
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Vet. Sci. , 19 February 2025
Sec. Veterinary Epidemiology and Economics
Volume 12 - 2025 | https://doi.org/10.3389/fvets.2025.1435920
This article is part of the Research Topic Economic Impact of Infections on the Farm Industry View all 4 articles
 Hnin Wai Phyu1,2
Hnin Wai Phyu1,2 Kamonpan Charoenkul1,2
Kamonpan Charoenkul1,2 Chanakarn Nasamran1,2
Chanakarn Nasamran1,2 Kitikhun Udom1,2
Kitikhun Udom1,2 Eaint Min Phyu2
Eaint Min Phyu2 Yu Nandi Thaw2
Yu Nandi Thaw2 Han Win Soe2
Han Win Soe2 Supassama Chaiyawong1,2
Supassama Chaiyawong1,2 Thant Nyi Lin3
Thant Nyi Lin3 Min Thein Maw4
Min Thein Maw4 Alongkorn Amonsin1,2*†
Alongkorn Amonsin1,2*†Porcine parainfluenza virus 1 (PPIV-1) is a paramyxovirus causing respiratory infections in pigs and has been reported worldwide. In this study, we conducted a cross-sectional survey of PPIV-1 in pig farms in Thailand and Myanmar from January 2022 to December 2023. Nasal swab samples from pigs in Thailand (n = 1,042) and Myanmar (n = 449) were collected from clinically healthy pigs and pigs with respiratory signs. PPIV-1 detection was carried out using the L gene-specific RT-PCR assay. Our results showed that 3.65% (38/1042) and 7.57% (34/449) were positive for PPIV-1 in Thailand and Myanmar, respectively. The viruses (n = 15) were subjected to whole genome sequencing (n = 4) and F and HN gene sequencing (n = 11). Genetic and phylogenetic analyses showed that Thai PPIV-1 (n = 7) was grouped into PPIV-1 lineage II (American lineage) and closely related to American and Chinese strains. On the other hand, one Thai PPIV-1 strain (n = 1) and Myanmar PPIV-1 (n = 7) belonged to lineage I (European lineage) and was closely related to European, Hong Kong (China), and South Korean strains. Our findings suggest that PPIV-1s from both lineages (I and II) are circulating in pigs in Thailand, and PPIV-1 of lineage I is circulating in pigs in Myanmar, suggesting high genetic diversity of PPIV-1 in the Southeast Asia region. This study is the first to report whole-genome sequences of PPIV-1 from pigs in Thailand and Myanmar. Our result provided insights and information about the current disease status and genetic diversity of PPIV-1 in pig farms, which will benefit further animal disease surveillance, prevention, and control.
Porcine parainfluenza virus 1 (PPIV-1), also known as porcine respirovirus 1 (PRV1), is a paramyxovirus that causes respiratory infections in pigs, first isolated in 2013. The PPIV-1 is an enveloped, negative-sense, single-stranded, non-segmented RNA virus within the family Paramyxoviridae, the genus Respirovirus. The genome size of PPIV-1 is around 15 kb. The viral genome consists of six open reading frames (ORFs) that encode the structural proteins, including nucleocapsid (N), phosphoprotein (P), matrix (M), fusion (F), haemagglutinin-neuraminidase (HN) and large (L) proteins (3’-N-P-M-F-HN-L-5′) (1, 2). Two significant surface glycoproteins, F and HN, are crucial in viral fusion between the viral envelope and cell membrane. They also play a role in viral attachment, entry into the host cells, and release from the cells (3, 4). F and HN glycoproteins are used for molecular epidemiological study of PPIV-1 (2). Based on F gene analysis, PPIV-1 can be classified into two distinct genetic lineages: lineage I (European lineage) and lineage II (American lineage) (5). The HN gene analysis also provided phylogenetic clustering similar to the results observed by the F gene and complete genome analysis (6).
PPIV-1 was first reported in nasopharyngeal and rectal swabs collected from slaughtered pigs in Hong Kong, China, 2009 and was first isolated in 2013. The genome of PPIV-1 was found to be closely related to human parainfluenza virus 1 (HPIV-1) and Sendai virus (SeV), which suggests that the virus has zoonotic potential (1). However, this speculation about the zoonotic potential of PPIV-1 was reported without experimental evidence of cross-species transmission of human infection. PPIV-1 has been distributed widely in pig farms across many countries, including Brazil, Chile, Germany, Netherlands, Italy, Poland, South Korea, and the USA (6–13). PPIV-1 has been isolated in pigs without or with respiratory symptoms such as coughing, sneezing, and nasal discharge (1, 10). The potential impact of PPIV-1 infection on pig farms and the swine industry is significant, as the virus causes a high morbidity rate, especially in nursery pigs. This results in economic losses due to decreased growth in younger pigs or increased mortality. PPIV-1 not only leads to economic losses in pig production but also has an important as zoonotic potential virus (1, 10, 14). PPIV-1 can co-infect with Swine Influenza Virus (SIV) and Porcine Reproductive and Respiratory Syndrome Virus (PRRSV), suggesting that the virus may be associated with a complex respiratory disease in pigs from multiple etiology, namely the porcine respiratory disease complex (PRDC) (6, 13). Although PPIV-1 has been reported in America, Europe, and some Asian countries, this virus has never been reported in pig farms in Thailand and Myanmar. Since there is limited information available regarding the epidemiology and current disease status of PPIV-1 in pig farms in Thailand and Myanmar, this study aimed to conduct a survey and genetically characterize PPIV-1 isolated in pig farms from Thailand and Myanmar.
In this study, a cross-sectional sample collection was carried out in 26 pig farms in ten provinces of Thailand, including Chainat, Chiangmai, Chonburi, Lopburi, Nakhon Pathom, Prachinburi, Phetchaburi, Ratchaburi, Saraburi, and Suphanburi from January 2022 to December 2023. In Myanmar, a cross-sectional sample collection was conducted in 19 pig farms located in two regions, Yangon and Nay Pyi Taw (Figure 1). The pig farms for sample collection were selected based on the criteria: pig farms being situated in high-density pig production areas and collaboration of pig farm owners. Approximately 10–30 samples were collected from different age groups of pigs per farm on each visit. In total, 1,491 nasal swab samples (NS) were collected from pigs of different ages and pigs with or without respiratory clinical symptoms. The nasal swab sample collected was conducted following the OFFLU guidelines (15). In brief, the swab was inserted into one nostril approximately 2–4 cm deep (depending on the age of the pigs) and gently swabbed using a circular motion for at least 5–10 s. Then, repeat the same procedure in the other nostril with the same swab. Age groups of pigs included the suckling group (≤ 4 weeks), nursery group (> 4–8 weeks), fattening group (> 8–20 weeks), and breeder (Gilt, Sow, Boar). Out of 1,491 samples, 1,042 NS were from Thailand, and 449 NS were from Myanmar. The nasal swab samples were placed in 2 mL viral transport media (MEM; Eagle Minimum Essential Medium, Hyclone™). In 1 liter of MEM, the powder is mixed with distilled water, with 1% Bovine serum albumin (BSA), 250 mg Gentamycin, 10 mL of L-glutamine, and 30 mL of NaHCO3. The collected samples were transported at 4°C to the laboratory within 24 h and kept in the freezer until tested, but no later than 4 weeks. The data relating to the collection date, age, sex, breed, and clinical status of the animals were recorded and used to select samples for the viral characterization step. For example, the PPIV-1 positive samples were selected first based on the location of the pig farms, followed by the age of the pigs, and their collection date. The study was carried out under the approval of the Chulalongkorn University Animal Cares and Uses Protocol (CU-VET IACUC 2331099), following the guidelines and regulations of CU-VET IACUC for all animal procedures. The study in Myanmar was conducted with permission from the Livestock Breeding and Veterinary Department (LBVD), Yangon and Nay Pyi Taw, Myanmar.
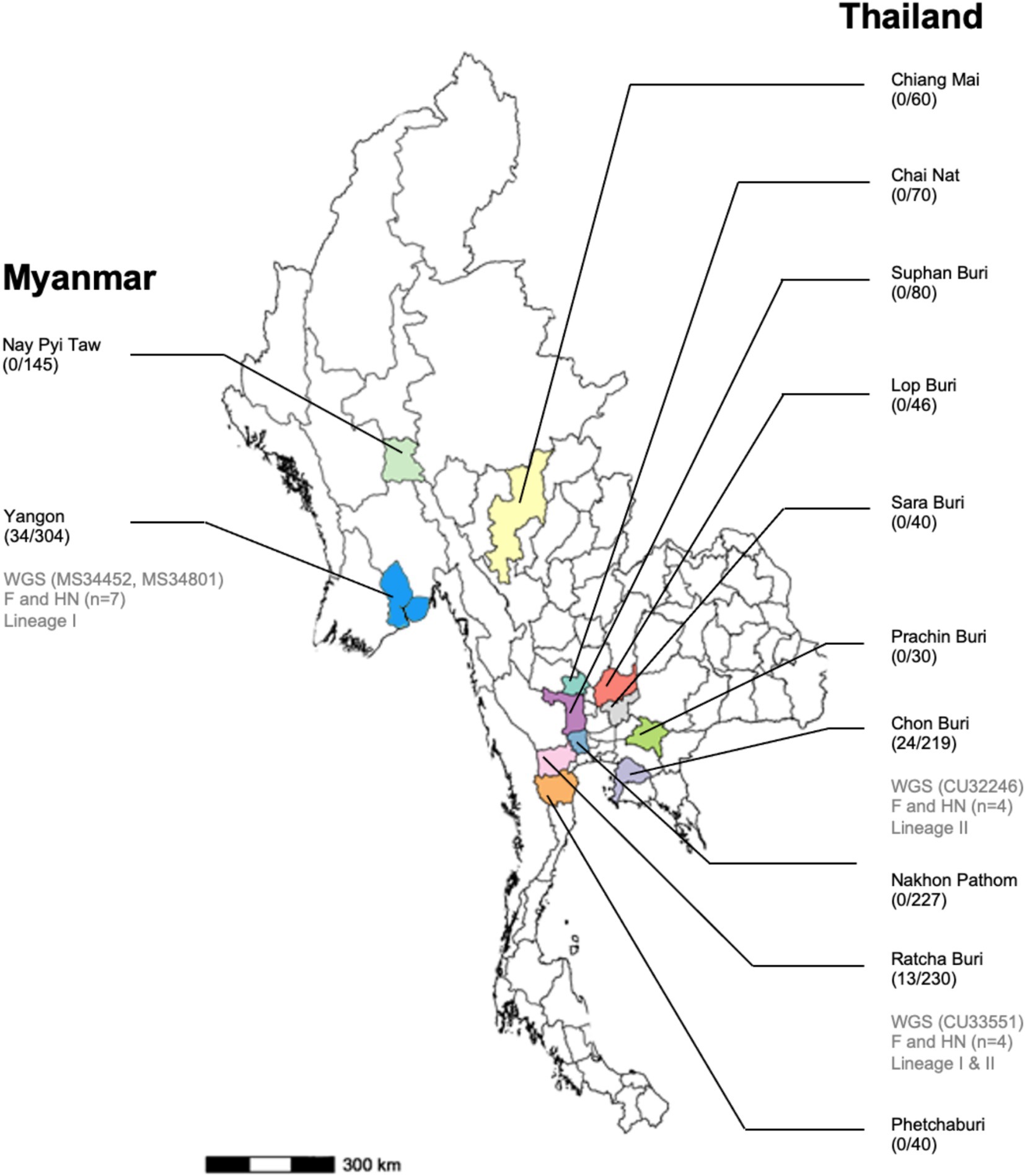
Figure 1. Map of Myanmar and Thailand shows the provinces/states where the samples were collected, the number of PPIV-positive/nasal samples tested, and the number of PPIV-characterized in this study.
RNA was extracted by aliquoting 200 μL of nasal swab samples and subjected to nucleic acid extraction using the Gene All® GENTi™ Viral DNA/RNA Extraction Kit (Gene All®; Lisbon, Portugal) on a GENTi™32 (GeneAll®) following the manufacturer’s recommendations. The RNA samples were tested for PPIV-1 using an RT-PCR assay specific to the L gene using a Super-Script III Platinum Taq One-step RT-PCR kit (Invitrogen, CA, United States). The L gene-specific primers (5’-TTTTGGGTTCAAGAGATTCTT-3′ and 5′- ACCTTTTGGTCTATG-TATAAAGA-3′), were used to detect PPIV-1, as described previously (1). In detail, the One-step RT-PCR reaction mixture included 3 μL of RNA, 12.5 μL of 2X SSIII buffer, 0.5 μM of each forward and reverse primer, and 1 U of Superscript III RT-Platinum Taq Polymerase and nuclease-free water to a final volume of 25 μL. The thermal cycling conditions contained at 55°C for 30 min for the cDNA synthesis step, at 94°C for 2 min for the initial denaturation step, and 35 cycles of denaturation at 94°C for 30 s, annealing at 47°C for 30 s and extension at 68°C for 1 min and final extension step at 68°C for 5 min. Positive samples for PPIV-1 were anticipated to yield a 154 bp PCR product when analyzed using 1.5% agarose gel electrophoresis. In this study, swine influenza virus (SIV) detection was also carried out in all nasal swab samples using real-time RT-PCR to investigate the co-infection of the virus in pigs (16).
In this study, the PPIV-1s were selected for whole genome sequencing (n = 4) and F and HN gene sequencing (n = 11). The PPIV-1 positive samples were selected based on the location of pig farms, the age of the pigs, their collection date, and the high yield of viral RNA by NanaDrop measurement. The samples were selected so that there was no overlap in location, age, and collection date. The samples were also selected based on the gel electrophoresis result, which showed a high yield of PCR products. Whole genome sequencing was performed by amplification of each gene of the viruses with newly designed primer sets using the Primer 3plus program and primers sets reported previously (Supplementary Table S1). In detail, the One-Step RT-PCR reactions were set up separately for each primer pair. The reaction mixture included 3 μL of RNA, 0.5 μL of each forward (10 μM) and reverse primer (10 μM), 12.5 μL of 2X Reaction Mix, 1 μL of SuperScript™ III RT/Platinum™ Taq Mix (cat no. 12574026) (InvitrogenTM; California, United States), and nuclease-free water to a final volume of 25 μL for each reaction. The amplified PCR products were pooled by mixing 20 uL per PCR reaction of one sample. The pooled PCR product (480 μL) was repeatedly loaded onto a single column to purify by using NucleoSpin® Gel and PCR Cleanup (MACHEREY-NAGEL, Düren, Germany) following the manufacturer’s instructions. The library was prepared using the Oxford Nanopore rapid sequencing kit V14 (SQK-RAD114) in accordance with the rapid sequencing protocol (Oxford Nanopore Technologies). A library was prepared per one round. In detail, 5 μL of the purified PCR product was mixed with 0.5 μL of fragmentation mix (FRA), incubated at 30°C for 2 min, then at 80°C for 2 min, and kept on ice. Then, 0.5 μL of diluted rapid adapter (RAP) was added and incubated at 25°C for 5 min. After that, 15 μL of sequencing buffer (SQB) and 10 μL of loading beads (LB) were added to complete the loading DNA library. The final library (at least 20 ng/μl), measured using a NanoDrop spectrophotometer (Thermo Fisher Scientific), was loaded into the flow cell (FLO-FLG114) R10.4.1 on the MinION Mk1b device and sequenced under MinKNOW software (v24.11.8) for 12 h (Oxford Nanopore Technologies, Oxford, United Kingdom). After sequencing, the qualified reads were converted from “Fast 5” into “Fastq” format by using the GPU version of the Nanopore Guppy basecaller (v6.5.7) tool. After basecalling, the EPI2ME platform (v5.2.3) was used to identify potential references for the sequencing data using wf-alignment workflow setting reference as all parainfluenza virus references in the NCBI database. The reads were aligned to references using Minimap2 (V2.28). The resulting alignments were then polished, and the consensus sequence was refined using Medaka (version 2.0.0). The nucleotide sequences were mapped to the reference (PPIV-1/S119N/Hong Kong/2009).
Phylogenetic and genetic analyses were performed by comparing the nucleotide sequences of Thai PPIV-1 and Myanmar PPIV-1 with those of reference viruses under the genus Respirovirus available from the GenBank database, including bovine parainfluenza virus 3, caprine parainfluenza virus 3, human parainfluenza virus 1, human parainfluenza virus 3 and murine respirovirus 1 (Sendai virus). The reference strains of PPIV-1s from different geographical regions were also included. Phylogenetic trees of the whole genome, F, and HN genes of PPIV-1s were performed using MEGA v.10.0 with a maximum likelihood method with the general time reversible gamma model with 1,000 bootstrap replicates. For genetic analysis, the nucleotide sequences and deduced amino acids of PPIV-1s were aligned and compared with those of reference viruses using MegAlign, DNASTAR software v.5.03 (DNASTAR Inc., Wisconsin, United States). An analysis of the multiple sequence alignment (MSA) of nucleotide and amino acids of Thai and Myanmar PPIV-1s and those of reference PPIV-1 strains was performed.
In this study, we conducted a cross-sectional survey in pig farms located in ten provinces of Thailand (n = 26 farms) and two regions in Myanmar (n = 19 farms) from January 2022 to December 2023. A total of 1,491 nasal swab samples (1,042 NS in Thailand and 449 NS in Myanmar) were collected and tested for PPIV-1 using L gene-specific RT-PCR. Our results showed that 3.65% (38/1042) and 7.57% (34/449) were PPIV-1 positive in Thailand and Myanmar, respectively (Table 1). In detail, PPIV-1 could be detected in 6 out of 26 (23.07%) tested farms in Thailand and 5 out of 19 (26.32%) tested farms in Myanmar. By location, the PPIV-1 positive samples were highly detected in Chonburi, Thailand (24/219; 10.95%) and Yangon, Myanmar (34/304; 11.18%). PPIV-1s were found in healthy pigs at 4.47% (61/1366) and in symptomatic pigs at 8.80% (11/125). In Thailand, PPIV-1 positive proportions were 3.68% (35/952) in healthy pigs and 3.33% (3/90) in symptomatic pigs, while in Myanmar, PPIV-1 was detected in pigs with respiratory signs at 22.86% (Supplementary Table S2). By age group, PPIV-1 was mainly found in nursery pigs (5–8 weeks) at 8.24% (62/752). By season, the positivity of PPIV-1 was highest in the summer season (28/295; 9.49%), which was comparable to the other two seasons in this study (Supplementary Table S2). In addition, the co-infection with SIV was observed in 10 PPIV-1-positive samples (10/72, 13.89%) (Table 1). It is noted that SIV positivity in nasal swab samples in Thailand and Myanmar was 15.26% (159/1042) and 1.56% (7/449), respectively.
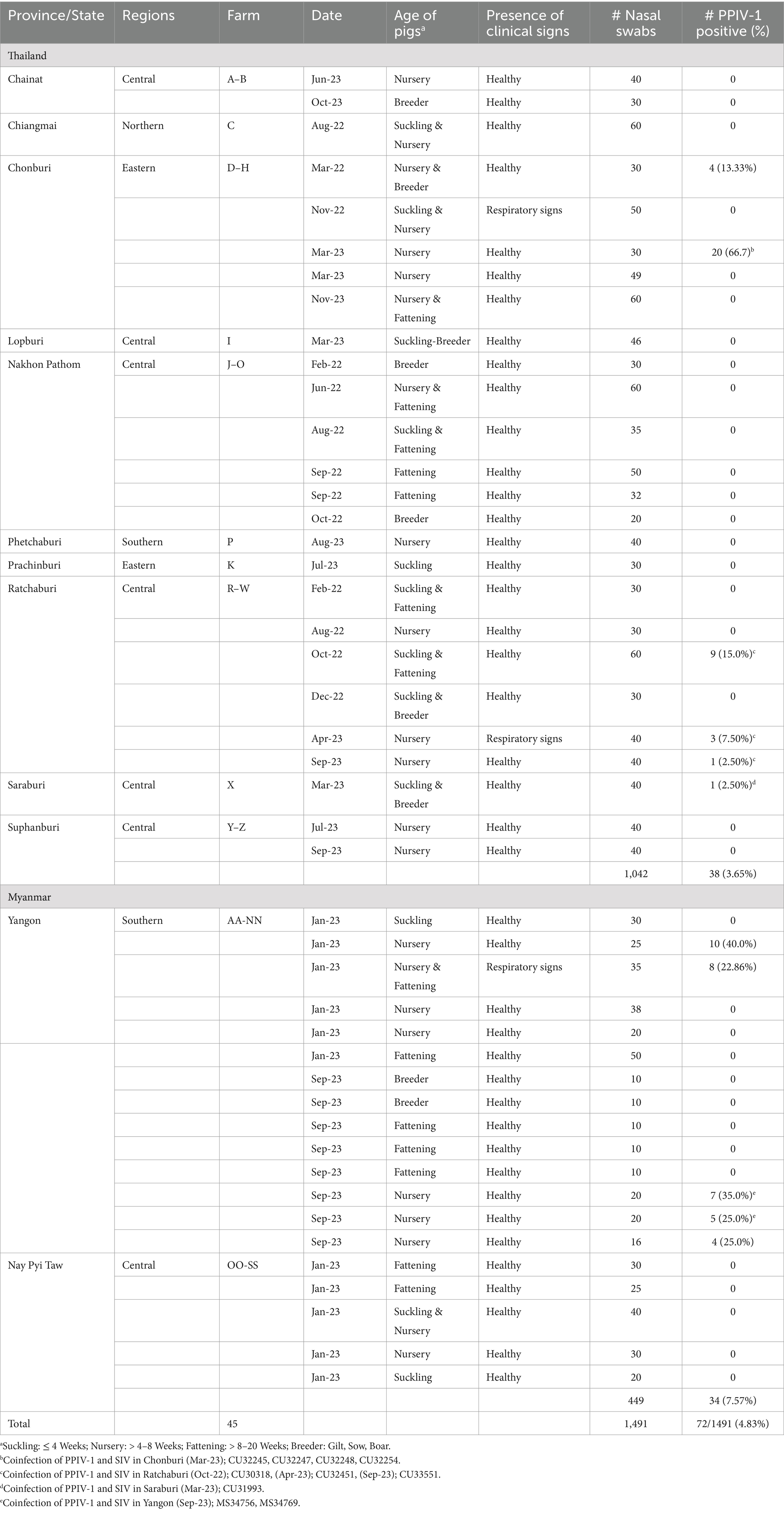
Table 1. Description of nasal swab samples collected from pig farms in Thailand and Myanmar in this study.
In this study, 15 out of 72 PPIV-1 positive samples were subjected to complete genome sequencing (n = 4) and F and HN gene sequencing (n = 11). This study successfully sequenced the whole genome of PPIV-1s (n = 4) with completed nucleotide sequences of all six major genes. The completed F and HN genes of PPIV-1 of PPIV-1 (n = 11) were also successfully sequenced. The nucleotide sequences of the viruses were deposited into the GenBank Database under the accession numbers (PP766830-833, and PP761466-487) (Table 2). Our results showed that the genome sizes of two Thai PPIV-1s are 15,194 bp – 15,312 bp, and those of Myanmar PPIV-1s are 15,384 bp −15,388 bp. The genome structure of the virus contained six major proteins encoding nucleocapsid (N), phosphoprotein (P), matrix (M), fusion (F), hemagglutinin-neuraminidase (HN), and large proteins (L) (3’-N/P/M/F/HN/L-5′). Notably, the lengths of all six major genes of Thai and Myanmar PPIV-1s are identical. However, due to the availability of the 3′ and 5′ nucleotide sequences, the overall genome length differs.
Phylogenetic analysis of the complete genome produced results similar to the F gene analysis, with Thai PPIV-1 being grouped into lineage II (American lineage) and showing close relations to American and Chinese strains. Conversely, Myanmar PPIV-1 was classified into lineage I (European lineage) and was closely related to European, Hong Kong (Chinese), and South Korean strains (Figure 2). It should be noted that both PPIV-1 lineages I and II have been reported in China. Based on the phylogenetic analysis of F genes, PPIV-1 can be classified into two genetic lineages: lineage I (European lineage) and lineage II (American lineage) (5). Our result revealed that 7 out of 7 out of 8 F gene sequences from Thai-PPIV-1 belong to Lineage II, while one sequence (CU32451), belongs to PPIV-1 Lineage I. This finding suggested that PPIV-1 of both lineages (I and II) are circulating in swine in Thailand. On the other hand, Myanmar PPIV-1 (n = 7) belonged to lineage I (European lineage) (Figure 3). For the HN gene, the phylogenetic analysis showed that Thai PPIV-1 clustered with lineage II (American lineage) (n = 7) and lineage I (European) (n = 1), whereas Myanmar PPIV-1 (n = 7) grouped with lineage I (European lineage) (Figure 4). The phylogenetic analyses of the N, P, M, and L genes indicated that Thai PPIV-1s also clustered with lineage II (American lineage), while Myanmar PPIV-1 was classified into lineage I (European lineage) (Figures 2, 3; Supplementary Figures S1–S4).

Figure 2. Phylogenetic tree based on whole genome sequences (WGS) of PPIV-1 in this study and reference strains. Blue circles represent Thai PPIV-1, and red triangles represent Myanmar PPIV-1 strains.

Figure 3. Phylogenetic tree based on F gene of PPIV-1 in this study and reference strains. Blue circles represent Thai PPIV-1, and red triangles represent Myanmar PPIV-1 strains.
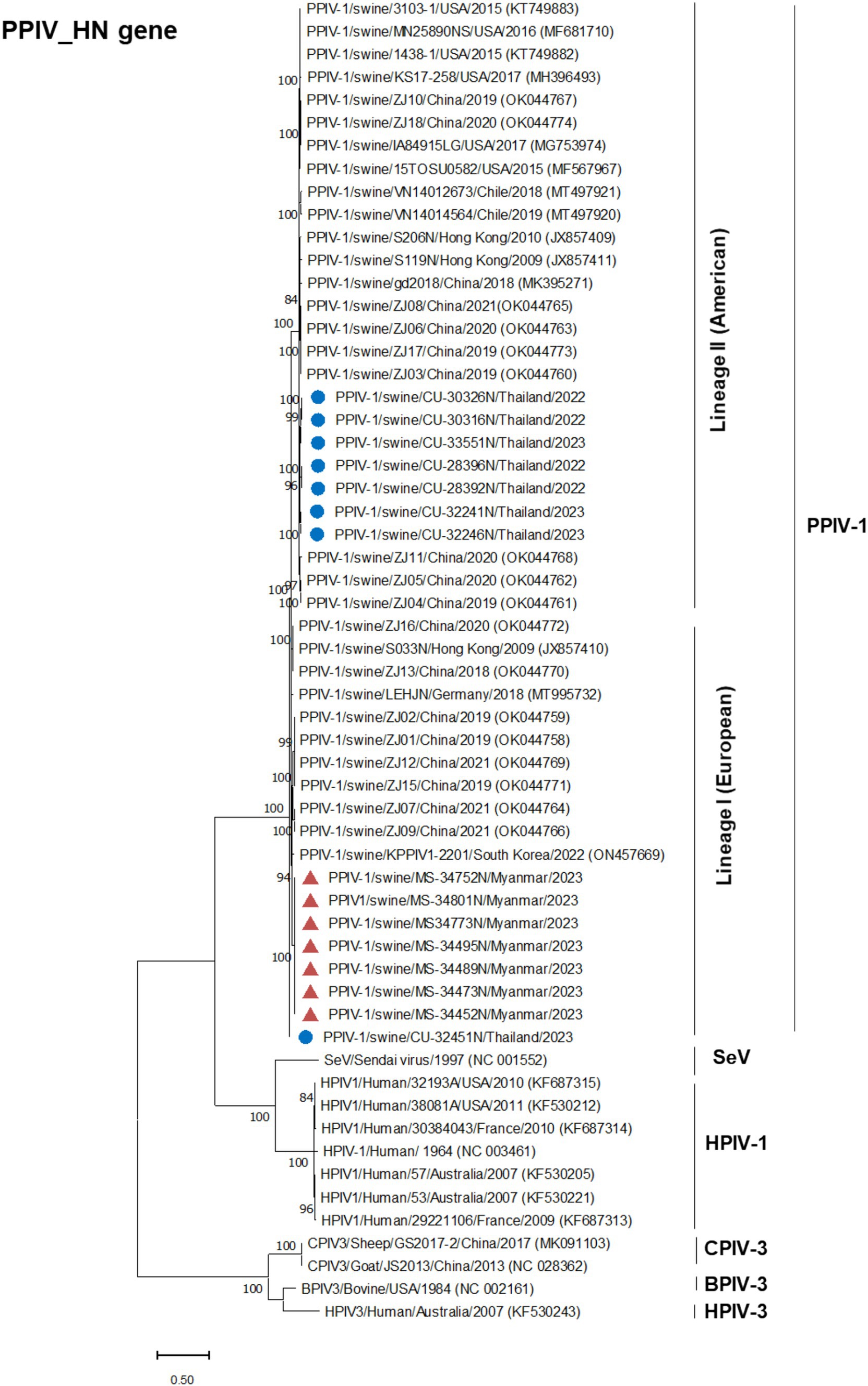
Figure 4. Phylogenetic tree based on HN gene of PPIV-1 in this study and reference strains. Blue circles represent Thai PPIV-1, and red triangles represent Myanmar PPIV-1 strains.
An analysis of the multiple sequence alignment (MSA) of the complete genome of PPIV-1 demonstrated that Thai PPIV-1s exhibited high nucleotide identities with reference PPIV-1s from China (ZJ10, ZJ11), the USA (IA8915), and Chile (VN14012673), showing identities of 95.46–96.91%, with highest % nucleotide identities to China PPIV-1 (ZJ11/2020, 96.91%) (Table 3). For F gene nucleotide comparison, Thai PPIV-1s possessed high nucleotide identity to Hong Kong PPIV-1s (S119N and S206N; 96.50–96.62%), China PPIV-1s (ZJ10 and ZJ 11; 95.99–96.31%) and the USA PPIV-1s (IA84915LG and VN14012673; 95.73–95.40%), respectively. When comparing Thai PPIV-1s with Myanmar PPIV-1s, Hong Kong, Germany, and South Korea strains, the nucleotide identities of the F gene showed 90.03–90.90% (Supplementary Table S3). The nucleotide identities of the HN gene of Thai PPIV-1s showed 94.98–95.74% with Chile, China, Hong Kong (China), and the USA strains (Supplementary Table S4).
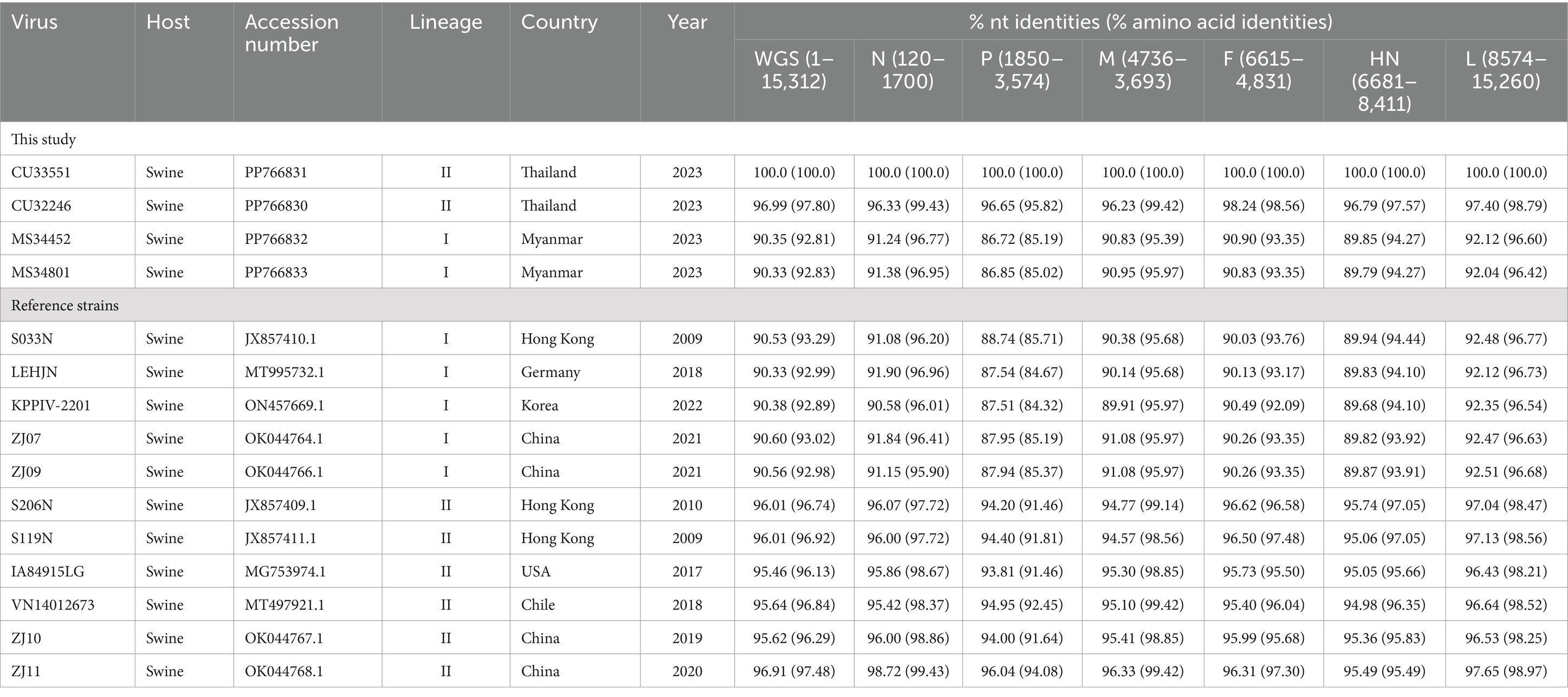
Table 3. Nucleotide (nt) and amino acid (aa) identities of the complete genome of Thai and Myanmar PPIV-1 with reference PPIV-1 strains.
Genetic analysis of the F gene showed that the F gene of PPIV-1s contains 1,674 nucleotides (558 amino acids). The twelve unique amino acid positions were identified, which could be used to classify the lineage or genotype-specific determinants. In particular, six strains of Thai PPIV-1s (lineage II, American lineage) had amino acid substitution, Proline to Serine at position 8 (P8S) of the F protein. Meanwhile, the Thai PPIV-1 strain (CU32451N) contained Leucine (L), which is similar to that of lineage I (European lineage) (Table 4).
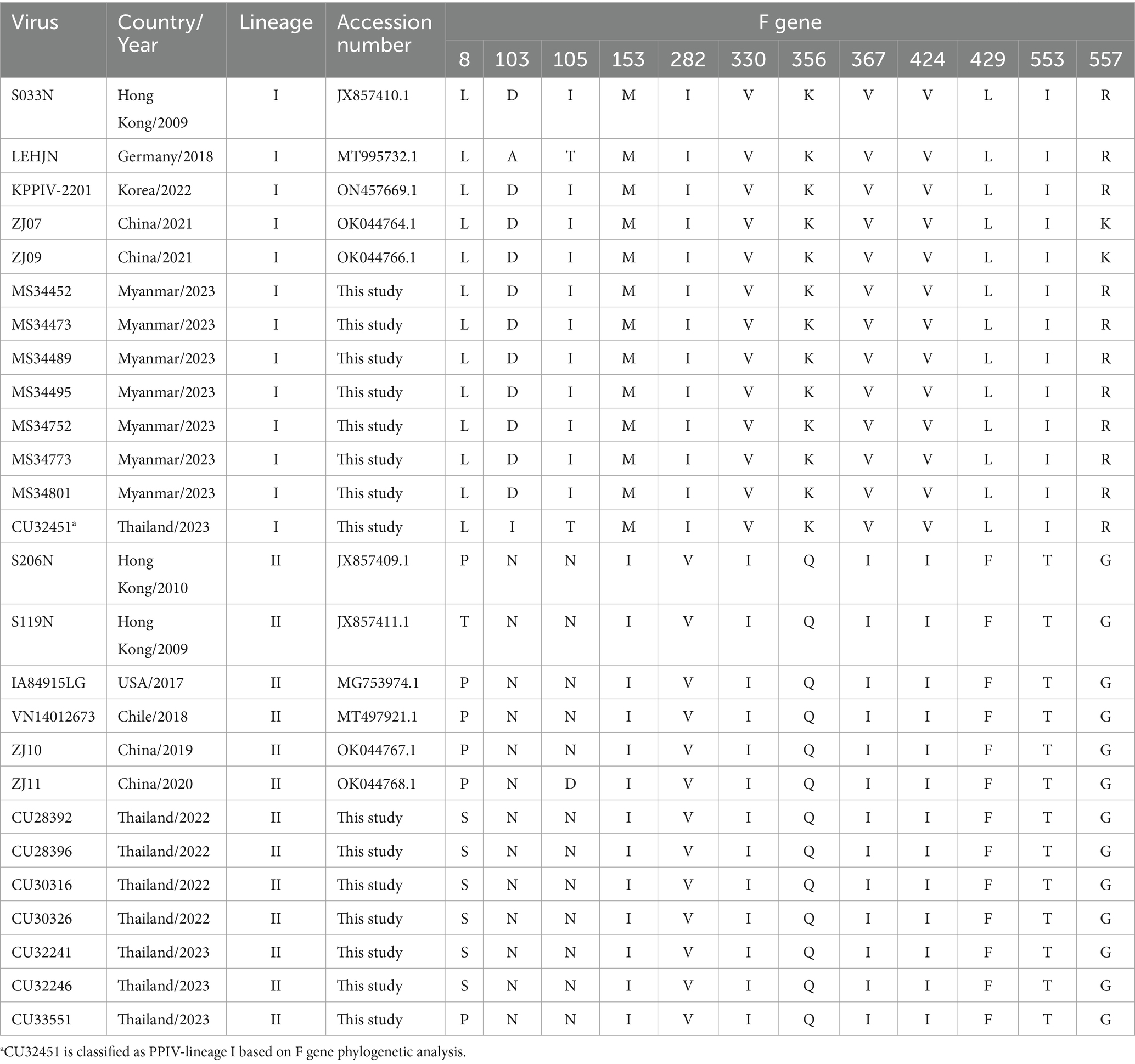
Table 4. Genetic analysis of the F gene among Thai and Myanmar PPIV-1s from this study and reference PPIV-1 strains.
This study is the first to report and genetically analyze Porcine Parainfluenza Virus 1 (PPIV-1) in pig farms in Thailand and Myanmar. Our study showed that the positivity of PPIV-1s was 4.83% (72/1491), with 3.65% (38/1042) in Thailand and 7.57% (34/449) in Myanmar, respectively. In this study, PPIV-1s were detected in healthy pigs and pigs with clinical signs at 4.47% (61/1366) and 8.80% (11/125), respectively. Our findings are comparable to those of the previous study in Hong Kong (China) (1). However, a higher positive proportion of PPIV-1 has been reported in Germany, Netherlands, Poland, South Korea, and the USA (6, 10, 11, 13). Similarly, in Myanmar, PPIV-1 was detected in pigs with respiratory signs at 22.86% (Supplementary Table S2). By age groups (suckling, nursery, fattening, and breeder), our findings showed that PPIV-1 was mostly detected in younger pigs (nursery and suckling pigs).
In the USA, a study reported a high PPIV-1 positive (43.3%) in samples submitted to a veterinary diagnostic laboratory (2). In Chile, samples from a swine influenza surveillance program of pigs 3–11 weeks of age with clinical symptoms were tested, and PPIV-1 was found to be 18.9% (7). In Europe, PPIV-1 was detected in respiratory samples of pigs in Germany and Netherlands (31.4%) (6), and Poland (76.7%) (13). In Italy, a study showed that 1.55% of pig samples tested were positive for PPIV-1 (12). In Asia, a survey in South Korea, PPIV-1 was detected in oral fluid samples (71.4%), suggesting PPIV-1 was widely distributed in Korean pig farms (11). In China, a study of PPIV-1 detection showed that 64.8% (201/310) of nasal swab samples were PPIV-1 positive (17).
In this study, the discrepancy in the occurrence of PPIV-1 compared to previous reports may be due to differences in the sampling strategy, type of samples, the clinical status of the animals, and the detection method used to detect the viruses. Our results showed that PPIV-1 was most commonly found in nursery pigs, consistent with previous studies (2, 11). PPIV-1 positivity was higher in nursery pigs than in suckling pigs and breeders. We also found that PPIV-1 can co-infect with SIV. Co-infection of PPIV-1 with SIV was observed in 13.89% of PPIV-1 positive samples (10/72). Similarly, the studies in Chile, Germany, and Poland showed co-infection between PPIV-1 and SIV (7, 13, 18). A previous study in Italy reported co-infection among PPIV-1, SIV, and PRRSV (12). However, the severity of clinical signs and the role of PPIV-1 as a primary or secondary pathogen in co-infected animals need further investigation.
Currently (as of April 2024), only 29 whole genome sequences of PPIV-1 are available in the GenBank database. Our result expands the PPIV-1 genomic information in the nucleotide database. This study has provided four complete genomes and eleven F and HN gene sequences of PPIV-1 from Thailand and Myanmar. The complete genome of Thai PPIV-1 (n = 2) and Myanmar PPIV-1 (n = 2), as well as F and HN gene sequences of Thai PPIV-1 (n = 6) and Myanmar PPIV-1 (n = 5) were deposited in the database. Based on the phylogenetic analysis of the complete genome, Thai PPIV-1s (n = 2) were grouped with lineage II of PPIV-1 (American lineage) and were closely related to American and Chinese strains. On the other hand, Myanmar PPIV-1s (n = 2) belonged to lineage I (European lineage) and were closely related to Hong Kong (China), German, and South Korean strains. These findings supported a high genetic diversity of PPIV-1 in the Southeast Asia region, as in the previous report. For the F gene, the phylogenetic analysis showed that Thai PPIV-1s (n = 7) were grouped with lineage II (American lineage) except for one Thai PPIV-1 strain (CU32451) (n = 1), which is closely related to the lineage I (European lineage), the prototype strain from Hong Kong (China) (S033N; JX857410.1). It should be noted that heterosubtypic recombination events between PPIV-1 lineage I and II had been observed in Chinese PPIV-1 (19). However, the recombination of Thai-PPIV-1 (CU32451) requires further investigation through whole genome analysis sequencing. On the other hand, Myanmar PPIV-1s (n = 7) belonged to lineage I (European lineage) and were closely related to Hong Kong (China), German, and South Korean strains. Genetic analysis of the F gene indicated that twelve distinct amino acid positions are specific to a particular lineage. For example, Myanmar PPIV-1 has the same amino acid pattern as PPIV-1 of lineage I, whereas those of the Thai PPIV-1 strains, except CU32451, are similar to PPIV-1 of lineage II. The differences in nucleotides that correspond to unique amino acid positions among lineages can be utilized to develop diagnostic tests for lineage identification in the future.
Due to the global trade of breeding pigs and pig-derived products, pigs can be carriers of transboundary emerging diseases, such as PPIV-1 (20). It is important to note that the pig industry and its products in Thailand heavily depend on imports and exports from the U.S. and European countries. Meanwhile, medium and small-scale pig farms in Myanmar primarily source their livestock through cross-border trade. This reliance can contribute to variations in the PPIV-1 lineage across the region. Hence, it is crucial to genetically characterize these viruses from various geographical regions to determine the origin of PPIV-1s. It is also important to monitor the introduction of PPIV-1 into Thailand and Myanmar.
In conclusion, this study is the first to report and genetically characterize the complete genome of PPIV-1s in pig farms located in Thailand and Myanmar. Phylogenetic analysis showed that Thai PPIV-1s are closely related to PPIV-1s from China and the USA. On the other hand, Myanmar PPIV-1s are closely related to PPIV-1s from Hong Kong (China), Germany, and South Korea. This research provides current disease status and genetic diversity information on PPIV-1 in pig farms located in Thailand and Myanmar. The surveillance and molecular epidemiology of PPIV-1 should be further investigated on a larger scale to monitor the geographical distribution, evolution, and potential zoonotic transmission of PPIV-1.
The authors declare that the data supporting the findings of this study are available in Supplementary Tables. The nucleotide sequence data that support the findings of this study are openly available in the GenBank database at https://www.ncbi.nlm.nih.gov/genbank/, under accession numbers # PP766830-833, PP761466-487.
The animal studies were approved by the Chulalongkorn University Animal Cares and Uses Protocol (CU-VET IACUC 2331099) and the Livestock Breeding and Veterinary Department (LBVD), Yangon and Nay Pyi Taw, Myanmar. The studies were conducted in accordance with the local legislation and institutional requirements. Written informed consent was not obtained from the owners for the participation of their animals in this study because verbal consent was obtained from all pig farm owners for sample collection in compliance with the ARRIVE guidelines.
HP: Investigation, Methodology, Writing – original draft, Writing – review & editing. KC: Investigation, Methodology, Writing – review & editing. CN: Formal analysis, Investigation, Methodology, Writing – review & editing. KU: Investigation, Writing – review & editing. EP: Investigation, Methodology, Writing – review & editing. YT: Investigation, Methodology, Writing – review & editing. HS: Investigation, Methodology, Writing – review & editing. SC: Formal analysis, Investigation, Methodology, Writing – review & editing. TL: Investigation, Methodology, Writing – review & editing. MM: Investigation, Methodology, Writing – review & editing. AA: Conceptualization, Formal analysis, Funding acquisition, Investigation, Project administration, Writing – original draft, Writing – review & editing.
The author(s) declare that financial support was received for the research, authorship, and/or publication of this article. Chulalongkorn University provided financial support to the Center of Excellence for Emerging and Re-emerging Infectious Diseases in Animals (CUEIDAs) and One Health Research Cluster. The Thailand Science Research and Innovation Fund Chulalongkorn University (HEAF68310011) supported this research.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
The Supplementary material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fvets.2025.1435920/full#supplementary-material
1. Lau, SK, Woo, PC, Wu, Y, Wong, AY, Wong, BH, Lau, CC, et al. Identification and characterization of a novel paramyxovirus, porcine parainfluenza virus 1, from deceased pigs. J Gen Virol. (2013) 94:2184–90. doi: 10.1099/vir.0.052985-0
2. Park, JY, Welch, MW, Harmon, KM, Zhang, J, Piñeyro, PE, Li, G, et al. Detection, isolation, and in vitro characterization of porcine parainfluenza virus type 1 isolated from respiratory diagnostic specimens in swine. Vet Microbiol. (2019) 228:219–25. doi: 10.1016/j.vetmic.2018.12.002
3. Henrickson, KJ. Parainfluenza Viruses. Clin Microbiol Rev. (2003) 16:242–64. doi: 10.1128/CMR.16.2.242-264.2003
4. Morrison, TG. Structure and function of a paramyxovirus fusion protein. Biochim Biophys Acta. (2003) 1614:73–84. doi: 10.1016/S0005-2736(03)00164-0
5. Stadejek, T, Cybulski, P, Gauger, PC, and Woźniak, A. European and American strains of porcine parainfluenza virus 1 (Ppiv-1) belong to two distinct genetic lineages. Pathogens. (2022) 11:375. doi: 10.3390/pathogens11030375
6. Schuele, L, Lizarazo-Forero, E, Cassidy, H, Strutzberg-Minder, K, Boehmer, J, Schuetze, S, et al. First detection of porcine Respirovirus 1 in Germany and the Netherlands. Transbound Emerg Dis. (2021) 68:3120–5. doi: 10.1111/tbed.14100
7. Agüero, B, Mena, J, Berrios, F, Tapia, R, Salinas, C, Dutta, J, et al. First report of porcine Respirovirus 1 in South America. Vet Microbiol. (2020) 246:108726. doi: 10.1016/j.vetmic.2020.108726
8. Dénes, L, Cságola, A, Schönhardt, K, Halas, M, Solymosi, N, and Balka, G. First report of porcine parainfluenza virus 1 (species porcine Respirovirus 1) in Europe. Transbound Emerg Dis. (2021) 68:1731–5. doi: 10.1111/tbed.13869
9. Lunardi, M, Darold, GM, Francescon, RR, and Alfieri, AA. First report of porcine Respirovirus 1 in Brazil. Microb Pathog. (2023) 182:106222. doi: 10.1016/j.micpath.2023.106222
10. Palinski, RM, Chen, Z, Henningson, JN, Lang, Y, Rowland, RR, Fang, Y, et al. Widespread detection and characterization of porcine parainfluenza virus 1 in pigs in the USA. J Gen Virol. (2016) 97:281–6. doi: 10.1099/jgv.0.000343
11. Park, J, Kim, HR, Kim, JM, Lee, KK, Kim, WI, Lyoo, YS, et al. First report of porcine Respirovirus 1 in South Korea. Transbound Emerg Dis. (2022) 69:4041–7. doi: 10.1111/tbed.14715
12. Sozzi, E, Leo, G, Bertasio, C, Alborali, GL, Salogni, C, Tonni, M, et al. Presence and characterisation of porcine Respirovirus 1 (Prv 1) in northern Italy. Pathogens. (2024) 13:85. doi: 10.3390/pathogens13010085
13. Woźniak, A, Cybulski, P, Denes, L, Balka, G, and Stadejek, T. Detection of porcine Respirovirus 1 (Prv 1) in Poland: incidence of co-infections with influenza a virus (Iav) and porcine reproductive and respiratory syndrome virus (Prrsv) in herds with a respiratory disease. Viruses. (2022) 14:148. doi: 10.3390/v14010148
14. Thibault, PA, Watkinson, RE, Moreira-Soto, A, Drexler, JF, and Lee, B. Zoonotic potential of emerging paramyxoviruses: knowns and unknowns. Adv Virus Res. (2017) 98:1–55. doi: 10.1016/bs.aivir.2016.12.001
15. OFFLU. Offlu-_Collection_of_Specimens_for_Detection_of_Influenza__December2017.Pdf (2017). Available at: https://www.offlu.org/wp-content/uploads/2021/01/OFFLU-_Collection_of_Specimens_for_Detection_of_Influenza__December2017.pdf.
16. Spackman, E, Senne, DA, Myers, T, Bulaga, LL, Garber, LP, Perdue, ML, et al. Development of a real-time reverse transcriptase Pcr assay for type a influenza virus and the avian H5 and H7 hemagglutinin subtypes. J Clin Microbiol. (2002) 40:3256–60. doi: 10.1128/JCM.40.9.3256-3260.2002
17. Li, Y, Sthal, C, Bai, J, Liu, X, Anderson, G, and Fang, Y. Development of a real-time Rt-Qpcr assay for the detection of porcine Respirovirus 1. J Virol Methods. (2021) 289:114040. doi: 10.1016/j.jviromet.2020.114040
18. Graaf-Rau, A, Hennig, C, Lillie-Jaschniski, K, Koechling, M, Stadler, J, Boehmer, J, et al. Emergence of swine influenza a virus, porcine Respirovirus 1 and swine Orthopneumovirus in porcine respiratory disease in Germany. Emerg Microbes Infect. (2023) 12:2239938. doi: 10.1080/22221751.2023.2239938
19. Zhao, J, Sun, J, Li, X, Xing, G, Zhang, Y, Lai, A, et al. Divergent viruses discovered in swine Alter the understanding of evolutionary history and genetic diversity of the Respirovirus genus and related porcine parainfluenza viruses. Microbiol Spectr. (2022) 10:e0024222. doi: 10.1128/spectrum.00242-22
Keywords: genetic characterization, Myanmar, pig, PPIV-1, Thailand
Citation: Phyu HW, Charoenkul K, Nasamran C, Udom K, Phyu EM, Thaw YN, Soe HW, Chaiyawong S, Lin TN, Maw MT and Amonsin A (2025) Genetic characterization of porcine parainfluenza virus 1 (PPIV-1) in pig farms: first report of PPIV-1 in Thailand and Myanmar. Front. Vet. Sci. 12:1435920. doi: 10.3389/fvets.2025.1435920
Received: 22 May 2024; Accepted: 04 February 2025;
Published: 19 February 2025.
Edited by:
Gianmarco Ferrara, University of Messina, ItalyReviewed by:
Ram Kumar, Guru Angad Dev Veterinary and Animal Sciences University, IndiaCopyright © 2025 Phyu, Charoenkul, Nasamran, Udom, Phyu, Thaw, Soe, Chaiyawong, Lin, Maw and Amonsin. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Alongkorn Amonsin, YWxvbmdrb3JuLmFAY2h1bGEuYWMudGg=
†ORCID: Alongkorn Amonsin, orcid.org/0000-0001-6769-4906
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.