
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Plant Sci. , 24 March 2025
Sec. Plant Breeding
Volume 16 - 2025 | https://doi.org/10.3389/fpls.2025.1552268
Grain size significantly affects rice yield and quality. Although several genes that regulate grain size have been identified, their mechanisms remain unclear. In this study, we characterized the swg5 mutant, which has a smaller plant height, shorter panicles, and smaller grains compared to the wild type (WT). MutMap resequencing and gene knockout analysis identified SWG5, a gene encoding the kinesin-13a protein, a new allele of SRS3 that positively regulates grain length and weight. RNA sequencing analyses revealed that the SWG5 allele is involved in diterpenoid biosynthesis, amino sugar metabolism, and pentose-glucuronate interconversions. Furthermore, young panicles of the swg5 mutant exhibited decreased sucrose invertase activity as well as reduced sugar and starch content. These findings indicate that SWG5/SRS3 plays a significant role in sugar metabolism, influencing grain size and weight in rice. This research provides valuable insights into breeding rice varieties with improved yield and grain quality.
Rice (Oryza sativa L.) is the most important food crop globally, feeding over half of the worldwide population (Zuo and Li, 2014). Enhancing rice yield is crucial for food security, economic development, and environmental sustainability (Hashim et al., 2024). Rice yield is determined by four main factors: effective panicle number, grain number per panicle, grain filling rate, and grain weight, with the latter being associated with grain size. Grain size significantly influences rice yield and appearance quality (Ren et al., 2023; Zhao et al., 2022). Thus, identifying genes that regulate grain size and understanding their mechanisms is essential for breeding high-yield, high-quality rice varieties.
In recent years, significant progress has been made in understanding the molecular mechanisms governing rice grain size. Researchers have identified key genes that influence the development of rice grain size, shape, and weight. These genes impact grain development through various pathways, including hormone metabolism, G protein signaling, mitogen-activated protein kinase (MAPK) signaling, and the ubiquitin–proteasome pathway (Li et al., 2018; Li et al., 2019). However, the molecular regulatory mechanisms of rice grain size are not yet fully understood.
Sugars are the main products of photosynthesis and are essential for rice growth and development. Mutations in the PHS8 gene lead to the accumulation of small sugars in the endosperm. This accumulation inhibits the expression of OsABI3 and OsABI5 in the ABA signaling pathway, affecting seed germination (Du et al., 2018). Knocking out the OsSTP15 gene reduces glucose efflux and enhances sugar transport from the source leaves to the stem base. As a result, levels of glucose, sucrose, and trehalose-6-phosphate increase at the stem base. This rise in sugar levels boosts cytokinin (CK) synthesis and signaling, promoting tillering and increasing rice yield (Li et al., 2024). Overexpressing the OsNAC23 gene raises 6-phospho-trehalose levels in source organs, improving photosynthesis and supporting the development of storage organs. This process leads to a high rice yield (Li et al., 2022). Recent studies have identified genes that regulate rice grain size through sugar signaling. For example, OsFRK3 decreases maltose levels while increasing sucrose and fructose levels, which improves grain size and weight (Zhang Z. et al., 2023). The GF14f gene negatively regulates grain size (Song et al., 2024; Zhang et al., 2019). Additionally, FGW1 interacts with GF14F to regulate grain width by affecting cell processes and sugar metabolism (Li et al., 2023). The SMS2 gene encodes vacuolar acid invertase. Mutations in this gene reduce the sugar content in rice grains, resulting in smaller grains (Huang et al., 2024).
Kinesin-13 proteins are important for plant development, stress responses, and the regulation of grain size. The SRS3 gene, which belongs to the kinesin-13 subfamily, encodes a protein made up of 819 amino acids. This protein includes a motor domain and a coiled-coil structure. SRS3 affects grain size by influencing the cell length in the spikelet hull (Kitagawa et al., 2010). An allelic variant of SRS3, SGL, regulates the height of rice plants through gibberellin (GA) signaling (Wu et al., 2014). Recent studies have shown that SRS3/BSH1 acts downstream of BRI1 (BRASSINOSTEROID INSENSITIVE 1) to negatively regulate brassinosteroid (BR) signaling and plant architecture in rice (Zhang et al., 2022). SRS3 also forms a complex with OsTUB1, which stabilizes the Na+ transporter OsHKT1;5. This stabilization helps maintain ion balance and enables rice plants to cope with salt stress (Chen et al., 2022). These studies demonstrated that kinesin-13 proteins significantly influence rice growth, development, and stress responses. However, the exact mechanisms by which they affect agronomic traits remain unclear.
In this study, we identified and characterized the rice mutant small and wide grain 5 (swg5), which exhibits a short grain length and increased grain width. Using MutMap technology, we cloned the SWG5 gene and found that it encodes the kinesin-13a protein, allelic to the SRS3 gene. Although SRS3 is known to regulate grain size, its molecular mechanism remains poorly understood. We further investigated the molecular mechanism of SWG5 in regulating grain size and weight in rice. Our results demonstrated that SWG5 positively regulates grain length and weight while negatively regulating grain width by influencing sucrose metabolism. This study enhances our understanding of the regulatory role of SWG5/SRS3 in determining grain size.
The wild-type material utilized in this study was the japonica rice variety ZH11 (WT). The swg5 mutant, characterized by small and wide grains, was derived from ZH11 treated with EMS. CRISPR-Cas9 technology was employed to generate the swg5-cr1 and swg5-cr2 mutants. All rice materials were cultivated in the natural field environments of Lingshui (N18°, E110°) and Sanya (N18°, E108°), Hainan Province, China, from 2022 to 2024.
During the maturity stage, 10 plants each from the WT and swg5 groups were selected to measure plant height and tiller number. Photographs of the plants were taken using a digital camera. After harvesting, the plants were stored indoors for three months. For analysis, 10 mature panicles from each group were assessed for panicle length, grain number, seed setting rate, and the number of primary and secondary branches. Additionally, mature seeds were measured for grain length, width, and weight using a ScanMaker i800 (MICROTEK, China). Photographs of the panicles and seeds were also taken.
A scanning electron microscope (S-3000N & Quorum PP3000T, Hitachi, Japan) was employed to capture images of the outer surface of the mature seed glume. ImageJ software (Schneider et al., 2012) was subsequently utilized to quantify and measure the cells on the glume surface. The protocols for observing and measuring cell number and size were consistent with those described in our previous study (Duan et al., 2015).
We crossed ZH11 with the swg5 mutant at the heading stage to generate an F2 population and cloned the SWG5 gene. From this F2 population, we selected 50 rice plants exhibiting the swg5 phenotype, pooled their DNA in equal amounts, and conducted whole-genome resequencing using NextSeq 550 (Illumina). We then used the MutMap method to isolate the SWG5 gene (Abe et al., 2012) and analyzed the SNP-to-INDEL ratio, consistent with our previous studies (Fang et al., 2016). The swg5 mutation was identified by PCR amplification of the SWG5 genomic region in both wild-type and swg5 mutant plants, followed by sequencing. cDNA was amplified by PCR and analyzed by electrophoresis to assess the effect of SWG5 on alternative splicing. The primer sequences are listed in Supplementary Table S1.
We employed CRISPR/Cas9 gene editing to modify the SWG5 gene in the japonica variety ZH11 wild type (WT) and elucidate the role of SWG5 in rice grain development. This procedure resulted in the creation of SWG5 knockout mutants swg5-1 (1-bp insertion) and swg5-2 (2-bp deletion). The design of the target gene, vector construction, and transformation methods were performed according to previously established protocols (Zhang et al., 2022).
Total RNA was extracted from 5-cm young panicles of ZH11 (WT) and swg5-cr1 using the RNA Prep Pure Plant Kit (Tiangen, Beijing). Each sample comprised 20 plants, with three biological replicates prepared for library construction. Sequencing was conducted at Guangzhou Kidio Biotechnology Co., Ltd., using the Illumina NovaSeq 4000 platform. Quality control was performed using Fastp (v0.12.4) to filter low-quality data and obtain clean readings. The filtered reads were quantified and aligned to the Nipponbare Reference IRGSP-1.0 from the Ensembl Plants database using HISAT2 (v2.2.1) (Chen et al., 2018; Kim et al., 2019). Gene expression levels were quantified using FPKM, which estimates the number of fragments per kilobase of transcript per million mapped reads. Differential expression analysis was conducted using DESeq2 software, with edgeR used for comparisons between samples (Liao et al., 2019; Love et al., 2014). Genes or transcripts with a false discovery rate (FDR) below 0.05, |log2(fold-change)| > 0.5, and an adjusted p-value < 0.05 were considered significant.
GO and KEGG enrichment analyses were performed on differentially expressed genes in the young panicles of ZH11 and swg5-cr1 using the clusterProfiler package in R (v4.3.2) to elucidate the mechanism of SWG5 in regulating grain size. These analyses followed the methods described in our previous study (Huang et al., 2024).
We randomly selected seven differentially expressed genes to validate the RNA-seq results and investigate the role of SWG5 in regulating rice grain size and weight through sugar signaling. We specifically chose three genes involved in sugar metabolism or synthesis (Os05g0247100/OsHI-XIP, Os07g0539300, and Os06g0356800) for confirmation by qRT-PCR. Specific primers were designed using the NCBI database, and the experiments were conducted in triplicate. The relative expression levels of the genes were calculated using the 2−ΔΔCT method. The primer sequences are listed in Supplementary Table S1.
At the heading stage, we collected samples from young panicles (5-10 cm) of ZH11, swg5-cr1, and swg5-cr2 to evaluate sugar and starch contents, as well as enzyme activity. Invertase activity [VIN (vacuolar invertase), CWIN (soluble cell wall invertase), CIN (cell wall invertase)] was measured using previously established methods (Morey et al., 2018). The sucrose and starch content in the young panicles (5–10 cm) was determined according to the protocols described in earlier studies (Li et al., 2013; Shu et al., 2014).
Data collation was performed using Microsoft Excel 2021, and statistical analyses were conducted using GraphPad Prism (v9.3.1.471). Two-tailed Student’s t-tests were utilized for comparisons between the two groups. One-way ANOVA was employed for comparisons among multiple groups. Graphs were generated using Adobe Photoshop CC 2020 and GraphPad Prism (v9.3.1.471).
We identified and characterized the rice mutant swg5, which exhibits reduced grain length and increased grain width, to better understand the genetic networks regulating grain size. At maturity, the swg5 mutant displayed a significantly shorter plant height compared to WT (Figures 1A, C). The panicle length (Figures 1B, D), grain number per panicle (Figure 1F), and seed setting rate (Figure 1G) were all significantly reduced in the swg5 mutant. At the same time, no significant changes were observed in the number of tillers (Figure 1E) or the primary and secondary branches (Figures 1H, I). Regarding leaf morphology, the flag leaf length of the swg5 mutant was significantly shorter than that of the WT, while the flag leaf width was significantly greater (Figures 1J, K).
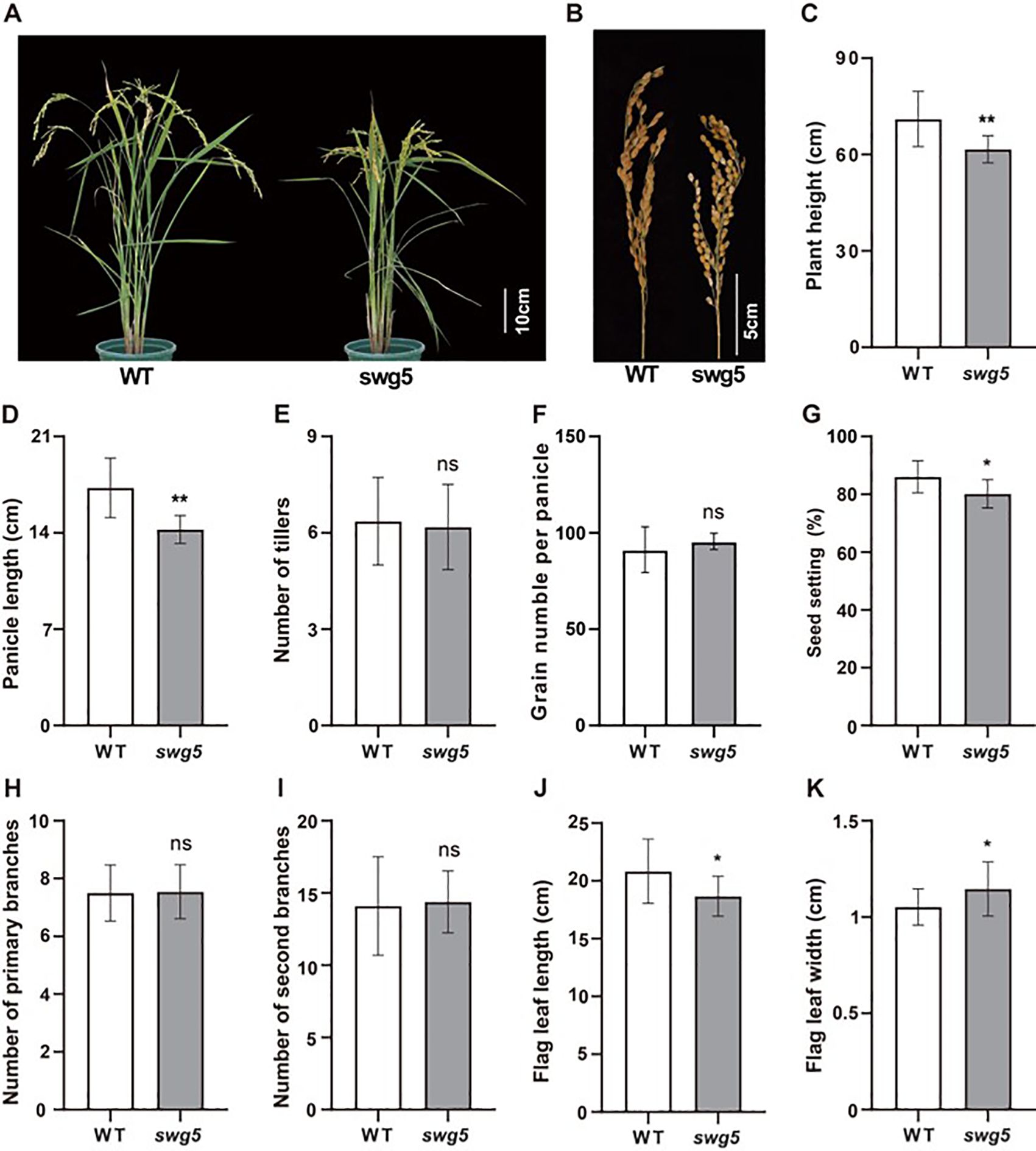
Figure 1. Phenotypic comparison of wild-type (WT) and swg5 rice plants. (A) Whole plants; bar = 10 cm. (B) Panicles; bar = 5 cm. (C) Plant height. (D) Panicle length. (E) Panicle number. (F) Grain number per panicle. (G) Setting rate. (H) Primary branch number. (I) Secondary branch number. (J) Flag leaf length. (K) Flag leaf width. Values are means ± SD. Student’s t-test was used to calculate the p-values: *P < 0.05, **P < 0.01. NS indicates no significant difference.
More importantly, we measured the mature seed traits of both the WT and swg5 mutants. The grain length (Figures 2A, D) and 1000 grain weight (Figure 2F) of the swg5 mutant were reduced by 22.54% and 9.56%, respectively, while the grain width (Figures 2B, E) increased by 9.35% compared to the WT. These results suggest that SWG5 has a pleiotropic effect on both grain size and plant architecture.
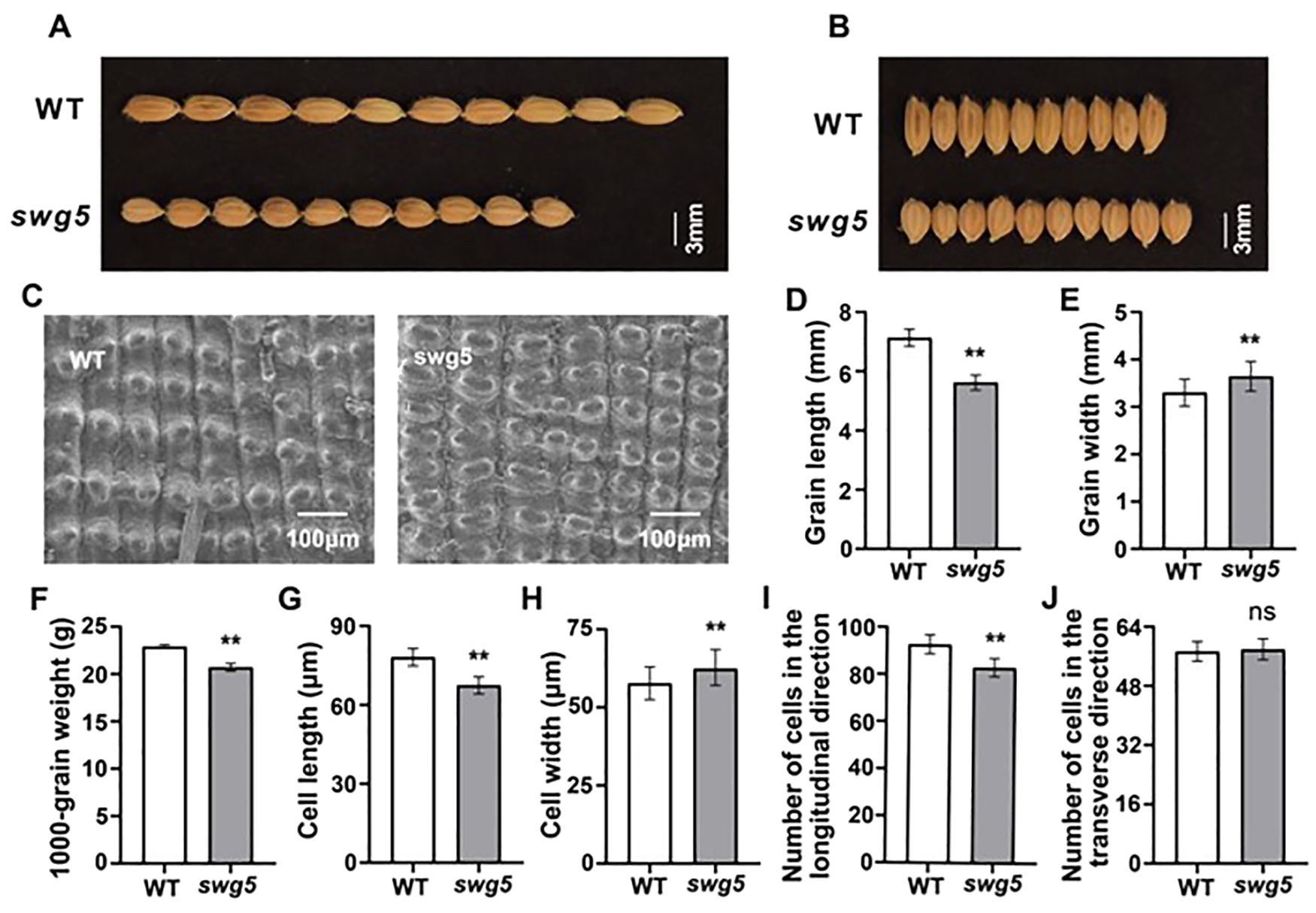
Figure 2. Morphological and histological analysis of spikelet hulls in WT and swg5 to evaluate grain size. (A-B) Grain shape morphology in WT and swg5; bar = 3 mm. (C) Scanning electron microscopy of the glume outer surfaces of mature seeds; scale bars = 100 µm. (D–F) Grain length (D) grain width (E) and 1000 grain weight (F) in WT and swg5. (G, H) Average length (G) and width (H) of the outer epidermal cells of lemmas. (I, J) Number of outer epidermal cells in the longitudinal (I) and transverse (J) planes. Values are means ± SD. Student’s t-test was used to calculate the p-values; **P < 0.01. NS indicates no significant difference.
We analyzed cell size and number in the spikelet hulls using scanning electron microscopy (SEM) to investigate the cellular basis for the short- and wide-grain traits in the swg5 mutant (Figure 2C). The SEM images of the outer epidermis of mature grains showed that cell length was shorter and cell width was wider in the swg5 mutant compared to WT (Figures 2G, H). Additionally, the number of cells with the same longitudinal length was significantly lower in the swg5 mutant (Figure 2I). However, there was no significant difference in the number of cells with the same transverse length (Figure 2J). Specifically, the number of outer epidermal cells in the longitudinal planes was reduced by 11.1% in the swg5 mutant. At the same time, the number of cells in the transverse planes showed no significant difference. These results suggest that the reduced grain length in the swg5 mutant is due to few longitudinal cells, while the increased grain width was caused by large cell size.
We crossed swg5 mutants with the wild-type variety ZH11 to produce F2 populations to isolate SWG5. The phenotype of F1 heterozygous plants resembled that of WT. Analysis of the F2 population revealed a near 3:1 segregation ratio (normal: short and wide grains) for grain size phenotypes, suggesting that swg5 is controlled by a single recessive gene. We isolated DNA from 50 F2 plants exhibiting the swg5 mutant phenotype for whole-genome sequencing, with the ZH11 genome resequenced as a control. Potential causal mutations were identified following the method described by Huang et al. (2017). Six SNPs were linked to the small and wide grain phenotype of the swg5 mutant. Of these, three occurred in coding regions but caused synonymous mutations. Notably, an SNP mutation was identified in the splice site region of LOC_Os05g06280, where a single base substitution from G to A was detected at site 787 in the intron, relative to the start codon (Figures 3A, B). This mutation in the splice site region primarily affects the splicing of precursor mRNA (pre-mRNA). Importantly, LOC_Os05g06280/SRS3, which encodes the kinesin-13a protein, has previously been reported to regulate rice grain length (Kitagawa et al., 2010; Wu et al., 2014). Therefore, we further sequenced and analyzed this mutation in LOC_Os05g06280. Sequence analysis revealed that the mutation, located near the start of exon 2, caused a 234-bp alternative splicing deletion in the coding sequence (Figures 3C, D). Additionally, gene expression analysis of WT and swg5 mutants indicated that LOC_Os05g06280 expression was significantly reduced in the mutant (Figure 3E). LOC_Os05g06280/SRS3 was considered the putative candidate gene for SWG5 based on these findings.
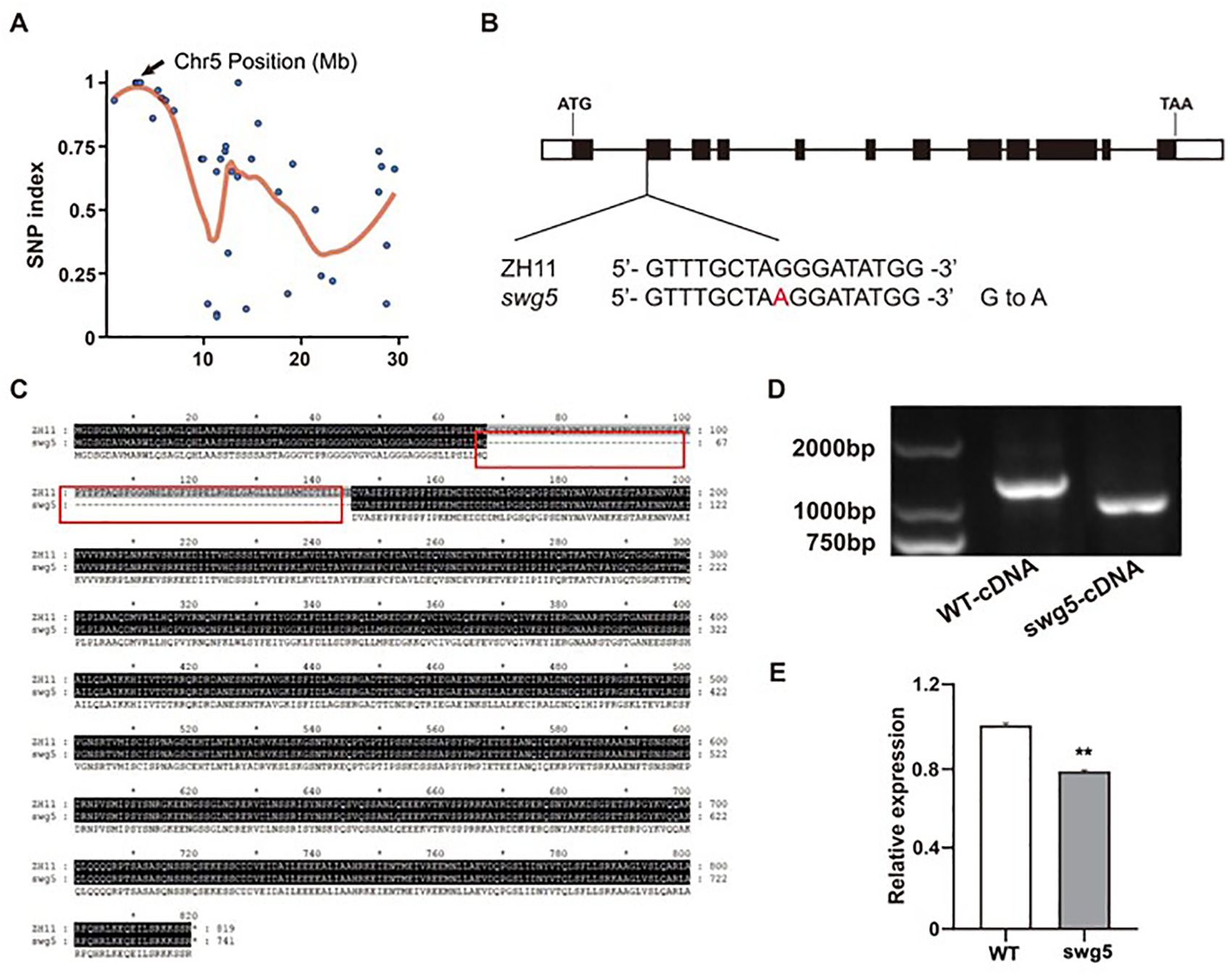
Figure 3. Identification of the swg5 mutation. (A) Cloning of the SWG5 gene using MutMap. (B) Structure of the SWG5 gene showing the mutation site. (C) Comparison of amino acids between WT and the swg5 mutant. (D) PCR analysis of the 234-bp splicing deletion in cDNA from three biological replicates. (E) Expression level of SWG5 in WT and swg5. Values are means ± SD. Student’s t-test was used to calculate the p-values; **P < 0.01.
We used the CRISPR/Cas9 system to knock SWG5 out in the ZH11 background to confirm that it was important for rice grain size. We created several transgenic plants with different mutations and selected two lines for analysis (Figures 4A–C): one with a 1-bp insertion (swg5-1) and the other with a 2-bp deletion (swg5-2). Grain size analysis showed that both mutants had a 20.63% and 20.6% reduction in grain length, while their grain widths increased by 13.31% and 12.16%, compared to ZH11 (Figure 4D–F). As a result, the grain shape index (grain length/width) decreased by 29.56% and 29.6% in the swg5-1 and swg5-2 mutants, respectively (Figure 4G). Thousand grain weight was reduced by 12.6% and 16.9%, respectively (Figure 4H). These results suggest that SWG5 plays a key role in grain development, positively regulating grain length and weight while negatively regulating grain width.
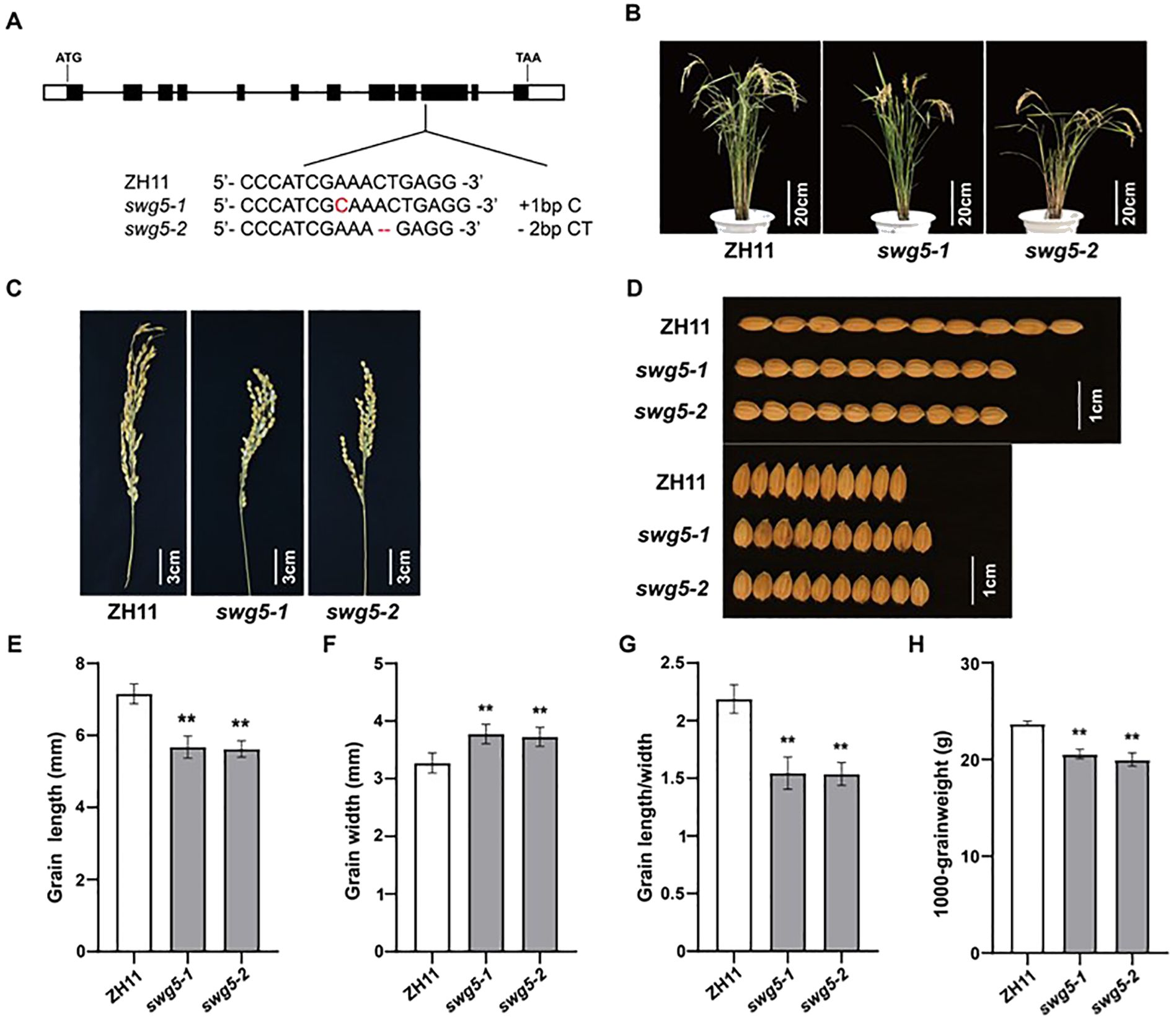
Figure 4. Gene knockouts of SWG5. (A) Mutant sequences of two homozygous mutants at the T2 generation. (B, C) Mature plants (B) and panicles (C) of ZH11, swg5-1, and swg5-2. Bars = 20 cm in (B) and 3 cm in (C). (D) Grain length and width phenotypes of ZH11, swg5-1, and swg5-2. Bars = 1 cm (E–H) Grain length (E) grain width (F) grain length/width ratio (G) and 1000-grain weight (H) in ZH11, swg5-1, and swg5-2. Values are means ± SD. Student’s t-test was used for comparisons (n = 10 spikelets; **P < 0.01).
We conducted RNA sequencing (RNA-seq) on young panicles of ZH11 and swg5-1 mutants to understand how SWG5 regulates grain size. Among 231 differentially expressed genes (DEGs), 38.52% (89) were upregulated, and 61.47% (142) were downregulated in the SWG5 knockout mutants compared to wild-type ZH11. We performed a functional analysis of these DEGs using GO and KEGG enrichment. The GO analysis identified 30 significantly enriched terms: 19 in biological processes, two in cellular components, and nine in molecular functions.
The top five biological process terms were cellular process, metabolic process, biological regulation, localization, and response to stimulus. In the cellular component category, the top two terms were cellular anatomical entity and protein-containing complex. The five high-ranking molecular function terms were catalytic activity, binding, transporter activity, transcription regulator activity, and molecular function regulator activity (Figure 5A). KEGG enrichment revealed that the five high-ranking metabolic pathways were diterpenoid biosynthesis, amino sugar and nucleotide sugar metabolisms, pentose and glucuronate interconversions, metabolic pathways, and the MAPK signaling pathway (Figure 5B). The remaining pathways had fewer DEGs than the previous ones, suggesting that SWG5 regulates grain size through these five key pathways. Notably, amino sugar and nucleotide sugar metabolism, along with pentose and glucuronate interconversions, are involved in sugar metabolism. These results suggest that SWG5 may regulate grain size by modulating sugar metabolism.
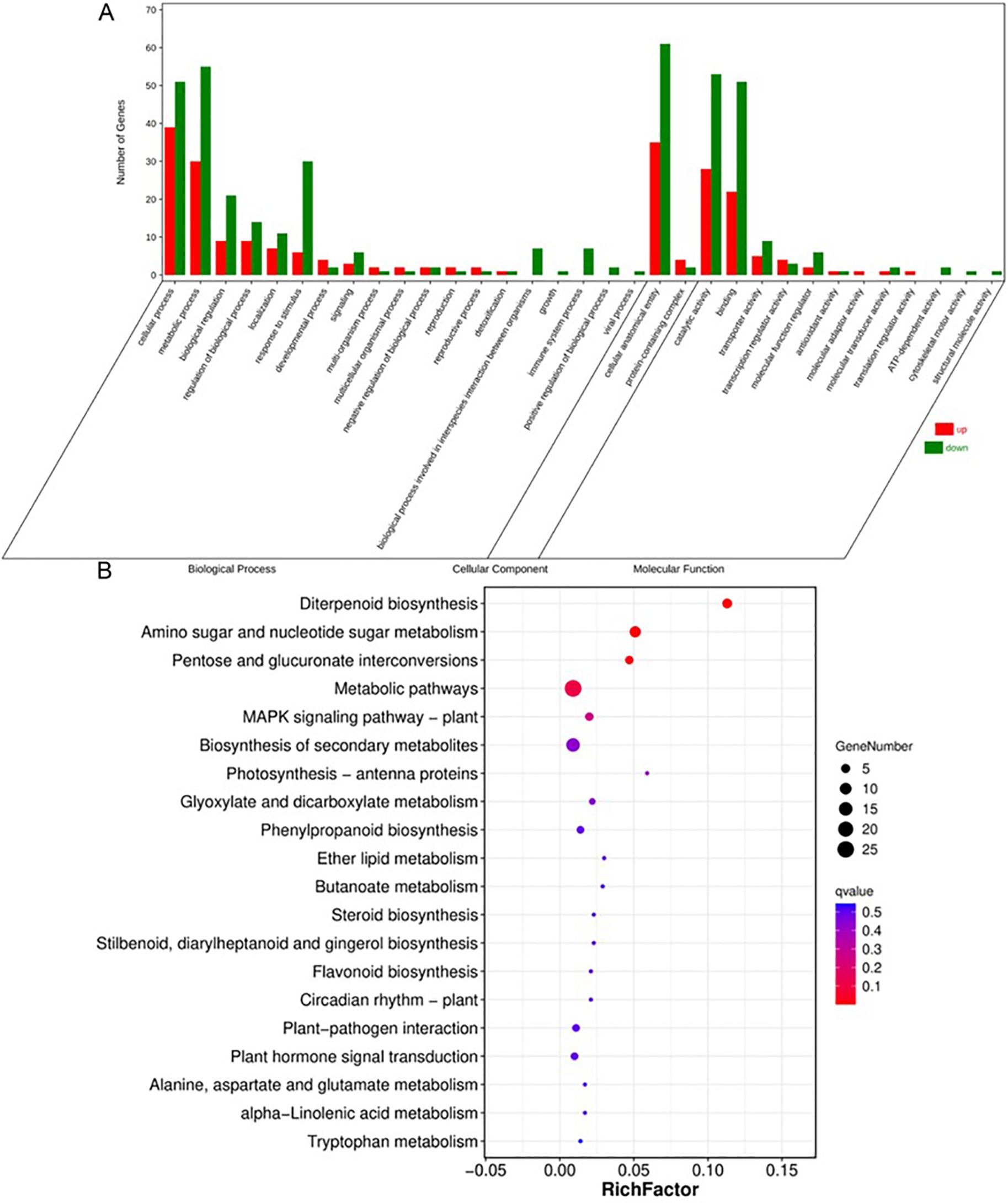
Figure 5. RNA-Seq analysis of grain size regulation by the SWG5 gene. (A) Gene ontology enrichment analysis of differentially expressed genes (DEGs). (B) KEGG pathway analysis of differentially expressed genes (DEGs).
We randomly selected 10 genes (both upregulated and downregulated) from these pathways for qRT-PCR to validate the RNA-seq results. Three genes involved in sugar metabolism, transport, and synthesis were chosen: Os05g0247100/OsHI-XIP (xylanase inhibitory protein), Os07g0539300 (glucan Endo-1,3-beta-glucosidase precursor), and Os06g0356800 (glycosyl hydrolase). Specific primers were designed using Primer Premier (version 5.0). The qRT-PCR results confirmed that the gene expression patterns were consistent with the RNA-seq data, validating the transcriptome analysis (Figure 6).
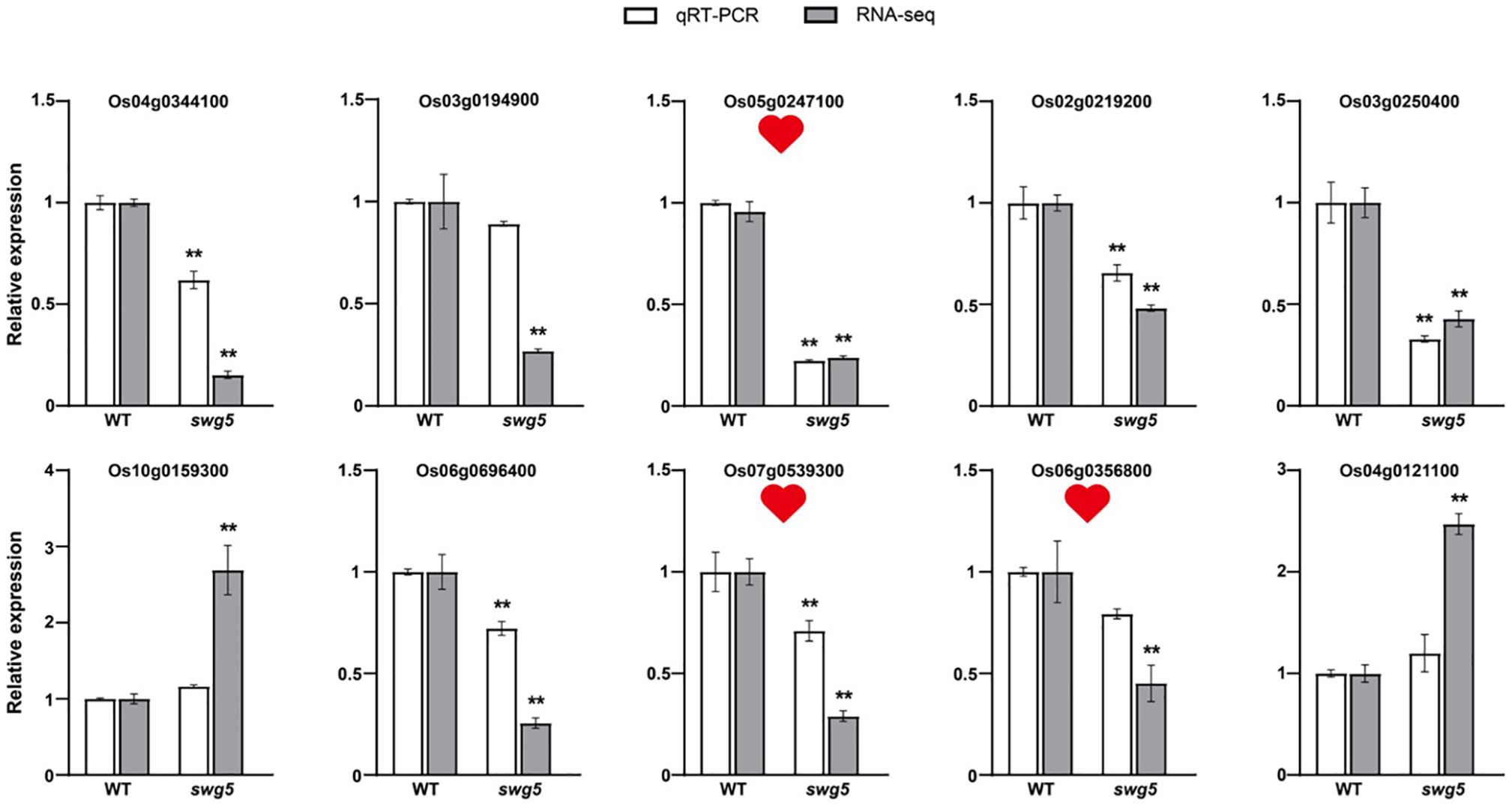
Figure 6. The differentially expressed genes between WT and SWG5 were analyzed using qRT-PCR. The white and gray bars represent transcriptome data and quantitative data, respectively. The three red hearts indicate the genes involved in sugar metabolism. Values are means ± SD. Student’s t-test was used to calculate the p-values; **P < 0.01.
To determine if SWG5 affects grain size and weight through sugar metabolism, we measured the activities of CIN, VIN, and CWIN, as well as sugar and starch content, in the young panicles of ZH11 and SWG5 knockout mutants (swg5-1 and swg5-2). The activities of CIN and CWIN were significantly lower in the SWG5 knockout mutants than in ZH11 (Figures 7A, C). In contrast, the VIN activity showed no significant difference (Figure 7B). Sucrose, fructose, and glucose levels were also significantly lower in the knockout mutants, although fructose content in swg5-2 was similar to ZH11 (Figures 7D–F). Additionally, the starch content in the mature seeds of SWG5 knockout mutants (swg5-1 and swg5-2) was significantly reduced compared to ZH11 (Figure 7G). These results suggest that SWG5 may regulate grain size by reducing invertase activity, which limits sucrose and starch accumulation.
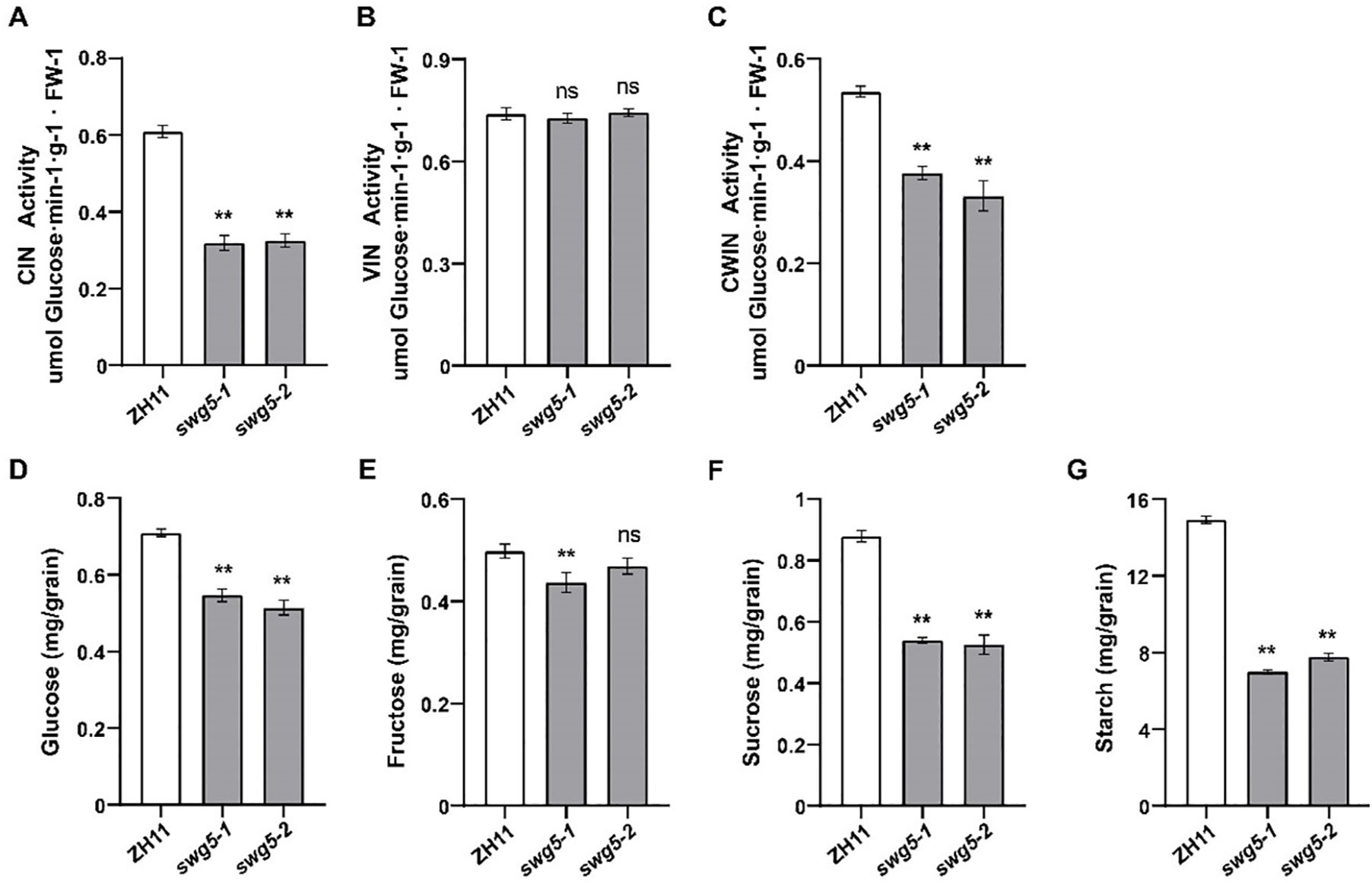
Figure 7. Comparison of enzyme activity and sugar and starch contents between ZH11, swg5-1, and swg5-2. (A–C) Activities of three invertase isoforms in young panicles of ZH11, swg5-1, and swg5-2 plants. (D–F) Levels of glucose, fructose, and sucrose in young panicles of ZH11, swg5-1, and swg5-2 plants. (G) Starch content in mature seeds of the ZH11, swg5-1, and swg5-2 plants. Values are means ± SD. Student’s t-test was used to calculate the p-values; **P < 0.01. NS indicates no significant difference.
Rice grain size affects both yield and quality (Harberd, 2015; Li et al., 2018), and understanding such regulation is crucial for improving rice varieties and seed development (Yan et al., 2024; Zhang J. et al., 2023). Advances in molecular biology and genomics have accelerated research in this area (Long et al., 2024; Ren et al., 2023). However, the exact molecular mechanisms remain unclear.
In this study, we identified the swg5 mutant from an EMS-mutagenized ZH11 population (Figure 1). Using MutMap, genetic complementation, and gene knockout techniques, we found that the candidate gene is LOC_Os05g06280, which encodes the kinesin-13a protein (Figures 3, 4). SWG5 is a new allelic mutant of genes such as SRS3, SGL, BHS1, and sar1 (Deng et al., 2015; Kitagawa et al., 2010; Wu et al., 2014; Zhang et al., 2022). These mutants share traits such as small grains, short panicles, reduced plant height, and short internodes. The SRS3 mutation occurs in the ninth exon of LOC_Os05g06280, with a single nucleotide change from C to T (Kitagawa et al., 2010). The BSH1 mutation is in the eighth exon, with four nucleotide changes from CTGC to TTTT (Zhang et al., 2022). The SGL mutation is in the noncoding region upstream of the ATG start codon and involves a single base deletion (Wu et al., 2014). The TCM2092 mutation occurs in the 11th intron, where a G to A change leads to abnormal splicing and truncates the SWG5 protein from 819 to 762 amino acids (Kitagawa et al., 2010). In our study, we identified an SWG5 mutation in the first intron, near the second exon, with a G to A change (Figure 3B). This mutation also causes abnormal splicing, leading to a truncated SWG5 protein of 819 amino acids. Additionally, the expression level of SWG5 was reduced (Figure 3E). These mutations affect various phenotypic traits. We suggest that these changes may impact gene expression or function, affecting development, metabolism, and cellular processes. Further investigation is needed to fully understand the role of SWG5 in rice.
Many genes related to rice grain morphology have been identified (Li et al., 2019; Ren et al., 2023). Most of these genes affect either grain length (Wang et al., 2024, 2015) or width (Liu et al., 2017; Song et al., 2007) by regulating cell growth or expansion. However, most of these genes affect only one aspect of grain morphology. Some genes, such as WTG1 and GW6 (Huang et al., 2017; Shi et al., 2020), affect both traits. In this study, cytological analysis showed that the SWG5 mutant had fewer longitudinal cells but no change in transverse cells (Figures 2I, J). This suggests that SWG5 specifically regulates the direction of cell division, reducing the number of longitudinal cells and shortening the lemma length.
Additionally, cell size also plays a role in grain size. Despite having fewer cells, the SWG5 mutant showed an increase in individual cell width, resulting in wider grains (Figures 2B, H). In conclusion, we propose that SWG5 promotes cell proliferation to control grain length while limiting transverse cell expansion to control grain width. The underlying mechanism should be explored in future studies.
We used CRISPR/Cas9 gene editing to knock out SWG5 in ZH11. The results showed that knocking out SWG5 reduced grain length and 1000-grain weight but increased grain width (Figure 4). Previous studies have also indicated that SWG5 plays a role in regulating salt stress tolerance (Chen et al., 2022). This result suggests that SWG5 not only controls grain size but may also help plants grow and adapt to stress.
We performed an RNA-seq analysis on young panicles (5–10 cm) of ZH11 and SWG5 knockout (KO) mutants to better understand how SWG5 regulates grain size. KEGG analysis revealed that differentially expressed genes (DEGs) were enriched in pathways related to diterpenoid biosynthesis, amino sugar and nucleotide sugar metabolism, pentose and glucuronate interconversions, metabolic pathways, and the MAPK signaling pathway (Figure 5B). These results suggest that SWG5 may regulate grain size through glucose metabolism.
Grain biomass accumulation depends on sucrose, which is converted into hexoses and other simple sugars as it moves from the leaf to the developing grain. These sugars are crucial for energy supply and assimilation (Koch, 2004; Wan et al., 2018). Sugar transporters are essential for carbon distribution, helping move sucrose from source leaves to growing tissues, while invertases break down sucrose (Li et al., 2017).
We further examined whether SWG5 affected sugar metabolism by measuring enzyme activity and sugar composition in ZH11 and swg5 mutants. Compared to ZH11, swg5 mutants showed significantly reduced activity of CIN and CWIN in young panicles (Figures 7A, C). However, VIN activity did not differ (Figure 7B). Sugar composition in the KO mutants was also altered, with low levels of sucrose and hexoses (glucose and fructose) (Figures 7D–F).
Starch, the primary carbohydrate in rice grains, is synthesized from hexose units derived from sucrose and serves as the main energy storage in grains. We measured the starch content in mature grains of the swg5 mutants, which was significantly lower (Figure 7G). This result suggests that the loss of SWG5 reduces invertase activity, affecting sucrose breakdown and hindering grain biomass accumulation, ultimately leading to a smaller grain size. Other studies have also shown that multiple genes involved in sugar metabolism regulate grain size (Deng et al., 2020; Xu et al., 2019), further supporting the importance of sugars in grain development.
Although this study shows that SWG5 plays a key role in regulating grain size, more research is needed. First, how SWG5 controls grain growth through sugar metabolism, especially its interactions with sugar enzymes or transporters, requires further study. Additionally, the interaction between SWG5 and other grain-size genes should be explored. Future work could involve creating double mutants of SWG5 and other grain-size genes to examine their combined effects and to better understand the mechanisms of grain-size regulation.
The original contributions presented in the study are publicly available. This data can be found here: NCBI, PRJCA034576.
WL: Investigation, Writing – original draft. GC: Investigation, Writing – original draft. YX: Investigation, Writing – original draft. XF: Investigation, Writing – original draft. LZ: Investigation, Writing – original draft. YT: Data curation, Formal Analysis, Writing – original draft. RX: Conceptualization, Funding acquisition, Project administration, Supervision, Writing – original draft, Writing – review & editing. YL: Conceptualization, Funding acquisition, Project administration, Supervision, Writing – original draft, Writing – review & editing. LW: Project administration, Writing – original draft, Writing – review & editing.
The author(s) declare that financial support was received for the research and/or publication of this article. This study was supported by the Project of Sanya Yazhou Bay Science and Technology City (SCKJ-JYRC-2024-28), the Natural Science Foundation of Hainan Province (322RC592), the National Natural Science Foundation of China (32360083 and 31960159), the Hainan Province Science and Technology Special Fund (Hainan Provincial Key Research and Development Project of China; ZDYF2021XDNY165), and the Department of Education of Guizhou Province Qianjiaoji (No. [2022] 311).
We would like to thank the editor and reviewers for their valuable suggestions, which have greatly improved the quality of the manuscript.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fpls.2025.1552268/full#supplementary-material
Abe, A., Kosugi, S., Yoshida, K., Natsume, S., Takagi, H., Kanzaki, H., et al. (2012). Genome sequencing reveals agronomically important loci in rice using MutMap. Nat. Biotechnol. 30, 174–178. doi: 10.1038/nbt.2095
Chen, G., Xuan, W., Zhao, P., Yao, X., Peng, C., Tian, Y., et al. (2022). OsTUB1 confers salt insensitivity by interacting with Kinesin13A to stabilize microtubules and ion transporters in rice. New Phytol. 235, 1836–1852. doi: 10.1111/nph.18282
Chen, S., Zhou, Y., Chen, Y., Gu, J. (2018). fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890. doi: 10.1093/bioinformatics/bty560
Deng, X., Han, X., Yu, S., Liu, Z., Guo, D., He, Y., et al. (2020). OsINV3 and its homolog, OsINV2, control grain size in rice. Int. J. Mol. Sci. 21, 6. doi: 10.3390/ijms21062199
Deng, Z. Y., Liu, L. T., Li, T., Yan, S., Kuang, B. J., Huang, S. J., et al. (2015). OsKinesin-13A is an active microtubule depolymerase involved in glume length regulation via affecting cell elongation. Sci. Rep. 5, 1. doi: 10.1038/srep09457
Du, L., Xu, F., Fang, J., Gao, S., Tang, J., Fang, S., et al. (2018). Endosperm sugar accumulation caused by mutation of 8/1 leads to pre-harvest sprouting in rice. Plant J. 95, 545–556. doi: 10.1111/tpj.13970
Duan, P., Ni, S., Wang, J., Zhang, B., Xu, R., Wang, Y., et al. (2015). Regulation ofOsGRF4 by OsmiR396 controls grain size and yield in rice. Nat. Plants 2, 1–5. doi: 10.1038/nplants.2015.203
Fang, N., Xu, R., Huang, L., Zhang, B., Duan, P., Li, N., et al. (2016). SMALL GRAIN 11 controls grain size, grain number and grain yield in rice. Rice 9, 1–11. doi: 10.1186/s12284-016-0136-z
Harberd, N. P. (2015). Shaping taste: the molecular discovery of rice genes improving grain size, shape and quality. J. Genet. Genom. 42, 597–599. doi: 10.1016/j.jgg.2015.09.008
Hashim, N., Ali, M. M., Mahadi, M. R., Abdullah, A. F., Wayayok, A., Mohd Kassim, M. S., et al. (2024). Smart farming for sustainable rice production: an insight into application, challenge, and future prospect. Rice Sci. 31, 47–61. doi: 10.1016/j.rsci.2023.08.004
Huang, J., Zhou, Z., Wang, Y., Yang, J., Wang, X., Tang, Y., et al. (2024). SMS2, a novel allele of OsINV3, regulates grain size in rice. Plants 13, 1219. doi: 10.3390/plants13091219
Huang, K., Wang, D., Duan, P., Zhang, B., Xu, R., Li, N., et al. (2017). WIDE AND THICK GRAIN 1, which encodes an otubain-like protease with deubiquitination activity, influences grain size and shape in rice. Plant J. 91, 849–860. doi: 10.1111/tpj.13613
Kim, D., Paggi, J. M., Park, C., Bennett, C., Salzberg, S. L. (2019). Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915. doi: 10.1038/s41587-019-0201-4
Kitagawa, K., Kurinami, S., Oki, K., Abe, Y., Ando, T., Kono, I., et al. (2010). A novel kinesin 13 protein regulating rice seed length. Plant Cell Physiol. 51, 1315–1329. doi: 10.1093/pcp/pcq092
Koch, K. (2004). Sucrose metabolism: regulatory mechanisms and pivotal roles in sugar sensing and plant development. Curr. Opin. Plant Biol. 7, 235–246. doi: 10.1016/j.pbi.2004.03.014
Li, Y., He, P., Wang, X., Chen, H., Ni, J., Tian, W., et al. (2023). FGW1, a protein containing DUF630 and DUF632 domains, regulates grain size and filling in Oryza sativa L. Crop J. 11, 1390–1400. doi: 10.1016/j.cj.2023.03.015
Li, M., Li, H., Zhu, Q., Liu, D., Li, Z., Chen, H., et al. (2024). Knockout of the sugar transporter OsSTP15 enhances grain yield by improving tiller number due to increased sugar content in the shoot base of rice (Oryza sativa L.). New Phytol. 241, 1250–1265. doi: 10.1111/nph.19411
Li, K., Liu, S., Tan, Y., Chao, N., Tian, X., Qi, L., et al. (2013). Optimized GC-MS method to simultaneously quantify acetylated aldose, ketose, and alditol for plant tissues based on derivatization in a methyl sulfoxide/1-methylimidazole system. J. Agric. Food Chem. 61, 4011–4018. doi: 10.1021/jf3053862
Li, Z., Wei, X., Tong, X., Zhao, J., Liu, X., Wang, H., et al. (2022). The OsNAC23-Tre6P-SnRK1a feed-forward loop regulates sugar homeostasis and grain yield in rice. Mol. Plant 15, 706–722. doi: 10.1016/j.molp.2022.01.016
Li, J., Wu, L., Foster, R., Ruan, Y.-L. (2017). Molecular regulation of sucrose catabolism and sugar transport for development, defence and phloem function. J. Integr. Plant Biol. 59, 322–335. doi: 10.1111/jipb.12539
Li, N., Xu, R., Duan, P., Li, Y. (2018). Control of grain size in rice. Control grain size rice Plant Reprod. 31, 237–251. doi: 10.1007/s00497-018-0333-6
Li, N., Xu, R., Li, Y. (2019). Molecular Networks of Seed Size Control in Plants. Mol. Networks seed size control plants Ann. Rev. Plant Biol. 70, 435–463. doi: 10.1146/annurev-arplant-050718-095851
Liao, Y., Smyth, G. K., Shi, W. (2019). The R package Rsubread is easier, faster, cheaper and better for alignment and quantification of RNA sequencing reads. Nucleic Acids Res. 47, e47. doi: 10.1093/nar/gkz114
Liu, J., Chen, J., Zheng, X., Wu, F., Lin, Q., Heng, Y., et al. (2017). GW5 acts in the brassinosteroid signalling pathway to regulate grain width and weight in rice. Nat. Plants 3, 1–7. doi: 10.1038/nplants.2017.43
Long, Y., Wang, C., Liu, C., Li, H., Pu, A., Dong, Z., et al. (2024). Molecular mechanisms controlling grain size and weight and their biotechnological breeding applications in maize and other cereal crops. J. Adv. Res. 62, 27–46. doi: 10.1016/j.jare.2023.09.016
Love, M. I., Huber, W., Anders, S. (2014). Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550. doi: 10.1186/s13059-014-0550-8
Morey, S. R., Hirose, T., Hashida, Y., Miyao, A., Hirochika, H., Ohsugi, R., et al. (2018). Genetic evidence for the role of a rice vacuolar invertase as a molecular sink strength determinant. Rice 11, 6. doi: 10.1186/s12284-018-0201-x
Ren, D., Ding, C., Qian, Q. (2023). Molecular bases of rice grain size and quality for optimized productivity. Sci. Bull. 68, 314–350. doi: 10.1016/j.scib.2023.01.026
Schneider, C. A., Rasband, W. S., Eliceiri, K. W. (2012). NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675. doi: 10.1038/nmeth.2089
Shi, C.-L., Dong, N.-Q., Guo, T., Ye, W.-W., Shan, J.-X., Lin, H.-X. (2020). A quantitative trait locus GW6 controls rice grain size and yield through the gibberellin pathway. Plant J. 103, 1174–1188. doi: 10.1111/tpj.14793
Shu, X., Sun, J., Wu, D. (2014). Effects of grain development on formation of resistant starch in rice. Food Chem. 164, 89–97. doi: 10.1016/j.foodchem.2014.05.014
Song, X.-J., Huang, W., Shi, M., Zhu, M.-Z., Lin, H.-X. (2007). A QTL for rice grain width and weight encodes a previously unknown RING-type E3 ubiquitin ligase. Nat. Genet. 39, 623–630. doi: 10.1038/ng2014
Song, Y., Yang, H., Zhu, W., Wang, H., Zhang, J., Li, Y. (2024). The Os14-3-3 family genes regulate grain size in rice. J. Genet. Genom. 51, 454–457. doi: 10.1016/j.jgg.2023.10.005
Wan, H., Wu, L., Yang, Y., Zhou, G., Ruan, Y.-L. (2018). Evolution of Sucrose Metabolism: The Dichotomy of Invertases and Beyond. Trends Plant Sci. 23 (2), 163–177. doi: 10.1016/j.tplants.2017.11.001
Wang, Y., Chen, W., Xing, M., Sun, J., Wang, S., Yang, Z., et al. (2024). Wild rice GL12 synergistically improves grain length and salt tolerance in cultivated rice. Nat. Commun. 15, 9453. doi: 10.1038/s41467-024-53611-9
Wang, Y., Xiong, G., Hu, J., Jiang, L., Yu, H., Xu, J., et al. (2015). Copy number variation at the GL7 locus contributes to grain size diversity in rice. Nat. Genet. 47, 944–948. doi: 10.1038/ng.3346
Wu, T., Shen, Y., Zheng, M., Yang, C., Chen, Y., Feng, Z., et al. (2014). Gene SGL,encoding a kinesin-like protein with transactivation activity, is involved in grain length and plant height in rice. Plant Cell Rep. 33, 235–244. doi: 10.1007/s00299-013-1524-0
Xu, X., Ren, Y., Wang, C., Zhang, H., Wang, F., Chen, J., et al. (2019). OsVIN2 encodes a vacuolar acid invertase that affects grain size by altering sugar metabolism in rice. Plant Cell Rep. 38, 1273–1290. doi: 10.1007/s00299-019-02443-9
Yan, L., Jiao, B., Duan, P., Guo, G., Zhang, B., Jiao, W., et al. (2024). Control of grain size and weight by the RNA-binding protein EOG1 in rice and wheat. Cell Rep. 43, 114856. doi: 10.1016/j.celrep.2024.114856
Zhang, Y., Dong, G., Chen, F., Xiong, E., Liu, H., Jiang, Y., et al. (2022). The kinesin-13 protein BR HYPERSENSITIVE 1 is a negative brassinosteroid signaling component regulating rice growth and development. Theor. Appl. Genet. 135, 1751–1766. doi: 10.1007/s00122-022-04067-2
Zhang, Z., Tan, J., Chen, Y., Sun, Z., Yan, X., Ouyang, J., et al. (2023). New fructokinase, OsFRK3, regulates starch accumulation and grain filling in rice. J. Agric. Food Chem. 71, 1056–1066. doi: 10.1021/acs.jafc.2c06783
Zhang, J., Zhang, X., Liu, X., Pai, Q., Wang, Y., Wu, X. (2023). Molecular network for regulation of seed size in plants. Int. J. Mol. Sci. 24, 10666. doi: 10.3390/ijms241310666
Zhang, Z., Zhao, H., Huang, F., Long, J., Song, G., Lin, W. (2019). The 14-3-3 protein GF14f negatively affects grain filling of inferior spikelets of rice (Oryza sativa L.). Plant J. 99, 344–358. doi: 10.1111/tpj.14329
Zhao, D., Zhang, C., Li, Q., Liu, Q. (2022). Genetic control of grain appearance quality in rice. Biotechnol. Adv. 60, 108014. doi: 10.1016/j.bioteChadv.2022.108014
Keywords: rice, SWG5, grain size, sugar signaling, grain weight
Citation: Lu W, Cai G, Xing Y, Fu X, Zhou L, Tang Y, Xu R, Li Y and Wu L (2025) SWG5 regulates grain size and weight via sugar metabolism-mediated signaling in rice. Front. Plant Sci. 16:1552268. doi: 10.3389/fpls.2025.1552268
Received: 27 December 2024; Accepted: 28 February 2025;
Published: 24 March 2025.
Edited by:
Cheng Shihua, China National Rice Research Institute (CAAS), ChinaReviewed by:
Yue Feng, Chinese Academy of Agricultural Sciences, ChinaCopyright © 2025 Lu, Cai, Xing, Fu, Zhou, Tang, Xu, Li and Wu. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Lian Wu, d3VsaWFuQGhhaW5hbnUuZWR1LmNu
†These authors have contributed equally to this work
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.