- 1Anhui Province Key Laboratory of Animal Nutritional Regulation and Health, College of Animal Science, Anhui Science and Technology University, Fengyang, China
- 2Nanjing Hongshan Forest Zoo, Nanjing, China
- 3MOE Joint International Research Laboratory of Animal Health and Food Safety, College of Veterinary Medicine, Nanjing Agricultural University, Nanjing, China
Objective: Entamoeba spp. are globally distributed zoonotic parasites that infect various hosts, among which non-human primates (NHPs) have been identified as one of the most common hosts of these parasites. Consequently, the infections of Entamoeba spp. in captive NHPs from Nanjing Hongshan Forest Zoo in China were investigated in order to assess their zoonotic potential.
Methods: A total of 120 fresh fecal samples, including 19 species of NHPs, were collected from four breeding bases of the zoo from May to June 2019. The infections of six species of Entamoeba spp. were detected by PCR using the 16S or 18S rDNA-specific primers, and the positive samples were sequenced and analyzed.
Results: Entamoeba spp. were detected as positive in 59 NHPs fecal samples (49.17%), including five Entamoeba species: Entamoeba histolytica (7.50%), E. dispar (22.50%), E. coli (22.50%), E. chattoni (10.00%) and E. nuttalli (1.67%). Infection with one Entamoeba species was more common (35%) than co-infections (13.33%) or infections with three Entamoeba species (0.83%). There was a significantly higher prevalence rate of Entamoeba spp. in the species Pongo pygmaeus and Macaca mulatta than in Papio sp., Mandrillus sphinx, and Saimiri sciureus.
Conclusion: Entamoeba spp. are highly prevalent in the NHPs raised in Nanjing Hongshan Forest Zoo. Therefore, attention should be paid to the development of containment strategies of Entamoeba spp. in this zoological garden.
Introduction
Entamoeba histolytica, the causative agent of amebiasis, is a protozoan parasite that infects about 10% of the world' population and causes 100,000 deaths per year globally (1, 2). The genus Entamoeba includes several species, such as E. dispar and E. moshkovskii, which are morphologically similar to E. histolytica (3). Although E. dispar and E. moshkovskii have no apparent invasive potential, these two species exhibit some pathogenicity (4, 5). E. coli, E. hatmanni, and E. nuttalli are considered non-pathogenic; however, those species are frequently found in the stools of several animals (6–8). As a reference method for parasitological diagnosis, the microscopic examination has been widely used to identify Entamoeba species, even though it lacks the ability to differentiate the infection caused by E. histolytica from non-pathogenic Entamoeba spp. (9). Therefore, molecular methods have been preferably used to identify and investigate the epidemiology of Entamoeba spp. in human and animal hosts (6, 7, 10, 11).
Among the hosts of Entamoeba spp., non-human primates (NHPs) have a close phylogenetic relationship with human, which increases the risk of Entamoeba spp. transmission between NHPs and humans. Previous studies showed that both captive and free-ranging NHPs are frequently infected with Entamoeba spp. (11, 12), and many Entamoeba spp. were identified in NHPs, especially E. histolytica (6, 13). Given that captive NHPs have more opportunities to contact with humans than free-ranging, they are a major concern group of host with a higher potential for zoonotic infections (6, 14).
Nanjing Hongshan Forest Zoo is one of the most distinctive zoological gardens in China with abundant NHPs animal resources and colorful theme activities, attracting millions of visitors every year. However, there is inadequate information on the epidemic of parasitic zoonoses within this zoological garden. The presence of zoonotic parasites poses a threat to the health of NHPs, which may in turn cause zoonotic transmission events between tourists and NHPs. Consequently, in this study, we aim to investigate the prevalence of Entamoeba spp. in NHPs in Nanjing Hongshan Forest Zoo to assess the potential for zoonotic transmission.
Materials and Methods
Collection of Stool Samples
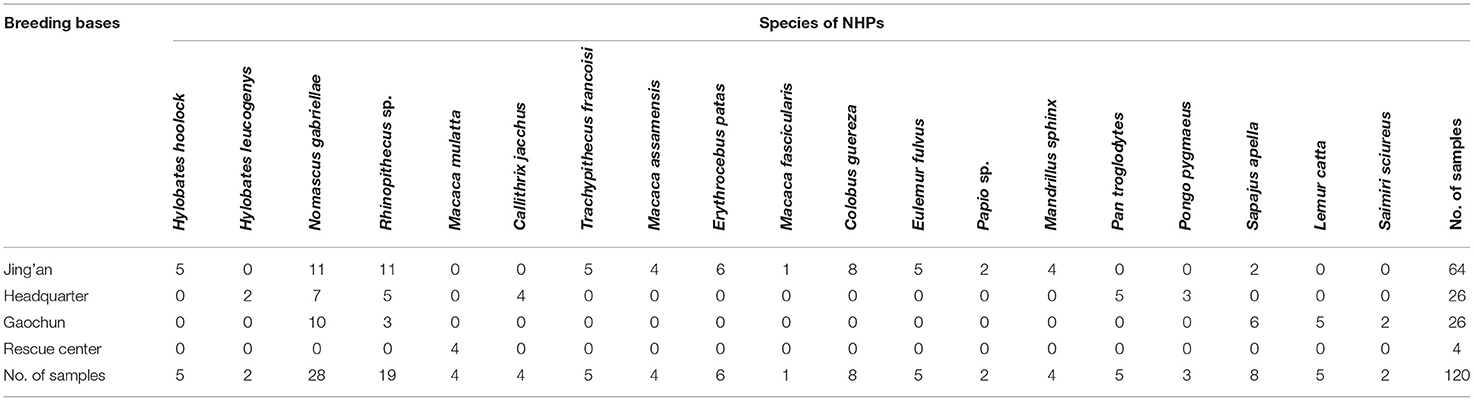
From May to June 2019, stool samples of 120 NHPs were collected from four breeding bases, including the headquarter (26/120), Gaochun base (26/120), Jing'an base (64/120), and rescue center (4/120) of Nanjing Hongshan Forest Zoo. There were 19 species included in 120 NHPs. About 15g fecal sample were collected immediately after defecation for each individual and placed in a plastic container marked with the name, species, gender, and breeding base. Fecal samples were then stored at 4°C for DNA extraction. The detailed information of the samples is shown in Table 1.
Detection of the Samples by Polymerase Chain Reaction
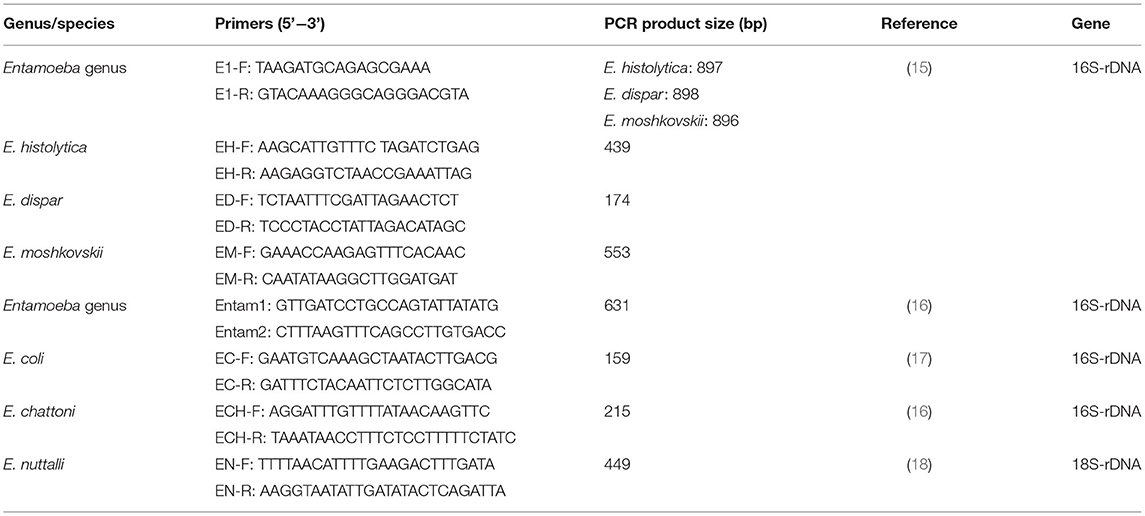
From each sample, 0.2 g of stool were used to isolate genomic DNA using the Stool DNA Kit (Tiangen Biotech, Beijing, China) according to the manufacturer's instructions. The genomic DNA was then detected by PCR assays based on the 16S or 18S rDNA sequence of six Entamoeba species according to previous studies (15–18). E. chattoni and E. nuttalli were detected by one round PCR, while E. histolytica, E. dispar, E. moshkovskii, and E. coli were detected by nested PCR using specific primers (Table 2). After the final PCR reaction, the products were examined by agarose gel electrophoresis, and the size of target fragments was validated by comparing them with DL2000 DNA Marker (TaKaRa, Dalian, China).
Sequence Analysis and Data Accessibility
Positive samples with the correct molecular size were purified and confirmed by two-directional sequencing using the specific PCR primers for each species by Sangon Biotech (Shanghai, China). The obtained nucleotide sequences were aligned with published international reference sequences available from the NCBI GenBank™ database using BLAST.
The assembled sequences obtained in this study were submitted to GenBank under accession number ON254797 for E. histolytica (9 samples), ON254798 (1 sample) and ON254799 (1 sample) for E. nuttalli, ON254800 for E. chattoni (12 samples), ON254801 (21 samples) and ON254802 (6 samples) for E. coli, and ON254803 for E. dispar (27 samples).
Statistical Analysis
The samples with the sequence that can be assigned to the reference sequences were defined as positive, and used to calculate the prevalence of Entamoeba spp. infections. The statistical analysis was performed to analyze the risk factors, including breeding bases and genders, for Entamoeba spp. infections in NHPs using SPSS 25.0. A chi-square test was used to analyze the non-metric variables in the form of frequency tables. P < 0.05 was considered significant.
Result
Entamoeba spp. Infection in NHPs
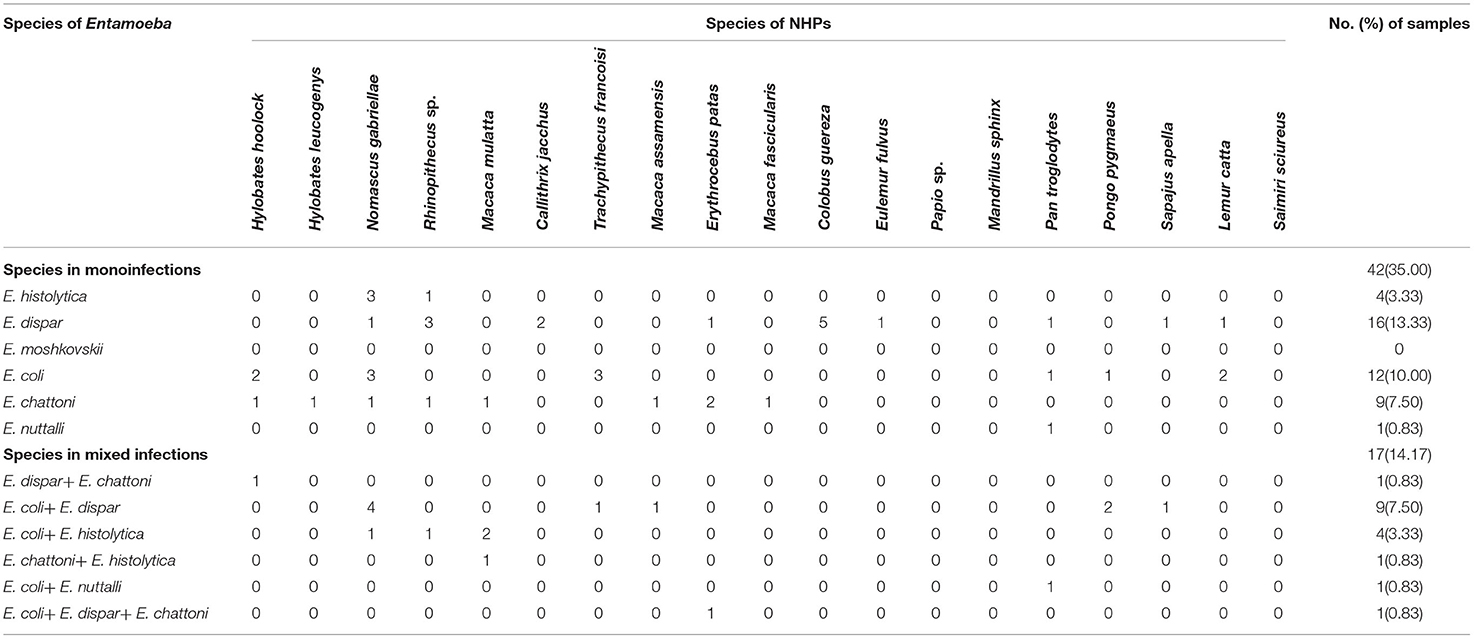
Among 120 stool samples from NHPs, 59 (49.17%) were detected as Entamoeba spp. positive. As shown in Table 3, the most common species detected were E. coli and E. dispar in 27 (22.50%) samples, respectively, followed by E. chattoni (10.00%, 12/120), E. histolytica (7.50%, 9/120) and E. nuttalli (1.67%, 2/120). None sample presented infection with E. moshkovskii. Statistical analysis showed that there was a significant difference in the infection rate of different Entamoeba spp. in the NHPs (X2 = 63.003, p = 0.000).
Multiple Infections
Among the 59 positive stool samples, 17 presented multiple infections (14.17%), from which 16 corresponded to co-infections and 1 to triple infection. Co-infections E. coli + E. dispar represented 52.94% (9/17) of the co-infections, and the only case of triple infection was with E. coli + E. dispar + E. chattoni (Table 3).
Multiple infections were detected in 9 among the 19 species of NHPs. The highest rate of multiple infection was revealed in M. mulatta (75.0%, 3/4), followed by P. pygmaeus (66.7%, 2/3). The multiple infections of the other species are shown in Table 3.
Risk Factors for the Infection of Entamoeba. spp in NHPs
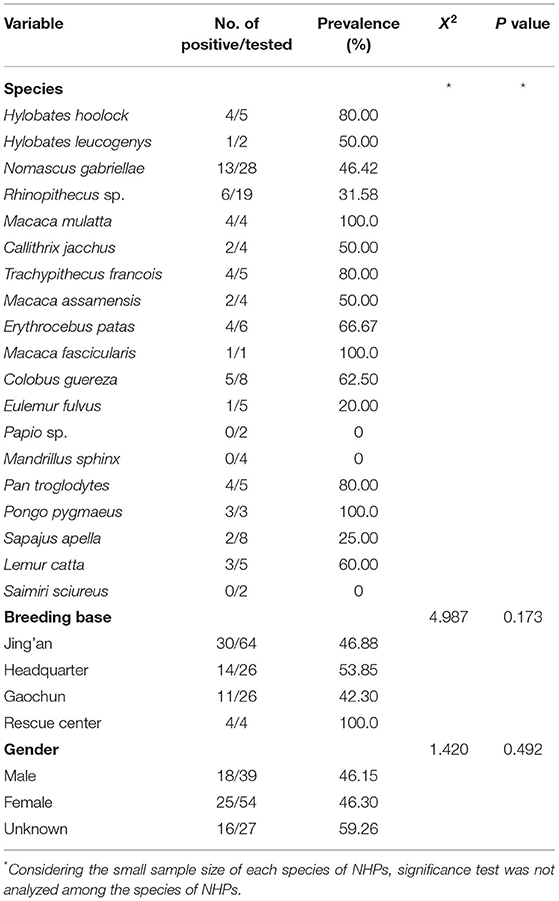
From the 19 species of NHPs, M. mulatta, M. fascicularis, and P. pygmaeus were all positive for Entamoeba spp. infections, while none of the fecal samples obtained from Papio sp., M. sphinx, and S. sciureus were detected as positive. The infection rates of other species of NHPs are shown in Table 4.
As shown in Table 4, all the samples from the rescue center presented Entamoeba infection, while the infection rates of the other three breeding bases were close to 50.0%. No statistical significance was observed among different bases (X2 = 4.987, p = 0.173).
To assess the effect of NHPs gender on Entamoeba spp. infection, the infection rates of the gender categories (male, female, and unknown) were analyzed. It should be noted that there was no significant association of Entamoeba spp. infection with gender (X2 = 1.420, p = 0.492) (Table 4). Nevertheless, no E. histolytica-positive samples were detected in the NHPs male category.
Discussion
The Entamoeba genus contains many species residing in the intestinal lumen, of which E. histolytica is the main species associated with pathological sequelae (9). Nevertheless, other species are also frequently detected in a range of animals, such as goat, pig, yak, mice, and alpacas (7, 19–22). In addition, both captive and free-ranging NHPs have been identified as common hosts of Entamoeba spp. (6, 11, 14).
The present study provides the first information on the epidemiological data of Entamoeba spp. infections in NHPs from Nanjing Hongshan Forest Zoo (49.17%, 59/120). The prevalence rate of Entamoeba spp. was lower than in the free-ranging NHPs in savanna woodland (Tanzania, Pan troglodytes, 79%) (23) and Taihangshan (China, Macaca mulatta tcheliensis, 89.96%) (11). Compared to zoological gardens housing captive NHPs, the positivity was higher than those in Ibadan (Nigeria, Erythrocebus patas, Cercocebus atys, Mandrillus sphinx, Cercopithecus sabaeus, 13.90%) (14), Belgium (Belgium, prosimians, New World monkeys, Old World monkeys and apes, 44%) (24), and the experimental macaques in Yunnan (China, M. mulatta, M. nemestrina and M. fascicularis, 9.31%) (13). These results indicate that free-ranging NHPs may have a much higher Entamoeba occurrence rate than captive ones. It is likely that captive NHPs are largely housed in zoological gardens and research facilities, and the captive management may interrupt the transmission of parasitic pathogens to hosts. Of note, the infection rate of Entamoeba spp. in captive NHPs from Nanjing Hongshan Forest Zoo is much higher than those reported in several epidemiological studies (seven NHPs species) (13, 14) suggesting that this attraction should strengthen the housing and husbandry practices to improve the health of NHPs.
Except for E. moshkovskii, the other five Entamoeba spp. species were found in this study, presenting different infection rates. Compared with the studies in Belgium and the Netherlands (four Entamoeba species among 36 NHPs species), China (three Entamoeba species among three NHPs species) and Italy (three Entamoeba species among nine NHPs species), more species of Entamoeba spp. were detected in Nanjing Hongshan Forest Zoo (6, 13, 25). The infection rate of E. histolytica was 7.50%, which is consistent with the data reported by the zoological gardens in Belgium and Netherlands (36 NHPs species, 8.1%) and the experimental macaques (M. mulatta, M. nemestrina and M. fascicularis, 6.38%) in Yunnan Province in China (6, 13). In contrast, no E. histolytica infection was detected in savanna woodland (Tanzania, Pan troglodytes), Taihangshan (China, Macaca mulatta tcheliensis), and an Italian zoological garden (Italy, nine NHP species) (11, 23, 25). Our results revealed higher diversity of Entamoeba species in NHPs in Nanjing Hongshan Forest Zoo China, and these NHPs may be potential sources for human infection with E. histolytica.
Entamoeba dispar and E. coli were the more prevalent species revealed in this study, consistent with other 13 zoos/parks in China (26). While in the NHPs from zoological gardens in the UK and the experimental NHPs from Japan and China, only E. dispar/E. coli was detected as the main species for infections (27–29). In addition, E. hartmanni was the most prevalent Entamoeba species shed in the NHPs in Belgium and Netherlands, and in savanna woodland chimpanzees (6, 23), and E. chattoni was the predominant species shed in macaques from wild Taihangshan and southwest China (11, 30). These results indicated that different regions had their specific prevalent species of Entamoeba, which might be related to the parameters including lifestyles, and the species of NHPs. The observation needs to be further studied. There is, to date, no report indicating the E. moshkovskii infections in NHPs (11, 25, 31). Likewise, no E. moshkovskii-positive sample was detected in the NHPs in this study. However, the cases of human infection with E. moshkovskii have been reported in Malaysia, Japan and Colombia (32–34), and E. moshkovskii has also been detected in animal hosts, such as cattles (UK), elephants (Namibia), and chelonian (Mexico) (35, 36). Further investigations are needed to demonstrate whether NHPs can be infected by E. moshkovskii.
The multiple infection rate of Entamoeba spp. in the NHPs in this study was 14.17%. It was lower than that in the other 13 zoos/parks in China (45.4%) and in the free-ranging Taihangshan Macaques (84.93%) (11, 25). The differences in the prevalence of multiple infections may be related to the method used for Entamoeba detection. Microscopic examination was used in those studies, which may not allow correct discriminations of Entamoeba species. Compared with the studies using molecular techniques, the multiple infection rate in this study was consistent with the studies in Tanzania (16.67%) (22), higher than that in Italian (0) (24), and lower than that in Belgium and the Netherlands (51.90%) (6, 22, 24). The difference in Entamoeba multiple infections in NHPs may be associated with the climate, the species of NHPs or breeding and management conditions, which need to be further studied. Entamoeba spp. was prevalent in all four breeding bases of the zoo, especially in the rescue center. It could attribute to the poor conditions in which the NHPs were raised before being rescued. Importantly, both males and females have a high infection rate, whereas no significant difference was observed (X2 = 1.42, p = 0.492), which was consistent with the finding in the experimental macaques in Yunnan (13). It indicates that the infection of Entamoeba spp. was not correlated to the gender of NHPs. However, all the E. histolytica-positive samples were detected from females, which warrants further investigations.
In the 19 species of NHPs, no M. sphinx and Papio sp. was confirmed with Entamoeba spp. infections. In a previous study, both the semi-free-ranging (100%) and captive (81.20%) M. sphinx presented a high infection rate of Entamoeba spp. (26, 37), and Papio sp. was verified to be a natural host of Entamoeba spp. (38). It is likely that our sampling size was too small to identify the positive infection of Entamoeba spp. in these two NHPs species. In Martinez et al.'s research, S. sciureus was not susceptible to E. histolytica (39). In our study, S. sciureus was found to be Entamoeba spp. negative, which could be explained by the small sample size. The infection rate of Entamoeba spp. ranged from 25% (Sapajus apella) to 100% (M. mulatta, Macaca fascicularis, and P. pygmaeus), indicating that the infection of Entamoeba spp. was widespread in most species of NHPs in this zoo.
In conclusion, this study revealed that Entamoeba spp. are widely prevalent in the captive NHPs in Nanjing Hongshan Forest Zoo. Furthermore, five species (E. coli, E. dispar, E. histolytica, E. chattoni, and E. nuttalli) were identified, with E. coli and E. dispar as the predominant species. The epidemiology of Entamoeba spp. infection is complex, and these captive NHPs could potentially participate in zoonotic transmission, and the Entamoeba spp. infection of other animals from this zoological garden. The prevention and control of Entamoeba spp. infection in this zoological garden should be strengthened to reduce disease exposure risk of public health.
Data Availability Statement
The datasets supporting the findings of this article are included within the article. Additional datasets generated for this study are available on reasonable request to the corresponding authors.
Ethics Statement
The animal study was reviewed and approved by the Animal Care and Welfare Committee of Anhui Science and Technology University, Fengyang, China. Written informed consent was obtained from the owners for the participation of their animals in this study.
Author Contributions
XL designed this study, performed data analysis, and drafted the manuscript. GB, MY, YYG, and YF helped to carry out the study. WL and YFG helped to design this study and perform data analysis. WC participated in the collection of experimental materials. ML critically revised the manuscript. All authors read and approved the final version of the manuscript.
Funding
This work was supported by the Modern Cattle and Goat Industrial Technology System Program of Anhui Province, the Anhui University Collaborative Innovation Project (GXXT-2019-035), the Anhui Provincial Natural Science Foundation (1808085MC84 and 1908085QC116), the School-Level Talent Introduction Project of Anhui Science and Technology University (DKYJ201902), the Natural Science Research Projects of Anhui Science and Technology University (2021lzryb08), and National College Students Innovative and Entrepreneurial Training Program (202110879064 and 202010879040).
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Publisher's Note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
References
1. Hung CC. Amoebiasis: current status in Australia. Med J Aust. (2007) 187:412–6. doi: 10.5694/j.1326-5377.2007.tb01293.x
2. World Health Organization. WHO/Pan American Health Organization/UNESCO Report of a Consultation of Experts on Amoebiasis. Wkly Epidemiol Rec. (1997) 72:97–9.
3. Lauren DM. Distinguishing between pathogenic and non-pathogenic species of Entamoeba. Lab Med. (2004) 35:613–5. doi: 10.1309/B81NPVAW8Y4BGY11
4. Jackson T. Entamoeba histolytica and Entamoeba dispar are distinct species, clinical, epidemiological and serological evidence. Int J Parasitol. (1998) 28:181. doi: 10.1016/S0020-7519(97)00177-X
5. Tengku SA, Norhayati M. Public health and clinical importance of amoebiasis in Malaysia: a review. Trop Biomed. (2011) 28:194–222. doi: 10.1371/journal.ppat.1002177
6. Levecke B, Dreesen L, Dorny P, Verweij JJ, Vercammen F, Casaert S, et al. Molecular identification of Entamoeba spp. in captive nonhuman primates. J Clin Microbiol. (2010) 48:2988–00. doi: 10.1128/JCM.00013-10
7. Al-Habsi K, Yang R, Ryan U, Jacobson C, Miller DW. Morphological and molecular characterization of an uninucleated cyst-producing Entamoeba spp. in captured rangeland goats in western Australia. Vet Parasitol. (2017) 235:41–6. doi: 10.1016/j.vetpar.2017.01.013
8. Matsubayashi M, Suzuta F, Terayama Y, Shimojo K, Yui T, Haritani M, et al. Ultrastructural characteristics and molecular identification of Entamoeba suis isolated from pigs with hemorrhagic colitis: implications for pathogenicity. Parasitol Res. (2014) 113:3023–8. doi: 10.1007/s00436-014-3965-y
9. Fotedar R, Stark D, Beebe N, Marriott D, Harkness J. Laboratory diagnostic techniques for Entamoeba species. Clin Microbiol Rev. (2007) 20:511–3. doi: 10.1128/CMR.00004-07
10. Roshdy HM, El-Kader NA, Tammam M, Fuentes I, Mohamed M, EI-Sheikh NA, et al. Molecular diagnosis of Entamoeba spp. versus microscopy in the Great Cairo. Acta Parasitol. (2017) 62:188–91. doi: 10.1515/ap-2017-0022
11. Zhang Q, Liu K, Wang C, Luo J, Lu J, He H. Molecular characterization of Entamoeba spp. in wild Taihangshan Macaques (Macaca mulatta tcheliensis) in China. Acta Parasitol. (2019) 64:228–31. doi: 10.2478/s11686-019-00026-y
12. David EB, Patti M, Coradi ST, Oliveira-Sequeira TCG, Ribolla PEM, Guimaraes S. Molecular typing of Giardia duodenalis isolates from nonhuman primates housed in a Brazilian zoo. Rev Inst Med Trop Saõ Paulo. (2014) 56:49–54. doi: 10.1590/S0036-46652014000100007
13. Pu LH, Li Z, Wu J, Zhang YL, Zou FC. Prevalence, molecular epidemiology and zoonotic risk of Entamoeba spp. from experimental macaques in Yunnan Province, southwestern China. Parasitol Res. (2020) 119:2733–40. doi: 10.1007/s00436-020-06762-9
14. Adetunji VE. Prevalence of gastro-intestinal parasites in primates and their keepers from two zoological gardens in Ibadan, Nigeria. Sokoto. J Vet Sci. (2014) 12:25–30. doi: 10.4314/sokjvs.v12i2.5
15. Khairnar K, Parija SC. A novel nested multiplex polymerase chain reaction (PCR) assay for differential detection of Entamoeba histolytica, E. moshkovskii and E. dispar DNA in stool samples. BMC Microbiol. (2007) 7:47. doi: 10.1186/1471-2180-7-47
16. Verweij JJ, Polderman AM, Clark CG. Genetic variation among human isolates of uninucleated cyst-producing Entamoeba species. J Clin Microbiol. (2001) 39:1644–6. doi: 10.1128/JCM.39.4.1644-1646.2001
17. Tachibana H, Yanagi T, Akatsuka A, Kobayashi S, Kanbara H, Tsutsumi V. Isolation and characterization of a potentially virulent species Entamoeba nuttalli from captive Japanese macaques. Parasitology. (2009) 136:1169–77. doi: 10.1017/S0031182009990576
18. Tachibana H, Yanagi T, Pandey K, Cheng XJ, Kobayashi S, Sherchand JB, et al. An Entamoeba spp. strain isolated from rhesus monkey is virulent but genetically different from Entamoeba histolytica. Mol Biochem Parasit. (2007) 153:107–14. doi: 10.1016/j.molbiopara.2007.02.006
19. Ren M, Yang F, Gou JM, Wang PX, Lin Q. First detection and molecular identification of Entamoeba in yaks from China. Acta Parasitol. (2020) 66. doi: 10.1007/s11686-020-00258-3
20. Hirashima Y, Manchanayake T, Yano T, Kitahara S, Koreeda T, Kamimura S, et al. Development of molecular diagnostic protocols for detecting three types of Entamoeba from diarrheal and asymptomatic pigs and environmental moist soils. Parasitol Res. (2017) 116:2001–7. doi: 10.1007/s00436-017-5483-1
21. Chikako S, Mamun K, Mami T, Dinesh M, Seiki K, Ali IKM, et al. Entamoeba moshkovskii is associated with diarrhea in infants and causes diarrhea and colitis in mice. J Infect Dis. (2012) 206:744–51. doi: 10.1093/infdis/jis414
22. Gao WW, Ma YT, Ma YY, Li RL, Zhu XQ. First report of Eimeria and Entamoeba infection in alpacas (Vicugna pacos) in Shanxi Province, northern China. Parasitol Res. (2021) 120:2031–5. doi: 10.1007/s00436-021-07157
23. Pomajbíková KJ, Cepicka I, Kalousová B, Jirku M, Stewart F, Levecke B, et al. Molecular identification of Entamoeba species in savanna woodland chimpanzees (Pan troglodytes schweinfurthii). Parasitology. (2016) 143:741–8. doi: 10.1017/S0031182016000263
24. Levecke B, Dorny P, Geurden T, Vercammen F, Vercruysse J. Gastrointestinal protozoa in non-human primates of four zoological gardens in Belgium. Vet Parasitol. (2007) 148:236–46. doi: 10.1016/j.vetpar.2007.06.020
25. Berrilli F, Prisco C, Friedrich KG, Cerbo PD, Cave D, Liberato CD. Giardia duodenalis assemblages and Entamoeba species infecting non-human primates in an Italian zoological garden: zoonotic potential and management traits. Parasit Vectors. (2011) 4:119. doi: 10.1186/1756-3305-4-199
26. Dong H, Li J, Qi M, Wang R, Yu F, Jian F, et al. Prevalence, molecular epidemiology, and zoonotic potential of Entamoeba spp. in nonhuman primates in China. Infect Genet Evol. (2017) 54:216–20. doi: 10.1016/j.meegid.2017.07.002
27. Regan CS, Yon L, Hossain M, Elsheikha HM. Prevalence of Entamoeba species in captive primates in zoological gardens in the UK. Peer J. (2014) 2:e492. doi: 10.7717/peerj.492
28. Tachibana H, Cheng XJ, Kobayashi S, Matsubayashi N, Gotoh S, Matsubayashi K. High prevalence of infection with Entamoeba dispar, but not E. histolytica, in captive macaques. Parasitol Res. (2001) 87:14–7. doi: 10.1007/s004360000289
29. Feng M, Yang B, Yang L, Fu Y, Tachibana H. High prevalence of Entamoeba infections in captive long-tailed macaques in China. Parasitol Res. (2011) 109:1093–7. doi: 10.1007/s00436-011-2351-2
30. Feng M, Cai J, Min X, Fu Y, Xu Q, Tachibana H, Cheng X. Prevalence and genetic diversity of Entamoeba species infecting macaques in southwest China. Parasitol Res. (2013) 112:1529–36. doi: 10.1007/s00436-013-3299-1
31. Elsheikha HM, Regan CS, Clark CG. Novel Entamoeba findings in nonhuman primates. Trends Parasitol. (2017) 1724:12. doi: 10.1016/j.pt.2017.12.008
32. Anuar TS, Al-Mekhlafi HM, Ghani M, Azreen SN, Salleh FM, Ghazali N, et al. First molecular identification of Entamoeba moshkovskii in Malaysia. Parasitol. (2012) 139:1521–5. doi: 10.1017/S0031182012001485
33. Zebardast N, Yeganeh F, Gharavi MJ, Abadi A, Tabaei SS, Haghighi A. Simultaneous detection and differentiation of Entamoeba histolytica, E. dispar, E. moshkovskii, Giardia lamblia and Cryptosporidium spp. in human fecal samples using multiplex PCR and qPCR-MCA. Acta Trop. (2016)162:233–8. doi: 10.1016/j.actatropica.2016.07.004
34. Consuelo LM, León C, Jairo F, Patricia R, Ligia M, Olivera MJ, et al. Molecular epidemiology of Entamoeba: first description of Entamoeba moshkovskii in a rural area from central Colombia. PLoS ONE. (2015) 10: e0140302. 0140302 doi: 10.1371/journal.pone.0140302
35. Jacob AS, Busby EJ, Levy A, Komm N, Clark CG. Expanding the Entamoeba universe: new hosts yield novel ribosomal lineages. J Eukaryotic Microbiol. (2016) 63:69–78. doi: 10.1111/jeu.12249
36. Garcia G, Ramos F, Perez RG, Yanez J, Estrada MS, Mendoza LH et al. Molecular epidemiology and genetic diversity of Entamoeba species in a chelonian collection. J Med Microbiol. (2014) 63:271–83. doi: 10.1099/jmm.0.061820-0
37. Setchell JM, Bedjabaga IB, Goossens B, Reed P, Wickings EJ, Knapp LA. Parasite prevalence, abundance, and diversity in a semi-free-ranging colony of Mandrillus sphinx. Int J Primatol. (2007) 28:1345–62. doi: 10.1007/s10764-007-9225-6
38. Myers BJ, Malherbe KH. Intestinal commensals and parasites of the South African baboon (Papio cynocephalus). Trans Am Microsc Soc. (1971) 90:80–3. doi: 10.2307/3224900
Keywords: molecular identification, non-human primates, zoological garden, Entamoeba spp., infection
Citation: Liu X, Bao G, Yue M, Fang Y, Gu Y, Li W, Gu Y, Cheng W and Lu M (2022) Prevalence and Molecular Identification of Entamoeba spp. in Non-human Primates in a Zoological Garden in Nanjing, China. Front. Vet. Sci. 9:906822. doi: 10.3389/fvets.2022.906822
Received: 29 March 2022; Accepted: 09 May 2022;
Published: 30 May 2022.
Edited by:
Yukifumi Nawa, Khon Kaen University, ThailandReviewed by:
Serena Cavallero, Sapienza University of Rome, ItalyLongxian Zhang, Henan Agricultural University, China
Copyright © 2022 Liu, Bao, Yue, Fang, Gu, Li, Gu, Cheng and Lu. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Xinchao Liu, bGl1eGNoQGFoc3R1LmVkdS5jbg==; Mingmin Lu, bWluZ21pbi5sdUBuamF1LmVkdS5jbg==
 Xinchao Liu
Xinchao Liu Guangbin Bao1
Guangbin Bao1 Wenchao Li
Wenchao Li Mingmin Lu
Mingmin Lu