
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Plant Sci. , 07 March 2025
Sec. Plant Pathogen Interactions
Volume 16 - 2025 | https://doi.org/10.3389/fpls.2025.1556928
This article is part of the Research Topic Unraveling Pathogen-Plant-Microbiome Interactions in Horticultural Crops Through Omics Approaches View all 14 articles
Late blight caused by the oomycete Phytophthora infestans poses a severe threat to global tomato (Solanum lycopersicum L.) production. While genetic resistance forms the cornerstone of disease control, the mechanisms underlying cultivar-specific resistance, particularly their interactions with rhizosphere microbiomes, remain poorly understood. To elucidate the mechanisms of tomato cultivar resistance to late blight and screen out antagonistic microorganisms against P. infestans, we investigated the microbial compositions in the rhizospheres of tomato cultivars with different late blight-resistance levels under both natural and P. infestans-inoculated conditions. Considerable differences in soil microbial diversity and composition of rhizospheres were found between late blight-resistant and -susceptible tomato cultivars. Under natural conditions, the resistant tomato cultivar exhibited higher bacterial diversity and lower fungal diversity than that of the susceptible cultivar. Additionally, after P. infestans inoculation, both the resistant and susceptible cultivars showed enrichment of microorganisms with potential antagonistic effects in the rhizospheres. Among them, bacterial genera, such as Pseudomonas, Azospirillum, and Acidovorax, and fungal genera, including Phoma, Arthrobotrys, Pseudallescheria, and Pseudolabrys, were enriched in the rhizospheres of the late blight-resistant tomato cultivar. In contrast, bacterial genera, including Flavobacterium, Pseudolabrys, and Burkholderia-Caballeronia-Paraburkholderia, and the Trichoderma fungal genus were enriched in the rhizospheres of the late blight-susceptible tomato cultivar. Simultaneously, the enrichment of pathogenic microorganisms, such as Neocosmospora and Plectosphaerella, was also detected in the rhizospheres of the susceptible tomato cultivar. Moreover, no enrichment of pathogenic microorganisms occurred in the late blight-resistant tomato cultivar after P. infestans inoculation. These findings suggest that these traits serve as effective defense mechanisms against pathogen invasion in resistant tomato cultivar. Overall, this study provides a comprehensive analysis of the rhizosphere microbial community structures in late blight-resistant and -susceptible tomato cultivars under natural conditions and their response following pathogen inoculation. Additionally, potential antagonistic microorganisms against late blight were also identified. The findings offer valuable insights for effective late blight management in tomatoes and contribute to the development of sustainable agricultural practices.
Tomato (Solanum lycopersicum L.) is a crop of considerable economic importance worldwide, and it continues to grow in terms of both cultivation areas and yields in China. However, late blight caused by the pathogenic Phytophthora infestans poses a serious threat to sustainable tomato production. This disease is highly destructive and causes large-scale crop losses, with severe economic consequences for agricultural production (Arafa et al., 2022). Currently, the main strategies for controlling late blight, both domestically and internationally, include physical control, chemical treatment, and the development of disease-resistant cultivars (Ivanov et al., 2021). However, traditional methods have several limitations. Physical control is cumbersome and inefficient; hence, implementation on a large scale is challenging. Although chemical control through pesticide application effectively suppresses diseases in the short term, prolonged use results in environmental pollution and potential risks to food safety (Liu et al., 2018). Furthermore, excessive pesticide use triggers pathogens to develop resistance, thereby enhancing their mutation rates and pathogenicity (Tang et al., 2021). Although breeding disease-resistant cultivars is an ideal long-term strategy, the process is time-consuming and often takes several years or even decades (Galeano Garcia et al., 2018). Therefore, low-cost, environmentally friendly, and sustainable solutions for controlling late blight are urgently required. Among these alternatives, biological control has gained notable attention because of its eco-friendly and sustainable nature. Antagonistic microorganisms and microbial inoculants are essential for controlling plant diseases in sustainable agriculture (Azcón-Aguilar and Barea, 1997). Therefore, exploring beneficial microbe-plant interactions and their application in agricultural production is critical because it may reduce the need for fertilizers and pesticides, which subsequently contributes to more sustainable agricultural practices (Qu et al., 2020).
The rhizosphere microbiome is often referred to as the “second genome” or “extended genotype” of the plant (Banerjee and van der Heijden, 2023). Rhizosphere microorganisms play a crucial role in promoting plant growth and development, particularly by facilitating the absorption of mineral nutrients, enhancing plant disease resistance, and improving tolerance to abiotic and biotic stressors (Trivedi et al., 2021; Bai et al., 2022; Zhang et al., 2019). Specifically, rhizosphere microorganisms promote nutrient uptake by releasing available nutrients into the soil via primary or secondary metabolites. In addition, rhizosphere microorganisms enhance plant disease resistance by inhibiting the growth of pathogens or inducing systemic resistance in plants (Stringlis et al., 2018). However, pronounced differences in rhizosphere microbial community structures have been observed among different genotypes, even within the same crop species (Walters et al., 2018). Stress tolerance is an important factor that influences rhizosphere microbial community structure. For instance, the varying resistance of kiwifruit to bacterial canker disease (Fu et al., 2024), and that of tobacco (Zhao et al., 2023; Xia et al., 2024; Yang et al., 2024), mulberry (Dong et al., 2021), and tomato (Choi et al., 2020) to bacterial wilt are linked to differences in microbial community composition.
Throughout their growth cycle, plants are exposed to various abiotic and biotic stresses, such as insect herbivory and pathogen infections. Plants have developed resistance mechanisms to cope with these challenges (Song et al., 2021; Liu et al., 2023). One strategy involves altering the rhizosphere microbial community in response to stress. Plants tend to recruit specific functional microbial groups to enhance their ability to withstand adverse environmental conditions (Chen et al., 2018). For example, plants may recruit nutrient-cycling microbes under nutrient-deficient conditions and disease-suppressing microbes during pathogen attacks (Xun et al., 2021). For instance, maize selectively enhances the colonization of phosphate-solubilizing microbes in the rhizosphere under low-phosphorus conditions (Shao et al., 2021), and rice recruits rhizosphere microorganisms to improve salt tolerance in saline soils (Lian et al., 2020). Similarly, wheat (Yin et al., 2021), maize (Xun et al., 2023), and cabbage (Ping et al., 2024) selectively enrich antagonistic microorganisms in their rhizospheres in response to pathogen attacks.
The exploration of specific functional microorganisms and the “bottom-up” regulatory mechanisms of the rhizosphere microbiome on plant phenotypic traits is of great significance for leveraging beneficial rhizosphere microbes to develop sustainable agriculture. In the case of late blight caused by P. infestans, previous research primarily focused on the genetic resistance of tomato cultivars and their direct interaction with the pathogen, few studies have directly linked the composition and function of rhizosphere microbiomes to the suppression of P. infestans. Unlike previous studies, which primarily focused on pathogen-host interactions, we not only investigated how differences in rhizosphere microbiomes between tomato cultivars with varying late-blight resistance under natural conditions, but also analyzed how they responded to pathogen inoculation. The aim of this study was to elucidate the mechanisms of soil microbiomes in the rhizospheres of different tomato cultivars with varying late-blight resistance and they how respond to pathogen attacks. Furthermore, we also wanted to identify antagonistic microorganisms for the biocontrol of late blight. Our results will contribute to the more precise and integrated management and bio-controlling of tomato late blight disease.
The experiment was conducted from October 2023 to January 2024 in a greenhouse at the Vegetable Teaching and Experimental Base of the Agricultural College, Guangxi University (108°17′25″E, 22°51′02″N). The following tomato cultivars were used in this study: ‘Wild Tomato A3’ and ‘Zixia’. The ‘Zixia’ cultivar is highly susceptible to late blight, whereas ‘Wild Tomato A3’ exhibits notable resistance to the disease.
Initially, the seed surfaces were sterilized, soaked, and allowed to germinate before being sown in seedling trays for 30 days of growth. The seedlings were cultivated in pots (30 cm in height, 25 cm in diameter) containing soil with the following properties: pH 6.33, organic matter content 10.56 g·kg-1, total nitrogen 0.89 g·kg-1, phosphorus 0.78 g·kg-1, and potassium 8.51 g·kg-1. Available nitrogen, phosphorus, and potassium levels were 20.27, 1.68, and 102.8 mg·kg-1, respectively. After 14 days of growth in the pots, the plants were inoculated with late blight pathogen Phytophthora infestans. The inoculum was prepared by culturing P. infestans on potato dextrose agar medium, and the sporangia were collected. The inoculation procedure involved spraying 10 mL of a 1 × 106 spores/mL solution evenly on each plant. Controls were sprayed with 10 mL of distilled water instead of the inoculum. The plants were then incubated under conditions conducive to disease development (16°C, 90%–95% relative humidity, and 12-h photoperiod). The following four treatment groups were investigated: late blight-resistant tomato cultivar without P. infestans inoculation (A3); tomato cultivar highly susceptible to late blight without P. infestans inoculation (ZX), late blight-resistant tomato cultivar with P. infestans inoculation (A3P), and tomato cultivar highly susceptible to late blight with P. infestans inoculation (ZXP).
The rhizosphere soil samples were collected on January 18, 2024. Three tomato plants were randomly selected for each treatment. With the tomato plant in the center, the surrounding soil was loosened using a small sterilized spatula, and the tomato plant was carefully uprooted. Large clods were shaken off and the soil attached to the tomato roots was collected as a rhizosphere soil sample (Yang et al., 2023). The rhizosphere soil samples were placed in pre-labeled sterile self-sealing bags and placed in an icebox with ice packs for transport to the laboratory. In the laboratory, impurities were removed from the rhizosphere soil samples, and the soil was sieved through a 2-mm stainless steel mesh for rhizosphere soil microbial analysis (Ling et al., 2015). Each treatment included three biological replicates for rhizosphere soil microbial analysis.
The total DNA was extracted from the soil samples using the Fast DNA® Spin Kit for Soil (MP Biomedicals, Thomas Irvine, CA, USA) according to the manufacturer’s instructions. The extracted DNA was examined on 1% agarose gel to verify extraction quality, followed by gel electrophoresis. The concentration and purity of the extracted DNA were determined using a NanoDrop 2000 spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA) (Bulgarelli et al., 2012). Following extraction, the DNA was stored at −20°C for further processing. PCR amplification was performed on an ABI GeneAmp® 9700 (ABI, Los Angeles, CA, USA) using specific primers and sequence types. Targeting the key regions within microbial genomes, we amplified the V3–V4 hypervariable region of the 16S rRNA gene for bacteria and the ITS1 region for fungi. This was achieved using specific primer pairs: 338F (5′-ACTA3PTACGGGAGGCAGCAG-3′) and 806R (5′-GGACTACHVGGGTWTCTAAT-3′) for bacterial samples (Lundberg et al., 2012), and ITS1F (5’-CTTGGTCATTTAGAGGAAGTAA-3’) and ITS2R (5’-GCTGCGTTCTTCATCGATGC-3’) for fungal samples (Adams et al., 2013). The PCR protocol involved an initial denaturation at 95°C for 3 min, followed by denaturation cycles (16S rRNA gene: 27 cycles; ITS gene: 35 cycles) at 95°C for 30 s, annealing at 55°C for 30 s, and extension at 72°C for 72 s. This was concluded with a final extension at 72°C for 10 min and termination at 4°C. The PCR products were detected and recovered using 2% agarose gel electrophoresis. The recovered products were purified using an AxyPrep DNA Gel Extraction Kit (Axygen Biosciences, Union City, CA, USA). The purified products were again detected using 2% agarose gel electrophoresis, and a Quantus™ Fluorometer (Promega, Madison, WI, USA) was subsequently used to ensure the integrity and purity of the samples.
Purified amplicons were pooled in equimolar amounts and were then paired-end sequenced on Illumina MiSeqPE300 (bacteria) and MiSeqPE250 (fungi) platforms (Illumina, San Diego, USA) by Majorbio Bio-Pharm Technology Co. Ltd. (Shanghai, China) according to standard protocols. The raw data for the soil bacterial and fungal sequences were deposited in the NCBI Sequence Read Archive database under the accession numbers PRJNA1154786 and PRJNA1154788, respectively.
Raw FASTQ files were demultiplexed using an in-house Perl script, quality-filtered using fastp version 0.19.6, and merged using FLASH version 1.2.7. The following criteria were used: (i) the reads were truncated at any site receiving an average quality score of <20 over a 50-bp sliding window, truncated reads shorter than 50 bp were discarded, and reads containing ambiguous characters were also discarded; (ii) only overlapping sequences longer than 10 bp were assembled according to their overlapped sequence, the maximum mismatch ratio in the overlapping region was 0.2, and reads that could not be assembled were discarded; (iii) samples were distinguished according to the barcode and primers, and the sequence direction was adjusted for exact barcode matching and two nucleotide mismatches in primer matching. The optimized sequences were then clustered into operational taxonomic units (OTUs) using UPARSE 7.1 with a 97% sequence similarity level. The most abundant sequence of each OTU was selected as the representative sequence. The taxonomy of each OTU representative sequence was analyzed using RDP Classifier version 2.11 against the 16S rRNA gene database (Release138 http://www.arb-silva.de) and fungi Unite database (Release 8.0 http://unite.ut.ee/index.php) using a confidence threshold of 0.7 (Zhou et al., 2024).
Bioinformatic analysis of the soil microbiota was performed using the Majorbio Cloud platform (https://cloud.majorbio.com). Based on OTU information, alpha diversity indices, including observed OTUs, Shannon index, and Chao richness, were calculated using Mothur v1.30.1. Similarities among the microbial communities in different samples were determined using principal coordinate analysis (PCoA) based on Bray–Curtis dissimilarity using the Vegan v2.5-3 package. Linear discriminant analysis (LDA) effect size (LEfSe) (http://huttenhower.sph.harvard.edu/LEfSe) was used to identify significantly abundant taxa (phylum to genera) of bacteria and fungi among the different groups (LDA score > 3.5, P < 0.05). FAPROTAX software (1.2.1) was used to predict the functions of the bacterial communities, whereas FUNGuild software (http://www.funguild.org/) was used to predict the functions of fungal communities.
As shown in Figure 1A, in the absence of P. infestans inoculation, the Shannon index of the soil bacterial communities in the rhizospheres of A3 was markedly higher than that of ZX. However, the Chao index of soil bacterial richness in the rhizospheres was not significantly different between A3 and ZX (Figure 1B). The Shannon and Chao indices of the soil fungal communities in the rhizospheres of A3 were lower than those of ZX; however, no significant differences were observed (Figures 1C, D).
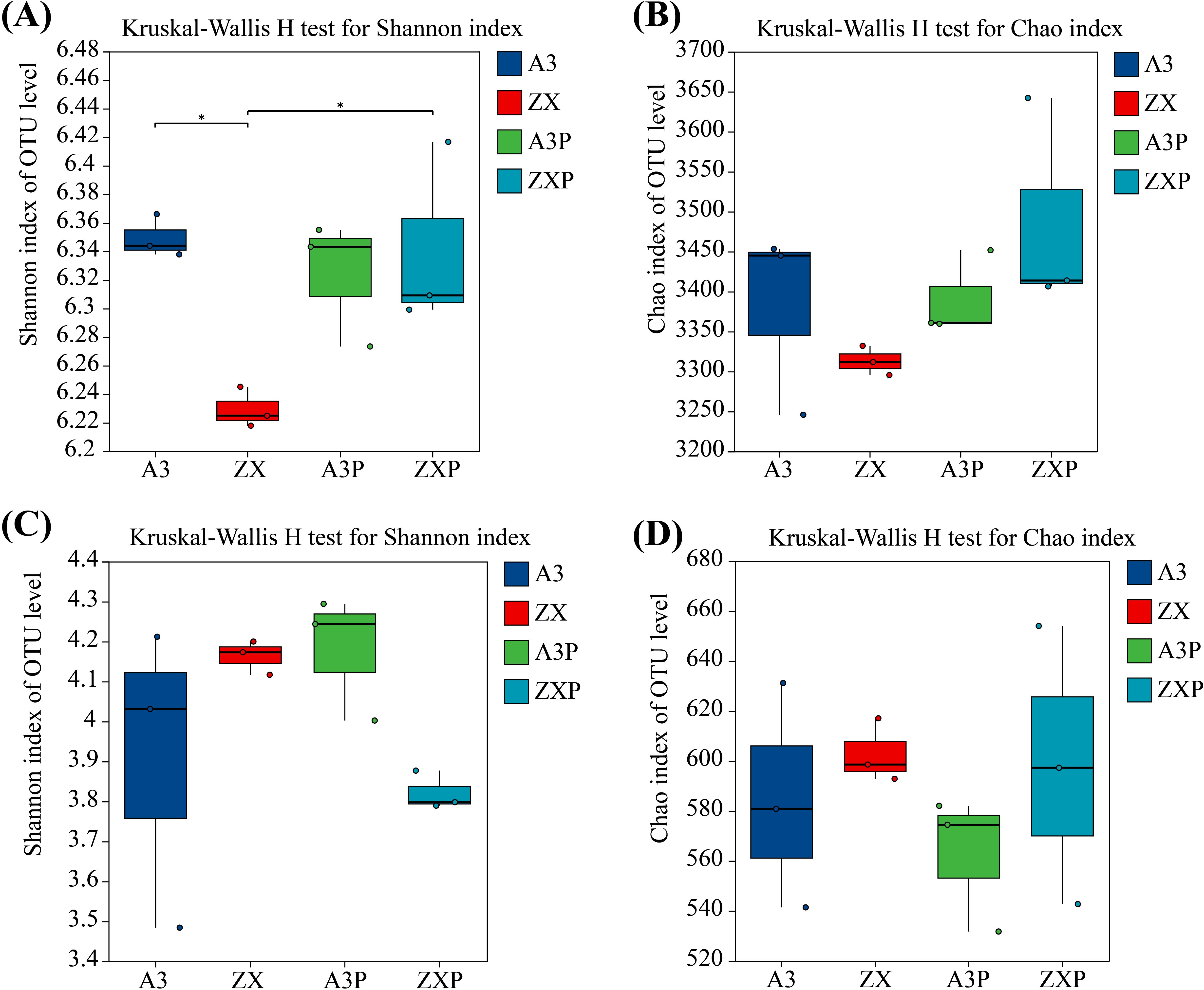
Figure 1. Alpha diversity index of the bacterial (A, B) and fungal (C, D) communities in the rhizospheres of A3, ZX, A3P, and ZXP plants. A3, late blight-resistant tomato cultivar without Phytophthora infestans inoculation; ZX, tomato cultivar highly susceptible to late blight without P. infestans inoculation; A3P, late blight-resistant tomato cultivar with P. infestans inoculation; ZXP, tomato cultivar highly susceptible to late blight with P. infestans inoculation.
Additionally, under P. infestans inoculation, the Shannon index of the soil bacterial and fungal communities in the rhizospheres of A3P was higher than that of ZXP; however, no significant differences were observed between them (Figures 1A, C). In addition, the Chao indices of soil bacterial and fungal richness in the rhizospheres were not significantly different between A3P and ZXP (Figures 1B, D).
These results indicated that under natural conditions, the bacterial diversity in the rhizospheres of the resistant cultivar (A3) was markedly higher than that of the susceptible cultivar (ZX). However, bacterial richness and fungal diversity and richness in the rhizospheres were not significantly different from each other. After P. infestans inoculation, bacterial diversity and richness in the rhizospheres of the resistant cultivar (A3P) decreased, whereas bacterial diversity in the rhizospheres of the susceptible cultivar (ZX) markedly increased. Moreover, after P. infestans inoculation, soil fungal diversity increased, whereas soil fungal richness decreased in the rhizospheres of the resistant cultivar (A3P). In contrast, soil fungal diversity decreased; however, their richness did not significantly change in the rhizospheres of the susceptible cultivar (ZXP).
Based on the Bray–Curtis distance, PCoA was used to visualize the compositional characteristics of the bacterial and fungal communities in the rhizospheres of A3, ZX, A3P, and ZXP plants. Distinct clustering of rhizospheric bacterial and fungal communities was observed among A3, ZX, A3P, and ZXP plants (Figure 2). Further analysis should be conducted to explore the specific differences in community composition.
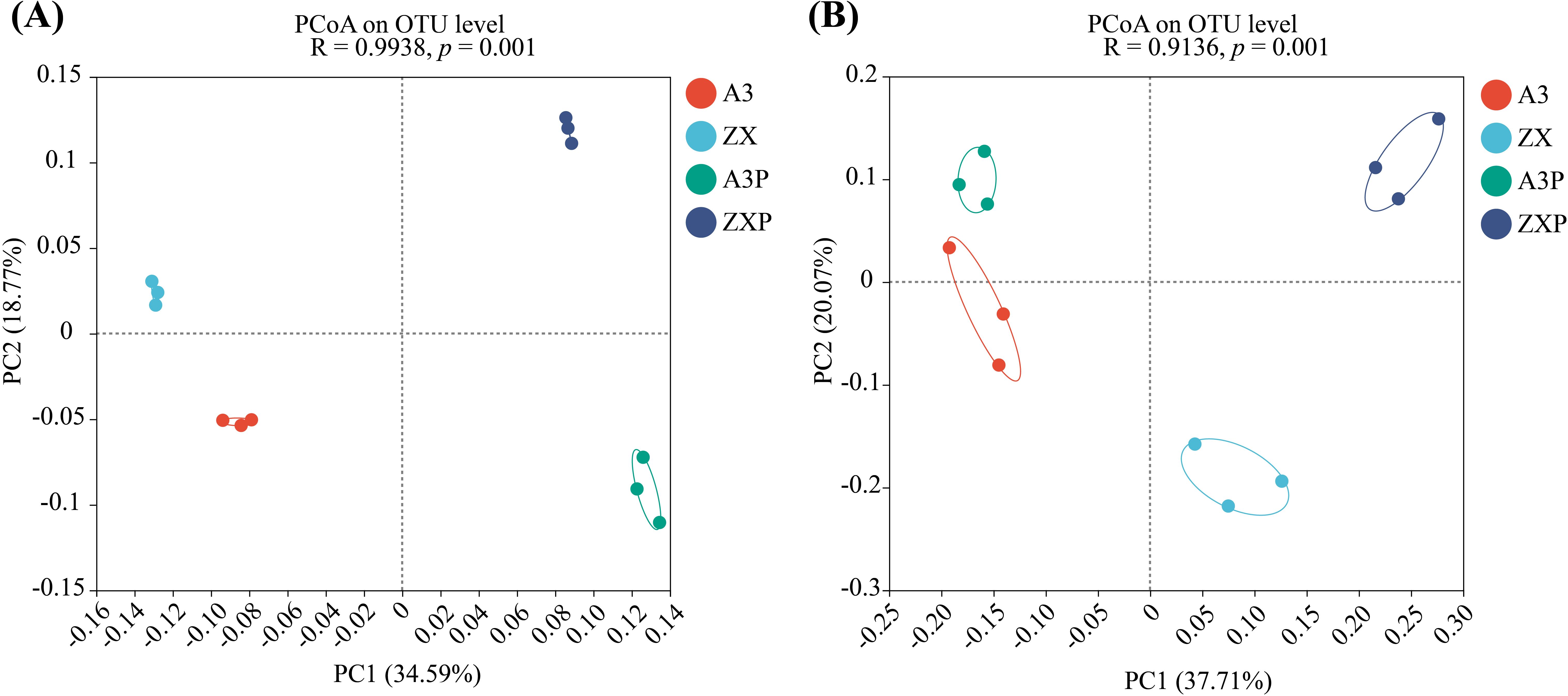
Figure 2. Beta diversity indexes of the (A) bacterial and (B) fungal communities in the rhizospheres of A3, ZX, A3P, and ZXP plants. A3, late blight-resistant tomato cultivar without Phytophthora infestans inoculation; ZX, tomato cultivar highly susceptible to late blight without P. infestans inoculation; A3P, late blight-resistant tomato cultivar with P. infestans inoculation; ZXP, tomato cultivar highly susceptible to late blight with P. infestans inoculation; PCoA, principal coordinate analysis; OTU, operational taxonomic unit; PC, principal component.
As shown in Figure 3A, at the phylum level, the top five dominant bacterial phyla (relative abundance ≥1%) were consistent, namely Proteobacteria, Actinobacteriota, Firmicutes, Patescibacteria, and Chloroflexi. However, other dominant bacterial phyla, such as Desulfobacterota, were the unique dominant soil bacterial phyla in the rhizospheres of A3P, and Myxococcota was absent in the rhizospheres of ZX.
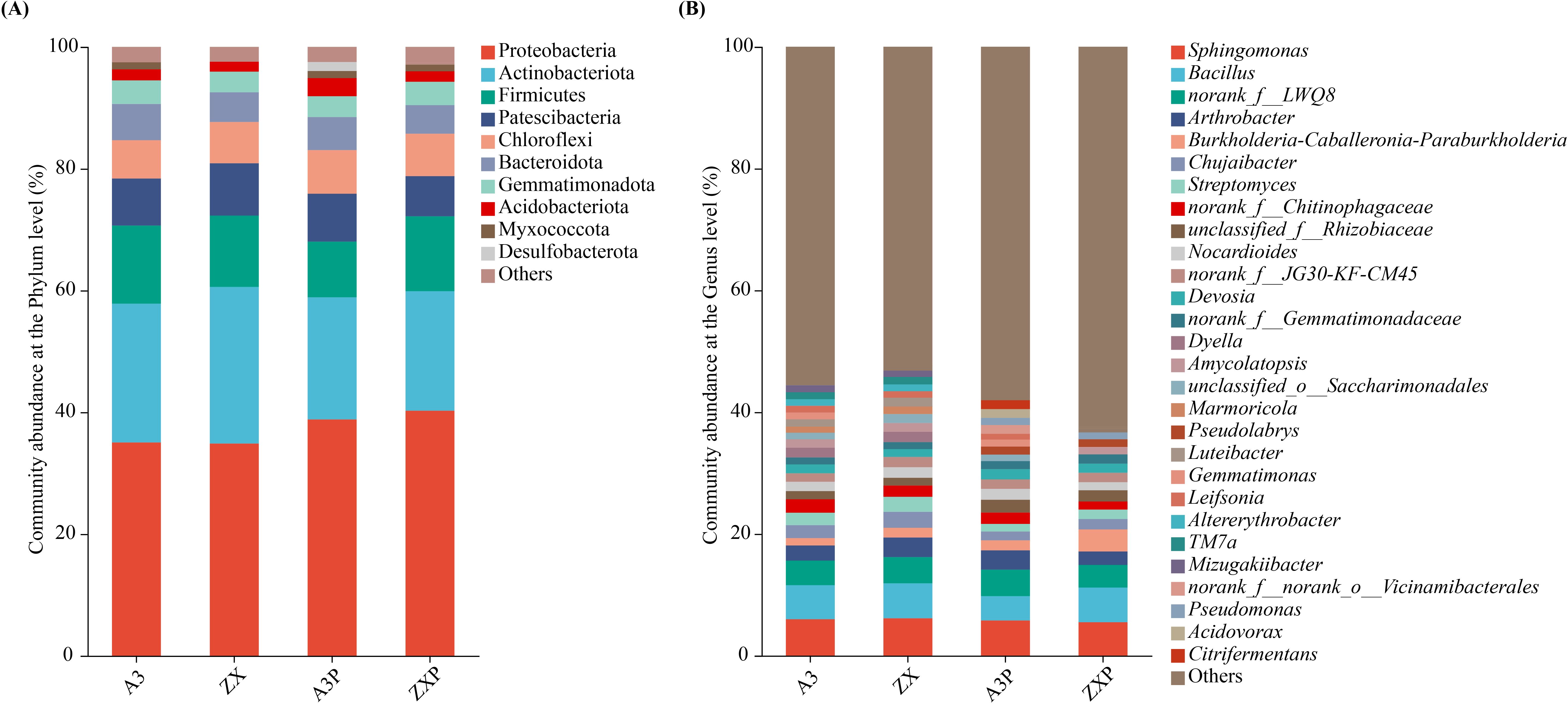
Figure 3. Rhizospheric bacterial compositions at the phylum (A) and genus (B) levels among A3, ZX, A3P, and ZXP. A3, late blight-resistant tomato cultivar without Phytophthora infestans inoculation; ZX, tomato cultivar highly susceptible to late blight without P. infestans inoculation; A3P, late blight-resistant tomato cultivar with P. infestans inoculation; ZXP, tomato cultivar highly susceptible to late blight with P. infestans inoculation.
As shown in Figure 3B, the top 10 dominant soil bacterial genera (relative abundance ≥1%) in the rhizospheres of A3 were Sphingomonas, Bacillus, norank_f:LWQ8, Arthrobacter, norank_f:Chitinophagaceae, Chujaibacter, Streptomyces, Dyella, Nocardioides, and Devosia. In contrast, Sphingomonas, Bacillus, norank_f:LWQ8, Arthrobacter, Chujaibacter, Streptomyces, norank_f:Chitinophagaceae, Nocardioides, norank_f:JG30-KF-CM45, and Dyella were the top 10 dominant soil bacterial genera in the rhizospheres of ZX. Additionally, Sphingomonas, norank_f:LWQ8, Bacillus, Arthrobacter, unclassified_f:Rhizobiaceae, norank_f:Chitinophagaceae, Nocardioides, Devosia, Burkholderia-Caballeronia-Paraburkholderia, and norank_f:JG30-KF-CM45 were the top 10 dominant soil bacterial genera in the rhizospheres of A3P. In contrast, Bacillus, Sphingomonas, norank_f:LWQ8, Burkholderia-Caballeronia-Paraburkholderia, Arthrobacter, unclassified_f:Rhizobiaceae, Chujaibacter, Streptomyces, norank_f:Gemmatimonadaceae, and norank_f:JG30-KF-CM45 were the top 10 dominant soil bacterial genera in the rhizospheres of ZXP.
These results suggest that soil bacterial composition in the rhizospheres of tomato cultivars with different resistance abilities to late blight could be altered by P. infestans inoculation. In particular, the relative abundances of Firmicutes and Patescibacteria decreased in the rhizospheres of A3P; however, they increased in the rhizospheres of ZXP. Moreover, Citrifermentans was uniquely enriched in the rhizospheres of A3P. In contrast, the relative abundance of Burkholderia-Caballeronia-Paraburkholderia markedly increased by 125.6% in ZXP compared to that in ZX. Moreover, Pseudolabrys and Pseudomonas were enriched in the rhizospheres of different tomato cultivars only after P. infestans inoculation.
As shown in Figure 4A, Ascomycota, Basidiomycota, unclassified_k:Fungi, Olpidiomycota, and Chytridiomycota were the top five common soil dominant fungal phyla (with relative abundance ≥1%) in the rhizospheres of tomato cultivars with varying resistance to P. infestans.
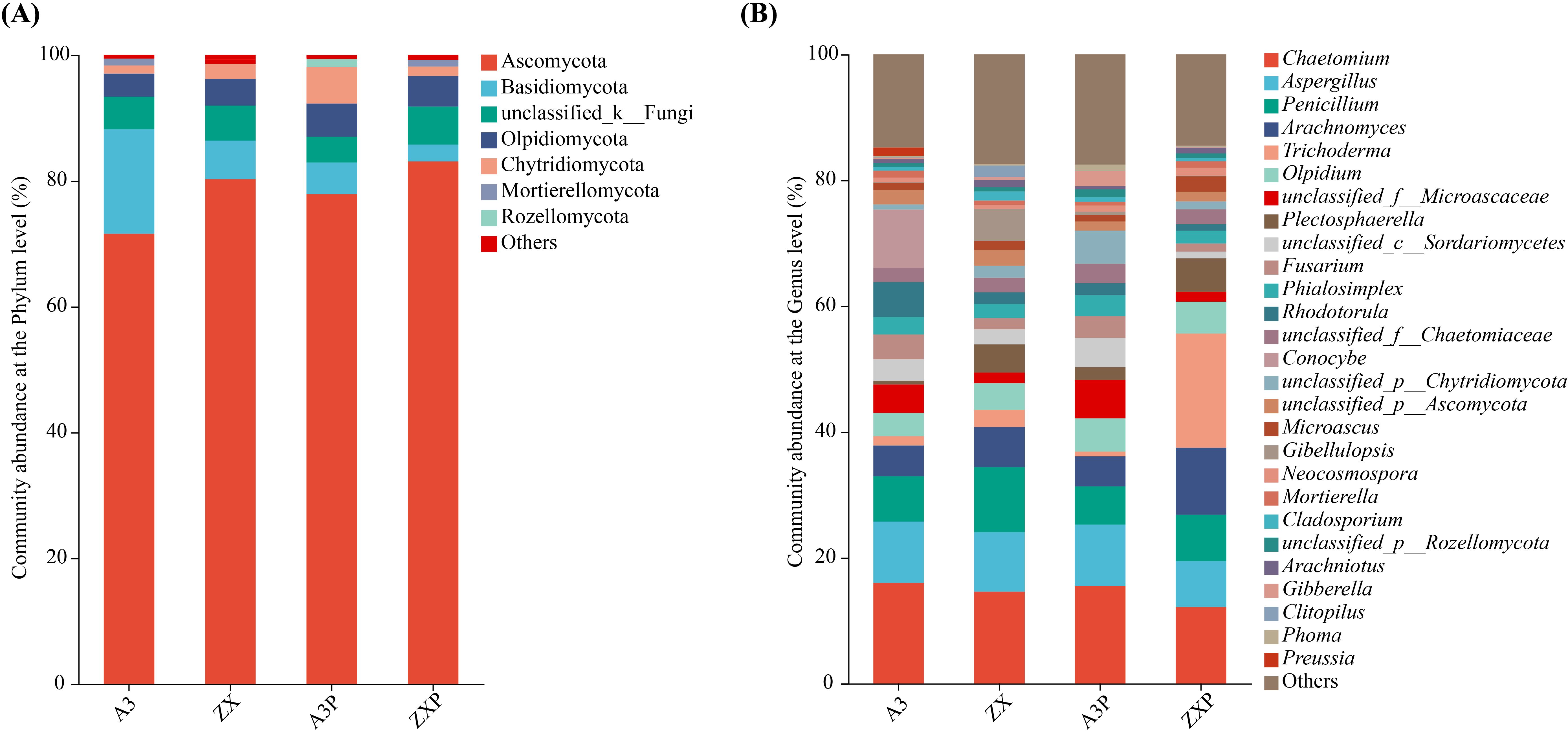
Figure 4. Rhizospheric fungal compositions at the phylum (A) and genus (B) levels among A3, ZX, A3P, and ZXP. A3, late blight-resistant tomato cultivar without Phytophthora infestans inoculation; ZX, tomato cultivar highly susceptible to late blight without P. infestans inoculation; A3P, late blight-resistant tomato cultivar with P. infestans inoculation; ZXP, tomato cultivar highly susceptible to late blight with P. infestans inoculation.
As shown in Figure 4B, the top 10 dominant soil fungal genera (with relative abundance ≥1%) in the rhizospheres of A3 were Chaetomium, Aspergillus, Conocybe, Penicillium, Rhodotorula, Arachnomyces, unclassified_f:Microascaceae, Fusarium, Olpidium, and unclassified_c:Sordariomycetes. In contrast, Chaetomium, Penicillium, Aspergillus, Arachnomyces, Gibellulopsis, Plectosphaerella, Olpidium, Trichoderma, unclassified_p:Ascomycota, and unclassified_c:Sordariomycetes were the top 10 dominant soil fungal genera in the rhizospheres of ZX. Additionally, Chaetomium, Aspergillus, unclassified_f:Microascaceae, Penicillium, Olpidium, unclassified_p:Chytridiomycota, Arachnomyces, unclassified_c:Sordariomycetes, Fusarium, and Phialosimplex were the top 10 dominant soil fungal genera in the rhizospheres of A3P. In contrast, Trichoderma, Chaetomium, Arachnomyces, Penicillium, Aspergillus, Plectosphaerella, Olpidium, Microascus, unclassified_f:Chaetomiaceae, and Phialosimplex were the top 10 dominant soil fungal genera in the rhizospheres of ZXP.
These results showed that the relative abundances of Ascomycota and Olpidiomycota increased, and the relative abundance of Basidiomycota decreased in the rhizospheres of A3P and ZXP. Furthermore, the relative abundance of Chytridiomycota markedly increased in the rhizospheres of A3P, whereas it declined in the rhizospheres of ZXP plants. Moreover, Rozellomycota is a unique soil-dominant fungal phylum in the rhizospheres of A3P. At the genus level, Conocybe was specifically enriched only in the rhizospheres of A3. However, after inoculation with P. infestans, Conocybe was absent. Gibellulopsis was enriched in the ZX rhizospheres. After P. infestans inoculation, the relative abundance of Trichoderma in ZXP rhizospheres increased by 607.75% compared to that in ZX rhizospheres.
As shown in Figure 5A, 6, 16, 14, and 9 bacterial taxa were identified as biomarkers in the A3, ZX, A3P, and ZXP rhizospheres, respectively. At the phylum level, Bacteroidetes, Actinobacteria, Desulfobacterota, and Proteobacteria were identified as biomarkers in the rhizospheres of A3, ZX, A3P, and ZXP, respectively. At the genus level, norank_f:Chitinophagaceae was identified as a biomarker in the A3 rhizospheres, and Streptomyces, Chujaibacter, and Dyella were identified as biomarkers of the ZX rhizospheres. Acidovorax, Pseudomonas, unclassified_f:Rhizobiaceae, and Azospirillum were identified as biomarkers in the A3P rhizospheres, whereas Flavobacterium, Pseudolabrys, and Burkholderia-Caballeronia-Paraburkholderia were identified as biomarkers in the rhizospheres of the ZXP plants.
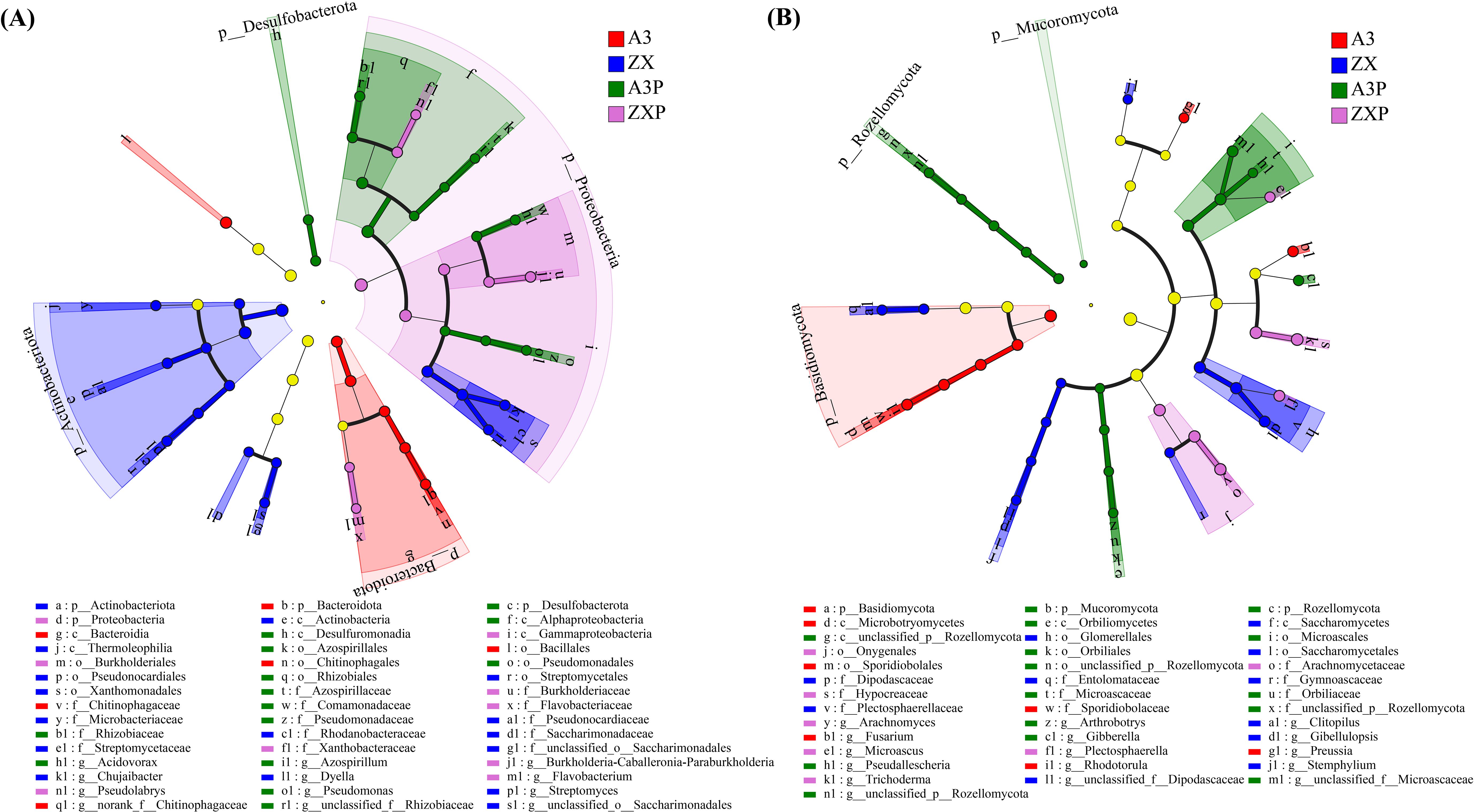
Figure 5. LEfSe analysis of significant abundance of rhizospheric bacteria (A) and fungi (B) among A3, ZX, A3P, and ZXP plants. A3, late blight-resistant tomato cultivar without Phytophthora infestans inoculation; ZX, tomato cultivar highly susceptible to late blight without P. infestans inoculation; A3P, late blight-resistant tomato cultivar with P. infestans inoculation; ZXP, tomato cultivar highly susceptible to late blight with P. infestans inoculation; LEfSe, linear discriminant analysis effect size.
As shown in Figure 5B, 7, 11, 15, and 7 fungal taxa were identified as biomarkers in the A3, ZX, A3P, and ZXP rhizospheres, respectively. Basidiomycota were markedly enriched in the rhizospheres of A3, whereas Mucoromycota and Rozellomycota were considerably enriched in the rhizospheres of A3P. At the genus level, Rhodotorula, Fusarium, and Preussia were significantly enriched in the rhizospheres of A3, whereas Gibellulopsis, Clitopilus, and Stemphylium were notable enriched in the rhizospheres of ZX. Gibberella, Arthrobotrys, and Pseudallescheria were markedly enriched in the A3P rhizospheres. In contrast, Trichoderma, Arachnomyces, Plectosphaerella, and Microascus were considerably enriched in the rhizospheres of the ZXP plants.
As shown in Figure 6, the bacterial communities in the A3, ZX, A3P, and ZXP rhizospheres exhibited the highest expression levels during chemoheterotrophy and aerobic_chemoheterotrophy. After inoculation with P. infestans, the expression levels of the following 10 functions markedly increased in both A3P and ZXP: knallgas_bacteria, dark_hydrogen_oxidation, nitrate_denitrification, nitrous_oxide_denitrification, nitritedenitrification, denitrification, nitriterespiration, nitrogenrespiration, ureolysis, and nitraterespiration. In addition, the expression levels of A3P were markedly higher than those of ZXP. The expression levels of plant pathogens, nitrogenfixation, and nitrate_reduction also showed notable increasing trends after inoculation, although there were no significant differences between A3P and ZXP. In A3P rhizospheres, the expression levels of cellulolysis and Mn oxidation were considerably lower than those in A3.
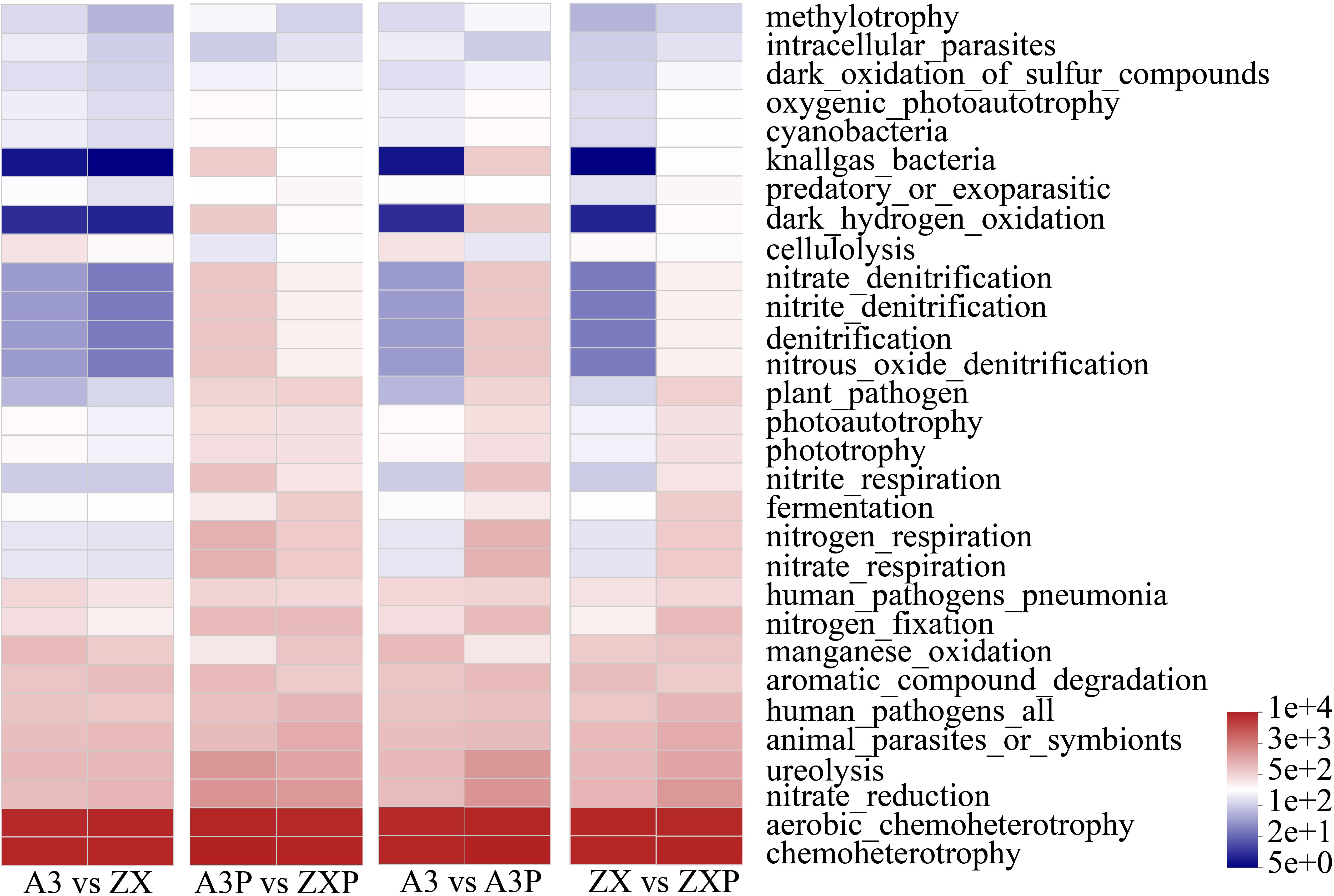
Figure 6. FAPROTAX functional prediction of rhizospheric bacteria in A3, ZX, A3P, and ZXP plants. A3, late blight-resistant tomato cultivar without Phytophthora infestans inoculation; ZX, tomato cultivar highly susceptible to late blight without P. infestans inoculation; A3P, late blight-resistant tomato cultivar with P. infestans inoculation; ZXP, tomato cultivar highly susceptible to late blight with P. infestans inoculation.
FUNGuild predicted the trophic and functional guilds of fungal communities in the A3, ZX, A3P, and ZXP rhizospheres. The rhizospheric fungal communities from all four treatments could be classified into ten guilds. Undefined Saprotroph, Dung Saprotroph, Animal Parasite-Fungal Parasite, Plant Pathogen, Animal Endosymbiont-Animal, Pathogen-Endophyte-Plant, and Pathogen-Undefined Saprotroph were common across all treatments. Among these, the relative abundance of Undefined Saprotrophs decreased considerably after pathogen inoculation, with a more pronounced reduction observed in the resistant varieties. The relative abundance of Plant Pathogens in the rhizospheres of ZX was markedly higher than that in A3, whereas the relative abundance in the rhizospheres of A3P was markedly higher than that in A3. However, the relative abundance in the rhizospheres of ZXP was considerably lower than that in the rhizospheres of ZX. Additionally, Dung Saprotroph-Plant Saprotroph was unique to the rhizospheres of A3, Plant Pathogen-Wood Saprotroph and Plant Saprotroph were specific to the rhizospheres of ZX, and fungal parasites were unique to the rhizospheres of A3P. The Undefined Saprotroph-Wood Saprotroph guild was only found in the rhizospheres of A3P and ZXP (Figure 7).
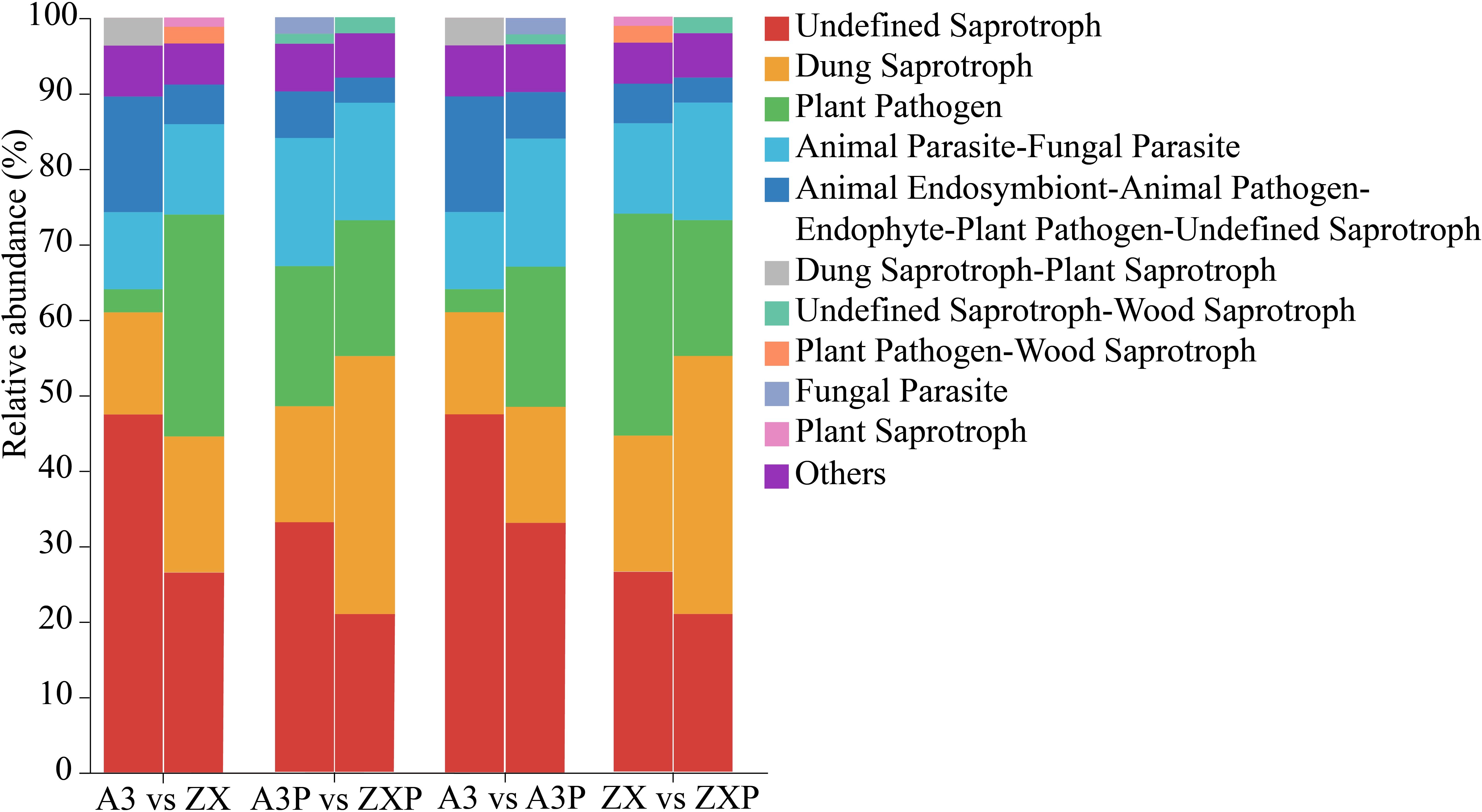
Figure 7. FUNGuild functional predictions of rhizospheric fungi in A3, ZX, A3P, and ZXP plants. A3, late blight-resistant tomato cultivar without Phytophthora infestans inoculation; ZX, tomato cultivar highly susceptible to late blight without P. infestans inoculation; A3P, late blight-resistant tomato cultivar with P. infestans inoculation; ZXP, tomato cultivar highly susceptible to late blight with P. infestans inoculation.
Soil microbes in the rhizosphere are closely associated with plant resistance to stress (Oyserman et al., 2022). Moreover, the inherent natural microbiome of a plant confers resistance at both the individual and community levels (Carrión et al., 2019; Li et al., 2022). According to the well-known “cry-for-help hypothesis,” plants under pathogen attack secrete specific root exudates that promote colonization of beneficial microbes in the rhizosphere (Rolfe et al., 2019). Furthermore, when plants are exposed to adverse conditions, such as drought, salinity, alkalinity, pathogens, or pests, rhizosphere microbes engage in symbiotic interactions that enhance their ability to adapt to these stresses (Philippot et al., 2013; Li et al., 2020). Therefore, understanding the assembly of the rhizosphere microbiome is crucial to elucidate its role in plant stress responses. Therefore, in this study, tomato cultivars with varying resistance to late blight were selected to analyze their natural rhizosphere microbiomes and their responses to pathogen inoculation.
First, we observed that soil bacterial diversity in the rhizospheres of the resistant tomato cultivar was considerably higher than that in the rhizospheres of the susceptible cultivar. In contrast, the soil fungal diversity in the rhizospheres of the susceptible cultivar was higher than that of the resistant cultivar. Similarly, bacterial diversity in Chinese wheat varieties resistant to wheat yellow mosaic virus is higher than that in susceptible varieties (Wu et al., 2021). Additionally, the rhizospheres of Fusarium head blight-resistant wheat varieties exhibited markedly higher bacterial diversity than that of susceptible varieties, whereas fungal diversity was lower in the rhizospheres of resistant varieties than in their susceptible counterparts (Li et al., 2023). Moreover, after P. infestans inoculation, the rhizosphere bacterial diversity of the susceptible cultivar markedly increased, whereas the changes in the resistant varieties were less pronounced. Soil bacterial diversity was considerably increased in the rhizospheres of diseased plants compared to that in healthy plants (Huang et al., 2019). Simply, P. infestans inoculation failed to overcome the defense mechanisms of the resistant tomato cultivar, leaving the plants unaffected by the pathogen. Consequently, the rhizosphere microbiomes of the resistant cultivar remained relatively stable. In contrast, the susceptible tomato cultivar was unable to resist pathogen infection, resulting in pronounced alterations in the rhizosphere microbial diversity. A more diverse and abundant rhizosphere bacterial community may be a key factor contributing to the inherent resistance of tomato cultivars to late blight.
Additionally, dominant soil bacterial genera, such as Dyella, Luteibacter, TM7a, Mizugakiibacter, Altererythrobacter, and Marmoricola, were lost in the rhizospheres of both late blight-resistant and -susceptible cultivars after P. infestans inoculation. This indicates that the soil microbial composition in the rhizospheres can be altered by P. infestans inoculation, regardless of the cultivar resistance. Some soil bacterial genera, such as Citrifermentans, norank_f:norank_o:Vicinamibacterales, and Acidovorax, were specifically enriched in the rhizospheres of the resistant tomato cultivar. Furthermore, LEfSe analysis identified Azospirillum, Pseudomonas, Acidovorax, and unclassified_f:Rhizobiaceae as biomarkers in the rhizospheres of the resistant tomato cultivar. Azospirillum synthesizes plant hormones (Iparraguirre et al., 2024) that promote plant growth and development (Lambrecht et al., 2000) and enhance plant stress resistance (Gray and Smith, 2005). Moreover, Acidovorax activates plant defense mechanisms and limits pathogen colonization (Emmenegger et al., 2023). Furthermore, Pseudomonas is a well-known plant growth-promoting rhizobacterium (PGPR) that inhibits fungal pathogens by releasing lytic enzymes, phenazines, and organic volatiles (De Vrieze et al., 2015; Zhang et al., 2015; Müller et al., 2016). Pseudomonas also induces plant defense mechanisms and promotes plant growth under stressful conditions (Kumar and Verma, 2018). In contrast, Pseudolabrys, Flavobacterium, and Burkholderia-Caballeronia-Paraburkholderia were identified as biomarkers in the rhizospheres of the susceptible tomato cultivar after P. infestans inoculation. Although Pseudolabrys is considered a PGPR that enhances plant growth and health (Peng et al., 2022), Flavobacterium also contributes to plant health (Lazcano et al., 2021; Zhou et al., 2022; Chen et al., 2023). The microbe can be isolated from soil and produces antimicrobial substances, such as hydrogen cyanide, chitinase, and siderophores (Belimov et al., 2005; Soltani et al., 2010; Kharade and McBride, 2014). Notably, some Flavobacterium species protect tomato plants from pathogenic infections by limiting sugar acquisition through efficient sugar metabolism (Kwak et al., 2018). Furthermore, Burkholderia-Caballeronia-Paraburkholderia exhibit plant growth-promoting abilities and antagonistic effects against pathogens (Dendooven et al., 2024). Our results suggest that both the resistant and susceptible tomato cultivars can recruit growth-promoting and biocontrol microorganisms into their rhizospheres upon P. infestans inoculation.
Additionally, fungal genera, including Gibberella, unclassified_p_Rozellomycota, and Phoma, were specifically enriched in the rhizospheres of the resistant tomato cultivar after P. infestans inoculation. In contrast, Neocosmospora was predominantly enriched in the rhizospheres of the susceptible tomato cultivar under the same conditions. Phoma protects crops from pathogens by inducing systemic resistance (Sultana et al., 2008), whereas Neocosmospora causes stem rot disease in various crops, including peanuts (Sun et al., 2012), mulberries (Zhu et al., 2024), dragon fruit (Zheng et al., 2018), and tomatoes (Kwon et al., 2017). LEfSe analysis further revealed that Gibberella, Arthrobotrys, and Pseudallescheria were identified as biomarkers in the rhizospheres of the resistant tomato cultivar inoculated with P. infestans. Arthrobotrys can control nematodes and induce plant defense responses against fungal pathogens (Bordallo et al., 2002). Pseudallescheria also produce bioactive secondary metabolites that combat various plant pathogens (Ko et al., 2010; Zhu et al., 2020). In contrast, Trichoderma, Arachnomyces, Plectosphaerella, and Microascus were identified as biomarkers in the rhizosphere of the susceptible tomato cultivar inoculated with P. infestans. Trichoderma and its secondary metabolites released into the rhizosphere influence plant growth and nutrition, induce systemic resistance, and contribute to pathogen biocontrol (Hermosa et al., 2012; Kotasthane et al., 2015; Zeilinger et al., 2016). Trichoderma has a strong ability to compete with pathogens for nutrients and space in the rhizosphere (Soltani et al., 2010). Additionally, Trichoderma produces antibiotics and hydrolytic enzymes, such as chitinase and β-1,3-glucanase, which partially degrade pathogen cell walls and lead to parasitism (Rai et al., 2016). In contrast, Plectosphaerella is considered to be responsible for diseases in various plants (Usami et al., 2015; Li et al., 2017; Yang L. et al., 2023).
Under natural conditions, soil bacterial composition and function in the rhizospheres of tomato cultivars with varying levels of resistance were similar, with no significant differences. However, after pathogen inoculation, the functional expression levels of the rhizosphere soil bacterial communities markedly increased for 10 bacterial functions: knallgas_bacteria, dark_hydrogen_oxidation, nitrate_denitrification, nitrous_oxide_denitrification, nitrite_denitrification, denitrification, nitrite_respiration, nitrogen_respiration, ureolysis, and nitrate_respiration. Notably, the expression levels in the rhizospheres of the resistant tomato cultivar were markedly higher than those in the susceptible tomato cultivar. In addition, under natural conditions, the relative abundance of pathotrophic fungi in the rhizospheres of susceptible tomato cultivar was considerably higher than that in the rhizospheres of resistant tomato cultivar. Conversely, after P. infestans inoculation, the relative abundance of saprotrophic fungi increased in the rhizospheres of the susceptible cultivar. Our results are consistent with those of previous studies that have reported an increase in the abundance of soil saprotrophic fungi in the rhizospheres of diseased plants (Wang et al., 2024).
These findings suggest that tomato resistance to late blight is closely linked to the functional profiles of the rhizosphere soil microbial communities. Under natural conditions, the low abundance of pathogenic microorganisms, combined with the highly expressed bacterial functions following pathogen inoculation, play a positive role in resisting pathogen invasion.
Under natural conditions, soil bacterial diversity in the rhizospheres of the late blight-resistant tomato cultivar was considerably higher than that in the susceptible cultivar, whereas fungal diversity was higher in the susceptible cultivar. Additionally, the low abundance of pathogenic microorganisms in the rhizospheres of the resistant cultivar coupled with the high functional expression of bacterial communities under P. infestans inoculation played a positive role in resisting pathogen invasion. Furthermore, Pseudomonas, Azospirillum, Acidovorax, Phoma, Arthrobotrys, Pseudallescheria, Pseudolabrys, Flavobacterium, Burkholderia-Caballeronia-Paraburkholderia, and Trichoderma were identified as functional microorganisms with potential roles in protection against late blight. In contrast, the enrichment of Neocosmospora and Plectosphaerella in the rhizospheres of the late blight-susceptible tomato cultivar may contribute to the exacerbation of late blight symptoms.
The datasets presented in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found below: https://www.ncbi.nlm.nih.gov/, PRJNA1206999 https://www.ncbi.nlm.nih.gov/, PRJNA1207005.
XZ: Data curation, Formal Analysis, Investigation, Software, Visualization, Writing – original draft, Writing – review & editing. LL: Data curation, Formal Analysis, Software, Supervision, Writing – original draft. KC: Data curation, Formal Analysis, Writing – original draft. YY: Data curation, Formal Analysis, Writing – original draft. LQ: Data curation, Formal Analysis, Writing – original draft. XL: Data curation, Formal Analysis, Writing – original draft. QL: Data curation, Formal Analysis, Writing – original draft. SY: Conceptualization, Funding acquisition, Project administration, Writing – original draft, Writing – review & editing.
The author(s) declare that financial support was received for the research, authorship, and/or publication of this article. This research was funded by the Innovation Project of Guangxi Graduate Education (YCSW2023038) and the Bama County Talent Science and Technology Project (Grant No. Barenke20220005).
The authors declare that this study was conducted in the absence of any commercial or financial relationships that could be construed as potential conflicts of interest.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
A3, late blight-resistant tomato cultivar without P. infestans inoculation; ZX, tomato cultivar highly susceptible to late blight without P. infestans inoculation; A3P, late blight-resistant tomato cultivar with P. infestans inoculation; ZXP, tomato cultivar highly susceptible to late blight with P. infestans inoculation; OTU, operational taxonomic unit; LDA, linear discriminant analysis; LEfSe, LDA effect size; PCoA, principal coordinate analysis; PCoA, principal coordinate analysis; PGPR, plant growth-promoting rhizobacterium.
Adams, R. I., Miletto, M., Taylor, J. W., Bruns, T. D. (2013). Dispersal in microbes: fungi in indoor air are dominated by outdoor air and show dispersal limitation at short distances. ISME J. 7, 1262–1273. doi: 10.1038/ismej.2013.28
Arafa, R. A., Kamel, S. M., Taher, D. I., Solberg, S.Ø., Rakha, M. T. (2022). Leaf extracts from resistant wild tomato can be used to control late blight (Phytophthora infestans) in the cultivated tomato. Plants 11, 1824. doi: 10.3390/plants11141824
Azcón-Aguilar, C., Barea, J. M. (1997). Arbuscular mycorrhizas and biological control of soil-borne plant pathogens – an overview of the mechanisms involved. Mycorrhiza 6, 457–464. doi: 10.1007/s005720050147
Bai, B., Liu, W., Qiu, X., Zhang, J., Zhang, J., Bai, Y. (2022). The root microbiome: Community assembly and its contributions to plant fitness. J. Integr. Plant Biol. 64, 230–243. doi: 10.1111/jipb.13226
Banerjee, S., van der Heijden, M. G. A. (2023). Soil microbiomes and one health. Nat. Rev. Microbiol. 21, 6–20. doi: 10.1038/s41579-022-00779-w
Belimov, A. A., Hontzeas, N., Safronova, V. I., Demchinskaya, S. V., Piluzza, G., Bullitta, S., et al. (2005). Cadmium-tolerant plant growth-promoting bacteria associated with the roots of Indian mustard (Brassica juncea L. Czern.). Soil Biol. Biochem. 37, 241–250. doi: 10.1016/j.soilbio.2004.07.033
Bordallo, J. J., Lopez-Llorca, L. V., Jansson, H.-B., Salinas, J., Persmark, L., Asensio, L. (2002). Colonization of plant roots by egg-parasitic and nematode-trapping fungi. New Phytol. 154, 491–499. doi: 10.1046/j.1469-8137.2002.00399.x
Bulgarelli, D., Rott, M., Schlaeppi, K., Ver Loren van Themaat, E., Ahmadinejad, N., Assenza, F., et al. (2012). Revealing structure and assembly cues for Arabidopsis root-inhabiting bacterial microbiota. Nature 488, 91–95. doi: 10.1038/nature11336
Carrión, V. J., Perez-Jaramillo, J., Cordovez, V., Tracanna, V., de Hollander, M., Ruiz-Buck, D., et al. (2019). Pathogen-induced activation of disease-suppressive functions in the endophytic root microbiome. Science 366, 606–612. doi: 10.1126/science.aaw9285
Chen, Y., Wang, J., Yang, N., Wen, Z., Sun, X., Chai, Y., et al. (2018). Wheat microbiome bacteria can reduce virulence of a plant pathogenic fungus by altering histone acetylation. Nat. Commun. 9, 3429. doi: 10.1038/s41467-018-05683-7
Chen, P., Zhang, J., Li, M., Fang, F., Hu, J., Sun, Z., et al. (2023). Synergistic effect of Bacillus subtilis and Paecilomyces lilacinus in alleviating soil degradation and improving watermelon yield. Front. Microbiol. 13. doi: 10.3389/fmicb.2022.1101975
Choi, K., Choi, J., Lee, P. A., Roy, N., Khan, R., Lee, H. J., et al. (2020). Alteration of bacterial wilt resistance in tomato plant by microbiota transplant. Front. Plant Sci. 11. doi: 10.3389/fpls.2020.01186
Dendooven, L., Pérez-Hernández, V., Navarro-Pérez, G., Tlalmis-Corona, J., Navarro-Noya, Y. E. (2024). Spatial and temporal shifts of endophytic bacteria in conifer seedlings of Abies religiosa (Kunth) Schltdl. & Cham. Microb. Ecol. 87, 90. doi: 10.1038/s41467-023-43793-z
De Vrieze, M., Pandey, P., Bucheli, T. D., Varadarajan, A. R., Ahrens, C. H., Weisskopf, L., et al. (2015). Volatile organic compounds from native potato-associated Pseudomonas as potential anti-oomycete agents. Front. Microbiol. 6. doi: 10.3389/fmicb.2015.01295
Dong, Z., Guo, Y., Yu, C., Zhixian, Z., Rongli, M., Deng, W., et al. (2021). The dynamics in microbial communities under bacterial wilt resistance by mulberry genotypes. Arch. Microbiol. 203, 1107–1121. doi: 10.1007/s00203-020-02098-1
Emmenegger, B., Massoni, J., Pestalozzi, C. M., Bortfeld-Miller, M., Maier, B. A., Vorholt, J. A. (2023). Identifying microbiota community patterns important for plant protection using synthetic communities and machine learning. Nat. Commun. 14, 7983. doi: 10.1038/s41467-023-43793-z
Fu, M., Chen, Y., Liu, Y.-X., Chang, X., Zhang, L., Yang, X., et al. (2024). Genotype-associated core bacteria enhance host resistance against kiwifruit bacterial canker. Hortic. Res. 11, uhae236. doi: 10.1093/hr/uhae236
Galeano Garcia, P., Neves dos Santos, F., Zanotta, S., Eberlin, M. N., Carazzone, C. (2018). Metabolomics of Solanum lycopersicum infected with Phytophthora infestans leads to early detection of late blight in asymptomatic plants. Molecules 23, 3330. doi: 10.3390/molecules23123330
Gray, E. J., Smith, D. L. (2005). Intracellular and extracellular PGPR: commonalities and distinctions in the plant–bacterium signaling processes. Soil Biol. Biochem. 37, 395–412. doi: 10.1016/j.soilbio.2004.08.030
Hermosa, R., Viterbo, A., Chet, I., Monte, E. (2012). Plant-beneficial effects of Trichoderma and of its genes. Microbiology 158, 17–25. doi: 10.1099/mic.0.052274-0
Huang, X., Zhou, X., Zhang, J., Cai, Z. (2019). Highly connected taxa located in the microbial network are prevalent in the rhizosphere soil of healthy plant. Biol. Fertil Soils 55, 299–312. doi: 10.1007/s00374-019-01350-1
Iparraguirre, J., Masciarelli, O., Villasuso, A. L., Piatti, D., Llanes, A. (2024). Macrocystis pyrifera alga extracts combined with Azospirillum argentinense improve growth and hormonal responses in zea mays plants under drought stress. J. Soil Sci. Plant Nutr. 24, 3209–3223. doi: 10.1007/s42729-024-01745-6
Ivanov, A. A., Ukladov, E. O., Golubeva, T. S. (2021). Phytophthora infestans: An overview of methods and attempts to combat late blight. J. Fungi 7, 1071. doi: 10.3390/jof7121071
Kharade, S. S., McBride, M. J. (2014). Flavobacterium johnsoniae chitinase chia is required for chitin utilization and is secreted by the type ix secretion system. J. Bacteriol. 196, 961–970. doi: 10.1128/jb.01170-13
Ko, W.-H., Tsou, Y.-J., Ju, Y.-M., Hsieh, H.-M., Ann, P.-J. (2010). Production of a fungistatic substance by pseudallescheria boydii isolated from soil amended with vegetable tissues and its significance. Mycopathologia 169, 125–131. doi: 10.1007/s11046-009-9237-1
Kotasthane, A., Agrawal, T., Kushwah, R., Rahatkar, O. V. (2015). In-vitro antagonism of Trichoderma spp. against Sclerotium rolfsii and Rhizoctonia solani and their response towards growth of cucumber, bottle gourd and bitter gourd. Eur. J. Plant Pathol. 141, 523–543. doi: 10.1007/s10658-014-0560-0
Kumar, A., Verma, J. P. (2018). Does plant—Microbe interaction confer stress tolerance in plants: A review? Microbiol. Res. 207, 41–52. doi: 10.1016/j.micres.2017.11.004
Kwak, M.-J., Kong, H. G., Choi, K., Kwon, S.-K., Song, J. Y., Lee, J., et al. (2018). Rhizosphere microbiome structure alters to enable wilt resistance in tomato. Nat. Biotechnol. 36, 1100–1109. doi: 10.1038/nbt.4232
Kwon, J.-H., Choi, O., Kang, B., Lee, Y., Park, J., Kang, D.-W., et al. (2017). Identification of Neocosmospora ipomoeae causing tomato stem rot in Korea. Australas Plant Dis. Notes 12, 34. doi: 10.1007/s13314-017-0254-5
Lambrecht, M., Okon, Y., Broek, A. V., Vanderleyden, J. (2000). Indole-3-acetic acid: a reciprocal signalling molecule in bacteria–plant interactions. Trends Microbiol. 8, 298–300. doi: 10.1016/S0966-842X(00)01732-7
Lazcano, C., Boyd, E., Holmes, G., Hewavitharana, S., Pasulka, A., Ivors, K. (2021). The rhizosphere microbiome plays a role in the resistance to soil-borne pathogens and nutrient uptake of strawberry cultivars under field conditions. Sci. Rep. 11, 3188. doi: 10.1038/s41598-021-82768-2
Li, P., Chai, A., Shi, Y., Xie, X., Li, B. (2017). First report of root rot caused by Plectosphaerella cucumerina on cabbage in China. Mycobiology 45, 110–113. doi: 10.5941/MYCO.2017.45.2.110
Li, Y., Li, Y., Yang, M., Chang, S. X., Qi, J., Tang, C., et al. (2020). Changes of microbial functional capacities in the rhizosphere contribute to aluminum tolerance by genotype-specific soybeans in acid soils. Biol. Fertil Soils 56, 771–783. doi: 10.1007/s00374-020-01451-2
Li, M., Pommier, T., Yin, Y., Wang, J., Gu, S., Jousset, A., et al. (2022). Indirect reduction of Ralstonia solanacearum via pathogen helper inhibition. ISME J. 16, 868–875. doi: 10.1038/s41396-021-01126-2
Li, H., Tang, M., Zheng, T., Yang, M., Wang, Y., Shuai, Y., et al. (2023). Differences in rhizosphere microbial community structure and composition in resistance and susceptible wheat to Fusarium head blight. Cell. Microbiol. 2023, 9963635. doi: 10.1155/2023/9963635
Lian, T., Huang, Y., Xie, X., Huo, X., Shahid, M. Q., Tian, L., et al. (2020). Rice SST variation shapes the rhizosphere bacterial community, conferring tolerance to salt stress through regulating soil metabolites. mSystems. 5. doi: 10.1128/msystems.00721-20
Ling, N., Song, Y., Raza, W., Huang, Q., Guo, S., Shen, Q. (2015). The response of root-associated bacterial community to the grafting of watermelon. Plant Soil 391, 253–264. doi: 10.1007/s11104-015-2399-3
Liu, Q., Cheng, L., Nian, H., Jin, J., Lian, T. (2023). Linking plant functional genes to rhizosphere microbes: a review. Plant Biotechnol. J. 21, 902–917. doi: 10.1111/pbi.13950
Liu, Y., Langemeier, M. R., Small, I. M., Joseph, L., Fry, W. E., Ristaino, J. B., et al. (2018). A risk analysis of precision agriculture technology to manage tomato late blight. Sustain 10, 3108. doi: 10.3390/su10093108
Lundberg, D. S., Lebeis, S. L., Paredes, S. H., Yourstone, S., Gehring, J., Malfatti, S., et al. (2012). Defining the core Arabidopsis thaliana root microbiome. Nature 488, 86–90. doi: 10.1038/nature11237
Müller, T., Behrendt, U., Ruppel, S., von der Waydbrink, G., Müller, M. E. H. (2016). Fluorescent Pseudomonads in the phyllosphere of wheat: potential antagonists against fungal phytopathogens. Curr. Microbiol. 72, 383–389. doi: 10.1007/s00284-015-0966-8
Oyserman, B. O., Flores, S. S., Griffioen, T., Pan, X., van der Wijk, E., Pronk, L., et al. (2022). Disentangling the genetic basis of rhizosphere microbiome assembly in tomato. Nat. Commun. 13, 3228. doi: 10.1038/s41467-022-30849-9
Peng, Z., Guo, X., Xiang, Z., Liu, D., Yu, K., Sun, K., et al. (2022). Maize intercropping enriches plant growth-promoting rhizobacteria and promotes both the growth and volatile oil concentration of Atractylodes lancea. Front. Plant Sci. 13. doi: 10.3389/fpls.2022.1029722
Philippot, L., Raaijmakers, J. M., Lemanceau, P., van der Putten, W. H. (2013). Going back to the roots: the microbial ecology of the rhizosphere. Nat. Rev. Microbiol. 11, 789–799. doi: 10.1038/nrmicro3109
Ping, X., Khan, R. A. A., Chen, S., Jiao, Y., Zhuang, X., Jiang, L., et al. (2024). Deciphering the role of rhizosphere microbiota in modulating disease resistance in cabbage varieties. Microbiome 12, 160. doi: 10.1186/s40168-024-01883-0
Qu, Q., Zhang, Z., Peijnenburg, W. J. G. M., Liu, W., Lu, T., Hu, B., et al. (2020). Rhizosphere microbiome assembly and its impact on plant growth. J. Agric. Food Chem. 68, 5024–5038. doi: 10.1021/acs.jafc.0c00073
Rai, S., Kashyap, P. L., Kumar, S., Srivastava, A. K., Ramteke, P. W. (2016). Identification, characterization and phylogenetic analysis of antifungal Trichoderma from tomato rhizosphere. SpringerPlus 5, 1939. doi: 10.1186/s40064-016-3657-4
Rolfe, S. A., Griffiths, J., Ton, J. (2019). Crying out for help with root exudates: adaptive mechanisms by which stressed plants assemble health-promoting soil microbiomes. Curr. Opin. Microbiol. 49, 73–82. doi: 10.1016/j.mib.2019.10.003
Shao, J., Miao, Y., Liu, K., Ren, Y., Xu, Z., Zhang, N., et al. (2021). Rhizosphere microbiome assembly involves seed-borne bacteria in compensatory phosphate solubilization. Soil Biol. Biochem. 159, 108273. doi: 10.1016/j.soilbio.2021.108273
Soltani, A.-A., Khavazi, K., Asadi-Rahmani, H., Omidvari, M., Dahaji, P. A., Mirhoseyni, H. (2010). Plant growth promoting characteristics in some Flavobacterium spp. isolated from soils of Iran. J. Agric. Sci. 2, 106. doi: 10.5539/jas.v2n4p106
Song, Y., Wilson, A. J., Zhang, X.-C., Thoms, D., Sohrabi, R., Song, S., et al. (2021). FERONIA restricts Pseudomonas in the rhizosphere microbiome via regulation of reactive oxygen species. Nat. Plants 7, 644–654. doi: 10.1038/s41477-021-00914-0
Stringlis, I. A., Yu, K., Feussner, K., de Jonge, R., Van Bentum, S., Van Verk, M. C., et al. (2018). MYB72-dependent coumarin exudation shapes root microbiome assembly to promote plant health. Proc. Natl. Acad. Sci. 115, E5213–E5222. doi: 10.1073/pnas.1722335115
Sultana, F., Hossain, M. M., Kubota, M., Hyakumachi, M. (2008). Elicitation of systemic resistance against the bacterial speck pathogen in Arabidopsis thaliana by culture filtrates of plant growth-promoting fungi. Can. J. Plant Pathol. 30, 196–205. doi: 10.1080/07060661.2008.10540535
Sun, W. M., Feng, L. N., Guo, W., Liu, D. Q., Yang, Z. H., Liu, L. F., et al. (2012). First report of Neocosmospora striata causing peanut pod rot in China. Plant Dis. 96, 146–146. doi: 10.1094/PDIS-06-11-0461
Tang, F. H. M., Lenzen, M., McBratney, A., Maggi, F. (2021). Risk of pesticide pollution at the global scale. Nat. Geosci. 14, 206–210. doi: 10.1038/s41561-021-00712-5
Trivedi, P., Leach, J. E., Tringe, S. G., Sa, T., Singh, B. K. (2021). Author Correction: Plant–microbiome interactions: from community assembly to plant health. Nat. Rev. Microbiol. 19, 72–72. doi: 10.1038/s41579-020-00490-8
Usami, T., Matsubara, C., Kashiwazaki, Y., Shito, T., Kanegae, Y., Ebihara, Y. (2015). Leaf and spathe spot of calla lily caused by an undescribed species of Plectosphaerella. J. Gen. Plant Pathol. 81, 291–296. doi: 10.1007/s10327-015-0597-8
Walters, W. A., Jin, Z., Youngblut, N., Wallace, J. G., Sutter, J., Zhang, W., et al. (2018). Large-scale replicated field study of maize rhizosphere identifies heritable microbes. Proc. Natl. Acad. Sci. 115, 7368–7373. doi: 10.1073/pnas.1800918115
Wang, F., Zhang, H., Liu, H., Wu, C., Wan, Y., Zhu, L., et al. (2024). Combating wheat yellow mosaic virus through microbial interactions and hormone pathway modulations. Microbiome 12, 200. doi: 10.1186/s40168-024-01911-z
Wu, C., Wang, F., Zhang, H., Chen, G., Deng, Y., Chen, J., et al. (2021). Enrichment of beneficial rhizosphere microbes in Chinese wheat yellow mosaic virus-resistant cultivars. Appl. Microbiol. Biotechnol. 105, 9371–9383. doi: 10.1007/s00253-021-11666-4
Xia, H., Shen, J., Riaz, M., Yang, H., Dong, Q., Zu, C., et al. (2024). Microbial regulatory mechanisms of disease-resistant tobacco varieties in the prevention and control of bacterial wilt disease. Appl. Soil Ecol. 202, 105598. doi: 10.1016/j.apsoil.2024.105598
Xun, W., Ren, Y., Yan, H., Ma, A., Liu, Z., Wang, L., et al. (2023). Sustained inhibition of maize seed-borne fusarium using a bacillus-dominated rhizospheric stable core microbiota with unique cooperative patterns. Adv. Sci. 10, 2205215. doi: 10.1002/advs.202205215
Xun, W., Shao, J., Shen, Q., Zhang, R. (2021). Rhizosphere microbiome: Functional compensatory assembly for plant fitness. Comput. Struct. Biotechnol. J. 19, 5487–5493. doi: 10.1016/j.csbj.2021.09.035
Yang, L., Gao, W., Huo, J., Zhang, C., Wang, Y. (2023). Identification of strawberry wilt caused by Plectosphaerella cucumerina in China. Plant Dis. 107, 3290. doi: 10.1094/PDIS-03-23-0544-PDN
Yang, L., Guo, Y., Yang, H., Li, S., Zhang, Y., Gao, C., et al. (2024). Distinct microbiota assembly and functional patterns in disease-resistant and susceptible varieties of tobacco. Front. Microbiol. 15. doi: 10.3389/fmicb.2024.1361883
Yang, S., Xiao, J., Liang, T., Tan, H. (2023). Response of bacterial compositions to the use of slow-release fertilizers with long-acting agents and synergists. Appl. Soil Ecol. 182, 104699. doi: 10.1016/j.apsoil.2022.104699
Yin, C., Casa Vargas, J. M., Schlatter, D. C., Hagerty, C. H., Hulbert, S. H., Paulitz, T. C. (2021). Rhizosphere community selection reveals bacteria associated with reduced root disease. Microbiome 9, 86. doi: 10.1186/s40168-020-00997-5
Zeilinger, S., Gruber, S., Bansal, R., Mukherjee, P. K. (2016). Secondary metabolism in Trichoderma – Chemistry meets genomics. Fungal Biol. Rev. 30, 74–90. doi: 10.1016/j.fbr.2016.05.001
Zhang, L., Khabbaz, S. E., Wang, A., Li, H., Abbasi, P. A. (2015). Detection and characterization of broad-spectrum antipathogen activity of novel rhizobacterial isolates and suppression of Fusarium crown and root rot disease of tomato. J. Appl. Microbiol. 118, 685–703. doi: 10.1111/jam.12728
Zhang, J., Liu, Y.-X., Zhang, N., Hu, B., Jin, T., Xu, H., et al. (2019). NRT1.1B is associated with root microbiota composition and nitrogen use in field-grown rice. Nat. Biotechnol. 37, 676–684. doi: 10.1038/s41587-019-0104-4
Zhao, W., Li, Y., Yang, C., Yang, Y., Hu, Y. (2023). Rhizosphere microbial community and metabolites of susceptible and resistant tobacco cultivars to bacterial wilt. J. Microbiol. 61, 389–402. doi: 10.1007/s12275-023-00012-0
Zheng, F., Xu, G., Zheng, F. Q., Ding, X. F., Xie, C. P. (2018). Neocosmospora rubicola causing stem rot of pitaya (Hylocereus costaricensis) in China. Plant Dis. 102, 2653–2653. doi: 10.1094/PDIS-09-17-1469-PDN
Zhou, X., Chen, S., Qiu, L., Liao, L., Lu, G., Yang, S. (2024). How rhizosphere microbial assemblage is influenced by dragon fruits with white and red flesh. Plants 13, 1346. doi: 10.3390/plants13101346
Zhou, X., Zhang, X., Ma, C., Wu, F., Jin, X., Dini-Andreote, F., et al. (2022). Biochar amendment reduces cadmium uptake by stimulating cadmium-resistant PGPR in tomato rhizosphere. Chemosphere 307, 136138. doi: 10.1016/j.chemosphere.2022.136138
Zhu, Z., Dong, Z., Mo, R., Liu, X., Zuo, Y., Hu, X., et al. (2024). First report of Neocosmospora mori causing root rot and stem blight of mulberry in Nanzhang, Hubei, China. Plant Dis. 108, 206. doi: 10.1094/PDIS-04-23-0661-PDN
Keywords: tomato, Solanum lycopersicum L., late blight, soil microbial community structure, rhizosphere, antagonistic microorganisms
Citation: Zhou X, Liao L, Chen K, Yin Y, Qiu L, Li X, Li Q and Yang S (2025) Diversity and composition of soil microbial communities in the rhizospheres of late blight-resistant tomatoes after Phytophthora infestans inoculation. Front. Plant Sci. 16:1556928. doi: 10.3389/fpls.2025.1556928
Received: 07 January 2025; Accepted: 17 February 2025;
Published: 07 March 2025.
Edited by:
Abhay K. Pandey, North Bengal Regional R & D Center, IndiaReviewed by:
Carolina N. Resendiz Nava, Autonomous University of Queretaro, MexicoCopyright © 2025 Zhou, Liao, Chen, Yin, Qiu, Li, Li and Yang. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Shangdong Yang, eXNkNzA2QGd4dS5lZHUuY24=
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.