
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Vet. Sci. , 26 March 2025
Sec. Animal Nutrition and Metabolism
Volume 12 - 2025 | https://doi.org/10.3389/fvets.2025.1469169
This article is part of the Research Topic From Cow to Shelf: Exploring Milk Fatty Acids as Biomarkers to Improve the Productivity and Sustainability of Dairy Systems View all articles
Introduction: This study was designed to determine the effects of diets containing different-quality forages and concentrate content on milk composition and lipogenic enzyme expression in the mammary glands of dairy cows using RNA sequencing.
Methods: Thirty Holstein cows were randomly assigned to one of three treatments: mixed forage consisted of hay, silage, and alfalfa forage to a concentrate ratio of 54:46 (MF); corn stover as forage and forage to a concentrate ratio of 35:65 (CSA); corn stover as forage and forage to a concentrate ratio the same as MF (CSB). Mammary tissue biopsies were performed to analyze lipogenic enzyme expression using RNA-seq.
Results: Treatments did not affect dry matter intake, milk protein, or lactose. The milk yield, fat content and saturated fatty acids (SFAs), and short- and medium-chain fatty acids (SMFAs) contents were lower in CSA and CSB than in MF. Still, the unsaturated FA and long-chain FA contents were higher than in MF. We used RNA-seq to compare analyses of three groups of mammary tissue in transcriptomics, Gene Ontology and KEGG-enriched pathways. Differentially expressed genes (DEGs) involved in lipid metabolic pathways and the FA biosynthesis pathway in MF were significantly higher than in CSB. In contrast, DEGs of FA extension and unsaturated FA synthesis pathway were significantly lower than in CSB.
Conclusion: Corn stover as a forage diet reduced the milk yield, fat content, SMFAs, SFAs, and the gene expression of mammary lipogenic enzymes in dairy cows.
Milk yield and its components are two of the most important economic traits influenced by nutrition, environmental factors, cow breed, cow age, and seasonality (1). Milk fat is a part of milk components that is a great concern for consumers (2), and its modulation through diet has been researched in many studies. Milk fat is the milk component most susceptible to diet manipulation in cows (3). Diets with different qualities of forages had distinct effects on milk fatty acid (FA) composition, being related to the expression of lipid metabolism-related genes in the milk FA composition. Therefore, diet may be key to milk FA-regulating mechanisms (4).
Recently, Chinese wildrye and alfalfa hay have been used as the main forages for feeding dairy cows in China. However, alfalfa hay is of high nutrition but in short supply. Large amounts of corn stover are produced yearly (5). Still, the low nutrient level in corn stover might result in lower livestock production if it is used as the main forage source. A diet with Chinese wildrye and alfalfa hay as the main forages increased the milk yield, milk fat and protein contents, and the expression of mammary lipogenic enzymes compared with corn stover as the forage fed to ruminants (6, 7). Low-quality forage often requires more concentrate to maintain the nutrition level of the diet. Applying a high ratio of concentrate in dairy cow diets may decrease milk fat content, especially when the concentrates contain a large amount of starch (8). Feeding different-quality diets also affects the mRNA abundance of milk lipid metabolism-related genes in the mammary glands of dairy cows (9, 10). Milk fat synthesis is a complex procedure involving a wide gene network with ambiguous regulation (11).
Along with the rapid development and cost reduction of next-generation sequencing, sequence-based assays of transcriptome RNA-seq have recently become a comprehensive and accurate tool for gene expression pattern analyses. RNA-seq enables analysis of the complexity of whole eukaryotic transcriptomes with less bias, greater dynamic range, lower frequency of false-positive signals, and higher reproducibility than microarray technology (12, 13). Several studies on bovine milk transcriptome have used RNA-seq techniques (14–16). However, no studies have been published on how different forage diets affect the bovine mammary gland transcriptome using RNA-seq. The present study aimed to identify genes and pathways involved in diets composed of different-quality forages on mammary lipogenic enzymes in mid-lactating dairy cows using RNA-seq technology. Comprehensive information on the molecular events of diet-affected milk fat will enhance our understanding of milk composition and lead to the identification of novel genes or transcripts that regulate milk fat.
This study was conducted in accordance with the guidelines set forth by the Animal Care and Ethics Committee of Inner Mongolia Agricultural University (NO. 2023016). A total of 30 multiparous Holstein dairy cows (body weight = 554 ± 21 kg, days in milk = 120 ± 24 d) were randomly assigned to one of three groups (10 cows per group) with a 60-day experimental period. The last three experimental days were those during which the sample collection took place. Cows were housed in individual stalls with free access to drinking water, fed TMR ad libitum, and milked twice daily at 06:00 and 18:00 h during this trial. Three TMRs with different forages and concentrate-to-forage ratios were designed. The diets (on DM base) were: [1] mixed forages including hay (4%), corn silage (27%), and alfalfa (23%) with 46% of concentrate (MF); [2] corn stover as forage formulated similar proximate nutrient levels with MF, and forage to a concentrate ratio of 35:65 (CSA); and [3] 54% of corn stover and 46% of concentrate (CSB).
The feed intake for each cow was measured daily and adjusted using the DM of TMR to determine the daily DM intake (DMI). Samples of TMR and orts were collected for three consecutive days and stored at −20°C before analysis of the nutrient components of diets (Tables 1, 2). The milk yield was recorded daily throughout the experiment. Milk samples were collected during the sampling period of the last 3 days. The samples were mixed thoroughly and stored at 4°C with preservatives until analysis of milk fat, protein, and lactose using MilkoScanTM Minor Type 78110 (FOSS Analytical A/S 69, DK 3400, Hillerod, Denmark) and somatic cell count (SCC) using ADAM SCC II (Nanoentek Inc., Seoul, Korea).
Milk FAs were extracted using a previously described procedure (6). FA methyl esters from milk and feed samples were analyzed using a Shimadzu GC 2014 system (Shimadzu, Kyoto, Japan) gas chromatography equipped with a 100 m HP 88 fused silica capillary column × 0.25 mm i.d., coated with 0.2 μm film thickness (Agilent J&W, Santa Clara, CA, USA).
Five cows from each treatment were randomly selected to perform mammary tissue biopsies after milking on the last day of the experiment. The biopsy was conducted according to a previously described procedure (17). The mammary tissue biopsies (~500 mg/animal) were rinsed in a 0.9% sterile saline solution and then frozen in liquid nitrogen. The samples were kept at −80°C until RNA extraction.
Total RNA was extracted from each frozen mammary biopsy using an RNA Prep Pure Tissue kit (Tiangen Biotech Co., Ltd., Beijing, China). The concentration and purity of isolated total RNA were determined using a BioTek Synergy H4 Hybrid Reader (BioTek Laboratory Instrument, Winooski, VT, USA). The mRNA integrity was evaluated by observing 18 S and 28 S ribosomal bands after agarose gel electrophoresis. Each tissue was performed in five replicates.
A total of 30 μg of RNA per cow was used as the input material for the RNA sample preparations. Sequencing libraries were generated using Solexa on the Illumina platform. The library construction procedure was conducted according to the method by Cui et al. (18). Oligo(dT)25 magnetic beads purified mRNA from total RNA. Then, divalent cations under 94°C denaturation in RNA fragmentation buffer fragmented the mRNAs. Random oligonucleotides were used to synthesize first-strand cDNA. RNase H and DNA polymerase I subsequently generated second-strand cDNA. After adenylation of the 3′ ends of DNA fragments, adapter oligonucleotides were ligated to prepare for gel purification recovery using a Gel Extraction Kit (Illumina, CA). The PCR primer selectively enriched DNA fragments with ligated adaptor molecules on both ends in a 12-cycle PCR reaction. Gel Extraction Kit QIA quick Gel and Mini Columm were used for gel purification and recovery and the extraction of DNA fragments of ~200 bp in length. The purified products were quantified using an Agilent high-sensitivity DNA assay on the Agilent 2100 Bioanalyzer system. Illumina sample clusters were prepared using Cluster Station according to the manufacturer's instructions. After cluster generation, an Illumina Hiseq 2000 platform sequenced the libraries.
A large amount of sample data was obtained using Solexa RNA paired-end sequencing. Given the impact of the Solexa data error rate on the results, quality preprocessing of the original data was performed. The sliding window method removed low-quality fragments: the quality threshold was 20 (error rate = 1%), the window size was 5 bp, and the length threshold was 35 bp. The length threshold of the sequence containing part N in the reads was 35 bp. After preprocessing, we obtained 121 million effective RNA-seq data. The data volume was 10 G, and the average length was 86.65 bp. The amount of sample sequencing data was ~5 G.
Effective reads of the three samples were combined for de novo splicing. The software Trinity, version trinityrnaseq_r2012-10-05, and the paired-end splicing method were used. Repeat the splicing sequence, and finally get 199,039 transcripts with a length >200 bp and a size of 216 Mb. Taking the longest transcript under each Loci (comp*_c*_) as Unigene (the software's Chrysalisclusters module), we obtained 148,900 unigenes, 110 Mb in size. The sample unigene was compared with the public data gene, and function annotation was conducted using the similarity of the gene. Annotations with a similarity >30% and e < 1e−5 were obtained, and all annotation details obtained were merged and compared with NR, SWISS-PROT, TrEMBL, Cdd, pfam, and eukaryotic complete genomes (COG) libraries. The unigene sample was compared with the public data gene, and function annotation was conducted using the similarity of genes, including NR, SWISS-PROT, TrEMBL, Cdd, pfam, COG classification, Gene Ontology (GO), and KEGG annotations. Annotation, differentially expressed genes (DEGs) analysis, and enrichment analysis were performed only for genes with a length ≥500 bp. GO, COG cluster analysis, and KEGG metabolic pathway analysis were conducted by comparing the predicted proteins in the mammary tissue of dairy cows. These analytical methods helped predict the effects of different roughage qualities on the functions of genes related to milk fat synthesis in the mammary glands of dairy cows from the overall transcriptome level.
qRT-PCR was performed on nine randomly selected differentially expressed milk fat synthesis-related genes using the total RNA used for RNA-seq to validate the repeatability and reproducibility of the gene expression data obtained using RNA sequencing in Holstein cows. RNA was converted into cDNA by using 2 μL of 100 ng total RNA, 2 μL of 5 × PrimeScript RT Master Mix, and 6 μL of DNase/RNase free water (TaKaRa, Tokyo, Japan) at 37°C for 15 min and then 85°C for 5 s. RT-qPCR was performed using 2 μL of diluted cDNA combined with 10 μL of 2 × SYBR premix Ex Taq II (TaKaRa, Tokyo, Japan), 0.4 μL each of 10 μM forward and reverse primers, and 7.2 μL DNase/RNase free water in 96 well microwell plates of the BIO-RAD IQ5 Multicolor RT-qPCR Detection System (USA). RT-qPCR reactions were performed 30 s with pre-denaturation at 95°C, followed by 40 cycles of 15 s denaturation at 95°C, 20 s annealing at 58°C and 30 s extension at 72°C. RT-qPCR analysis was performed using the 2−ΔΔCT method with β-actin as the reference gene. The primer sequences and annealing temperature are summarized in Table 3. The primers were designed according to Zhang et al. (17) and were synthesized by Shanghai Sangon Biological Engineering and Technology Service Co. Ltd. (Shanghai, China).
DMI and milk performance were analyzed using a two-sided Student's t-test with the SAS 9.0 software package (SAS Inst. Inc., Cary, NC, USA). A 95% confidence interval was used in all cases as the default for the hypothesis test. The assumptions of normality were validated using the univariate procedure, and the results were deemed significant when P < 0.05. Differential expression genes (DEGs) were identified using value of logarithmic transformed fold-change [log(FC)] ≥ 1 or < -1.
There were no differences in DMI between the dietary treatments (Table 4). The milk yield was lower in cows fed the CSB diet, and milk fat content was lower in cows fed the CSA diet than in cows fed the MF diet (P < 0.05). Milk protein and lactose contents were not affected after cows were fed different diets. SCC was not affected by diet and remained within the normal range. Compared with cows fed the MF diet, saturated FAs (SFAs) and short- and medium-chain FAs (SMFAs) in milk were lower (P < 0.05) in cows fed the CSA and CSB diet, whereas unsaturated FA (UFA) and long-chain FAs (LCFAs) in milk fat were higher (P < 0.05).
We sequenced cDNA libraries of three mammary gland samples from 15 Holstein cows with CSA, CSB, and MF diets. We acquired 49,969,014–54,513,152 paired-end reads of 86.65 bp in length per sample. As a result, the total read length was 10 gigabases (Gb) for the three samples. The sample unigene was compared with the public data gene, and function annotation was performed using gene similarity. The similarity with the NR database was 30.35%. Among the 148,900 unigenes obtained in NR annotations, 91.45% were the 10 species with the largest number of Unigene annotations, and 78% of unigenes matched the protein sequence of Bos taurus (Figure 1), which also verified the reliability of our sequencing data (17).
Clusters of orthologous groups for COG functional classification prediction of genes found that 18,828 unigenes were annotated into 25 COG classifications. Among them, the largest number of unigenes, about 6,518, were related to the function of signal transduction mechanisms, and 877 unigenes were related to lipid transport and metabolic functions (Figure 2). There were 469 gene expressions that significant differed in the fat transcript and metabolism between treatments (Table 5).
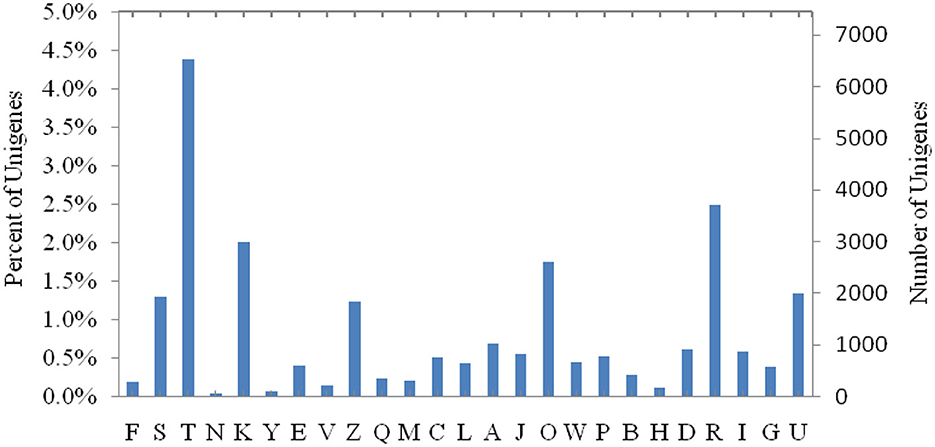
Figure 2. COG functional categories. [A] RNA processing and modification. [B] Chromatin structure and dynamics. [C] Energy production and conversion. [D] Cell cycle control, cell division, and chromosome partitioning. [E] Amino acid transport and metabolism. [F] Nucleotide transport and metabolism. [G] Carbohydrate transport and metabolism. [H] Coenzyme transport and metabolism. [I] Lipid transport and metabolism. [J] Translation, ribosomal structure, and biogenesis. [K] Transcription. [L] Replication, recombination, and repair. [M] Cell wall/membrane/envelope biogenesis. [N] Cell motility. [O] Posttranslational modification, protein turnover, and chaperones. [P] Inorganic ion transport and metabolism. [Q] Secondary metabolite biosynthesis, transport, and catabolism. [R] General function prediction only. [S] Function unknown. [T] Signal transduction mechanisms. [U] Intracellular trafficking, secretion, and vesicular transport. [V] Defense mechanisms. [W] Extracellular structures. [Y] Nuclear structure. [Z] Cytoskeleton.
The results of unigene based on blast-uniprot (i.e., combined with SWISS-PROT and TrEMBL results) were obtained, and the obtained uniprot number was used to compare the GO term. The GO function classification and prediction of unigene showed that 211,947 unigenes were classified as GO, of which 96,295 were classified as biological processes and 77,729 as cellular components. A total of 37,950 were classified as molecular functions. The functions of the sample genes clustered in the cellular and metabolic processes at 18,899 and 16,358, respectively, in the biological process classification. There were 680 differentially expressed functions in the biological process among the samples. The cellular component was concentrated in the cell and cell parts, and 149 differentially expressed proteins were found in the cellular component among the three samples. In the classification of molecular functions, 17,970 binding and 11,509 catalytic activities were concentrated in the catalytic activities. The samples had 350 differentially expressed molecular functions (Figure 3). The differentially expressed genes of the samples were primarily involved in biological processes, cell components, and molecular functions, and the term related to membrane composition was the most significantly enriched GO. Second, the differentially expressed genes were significantly correlated with biological functions integral to the membrane, intracellular membrane, transport, and plasma membrane.
KEGG pathway analysis of the genes revealed the metabolic pathways involved in Unigene in the mammary glands of dairy cows and the functions of genes. A total of 23,095 transcripts were annotated, with 1,091 enzymes involved in 302 metabolic pathways. The number of transcripts involved in the cancer pathway was the highest, with 862, followed by the MAPK signaling pathway, with 674, and Focal adhesion reached 663. The most significantly different genes among the groups were in the cancer pathway. Different dietary patterns significantly affected milk fat and FA composition. Therefore, we summarized the milk fat anabolic pathways. There were 935 transcripts obtained from all transcript libraries that participated in 16 lipid metabolism pathways (Table 6).
The expression abundance values of sample transcripts were used for differential expression analysis, including fold change analysis, Fisher test, and chisq test. The final results are based on fold changes, as shown in Table 7. Comparative analysis was performed based on the expression abundance values (RPKM) of the transcripts in different groups (Figure 4). The number of up-regulated genes were 45,399, and the number of down-regulated genes were 15,750 identified in the mammary tissue between CSA_VS_CSB; The number of up-regulated genes were 19,808, and the number of down-regulated genes were 40,673 identified in the mammary tissue between CSA_VS_MF; The number of up-regulated genes were 14,049, and the number of down-regulated genes were 74,111 identified in the mammary tissue between CSB_VS_MF. The order of the number of up-regulated expressed genes from low to high after all transcripts were compared between the three groups was CSB < CSA < MF.
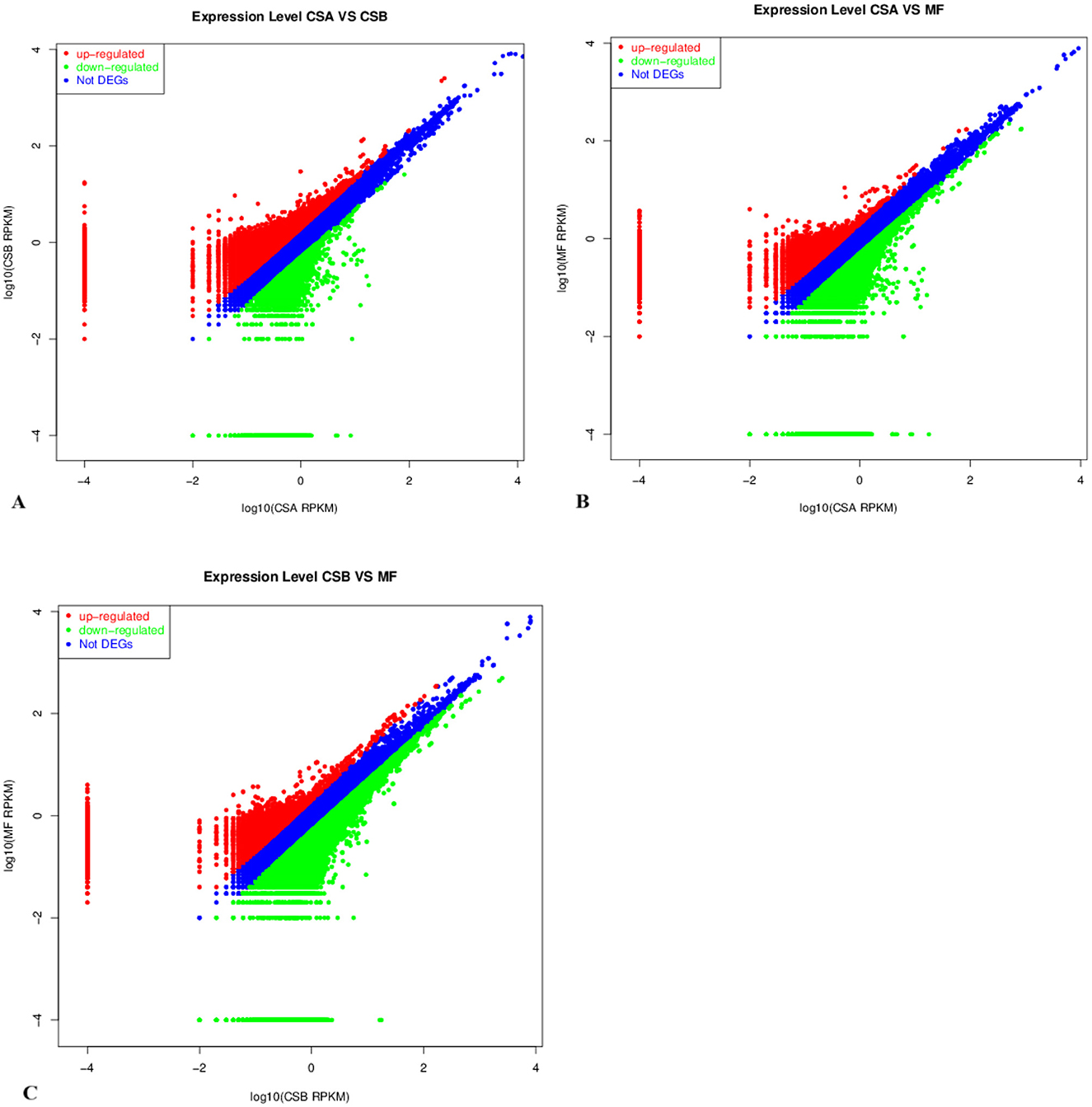
Figure 4. The DEGs in the mammary tissue of cows with different roughage diets. (A) DEGs identified in the mammary tissue between CSA_VS_CSB. (B) DEGs identified in the mammary tissue between CSA_VS_MF. (C) DEGs identified in the mammary tissue between CSB_VS_MF. The red and green dots represent the up-regulated and down-regulated gene (P ≤ 0.01), respectively; the blue dots represent the transcripts whose expression levels did not reach statistical significance (P > 0.01). MF, with hay, corn silage, and alfalfa mixed as forage; CSA, with only corn stalk as forage and with the same level of proximate nutrients to MF; CSB, with only corn stalk as forage and with the same proportion of forage to MF.
After comparing the three groups, the classification reaching the most differences in the annotation in GO analysis was membrane. The KEGG pathway analysis showed that the calcium signaling pathway was the most downregulated, and the cancer pathway was the most upregulated in CS2 than MF. Focal adhesion was the most downregulated, and the cancer pathway was the most upregulated in CSA. Compared with CSB, ABC transporters were the most downregulated pathway, focal adhesion was the most upregulated pathway, and cancer was the second pathway in CSA.
By KEGG analyzing the genes involved in FA synthesis in the mammary glands of dairy cows, 935 transcripts were found to be involved in 16 pathways of lipid metabolism, and 62 genes were found to be significant. DEGs identified in the mammary tissue involved in important lipid metabolism pathways between treatments are shown in Table 8. In the MF group, the differentially expressed genes of the ether lipid metabolism (ko00565), unsaturated FA biosynthesis pathway (ko01040), and FA elongation (ko00062) pathway were significantly lower than those in the CSB group (P < 0.05). These differentially expressed genes included Ethanolamine phosphotransferase (EPT)1, Lysophosphatidylcholine acyltransferase 1 (LPCAT1), peroxisomal FA β-oxidation Acyl-CoA Oxidase 1, 3(ACOX1, ACOX 3), Hydroxyacyl-CoA dehydrogenase trifunctional multienzyme complex subunit β(HADHB), Fatty Acid Desaturase 2(FADS2), Stearoyl-CoA Desaturase (SCD), and Elongation of Very Long-chain Fatty Acid-like Fatty Acid Elongase 4, 5, 6, 7 (ELOVL4, ELOVL5, ELOVL6, ELOVL7). At the same time, the differentially expressed genes of the FA biosynthesis pathway (ko00061) in the MF group were higher than those in the CSB group (P < 0.05). These differentially expressed genes included de novo synthesis Acetyl-CoA Carboxylase (ACAC), Fatty Acid Synthase (FASN), and Fatty-acid synthesis II dehydrogenases bacterial 3-ketoacyl-ACP reductase (fabG). A map of metabolic pathways involved in differentially expressed genes can be obtained by comparing the differentially expressed genes to the KEGG database. The pathway in which transcripts are most involved in glycerophospholipid metabolism and the most differentially significant genes are produced (Figure 5).
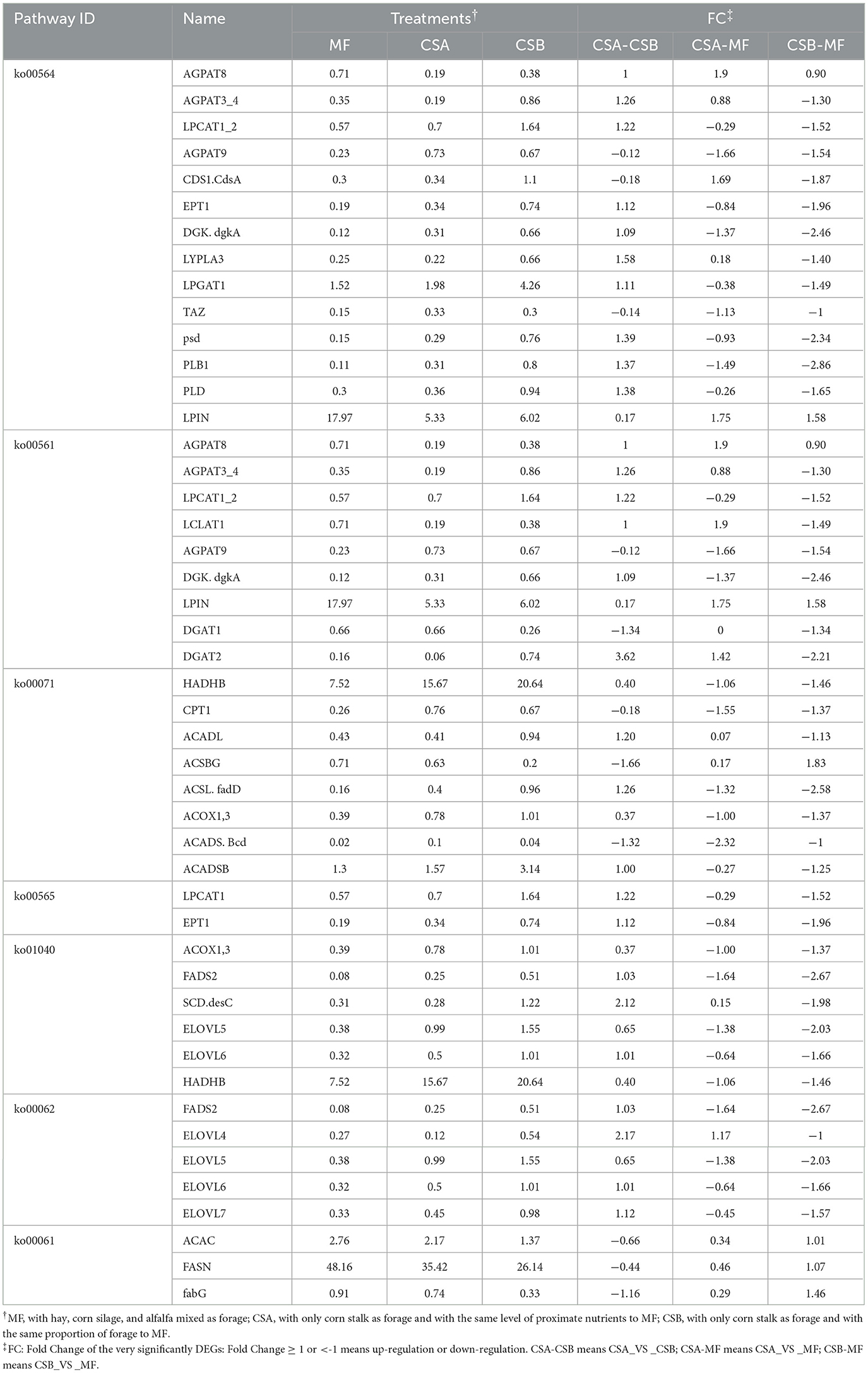
Table 8. Differential expressed genes (DEGs) identified in the mammary tissue in main lipid metabolism pathways between treatments.
The differential expression of nine selected differentially expressed genes, such as ACACA, SCD, FASN, LPL, PPARA, ACSL1, CD36, FABP3, and LPIN, was validated using RT-qPCR to provide additional evidence for the gene expression profiles recorded at the transcriptome level. The relative expression levels of the selected genes verified using RT-qPCR in a further cohort of samples agreed with the RNA-seq data, although this magnitude varied (Figure 6).
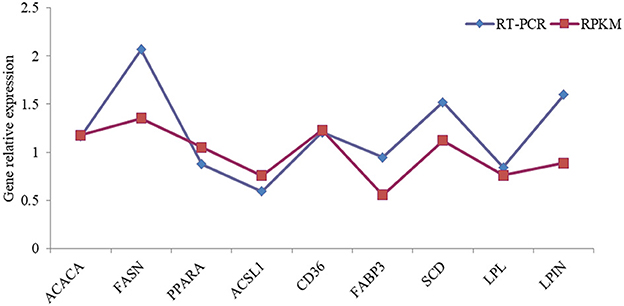
Figure 6. The comparison of high-throughput sequencing and RT-PCR relative expression for fatty acid synthesis-related genes in dairy cows. ACACA, Acetyl-CoA Carboxylase; FASN, Fatty Acid Synthase SCD, Stearoyl-CoA Desaturase; LPL, lipoprotein lipase; PPARA, peroxisome proliferators-activated receptors α; ACSL1, Acyl-CoA Synthetase Long Chain Family Member 1; CD36, luster of differentiation 36; FABP3, Fatty Acid Binding Protein 3; LPIN, Lipid phosphate phosphohydrolase.
Multiple mechanisms regulate the DMI of ruminants, but the change in dietary starch concentration was ineffective in the DMI of dairy cows (19, 20). Feeding similar NDF content to corn silage-based diets using alfalfa hay, wheat straw, and corn stover can produce similar animal performance. Still, forage with high NDF decreased the milk yield in dairy cows (21). In our study, the CSB diet with higher NDF and lower net energy decreased milk yield compared with the MF and CSA diets fed to dairy cows. As the main forage, the alfalfa hay produced a higher milk yield than the corn stover forage to feed cows. Still, DMI, milk protein, and lactose showed no difference between these treatments (7, 22). A high starch level diet was associated with “Milk fat depression” (8, 23). Still, different dietary starch levels did not affect DMI, milk protein, or lactose in the early lactation of dairy cows and goats (24, 25). Cabrita et al. (26) showed that diets with the same crude protein levels and different starch concentrations resulted in no change in the milk lactose of dairy cows. The milk lactose content could not be manipulated using a diet except under extreme and unusual feeding situations (27). In our study, a single corn stover forage diet (CSA and CSB) decreased milk yield, and a high concentration in the CSA reduced milk fat content of dairy cows was obtained compared to a mixed forage diet.
SMFAs in milk decreased in CSA and CSB, which agreed with Chilliard et al. (28) that cows fed a high dietary C18 FA could decrease medium-chain FA secretion and concentration through dilution with a large amount of LCFA. LCFAs in milk were obtained from arterial blood, and these LCFAs were derived from diets. Our results showed that cows fed with the CSA and CSB diets produced a high amount of UFAs in milk, supporting the hypothesis that cows fed with a high amount of UFA diet produced relatively high proportions of UFAs in milk (29).
The proliferation, differentiation, and survival of normal cells were regulated by interlinked pathways that transmit and integrate signals from growth factors, hormones, and cell–cell and cell–matrix interactions. These pathways can turn into cancer pathways by altering component proto-oncogenes and tumor suppressors. Focal adhesions are dynamic multi-protein complexes that participate in extracellular matrix adhesion and play an important role in translating extracellular matrix stiffness signals into intracellular chemical responses. Focal adhesion dysfunction is critical in tumor cell invasion and metastasis (30). Therefore, the focal adhesion pathway is closely related to the cancer pathway. The activation of MAPK signaling pathways is usually caused by external stimuli such as bacterial infections, inflammatory responses, and oxidative stress (31). Under normal circumstances, the activation of pathways helps regulate immune responses, inflammatory responses, and cellular protective mechanisms. The results indicated cows in CSB group fed diets with low nutrient levels were more involved in the cancer pathway and focal adhesion. In such cases, cows need to use more body mobilization to protect themselves against oxidative stress, inflammation and even more serious diseases. At the same time, the expression of related genes was significantly increased compared to the diet with high nutrient levels.
The ABC family, which is present in all domains of life, is widely acknowledged as one of the largest transporter families (32, 33). ABC transporters have garnered considerable interest due to their capacity to transport diverse substances across cellular membranes and actively facilitate their elimination (34, 35). ABC transport participates in the absorption, accumulation, and excretion of many toxic substances and plays an important role in the defense process of body tissues. The genes of the ABC transport pathway in the high-concentrate corn stover group were more upregulated than those in the low-concentrate corn stover group. This result suggests that a high-concentrate diet may lead to more toxic substances and more genes involved in the defense process. The number of differentially expressed genes in triglyceride metabolism in the MF group was higher than that in the CSA group. This result was consistent with previous results that milk fat content in the MF group was significantly higher than in the CSA group.
Milk fat is the major factor determining the organoleptic quality and the commercial price of milk. LPCAT1 is a key enzyme in the lipid remodeling pathway known as the Lands cycle (36). LPCAT1 may be regulated by the cholesterol biosynthesis pathway (37). In this study, EPT1 and LPCAT1 were higher in the CSB group, showing that milk fat content was lower than in the MF group. Elongation of very long-chain FA (ELOV) proteins is widely present in the genomes of animals, plants, and microorganisms, regulating the length of FA (38). Elovl4 plays a crucial role in the biosynthesis of the so-called “very long-chain (>C24) polyunsaturated fatty acids,” compounds that accumulate in vertebrate photoreceptors that can contain up to C44 (39). In teleosts, most species studied have a single Elovl5 gene that can elongate C18 and C20 PUFA substrates (40). Elovl6 participates in the elongation of SFA and MUFA up to C18, while Elovl7 elongates SFAs and MUFAs of 18–22 carbons (41).
The FADS2 gene encodes delta-6 desaturase (D6D) and is a member of the FA desaturase gene family. D6D is the key enzyme that catalyzes the transformation of linoleic acid and α-linolenic acid to long-chain polyunsaturated FA (42). SCD is the main enzyme involved in monounsaturated FA synthesis in ruminants. A diet with high C18:1c9 inhibited SCD mRNA expression in the mammary glands of mice (43). The SCD promoter activity was decreased by C18:1c9 in MAC T cells (44). These studies agreed with the current results regarding the changes in the milk FA composition and lipid metabolism pathways. Differentially expressed genes in the CSB diet with high UFA, LCFA, and C18:1c9 are reflected in the dietary FA profile. This may decrease SCD and FADS2 expression and increase ELOVL4, ELOVL5, ELOVL6, and ELOVL7 expressions in mammary glands compared with MF. FabG was identified as an essential gene involved in fatty acid biosynthesis (45). ACAC and FASN are key enzymes involved in the de novo synthesis of FA in mammary tissues (46). High de novo FA synthesis in mammary glands increases SMFA content and milk fat (47). The differentially expressed gene results might explain the high amounts of SMFA in the milk fat of cows fed the MF diet.
According to the results of differential gene expression levels of lipid metabolic pathways, the expression levels of genes in the FA biosynthesis pathway in the MF group were significantly higher than in the CSB group. The gene expression of lipid metabolism changed with different diets has been previously described by Zhang et al. (17). The result was consistent with the significantly high milk fat content in the MF and CSB groups. The differentially expressed genes in the FA extension pathway and unsaturated FA synthesis in the MF group were significantly lower than those in CSB, which was consistent with the result that the contents of LCFA and UFA in MF were significantly lower than those in CSB.
The RNA sequencing data demonstrated that cows fed a low nutritional diet (CSB) had more genes involved in the cancer pathway. Feeding a better nutrition diet (MF) to cows caused the cells to become more active. Cows fed a high-concentrate ratio diet (CSA) had more genes involved in the detoxification defense pathway. According to the analysis of lipid metabolic pathways using KEGG, the DEGs of the FA biosynthesis pathway such as ACAC, FASN and fabG in the MF group were significantly higher than that in the CS group. In contrast, the DEGs of the FA extension pathway and the unsaturated FA synthesis pathway such as ELOVL4, 5, 6, 7 and FADS2, SCD were significantly lower than those in the CS group, consistent with milk fat content and composition results in dairy cows.
The original contributions presented in the study are included in the article/Supplementary material, further inquiries can be directed to the corresponding author.
The animal study was approved by the Animal Care and Ethics Committee of Inner Mongolia Agricultural University. The study was conducted in accordance with the local legislation and institutional requirements.
HZ: Investigation, Writing – original draft. CA: Funding acquisition, Writing – review & editing. K-E: Data curation, Formal analysis, Writing – review & editing. ND: Investigation, Validation, Visualization, Writing – review & editing.
The author(s) declare that financial support was received for the research and/or publication of this article. This research was supported by grants from the National Key Basic Research Program of China (No. 2011CB100803) and the Inner Mongolia Natural Science Foundation (2023LHMS03060).
The authors acknowledge the Inner Mongolia Dairy Association Science and Technology Limited Company (Hohhot, China) for the animals supplied.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fvets.2025.1469169/full#supplementary-material
1. Chamekh L, Khorchani T, Dbara M, Hammadi M, Yahyaoui MH. Factors affecting milk yield and composition of tunisian camels (Camelus dromedarius) over complete lactation. Trop Anim Health Prod. (2020) 52:3187–94. doi: 10.1007/s11250-020-02344-0
2. Vargas-Bello-Pérez E, Faber I, Osorio JS, Stergiadis S. Consumer knowledge and perceptions of milk fat in Denmark, the United Kingdom, and the United States. J Dairy Sci. (2020) 103:4151–63. doi: 10.3168/jds.2019-17549
3. Moate PJ, Chalupa W, Boston RC, Lean IJ. Milk fatty acids II: prediction of the production of individual fatty acids in bovine milk. J Dairy Sci. (2008) 91:1175–88. doi: 10.3168/jds.2007-0226
4. Mach N, Goselink RMA, Van Baal J, Kruijt L, Van Vuuren AM, Smits MA. Relationship between milk fatty acid composition and the expression of lipogenic genes in the mammary gland of dairy cows. Livest Sci. (2013) 151:92–6. doi: 10.1016/j.livsci.2012.10.014
5. Pang YZ, Liu YP Li XJ, Wang KS, Yuan HR. Improving biodegradability and biogas production of corn stover through sodium hydroxide solid state pretreatment. Energy Fuels. (2008) 22:2761–6. doi: 10.1021/ef800001n
6. Zhang H, Ao CJ, Khas-Erdene LWS, Song LW, Zhang XF. Effects of different model diets on milk composition and expression of genes related to fatty acid synthesis in the mammary gland of lactating dairy goats. J Dairy Sci. (2015) 98:4619–28. doi: 10.3168/jds.2013-7097
7. Zhu W, Fu Y, Wang B, Wang C, Ye JA, Wu YM, et al. Effects of dietary forage sources on rumen microbial protein synthesis and milk performance in early lactating dairy cows. J Dairy Sci. (2013) 96:1727–34. doi: 10.3168/jds.2012-5756
8. Bernard L, Toral PG, Chilliard Y. Comparison of mammary lipid metabolism in dairy cows and goats fed diets supplemented with starch, plant oil, or fish oil. J Dairy Sci. (2017) 100:9338–51. doi: 10.3168/jds.2017-12789
9. Bernard L, Leroux C, Chilliard Y. Expression and nutritional regulation of lipogenic genes in the ruminant lactating mammary gland. Adv Exp Med Biol. (2008) 606:67–108. doi: 10.1007/978-0-387-74087-4_2
10. Kadegowda AKG, Bionaz M, Piperova LS, Erdman RA, Loor JJ. Peroxisome proliferator-activated receptor-gamma activation and long-chain fatty acids alter lipogenic gene networks in bovine mammary epithelial cells to various extents. J Dairy Sci. (2009) 92:4276–89. doi: 10.3168/jds.2008-1932
11. Panagiota K, Alexandros M, Christina M, Eleni T. Effect of whole sesame seeds dietary inclusion levels on transcriptional signatures of lipid metabolism in mammary gland of goats. Small Rumin Res. (2024) 236:107294. doi: 10.1016/j.smallrumres.2024.107294
12. Croucher NJ, Thomson NR. Studying bacterial transcriptomes using RNA-seq. Curr Opin Microbiol. (2010) 13:619–24. doi: 10.1016/j.mib.2010.09.009
13. Marioni JC, Mason CE, Mane SM, Stephens M, Gilad Y. RNA-seq: an assessment of technical reproducibility and comparison with gene expression arrays. Genome Res. (2008) 18:1509–17. doi: 10.1101/gr.079558.108
14. Cánovas A, Rincon G, Islas-Trejo A, Wickramasinghe S, Medrano JF. SNP discovery in the bovine milk transcriptome using RNA-Seq technology. Mamm Genome. (2010) 21:592–8. doi: 10.1007/s00335-010-9297-z
15. Wickramasinghe S, Hua S, Rincon G, Islas-Trejo A, German JB, Lebrilla CB, et al. Transcriptome profiling of bovine milk oligosaccharide metabolism genes using RNA-sequencing. PLoS ONE. (2011) 6:e18895. doi: 10.1371/journal.pone.0018895
16. Wickramasinghe S, Rincon G, Islas-Trejo A, Medrano JF. Transcriptional profiling of bovine milk using RNA sequencing. BMC Genomics. (2012) 13:45. doi: 10.1186/1471-2164-13-45
17. Zhang H, Ao C, Khas-Erdene N, Dan N. Effects of isonitrogenous and isocaloric total mixed ration composed of forages with different quality on milk fatty acid composition and gene expression of mammary lipogenic enzymes in mid-lactating dairy cows. Anim Sci J. (2019) 90:214–21. doi: 10.1111/asj.13154
18. Cui XG, Hou YL, Yang SH, Xie Y, Zhang SL, Zhang Y, et al. Transcriptional profiling of mammary gland in Holstein cows with extremely different milk protein and fat percentage using RNA sequencing. BMC Genomics. (2014) 15:226. doi: 10.1186/1471-2164-15-226
19. Dann HM, Tucker HA, Cotanch KW, Krawczel PD, Mooney CS, Grant RJ, et al. Evaluation of lower-starch diets for lactating Holstein dairy cows. J Dairy Sci. (2014) 97:7151–61. doi: 10.3168/jds.2014-8341
20. Fredin SM, Akins MS, Ferraretto LF, Shaver RD. Effects of corn-based diet starch content and neutral detergent fiber source on lactation performance, digestibility, and bacterial protein flow in dairy cows. J Dairy Sci. (2015) 98:554–65. doi: 10.3168/jds.2014-8592
21. Eastridge ML, Starkey RA, Gott PN, Oelker ER, Sewell AR, Mathew B, et al. Dairy cows fed equivalent concentrations of forage neutral detergent fiber from corn silage, alfalfa hay, wheat straw, and corn stover had similar milk yield and total tract digestibility. Anim Feed Sci Technol. (2017) 225:81–6. doi: 10.1016/j.anifeedsci.2017.01.013
22. Wang B, Mao SY, Yang HJ, Wu YM, Wang JK Li SL, et al. Effects of alfalfa and cereal straw as a forage source on nutrient digestibility and lactation performance in lactating dairy cows. J Dairy Sci. (2014) 97:7706–15. doi: 10.3168/jds.2014-7961
23. Ferraretto LF, Shaver RD, Bertics SJ. Effect of dietary supplementation with live-cell yeast at two dosages on lactation performance, ruminal fermentation, and total-tract nutrient digestibility in dairy cows. J Dairy Sci. (2012) 95:4017–28. doi: 10.3168/jds.2011-5190
24. Piccioli-Cappelli F, Loor JJ, Seal CJ, Minuti A, Trevisi E. Effect of dietary starch level and high rumen-undegradable protein on endocrine-metabolic status, milk yield, and milk composition in dairy cows during early and late lactation. J Dairy Sci. (2014) 97:7788–803. doi: 10.3168/jds.2014-8336
25. McCarthy MM, Yasui T, Ryan CM, Pelton SH, Mechor GD, Overton TR. Metabolism of early-lactation dairy cows as affected by dietary starch and monensin supplementation. J Dairy Sci. (2015) 98:3351–65. doi: 10.3168/jds.2014-8821
26. Cabrita AR, Bessa RJ, Alves SP, Dewhurst RJ, Fonseca AJ. Effects of dietary protein and starch on intake, milk production, and milk fatty acid profiles of dairy cows fed corn silage-based diets. J Dairy Sci. (2007) 90:1429–39. doi: 10.3168/jds.S0022-0302(07)71628-4
27. Jenkins TC, McGuire MA. Major advances in nutrition: impact on milk composition. J Dairy Sci. (2006) 89:1302–10. doi: 10.3168/jds.S0022-0302(06)72198-1
28. Chilliard Y, Ferlay A. Dietary lipids and forages interactions on cow and goat milk fatty acid composition and sensory properties. Reprod Nutr Dev. (2004) 44:467–92. doi: 10.1051/rnd:2004052
29. Zened A, Enjalbert F, Nicot MC, Troegeler-Meynadier A. Starch plus sunflower oil addition to the diet of dry dairy cows results in a trans-11 to trans-10 shift of biohydrogenation. J Dairy Sci. (2013) 96:451–9. doi: 10.3168/jds.2012-5690
30. Paluch EK, Aspalter IM, Sixt M. Focal adhesion-independent cell migration. Annu Rev Cell Dev Biol. (2016) 32:469–90. doi: 10.1146/annurev-cellbio-111315-125341
31. Liu XC, Xu R, Miao YT, Zang YN, Huang YX, Zheng YY, et al. Exploring the immunomodulatory mechanisms of Osmunda japonica thunb polysaccharides: activation of MAPK and NF-κB signaling pathways. Food Bioscience. (2025) 63:105695. doi: 10.1016/j.fbio.2024.105695
32. Gott RC, Kunkel GR, Zobel ES, Lovett BR, Hawthorne DJ. Implicating ABC transporters in insecticide resistance: research strategies and a decision framework. J Econ Entomol. (2017) 110:667–77. doi: 10.1093/jee/tox041
33. Thomas C, Tampé R. Structural and mechanistic principles of ABC transporters. Annu Rev Biochem. (2020) 89:605–36. doi: 10.1146/annurev-biochem-011520-105201
34. Jin M, Liao C, Chakrabarty S, Zheng W, Wu K, Xiao Y. Transcriptional response of ATP-binding cassette (ABC) transporters to insecticides in the cotton bollworm, Helicoverpa armigera. Pestic Biochem Physiol. (2019) 154:46–59. doi: 10.1016/j.pestbp.2018.12.007
35. Ju D, Dewer Y, Zhang SP, Hu C, Li PR, Yang XQ. Genome-wide identification, characterization, and expression profiling of ATP-binding cassette (ABC) transporter genes potentially associated with abamectin detoxification in Cydia pomonella. Ecotoxicol Environ Saf. (2022) 230:113152. doi: 10.1016/j.ecoenv.2021.113152
36. Wang B, Tontonoz P. Phospholipid remodeling in physiology and disease. Annu Rev Physiol. (2019) 81:165–88. doi: 10.1146/annurev-physiol-020518-114444
37. Tao MY, Luo J, Gu T, Yu XJ, Song Z, Jun YL, et al. LPCAT1 reprogramming cholesterol metabolism promotes the progression of esophageal squamous cell carcinoma. Cell Death Dis. (2021) 12:845. doi: 10.1038/s41419-021-04132-6
38. Guillou H, Zadravec D, Martin PGP, Jacobsson A. The key roles of elongases and desaturases in mammalian fatty acid metabolism: insights from transgenic mice. Prog Lipid Res. (2010) 49:186–99. doi: 10.1016/j.plipres.2009.12.002
39. Serrano R, Navarro JC, Portolés T, Sales C, Beltrán J, Monroig Ó, et al. Identification of new, very long-chain polyunsaturated fatty acids in fish by gas chromatography coupled to quadrupole/time-of-flight mass spectrometry with atmospheric pressure chemical ionization. Anal Bioanal Chem. (2021) 413:1039–46. doi: 10.1007/s00216-020-03062-0
40. Castro LFC, Tocher DR, Monroig O. Long-chain polyunsaturated fatty acid biosynthesis in chordates: insights into the evolution of fads and elovl gene repertoire. Prog Lipid Res. (2016) 62:25–40. doi: 10.1016/j.plipres.2016.01.001
41. Deák F, Anderson RE, Fessler JL, Sherry DM. Novel cellular functions of very long chain-fatty acids: insight from ELOVL4 mutations. Front Cell Neurosci. (2019) 13:428. doi: 10.3389/fncel.2019.00428
42. Wang GJ, Tian Y, Zhang HY. Research progress of fatty acid desaturase 2 gene in glycolipid metabolism. Zhongguo Yi Xue Ke Xue Yuan Xue Bao. (2022) 44:158−63.
43. Singh K, Hartley DG, McFadden TB, Mackenzie DDS. Dietary fat regulates mammary stearoyl coA desaturase expression and activity in lactating mice. J Dairy Res. (2004) 71:1–6. doi: 10.1017/S0022029903006502
44. Keating AF, Kennelly JJ, Zhao FQ. Characterization and regulation of the bovine stearoyl-CoA desaturase gene promoter. Biochem Biophys Res Commun. (2006) 344:233–40. doi: 10.1016/j.bbrc.2006.03.133
45. Metris A, Reuter M, Gaskin DJH, Baranyi J, van Vliet AHM. In vivo and in silico determination of essential genes of Campylobacter jejuni. BMC Genomics. (2011) 12:535. doi: 10.1186/1471-2164-12-535
46. Bionaz M, Loor JJ. ACSL1, AGPAT6, FABP3, LPIN1, and SLC27A6 are the most abundant isoforms in bovine mammary tissue and their expression is affected by stage of lactation. J Nutr. (2008) 138:1019–24. doi: 10.1093/jn/138.6.1019
Keywords: dairy cow, forage quality, lipid metabolic, milk fat, RNA sequencing
Citation: Zhang H, Ao C, Khas-Erdene and Dan N (2025) Transcriptional profiling of milk fat synthesis in mammary gland of dairy cows with different forage quality using RNA sequencing. Front. Vet. Sci. 12:1469169. doi: 10.3389/fvets.2025.1469169
Received: 23 July 2024; Accepted: 17 February 2025;
Published: 26 March 2025.
Edited by:
Juliana Petrini, Clinica do Leite Ltda, BrazilReviewed by:
Bianca Castiglioni, National Research Council (CNR), ItalyCopyright © 2025 Zhang, Ao, Khas-Erdene and Dan. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Changjin Ao, Y2hhbmdqaW5hb0BhbGl5dW4uY29t
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.