
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Plant Sci. , 20 March 2025
Sec. Plant Symbiotic Interactions
Volume 16 - 2025 | https://doi.org/10.3389/fpls.2025.1539336
This article is part of the Research Topic Cross-Kingdom Communications Among Plants, Fungi and Bacteria: From Molecules to Ecological Factors View all 10 articles
 Yanan Quan1,2,3
Yanan Quan1,2,3 Xiuwen Gan1,2,3
Xiuwen Gan1,2,3 Shiyun Lu1,2,3
Shiyun Lu1,2,3 Xiaodong Shi1,2,3
Xiaodong Shi1,2,3 Mingsheng Bai1,2,3
Mingsheng Bai1,2,3 Yin Lin1,2,3
Yin Lin1,2,3 Yufei Gou1,2,3
Yufei Gou1,2,3 Hong Zhang1,2,3
Hong Zhang1,2,3 Xinyue Zhang1,2,3
Xinyue Zhang1,2,3 Jiayuan Wei1,2,3
Jiayuan Wei1,2,3 Tianyu Chang1,2,3
Tianyu Chang1,2,3 Jingyu Li1,2,3
Jingyu Li1,2,3 Jianli Liu1,2,3*
Jianli Liu1,2,3*Introduction: Tetraena mongolica was established in the West Ordos Region of northwest China approximately 140 million years ago. It plays an irreplaceable role in maintaining local ecosystem stability.
Methods: This study aimed to evaluate the effects of planting T. mongolica on soil nutrition and microbial communities by comparing the root zone soil (Rz_soil) and bare soil (B_soil) across three different plant communitie.
Results: The results showed that T. mongolica decreased soil pH and Na+ while increasing available potassium, soil organic matter, organic carbon, total nitrogen, and potassium. T. mongolica significantly improved the diversity indices (Sobs and Ace), as well as the richness index (Chao), of bacterial and fungal communities across three plant communities. Meanwhile, the relative abundances of Rubrobacter and norank_c_Actinobacteria in the bacterial communities declined significantly in the Rz_soil compared with the B_soil across all three plant communities. In contrast, the relative abundances of Fusarium and Penicillium were higher, whereas those of Monosporascus and Darksidea were lower in Rz_soil than in B_soil in the two plant communities. T. mongolica decreased the soil bacterial co-occurrence networks while increasing the soil fungal co-occurrence networks.
Discussion: These results provide a new perspective to understand the role of T. Mongolica in the desert ecosystems.
Soil is the foundation of terrestrial ecosystems, and serves as a crucial site for material cycling and energy conversion between biotic and abiotic environments (Lehmann et al., 2020; Creamer et al., 2022). It is a multicomponent complex characterized by intricate physical, chemical, and biological properties, providing essential growth medium and conditions, such as nutrients, water, and air, which are necessary to support plant life (Zhang et al., 2021; Hartmann and Six, 2023). Soil properties are influenced by various environmental factors including temperature, moisture, oxygen levels, and organisms (Crowther et al., 2019; Zheng et al., 2019; Patel et al., 2021; Feng et al., 2022). Plants are the most significant biological factors affecting soil structure and characteristics (de Vries et al., 2023). Plants influence several soil properties, including organic matter transformation, water cycling, and community composition through mechanisms, such as root exudates, litter decomposition, and regulation of the field microclimate. In this interaction, plants first alter soil properties and, in turn, affect plant growth, known as “plant-soil feedback” (Beckman et al., 2023; de Vries et al., 2023; Ali et al., 2024; Zou et al., 2024). Microorganisms are an important component of soil (Fierer, 2017) and are also involved in plant-soil feedback processes (Saleem et al., 2019; Semchenko et al., 2022). Microorganisms play crucial roles in soil nutrient cycling (Zhang et al., 2021). On the one hand, microorganisms can convert soil nutrients into forms that plants can utilize (Philippot et al., 2024). On the other hand, they also can decompose and mineralize plant litter and root exudates into soil components (Chen et al., 2021; Coban et al., 2022). Therefore, soil microorganisms play a bridging role in “plant-soil feedback.”
The West Ordos Region of Inner Mongolia, located in Northwest China on the eastern edge of the Asian desert, features a natural geographical landscape dominated by steppification deserts. The region is arid and experiences minimal rainfall, a significant water deficit, soil infertility, and a lack of essential nutrients. The flora primarily consists of xerophytes, superxerophytes, salt-tolerant shrubs, and semi-shrubs (Guo et al., 2024b). The West Ordos Region is also known as the “refuge” of ancient Mediterranean relict plants. Because of edge effects and paleogeography, most keystone species and dominant species of the plant community in the West Ordos region are ancient Mediterranean relict species (Li et al., 2018). Tetraena mongolica is a strong xerophytic succulent leafy deciduous shrub that belongs to the Family Zygophyllaceae. This species is a unique example of an ancient relict plant in China, often referred to as the “living fossil”. It is classified as a nationally endangered and rare protected species and represents one of the characteristic genera of the Mongolian Plateau and Central Asia (Cheng et al., 2020; Liu et al., 2023a). T. mongolica is exclusively found in western Etuoke County, Wuhai City, within the Inner Mongolia Autonomous Region, and in Shizuishan City of the Ningxia Hui Autonomous Region. As a keystone species in desert ecosystems, T. mongolica plays an irreplaceable role in maintaining the local ecosystem stability and protecting the ecological environment (Liu et al., 2023b).
Currently, research on T. mongolica has primarily focused on the physical geography of its distribution (Guo et al., 2024a), biological characteristics, eco-physiological adaptations, chemical composition (Wu et al., 2020), and genetic structure (Cheng et al., 2020; Dang et al., 2020). However, a systematic analysis of the local soil altered by T. mongolica is lacking. This study aimed to assess the effects of T. mongolica cultivation on soil nutrition and microbial communities. These findings provide a new perspective to understand the role of T. Mongolica in the desert ecosystems.
The study area is situated in the Gander Mountain Core Protection Area of T. mongolica, located in the Hainan District of Wuhai City, Inner Mongolia, China (106°87′–106°89′E, 39°52′–39°55′N). This region falls within a temperate zone and experiences a continental climate characterized by cold winters, hot summers, minimal rainfall, strong winds, sandy terrain, and high thermal energy. The landscape is predominantly harsh and desert like.
In August 2019, three distinct desert plant communities in a protected area were selected as research sites. These included the community of T. mongolica, Reaumuria songarica, Salsola passerine, and Stipa capillata (Tm_Rs_Sp_S); T. mongolica and S. capillata (Tm_S); and the community dominated by T. mongolica (Tm) (Supplementary Figure S1). A large sample plot measuring 100 × 100 m was established for each plant community type, with approximately 1.2 km separating each plot. Within each large sample plot, five small plots (10 × 10 m) were arranged in a five-point pattern. All T. mongolica plants in the small plots were selected for soil collection. After removing 3 cm of topsoil, root zone soil samples (Rz_Soil) were collected at a radial distance of 5 cm from the plants at depths of 0–20 cm in four cardinal directions: southeast, southwest, northwest, and northeast. Simultaneously, a bare soil samples (B_soil) was collected from the area closest to T. mongolica that was devoid of roots. All the samples from the same small plot were combined into a single sample. Thirty soil samples were collected at each sampling site. All samples were stored on ice in sterile zip bags until arrival at the laboratory. Each sample was divided into two portions: one was stored at -80°C for molecular biology research, and the other was air-dried and sieved to determine its physical and chemical properties.
pH was measured using a calcium chloride extraction pH-sensitive electrode. Soil organic matter (SOM) content was measured using the potassium dichromate external heating method. Organic carbon (OC) was quantified using the potassium dichromate oxidation-external heating method. Total nitrogen (TN) content was determined via Kjeldahl nitrogen determination after the elimination of concentrated sulfuric acid and hydrogen peroxide. Organic nitrogen (ON) was measured using the hydrochloric acid hydrolysis-distillation method. Ammonium-nitrogen (NH4-N) was analyzed using the indigo blue colorimetric method after extracting the potassium chloride solution. Nitrate-nitrogen (NO3-N) was determined using the dual-wavelength colorimetric method after the extraction of the potassium chloride solution. The total phosphorus (TP) content was determined using the molybdenum-antimony resistance colorimetric method after digestion with concentrated sulfuric acid and hydrogen peroxide. The available phosphorus (AP) content was determined using the molybdate blue colorimetric method after extraction with a 0.5 M NaHCO3 solution. The total potassium (TK) content was measured using sodium hydroxide melt-flame spectrophotometry, and the available potassium (AK) was evaluated using ammonium acetate extraction-flame spectrophotometry. Water-soluble Ca2+, Mg2+, Na+, and K+ were quantified using deionized water extraction-flame spectrophotometry. Carbonate (CO32-) was determined via deionized water extraction, followed by acid-alkali neutralization titration. Sulfate (SO42-) was measured using deionized water extraction-barium sulfate turbidimetry, and chloride (Cl-) was determined through deionized water extraction-silver nitrate titration. These standard methods were followed, as described by Bao (2000) and Lu (2000).
Total microbial genomic DNA was extracted from approximately 200 mg of soil using the E.Z.N.A.® soil DNA Kit (Omega Bio-tek, Norcross, GA, US. PCR amplification of bacterial 16S rRNA and fungal ITS genes was conducted using Illumina-overhang-added primer pairs targeting the bacterial V4 region (515FmodF: 5’-GTGYCAGCMGCCGCGGTAA-3’ and 806RmodR: 5’-GGACTACNVGGGTWTCTAAT-3’) (Sampson et al., 2016; Walters et al., 2016) and fungal ITS1 region (ITS1F: 5’-CTTGGTCATTTAGAGGAAGTAA-3’ and ITS2R: 5’-GCTGCGTTCTTCATCGATGC-3’) (Sampson et al., 2016).
Purified amplicons were pooled in equimolar amounts and paired-end sequenced on an Illumina MiSeq PE300 platform (Illumina, San Diego, CA, USA) following the standard protocols established by Majorbio Bio-Pharm Technology Co. Ltd. (Shanghai, China). The sequence reads were deposited in the NCBI Sequence Read Archive (SRA) database under accession numbers PRJNA1183314 and PRJNA1183552.
Bacterial and fungal OTUs were annotated using the Greengenes database (v13.8) (DeSantis et al., 2006) and the UNITE database (v8.2) (Nilsson et al., 2019), respectively, and reads that were not classified as bacterial or fungal were excluded. Bioinformatic analysis of soil bacteria and fungi was conducted using the Majorbio Cloud platform (https://cloud.majorbio.com). Microbial alpha diversity metrics, including Sobs, Shannon index, Simpson index, Heip evenness, Ace, and Chao1 richness, were estimated using Mothur v1.30.1 (Schloss et al., 2009). Both bacterial and fungal beta diversity analyses were performed using non-metric multidimensional scaling analysis (NMDS) with QIIME 2 (Bolyen et al., 2019) to calculate the distance matrix and the Vegan v2.5-3 package for analysis and plotting. Student’s t-test was used to evaluate differences in the relative abundance of microbial taxonomic groups. Venn diagram was created using the Venn diagram package in R (v1.6.20). Co-occurrence networks were constructed to explore internal community relationships across the samples (Barberán et al., 2012). A correlation between two nodes was considered statistically robust if Spearman’s correlation coefficient was >0.6 or <-0.6, with a P-value of <0.05. The functions of bacterial communities were predicted using the FAPROTAX tool (Louca et al., 2016). The functions of fungal communities were predicted using FUNGuild (http://www.funguild.org/). Student’s t-test was used to evaluate the differences in the functions of the microbial communities. Canonical correspondence analysis (CCA) and redundancy analysis (RDA) were performed using the Vegan v2.5-3 package to investigate the effects of soil physicochemical properties on the structure of the soil bacterial and fungal communities. The non-parametric permutational multivariate analysis of variance (PERMANOVA) test was used to assess the percentage of variation explained by the treatment, along with its statistical significance, using the vegan v2.5-3 package.
Various physicochemical characteristics of the samples were measured in the Rz_soil of T. mongolica in the three different plant communities (Table 1). In these communities, the soil pH in the T. mongolica Rz_soil decreased, whereas AK increased when compared with B_soil, which was sampled from areas without plant roots closest to T. mongolica plants. In both plant communities, SOM, OC, TN, nitrogen-to-phosphorus ratio (N/P), and K+ increased with the establishment of the T. mongolica plantations. Conversely, NH4-N and Na+ levels decreased in the T. mongolica plantations. NO3-N, Mg2+, and CO32- levels increased in T. mongolica plantations in only one plant community. ON, AP, and organophosphorus (OP) decreased in T. mongolica plantations in only one community. Across all three plant communities, the T. mongolica plantation had no significant effects on TP, TK, carbon-to-nitrogen ratio (C/N), carbon-to-phosphorus ratio (C/P), or Cl-. Ca2+ and SO42- levels increased in one community, decreased in the other, and showed no significant differences in the third community.

Table 1. Soil physicochemical properties in Rz_soil and B_soil in three plant communities in which T. mongolica is the dominant or keystone specie.
The indices of α-diversity, including Sobs, Shannon, Chao, Pielou, and Coverage, were calculated to quantify the diversity, richness, evenness, and sequencing depth of the microbial communities in the three plant communities. The coverage index for all the microbial communities was approximately 0.9, indicating that the sequencing capability was acceptable (Table 2). T. mongolica significantly improved the Sobs and Ace diversity indices, as well as the Chao richness index, of bacterial and fungal communities in the Rz_soil compared with the B_soil. Shannon diversity and Heip evenness indices of the bacterial communities in the Rz_soil were significantly higher in the two plant communities, whereas the fungal communities showed significantly higher indices in only one plant community. However, there were no significant effects on the Simpson diversity index for either the bacterial or fungal communities across the three plant communities.
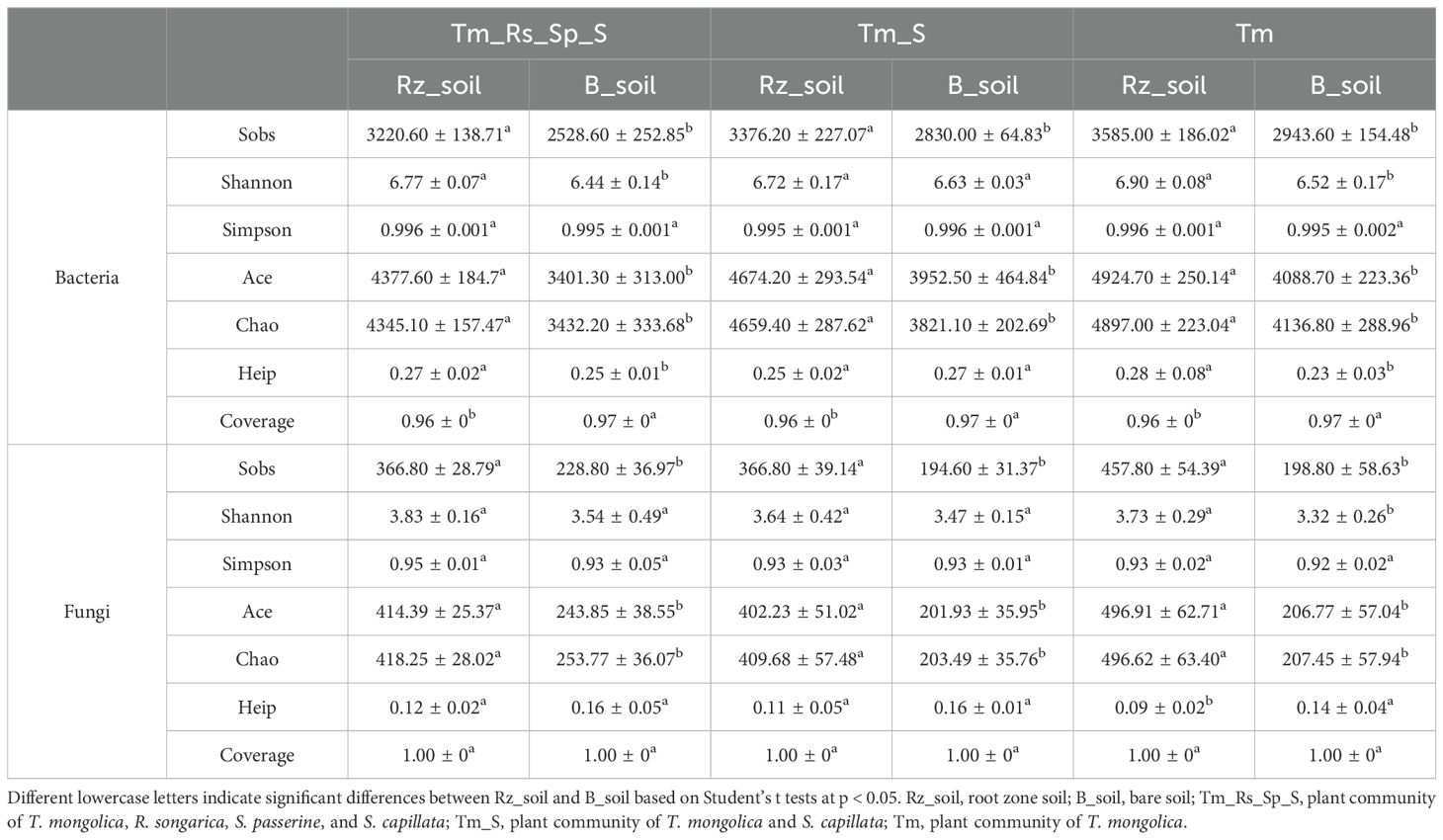
Table 2. α-diversity indices of bacterial and fungal communities in Rz_soil and B_soil in three plant communities in which T. mongolica is the dominant or keystone specie.
All samples from Rz_soil and B_soil were positioned in distinct regions along the coordinate axis in the NMDS figure based on the Bray-Curtis method (stress <0.2). The bacterial and fungal communities in Rz_soil were separated from those in B_soil along the x-axis across the three plant communities (Figure 1). This finding demonstrates that T. mongolica plantations significantly altered the bacterial and fungal communities. However, the bacterial communities in the two soil types and the fungal communities in the Rz_soil were not distinctly different, and only the fungal communities in the B_soil exhibited a clear differentiation. This suggests that although the fungal communities in B_soil were distinct, the bacterial communities in B_soil showed less variation. In the T. mongolica plantation, both the bacterial and fungal communities in the Rz_soil exhibited minimal differences.
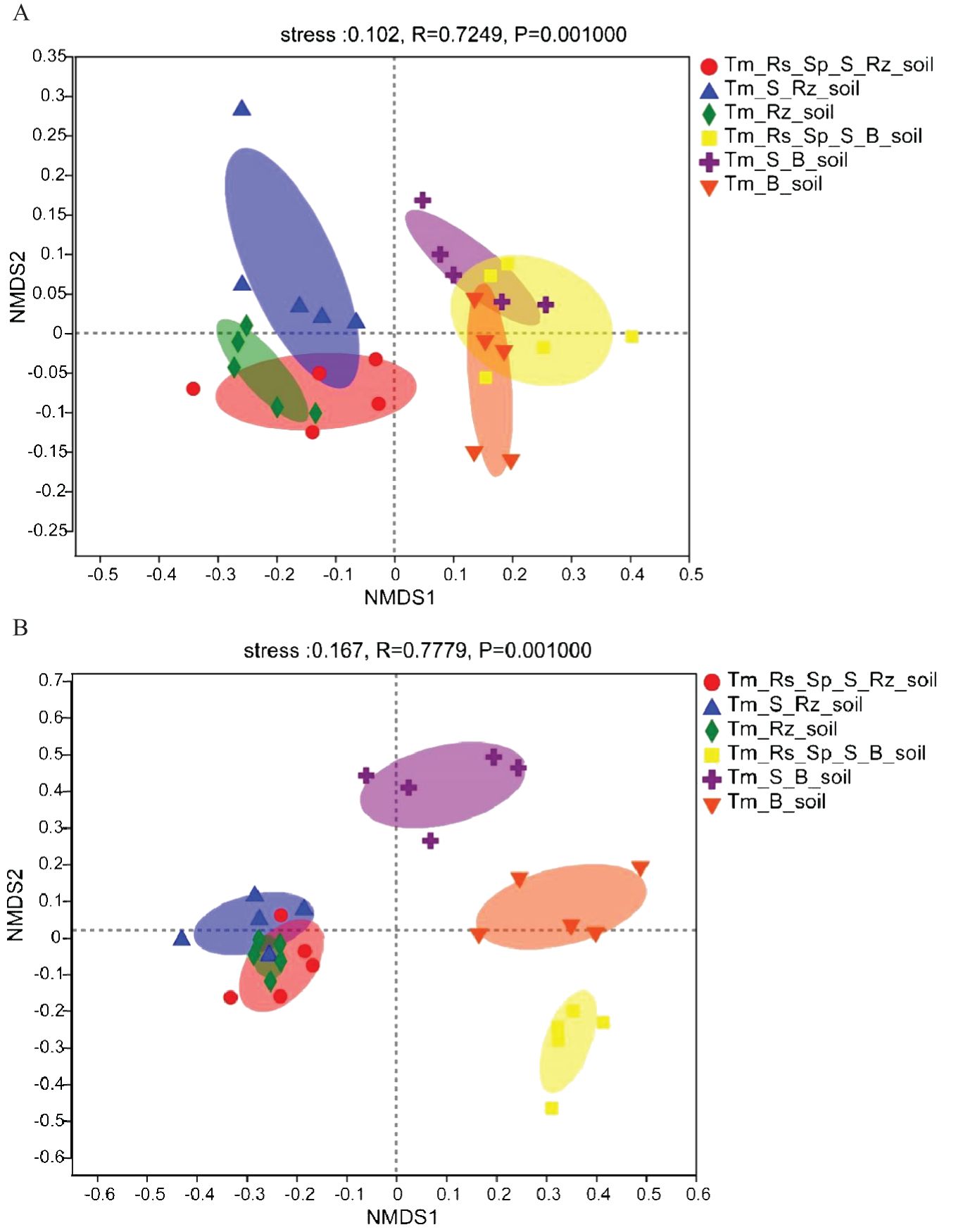
Figure 1. Non-metric multidimensional scaling (NMDS) analysis and plot of bacterial and fungal communities in Rz_soil and B_soil in three plant communities based on the bray–curtis distance. (A) The distribution of bacterial communities in Rz_soil and B_soil in three plant communities. (B) The distribution of fungal communities in Rz_soil and B_soil in three plant communities. Tm_Rs_Sp_S_Rz_soil, root zone soil in plant community of T. mongolica, R. songarica, S. passerine, and S. capillata; Tm_Rs_Sp_S_B_soil, bare soil in plant community of T. mongolica, R. songarica, S. passerine, and S. capillata; Tm_S_Rz_soil, root zone soil in plant community of T. mongolica and S. capillata; Tm_S_B_soil, bare soil in plant community of T. mongolica and S. capillata; Tm_Rz_soil, root zone soil in plant community of T. mongolica; Tm_B_soil, bare soil in plant community of T. mongolica.
T. mongolica did not significantly affect the number and diversity of bacterial and fungal phyla with a relative abundance of >1% in the B_soil and Rz_soil across the three plant communities. Actinobacteria was the dominant phylum in all samples. The relative abundances of Proteobacteria, Chloroflexi, Acidobacteria, Planctomycetes, Gemmatimonadetes, Verrucomicrobia, and Bacteroidetes were ranked from the second-most dominant phylum to the subsequent phyla (Supplementary Figure S2A). However, the relative abundance of these phyla varied between B_soil and Rz_soil in the three plant communities. Notably, the relative abundance of Bacteroidetes was significantly higher in the Rz_soil than in the B_soil across all three plant communities. Conversely, the relative abundance of Actinobacteria decreased, whereas the relative abundance of Proteobacteria increased significantly between Rz_soil and B_soil in Tm_Rs_Sp_S and Tm plant communities. These changes were not significant in the Tm_S community of the T. mongolica plantation. In addition, the relative abundance of Planctomycetes was greater in Rz_soil than in B_soil; however, this was only observed in the Tm_Rs_Sp_S plant community. In contrast, the relative abundance of Gemmatimonadetes was lower in Rz_soil than in B_soil, specifically in the Tm_S plant community (Supplementary Figures S2B–D).
Ascomycota was the dominant phylum in all samples. Basidiomycota, unclassified_k:fungi, and Mortierellomycota exhibited higher relative abundance (Supplementary Figure S3A). The relative abundance of phyla also varied between B_soil and Rz_soil across the three plant communities. Specifically, the relative abundance of Ascomycota increased, whereas the relative abundance of unclassified_k: fungi decreased significantly between Rz_soil and B_soil in the plant communities Tm_Rs_Sp_S and Tm. However, this change was not significant in the Tm_S plant community of the T. mongolica plantation. The relative abundances of Glomeromycota and Calcarisporiellomycota were lower in Rz_soil than in B_soil; however, this was only observed in the plant communities Tm_S and Tm (Supplementary Figures S3B–D).
T. mongolica did not alter the dominant bacterial genera in the Rz_soil and B_soil across the three plant communities. Rubrobacter and norank_c:Actinobacteria were the dominant genera in all the soil samples from the three plant communities. However, the relative abundances of these two genera also changed between the B_soil and Rz_soil (Figure 2A). The relative abundances of Rubrobacter and norank_c:Actinobacteria declined significantly in the Rz_soil compared with those in the B_soil across the three plant communities (Figures 2B–D).
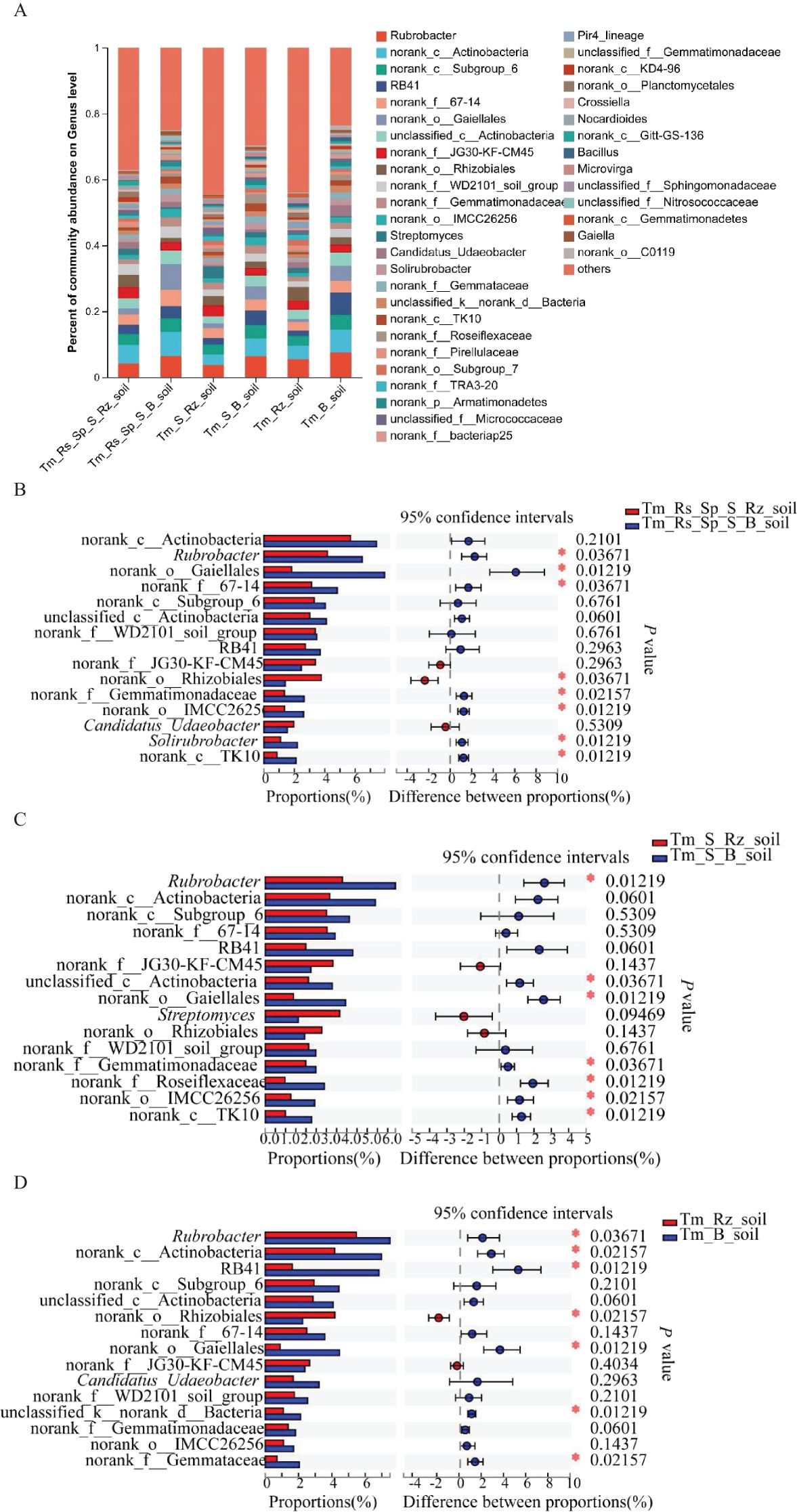
Figure 2. Relative abundance and differences of bacteria at genus level in Rz_soil and B_soil in three plant communities. (A) Genus-level bacterial communities’ composition. (B–D) Different bacterial genera in top 15 genera between B_soil and Rz_soil in the three plant communities. Tm_Rs_Sp_S_Rz_soil, root zone soil in plant community of T. mongolica, R. songarica, S. passerine, and S. capillata; Tm_Rs_Sp_S_B_soil, bare soil in plant community of T. mongolica, R. songarica, S. passerine, and S. capillata; Tm_S_Rz_soil, root zone soil in plant community of T. mongolica and S. capillata; Tm_S_B_soil, bare soil in plant community of T. mongolica and S. capillata; Tm_Rz_soil, root zone soil in plant community of T. mongolica; Tm_B_soil, bare soil in plant community of T. mongolica. * indicates a significant difference between Rz_soil and B_soil based on Student’s t tests at p < 0.05; ** indicates a significant difference at p < 0.01.
T. mongolica altered the dominant fungal genera in the Rz_soil and B_soil across three plant communities. Unclassified p:Ascomycota was the dominant genus in B_soil in Tm_Rs_Sp_S and Tm_S plant communities. Unclassified f_Ceratobasidiaceae emerged as the dominant genus in the B_soil of the Tm plant community. Fusarium was the predominant genus in all Rz_soils of the three plant communities (Figure 3A). The relative abundances of Fusarium, unclassified f:Pleosporales_fam_Incertae_sedis, unclassified f:Nectriaceae, and unclassified o:Hypocreales were significantly higher in the Rz_soil than in the B_soil in the Tm_Rs_Sp_S plant community. Conversely, the relative abundances of unclassified p:Ascomycota, Monosporascus, unclassified f_Ceratobasidiaceae, unclassified k:Fungi, and Darksidea were lower in the Rz_soil than in the B_soil. In the Tm_S plant community, the relative abundances of Penicillium, Acremonium, unclassified f:Pleosporales_fam_Incertae_sedis, and unclassified o:Hypocreales were also higher in the Rz_soil than in the B_soil.
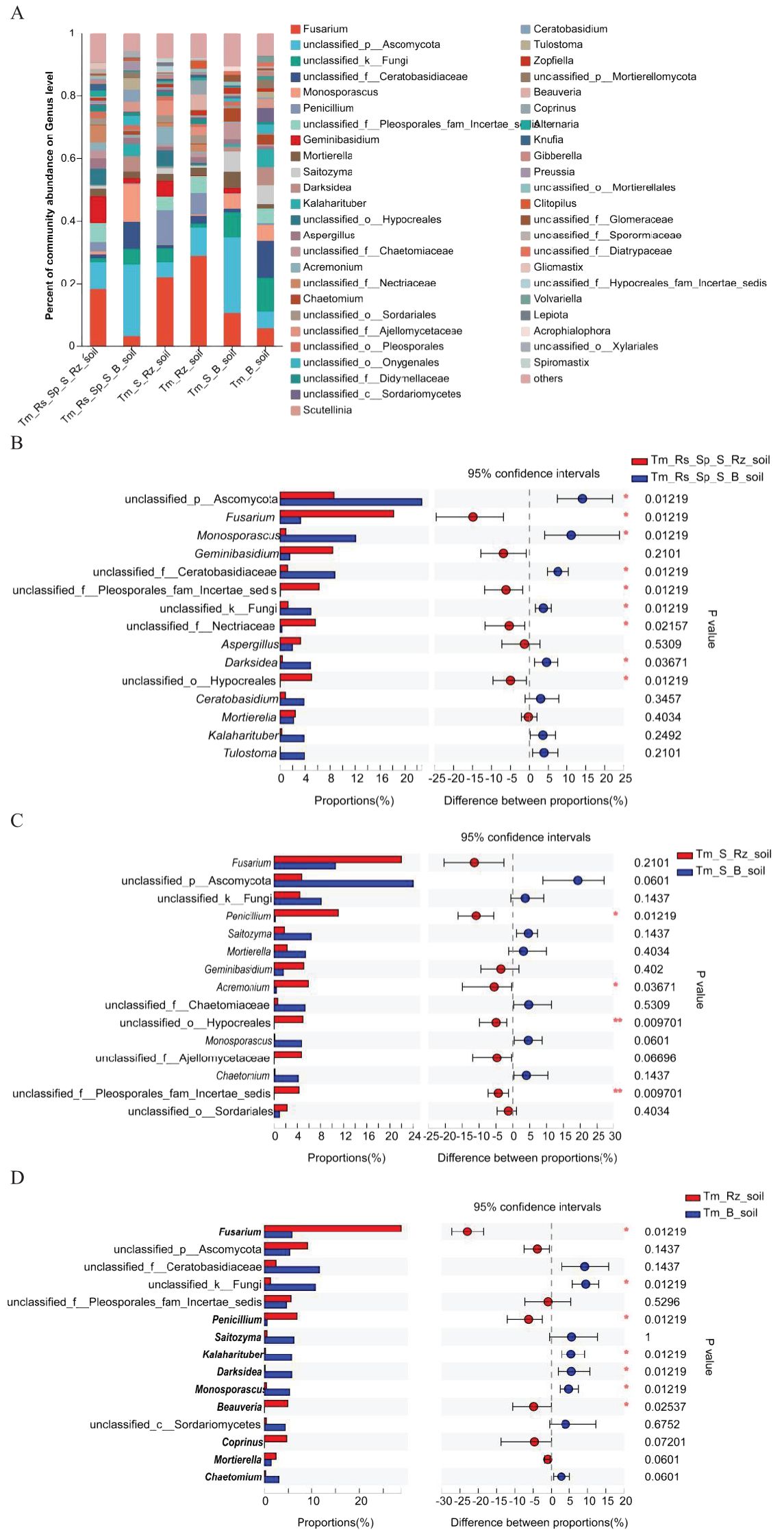
Figure 3. Relative abundance and differences of fungi at genus level in Rz_soil and B_soil in three plant communities. (A) Genus-level fungal communities’ composition. (B–D) Different fungal genera in top 15 genera between B_soil and Rz_soil from the three plant communities. Tm_Rs_Sp_S_Rz_soil, root zone soil in plant community of T. mongolica, R. songarica, S. passerine, and S. capillata; Tm_Rs_Sp_S_B_soil, bare soil in plant community of T. mongolica, R. songarica, S. passerine, and S. capillata; Tm_S_Rz_soil, root zone soil in plant community of T. mongolica and S. capillata; Tm_S_B_soil, bare soil in plant community of T. mongolica and S. capillata; Tm_Rz_soil, root zone soil in plant community of T. mongolica; Tm_B_soil, bare soil in plant community of T. mongolica. * indicates a significant difference between Rz_soil and B_soil based on Student’s t tests at p < 0.05; ** indicates a significant difference at p < 0.01.
The relative abundances of Fusarium, Penicillium, and Beauveria were higher, whereas those of unclassified_k:Fungi, Kalaharituber, Monosporascus, and Darksidea were lower in Rz_soil than in B_soil from the Tm plant community (Figures 3B–D). At the phylum level, the relative abundance of Ascomycota increased, whereas that of unclassified_k _fungi decreased significantly in the Rz_soil and B_soil across the two plant communities. At the genus level, the relative abundances of Fusarium and Penicillium were higher, whereas those of Monosporascus and Darksidea were lower in the Rz_soil than in the B_soil in both plant communities.
In all three plant communities, the specific OTUs of bacteria and fungi in the Rz_soils were more abundant than those in the B_soils. Thus, the T. mongolica plantation enhanced the OTUs in the Rz_soil compared with the B_soil. In the bacterial communities, OTUs were more common than specific OTUs in both Rz_soil and B_soil. However, in the fungal communities, common OTUs were higher in the B_soil and lower in the Rz_soil (Supplementary Figure S4).
Network analysis results indicated that the number of nodes, total edges, average degree, and density of the bacterial co-occurrence network in T. mongolica Rz_soil were lower than those in B_soil within Tm_Rs_Sp_S and Tm_S plant communities. However, in the Tm plant community, the co-occurrence network parameters in T. mongolica Rz_soil were higher than those in the B_soil. In contrast, for the fungal co-occurrence network, the parameters in T. mongolica Rz_soil exceeded those in B_soil across all three plant communities (Figures 4, 5; Supplementary Table S1). For the bacterial co-occurrence network, the percentage of negative edges was similar to that of the positive edges. However, for the fungal co-occurrence network, the percentage of negative edges was considerably lower than that of positive edges. Therefore, T. mongolica appeared to decrease soil bacterial co-occurrence networks and enhance soil fungal co-occurrence networks.
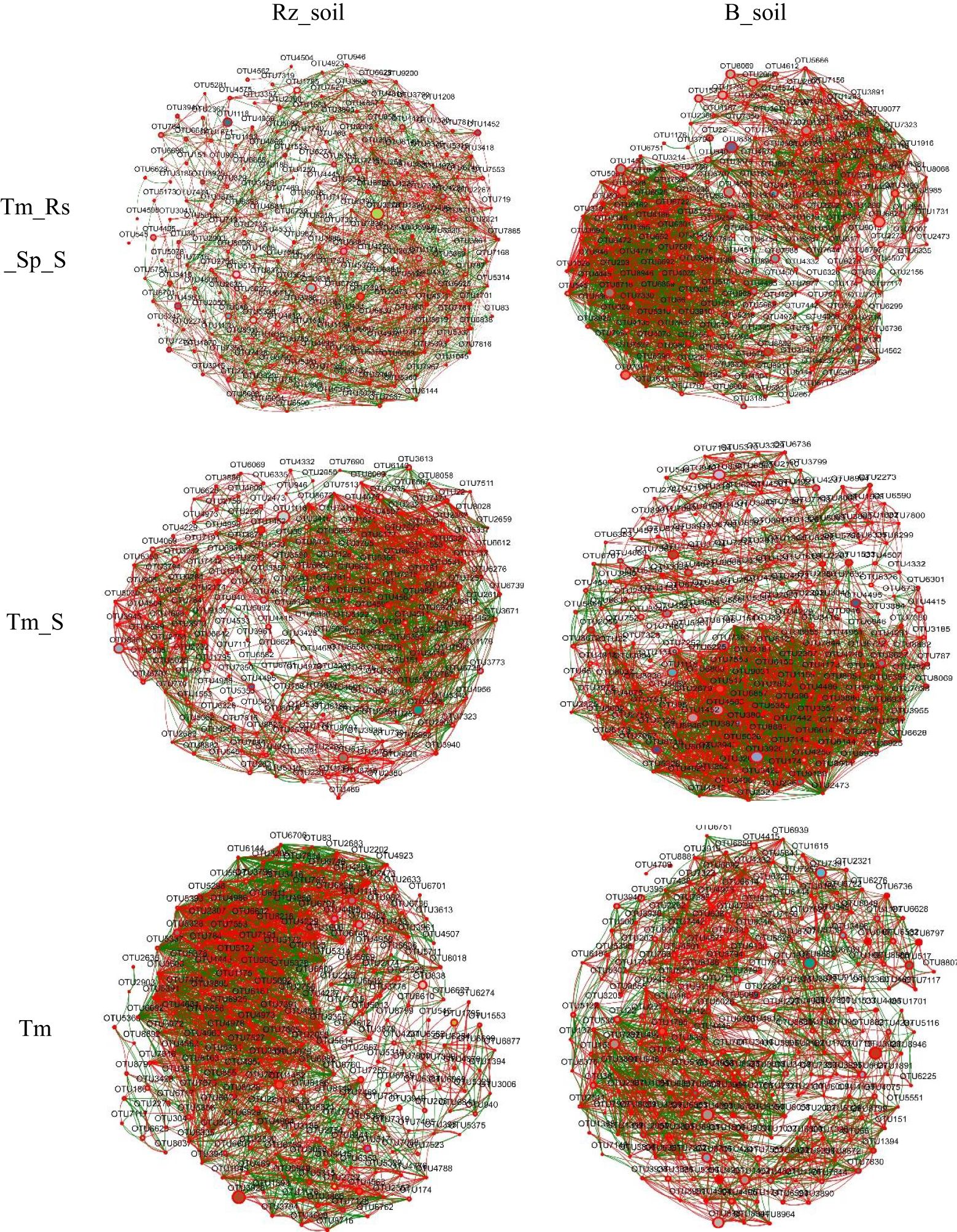
Figure 4. Bacterial co-occurrence networks between B_soil and Rz_soil in the three plant communities based on Spearman’s correlation coefficient (r) (|r| ≥ 0.5, p <0.05). The red line indicates positive correlation and the green line indicates negative correction. Rz_soil, root zone soil; B_soil, bare soil; Tm_Rs_Sp_S, plant community of T. mongolica, R. songarica, S. passerine, and S. capillata; Tm_S, plant community of T. mongolica and S. capillata; Tm, plant community of T. mongolica.
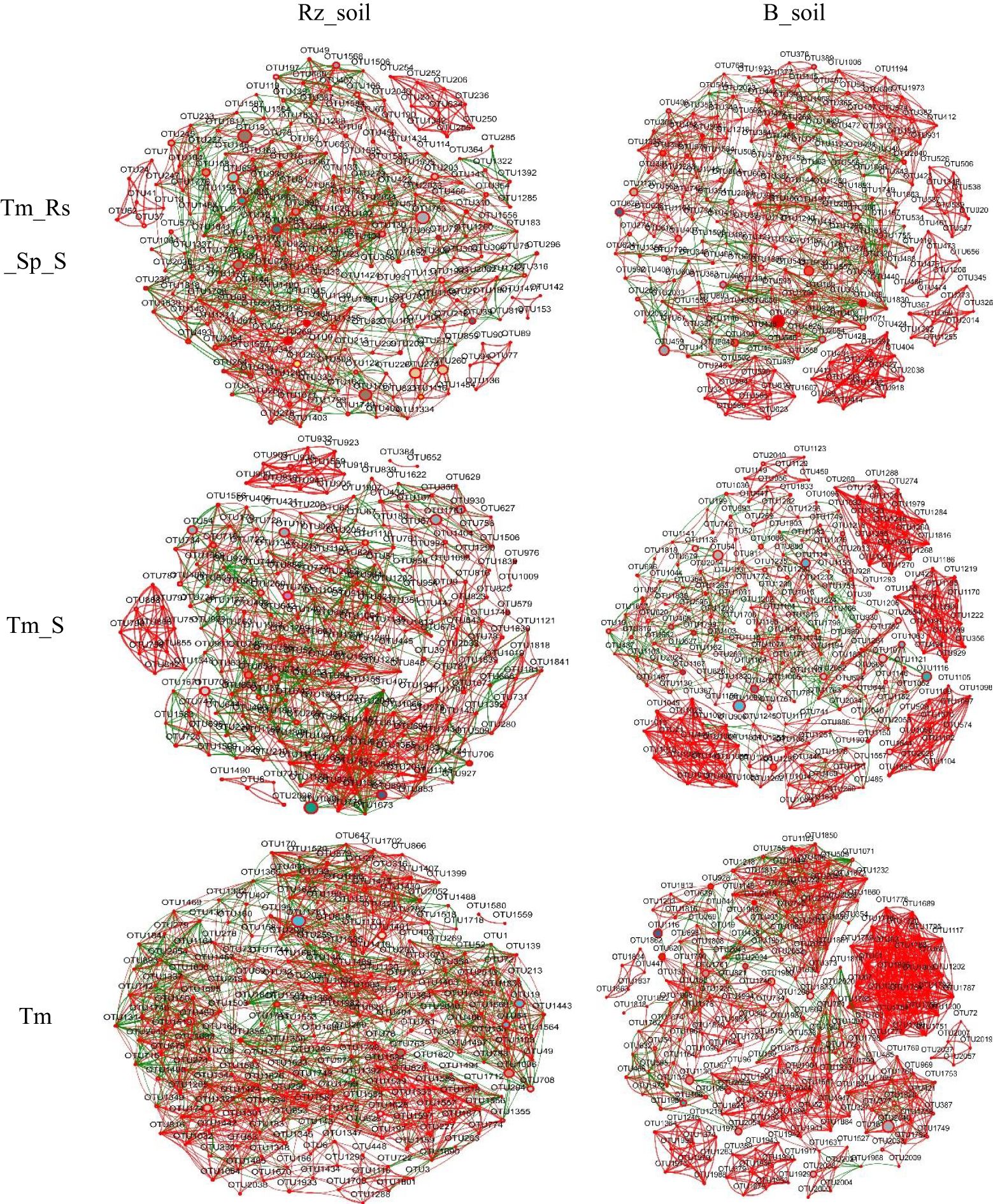
Figure 5. Fungal co-occurrence networks between B_soil and Rz_soil in the three plant communities based on Spearman’s correlation coefficient (r) (|r| ≥ 0.5, p <0.05). The red line indicates positive correlation and the green line indicates negative correction. Rz_soil, root zone soil; B_soil, bare soil; Tm_Rs_Sp_S, plant community of T. mongolica, R. songarica, S. passerine, and S. capillata; Tm_S, plant community of T. mongolica and S. capillata; Tm, plant community of T. mongolica.
Functional roles of the bacterial community were predicted using the FAPROTAX tool (Figure 6 A). T. mongolica enhanced the functional capabilities of bacterial communities, including denitrification (nitrate respiration, nitrate denitrification, nitrite denitrification, and nitrous oxide denitrification), ureolysis, dark hydrogen oxidation, methylotrophy (methylotrophy and methanol oxidation), organic matter decomposition (xylanolysis, chitinolysis, and cellulolysis), aromatic compound degradation, and photoconversion (photoheterotrophy, anoxygenic photoautotrophy, and anoxygenic photoautotrophy S oxidizing) (Figures 6B–D). Consequently, T. mongolica facilitated soil functions related to energy acquisition through denitrification, methylotrophy, hydrogen oxidation, and photoconversion, while also being influenced by organic matter decomposition, aromatic compound degradation, and photoconversion.
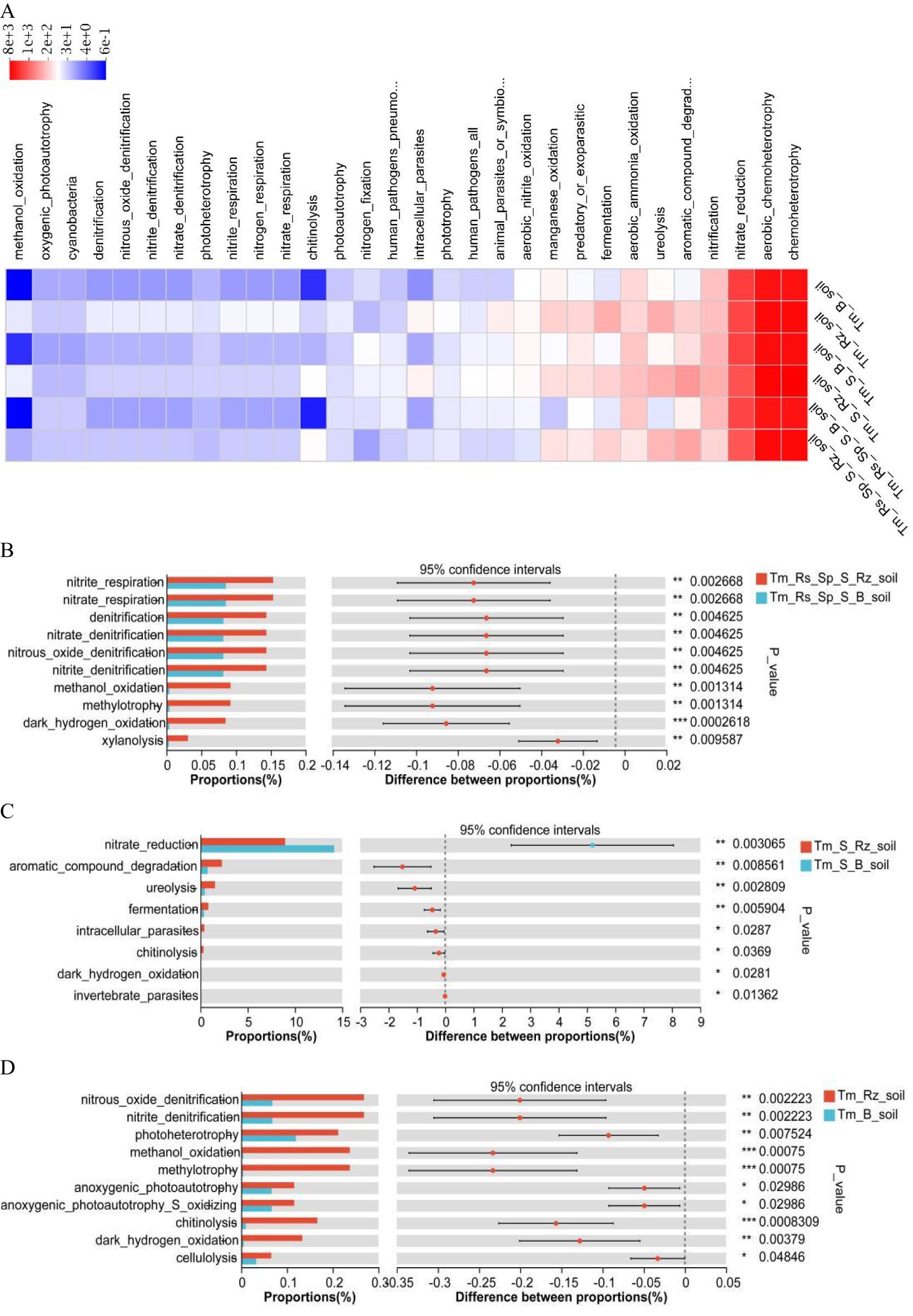
Figure 6. Bacterial community function predicted using the FAPROTAX tool in Rz_soil and B_soil in three plant communities. (A) Bacterial community function. (B–D) Bacterial community function in B_soil and Rz_soil in the three plant communities. Tm_Rs_Sp_S_Rz_soil, root zone soil in plant community of T. mongolica, R. songarica, S. passerine, and S. capillata; Tm_Rs_Sp_S_B_soil, bare soil in plant community of T. mongolica, R. songarica, S. passerine, and S. capillata; Tm_S_Rz_soil, root zone soil in plant community of T. mongolica and S. capillata; Tm_S_B_soil, bare soil in plant community of T. mongolica and S. capillata; Tm_Rz_soil, root zone soil in plant community of T. mongolica; Tm_B_soil, bare soil in plant community of T. mongolica. * indicates a significant difference between Rz_soil and B_soil based on Student’s t tests at p < 0.05; ** indicates a significant difference at p < 0.01; *** indicates a significant difference based on Student’s t tests at p < 0.001; **** indicates a significant difference based on Student’s t tests at p < 0.0001.
The function of the fungal community was predicted using the FUNGuild tool. T. mongolica increased the relative abundance of undefined saprotrophs, animal pathogens, endophytes, lichen parasites, plant pathogens, soil saprotrophs, and wood saprotrophs whereas decreasing the relative abundance of plant pathogens within fungal communities. Thus, T. mongolica promoted the growth of saprotrophic fungi and reduced the prevalence of pathogenic fungi (Figure 7A). However, these changes were not significantly different between Rz_soil and B_soil across the three plant communities (Figures 7B–D).
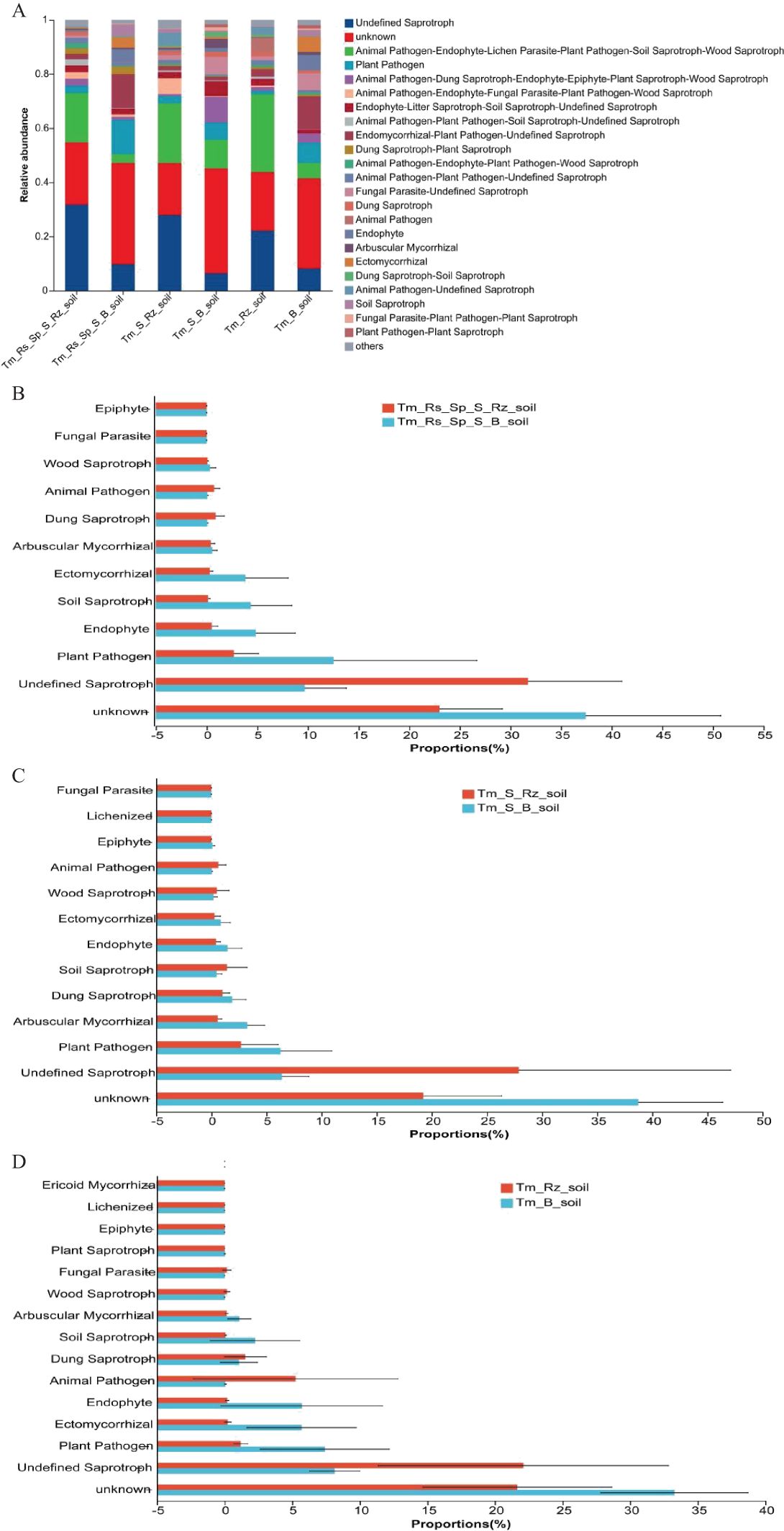
Figure 7. Fungal community function predicted using the FunGuild tool in Rz_soil and B_soil in three plant communities. (A) Fungal community function. (B–D) Fungal community function in B_soil and Rz_soil in the three plant communities. Error bars represent standard deviations (n = 5). Tm_Rs_Sp_S_Rz_soil, root zone soil in plant community of T. mongolica, R. songarica, S. passerine, and S. capillata; Tm_Rs_Sp_S_B_soil, bare soil in plant community of T. mongolica, R. songarica, S. passerine, and S. capillata; Tm_S_Rz_soil, root zone soil in plant community of T. mongolica and S. capillata; Tm_S_B_soil, bare soil in plant community of T. mongolica and S. capillata; Tm_Rz_soil, root zone soil in plant community of T. mongolica; Tm_B_soil, bare soil in plant community of T. mongolica.
The effects of soil chemical properties on the bacterial and fungal communities were assessed using CCA and RDA. Among the 19 chemical properties examined, pH, TK, NH4-N, OP, AK, and K+ had a more significant effect on the bacterial community than the other properties. Specifically, pH, TK, NH4-N, and OP were positively correlated with B_soil whereas AK and K were negatively correlated with Rz_soil. OM, pH, TK, TN, AK, AP, OC, and K+ significantly affected the fungal community. pH was the most influential factor, showing a positive correlation with B_soil and a negative correlation with Rz_soil (Figure 8).

Figure 8. Canonical correspondence analysis (CCA) or redundancy analysis (RDA) of bacterial and fungal communities constrained by soil physicochemical properties. (A) RDA of the bacterial community with soil physicochemical properties. (B) CCA of the fungal community with soil physicochemical properties. SOM, soil organic matter; OC, organic carbon; TN, total nitrogen; ON, organic nitrogen; NH4-N, ammonium-nitrogen; NO3-N, nitrate-nitrogen; TP, total phosphorus; AP, available phosphorus; OP, organic phosphorus; TK, total potassium; AK, available potassium; Ca2+, calcium ion; Mg2+, magnesium ion; Na+, sodium ion; K+, potassium ion; CO32-, carbonate; SO42-, sulfate; Cl-, chloride. Tm_Rs_Sp_S_Rz_soil, root zone soil in plant community of T. mongolica, R. songarica, S. passerine, and S. capillata; Tm_Rs_Sp_S_B_soil, bare soil in plant community of T. mongolica, R. songarica, S. passerine, and S. capillata; Tm_S_Rz_soil, root zone soil in plant community of T. mongolica and S. capillata; Tm_S_B_soil, bare soil in plant community of T. mongolica and S. capillata; Tm_Rz_soil, root zone soil in plant community of T. mongolica; Tm_B_soil, bare soil in plant community of T. mongolica.
Plants have a significant influence on the soil. This influence is multifaceted, encompassing the enhancement of the physical, chemical, and biological properties of the soil as well as the cycling and utilization of nutrients and soil microbes. Collectively, these factors promote the stability and sustainable development of ecosystems (Kranz et al., 2020; Liu et al., 2020; Furey and Tilman, 2021). This study demonstrated that T. mongolica reduced soil pH by decreasing Na+ and K+ content, increased AK, SOM, OC, and TN in desert soil. As a keystone species in desert communities, T. mongolica plays an irreplaceable role in maintaining local ecosystem stability and protecting the ecological environment through “plant-soil feedback.”
During plant growth, both acidic and alkaline substances are released, which can influence the acidity or alkalinity of soil. For instance, certain plants release organic acids during their growth process, resulting in increased soil acidity, whereas other plants may enhance soil alkalinity through root exudates (Tibbett et al., 2019). Yan et al. (2020) observed that tea plantations contribute to soil acidification, with pH levels decreasing by 0.47 to 1.43. In the fruit and vegetable systems, the pH decreased by 0.40 to 1.08, and in the cereal systems, it decreased by 0.30 to 0.89. Zhou et al. (2017) reported the pH was 4.58 in 78-year-old forest plantations of slash pine, 5.74 in Hoop pine, 6.01 in Kauri pine, and 4.49 in Eucalyptus. Jin et al. (2022) demonstrated that hickory plantations increased the pH of relatively acidic soils, but decreased the pH of relatively alkaline soils. The T. mongolica plantation also reduced the pH by 0.5 across the three plant communities. The environment in which T. mongolica thrives is the irrigation area in the middle and upper reaches of the Yellow River, characterized by long-term dryness, minimal rainfall, and high evaporation rates. Soil salinity is primarily composed of sulfates and chlorides. An arid climate and improper irrigation practices have led to the secondary salinization of the soil. The concentrations of water-soluble neutral salts, such as NaCl and Na2SO4, in the soil exceed 0.1%, whereas the concentrations of alkaline salts, such as Na2CO3 and NaHCO3, exceed 5% (Li et al., 2023). Consequently, the soil is classified as severely saline-alkali. Our findings indicate that T. mongolica significantly decreased soil salinity (Table 1). Na+ concentration decreased significantly in the T. mongolica plantation across both plant communities. However, the concentrations of Cl-, SO42-, and CO32- were not reduced. Therefore, T. mongolica lowered soil pH by decreasing Na+ concentrations.
Plants can loosen soil, increase soil porosity, reduce soil compaction, and enhance soil physical properties via root activity. Plant residues and root exudates can increase the organic matter content of soil (Oleghe et al., 2019). Changes in soil acidity or alkalinity can influence the solubility and availability of mineral elements, thereby affecting their absorption and utilization by plants (Zhao et al., 2021). Our findings indicate that T. mongolica increased SOM and OC in the Rz_soil of T. mongolica compared with B_soil. We hypothesized that T. mongolica enhances C content in the soil through root sediments, root secretions, and litter. TN also increased; however, ON and NH4-N levels decreased. We speculate that T. mongolica contributes to the N content in the soil through nitrogen fixation and recovery. In addition, T. mongolica appeared to accelerate the decomposition of ON and absorption of NH4-N; however, it did not affect TP or AP transformation. In only one plant community was P absorption enhanced, leading to reduced AP levels. Although, T. mongolica did not affect TK, it increased AK by accelerating K release.
The organic matter produced by the T. mongolica plantation not only provides abundant carbon and energy sources for soil microorganisms but also enhances soil structure and increases soil biodiversity. In this study, we found that the bacterial and fungal communities in Rz_soil were distinct from those in B_soil across the three plant communities within the T. mongolica plantation. T. mongolica significantly improved the diversity index (Sobs) and the richness index (Chao) of both bacterial and fungal communities in the three plant communities. In this study, Rubrobacter and norank_c_Actinobacteria were the dominant genera across all the soil samples collected from the three plant communities. The presence of T. mongolica did not alter the dominant bacterial genera in the Rz_soil or B_soil of these communities. However, T. mongolica plantations affected the relative abundances of certain genera. Specifically, the relative abundance of Rubrobacter and norank_c_Actinobacteria significantly declined in the Rz_soil compared with that in the B_soil across the three plant communities. Both Rubrobacter and norank_c_Actinobacteria belong to the phylum Actinobacteria, which is known for its potential to resist plant diseases, promote plant growth, and enhance salt tolerance in plants (Passari et al., 2019; Xiong et al., 2019). The small perennial shrub T. mongolica decreases the abundance of beneficial bacteria in long-term plantations, a phenomenon commonly observed in crop cultivation (Schlatter et al., 2017; Lee et al., 2021).
In this study, we found that the unclassified phylum Ascomycota was the dominant genus in B_soils in Tm_Rs_Sp_S and Tm_S. The unclassified family Ceratobasidiaceae emerged as the dominant genus in the B_soil from the Tm plant community. Fusarium was the predominant genus in all Rz soils across all three plant communities. Fusarium is a potential fungal pathogen (van Agtmaal et al., 2017). The increased abundance of Fusarium, a potential plant fungal pathogen, requires the recruitment of antagonistic bacteria to maintain plant health. Additionally, T. mongolica altered the relative abundance of the dominant bacterial genera Rubrobacter and norank_c_Actinobacteria in both Rz_soils and B_soils in the three plant communities. The relative abundances of Rubrobacter and norank_c_Actinobacteria were significantly lower in Rz_soil than in B_soil across the three plant communities. Consequently, T. mongolica poses a high risk for root disease outbreaks.
The high-throughput sequencing technology employed in this study is capable of delineating the species and relative abundance of microbial community structure. However, it is crucial to acknowledge that relative abundance alone may not provide a comprehensive reflection of microbial quantities, particularly when assessing microbial community. To address this limitation, absolute abundance can be suggested. Jiang et al. (2019) observed that the relative abundance of Proteobacteria remained statistically unchanged in fertilizer application experiments conducted on tomato crops in coastal saline-alkali soil. Nonetheless, when absolute abundance quantification methods can be utilized, a significant escalation in the absolute abundance of Proteobacteria was detected. Similar findings have been documented in the studies by Yang et al. (2018) and Lou et al. (2018). Consequently, absolute quantitation of microbiota abundance is essential.
Methods of absolute quantitation of microbiota abundance encompass a variety of techniques, including the use of spike-in bacteria as documented by Stammler et al. (2016), the quantification of bacterial DNA through Flow Cytometry (FCM) as outlined by Vandeputte et al. (2017), and the integrated high-throughput absolute abundance quantification (iHAAQ) approach, which merges high-throughput sequencing with quantitative Polymerase Chain Reaction (qPCR), as introduced Lou et al. (2018). Additionally, Tkacz et al. (2018) presented an absolute quantitation method utilizing synthetic spike DNA. Guo et al. (2020) employed host-associated quantitative abundance profiling (HA-QAP) to delineate variations in the microbial load within the root microbiome. Additionally, Zhang et al. (2022) assessed two absolute microbiome profiling (AMP) methods in soil microbiota quantitative research, spike-AMP and qPCR-AMP, concluding that qPCR-AMP is superior for quantitative assessment of soil microbial communities. Wang et al. (2020) proposed an “amplification-selection” model for rhizosphere microbiome assembly, which utilizes synthetic chimeric spikes in plasmids for microbiome profiling. They hypothesized that plant root exudates can ‘feed’ soil microorganisms, nutrition-poor bulk soil is seen as “countryside”, while rhizosphere soil is as “metropolis”, where microbial communities can find more opportunities for growth and reproduction. The microorganisms in rhizosphere soil undergo amplification prior to the selection by the root system.
In this study, we observed T. mongolica notably enhanced the diversity indices (Sobs and Ace), as well as the richness index (Chao), of both bacterial and fungal communities across three plant communities. Moreover, the relative abundance of Bacteroidetes and Ascomycota in the Rz_soils was significantly higher than that in the B_soils. These findings aligned with “two-step or multiple-step selection” model, as inferred from the relative abundance data obtained through amplicon-based high-throughput sequencing. Therefore, absolute quantitation of microbiota abundance should be recommended in microbial ecology.
In this study, pH, TK, NH4-N, OP, AK, and K had a more significant effect on the bacterial community in the soil than the other chemical properties. The pH, TK, NH4-N, and OP were positively correlated with B_soil, whereas AK and K were negatively correlated with Rz_soil. For the fungal community, pH had the most substantial effect, showing a positive correlation with B_soil and a negative correlation with Rz_soil. Zhou et al. (2017) asserted that pH and vegetation are the primary factors influencing soil bacterial diversity and composition in the chronosequence of rubber trees (Hevea brasiliensis) plantations. Lauber et al. (2009) reported that soil pH significantly affects the structure of soil bacterial communities on a continental scale. Different microorganisms thrive within specific pH ranges, which are conducive to their growth and reproduction. Consequently, soil pH can influence the activity of soil microorganisms; if the pH of the soil solution falls outside the appropriate range, microbial activity is inhibited. Furthermore, the pH of the soil solution can alter the solubility of minerals, thereby affecting the nutrient availability in the soil and the activity of soil microorganisms (Philippot et al., 2024).
Soil enzymes play a crucial role in soil organic matter degradation, mineralization, and nutrient cycling. Their activities significantly influence soil nutrient content (Burns et al., 2013; Dotaniya et al., 2019). By measuring the soil enzyme activity, we can gain a deeper understanding of how T. mongolica affects soil properties from a protein perspective. Unfortunately, the soil enzyme activity between Rz_soil and B_soil in soil samples from three plant communities. It is one of further research work.
Microorganism in soil is another biological factor that affects soil nutrition (Zhang et al., 2021; Coban et al., 2022; Philippot et al., 2024). The high-throughput sequencing technology is common method for reveal microorganism in soil (Nkongolo and Narendrula-Kotha, 2020). In this study, throughput sequencing technology based on amplicon was used to determine microbial communities. However, metagenomic sequencing technology, which is superior to amplicon sequencing, can provide insights into bacteria and fungi involved in biogeochemical cycles, not only soil microbial community composition, but also their functional genes and metabolic pathways. This is particularly for the C cycle (including CO2 fixation and respiration), N cycle (encompassing nitrification, denitrification, and N2 fixation), P cycle, and S cycle (including sulfur assimilation, anaerobic sulfate respiration, and sulfide oxidation) (Simon and Daniel, 2011; Scholz et al., 2012). Further research in this area is required.
T. mongolica is rooted in the West Ordos Region of Inner Mongolia, northwest China, since the ancient Mediterranean period, approximately 140 million years ago. This study found that T. mongolica plantations decreased the soil pH and increased the nutrient content. Additionally, T. mongolica plantations altered community composition, co-occurrence networks, and ecological functions. In conclusion, as a keystone species in desert ecosystems, T. mongolica plantations significantly influence desert soil properties and microbial communities and play an irreplaceable role in local ecosystem stability. These findings offer a new perspective to understand the role of T. Mongolica in the desert ecosystems.
The datasets presented in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found in the article/Supplementary Material.
YQ: Writing – original draft. XG: Writing – original draft. SL: Data curation, Writing – original draft. XS: Data curation, Writing – original draft. MB: Investigation, Writing – original draft. YL: Data curation, Writing – original draft. YG: Data curation, Writing – original draft. HZ: Writing – original draft. XZ: Data curation, Writing – original draft. JW: Investigation, Writing – original draft. TC: Investigation, Writing – original draft. JYL: Funding acquisition, Writing – review & editing. JLL: Funding acquisition, Writing – review & editing.
The author(s) declare that financial support was received for the research and/or publication of this article. This research was supported by the National Natural Science Foundation of China (32360350; 31960346), Science and Technology Leading Talent Project of Ningxia (2022GKLRLX06), and The Talent Highland for the Research and Development of Engineering Technologies for Ecological Restoration in Arid Areas (2023RCGD07).
The authors thank the respectful editors and reviewers for their valuable and constructive comments and suggestions that helped strengthen this paper. We would like to appreciate KetengEdit (www.ketengedit.com) for its linguistic assistance.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fpls.2025.1539336/full#supplementary-material
Ali, H. E., Al-Wahaibi, A. M., Shahid, M. S. (2024). Plant-soil feedback and plant invasion: effect of soil conditioning on native and invasive Prosopis species using the plant functional trait approach. Front. Plant Sci. 15. doi: 10.3389/fpls.2024.1321950
Barberán, A., Bates, S. T., Casamayor, E. O., Fierer, N. (2012). Using network analysis to explore co-occurrence patterns in soil microbial communities. ISME J. 6, 343–351. doi: 10.1038/ismej.2011.119
Beckman, N. G., Dybzinski, R., Tilman, D. (2023). Short-term plant-soil feedback experiment fails to predict outcome of competition observed in long-term field experiment. Ecology. 104, e3883. doi: 10.1002/ecy.3883
Bolyen, E., Rideout, J. R., Dillon, M. R., Bokulich, N. A., Abnet, C. C., Al-Ghalith, G. A., et al. (2019). Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857. doi: 10.1038/s41587-019-0209-9
Burns, R. G., DeForest, J. L., Marxsen, J., Sinsabaugh, R. L., Stromberger, M. E., Wallenstein, M. D., et al. (2013). Soil enzymes in a changing environment: current knowledge and future directions. Soil Biol. Biochem. 58, 216–234. doi: 10.1016/j.soilbio.2012.11.009
Chen, Q. L., Hu, H. W., He, Z. Y., Cui, L., Zhu, Y. G., He, J. Z. (2021). Potential of indigenous crop microbiomes for sustainable agriculture. Nat. Food. 2, 233–240. doi: 10.1038/s43016-021-00253-5
Cheng, J., Kao, H., Dong, S. (2020). Population genetic structure and gene flow of rare and endangered Tetraena mongolica Maxim. revealed by reduced representation sequencing. BMC Plant Biol. 20, 1–13. doi: 10.1186/s12870-020-02594-y
Coban, O., De Deyn, G. B., van der Ploeg, M. (2022). Soil microbiota as game-changers in restoration of degraded lands. Science. 375, abe0725. doi: 10.1126/science.abe0725
Creamer, R., Barel, J., Bongiorno, G., Zwetsloot, M. (2022). The life of soils: Integrating the who and how of multifunctionality. Soil Biol. Biochem. 166, 108561. doi: 10.1016/j.soilbio.2022.108561
Crowther, T. W., van den Hoogen, J., Wan, J., Mayes, M. A., Keiser, A. D., Mo, L., et al. (2019). The global soil community and its influence on biogeochemistry. Science. 365, eaav0550. doi: 10.1126/science.aav0550
Dang, Z., Huang, L., Jia, Y., Lockhart, P. J., Fong, Y., Tian, Y. (2020). Identification of genic SSRs provide a perspective for studying environmental adaptation in the endemic shrub Tetraena mongolica. Genes (Basel). 11, 322. doi: 10.3390/genes11030322
DeSantis, T. Z., Hugenholtz, P., Larsen, N., Rojas, M., Brodie, E. L., Keller, K., et al. (2006). Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl. Environ. Microbiol. 72, 5069–5072. doi: 10.1128/AEM.03006-05
de Vries, F., Lau, J., Hawkes, C., Semchenko, M. (2023). Plant-soil feedback under drought: does history shape the future? Trends Ecol. Evol. 38, 708–718. doi: 10.1016/j.tree.2023.03.001
Dotaniya, M. L., Aparna, K., Dotaniya, C. K., Singh, M., Regar, K. L. (2019). Role of soil enzymes in sustainable crop production (New York: Academic Press), 569–589.
Feng, Z., Wang, L., Peng, Q., Li, J., Liang, T. (2022). Effect of environmental factors on soil properties under different land use types in a typical basin of the North China Plain. J. Clean. Prod. 344, 131084. doi: 10.1016/j.jclepro.2022.131084
Fierer, N. (2017). Embracing the unknown: disentangling the complexities of the soil microbiome. Nat. Rev. Microbiol. 15, 579–590. doi: 10.1038/nrmicro.2017.87
Furey, G. N., Tilman, D. (2021). Plant biodiversity and the regeneration of soil fertility. Proc. Natl. Acad. Sci. U.S.A. 118, e2111321118. doi: 10.1073/pnas.2111321118
Guo, P., Lu, Q., Li, S. (2024b). Productivity and species diversity of plant communities are higher inside than outside the West Ordos National Nature Reserve, Northern China. Plants (Basel). 13, 660. doi: 10.3390/plants13050660
Guo, J., Zhang, M., Bi, Y., Zhao, Z., Wang, R., Li, M. (2024a). Spatiotemporal distribution prediction of the relict and endangered plant Tetraena mongolica in inner Mongolia, China under climate change. Sci. Rep. 14, 28478. doi: 10.1038/s41598-024-79088-6
Guo, X., Zhang, X., Qin, Y., Liu, Y. X., Zhang, J., Zhang, N., et al. (2020). Host-associated quantitative abundance profiling reveals the microbial load variation of root microbiome. Plant Commun. 1, 100003. doi: 10.1016/j.xplc.2019.100003
Hartmann, M., Six, J. (2023). Soil structure and microbiome functions in agroecosystems. Nat. Rev. Earth Env. 4, 4–18. doi: 10.1038/s43017-022-00366-w
Jiang, S. Q., Yu, Y. N., Gao, R. W., Wang, H., Zhang, J., Li, R., et al. (2019). High-throughput absolute quantification sequencing reveals the effect of different fertilizer applications on bacterial community in a tomato cultivated coastal saline soil. Sci. Total Environ. 687, 601–609. doi: 10.1016/j.scitotenv.2019.06.105
Jin, J., Huang, X., Wu, J., Zhao, W., Fu, W. (2022). A 10-year field experiment proves the neutralization of soil pH in Chinese hickory plantation of southeastern China. J. Soils Sed. 22, 2995–3005. doi: 10.1007/s11368-022-03290-0
Kranz, C. N., McLaughlin, R. A., Johnson, A., Miller, G., Heitman, J. L. (2020). The effects of compost incorporation on soil physical properties in urban soils–A concise review. J. Environ. Manage. 261, 110209. doi: 10.1016/j.jenvman.2020.110209
Lauber, C. L., Hamady, M., Knight, R., Fierer, N. (2009). Pyrosequencing-based assessment of soil pH as a predictor of soil bacterial community structure at the continental scale. Appl. Environ. Microbiol. 75, 5111–5120. doi: 10.1128/AEM.00335-09
Lee, S. M., Kong, H. G., Song, G. C., Ryu, C. M. (2021). Disruption of Firmicutes and Actinobacteria abundance in tomato rhizosphere causes the incidence of bacterial wilt disease. ISME J. 15, 330–347. doi: 10.1038/s41396-020-00785-x
Lehmann, J., Bossio, D. A., Kogel-Knabner, I., Rillig, M. C. (2020). The concept and future prospects of soil health. Nat. Rev. Earth Environ. 1, 544–553. doi: 10.1038/s43017-020-0080-8
Li, E., Huang, Y., Chen, H., Zhang, J. (2018). Floristic diversity analysis of the Ordos Plateau, a biodiversity hotspot in arid and semi-arid areas of China. Folia. Geobot. 53, 405–416. doi: 10.1007/s12224-018-9331-6
Li, T., Zhang, Q., Wang, G., Singh, V. P., Zhao, J., Sun, S., et al. (2023). Ecological degradation in the Inner Mongolia reach of the Yellow River Basin, China: Spatiotemporal patterns and driving factors. Ecol. Indicators. 154, 110498. doi: 10.1016/j.ecolind.2023.110498
Liu, S., Wang, J., Pu, S., Blagodatskaya, E., Kuzyakov, Y., Razavi, B. S. (2020). Impact of manure on soil biochemical properties: A global synthesis. Sci. Total Environ. 745, 141003. doi: 10.1016/j.scitotenv.2020.141003
Liu, Z., Wang, C., Yang, X., Liu, G., Cui, Q., Indree, T., et al. (2023b). The relationship and influencing factors between endangered plant tetraena mongolica and soil microorganisms in West Ordos Desert Ecosystem, Northern China. Plants (Basel). 12, 1048. doi: 10.3390/plants12051048
Liu, B., Zhao, X., Wang, Z., Liu, H., Huang, X., Yang, P. (2023a). Chromosome-level genome assembly of the endangered plant Tetraena mongolica. DNA Res. 30, dsad004. doi: 10.1093/dnares/dsad004
Lou, J., Yang, L., Wang, H., Wu, L., Xu, J. (2018). Assessing soil bacterial community and dynamics by integrated high-throughput absolute abundance quantification. PeerJ. 6, e4514. doi: 10.7717/peerj.4514
Louca, S., Parfrey, L. W., Doebeli, M. (2016). Decoupling function and taxonomy in the global ocean microbiome. Science. 353, 1272–1277. doi: 10.1126/science.aaf4507
Lu, R. (2000). Methods for soil agrochemistry analysis (Beijing: China Agricultural Science and Technology Press).
Nilsson, R. H., Larsson, K.-H., Taylor, A. F. S., Bengtsson-Palme, J., Jeppesen, T. S., Schigel, D., et al. (2019). The UNITE database for molecular identification of fungi: handling dark taxa and parallel taxonomic classifications. Nucleic Acids Res. 47, D259–D264. doi: 10.1093/nar/gky1022
Nkongolo, K. K., Narendrula-Kotha, R. (2020). Advances in monitoring soil microbial community dynamic and function. J. Appl. Genet. 61, 249–263. doi: 10.1007/s13353-020-00549-5
Oleghe, E., Naveed, M., Baggs, E., Hallett, P. (2019). Residues with varying decomposability interact differently with seed or root exudate compounds to affect the biophysical behaviour of soil. Geoderma. 343, 50–59. doi: 10.1016/j.geoderma.2019.02.023
Passari, A. K., Upadhyaya, K., Singh, G., Abdel-Azeem, A. M., Thankappan, S., Uthandi, S., et al. (2019). Enhancement of disease resistance, growth potential, and photosynthesis in tomato (Solanum lycopersicum) by inoculation with an endophytic actinobacterium, Streptomyces thermocarboxydus strain BPSAC147. PloS One 14, e0219014. doi: 10.1371/journal.pone.0219014
Patel, K. F., Fansler, S. J., Campbell, T. P., Bond-Lamberty, B., Smith, A. P., RoyChowdhury, T., et al. (2021). Soil texture and environmental conditions influence the biogeochemical responses of soils to drought and flooding. Nat. Rev. Earth Environ. 2, 127. doi: 10.1038/s43247-021-00198-4
Philippot, L., Chenu, C., Kappler, A., Rillig, M. C., Fierer, N. (2024). The interplay between microbial communities and soil properties. Nat. Rev. Microbiol. 22, 226–239. doi: 10.1038/s41579-023-00980-5
Saleem, M., Hu, J., Jousset, A. (2019). More than the sum of its parts: microbiome biodiversity as a driver of plant growth and soil health. Annu. Rev. Ecol. Evol. S. 50, 145–168. doi: 10.1146/annurev-ecolsys-110617-062605
Sampson, T. R., Debelius, J. W., Thron, T., Janssen, S., Shastri, G. G., Ilhan, Z. E., et al. (2016). Gut microbiota regulate motor deficits and neuroinflammation in a model of Parkinson’s disease. Cell. 167, 1469–1480. doi: 10.1016/j.cell.2016.11.018
Schlatter, D., Kinkel, L., Thomashow, L., Weller, D., Paulitz, T. (2017). Disease suppressive soils: new insights from the soil microbiome. Phytopathology. 107, 1284–1297. doi: 10.1094/PHYTO-03-17-0111-RVW
Schloss, P. D., Westcott, S. L., Ryabin, T., Hall, J. R., Hartmann, M., Hollister, E. B., et al. (2009). Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 75, 7537–7541. doi: 10.1128/AEM.01541-09
Scholz, M. B., Lo, C.-C., Chain, P. S. (2012). Next generation sequencing and bioinformatic bottlenecks: the current state of metagenomic data analysis. Curr. Opin. Biotechnol. 23, 9–15. doi: 10.1016/j.copbio.2011.11.013
Semchenko, M., Barry, K. E., de Vries, F. T., Mommer, L., Moora, M., Macia-Vicente, J. G. (2022). Deciphering the role of specialist and generalist plant-microbial interactions as drivers of plant-soil feedback. New Phytol. 234, 1929–1944. doi: 10.1111/nph.18118
Simon, C., Daniel, R. (2011). Metagenomic analyses: past and future trends. Appl. Environ. Microbiol. 77, 1153–1161. doi: 10.1128/AEM.02345-10
Stammler, F., Glasner, J., Hiergeist, A., Holler, E., Weber, D., Oefner, P. J., et al. (2016). Adjusting microbiome profiles for differences in microbial load by spike-in bacteria. Microbiome. 4, 28. doi: 10.1186/s40168-016-0175-0
Tibbett, M., Gil-Martínez, M., Fraser, T., Green, I. D., Duddigan, S., De Oliveira, V. H., et al. (2019). Long-term acidification of pH neutral grasslands affects soil biodiversity, fertility and function in a heathland restoration. Catena. 180, 401–415. doi: 10.1016/j.catena.2019.03.013
Tkacz, A., Hortala, M., Poole, P. S. (2018). Absolute quantitation of microbiota abundance in environmental samples. Microbiome. 6, 110. doi: 10.1186/s40168-018-0491-7
van Agtmaal, M., Straathof, A., Termorshuizen, A., Teurlincx, S., Hundscheid, M., Ruyters, S., et al. (2017). Exploring the reservoir of potential fungal plant pathogens in agricultural soil. Appl. Soil. Ecol. 121, 152–160. doi: 10.1016/j.apsoil.2017.09.032
Vandeputte, D., Kathagen, G., D’Hoe, K., Vieira-Silva, S., Valles-Colomer, M., Sabino, J., et al. (2017). Quantitative microbiome profiling links gut community variation to microbial load. Nature. 551, 507–511. doi: 10.1038/nature24460
Wang, X., Wang, M., Xie, X., Guo, S., Zhou, Y., Zhang, X., et al. (2020). An amplification-selection model for quantified rhizosphere microbiota assembly. Sci. Bull. 65, 983–986. doi: 10.1016/j.scib.2020.03.005
Walters, W., Hyde, E. R., Berg-Lyons, D., Ackermann, G., Humphrey, G., Parada, A., et al (2016). Improved bacterial 16S rRNA gene (V4 and V4-5) and fungal internal transcribed spacer marker gene primers for microbial community surveys. mSystems. 1, e00009-15. doi: 10.1128/mSystems.00009-15
Wu, Z., Wei, W., Cheng, K., Zheng, L., Ma, C., Wang, Y. (2020). Insecticidal activity of triterpenoids and volatile oil from the stems of Tetraena mongolica. Pestic. Biochem. Physiol. 166, 104551. doi: 10.1016/j.pestbp.2020.02.017
Xiong, Y. W., Gong, Y., Li, X. W., Chen, P., Ju, X. Y., Zhang, C. M., et al. (2019). Enhancement of growth and salt tolerance of tomato seedlings by a natural halotolerant actinobacterium Glutamicibacter halophytocola KLBMP 5180 isolated from a coastal halophyte. Plant Soil. 445, 307–322. doi: 10.1007/s11104-019-04310-8
Yan, P., Wu, L., Wang, D., Fu, J., Shen, C., Li, X., et al. (2020). Soil acidification in Chinese tea plantations. Sci. Total Environ. 715, 136963. doi: 10.1016/j.scitotenv.2020.136963
Yang, L., Lou, J., Wang, H., Wu, L., Xu, J. (2018). Use of an improved high-throughput absolute abundance quantification method to characterize soil bacterial community and dynamics. Sci. Total Environ. 633, 360–371. doi: 10.1016/j.scitotenv.2018.03.201
Zhang, N., Nunan, N., Hirsch, P. R., Sun, B., Zhou, J., Liang, Y. (2021). Theory of microbial coexistence in promoting soil–plant ecosystem health. Biol. Fertility Soils. 57, 897–911. doi: 10.1007/s00374-021-01586-w
Zhang, M., Zhang, L., Huang, S., Li, W., Zhou, W., Philippot, L., et al. (2022). Assessment of spike-AMP and qPCR-AMP in soil microbiota quantitative research. Soil Biol. Biochem. 166, 108570. doi: 10.1016/j.soilbio.2022.108570
Zhao, M., Zhao, J., Yuan, J., Hale, L., Wen, T., Huang, Q., et al. (2021). Root exudates drive soil-microbe-nutrient feedbacks in response to plant growth. Plant Cell Environ. 44, 613–628. doi: 10.1111/pce.13928
Zheng, Q., Hu, Y., Zhang, S., Noll, L., Böckle, T., Dietrich, M., et al. (2019). Soil multifunctionality is affected by the soil environment and by microbial community composition and diversity. Soil Biol. Biochem. 136, 107521. doi: 10.1016/j.soilbio.2019.107521
Zhou, X., Guo, Z., Chen, C., Jia, Z. (2017). Soil microbial community structure and diversity are largely influenced by soil pH and nutrient quality in 78-year-old tree plantations. Biogeosciences. 14, 2101–2111. doi: 10.5194/bg-14-2101-2017
Keywords: Tetraena mongolica, soil nutrition, bacterial communities, fungi communities, root zone soil, bare soil
Citation: Quan Y, Gan X, Lu S, Shi X, Bai M, Lin Y, Gou Y, Zhang H, Zhang X, Wei J, Chang T, Li J and Liu J (2025) The relict plant Tetraena mongolica plantations increase the nutrition and microbial diversity in desert soil. Front. Plant Sci. 16:1539336. doi: 10.3389/fpls.2025.1539336
Received: 04 December 2024; Accepted: 25 February 2025;
Published: 20 March 2025.
Edited by:
Xianan Xie, South China Agricultural University, ChinaReviewed by:
Fang Song, Hubei Academy of Agricultural Science, ChinaCopyright © 2025 Quan, Gan, Lu, Shi, Bai, Lin, Gou, Zhang, Zhang, Wei, Chang, Li and Liu. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Jianli Liu, bGpsNzUyM0AxMjYuY29t
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.