
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
BRIEF RESEARCH REPORT article
Front. Synth. Biol. , 26 March 2025
Sec. Metabolic Engineering
Volume 3 - 2025 | https://doi.org/10.3389/fsybi.2025.1473338
Limosilactobacillus reuteri strain DSM 20016 is specialised to colonize the human gut for much longer than other L. reuteri strains and most other Lactobacillaceae family members. These adaptations, along with its safe-to-consume food status and public acceptance as a probiotic, make it an attractive chassis for synthetic biology endeavours aimed at introducing novel functions into the gut microbiome, including feedback systems for sensing disease state and therapeutic applications for remedying chronic disorders. Here, we perform whole-genome sequencing and present a novel variant of L. reuteri DSM 20016 (now denoted “LAD4” in this work; DSMZ repository number 116333) with mutations that disrupt DNA restriction-modification and cell wall regulation; these appear to enable increased uptake of the PAMβ1-origin low copy-number plasmid pTRKH3. Additional mutations include genes involved in protein degradation ability, alkaline shock responses, and a mobile genetic element transfer. One of these mutations, or some combination of them, enables stable, consistent production and detection, without the need to buffer media, of the acid-resistant reporter protein mCherry2. This novel variant, in combination with the pTRKH3 plasmid backbone, will enable researchers to more easily utilize this uniquely positioned microbe, which was previously limited by inconsistent reporter protein production and unreliable growth characteristics.
Members of the Lactobacillales order of bacteria, also known as lactic acid bacteria (LAB), have a wide range of medical (Jansson et al., 2019; Nikawa et al., 2004), therapeutic (Urbańska et al., 2016; Yang et al., 2021; Bermúdez-Humarán et al., 2011; Mao et al., 2018), and industrial (Sauer et al., 2017) applications. They are also integral to the production of a variety of fermented foods, including coffee, chocolate, cheese, and yoghurt (Hutkins, 2007).
Interest in the genetic manipulation of LAB has increased due to their many useful properties and potential applications. However, the breadth of phenotypes and genotypes among LAB has led to a diverse collection of species being selected as targets for research. This has resulted in there being no clear model organism for LAB studies. While some species are undoubtedly favoured over others, no species has experienced the level of development of E. coli or S. cerevisiae for Gram-negative bacteria and fungi, respectively. The lack of tractable model organisms has hindered LAB research on multiple fronts.
Limosilactobacillus reuteri is generally regarded as a safe-for-consumption (GRAS) probiotic species of LAB (Mu et al., 2018) that has been consistently isolated from a broad range of vertebrate gastrointestinal tracts, including those of fish (Ahmad et al., 2022), avian, and mammalian hosts (Walter et al., 2011); individual L. reuteri strains have evolved specific adaptations that allow for the effective colonization of specific hosts (MacKenzie et al., 2010). Human-associated L. reuteri strains are particularly attractive as a chassis for human interventions, either as in situ producers of therapeutic molecules (Hendrikx et al., 2019) or as whole-cell biosensors for disease-state detection (Lubkowicz et al., 2018). L. reuteri DSM20016 (strain designation F275) is particularly well adapted, via host-specific mucosal binding proteins (MacKenzie et al., 2010), to the human gastrointestinal (GI) tract, and it can persist in the human gut for extended periods of time (Reuter, 2001) relative to traditional probiotic strains (Lahti et al., 2013; Wang et al., 2015). This makes it well suited to human intestinal applications in biosensing and therapeutics, including interventions used to treat intestinal disorders such as inflammatory bowel diseases.
This strain has been observed to suffer from inconsistent behaviour with regard to the transformation of exogenous DNA and the expression of heterologous proteins (Duggan and McMillen, 2023; Team Oxford, 2019), which may hinder its adoption. Here, we have identified and characterized a mutant strain of L. reuteri DSM20016 (termed “LAD4”) with substantially improved properties that enable consistent transformation with the otherwise incompatible pTRKH3 plasmid and generates more reliable protein expression. These characteristics make L. reuteri LAD4 particularly promising as a chassis for LAB work in synthetic biology, and it has the potential to become a more widely adopted representative for this group of bacteria. Use of this strain (available from DSMZ as DSM 116333 or upon request) will provide a solid foundation for researchers wishing to carry out synthetic biology work in a human-associated lactic acid bacterium.
Plasmid pNZ123 was a generous gift from the Mahadevan laboratory and was altered via Gibson assembly to be resistant to erythromycin instead of chloramphenicol and to accept modular cloning (MoClo) parts; the mCherry2 open reading frame (ORF) was then inserted via GoldenGate MoClo assembly. pTRKH3_ermGFP was acquired from Addgene, the existing GFP ORF was replaced via Gibson assembly to accept MoClo parts, and the mCherry2 ORF was then inserted via GoldenGate MoClo assembly. The mCherry2 ORF consisted of constitutive promoter Pbs3 designed for protein production in a wide range of organisms (Yang et al., 2018) as well as universal RBS part BBa_K2918014, mCherry2 codon optimised for L. acidophilus using IDT’s codon optimisation web-tool, and a BBaB0015 double terminator (sequences available in the Supporting Information). The plasmid was harvested from E. coli DH10β before transformation into L. reuteri DSM20016.
L. reuteri DSM20016 was grown in De–Man Rogosa Sharpe (MRS) media at 37 °C in a static benchtop incubator for 16 h. MRS agar plates were made by adding 15 g/L agar to MRS broth prior to autoclaving. Growth on MRS agar was carried out in anaerobic chambers for 2–3 days or until colonies were visible. Transformation of both L. reuteri DSM20016 and LAD4 was based on a protocol by Berthier et al. (1996), with some optimisation and critical modifications previously published, including not incubating plasmid DNA with electrocompetent L. reuteri at all prior to electroporation and growth in an anaerobic environment after plating (Duggan and McMillen, 2023). Antibiotic concentrations in both liquid and solid media were 250 μg/mL erythromycin or 35 μg/mL chloramphenicol for E. coli and 10 μg/mL erythromycin or 25 μg/mL chloramphenicol for L. reuteri.
Colonies were selected based on two factors: an OD600 of over 1, indicating successful growth post-picking, and a RFU value over 100, indicating effective heterologous protein production. Seven of the DSM20016 colonies tested met these criteria and were inoculated into 200 mL MRS broth via a 1:1000 dilution with no selection. After 16 h, cultures were transferred into 200 mL of fresh MRS via a 1:100 dilution. This transfer was repeated a third time before 200-µL aliquots were plated onto selective and non-selective MRS agar, and glycerol stocks of the seven sub-strains were created. No colonies were observed on any of the selective plates, indicating that all sub-strains had lost the resistance-conferring plasmid.
Sub-strains were retransformed with the same pTRKH3-mCherry2 plasmid using both dam-/dcm-plasmid and the DH10β methylation pattern plasmid. After 2 days, colonies for both conditions were counted, and as many as possible (up to 88) were inoculated into selective filter-sterilised MRS media. After 16 h, 200-µL aliquots were transferred to clear-bottom microplates for measurement. LAD4 behaviour was corroborated across three repeats (see S1 and S2).
Prior to quantifying fluorescence and OD600, cells were grown for 24 h in filter-sterilised MRS, and 200-µL aliquots from these cultures were put into Corning black-walled 96-well optical plates for measurement. A Tecan™ Infinite® M1000 Pro plate reader was then used to measure for OD600 and fluorescence at the mCherry2 excitation:emission (589:610 nm) wavelengths. Gain was set at 180.
Genomic extraction for L. reuteri was performed using the Qiagen MagAttract HMW DNA Kit, following their protocol for extraction from Gram-positive organisms (QIAGEN, 2023) with the following modifications.
After step 1 of the procedure, the pellet was resuspended and washed in standard P1 buffer and centrifuged again before moving on to step 2. This was to remove lactic acid produced by L. reuteri during growth and mitigates interference with later steps.
At step 3, 20 µL from a 100 mg/ml lysozyme stock was added along with 6 µL from a 20 mg/ml mutanolysin stock. The dual action of both is required to efficiently lyse L. reuteri (Duggan and McMillen, 2023).
At step 4, 30 min was found to be ample time to lyse the cells. Longer lysis incubations result in progressive genomic DNA degradation (see Supplementary Figure S3).
Primers for pTRKH3 plasmid, containing constitutive mCherry2 expression, were used (forward: cacccgttctcggagca, reverse: ctacgagttgcatgataaagaagaca) using a protocol previously published with non-standard modifications, including sampling from liquid culture, a NaOH boil, and keeping the samples chilled at all reasonable times (Duggan and McMillen, 2023). Expected band lengths should be ∼1200 base pairs.
The assay was modelled on a protocol by Barrick Jeffrey (2022) with the following modifications. The antibiotic used was changed to carbenicillin due to L. reuteri’s resistance to suggested antibiotics. On Day −2, cells were revived from glycerol stock and grown in 25 mL non-delective MRS media overnight at 37 °C with no shaking. On Day −1, 100 µL of the overnight growths were reinoculated into 10 mL fresh MRS media, mixed gently via swirling, and then had 250 µL inoculated into 25 mL fresh MRS media for a second overnight growth at 37 °C with no shaking. On Day 0, the overnight cultures were each diluted down to ∼5,000 cells/mL and separated into as many aliquots as were to be plated and were allowed to grow overnight for exactly 24 h at 37 °C, no shaking. On Day 1, the non-selective conditions were diluted by a factor of 10–6, and 200 µL was plated. For the conditions being spread onto selective plates, 200 µL was plated directly from the overnight growth with no dilution. Plates were then incubated anaerobically for 2 days (non-selective conditions) and 4 days (selective conditions) at 37 °C before being counted. Analysis was performed using fluxxer.R, as described by Barrick Jeffrey (2022).
Genomic DNA extracted from our L. reuteri DSM20016 reference strain and the L. reuteri LAD4 mutant were sequenced at SeqCenter (Pittsburgh, United States) using the Illumina NextSeq platform with 151-bp, paired-end reads. Raw reads were first processed using Trimmomatic v.0.39 to remove Nextera PE adapter sequences and low-quality reads (ILLUMINACLIP–NexteraPE-PE.fa, Seed Mismatch = 2, Palindromic Clip Threshold = 30, Simple Clip Threshold = 10; SLIDINGWINDOW–Window Size = 4, Required Quality = 5; AVGQUAL – 20; MINLEN – 100) (Bolger et al., 2014). Quality control was then performed on all filtered sequences with fastQC v0.11.9 using default settings (Andrews, 2010). The complete genome sequencing results have been deposited in the Harvard Dataverse repository (https://doi.org/10.7910/DVN/5YYAXA).
Processed reads were analysed using breseq v0.37.1 with default settings in order to map reads to the L. reuteri DSM20016 reference genome (ASM1682v1) and identify mutations (Deatherage and Barrick, 2014). More than 99% of reads from both our L. reuteri DSM20016 and L. reuteri LAD4 strains aligned successfully with the reference genome, with average coverages of 295× and 446×, respectively. Focusing on the high-confidence mutations identified by breseq, we first eliminated five mutations that were called in both isolates to develop a mutational profile that differentiated our L. reuteri LAD4 strain from our L. reuteri DSM20016 strain. This left us with the seven mutations spanning thirteen genes present only in L. reuteri LAD4 that are discussed in this study.
Interpro scan and affiliate databases (Paysan-Lafosse et al., 2023; Thomas et al., 2022) were used to identify putative domain and family associations for mutated genes in LAD4. UniProt BLAST (Consortium, 2023) was used to find gene homologs with more complete annotations. PDBe was used to find and analyse mutation positions in crystal structures when applicable (Armstrong et al., 2020). BioCyc and affiliate databases were used to investigate potential gene product interactions (Caspi et al., 2016).
Although L. reuteri DSM20016 has been manipulated and utilized previously (Lubkowicz et al., 2018; Lizier et al., 2010), attempts to replicate these manipulations both by ourselves and by others have been unsuccessful (Team Oxford, 2019). In our research, we have found this is largely a result of L. reuteri DSM20016 suffering from low transformation efficiencies with the previously utilized low copy-number pTRKH3 plasmid (Lubkowicz et al., 2018; Lizier et al., 2010) (∼80 CFU/μg DNA at 80 nM), necessitating the plating of entire transformant solutions to increase the chances of yielding plasmid-positive colonies, averaging only 1.5 colonies per plate (Duggan and McMillen, 2023). Even when colonies were picked successfully, they were found to have only a roughly 25% chance of growing effectively in liquid culture, and among those with successful growth, detection of constitutively produced heterologous protein, inferred via fluorescence, was unpredictable (Duggan and McMillen, 2023). Since enhanced green fluorescent protein (EGFP) is sensitive to the low pH environment common in L. reuteri DSM20016 (Lizier et al., 2010), we chose mCherry2 as a reporter protein due to its significantly higher acid tolerance (Duggan and McMillen, 2023), in an attempt to prevent the possible acidic denaturing of GFP from causing inconsistent fluorescence levels.
Control plates were also included in transformations, with antibiotic selection but without any plasmid included in the electroporation step. These plates yielded zero colonies, suggesting that all colonies chosen under antibiotic selection did carry the antibiotic resistance plasmid containing constitutive mCherry2. As fluorescence readings could not be used to reliably confirm plasmid presence, colony PCR was carried out on 48 post-transformation colonies and was found to be sporadic and unreliable (Figure 1). Colony PCR also suggested that colonies failing to yield growth when inoculated into liquid culture could still possess the correct resistance-conferring plasmid.
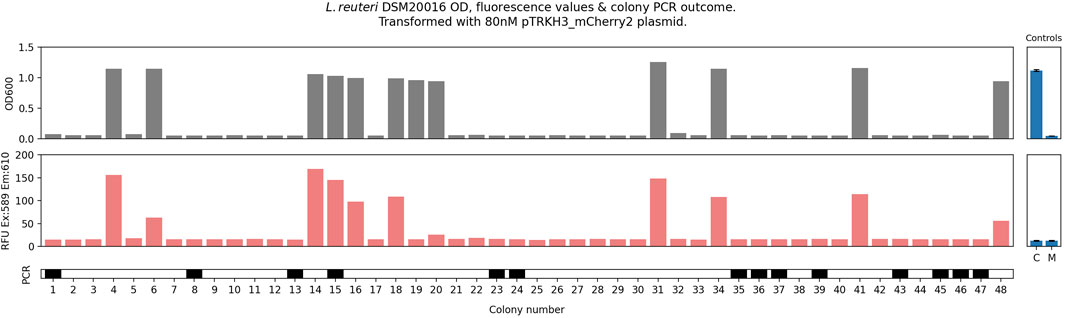
Figure 1. Electrocompetent L. reuteri (100 μL) was transformed with 80 nM pTRKH3_mCherry2 plasmid; 800 µL of recovery was plated. This was repeated eight times (32 plates). All colonies (48) were picked and grown for 24 h, after which 200-µL aliquots were measured for optical density at 600 nm (OD600) and fluorescence (excitation at 589 nm, emission at 610 nm, reported in raw fluorescence units—RFU). 5-μL aliquots were subject to colony PCR to confirm plasmid uptake, with a solid bar indicating that a positive PCR band was seen. Positive and negative PCR controls gave the expected results. Controls: C, cell only with no plasmid or selection; M, media only, no cells. This is adapted from Duggan and McMillen, 2023.
These characteristics greatly hinder experimental replicability and make wild-type (WT) DSM 20016 unsuitable for high-throughput experiments. They also present a fundamental barrier to its use in synthetic biology applications, which often rely on the fine-tuning of consistent, predictably controlled gene products or pathways.
The pSH71-origin plasmid pNZ123 was also tested (De Vos, 1987) and found to be far more compatible for WT DSM 20016 transformation (Supplementary Figure S4). When transformed with pNZ-mCherry2 and plated, L. reuteri gave a dual-phenotype of both slow- and fast-growing colonies (Supplementary Figure S5); this phenotypic variance has been observed in other L. reuteri strains (Ortiz-Velez et al., 2018). However, even when only one phenotype was selected for measurement, either inoculation growth or protein production was inconsistent under a constitutive promoter (Supplementary Figures S6, S7), frustrating efforts to obtain consistent replicability.
We transformed L. reuteri DSM20016 with pTRKH3-mCherry and selected all colonies that exhibited both inoculation survival and at least half of maximally observed mCherry2 production (seven in total, labelled LAD1 to LAD7). We then cured them of their plasmid and retransformed, along with a WT control. Some debate exists as to the methylation pattern requirements of DNA when transforming into L. reuteri DSM20016 (Ahrne et al., 1992); to further elucidate this, all LADs were transformed with plasmid isolated from both E. coli DH10β and a Dam−/Dcm− knockout E. coli (Dam Dcm Competent E, 2023). No significant preference for plasmid DNA methylation was noted (Figure 2A), further corroborating our own previous observations (Duggan and McMillen, 2023).
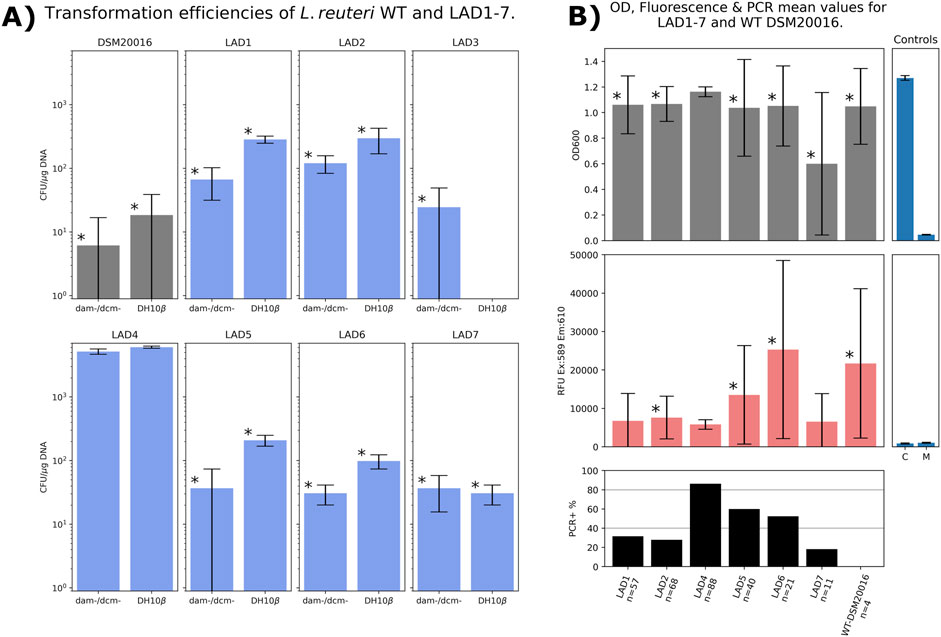
Figure 2. (A) Transformation efficiencies of WT DSM20016 alongside LADs 1 through 7, all tested with 80 nM non-methylated (dam-/dcm-) or methylated (DH10β) pTRKH3_mCherry2 plasmid. All LAD3 colonies failed to grow after picking and so were excluded from further measurements. Bars marked “*” denote results significantly different (t-test p = <0.05) from those obtained with LAD4, compared pairwise within each methylation condition (LAD4 non-methylated vs. each strain; LAD4 methylated vs. each strain). (B) As many colonies as could be picked (up to 88) were measured for optical density at 600 nm (OD600) and fluorescence (excitation at 589 nm, emission at 610 nm, reported in raw fluorescence units, RFU) and subjected to colony PCR for plasmid confirmation. Positive and negative PCR controls gave the expected results. Controls: C, cell only with no plasmid or selection; M, media only. Bars marked “*” denote results with significant differences (p = <0.05) in pairwise comparisons with LAD4.
Six of the LAD strains exhibited slightly higher transformation efficiencies, and all seven showed higher levels of inoculation-survival than average among the initial 48 strains examined. However, only one strain, LAD4, exhibited consistent colony survival, significantly heightened transformation efficiency, and relatively low but highly consistent constitutive mCherry2 production (see Figure 2).
An examination of 88 colonies of LAD4 (Figure 3) confirmed that it possessed notable characteristics not usually found in L. reuteri DSM20016: significantly (p = <0.05) improved transformation efficiencies (5600 CFU/μg DNA, vs. ∼80 CFU/μg for the WT, or averaging 100 CFU/μg DNA for other LAD strains); significantly (p = <0.05) improved liquid culture growth of all picked colonies (100% for LAD4 vs. ∼25% success in the WT or averaging 66% in other LAD strains); detectable heterologous protein expression with substantially greater consistency than in other strains (RFU coefficient of variation (CV: standard deviation divided by mean) of 0.21, compared to a CV of 1.17 for the WT, and CVs ranging from 0.73 to 1.11 in the other LAD strains—see Figure 1); correct bands when examined using colony PCR in ∼80% of replicates. Across several repeats (Supplementary Figures S1 and S2), LAD4 demonstrated far greater consistency in plasmid transformation and protein expression when transformed with pTRKH3 plasmid than seen previously in WT-DSM20016. The LAD4-pTRKH3 chassis–plasmid combination functions as a robust, effective system for introducing heterologous DNA and obtaining consistent heterologous protein expression L. reuteri DSM20016, suggesting that it could play a helpful role in synthetic biology applications.
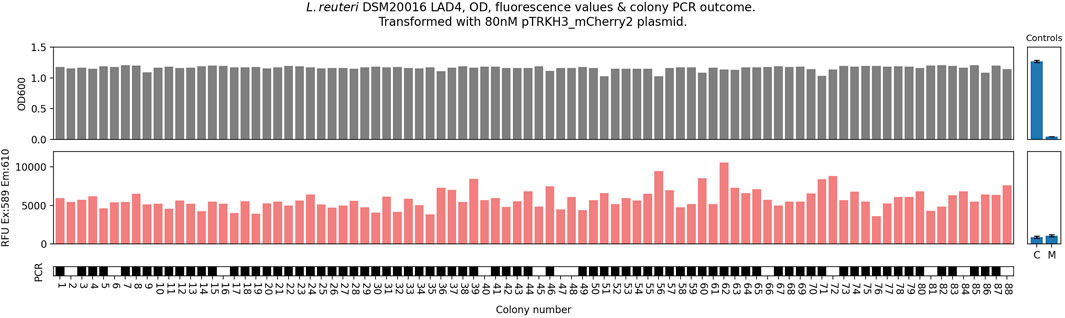
Figure 3. L. reuteri DSM20016, sub-strain LAD4 colony behaviour when cured and retransformed with pTRKH3-mCherry2 plasmid, picked, grown in filtered MRS and measured for OD600 and mCherry Ex/Em values, and colony PCR, with a solid block indicating that a correct colony PCR band was observed. Positive and negative PCR controls gave the expected results.
To investigate the underlying causes of these positive features, we performed whole-genome sequencing on both the WT L. reuteri DSM200016 strain and the L. reuteri LAD4 mutant strain. Both were compared against the L. reuteri DSM200016 reference genome (GenBank accession: GCA_000016825.1) to distinguish background mutations from our WT-DSM20016, potentially accrued since receipt from the repository, from mutations specific to LAD4.
Twelve mutations were identified, five of which were present in both strains and seven of which were specific to LAD4. The seven mutations spanned thirteen genes (Figure 4; Table 1).
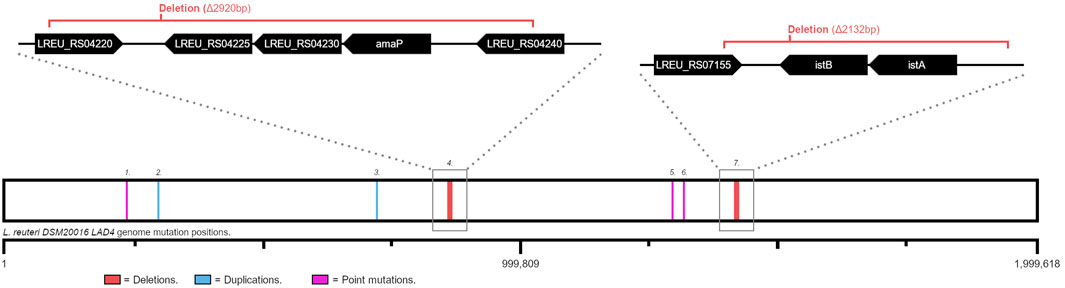
Figure 4. Each LAD4 specific mutation position mapped onto the L. reuteri DSM20016 genome. Each mutation number corresponds to a numbered entry in Table 1. Blue lines: base pair duplication events; pink lines: point mutations; red lines (with expanded regions): deletions of the regions covered by the red brackets. Expanded genome track genes are not to scale.
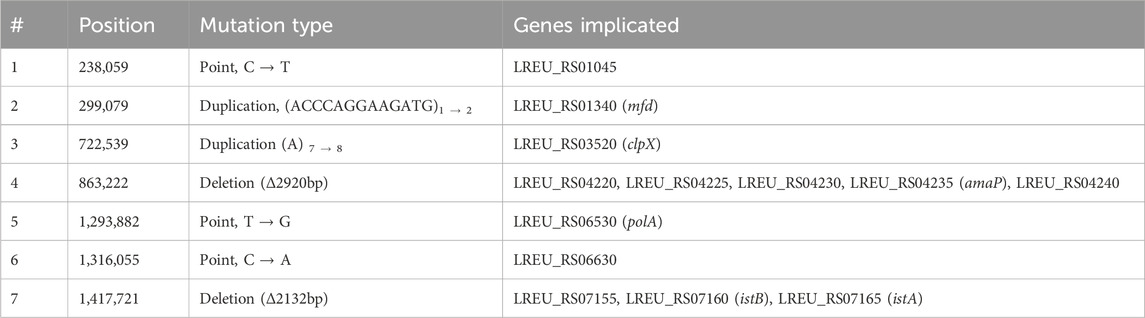
Table 1. Mutation details for LAD4 specific genome alterations. Entries correspond to numbered positions in Figure 4. Gene sequences available in Supporting Information.
Of the thirteen genes mutated only in LAD4, seven are associated with DNA repair, maintenance, replication, or modification; three are associated with alkaline stress response; two are associated with protein production, folding, or degradation; and one is a sugar metabolism gene.
The mutations associated with DNA processing and repair are as follows. LREU_RS01045 (mutation #1 in Figure 4; Table 1) encodes a putative site-specific DNA methyltransferase that likely uses cofactor S-adenosyl-L-methionine (SAM) as a methyl-group donor (IPR029063) (Paysan-Lafosse et al., 2023) and may be a component of a type-II restriction modification system in L. reuteri that methylates host DNA to protect it from type-II restriction enzyme action (IPR029063) (Paysan-Lafosse et al., 2023; Roberts et al., 2023). The mutation results in 44 residues being omitted from the C-terminus. According to the AlphaFold predicted structure (Jumper et al., 2021), these deleted residues comprise two alpha helix structures, one of which is functionally considered highly conserved for SAM-methyltransferases (Martin and McMillan, 2002). This deletion is thus likely to have destroyed or strongly limited the functionality of the DNA methyltransferase, although it is unclear what effect this would have on the overall restriction modification system.
LREU_RS01340 (mfd) (mutation #2 in Figure 4; Table 1) produces the protein Mfd. When RNA polymerase encounters DNA damage, it stalls. Mfd recognises the stall and binds to the RNA polymerase-DNA complex; Mfd then translocates along the DNA strand, pushing RNA polymerase to disassociate from it. Mfd also recruits global genome repair (GGR) machinery to the site, enabling repair (Hanawalt and Spivak, 2008). The Mfd protein is essential for the transcription coupled repair (TCR) of DNA damage, but its absence may not be fatal: GGR only requires Mfd when the site is blocked by stalled polymerase. If the site is accessible, GGR machinery will still bind without issue. In LAD4, the mutation in Mfd is a duplication of 13 bases within the gene, which shifts the reading frame and introduces a premature stop codon, resulting in the loss of 72% of the protein and thus likely a loss of Mfd function. The loss of this gene is expected to result in the loss of TCR, with the bacteria relying solely on GGR to maintain genome integrity.
LREU_RS06530 (polA) (mutation # 5 in Figure 4; Table 1) contains a point mutation (T204P), changing an amino acid at the end of an α-helix from threonine to proline. polA is found ubiquitously in all organisms and encodes DNA polymerase I (Pol I) in bacteria. Bacteria have multiple DNA polymerases, each with differing functions, but Pol I is concerned with the initiation of DNA replication and the repair of DNA damage due to proofreading and bidirectional exonuclease activity (Allen et al., 2010).
The Pol I protein is highly modular. Its N-terminal region, which is responsible for 5’ – 3’ exonuclease activity, can even be removed without disrupting polymerase activity (Allen et al., 2010). This N-terminal fragment of Pol I is responsible for the cleavage and removal of damaged DNA, enabling the polymerase fragment to repair the now single-stranded region (Allen et al., 2010), as well as the cleavage of RNA from RNA–DNA junctions—a function essential to processing Okazaki fragments during replication (Allen et al., 2010). It is within this N-terminal fragment that the point mutation arises.
It is unclear whether this mutation is significant enough to alter 5’–3’ exonuclease activity. If activity is disrupted, it would potentially impair genomic replication on the reverse strand or impair the Pol I mediated lesion excision and possibly increase the rate at which genomic mutations occur.
LREU_RS06630 (mutation #6 in Figure 4; Table 1) suffers a point mutation (A315S), changing an alanine to a serine. Examination of the InterPro protein database predicts homology between this gene and the phosphoesterase family, with similarity to an E. coli exonuclease SbcD (Paysan-Lafosse et al., 2023). Domain-level homology to calcineurin-like phosphoesterase type ApaH and homology to the N-terminal metallophosphatase domain of the Mre11 nuclease (Paysan-Lafosse et al., 2023) also emerge from the InterPro database. Mre11 is required for double-strand break repair in yeast, with the E. coli homolog SbcD performing a similar function. PANTHER classification further indicates homology to an ATP-dependent double-strand DNA exonuclease component involved in DNA repair (Thomas et al., 2022).
Inferring from similarity to various other domains, it seems likely that this gene is involved in DNA repair, most likely double-strand break repair. It is unclear if the point mutation would alter the protein’s activity, although crystal structures of the E. coli SbcD protein show this portion of the protein to be distal to the homodimer-DNA binding active sites (Armstrong et al., 2020), potentially lessening the mutation’s impact on function.
LREU_RS07155 (mutation #7 in Figure 4; Table 1) is a pseudogene, having already suffered a premature stop codon mutation in the L. reuteri DSM20016 reference genome. The deletion event in LAD4 removes a portion of the pseudogene occurring after the premature stop codon, meaning that this mutation does not have an impact on a functional protein.
Analysis via InterPro revealed that the gene’s original protein product likely contained a helix-turn-helix domain (Paysan-Lafosse et al., 2023), indicating potential DNA-binding activity, as well as a larger domain with homology to the DNA transposase InsF (Thomas et al., 2022). A BLAST search revealed several other Lactobacillaceae family members with genes showing 70% or greater identity with the pseudogene; these genes are annotated as IS3 family transposases. Analysis of these related proteins revealed that the catalytic core of the putative transposase was disrupted by the initial premature stop codon.
LREU_RS07160 (istB) (mutation #7 in Figure 4; Table 1) has been completely deleted in LAD4. istB is one component of a two-gene transposase operon (Arias-Palomo and Berger, 2015). Five istB dimers bind to dsDNA target sites and induce a 180° bend, allowing efficient insertion of new DNA by istA (Arias-Palomo and Berger, 2015).
LREU_RS07165 (istA) (mutation #7 in Figure 4; Table 1) has also been deleted in LAD4. istA binds to transposon ends, recognises DNA-IstB complex, and binds to it, removing istB from the DNA strand in the process (Arias-Palomo and Berger, 2015). It then carries out strand transfer and insertion of the mobile genetic element.
Both istA and istB, along with small flanking regions, make up insertion sequence 21 (IS21). This is implicated in a number of mobile genetic elements and is itself mobile (Arias-Palomo and Berger, 2015; Spínola-Amilibia et al., 2023). The deletion of these genes likely has little impact on the essential functioning of L. reuteri DSM20016, as they appear to be concerned only with their own propagation. However, IS21 does not exhibit insertion target sequence specificity (Spínola-Amilibia et al., 2023), implying that the deletion of IS21 could potentially result in a more stable DSM 20016 strain, less susceptible to the disruption of introduced genetic elements to increase fitness or escape cell stress.
The mutations associated with alkaline stress response are as follows. LREU_RS04225 (mutation #4 in Figure 4; Table 1) is entirely deleted in L. reuteri LAD4. Homology analysis indicates the gene encoded an alkaline-shock protein, conserved within Gram-positive bacteria (Müller et al., 2014) belonging to the Asp23 family (Paysan-Lafosse et al., 2023). In S. aureus, Asp23 is bound to the cell membrane by the anchor protein AmaP. Upon alkaline shock, Asp23 dissociates and becomes soluble within the cell (Müller et al., 2014). Deletion of Asp23 in S. aureus resulted in the upregulation of cell wall stress response genes also upregulated during vancomycin cell-wall challenge, including VreSR, a cell wall stress sensor, and PrsA (McCallum et al., 2006), along with several genes of unknown function (Müller et al., 2014).
LREU_RS04230 (mutation #4 in Figure 4; Table 1) is entirely deleted. The gene shares homology to a domain of unknown function (DUF)223 family proteins (Paysan-Lafosse et al., 2023). DUF223 is within the Asp23 operon between the putative Asp23 and AmaP. The DUF223 proteins are present in many species, suggesting a conserved role, but they are not currently sufficiently characterized for us to suggest a specific effect of this deletion.
LREU_RS04235 (amaP) (mutation #4 in Figure 4; Table 1) is entirely deleted in LAD4. The protein AmaP likely acts in concert with LREU_RS04225 to enable the Asp23 operon alkaline-shock response. AmaP may well localise LREU_RS04225 to the cell wall under normal conditions, releasing it under stress (Müller et al., 2014). The deletion of amaP also demonstrably upregulates cell wall stress response genes in S. aureus (Müller et al., 2014). Deletion of the entire operon in L. reuteri LAD4 may well have a similar effect.
The two mutations related to the post-translational modification and degradation of proteins are the following. LREU_RS03520 (clpX) (mutation #3 in Figure 4; Table 1) harbours an insertion in LAD4: a region of seven adenine bases has an eighth adenosine added. This shifts the reading frame, resulting in a premature stop codon and the loss of 94% of the protein product. ClpX forms a hexamer, and the deletion of most of the gene undoubtedly destroys its ability to function correctly. ClpX is a member of the Clp family of ATPases; although most members of the family are upregulated during heat shock, ClpX is not and has a somewhat variable role across organisms (Frees et al., 2007). Several members of this family, including ClpX, are capable of guiding proteins into the proteolytic ClpP complex for degradation (Frees et al., 2007). In the Gram-positive model organism B. subtilis, ClpX was found to be five times more numerous than any other ClpP associated chaperone protein (Frees et al., 2007). ClpX can aid in the refolding of protein aggregates or inclusion bodies without degradation if unbound to ClpP (Frees et al., 2007).
ClpX deletion in B. anthracis can result in thinner cell walls (Zou et al., 2021), potentially presenting less of an issue for transiting macromolecules such as DNA during transformation (Aune and Aachmann, 2010). In both B. subtilis and Streptococcus mutans, ClpX deletion was shown to decrease naturally acquired competency (Pandey and Biswas, 2022).
LREU_RS04220 (mutation #4 in Figure 4; Table 1) is severely truncated in LAD4 because of a deletion in this section of the genome: in the 164 residue gene, 155 residues are deleted from the C-terminus, while six residues are added from the other side of the deletion before a stop codon is encountered. This gene likely produces an acetyl-CoA-acyltransferase, as suggested by homology to the GCN5-related N-acetyltransferase family (Paysan-Lafosse et al., 2023). This group of enzymes catalyses the transfer of an acetyl moiety from an acetyl-CoA donor to a target molecules primary amine group (Burckhardt and Escalante-Semerena, 2020). This protein family has broad function and can be involved with many cellular processes, preventing us from predicting the effect of this gene deletion without further investigation.
Finally, one mutation relates to the sugar metabolism. LREU_RS04240 (mutation #4 in Figure 4; Table 1) suffered substantial deletion in LAD4, losing 319 amino acid residues from the C-terminus of the originally 509 residue protein product. It likely encodes an FGGY family carbohydrate kinase (Paysan-Lafosse et al., 2023). Members of this family have been found to perform ATP-dependent phosphorylation on a single sugar, with the specific molecular target varying across at least nine sugars for different family members (Zhang et al., 2011). The loss of this gene’s function would likely result in impaired utilization of a single sugar carbon source; however, bacteria often possess multiple FGGY carbohydrate kinase genes, allowing for the utilization of a variety of possible sugar sources. The result of this gene disruption is thus likely to cause impaired growth only under specific sugar-limiting conditions.
In summary, we cannot make definitive conclusions about the connections between these observed mutations and the useful phenotype of the LAD4 strain when transformed with pTRKH3 plasmid, but several themes have emerged, suggesting targets for further inquiry. Three of the LAD4-specific mutations appear to have cell wall altering phenotypes (ClpX, LREU_RS04225, and AmaP), which could offer some explanation for the dramatic increase in transformation efficiency seen in this strain—altering cell wall physiology can encourage exogenous DNA uptake (Aune and Aachmann, 2010). However, this behaviour, along with LAD4’s greater PCR success, could also be explained via the mutation in the type-II restriction modification associated gene LREU_RS01045; no other restriction modification genes suffered mutations in LAD4, indicating that the endonuclease may still be functional but unable to correctly target DNA for destruction due to the LREU_RS01045 dysfunction. The deletion of ClpX would presumably slow the rate of protein degradation, potentially explaining the strain’s ability to consistently accumulate expressed exogenous protein. The IS21 mobile element occurs twice within DSM 20016, and deletion of one copy could serve to slow the process by which the strain mutates and disrupts artificially introduced DNA elements via IS21 insertion.
The majority of mutations in the novel LAD4 strain pertain to DNA replication and repair machinery, similar to those seen in E. coli long-term evolution experiments (Blount et al., 2012), where some strains with mutations in DNA replication and repair machinery (like Mfd) displayed increased adaptability to aerobic citrate utilization. However, a fluctuation assay revealed the novel strain to have no significant difference in mutation rates when compared to DSM 20016 (LAD4 rate: 1.152e-06, DSM 20016 rate: 1.354e-06, t-test p-value: 0.244).
The identification of these mutations provides a strong starting point for further investigation of LAD4’s useful phenotypic properties. Meanwhile, those useful properties are available to researchers who seek to work with a lactic acid bacterium well adapted for consistent behaviour in terms of the incorporation of heterologous DNA and the expression of heterologous proteins.
The datasets presented in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found below: https://doi.org/10.7910/DVN/5YYAXA, 1.
AD: Conceptualization, Methodology, Writing–review and editing, Data curation, Formal analysis, Investigation, and Writing–original draft. MD: Conceptualization, Data curation, Formal analysis, Methodology, Software, Validation, and Writing–review and editing. DM: Conceptualization, Funding acquisition, Methodology, Project administration, Resources, Supervision, Writing–review and editing, Data curation, and Formal analysis.
The authors declare that financial support was received for the research, authorship, and/or publication of this article. This work was supported by the Natural Sciences and Engineering Research Council (NSERC) of Canada, Discovery grant number RGPIN-2017-06795. The funder played no direct role in the research but supported a general research program on the development of approaches and tools in microbial synthetic biology. This study was partially supported by the New Frontiers in Research Fund (NFRF), Exploration grant number NFRFE-2019-00742. The funder played no direct role in the research but supported a research program on the development of engineered microbes with potential for deployment in the gastrointestinal tract.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declared that they were an editorial board member of Frontiers at the time of submission. This had no impact on the peer review process and the final decision.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fsybi.2025.1473338/full#supplementary-material
LAB, Lactic acid bacteria; GRAS, Generally regarded as safe; GI, Gastrointestinal; Ex/Em, Excitation emission; IBD, Inflammatory bowel disease; WT, Wild type; SAM, S-adenosyl-L-methionine; GGR, Global genome repair; TCR, Transcription coupled repair; IS21, Insertion sequence 21; DNA, Deoxyribose nucleic acid; DUF, Domain of unknown function; ORF, Open reading frame; CV, coefficient of variance.
Ahmad, W., Nasir, A., Sattar, F., Ashfaq, I., Chen, M. H., Hayat, A., et al. (2022). Production of bimodal molecular weight levan by a Lactobacillus reuteri isolate from fish gut. Folia Microbiol. (Praha) 67, 21–31. doi:10.1007/s12223-021-00913-w
Ahrne, S., Molin, G., and Axelsson, L. (1992). Transformation of Lactobacillus reuteri with electroporation: studies on the erythromycin resistance plasmid pLUL631. Curr. Microbiol. 24, 199–205. doi:10.1007/bf01579282
Allen, W. J., Li, Y., and Waksman, G. (2010). Bacterial DNA polymerase I. eLS. doi:10.1002/9780470015902.A0001043.PUB2
Andrews, S. (2010). FastQC: a quality control tool for high throughput sequence data. Babraham Bioinforma. Available online at: https://www.bioinformatics.babraham.ac.uk/projects/fastqc/.
Arias-Palomo, E., and Berger, J. M. (2015). An atypical AAA+ ATPase assembly controls efficient transposition through DNA remodeling and transposase recruitment. Cell 162, 860–871. doi:10.1016/j.cell.2015.07.037
Armstrong, D. R., Berrisford, J. M., Conroy, M. J., Gutmanas, A., Anyango, S., Choudhary, P., et al. (2020). PDBe: improved findability of macromolecular structure data in the PDB. Nucleic Acids Res. 48, D335-D343–D343. doi:10.1093/nar/gkz990
Aune, T. E. V., and Aachmann, F. L. (2010). Methodologies to increase the transformation efficiencies and the range of bacteria that can be transformed. Appl. Microbiol. Biotechnol. 85, 1301–1313. doi:10.1007/s00253-009-2349-1
Barrick Jeffrey (2022). Fluctuation tests. Barrick Lab. Protoc. Available online at: https://barricklab.org/twiki/bin/view/Lab/ProtocolsFluctuationTests#References.
Bermúdez-Humarán, L. G., Kharrat, P., Chatel, J. M., and Langella, P. (2011). Lactococci and lactobacilli as mucosal delivery vectors for therapeutic proteins and DNA vaccines. Microb. Cell Fact. 10, S4. doi:10.1186/1475-2859-10-S1-S4
Berthier, F., Zagorec, M., Champomier-Verg, M., Ehrlich, D., and Morel-Devillel, F. (1996). Efficient transformation of Lactobacillus sake by electroporation. Microbiol. (N Y) 142, 1273–1279.
Blount, Z. D., Barrick, J. E., Davidson, C. J., and Lenski, R. E. (2012). Genomic analysis of a key innovation in an experimental Escherichia coli population. Nature 489 (7417 489), 513–518. doi:10.1038/nature11514
Bolger, A. M., Lohse, M., and Usadel, B. (2014). Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120. doi:10.1093/bioinformatics/btu170
Burckhardt, R. M., and Escalante-Semerena, J. C. (2020). Small-molecule acetylation by GCN5-related N-acetyltransferases in bacteria. Microbiol. Mol. Biol. Rev. 84, e00090. doi:10.1128/MMBR.00090-19
Caspi, R., Billington, R., Ferrer, L., Foerster, H., Fulcher, C. A., Keseler, I. M., et al. (2016). The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of pathway/genome databases. Nucleic Acids Res. 44, D471–D480. doi:10.1093/nar/gkv1164
Consortium, T. U. (2023). UniProt: the universal protein knowledgebase in 2023. Nucleic Acids Res. 51, D523–D531. doi:10.1093/nar/gkac1052
Dam Dcm Competent E (2023). Coli | NEB. Available online at: https://international.neb.com/products/c2925-dam-dcm-competent-e-coli#Product%20Information.
Deatherage, D. E., and Barrick, J. E. (2014). Identification of mutations in laboratory evolved microbes from next-generation sequencing data using breseq. Methods Mol. Biol. 1151, 165–188. doi:10.1007/978-1-4939-0554-6_12
De Vos, W. M. (1987). Gene cloning and expression in lactic streptococci. FEMS Microbiol. Rev. 3, 281–295. doi:10.1111/j.1574-6968.1987.tb02466.x
Duggan, A., and McMillen, D. (2023). Methods for electroporation and transformation confirmation in Limosilactobacillus reuteri DSM20016. J. Vis. Exp. (196), e65463. doi:10.3791/65463
Frees, D., Savijoki, K., Varmanen, P., and Ingmer, H. (2007). Clp ATPases and ClpP proteolytic complexes regulate vital biological processes in low GC, Gram-positive bacteria. Mol. Microbiol. 63, 1285–1295. doi:10.1111/j.1365-2958.2007.05598.x
Hanawalt, P. C., and Spivak, G. (2008). Transcription-coupled DNA repair: two decades of progress and surprises. Nat. Rev. Mol. Cell Biol. 9 (12 9), 958–970. doi:10.1038/nrm2549
Hendrikx, T., Duan, Y., Wang, Y., Oh, J. H., Alexander, L. M., Huang, W., et al. (2019). Bacteria engineered to produce IL-22 in intestine induce expression of REG3G to reduce ethanol-induced liver disease in mice. Gut 68, 1504–1515. doi:10.1136/gutjnl-2018-317232
Hutkins, R. W. (2007). Microbiology and Technology of fermented foods. Microbiol. Technol. Fermented Foods 1–473. doi:10.1002/9780470277515
Jansson, P. A., Curiac, D., Lazou Ahrén, I., Hansson, F., Martinsson Niskanen, T., Sjögren, K., et al. (2019). Probiotic treatment using a mix of three Lactobacillus strains for lumbar spine bone loss in postmenopausal women: a randomised, double-blind, placebo-controlled, multicentre trial. Lancet Rheumatol. 1, e154–e162. doi:10.1016/S2665-9913(19)30068-2
Jumper, J., Evans, R., Pritzel, A., Green, T., Figurnov, M., Ronneberger, O., et al. (2021). Highly accurate protein structure prediction with AlphaFold. Nature 596 (7873 596), 583–589. doi:10.1038/s41586-021-03819-2
Lahti, L., Salonen, A., Kekkonen, R. A., Salojärvi, J., Jalanka-Tuovinen, J., Palva, A., et al. (2013). Associations between the human intestinal microbiota, Lactobacillus rhamnosus GG and serum lipids indicated by integrated analysis of high-throughput profiling data. PeerJ 2013, e32. doi:10.7717/peerj.32
Lizier, M., Sarra, P. G., Cauda, R., and Lucchini, F. (2010). Comparison of expression vectors in Lactobacillus reuteri strains. FEMS Microbiol. Lett. 308, 8–15. doi:10.1111/j.1574-6968.2010.01978.x
Lubkowicz, D., Ho, C. L., Hwang, I. Y., Yew, W. S., Lee, Y. S., and Chang, M. W. (2018). Reprogramming probiotic Lactobacillus reuteri as a biosensor for Staphylococcus aureus derived AIP-I detection. ACS Synth. Biol. 7, 1229–1237. doi:10.1021/acssynbio.8b00063
MacKenzie, D. A., Jeffers, F., Parker, M. L., Vibert-Vallet, A., Bongaerts, R. J., Roos, S., et al. (2010). Strain-specific diversity of mucus-binding proteins in the adhesion and aggregation properties of Lactobacillus reuteri. Microbiol. (N Y) 156, 3368–3378. doi:10.1099/mic.0.043265-0
Mao, N., Cubillos-Ruiz, A., Cameron, D. E., and Collins, J. J. (2018). Probiotic strains detect and suppress cholera in mice. Sci. Transl. Med. 10, eaao2586. doi:10.1126/scitranslmed.aao2586
Martin, J. L., and McMillan, F. M. (2002). SAM (dependent) I AM: the S-adenosylmethionine-dependent methyltransferase fold. Curr. Opin. Struct. Biol. 12, 783–793. doi:10.1016/s0959-440x(02)00391-3
McCallum, N., Spehar, G., Bischoff, M., and Berger-Bächi, B. (2006). Strain dependence of the cell wall-damage induced stimulon in Staphylococcus aureus. Biochimica Biophysica Acta (BBA) - General Subj. 1760, 1475–1481. doi:10.1016/j.bbagen.2006.06.008
Mu, Q., Tavella, V. J., and Luo, X. M. (2018). Role of Lactobacillus reuteri in human health and diseases. Front. Microbiol. 9, 757. doi:10.3389/fmicb.2018.00757
Müller, M., Reiß, S., Schlüter, R., Mäder, U., Beyer, A., Reiß, W., et al. (2014). Deletion of membrane-associated Asp23 leads to upregulation of cell wall stress genes in Staphylococcus aureus. Mol. Microbiol. 93, 1259–1268. doi:10.1111/mmi.12733
Nikawa, H., Makihira, S., Fukushima, H., Nishimura, H., Ozaki, Y., Ishida, K., et al. (2004). Lactobacillus reuteri in bovine milk fermented decreases the oral carriage of mutans streptococci. Int. J. Food Microbiol. 95, 219–223. doi:10.1016/j.ijfoodmicro.2004.03.006
Ortiz-Velez, L., Ortiz-Villalobos, J., Schulman, A., Oh, J. H., van Pijkeren, J. P., and Britton, R. A. (2018). Genome alterations associated with improved transformation efficiency in Lactobacillus reuteri. Microb. Cell Fact. 17, 138. doi:10.1186/s12934-018-0986-8
Pandey, S. D., and Biswas, I. (2022). Clp ATPases differentially affect natural competence development in Streptococcus mutans. Microbiologyopen 11, e1288. doi:10.1002/mbo3.1288
Paysan-Lafosse, T., Blum, M., Chuguransky, S., Grego, T., Pinto, B. L., Salazar, G. A., et al. (2023). InterPro in 2022. Nucleic Acids Res. 51, D418–D427. doi:10.1093/nar/gkac993
QIAGEN (2023). MagAttract HMW DNA handbook - QIAGEN. Available online at: https://www.qiagen.com/us/resources/resourcedetail?id=3e331b6c-0742-483b-b74f-a7be2d4c1116&lang=en.
Reuter, G. (2001). The Lactobacillus and Bifidobacterium microflora of the human intestine: composition and succession. Curr. Issues Intest. Microbiol. 2, 43–53.
Roberts, R. J., Vincze, T., Posfai, J., and Macelis, D. (2023). REBASE: a database for DNA restriction and modification: enzymes, genes and genomes. Nucleic Acids Res. 51, D629–D630. doi:10.1093/nar/gkac975
Sauer, M., Russmayer, H., Grabherr, R., Peterbauer, C. K., and Marx, H. (2017). The efficient clade: lactic acid bacteria for industrial chemical production. Trends Biotechnol. 35, 756–769. doi:10.1016/j.tibtech.2017.05.002
Spínola-Amilibia, M., Araújo-Bazán, L., de la Gándara, Á., Berger, J. M., and Arias-Palomo, E. (2023). IS21 family transposase cleaved donor complex traps two right-handed superhelical crossings. Nat. Commun. 14, 2335. doi:10.1038/s41467-023-38071-x
Team Oxford (2019). Oxford IGEM team/results - 2019. Available online at: https://2019.igem.org/Team:Oxford/Results.
Thomas, P. D., Ebert, D., Muruganujan, A., Mushayahama, T., Albou, L. P., and Mi, H. (2022). PANTHER: making genome-scale phylogenetics accessible to all. Protein Sci. 31, 8–22. doi:10.1002/pro.4218
Urbańska, M., Gieruszczak-Białek, D., and Szajewska, H. (2016). Systematic review with meta-analysis: Lactobacillus reuteri DSM 17938 for diarrhoeal diseases in children. Alimentary Pharmacol. Ther. 43, 1025–1034. doi:10.1111/apt.13590
Walter, J., Britton, R. A., and Roos, S. (2011). Host-microbial symbiosis in the vertebrate gastrointestinal tract and the Lactobacillus reuteri paradigm. Proc. Natl. Acad. Sci. U. S. A. 108, 4645–4652. doi:10.1073/pnas.1000099107
Wang, C., Nagata, S., Asahara, T., Yuki, N., Matsuda, K., Tsuji, H., et al. (2015). Intestinal microbiota profiles of healthy pre-school and school-age children and effects of probiotic supplementation. Ann. Nutr. Metab. 67, 257–266. doi:10.1159/000441066
Yang, C., Liang, L., Lv, P., Liu, L., Wang, S., Wang, Z., et al. (2021). Effects of non-viable Lactobacillus reuteri combining with 14-day standard triple therapy on Helicobacter pylori eradication: a randomized double-blind placebo-controlled trial. Helicobacter 26, e12856. doi:10.1111/hel.12856
Yang, S., Liu, Q., Zhang, Y., Du, G., Chen, J., and Kang, Z. (2018). Construction and characterization of broad-spectrum promoters for synthetic biology. ACS Synth. Biol. 7, 287–291. doi:10.1021/acssynbio.7b00258
Zhang, Y., Zagnitko, O., Rodionova, I., Osterman, A., and Godzik, A. (2011). The FGGY carbohydrate kinase family: insights into the evolution of functional specificities. PLoS Comput. Biol. 7, e1002318. doi:10.1371/journal.pcbi.1002318
Keywords: Limosilactobacillus reuteri, Lactobacillus reuteri, DSM 20016, DSM 116333, transformation, heterologous protein production, non-model organisms
Citation: Duggan AD, Dillon MM and McMillen DR (2025) A promising novel strain of L. reuteri DSM20016 as a chassis for synthetic biology applications. Front. Synth. Biol. 3:1473338. doi: 10.3389/fsybi.2025.1473338
Received: 30 July 2024; Accepted: 24 February 2025;
Published: 26 March 2025.
Edited by:
Thomas Mansell, Iowa State University, United StatesReviewed by:
Lichao Sun, Beijing Institute of Technology, ChinaCopyright © 2025 Duggan, Dillon and McMillen. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: David R. McMillen, ZGF2aWQubWNtaWxsZW5AdXRvcm9udG8uY2E=
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.