
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
REVIEW article
Front. Plant Sci. , 12 February 2025
Sec. Aquatic Photosynthetic Organisms
Volume 16 - 2025 | https://doi.org/10.3389/fpls.2025.1540914
 Thomas Barske
Thomas Barske Martin Hagemann*
Martin Hagemann*Cyanobacteria are the only prokaryotes capable of performing oxygenic photosynthesis. To thrive under environmental fluctuations, photosynthesis and metabolic activities needs to be adjusted. Previous studies showed that the acclimation of primary carbon metabolism to fluctuating carbon/nitrogen levels is mainly regulated at post-transcriptional level including diverse posttranslational modifications (PTMs). Protein phosphorylation is regarded as main PTM in the sensing and balancing metabolic changes. In this review we aim to summarize the knowledge on serine/threonine-specific protein phosphorylation among cyanobacteria. Phosphoproteome studies identified several hundred phosphoproteins bearing many more specific phosphorylation sites. On the other hand, only relatively few serine/threonine-specific protein kinases were annotated in cyanobacterial genomes, for example 12 in the model cyanobacterium Synechocystis sp. PCC 6803. Systematic mutation of the kinase-encoding genes revealed first insights into their specific functions and substrates. Future research is needed to address how a limited number of protein kinases can specifically modify hundreds of phosphoproteins and to uncover their roles in the regulatory networks of cyanobacterial metabolism.
Cyanobacteria are the only prokaryotes performing oxygenic photosynthesis, i.e. they assimilate all organic matter from inorganic nutrients using light energy to produce the necessary energy and reducing power thereby releasing O2. In fact, ancient cyanobacteria invented this process at least 2.7 billon years ago, which had a profound impact on the earth geochemistry (Hohmann-Marriott and Blankenship, 2011). Furthermore, chloroplasts found in the kingdom of plantae are derivatives from a single endosymbiotic event approximately 1.5 billion years ago, in which a free-living cyanobacterium was engulfed by an heterotrophic eukaryotic cell (e.g., Weber et al., 2006). Since then, cyanobacteria continued to evolve and through their outstanding capability to adapt to changing environmental conditions inhabited all kinds of diverse habitats in which some light is available (Houmard, 1995). Today, cyanobacteria contributing to almost one quarter of globally fixed CO2 and additionally are important assimilators of atmospheric N2 (Kasting and Siefert, 2002; Durall and Lindblad, 2015). Lately, through increasing concern about climate change and the exploration for sustainable energy sources raised much interest in photoautotrophic cyanobacteria as chassis to produce biofuels and chemical feedstock (Angermayr et al., 2009; Ducat et al., 2011; Hagemann and Hess, 2018). Thus, bioengineered cyanobacteria were generated to manufacture products such as ethanol (Deng and Coleman, 1999; Kopka et al., 2017), isobutyraldehyde and isobutanol (Atsumi et al., 2009), fatty acids (Liu et al., 2011), sucrose (Ducat et al., 2012; Qiao et al., 2018), and isoprene (Lindberg et al., 2010; Pade et al., 2016). However, production yields remained low making the production of bio-compounds not yet economically feasible (Hagemann and Hess, 2018).
In order to improve the application of cyanobacteria, a deeper understanding of the primary carbon metabolism and its regulation is necessary. As in all other oxygenic phototrophs, cyanobacteria assimilate CO2 into organic matter through the Calvin-Benson-Bassham (CBB) cycle, with ribulose 1,5-bisphosphate carboxylase/oxygenase (Rubisco) as the key CO2-fixing enzyme. To compensate for the low CO2 affinity of Rubisco and its side reaction with O2 (Tcherkez et al., 2006), which leads to the production of the toxic byproduct 2-phosphoglycolate that is salvaged in the photorespiratory cycle (Eisenhut et al., 2008a), cyanobacteria evolved the inorganic carbon (Ci, CO2 and bicarbonate)-concentrating mechanism (CCM), which increases CO2 levels in the proximity of Rubisco. The CCM in Synechocystis sp. PCC 6803 comprises three bicarbonate uptake transporters: (i) the constitutively expressed Na+-bicarbonate symporter BicA (Price et al., 2004), (ii) the low Ci induced ABC-type transporter BCT1 (Omata et al., 1999), and (iii) Na+-bicarbonate symporter SbtA (Shibata et al., 2002). Additionally, CO2 can be converted into bicarbonate by either the constitutively expressed Ndh1-4 or the low Ci-induced Ndh1-3 complex (Shibata et al., 2001). Rubisco together with carbonic anhydrase are confined to the carboxysome, in which the accumulated bicarbonate is converted into CO2 thereby saturating Rubisco carboxylation and largely inhibiting its oxygenase reaction (Rae et al., 2013; Hagemann et al., 2021).
The fixed carbon is then channeled into different metabolic routes to produce cyanobacterial biomass and reserve polymers (reviewed in Lucius and Hagemann, 2024). Previous studies revealed that cyanobacteria such as Synechocystis sp. PCC 6803 cells undergo a global metabolic reprogramming when cultivated under different C/N ratios, e.g., after shifts from high CO2 (5%, HC) into ambient air (0.04% CO2, LC) (Eisenhut et al., 2008b). Interestingly, the distinct metabolic signature is similar to metabolic changes in the model plant Arabidopsis thaliana under low versus high photorespiratory flux (Orf et al., 2016a). However, shifts from HC to LC do not cause significant changes in transcript levels for enzymes involved in primary carbon metabolism (Klähn et al., 2015). Likewise, proteomic studies revealed that carbon metabolism proteins respond more strongly to light changes, but barely to different Ci availability (Jahn et al., 2018; Spät et al., 2021; Barske et al., 2023). Such findings point toward biochemical control rather than transcriptional regulation to enable a quick acclimation of carbon partitioning without comparatively high energetic costs for proteomic responses (Jablonsky et al., 2016). In contrast, the expression of CCM-related genes is under control of transcription factors, namely NdhR (Figge et al., 2001; Wang et al., 2004), CmpR (Omata et al., 2001), and CyAbrB2 (Shalev-Malul et al., 2008; Orf et al., 2016b).
In many studies the unicellular cyanobacterial strain Synechocystis sp. PCC 6803 (hereafter Synechocystis) have been used as model. This strain was isolated from a freshwater pond in Berkeley and became part of the Pasteur Culture Collection (PCC) in 1968 (Rippka et al., 1979; Zavřel et al., 2017). Although being classified as a freshwater cyanobacterium, Synechocystis can be found in a variety of different habitats e.g., coastal areas and even areas of high salinity (Reed and Stewart, 1985; Pattanayak et al., 2015). Synechocystis can perform different modes of lifestyle i.e., photoautotrophy, mixotrophy, and light-activated heterotrophy on external glucose (reviewed in Lucius and Hagemann, 2024). As a non-diazotrophic cyanobacterium, it can only grow with combined nitrogen sources, usually with nitrate. Synechocystis was the first organism performing oxygenic photosynthesis with a fully sequenced genome (Kaneko et al., 1996; Kaneko and Tabata, 1997). Through its natural competency to take up foreign DNA and incorporate it into its own genome by homologous recombination (Grigorieva and Shestakov, 1982), Synechocystis is accessible to genetic manipulation. The early available genome sequence and the genetic systems permitted to generate and characterize a large collection of specific mutants of Synechocystis, including collection of different protein kinase-encoding genes. Moreover, new proteomic technologies have been applied to this model strain that uncovered a high number of phosphoproteins that can regulate cyanobacterial metabolism under different growth conditions. Therefore, our review is mostly dealing with results obtained from studying Synechocystis and protein-kinase-defective mutants of this strain.
To accommodate appropriate metabolic fluxes, the primary C-metabolism needs to be able to flexible acclimated itself towards changing environmental conditions, which is mainly performed through biochemical, post-transcriptional regulation (Jablonsky et al., 2016). Because some metabolic routes involve similar enzymes, which can work in opposite directions (reviewed in Lucius and Hagemann, 2024), a multilayered regulatory system is needed to effectively respond to changes in environment and to avoid futile cycles within the metabolic network. However, regulatory mechanisms, particularly on post-transcriptional level of the primary C-metabolism are scarcely understood among cyanobacteria.
During the last years, a few regulatory circuits have been identified that somehow regulate the carbon metabolism in Synechocystis and likely other cyanobacteria. The RNA polymerase σ factor SigE can act as a positive regulator of genes involved in carbohydrate catabolism in dark-exposed cells living in a heterotrophic lifestyle and shows a circadian oscillation reaching its peak in light/dark transition (Osanai et al., 2005, 2011). Further mutant studies identified the histidine kinase (Hik) 8 (encoded by sll0750), an orthologue to circadian clock protein SasA, to play a role in the control of the C-metabolism (Osanai et al., 2015; Huang et al., 2023). Likely Hik8 interacts with response regulator (Rre) 37 that is encoded by the gene sll1330 (Osanai et al., 2014). A mutant defective in Rre37 is no longer capable of light-activated heterotrophic growth (Osanai et al., 2014). Furthermore, Hik 37 (slr0110) seem to be involved in glucose-mediated catabolism (Gao et al., 2014). The transcription factor RpaA is somehow involved in SigE degradation in the dark and stimulates transcription of enzymes of glycogen and glucose metabolism (Iijima et al., 2015; Köbler et al., 2018). Together with the clock complex KaiAB1C1-SasA, RpaA also affects the switch from autotrophy in the light to the usage of stored carbon in the dark (Scheurer et al., 2021).
Posttranslational modifications i.e. covalent modifications of amino acid side chains, protein-protein interaction and effector-metabolite-binding become of interest as regulatory mechanism in bacterial carbon allocation and have been reported to play an important role in coordinating glycolytic fluxes in animal and plant cells (e.g., Zaffagnini et al., 2013; van Heerden et al., 2015). GlnB (PII) is one of the best characterized regulatory proteins in cyanobacteria. By binding 2-oxoglutarate (2OG), the precursor for ammonia assimilation in cyanobacteria, ATP and/or ADP, PII is able to integrate information about the C/N balance and energy state of the cell and adjust C- and N-fluxes accordingly (reviewed in Forchhammer et al., 2022). Recently, PII has been demonstrated to interact with the small protein PirC (Orthwein et al., 2021). PirC is said to be released from PII under N-limitation sensed by increased cellular 2OG levels and interacts with the phosphoglycerate mutates 1 (Pgam1) and thus blocking fluxes into the lower glycolysis and thereby favoring anabolic glycolytic routes and the accumulation of glycogen. Furthermore, PII can be phosphorylated at Ser49 in Synechocystis (reviewed in Forchhammer et al., 2022) and the PII phosphorylation state responds to different C/N ratios (e.g., Schwarz et al., 2014). Recently, it was shown that absence of the carbon-metabolism-regulating Hik8 impacts also PII phosphorylation (Huang et al., 2023). The small, disordered protein CP12 known to bind the CBB enzymes GapDH2 and PRK under oxidative conditions is modulating CBB and OPP activity under redox changing conditions (Gurrieri et al., 2021; Lucius et al., 2022). In addition to proteins sensing the redox or metabolic state inside the cyanobacteria, proteome studies revealed an increasing number of post-translational protein modifications (PTMs) on many enzymes involved in the primary C and N metabolism. Among them, protein phosphorylation is regarded to play a central role in the signal recognition and regulation of cellular activities among cyanobacteria as has been shown in many other bacteria (Maček et al., 2019).
Reversible protein phosphorylation and dephosphorylation is one of the most important PTMs that is catalyzed by protein kinases and phosphatases, respectively. Protein kinases are defined as enzymes that transfer a phosphate group onto an amino acid (AA) side chain in a target protein (Cozzone, 1993). Generally, protein kinases using the γ-phosphate of ATP as phosphate group donor but additionally GTP and PEP were shown to serve as phosphate group suppliers (Hunter, 2012). According to the targeted AA, the Nomenclature Committee of International Union of Biochemist classified protein kinases into 5 groups: (i) AA with alcohol groups as acceptors such as serine and threonine forming phosphate esters, (ii) AA with phenolic groups as acceptors namely tyrosine forming phosphate esters, (iii) basic AA such as histidine, arginine and lysine producing phosphoramidates, (iv) AA with acyl groups as acceptors such as aspartate and glutamic acid generating mixed phosphate-carboxylate acid anhydrites, and (v) cysteine residues as acceptor that produce thioesters (Hunter, 2012; Cozzone, 1993).
In bacteria, protein phosphorylation by different protein kinase classes (Figure 1) is considered as a signal transduction device that links impulses from environmental conditions with the regulation of essential physiological processes (Kobir et al., 2011). Such signals are often transmitted via histidine autophosphorylation and aspartate phosphorylation in two component systems (TCS), which represents the most common type of protein phosphorylation signaling in bacteria and marks the most abundant form of p-events in bacteria (Maček et al., 2019). TCS can be found in all bacterial species and comprise of a signal sensing histidine kinase (Hik). Often membrane associated, upon signal perception Hik use ATP to auto-phosphorylate themselves on a histidine residue which in turn can transfer the phosphate onto an aspartate residue on a response regulator (Rre). The Rre can usually bind on specific promoter sequences thereby translating the sensed signal into a stress-specific response (Stock et al., 2000; Hirakawa et al., 2020). Hence, TCS mediated signal cascades lead to transcriptional changes in most cases (Gross et al., 1989).
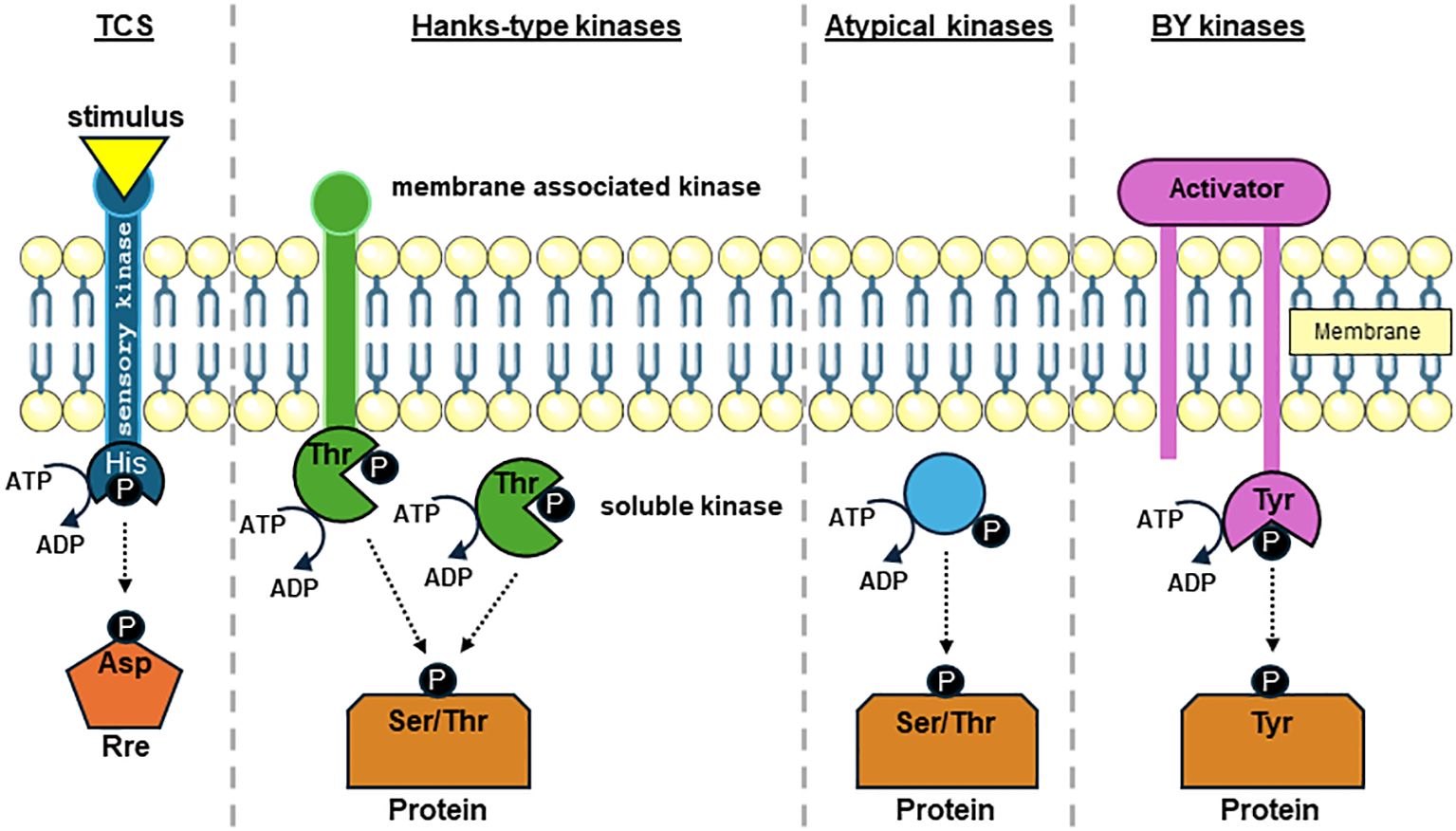
Figure 1. Overview of protein-kinase-mediated phosphate groups transfer to amino acid side chains in bacteria. Phosphate group transfer to side chains of histidine (His), aspartic acid (Asp), serine (Ser), threonine (Thr), and tyrosine (Tyr) residues by prototypic protein kinases using the γ-phosphate of ATP as phosphate group donor is displayed. Two component system (TCS) consist usually of a membrane associated His kinase (Hik) and a response regulator (Rre) that commonly affects the transcription of target genes. Upon sensing an environmental stimulus, autophosphorylation occurs on a His sidechain, which subsequently is transferred to an Asp in the Rre. Bacterial Hanks-kinases can either harbor transmembrane domains or are soluble within the cytosol. Hanks-type kinases can catalyze the phosphate group transfer to both Ser and Thr sidechains. Atypical Ser and Thr protein kinases share no or only marginal similarity to Hanks-kinases. Bacterial tyrosine (BY) kinases require an activator protein to stabilize the kinase complex. When activated, BY kinase autophosphorylate on a Tyr residue and transfers the phosphate group to a Tyr residue on the target protein.
Phospho-esters on serine, threonine and tyrosine are the second most common form of protein phosphorylation in bacteria (Maček et al., 2019). Protein phosphorylation upon serine and threonine residues are commonly catalyzed by Hanks-type kinases (Figure 1) that share a strong similarity to kinases found in eukaryotes (Hanks et al., 1988; Hanks, 2003; Pereira et al., 2011; Mijakovic et al., 2016; Maček et al., 2019). Hanks-type kinases in bacteria can be membrane bound or can exist as soluble proteins. It should be noted that in addition to Hanks-type kinases other kinases have been described phosphorylating serine and threonine residues termed atypical kinases (Pereira et al., 2011; Mijakovic et al., 2016). Unlike serine and threonine phosphorylation no eukaryotic-like tyrosine kinases were identified in bacteria (Grangeasse et al., 2012). The majority of tyrosine residue phosphorylation is carried out by the protein kinase family bacterial protein-tyrosine kinases (BY-kinases; Figure 1). BY-kinases often sense extracellular signals involving an activator protein part (Grangeasse et al., 2012; Mijakovic et al., 2016). Protein arginine phosphorylation was discovered recently in B. subtilis, where it was proven to affect factors in the stress response system. This view was only recently expanded by reports of arginine phosphorylation in S. aureus (Fuhrmann et al., 2016; Elsholz et al., 2012; Schmidt et al., 2014; Junker et al., 2018). The cysteine thiol-group is not only prone to oxidative modification but can also undergo phosphorylation (Mijakovic et al., 2016). Cysteine phosphorylation has been shown to play a regulatory role in the control of transcription factors in S. aureus (Sun et al., 2012).
Finally, protein phosphorylation is often involved in transcriptional regulation using alternative sigma factors in bacteria. In this process anti-sigma factors involved in partner switching systems such as RsbW and SpoIIAB are serine kinase, which are highly conserved among bacterial species and can specifically activate/inactivate alternative sigma factors under specific growth conditions. Such partner switching system belong to the abundant of phosphorylation-events in bacteria (e.g., Bouillet et al., 2018). The kinase activity of RsbW-like proteins such as Slr1861 or PmgA of Synechocystis have been verified by in vitro phosphorylation assays (Shi et al., 1999; Nakamura et al., 2024). Several proteins involved in sigma factor partner switching systems have been identified as phosphoproteins in Synechocystis (Supplementary Table 1).
Cyanobacterial diversity and ability to adapt to changing environmental conditions is highlighted by their complexity of signal perception systems (Zorina, 2013). Thus, rapid signal transmission from a receptor to a receiver is essential for an adequate response to stress. Such dynamics is possible through reversible PTMs for example in form of protein phosphorylation (Maček and Mijakovic, 2011). First direct evidence of protein phosphorylation in cyanobacteria was obtained through [32P] orthophosphate in vivo and cell-free in vitro labeling experiments in Calothrix sp. PCC 7601 (Schuster et al., 1984), Synechococcus sp. PCC 6301 (Allen et al., 1985), Anabaena sp. PCC 7120 (Mann et al., 1991), Synechocystis (Bloye et al., 1992; Hagemann et al., 1993), and in Synechococcus elongatus PCC 7942 (Forchhammer and Tandeau De Marsac, 1994). These studies showed phosphorylation of many protein bands, which were dynamic under altering growth conditions. However, in most cases the nature of the phosphorylated protein(s) remained enigmatic. The overall physiological importance of protein phosphorylation in cyanobacteria and their role in regulatory processes e.g., nitrogen metabolism, carbon metabolism, cell motility or osmotic stress has been widely acknowledged in later studies as outlined below.
Even though histidine and aspartate phosphorylation usually account for the majority of p-events in bacteria, their thermodynamic instability renders the in vivo detection challenging in phosphoproteomic studies (Maček et al., 2019). The mostly applied extraction of phosphorylated peptides is carried out under acidic conditions (pH <4) and thus favoring the detection of p-events on serine, threonine and tyrosine residues instead of phosphorylated histidine and aspartate residues through their low chemically stability in a lower pH (Maček and Mijakovic, 2011; Mijakovic and Maček, 2012; Maček et al., 2019). Depending for long on quantifying relative proteins levels of perturbed biological systems in 2D-gels, early global phosphoproteome studies relied heavily on this technology to identify p-events (Maček and Mijakovic, 2011). Technical advances in gel-free and mass-spectroscopy (MS)-based approaches resulted in the first published in vivo phosphoproteome of Bacillus subtilis in an exponential growth phase (Maček et al., 2007a) and of E. coli (Maček et al., 2007b), which highly increased the number of detected phosphoproteins. Similar developments occurred among cyanobacteria.
2D-gel-based and gel-free phosphoproteomic experiments revealed an ever-increasing number of phosphoproteins in Synechocystis. The first 2D-gel based phosphoproteome study was performed under fluctuating salinity condition (Mikkat et al., 2014). To identify phosphoproteins, the Synechocystis proteins were first separated by isoelectric focusing (first dimension) and then in large SDS-gels (second dimension). This separation technique discriminated many proteins and was limited to about 500 proteins visible in separate spots. Protein phosphorylation could initially be identified by specific dyes, which was subsequently verified by MS-based estimation of peptide masses that increased by 80 Da when a phosphate group was added (for more details see Mikkat et al., 2014). This study was able to identify 32 phosphoproteins such as GlnB, Kai proteins and a great number of proteins taking part in the primary C-metabolism (e.g., Pgm, Eno, Gap2). The first gel-free LC-MS-based comparative global phosphoproteome study analyzed the acclimation of Synechocystis towards N starvation (Spät et al., 2015). In this approach, the entire proteome is digested with different proteases and then the defined protein fragments are separated by LC techniques. Proteins can subsequently be identified by peptide-mass-fingerprinting via MS. This technique compares the sizes of in silico produced proteolytic fragment patterns with the in vitro measured peptide sizes, thereby permitting precise protein identification. Again, phosphorylated peptides show an increased mass of 80 Da. To improve the coverage and identification of phosphoproteins, phosporylated peptides are usually enriched by metal-affinity chromatography before the LC separation (for more details see Spät et al., 2015). This study showed an overall increase in p-events under low N conditions, among many proteins were involved in CCM, the primary C-metabolism and the central regulator of C/N partitioning GlnB (PII). Similarly, a snapshot phosphoproteome experiment under variating light conditions was performed and discovered that many photosynthesis related proteins undergo changes in p-occupancy in changing light qualities (Chen et al., 2015). Furthermore, this study showed through side specific mutations that the phosphorylation of phycocyanin β-subunit CpcB is of importance, e.g. in state transition. Another phosphoproteome study investigated Synechocystis cells acclimated to different carbon conditions with the emphasis on evaluating protein phosphorylation in relation with C-fixation, photosynthesis and photoprotection (Angeleri et al., 2016). Alterations in the proteome and phosphoproteome levels under different growth conditions were also analyzed by Toyoshima et al. (2020). The authors compared photoautotrophic, photoheterotrophic, heterotrophic growth in the presence of light and mixotrophic condition together with growth under N-starvation (24 h, 48 h) and revealed that relatively small alterations in the proteome can be accompanied with vast deviations in overall protein phosphorylation status in the cell and highlights the underlying significance of reversible protein phosphorylation in acclimation processes. We analyzed recently changes in the proteome and phosphoproteome of the Synechocystis wild-type and selected kinase mutants under different CO2 levels (Spät et al., 2021; Barske et al., 2023). These studies showed that the abundances of enzymes involved in the primary C metabolism remained similar under high and low CO2, however, several proteins showed marked changes in their phosphorylation.
Summarizing the phosphoproteome attempts with Synechocystis (Mikkat et al., 2014; Spät et al., 2015, 2018, 2021; Chen et al., 2015; Angeleri et al., 2016; Toyoshima et al., 2020; Barske et al., 2023) resulted in a list of at least 481 phosphoproteins (Supplementary Table 1), i.e., more than 10% of the entire Synechocystis proteome can be phosphorylated under specific growth conditions. Furthermore, many of the identified phosphoproteins displayed more than one phosphorylated site, hence, the total number of phosphorylation events (p-events) on Synechocystis is much higher (see also Spät et al., 2023 for a comprehensive overview on proteomic and phosphoproteomic data obtained with Synechocystis). Similarly high numbers of phosphoproteins were detected through a global phosphoproteome study with marine cyanobacterium Synechococcus sp. PCC 7002 where 410 p-events on 245 proteins could be detected (Yang et al., 2013). Moreover, Liang et al. (2021) analyzed the phosphoproteome of Nostoc flagelliforme in response to dehydration. The authors were able to detect 271 phosphoproteins with 1168 phosphorylation sites. Among them, many showed changed phosphorylation under dehydration, especially on proteins known to be involved in signal transduction and response to reactive oxygen species (ROS).
The physiological relevance of the identified p-events remains in most cases elusive. Clear evidence was provided for the importance of protein phosphorylation sites in phosphoglucomutase 1 (Pgm1), which revealed the role of posttranslational modification on serine 47 (S47) during nitrogen starvation and its concomitant role in modulating its activity (Doello et al., 2022). Additionally, it was discovered that regulatory proteins such as PII (Forchhammer and Tandeau De Marsac, 1994) and CP12 (Spät et al., 2018, 2021) are prone to protein phosphorylation and furthermore presenting changes in their phosphorylation status under changing environmental conditions (Forchhammer and Tandeau De Marsac, 1994; Spät et al., 2018). Another early identified phosphoprotein is KaiC, the clock protein involved in the circadian rhythm of Synechococcus elongatus PCC 7942 (Nakajima et al., 2005) and many other cyanobacteria including Synechocystis (e.g., Köbler et al., 2024). Here, the rate of phosphorylation and dephosphorylation determines the phase of the circadian clock, which is sensed by certain output proteins (Golden et al., 1998). These examples indicate the significance of posttranslational modification in form of protein phosphorylation as a hallmark in metabolic regulation in Synechocystis and likely in other cyanobacteria.
Even though phosphorylation on serine, threonine and tyrosine residues were known to occur in cyanobacteria, studies on kinases creating phospho-monoesters were initially largely neglected (Mann, 1994). This changed after the discovery of a Hanks-type Ser/Thr kinase in Myxococcus xanthus (Muñoz-Dorado et al., 1991). Subsequently, similar PCR-based strategies were also employed in cyanobacteria such as Anabaena sp. PCC 7120 and resulted in the discovery of the first cyanobacterial Hanks-kinase (Zhang, 1993, 1996) with an increasing number of similar kinases and phosphatases found in other cyanobacteria ever since (Zhang et al., 2005). The available genome sequences revealed that the distribution of Ser/Thr and Tyr kinases and phosphatases is rather uneven among cyanobacteria and can vary from 0, detected in some Prochlorococcus strains, to up 56 encoding genes in the nitrogen-fixing strain N. punctiforme PCC 73102 (Zhang et al., 2005; Zorina, 2013). Interestingly, freshwater cyanobacteria seem to harbor a larger number of Ser/Thr and Tyr kinases and phosphatases compared to marine cyanobacteria, while no clear correlation between genome size and number of Ser/Thr and Tyr kinases and phosphatases could be made (Zhang et al., 2005).
Soon after the first cyanobacterial genome sequence of Synechocystis was released (Kaneko et al., 1996), it was searched for Ser/Thr and Tyr-specific protein kinases and phosphatases, which bear similarities to Hanks-kinases and Hanks-phosphatases. The search revealed that Synechocystis possesses 12 Ser/Thr kinases, one Tyr-kinases and 7 phosphatases (Zhang et al., 1998; Leonard et al., 1998; Shi et al., 1998). The 12 kinases can be divided into “serine/threonine-protein N2-like kinases” - PKN2 and “activity of BC1 complex” kinases - ABC1 (Leonard et al., 1998), respectively (Table 1). The PKN2 group comprises the protein kinases SpkA-G and share strong structural similarity to Hanks-kinases (Leonard et al., 1998; Zhang et al., 2007). Protein kinase activity could be verified for SpkA-F using artificial substrates such as histone, MBP and casein as well as autophosphorylation activity with exception of SpkE (Kamei et al., 2001, 2002, 2003). However, Zorina et al. (2014) detected protein kinase activity for SpkE with casein and other substrates as well, hence, all annotated PKN2-type kinases with the exception of SpkG are principally active enzymes. Protein kinases SpkH-L belong to atypical ABC1 protein kinase family (Leonard et al., 1998). Only the catalytic activity of the SpkH was recently confirmed (Zorina et al., 2023). Several groups established collections of protein kinase mutants of Synechocystis, which were screened regarding phenotypic alterations and sometimes specific protein substrates during the last years (Kamei et al., 2002; Laurent et al., 2008; Zorina et al., 2011; Mata-Cabana et al., 2012; Barske et al., 2023). The combination of such screening attempts with subsequent phosphoproteomic experiments and physiological measurements permitted the functional characterization of several annotated Spk’s in Synechocystis during the last years (Figure 2). Basic, kinase-specific features are summarized in Table 1 and discussed in the next paragraphs.
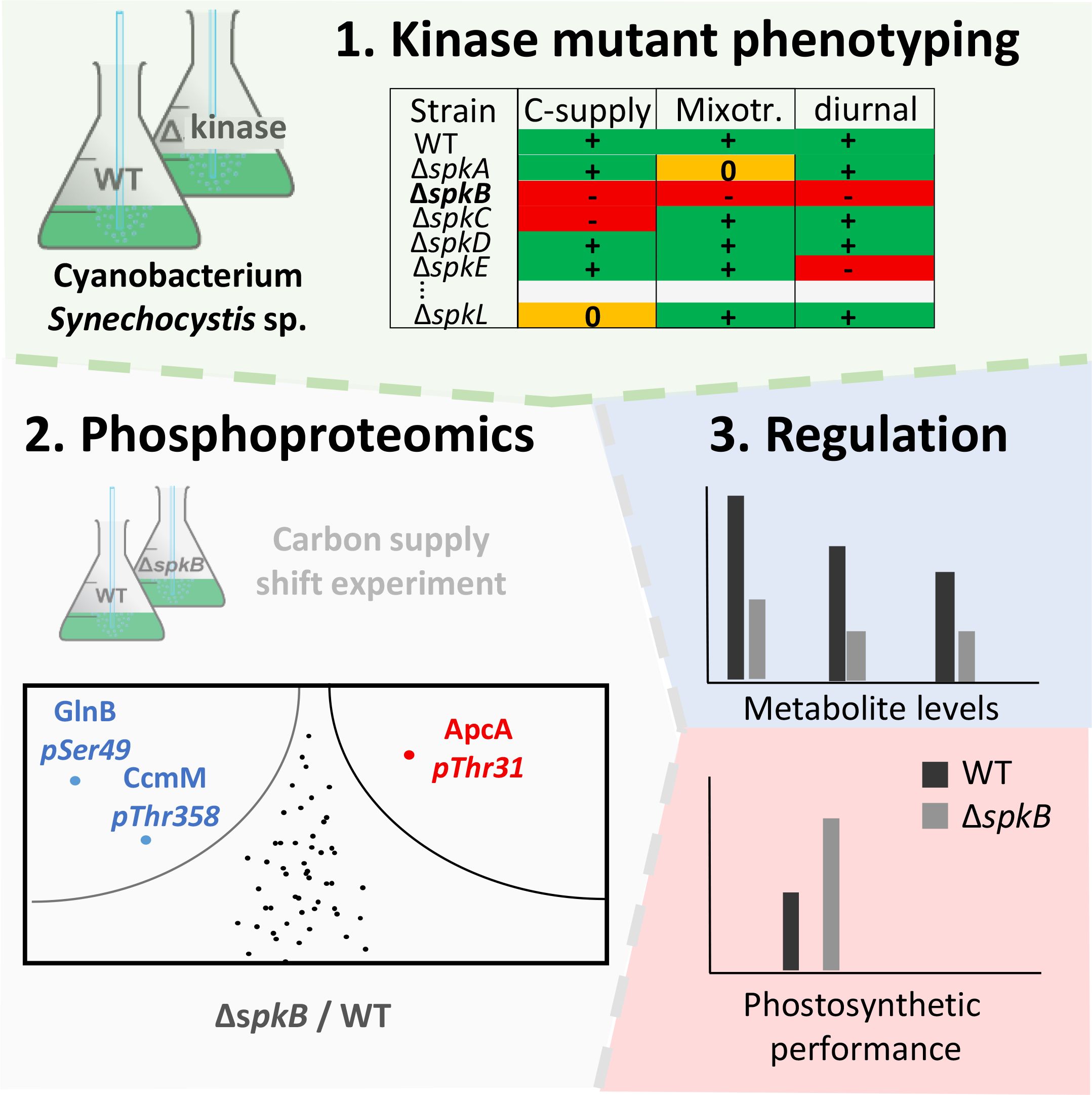
Figure 2. Strategy for the in depth characterization of serine/threonine-specific protein kinases (Spk’s) combining the screening of mutant collections under different growth conditions, applying phosphoproteomics, and subsequent physiological characterizations (e.g., Barske et al., 2023).
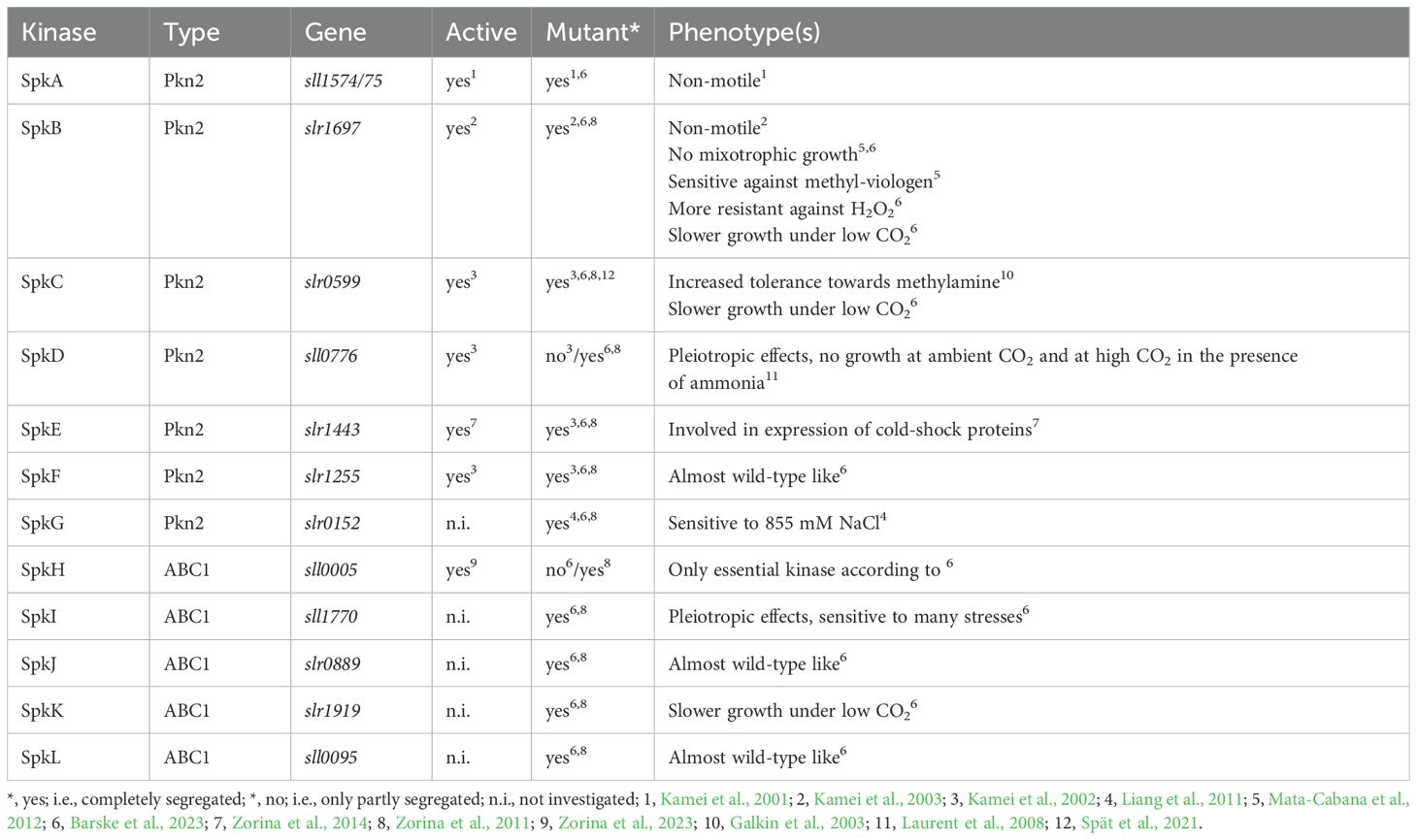
Table 1. Overview on annotated Ser/Thr-specific protein kinases (Spk’s) in Synechocystis sp. PCC 6803 and some related features.
The spkA encoding sequence is found on two separate genes in the primarily sequenced glucose-tolerant Synechocystis wild type (Table 1), whereas it forms a continuous gene in the original glucose-sensitive wild type from the Pasteur Culture Collection. Inactivation of the wild-type spkA gene provided evidence of its role in cell motility, because the null mutant ΔspkA showed no colony movement under lateral illumination (Kamei et al., 2001). Though retaining pili, the authors proposed that SpkA might not be essential for pili biogenesis but can somehow influence gliding motility towards a light source. This view was further expanded by a gene transcriptome analysis with ΔspkA in which expression changes for pilin encoding operons were detected. Some of them were subsequently verified in Northern-blot experiments showing reduced level of pilA9 and enhanced quantities of pilA5 together with the observation of missing thick pili (Panichkin et al., 2006). Our recent survey of phenotypic variations among spk mutants revealed slower growth of ΔspkA under mixotrophic and salt stress conditions compared to wild type (Barske et al., 2023).
Initial reports for the SpkB-deficient mutant ΔspkB described a similar phenotype as ΔspkA with a strongly reduced gliding motility (Kamei et al., 2003). Additionally, it was demonstrated that the absence of SpkB resulted in a redox-sensitive phenotype, because in contrast to wild type the mutant was unable to tolerate elevated concentrations of menadion, methylviologen or high light doses, i.e. treatments that induce high intracellular levels of ROS. Furthermore, 32P-labelling of protein extracts revealed a decrease of one of the most prominent phosphorylated protein bands, which mainly contained GlyS (glycyl-tRNA synthetase subunit beta) and thus might represent one specific substrate of SpkB (Mata-Cabana et al., 2012).
Our recent experiments with the mutant ΔspkB verified the non-motile phenotype, which is correlated by the differential accumulation of many proteins associated with the cell surface including pili subunits (Barske et al., 2023). The most striking differences of ΔspkB compared to the Synechocystis wild type were related to carbon metabolism, because mutant cells grew significantly slower under low CO2, while high CO2 conditions complemented the phenotype. Furthermore, the mutant ΔspkB was sensitive to glucose additions in the light and under diurnal light rhythms, which is correlated with changes in glycogen accumulation (Barske et al., 2023). The detailed analysis of the proteome revealed only few alterations in protein abundances, none of them is directly correlated with the changed ability to grow at ambient CO2 or in the presence of external glucose. However, the phosphoproteome revealed that two proteins were not phosphorylated anymore in the mutant. To this end, phosphorylation of the proteins Sll1545 and Slr0483 was only detected in wild-type samples. Slr1545 is the glutathione S-transferase, Gst1, which plays an important role in the redox regulation of proteins among cyanobacteria (Kammerscheit et al., 2019). Its expression change could be related to the observed differences in ROS-tolerance of ΔspkB. The Slr0483 protein is a membrane protein of unknown function that bears a CAAD domain (cyanobacterial aminoacyl-tRNA synthetase appended domain, PMID: 18775859). In addition to these proteins without any detected phosphorylation in ΔspkB, a few p-events were identified with significantly diminished phosphorylation (Barske et al., 2023). Among them, the (auto)phosphorylation of SpkF at T24 was significantly diminished under ambient CO2 in the mutant ΔspkB accompanied with generally reduced SpkF levels in this strain. Furthermore, the phosphorylation of the PII (GlnB) protein at S49 was strongly reduced in ΔspkB compared to the WT when grown at high CO2 and especially when shifted for 3 h to ambient CO2. The PII protein is the master regulator of many aspects in the C/N homeostasis in cyanobacteria and other organisms as well (reviewed in Forchhammer et al., 2022). But no protein involved in the CCM or the glucose/glycogen metabolism showed any significant changes in abundance and phosphorylation levels, which make it difficult to explain the observed phenotypic alteration of mutant ΔspkB under different carbon conditions (Barske et al., 2023). Finally, it should be noted that our study detected only non-phosphorylated GlyS in extracts of the wild type and the mutant ΔspkB, while a previous study identified GlyS as substrate for SpkB (Mata-Cabana et al., 2012).
Several studies were conducted on the characterization of SpkC, which is encoded by slr0599. Initially it was reported that mutant ΔspkC shows an increased tolerance toward the toxic compounds methylamine and methionine-sulfoximine in low light, while several other kinase mutants behaved like wild type in this study (Galkin et al., 2003). A microarray experiment with wild-type cells exposed to the inhibitors DCMU and DBMIB in low light characterized an induction of spkC transcription (Hihara et al., 2003). Zorina et al. (2011) analyzed the mutant ΔspkC and several others in a 2D-gel-based phosphoproteome study under standard and heat shock conditions. They found rather minor changes in the overall proteome but several changes in the phosphorylation of protein spots, among them some heat-shock proteins including GroES. Subsequently, they used recombinant GroES as substrate in in vitro phosphorylation assays with crude extracts from several spk mutants. In contrast to wild type and other spk mutants, extracts from mutant ΔspkC and ΔspkF as well as ΔspkK showed no GroES phosphorylation, whereas crude protein extracts from the corresponding complementation strains were able to phosphorylate GroES. According to these results they proposed a cascade of SpkC/F/K in phosphorylating the heat-shock protein GroES (Zorina et al., 2011).
More recently, we re-investigated the phosphoproteome of the mutant ΔspkC under high and low CO2, because it showed a diminished growth after shifts from high to ambient CO2 levels (Spät et al., 2021). Overall, more than 2500 proteins were quantified in our study, equivalent to approximately 70% of the Synechocystis theoretical proteome. Proteins with changing abundances under different CO2 levels are often involved in the CCM or the nitrogen metabolism, whereas enzymes related to primary carbon metabolism showed almost no changes in their abundances. Interestingly, among the few proteins with changed abundances the bicarbonate transporter SbtA and some other low-CO2-induced proteins were less strongly accumulated in mutant ΔspkC than in wild type, which is consistent with the slower growth of the mutant at ambient conditions (Spät et al., 2021). Furthermore, 105 phospho-proteins harboring over 200 site-specific phosphorylation events were identified. Subunits of the bicarbonate transporter BCT1 and the redox switch protein CP12 were among phosphoproteins with reduced phosphorylation levels at lower CO2, whereas the serine/threonine protein kinase SpkC revealed increased phosphorylation levels, which supports its possible regulatory involvement in the acclimation towards changing CO2 conditions. To identify potential target proteins of SpkC-mediated phosphorylation, we searched for phosphoproteins that were reproducibly identified in wild type but were never detected in the mutant ΔspkC. According to this attempt, at least four potential phosphorylation targets of SpkC were identified. Among them, the phosphorylation in the subunit CmpB of the ATP-dependent bicarbonate transporter BCT1 was always absent in ΔspkC, whereas it was detectable in every replicate from wild type. This phosphorylation change might be directly connected to lowered growth of mutant ΔspkC under lowered CO2. In addition, phosphorylation of the DnaJ-like protein Sll1384, of Slr1619, and of the response-regulator-like protein Slr6040 on plasmid pSYSX occurred exclusively in the wild type. Collectively, our data make it likely that SpkC is somehow involved in the sensing/regulation of the acclimation of cyanobacteria towards limiting CO2 conditions. In this regard it is interesting to note that SpkC was identified as integral protein in the Synechocystis plasma membrane (Liberton et al., 2016).
In an early study it was shown that SpkD might be essential for Synechocystis to grow under standard laboratory conditions, because only partially deletion of the spkD-encoding sll0776 gene was achieved (Kamei et al., 2002). However, in later studies it was possible to receive completely segregated ΔspkD mutants (Laurent et al., 2008; Barske et al., 2023), which make the essential character of the kinase questionable. These different results can be related to slight differences in the wild-type strains of Synechocystis used in the corresponding studies, however, it cannot be excluded that still unknown suppressor mutations allowed the segregation in the later studies. Laurent et al. (2008) observed a clear upregulation of spkD expression under low CO2 compared to high CO2 and concomitantly ΔspkD was unable to grow under ambient air. The authors suggested that SpkD is involved in adjustments of TCA cycle metabolites, because the supplementation of alternative organic carbon sources such as glucose, phosphoglyceraldehyde and pyruvate could not rescue the mutant phenotype, whereas the external addition of TCA metabolites e.g., acetyl CoA, succinate, and 2OG were able to revert it (Laurent et al., 2008). However, in our recent survey of many spk mutants, we did not observe very clear alterations in mutant ΔspkD compared to wild type, including its wild-type-like growth under low and elevated CO2 levels (Barske et al., 2023). Recent transcriptome analysis found spkD upregulated in the lexA mutant (Kizawa et al., 2016). Moreover, phosphoproteome studies under different light (Chen et al., 2015) conditions and our results under changing Ci availability (Spät et al., 2021) identified SpkD as a phosphoprotein. Finally, SpkD together with SpkG were hypothesized to influence the expression of fatty acid desaturases, i.e., they might play a role in the regulation of PUFA contents (Chen et al., 2021).
The PKN2-type kinase SpkE is encoded by the gene slr1443 in Synechocystis and was verified to be an active protein kinase (Table 1). An early study reported that SpkE seems be required for post-translational modification of pili proteins after biogenesis (Kim et al., 2004). Another study provided some evidence that SpkE might be involved in cold shock response (Zorina et al., 2014). Cells of the mutant ΔspkE showed the most prominent change in the expression of cold-shock proteins when shifted from 32°C to 22°C for 30 min. In addition to the protein synthesis pattern, the 2D-gel-based approach showed also significant changes in the protein phosphorylation patterns. Some of these changes were also observed in the mutant Δhik33, a sensory histidine-kinase proven to be involved in cold shock and other stress responses. These finding led to the hypothesis that SpkE might be an additional component in cold stress responses in Synechocystis (Zorina et al., 2014). In our study, the mutant ΔspkE showed diminished growth under diurnal conditions, but the mechanism underlying this phenotype was not further investigated (Barske et al., 2023).
Induction of spkF expression was observed in a transcriptome study analyzing ethanol resistance in Synechocystis (Wang et al., 2012). Furthermore, SpkF has been shown to be prone to modulation by phosphorylation upon N-starvation (Spät et al., 2015) with a reported transiently increase in phosphorylation of SpkF when Synechocystis resuscitates after chlorosis (Spät et al., 2018) and under changes in Ci availability (Spät et al., 2021). Our recent phosphoproteome study identified a diminished (auto)phosphorylation of SpkF at T24 under ambient CO2 in the mutant ΔspkB accompanied with generally reduced SpkF levels in this strain, however, the ability of mutant ΔspkF to grow at elevated or ambient CO2 conditions was not changed (Barske et al., 2023). A global proteome study identified SpkF as integral proteins in the plasma membrane (Liberton et al., 2016).
In contrast to most other PKN2-type kinases, the enzymatic activity of SpkG has not been verified. The spkG gene is transcribed as last gene in the photosystem II assembly protein operon (slr0144-slr0152), in which several phosphoproteins have been identified (Angeleri et al., 2016). An upcoming study provided indirect evidence that SpkG might represent an active protein kinase, because in the mutant ΔspkG the phosphorylation of some proteins, especially ferredoxin 5 (Fd5) did not occur anymore, hence, it could be concluded that SpkG might be specifically involved in Fd5 phosphorylation (Angeleri et al., 2018). Moreover, spkG expression was strongly induced when Synechocystis was exposed to elevated salt concentrations and, accordingly, growth of ΔspkG was impaired in the presence of 855 mM NaCl (Liang et al., 2011), whereas the mutant ΔspkG grew like wild type on plates supplemented with 500 mM NaCl (Barske et al., 2023). The possible involvement of SpkG in the sensing of salt stress was supported by transcriptome studies, which showed that some high-salt-induced genes are less strongly expressed in the mutant ΔspkG (Liang et al., 2011). More recently, SpkG together with SpkD were hypothesized to influence the expression of fatty acid desaturases, i.e. they might play a role in the regulation of PUFA contents (Chen et al., 2021). Finally, transcriptomics showed that in H2O2-treated wild-type cells spkG was found to be downregulated (Li et al., 2004).
In our survey of different spk mutants, we were not able to completely segregate the mutation of sll0005 encoding SpkH despite several attempts (Barske et al., 2023), whereas Zorina et al. (2011) reported a complete knock out of this gene. As mentioned above, the different results can be related to slight differences in the wild type strains of Synechocystis used in the corresponding studies, however, it cannot be excluded that still unknown suppressor mutations allowed the segregation in the earlier study. Recently, its enzymatic activity was verified, because recombinant SpkH protein was able to phosphorylate some typical protein kinase substrate proteins (Zorina et al., 2023). The spkH expression was reported to be light induced and significantly downregulated after addition of salt (Allakhverdiev et al., 2002). Correspondingly, spkH was also found to be highly expressed in Synechocystis under osmotic stress in from of supplemented sorbitol (Paithoonrangsarid et al., 2004).
Among the ABC1-type kinases, SpkI received most attentions in Synechocystis (Irmler and Forchhammer, 2001; Huang et al., 2002; Wang et al., 2022). This high interest is related to the localization of the SpkI encoding gene sll1770, which is situated upstream of the gene for the PII phosphatase pphA. Hence, it was initially suspected that SpkI might be involved in PII phosphorylation, because the PII-specific kinase is still not identified. But it was proven that SpkI is non-essential for the phosphorylation of the PII protein (Irmler and Forchhammer, 2001). A global gene expression study displayed a spkI induction under UV-B and intense light intensities (Huang et al., 2002). More recent reports highlighted a wild-type-like growth accompanied with a higher NPQ capacity in ΔspkI under standard growth condition (Wang et al., 2022). However, the mutant showed reduced growth and decreased net photosynthesis in a high salt environment. Moreover, reduced levels of major photosynthetic protein were detected while fluorescence measurements revealed modification in photosystem I and Cytb6f-complex together with impaired QA and state transition. Hence the authors suggested a central role of SpkI in maintaining photosynthesis during salt acclimation (Wang et al., 2022). A reduced capability of mutant ΔspkI to grow under diverse stress conditions such as changes in the CO2 level, diurnal rhythms, and salt stress was also found in our recent survey of kinase mutants (Barske et al., 2023). This pleiotropic phenotype might be related to the important role of SpkI in regulation of photosynthesis.
To our knowledge, no conclusive data are available for SpkJ.
SpkK could be located to the thylakoid membrane (Baers et al., 2019) and is proposed to act in a cascade together with SpkC and SpkF in the phosphorylation of the heat shock protein GroES (Zorina et al., 2011). We reported a slower growth of mutant ΔspkK when transferred from high to ambient CO2 conditions (Barske et al., 2023).
The function of SpkL has not been analyzed in great detail. The spkL gene showed lower expression under iron starvation (Hernández-Prieto et al., 2012).
Much progress has been made in the field of cyanobacterial phosphoproteins and protein kinases. Especially the technical advances in phosphoproteomics resulted in an increasing number of identified phosphoproteins in Synechocystis (Supplementary Table 1) and other cyanobacteria as well. However, it must be mentioned that compared to the MS-based protein identification and quantification, which permits the quantitative detection of approximately 75% of the theoretical Synechocystis proteome (Spät et al., 2023), the detection and quantification of phosphoproteins is much less reliable. In many cases, phosphorylated peptides need to be enriched through specific affinity media, which make the method less reproducible and difficult to absolute quantification. In our recent studies, we applied identical growth, protein extraction and proteome methods, which resulted in a consistent list of quantified proteins (almost 95% reproduction) in the Synechocystis wild type and selected mutants in the two independent studies (Spät et al., 2021; Barske et al., 2023). In contrast, the list of identified phosphoproteins and their CO2-dependent changes in phosphorylation levels was much less reproducible, only approximately 50% of the phosphorylated proteins were found in the two studies. Furthermore, the detection limit of present-day MS methods improved many times, which makes it possible to detect also rare p-events, i.e. phosphorylated amino acid residues with less than 1% occupancy. But, the physiological meaning of such rare protein phosphorylation is highly questionable.
Nevertheless, the high number of detected phosphoproteins, many of them showed changes in the phosphorylation under different growth conditions, is consistent with the assumption that protein phosphorylation represents the dominant PTM involved in regulation of metabolism and stress acclimation. However, only in a relatively few cases, we have solid biochemical or physiological evidence that the changes in protein phosphorylation indeed affected enzyme activities or photosynthetic performance (examples are mentioned and discussed above). In the majority of cases, we can only speculate which or even if any function is related to the observed protein phosphorylation. Obviously, many more studies are needed in which specific protein variants with and without existing phosphorylation sites are studied in vivo and in vitro in detail. Such studies should include the identification of the responsible protein kinases and also phosphatases. In most cases the responsible kinases for a specific p-event are unknown. Even many proposed functions for the studied protein kinases are only evidence based. In vitro assays proofing the direct interaction with their claimed targets remains to be conducted in future experiments, for example with recombinant SpkB to verify its specific involvement in the phosphorylation of GlnB as proposed by Barske et al. (2023).
One of the most obvious open question is related to the large discrepancy between the high number of phosphoproteins and the even higher number of p-events (Table 1; Spät et al., 2023) and the much smaller number of Ser/Thr protein kinases in Synechocystis (Table 1). Similar large deviations between hundreds of p-events and small numbers of annotated protein kinases have been reported for other bacteria as well, which initiated attempts to use computational predictions to analyze the kinase/substrate interactions (reviewed in Grunfeld et al., 2024). Generally, this situation makes it difficult to assume very specific kinase/protein substrate interactions in bacteria such as Synechocystis. Hence, it is generally assumed that bacterial Spks have a rather relaxed substrate specificity and the same kinase can act in different regulatory mechanisms together with other proteins (Kobir et al., 2011; Grunfeld et al., 2024). Moreover, in addition to kinase mediated p-events, kinase independent p-events are possible, as it has recently been shown for the phosphorylation of PGM1 and PGM2 (Doello et al., 2022; Neumann et al., 2022). Those events could vastly increase the total number of p-events. Furthermore, the complete picture of Ser/Thr protein kinases in Synechocystis should be re-evaluated. Many bacterial Ser/Thr protein kinases resemble Hanks-type kinases which is also true for Synechocystis (Zhang et al., 1998; Leonard et al., 1998; Shi et al., 1998). Nevertheless, there is a chance of evolutionary unrelated protein kinases such as the isocitrate dehydrogenase kinase/phosphatase found in E. coli (Zheng and Jia, 2010) or the non-enzymatic acetyl phosphate dependent phosphorylation which was reported in the bacteria Streptococcus pneumoniae (Kaiser et al., 2020) and B. subtilis (Cairns et al., 2015). Hence, many more efforts are necessary to uncover the complex role of protein phosphorylation in stress acclimation and metabolic control in cyanobacteria.
TB: Writing – original draft. MH: Writing – original draft, Writing – review & editing.
The author(s) declare financial support was received for the research, authorship, and/or publication of this article. Our work on the regulation of carbon metabolism was supported by the Deutsche Forschungsgemeinschaft (DFG, grant HA2002/23-1) in the frame of the research consortium SCyCode (FOR2816) and funds of the University Rostock, Germany.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fpls.2025.1540914/full#supplementary-material
Allakhverdiev, S. I., Nishiyama, Y., Miyairi, S., Yamamoto, H., Inagaki, N., Kanesaki, Y., et al. (2002). Salt stress inhibits the repair of photodamaged photosystem II by suppressing the transcription and translation of psbA genes in Synechocystis. Plant Physiol. 130, 1443–1453. doi: 10.1104/pp.011114
Allen, J. F., Sanders, C. E., Holmes, N. G. (1985). Correlation of membrane protein phosphorylation with excitation energy distribution in the cyanobacterium Synechococcus 6301. FEBS Lett. 193, 271–275. doi: 10.1016/0014-5793(85)80167-8
Angeleri, M., Muth-Pawlak, D., Aro, E. M., Battchikova, N. (2016). Study of O-phosphorylation sites in proteins involved in photosynthesis-related processes in synechocystis sp. Strain PCC 6803: application of the SRM approach. J. Proteome Res. 15, 4638–4652. doi: 10.1021/acs.jproteome.6b00732
Angeleri, M., Zorina, A., Aro, E. M., Battchikova, N. (2018). ). Interplay of SpkG kinase and the Slr0151 protein in the phosphorylation of ferredoxin 5 in Synechocystis sp. strain PCC 6803. FEBS Lett. 592, 411–421. doi: 10.1002/1873-3468.12970
Angermayr, S. A., Hellingwerf, K. J., Lindblad, P., Teixeira de Mattos, M. J. (2009). Energy biotechnology with cyanobacteria. Curr. Opin. Biotechnol. 20, 257–263. doi: 10.1016/J.COPBIO.2009.05.011
Atsumi, S., Higashide, W., Liao, J. C. (2009). Direct photosynthetic recycling of carbon dioxide to isobutyraldehyde. Nat. Biotechnol. 27, 1177–1180. doi: 10.1038/nbt.1586
Baers, L. L., Breckels, L. M., Mills, L. A., Gatto, L., Deery, M. J., Stevens, T. J., et al. (2019). Proteome mapping of a cyanobacterium reveals distinct compartment organization and cell-dispersed metabolism. Plant Physiol. 181, 1721–1738. doi: 10.1104/pp.19.00897
Barske, T., Spät, P., Schubert, H., Walke, P., Maček, B., Hagemann, M. (2023). The role of serine/threonine-specific protein kinases in cyanobacteria - spkB is involved in acclimation to fluctuating conditions in synechocystis sp. PCC 6803. Mol. Cell. Proteomics 22, 100656. doi: 10.1016/j.mcpro.2023.100656
Bloye, S. A., Silman, N. J., Mann, N. H., Carr, N. G. (1992). Bicarbonate concentration by Synechocystis PCC6803: Modulation of protein phosphorylation and inorganic carbon transport by glucose. Plant Physiol. 99, 601–606. doi: 10.1104/pp.99.2.601
Bouillet, S., Arabet, D., Jourlin-Castelli, C., Méjean, V., Lobbi-Nivol, C. (2018). Regulation of σ factors by conserved partner switches controlled by divergent signalling systems. Environ. Microbiol. Rep. 10, 127–139. doi: 10.1111/1758-2229.12620
Cairns, L. S., Martyn, J. E., Bromley, K., Stanley-Wall, N. R. (2015). An alternate route to phosphorylating DegU of Bacillus subtilis using acetyl phosphate. BMC Microbiol. 15, 1–12. doi: 10.1186/s12866-015-0410-z
Chen, G., Cao, Y., Zhong, H., Wang, X., Li, Y., Cui, X., et al. (2021). Serine/threonine kinases play important roles in regulating polyunsaturated fatty acid biosynthesis in synechocystis sp. PCC6803. Front. Bioengineering Biotechnol. 9. doi: 10.3389/fbioe.2021.618969
Chen, Z., Zhan, J., Chen, Y., Yang, M., He, C., Ge, F., et al. (2015). Effects of Phosphorylation of b Subunits of Phycocyanins on State Transition in the Model Cyanobacterium Synechocystis sp. PCC 6803. Plant Cell Physiol. 56, 1997–2013. doi: 10.1093/pcp/pcv118
Cozzone, A. J. (1993). ATP-dependent protein kinases in bacteria. J. Cell. Biochem. 51, 7–13. doi: 10.1002/jcb.240510103
Deng, M., Coleman, J. R. (1999). Ethanol synthesis by genetic engineering in cyanobacteria. Appl. Environ. Microbiol. 65, 523–528. doi: 10.1128/AEM.65.2.523-528.1999
Doello, S., Neumann, N., Forchhammer, K. (2022). Regulatory phosphorylation event of Phosphoglucomutase 1 tunes its activity to regulate glycogen metabolism. FEBS J. 289, 6005–6020. doi: 10.1111/febs.16471
Ducat, D. C., Avelar-Rivas, J. A., Way, J. C., Silvera, P. A. (2012). Rerouting carbon flux to enhance photosynthetic productivity. Appl. Environ. Microbiol. 78, 2660–2668. doi: 10.1128/AEM.07901-11/ASSET/BC9BEAB8-0E0B-4DDC-8351-99352EE89293/ASSETS/GRAPHIC/ZAM9991031310006.JPEG
Ducat, D. C., Way, J. C., Silver, P. A. (2011). Engineering cyanobacteria to generate high-value products. Trends Biotechnol. 29, 95–103. doi: 10.1016/j.tibtech.2010.12.003
Durall, C., Lindblad, P. (2015). Mechanisms of carbon fixation and engineering for increased carbon fixation in cyanobacteria. Algal Res. 11, 263–270. doi: 10.1016/j.algal.2015.07.002
Eisenhut, M., Huege, J., Schwarz, D., Bauwe, H., Kopka, J., Hagemann, M. (2008b). Metabolome phenotyping of inorganic carbon limitation in cells of the wild type and photorespiratory mutants of the cyanobacterium Synechocystis sp. Strain PCC 6803. Plant Physiol. 148, 2109–2120. doi: 10.1104/pp.108.129403
Eisenhut, M., Ruth, W., Haimovich, M., Bauwe, H., Kaplan, A., Hagemann, M. (2008a). The photorespiratory glycolate metabolism is essential for cyanobacteria and might have been conveyed endosymbiontically to plants. Proc. Natl. Acad. Sci. U.S.A. 105, 17199–17204. Available at: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2579401/pdf/zpq17199.pdf.
Elsholz, A. K. W., Turgay, K., Michalik, S., Hessling, B., Gronau, K., Oertel, D., et al. (2012). Global impact of protein arginine phosphorylation on the physiology of Bacillus subtilis. Proc. Natl. Acad. Sci. U.S.A. 109, 7451–7456. doi: 10.1073/pnas.1117483109
Figge, R. M., Cassier-Chauvat, C., Chauvat, F., Cerff, R. (2001). Characterization and analysis of an NAD(P)H dehydrogenase transcriptional regulator critical for the survival of cyanobacteria facing inorganic carbon starvation and osmotic stress. Mol. Microbiol. 39, 455–469. doi: 10.1046/j.1365-2958.2001.02239.x
Forchhammer, K., Selim, K. A., Huergo, L. F. (2022). New views on PII signaling: from nitrogen sensing to global metabolic control. Trends Microbiol. 30, 722–735. doi: 10.1016/J.TIM.2021.12.014
Forchhammer, K., Tandeau De Marsac, N. (1994). The PII Protein in the Cyanobacterium Synechococcus sp. Strain PCC 7942 Is Modified by Serine Phosphorylation and Signals the Cellular N-Status. J. Bacteriology 176, 84–91.
Fuhrmann, J., Subramanian, V., Kojetin, D. J., Thompson, P. R. (2016). Activity-based profiling reveals a regulatory link between oxidative stress and protein arginine phosphorylation. Cell Chem. Biol. 23, 967–977. doi: 10.1016/J.CHEMBIOL.2016.07.008
Galkin, A. N., Mikheeva, L. E., Shestakov, S. V. (2003). The insertional inactivation of genes encoding eukaryotic-type serine/threonine protein kinases in the cyanobacterium Synechocystis sp. PCC 6803. Microbiology 72, 52–57. doi: 10.1023/A:1022226006358
Gao, L., Shen, C., Liao, L., Huang, X., Liu, K., Wang, W., et al. (2014). Functional proteomic discovery of Slr0110 as a central regulator of carbohydrate metabolism in Synechocystis species PCC6803. Mol. Cell. Proteomics 13, 204–219. doi: 10.1074/MCP.M113.033803
Golden, S. S., Johnson, C. H., Kondo, T. (1998). The cyanobacterial circadian system: a clock apart. Curr. Opin. Microbiol. 1, 669–673. doi: 10.1016/s1369-5274(98)80113-6
Grangeasse, C., Nessler, S., Mijakovic, I. (2012). Bacterial tyrosine kinases: Evolution, biological function and structural insights. Philos. Trans. R. Soc. B: Biol. Sci. 367, 2640–2655. doi: 10.1098/rstb.2011.0424
Grigorieva, G., Shestakov, S. (1982). Transformation in the cyanobacterium Synechocystis sp. 6803. FEMS Microbiol. Lett. 13, 367–370. doi: 10.1111/j.1574-6968.1982.tb08289.x
Gross, R., Aricò, B., Rappuoli, R. (1989). Families of bacterial signal-transducing proteins. Mol. Microbiol. 3, 1661–1667. doi: 10.1111/j.1365-2958.1989.tb00152.x
Grunfeld, N., Levine, E., Libby, E. (2024). Experimental measurement and computational prediction of bacterial Hanks-type Ser/Thr signaling system regulatory targets. Mol. Microbiol. 122, 152–164. doi: 10.1111/mmi.15220
Gurrieri, L., Fermani, S., Zaffagnini, M., Sparla, F., Trost, P. (2021). Calvin–Benson cycle regulation is getting complex. Trends Plant Sci. 26, 898–912. doi: 10.1016/j.tplants.2021.03.008. Elsevier Ltd.
Hagemann, M., Golldack, D., Biggins, J., Erdmann, N. (1993). Salt-dependent protein phosphorylation in the cyanobacterium Synechocystis PCC 6803. FEMS Microbiol. Lett. 113, 205–209. doi: 10.1111/j.1574-6968.1993.tb06515.x
Hagemann, M., Hess, W. R. (2018). Systems and synthetic biology for the biotechnological application of cyanobacteria. Curr. Opin. Biotechnol. 49, 94–99. doi: 10.1016/j.copbio.2017.07.008
Hagemann, M., Song, S., Brouwer, E.-M. (2021). “Inorganic carbon assimilation in cyanobacteria: mechanisms, regulation, and engineering,” in Cyanobacteria Biotechnology. Eds. Nielsen, J., Lee, S., Stephanopoulos, G., Husdon, P. (Weinheim, Germany: John Wiley and Sons), 1–31.
Hanks, S. K. (2003). Genomic analysis of the eukaryotic protein kinase superfamily: A perspective. Genome Biol. 4, 111. doi: 10.1186/gb-2003-4-5-111
Hanks, S. K., Quinn, A. M., Hunter, T. (1988). The protein kinase family: Conserved features and deduced phylogeny of the catalytic domains. Science 241, 42–52. doi: 10.1126/science.3291115
Hernández-Prieto, M. A., Schön, V., Georg, J., Barreira, L., Varela, J., Hess, W. R., et al. (2012). Iron deprivation in synechocystis: inference of pathways, non-coding RNAs, and regulatory elements from comprehensive expression profiling. G3 2, 1475–1495. doi: 10.1534/g3.112.003863
Hihara, Y., Sonoike, K., Kanehisa, M., Ikeuchi, M. (2003). DNA microarray analysis of redox-responsive genes in the genome of the cyanobacterium Synechocystis sp. strain PCC 6803. J. Bacteriology 185, 1719–1725. doi: 10.1128/JB.185.5.1719-1725.2003
Hirakawa, H., Kurushima, J., Hashimoto, Y., Tomita, H. (2020). Progress overview of bacterial two-component regulatory systems as potential targets for antimicrobial chemotherapy. Antibiotics 9, 1–15. doi: 10.3390/antibiotics9100635
Hohmann-Marriott, M. F., Blankenship, R. E. (2011). Evolution of photosynthesis. Annu. Rev. Plant Biol. 62, 515–548. doi: 10.1146/annurev-arplant-042110-103811
Houmard, J. (1995). “How do cyanobacteria perceive and adjust to their environment?,” in Molecular Ecology of Aquatic Microbes. Ed. Joint, I. (Germany: Springer Berlin), 153–170.
Huang, C., Duan, X., Ge, H., Xiao, Z., Zheng, L., Wang, G., et al. (2023). Parallel proteomic comparison of mutants with altered carbon metabolism reveals Hik8 regulation of P(II) phosphorylation and glycogen accumulation in a cyanobacterium. Mol. Cell. Proteomics 22, 100582. doi: 10.1016/j.mcpro.2023.100582
Huang, L., McCluskey, M. P., Ni, H., LaRossa, R. A. (2002). Global gene expression profiles of the cyanobacterium Synechocystis sp. strain PCC 6803 in response to irradiation with UV-B and white light. J. Bacteriology 184, 6845–6858. doi: 10.1128/JB.184.24.6845-6858.2002
Hunter, T. (2012). Why nature chose phosphate to modify proteins. Phil. Trans. R. Soc B 367, 2513–2516. doi: 10.1098/rstb.2012.0013
Iijima, H., Shirai, T., Okamoto, M., Kondo, A., Yokota Hirai, M., Osanai, T. (2015). Changes in primary metabolism under light and dark conditions in response to overproduction of a response regulator RpaA in the unicellular cyanobacterium Synechocystis sp. PCC 6803. Front. Microbiol. 6. doi: 10.3389/fmicb.2015.00888
Irmler, A., Forchhammer, K. (2001). A PP2C-type phosphatase dephosphorylates the PII signaling protein in the cyanobacterium Synechocystis PCC 6803. Proc. Natl. Acad. Sci. U.S.A. 98, 12978–12983. doi: 10.1073/pnas.231254998
Jablonsky, J., Papacek, S., Hagemann, M. (2016). Different strategies of metabolic regulation in cyanobacteria: From transcriptional to biochemical control. Sci. Rep. 6, 33024. doi: 10.1038/srep33024
Jahn, M., Vialas, V., Karlsen, J., Maddalo, G., Edfors, F., Forsström, B., et al. (2018). Growth of cyanobacteria is constrained by the abundance of light and carbon assimilation proteins. Cell Rep. 25, 478–486.e8. doi: 10.1016/j.celrep.2018.09.040
Junker, S., Maa, S., Otto, A., Michalik, S., Morgenroth, F., Gerth, U., et al. (2018). Spectral library based analysis of arginine phosphorylations in staphylococcus aureus. Mol. Cell. Proteomics 17, 335–348. doi: 10.1074/MCP.RA117.000378
Kaiser, S., Hoppstädter, L. M., Bilici, K., Heieck, K., Brückner, R., Siemens Chair, W. (2020). Control of acetyl phosphate-dependent phosphorylation of the response regulator CiaR by acetate kinase in Streptococcus pneumoniae. Microbiology 166, 411–421. doi: 10.1099/mic.0.000894
Kamei, A., Yoshihara, S., Yuasa, T., Geng, X., Ikeuchi, M. (2003). Biochemical and functional characterization of a eukaryotic-type protein kinase, SpkB, in the cyanobacterium, Synechocystis sp. PCC 6803. Curr. Microbiol. 46, 296–301. doi: 10.1007/s00284-002-3887-2
Kamei, A., Yuasa, T., Geng, X., Ikeuchi, M. (2002). Biochemical Examination of the Potential Eukaryotic-type Protein Kinase Genes in the Complete Genome of the Unicellular Cyanobacterium Synechocystis sp. PCC 6803. DNA Res. 9, 71–78.
Kamei, A., Yuasa, T., Orikawa, K., Geng, X. X., Ikeuchi, M. (2001). A eukaryotic-type protein kinase, SpkA, is required for normal motility of the unicellular cyanobacterium Synechocystis sp. strain PCC 6803. J. Bacteriology 183, 1505–1510. doi: 10.1128/JB.183.5.1505-1510.2001
Kammerscheit, X., Chauvat, F., Cassier-Chauvat, C. (2019). First in vivo evidence that Glutathione-S-transferase operates in photo-oxidative stress in cyanobacteria. Front. Microbiol. 10. doi: 10.3389/fmicb.2019.01899
Kaneko, T., Sato, S., Kotani, H., Tanaka, A., Asamizu, E., Nakamura, Y., et al. (1996). Sequence analysis of the genome of the unicellular cyanobacterium synechocystis sp. Strain PCC6803. II. Sequence determination of the entire genome and assignment of potential protein-coding regions (Supplement). DNA Res. 3, 185–209. doi: 10.1093/dnares/3.3.185
Kaneko, T., Tabata, S. (1997). Complete genome structure of the unicellular cyanobacterium Synechocystis sp. PCC6803. Plant Cell Physiol. 38, 1171–1176. doi: 10.1093/oxfordjournals.pcp.a029103
Kasting, J. F., Siefert, J. L. (2002). Life and the evolution of Earth’s atmosphere. Science 296, 1066–1068. doi: 10.1126/science.1071184
Kim, Y. H., Park, Y. M., Kim, S. J., Park, Y., Choi, J. S., Chung, Y. H. (2004). The role of Slr1443 in pilus biogenesis in Synechocystis sp. PCC 6803: Involvement in post-translational modification of pilins. Biochem. Biophys. Res. Commun. 315, 179–186. doi: 10.1016/j.bbrc.2004.01.036
Kizawa, A., Kawahara, A., Takimura, Y., Nishiyama, Y., Hihara, Y. (2016). RNA-seq profiling reveals novel target genes of LexA in the cyanobacterium synechocystis sp. PCC 6803. PCC 6803. Front. Microbiol. 7. doi: 10.3389/fmicb.2016.00193
Klähn, S., Orf, I., Schwarz, D., Matthiessen, J. K. F., Kopka, J., Hess, W. R., et al. (2015). Integrated transcriptomic and metabolomic characterization of the low-carbon response using an ndhR mutant of Synechocystis sp. PCC 6803. Plant Physiol. 169, 1540–1556. doi: 10.1104/pp.114.254045
Kobir, A., Shi, L., Boskovic, A., Grangeasse, C., Franjevic, D., Mijakovic, I. (2011). Protein phosphorylation in bacterial signal transduction. Biochim. Biophys. Acta - Gen. Subj. 1810, 989–994. doi: 10.1016/j.bbagen.2011.01.006
Köbler, C., Schmelling, N. M., Wiegard, A., Pawlowski, A., Pattanayak, G. K., Spät, P., et al. (2024). Two KaiABC systems control circadian oscillations in one cyanobacterium. Nat. Commun. 15, 7674. doi: 10.1038/s41467-024-51914-5
Köbler, C., Schultz, S. J., Kopp, D., Voigt, K., Wilde, A. (2018). The role of the Synechocystis sp. PCC 6803 homolog of the circadian clock output regulator RpaA in day–night transitions. Mol. Microbiol. 110, 847–861. doi: 10.1111/mmi.14129
Kopka, J., Schmidt, S., Dethloff, F., Pade, N., Berendt, S., Schottkowski, M., et al. (2017). Systems analysis of ethanol production in the genetically engineered cyanobacterium Synechococcus sp. PCC 7002. Biotechnol. Biofuels 10, 56. doi: 10.1186/s13068-017-0741-0
Laurent, S., Jang, J., Janicki, A., Zhang, C. C., Bédu, S. (2008). Inactivation of spkD, encoding a Ser/Thr kinase, affects the pool of the TCA cycle metabolites in Synechocystis sp. strain PCC 6803. Microbiology 154, 2161–2167. doi: 10.1099/mic.0.2007/016196-0
Leonard, C. J., Aravind, L., Koonin, E. V. (1998). Novel families of putative protein kinases in bacteria and archaea: Evolution of the “eukaryotic” protein kinase superfamily. Genome Res. 8, 1038–1047. doi: 10.1101/gr.8.10.1038
Li, H., Singh, A. K., McIntyre, L. M., Sherman, L. A. (2004). Differential gene expression in response to hydrogen peroxide and the putative PerR regulon of Synechocystis sp. strain PCC 6803. J. Bacteriology 186, 3331–3345. doi: 10.1128/JB.186.11.3331-3345.2004
Liang, W., Yan, F., Wang, M., Li, X., Zhang, Z., Ma, X., et al. (2021). Comprehensive Phosphoproteomic Analysis of Nostoc flagelliforme in Response to Dehydration Provides Insights into Plant ROS Signaling Transduction. ACS Omega 6, 13554–13566. doi: 10.1021/acsomega.0c06111
Liang, C., Zhang, X., Chi, X., Guan, X., Li, Y., Qin, S., et al. (2011). Serine/threonine protein kinase SpkG is a candidate for high salt resistance in the unicellular cyanobacterium Synechocystis sp. PCC 6803. PloS One, 6, e18718. doi: 10.1371/journal.pone.0018718
Liberton, M., Saha, R., Jacobs, J. M., Nguyen, A. Y., Gritsenko, M. A., Smith, R. D., et al. (2016). Global proteomic analysis reveals an exclusive role of thylakoid membranes in bioenergetics of a model cyanobacterium. Mol. Cell. Proteomics 15, 2021–2032. doi: 10.1074/MCP.M115.057240
Lindberg, P., Park, S., Melis, A. (2010). Engineering a platform for photosynthetic isoprene production in cyanobacteria, using Synechocystis as the model organism. Metab. Eng. 12, 70–79. doi: 10.1016/j.ymben.2009.10.001
Liu, X., Sheng, J., Curtiss, R. (2011). Fatty acid production in genetically modified cyanobacteria. Proc. Natl. Acad. Sci. U.S.A. 108 (17), 6899–6904. doi: 10.1073/PNAS.1103014108/SUPPL_FILE/PNAS.1103014108_SI.PDF
Lucius, S., Hagemann, M. (2024). The primary carbon metabolism in cyanobacteria and its regulation. Front. Plant Sci. 15, 1417680. doi: 10.3389/fpls.2024.1417680
Lucius, S., Theune, M., Arrivault, S., Hildebrandt, S., Mullineaux, C. W., Gutekunst, K., et al. (2022). CP12 fine-tunes the Calvin-Benson cycle and carbohydrate metabolism in cyanobacteria. Front. Plant Sci. 13. doi: 10.3389/FPLS.2022.1028794/FULL
Maček, B., Forchhammer, K., Hardouin, J., Weber-Ban, E., Grangeasse, C., Mijakovic, I. (2019). Protein post-translational modifications in bacteria. Nat. Rev. Microbiol. 17, 651–664. doi: 10.1038/s41579-019-0243-0
Maček, B., Gnad, F., Soufi, B., Kumar, C., Olsen, J., Mijakovic, I., et al. (2007b). Phosphoproteome analysis of E. coli reveals evolutionary conservation of bacterial Ser/Thr/Tyr phosphorylation. Mol. Cell. Proteomics 7, 299–307. doi: 10.1074/mcp.M700311-MCP200
Maček, B., Mijakovic, I. (2011). Site-specific analysis of bacterial phosphoproteomes. Proteomics 11, 3002–3011. doi: 10.1002/pmic.201100012
Maček, B., Mijakovic, I., Olsen, J., Gnad, F., Kumar, C., Jensen, P. R., et al. (2007a). The serine/threonine/tyrosine phosphoproteome of the model bacterium Bacillus subtilis. Mol. Cell. Proteomics 6, 697–707. doi: 10.1074/mcp.M600464-MCP200
Mann, N. H. (1994). Protein phosphorylation in cyanobacteria. Microbiology 140, 3207–3321. doi: 10.1099/13500872-140-12-3207
Mann, N. H., Rippka, R., Herdman, M. (1991). Regulation of protein phosphorylation in the cyanobacterium Anabaena strain PCC 7120. J. Gen. Microbiol. 137, 1. doi: 10.1099/00221287-137-2-331
Mata-Cabana, A., García-Domínguez, M., Florencio, F. J., Lindahl, M. (2012). Thiol-based redox modulation of a cyanobacterial eukaryotic-type serine/threonine kinase required for oxidative stress tolerance. Antioxidants Redox Signaling 17, 521–533. doi: 10.1089/ars.2011.4483
Mijakovic, I., Grangeasse, C., Turgay, K. (2016). Exploring the diversity of protein modifications: special bacterial phosphorylation systems. FEMS Microbiol. Rev. 3, 398–417. doi: 10.1093/femsre/fuw003
Mijakovic, I., Maček, B. (2012). Impact of phosphoproteomics on studies of bacterial physiology. FEMS Microbiol. Rev. 36, 877–892. doi: 10.1111/j.1574-6976.2011.00314.x
Mikkat, S., Fulda, S., Hagemann, M. (2014). A 2D gel electrophoresis-based snapshot of the phosphoproteome in the cyanobacterium Synechocystis sp. strain PCC 6803. Microbiology 160, 296–306. doi: 10.1099/mic.0.074443-0
Muñoz-Dorado, J., Inouye, S., Inouye, M. (1991). A gene encoding a protein serine/threonine kinase is required for normal development of M. xanthus, a gram-negative bacterium. Cell 67, 995–1006. doi: 10.1016/0092-8674(91)90372-6
Nakajima, M., Imai, K., Ito, H., Nishiwaki, T., Murayama, Y., Iwasaki, H., et al. (2005). Reconstitution of circadian oscillation of cyanobacterial KaiC phosphorylation in vitro. Science 308, 414–415. doi: 10.1126/science.1108451
Nakamura, R., Takahashi, Y., Tachibana, S., Terada, A., Suzuki, K., Kondo, K., et al. (2024). Partner-switching components PmgA and Ssr1600 regulate high-light acclimation in Synechocystis sp. PCC 6803. Plant Physiol. 196, 621–633. doi: 10.1093/plphys/kiae323
Neumann, N., Friz, S., Forchhammer, K. (2022). Glucose-1,6-bisphosphate, a key metabolic regulator, is synthesized by a distinct family of α-phosphohexomutases widely distributed in prokaryotes. mBio 13, e0146922. doi: 10.1128/mbio.01469-22
Omata, T., Gohta, S., Takahashi, Y., Harano, Y., Maeda, S. I. (2001). Involvement of a CbbR homolog in low CO2-induced activation of the bicarbonate transporter operon in cyanobacteria. J. Bacteriology 183, 1891–1898. doi: 10.1128/JB.183.6.1891-1898.2001
Omata, T., Price, G. D., Badger, M. R., Okamura, M., Gohta, S., Ogawa, T. (1999). Identification of an ATP-binding cassette transporter involved in bicarbonate uptake in the cyanobacterium Synechococcus sp. strain PCC 7942. Proc. Natl. Acad. Sci. U.S.A. 96, 13571–13576. doi: 10.1073/pnas.96.23.13571
Orf, I., Schwarz, D., Kaplan, A., Kopka, J., Hess, W. R., Hagemann, M., et al. (2016b). CyAbrB2 contributes to the transcriptional regulation of low CO2 acclimation in Synechocystis sp. PCC 6803. Plant Cell Physiol. 57, 2232–2243. doi: 10.1093/pcp/pcw146
Orf, I., Timm, S., Bauwe, H., Fernie, A. R., Hagemann, M., Kopka, J., et al. (2016a). Can cyanobacteria serve as a model of plant photorespiration? - a comparative meta-analysis of metabolite profiles. J. Exp. Bot. 67, 2941–2952. doi: 10.1093/jxb/erw068
Orthwein, T., Scholl, J., Spät, P., Lucius, S., Koch, M., Maček, B., et al. (2021). The novel PII-interactor PirC identifies phosphoglycerate mutase as key control point of carbon storage metabolism in cyanobacteria. Proc. Natl. Acad. Sci. U.S.A. 118, 2019988118. doi: 10.1073/pnas.2019988118/-/DCSupplemental
Osanai, T., Kanesaki, Y., Nakano, T., Takahashi, H., Asayama, M., Shirai, M., et al. (2005). Positive regulation of sugar catabolic pathways in the cyanobacterium Synechocystis sp. PCC 6803 by the group 2 σ factor SigE. J. Biol. Chem. 280, 30653–30659. doi: 10.1074/JBC.M505043200
Osanai, T., Oikawa, A., Azuma, M., Tanaka, K., Saito, K., Hirai, M. Y., et al. (2011). Genetic engineering of group 2 σ factor SigE widely activates expressions of sugar catabolic genes in Synechocystis species PCC 6803. J. Biol. Chem. 286, 30962–30971. doi: 10.1074/jbc.M111.231183
Osanai, T., Oikawa, A., Numata, K., Kuwahara, A., Iijima, H., Doi, Y., et al. (2014). Pathway-level acceleration of glycogen catabolism by a response regulator in the cyanobacterium Synechocystis species PCC 6803. Plant Physiol. 164, 1831–1841. doi: 10.1104/pp.113.232025
Osanai, T., Shirai, T., Iijima, H., Kuwahara, A., Suzuki, I., Kondo, A., et al. (2015). Alteration of cyanobacterial sugar and amino acid metabolism by overexpression hik8, encoding a KaiC-associated histidine kinase. Environ. Microbiol. 17, 2430–2440. doi: 10.1111/1462-2920.12715
Pade, N., Erdmann, S., Enke, H., Dethloff, F., Dühring, U., Georg, J., et al. (2016). Insights into isoprene production using the cyanobacterium Synechocystis sp. PCC 6803. Biotechnol. Biofuels 9, 1–16. doi: 10.1186/S13068-016-0503-4/TABLES/2
Paithoonrangsarid, K., Shoumskaya, M. A., Kanesaki, Y., Satoh, S., Tabata, S., Los, D. A., et al. (2004). Five histidine kinases perceive osmotic stress and regulate distinct sets of genes in Synechocystis. J. Biol. Chem. 279, 53078–53086. doi: 10.1074/JBC.M410162200
Panichkin, V. B., Arakawa-Kobayashi, S., Kanaseki, T., Suzuki, I., Los, D. A., Shestakov, S. V., et al. (2006). Serine/threonine protein kinase SpkA in Synechocystis sp. strain PCC 6803 is a regulator of expression of three putative pilA operons, formation of thick pili, and cell motility. J. Bacteriology 188, 7696–7699. doi: 10.1128/JB.00838-06
Pattanayak, G. K., Korbinian, R., Osanai, T., Iijima, H., Nakaya, Y., Kuwahara, A., et al. (2015). Seawater cultivation of freshwater cyanobacterium Synechocystis sp. PCC 6803 drastically alters amino acid composition and glycogen metabolism. Front. Microbiol. 6. doi: 10.3389/fmicb.2015.00326
Pereira, S. F. F., Goss, L., Dworkin, J. (2011). Eukaryote-like serine/threonine kinases and phosphatases in bacteria. Microbiol. Mol. Biol. Rev. 75, 192–212. doi: 10.1128/mmbr.00042-10
Price, G. D., Woodger, F. J., Badger, M. R., Howitt, S. M., Tucker, L. (2004). Identification of a SulP-type bicarbonate transporter in marine cyanobacteria. Proc. Natl. Acad. Sci. U.S.A. 101, 18228–18233. doi: 10.1073/PNAS.0405211101
Qiao, C., Duan, Y., Zhang, M., Hagemann, M., Luo, Q., Lu, X. (2018). Effects of lowered and enhanced glycogen pools on salt-induced sucrose production in a sucrose-secreting strain of Synechococcus elongatus PCC 7942. Appl. Environ. Microbiol. 84, e02023–e02017. doi: 10.1128/AEM.02023-17
Rae, B. D., Long, B. M., Badger, M. R., Price, G. D. (2013). Functions, compositions, and evolution of the two types of carboxysomes: polyhedral microcompartments that facilitate CO2 fixation in cyanobacteria and some proteobacteria. Microbiol. Mol. Biol. Rev. 77, 357–379. doi: 10.1128/MMBR.00061-12
Reed, R. H., Stewart, W. D. P. (1985). Osmotic adjustment and organic solute accumulation in unicellular cyanobacteria from freshwater and marine habitats. Mar. Biol. 88, 1–9. doi: 10.1007/BF00393037
Rippka, R., Deruelles, J., Herdman, M., Waterbury, J. B., Stanier, R. Y. (1979). Generic assignments, strain histories and properties of pure cultures of cyanobacteria. J. Gen. Microbiol. 111, 1–61. doi: 10.1099/00221287-111-1-1
Scheurer, N. M., Rajarathinam, Y., Timm, S., Köbler, C., Kopka, J., Hagemann, M., et al. (2021). Homologs of circadian clock proteins impact the metabolic switch between light and dark growth in the cyanobacterium synechocystis sp. PCC 6803. Front. Plant Sci. 12. doi: 10.3389/fpls.2021.675227
Schmidt, A., Trentini, D. B., Spiess, S., Fuhrmann, J., Ammerer, G., Mechtler, K., et al. (2014). Quantitative phosphoproteomics reveals the role of protein arginine phosphorylation in the bacterial stress response. Mol. Cell. Proteomics 13, 537–550. doi: 10.1074/mcp.M113.032292
Schuster, G., Owens, G. C., Cohen, Y., Ohad, I. (1984). Thylakoid polypeptide composition and light-independent phosphorylation of the chlorophyll a,b-protein in prochloron, A prokaryote exhibiting oxygenic photosynthesis. Biochim. Biophys. Acta 767, 596–605. doi: 10.1016/0005-2728(84)90061-6
Schwarz, D., Orf, I., Kopka, J., Hagemann, M. (2014). Effects of Inorganic Carbon Limitation on the Metabolome of the Synechocystis sp. PCC 6803 Mutant Defective in glnB Encoding the Central Regulator PII of Cyanobacterial C/N Acclimation. Metabolites 4, 232–247. doi: 10.3390/metabo4020232
Shalev-Malul, G., Lieman-Hurwitz, J., Viner-Mozzini, Y., Sukenik, A., Gaathon, A., Lebendiker, M., et al. (2008). An AbrB-like protein might be involved in the regulation of cylindrospermopsin production by Aphanizomenon ovalisporum. Environ. Microbiol. 10, 988–999. doi: 10.1111/j.1462-2920.2007.01519.x
Shi, L., Bischoff, K. M., Kennelly, P. J. (1999). The icfG Gene Cluster of Synechocystis sp. Strain PCC 6803 Encodes an Rsb/Spo-Like Protein Kinase, Protein Phosphatase, and Two Phosphoproteins. J. Bacteriology 181, 4761–4767. doi: 10.1128/jb.181.16.4761-4767.1999
Shi, L., Potts, M., Kennelly, P. J. (1998). The serine, threonine, and/or tyrosine-specific protein kinases and protein phosphatases of prokaryotic organisms: a family portrait. FEMS Microbiol. Rev. 22, 229–253. doi: 10.1111/j.1574-6976.1998.tb00369.x
Shibata, M., Katoh, H., Sonoda, M., Ohkawa, H., Shimoyama, M., Fukuzawa, H., et al. (2002). Genes essential to sodium-dependent bicarbonate transport in cyanobacteria: Function and phylogenetic analysis. J. Biol. Chem. 277, 18658–18664. doi: 10.1074/jbc.M112468200
Shibata, M., Ohkawa, H., Kaneko, T., Fukuzawa, H., Tabata, S., Kaplan, A., et al. (2001). Distinct constitutive and low-CO2-induced CO2 uptake systems in cyanobacteria: Genes involved and their phylogenetic relationship with homologous genes in other organisms. Proc. Natl. Acad. Sci. U.S.A. 98, 11789–11794. doi: 10.1073/pnas.191258298
Spät, P., Barske, T., Maček, B., Hagemann, M. (2021). Alterations in the CO2 availability induce alterations in the phosphoproteome of the cyanobacterium Synechocystis sp. PCC 6803. New Phytol. 231, 1123–1137. doi: 10.1111/nph.17423
Spät, P., Klotz, A., Rexroth, S., Maček, B., Forchhammer, K. (2018). Chlorosis as a developmental program in cyanobacteria: The proteomic fundament for survival and awakening. Mol. Cell. Proteomics 17, 1650–1669. doi: 10.1074/MCP.RA118.000699
Spät, P., Krauspe, V., Hess, W. R., Maček, B., Nalpas, N. (2023). Deep proteogenomics of a photosynthetic cyanobacterium. J. Proteome Res. 22, 1969–1983. doi: 10.1021/acs.jproteome.3c00065
Spät, P., Maček, B., Forchhammer, K. (2015). Phosphoproteome of the cyanobacterium Synechocystis sp. PCC 6803 and its dynamics during nitrogen starvation. Front. Microbiol. 6. doi: 10.3389/fmicb.2015.00248
Stock, A. M., Robinson, V. L., Goudreau, P. N. (2000). TWO-COMPONENT SIGNAL TRANSDUCTION. Annu. Rev. Biochem. 69, 183–215. doi: 10.1146/annurev.biochem.69.1.183www.annualreviews.org
Sun, F., Ding, Y., Ji, Q., Liang, Z., Deng, X., Wong, C. C. L., et al. (2012). Protein cysteine phosphorylation of SarA/MgrA family transcriptional regulators mediates bacterial virulence and antibiotic resistance. Proc. Natl. Acad. Sci. U.S.A. 109, 15461–15466. doi: 10.1073/pnas.1205952109
Tcherkez, G. G. B., Farquhar, G. D., Andrews, T. J. (2006). Despite slow catalysis and confused substrate specificity, all ribulose bisphosphate carboxylases may be nearly perfectly optimized. Proc. Natl. Acad. Sci. U.S.A. 103, 7246–7251. doi: 10.1073/pnas.0600605103
Toyoshima, M., Tokumaru, Y., Matsuda, F., Shimizu, H. (2020). Assessment of Protein Content and Phosphorylation Level in Synechocystis sp. PCC 6803 under Various Growth Conditions Using Quantitative Phosphoproteomic Analysis. Molecules (Basel Switzerland) 25, 3528. doi: 10.3390/molecules25163582
van Heerden, J. H., Bruggeman, F. J., Teusink, B. (2015). Multi-tasking of biosynthetic and energetic functions of glycolysis explained by supply and demand logic. BioEssays 37, 34–45. doi: 10.1002/bies.201400108
Wang, J., Chen, L., Huang, S., Liu, J., Ren, X., Tian, X., et al. (2012). RNA-seq based identification and mutant validation of gene targets related to ethanol resistance in cyanobacterial Synechocystis sp. PCC 6803. Biotechnol. Biofuels 5, 89. doi: 10.1186/1754-6834-5-89
Wang, X., Ge, H., Zhang, Y., Wang, Y., Zhang, P. (2022). Ser/Thr Protein Kinase SpkI Affects Photosynthetic Efficiency in Synechocystis sp. PCC 6803 upon Salt Stress. Life 12, 713. doi: 10.3390/life12050713
Wang, H. L., Postier, B. L., Burnap, R. L. (2004). Alterations in Global Patterns of Gene Expression in Synechocystis sp. PCC 6803 in Response to Inorganic Carbon Limitation and the Inactivation of ndhR, a LysR Family Regulator. J. Biol. Chem. 279, 5739–5751. doi: 10.1074/jbc.M311336200
Weber, A. P. M., Linka, M., Bhattacharya, D. (2006). Single, ancient origin of a plastid metabolite translocator family in Plantae from an endomembrane-derived ancestor. Eukaryotic Cell 5, 609–612. doi: 10.1128/EC.5.3.609-612.2006
Yang, M. K., Qiao, Z. X., Zhang, W. Y., Xiong, Q., Zhang, J., Li, T., et al. (2013). ). Global phosphoproteomic analysis reveals diverse functions of serine/threonine/tyrosine phosphorylation in the model cyanobacterium Synechococcus sp. strain PCC 7002. J. Proteome Res. 12, 1909–1923. doi: 10.1021/pr4000043
Zaffagnini, M., Fermani, S., Costa, A., Lemaire, S. D., Trost, P., Lindermayr, C. (2013). Plant cytoplasmic GAPDH: redox post-translational modifications and moonlighting properties. Front. Plant Sci. 4. doi: 10.3389/fpls.2013.00450
Zavřel, T., Očenášová, P., Červeny, J. (2017). Phenotypic characterization of Synechocystis sp. PCC 6803 substrains reveals differences in sensitivity to abiotic stress. PloS One 12, 1–21. doi: 10.1371/journal.pone.0189130
Zhang, C. C. (1993). A gene encoding a protein related to eukaryotic protein kinases from the filamentous heterocystous cyanobacterium Anabaena PCC 7120. Proc. Natl. Acad. Sci. U.S.A. 90, 11840–11844. Available at: https://www.pnas.org.
Zhang, C. C. (1996). Bacterial signalling involving eukaryotic-type protein kinases. Mol. Microbiol. 20, 9–15. doi: 10.1111/j.1365-2958.1996.tb02483.x
Zhang, C.-C., Gonzalez, L., Phalip, V. (1998). SURVEY AND SUMMARY Survey, analysis and genetic organization of genes encoding eukaryotic-like signaling proteins on a cyanobacterial genome. Nucleic Acids Res. 26, 3619–3625.
Zhang, C.-C., Jang, J., Sakr, S., Wang, L. (2005). Protein phosphorylation on Ser, Thr and Tyr residues in cyanobacteria. J. Mol. Microbiol. Biotechnol. 9, 154–166. doi: 10.1159/000089644
Zhang, X., Zhao, F., Guan, X., Yang, Y., Liang, C., Qin, S. (2007). Genome-wide survey of putative Serine/Threonine protein kinases in cyanobacteria. BMC Genomics 8, 395. doi: 10.1186/1471-2164-8-39
Zheng, J., Jia, Z. (2010). Structure of the bifunctional isocitrate dehydrogenase kinase/phosphatase. Nature 465, 961–965. doi: 10.1038/nature09088
Zorina, A. (2013). Eukaryotic protein kinases in cyanobacteria. Russian J. Plant Physiol. 60, 589–596. doi: 10.1134/s1021443713040195
Zorina, A. A., Bedbenov, V. S., Novikova, G. V., Panichkin, V. B., Los, D. A. (2014). Involvement of serine/threonine protein kinases in the cold stress response in the cyanobacterium Synechocystis sp. PCC 6803: Functional characterization of SpkE protein kinase. Mol. Biol. 48, 390–398. doi: 10.1134/S0026893314030212
Zorina, A. A., Novikova, G. V., Gusev, N. B., Leusenko, A. V., Los, D. A., Klychnikov, O. I. (2023). SpkH (Sll0005) from Synechocystis sp. PCC 6803 is an active Mn2+-dependent Ser kinase. Biochimie 213, 114–122. doi: 10.1016/j.biochi.2023.05.006
Keywords: Synechocystis, proteomics, kinase, mutant, environment
Citation: Barske T and Hagemann M (2025) The regulatory impact of serine/threonine-specific protein phosphorylation among cyanobacteria. Front. Plant Sci. 16:1540914. doi: 10.3389/fpls.2025.1540914
Received: 06 December 2024; Accepted: 22 January 2025;
Published: 12 February 2025.
Edited by:
Rei Narikawa, Shizuoka University, JapanReviewed by:
Amit Srivastava, The Czech Academy of Sciences, CzechiaCopyright © 2025 Barske and Hagemann. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Martin Hagemann, bWFydGluLmhhZ2VtYW5uQHVuaS1yb3N0b2NrLmRl
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.