93%, Figure 2H). Only one miRNA, miR-497-5p, was downregulated with a mean difference of -0.57 (WMD -0.57, 95% CI: -0.98 to -0.17, p = 0.005, I2 =85.6%, Figure 2F). The other two miRNAs, miR-148a-3p (WMD 7.09, 95% CI: -2.15 to 16.34, p = 0.133, I2 =99.7%, Figure 2E) and miR-122-5p (WMD -7.92, 95% CI: -21.21 to 5.37, p = 0.243, I2 =99.8%, Figure 2I), were observed to not have significant differences in expression.
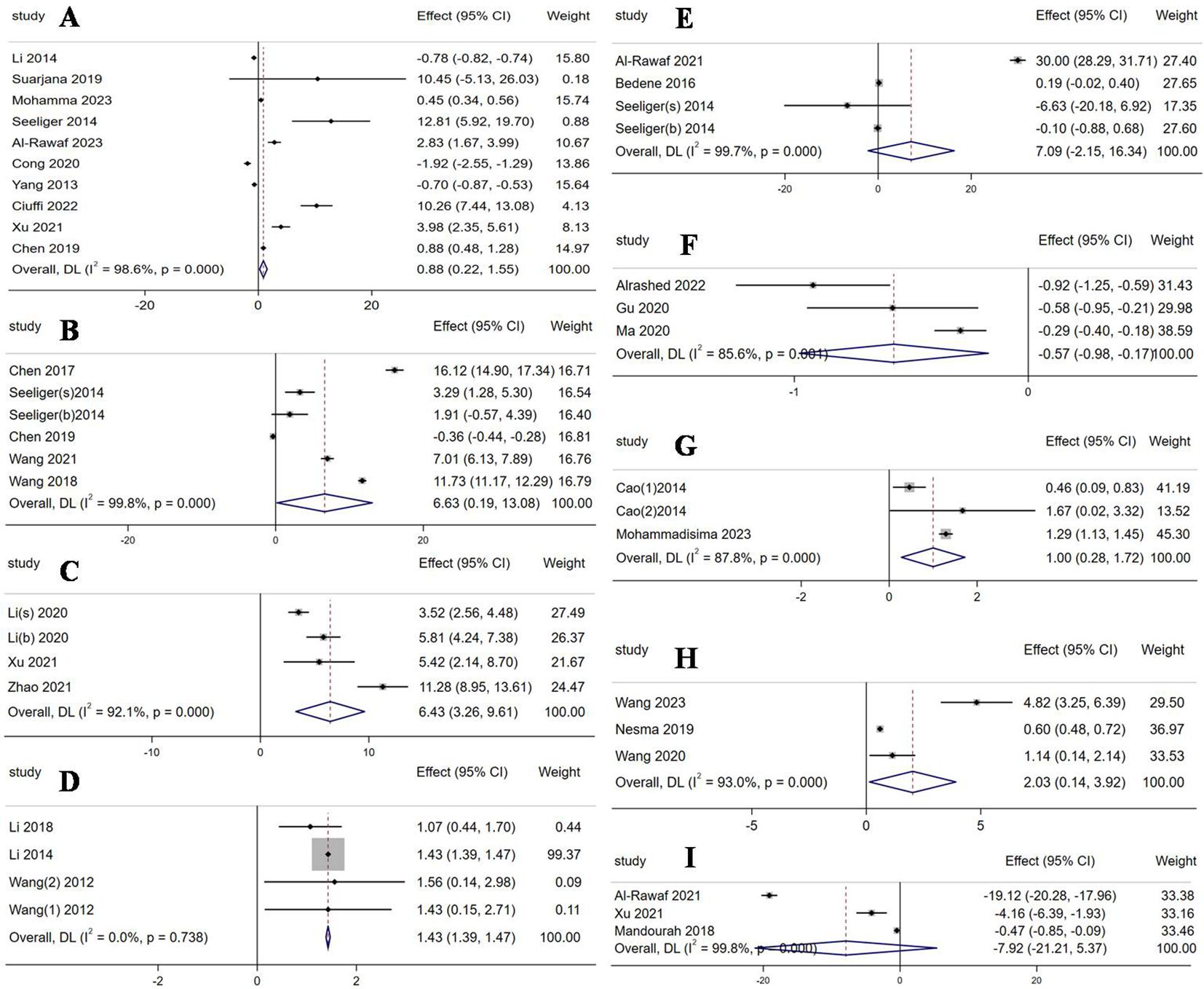
Figure 2. Forest plot of the expression of each circulating miRNA. (A) miR-21-5p; (B) miR-125b-5p; (C) miR-483-5p; (D) miR-133a; (E) miR-148a-3p; (F) miR-497-5p; (G) miR-422a; (H) miR-214-3p; (I) miR-122-5p.
3.4 Subgroup analysis and meta-regression analysis
In order to explore the source of heterogeneity, a subgroup analysis should be carried out for miR-21-5p, miR-125b-5p, miR-483-5p, miR-148a-3p, miR-497-5p, miR-422a, miR-214-3p, and miR-122-5p. Due to a lack of publications, only miR-21-5p was analyzed based on the sample sources (Supplementary Figure 1). The result showed that miR-21-5p was significantly upregulated in serum samples (WMD 5.66, 95% CI: 2.37 to 8.96, p = 0.001, I2 = 93.5%), but was observed to have no significant expression differences in plasma samples (WMD 0.03, 95% CI: -0.92 to 0.98, p = 0.948, I2 = 99.4%). The result of the subgroup analysis, as well as a subsequent meta-regression analysis (p = 0.058), indicated that the sample source was not a potential source of heterogeneity for miR-21-5p.
3.5 Sensitivity analysis and publication bias test
After excluding any individual study, a sensitivity analysis revealed that the results for the miRNAs in the present study were stable (Supplementary Figure 2). All the studies included in the present meta‐analysis were symmetrically distributed in a funnel plot (Supplementary Figure 3). The p-values of Begg’s and Egger’s tests are shown in Supplementary Table 2 (p > 0.05 for all), which indicated the absence of significant publication bias in the included studies.
4 Discussion
In the present meta-analysis, nine miRNAs were differentially expressed in more than one study for OP. Among these miRNAs, miR-21-5p, miR-125b-5p, miR-483-5p, miR-133a, miR-422a, and miR-214-3p were significantly upregulated, and miR-497-5p was significantly downregulated.
The dysregulation of miR-21-5p in OP has received a lot of attention from researchers in recent years. Notably, the direction of regulation for miR-21-5p was contradictory in the studies included in the meta-analysis, with seven studies reporting upregulation (15–21) and three studies reporting downregulation (22–24). This could be due to miR-21-5p being closely associated with both osteogenic and osteoclastic differentiation through a variety of mechanisms. miR-21-5p has been reported to promote osteogenesis by targeting the SOX2 (25), PLAP-1 (26), ACVR2B (27), ERK-MAPK (24, 28), Smad7-Smad1/5/8-Runx2 (29–31), PI3K/β-catenin (32), and PTEN/PI3K/Akt/HIF-1α pathways (33). The regulatory effects of miR-21-5p on osteoclasts are complex and involve multiple mechanisms. Sugatani et al. proposed a new molecular mechanism for osteoclastogenesis, namely the C-Fos/miR-21/PDCD4 positive feedback loop. C-Fos upregulated miR-21 expression and inhibited PDCD4 expression, which in turn promoted osteoclastogenesis (34). Subsequent studies also confirmed that miR-21 directly regulates osteoclast function by targeting PDCD4 (35–37). In addition, PTEN (38, 39), SKP2 (40), OPG (41), and FasL (42) have also been shown to be effective targets of miR-21-5p in promoting osteoclast differentiation. However, an inhibitory effect of miR-21-5p on osteoclast differentiation has also been reported, for instance, Huang et al. found that miR-21-5p was significantly decreased during osteoclast differentiation and that miR-21-5p inhibited osteoclast differentiation by acting on its target gene SKP2 (40). In juvenile idiopathic arthritis, miR-21-5p could inhibit the production of osteoclasts from rheumatoid arthritis fibroblast-like synovial cells induced by M-CSF (43). Unsatisfactorily, in the aforementioned studies, miR-21-5p promoted both osteogenic and osteoclastic differentiation, which is contradictory. This prevented us from accurately describing the mechanistic role of miR-21-5p in the pathogenesis of osteoporosis, so further studies are needed. Nevertheless, the results of this meta-analysis indicated that the level of miR-21-5p was upregulated in patients with osteoporosis, which was consistent with the result of a previously reported meta-analysis based on the robust rank aggregation method (13). Based on this finding, it could be considered that the expression level of miR-21-5p is related to the occurrence of osteoporosis, suggesting that miR-21-5p might be a promising biomarker for the diagnosis of osteoporosis. However, it should be noted that miR-21-5p was also associated with cardiovascular diseases (44), cancer (45), and other bone diseases (46), so a differential diagnosis is required in clinical practice.
For the expression level of miR-125b-5p in osteoporosis patients, four included studies reported upregulation (17, 47–49) and only one study reported downregulation (21). Wang et al. discovered that overexpression of miR-125b-5p was responsible for the development of postmenopausal osteoporosis and promoted its progression through the TRAF6 gene via the JAK2/STAT3 pathway (47). Xue et al. found that miR-125b attenuated the osteoblastic differentiation of periodontal ligament cells by targeting NKIRAS2 and enhancing NF-κB signaling (50). Wang et al. demonstrated that miR-125b-5p regulated the osteogenic differentiation of human mesenchymal stem cells by targeting BMPR1b and that inhibiting miR-125b-5p expression could enhance the capacity of bone defect repair in vivo (51). Huang et al. observed that the overexpression of miR-125b-5p inhibited osteoblastic differentiation by directly targeting Cbfβ and indirectly acting on Runx2 at the early stage of osteoblastic differentiation (52). On the contrary, a study from Japan showed that miR-125b-5p inhibited osteoclast formation by targeting Prdm1, encoding a transcriptional repressor of anti-osteoclastogenesis factors (53). Chen et al. found that irisin can upregulate the expression level of miR-125b-5p by targeting SIPA1L2, which regulated the Rap1/PI3K/AKT axis and finally increased the expression levels of the chondrogenic differentiation genes COL2A1, ACAN, and SOX9 (54). Overall, the available fundamental research showed that miR-125b-5p was more inclined to inhibit osteogenesis differentiation. The level of miR-125b-5p was consistently upregulated in patients with osteoporosis in the present meta-analysis. This suggests that miR-125b-5p might be a potential biomarker for the diagnosis of osteoporosis. However, it should be noted that miR-125b-5p has also been associated with stroke (55), Alzheimer’s disease (56), and cancer (57), so a differential diagnosis is required.
In the present study, miR-483-5p was evaluated in three studies (20, 58, 59), and all the studies indicated it was upregulated in osteoporosis. Li et al. revealed that miR-483-5p is involved in the pathogenesis of osteoporosis by reducing the apoptosis of osteoclasts (58). Zhao et al. also indicated that miR-483-5p could inhibit osteogenic differentiation by inhibiting SATB2 and activating the PI3K/AKT pathway (59). Peng et al. demonstrated that miR-483-5p regulated the RAS/MEK/ERK signaling pathway by targeting RPL31 and inhibiting its expression, thereby playing an inhibitory role in osteogenic differentiation (60). In light of these findings, miR-483-5p could be used as a diagnostic marker for osteoporosis. However, a differential diagnosis is necessary based on the association of miR-483-5p with other diseases (61, 62).
miR-133a was overexpressed in all the included studies (22, 63, 64). Wang et al. revealed that the overexpression of miR-133a suppressed osteoblast differentiation of bone marrow mesenchymal stem cells and silencing miR-133a resulted in positive effects on glucocorticoid-treated mesenchymal stem cells and on bone loss in glucocorticoid-induced osteoporosis animal models through the MAPK/ERK signaling pathway by targeting FGFR1 (65, 66). Li et al. found that miR-133a knockdown altered the levels of osteoclastogenesis-related factors in serum, increased lumbar spine BMD, and changed bone histomorphology in ovariectomized rats (63). In contrast, Zhou et al. suggested that miR-133a in osteoblasts significantly alleviated bone loss and microstructural and biomechanical properties in mice with mechanical unloading, contributing to osteopenia alleviation. Furthermore, miR-133a could also restrain osteoclastogenesis (67). However, in the present article, the direction of regulation for miR-133a in different studies was consistent, which meant that miR-133a might be a promising biomarker for osteoporosis diagnosis. Of course, a differential diagnosis is also required (68, 69).
Based on three studies, miR-497-5p was downregulated in osteoporotic patients with agreement on the direction of change (70–72). Zhao et al. showed that miR-497-5p enhanced osteogenic differentiation by repressing HMGA2 and impairing the JNK signaling pathway based on the MC3T3-E1 cell line (73). Lu et al. discovered the downregulation of SNHG1 and HIF1AN, in contrast with an elevation in miR-497-5p levels throughout osteogenic differentiation. By influencing miR-497-5p, SNHG1 exhibited the ability to modulate HIF1AN, thereby inhibiting osteogenic differentiation (74). Gu et al. suggested that miR-497-5p upregulation promoted osteoblast viability and collagen synthesis by activating the TGF-β1/Smads signaling pathway (70). Taken together, the discoveries of the mechanism studies were consistent with the result of the present meta-analysis, which suggested that miR-497-5p might be a novel reference for the diagnosis of osteoporosis. Notably, like other miRNAs, miR-497-5p was also involved in the development of other diseases (75–77).
For the expression direction of miR-422a, all the studies reported upregulation (16, 78). Baloun et al. observed that the serum concentration of miR-422a was positively correlated with markers of bone remodeling (β-CTX and P1NP), suggesting a role in the pathogenesis of osteoporosis (79). Cao et al. found significant upregulation of miR-422a in the low BMD group compared with the high BMD group using RT-qPCR analysis (P = 0.029). Furthermore, through bioinformatic target gene and RT-qPCR analyses, Cao et al. identified several potential target genes (CBL, CD226, IGF1, PAG1, and TOB2) of miR-422a that inhibit osteoclastogenesis and the expression of these genes correlated negatively with miR-422a expression (78). In our meta-analysis, the expression level of miR-422a was upregulated in osteoporosis patients. However, there are insufficient studies on the specific mechanism of miR-422a in the pathogenesis of osteoporosis. Therefore, further basic and clinical studies were required to confirm whether miR-422a could be a potential biomarker for osteoporosis diagnosis.
Consistent results for the direction of regulation were also found for miR-214-3p, and its expression level was elevated in patients with osteoporosis (80–82). miR-214-3p had been reported to suppress osteogenic differentiation by targeting Osterix (83), ATF4 (84, 85), and FGFR1 (86), and promote osteoclastogenesis by targeting phosphatase and tensin homolog (PTEN) (87). Furthermore, downregulation of miR-214-3p by Circ-ITCH (88), PTENP1 (82), and Nrf2 (89) has been reported to promote osteogenic differentiation and inhibit osteoclast differentiation, attenuating osteoporosis. These discoveries revealed that miR-214-3p has a crucial role in osteoporosis and might be a promising biomarker for the diagnosis of osteoporosis. Notably, a differential diagnosis is necessary due to the association of miR-214-3p with other diseases (90, 91).
In this meta-analysis, no significant expression differences were found for miR-122-5p (20, 92, 93) and miR-148a-3p (17, 92, 94) between patients with osteoporosis and healthy individuals. Only two studies reported an inhibitory effect of miR-122-5p on osteoblast proliferation/differentiation in osteoporosis (95, 96). Although miR-148a-3p has been reported to prevent osteoblast differentiation and bone remodeling in several fundamental studies (97–100), its expression direction in clinical studies was contradictory (17, 92, 94). Therefore, further studies are still required to confirm whether these two miRNAs can be used as diagnostic biomarkers for osteoporosis.
There were several limitations in our meta-analysis. First, although we had comprehensively searched enough databases, the number of studies and sample sizes of the studies available for meta-analysis were still small due to miRNA not being routinely used for the diagnosis of osteoporosis in clinical practice, which might impact the statistical power and generalizability of the results. Studies with larger sample sizes and deeper data analyses are needed to validate our findings. Second, data was partially obtained from bar charts or scatter charts that might not be accurate. Third, there was evident heterogeneity, reflected in the wide variation in miRNA expression in the osteoporosis population, limiting their use as diagnostic biomarkers for osteoporosis.
In conclusion, even with the limitations of the meta-analysis, it can be argued that miR-21-5p, miR-125b-5p, miR-483-5p, miR-133a, miR-148a-3p, miR-497-5p, miR-422a, miR-214-3p, and miR-122-5p are associated with osteoporosis and could be potential diagnostic biomarkers for osteoporosis. In the future, well-designed exhaustive studies should be conducted to validate the diagnostic value of these miRNAs for osteoporosis.
Data availability statement
The original contributions presented in the study are included in the article/Supplementary Material. Further inquiries can be directed to the corresponding authors.
Author contributions
JG: Writing – original draft. XZ: Writing – original draft. JD: Data curation, Formal analysis, Methodology, Writing – original draft. HZ: Data curation, Formal analysis, Methodology, Writing – original draft. XZ: Data curation, Formal analysis, Methodology, Writing – original draft. JJ: Data curation, Formal analysis, Methodology, Writing – original draft. WC: Conceptualization, Writing – review & editing.
Funding
The author(s) declare financial support was received for the research, authorship, and/or publication of this article. This study was supported by the Shandong Provincial Traditional Chinese Medicine Science and Technology project (Grant No. M-2023295); the Shandong Pharmaceutical Society Rational Drug Use project (Grant No. hlyy2024-03); the Shandong Medical Staff Science and Technology Innovation Plan project (Grant No. SDYWZGKCJHLH2023029); and the Tai’an City Science and Technology Innovation Development project (Grant No. 2022NS347, 2022ZC396, 2022NS232, 2023NS453, 2023NS389).
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Publisher’s note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Supplementary material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fendo.2024.1481649/full#supplementary-material
References
1. Wright NC, Looker AC, Saag KG, Curtis JR, Delzell ES, Randall S, et al. The recent prevalence of osteoporosis and low bone mass in the United States based on bone mineral density at the femoral neck or lumbar spine. J Bone Miner Res. (2014) 29:2520–6. doi: 10.1002/jbmr.2269
2. Yaacobi E, Sanchez D, Maniar H, Horwitz DS. Surgical treatment of osteoporotic fractures: An update on the principles of management. Injury. (2017) 48:S34–40. doi: 10.1016/j.injury.2017.08.036
3. Ensrud KE, Kats AM, Boyd CM, Diem SJ, Schousboe JT, Taylor BC, et al. Association of disease definition, comorbidity burden, and prognosis with hip fracture probability among late-life women. JAMA Intern Med. (2019) 179:1095–103. doi: 10.1001/jamainternmed.2019.0682
4. Camacho PM, Petak SM, Binkley N, Diab DL, Eldeiry LS, Farooki A, et al. American association of clinical endocrinologists/american college of endocrinology clinical practice guidelines for the diagnosis and treatment of postmenopausal osteoporosis-2020 update. Endocr Pract. (2020) 26:1–46. doi: 10.4158/gl-2020-0524suppl
5. LeBoff MS, Greenspan SL, Insogna KL, Lewiecki EM, Saag KG, Singer AJ, et al. The clinician's guide to prevention and treatment of osteoporosis. Osteoporos Int. (2022) 33:2049–102. doi: 10.1007/s00198-021-05900-y
6. Link TM, Heilmeier U. Bone quality-beyond bone mineral density. Semin Musculoskelet Radiol. (2016) 20:269–78. doi: 10.1055/s-0036-1592365
7. Rumancik S, Routh RH, Pathak RD, Burshell AL, Nauman EA. Assessment of bone quantity and distribution in adult lumbar scoliosis: new dual-energy x-ray absorptiometry methodology and analysis. Spine (Phila Pa 1976). (2005) 30:434–9. doi: 10.1097/01.brs.0000153344.06682.b2
8. Girardi FP, Parvataneni HK, Sandhu HS, Cammisa FP Jr, Grewal H, Schneider R, et al. Correlation between vertebral body rotation and two-dimensional vertebral bone density measurement. Osteoporos Int. (2001) 12:738–40. doi: 10.1007/s001980170049
9. Shyu AB, Wilkinson MF, van Hoof A. Messenger RNA regulation: to translate or to degrade. EMBO J. (2008) 27:471–81. doi: 10.1038/sj.emboj.7601977
10. Gebert LFR, MacRae IJ. Regulation of microRNA function in animals. Nat Rev Mol Cell Biol. (2019) 20:21–37. doi: 10.1038/s41580-018-0045-7
11. Zhao W, Shen G, Ren H, Liang D, Yu X, Zhang Z, et al. Therapeutic potential of MicroRNAs in osteoporosis function by regulating the biology of cells related to bone homeostasis. J Cell Physiol. (2018) 233:9191–208. doi: 10.1002/jcp.26939
12. Ciuffi S, Donati S, Marini F, Palmini G, Luzi E, Brandi ML, et al. Circulating MicroRNAs as novel biomarkers for osteoporosis and fragility fracture risk: is there a use in assessment risk? Int J Mol Sci. (2020) 21:6927–48. doi: 10.3390/ijms21186927
13. Pala E, Denkçeken T. Differentially expressed circulating miRNAs in postmenopausal osteoporosis: a meta-analysis. Biosci Rep. (2019) 39:BSR20190667. doi: 10.1042/bsr20190667
14. Liao XZ, Fu YH, Ma JY, Zhu WG, Yuan P. Non-vitamin K antagonist oral anticoagulants versus warfarin in patients with atrial fibrillation and peripheral artery disease: a systematic review and meta-analysis. Cardiovasc Drugs Ther. (2020) 34:391–9. doi: 10.1007/s10557-020-06962-6
15. Al-Rawaf HA, Gabr SA, Iqbal A, Alghadir AH. MicroRNAs as potential biopredictors for premenopausal osteoporosis: a biochemical and molecular study. BMC Womens Health. (2023) 23:481. doi: 10.1186/s12905-023-02626-3
16. Mohammadisima N, Farshbaf-Khalili A, Ostadrahimi A. Up-regulation of plasma miRNA-21 and miRNA-422a in postmenopausal osteoporosis. PloS One. (2023) 18:e0287458. doi: 10.1371/journal.pone.0287458
17. Seeliger C, Karpinski K, Haug AT, Vester H, Schmitt A, Bauer JS, et al. Five freely circulating miRNAs and bone tissue miRNAs are associated with osteoporotic fractures. J Bone Miner Res. (2014) 29:1718–28. doi: 10.1002/jbmr.2175
18. Suarjana IN, Isbagio H, Soewondo P, Rachman IA, Sadikin M, Prihartono J, et al. The role of serum expression levels of microrna-21 on bone mineral density in hypostrogenic postmenopausal women with osteoporosis: study on level of RANKL, OPG, TGFβ-1, sclerostin, RANKL/OPG ratio, and physical activity. Acta Med Indones. (2019) 51:245–52.
19. Ciuffi S, Marini F, Fossi C, Donati S, Giusti F, Botta A, et al. Circulating MicroRNAs as biomarkers of osteoporosis and fragility fractures. J Clin Endocrinol Metab. (2022) 107:2267–85. doi: 10.1210/clinem/dgac293
20. Xu J, Chen Y, Yu D, Zhang L, Dou X, Wu G, et al. Evaluation of the cargo contents and potential role of extracellular vesicles in osteoporosis. Aging (Albany NY). (2021) 13:19282–92. doi: 10.18632/aging.203264
21. Chen Z, Bemben MG, Bemben DA. Bone and muscle specific circulating MicroRNAs in postmenopausal women based on osteoporosis and sarcopenia status. Bone. (2019) 120:271–8. doi: 10.1016/j.bone.2018.11.001
22. Li H, Wang Z, Fu Q, Zhang J. Plasma miRNA levels correlate with sensitivity to bone mineral density in postmenopausal osteoporosis patients. Biomarkers. (2014) 19:553–6. doi: 10.3109/1354750x.2014.935957
23. Cong C, Tian J, Gao T, Zhou C, Wang Y, Cui X, et al. lncRNA GAS5 is upregulated in osteoporosis and downregulates miR-21 to promote apoptosis of osteoclasts. Clin Interv Aging. (2020) 15:1163–9. doi: 10.2147/cia.S235197
24. Yang N, Wang G, Hu C, Shi Y, Liao L, Shi S, et al. Tumor necrosis factor α suppresses the mesenchymal stem cell osteogenesis promoter miR-21 in estrogen deficiency-induced osteoporosis. J Bone Miner Res. (2013) 28:559–73. doi: 10.1002/jbmr.1798
25. Trohatou O, Zagoura D, Bitsika V, Pappa KI, Antsaklis A, Anagnou NP, et al. Sox2 suppression by miR-21 governs human mesenchymal stem cell properties. Stem Cells Transl Med. (2014) 3:54–68. doi: 10.5966/sctm.2013-0081
26. Li C, Li C, Yue J, Huang X, Chen M, Gao J, et al. miR-21 and miR-101 regulate PLAP-1 expression in periodontal ligament cells. Mol Med Rep. (2012) 5:1340–6. doi: 10.3892/mmr.2012.797
27. Wei F, Liu D, Feng C, Zhang F, Yang S, Hu Y, et al. microRNA-21 mediates stretch-induced osteogenic differentiation in human periodontal ligament stem cells. Stem Cells Dev. (2015) 24:312–9. doi: 10.1089/scd.2014.0191
28. Mei Y, Bian C, Li J, Du Z, Zhou H, Yang Z, et al. miR-21 modulates the ERK-MAPK signaling pathway by regulating SPRY2 expression during human mesenchymal stem cell differentiation. J Cell Biochem. (2013) 114:1374–84. doi: 10.1002/jcb.24479
29. Song Q, Zhong L, Chen C, Tang Z, Liu H, Zhou Y, et al. miR-21 synergizes with BMP9 in osteogenic differentiation by activating the BMP9/Smad signaling pathway in murine multilineage cells. Int J Mol Med. (2015) 36:1497–506. doi: 10.3892/ijmm.2015.2363
30. Li H, Yang F, Wang Z, Fu Q, Liang A. MicroRNA-21 promotes osteogenic differentiation by targeting small mothers against decapentaplegic 7. Mol Med Rep. (2015) 12:1561–7. doi: 10.3892/mmr.2015.3497
31. Li X, Guo L, Liu Y, Su Y, Xie Y, Du J, et al. MicroRNA-21 promotes osteogenesis of bone marrow mesenchymal stem cells via the Smad7-Smad1/5/8-Runx2 pathway. Biochem Biophys Res Commun. (2017) 493:928–33. doi: 10.1016/j.bbrc.2017.09.119
32. Meng YB, Li X, Li ZY, Zhao J, Yuan XB, Ren Y, et al. microRNA-21 promotes osteogenic differentiation of mesenchymal stem cells by the PI3K/β-catenin pathway. J Orthop Res. (2015) 33:957–64. doi: 10.1002/jor.22884
33. Yang C, Liu X, Zhao K, Zhu Y, Hu B, Zhou Y, et al. miRNA-21 promotes osteogenesis via the PTEN/PI3K/Akt/HIF-1α pathway and enhances bone regeneration in critical size defects. Stem Cell Res Ther. (2019) 10:65. doi: 10.1186/s13287-019-1168-2
34. Sugatani T, Vacher J, Hruska KA. A microRNA expression signature of osteoclastogenesis. Blood. (2011) 117:3648–57. doi: 10.1182/blood-2010-10-311415
35. Zhang Y, Tian Y, Yang X, Zhao Z, Feng C, Zhang Y. MicroRNA−21 serves an important role during PAOO−facilitated orthodontic tooth movement. Mol Med Rep. (2020) 22:474–82. doi: 10.3892/mmr.2020.11107
36. Xu Z, Liu X, Wang H, Li J, Dai L, Li J, et al. Lung adenocarcinoma cell-derived exosomal miR-21 facilitates osteoclastogenesis. Gene. (2018) 666:116–22. doi: 10.1016/j.gene.2018.05.008
37. Hu CH, Sui BD, Du FY, Shuai Y, Zheng CX, Zhao P, et al. miR-21 deficiency inhibits osteoclast function and prevents bone loss in mice. Sci Rep. (2017) 7:43191. doi: 10.1038/srep43191
38. Zhao Q, Liu C, Xie Y, Tang M, Luo G, Chen X, et al. Lung Cancer Cells Derived Circulating miR-21 Promotes Differentiation of Monocytes into Osteoclasts. Onco Targets Ther. (2020) 13:2643–56. doi: 10.2147/ott.S232876
39. Wang S, Liu Z, Wang J, Ji X, Yao Z, Wang X. miR−21 promotes osteoclastogenesis through activation of PI3K/Akt signaling by targeting Pten in RAW264. 7 Cells Mol Med Rep. (2020) 21:1125–32. doi: 10.3892/mmr.2020.10938
40. Huang Y, Yang Y, Wang J, Yao S, Yao T, Xu Y, et al. miR-21-5p targets SKP2 to reduce osteoclastogenesis in a mouse model of osteoporosis. J Biol Chem. (2021) 296:100617. doi: 10.1016/j.jbc.2021.100617
41. Pitari MR, Rossi M, Amodio N, Botta C, Morelli E, Federico C, et al. Inhibition of miR-21 restores RANKL/OPG ratio in multiple myeloma-derived bone marrow stromal cells and impairs the resorbing activity of mature osteoclasts. Oncotarget. (2015) 6:27343–58. doi: 10.18632/oncotarget.4398
42. Sugatani T, Hruska KA. Down-regulation of miR-21 biogenesis by estrogen action contributes to osteoclastic apoptosis. J Cell Biochem. (2013) 114:1217–22. doi: 10.1002/jcb.24471
43. Li HW, Zeng HS. Regulation of JAK/STAT signal pathway by miR-21 in the pathogenesis of juvenile idiopathic arthritis. World J Pediatr. (2020) 16:502–13. doi: 10.1007/s12519-019-00268-w
44. Xu L, Tian L, Yan Z, Wang J, Xue T, Sun Q. Diagnostic and prognostic value of miR-486-5p, miR-451a, miR-21-5p and monocyte to high-density lipoprotein cholesterol ratio in patients with acute myocardial infarction. Heart Vessels. (2023) 38:318–31. doi: 10.1007/s00380-022-02172-2
45. Wang JH, Bai ZZ, Niu XD, Zhu CL, Liang T, Hu YL, et al. Serum extracellular vesicle-derived miR-21-5p and miR-26a-5p as non-invasive diagnostic potential biomarkers for gastric cancer: A preliminary study. Int J Biol Markers. (2024) 39:217–25. doi: 10.1177/03936155241261390
46. Chen C, Liu YM, Fu BL, Xu LL, Wang B. MicroRNA-21: an emerging player in bone diseases. Front Pharmacol. (2021) 12:722804. doi: 10.3389/fphar.2021.722804
47. Wang G, Zhang L, Yan C, Wang F, Zhang Y. Overexpression of miR125b promotes osteoporosis through miR-125b-tRAF6 pathway in postmenopausal ovariectomized rats. Diabetes Metab Syndr Obes. (2021) 14:671–82. doi: 10.2147/dmso.S288338
48. Wang C, He H, Wang L, Jiang Y, Xu Y. Reduced miR-144-3p expression in serum and bone mediates osteoporosis pathogenesis by targeting RANK. Biochem Cell Biol. (2018) 96:627–35. doi: 10.1139/bcb-2017-0243
49. Chen H, Jiang H, Can D, Xu H, Zhang K, Guo S. Evaluation of microRNA 125b as a potential biomarker for postmenopausal osteoporosis. Trop J Pharm Res. (2017) 16:641–7. doi: 10.4314/tjpr.v16i3.20
50. Xue N, Qi L, Zhang G, Zhang Y. miRNA-125b regulates osteogenic differentiation of periodontal ligament cells through NKIRAS2/NF-κB pathway. Cell Physiol Biochem. (2018) 48:1771–81. doi: 10.1159/000492350
51. Wang H, Xie Z, Hou T, Li Z, Huang K, Gong J, et al. MiR-125b regulates the osteogenic differentiation of human mesenchymal stem cells by targeting BMPR1b. Cell Physiol Biochem. (2017) 41:530–42. doi: 10.1159/000457013
52. Huang K, Fu J, Zhou W, Li W, Dong S, Yu S, et al. MicroRNA-125b regulates osteogenic differentiation of mesenchymal stem cells by targeting Cbfβ in vitro. Biochimie. (2014) 102:47–55. doi: 10.1016/j.biochi.2014.02.005
53. Ito S, Minamizaki T, Kohno S, Sotomaru Y, Kitaura Y, Ohba S, et al. Overexpression of miR-125b in Osteoblasts Improves Age-Related Changes in Bone Mass and Quality through Suppression of Osteoclast Formation. Int J Mol Sci. (2021) 22:6745–57. doi: 10.3390/ijms22136745
54. Chen T, Peng Y, Hu W, Shi H, Li P, Que Y, et al. Irisin enhances chondrogenic differentiation of human mesenchymal stem cells via Rap1/PI3K/AKT axis. Stem Cell Res Ther. (2022) 13:392. doi: 10.1186/s13287-022-03092-8
55. Loggini A, Hornik J, Hornik A. The role of MicroRNAs as super-early biomarkers in acute ischemic stroke: A systematic review. Clin Neurol Neurosurg. (2024) 244:108416. doi: 10.1016/j.clineuro.2024.108416
56. McKeever PM, Schneider R, Taghdiri F, Weichert A, Multani N, Brown RA, et al. MicroRNA expression levels are altered in the cerebrospinal fluid of patients with young-onset alzheimer's disease. Mol Neurobiol. (2018) 55:8826–41. doi: 10.1007/s12035-018-1032-x
57. Papanota AM, Karousi P, Kontos CK, Artemaki PI, Liacos CI, Papadimitriou MA, et al. A cancer-related microRNA signature shows biomarker utility in multiple myeloma. Int J Mol Sci. (2021) 22:13144–61. doi: 10.3390/ijms222313144
58. Li K, Chen S, Cai P, Chen K, Li L, Yang X, et al. MiRNA-483-5p is involved in the pathogenesis of osteoporosis by promoting osteoclast differentiation. Mol Cell Probes. (2020) 49:101479. doi: 10.1016/j.mcp.2019.101479
59. Zhao F, Xu Y, Ouyang Y, Wen Z, Zheng G, Wan T, et al. Silencing of miR-483-5p alleviates postmenopausal osteoporosis by targeting SATB2 and PI3K/AKT pathway. Aging (Albany NY). (2021) 13:6945–56. doi: 10.18632/aging.202552
60. Peng H, Yu Y, Gu H, Qi B, Yu A. MicroRNA-483-5p inhibits osteogenic differentiation of human bone marrow mesenchymal stem cells by targeting the RPL31-mediated RAS/MEK/ERK signaling pathway. Cell Signal. (2022) 93:110298. doi: 10.1016/j.cellsig.2022.110298
61. Nagaraj S, Want A, Laskowska-Kaszub K, Fesiuk A, Vaz S, Logarinho E, et al. Candidate alzheimer's disease biomarker miR-483-5p lowers TAU phosphorylation by direct ERK1/2 repression. Int J Mol Sci. (2021) 22:3653–71. doi: 10.3390/ijms22073653
62. Gallo W, Esguerra JLS, Eliasson L, Melander O. miR-483-5p associates with obesity and insulin resistance and independently associates with new onset diabetes mellitus and cardiovascular disease. PloS One. (2018) 13:e0206974. doi: 10.1371/journal.pone.0206974
63. Li Z, Zhang W, Huang Y. MiRNA-133a is involved in the regulation of postmenopausal osteoporosis through promoting osteoclast differentiation. Acta Biochim Biophys Sin (Shanghai). (2018) 50:273–80. doi: 10.1093/abbs/gmy006
64. Wang Y, Li L, Moore BT, Peng XH, Fang X, Lappe JM, et al. miR-133a in human circulating monocytes: a potential biomarker associated with postmenopausal osteoporosis. PloS One. (2012) 7:e34641. doi: 10.1371/journal.pone.0034641
65. Wang G, Wan L, Zhang L, Yan C, Zhang Y. MicroRNA-133a regulates the viability and differentiation fate of bone marrow mesenchymal stem cells via MAPK/ERK signaling pathway by targeting FGFR1. DNA Cell Biol. (2021) 40:1112–23. doi: 10.1089/dna.2021.0206
66. Wang G, Wang F, Zhang L, Yan C, Zhang Y. miR-133a silencing rescues glucocorticoid-induced bone loss by regulating the MAPK/ERK signaling pathway. Stem Cell Res Ther. (2021) 12:215. doi: 10.1186/s13287-021-02278-w
67. Zhou Y, Chen X, Zhu Z, Bi D, Ma S. miR-133a delivery to osteoblasts ameliorates mechanical unloading-triggered osteopenia progression in vitro and in vivo. Int Immunopharmacol. (2021) 97:107613. doi: 10.1016/j.intimp.2021.107613
68. Li S, Zhi F, Hu M, Xue X, Mo Y. miR-133a is a potential target for arterial calcification in patients with end-stage renal disease. Int Urol Nephrol. (2022) 54:217–24. doi: 10.1007/s11255-021-02906-7
69. Ferreira LR, Frade AF, Santos RH, Teixeira PC, Baron MA, Navarro IC, et al. MicroRNAs miR-1, miR-133a, miR-133b, miR-208a and miR-208b are dysregulated in Chronic Chagas disease Cardiomyopathy. Int J Cardiol. (2014) 175:409–17. doi: 10.1016/j.ijcard.2014.05.019
70. Gu Z, Xie D, Huang C, Ding R, Zhang R, Li Q, et al. MicroRNA-497 elevation or LRG1 knockdown promotes osteoblast proliferation and collagen synthesis in osteoporosis via TGF-β1/Smads signalling pathway. J Cell Mol Med. (2020) 24:12619–32. doi: 10.1111/jcmm.15826
71. Alrashed MM, Alshehry AS, Ahmad M, He J, Wang Y, Xu Y. miRNA let-7a-5p targets RNA KCNQ1OT1 and participates in osteoblast differentiation to improve the development of osteoporosis. Biochem Genet. (2022) 60:370–81. doi: 10.1007/s10528-021-10105-3
72. Ma J, Lin X, Chen C, Li S, Zhang S, Chen Z, et al. Circulating miR-181c-5p and miR-497-5p Are Potential Biomarkers for Prognosis and Diagnosis of Osteoporosis. J Clin Endocrinol Metab. (2020) 105:1445–60. doi: 10.1210/clinem/dgz300
73. Zhao H, Yang Y, Wang Y, Feng X, Deng A, Ou Z, et al. MicroRNA-497-5p stimulates osteoblast differentiation through HMGA2-mediated JNK signaling pathway. J Orthop Surg Res. (2020) 15:515. doi: 10.1186/s13018-020-02043-4
74. Lu Y, Pan K, Zhang Y, Peng J, Cao D, Li X. The mechanism of lncRNA SNHG1 in osteogenic differentiation via miR-497-5p/ HIF1AN axis. Connect Tissue Res. (2024) 65:63–72. doi: 10.1080/03008207.2023.2281321
75. Zhu W, Zhang H, Gao J, Xu Y. Silencing of miR-497-5p inhibits cell apoptosis and promotes autophagy in Parkinson's disease by upregulation of FGF2. Environ Toxicol. (2021) 36:2302–12. doi: 10.1002/tox.23344
76. Chen T, Zhang X, Qian W, Zhou R, Su M, Ma Y. Serum miR-497-5p serves as a diagnostic biomarker for acute coronary syndrome and predicts the occurrence of major adverse cardiovascular events after percutaneous coronary intervention. Bioengineered. (2022) 13:8266–76. doi: 10.1080/21655979.2022.2051885
77. Chen X, Zhou L, Han Y, Lin S, Zhou L, Wang W, et al. miR-497-5p expression and biological activity in gastric cancer. J Cancer. (2024) 15:3995–4006. doi: 10.7150/jca.90087
78. Cao Z, Moore BT, Wang Y, Peng XH, Lappe JM, Recker RR, et al. miR-422a as a potential cellular microRNA biomarker for postmenopausal osteoporosis. PloS One. (2014) 9:e97098. doi: 10.1371/journal.pone.0097098
79. Baloun J, Pekacova A, Wenchich L, Hruskova H, Senolt L, Svec X, et al. Menopausal transition: prospective study of estrogen status, circulating MicroRNAs, and biomarkers of bone metabolism. Front Endocrinol (Lausanne). (2022) 13:864299. doi: 10.3389/fendo.2022.864299
80. Wang C, Wang P, Li F, Li Y, Zhao M, Feng H, et al. Adenovirus-associated anti-miRNA-214 regulates bone metabolism and prevents local osteoporosis in rats. Front Bioeng Biotechnol. (2023) 11:1164252. doi: 10.3389/fbioe.2023.1164252
81. Mohamad N, Nabih ES, Zakaria ZM, Nagaty MM, Metwaly RG. Insight into the possible role of miR-214 in primary osteoporosis via osterix. J Cell Biochem. (2019) 120:15518–26. doi: 10.1002/jcb.28818
82. Wang CG, Wang L, Yang T, Su SL, Hu YH, Zhong D. Pseudogene PTENP1 sponges miR-214 to regulate the expression of PTEN to modulate osteoclast differentiation and attenuate osteoporosis. Cytotherapy. (2020) 22:412–23. doi: 10.1016/j.jcyt.2020.04.090
83. Shi K, Lu J, Zhao Y, Wang L, Li J, Qi B. MicroRNA-214 suppresses osteogenic differentiation of C2C12 myoblast cells by targeting Osterix. Bone. (2013) 55:487–94. doi: 10.1016/j.bone.2013.04.002
84. Wang X, Guo B, Li Q, Peng J, Yang Z, Wang A, et al. miR-214 targets ATF4 to inhibit bone formation. Nat Med. (2013) 19:93–100. doi: 10.1038/nm.3026
85. Zhang K, Liu X, Tang Y, Liu Z, Yi Q, Wang L, et al. Fluid shear stress promotes osteoblast proliferation and suppresses mitochondrial-mediated osteoblast apoptosis through the miR-214-3p-ATF4 signaling axis. Physiol Res. (2022) 71:527–38. doi: 10.33549/physiolres.934917
86. Yang L, Ge D, Cao X, Ge Y, Chen H, Wang W, et al. MiR-214 attenuates osteogenic differentiation of mesenchymal stem cells via targeting FGFR1. Cell Physiol Biochem. (2016) 38:809–20. doi: 10.1159/000443036
87. Zhao C, Sun W, Zhang P, Ling S, Li Y, Zhao D, et al. miR-214 promotes osteoclastogenesis by targeting Pten/PI3k/Akt pathway. RNA Biol. (2015) 12:343–53. doi: 10.1080/15476286.2015.1017205
88. Zhong D, Xu GZ, Wu JZ, Liu H, Tang JY, Wang CG. Circ-ITCH sponges miR-214 to promote the osteogenic differentiation in osteoporosis via upregulating YAP1. Cell Death Dis. (2021) 12:340. doi: 10.1038/s41419-021-03586-y
89. Hong J, Shi Z, Li C, Ji X, Li S, Chen Y, et al. Virtual screening identified natural Keap1-Nrf2 PPI inhibitor alleviates inflammatory osteoporosis through Nrf2-mir214-Traf3 axis. Free Radic Biol Med. (2021) 171:365–78. doi: 10.1016/j.freeradbiomed.2021.05.020
90. Dong H, Yan J, Huang P, Wang X, Zhang R, Zhang C, et al. miR-214-3p promotes the pathogenesis of Parkinson's disease by inhibiting autophagy. BioMed Pharmacother. (2024) 171:116123. doi: 10.1016/j.biopha.2024.116123
91. Liu L, Xiao W, Yang Z, Wang Q, Yi J. Human umbilical cord mesenchymal stem cell-derived exosomal miR-214-3p regulates the progression of gallbladder cancer by regulating ACLY/GLUT1. Adv Clin Exp Med. (2024) 33:499–510. doi: 10.17219/acem/169976
92. Al-Rawaf HA, Alghadir AH, Gabr SA. Circulating microRNA expression, vitamin D, and hypercortisolism as predictors of osteoporosis in elderly postmenopausal women. Dis Markers. (2021) 2021:3719919. doi: 10.1155/2021/3719919
93. Mandourah AY, Ranganath L, Barraclough R, Vinjamuri S, Hof RV, Hamill S, et al. Circulating MicroRNAs as potential diagnostic biomarkers for osteoporosis. Sci Rep. (2018) 8:8421. doi: 10.1038/s41598-018-26525-y
94. Bedene A, Mencej Bedrač S, Ješe L, Marc J, Vrtačnik P, Preželj J, et al. MiR-148a the epigenetic regulator of bone homeostasis is increased in plasma of osteoporotic postmenopausal women. Wien Klin Wochenschr. (2016) 128:519–26. doi: 10.1007/s00508-016-1141-3
95. Xu X, Chen Y, Tan B, Wang D, Yuan Z, Wang F. Circular RNA circ_0011269 sponges miR-122 to regulate RUNX2 expression and promotes osteoporosis progression. J Cell Biochem. (2020) 121:4819–26. doi: 10.1002/jcb.29709
96. Meng YC, Lin T, Jiang H, Zhang Z, Shu L, Yin J, et al. miR-122 exerts inhibitory effects on osteoblast proliferation/differentiation in osteoporosis by activating the PCP4-mediated JNK pathway. Mol Ther Nucleic Acids. (2020) 20:345–58. doi: 10.1016/j.omtn.2019.11.038
97. Yuan H, Xu X, Feng X, Zhu E, Zhou J, Wang G, et al. A novel long noncoding RNA PGC1β-OT1 regulates adipocyte and osteoblast differentiation through antagonizing miR-148a-3p. Cell Death Differ. (2019) 26:2029–45. doi: 10.1038/s41418-019-0296-7
98. Liu N, Sun Y. microRNA-148a-3p-targeting p300 protects against osteoblast differentiation and osteoporotic bone reconstruction. Regener Med. (2021) 16:435–49. doi: 10.2217/rme-2020-0006
99. Pan B, Zheng L, Liu S, Fang J, Lou C, Hu X, et al. MiR-148a deletion protects from bone loss in physiological and estrogen-deficient mice by targeting NRP1. Cell Death Discovery. (2022) 8:470. doi: 10.1038/s41420-022-01261-5
Keywords: microRNA, expression, diagnosis, osteoporosis, meta-analysis
Citation: Gao J, Zhang X, Ding J, Zhang H, Zhang X, Jiang J and Chen W (2024) The characteristic expression of circulating MicroRNAs in osteoporosis: a systematic review and meta-analysis. Front. Endocrinol. 15:1481649. doi: 10.3389/fendo.2024.1481649
Received: 16 August 2024; Accepted: 18 November 2024;
Published: 16 December 2024.
Edited by:
Maria Felicia Faienza, University of Bari Aldo Moro, ItalyReviewed by:
Zhongyang Sun, Nanjing University, ChinaDonatella Mentino, University of Bari Aldo Moro, Italy
Copyright © 2024 Gao, Zhang, Ding, Zhang, Zhang, Jiang and Chen. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Juan Jiang, amlhbmd3ZW5qaW5nMjAwMEAxMjYuY29t; Wenwen Chen, d2VuLTg2MDUyMUAxNjMuY29t
†These authors have contributed equally to this work
‡ORCID: Juan Jiang, orcid.org/0009-0008-2821-0954
 Jie Gao1†
Jie Gao1† Wenwen Chen
Wenwen Chen