
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Plant Sci. , 27 March 2025
Sec. Plant Bioinformatics
Volume 16 - 2025 | https://doi.org/10.3389/fpls.2025.1541320
Introduction: The genus Impatiens L. ( Balsaminaceae) is one of the three most important bedding plant genera globally, valued for its medicinal, ornamental, and economic properties. However, the morphological overlap among species and the lack of genomic data have limited our understanding of their molecular phylogenetic relationships.
Methods: This study involved the sequencing of the chloroplast genomes of 9 Impatiens species, including Impatiens lateristachys, Impatiens siculifer var. porphyrea, Impatiens apalophylla, Impatiens pritzelii, Impatiens menghuochengensis, Impatiens membranifolia, Impatiens qingchengshanica, Impatiens aquatilis, and Impatiens racemosa. The study evaluated sequence divergence by comparing genomic features, repeat sequences, codon usage, IR expansion and contraction, sequence alignment, and selective pressures. It then constructed phylogenetic relationships using the maximum likelihood method, revealing the evolutionary relationships among these species.
Results: The results indicated that the chloroplast genome sizes ranged from 151, 784 bp (I. racemosa) to 152, 628 bp (I. apalophylla), encoding between 108-115genes[77 to 81 protein-coding genes, 27 to 30 tRNA genes, and 4 rRNA genes]. Additionally, A detailed analysis was performed on the characteristics of repeat sequences, codon preferences, and IR region. Coding sections were more conserved than non-coding regions, and the IR regions were more conserved than the LSC and SSC regions, according to sequence variation and mutation hotspot analyses. The 9 species of Impatiens were classified into subgenus Clavicarpa and subgenus Impatiens, including the sections Impatiens and Racemosae, according to the phylogenetic tree.
Discussion: This study presents the chloroplast genomes of 9 species within the genus Impatiens, marking a novel attempt at using phylogenetic analysis to determine the taxonomic positions of Impatiens species. It provides new molecular evidence for the systematic and evolutionary studies of Impatiens species.
The genus Impatiens L. was established by the botanist Carl Linnaeus in 1753 (Linnaeus, 1753). It belongs to the order Ericales, which includes only two genera: Hydrocera and Impatiens. Impatiens plants are annual or perennial herbaceous species within the genus Impatiens of the Balsaminaceae. They hold significant ornamental, economic, and medicinal value (Yu, 2008; Gao, 2012). The genus Impatiens has a broad distribution, primarily across tropical and subtropical regions, extending into tropical Africa, Southwest Asia, southern China, Europe, Russia, and temperate areas of North America (Grey-Wilson, 1989; Yu, 2012). Impatiens species exhibit diverse morphology and flower colors and are recognized as among the top three ornamental plants for flowerbeds and borders. Notably, I. hawkeri and I. walleriana are frequently utilized in horticulture, while many other species remain in the wild or are yet to be fully developed (Yu, 2012). China, particularly the karst regions, is recognized as the origin and center of diversification for the Balsaminaceae family. In Guizhou, Yunnan, and Guangxi, there are about 250 wild Impatiens species known to exist, many of these are utilized as supplements or medicines. In ancient China, Impatiens was known as “zhijiahua” and was ground into a paste for coloring nails (Cai et al., 2013). It was also referred to as “tougucao” and “jixingzi,” annual herbs used to cure onychomycosis, paronychia, rheumatism, beriberi, bruising, discomfort, and warts (Jiang et al., 2017; Kim C. S. et al., 2017).
The stems of Impatiens species are typically fleshy and succulent, characterized by thick leaves and delicate, membranous, fragile flowers. Floral parts often overlap and adhere during specimen pressing, making their separation and reconstruction difficult (Grey-Wilson, 1989). At the macroscopic level, the flowers, capsules, and seeds of Impatiens exhibit significant diversity. Consequently, the shape and size of the sepals, petals, capsules, and seeds are critical for species identification within this genus (Chen, 2001). In contrast to macroscopic features, the microstructural characteristics of pollen and seeds are less affected by environmental conditions (Janssens et al., 2012). Currently, the analysis of microstructural traits, such as pollen, leaf epidermis, and seed coat, has served as an important reference for taxonomy and phylogeny discussions (Guo, 2017; Shui et al., 2011; Pechimuthu et al., 2024). Molecular phylogenetics has significantly advanced our understanding of relationships within Impatiens. Fujihashi et al. (2002) published the first molecular phylogeny of Impatiens. This study, based on ITS (Internal Transcribed Spacer) sequences from 111 species, provided significant phylogenetic insights through a second independent molecular analysis (Yuan et al., 2004). Existing molecular classification studies have primarily focused on several chloroplast regions, including coding genes such as matK, rbcL, and trnK, as well as intergenic spacers like atpB-rbcL and trnL-trnF (Shajitha et al., 2016; Janssens et al., 2006). These studies have limited datasets and focus on a small number of samples with distinct regional characteristics.
For example, Rahelivololona et al. (2018) performed molecular phylogenetic analysis of 33 Impatiens species from Madagascar using nuclear and plastid data. However, molecular phylogenetic studies alone cloud not determine the distribution of morphological traits across evolutionary branches, nor can they fully capture the species diversity and complex phylogenetic relationships within Impatiens. In 2012, Yu et al. classified Impatiens into eight groups based on morphological traits and published the book “Balsaminaceae of China” (Yu, 2012). In 2016, Yu et al. proposed a new classification of Impatiens using sequencing data from three major genetic regions—nuclear ITS, chloroplast atpB-rbcL, and trnL-F—dividing the genus into two subgenera and seven groups. This classification provided significant data support for Impatiens resource taxonomy (Yu et al., 2016). To date, all molecular data are still derived from short sequences, many of which originate from samples with distinct regional characteristics. This limits the inferences for classification and phylogeny. For species exhibiting morphological diversity and controversial classification, molecular data alone are insufficient to provide conclusive evidence. Therefore, comprehensive studies integrating both morphological and molecular data are urgently needed to support the taxonomy and phylogeny of Impatiens. This study aims to resolve the phylogenetic relationships of Impatiens at higher resolution by using complete chloroplast genome sequences, thereby advancing taxonomy and updating research on the genus.
Chloroplasts are commonly found in the cytoplasm of higher plants and are responsible for synthesizing proteins, fatty acids, starch, pigments, and other compounds. They contain an independent and complete semi-autonomous genetic system known as the chloroplast genome (Janssens et al., 2016). Due to their ability to self-replicate, maternal inheritance, and relative conservatism, chloroplast genomes have become a significant tool in systematic research (Li et al., 2018). The chloroplast genomes of most angiosperms consist of a pair of inverted repeat sequences (IR), a large single-copy region (LSC), and a small single-copy region (SSC), with lengths ranging from 115 kb to 165 kb (Wang et al., 2024; Sun et al., 2024; Huang et al., 2024a). These genomes typically contain 110 to 113 genes, which are involved in gene expression (such as tRNA and rRNA genes), photosynthesis, and other metabolic functions (Zhou et al., 2020). The highly conserved nature of chloroplast genomes provides valuable molecular evidence for phylogenetic analysis and plant classification. Moreover, chloroplast genomes are useful for molecular markers, genetic modification, and plant barcode recognition (Gu et al., 2018). A detailed comparison of the chloroplast genomes of Impatiens species can deepen our understanding of their phylogenetic relationships and provide important insights for classification and evolutionary studies. Despite previous studies that utilized certain chloroplast genes (such as matK, rbcL, trnK, trnL-trnF, and atpB-rbcL) to investigate the phylogeny of Impatiens species, challenges remain (Shajitha et al., 2016; Yu et al., 2016). Current research primarily focuses on species that are unique to specific regions, leading to classification controversies for some morphologically complex species (such as I. lateristachys) due to unresolved phylogenetic relationships. The existing molecular data are based solely on a limited number of short gene sequences, which constrains their ability to resolve complex phylogenetic relationships.
Using entire chloroplast genomes, this study presented the chloroplast genomes of 9 species in the genus Impatiens, marking a novel attempt at phylogenetic analysis to determine the taxonomic position of Impatiens species. The objectives of this study were: (i) to conduct a comprehensive analysis of the chloroplast genomes of Impatiens species, including basic structural information of the chloroplast genomes, characteristics of repetitive sequences, codon usage preferences, region of inverted repeats (IR) expanding and contracting, comparative genomic differences, mutation hotspots, and analysis of selective pressures among species. (ii) to further understand the phylogenetic relationships of Impatiens species. (iii) to construct a phylogenetic tree based on complete chloroplast genomes and conduct taxonomic analysis. Research on the phylogeny, taxonomy, population genetics, and genetic engineering of Impatiens species can use this paper as a reference. It offers important information for the systematics and evolutionary studies of Impatiens species.
9 Impatiens species samples were gathered from different sites (Supplementary Table S1) and kept at Southwest Forestry University’s Plant Laboratory in Kunming, Yunnan Province. Fresh leaves from Province were gathered, put right away in liquid nitrogen, and then kept at -80℃ until analysis. Genomic DNA was extracted using the Omega DNA Extraction Kit (Beijing, China). A spectrophotometer was used to measure the concentration of roughly 5-10 μg of genomic DNA, and electrophoresis on a 1.5% agarose gel was used to confirm the integrity of the DNA (Doyle and Doyle, 1990).
Pairwise sequencing of the chloroplast genomes of 9 Impatiens species was carried out utilizing the Illumina NovaSeq 6000 platform. Adapter sequences and paired-end reads with a N content greater than 10% of the total read length were eliminated from the raw data during processing. Clean sequencing data was obtained by excluding paired-end reads that had low-quality bases (Q < 5) that accounted for more than 50% of the read length.
The chloroplast genomes of 9 Impatiens species were assembled using GetOrganelle v1.7.7.0 (Jin et al., 2020) with default parameters, resulting in complete circular chloroplast genome sequences. The assembled FASTA files were submitted to the online annotation tool Cpgavas2 (https://www.herbalgenomics.org/cpgavas2) (Shi et al., 2019) to obtain relevant sequence information for the chloroplast genomes.
The web program Bioinformatics Cloud (http://cloud.genepioneer.com:9929) was used to determine the GC content of four sections of the chloroplast genomes of 9 Impatiens species (Tong et al., 2022). To create chloroplast genome maps, the annotated GBF data were put into the web program Chloroplot (https://irscope.shinyapps.io/Chloroplot/) (Zheng et al., 2020).
The chloroplast genomes of 9 Impatiens species were subjected to simple sequence repeat (SSR) analysis using the web application MISA (https://webblast.ipk-gatersleben.de/misa/index.php) (Thiel et al., 2003). The parameters for repeat units of one to six nucleotides were set to 10, 6, 4, 3, 3, and 3, with a minimum distance of 100 bp between two SSRs. The online software REPuter (https://bibiserv.cebitec.uni-bielefeld.de/reputer) (Kurtz et al., 2001) was utilized to analyze the scattered repeats in the cpDNA of 9 Impatiens species. The parameters were set as follows: the minimum repeat size was set to 30 bp, the Hamming distance was set to 3, and the sequence identity was set to 90%.
Codon usage frequencies of synonymous codons in the cpDNA of 9 Impatiens species were calculated using CodonW 1.4.4 (Langmead et al., 2009) with default parameters. A codon heatmap was generated and enhanced using TBtools.
The boundary regions of the large single-copy (LSC), inverted repeat (IR), and small single-copy (SSC) regions of the cpDNA of 9 Impatiens species were compared using the CPJSdraw boundary mapping tool available on the online platform GenePioneer (http://cloud.genepioneer.com:9929). This analysis focused on the variations in the positions of these regions. The online tool mVISTA (http://genome.lbl.gov/vista/index.shtml) (Frazer et al., 2004) was used to visualize the cpDNA of 9 Impatiens species, taking Hydrocera triflora as a reference sequence under the Shuffle-LAGAN model. This comparison focused on the differences among exons, introns, non-coding regions, and coding regions within the chloroplast genomes, thereby highlighting conserved and variable regions among species.
MAFFT software (Katoh and Standley, 2013) was used to align the FASTA files of the 9 species, followed by manual correction using MEGA7 (Kumar et al., 2016). Nucleotide diversity analysis was conducted using DnaSP5 software (Rozas et al., 2017). Using Hydrocera triflora as the reference species, the CPStools (Huang et al., 2024b) package was downloaded and installed via Python software. The Ka/Ks analysis module of CPStools was then employed to import GenBank (GB) files for analysis. Subsequently, a clustering heatmap of Ka/Ks values was generated using TBtools (Chen et al., 2023) software to visually present the results.
The chloroplast genome sequences of 24 species from the Ericales order were selected, including the 9 Impatiens species newly sequenced in this study and an additional 15 species downloaded from NCBI. These 14 Ericales species comprised 11 from the Impatiens genus, one from the Hydrocera genus, and one species each from the Primulaceae and Actinidiaceae (Supplementary Table S12). Using the Primulaceae and Actinidiaceae as outgroups, chloroplast genome sequences were aligned using the online MAFFT tool (https://www.ebi.ac.uk/Tools/msa/mafft/). After successful alignment, Gblocks (http://www.phylogeny.fr/one_task.cgi?task_type=gblocks) was used to trim the conserved regions. Phylogenetic trees were then reconstructed using the maximum likelihood method implemented in IQtree (Nguyen et al., 2015), with default parameters (1000 iterations, 1000 bootstraps, and model selection). The best-fit model GTR+F+I+R5 was used to construct the tree, and the results were visualized and refined using Figtree software (http://tree.bio.ed.ac.uk/software/figtree).
The chloroplast genomes of the 9 Impatiens species analyzed were typical circular DNA molecules, with GC content in each region consistent with previously published data on Impatiens chloroplast genomes. The chloroplast genomes had an average GC content of 37%, with sizes ranging from 151, 784 bp in I. racemosa to 152,628 bp in I. apalophylla. The large single-copy (LSC) region sizes ranged from 82, 995 bp in I. lateristachys to 83, 480 bp in I. siculifer var. porphyrea), with a GC content of 34% to 35%. The small single-copy (SSC) regions exhibited GC content of 29-30%, with sizes ranging from 17, 253 bp in I. pritzelii to 17, 893 bp in I. qingchengshanica. The inverted repeat (IR) region sizes ranged from 25, 535 bp in I. racemosa to 25, 883 bp in I. aquatilis, with a GC content of 43%. The average AT content across the chloroplast genomes of the 9 Impatiens species was 63.12%, while the average GC content was 36.88%. No significant interspecific difference in GC content were observed among the basic structural regions (LSC, IR, and SSC). Additionally, the lowest GC contents were 29.26% in the SSC, 34.32% in the LSC, and 43.04% in the IR regions. The LSC and IR sections had significantly higher GC content compared to the SSC region (Table 1, Figure 1; Supplementary Figure S1). These results indicated that the 9 Impatiens species differed in the lengths of their chloroplast genomes and their GC content.
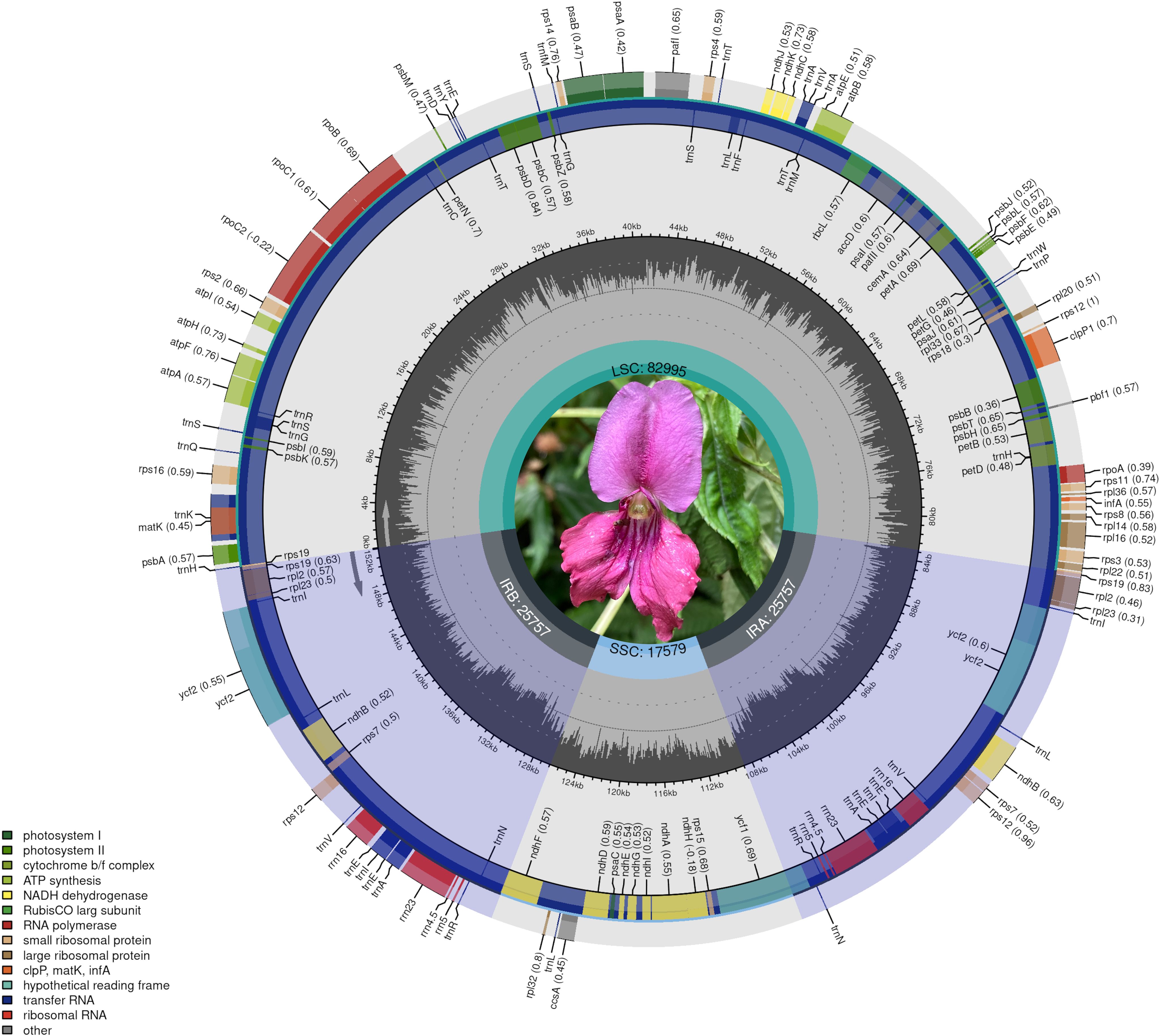
Figure 1. Gene map of I. lateristachys. The genes located outside the map are transcribed in a clockwise direction, while those inside are transcribed counterclockwise.
The chloroplast genomes of the 9 Impatiens species were divided into four major groups according on gene function. The first group included genes involved in self-replication, such as tRNA-coding genes, rRNA-coding genes, RNA polymerase subunit-coding genes, as well as ribosomal small subunit protein-coding genes and ribosomal large subunit protein-coding genes. ATP synthase, photosystem I and II, cytochrome b/f complex, NADH dehydrogenase, and ribulose-1, 5-bisphosphate carboxylase genes were among the genes in the second group that were involved in photosynthesis. The third group consisted of biosynthesis-related genes, including mature enzyme genes, envelope protein genes, and ATP-dependent protease genes. The fourth group included genes of unknown function, primarily presumed chloroplast open reading frames (Supplementary Table S2–S9).
For instance, the chloroplast genome of I. lateristachys (Table 2) contained 113 genes in total, including 79 genes that coded for proteins, 30 genes that coded for transfer RNA (tRNA), and 4 genes that coded for ribosomal RNA (rRNA) (Supplementary Table S10). The first group of self-replication-related genes comprised 25 genes, including 9 ribosomal large subunit protein-coding genes, 12 ribosomal small subunit protein-coding genes, and 4 RNA polymerase subunit-coding genes. The second group involved 43 genes related to photosynthesis, including 5 genes for photosystem I, 14 genes for photosystem II, 6 genes for ATP synthase, 6 genes for the cytochrome b/f complex, 11 genes for NADH dehydrogenase, and 1 gene for ribulose-1, 5-bisphosphate carboxylase. The third group of biosynthesis-related genes comprised 6 genes, including mature enzyme genes, envelope protein genes, and ATP-dependent protease genes. The fourth group contained 5 genes of unknown function, 3 of which were presumed to be chloroplast open reading frames.
SSR analysis was performed on 9 Impatiens species, identifying 834 SSR sequences. Of these, 709 were mononucleotide (85.0%), 44 were dinucleotides (5.3%), 32 were trinucleotides (3.8%), 48 were tetranucleotides (5.8%), and 1 was a hexanucleotide (0.1%), with no pentanucleotides were detected. Mononucleotide sequences were the most abundant, followed by tetranucleotides, while hexanucleotides, detected only in I. pritzelii, were the least common. I. menghuochengensis had the most, while I. lateristachys had the fewest, at just 74 (Supplementary Table S11). As shown in Figure 2, in the 9 species of Impatiens, SSR repeat sequences included A/T or A/C/T as mononucleotide repeats, AT/TA as dinucleotide repeats, with AT detected solely in I. menghuachengensis, AAT, GAA, TAT, TTA, and TAA for trinucleotide repeats, AATT, ATCT, TATT, TTCA, TTCT, ATA, TTTC, AAAT, AATA, TAAA, ATGA, ATTA, and TTAT for tetranucleotide repeats, and the hexanucleotide repeat TAAGTA found exclusively in I. pritzelii. The analysis revealed that polyA and polyT comprised the majority of simple repeat sequences, with few polyG or polyC sequences. This also explained the observed A/T base preference in cpDNA, and the findings are consistent with those from simple repeat sequences in the cpDNA of other species.
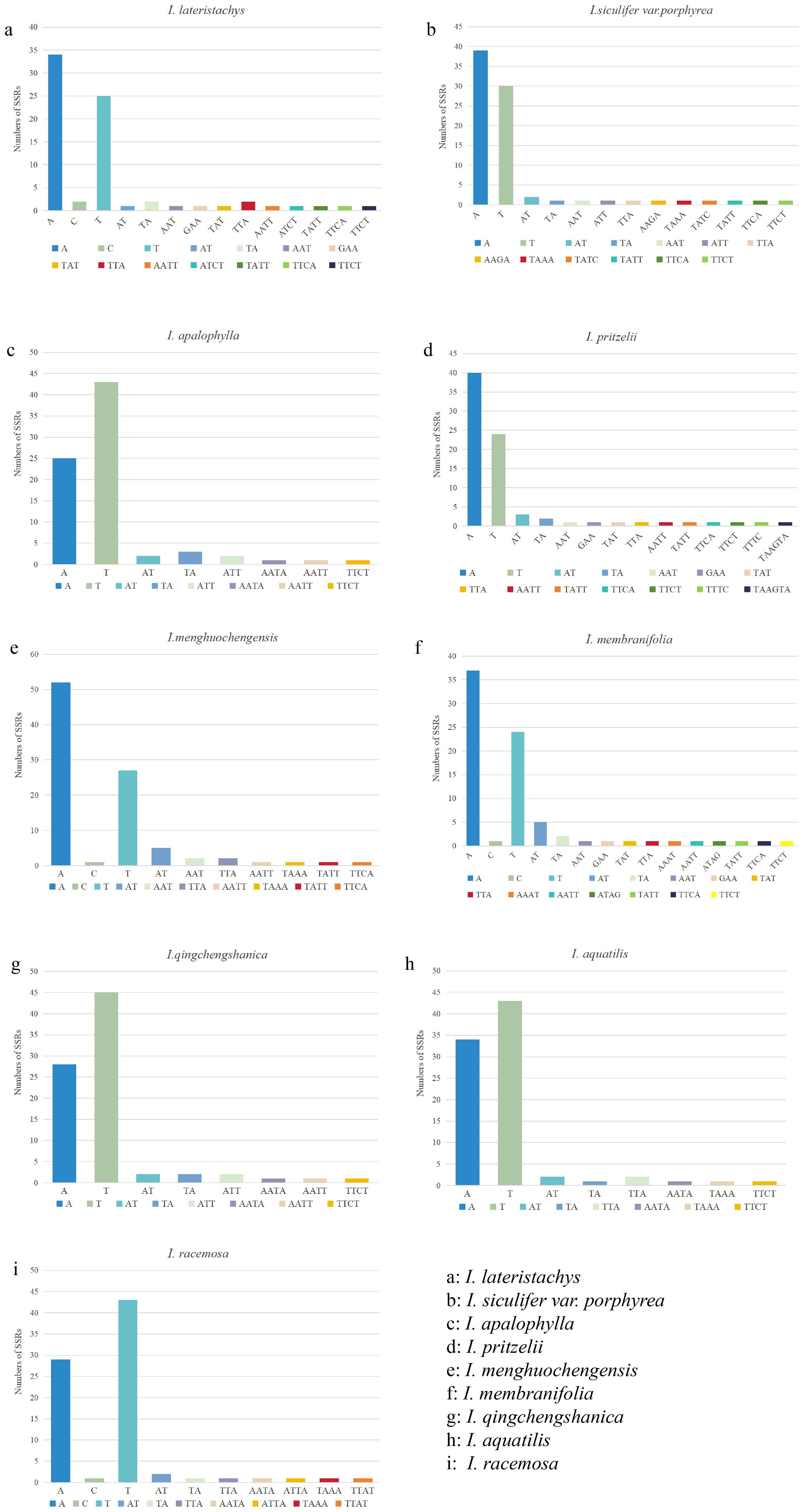
Figure 2. Number of SSR motifs identified in various class types. The Y-axis represents the number of SSRs, while the X-axis represents the types of SSRs. (a): I. lateristachys, (b): I. siculifer var. porphyrea, (c): I. apalophylla, (d): I. pritzelii, (e): I. menghuochengensis, (f): I. membranifolia, (g): I. qingchengshanica, (h): I. aquatilis, (i): I. racemosa.
This study statistically examined scattered repeat sequences in the chloroplast genomes of 9 Impatiens species (Figure 3). A total of 183 pairs of repeat sequences were identified, including 10 pairs of reverse repeats (R), 89 pairs of palindromic repeats (P), and 84 pairs of forward repeats (F). I. pritzelii possessed 23 repeats, including 11 forward and 12 palindromic repeats, making it the species with the most repeats. In contrast, the fewest repetitions were found in I. apalophylla. Approximately 56.59% of the total scattered repeats were palindromic repeats, which were most abundant in I. pritzelii and I. aquatilis samples, forward repeats constituted about 37.91%. Reverse repeats were detected in I. lateristachys, I. siculifer var. porphyrea, and I racemosa. No complementary repeat sequences were identified among the 9 Impatiens species.

Figure 3. Scattered repeats of chloroplast genomes in Impatiens and their numbers. F: forward repeats, P: palindromic repeats, R: reverse repeats.
This study analyzed codon usage bias in the chloroplast genomes of 9 Impatiens species, revealing the use of 64 codons. In protein-coding genes, the number of codons ranged from 50,594 in I. racemosa to 50,876 in I. apalophylla (Table 3). As in most angiosperms, the most prevalent codon among these 9 Impatiens species encoded leucine (Leu), while the codon UUU, which encodes phenylalanine (Phe), was the most frequent, ranging from 2,296 (I. apalophylla) to 2,433 (I. qingchengshanica). In contrast, the codons GCG (for alanine) and CGC (for arginine) were the least abundant, with counts ranging from 217 (I. racemosa) to 230 (I. qingchengshanica). The termination codon UAA was the most abundant (Table 3).
Among these 9 Impatiens species, 35-37 types of codons with an RSCU value ≥ 1.00 were identified, predominantly those ending with A/U, while those ending with C/G were relatively scarce. The arginine (Arg)-encoding codon AGA had the greatest RSCU score (Arg) (Figure 4). Additionally, the RSCU values of the chloroplast genomes predominantly ended with A/U, consistent with the characteristic low GC content of these genomes. The findings showed that the 9 Impatiens species shared a significant degree of similarity in codon usage and amino acid frequencies.

Figure 4. Cluster heatmap analysis of synonymous codons in the chloroplast genome of Impatiens. The RSCU values range from 0.3 to 2.4, corresponding to a color gradient from blue to red.
In this study, the chloroplast genomes of 9 Impatiens species were analyzed, focusing on the IR border regions. All genes and sequence lengths exhibited a tetrad structure and were highly conserved. However, differences were observed in the boundary regions, with contractions and expansions of the IR boundaries leading to variations in structure and size. The lengths of the IR regions in the 9 Impatiens species ranged from 25,535 bp (I. racemosa) to 25,883bp (I. aquatilis). The LSC region ranged from 80 to 118 bp, and the IR region overlap ranged from of 161 to 199bp. The rps19 gene was located at the intersection of the LSC and IR sections in most species. The entire rpl22 gene was located in the LSC region, near the IR boundary. In most Impatiens species, the ndhF gene was located at the boundary between the SSC and IR regions, with in half of the species, it was entirely within the SSC region. The ycf1 gene spanned both the SSC and IR regions in all 9 Impatiens species, with lengths ranging from 4, 165 to 4,545 bp in the SSC region and 313 to 1,325 bp in the IR region. Copies of the ycf1 gene were present in all species except I. qingchengshanica, and I. racemosa. The trnN gene was entirely located in the IR region in most species, while the trnH gene was located entirely in the LSC region. Gene contractions and expansions at the IR/SSC boundary varied among species (Figure 5).

Figure 5. Compares the boundaries of the large single-copy region (LSC), small single-copy region (SSC), and inverted repeat region (IR) in the chloroplast genomes of 9 species. The distances between the gene ends and the border points are shown by the numbers above the gene features. JLB (IRb/LSC), JSA (SSC/IRa), JSB (IRb/SSC), and JLA (IRa/LSC) denote the junction sites between the quadripartite regions of the genome.
To investigate the differentiation of chloroplast genome sequences in Hydrocera triflora and other Impatiens species, highly variable regions were identified using the mVISTA software. Sequence homology across the entire chloroplast genome was analyzed using Hydrocera triflora as the reference genome, with results shown in the sequence homology plots (Figure 6). The findings showed a high degree of similarity between the chloroplast genomes of the 10 species, with strong conservation, high collinearity, and significant homology, reflecting a high degree of similarity. However, certain differences were observed, with varying mutation rates among the IR, SSC, and LSC regions, where the IR region was found to be more conserved. Coding regions showed higher conservation compared to non-coding regions. However, high divergence was observed in both intergenic spacers and coding genes, such as matK, psbK, rps16, petN, trnC-GCA, rpoB, rps18, rpl33, ycf3, ndhE, ndhG, ycf1, and trnR-ACG. Among the intergenic regions. The largest variations among intergenic regions were observed in psbI-atpA, trnC-GCA-petN, ndhG-ndhI, and psbM-psbD.
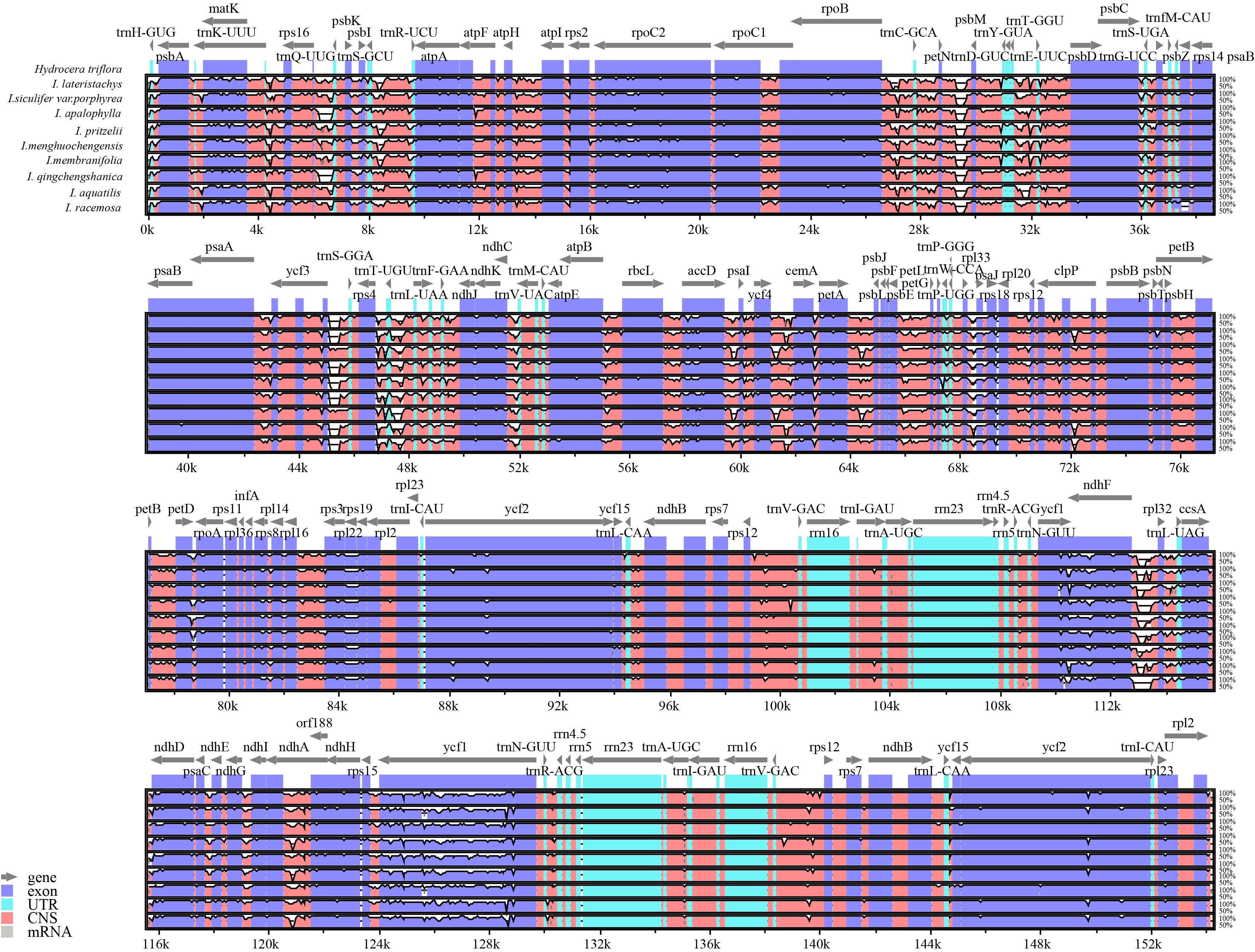
Figure 6. Compares the chloroplast genomes of 9 Impatiens species with that of Hydrocera triflora using the mVISTA method. The x-axis represents the positions within the chloroplast genome, and the y-axis represents the percentage of identity, ranging from 50% to 100%.
Various hotspots were utilized to identify closely related species, providing crucial evidence for species identification through comparisons of gene distribution with the results of sliding window analysis. Nucleotide polymorphism analysis showed that sequence divergence in the LSC and SSC single-copy regions was significantly higher than in the inverted repeat region. The analysis indicated that nucleotide values in intergenic regions were higher than those in coding regions, suggesting greater divergence intergenic regions (Figure 7). The average nucleotide diversity across the 9 Impatiens species was 0.021856. The nucleotide diversity was highest in rrn23 (0.09015) and ndhG (0.07973). Ten highly divergent hotspots were identified, including trnK-UUU, psbI-atpA, atpI, trnC-GCA-petN, and rps18 in the LSC region, and ndhE, ndhG, trnR-ACG, ycf1, and rrn23 in the SSC region. The IR region exhibited strong conservation, with no highly dispersed hotspots observed. Compared to the LSC region, the SSC region displayed more pronounced, highly dispersed hotspots, indicating greater variability. The hotspots identified by the mVISTA program were comparable to regions exhibiting the largest variations in Pi values. These distinct molecular markers could serve as valuable markers for phylogenetic analysis and species identification when combining results from DnaSP and mVISTA.
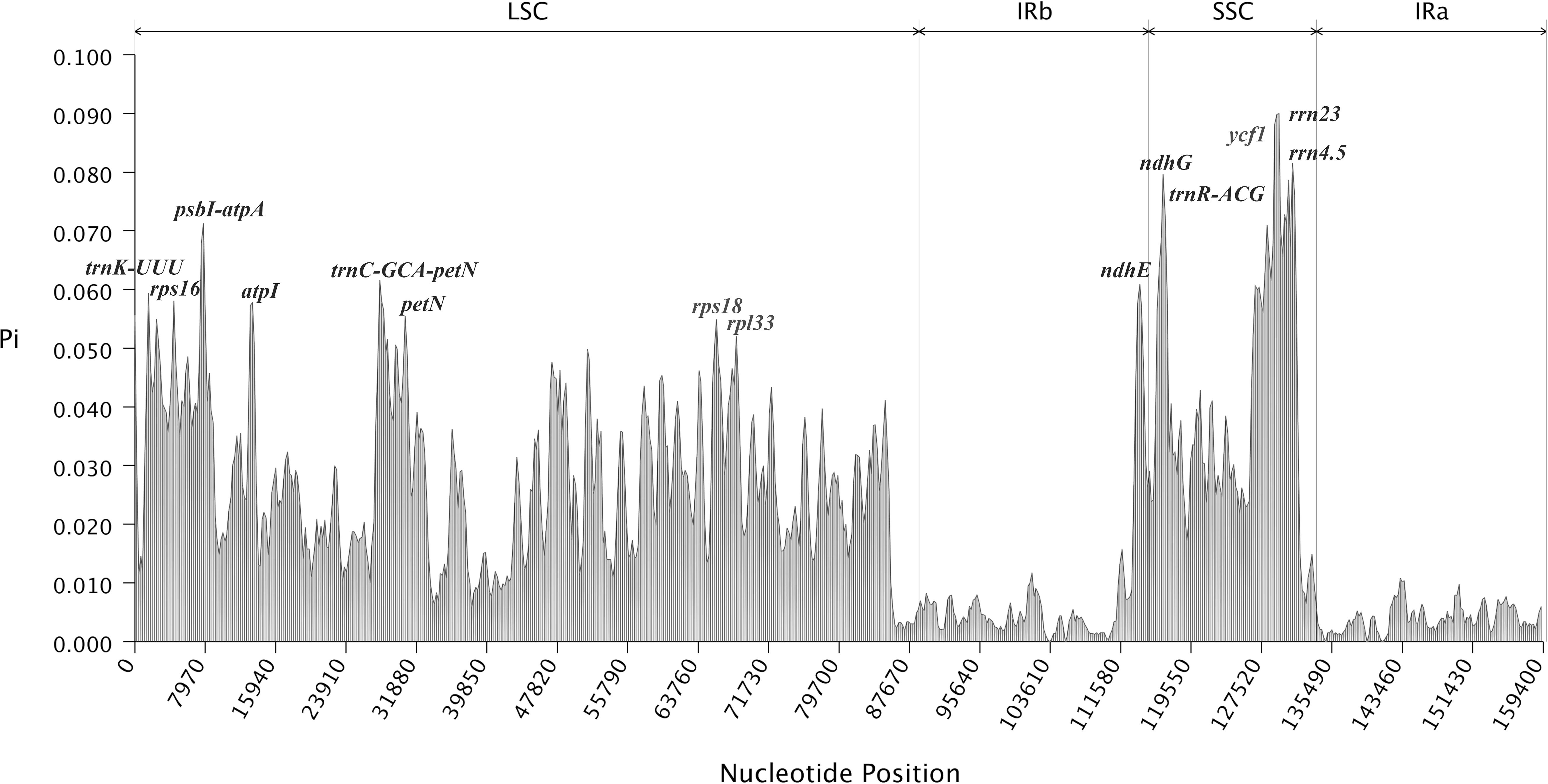
Figure 7. Mutational hotspot analysis. x-axis indicates regions of the chloroplast genome and y-axis indicates the nucleotide diversity of each region. Window size: 600 bp, step size: 200 bp.
To investigate the selective pressures acting on the chloroplast genome, we used Hydrocera triflora as a reference species and analyzed the selective and evolutionary differences among 9 species of Impatiens. We calculated the synonymous (Ks) and nonsynonymous (Ka) substitution rates, as well as the average Ka/Ks ratio (ω) for 74 protein-coding genes. We calculated the synonymous (Ks) and nonsynonymous (Ka) substitution rates, as well as the average Ka/Ks ratio (ω) for 74 protein-coding genes. Among these, 58 genes were selected for clustering heatmap analysis (Figure 8). Our results indicated that most genes were under purifying selection, with only a few genes showing evidence of positive selection. Notably, no genes exhibited neutral selection. Specifically, the psbK gene in I. lateristachys, I. pritzelii, I. menghuochengensis, and I. racemosa, the rps18 gene in I. siculifer var. porphyrea, I. menghuochengensis, I. aquatilis, and I. racemosa, and the rpl12 gene in I. pritzelii and I. racemosa all had Ka/Ks ratios greater than 1, suggesting that these genes were under positive selection.
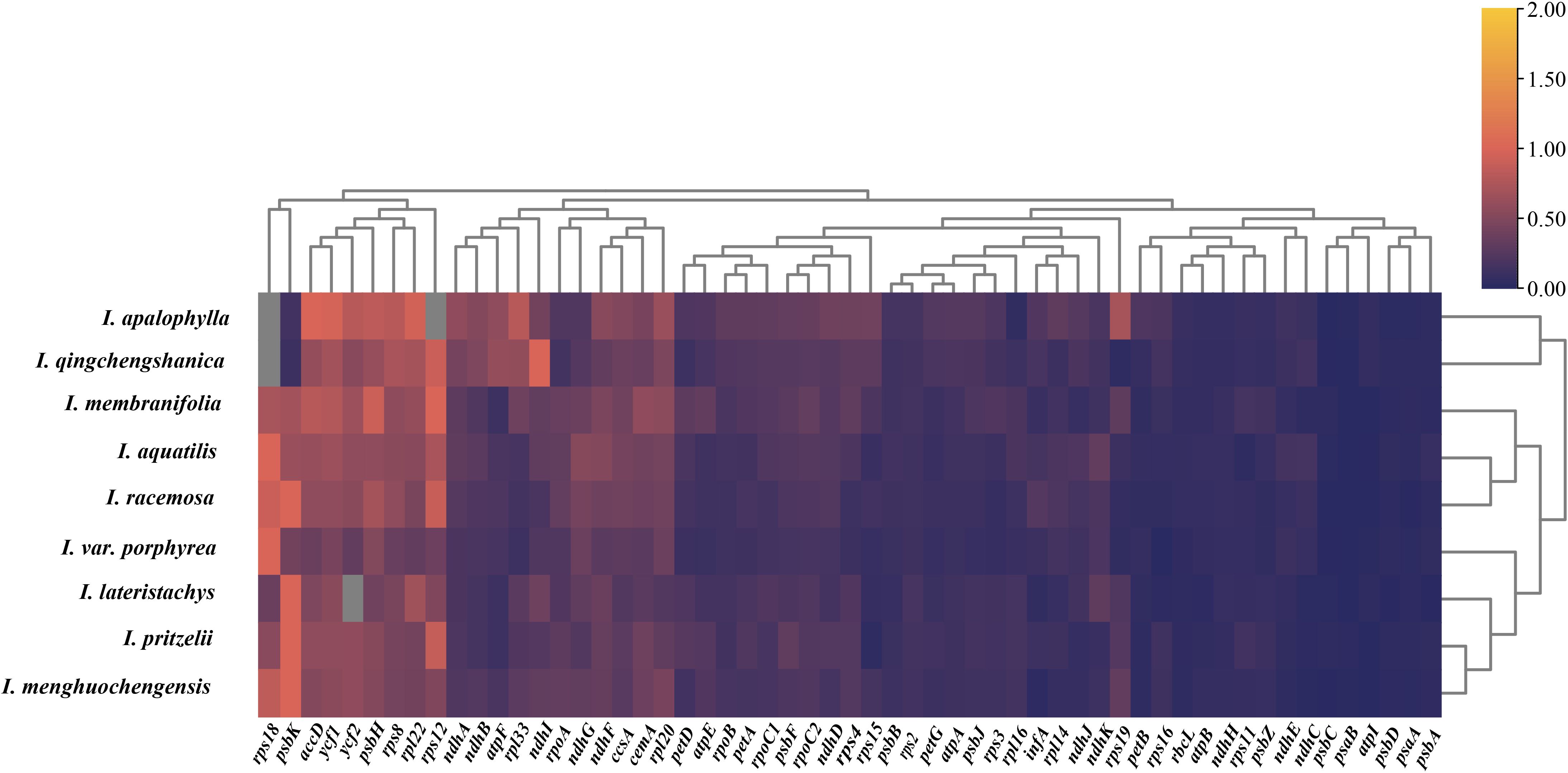
Figure 8. Selective pressure analysis results. A clustering heatmap of Ka/Ks values for the chloroplast genomes of 9 species with Hydrocera triflora as the reference species, the Ka/Ks values range from 0 to 2, corresponding to a color gradient from dark purple to orange.
This study was based on the complete chloroplast genome sequences of 24 plants from the Ericales order, including 21 Impatiens species, one Hydrocera species, and two outgroup species (one each from the Primulaceae and Actinidiaceae). A phylogenetic tree was constructed using maximum likelihood methods in IQtree, and the results were visualized and refined with FigTree, yielding a highly supported topology. These species have been extensively studied for their evolutionary relationships through morphology, palynology, and molecular markers, providing a robust foundation for our analysis. The selection of these species aims to illuminate the evolutionary history and taxonomic placement of Impatiens plants, validate previous findings, and elucidate their phylogenetic relationships through chloroplast genomic analysis.
The evolutionary tree consisted of two primary branches. One branch included plants from the Balsaminaceae family, comprising 20 species of Impatiens and Hydrocera triflora, while the other branch included species from other families (Figure 9). Further analysis revealed that the Balsaminaceae family was divided into two distinct branches: the genera Hydrocotyle and Impatiens, consistent with previous classification studies. The genus Impatiens was further divided into two subgenera: Clavicarpa and Impatiens. I. qingchengshanica and I. apalophylla, along with I. guizhouensis and I. omeiana, form a group classified as the subgenus Clavicarpa, while the remaining Impatiens species are classified under the subgenus Impatiens. In the subgenus Clavicarpa, species such as I. qingchengshanica and I. apalophylla possess ovaries with four carpels, each containing one ovule. Their capsules are hammer-shaped, and the pollen germination grooves are trichotomous, showing a triangular view from the apex. The subgenus Impatiens was divided into three sections: sect. Racemosae, sect. Uniflorae, and sect. Impatiens. I. aquatilis, I. racemosa, and I. siculifer var. porphyrea cluster with I. cyanantha and I. uliginosa, characterized by ovaries with five carpels, linear capsules, racemose and many-flowered inflorescences, two laterally situated sepals, and oval seeds, classifying them in the sect. Racemosae. I. chlorosepala was closely related to I. mengtszeana, and both were classified in the sect. Uniflorae, characterized by ovaries with five carpels, spindle-shaped capsules, clustered inflorescences, and oval seeds. I. menghuochengensis, I. lateristachys, and I. membranifolia cluster with other Impatiens species in the sect. Impatiens, characterized by ovaries with five carpels, linear capsules, few-flowered racemose inflorescences, and oval seeds.
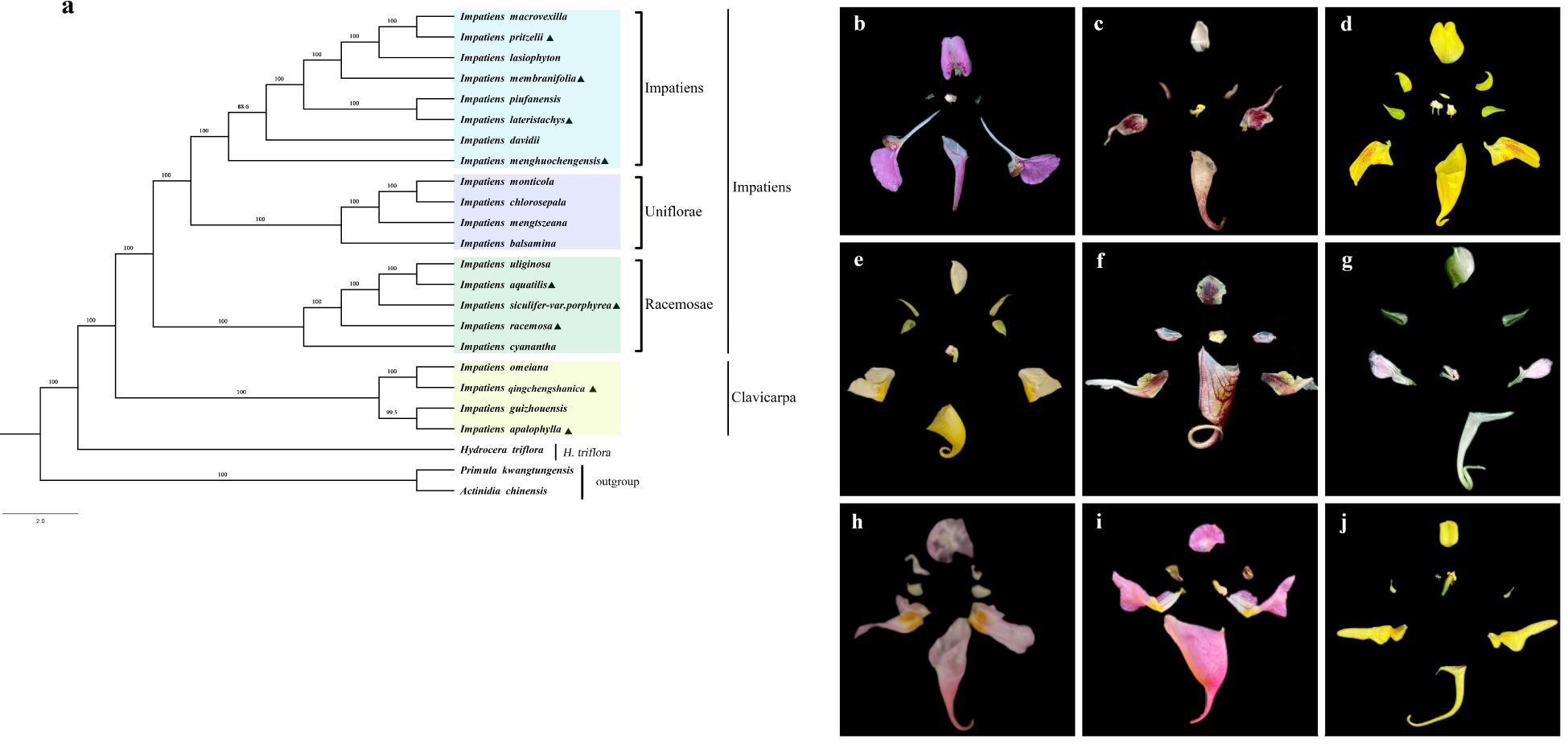
Figure 9. Phylogenetic tree was constructed based on the chloroplast genome sequences of 21 Impatins species and three other related species using the Maximum Likelihood (ML) method. (a):Triangles stand for the species that our study has recently sequenced. and the colored blocks correspond to the different groups indicated. (b-j): The Anatomical Structure of the Floral Parts of 9 Impatiens Species. (b): I. lateristachys, (c): I. siculifer var. porphyrea, (d): I. apalophylla, (e): I. pritzelii, (f): I. menghuochengensis, (g): I. membranifolia, (h): I. qingchengensis, (i): I. aquatilis, (j): I. racemosa.
The circular structure of the chloroplast genomes of the 9 Impatiens species studied consists of a quadripartite structure, including two Inverted Repeat (IR) regions, one Large Single Copy (LSC) region, and one Small Single Copy (SSC) region. This structure is similar to that of other species in the Ericales, such as Alniphyllum (Ebenaceae), Primula (Primulaceae), and Camellia (Theaceae) (Han et al., 2017). It is also consistent with the chloroplast genome structures of other previously published Impatiens species (Luo et al., 2021).
Chloroplast genomes in most photosynthetic species range from 115 to 165 kb in size (Jansen et al., 2005). The chloroplast genome sizes in the Impatiens species studied ranged from 151, 784 bp (I. racemosa) to 152,628 bp (I. apalophylla), falling within the size range of published angiosperm chloroplast genomes (Dong et al., 2021; Li et al., 2024; Liu et al., 2024). Among the 9 Impatiens species, the largest variation in chloroplast genome size was 844 bp. The LSC region ranged from 82, 995 bp (I. lateristachys) to 83, 480 bp (I. siculifer var.porphyrea), the SSC region ranges from 17, 253 bp (I. pritzelii) to 17, 893 bp (I. qingchengshanica), and the IR region ranged from 25, 535 bp (I. racemosa) to 25, 883 bp (I. aquatilis). Comparative analysis indicated that the interspecific genomes within the genus Impatiens were relatively conserved, with size differences primarily arising from variations in the LSC, contractions and expansions of the IR region, as well as gene insertions and deletions. The chloroplast genomes of plants can contain between 63 and 209 genes, but most are concentrated in the range of 110 to 130 genes (Jansen et al., 2005). With gene functions and GC content in line with earlier research findings (Jansen et al., 2005), the chloroplast genomes of 9 Impatiens species were found to include between 108 and 117 genes (including 77 to 81 protein-coding genes, 28 to 32 tRNA genes, and 4 rRNA genes).
SSRs, also known as microsatellite sequences in chloroplast genomes, are repetitive units consisting of one to six nucleotides, commonly found in eukaryotic cells. They are widely used in phylogenetic analyses, species identification, and genetic diversity assessments (Zhao et al., 2022). Genomic analyses of various chloroplasts have shown that indels and substitutions are likely induced by repetitive sequences (Gu et al., 2018). These sequences influence interspecies variations in copy numbers and play a crucial role in the stability and rearrangement of chloroplast genome sequences (Park et al., 2018). A total of 832 SSR sequences were identified in the cpDNA of 9 Impatiens species, including mononucleotides, dinucleotides, trinucleotides, and tetranucleotides. Pentanucleotides were not detected, while hexanucleotides (TAAGTA) were observed only in I. pritzelii. Previous studies on SSR motifs in eight Ericaceae species revealed hexanucleotide repeats in Ardisia polysticta, a member of the Primulaceae family (Li et al., 2018). This finding aligns with our results, which similarly demonstrate that not all SSR types were identified in every species. Previous studies on SSR motifs in eight Ericaceae species revealed hexanucleotide repeats in Ardisia polysticta, a member of the Primulaceae family (Li et al., 2018). This finding aligns with our results, which similarly demonstrate that not all SSR types were identified in every species. Mononucleotide repeats predominated, leading to a significantly lower GC content in the cpDNA of this genus compared to AT content. This pattern is consistent with findings in most plants, where A/T base repeats are the most abundant, as seen in other species (Li et al., 2022). Numerous cpSSR fragments were found in the chloroplast genomes of Impatiens, mainly composed of A/T mononucleotide repeat sequences, as well as polyadenine (polyA) and polythymidine (polyT). This high abundance of A/T bases is typical in plant chloroplast genomes (Ebert and Peakall, 2009). The development of SSR molecular markers provides offers valuable tools for further elucidating the phylogenetic and evolutionary relationships of Impatiens species.
Codon bias refers to the unequal usage of synonymous codons to encode the same amino acid across different organisms. This phenomenon has evolved over time due to various factors, including gene function, environmental selection, and gene expression levels. Species that are closely related exhibit similar codon usage patterns, offering valuable insights into interspecies evolution, exogenous gene expression, and genetic diversity (Wang et al., 2017). In this study, the 9 Impatiens species encoded between 50,594 (I. racemosa) and 50,876 (I. apalophylla) codons, with 35 to 37 codons showing an RSCU value of ≥ 1.00, mostly ending with A/U and a smaller number with C/G. This pattern aligns with the codon usage analyses in the chloroplast genomes of other higher plants (Gao et al., 2023). The GC content at the third codon position (GC3) plays a crucial role in influencing codon usage bias. Genomes with high GC content tend to favor codons enriched in G and C, while those with low GC content preferentially use codons rich in A and U (Jia et al., 2025). In this study, the average GC content of the chloroplast genome was approximately 37%, classifying it as low-GC. Therefore, codon usage in these chloroplast genomes predominantly favors codons with higher A and U content. This finding supports observations from previous studies. The genomes of the species studied exhibited a higher frequency of codons encoding leucine, consistent with previous reports (Yang et al., 2018; Guo et al., 2018; Wang et al., 2017). This study identified multiple codons encoding alanine (Ala), including GCA, GCT, GCC, and GCG, consistent with previous research (Sharp and Li, 1986). Previous studies have shown that codon usage bias is closely linked to factors such as mRNA stability and tRNA recognition efficiency (Hanson and Coller, 2018). In this study, we found that the frequently used codon AGA corresponds to tRNAs with higher abundance. This correspondence may affect mRNA stability by modulating tRNA recruitment and ribosomal translation. However, the precise molecular mechanisms and regulatory pathways underlying these relationships need further investigation for full elucidation. Given the highest degree of codon usage similarity among the 9 species, it is possible that these species encountered comparable environmental stresses in their biological niches.
The IR region contracts and expands at the boundaries of the LSC and SSC regions. These processes are key drivers of gene length variation. This occurs despite the generally high conservation of chloroplast genomes in terrestrial angiosperms (Zhu et al., 2016). Within chloroplast genomes, the IR region is generally regarded as the most conserved. However, variations in genome size, often caused by expansions and contractions in different plant lineages, make these regions useful for studying plant phylogenetic classifications (Wang et al., 2008). Events of contraction and expansion at the four boundary regions enhance our understanding of genome evolution and taxonomic hierarchies at or above the genus level. Aligning coding and non-coding sequences also aids in identifying mutation sites, offering essential data for studying interspecific and intrageneric phylogenetics and species evolution (Tang, 2017).
In this study of Impatiens species, genes at the IR/SC boundary underwent contraction and expansion, with the rps19 gene at the junction of the LSC and IR regions showing duplication. Most other chloroplast genomes also exhibited this characteristic (Su et al., 2014; Lee et al., 2015). Previous comparative analyses of the inverted repeat (IR) regions among Ericaceae, Balsaminaceae, and other families have reported the duplication of the rps19 gene only in Balsaminaceae species. Consistent with these findings, our study also identified the duplication of the rps19 gene. The ndhF gene was located at the intersection of the SSC and IR sections, while the rpl22 gene was entirely within the LSC region, close to the IR boundary. Previous studies have highlighted the crucial role of the ycf1 gene in plant viability. In this study, the ycf1 gene extended into the SSC region, varying in length across genomes, and spanning both the SSC and IR regions (Dong et al., 2015). Differences in the length and distribution of the ycf1 gene, along with positional variations of the rps19 and ndhF genes, contributed to length discrepancies within the Impatiens genus. Due to IR shrinkage, angiosperm chloroplast genomes often show variability in the number of duplicated genes, with duplication differences across species (Menezes et al., 2018). Copies of some pseudogenes, such as the ycf1 gene, were retained at the boundaries, as seen in the Withania somnifera (Mehmood et al., 2020). These variations are crucial for understanding the evolution of chloroplast genomes and genomic structures (Kim K. et al., 2017). These analyses thus enhance our understanding of the genetic structure and evolutionary dynamics of Impatiens species.
Using Hydrocera triflora as a reference, mVISTA analysis revealed high sequence collinearity, low variation, and high segment sequence similarity. Notably, the LSC and SSC regions exhibited considerably more variance than the IR region, with non-coding regions showing significantly higher variation than coding regions. This may be related to selective pressures, where lower selective pressure led to structural variation, while higher selective pressure resulted in greater structural stability (Huang et al., 2024c; Zhang, 2022). Therefore, primers for species identification can be created based on polymorphic regions near these boundaries.
Nucleotide polymorphisms analysis is an indicator of the degree of polymorphism and variation in nucleotide sequences between species. Regions with high variability may serve as molecular markers for population genetics studies. The chloroplast genome contains a number of mutation hotspots that have been verified as possible molecular markers, laying the foundation for studying phylogeny and relationships among plants (Dong et al., 2021). Luo identified mutational hotspots in six Impatiens species, including rps4-ndhJ, rpl32-ccsA, trnK-UUU-rps16, rpoB-petN, trnG-GCC, atpH-atpL, accD-psaI, ndhF, and ycf1 as potential molecular markers (Luo et al., 2021). In this study, 10 high-mutation hotspots were identified in the newly sequenced species: trnK-UUU, psbI-atpA, atpI, trnC-GCA-petN, rps18, ndhE, ndhG, trnR-ACG, ycf1, and rrn23, all sharing similar mutation characteristics. This study also found that the IR region was more conserved than the LSC and SSC regions, consistent with findings in other angiosperms (Lu et al., 2017; Wu et al., 2021). Divergence hotspots in the chloroplast genome have been widely used to determine the species of closely related plants (Dong et al., 2021; Bi et al., 2018). Therefore, we proposed that these ten highly variable regions could serve as DNA barcodes for Impatiens and be used in intra-species phylogeographic studies. External factors that influence a species’ evolutionary process and promote environmental adaption are known as selective pressure. When ω>1, advantageous mutations are selected, when ω =1, the species undergoes neutral selection, and when 0<ω<1, purifying selection occurs. A smaller ω indicates stronger negative selection pressure and greater conservation of the amino acid sequence (Fiz-Palacios et al., 2011). Analysis of selective pressure across 9 species revealed that most were under purifying selection, indicating a high level of conservation. Purifying selection was observed in most plants, suggesting a highly preserved evolutionary history (Huang et al., 2021). Purifying selection helped to prevent mutations (Wu et al., 2020), eliminating deleterious mutations and preserving conserved gene functions (Huang et al., 2021). In Impatiens, purifying selection may explain the high level of interspecific conservatism observed in the chloroplast genome. The adaptive evolution identified in this study may explain the observed diversity within the genus Impatiens, including habitat variation and morphological differences. Future research could further elucidate the environmental or functional drivers underlying the positive selection of these chloroplast genes, deepening our understanding of the adaptation and evolution of Impatiens species.
Chloroplast genomes are maternally inherited and exhibit low rates of base substitution and structural rearrangements, making them valuable tools for studying phylogenetic relationships (Fang et al., 2021). The genus Impatiens includes a diverse range of species with complex and varied morphological characteristics, which pose significant challenges for phylogenetic analysis and identification. In this study, chloroplast genome sequences of 24 species were analyzed using maximum likelihood methods for phylogenetic analysis. The results indicated that the genus Impatiens was mainly divided into two evolutionary branches, consistent with prior studies (Janssens et al., 2012; Li et al., 2018). Based on the phylogenetic analysis of chloroplast genome sequences, this study supported the classification of the genus Impatiens into two subgenera (Clavicarpa and Impatiens) by Yu et al. (2016). The newly sequenced 9 species were categorized into different subgenera and further divided into three sections: sect. Impatiens, sect. Racemosae, and sect. Uniflorae. The study found that I. qingchengshanica and I. apalophylla were closely related to I. omeiana and I. guizhouensis, representing the basal lineage of Impatiens. A notable characteristic was the presence of three pollen apertures, which Lu considered to be a primitive pollen type (Lu, 1991), as this pollen type was found in the genus Hydrocera and subgenera Clavicarpa, aligning with the classifications by Zhang et al. (2023) and Xia (2020). I. siculifer var. porphyrea was considered as a variety of I. siculifer, sharing similar morphological characteristics. This study placed it in the sect. Racemosae, consistent with Yu et al. (2016) classifications based on morphology and molecular data. The sect. Impatiens included previously unpublished species I. menghuochengensis and I. membranifolia, as well as classified species,I. pritzelii, and I. lateristachys. Both I. menghuochengensis and I. membranifolia exhibit four pollen apertures and share morphological traits with the sect. Impatiens, a classification supported by chloroplast genomic data. Morphologically, I. chlorosepala resembled I. mengtszeana, and genomic analyses also revealed a close phylogenetic relationship between the two. In 2012, I. lateristachys was classified into the sect. Fusicarpa based on morphological taxonomy (Yu, 2012). However, it was reclassified into the sect. Impatiens in a 2016 revision (Yu et al., 2016), a classification supported by our results. Traditional morphological classification methods are often constrained by the similarity or diversity of morphological traits, which can obscure the accurate depiction of species’ taxonomic status. In contrast, this study leverages the sequence conservation of chloroplast genomes to provide robust molecular markers for phylogenetic analysis, thereby more precisely elucidating the taxonomic position of I. lateristachys. In this study, I. pritzelii was closely related to I. macrovexilla and I. membranifolia, with a high support value (BS=100). According to Yu et al. (2016), I. pritzelii was classified in the Clavicarpa subgenus, which was inconsistent with our results. We speculated that the chloroplast genome length of I. pritzelii was similar to that of I. membranifolia, with only minor differences in IR expansion and contraction, further supporting their phylogenetic relationship from a genomic perspective. This indicated that the classification results are integrative, combining both macroscopic morphological features and molecular data. Using the entire chloroplast genome sequences, our classification aligns with Yu et al. (2016) classification, demonstrating a high congruence between genomic and morphological classifications in phylogenetic relationships. This study employed complete chloroplast genome sequences for classification, which showed a high consistency with Yu et al. (2016) classification. This congruence highlights the strong alignment between genomic and morphological classifications in elucidating phylogenetic relationships.
The chloroplast genomes used in this study provided extensive genetic information and high-resolution evolutionary signals, facilitating precise phylogenetic analysis of Impatiens species. Their maternal inheritance and structural stability make them ideal for clarifying kinship among Impatiens species, while their moderate evolutionary rate effectively reveals evolutionary events between these species. Chloroplast genome analysis identified positively selected genes such as psbK, rps18, and rpl12, potentially linked to the adaptive evolution of Impatiens species in diverse environments. Additionally, the conservation and diversity of chloroplast genomes offer essential criteria for identifying and conserving Impatiens species, further advancing taxonomic and ecological studies in this genus.
This study examines the chloroplast genomes of 9 Impatiens species, revealing high similarity in structure, size, GC content, gene number, and function, highlighting the conserved nature of these genomes. We identified expansions and contractions within the inverted repeat (IR) regions and found ten divergent areas that could serve as markers for phylogenetic classification and species identification. Most species showed purifying selection, leading to highly conserved genomic sequences. The phylogenetic tree, constructed using the maximum likelihood method, supports the classification and evolutionary relationships of Impatiens species. These genomic data provide a solid foundation for understanding the evolutionary dynamics within the Impatiens genus. Future research should integrate morphological traits, molecular studies, and genomic analyses to address classification and evolutionary questions in Impatiens.
New sequenced and other published chloroplast genome 716 sequences can be found in GenBank (https://www.ncbi.nlm.nih.gov/genbank/) with the accession numbers present in Supplementary Table S1.
MY: Data curation, Writing – original draft, Formal analysis. WL: Data curation, Writing – original draft, Formal analysis. JZ: Data curation, Writing – original draft, Formal analysis. HM: Writing – original draft, Software. XH: Writing – original draft, Software. MH: Writing – review & editing. HH: Conceptualization, Writing – review & editing.
The author(s) declare that financial support was received for the research and/or publication of this article. This study was supported by the National Natural Science Foundation of China (grant number 32060364, 32060366), the Project of High-level Introduction talents in Yunnan Province, and First-rate Discipline Landscape Architecture Construction Project of Yunnan Province, China.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fpls.2025.1541320/full#supplementary-material
Bi, Y., Zhang, M., Xue, J., Dong, R., Du, Y., Zhang, X. (2018). Chloroplast genomic resources for phylogeny and DNA barcoding: a case study on Fritillaria. Sci. Rep. 8, 1184. doi: 10.1038/s41598-018-19591-9
Cai, X. Z., Yi, R. Y., Zhuang, Y. H., Cong, Y. Y., Kuang, R. P., Liu, K. M. (2013). Seed coat micromorphology characteristics of Impatiens L. and its systematic significance. Acta Hortic. Sin. 40, 1337. doi: 10.16420/j.issn.0513-353x.2013.07.014
Chen, Y. (2001). “Balsaminaceae,” in Flora Reipublicae Popularis Sinica, vol. 47. (Science Press, Beijing), 1–243.
Chen, C., Wu, Y., Li, J., Wang, X., Zeng, Z., Xu, J., et al. (2023). TBtools-II: A “one for all, all for one” bioinformatics platform for biological big-data mining. Mol. Plant 16, 1733–1742. doi: 10.1016/j.molp.2023.09.010
Dong, W., Xu, C., Li, C., Sun, J., Zuo, Y., Shi, S., et al. (2015). ycf1, the most promising plastid DNA barcode of land plants. Sci. Rep. 5, 1–5. doi: 10.1038/srep08348
Dong, S., Ying, Z., Yu, S., Wang, Q., Liao, G., Ge, Y., et al. (2021). Complete chloroplast genome of Stephania tetrandra (Menispermaceae) from Zhejiang Province: Insights into molecular structures, comparative genome analysis, mutational hotspots and phylogenetic relationships. BMC Genomics 22, 1–20. doi: 10.1186/s12864-021-08193-x
Doyle, J. J., Doyle, J. L. (1990). A rapid DNA isolation procedure for small quantities of leaf tissue. Focus 12, 13–15.
Ebert, D., Peakall, R. O. D. (2009). Chloroplast simple sequence repeats (cpSSRs): technical resources and recommendations for expanding cpSSR discovery and applications to a wide array of plant species. Mol. Ecol. Resour. 9, 673–690. doi: 10.1111/j.1755-0998.2008.02319.x
Fang, Y., Hu, S. S., Zhang, D. Q., Lv, J., Li, R., Qing, R. W., et al. (2021). Assembly and characteristic analysis of Chara globularis chloroplast whole genome. J. Sichuan. Univ. (Natural. Sci. Edition). 58, 046004. doi: 10.19907/j.0490-6756.2021.046004
Fiz-Palacios, O., Schneider, H., Heinrichs, J., Savolainen, V. (2011). Diversification of land plants: insights from a family-level phylogenetic analysis. BMC Evol. Biol. 11, 1–10. doi: 10.1186/1471-2148-11-341
Frazer, K. A., Pachter, L., Poliakov, A., Rubin, E. M., Dubchak, I. (2004). VISTA: computational tools for comparative genomics. Nucleic Acids Res. 32, W273–W279. doi: 10.1093/nar/gkh458
Fujihashi, H., Akiyama, S., Ohba, H. (2002). Origin and relationships of the Sino-Himalayan impatiens (Balsaminaceae) based on molecular phylogenetic analysis, chromosome numbers and gross morphology. J. Jap. Bot. 77, 284–295.
Gao, M. (2012). Morphological and molecular biological studies on native Impatiens species in China (Beijing: Beijing Forestry University).
Gao, S.-Y., Li, Y.-Y., Yang, Z.-Q., Dong, K.-H., Xia, F.-S. (2023). Codon usage bias analysis of the chloroplast genome of Bothriochloa ischaemum. Acta Pratacult. Sin. 32, 85. doi: 10.11686/cyxb2022332
Grey-Wilson, C. A. (1989). revision of sumatran impatiens: studies in balsaminaceae: VIII. Kew Bull., 67–106. doi: 10.2307/4114646
Gu, C., Tembrock, L. R., Zheng, S., Wu, Z. (2018). The complete chloroplast genome of Catha edulis: a comparative analysis of genome features with related species. Int. J. Mol. Sci. 19, 525. doi: 10.3390/ijms19020525
Guo, H. (2017). Systematic study of the racemose inflorescence group in Impatiens (Shanxi: Shanxi Normal University).
Guo, S., Guo, L., Zhao, W., Xu, J., Li, Y., Zhang, X., et al. (2018). Complete chloroplast genome sequence and phylogenetic analysis of Paeonia ostii. Molecules 23, 246. doi: 10.3390/molecules23020246
Han, Y., Huang, K., Liu, Y., Jiao, T., Ma, G., Qian, Y., et al. (2017). Functional analysis of two flavanone-3-hydroxylase genes from Camellia sinensis: a critical role in flavonoid accumulation. Genes 8, 300. doi: 10.3390/genes8110300
Hanson, G., Coller, J. (2018). Codon optimality, bias and usage in translation and mRNA decay. Nat. Rev. Mol. Cell Biol. 19, 20–30. doi: 10.1038/nrm.2017.91
Huang, Y., Jin, X. J., Zhang, C. Y., Li, P., Meng, H. H., Zhang, Y. H. (2024c). Plastome evolution of Engelhardia facilitates phylogeny of Juglandaceae. BMC Plant Biol. 24, 634. doi: 10.1186/s12870-024-05293-0
Huang, K., Li, B., Chen, X., Qin, C., Zhang, X. (2024a). Comparative and phylogenetic analysis of chloroplast genomes from ten species in Quercus section Cyclobalanopsis. Front. Plant Sci. 15, 1430191. doi: 10.3389/fpls.2024.1430191
Huang, R., Xie, X., Chen, A., Li, F., Tian, E., Chao, Z. (2021). The chloroplast genomes of four Bupleurum (Apiaceae) species endemic to Southwestern China, a diversity center of the genus, as well as their evolutionary implications and phylogenetic inferences. BMC Genomics 22, 1–15. doi: 10.1186/s12864-021-08008-z
Huang, L., Yu, H., Wang, Z., Xu, W. (2024b). CPStools: A package for analyzing chloroplast genome sequences. iMetaOmics 1, e25. doi: 10.1002/imo2.v1.2
Jansen, R. K., Raubeson, L. A., Boore, J. L., dePamphilis, C. W., Chumley, T. W., Haberle, R. C., et al. (2005). Methods for obtaining and analyzing whole chloroplast genome sequences. Methods Enzymol. 395, 348–384. doi: 10.1016/S0076-6879(05)95020-9
Janssens, S., Geuten, K., Yuan, Y.-M., Song, Y., Küpfer, P., Smets, E. (2006). Phylogenetics of Impatiens and Hydrocera (Balsaminaceae) using chloroplast atpB-rbcL spacer sequences. Syst. Bot. 31, 171–180. doi: 10.1600/036364406775971796
Janssens, S. B., Wilson, Y. S., Yuan, Y.-M., Nagels, A., Smets, E. F., Huysmans, S. (2012). A total evidence approach using palynological characters to infer the complex evolutionary history of the Asian Impatiens (Balsaminaceae). Taxon 61, 355–367. doi: 10.1002/tax.2012.61.issue-2
Janssens, S. B., Groeninckx, I., De Block, P. J., Verstraete, B., Smets, E. F., Dessein, S. (2016). Biogeography and evolution of the Madagascan endemics of the Spermacoceae tribe (Rubiaceae). Mol. Phylog. Evol. 95, 58–66. doi: 10.1016/j.ympev.2015.10.024
Jia, X., Wei, J., Chen, Y., Zeng, C., Deng, C., Zeng, P., et al. (2025). Codon usage patterns and genomic variation analysis of chloroplast genomes provides new insights into the evolution of Aroideae. Sci. Rep. 15, 4333. doi: 10.1038/s41598-025-88244-5
Jiang, H. F., Zhuang, Z. H., Hou, B. W., Shi, B. J., Shu, C. J., Chen, L., et al. (2017). Adverse effects of hydroalcoholic extracts and the major components in the stems of Impatiens balsamina L. @ on Caenorhabditis elegans. Evid. Based. Complement. Altern. Med. 2017, 4245830. doi: 10.1155/2017/4245830
Jin, J.-J., Yu, W.-B., Yang, J.-B., Song, Y., dePamphilis, C. W., Yi, T.-S., et al. (2020). GetOrganelle: a fast and versatile toolkit for accurate de novo assembly of organelle genomes. Genome Biol. 21, 1–31. doi: 10.1186/s13059-020-02154-5
Katoh, K., Standley, D. M. (2013). MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780. doi: 10.1093/molbev/mst010
Kim, C. S., Bae, M., Oh, J., Subedi, L., Suh, W. S., Choi, S. Z., et al. (2017). Anti-neurodegenerative biflavonoid glycosides from Impatiens balsamina. J. Nat. Prod. 80, 471–478. doi: 10.1021/acs.jnatprod.6b00981
Kim, K., Nguyen, V. B., Dong, J., Wang, Y., Park, J. Y., Lee, S.-C., et al. (2017). Evolution of the Araliaceae family inferred from complete chloroplast genomes and 45S nrDNAs of 10 Panax-related species. Sci. Rep. 7, 4917. doi: 10.1038/s41598-017-05218-y
Kumar, S., Stecher, G., Mega, K. T. (2016). Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 33, 1870–1874. doi: 10.1093/molbev/msw054
Kurtz, S., Choudhuri, J. V., Ohlebusch, E., Schleiermacher, C., Stoye, J., REPuter, R. G. (2001). The manifold applications of repeat analysis on a genomic scale. Nucleic Acids Res. 29, 4633–4642. doi: 10.1093/nar/29.22.4633
Langmead, B., Trapnell, C., Pop, M., Salzberg, S. L. (2009). Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, 1–10. doi: 10.1186/gb-2009-10-3-r25
Lee, M., Park, J., Lee, H., Sohn, S.-H., Lee, J. (2015). Complete chloroplast genomic sequence of Citrus platymamma determined by combined analysis of Sanger and NGS data. Horticult. Environ. Biotechnol. 56, 704–711. doi: 10.1007/s13580-015-0061-x
Li, B., Li, Y., Li, Z., Xiang, M., Dong, X., Tang, X. (2022). The complete chloroplast genome of Impatiens mengtszeana (Balsaminaceae), an endemic species in China. Mitochondrial. DNA Part B. 7, 367–369. doi: 10.1080/23802359.2021.1994894
Li, Z.-Z., Saina, J. K., Gichira, A. W., Kyalo, C. M., Wang, Q.-F., Chen, J.-M. (2018). Comparative genomics of the balsaminaceae sister genera Hydrocera triflora and Impatiens pinfanensis. Int. J. Mol. Sci. 19, 319. doi: 10.3390/ijms19010319
Li, Q.-Q., Zhang, Z.-P., Aogan, J., Wen, J. (2024). Comparative chloroplast genomes of Argentina species: genome evolution and phylogenomic implications. Front. Plant Sci. 15, 1349358. doi: 10.3389/fpls.2024.1349358
Liu, J., Zang, E., Tian, Y., Zhang, L., Li, Y., Shi, L., et al. (2024). Comparative chloroplast genomes: insights into the identification and phylogeny of rapid radiation genus Rhodiola. Front. Plant Sci. 15, 1404447. doi: 10.3389/fpls.2024.1404447
Lu, Y.-Q. (1991). Pollen morphology of Impatiens L.(Balsaminaceae) and its taxonomic implications. J. Systematics Evol. 29, 352.
Lu, R. S., Li, P., Qiu, Y. X. (2017). The complete chloroplast genomes of three Cardiocrinum (Liliaceae) species: comparative genomic and phylogenetic analyses. Front. Plant Sci. 7, 2054. doi: 10.3389/fpls.2016.02054
Luo, C., Huang, W., Sun, H., Yer, H., Li, X., Li, Y., et al. (2021). Comparative chloroplast genome analysis of Impatiens species (Balsaminaceae) in the karst area of China: insights into genome evolution and phylogenomic implications. BMC Genomics 22, 1–18. doi: 10.1186/s12864-021-07807-8
Mehmood, F., Abdullah, Shahzadi, I., Ahmed, I., Waheed, M. T., Mirza, B. (2020). Characterization of Withania somnifera chloroplast genome and its comparison with other selected species of Solanaceae. Genomics 112, 1522–1530. doi: 10.1016/j.ygeno.2019.08.024
Menezes, A. P. A., Resende-Moreira, L. C., Buzatti, R. S. O., Nazareno, A. G., Carlsen, M., Lobo, F. P., et al. (2018). Chloroplast genomes of Byrsonima species (Malpighiaceae): comparative analysis and screening of high divergence sequences[J. Sci. Rep. 8, 2210. doi: 10.1038/s41598-018-20189-4
Nguyen, L. T., Schmidt, H. A., von Haeseler, A., Minh, B. Q. (2015). IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274. doi: 10.1093/molbev/msu300
Park, M., Park, H., Lee, H., Lee, B.-h., Lee, J. (2018). The complete plastome sequence of an Antarctic bryophyte Sanionia uncinata (Hedw.) Loeske. Int. J. Mol. Sci. 19, 709. doi: 10.3390/ijms19030709
Pechimuthu, M., Erayil, A. R., Thangavelu, M. (2024). Taxonomical implications of foliar epidermal anatomy of Impatiens L. species (Balsaminaceae) in the Nilgiris, Southern Western Ghats, India. Flora 318, 152573. doi: 10.1016/j.flora.2024.152573
Rahelivololona, E. M., Fischer, E., Janssens, S. B., Razafimandimbison, S. G. (2018). Phylogeny, infrageneric classification and species delimitation in the Malagasy Impatiens (Balsaminaceae). PhytoKeys . 110, 51. doi: 10.3897/phytokeys.110.28216
Rozas, J., Ferrer-Mata, A., Sánchez-DelBarrio, J. C., Guirao-Rico, S., Librado, P., Ramos-Onsins, S. E., et al. (2017). DnaSP 6: DNA sequence polymorphism analysis of large data sets. Mol. Biol. Evol. 34, 3299–3302. doi: 10.1093/molbev/msx248
Shajitha, P. P., Dhanesh, N. R., Ebin, P. J., Laly, J., Aneesha, D., Reshma, J., et al. (2016). A combined chloroplast atpB-rbcL and trnL-F phylogeny unveils the ancestry of balsams (Impatiens spp.) in the Western Ghats of India. 3. Biotech. 6, 1–5. doi: 10.1007/s13205-016-0574-8
Sharp, P. M., Li, W. H. (1986). An evolutionary perspective on synonymous codon usage in unicellular organisms. J. Mol. Evol. 24, 28–38. doi: 10.1007/BF02099948
Shi, L., Chen, H., Jiang, M., Wang, L., Wu, X., Huang, L., et al. (2019). CPGAVAS2, an integrated plastome sequence annotator and analyzer. Nucleic Acids Res. 47, W65–W73. doi: 10.1093/nar/gkz345
Shui, Y. M., Janssens, S., Huang, S. H., Chen, W. H., Yang, Z. G. (2011). Three new species of Impatiens L. from China and Vietnam: preparation of flowers and morphology of pollen and seeds. Syst. Bot. 36, 428–439. doi: 10.1600/036364411X569615
Su, H.-J., Hogenhout, S. A., Al-Sadi, A. M., Kuo, C.-H. (2014). Complete chloroplast genome sequence of Omani lime (Citrus aurantiifolia) and comparative analysis within the rosids. PloS One 9, e113049. doi: 10.1371/journal.pone.0113049
Sun, W., Wei, Z., Gu, Y., Wang, T., Liu, B., Yan, Y. (2024). Chloroplast genome structure analysis of Equisetum unveils phylogenetic relationships to ferns and mutational hotspot region. Front. Plant Sci. 15, 1328080. doi: 10.3389/fpls.2024.1328080
Tang, P. (2017). Evolution of chloroplast genome and re-construction of phylogenetic relationships among the Actinidia (Wuhan: Wuhan Botanical Garden Chinese Academy of Sciences).
Thiel, T., Michalek, W., Varshney, R., Graner, A. (2003). Exploiting EST databases for the development and characterization of gene-derived SSR-markers in barley (Hordeum vulgare L.). Theor. Appl. Genet. 106, 411–422. doi: 10.1007/s00122-002-1031-0
Tong, Y. H., Zheng, Q., Du, X. M., Feng, S. L., Zhou, L., Ding, C. B., et al. (2022). Analysis of chloroplast genome sequence of Camellia polyodonta. J. Plant Resour. Environ. 31, 27–36. doi: 10.3969/j.issn.1674-7895.2022.05.04
Wang, R.-J., Cheng, C.-L., Chang, C.-C., Wu, C.-L., Su, T.-M., Chaw, S.-M. (2008). Dynamics and evolution of the inverted repeat-large single copy junctions in the chloroplast genomes of monocots. BMC Evol. Biol. 8, 1–14. doi: 10.1186/1471-2148-8-36
Wang, C., Li, Y., Yang, G., Zhang, W., Guo, C. (2024). Comparative analysis of chloroplast genomes and phylogenetic relationships in the endemic Chinese bamboo Gelidocalamus (Bambusoideae). Front. Plant Sci. 15, 1470311. doi: 10.3389/fpls.2024.1470311
Wang, W., Yu, H., Wang, J., Lei, W., Gao, J., Qiu, X., et al. (2017). The complete chloroplast genome sequences of the medicinal plant Forsythia suspensa (Oleaceae). Int. J. Mol. Sci. 18, 2288. doi: 10.3390/ijms18112288
Wu, L., Cui, Y., Wang, Q., Xu, Z., Wang, Y., Lin, Y., et al. (2021). Identification and phylogenetic analysis of five Crataegus species (Rosaceae) based on complete chloroplast genomes. Planta 254, 1–12. doi: 10.1007/s00425-021-03667-4
Wu, Z., Liao, R., Yang, T., Dong, X., Lan, D., Qin, R., et al. (2020). Analysis of six chloroplast genomes provides insight into the evolution of Chrysosplenium (Saxifragaceae). BMC Genomics 21, 1–14. doi: 10.1186/s12864-020-07045-4
Xia, C.-Y. (2020). “Phylogeny study and taxonomic revision of Impatiens subg,” in Clavicarpa (Chongqing, China: Southwest University).
Yang, Y., Zhu, J., Feng, L., Zhou, T., Bai, G., Yang, J., et al. (2018). Plastid genome comparative and phylogenetic analyses of the key genera in Fagaceae: highlighting the effect of codon composition bias in phylogenetic inference. Front. Plant Sci. 9, 82. doi: 10.3389/fpls.2018.00082
Yu, S.-X. (2008). Revision of the taxonomy of Impatiens species in Guangxi: Discussion on the phylogeny of native Impatiens species in China (Beijing: Institute of Botany, Chinese Academy of Sciences).
Yu, S.-X., Janssens, S. B., Zhu, X.-Y., Lidén, M., Gao, T. G., Wang, W. (2016). Phylogeny of Impatiens (Balsaminaceae): integrating molecular and morphological evidence into a new classification. Cladistics 32, 179–197. doi: 10.1111/cla.2016.32.issue-2
Yuan, Y.-M., Song, Y., Geuten, K., Rahelivololona, E., Wohlhauser, S., Fischer, E., et al. (2004). Phylogeny and biogeography of Balsaminaceae inferred from ITS sequence data. Taxon 53, 391–403. doi: 10.2307/4135617
Zhang, Z. (2022). KaKs_Calculator 3.0: calculating selective pressure on coding and non-coding sequences. Genomics. Proteomics Bioinf. 20, 536–540. doi: 10.1016/j.gpb.2021.12.002
Zhang, X., Zhao, Q. Y., Gu, Z. J., Huang, H. Q., Yan, B., Huang, M. J. (2023). Studies on pollen micromorphology of Impatiens plants in Southwest Sichuan.
Zhao, R., Yin, S. Y., Jiang, C. H., Xue, J. N., Liu, C., Cai, X. H., et al. (2022). Comparison of chloroplast genomes of medicinal plants in Aristolochiaceae. Zhongguo. Zhong. yao. za. zhi= Zhongguo. Zhongyao. Zazhi=. China J. Chin. Mater. Med. 47, 2932–2937. doi: 10.19540/j.cnki.cjcmm.20211215.101
Zheng, S., Poczai, P., Hyvönen, J., Tang, J., Amiryousefi, A. (2020). Chloroplot: an online program for the versatile plotting of organelle genomes. Front. Genet. 11, 576124. doi: 10.3389/fgene.2020.576124
Zhou, T., Zhu, H., Wang, J., Xu, Y., Xu, F., Wang, X. (2020). Complete chloroplast genome sequence determination of Rheum species and comparative chloroplast genomics for the members of Rumiceae. Plant Cell Rep. 39, 811–824. doi: 10.1007/s00299-020-02532-0
Keywords: Impatiens, chloroplast genome, comparative analysis, phylogenetic relationships, taxonomic study
Citation: Yang M, Lan W, Zhong J, Ma H, Huang X, Huang M and Huang H (2025) Phylogenetic analysis of nine Impatiens species from subgenus Clavicarpa and subgenus Impatiens (Sect. Impatiens and Sect. Racemosae) based on chloroplast genomes. Front. Plant Sci. 16:1541320. doi: 10.3389/fpls.2025.1541320
Received: 07 December 2024; Accepted: 27 February 2025;
Published: 27 March 2025.
Edited by:
Linchun Shi, Chinese Academy of Medical Sciences and Peking Union Medical College, ChinaReviewed by:
Jing Li, Integrated DNA Technologies, United StatesCopyright © 2025 Yang, Lan, Zhong, Ma, Huang, Huang and Huang. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Meijuan Huang, bWVpanVhbmh1YW5nQHN3ZnUuZWR1LmNu; Haiquan Huang, aGFpcXVhbl9odWFuZ0Bzd2Z1LmVkdS5jbg==
†These authors have contributed equally to this work
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.