
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Plant Sci. , 18 March 2025
Sec. Plant Abiotic Stress
Volume 16 - 2025 | https://doi.org/10.3389/fpls.2025.1544879
This article is part of the Research Topic Advanced Breeding for Abiotic Stress Tolerance in Crops, Volume II View all 18 articles
Introduction: N6-methyladenosine (m6A) is the most prevalent posttranscriptional modification in eukaryotic mRNAs. AlkB homologs (ALKBHs) are involved in plant responses to stress by modulating m6A methylation. However, homologous genes in wheat remain largely uncharacterized.
Methods and results: In this study, 30 ALKBH genes were identified in wheat, and analyzed their physicochemical properties. The phylogenetic analysis allowed the classification of these genes into seven distinct subfamilies. Additionally, their conserved domains, motif compositions, gene structures, chromosomal localization, and synteny, and the predicted cis-acting elements within their promoters were examined. Expression analysis revealed that TaALKBH9B-5 exhibited the highest expression and its demethylase activity was investigated. Furthermore, TaALKBH9B-5 was significantly upregulated in response to abscisic acid treatment and cold stress, indicating a positive regulatory trend.
Discussion: In conclusion, this study provides a comprehensive genomic assessment of the TaALKBH gene family and offers a theoretical framework for understanding the role of TaALKBH9B in the response to abiotic stress in wheat.
N6-methyladenosine (m6A) represents one of the most abundant and widespread post-transcriptional RNA modifications in eukaryotes (Zhang et al., 2022). In plants, the methylation levels of m6A in messenger RNAs (mRNAs) are modulated by three principal components: writers, erasers, and readers. The writer complex, which includes methyltransferases such as MTA (human homolog of METTL3), MTB (METTL14), FIP37 (WTAP), VIRILIZER, HAKAI, and FIONA1, facilitates the addition of m6A to RNA (Yue et al., 2022b; Růžička et al., 2017; Wang et al., 2022; Yue et al., 2022a). m6A reader proteins, including ECT2, ECT3, ECT4, and CPSF30-L, possess a YTH domain that recognizes and binds to m6A-modified mRNAs, thereby regulating gene expression and impacting various biological functions (Wei et al., 2018; Arribas-Hernández et al., 2018; Hou et al., 2021). Demethylases from the AlkB homolog (ALKBH) family function as erasers, removing m6A from mRNAs (Martínez-Pérez et al., 2017). A substantial body of evidence suggests that RNA modifications critically regulate mRNAs, influencing plant growth, development, and stress response (Zhou et al., 2019; Amara et al., 2022). In a recent report, researchers have discovered that the natural variation in the wheat demethylase TaMTB (SNP176A/C) influences virus infection; however, it does not affect wheat growth and yield (Zhang et al., 2022). Furthermore, the wheat reading protein TaECT9 exhibits a significant upregulation in response to drought stress (Pan et al., 2024). To date, there have been no documented studies concerning the members of the ALKBH gene family in wheat.
The ALKBH family represents the sole group of enzymes in plants recognized for their m6A demethylase activity. The first member of this family, ALKBH, was identified in 1983 as a DNA repair protein in Escherichia coli (Kataoka et al., 1983). Following this discovery, the obesity-associated protein (FTO) (Jia et al., 2011) and ALKBH5 (Zheng et al., 2013) were identified as m6A demethylases, demonstrating that m6A methylation is reversible and influences gene expression. Since that time, numerous ALKBH family genes have been identified across various plant species, elucidating their roles in the regulation of growth and developmental processes. In Arabidopsis thaliana, AtALKBH10B has been demonstrated to demethylate m6A from mRNAs and to facilitate m6A demethylation at the FLOWERING LOCUS T (FT), SQUAMOSA PROMOTER BINDING PROTEIN-LIKE 3 (SPL3), and SPL9 during the flowering process, thereby specifically regulating the transition to flowering (Duan et al., 2017). In tomatoes, SlALKBH2 interacts with the SlDML2 transcript, influencing the ripening of tomato fruit (Zhou et al., 2019). Furthermore, an expanding body of research has underscored the significant role of ALKBH family members in plant responses to abiotic stress. Studies on Arabidopsis have demonstrated that alkbh6 mutant plants exhibit elevated levels of m6A and enhanced tolerance to salt, drought, and heat stress. This resilience is attributed to the downregulation of abscisic acid (ABA)-related genes mediated by AtALKBH6-induced m6A demethylation (Huong et al., 2020). Compared with wild-type plants, transgenic plants that overexpress ALKBH8B presented a reduced overall m6A level and exhibited greater salt tolerance (Huong et al., 2022). The alkbh9c mutant presented delayed seed germination under salt, drought, and low-temperature stress and under ABA treatment. While salt stress or ABA treatment inhibits the growth of mutant seedlings, drought conditions enhance it. Under dehydration stress, the m6A levels of several positive effectors associated with drought stress in the alkbh10b mutant were found to be elevated, although their transcriptional abundance was lower in the mutant compared to transgenic lines (Han et al., 2023). During seed germination, the ALKBH10B mutant presented reduced osmotic pressure and salt stress tolerance, resulting in a salt-sensitive phenotype. In cotton, the silencing of GhALKBH10 enhances antioxidant capacity and reduces cytoplasmic Na+ levels, thereby improving salt stress tolerance (Cui et al., 2022). Furthermore, the demethylase GhALKBH10B has been shown to increase drought tolerance at the seedling stage by decreasing the m6A level and the mRNA stability of genes associated with the ABA signaling pathway and calcium signaling (Zhang et al., 2024). In tomatoes, the Slalkbh10b mutant showed increased drought and salt stress tolerance, characterized by improved water retention, higher photosynthetic product, and proline levels, and reduced reactive oxygen species levels and cellular damage (Shen et al., 2023).
Wheat is one of the most essential food crops worldwide. However, throughout its growth cycle, wheat encounters various environmental challenges, including elevated temperatures, drought conditions, and high salinity, which impede normal growth and development, ultimately resulting in reduced yields. Numerous RNA modifications, particularly m6A methylation, are crucial for the response of plants to abiotic stresses. Despite this significance, research on m6A demethylases in wheat remains limited, with investigations into members of the ALKBH family being even scarcer. Therefore, there is an urgent need to identify and analyze the function of the wheat TaALKBH gene in stress resistance. The availability of several wheat reference genomes (http://www.wheatgenome.org/) provides a substantial platform for gene identification. In this study, 30 candidate genes belonging to the TaALKBH family were identified and categorized into seven distinct groups. Their evolutionary relationships, chromosomal locations, gene structures, and cis-regulatory elements were thoroughly analyzed. Furthermore, the tissue-specific expression patterns of the seven ALKBH subfamilies were preliminarily examined. TaALKBH9B-5 was identified as a potential m6A demethylase, and its expression under abiotic stress was investigated. This study elucidates the ALKBH family genes associated with m6A demethylases in wheat and establishes a foundation for future research into their biological functions.
The protein sequences of Arabidopsis ALKBH family members (AT1G11780, AT3G14140, AT3G14160, AT5G01780, AT2G22260, AT4G20350, AT4G02485, AT1G31600, AT4G36090, AT2G17970, AT1G48980, AT1G14710, AT2G48080, and AT4G02940) were downloaded from the National Center for Biotechnology Information (NCBI) database (https://www.ncbi.nlm.nih.gov, accessed on October 14, 2024) and used as Blastp templates to identify all ALKBH proteins in wheat, rice, and maize. Genomic data for wheat, rice, and maize were obtained from the Ensembl Plants Database (https://plants.ensembl.org/Triticum_aestivum/Info/Index, accessed on October 14, 2024. 30 wheat homologs with Eval < 10−6 and %ID > 60 were screened, along with 9 rice homologs and 11 maize homologs meeting the same criteria (Supplementary Table S1). All candidate proteins were further analyzed using the Pfam database (https://pfam.xfam.org/, accessed on October 14, 2024) (Finn et al., 2014). Detailed information on the TaALKBHs, including the amino acid counts, chromosome positioning, and coding sequence lengths, was obtained from the Ensembl Plants Database (Bolser et al., 2015). The molecular weight, theoretical isoelectric point, and subcellular localization predictive of each TaALKBH protein were obtained using CELLO (http://cello.life.nctu.edu.tw/, accessed on October 14, 2024) (Yu et al., 2004).
The phylogenetic tree of the wheat ALKBH gene family protein sequences was constructed using MEGA-X (Version: 10.2.6) software with the maximum likelihood method (ML) and 1000 bootstrap replicates (Kumar et al., 2018). The resultant analysis was visualized using TBtools.
The identified TaALKBH, OsALKBH, ZmALKBH and AtALKBH protein sequences were downloaded from the Ensembl Plants Database and imported into MEGA-X, and multiple sequence alignments were performed using the MUCLE analysis method, a rootless phylogenetic tree (1000 step repeats) was constructed by maximum likelihood method. A graphical representation of the generated developmental tree was beautified using the online tool ITOL [https://itol.embl.de//, (accessed October 14, 2024)].
The SWISS-MODEL (https://swissmodel.expasy.org/, accessed on 9 January 2025) was used to predict the structures of the TaALKBH proteins. A gene-building protein model was randomly selected from each subfamily for each species.
The conserved structural domains of the TaALKBH protein sequences were identified using the Batch CD-Search program (Marchler-Bauer and Bryant, 2004). Conserved motifs of the TaALKBH proteins were identified using the online service platform MEME [https://meme-suite.org/meme/, (accessed 24 October 2024)] with up to 15 motifs (Zheng et al., 2022). Gene structure analysis of GFF data downloaded from Ensembl Plants was performed to identify UTRs, exons, and introns. The 2000 base pair region upstream of the ATG of each TaALKBH gene was analyzed in cis-element analysis. The PlantCARE online platform [https://bioinformatics.psb.ugent.be/, (adopted on 25 October 2024)] was used for prediction (Yang et al., 2023). The results of the above analyses were visualized using TBtools.
The physical location of each TaALKBH gene in the annotation of the wheat genome in the Ensembl Plants database was used to determine chromosomal mapping. The GFF data containing all chromosome length information of wheat provided by the Ensembl Plants database were used to extract the position information and gene density map of the TaALKBH gene on chromosomes. The three datasets were then uploaded to TBtools for chromosomal position analysis of TaALKBHs. The One-Step MCScanX-Super Fast program integrated with TBtools was used to analyze the replication pattern and collinearity between the TaALKBHs (Chen et al., 2020). The Ka/Ks ratio was calculated using TBtools software, and the scattering time (T) (Hurst, 2002)was calculated based on Mya.
The materials used in this study, including Yangmai 158, were provided by Dr. Yang Jian. The wheat seeds were grown in an artificial greenhouse at 25°C for a 16 h light/8 h dark cycle, and when the wheat was grown to the three-leaf stage, the rhizomes and leaves of wheat were taken for RT–qPCR analysis.
The total RNA of the Wheat samples was extracted using the HiPure Plant RNA Mini Kit (Magen, Guangzhou, China). First-strand cDNA was synthesized using the First-Strand cDNA Synthesis Kit (Vazyme, Nanjing, China). The RT–qPCR was carried out using ChamQ Universal SYBR Green Master Mix (Vazyme, Nanjing, China) on an Applied Biosystems Quant Studio 5 Flex system (Applied Biosystems, Foster City, CA, USA), and the relative expression levels of the assayed genes were calculated using the 2−ΔΔCt method (Livak and Schmittgen, 2001) and the T. aestivum cell division cycle (TaCDC) gene (accession number: XM_020313450) was used as an internal reference gene (Zhang et al., 2019). Each treatment had three biological replicates and three technical replicates. The primer sequences used in this study are listed in Supplementary Table S2.
Wheat plants were divided into three tissue types: leaves, stems, and roots (Huang et al., 2024). One TaALKBH gene was randomly selected from each of the seven groups to analyze its expression in three different wheat tissues. Collect 3 replicates of each wheat tissue sample and store at -80°C until total RNA is extracted. Gene expression was determined by RT–qPCR. RT–qPCR reaction conditions: 95°C for 5 min, 1 cycle; This was followed by 40 cycles at 95°C 15 s, 58°C 20 s, and 72°C 30 s. It is then cycled at 72°C for 8 min and a single spot fluorescence detection is performed at 72°C. The results of tissue-specific expression analysis were displayed using GraphPad Prism 9. 0. 0.
The full length of TaALKBH9B-5 was cloned from Yangmai 158 cDNA. 35S: TaALKBH9B-5 was constructed using gateway technology. The primer pairs listed in Supplementary Table S2 were used for the first PCR. A second PCR was performed using primers attB1 and attB2 and the amplification product from the first PCR as a template. These amplified products are introduced into pDONR207 by a BP reaction (BP Clonase Reaction), and then these fragments are further transferred to the target vector, respectively. Full-length TaALKBH9B-5 was inserted into the pGEX4T-2 (a prokaryotic expression vector encoding GST) using the ClonExpress MultiS One-Step Cloning Kit (Vazyme, China). The primer sequences used in this study are listed in Supplementary Table S2.
Recombinant plasmid TaALKBH9B-5-GST and pGEX4T-2 were transformed into the Escherichia coli strain Rosetta-gami (DE3). The proteins were purified following the manufacturer’s instructions for Ni-NTA 6FF Sefinose (TM) Resin Kit (BBI C600332). 35S: TaALKBH9B-5 recombinant plasmid was transfected into wheat protoplasts using a PEG-mediated transformation method (Bio-Rad, Hercules, CA, USA). The transfected mesophyll protoplasts were cultured in W5 solution at 16°C for 18 h in the dark. Then TaALKBH9B-5-GFP protein was purified by GFP-Trap ® M2 magnetic Agarose beads (ChromoTek).
The optimal VIGS fragment sequence (300 bp) for TaALKBH9B-5 was obtained using the online VIGS tool [https://solgenomics.net/, (accessed November 1, 2024)]. Insert the fragment into the pBSMVγ vector. The plasmids pBSMVα, pBSMVβ, pBSMVγ, pBSMVγ: TaALKBH9B-5, and pBSMVγ: TaPDS were linearized using specific restriction enzymes, respectively. Linearized plasmids were transcribed to RNA using the Ribo MAXTM Large Scale RNA Production Systems-T7 and the Ribo m7G Cap Analog (Promega). The transcripts were mixed with FES buffer (0.06 M potassium phosphate, 0.1 M glycine, 1% bentonite, 1% sodium pyrophosphate decahydrate, 1% celite, pH 8.5) at 1: 1: 1: 7 (Zhang et al., 2019). Then 10 μL of the mixed transcript was inoculated with the second leaf of the wheat plant at the two-leaf stage by friction (Yang et al., 2020). Wheat plants inoculated with FES buffer were used as a negative control, and plants inoculated with the phytoene desaturase gene exhibiting a typical photobleaching phenotype were used as positive controls. Inoculated wheat seedlings were grown for 24 h under dark conditions of 25°C and 70% relative humidity. After 7 days of growth under 16 h light/8 h dark photoperiod, total RNA was extracted from the hearted leaves and stored at -80°C for detection. The primer sequences used in this study are listed in Supplementary Table S2.
The nucleotides of the extracted treatment and control groups were sequentially diluted and spotted on the Hybond-N+ membrane. UV crosslinking was then performed at UV 254 nm at 0.12 J/cm2. After blocking with 5% skim milk for 1 h, the membrane was incubated overnight at 4°C in TBST buffer containing anti-m6A antibody (synaptic system). Subsequently, the membrane was washed 3 times with TBST buffer and conjugated with HRP-conjugated secondary antibody (1: 5000, Abbkine Scientific Co., Ltd., California, USA, Cat. No. A21010) for 1 h. Visualized using ImmobilonTM Western HPR substrate Luminol Regeant (Merck Millipore). Further stained the membrane with methylene blue (G1301; Solarbio) to quantify RNA within the membrane (Zhang et al., 2022).
For the cold and heat treatments, wheat plants at the three-leaf stage (Yangmai 158) were exposed to 4°C or 42°C for 12 h. To induce drought stress, the seedlings were transferred to a Hoagland liquid culture supplemented with 20% polyethylene glycol (PEG, Sangon Biotech, Shanghai, China) and incubated for 12 h (Wu et al., 2023). For salinity, the plants were grown in Hoagland liquid culture containing 150 mM NaCl (Sangon Biotech, Shanghai, China) for 12 h. For hormone treatments, wheat plants at the three-leaf stage were treated with 100 μmol L−1 abscisic acid (ABA) and 100 μmol L−1 MeJA as exogenous sprays, as described in Yu et al (Yu et al., 2019). Wheat treated with distilled water was used as a control. Three biological replicates of the samples were collected at five different time points (0 h, 2 h, 4 h, 6 h, and 12 h). All samples were immediately frozen in liquid nitrogen after harvesting and were stored at −80°C until use. The expression of each gene was determined using RT–qPCR. The results were analyzed using GraphPad Prism 9. 0. 0.
Statistical analyses were performed using GraphPad Prism 9. 0. 0, and significant differences in the measured parameters were inferred according to Student’s t-test. Significant differences between the two groups of data were evaluated for comparisons (p < 0.05).
To identify members of the wheat ALKBH gene family, the BLASTP approach was utilized and employed 14 ALKBH protein sequences from Arabidopsis as a reference. Whole-genome comparisons were conducted for wheat, and the results were further validated through reciprocal BLASTP analysis (Supplementary Table S1). Based on these analyses, 30 genes were identified with a similarity greater than 60% between wheat and Arabidopsis. Table 1 provides a summary of the gene IDs, protein sizes, molecular weights, isoelectric points, and subcellular localization predictions. The sizes of the TaALKBH proteins varied from 255 to 649 aa, the molecular weights varied from 28.82 to 69.07 kDa, and the isoelectric points ranged from 4.97 to 10.33. The protein encoded by TraesCS4D02G230700 was the longest and presented the highest molecular weight (69.07 kDa), whereas TraesCS7D02G217000 encoded the shortest protein with the lowest molecular weight (28.82 kDa) (Table 1).
A phylogenetic tree was constructed utilizing MEGA-X to examine the phylogenetic relationships of ALKBH proteins across various species. The analysis encompassed 14 protein sequences from Arabidopsis (diploid), 30 from wheat (hexaploid), 9 from rice (diploid), and 11 from maize (diploid) (Figure 1A). The TaALKBH, OsALKBH, and ZmALKBH proteins exhibited high homology with the Arabidopsis AtALKBH proteins. Consistent with expectations, the ALKBH proteins from these four species were organized into seven distinct branches: ALKBH1, ALKBH2, ALKBH6, ALKBH7, ALKBH8, ALKBH9, and ALKBH10.
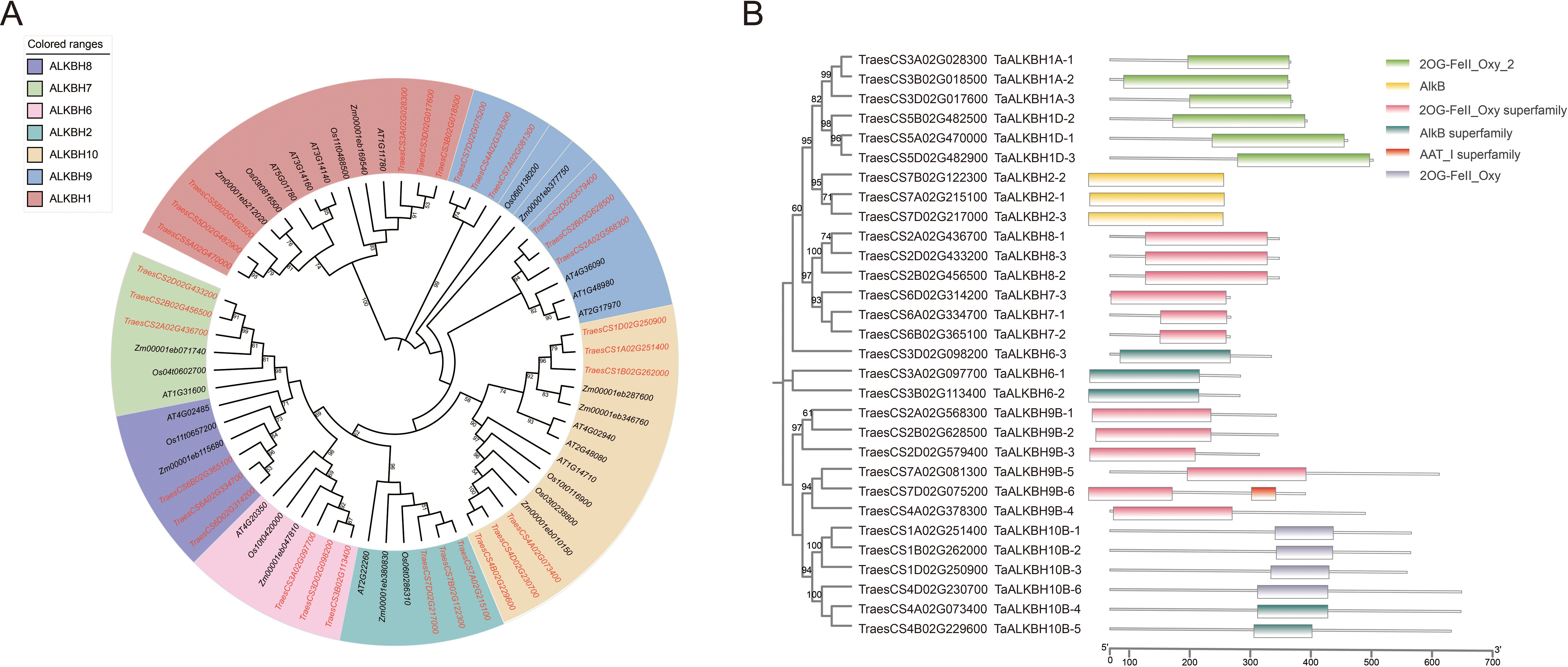
Figure 1. Phylogenetic and conserved-domain analyses of TaALKBHs. (A) Phylogenetic analysis of ALKBH family proteins in Arabidopsis thaliana, Triticum aestivum, Oryza sativa, and Zea mays. An unrooted phylogenetic tree was constructed via the maximum likelihood method implemented in MEGA-X software, with 1000 bootstrap replicates. Distinct colors are utilized to denote different groups. (B) Conserved-domain analysis of TaALKBHs. The various domains are depicted using boxes of distinct colors.
These TaALKBHs were named based on their homologous sequences in Arabidopsis (Figure 1B). All TaALKBH proteins exhibited conserved domains, which included 2OG-Fe (II)_Oxy, 2OGFe (II)_Oxy2, the 2OG-Fe (II)_Oxy superfamily, ALKB, and the AlkB superfamily (Figure 1B). The homologous proteins within each subfamily displayed identical conserved motifs, implying a potential for functional redundancy.
Conserved motifs are indicative of gene functions. An analysis of the amino acid sequences of the 30 TaALKBH proteins was conducted to identify conserved motifs using MEME. As illustrated in Figure 2A, a total of 15 highly conserved motifs were identified within these proteins. Notably, Motif 1 and Motif 5 were present across all subfamilies, while the ALKBH10 subfamily presented the greatest number of motifs. Additionally, Motif 7 was found to be specific to the ALKBH9B subfamily. The observation that members of the same subfamily displayed similar patterns of conserved motifs suggests functional similarities among these proteins (Figure 2A). Due to the similar conserved motifs among the genes within each subfamily, their subcellular localization was further analyzed. The predicted results showed that the same subfamily genes had similar subcellular localization (Supplementary Table S3). Furthermore, an analysis of the exon-intron structure offered insights into the evolutionary dynamics of gene families. Examination of the genomic DNA sequences of TaALKBHs revealed that genes within the same subfamily generally possess a similar number of exons, although variations in the lengths of exons and introns were noted (Figure 2B).
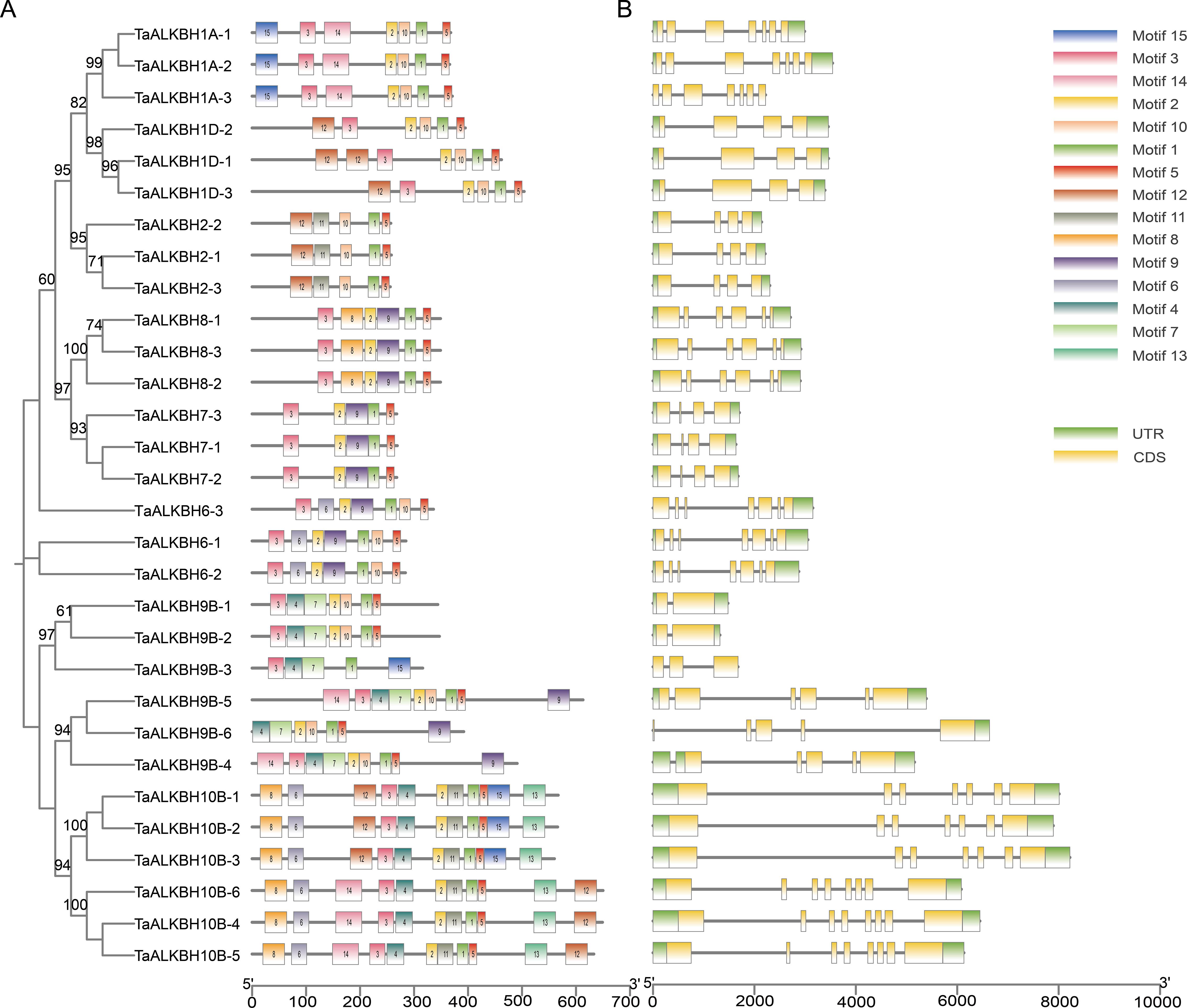

Figure 2. Analysis of conserved motifs and gene structures in TaALKBHs. (A) identification of conserved motifs in TaALKBHs. (B) distribution of gene structures in TaALKBHs.
The tertiary structure models of proteins are fundamentally associated with their functional capabilities. To construct tertiary structure models of the wheat TaALKBH protein, a representative from each subfamily was randomly selected for modeling purposes. The tertiary structure models of the ALKBH protein from Arabidopsis were utilized as a reference. The results indicated that the tertiary structure models of ALKBH1, ALKBH2, ALKBH6, ALKBH7, ALKBH8, and ALKBH10 from both species displayed significant similarities, whereas distinct differences were noted in the case of ALKBH9 (Figure 3). This result revealed the structural diversity of the ALKBH in the two species.
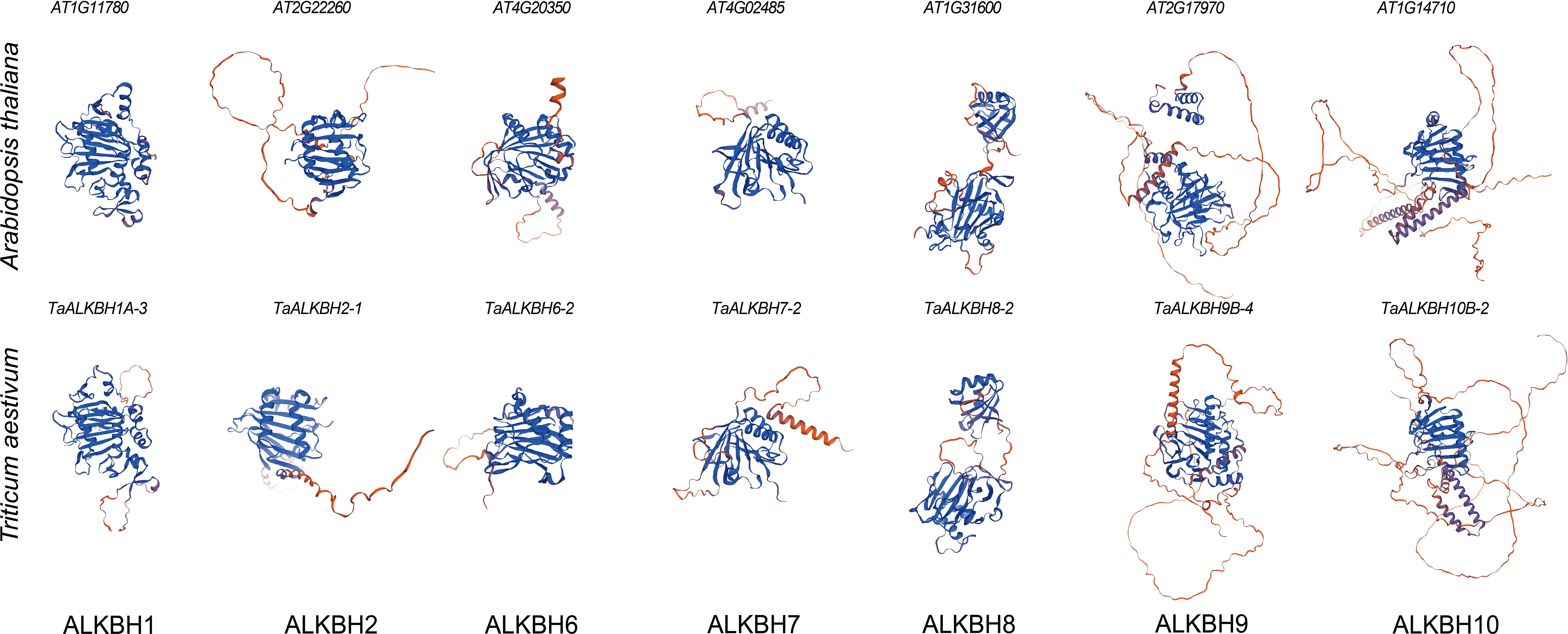
Figure 3. Tertiary structure models of ALKBH protein in Arabidopsis and wheat. Randomly selected a protein from each subfamily and used SWISS-MODEL for structural prediction. Based on QMEAN and GMQE, the model with the optimum results was selected.
The TaALKBH genes exhibited a random distribution across the 21 wheat chromosomes, with a majority situated in either the proximal or distal region (Figure 4A). Specifically, chromosomes 2A, 2B, 2D, 3A, 3B, 3D, 7A, 7B, and 7D each harbored two ALKBH genes, whereas chromosomes 1A, 1B, 1D, 4A, 4B, 4D, 5A, 5B, 5D, 6A, 6B, and 6D each contained a single gene. Regarding the gene density of the whole wheat genome, TaALKBH1A-1, TaALKBH1A-2, TaALKBH1A-3, TaALKBH9B-1, TaALKBH9B-2, and TaALKBH9B-3 were located in low-gene-density telomeric regions, which may indicate their role in gene duplication and evolutionary processes (Zhao et al., 2024). Gene duplication events were assessed via BLASTP and MCScanX, and the percentage of wheat genes was incorporated into the collinearity analysis. Among the 30 TaALKBH gene members, 28 segmental duplication pairs were identified (Figure 4B). In the context of genetics, the Ka/Ks ratio serves as an indicator of selective pressure acting on protein-coding genes (Hurst, 2002). Chromosomal covariance and synteny analyses revealed 28 putative paralogous genes (Ta-Ta) in the wheat genome. The Ka/Ks ratios for these genes ranged from 0.0505 to 0.5135. All the duplicated TaALKBH gene pairs presented Ka/Ks values of less than 1, suggesting a lack of strong positive selection. However, one gene pair presented a Ka/Ks value exceeding 0.5, indicating weak positive selection. A majority of the gene pairs presented Ka/Ks values less than 0.5, implying that the pairs had undergone purifying selection. The divergence time (T) was estimated via the formula million years ago (Mya). By this analysis, 28 pairs of homozygous (Ta-Ta) individuals were identified, with T values ranging from 2.342 to 13.241 Mya (Table 2).
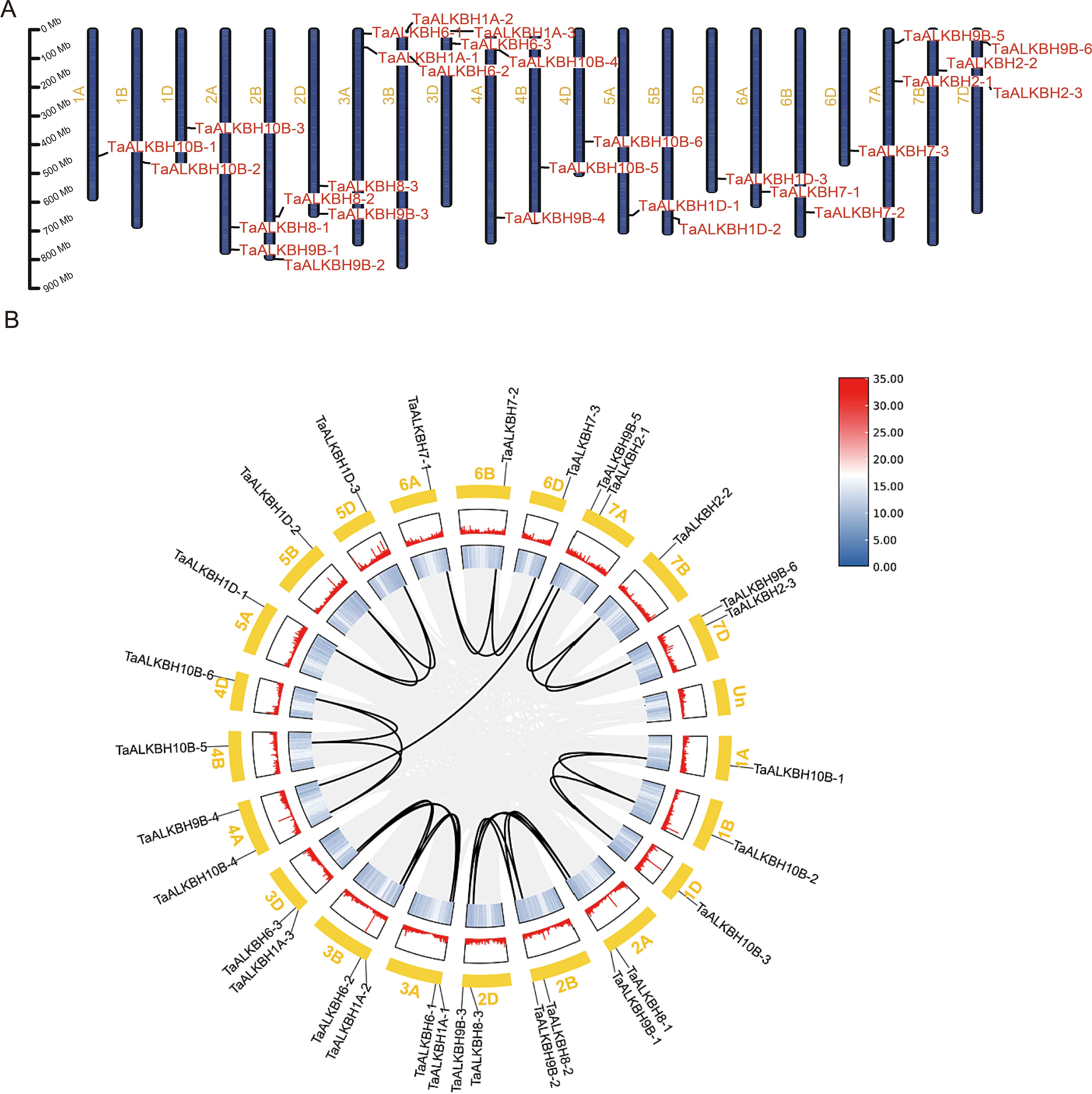
Figure 4. Genomic localization and collinearity analysis of TaALKBHs. (A) Distribution of TaALKBHs on chromosomes. The leftmost scale shows the chromosome length. Chromosomes are represented by green bars. (B) Concentric circles, from outer to inner, show (1) Triticum aestivum chromosomes, (2) GC content, (3) gene density, and (4) syntenic blocks; the 28 black lines represent collinear pairs of TaALKBHs.
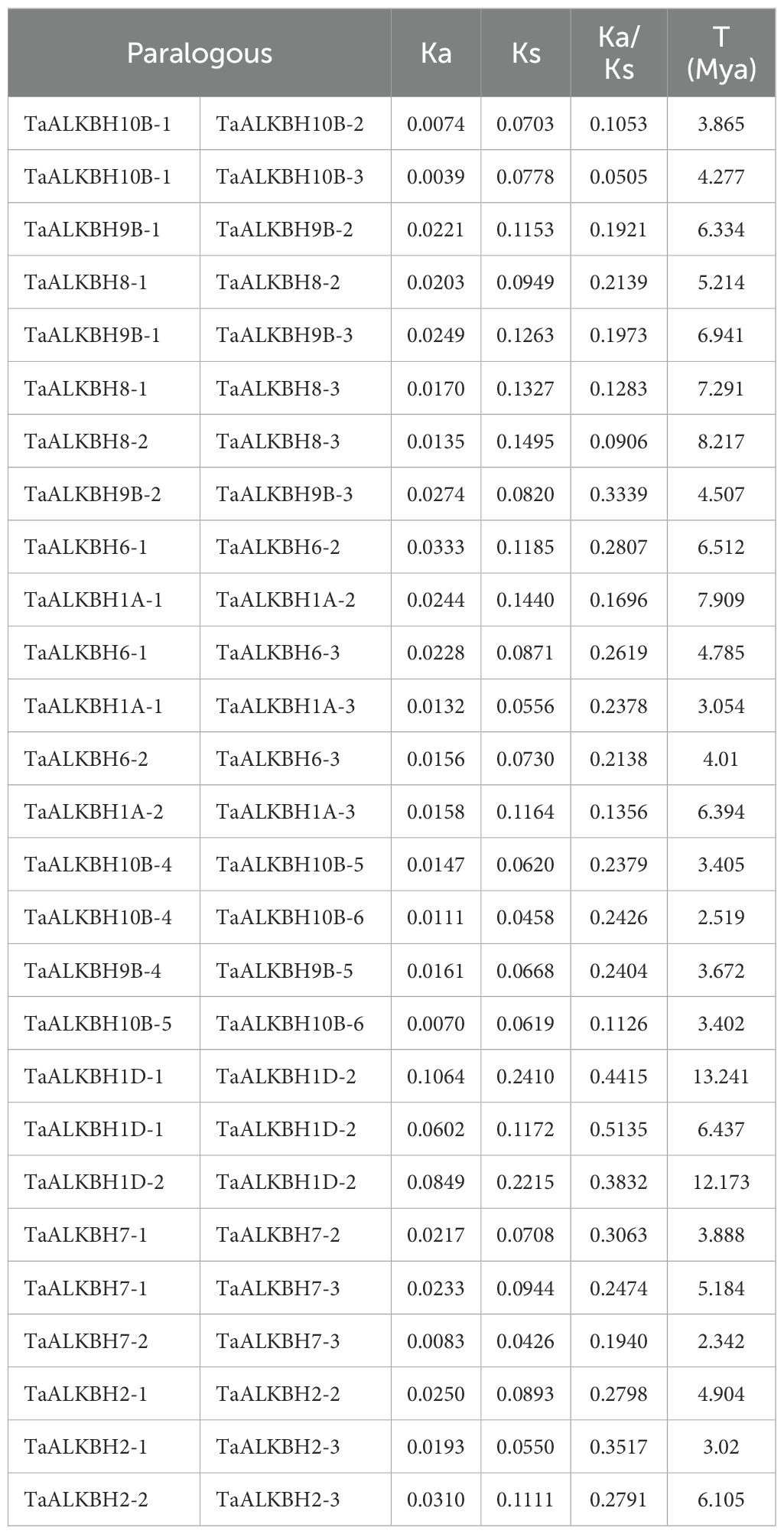
Table 2. Ks, Ka, and Ka/Ks values calculated for paralogous ALKBH gene pairs (T. aestivum–T. aestivum).
To improve the understanding of the potential involvement of TaALKBHs, cis-acting elements of 30 TaALKBHs were predicted through PlantCARE service using the 2000 bp promoter regions. This analysis revealed 23 functional classes of cis-acting elements associated with TaALKBHs. Depending on the different response conditions, these cis-acting elements could be categorized into five groups: light response, hormone response, environmental stress-related, developmental response, and miscellaneous elements (Figure 5A). Notably, hormone-response elements were present in all the promoters of the TaALKBHs, constituting 40.08% of the total identified cis-acting elements (Figure 5B). Among the hormone response elements, those associated with ABA, methyl jasmonate (MeJA), auxin, salicylic acid, and gibberellin presented a relatively high prevalence. The most frequently identified elements related to environmental stress were the low-temperature response elements. Furthermore, cis-acting elements related to developmental responses, as well as defensive and stress responses, were also detected. The significant abundance of light, MeJA, auxin, and gibberellin response elements in the promoters of the TaALKBHs suggest that the expression levels of these genes may be modulated by light exposure and various plant hormones, potentially influencing wheat development and response to environmental stimuli.
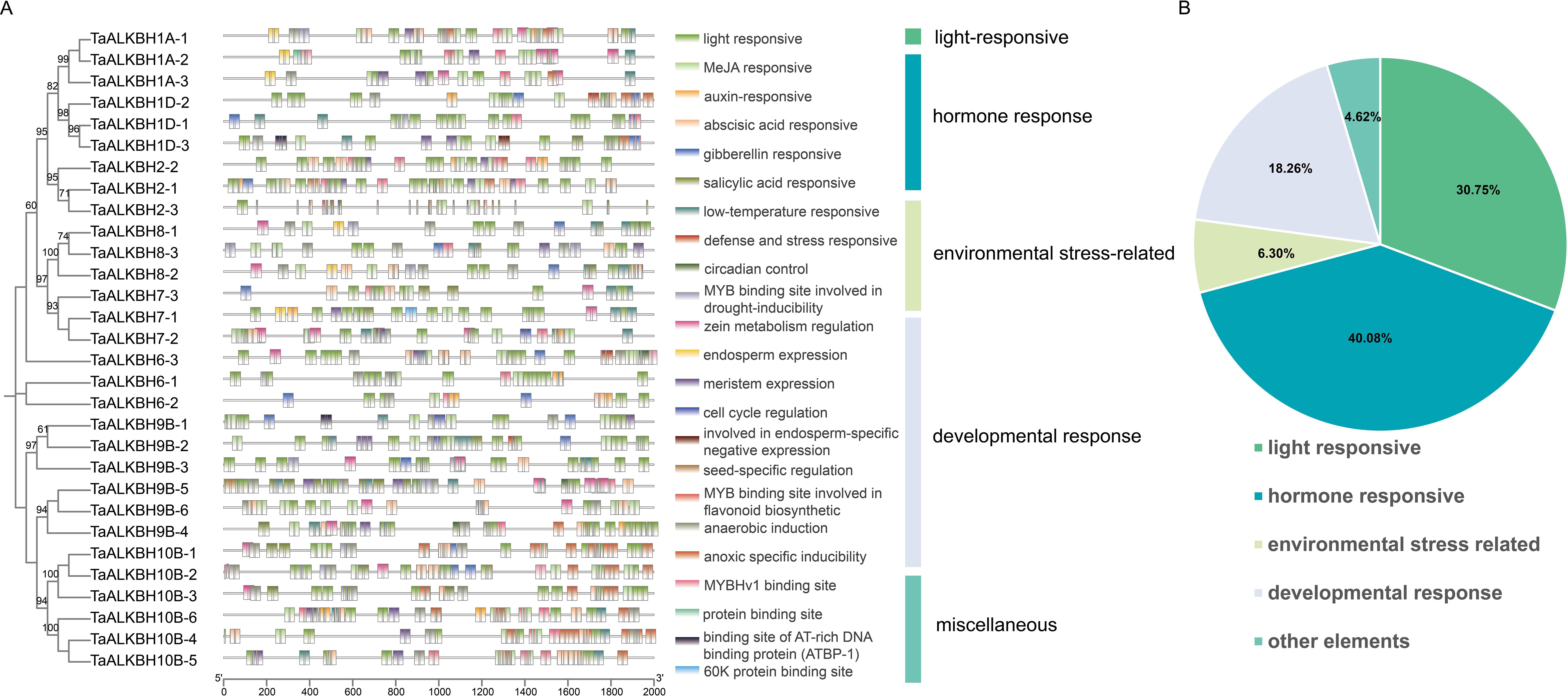
Figure 5. Functions of cis‐acting elements in the promoters of TaALKBHs. (A) Phylogenetic analysis of TaALKBHs. Cis‐acting element distribution in the TaALKBH promoters. Different cis-acting elements are represented by different colors. (B) The classification of the cis‐elements and the proportions of different types of cis‐elements. The 953 cis-elements were divided into five groups, including 293 light-responsive elements, 382 hormone responsive elements, 60 environmental stress-related elements, 174 developmental responsive elements, and 44 other elements.
To enhance the understanding of the functional roles of TaALKBHs, it is essential to delineate their expression profiles across different tissues. In wheat at the three-leaf stage, the plant is categorized into three distinct parts: roots, stems, and leaves. The physicochemical characteristics of each member of the ALKBH subfamily, as detailed in Table 1, correspond with the similarities identified in the conserved domains among the members of this subfamily, as depicted in Figure 1B. Additionally, the distribution of gene structure, and the conserved motifs present in each member of the ALKBH subfamily, as illustrated in Figure 2, demonstrate significant similarities. The expression patterns of each subfamily are hypothesized to exhibit similarities based on these results. Results from RT-qPCR analysis revealed that all members of the ALKBH9 subfamily, which comprises the largest number of members, exhibited high expression levels in both stems and leaves (Figure 6A). Consequently, the expression patterns observed in the members of each subfamily are considered somewhat representative of the entire subfamily. To conduct tissue-specific differential expression analysis, one gene was randomly selected from each group (Figure 6B). These results indicated that the expression and distribution patterns of TaALKBHs across the seven groups displayed significant variability. Notably, there was no substantial difference in the expression levels of TaALKBH1D-1 and TaALKBH7-3 in either rhizomes or leaves. In contrast, TaALKBH2-3, TaALKBH6-2, and TaALKBH8-2 exhibited the highest expression levels in the stems. Additionally, the expression levels of TaALKBH9B-5 and TaALKBH10B-4 in the roots and stems were significantly higher than those observed in the roots alone, with TaALKBH9B-5 demonstrating the most pronounced expression. These results imply that the expression patterns of TaALKBHs are tissue-specific and may correlate with the various developmental stages of the plant.
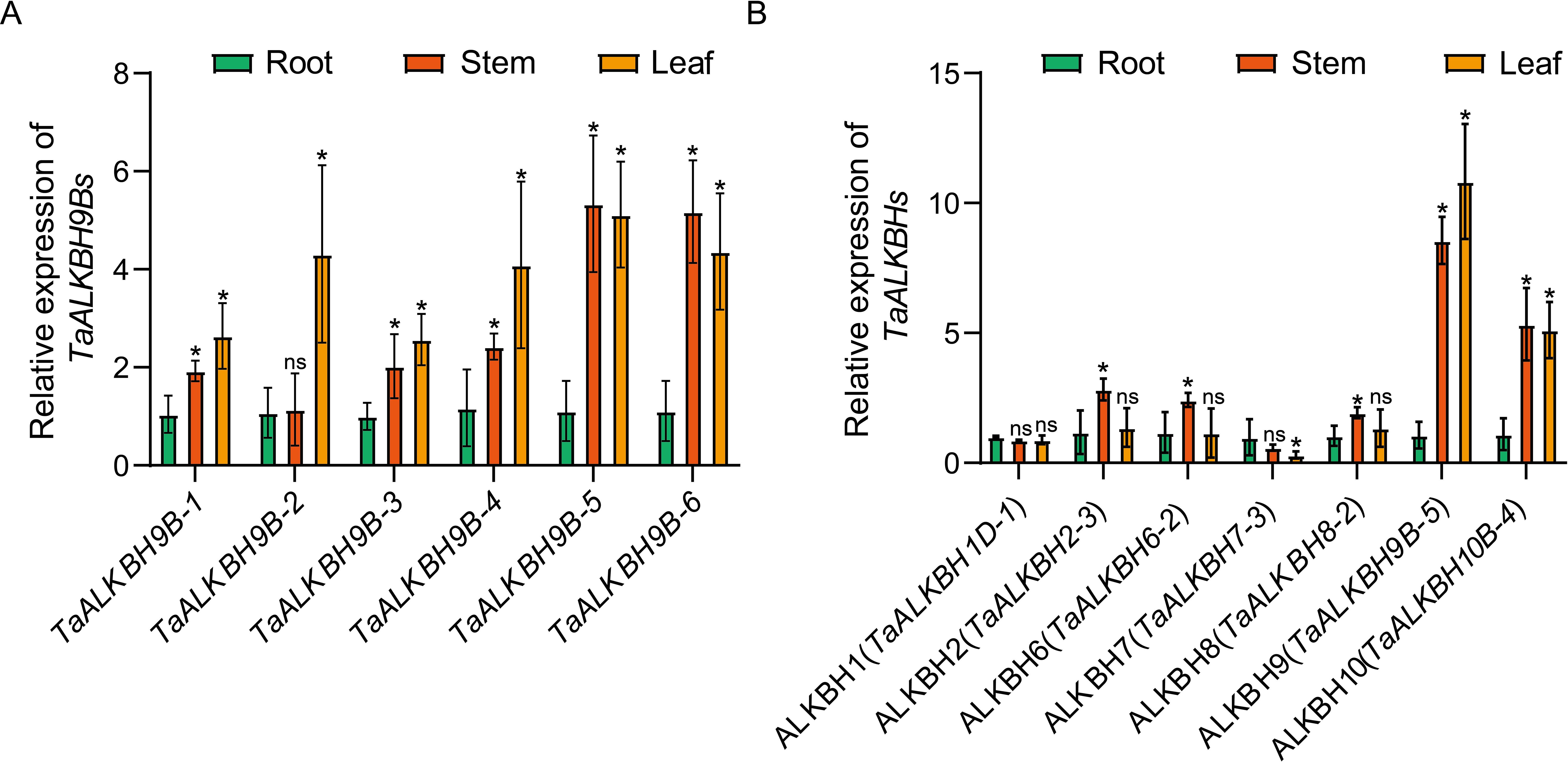
Figure 6. Tissue-specific differential expression of TaALKBHs. (A) Relative expression levels of TaALKBH9Bs genes in roots stems, and leaves. (B) Relative expression levels of TaALKBHs. The genes were randomly selected from each subfamily. All of the results shown were normalized to TaCDC expression as an internal control. Each value represents the mean ± SE of three replicates, and the asterisks represent significant differences between the test group and the control group (* p < 0.05, ns indicating no significant difference, Student’s).
In Arabidopsis, AtALKBH9B exhibits demethylase activity (Martínez-Pérez et al., 2017). TaALKBH9B-5 exhibited the most significant variation in tissue-specific expression profiles, it was selected for subsequent validation. Subsequently, the demethylase activity of TaALKBH9Bs was investigated. First, the TaALKBH9B-5 was inserted into GFP (a vector harboring the green fluorescent protein gene) and overexpressed in wheat cells through protoplast transformation, with GFP overexpression as a control. Total RNA was extracted using alkaline lysis, and m6A methylation levels were assessed through m6A spot hybridization. As shown in Figure 7A, cells overexpressing TaALKBH9B-5 presented lower levels of m6A methylation in total RNA than the control cells (Figure 7A). BSMV-mediated gene silencing techniques were employed to silence TaALKBH9B-5 in wheat. The results indicated that the overall m6A methylation level of total RNA in wheat plants with TaALKBH9B-5 silenced was greater than that in the control group (Figure 7B).
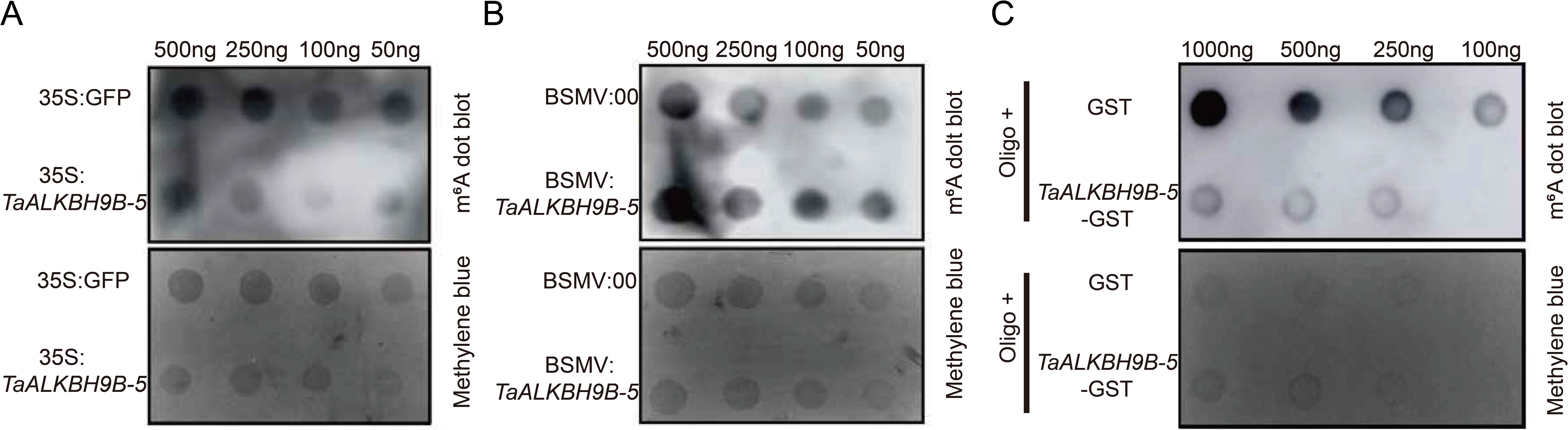
Figure 7. Identification of the m6A demethylase activity of TaALKBH9B-5. (A) RNA dot blot analysis of m6A-modified RNA in the total RNA of 35S: TaALKBH9B-5-GFP-transfected wheat protoplasts. 35S: GFP was used to transform the total RNA of the protoplasts as a negative control. (B) RNA dot blot analysis of m6A-modified RNA in the total RNA after BSMV-mediated silencing of TaALKBH9B-5. The total RNA from BSMV: 00 was used as a negative control. (C) RNA dot blot analysis of m6A-modified RNA in vitro. GST and TaALKBH-5-GST were purified from E. coli. The GST protein was used as a negative control. The membrane was further stained with methylene blue to quantify the RNA within the membrane.
Previous studies have reported that ALKBH9B and ALKBH10B exhibit in vitro demethylation activities. To obtain the TaALKBH9B-5 protein with in vitro activity, the TaALKBH9B-5 was inserted into pGEX4T-2 (a prokaryotic expression vector encoding GST). TaALKBH9B-5-GST was coincubated with the GST vector and Oligo-. Compared with that in the GST group, the methylation level in the TaALKBH9B-5-GST supplementation group was lower. These findings suggest that TaALKBH9B-5 functions as an m6A demethylase in wheat (Figure 7C).
The analysis of cis-acting elements indicated that the expression of TaALKBH9B-5 may be modulated by abiotic stress. To further elucidate the role of TaALKBH9B-5 in the response of wheat to abiotic stress, the relative expression levels of this gene were assessed in wheat leaves at the three-leaf stage under both normal conditions and various stress conditions, including high temperature, low temperature, drought, and salinity (Figures 8A–D). In conclusion, the expression of TaALKBH9B-5 changed significantly under the four stress treatments. First, the expression of TaALKBH9B-5 was down-regulated considerably in wheat leaves after 2 hours of treatment at 42°C (Figure 8A). Following 4 hours of treatment at 4°C, the expression of TaALKBH9B-5 was significantly up-regulated in wheat leaves; however, after 12 hours, the expression level of TaALKBH9B-5 was slightly lower than the control group (Figures 8B). Additionally, after 4 hours of treatment with 20% PEG4000, the expression of TaALKBH9B-5 was significantly down-regulated (Figure 8C), whereas TaALKBH9B-5 was down-regulated considerably after 2 hours of treatment with 150 mM NaCl. These results suggest that TaALKBH9B-5 may mediate the demethylation of RNA m6A and participate in the response of wheat to abiotic stresses.
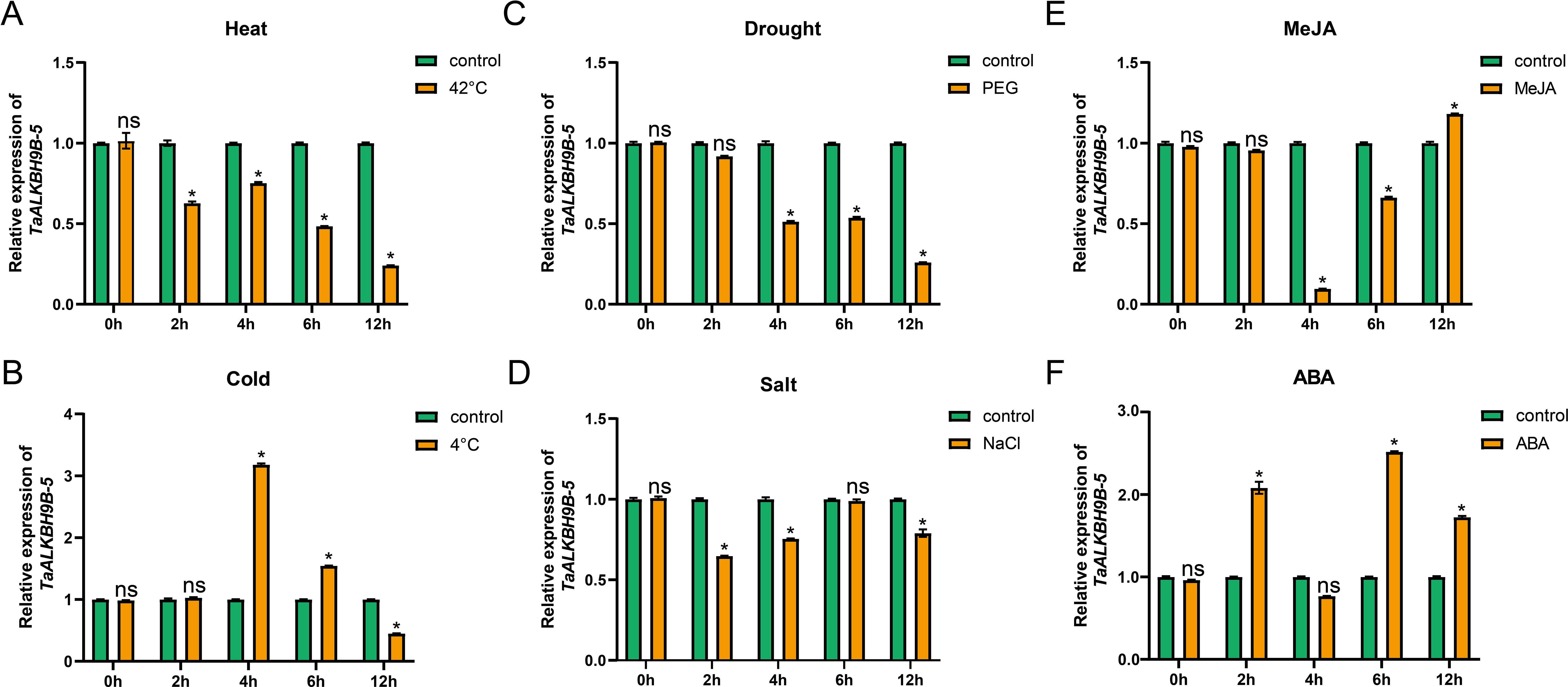
Figure 8. Relative expression level of TaALKBH9B-5 under abiotic stress. (A-D) The relative expression levels of TaALKBH9B-5 genes in leaves under heat (A), cold (B), polyethylene glycol [PEG, (C)], and salt (D) stress were quantified using real-time quantitative PCR (RT–qPCR). (E, F) RT-qPCR of TaALKBH9B-5 genes in leaves under MeJA (E) and ABA (F) treatment. All of the results shown were normalized to TaCDC expression as an internal control Each value represents the mean ± SE of three replicates, and the asterisks represent significant differences between the test group and the control group (* p < 0.05, ns indicating no significant difference, Student’s).
Recent studies underscore the critical role of plant hormones in the regulation of growth and development. Through gene family analysis, hormone-responsive elements have been identified. To assess hormone responsiveness, three-leaf-stage wheat plants were used for treatments with MeJA and ABA. The cis-acting element analysis revealed the highest number of associated responsive elements (Supplementary Table S4). The expression of TaALKBH9B-5 was measured (Figures 8E, F). The findings indicated that TaALKBH9B-5 exhibited a strong response to both MeJA and ABA; however, the nature of these responses was distinct. Specifically, the expression level of TaALKBH9B-5 decreased significantly following 4 hours of MeJA treatment (Figures 8E), while it increased markedly after 2 hours of ABA treatment (Figures 8F).
The ALKBH gene family is integral to plant growth, development, and responses to abiotic stressors through regulating RNA m6A demethylation (Tang et al., 2022). The advancement of whole-genome sequencing technologies has significantly improved the accessibility of numerous reference genomes, thereby facilitating the analysis of gene families. ALKBH genes have been identified across plant species, including sweet orange, poplar, soybean, and potato (Huang et al., 2024; Zhao et al., 2022; Li et al., 2024). However, the presence of the ALKBH gene family in wheat has not yet been documented. In the present study, a total of 30 TaALKBH genes were identified by comparing the wheat reference genome from the Ensembl Plants database with 14 AtALKBH protein sequences from Arabidopsis, employing a BLASTP-based methodology (Supplementary Table S1). A phylogenetic tree was constructed that included the 30 TaALKBH protein sequences alongside OsALKBHs, ZmALKBHs, and AtALKBHs (Figure 1A), revealing a high degree of homology among the protein amino acids of these species. Furthermore, the TaALKBHs were categorized into seven distinct subfamilies, corresponding to the ALKBH proteins identified in Arabidopsis, specifically ALKBH1, ALKBH2, ALKBH6, ALKBH7, ALKBH8, ALKBH9, and ALKBH10 (Figure 1A). The conserved structural domain 2OG-Fe (II)-Oxy suggests these proteins may possess RNA demethylase activity (Figure 1B). Members of each subfamily exhibit similar gene structures and conserved motifs, indicating a shared evolutionary origin and functional similarities (Figure 2). Most TaALKBHs contain motif 1 and motif 5, which are likely catalytic domains associated with demethylase activity (Figure 2A). Significant differences in gene structure among various subfamilies imply potential functional divergence (Figure 2B). However, variations in gene structure within certain subfamilies may be attributed to self-mutation events within the wheat genome. These structural alterations could involve changes or loss of gene content in the intron-exon regions, potentially affecting gene functionality. The ALKBH proteins of Arabidopsis and wheat exhibit similar structural characteristics within the same subfamily, whereas proteins from different subfamilies within the same species display considerable structural variation, indicating a diverse structural composition of the ALKBH family across different species (Figure 3).
In biological evolution, gene duplication events serve as important evolutionary mechanisms that facilitate alterations in gene function and contribute to gene evolution (Huang et al., 2015). The primary evolutionary patterns observed in land plants include segmental duplication, tandem duplication, and transposition events (Zhang, 2003; Cannon et al., 2004). Notably, evolutionary patterns of identical gene families can vary across species (Qiao et al., 2018; Xu et al., 2020). In the present study, 28 pairs of TaALKBHs that underwent gene duplication were examined (Table 2). The majority of these pairs resulted from genome-wide duplication events, which contributed to the amplification of TaALKBHs (Figure 4). To assess the selection pressures acting on TaALKBHs and to identify the selection pressures influencing the duplicated gene pairs, Ka, Ks, and Ka/Ks ratios were calculated for each homologous gene (Table 2) (Hurst, 2002). The Ka/Ks values for all duplicated TaALKBH gene pairs were less than 1, suggesting the absence of strong positive selection for these gene pairs. These findings underscore the dynamic expansion of TaALKBHs and suggest the potential functional diversity or redundancy within wheat. Therefore, the comprehensive identification and analysis of TaALKBHs may provide valuable insights into the functions and mechanisms of RNA m6A methylation in plant growth and development.
The analysis of cis-acting elements indicated that all TaALKBHs may respond to light, phytohormones, plant growth and development signals, and environmental stresses (Figure 5). Among these genes, TaALKBH9B-5 presented the most significant differential expression between stems and leaves (Figure 6B). Previous studies have reported that ALKBH9B possesses m6A methyltransferase activity; however, few studies have examined TaALKBH9B in wheat. In our investigation, the m6A dot spot analysis results revealed that the RNA m6A level in wheat tissues overexpressing TaALKBH9B-5 was lower than that in the control group (Figure 7A). Conversely, the RNA m6A level in wheat with TaALKBH9B-5 silenced via BSMV-mediated gene silencing was also lower than that in the control (Figure 7B), suggesting that TaALKBH9B-5 can remove m6A from mRNA. In vitro experiments confirmed this finding (Figure 7C). Collectively, these results indicate that TaALKBH9B-5 functions as an m6A demethylase in wheat. In a related study, overexpression of PagALKBH9B and PagALKBH10B increased the salt tolerance of transgenic lines by mitigating H2O2 accumulation and oxidative damage through the increased activities of superoxide dismutase (SOD), peroxidase (POD), and catalase (CAT) while also bolstering protection against chlorophyll a/b (Zhao et al., 2022). In Arabidopsis, SG-localized AtALKBH9B selectively demethylates the heat-activated retrotransposon element Onsen, facilitating its release from spatial confinement and enabling its mobility (Fan et al., 2023). Silencing of ALKBH9B in Arabidopsis has been shown to result in hypersensitivity to ABA treatment during seed germination and early seedling development. The m6A residues in the transcripts of ABA INSENSITIVE 1 (ABI1) and BRI1-EMS-SUPPRESSOR 1 (BES1) were removed by ALKBH9B following ABA treatment, thereby affecting the stability of these mRNAs (Tang et al., 2022). In this study, cis-acting element analysis revealed that TaALKBH9B-5 is also responsive to ABA, cold, and drought conditions (Figure 5). The relative expression levels of TaALKBH9B-5 also changed under high-temperature, low-temperature, drought, and salinity conditions, but the expression levels of TaALKBH9B-5 varied among the different stress treatments (Figure 8). The expression of TaALKBH9B-5 was significantly downregulated after treatment with heat, PEG4000, or NaCl but upregulated after cold treatment (Figures 8A–D). Additionally, the expression of TaALKBH9B-5 was significantly upregulated after treatment with ABA (Figure 8F). Drought and salt stress were often associated with ABA, but TaALKBH9B-5 did not respond consistently to them in these results. This may be because TaALKBH9B-5 is mainly expressed in leaves, while drought stress and salt stress mainly affect plant roots, and the expression patterns of TaALKBH9B-5 in different tissues are different and play different roles (Hu et al., 2021). These findings suggest that TaALKBH9B-5 has considerable potential applications in regulating plant growth, development, and managing abiotic stress responses by mediating the demethylation of RNA m6A.
In this study, an analysis of the TaALKBH gene family was performed and identified. TaALKBH9B-5 as an m6A demethylase demonstrated a significant upregulation in response to both cold stress and abscisic acid treatment. A total of 30 ALKBH genes were identified in the wheat genome. Phylogenetic tree analysis revealed the presence of seven subfamilies. Members within each subfamily exhibited a high degree of amino acid sequence, gene structure, and conserved domain. The TaALKBH genes were found to be randomly distributed across 21 chromosomes of wheat. Segmental duplication was identified as the primary mechanism responsible for the expansion of the TaALKBH gene family. Predictions regarding the cis-acting elements in the promoters of each gene member indicated that a majority contained multiple hormone response elements and stress response elements. Tissue-specific analysis of the seven subfamilies revealed TaALKBH9B-5 exhibiting higher expression levels in stems and leaves than in roots. Dot blot analysis indicated that TaALKBH9B-5 could reduce m6A methylation levels under both in vivo and in vitro conditions, suggesting its potential role as an m6A demethylase in wheat. TaALKBH9B-5 was significantly upregulated in response to ABA treatment and cold stress. These findings establish a foundation for further investigation into the molecular mechanisms underlying TaALKBH-mediated RNA m6A demethylation and its involvement in plant growth, development, and responses to abiotic stresses.
The original contributions presented in the study are included in the article/Supplementary Material. Further inquiries can be directed to the corresponding author.
PW: Conceptualization, Data curation, Formal Analysis, Investigation, Methodology, Writing – original draft, Writing – review & editing. TZ: Investigation, Methodology, Project administration, Supervision, Writing – original draft, Writing – review & editing. ZW: Data curation, Formal Analysis, Investigation, Writing – review & editing. LY: Investigation, Writing – review & editing. PL: Investigation, Writing – review & editing. JY: Data curation, Methodology, Writing – review & editing. BS: Funding acquisition, Project administration, Resources, Validation, Visualization, Writing – review & editing.
The author(s) declare that financial support was received for the research and/or publication of this article. This work was supported by National Key R&D Program of China (2023YFD1400800), the China Agriculture Research System from the Ministry of Agriculture of the China (CARS-03); and was also supported by a HAU grant for collaborative crop science research (CCSR2022-2).
We thank Dawei Li for providing the BSMV-based gene vectors.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fpls.2025.1544879/full#supplementary-material
Amara, U., Shoaib, Y., Kang, H. (2022). ALKBH9C, a potential RNA m6A demethylase, regulates the response of Arabidopsis to abiotic stresses and abscisic acid. Plant Cell Environ. 45, 3566–3581. doi: 10.1111/pce.14447
Arribas-Hernández, L., Bressendorff, S., Hansen, M. H., Poulsen, C., Erdmann, S., Brodersen, P. (2018). An m6A-YTH module controls developmental timing and morphogenesis in arabidopsis. Plant Cell 30, 952–967. doi: 10.1105/tpc.17.00833
Bolser, D. M., Kerhornou, A., Walts, B., Kersey, P. (2015). Triticeae resources in ensembl plants. Plant Cell Physiol. 56, e3. doi: 10.1093/pcp/pcu183
Cannon, S. B., Mitra, A., Baumgarten, A., Young, N. D., May, G. (2004). The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol. 4, 10. doi: 10.1186/1471-2229-4-10
Chen, C., Chen, H., Zhang, Y., Thomas, H. R., Frank, M. H., He, Y., et al. (2020). TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol. Plant 13, 1194–1202. doi: 10.1016/j.molp.2020.06.009
Cui, C., Ma, Z., Wan, H., Gao, J., Zhou, B. (2022). GhALKBH10 negatively regulates salt tolerance in cotton. Plant Physiol. Biochem. 192, 87–100. doi: 10.1016/j.plaphy.2022.09.029
Duan, H.-C., Wei, L.-H., Zhang, C., Wang, Y., Chen, L., Lu, Z., et al. (2017). ALKBH10B is an RNA N6-methyladenosine demethylase affecting arabidopsis floral transition. Plant Cell 29, 2995–3011. doi: 10.1105/tpc.16.00912
Fan, W., Wang, L., Lei, Z., Li, H., Chu, J., Yan, M., et al. (2023). m6A RNA demethylase AtALKBH9B promotes mobilization of a heat-activated long terminal repeat retrotransposon in Arabidopsis. Sci. Adv. 9, eadf3292. doi: 10.1126/sciadv.adf3292
Finn, R. D., Bateman, A., Clements, J., Coggill, P., Eberhardt, R. Y., Eddy, S. R., et al. (2014). Pfam: the protein families database. Nucleic Acids Res. 42, D222–D230. doi: 10.1093/nar/gkt1223
Han, R., Shoaib, Y., Cai, J., Kang, H. (2023). ALKBH10B-mediated m6A demethylation is crucial for drought tolerance by affecting mRNA stability in Arabidopsis. Environ. Exp. Bot. 209, 105306. doi: 10.1016/j.envexpbot.2023.105306
Hou, Y., Sun, J., Wu, B., Gao, Y., Nie, H., Nie, Z., et al. (2021). CPSF30-L-mediated recognition of mRNA m6A modification controls alternative polyadenylation of nitrate signaling-related gene transcripts in Arabidopsis. Mol. Plant 14, 688–699. doi: 10.1016/j.molp.2021.01.013
Hu, W., Ren, Q., Chen, Y., Xu, G., Qian, Y. (2021). Genome-wide identification and analysis of WRKY gene family in maize provide insights into regulatory network in response to abiotic stresses. BMC Plant Biol. 21, 427. doi: 10.1186/s12870-021-03206-z
Huang, Y., Cao, L., Chen, T., Chang, X., Fang, Y., Wu, L. (2024). Genome-wide identification of the ATP-dependent zinc metalloprotease (FtsH) in Triticeae species reveals that TaFtsH-1 regulates cadmium tolerance in Triticum aestivum. PloS One 19, e0316486. doi: 10.1371/journal.pone.0316486
Huang, Z., Duan, W., Song, X., Tang, J., Wu, P., Zhang, B., et al. (2015). Retention, molecular evolution, and expression divergence of the auxin/indole acetic acid and auxin response factor gene families in brassica rapa shed light on their evolution patterns in plants. Genome Biol. Evol. 8, 302–316. doi: 10.1093/gbe/evv259
Huong, T. T., Ngoc, L. N. T., Kang, H. (2020). Functional characterization of a putative RNA demethylase ALKBH6 in arabidopsis growth and abiotic stress responses. Int. J. Mol. Sci. 21, 6707. doi: 10.3390/ijms21186707
Huong, T. T., Yang, Z., Ngoc, L. N. T., Kang, H. (2022). ALKBH8B, a putative RNA demethylase, plays a role in the response of arabidopsis to salt stress and abscisic acid. J. Plant Biol. 65, 319–330. doi: 10.1007/s12374-022-09351-8
Hurst, L. D. (2002). The Ka/Ks ratio: diagnosing the form of sequence evolution. Trends Genet. 18, 486–487. doi: 10.1016/S0168-9525(02)02722-1
Jia, G., Fu, Y., Zhao, X., Dai, Q., Zheng, G., Yang, Y., et al. (2011). N6-Methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nat. Chem. Biol. 7, 885–887. doi: 10.1038/nchembio.687
Kataoka, H., Yamamoto, Y., Sekiguchi, M. (1983). A new gene (alkB) of Escherichia coli that controls sensitivity to methyl methane sulfonate. J. Bacteriol. 153, 1301–1307. doi: 10.1128/jb.153.3.1301-1307.1983
Kumar, S., Stecher, G., Li, M., Knyaz, C., Tamura, K. (2018). MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 35, 1547–1549. doi: 10.1093/molbev/msy096
Li, Y., Dong, X., Ma, J., Sui, C., Jian, H., Lv, D. (2024). Genome-wide identification and expression analysis of the ALKB homolog gene family in potato (Solanum tuberosum L.). Int. J. Mol. Sci. 25, 10984. doi: 10.3390/ijms252010984
Livak, K. J., Schmittgen, T. D. (2001). Analysis of relative gene expression data using real-time quantitative PCR and the 2–ΔΔCT method. Methods 25, 402–408. doi: 10.1006/meth.2001.1262
Marchler-Bauer, A., Bryant, S. H. (2004). CD-Search: protein domain annotations on the fly. Nucleic Acids Res. 32, W327–W331. doi: 10.1093/nar/gkh454
Martínez-Pérez, M., Aparicio, F., López-Gresa, M. P., Bellés, J. M., Sánchez-Navarro, J. A., Pallás, V. (2017). Arabidopsis m6A demethylase activity modulates viral infection of a plant virus and the m6A abundance in its genomic RNAs. Proc. Natl. Acad. Sci. U.S.A. 114, 10755–10760. doi: 10.1073/pnas.1703139114
Pan, Y., Jia, Y., Liu, W., Zhao, Q., Pan, W., Jia, Y., et al. (2024). Transcriptome-wide m6A methylation profile reveals its potential role underlying drought response in wheat (Triticum aestivum L.). Planta 260, 65. doi: 10.1007/s00425-024-04491-2
Qiao, X., Yin, H., Li, L., Wang, R., Wu, J., Wu, J., et al. (2018). Different modes of gene duplication show divergent evolutionary patterns and contribute differently to the expansion of gene families involved in important fruit traits in pear (Pyrus bretschneideri). Front. Plant Sci. 9. doi: 10.3389/fpls.2018.00161
Růžička, K., Zhang, M., Campilho, A., Bodi, Z., Kashif, M., Saleh, M., et al. (2017). Identification of factors required for m6A mRNA methylation in Arabidopsis reveals a role for the conserved E3 ubiquitin ligase HAKAI. New Phytol. 215, 157–172. doi: 10.1111/nph.14586
Shen, H., Zhou, Y., Liao, C., Xie, Q., Chen, G., Hu, Z., et al. (2023). The alkB homolog slALKBH10B negatively affects drought and salt tolerance in solanum lycopersicum. Int. J. Mol. Sci. 25, 173. doi: 10.3390/ijms25010173
Tang, J., Yang, J., Lu, Q., Tang, Q., Chen, S., Jia, G. (2022). The RNA N6-methyladenosine demethylase ALKBH9B modulates ABA responses in Arabidopsis. J. Integr. Plant Biol. 64, 2361–2373. doi: 10.1111/jipb.13394
Wang, C., Yang, J., Song, P., Zhang, W., Lu, Q., Yu, Q., et al. (2022). FIONA1 is an RNA N6-methyladenosine methyltransferase affecting Arabidopsis photomorphogenesis and flowering. Genome Biol. 23, 40. doi: 10.1186/s13059-022-02612-2
Wei, L.-H., Song, P., Wang, Y., Lu, Z., Tang, Q., Yu, Q., et al. (2018). The m6A Reader ECT2 Controls Trichome Morphology by Affecting mRNA Stability in Arabidopsis. Plant Cell 30, 968–985. doi: 10.1105/tpc.17.00934
Wu, M., Tu, A., Feng, H., Guo, Y., Xu, G., Shi, J., et al. (2023). Genome-wide identification and analysis of the ABCF gene family in triticum aestivum. Int. J. Mol. Sci. 24, 16478. doi: 10.3390/ijms242216478
Xu, W., Zhao, Y., Chen, S., Xie, J., Zhang, D. (2020). Evolution and functional divergence of the fructokinase gene family in populus. Front. Plant Sci. 11. doi: 10.3389/fpls.2020.00484
Yang, J., Wei, H., Hou, M., Chen, L., Zou, T., Ding, H., et al. (2023). ZmSPL13 and ZmSPL29 act together to promote vegetative and reproductive transition in maize. New Phytol. 239, 1505–1520. doi: 10.1111/nph.19005
Yang, J., Zhang, T., Li, J., Wu, N., Wu, G., Yang, J., et al. (2020). Chinese wheat mosaic virus-derived vsiRNA-20 can regulate virus infection in wheat through inhibition of vacuolar-(H+)-PPase induced cell death. New Phytol. 226, 205–220. doi: 10.1111/nph.16358
Yu, X., Han, J., Wang, E., Xiao, J., Hu, R., Yang, G., et al. (2019). Genome-wide identification and homoeologous expression analysis of PP2C genes in wheat (Triticum aestivum L.). Front. Genet. 10. doi: 10.3389/fgene.2019.00561
Yu, C.-S., Lin, C.-J., Hwang, J.-K. (2004). Predicting subcellular localization of proteins for Gram-negative bacteria by support vector machines based on n-peptide compositions. Protein Sci. 13, 1402–1406. doi: 10.1110/ps.03479604
Yue, J., Wei, Y., Sun, Z., Chen, Y., Wei, X., Wang, H., et al. (2022a). AlkB RNA demethylase homologues and N6-methyladenosine are involved in Potyvirus infection. Mol. Plant Pathol. 23, 1555. doi: 10.1111/mpp.13239
Yue, J., Wei, Y., Zhao, M. (2022b). The reversible methylation of m6A is involved in plant virus infection. Biol. (Basel). 11, 271. doi: 10.3390/biology11020271
Zhang, J. (2003). Evolution by gene duplication: an update. Trends Ecol. Evol. 18, 292–298. doi: 10.1016/S0169-5347(03)00033-8
Zhang, D., Li, M., Chen, C., Wang, Y., Cheng, Z., Li, W., et al. (2024). Downregulation of GhALKBH10B improves drought tolerance through increasing the stability of photosynthesis related- and ABA signaling pathway genes in cotton. Environ. Exp. Bot. 220, 105687. doi: 10.1016/j.envexpbot.2024.105687
Zhang, T., Liu, P., Zhong, K., Zhang, F., Xu, M., He, L., et al. (2019). Wheat yellow mosaic virus NIb interacting with host light induced protein (LIP) facilitates its infection through perturbing the abscisic acid pathway in wheat. Biol. (Basel). 8, 80. doi: 10.3390/biology8040080
Zhang, T., Shi, C., Hu, H., Zhang, Z., Wang, Z., Chen, Z., et al. (2022). N6-methyladenosine RNA modification promotes viral genomic RNA stability and infection. Nat. Commun. 13, 6576. doi: 10.1038/s41467-022-34362-x
Zhao, Y., Guo, Q., Cao, S., Tian, Y., Han, K., Sun, Y., et al. (2022). Genome-wide identification of the AlkB homologs gene family, PagALKBH9B and PagALKBH10B regulated salt stress response in Populus. Front. Plant Sci. 13. doi: 10.3389/fpls.2022.994154
Zhao, J., Yang, T., Liu, P., Liu, H., Zhang, H., Guo, S., et al. (2024). Genome-Wide Identification of the Soybean AlkB Homologue Gene Family and Functional Characterization of GmALKBH10Bs as RNA m6A Demethylases and Expression Patterns under Abiotic Stress. Plants 13, 2491. doi: 10.3390/plants13172491
Zheng, G., Dahl, J. A., Niu, Y., Fedorcsak, P., Huang, C.-M., Li, C. J., et al. (2013). ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol. Cell 49, 18–29. doi: 10.1016/j.molcel.2012.10.015
Zheng, L., Wan, Q., Wang, H., Guo, C., Niu, X., Zhang, X., et al. (2022). Genome-wide identification and expression of TIFY family in cassava (Manihot esculenta Crantz). Front. Plant Sci. 13. doi: 10.3389/fpls.2022.1017840
Keywords: AlkB homologs, wheat, m6A RNA methylation, demethylase, abiotic stress
Citation: Wang P, Zhang T, Wu Z, Yu L, Liao P, Yang J and Sun B (2025) Genome-wide identification and characterization of ALKB homolog gene family in wheat (Triticum aestivum L.). Front. Plant Sci. 16:1544879. doi: 10.3389/fpls.2025.1544879
Received: 13 December 2024; Accepted: 28 February 2025;
Published: 18 March 2025.
Edited by:
Tianlun Zhao, Zhejiang University, ChinaReviewed by:
Lirong Wei, Nanjing Agricultural University, ChinaCopyright © 2025 Wang, Zhang, Wu, Yu, Liao, Yang and Sun. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Bingjian Sun, c3VuYmluZ2ppYW5AaGVuYXUuZWR1LmNu
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.