
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
REVIEW article
Front. Plant Sci. , 25 March 2025
Sec. Plant Abiotic Stress
Volume 16 - 2025 | https://doi.org/10.3389/fpls.2025.1560204
This article is part of the Research Topic Plant Ecophysiology: Responses to Climate Changes and Stress Conditions View all 31 articles
Global climate change, characterized by increased frequency and intensity of extreme temperature events, poses significant challenges to plant survival and crop productivity. While considerable research has elucidated plant responses to temperature stress, the molecular mechanisms, particularly those involved in temperature sensing, remain incompletely understood. Thermosensors in plants play a crucial role in translating temperature signals into cellular responses, initiating the downstream signaling cascades that govern adaptive processes. This review highlights recent advances in the identification and classification of plant thermosensors, exploring their physiological roles and the biochemical mechanisms by which they sense temperature changes. We also address the challenges in thermosensor discovery and discuss emerging strategies to uncover novel thermosensory mechanisms, with implications for improving plant resilience to temperature stress in the face of a rapidly changing climate.
Plants, as sessile organisms, are constantly exposed to a range of environmental stress, including heat, cold, salinity and drought. These abiotic stresses are major limiting crop growth, productivity, and quality, contributing significantly to global food insecurity (He et al., 2018; Sivaji, 2021). With the escalating effects of climate change, heat stress has emerged as one of the most pressing challenges to agricultural productivity (Janni et al., 2020). For instance, studies predict that wheat production could decrease by 6% for each degree Celsius increase in global temperature (Asseng et al., 2015; Zaveri and D, 2019). Moreover, in crops like cereals, peas, lentils, and chickpeas, even brief episodes of heat stress (above 24°C) during the reproductive phase can negatively impact floral fertility, while sustained temperatures of 35°C or higher can result in total crop failure. The impact of heat stress on crop yield varies across species and geographic regions, with reductions ranging from 40% to 85% (Janni et al., 2020).
Unpredicted variability in temperature is also associated with frequent extreme low-temperature events. There is evidence that the Arctic region has warmed more than twice as fast as the global average, a phenomenon referred to as Arctic amplification (Cohen et al., 2014). This accelerated warming has led to significant reductions in Arctic sea ice and spring snow cover, which coincides with a period of ostensibly more frequent extreme weather events across the Northern Hemisphere mid-latitudes, including severe winters (Cohen et al., 2014). Low temperatures pose a widespread environmental stress that significantly inhibits agricultural productivity worldwide by impeding plant growth and development (Mishra et al., 2019). For example, temperatures below 15°C can impair soybean’s growth and development, leading to potential yield loss (Gass et al., 1996). Cold stress also damages flag leaves and spikes, affects the grain number per spike and grain filling rate, leading to a substantial reduction in final wheat production (Fuller et al., 2007; Thakur et al., 2010). A research investigation carried out by the International Rice Research Institute demonstrated a 10% reduction in rice grain yields per 1°C rise in the minimum temperature during the dry season’s growing period (Peng et al., 2004). Given these challenges, understanding the genetic, molecular, and physiological components involved in temperature stress sensing and response is critical for advancing plant biology and improving crop resilience.
Plants have developed intricate mechanisms to perceive environmental temperature signals, regulate growth and reproduction, and even store temperature memory to optimize agronomic traits (Ding and Yang, 2022). Numerous studies have explored plant temperature sensing, responses, and associated signaling pathways. In particular, the identity of plant thermosensors located at the start of the signaling pathways has long been sought (Noguchi and Kodama, 2022). As early as the year 2000, Suzuki and colleagues identified the cyanobacterium histidine kinase 33 (Hik33) is the thermosensor that regulates desaturase gene expression in response to temperature downshifts (Suzuki et al., 2000). The identification of plant thermosensors has gained significant attention since the discovery of CHILLING TOLERANCE DIVERGENCE 1 (COLD1) in 2015 (Ma et al., 2015). COLD1 has been identified as a key player in sensing cold temperatures, as demonstrated by Ma et al. (2015). The study suggests that COLD1 may initiate cold signaling through its physical interaction with RGA1, leading to Ca2+ influx into the cytoplasm, which triggers downstream responses to chilling stress. In contrast, phytochrome B (phyB) functions as a thermosensor, primarily sensing warm temperatures (Jung et al., 2016; Legris et al., 2016). phyB undergoes conformational changes in sensing temperature fluctuations, typically within the range of 15–30°C, thereby regulating plant growth and development in accordance with environmental conditions (Jung et al., 2016; Legris et al., 2016). While significant progress has been made in identifying and characterizing plant thermosensors like COLD1 and phyB, it is important to clarify the precise roles of these proteins to avoid misinterpretation of their functions. However, the term “thermosensor” is occasionally misused. A typical example is H2A.Z, which plays roles in both transcriptional activation and repression. Although H2A.Z is rapidly displaced from nucleosomes in response to elevated temperature (Kumar and Wigge, 2010; Cortijo et al., 2017), this displacement is not a direct thermosensing mechanism. Cortijo et al. (2017) demonstrated that H2A.Z eviction does not occur at high temperatures in vitro or in reconstituted nucleosome systems. Instead, the rapid eviction of H2A.Z at the +1 position under high-temperature conditions depends on the recruitment of heat shock factor A1a (HSFA1a) and potentially other HSFA1 transcription factors to heat shock elements near transcription start sites (Cortijo et al., 2017). Additionally, the deacetylation of histone H3, regulated by POWERDRESS (PWR) and HISTONE DEACETYLASE 9 (HDA9), regulates genes that are also modulated by H2A.Z dynamics, suggesting a potential role for these factors in regulating H2A.Z deposition in plants (Tasset et al., 2018). Together, these findings highlight the need to study how upstream factors such as HSFA1a, PWR, and HDA9 are regulated by temperature, in order to better understand the thermosensing mechanisms that control H2A.Z’s response to temperature changes. To promote more rigorous research on plant thermosensors, several criteria have been proposed (Vu et al., 2019): (1) temperature fluctuations directly alter the activity or conformation of the sensor, (2) these changes are crucial for the sensor to interpret and transmit the temperature stimulus, and (3) the sensor’s ability to detect temperature has a direct effect on the plant’s temperature responses.
From a thermodynamic standpoint, temperature changes can disrupt the equilibrium of biological systems (Reynolds et al., 1992). In nature, temperature fluctuations can have profound effects on living cells by altering the properties and activities of biomolecules such as nucleic acids, proteins, and lipids. Many organisms, including plants, have evolved thermosensors that exploit these properties. In microorganisms and animals, thermosensors are generally classified into four categories based on their chemical nature: DNA, RNA, protein, and plasma membrane-associated protein-based sensors (Sengupta and Garrity, 2013). DNA and RNA thermosensors sense temperature changes through alterations in their secondary and tertiary structures. For example, DNA can bend at low temperatures, and melting of this bent structure promotes transcription as temperature rises (Falconi et al., 1998). In Shigella, the DNA bend of the virF gene relaxes, releasing the histone-like nucleoid-structuring repressor and enabling transcription factors to bind, thereby activating virF gene transcription at temperatures above 32°C (Falconi et al., 1998). Similarly, RNA molecules may form stem-loop structures that inhibit translation, which is disrupted at higher temperatures, allowing translation to proceed (Cortijo et al., 2017; Chung et al., 2023). Protein and membrane-associated protein thermosensors undergo conformational changes or alterations in activity in response to temperature shifts (D. Zhang et al., 2019; Chung et al., 2023). In B. subtilis, the membrane-associated histidine kinase DesK, which senses temperature-dependent changes in membrane thickness and activates a pathway to restore membrane fluidity at low temperature by regulating the expression of genes involved in membrane adaptation (Inda et al., 2016).
So far, three major types of thermosensors in plants have been identified: RNA-based thermosensors, protein-based thermosensors, and plasma membrane-associated protein-based thermosensors, including photoreceptors, Ca2+-permeable channels, and other proteins. This review summarizes recent findings on plant thermosensors, drawing comparisons with thermosensors in other organisms. Understanding the molecular mechanisms underlying plant temperature sensing and response will provide critical insights for the genetic engineering of temperature-stress-tolerant crops, helping mitigate the impact of climate change on global crop yields.
Ribonucleic acid (RNA) is a single-stranded nucleic acid molecule and made up of ribonucleotides. RNA polymerase synthesizes RNA from DNA that is functional for protein-coding (messenger RNA, mRNA) or non-coding RNA genes. Because of these functions, RNA molecules are of the following types: mRNA, rRNA, tRNA, snRNA, snoRNA, miRNA, and lncRNA (Cech and Steitz, 2014). RNA molecules are capable of adopting secondary and tertiary structures that are highly sensitive to temperature, allowing them to function as thermosensors. These structures, which are energetically favorable at specific temperatures, regulate RNA stability, pre-mRNA splicing, and translation efficiency, all of which are influenced by temperature changes (Mizushima et al., 1997). Typically, RNA secondary structures are stabilized at lower temperatures and disrupted at higher temperatures, which in turn affects ribosome binding and translation efficiency (Chełkowska-Pauszek et al., 2021; Georgakopoulos-Soares et al., 2022). Moreover, the 5′-untranslated regions (UTRs) of mRNAs may harbor thermolabile stem-loop structures that exhibit temperature-dependent conformational changes, which in turn modulate translation (Di Martino et al., 2016). Recent studies have demonstrated that lncRNA FLINC is down-regulated at higher ambient temperature and affects ambient temperature-mediated flowering in Arabidopsis (Severing et al., 2018). Additionally, miR156 was highly upregulated, while squamosa promoter binding protein-like (SPL) was downregulated, which further induced FLOWERING LOCUST (FT) and FRUITFULL expression in Arabidopsis under heat stress (Kim et al., 2012; Stief et al., 2014). Therefore, it is speculated that non-coding RNAs could also be used as a thermoresensor to mediate plant responses to temperature fluctuations.
RNA-based thermosensors offer the advantage of rapid response to temperature fluctuations due to their direct conformational changes, enabling swift modulation of downstream signaling pathways. Previous studies have shown that some RNA thermosensors, which control heat shock and virulence genes, function like molecular zippers, reversibly opening and closing in response to ambient temperature shifts (Falconi et al., 1998). These RNA-based mechanisms have been observed in bacteria, and similar temperature-sensitive regulation is found in eukaryotes. For instance, mRNA 5′-UTR secondary structures are implicated in heat shock responses in Drosophila (Coleman-Derr and Zilberman, 2012) and Trypanosomes (Kumar and Wigge, 2010). In plants, several RNA thermosensors have been proposed to modulate responses to heat stress and other temperature-related environmental cues (Table 1).
A thermosensor based on secondary structure changes in the PHYTOCHROME-INTERACTING FACTOR 7 (PIF7) mRNA has been identified in Arabidopsis (Chung et al., 2020). A stem-loop structure within the 5′-untranslated region (UTR) of PIF7 mRNA undergoes conformational shifts in response to temperature. At low temperatures, the AUG-proximal hairpin structure is more compact, while at higher temperatures, the hairpin structure relaxes and partially unfolds, enhancing PIF7 mRNA translation (Figure 1A). This reversible conformational change results in the accumulation of PIF7 proteins, which activate the thermomorphogenesis pathway by inducing the transcription of key genes, such as the auxin biosynthesis gene YUCCA8, promoting hypocotyl and petiole elongation. Consequently, PIF7 mRNA functions as an RNA-based thermosensor that mediates plant responses to temperature fluctuations (Chung et al., 2020).
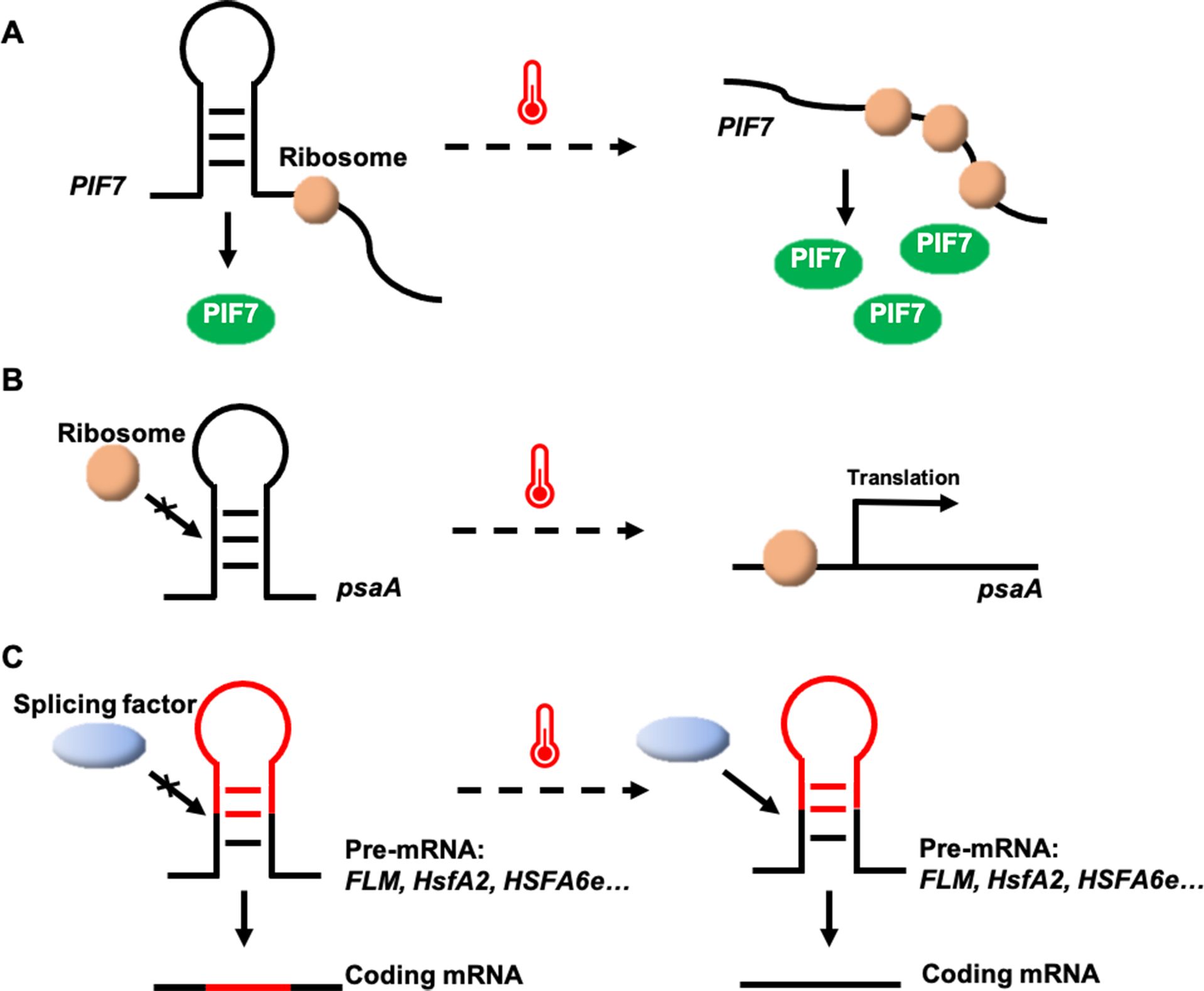
Figure 1. Potential thermosensing mechanisms in RNA thermosensors. (A) The heat-sensitive hairpin structure in the PIF7 mRNA 5′-UTR becomes a looser conformation at high temperatures, enhancing protein translation. (B) psaA mRNA senses temperature rising to unfold the hairpin structure within its 5′-UTR and facilitates psaA translation. (C) Temperature changes may affect alternative splicing by altering RNA secondary structure.
Structured RNAs with fundamental sensory and regulatory potential have been discovered in all kingdoms of life (Waldminghaus et al., 2008). RNA thermosensors have been identified not only in plants, but also in algae (K. P. Chung et al., 2023). In the unicellular green alga Chlamydomonas reinhardtii, an RNA thermosensor controls translation of the chloroplastic Photosystem I P700 chlorophyll a apoprotein A1 (psaA) mRNA (Chung et al., 2023). The 5′-UTR of psaA forms a hairpin-type secondary structure that masks the Shine-Dalgarno sequence (the prokaryotic ribosome-binding site) at 25°C, but melts at elevated temperatures of 40°C, increasing accessibility of the Shine-Dalgarno sequence, initiating and enhancing protein synthesis (Figure 1B). It is a valuable tool for inducible transgene expression from the Chlamydomonas plastid genome, in that a simple temperature shift of the algal culture can greatly increase recombinant protein yields (Chung et al., 2023). Utilizing the temperature-dependent regulatory mechanism of the 5′-UTR of the psaA gene, it is possible to introduce this element into crop genomes through genetic engineering to enhance crop resilience under high-temperature conditions.
In Arabidopsis, the MADS-box transcription factor gene FLOWERING LOCUS M (FLM), a key component of the thermosensory flowering time pathway, undergoes temperature-dependent alternative splicing (Posé et al., 2013; Airoldi et al., 2015). FLM produces two primary isoforms, FLM-β and FLM-δ, which differ in the presence of exon 2 or exon 3 (Posé et al., 2013). The overexpression of these variants has opposite effects on flowering, leading to late flowering (overexpression of FLM-δ) or early flowering (overexpression of FLM-β) (Capovilla et al., 2017). Both FLM-β and FLM-δ proteins compete for interaction with the floral repressor SHORT VEGETATIVE PHASE (SVP) to form an SVP-FLM heterocomplex to regulate the transcription of flowering–related genes, such as FLOWERING LOCUS T (FT) and SUPPRESSOR OF OVEREXPRESSION OF CONSTANS1 (SOC1) (Posé et al., 2013). FLM-β transcript levels decrease and FLM-δ transcript levels increase at high temperatures (Figure 1C). At low temperatures, the SVP-FLM-β complex predominates, actively repressing flowering. However, at elevated temperatures, the SVP-FLM-δ complex becomes more prevalent, acting as a dominant-negative activator of flowering (Posé et al., 2013; Sureshkumar et al., 2016; Lutz et al., 2017). Although it is not yet known how temperature regulates the alternative splicing of FLM, this splicing regulation pathway enables the plant to rapidly sense and respond to fluctuations in ambient temperature, facilitating adaptive flowering time responses.
In Arabidopsis, temperature variations also trigger alternative splicing of the heat shock transcription factor A2 (HsfA2) gene, producing different mRNA isoforms (Liu et al., 2013). HsfA2 contains only one 324-nucleotide intron, which is able to splice normally to form a full-length protein at 22°C. Under severe heat stress, a 5′-splice site in the HsfA2 pre-mRNA is activated, leading to the production of HsfA2-III (Figure 1C), a truncated isoform, S-HsfA2, which binds to the heat shock element (HSE) in the HsfA2 promoter and activates its own transcription, enhancing heat tolerance (Liu et al., 2013). Similar alternative splicing of HsfA2 has been observed in tomato. A study comparing wild relatives and domesticated tomato species revealed that efficient pre-mRNA splicing in wild species leads to an accumulation of HsfA2-II and suppression of HsfA2-I, thereby promoting better adaptation to heat stress (Hu et al., 2020). Further studies have shown that temperature-dependent structural changes at the 3′-splice site of the intron regulate the alternative splicing of HsfA2 pre-mRNA, suggesting that the secondary structures at this site mediate splicing in response to heat (Broft et al., 2022).
In wheat, the heat shock transcription factor gene TaHSFA6e also undergoes alternative splicing in response to heat stress, generating two major functional transcripts: TaHSFA6e-II and TaHSFA6e-III (Figure 1C) (Wen et al., 2023). TaHSFA6e-III contains a 14-amino acid peptide at its C-terminal (AHA motif), produced by alternative splicing, which enhances the transcriptional activity of three downstream heat shock protein 70 (TaHSP70) genes more effectively than TaHSFA6e-II, improving thermotolerance (Wen et al., 2023). This study highlights a heat-responsive pathway in wheat, where alternative splicing of TaHSFA6e regulates the transcriptional efficiency of heat shock proteins, which are localized in stress granules and play a role in translation re-initiation under heat stress conditions (Wen et al., 2023).
Although plants perceive temperature changes through the modulation of alternative splicing of mRNAs (as exemplified by FLM, HsfA2, and HSF6e), the ultimate transduction of temperature signals occurs through the translation of mRNA into proteins.
Current evidence reveals that HSFA9 belonging to class HSFA acts as an important hub in mediating embryogenesis, germination, photomorphogenesis, and stress protection (Kotak et al., 2007; Zinsmeister et al., 2020; Wang et al., 2024). Recent studies have identified that the RtHSFA9 genes make crucial contributions to the thermal adaption of R. tomentosa by positively regulating the RtHSFA2a, RtHSFA2b, and RtHSP genes (Li et al., 2024). RtHSFA9a dramatically enhances plant heat stress tolerance by positively regulating the transcription of RtHSFA2a, RtHSFA2b, and some RtHSPs, such as RtHSP21.8 and RtHSP70. RtHSFA9b and RtHSFA9c can activate the expression of RtHSFA2b and some RtHSP genes, consequently taking part in thermal adaption in R. tomentosa. In comparison with RtHSFA9b, RtHSFA9c has higher transcription activity in regulating RtHSFA2b and RtHSP genes and therefore confers promising thermotolerance to plants (Li et al., 2024). However, the mechanisms by which RtHSFA9s perceive accept temperature signals and the subsequent effects of temperature on RtHSFA9s remain to be elucidated.
Proteins, as complex biomolecules composed of thousands of interacting atoms, are inherently sensitive to temperature changes, similar to nucleic acids. Temperature fluctuations can induce significant conformational alterations in proteins, enabling them to regulate various cellular processes, including enzymatic reactions and protein-protein interactions, in response to environmental temperature shifts (Peterson et al., 2007). These temperature-dependent changes in protein structure can also affect the assembly and disassembly of protein complexes, thus modulating cellular functions. Additionally, temperature-induced alterations in protein localization serve as another mechanism through which temperature signals are transduced within the cell (Somero, 1995). In plants, a variety of protein-based and protein-membrane-based thermosensors have been identified, highlighting the importance of protein conformational changes and interactions in plant responses to temperature stress (Table 2).
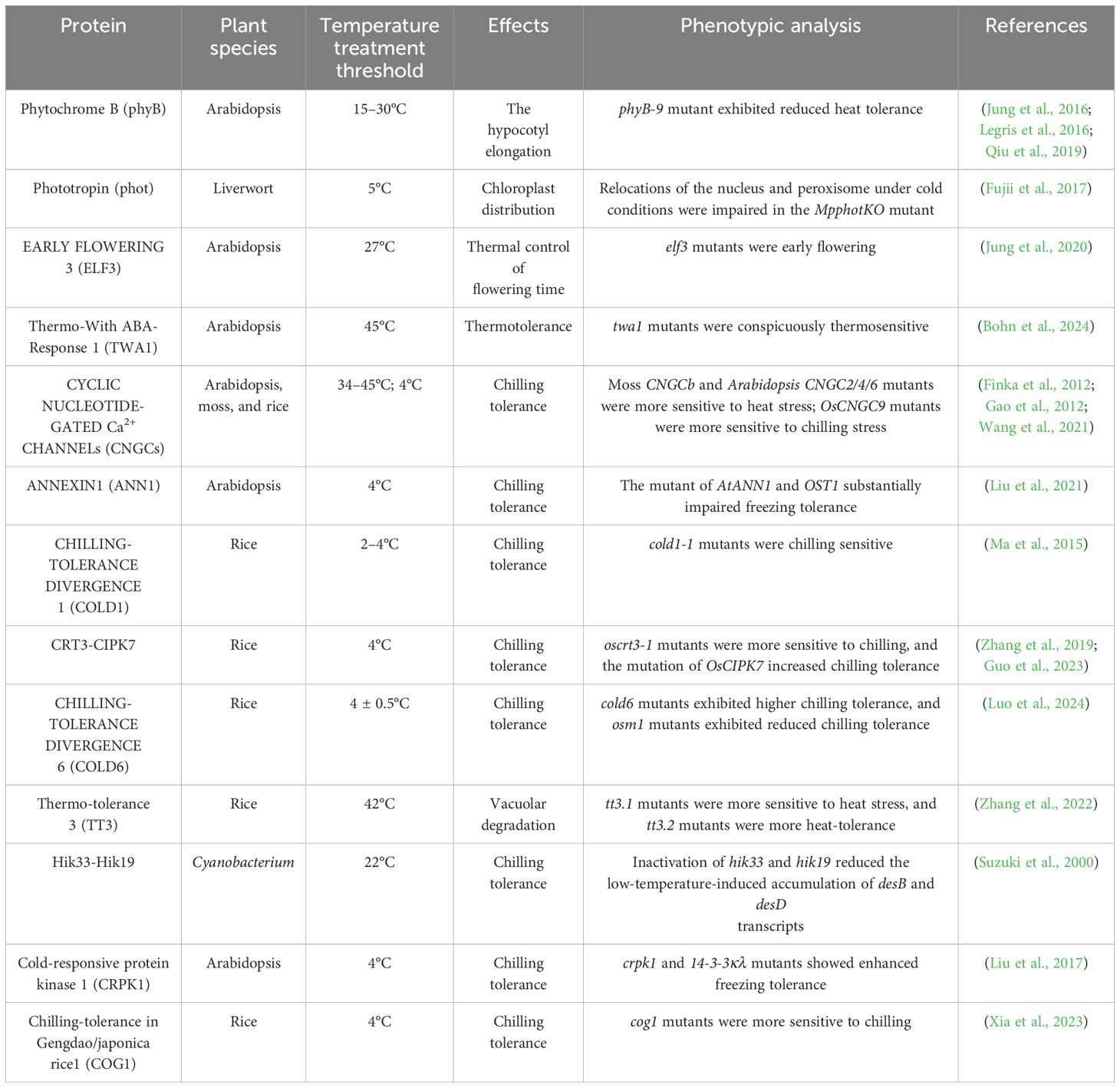
Table 2. List of various protein and plasma membrane-associated protein-based thermosensors discussed in this review. .
In natural environments, plants sensing temperature changes are closely related to sunlight. Light and temperature in plants are perceived through a common receptor, phytochrome B (phyB) (Jung et al., 2016; Legris et al., 2016). phyB exists in two states: a red-light-absorbing, biologically inactive Pr state, and a far-red-light-absorbing, biologically active Pfr state (Quail et al., 1995; Burgie and Vierstra, 2014). PhyB participates in temperature perception through its temperature-dependent reversion from the active Pfr state to the inactive Pr state. Increased rates of thermal reversion to warmer environments reduce both the abundance of the biologically active Pfr-Pfr dimer pool of phyB and the size of the associated nuclear bodies, which releases the repression of phyB on the PHYTOCHROME-INTERACTING FACTOR 4 (PIF4) (Figure 2A). As a result, PIF4 accumulates and promotes hypocotyl elongation by enhancing the expression of auxin biosynthesis genes (Jung et al., 2016; Legris et al., 2016; Qiu et al., 2019). Although there are five phytochrome isoforms (phyA–E) in Arabidopsis, thermal reversion is an order of magnitude faster in phyB than in the others (Burgie et al., 2021). This feature is also found in potato and maize, suggesting that phyB serves as the primary thermosensor among the phytochrome family in many plant species (Burgie et al., 2021; Noguchi and Kodama, 2022).
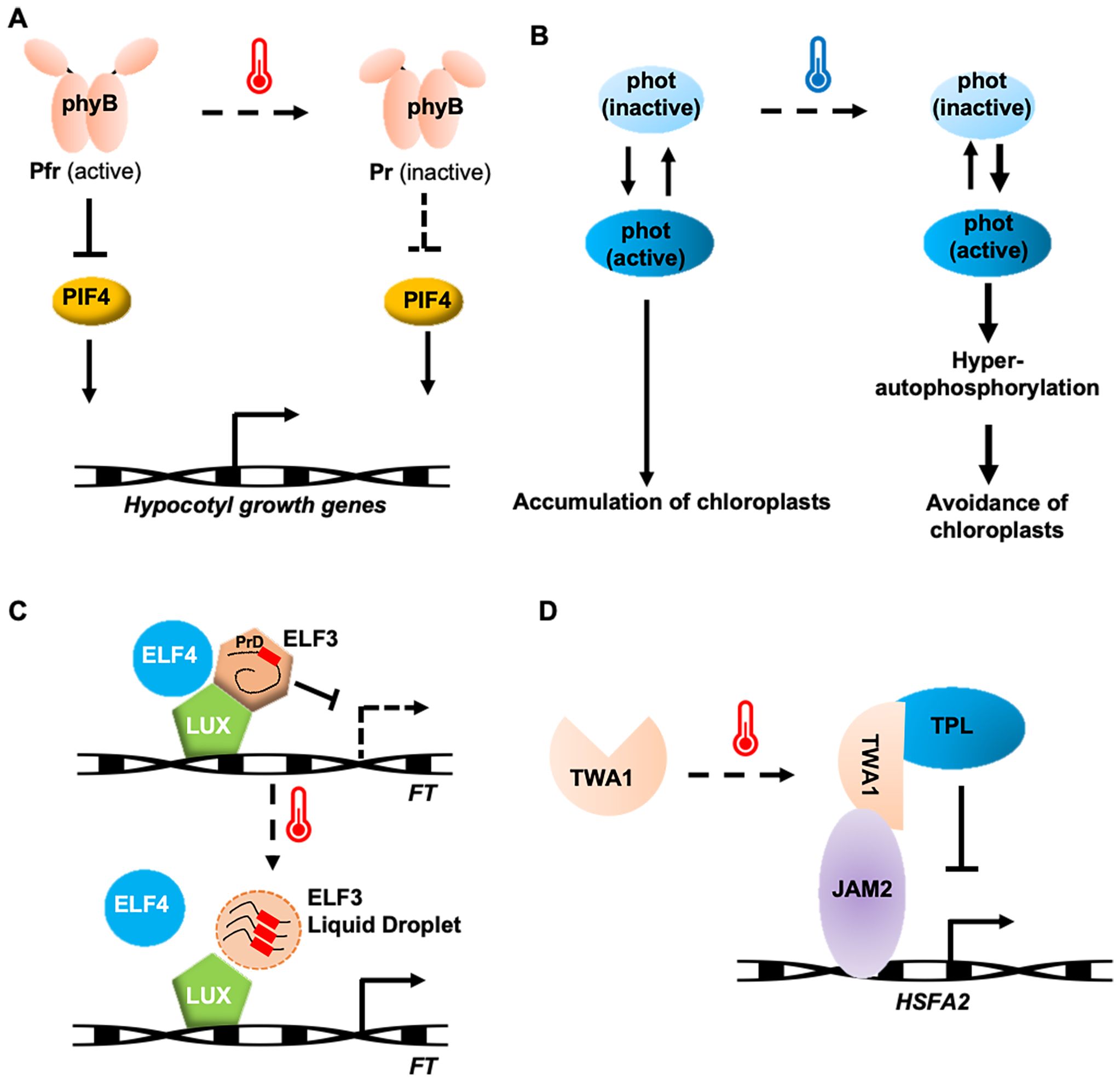
Figure 2. Potential thermosensing mechanisms in protein thermosensors. (A) phyB involved in Arabidopsis temperature perception and heat-tolerance formation. (B) A phototropin in liverwort functions as thermosensors by preventing the inactivation of its active forms at low temperature. (C) ELF3 responds to temperature by phase separation. (D) TWA1-mediated transcriptional repression by interacting with JAM2 and TPL. Arrows indicate positive regulation and T-bars indicate negative regulation.
phyB is also involved in the cold stress response. The key transcription factors mediating cold acclimation, C-REPEAT BINDING FACTORS (CBFs), are rapidly induced by cold stress and bind to CRT/DRE DNA regulatory elements in the promoters of a subset of cold-regulated (COR) genes; expression of these COR genes renders plants able to tolerate freezing stress (Jaglo-Ottosen et al., 1998; Gilmour et al., 2000). CBFs could interact with PIF3 under cold stress, which attenuates the mutual degradation of the PIF3-phyB complex. Cold-stabilized phyB acts downstream of CBFs to positively regulate freezing tolerance by modulating the expression of stress-responsive genes as well as growth-related genes (Jiang et al., 2020).
Phototropin (phot) is a blue light receptor involved in processes such as phototropism, stomatal opening, and chloroplast repositioning (Christie, 2007), and is the only reported thermal sensor of cold responses in plants to date. Phototropin contains two light-sensitive LOV (Light, Oxygen, or Voltage) domains at the N-terminus and a serine/threonine kinase domain at the C-terminus. The LOV domain exists in both active and inactive forms, determined by whether it combines with FMN (flavin mononucleotide). Under blue light, a covalent cysteine bridge forms between LOV2 and FMN, converting the inactive form into the active form. Upon activation, the phototropin protein undergoes a conformational change, and its kinase domain undergoes subsequent autophosphorylation. Similar to phyB, phototropins also undergo thermal reversion from an active to inactive form, but this process slows down at lower temperatures. This results in the accumulation of the active form of LOV2, increasing autophosphorylation, which determines the accumulation response or cold-avoidance response in liverwort (Figure 2B). Phototropin perceives both blue light and temperature, using this information to optimize chloroplast positioning for efficient photosynthesis (Fujii et al., 2017).
Like phototropin and phytochromes in plants, other photoreceptors (e.g., blue-light receptors cryptochromes and UV Resistance Locus 8) also display temperature-dependent lifetimes (Findlay and Jenkins, 2016; Pooam et al., 2021). Therefore, photoreceptors like phyB has already been demonstrated to act as thermosensors (Li and Song, 2024).
Temperature changes are part of the plant’s circadian clock perception. The evening complex, consisting of EARLY FLOWERING 3 (ELF3), a large scaffold protein, ELF4, a small α-helical protein, and LUX ARRYTHMO (LUX), a DNA-binding protein, plays a key role in temperature sensing (Box et al., 2015; Nieto et al., 2015). ELF3 contains a PrD domain with a high proportion of polyQ, which varies in length among species and serves as a tunable thermosensor. Increasing the polyQ length enhances thermal responsiveness (Ezer et al., 2017). At 22°C, ELF3 diffuses within the cell and binds to DNA, inhibiting transcription. At 27°C, ELF3 aggregates into a punctate pattern, undergoing phase separation, which prevents ELF3 from binding to flowering genes, thus allowing these genes to be expressed and promoting growth and flowering (Figure 2C) (Jung et al., 2020). The temperature sensitivity of ELF3 is modulated by ELF4, suggesting that ELF4 stabilizes ELF3’s function. ELF3 undergoes reversible phase transition through liquid-liquid phase separation (LLPS), making it an ideal temperature sensor (Jung et al., 2020).
A recently identified temperature-sensing transcriptional co-regulator in Arabidopsis, THERMO-WITH ABA-RESPONSE 1 (TWA1), is predicted to be an intrinsically disordered protein with a key thermosensory role, functioning through its amino-terminal highly variable region (Bohn et al., 2024). At elevated temperatures, TWA1 accumulates in nuclear subdomains and undergoes conformational changes, allowing it to interact with JASMONATE-ASSOCIATED MYC-LIKE (JAM) transcription factors and TOPLESS (TPL) and TOPLESS-RELATED (TPR) proteins to form repressor complexes (Figure 2D). The transcriptional upregulation of HSFA2 and heat shock proteins depends on TWA1, with orthologues of TWA1 providing different temperature thresholds, consistent with its role in early heat stress signaling (Bohn et al., 2024). Interestingly, specific amino acid changes in the amino-terminal region of TWA1 orthologues are linked to different temperature thresholds. Overexpression of TWA1 could improve crop heat tolerance without negatively affecting plant growth or yield (Bohn et al., 2024), highlighting its potential for application in crop breeding strategies.
The plasma membrane, which is one of the most thermally sensitive macromolecular structures in the cell, has long been proposed as a primary candidate for temperature sensing in plants (Balogh et al., 2013; Niu and Xiang, 2018). Subtle changes in temperature can affect various properties of cellular membranes, including fluidity, thickness, permeability, and packing, and thus affect the clustering of important membrane proteins, which are sensitive to ambient conditions (Escribá et al., 2008; Niu and Xiang, 2018). Although membrane lipids lack catalytic ability on their own, changes in their physical state strongly affect the folding, mobility, and activity of integral or membrane-associated proteins (Hayes et al., 2021; Kerbler and Wigge, 2023). To date, several plasma membrane-associated proteins have been proposed as potential thermosensors, including CNGCs, ANN1, COLD1, CRT3-CIPK7, COLD6, TT3, Hik33-Hik19, CRPK1, and COG1 (Table 2).
Extreme temperature changes can negatively affect membrane fluidity, with cold stress reducing fluidity and heat stress increasing it (Sangwan et al., 2002). Temperature fluctuations are first sensed by plasma membrane CYCLIC NUCLEOTIDE-GATED Ca2+ CHANNELs (CNGCs), which encode components of the membrane’s cyclic nucleotide-gated Ca2+ channels (Falcone et al., 2004). In plants, CNGCs function in signaling pathways that may be tied to their ability to transport Ca2+ rather than other cations into plant cells (Jha et al., 2016). It has been reported that CNGCb gene from Physcomitrella patens and its Arabidopsis thaliana ortholog CNGC2/4 act as the primary thermosensors of land plant cells (Finka et al., 2012). Additionally, AtCNGC6 also mediates heat-induced Ca2+ influx (Gao et al., 2012). Under heat conditions, the channels quickly open, allowing periplasmic Ca2+ to enter and bind calmodulins (CaMs) associated with the cytosolic C-terminal domain of the CNGCs (Niu et al., 2020). Ca2+ binding to CNGC-bound CaMs initiates a specific signaling cascade that activates kinases, which then phosphorylate and activate heat shock transcription factor A (HSFA), promoting the expression of HSP genes (Figure 3) (Guo et al., 2016; Ohama et al., 2016; Guihur et al., 2022).
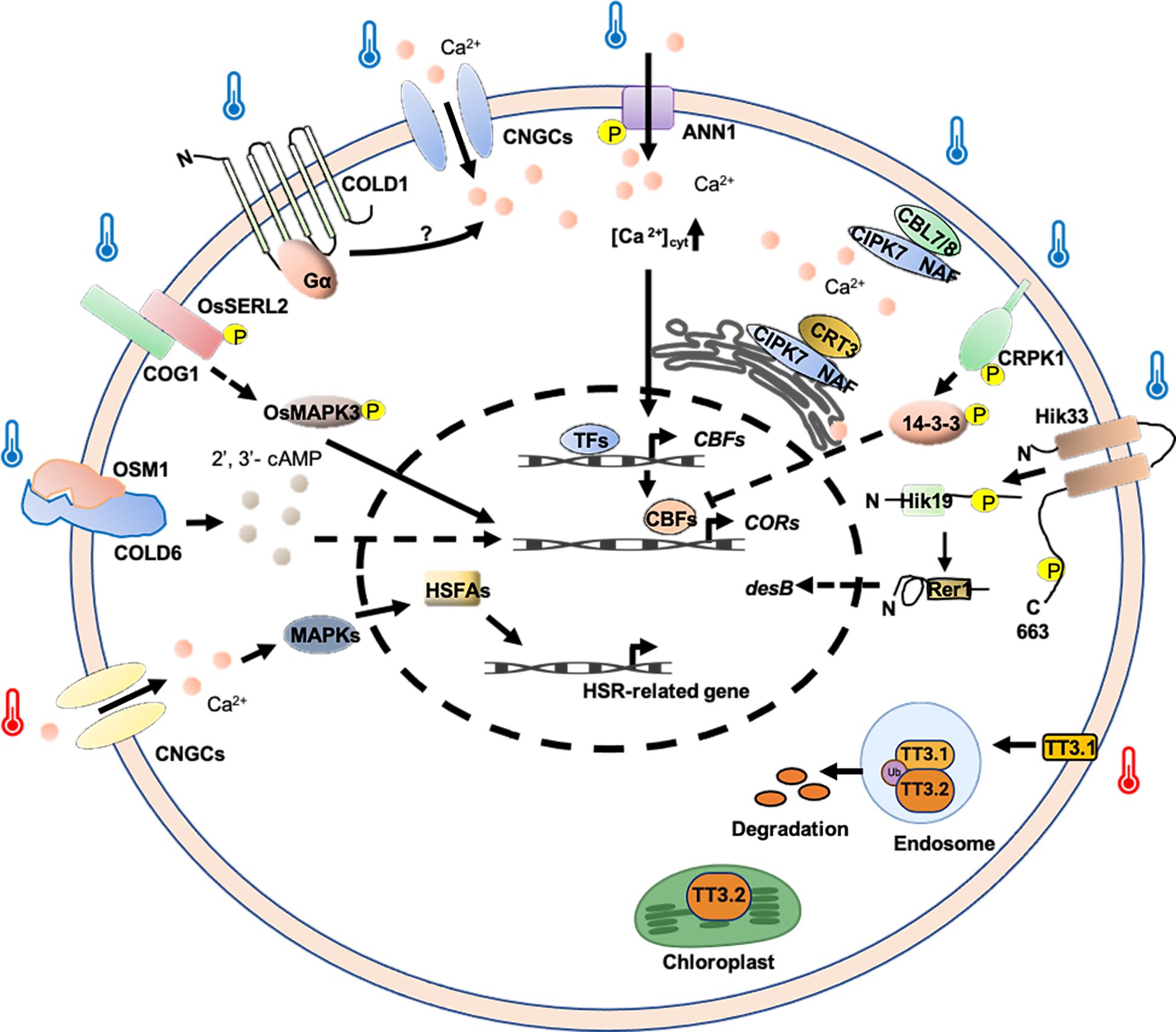
Figure 3. Potential thermosensing mechanisms in plasma membrane-associated protein-based thermosensors. Ca2+ and 2′, 3′-cAMP are important molecules involved in regulation temperature-stress response. Cold and heat-induced Ca2+ signatures may be decoded by Ca2+ sensors or Ca2+ -related proteins and thus regulate COR and HSR gene expression. TT3.1 senses high temperature to translocate to the endosomes, where it degrades protein TT3.2. TT3.2 degradation boosts chloroplast function at high temperatures. COLD6 interacted with cold-induced OSM1 to trigger an increase in the level of 2’, 3’-cAMP to promote chilling tolerance. TT3.1 proteins translocate from the PM to endosomes, ubiquitinating the chloroplast precursor protein TT3.2 to prevent chloroplast thylakoid damage and improving heat tolerance. Hik33 kinase domain might be phosphorylated under cold conditions, and then transferred to Hik19, and finally to Rer1, regulating the expression of the desB to adapt to cold stress. CRPK1 was activated by cold stress, phosphorylating 14-3-3 proteins and triggering 14-3-3 proteins to translocate into the nucleus to attenuate the CBF signaling. The COG1-OsSERL2 complex causes the activation of OsMAPK3 to transmit cold signal from the membrane to the cytoplasm, enhancing cold tolerance. Arrows indicate positive regulation and T-bars indicate negative regulation.
In rice, a loss-of-function mutant of OsCNGC9, named cds1 (cell death and susceptible to blast 1), is more sensitive to chilling stress (Wang et al., 2021). In this case, OsCNGC9 is phosphorylated and activated by OsSAPK8, a member of sucrose non-fermenting 1-related protein kinases (SnRK) family, triggering Ca2+ influx and the activation of cold stress-related genes. Furthermore, OsDREB1A, a rice dehydration-responsive element-binding transcription factor, positively regulates the transcriptional expression of OsCNGC9. Thus, OsSAPK8-mediated OsCNGC9 phosphorylation and OsDREB1A-mediated OsCNGC9 expression form a potential positive feedback loop, leading to enhanced OsCNGC9-mediated Ca2+ influx, the expression of cold stress-related calcium-dependent protein kinase genes (CPKs) and OsDREB1A, and chilling tolerance in rice (Figure 3) (Wang et al., 2021). Additionally, it was shown that rice CNGC14 and CNGC16 are involved in promoting tolerance toward heat and chilling stresses, and are regulators of Ca2+ signals in response to temperature stress (Cui et al., 2020).
Apart from CNGCs, Annexin 1 (ANN1) also plays a crucial role in increasing cytosolic Ca2+ during cold stress. In Arabidopsis, ANN1 mediates cold-triggered Ca2+ influx and contributes to freezing tolerance. Under normal conditions, the serine/threonine protein kinase Open Stomata 1 (OST1) interacts with PP2C, which inhibits OST1’s kinase activity. AtANN1 is localized both in the cytosol and at the plasma membrane. Upon cold stress, OST1 is activated and phosphorylates AtANN1. This phosphorylation enhances AtANN1’s Ca2+ transport activity and promotes its Ca2+-binding activity (Figure 3). The dual role of phosphorylation leads to an increase in cold-induced cytosolic Ca2+, which indirectly facilitates the expression of CBFs and CORs, positively regulating plant freezing tolerance (Liu et al., 2021). Although these findings provide important insights into the molecular mechanisms of plant responses to cold stress, several questions remain to be addressed. For instance, is the specific mechanism of AtANN1 consistent across different cellular environments? How do the interactions between OST1 and other signaling pathways influence the function of AtANN1? Future research should focus on these aspects.
In rice, CHILLING-TOLERANCE DIVERGENCE 1 (COLD1) plays an essential role in cold adaptation by regulating G-protein signaling. COLD1 is localized at the plasma membrane and endoplasmic reticulum (ER) and is involved in sensing cold temperatures, triggering Ca2+ signaling for chilling tolerance. A single-nucleotide mutation in COLD1 confers chilling tolerance to japonica rice, originating from the Chinese wild populations of Oryza rufipogon. COLD1 interacts with the G-protein α subunit (RGA1), activating Ca2+ channels and enhancing G-protein GTPase activity, thereby enhancing stress-related genes (OsDREB1A, OsDREB1B, OsDREB1C, and OsAP2) to promote chilling tolerance (Figure 3) (Ma et al., 2015). Recently, researchers have further revealed that COLD1 regulates the low temperature tolerance of rice by regulating the metabolism of downstream vitamin E-vitamin K1 (Luo et al., 2021). However, the molecular mechanism of COLD1 sensing low temperature and the Ca2+ channels activated by COLD1 remain unclear. In the future, structural biology and other means are needed to further elucidate the principle and conformational changes of low-temperature activation of COLD1.
Additionally, a single-nucleotide polymorphism (SNP2738) in ZmCOLD1 confers chilling tolerance and is associated with maize adaptation during speciation from teosinte (Zhou et al., 2023). ZmCOLD1 not only influences the influx of extracellular Ca2+, but also the levels of abscisic acid, gibberellic acid, and indole-3-acetic acid, during the germination stage, promoting chilling tolerance under low temperatures (Zhou et al., 2023).
In rice, the endoplasmic reticulum (ER)-localized Ca2+-binding protein calreticulin (CRT) plays a crucial role in various processes, including Ca2+ signaling (Michalak et al., 1999). CRT3 is a plant‐specific family member and differs from the CRT1/2 group that plays a role in SA‐dependent immune responses (Jia et al., 2009; Qiu et al., 2012). The CBL-INTERACTING PROTEIN KINASE (CIPK) is a calmodulin-interacting kinase that positively regulates cold stress responses, potentially through its kinase activity (D. Zhang et al., 2019). Under normal conditions, the interaction between OsCRT3 and OsCIPK7 is characterized by a low binding affinity. The kinase activity of OsCIPK7 is constrained by its intrinsic auto-inhibitory NAF/FISL domain, concomitantly with the maintenance of a diminished cytosolic Ca2+ concentration ([Ca2+]cyt). Under chilling stress, cold signaling induces conformational changes in OsCRT3, enhancing its binding affinity with OsCIPK7 and consequently activating its kinase activity (Figure 3). At the same time, OsCRT3, localized to the endoplasmic reticulum (ER), facilitates the increase in cold-induced [Ca2+]cyt, a signal that is detected by Ca2+ sensor proteins OsCBL7/8 (Figure 3). These proteins specifically interact with OsCIPK7 on the plasma membrane (PM). This interaction suggests a regulatory mechanism activated by chilling stress in rice, involving OsCRT3, OsCIPK7, Ca2+, and its sensor proteins OsCBL7/OsCBL8 (Guo et al., 2023).
As previously discussed, secondary messengers serve as critical upstream components in mediating signaling pathways, directly responding to signals from temperature stress sensors. Although it is well-established that temperature sensors initiate Ca2+ signaling pathways to mediate cold tolerance in cells, the mechanisms by which sensors interact with other secondary messengers remain poorly understood. Recent studies have identified 2′, 3′-cAMP as a secondary messenger crucial for chilling tolerance in crops (Luo et al., 2024). CHILLING-TOLERANCE DIVERGENCE 6 (COLD6), encoded by a major quantitative trait locus (QTL) gene in rice, interacts with RGA1 at the plasma membrane under normal conditions. Upon exposure to chilling, cold-induced osmotin protein (OSM1) binds to COLD6, displacing RGA1, which triggers the elevation of 2′, 3′-cAMP levels, promoting the expression of cold- responsive genes (Figure 3) (Luo et al., 2024). This elevation enhances chilling tolerance, providing a new insight into plant responses to cold environments and offering a potential target for molecular breeding aimed at improving cold tolerance in crops.
In rice, Thermo-tolerance 3 (TT3) has been identified, which modulates temperature perception based on protein localization and interaction (Zhang et al., 2022). TT3 is a quantitative trait locus (QTL) comprising two genes: TT3.1 and TT3.2. TT3.1 is a positive regulator of high-temperature stress, acting upstream of TT3.2, while TT3.2 plays a negative role in heat stress response. Under high-temperature stress, the heat stress‐induced plasma membrane (PM)‐localized E3 ligase TT3.1 proteins translocate from the PM to endosomes, where it ubiquitinates the chloroplast precursor protein TT3.2 (Figure 3). Subsequently, TT3.2 undergoes vacuolar degradation, preventing chloroplast thylakoid damage and improving heat tolerance (Zhang et al., 2022). However, the mechanism on how TT3.2 accumulation leads to chloroplast damage under heat stress conditions awaits further investigation. Given that the antagonistic effect of TT3.1 on TT3.2 confers protection to chloroplasts under heat stress conditions, overexpression of TT3.1 or targeted editing/silencing of TT3.2 could be employed as a strategic approach to enhance thermotolerance in rice.
Temperature also affects the activity and function of membrane proteins. Evidence suggests that the cyanobacterium histidine kinase 33 (Hik33) is the thermosensor that regulates desaturase gene expression in response to temperature downshifts (Suzuki et al., 2000). Hik33 contains 663 amino acid residues and the strongly conserved histidine kinase domain is located near the C-terminus. Hik33 is predicted to span the plasma membrane twice and forms a dimer, whose structure and activity may be influenced by the physical characteristics of lipids in the plasma membrane, such as their fluidity (or the extent of molecular motion), which is controlled by temperature and the extent of unsaturation of the fatty acids (Lau et al., 1997; Yaku and Mizuno, 1997; Singh et al., 1998; Suzuki et al., 2000). When the temperature is decreased (from 34 to 22°C), the histidine residue in the Hik33 kinase domain may be phosphorylated. A phosphate group is then transferred to Hik19, and finally to Rer1, which regulates the expression of the fatty-acid-desaturase genes B (desB) to adapt to cold stress (Figure 3) (Suzuki et al., 2000). Further, Hik33 has been shown to regulate the expression of osmotic stress-inducible genes and also to bind to certain chemicals, indicating that it may function as a multifunctional sensor for a variety of stresses (Jeon et al., 2010).
In addition to algae, there are similar cases in higher plants. In Arabidopsis, cold stress could activate the plasma membrane-localized protein kinase CRPK1 (cold-responsive protein kinase 1), which phosphorylates 14-3-3 proteins in the cytoplasm, thereby triggering 14-3-3 proteins to translocate into the nucleus (Liu et al., 2017). In the nucleus, phosphorylated 14-3-3 proteins form a protein complex with CBF proteins to promote the 26S proteasome-mediated degradation of CBF proteins, thus attenuating the CBF signaling (Figure 3). These results reveal that CRPK1 and 14-3-3 proteins act upstream of CBF proteins to negatively regulate plant freezing tolerance (Liu et al., 2017). Additional investigations are required to explore whether manipulating CRPK1 or 14-3-3 proteins could improve cold tolerance without compromising other vital physiological processes.
Xia et al. (2023) characterized Chilling-tolerance in Gengdao/japonica rice 1 (COG1) as a major gene identified in a QTL for positive regulation of chilling tolerance in japonica rice, which encodes a cold-induced LRR-RLP located in the plasma membrane and ER. Mechanistic studies have found that COG1 targets and activates the kinase somatic embryogenesis receptor kinase-like 2 (OsSERL2) in a cold-induced manner, promoting chilling tolerance (Figure 3). Furthermore, the cold signal transmitted by COG1-OsSERL2 activates OsMAPK3 in the cytoplasm, ultimately inducing a cold-tolerance response (Xia et al., 2023). This study not only advances our understanding of the molecular mechanisms underlying cold stress responses in rice but also highlights the potential of COG1 as a target for breeding cold-tolerant crop varieties. However, further research is needed to explore the broader applicability of these findings across diverse rice cultivars and environmental conditions, as well as to investigate potential trade-offs between chilling tolerance and other agronomic traits.
Earth is undergoing an unprecedented period of global climate change, characterized by frequent extreme temperature events. Dramatic temperature fluctuations induced by climate change inhibit plant growth, posing a significant threat to crop production and the sustainability of agricultural systems and food security. In response to this challenge, a range of plant thermosensors have been identified, shedding light on how plants sense and respond to temperature fluctuations (Ding and Yang, 2022; Noguchi and Kodama, 2022). RNA-based thermosensors have been identified frequently. For instance, temperature changes have the capacity to modulate not only the RNA structure, exemplified by PIF7 and psaA, but also influence the alternative splicing of mRNAs, including but not limited to FLM, HsfA2, and HsfA6e. These events triggered by extreme temperature play a pivotal role in temperature and regulating the expression of genes implicated in downstream temperature stress responses.
In contrast to RNA-based thermosensors, protein-based thermosensors have also been extensively characterized in plants, playing a crucial role in mediating temperature perception and signaling. As photoreceptors, phyB and phot undergo thermal reversion to relay downstream signaling perception of temperature change (Jung et al., 2016; Legris et al., 2016; Fujii et al., 2017). ELF3 and TWA1 sense temperature fluctuation through liquid-liquid phase separation (Fujii et al., 2017; Bohn et al., 2024). The fluidity and permeability of cellular phospholipid membranes are altered by extreme temperature changes in plants (Sangwan et al., 2002), suggesting that membrane-localized channels or receptors might sense temperature stress signals. The rapid increases in [Ca2+]cyt concentration triggered by extreme temperature are important in mediating downstream temperature stress-related gene expression (Wu et al., 2022; Li et al., 2023). Ca2+ signaling-related proteins, such as CNGCs, ANN1, COLD1, and CRT3-CIPK7, play important roles in temperature stress response by regulating Ca2+ level. In addition to the concentration of [Ca2+]cyt, temperature also affects the concentration of 2′, 3′-cAMP (Luo et al., 2024). Consequently, signaling proteins associated with 2′, 3′-cAMP, such as COLD, fulfill critical functions in the sensing temperature stress in plants. Under heat stress, TT3 modulates temperature perception based on protein localization and interaction (Zhang et al., 2022). The lower temperature affects the function of the membrane-located thermosensors. As the temperature decrease, the function of Hik33, CRPK1, and COG1 are activated, thereby transmitting the temperature signal to downstream components (Suzuki et al., 2000; Liu et al., 2017; Xia et al., 2023).
Notably, the interplay between different thermoreceptors, such as PIF7 and phyB in Arabidopsis, has provided critical insights into the molecular mechanisms of temperature perception. PIF7, an RNA thermosensor, is regulated by a hairpin structure within its 5′-UTR. Upon exposure to high temperatures, this structure partially unfolds, enabling translation initiation and leading to an increase in PIF7 protein levels. Concurrently, high temperatures promote the reversion of phyB from its active Pfr state to its inactive Pr state, a process known as thermal reversion. This allows PIF4 and PIF7 to activate thermomorphogenesis genes, demonstrating how plants integrate multiple thermosensors to regulate growth and development.
To cope with unfavorable changes in climate and an increasing global population, it is crucial to cultivate productive, climate-resilient crops. Advances in gene discovery have enabled the use of genetic modification techniques to enhance crop traits (Ding and Yang, 2022). Improving cold tolerance in rice is crucial for minimizing yield loss (Dong et al., 2025). As mentioned earlier, COLD1 and COG1 overexpression lines (Ma et al., 2015; Xia et al., 2023), and cold6 mutants exhibited higher chilling tolerance (Luo et al., 2024), which suggests that the potential of either genetic or transgenic approaches to improve chilling tolerance for rice breeding. Besides, the overexpression of TT3.1 or knockout of TT3.2 lines were more heat-tolerant and increased yield by more than 2.5 times (Zhang et al., 2022). Interestingly, TT3.1 and TT3.2 are conserved in other major crops such as maize and wheat (Li and Liu, 2022), which would be meaningful to know whether these orthologous genes could be used for breeding heat tolerant crops. From the discussion, one may conclude that there is great potential to apply the identified plant thermosensors to improve crop temperature stress tolerance and thus increase yields. Further, it is worth exploring how to efficiently apply the thermosensors identified in model plants to crop improvement. The present understanding of individual molecular components in temperature responses may be established. However, the interactive and coordinated mechanisms among multiple molecules remain unclear. Such molecular coordination may play crucial roles in various biological processes. Therefore, further research is needed to understand how multiple molecules coordinate temperature responses during the same stress or developmental processes.
Currently, few DNA thermosensors have been characterized, and most reported thermosensors are RNA, protein, and plasma membrane-associated protein-based thermosensors in plants. RNA can sense temperature changes through changes in secondary structure, while proteins can regulate their function through heat-induced conformational changes. Comparatively speaking, the structural changes of DNA are limited by its double helix conformation, and the regulatory mode is relatively limited (Singh et al., 2022). The potential of DNA as a temperature sensor has not been fully explored, possibly also due to a lack of relevant experimental tools or methods. While DNA-based thermosensors are less commonly discussed, recent studies have begun to uncover their potential roles in plant responses to temperature changes. For instance, temperature-induced changes in DNA methylation patterns and chromatin remodeling have been implicated in regulating gene expression under thermal stress (Song et al., 2021; Talarico et al., 2024). Thus, the potential role of epigenetic mechanisms—such as DNA methylation, RNA methylation, chromatin structure modifications, loss of imprinting, and non-coding RNAs—in temperature sensing remains largely unexplored. Understanding the full spectrum of temperature perception pathways is crucial for improving plant growth and development, and identifying additional thermosensors could play a pivotal role in enhancing crop stress resistance (Raza et al., 2024).
Genetic redundancy and lethality present challenges in identifying new thermosensors. Nevertheless, molecular genetic approaches, coupled with innovative bioimaging techniques such as Ca2+ imaging-based forward genetic screens and fluorescence-based Ca2+ indicators, are helping to uncover new temperature sensors (Sun et al., 2021; Jin et al., 2024). Moreover, the integration of protein-protein interaction studies, multi-omics techniques (including genomics, proteomics, metabolomics, lipidomics, glycomics, and transcriptomics), bioinformatics, and advanced microscopy is expected to accelerate the discovery of novel thermoreceptors. The diversity of plant responses to temperature anticipates that many new thermosensors and eventually novel sensing mechanisms will be uncovered soon (Casal et al., 2024). Ultimately, translating these findings into practical applications for improving crop resilience to temperature stress through breeding and cultivation techniques will be critical to sustaining global food security in the face of climate change.
TZ: Writing – original draft, Writing – review & editing, Data curation, Investigation. XC: Writing – original draft, Investigation. CL: Writing – original draft. YL: Writing – original draft. CP: Writing – review & editing. GL: Writing – review & editing, Funding acquisition, Supervision.
The author(s) declare that financial support was received for the research and/or publication of this article. This work was supported by the Ningbo 2035 Key Technology Project of Yongjiang Science and Technology Innovation, Grant Number: 2024Z269, the National Natural Science Foundation of China, Grant Number: 31972417, the Zhejiang Province Key Research and Development Special Project for Breeding, Grant Number: 2021C02065, the Haerbin Key Science and Technology Project, Grant Number: 2021ZSZZNS07, the Starry Night Science Fund of Zhejiang University Shanghai Institute for Advanced Study, Grant Number: SN-ZJU-SIAS-0011, and the Zhejiang Provincial “Three Rural Nine Parties” Science and Technology Cooperation Plan Project, Grant Number: 2025SNJF003.
Author TZ was employed by the company Zhejiang Seed Industry Group Xinchuang Bio-breeding Co., Ltd.
The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Airoldi, C. A., McKay, M., Davies, B. (2015). MAF2 is regulated by temperature-dependent splicing and represses flowering at low temperatures in parallel with FLM. PloS One 10, e0126516. doi: 10.1371/journal.pone.0126516
Asseng, S., Ewert, F., Martre, P., Rötter, R. P., Lobell, D. B., Cammarano, D., et al. (2015). Rising temperatures reduce global wheat production. Nat. Climate Change 5, 143–147. doi: 10.1038/nclimate2470
Balogh, G., Péter, M., Glatz, A., Gombos, I., Török, Z., Horváth, I., et al. (2013). Key role of lipids in heat stress management. FEBS Lett. 587, 1970–1980. doi: 10.1016/j.febslet.2013.05.016
Bohn, L., Huang, J., Weidig, S., Yang, Z., Heidersberger, C., Genty, B., et al. (2024). The temperature sensor TWA1 is required for thermotolerance in Arabidopsis. Nature 629, 1126–1132. doi: 10.1038/s41586-024-07424-x
Box, M. S., Huang, B. E., Domijan, M., Jaeger, K. E., Khattak, A. K., Yoo, S. J., et al. (2015). ELF3 controls thermoresponsive growth in Arabidopsis. Curr. Biol. 25, 194–199. doi: 10.1016/j.cub.2014.10.076
Broft, P., Rosenkranz, R., Schleiff, E., Hengesbach, M., Schwalbe, H. (2022). Structural analysis of temperature-dependent alternative splicing of HsfA2 pre-mRNA from tomato plants. RNA Biol. 19, 266–278. doi: 10.1080/15476286.2021.2024034
Burgie, E. S., Gannam, Z. T. K., McLoughlin, K. E., Sherman, C. D., Holehouse, A. S., Stankey, R. J., et al. (2021). Differing biophysical properties underpin the unique signaling potentials within the plant phytochrome photoreceptor families. Proc. Natl. Acad. Sci. U.S.A. 118(22), e2105649118. doi: 10.1073/pnas.2105649118
Burgie, E. S., Vierstra, R. D. (2014). Phytochromes: an atomic perspective on photoactivation and signaling. Plant Cell 26, 4568–4583. doi: 10.1105/tpc.114.131623
Capovilla, G., Symeonidi, E., Wu, R., Schmid, M. (2017). Contribution of major FLM isoforms to temperature-dependent flowering in Arabidopsis thaliana. J. Exp. Bot. 68, 5117–5127. doi: 10.1093/jxb/erx328
Casal, J. J., Murcia, G., Bianchimano, L. (2024). Plant thermosensors. Annu. Rev. Genet. 58, 135–158. doi: 10.1146/annurev-genet-111523-102327
Cech, T. R., Steitz, J. A. (2014). The noncoding RNA revolution-trashing old rules to forge new ones. Cell 157, 77–94. doi: 10.1016/j.cell.2014.03.008
Chełkowska-Pauszek, A., Kosiński, J. G., Marciniak, K., Wysocka, M., Bąkowska-Żywicka, K., Żywicki, M. (2021). The role of RNA secondary structure in regulation of gene expression in bacteria. Int. J. Mol. Sci. 22(15), 7845. doi: 10.3390/ijms22157845
Christie, J. M. (2007). Phototropin blue-light receptors. Annu. Rev. Plant Biol. 58, 21–45. doi: 10.1146/annurev.arplant.58.032806.103951
Chung, B. Y. W., Balcerowicz, M., Di Antonio, M., Jaeger, K. E., Geng, F., Franaszek, K., et al. (2020). An RNA thermoswitch regulates daytime growth in Arabidopsis. Nat. Plants 6, 522–532. doi: 10.1038/s41477-020-0633-3
Chung, K. P., Loiacono, F. V., Neupert, J., Wu, M., Bock, R. (2023). An RNA thermometer in the chloroplast genome of Chlamydomonas facilitates temperature-controlled gene expression. Nucleic Acids Res. 51, 11386–11400. doi: 10.1093/nar/gkad816
Cohen, J., Screen, J. A., Furtado, J. C., Barlow, M., Whittleston, D., Coumou, D., et al. (2014). Recent Arctic amplification and extreme mid-latitude weather. Nat. Geosci. 7, 627–637. doi: 10.1038/ngeo2234
Coleman-Derr, D., Zilberman, D. (2012). Deposition of histone variant H2A.Z within gene bodies regulates responsive genes. PloS Genet. 8, e1002988. doi: 10.1371/journal.pgen.1002988
Cortijo, S., Charoensawan, V., Brestovitsky, A., Buning, R., Ravarani, C., Rhodes, D., et al. (2017). Transcriptional regulation of the ambient temperature response by H2A.Z nucleosomes and HSF1 transcription factors in arabidopsis. Mol. Plant 10, 1258–1273. doi: 10.1016/j.molp.2017.08.014
Cui, Y., Lu, S., Li, Z., Cheng, J., Hu, P., Zhu, T., et al. (2020). CYCLIC NUCLEOTIDE-GATED ION CHANNELs 14 and 16 promote tolerance to heat and chilling in rice. Plant Physiol. 183, 1794–1808. doi: 10.1104/pp.20.00591
Di Martino, M. L., Falconi, M., Micheli, G., Colonna, B., Prosseda, G. (2016). The multifaceted activity of the virF regulatory protein in the shigella lifestyle. Front. Mol. Biosci. 3. doi: 10.3389/fmolb.2016.00061
Ding, Y., Yang, S. (2022). Surviving and thriving: How plants perceive and respond to temperature stress. Dev. Cell 57, 947–958. doi: 10.1016/j.devcel.2022.03.010
Dong, J., Zhang, S., Hu, H., Wang, J., Li, R., Wu, J., et al. (2025). Natural variation in CTF1 conferring cold tolerance at the flowering stage in rice. Plant Biotechnol. J. doi: 10.1111/pbi.14600
Escribá, P. V., González-Ros, J. M., Goñi, F. M., Kinnunen, P. K., Vigh, L., Sánchez-Magraner, L., et al. (2008). Membranes: a meeting point for lipids, proteins and therapies. J. Cell Mol. Med. 12, 829–875. doi: 10.1111/j.1582-4934.2008.00281.x
Ezer, D., Jung, J.-H., Lan, H., Biswas, S., Gregoire, L., Box, M. S., et al. (2017). The evening complex coordinates environmental and endogenous signals in Arabidopsis. Nat. Plants 3, 17087. doi: 10.1038/nplants.2017.87
Falcone, D. L., Ogas, J. P., Somerville, C. R. (2004). Regulation of membrane fatty acid composition by temperature in mutants of Arabidopsis with alterations in membrane lipid composition. BMC Plant Biol. 4, 17. doi: 10.1186/1471-2229-4-17
Falconi, M., Colonna, B., Prosseda, G., Micheli, G., Gualerzi, C. O. (1998). Thermoregulation of Shigella and Escherichia coli EIEC pathogenicity. A temperature-dependent structural transition of DNA modulates accessibility of virF promoter to transcriptional repressor H-NS. EMBO J. 17, 7033–7043. doi: 10.1093/emboj/17.23.7033
Findlay, K. M., Jenkins, G. I. (2016). Regulation of UVR8 photoreceptor dimer/monomer photo-equilibrium in Arabidopsis plants grown under photoperiodic conditions. Plant Cell Environ. 39, 1706–1714. doi: 10.1111/pce.12724
Finka, A., Cuendet, A. F., Maathuis, F. J., Saidi, Y., Goloubinoff, P. (2012). Plasma membrane cyclic nucleotide gated calcium channels control land plant thermal sensing and acquired thermotolerance. Plant Cell 24, 3333–3348. doi: 10.1105/tpc.112.095844
Fujii, Y., Tanaka, H., Konno, N., Ogasawara, Y., Hamashima, N., Tamura, S., et al. (2017). Phototropin perceives temperature based on the lifetime of its photoactivated state. Proc. Natl. Acad. Sci. U.S.A. 114, 9206–9211. doi: 10.1073/pnas.1704462114
Fuller, M. P., Fuller, A. M., Kaniouras, S., Christophers, J., Fredericks, T. (2007). The freezing characteristics of wheat at ear emergence. Eur. J. Agron. 26, 435–441. doi: 10.1016/j.eja.2007.01.001
Gao, F., Han, X., Wu, J., Zheng, S., Shang, Z., Sun, D., et al. (2012). A heat-activated calcium-permeable channel–Arabidopsis cyclic nucleotide-gated ion channel 6–is involved in heat shock responses. Plant J. 70, 1056–1069. doi: 10.1111/j.1365-313X.2012.04969.x
Gass, T., Schori, A., Fossati, A., Soldati, A., Stamp, P. (1996). Cold tolerance of soybean (Glycine max (L.) Merr.) during the reproductive phase. Eur. J. Agron. 5, 71–88. doi: 10.1016/S1161-0301(96)02011-4
Georgakopoulos-Soares, I., Parada, G. E., Hemberg, M. (2022). Secondary structures in RNA synthesis, splicing and translation. Comput. Struct. Biotechnol. J. 20, 2871–2884. doi: 10.1016/j.csbj.2022.05.041
Gilmour, S. J., Sebolt, A. M., Salazar, M. P., Everard, J. D., Thomashow, M. F. (2000). Overexpression of the Arabidopsis CBF3 transcriptional activator mimics multiple biochemical changes associated with cold acclimation. Plant Physiol. 124, 1854–1865. doi: 10.1104/pp.124.4.1854
Guihur, A., Rebeaud, M. E., Goloubinoff, P. (2022). How do plants feel the heat and survive? Trends Biochem. Sci. 47, 824–838. doi: 10.1016/j.tibs.2022.05.004
Guo, M., Liu, J. H., Ma, X., Luo, D. X., Gong, Z. H., Lu, M. H. (2016). The plant heat stress transcription factors (HSFs): structure, regulation, and function in response to abiotic stresses. Front. Plant Sci. 7. doi: 10.3389/fpls.2016.00114
Guo, X., Zhang, D., Wang, Z., Xu, S., Batistič, O., Steinhorst, L., et al. (2023). Cold-induced calreticulin OsCRT3 conformational changes promote OsCIPK7 binding and temperature sensing in rice. EMBO J. 42, e110518. doi: 10.15252/embj.2021110518
Hayes, S., Schachtschabel, J., Mishkind, M., Munnik, T., Arisz, S. A. (2021). Hot topic: Thermosensing in plants. Plant Cell Environ. 44, 2018–2033. doi: 10.1111/pce.13979
He, M., He, C. Q., Ding, N. Z. (2018). Abiotic stresses: general defenses of land plants and chances for engineering multistress tolerance. Front. Plant Sci. 9. doi: 10.3389/fpls.2018.01771
Hu, Y., Mesihovic, A., Jiménez-Gómez, J. M., Röth, S., Gebhardt, P., Bublak, D., et al. (2020). Natural variation in HsfA2 pre-mRNA splicing is associated with changes in thermotolerance during tomato domestication. New Phytol. 225, 1297–1310. doi: 10.1111/nph.16221
Inda, M. E., Oliveira, R. G., de Mendoza, D., Cybulski, L. E. (2016). The single transmembrane segment of minimal sensor desK senses temperature via a membrane-thickness caliper. J. Bacteriol. 198, 2945–2954. doi: 10.1128/jb.00431-16
Jaglo-Ottosen, K. R., Gilmour, S. J., Zarka, D. G., Schabenberger, O., Thomashow, M. F. (1998). Arabidopsis CBF1 overexpression induces COR genes and enhances freezing tolerance. Science 280, 104–106. doi: 10.1126/science.280.5360.104
Janni, M., Gullì, M., Maestri, E., Marmiroli, M., Valliyodan, B., Nguyen, H. T., et al. (2020). Molecular and genetic bases of heat stress responses in crop plants and breeding for increased resilience and productivity. J. Exp. Bot. 71, 3780–3802. doi: 10.1093/jxb/eraa034
Jeon, J., Kim, N. Y., Kim, S., Kang, N. Y., Novák, O., Ku, S. J., et al. (2010). A subset of cytokinin two-component signaling system plays a role in cold temperature stress response in Arabidopsis. J. Biol. Chem. 285, 23371–23386. doi: 10.1074/jbc.M109.096644
Jha, S. K., Sharma, M., Pandey, G. K. (2016). Role of cyclic nucleotide gated channels in stress management in plants. Curr. Genomics 17, 315–329. doi: 10.2174/1389202917666160331202125
Jia, X. Y., He, L. H., Jing, R. L., Li, R. Z. (2009). Calreticulin: conserved protein and diverse functions in plants. Physiol. Plant 136, 127–138. doi: 10.1111/j.1399-3054.2009.1223.x
Jiang, B., Shi, Y., Peng, Y., Jia, Y., Yan, Y., Dong, X., et al. (2020). Cold-induced CBF-PIF3 interaction enhances freezing tolerance by stabilizing the phyB thermosensor in arabidopsis. Mol. Plant 13, 894–906. doi: 10.1016/j.molp.2020.04.006
Jin, S., Wei, M., Wei, Y., Jiang, Z. (2024). Insights into Plant Sensory Mechanisms under Abiotic Stresses. Plants (Basel). 13 (14), 1907. doi: 10.3390/plants13141907
Jung, J. H., Barbosa, A. D., Hutin, S., Kumita, J. R., Gao, M., Derwort, D., et al. (2020). A prion-like domain in ELF3 functions as a thermosensor in Arabidopsis. Nature 585, 256–260. doi: 10.1038/s41586-020-2644-7
Jung, J. H., Domijan, M., Klose, C., Biswas, S., Ezer, D., Gao, M., et al. (2016). Phytochromes function as thermosensors in Arabidopsis. Science 354, 886–889. doi: 10.1126/science.aaf6005
Kerbler, S. M., Wigge, P. A. (2023). Temperature sensing in plants. Annu. Rev. Plant Biol. 74, 341–366. doi: 10.1146/annurev-arplant-102820-102235
Kim, J. J., Lee, J. H., Kim, W., Jung, H. S., Huijser, P., Ahn, J. H. (2012). The microRNA156-SQUAMOSA PROMOTER BINDING PROTEIN-LIKE3 module regulates ambient temperature-responsive flowering via FLOWERING LOCUS T in Arabidopsis. Plant Physiol. 159, 461–478. doi: 10.1104/pp.111.192369
Kotak, S., Vierling, E., Bäumlein, H., von-Koskull-Döring, P. (2007). A novel transcriptional cascade regulating expression of heat stress proteins during seed development of Arabidopsis. Plant Cell 19, 182–195. doi: 10.1105/tpc.106.048165
Kumar, S. V., Wigge, P. A. (2010). H2A.Z-containing nucleosomes mediate the thermosensory response in Arabidopsis. Cell 140, 136–147. doi: 10.1016/j.cell.2009.11.006
Lau, P. C., Wang, Y., Patel, A., Labbé, D., Bergeron, H., Brousseau, R., et al. (1997). A bacterial basic region leucine zipper histidine kinase regulating toluene degradation. Proc. Natl. Acad. Sci. U.S.A. 94, 1453–1458. doi: 10.1073/pnas.94.4.1453
Legris, M., Klose, C., Burgie, E. S., Rojas, C. C., Neme, M., Hiltbrunner, A., et al. (2016). Phytochrome B integrates light and temperature signals in Arabidopsis. Science 354, 897–900. doi: 10.1126/science.aaf5656
Li, J. Y., Liu, J. X. (2022). TT3.1: a journey to protect chloroplasts upon heat stress. Stress Biol. 2, 27. doi: 10.1007/s44154-022-00051-4
Li, J., Song, Y. (2024). Plant thermosensors. Plant Sci. 342, 112025. doi: 10.1016/j.plantsci.2024.112025
Li, H., Yang, L., Fang, Y., Wang, G., Liu, T. (2024). RtHSFA9s of rhodomyrtus tomentosa positively regulate thermotolerance by transcriptionally activating rtHSFA2s and rtHSPs. Life (Basel). 14 (12), 1591. doi: 10.3390/life14121591
Li, J. Y., Yang, C., Xu, J., Lu, H. P., Liu, J. X. (2023). The hot science in rice research: How rice plants cope with heat stress. Plant Cell Environ. 46, 1087–1103. doi: 10.1111/pce.14509
Liu, Q., Ding, Y., Shi, Y., Ma, L., Wang, Y., Song, C., et al. (2021). The calcium transporter ANNEXIN1 mediates cold-induced calcium signaling and freezing tolerance in plants. EMBO J. 40, e104559. doi: 10.15252/embj.2020104559
Liu, Z., Jia, Y., Ding, Y., Shi, Y., Li, Z., Guo, Y., et al. (2017). Plasma membrane CRPK1-mediated phosphorylation of 14-3-3 proteins induces their nuclear import to fine-tune CBF signaling during cold response. Mol. Cell 66, 117–128.e115. doi: 10.1016/j.molcel.2017.02.016
Liu, J., Sun, N., Liu, M., Liu, J., Du, B., Wang, X., et al. (2013). An autoregulatory loop controlling Arabidopsis HsfA2 expression: role of heat shock-induced alternative splicing. Plant Physiol. 162, 512–521. doi: 10.1104/pp.112.205864
Luo, W., Huan, Q., Xu, Y., Qian, W., Chong, K., Zhang, J. (2021). Integrated global analysis reveals a vitamin E-vitamin K1 sub-network, downstream of COLD1, underlying rice chilling tolerance divergence. Cell Rep. 36, 109397. doi: 10.1016/j.celrep.2021.109397
Luo, W., Xu, Y., Cao, J., Guo, X., Han, J., Zhang, Y., et al. (2024). COLD6-OSM1 module senses chilling for cold tolerance via 2’,3’-cAMP signaling in rice. Mol. Cell. 84(21), 4224–4238.e9. doi: 10.1016/j.molcel.2024.09.031
Lutz, U., Nussbaumer, T., Spannagl, M., Diener, J., Mayer, K. F., Schwechheimer, C. (2017). Natural haplotypes of FLM non-coding sequences fine-tune flowering time in ambient spring temperatures in Arabidopsis. Elife 6, e22114. doi: 10.7554/eLife.22114
Ma, Y., Dai, X., Xu, Y., Luo, W., Zheng, X., Zeng, D., et al. (2015). COLD1 confers chilling tolerance in rice. Cell 160, 1209–1221. doi: 10.1016/j.cell.2015.01.046
Michalak, M., Corbett, E. F., Mesaeli, N., Nakamura, K., Opas, M. (1999). Calreticulin: one protein, one gene, many functions. Biochem. J. 344 Pt 2, 281–292. doi: 10.1042/bj3440281
Mishra, K. B., Mishra, A., Kubásek, J., Urban, O., Heyer, A. G., Govindjee (2019). Low temperature induced modulation of photosynthetic induction in non-acclimated and cold-acclimated Arabidopsis thaliana: chlorophyll a fluorescence and gas-exchange measurements. Photosynth. Res. 139, 123–143. doi: 10.1007/s11120-018-0588-7
Mizushima, T., Kataoka, K., Ogata, Y., Inoue, R., Sekimizu, K. (1997). Increase in negative supercoiling of plasmid DNA in Escherichia coli exposed to cold shock. Mol. Microbiol. 23, 381–386. doi: 10.1046/j.1365-2958.1997.2181582.x
Nieto, C., López-Salmerón, V., Davière, J. M., Prat, S. (2015). ELF3-PIF4 interaction regulates plant growth independently of the Evening Complex. Curr. Biol. 25, 187–193. doi: 10.1016/j.cub.2014.10.070
Niu, W. T., Han, X. W., Wei, S. S., Shang, Z. L., Wang, J., Yang, D. W., et al. (2020). Arabidopsis cyclic nucleotide-gated channel 6 is negatively modulated by multiple calmodulin isoforms during heat shock. J. Exp. Bot. 71, 90–104. doi: 10.1093/jxb/erz445
Niu, Y., Xiang, Y. (2018). An overview of biomembrane functions in plant responses to high-temperature stress. Front. Plant Sci. 9. doi: 10.3389/fpls.2018.00915
Noguchi, M., Kodama, Y. (2022). Temperature sensing in plants: on the dawn of molecular thermosensor research. Plant Cell Physiol. 63, 737–743. doi: 10.1093/pcp/pcac033
Ohama, N., Kusakabe, K., Mizoi, J., Zhao, H., Kidokoro, S., Koizumi, S., et al. (2016). The transcriptional cascade in the heat stress response of arabidopsis is strictly regulated at the level of transcription factor expression. Plant Cell 28, 181–201. doi: 10.1105/tpc.15.00435
Peng, S., Huang, J., Sheehy, J. E., Laza, R. C., Visperas, R. M., Zhong, X., et al. (2004). Rice yields decline with higher night temperature from global warming. Proc. Natl. Acad. Sci. U.S.A. 101, 9971–9975. doi: 10.1073/pnas.0403720101
Peterson, M. E., Daniel, R. M., Danson, M. J., Eisenthal, R. (2007). The dependence of enzyme activity on temperature: determination and validation of parameters. Biochem. J. 402, 331–337. doi: 10.1042/bj20061143
Pooam, M., Dixon, N., Hilvert, M., Misko, P., Waters, K., Jourdan, N., et al. (2021). Effect of temperature on the Arabidopsis cryptochrome photocycle. Physiol. Plant 172, 1653–1661. doi: 10.1111/ppl.13365
Posé, D., Verhage, L., Ott, F., Yant, L., Mathieu, J., Angenent, G. C., et al. (2013). Temperature-dependent regulation of flowering by antagonistic FLM variants. Nature 503, 414–417. doi: 10.1038/nature12633
Qiu, Y., Li, M., Kim, R. J., Moore, C. M., Chen, M. (2019). Daytime temperature is sensed by phytochrome B in Arabidopsis through a transcriptional activator HEMERA. Nat. Commun. 10, 140. doi: 10.1038/s41467-018-08059-z
Qiu, Y., Xi, J., Du, L., Roje, S., Poovaiah, B. W. (2012). A dual regulatory role of Arabidopsis calreticulin-2 in plant innate immunity. Plant J. 69, 489–500. doi: 10.1111/j.1365-313X.2011.04807.x
Quail, P. H., Boylan, M. T., Parks, B. M., Short, T. W., Xu, Y., Wagner, D. (1995). Phytochromes: photosensory perception and signal transduction. Science 268, 675–680. doi: 10.1126/science.7732376
Raza, A., Zaman, Q. U., Hu, Z. (2024). Leveraging a new thermosensor for heat-smart future agriculture. Plant Commun. 5, 101007. doi: 10.1016/j.xplc.2024.101007
Reynolds, C. A., King, P. M., Richards, W. G. (1992). Free energy calculations in molecular biophysics. Mol. Phys. 76, 251–275. doi: 10.1080/00268979200101321
Sangwan, V., Orvar, B. L., Beyerly, J., Hirt, H., Dhindsa, R. S. (2002). Opposite changes in membrane fluidity mimic cold and heat stress activation of distinct plant MAP kinase pathways. Plant J. 31, 629–638. doi: 10.1046/j.1365-313x.2002.01384.x
Sengupta, P., Garrity, P. (2013). Sensing temperature. Curr. Biol. 23, R304–R307. doi: 10.1016/j.cub.2013.03.009
Severing, E., Faino, L., Jamge, S., Busscher, M., Kuijer-Zhang, Y., Bellinazzo, F., et al. (2018). Arabidopsis thaliana ambient temperature responsive lncRNAs. BMC Plant Biol. 18, 145. doi: 10.1186/s12870-018-1362-x
Singh, M., Berger, B., Kim, P. S., Berger, J. M., Cochran, A. G. (1998). Computational learning reveals coiled coil-like motifs in histidine kinase linker domains. Proc. Natl. Acad. Sci. U.S.A. 95, 2738–2743. doi: 10.1073/pnas.95.6.2738
Singh, A., Maity, A., Singh, N. (2022). Structure and dynamics of dsDNA in cell-like environments. Entropy. (Basel). 24 (11), 1587. doi: 10.3390/e24111587
Sivaji, M. (2021). “Abiotic stress-induced molecular and physiological changes and adaptive mechanisms in plants,” in Abiotic Stress in Plants (pp. Ch. 15). Eds. Shah, F., Shah, S., Yajun, C., Chao, W., Depeng, W. (IntechOpen, Rijeka).
Somero, G. N. (1995). Proteins and temperature. Annu. Rev. Physiol. 57, 43–68. doi: 10.1146/annurev.ph.57.030195.000355
Song, Z. T., Liu, J. X., Han, J. J. (2021). Chromatin remodeling factors regulate environmental stress responses in plants. J. Integr. Plant Biol. 63, 438–450. doi: 10.1111/jipb.13064
Stief, A., Altmann, S., Hoffmann, K., Pant, B. D., Scheible, W. R., Bäurle, I. (2014). Arabidopsis miR156 Regulates Tolerance to Recurring Environmental Stress through SPL Transcription Factors. Plant Cell 26, 1792–1807. doi: 10.1105/tpc.114.123851
Sun, S., Zhang, X., Chen, K., Zhu, X., Zhao, Y. (2021). Screening for Arabidopsis mutants with altered Ca(2+) signal response using aequorin-based Ca(2+) reporter system. STAR. Protoc. 2, 100558. doi: 10.1016/j.xpro.2021.100558
Sureshkumar, S., Dent, C., Seleznev, A., Tasset, C., Balasubramanian, S. (2016). Nonsense-mediated mRNA decay modulates FLM-dependent thermosensory flowering response in Arabidopsis. Nat. Plants 2, 16055. doi: 10.1038/nplants.2016.55
Suzuki, I., Los, D. A., Kanesaki, Y., Mikami, K., Murata, N. (2000). The pathway for perception and transduction of low-temperature signals in Synechocystis. EMBO J. 19, 1327–1334. doi: 10.1093/emboj/19.6.1327
Talarico, E., Zambelli, A., Araniti, F., Greco, E., Chiappetta, A., Bruno, L. (2024). Unravelling the epigenetic code: DNA methylation in plants and its role in stress response. Epigenomes 8 (3), 30. doi: 10.3390/epigenomes8030030
Tasset, C., Singh Yadav, A., Sureshkumar, S., Singh, R., van der Woude, L., Nekrasov, M., et al. (2018). POWERDRESS-mediated histone deacetylation is essential for thermomorphogenesis in Arabidopsis thaliana. PloS Genet. 14, e1007280. doi: 10.1371/journal.pgen.1007280
Thakur, P., Kumar, S., Malik, J. A., Berger, J. D., Nayyar, H. (2010). Cold stress effects on reproductive development in grain crops: An overview. Environ. Exp. Bot. 67, 429–443. doi: 10.1016/j.envexpbot.2009.09.004
Vu, L. D., Gevaert, K., De Smet, I. (2019). Feeling the heat: searching for plant thermosensors. Trends Plant Sci. 24, 210–219. doi: 10.1016/j.tplants.2018.11.004
Waldminghaus, T., Kortmann, J., Gesing, S., Narberhaus, F. (2008). Generation of synthetic RNA-based thermosensors. Biol. Chem. 389, 1319–1326. doi: 10.1515/bc.2008.150
Wang, J., Ren, Y., Liu, X., Luo, S., Zhang, X., Liu, X., et al. (2021). Transcriptional activation and phosphorylation of OsCNGC9 confer enhanced chilling tolerance in rice. Mol. Plant 14, 315–329. doi: 10.1016/j.molp.2020.11.022
Wang, X., Zhu, Y., Tang, L., Wang, Y., Sun, R., Deng, X. (2024). Arabidopsis HSFA9 acts as a regulator of heat response gene expression and the acquisition of thermotolerance and seed longevity. Plant Cell Physiol. 65, 372–389. doi: 10.1093/pcp/pcad164
Wen, J., Qin, Z., Sun, L., Zhang, Y., Wang, D., Peng, H., et al. (2023). Alternative splicing of TaHSFA6e modulates heat shock protein-mediated translational regulation in response to heat stress in wheat. New Phytol. 239, 2235–2247. doi: 10.1111/nph.19100
Wu, N., Yao, Y., Xiang, D., Du, H., Geng, Z., Yang, W., et al. (2022). A MITE variation-associated heat-inducible isoform of a heat-shock factor confers heat tolerance through regulation of JASMONATE ZIM-DOMAIN genes in rice. New Phytol. 234, 1315–1331. doi: 10.1111/nph.18068
Xia, C., Liang, G., Chong, K., Xu, Y. (2023). The COG1-OsSERL2 complex senses cold to trigger signaling network for chilling tolerance in japonica rice. Nat. Commun. 14, 3104. doi: 10.1038/s41467-023-38860-4
Yaku, H., Mizuno, T. (1997). The membrane-located osmosensory kinase, EnvZ, that contains a leucine zipper-like motif functions as a dimer in Escherichia coli. FEBS Lett. 417, 409–413. doi: 10.1016/s0014-5793(97)01329-x
Zaveri, E., Lobell, D. B. (2019). The role of irrigation in changing wheat yields and heat sensitivity in India. Nat. Commun. 10, 4144. doi: 10.1038/s41467-019-12183-9
Zhang, D., Guo, X., Xu, Y., Li, H., Ma, L., Yao, X., et al. (2019). OsCIPK7 point-mutation leads to conformation and kinase-activity change for sensing cold response. J. Integr. Plant Biol. 61, 1194–1200. doi: 10.1111/jipb.12800
Zhang, H., Zhou, J. F., Kan, Y., Shan, J. X., Ye, W. W., Dong, N. Q., et al. (2022). A genetic module at one locus in rice protects chloroplasts to enhance thermotolerance. Science 376, 1293–1300. doi: 10.1126/science.abo5721
Zhou, Y., Zhang, H., Zhang, S., Zhang, J., Di, H., Zhang, L., et al. (2023). The G protein-coupled receptor COLD1 promotes chilling tolerance in maize during germination. Int. J. Biol. Macromol. 253, 126877. doi: 10.1016/j.ijbiomac.2023.126877
Keywords: thermosensor, temperature, stress, plants, crops
Citation: Zhu T, Cheng X, Li C, Li Y, Pan C and Lu G (2025) Decoding plant thermosensors: mechanism of temperature perception and stress adaption. Front. Plant Sci. 16:1560204. doi: 10.3389/fpls.2025.1560204
Received: 14 January 2025; Accepted: 03 March 2025;
Published: 25 March 2025.
Edited by:
Raul Antonio Sperotto, Federal University of Pelotas, BrazilReviewed by:
Klára Kosová, Crop Research Institute (CRI), CzechiaCopyright © 2025 Zhu, Cheng, Li, Li, Pan and Lu. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Gang Lu, Z2x1QHpqdS5lZHUuY24=
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.