
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Plant Sci. , 18 March 2025
Sec. Crop and Product Physiology
Volume 16 - 2025 | https://doi.org/10.3389/fpls.2025.1536178
This article is part of the Research Topic Elucidating the Molecular, Physiological, and Biochemical Mechanisms Underlying Stress Responses in Crop Plants View all 12 articles
 Syed Salman Hashmi1†
Syed Salman Hashmi1† Lubna1†
Lubna1† Saqib Bilal1
Saqib Bilal1 Rahmatullah Jan2
Rahmatullah Jan2 Saleem Asif3
Saleem Asif3 Ashraf M. M. Abdelbacki4
Ashraf M. M. Abdelbacki4 Kyung-Min Kim3*
Kyung-Min Kim3* Ahmed Al-Harrasi1
Ahmed Al-Harrasi1 Sajjad Asaf1*
Sajjad Asaf1*ATP-binding cassette (ABC) transporters are integral membrane proteins involved in the active transport of various substrates, including heavy metals, across cellular membrane. In this study, we performed a genome-wide analysis and explored the expression profiles of ABC transporter genes in Solanum lycopersicum to identify their role in cadmium (Cd) stress tolerance. Several techniques were employed to determine the regulatory role of ABC transporters. A total of 154 ABC transporter genes were identified in the genome of S. lycopersicum, located on all 12 chromosomes. Comparative phylogenetic analysis between S. lycopersicum and Arabidopsis thaliana revealed several orthologous gene pairs, which were duly supported by the structural analysis of the genes by studying the exon-intron pattern and motif analysis. Collinearity analysis revealed multiple gene duplication events owing to intra-chromosomal and inter-chromosomal mutations. The cis-regulatory analysis identified several hormone-responsive elements suggesting that ABCs are actively involved in transporting hormones like ABA, SA, MeJA, auxin, and gibberellin. These hormones are known to combat a number of stress conditions, hence validating the role of ABCs in Cd stress. Under Cd stress, expression profiling demonstrated that several SlABCs exhibit significant transcriptional changes, indicating their involvement in Cd transport, sequestration, and detoxification mechanisms. Specific genes, including Groups 3 and 5 members, were upregulated under Cd exposure, suggesting their functional roles in mitigating Cd toxicity. The study revealed differential expressions of various SlABC genes encoding ATP binding cassette transporters, including the upregulation of several genes like Solyc08g067620.2, Solyc08g067610.3, Solyc12g019640.2, Solyc06g036240.2, and Solyc05g053610.2 in response to different concentrations of Cd. This study comprehensively explains the ABC transporter gene family in S. lycopersicum, emphasizing their critical roles in Cd stress tolerance. This study could prove useful in combating Cd stress not only in S. lycopersicum but also in other fleshy fruit plants; however, further advanced studies on specific pathways that lead to differential expression of the ABC genes are required to understand the mechanism behind tolerance to heavy metals fully.
S. lycopersicum, a member of the Solanaceae family, commonly known as the tomato plant, is widely cultivated for its edible product, i.e., tomato. It is among the most cultivated vegetables across the globe, with estimated production of 187 million tons (Arshad et al., 2024). It is widely used for making a variety of dishes and sauces. Lycopene, phenolics, flavonoids, carotenoids, gallic acid, quercetin, vitamin A, vitamin B, and vitamin C make S. lycopersicum an important medicinal plant. These bioactive compounds have potential anti-cancer, anti-inflammatory, radio-protective, antibacterial, and antifungal activity. S. lycopersicum also possess the potential to alleviate symptoms of metabolic syndromes like diabetes, hypertension, diabetes-induced testicular injuries, and hypercholesterolemia (Kumar et al., 2021; Mallick, 2021; Collins et al., 2022). In nature, plants have to cope with numerous environmental conditions that induce stress. The stress may be induced by living organisms, i.e., viruses, fungi, bacteria, insects, and other pathogens (biotic), or their non-living environment, i.e., drought, salinity, floods, nutrient deficiency, heavy metals (HMs), and extreme temperatures (abiotic). Since plants are sessile, escaping biotic and abiotic stress is not an option, and prolonged exposure may have a detrimental impact on plant growth and development (Iqbal et al., 2021; Zhang et al., 2023).
The presence of HMs in large quantities in the soil is a matter of great concern globally (Kaur et al., 2021). Any metal with an atomic density higher than 4 g.cm-3 is considered a heavy metal e,g. nickel (Ni), lead (Pb), copper (Cu), mercury (Hg), zinc (Zn), aluminum (Al), chromium (Cr), and cadmium (Cd). The levels of HM pollution in our environment is rising at an alarming rate due to natural factors (acid rain and volcanic eruptions etc.) and human activities (mining, agriculture and industries) (Feng et al., 2021). HMs pose a long-term threat to the natural environment and human survival because of their non-degradable nature, long-term presence, and invisibility in the environment (Suman et al., 2018). Within cells, HMs affect photosynthesis, respiration, mineral nutrition, enzyme reactions, and many other physiological processes (Pourrut et al., 2011). The excessive accumulation of HMs in plant tissues can reduce root length, plant biomass, seed germination, and chlorophyll synthesis (Shahid et al., 2014). The inhibition of root growth, especially at the tip, is susceptible to certain HMs such as Al, Cu, Cd, and Hg (Gupta et al., 2014). The effects of HMs on plants are multifaceted, mainly including plant growth, plant dry weight, net photosynthesis, and stomatal conductance etc (Xiong et al., 2019). Plants, when exposed to HMs, result in metabolic cascades involving a plethora of genes. This results in the production of biochemicals that help plants endure HM stress (Khan et al., 2023a). These biochemicals are readily transported by membrane transport proteins, including transmembrane proteins like ATP-binding cassette (ABC) transporters (Ofori et al., 2018).
The ABC transporter factor family is found in various plant species, with different numbers of genes identified in each species. For example, Arabidopsis thaliana has 127 ABC genes, Oryza sativa L. has 123, and Hordeum vulgare has 131 (Guo et al., 2022). Initially, these transporters were identified for their role in cell detoxification by removing harmful chemicals from the cytoplasm. However, further research has demonstrated that ABC transporters are involved in a wide range of biological functions in plants, primarily situated in cellular membranes (Do et al., 2018). These genes have diverse functions, including transporting hormones, lipids, metals, pathogens, antibiotics, secondary metabolites, and other substances. They play a vital role in plant-pathogen interactions and ion channel regulation (Guo et al., 2022). Apart from this, it also affects seed dormancy, grain shape and size regulation, accumulation of anti-nutritional factors in seeds, transport of abscisic acid (ABA) and other phytohormones, growth and differentiation, nutrient uptake, redox balance, and endosperm weakening (Kang et al., 2011; Do et al., 2018).
ABC also plays a critical role in a variety of stress conditions. In a study by An et al. (An et al., 2019), Betula halophile, a plant with the ability to survive extreme drought and salinity stress was reported to contain 15 diverse ABC transporter genes. The ABC family transporter HMT1 plays a significant role in the uptake of HMs through chelation and their subsequent partitioning into the vacuole. Similarly, AtABCC1 and AtABCC2 are crucial for metal uptake and sequestration, facilitating the transport of heavy metal-phytochelatin complexes into the vacuole, especially for metals like arsenic [As(III)], Cd [Cd(II)], and mercury [Hg(II)] (Park et al., 2009; Yang and Murphy, 2009; Song et al., 2010). ABC transporters protect tissues from metal toxicity in plant roots, facilitate auxin transport, and contribute to suberin deposition. In leaves, these transporters play role in ABA import, cuticle formation, and ion transport (Do et al., 2018). ABC transporters are vital for protecting pollen from environmental stresses and ensuring proper fertilization and seed development (Luo et al., 2020). In the shoots, they are involved in auxin transport, transporting fatty acyl CoA to peroxisomes, forming the cuticle, and generating plastid lipids (Do et al., 2018). They also contribute to forming the exine, the outer pollen wall, and aid in accumulating minerals, lipids, and cutin in seeds (Kang et al., 2011). Additionally, tonoplastic ABC transporters such as ABCB27 and OsALS1 have been linked to Al tolerance. The role of AtABCC1 and AtABCC2 in the vacuolar sequestration of Cd was effectively demonstrated using a fluorescent probe sensitive to Cd (Song et al., 2014). Furthermore, AtABCB25, a mitochondrial ABC transporter involved in synthesizing iron-sulfur clusters, was upregulated in the roots of Cd-treated plants. Overexpression of AtABCB25 was shown to enhance Cd resistance (Song et al., 2014). The findings strongly indicate that ABC transporters likely play a crucial role in regulating the absorption and movement of both vital nutrients and HMs within organism. This suggests that they could influence how much nutrients or HMs accumulate in tissues. ABCs potentially impact nutrient utilization on nutrient-poor soils and the ability of plants to detoxify HMs through phytoremediation.
A genome-wide analysis is the comprehensive identification of all genes of the respective family including their family members and organization of their information. This approach provides essential information, such as evolutionary history, diversity, and relationship among genes and proteins, which serve as useful fundamental resources for further investigations. Genome-wide analyses of ABC transporters in Arabidopsis (Sánchez-Fernández et al., 2001), rice (Garcia et al., 2004), maize (Pang et al., 2013), Lotus japonicas (Sugiyama et al., 2006), grape (Çakır and Kılıçkaya, 2013), pineapple (Chen et al., 2017), and Hevea brasiliensis (Zhiyi et al., 2015) have already been performed. Ofori et al (Ofori et al., 2018). carried out a genome-wide analysis of a tomato plant and identified 154 genes in its genome that encode for ABC proteins. The study also reports a phylogenetic analysis of these genes, providing valuable insights into the evolutionary history of ABC proteins found in S. lycopersicum. The study proved to be a gateway for further insights into ABC protein studies in S. lycopersicum and the family Solanaceae; however, to date, no data is available on the involvement and differential expression of the aforementioned genes in S. lycopersicum plants when exposed to HMs. Additionally, further focused research is needed to better understand how ABC transporters function in nutrient uptake, transport, and maintaining internal balance, as well as their role in detoxifying HMs. In this regard, we conducted an in-depth exploration of the ABC gene family in S. lycopersicum through a combination of in-silico analysis and experimental approaches. First, we performed a genome-wide analysis of the S. lycopersicum ABC genes and followed up with a greenhouse experiment to examine how these genes respond to varying concentrations of Cd. Additionally, we utilized RNA-Seq to analyze gene expression patterns and further validated our findings through qRT-PCR (Real-Time Quantitative Reverse Transcription Polymerase Chain Reaction). The goal of this research was to better understand the role of ABC genes in S. lycopersicum’s response to Cd stress, integrating both computational and experimental data for a comprehensive analysis.
The current research was conducted on the Tomato plant (S. lycopersicum c v. Yegwang). Seeds were surface sterilized according to previously reported protocols (Khan et al., 2023b). The seeds were properly washed with distilled water (dH2O) thrice and then germinated at room temperature (28 ± 2°C; humidity 60 ± 5%) in total darkness for 2-3 days with the help of sterile filter paper. Seedlings were transferred to pots containing autoclaved soil which was composed of coco peat moss (10–15%), coco peat (45–50%), pertile (35–40), NH+ (ca.0.09 mg/g), KO (c.0.1 mg/g), zeolite (6-8%) with NO3 (ca. 0.205 mg/g), and PO (ca.0.35 mg/g) in a greenhouse at a constant temperature (28 ± 2°C) under a 16/8 hr light and dark period, respectively. Three sets with 5 replicates per set were used for the experiment. Set 1 was kept as control, set 2 contained plants subjected to 1mM Cd stress, and set 3 contained plants subjected to 2mM Cd stress. The concentrations for Cd stress were selected based on the previous report (Khan et al., 2023a) and Cd stress was induced three days after transferring the plantlets into pots by soil irrigation method every three days. For the analyses, leaf samples from each replicate of all the sets were collected, and with the help of liquid nitrogen, the samples were immediately dried and stored at -80°C for RNA extraction.
Leaf samples freeze dried with liquid nitrogen were utilized to extract RNA according to previously reported protocols (Khan et al., 2023a). Briefly, The RNA obtained was diluted until the RNA quantity for each sample reached 100 ng. Three replicates from every treatment were carefully selected and sequenced for subsequent analysis. For RNA-seq analysis, the sequencing platform (Illumina HiSeq2000) was used as per the manufacturer’s procedure, resulting in 51-bp single-end reads. An efficient computational pipeline was used to find differences in gene regulation between the control and plants subjected to Cd stress. The quality was assessed through FastQC. Data was trimmed with the help of TrimGalore. To align and index the data to reference genome, HISAT2 was exploited. Feature Count (subread_v2.0.2) carried out read count quantification, and differential gene analysis was carried out with the help of DESeq2 in the R program. For DEG analysis, p-values were adjusted for multiple testing using the Benjamini-Hochberg procedure to control the false discovery rate (FDR). Genes were considered differentially expressed if they exhibited a log2 fold change (log2FC) greater than ±2, corresponding to a two-fold change in expression.
The S. lycopersicum and A. thaliana genes that encodes ATP binding cassettes (ABC) transporters were retrieved from Solgenomics (https://solgenomics.net accessed on 15 July 2023) and TAIR databases (https://www.arabidopsis.org/index.jsp accessed on 15 July 2023). The corresponding genome sequences and protein sequences were downloaded from Solanaceae Genomics Network (SGN, https://solgenomics.net/, accessed on 15 July 2023), and Arabidopsis Information Resource (TAIR, https://www.arabidopsis.org/index.jsp, accessed on 15 July 2023). Consequently, 154 ABC genes from S. lycopersicum and 130 ABC genes from Arabidopsis were identified. For valid identification and confirming the presence of the respective domains of the obtained protein sequences, the Simple Modular Architecture Research Tool (SMART, http://smart.embl.de/, accessed on 15 July 2023) was used at E-value <10-5. Only sequences containing the ABC transporter domain were retained, while sequences lacking this domain were excluded from further analysis. This validated dataset was then used for subsequent analysis
In order to classify the retrieved genes in S. lycopersicum (SlABCs), a phylogenetic tree was constructed by employing the Maximum likelihood (ML) technique with the help of software (MEGA 11), keeping the bootstrap replicates (BS) at 1000 times. The procedure above was repeated to construct a combined phylogenetic tree of 154 SlABCs with 130 ABCs from A. thaliana (AtABCs) as well, so as to determine the evolutionary relationship among the genes. The annotations of the resulting phylogenetic trees were performed using Version 1.4.4 of the FigTree (https://mybiosoftware.com/figtree-1-3-1-produce-figures-phylogenetic-trees.html#google_vignette).
Gene Structure Display Server (GSDS), an online tool (https://gsds.gao-lab.org/), was employed to investigate the gene structure, i.e., the number, structure, and organization of exons and introns in the identified SlABCs. In order to do so, the coding sequence (CDS) of SlABCs was aligned with the genomic sequence. For delving into the details of conserved Motifs in SlABCs, another online tool i.e. MEME (Multiple EM for Motif Elicitation) version 5.4.1 (https://meme-suite.org/meme/doc/meme.html) was employed by adjusting the motif number (maximum) to 20, keeping the optimum motif width between 6 to 200 amino acid (aa) residue and fixing the maximum repetitions to “any.”
The localization and distribution of SlABCs on the chromosomes of S. lycopersicum was accomplished by genome-wide collinearity analysis using TBtools software. The gene data required for this purpose was downloaded from the NCBI database.
The selected duplicate gene pairs were subjected to TBtools to compute the Ks and Ka values. The determined values were then used to determine the Ka/Ks ratio. The formula t = Ks/2r was used to calculate the divergence time with r being the representative value (1.5 × 10-8) as reported in the literature (Ali et al., 2024).
An online tool called OrthoVenn2 (https://orthovenn2.bioinfotoolkits.net/home) was utilized for identification of orthologous groups among the ABC transporter encoding genes of S. lycopersicum and A. thaliana by analyzing their full-length sequences as reported by (Abiraami et al., 2023). For the mapping and analysis of Cis-regulatory modules, PlantCARE, an online website (http://bioinformatics.psb.ugent.be/webtools/plantcare/html) was explored using 2000 bp sequence upstream of SlABCs from S. lycopersicum genome with the help of a software (TBtools).
The protein sequences of 154 SlABCs were acquired in FASTA format and uploaded online to version 11.5 of STRING’s server (https://string-db.org/) to predict the nature of interaction among different proteins encoded by SlABCs and determine the putative function of respective proteins. The network construction was done with the help of selected genes (having significant confidence scores), whereas non-interactive genes were skimmed from the network.
For the prediction of annotated cellular components, molecular functioning and biological processes involving SlABCs, the enrichment analysis of Gene Ontology (GO) categories was performed with the help of online software agriGo (http://bioinfo.cau.edu.cn/agriGO/). Algorithm TopGo ‘elim’ was exploited for the said purpose using the loci of SlABCs, which were mined online from the SolGenomics website. Significantly enriched GO Terms (having False Discovery Rate (FDR) and P-value less than 0.05) were considered for the current study, as reported in literature (Khan et al., 2023b).
In order to authenticate the results of the RNA-Seq dataset, 10 genes were selected based on their significant differential expression observed under cadmium (Cd) stress conditions in the RNA-Seq analysis. These genes were chosen to represent the most biologically relevant responses to Cd stress. The selected genes were subjected to qRT-PCR. RNA dilution (0.1µg/µl final concentration) was carried out, followed by the cDNA production with the help of PCR BIOSYSTEMS’s qPCRBIO cDNA Synthesis Kits. qRT-PCR was conducted using BioFACTTM 2× Real-Time PCR Master Mix (Including SYBR® Green I). A total of 20 μl sample containing master mix (10 μl), RNase free water (8 μl), cDNA, and primer (1 μl each) was utilized to carry out PCR reaction with the help of PCR machine (Step One Plus Real-Time PCR System, Life Technologies Holdings Pte Ltd., Singapore). The primers for selected genes were designed with the help of an online tool, i.e., Primer3 (https://bioinfo.ut.ee/primer3-0.4.0/), the details of which are available in (Supplementary Table S1). A housekeeping gene, i.e., Actin, was used as an internal control for gene expression normalization. To analyze expression levels of selected genes from control plants and those subjected to Cd stress. Two technical and three biological repeats were employed for each sample in order to minimize experimental error. The conditions for the PCR reaction were as follows: 10 min at 94°C, followed by 35 cycles at 94°C (45 s), 65°C (45 s), and 72°C (1 min), with an extension step at 72°C (10 min). The gene amplification threshold was set at 0.1. Each sample was run three times with three different replicates.
The phylogenetic tree was constructed using the Maximum Likelihood (ML) method for in-depth analysis of the relationship between the SlABCs. As evident in Figure 1, the results depicted that the identified SlABCs belong to 15 different groups, each labeled and designated by a specified color. More than half of the genes (55.8%) were grouped into 3 out of 15 groups, with Groups 5 and 6 having the maximum number of SlABCs (29 each), closely followed by Group 8 (28 SlABCs). The rest of the 12 groups collectively had 68 SlABCs (44.1%), with Group 11 having 13 SlABCs, Group 10 having 11 SlABCs, Group 1 having 9 SlABCs, Group 3 with 8 SlABCs, Group 13 with 5 SlABCs, Group 7, Group 9 and Group 14 having 4 SlABCs apiece and Group 2 and Group 15 with 3 SlABCs each. Groups 2 and 12 had the least SlABCs, i.e., 2 SlABCs each. A bootstrap value of over 99% in the majority of the clades validates the results above.
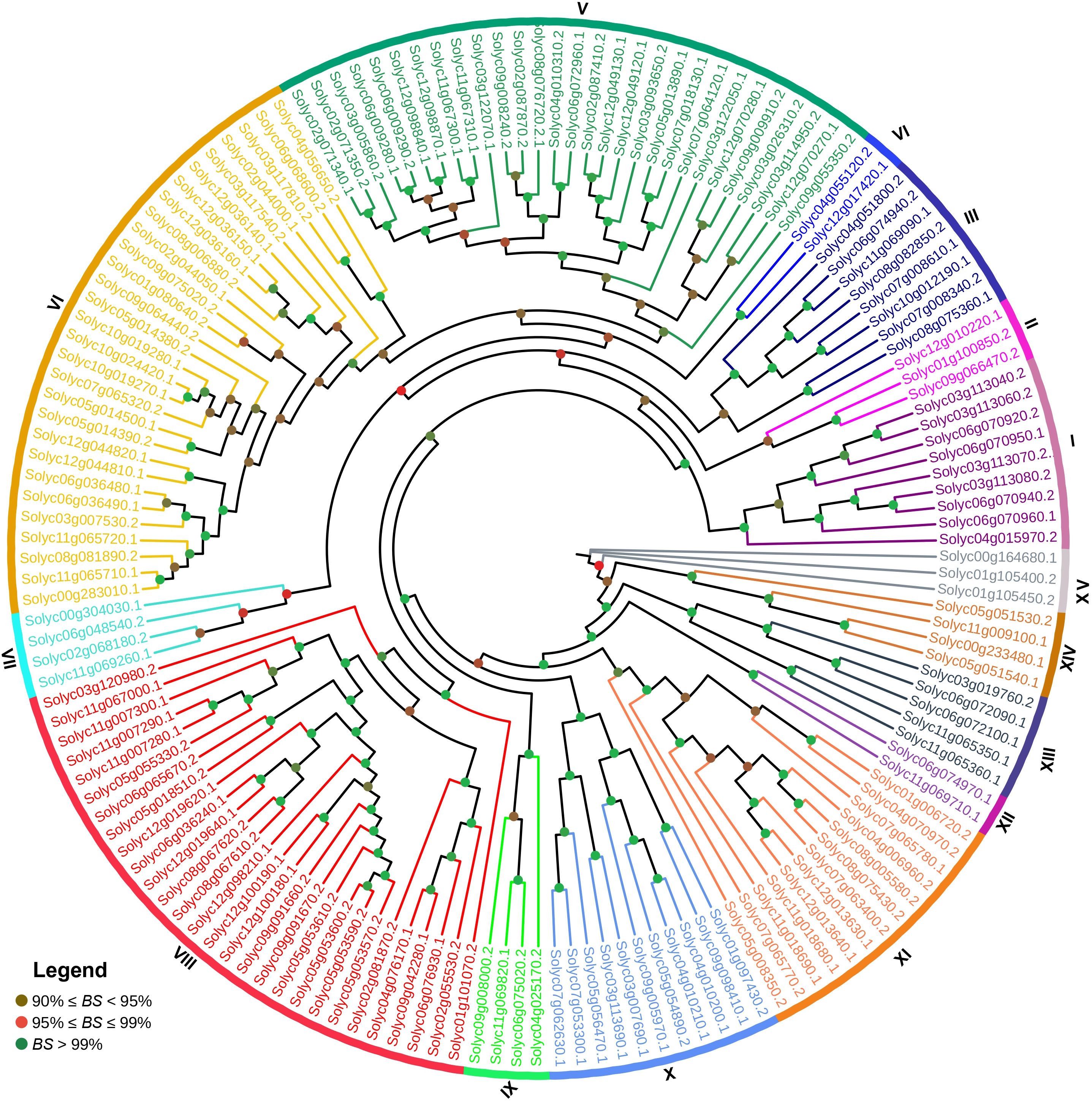
Figure 1. The phylogenetic tree of 154 SlABCs constructed using Maximum Likelihood (ML) method with the help of MEGA-11 software with 1000 bootstrap replicates. The major 15 phylogenetic groups are designated as 1-15, respectively.
To further understand the evolutionary relationship of the SlABCs with AtABCs, a combined phylogenetic tree of 154 SlABCs and 130 ABC genes belonging to A. thaliana (AtABCs) was constructed using the aforementioned method. The resulting combined phylogenetic tree, as evident in Figure 2, revealed 13 groups named A-M respectively. Apart from Group C and D, which only contained AtABCs (7 and 2 respectively), and L and M, which only contained SlABCs (5 and 2 respectively), the rest of the 9 groups contained ABC genes from both S. lycopersicum and A. thaliana. Group F contained the highest number of genes i.e. 102 (46 AtABCs and 56 SlABCs) followed by Group E with 57 (30 AtABCs and 27 SlABCs), Group G with 50 (18 AtABCs and 32 SlABCs), Group A with 23 (12 AtABCs and 11 SlABCs), Group I with 19 (7 AtABCs and 12 SlABCs), Group K with 9 (4 AtABCs and 5 SlABCs), Group B with 3 (2 AtABCs and 1 SlABC), Group J with 3 (1 AtABC and 2 SlABCs) and Group H with 2 (1 AtABC and 1 SlABC). Bootstrap value of over 95%-99% in all the clades validates the abovementioned results.

Figure 2. A joint phylogenetic tree was constructed by aligning 154 SlABCs and 130 A. thaliana ABCs using the Maximum Likelihood (ML) method with the help of MEGA-11 software with 1000 bootstrap replicates. The resulting thirteen groups, labeled A-M respectively, are shown in different colors.
A total of 154 ABC genes were identified in S. lycopersicum, which were unequally distributed on chromosomes of S. lycopersicum (Supplementary Table S2). Maximum number of SlABCs were present on chromosome 12 (20 SlABCs) followed by chromosome 6 (19 SlABCs), chromosome 3 and 11 (17 SlABCs each), chromosome 5 (15 SlABCs), chromosome 9 (12 SlABCs), chromosome 4 (11 SlABCs), chromosome 7 (10 SlABCs each), chromosome 2 and 8 (9 SlABCs each), chromosome 1 (7 SlABCs) and chromosome 10 (4 SlABCs). The precise location of 4 genes, however, was not identified. Apart from chromosomal distribution, other physical properties of the proteins encoded by SlABCs, like isoelectric point, molecular weight, alternative isoforms, number of exons and motifs, etc., also showed diversity (Supplementary Table S2). The average pI value was recorded to be 7.64, with Solyc01g105400.2 showing the lowest pI value (4.31) and Solyc00g304030.1 displaying the highest value (10.43). Much variation was also noted in encoded proteins and gene length. The number of amino acids ranged from 53 to 1,909 whereas the gene length varied from 244 to 35,740 bp in 154 identified SlABCs. The predicted molecular weight ranged between 5.9 kDa and 213.17 kDa; the average molecular weight was 96.85 kDa. Subcellular localization revealed that ABC proteins were located in almost all major organelles, including plastid, vacuole, nucleus mitochondria, endoplasmic reticulum, chloroplast, and Golgi complex (Supplementary Table S2).
A collinearity analysis was carried out to analyze the evolutionary relationship and duplication events in SlABCs. Duplication of genes is critical in increasing a gene family’s size. A number of SlABCs were found to be highly conserved on all the 12 chromosomes of S. lycopersicum suggesting multiple duplication events within the S. lycopersicum genome. The duplication included both intra-chromosomal and inter-chromosomal mutations (Figure 3A). Synteny analysis (Figure 3B) for identification of locus relationship among S. lycopersicum and A. thaliana genes showed the presence of 8 homologous genes. These genes were located on 7 out of the 12 chromosomes of S. lycopersicum. Interestingly, the gene on chromosome 6 of S. lycopersicum had homologous genes on chromosome 1. 2. 3, and 4 of A. thaliana, whereas chromosome 12 had 2 different genes having homologous genes on chromosome 3 and 4 of A. thaliana.

Figure 3. (A) Chromosomal positions and inter-chromosomal groups of duplicated SlABCs pairs were mapped on chromosomes (Chr1–Chr12) of S. lycopersicum. The lines represent the network zone of duplication among SlABCs, (B) Synteny analysis for identification of locus relationship among S. lycopersicum and A. thaliana genes.
In order to estimate the balance between different types of mutations, the Ks and Ka values of selected homologous gene pairs were calculated. The values were used to calculate the ratio between synonymous and non-synonymous values, the mean for which was 0.411. The results showed that the homologous genes evolved through purifying selection, suggesting that mutations that could deteriorate the genes were removed. The whole process was estimated to have occurred approximately 9 to 79 mya depending upon the homologous gene pair in question (Table 1).
Examining gene structure and motif composition offers valuable insights into evolutionary history and functional diversity. To explore these aspects, the conserved motifs and exon-intron organization of the 154 SlABC genes in S. lycopersicum were analyzed (Figures 4, 5). The number of exons in SlABC genes ranged from 1 to 40, while introns varied between 0 and 39, indicating considerable structural diversity within this gene family. MEME software identified 20 distinct conserved motifs, reflecting potential functional specialization among SlABC genes. Among these motifs, several were identified as Nucleotide Binding Domains (NBDs), crucial for ATP binding and hydrolysis. Specifically, Motif 2 contained the Walker A motif, a key ATP-binding site, while Motif 5 harbored an ABC transporter signature motif essential for transporter function. Additionally, Motif 12 was associated with the Walker B motif, which plays a fundamental role in ATP hydrolysis. Motif 13 was identified as another ATP-binding region commonly found in ABC transporters. The analysis further revealed that full-length ABC transporters predominantly contained two NBDs, whereas half-size ABC transporters had only one. The position of these domains varied across different genes, appearing at either the N-terminal, C-terminal, or in a duplicated arrangement, depending on the specific subfamily.

Figure 4. Schematic distribution of 20 conserved motifs in SlABCs using MEME Analysis. The motif consensus along with motif symbol is provided in the legend.

Figure 5. Exon–intron pattern analysis of SlABCs. The blue box represents upstream/downstream, the yellow box indicates exons, whereas the lines represent introns.
The conservation of these motifs across multiple SlABC genes highlights their strong functional preservation, particularly in ATP-dependent substrate transport. These findings suggest that despite sequence variations, the core functional components of the SlABC gene family remain highly conserved.
A total of 42 ortholog clusters (1-42) were identified which are given in Supplementary Figure S1. The orange dots represent A. thaliana ABC genes (AtABCs), whereas the blue dots represent SlABCs. The 42 designated clusters show different configurations, including many to many (eight clusters i.e. 1-7 and 10), many to one (six clusters i.e., 8-9, 13-16), one to many (5 clusters i.e. 11-12, 17-19) and one to one (23 clusters, i.e. 20-42) specific ABC members. Interestingly, the orthologs belonging to the same cluster had the same number of motifs, For example, all genes in cluster 2 had the same number of motifs (16) and a similar motif sequence. Similarly, genes belonging to clusters 11 and 12 had seven motifs with a similar motif sequence. Similarly, the number of exons and introns within the same cluster was also similar. For example, all the genes in Cluster 4 had 18 exons and 17 introns, whereas genes in Cluster 5 had a single exon with no introns. The number and sequence of SlABC genes in clusters 4 and 5, respectively, are also similar. The same pattern was observed for most of the remaining clusters as well. The above discussion validates the fact that the orthologs have evolved from a common ancestral gene.
According to Figure 2, the combined phylogenetic tree divided SlABCs and AtABCs into 13 groups. Except for Group B, which did not have any orthologs, all the other groups have a number of orthologs i.e. Group F contains 19 orthologs, Group E contained a total of 13 orthologs, Group G have six orthologs, Group A and Group I have five orthologs each, Group K contains two orthologs and Group J and Group H have one ortholog each. Although A. thaliana and S. lycopersicum belong to different families that diverged millions of years ago, most of the ABCs have orthologous relationships, suggesting that the nature and functions of AtABCs and SlABCs would show a similar pattern.
Various hormones play a critical role in enabling the plant to combat different biotic and abiotic stress conditions. In this regard, 137 SlABCs were found to play a role in hormone-responsive cis-regulatory elements (Figure 6). A total of 1150 cis-regulatory modules comprising 15 different elements were identified in which ABRE, an ABA-responsive element, was the most repeated cis-regulatory element present (279) found in 114 SlABCs followed by CGTCA-motif (155), a Methyl jasmonate (MeJA) responsive element and TGACG-motif (155) which is involved in the regulation of salicylic acid (SA) and MeJA signaling pathways which were identified in 54 and 53 SlABCs respectively. The number of cis-regulatory modules found in individual SlABCs showed tremendous diversity and ranged between 1-24 with Solyc12g098870.1 having the most number of cis-regulatory modules (7 TGACG-motifs and CGTCA-motifs, 6 ABRE and 1 O2-site, TC-rich repeats and TATC-box) and two SlABCs (Solyc11g018680.1 and Solyc09g042280.1) having only one cis-regulatory module. Interestingly, only one SlABC (Solyc11g065350.1) contained the SARE motif, a SA-responsive element. From the above discussion, it is evident that all of the SlABCs contain at least one stress-responsive cis-regulatory element, suggesting their role in combating a variety of biotic and abiotic stress to which plant is constantly exposed due to their sessile nature.
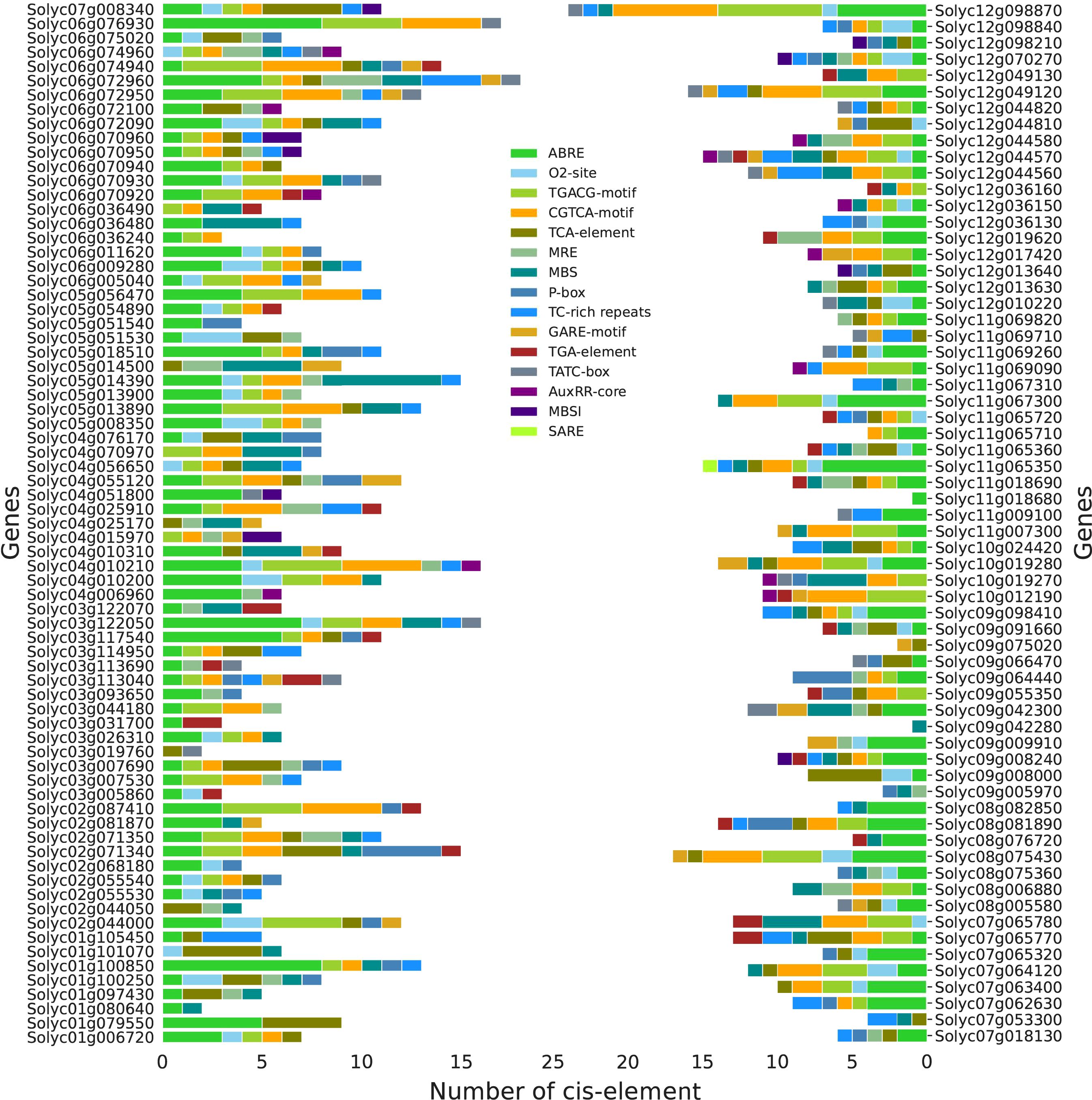
Figure 6. Identified Cis-regulatory modules in SlABCs. The legend in the middle shows the designated color for each regulatory module whereas the legend at the bottom displays the number of regulatory elements in individual SlABCs.
Protein-protein interactions (PPI) offer valuable insights into the biological functions of proteins, as those involved in similar pathways often interact with each other. To better understand the functional associations of SlABC transporters, a protein-protein interaction (PPI) network was constructed using STRING (Figure 7A). The analysis revealed a well-connected network, where several SlABC transporters formed central hubs, suggesting their significant role in intracellular transport and regulatory processes. The functional enrichment of interacting proteins highlighted key biological processes, including ATP-dependent transport, xenobiotic efflux, lipid transport, and long-chain fatty acid metabolism. Notably, the presence of ABC-type xenobiotic transporters suggests that some SlABC proteins may be involved in detoxification, while interactions with lipid transporters indicate a potential role in maintaining membrane stability and homeostasis. Gene ontology (GO) enrichment further confirmed associations with transmembrane transport, ATP binding, and efflux mechanisms, emphasizing their importance in energy-dependent substrate transport.
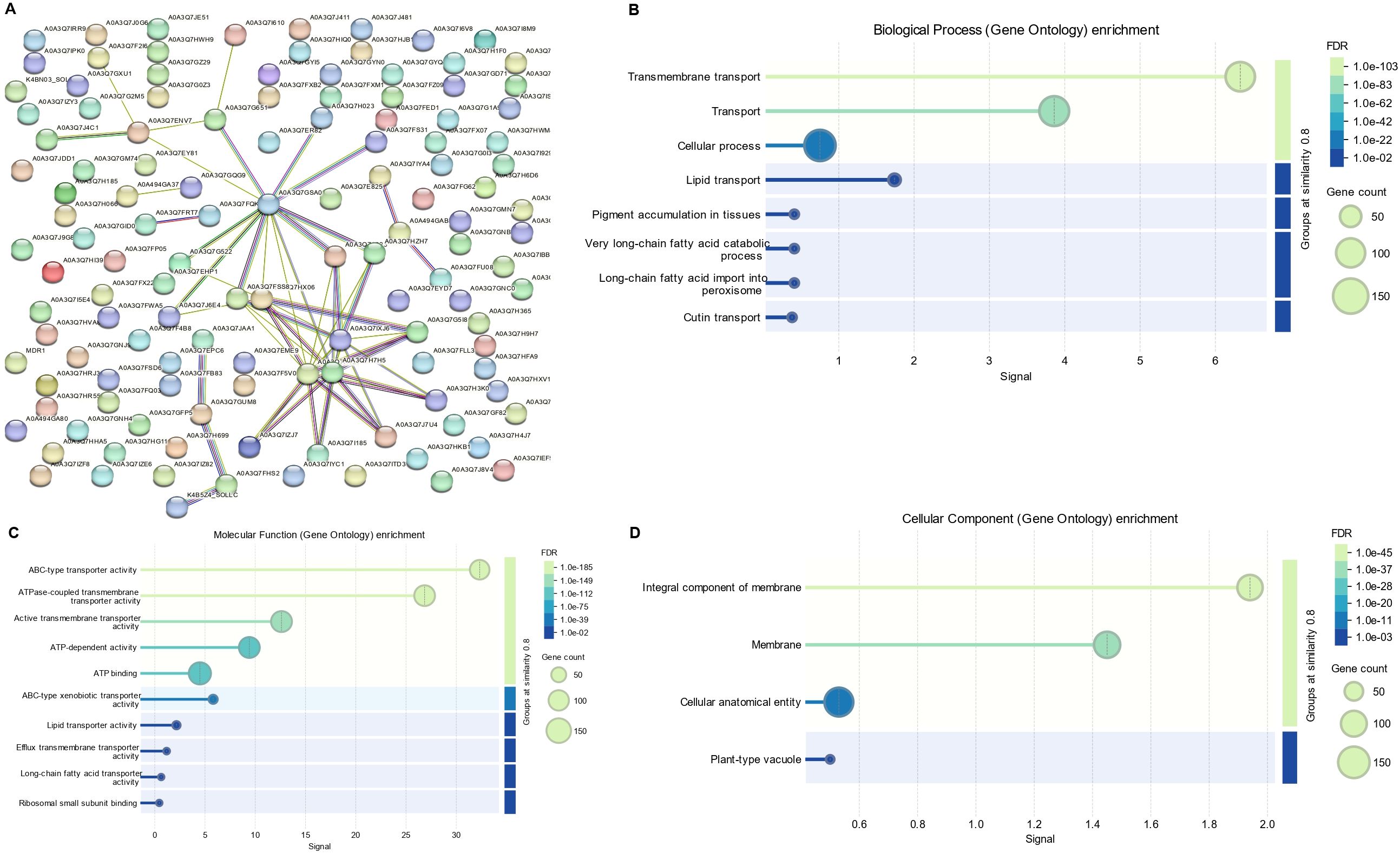
Figure 7. (A) Network of Protein-protein interaction among SlABCs based on their available information. The prediction of interacting network was carried out using STRING analysis. Different line colors represent the types of evidence for the associations (B) Gene ontology (GO) enrichment of SlABCs representing biological process (C) Gene ontology (GO) enrichment of SlABCs representing molecular functions (D) Gene ontology (GO) enrichment of SlABCs representing cellular components. The sizes of the circles indicate the number of genes in each category, while the x-axis bars indicate fold enrichment.
These findings prove that SlABC transporters are functionally diverse and play crucial roles in cellular transport, stress adaptation, and metabolic processes in S. lycopersicum. The PPI network suggests that certain SlABC transporters act as key regulators of nutrient transport and environmental stress responses, potentially contributing to overall plant resilience and development.
To predict the annotated function of the identified SlABCs, Gene Ontology (GO) enrichment analysis was carried out through online GO tools. As evident in Figure 7, the SlABCs were involved in a number of biological (Figure 7B), molecular (Figure 7C), and cellular components (Figure 7D). Key biological processes include transport (GO:0006810), localization (GO:0051179), the establishment of localization (GO:0051234), and transmembrane transport (GO:0055085), whereas molecular functions included a wide range of activities like active transmembrane transporter activity (GO:0022804), ATP binding (GO:0005524), adenyl ribonucleotide binding (GO:0032559), purine ribonucleoside triphosphate binding (GO:0035639), transmembrane transporter activity (GO:0022857), ribonucleotide binding (GO:0032553), carbohydrate derivative binding (GO:0097367), anion binding (GO:0043168) and ABC-type transporter activity (GO:0140359) etc.
Plants in their natural habitat had to face a number of extreme conditions like drought, salinity, and heavy metal-induced stress. A variety of ABC isoforms are well known for their role in the reclamation of heavy metal-induced stress; however, the response of SlABCs to Cd-induced stress has not been explored to the best of our knowledge. In order to address the existing research gap, the S. lycopersicum plantlets were exposed to two different concentrations of Cd (1 mM and 2 mM) and expression levels of targeted SlABCs was evaluated using RNA-seq analysis (Figure 8). About 105 selected SlABCs were found to be expressed during Cd stress, with several genes showing differential expression patterns. 15 SlABCs were expressed in response to 1mM Cd stress alone, 8 SlABCs were expressed in response to 2 mM Cd, and 50 SlABCs showed expression in both 1mM and 2mM Cd stress (Figure 8B). In the case of 1mM Cd stress, 10 SlABCs were upregulated, whereas 5 SlABCs were downregulated. Similarly, 3 SlABCs were upregulated, and 5 SlABCs were downregulated in response to 2mM Cd stress, whereas 35 SlABCs were upregulated and 15 SlABCs were downregulated in response to both 1mM and 2mM Cd stress (Figure 8C).
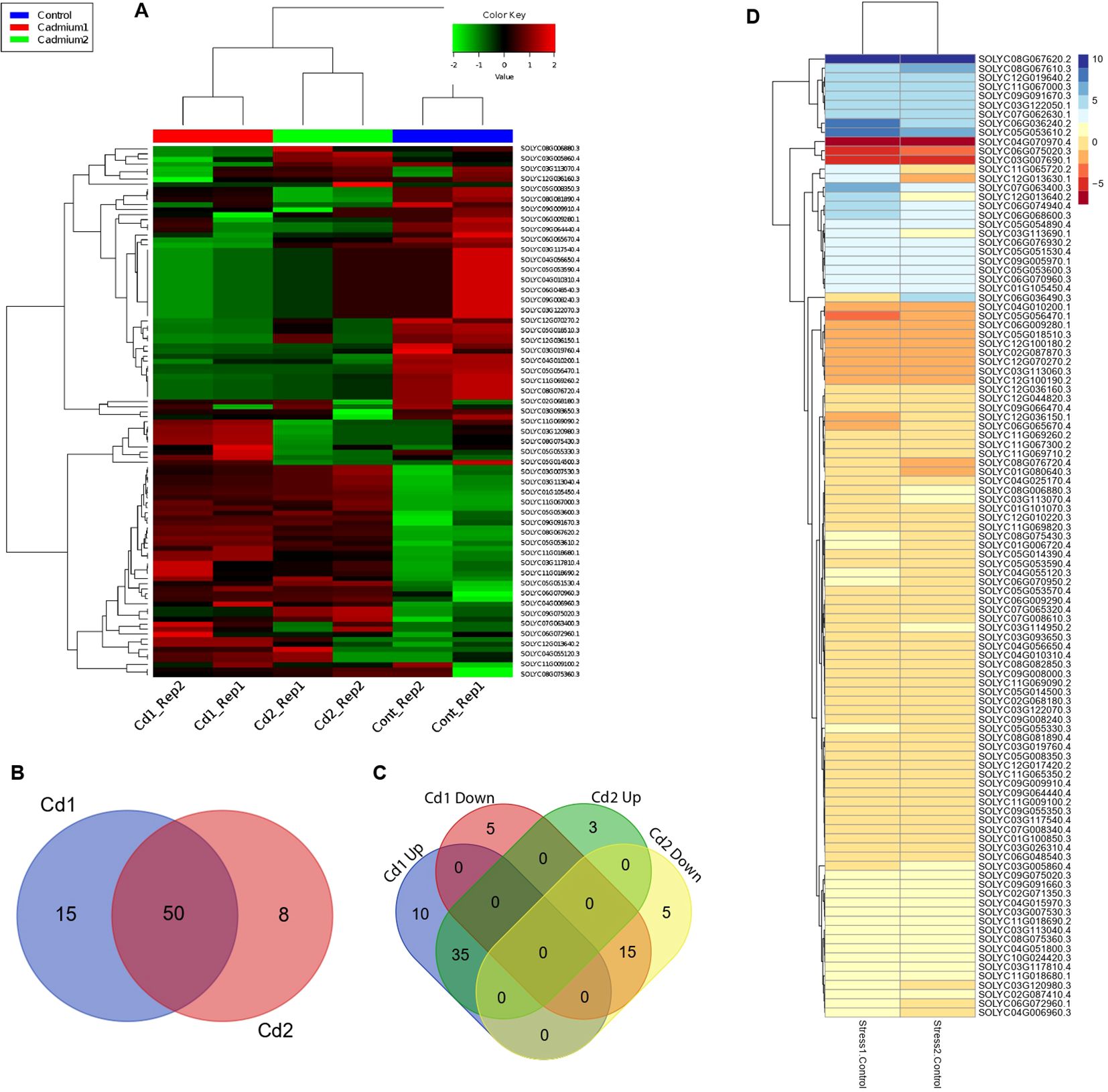
Figure 8. (A) Hierarchical clustering indicate the substantial difference in SlABCs genes induced by 1 mM and 2 mM Cd stress (B) Venn Diagram showing the number of SlABCs that were upregulated in response to Cd1 (1mM Cd) and Cd2 (2mM Cd) (C) Venn diagram showing the number of differentially expressed SlABC gene in plants exposed to Cd1 (1mM Cd) and Cd2 (2mM Cd) (D) Heatmap demonstrating the differentially expressed gene pattern. Expression changes in the genes are represented by the color scheme displayed in the legend.
Using specific primers, the RNA-seq-based gene expression data was validated by subjecting 10 SlABCs, i.e., Solyc03g007690.1, Solyc04g010200.1, Solyc04g070970.4, Solyc05g053610.2, Solyc05g056470.1, Solyc06g036240.2, Solyc06g075020.3, Solyc07g063400.3, Solyc08g067610.3, and Solyc08g067620.2 to qRT-PCR for analyzing the relative gene expression levels. There was a strong correlation between gene expression results obtained by both RT-PCR and qRT-PCR. For instance, 5 SlABCs i.e. Solyc03g007690.1, Solyc04g010200.1, Solyc04g070970.4, Solyc05g056470.1 and Solyc06g075020.3 were downregulated in both RNA-seq (Figure 8D) and qRT-PCR (Figure 9) data whereas the other 5 SlABCs i.e. Solyc05g053610.2, Solyc06g036240.2, Solyc07g063400.3, Solyc08g067610.3, and Solyc08g067620.2 were all upregulated in both RNA-seq and qRT-PCR based gene expression analysis thereby validating the accuracy our results.
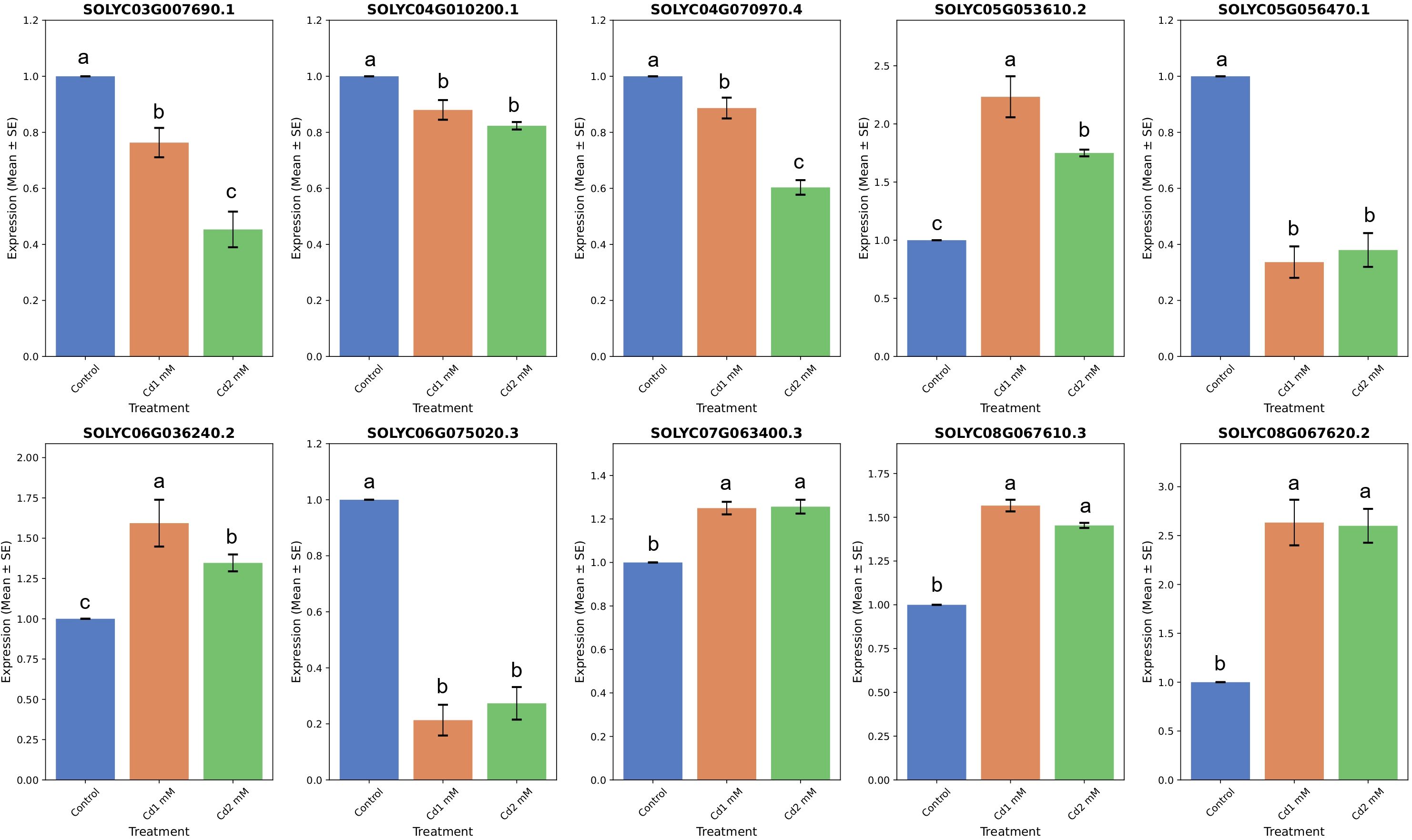
Figure 9. qRT-PCR based quantification of selected SlABCs for the validation of RT-PCR results. The x-axis displays the type of treatment, whereas the y-axis displays the gene expression level. The data is expressed in terms of Mean ± SE. Different letters over bar indicates significant difference in treatment according to Duncan's multiple range test DMRT p≤ 0.05).
Plants are among organisms that are extremely sensitive to any change in the environment. Change in temperature, humidity, salinity, rainfall patterns radiation etc., negatively impacts the growth of plants (Georgieva and Vassileva, 2023). This incurs a number of problems as plants are exposed to a number of biotic and abiotic stress conditions throughout their life (Nawaz et al., 2023). Due to the sessile nature of plants, abiotic stress factors like extreme temperature, salinity, drought, nutrient deficiency, floods, and elevated levels of HMs are detrimental to plants’ growth, survival, and overall yields (Zhang et al., 2023). Plants respond to abiotic stress by involving a number of complex biochemical pathways controlled by a plethora of genes (Javaid et al., 2022). Among them, ABCs are well known for their significant role in the management of abiotic stress (Dahuja et al., 2021). In a detailed study on Pyrus bretchneideri, 82 ABC genes were found to be involved in salt stress, and 91 genes were identified as drought-responsive genes, proving the significant role of ABC genes in response to abiotic stress (Kou et al., 2024). Yu et al. reported the upregulation of ABCB genes in response to different abiotic stresses like drought, salinity, and cold (Yu et al., 2017). Another study revealed the regulation of 51 ABCs upon exposure to abiotic stress (Nguyen et al., 2014). In Glycyrrhiza glabra, 7 out of 9 ABCs studied showed upregulation in response to drought stress (Devi et al., 2024) suggesting active participation of ABC genes in abiotic stress conditions. ABCs have also been reported for their role in heavy metal tolerance. Guo et al. reported the regulation of ABCs in response to different concentrations of lead (Guo et al., 2022). Arsenic toxicity in rice was reported to be mitigated by the active transport of arsenic to the vacuole for detoxification by ABCs (Rekha et al., 2021). In rice, 47 ABCs were upregulated in response to HMs (Nguyen et al., 2014).
Although not much work has been done on the role of ABCs in mitigating abiotic stress in S. lycopersicum, the above literature strongly suggests that they play a vital role in managing abiotic stress in plants. The current study, therefore, explored the role of ABC genes in promoting heavy metal tolerance in S. lycopersicum. A total of 154 SlABCs were identified which belong to 15 different groups based on phylogenetic analysis. The identified SlABCs were distributed across all 12 chromosomes of S. lycopersicum. Within the cell, these SlABCs were found to be distributed across all the major organelles of the cells like vacuole, nucleus, mitochondria, endoplasmic reticulum and golgi complexes. Both Golgi bodies and the endoplasmic reticulum plays a vital roles in the transportation of materials within the cells (Mohan et al., 2023). One of the main mechanism involved in HMs tolerance is its sequestration into vacuole to avoid oxidative damage caused by HMs (Zhang et al., 2018). The presence of SlABCs in all of these vital organelles may therefore enable the movement of Cd with the help of various cellular organelles into the vacuole.
Our study revealed the presence of 1150 hormone-responsive elements in SlABCs, including ABA, SA, and MeJA-responsive elements, all of which have been reported for their active participation in the regulation of heavy metal stress (Pompeu et al., 2017; Guo et al., 2018; Lu et al., 2020; Gupta and Seth, 2021; Kamran et al., 2021; Wei et al., 2021; Fan et al., 2022; Wei et al., 2022; Sun et al., 2023; Wu et al., 2023; Qin et al., 2024) suggesting that SlABCs did play critical role in Cd tolerance by hormonal transport. In a major study, transcriptome analysis of 29,921 differentially expressed genes revealed that the majority of these genes were responsible for signal transduction and transport of plant hormones and antioxidant processes (Li et al., 2023) which further signifies the importance of these hormones in combating HMs stress. GO analysis of the SlABCs revealed their involvement in a number of biological, molecular, and cellular functions, including the establishment of localization, ATP binding activity, transmembrane transport, and ABC-type transporter activity. Previous studies on soybean also revealed active involvement of ABC transporters in establishment of localization, ATPase activity, cellular transport and nucleotide binding (Naaz et al., 2023). The protein-protein interaction analysis (Figure 7A) further supports the functional significance of SlABC transporters, revealing their involvement in ATP-driven transport, xenobiotic detoxification, and lipid metabolism. The identification of central hub proteins suggests that some SlABC genes play key regulatory roles, particularly in stress adaptation and intracellular transport processes in S. lycopersicum.
Differential expression of ABCs in response to Cd in many plants has been reported previously. In rice, a G-type ABC transporter OsABCG36 was upregulated in root samples exposed to Cd. When OsABCG36 was knocked out, the plants demonstrated higher sensitivity to Cd stress (Fu et al., 2019). Similar results were reported in another study conducted on A. thaliana. The ABCG36 gene from poplar (PtoABCG36) was transformed into A. thaliana. The resulting transgenic A. thaliana resulted in improved Cd accumulation and resistance to Cd stress (Wang et al., 2019). Naaz et al (Naaz et al., 2023), explored potential role of ABCs from soybean (GmABCs) in mitigation of Cd stress. Relative expression analysis of 10 GmABCs revealed up to 50-folds greater expression in plants exposed to Cd as compared to control. Out of these, 7 GmABCs were observed to be upregulated and 3 GmABCs were downregulated upon exposure to Cd. In an experiment on A. thaliana, an AtABCC3 defective and overexpressed variety were subjected to different Cd concentrations. Whereas AtABCC3 defective variety showed increased sensitivity to Cd even at low concentrations, the AtABCC3 overexpression resulted in Cd sequestration into vacuole offering Cd tolerance (Brunetti et al., 2015). Similar results were reported for AtABCC1 and AtABCC2 defective varieties (Park et al., 2012), which signifies the role of ABCs in conferring Cd tolerance.
In the current study, RNA-seq data analysis revealed the regulation of a number of SlABCs in response to Cd stress. Both 1 mM and 2mM Cd concentration resulted in regulation of SlABCs which indicates its potential involvement in Cd stress. About 10 highly expressed SlABCs were subjected to qRT-PCR, which showed expression patterns similar to those in RNA-seq, which validates the accuracy of our results. Bovet et al (Bovet et al., 2005). explored ABC genes in A. thaliana for their possible role in Cd tolerance. A number of ABCs were reportedly expressed in plants subjected to Cd stress, highlighting the possible role of AtABCs in Cd stress. Our study revealed a number of orthologs between S. lycopersicum and A. thaliana (Supplementary Figure S1), validating the role of SlABCs in Cd stress. Expression data in our study revealed Solyc08g006880.3 upregulation in response to Cd stress, which is an ortholog of AT1G30400.1 and AT2G34660.1 (Figure 2; Supplementary Figure S1). Both these AtABCs are known to be involved in the detoxification of Cd (Park et al., 2012). Upregulation of Solyc03g120980.3 in plants exposed to Cd was also observed, which is in the same phylogenetic clade (Figure 2) as AT1G59870.1, a gene previously reported for its role in Cd tolerance in A. thaliana (Kim et al., 2007). Solyc12g044820.3, expressed in response to Cd stress, shares the same clade with AT3G21250.4, previously reported in Cd tolerance in A. thaliana (Gaillard et al., 2008). Solyc05g014390.2 and Solyc07g065320.2 were also expressed in response to different concentrations of Cd. Both these SlABCs are present in the same clade as AT3G13080.1 (Figure 2), which has been previously reported for its active involvement in the transport of and detoxification of Cd (Bovet et al., 2005), thus validating our results.
Despite useful insights on the role of ABC transporter proteins in mitigating Cd stress in S. lycopersicum, there are certain limitations. The experimental analysis was conducted in a controlled environment in growth room with specific conditions however, in the natural environment, plants have to cope with variety of adverse conditions like temperature variation, scarcity or abundance of water, salinity, radiations and availability of nutrients (Mareri et al., 2022). Such adverse conditions, combined with Cd stress may affect the differential expression of SlABCs. These factors also need to be brought under consideration in future. For comprehensive understanding of complex physiological, biochemical and genetic pathways involved in mitigation of Cd stress in S. lycopersicum, further research exploring influence of epigenetic changes is of paramount importance. With comprehensive knowledge of how the differential gene expression may be effected in the natural environment, tailor-made plant breeding strategies could be employed for production of cultivars with high yields despite unfavorable conditions.
This study identified 154 S. lycopersicum ABC transporter genes (SlABCs), categorized into 15 groups, with evolutionary conservation and functional diversity. Chromosomal mapping revealed an uneven distribution, with the highest density on chromosome 12. Gene structure and motif analyses highlighted their complexity and diverse functions. Synteny and collinearity analyses demonstrated that duplication events and purifying selection significantly shaped this gene family. Cis-regulatory element analysis linked SlABCs to stress responses, particularly ABA, MeJA, and SA pathways. RNA-Seq data revealed differential expressions of several SlABCs under Cd stress, with specific genes showing significant upregulation. qRT-PCR further validated these findings, confirming their involvement in Cd detoxification. With these findings, further research into genome editing targeting the highly expressed SlABCs could be employed for production of varieties with superior yields. Gene Ontology enrichment and protein interaction analyses supported their roles in transmembrane transport and cellular homeostasis. Overall, SlABCs are critical in stress adaptation, particularly in heavy metal detoxification, offering potential targets for genetic and biotechnological interventions to enhance stress resilience in S. lycopersicum and other crops. Insights from the current study could prove useful for plant breeders to develop cultivars that could withstand high levels of HMs including Cd.
The data presented in the study are deposited in the National Center for Biotechnology Information (NCBI) repository, accession number PRJNA913645.
SH: Data curation, Formal Analysis, Writing – original draft. L: Conceptualization, Writing – original draft, Writing – review & editing. SB: Conceptualization, Methodology, Writing – review & editing. RJ: Investigation, Methodology, Writing – review & editing. SAsi: Data curation, Methodology, Writing – original draft, Writing – review & editing. AMMA: Resources, Writing – review & editing. AA-H: Resources, Supervision, Writing – review & editing. K-MK: Supervision, Validation, Writing – review & editing. SAsa: Conceptualization, Data curation, Writing – original draft.
The author(s) declare that no financial support was received for the research and/or publication of this article.
This work was carried out with the support of the “Cooperative Research Program for Agriculture Science and Technology Development (Project No. RS-2024-00348677)”, Rural Development Administration, Republic of Korea. The authors extend their appreciation to the Researchers Supporting Project number (RSPD2025R978), King Saud University, Riyadh, Saudi Arabia.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fpls.2025.1536178/full#supplementary-material
Supplementary Table 1 | Details of Primers designed for selected genes
Supplementary Table 2 | Different characteristic features of SlABCs.
Supplementary Figure 1 | Ortholog clusters outlining the AtABCs (orange) and SlABCs (blue). Different clusters are labelled as 1-42 respectively.
Abiraami, T. V., Sanyal, R. P., Misra, H. S., Saini, A. (2023). Genome-wide analysis of bromodomain gene family in Arabidopsis and rice. Front. Plant Sci. 14. doi: 10.3389/fpls.2023.1120012
Ali, M., Shafiq, M., Haider, M. Z., Sami, A., Alam, P., Albalawi, T., et al. (2024). Genome-wide analysis of NPR1-like genes in citrus species and expression analysis in response to citrus canker (Xanthomonas axonopodis pv. citri). Front. Plant Sci. 15. doi: 10.3389/fpls.2024.1333286
An, L., Ma, Q., Du, J., Yu, M., Li, F., Luan, J., et al. (2019). Preliminary classification of the ABC transporter family in betula halophila and expression patterns in response to exogenous phytohormones and abiotic stresses. Forests 10, 722. doi: 10.3390/f10090722
Arshad, A., Cîmpeanu, S. M., Jerca, I. O., Sovorn, C., Ali, B., Badulescu, L. A., et al. (2024). Assessing the growth, yield, and biochemical composition of greenhouse cherry tomatoes with special emphasis on the progressive growth report. BMC Plant Biol. 24, 1002. doi: 10.1186/s12870-024-05701-5
Bovet, L., Feller, U., Martinoia, E. (2005). Possible involvement of plant ABC transporters in cadmium detoxification: a cDNA sub-microarray approach. Environ. Int. 31, 263–267. doi: 10.1016/j.envint.2004.10.011
Brunetti, P., Zanella, L., De Paolis, A., Di Litta, D., Cecchetti, V., Falasca, G., et al. (2015). Cadmium-inducible expression of the ABC-type transporter AtABCC3 increases phytochelatin-mediated cadmium tolerance in Arabidopsis. J. Exp. Bot. 66, 3815–3829. doi: 10.1093/jxb/erv185
Çakır, B., Kılıçkaya, O. (2013). Whole-genome survey of the putative ATP-binding cassette transporter family genes in Vitis vinifera. PloS One 8, e78860. doi: 10.1371/journal.pone.0078860
Chen, P., Li, Y., Zhao, L., Hou, Z., Yan, M., Hu, B., et al. (2017). Genome-wide identification and expression profiling of ATP-binding cassette (ABC) transporter gene family in pineapple (Ananas comosus (L.) merr.) reveal the role of acABCG38 in pollen development. Front. Plant Sci. 8, 2150. doi: 10.3389/fpls.2017.02150
Collins, E. J., Bowyer, C., Tsouza, A., Chopra, M. (2022). Tomatoes: an extensive review of the associated health impacts of tomatoes and factors that can affect their cultivation. Biol. (Basel) 11. doi: 10.3390/biology11020239
Dahuja, A., Kumar, R.R., Sakhare, A., Watts, A., Singh, B., Goswami, S., et al. (2021). Role of ATP-binding cassette transporters in maintaining plant homeostasis under abiotic and biotic stresses. Physiol. Plant 171, 785–801. doi: 10.1111/ppl.v171.4
Devi, R., Goyal, P., Verma, B., Hussain, S., Chowdhary, F., Arora, P., et al. (2024). A transcriptome-wide identification of ATP-binding cassette (ABC) transporters revealed participation of ABCB subfamily in abiotic stress management of Glycyrrhiza glabra L. BMC Genomics 25, 315. doi: 10.1186/s12864-024-10227-z
Do, T. H. T., Martinoia, E., Lee, Y. (2018). Functions of ABC transporters in plant growth and development. Curr. Opin. Plant Biol. 41, 32–38. doi: 10.1016/j.pbi.2017.08.003
Fan, S., Wu, H., Gong, H., Guo, J. (2022). The salicylic acid mediates selenium-induced tolerance to drought stress in tomato plants. Scientia Hortic. 300, 111092. doi: 10.1016/j.scienta.2022.111092
Feng, Z., Ji, S., Ping, J., Cui, D. (2021). Recent advances in metabolomics for studying heavy metal stress in plants. TrAC Trends Analytical Chem. 143, 116402. doi: 10.1016/j.trac.2021.116402
Fu, S., Lu, Y., Zhang, X., Yang, G., Chao, D., Wang, Z., et al. (2019). The ABC transporter ABCG36 is required for cadmium tolerance in rice. J. Exp. Bot. 70, 5909–5918. doi: 10.1093/jxb/erz335
Gaillard, S., Jacquet, H., Vavasseur, A., Leonhardt, N., Forestier, C. (2008). AtMRP6/AtABCC6, an ATP-binding cassette transporter gene expressed during early steps of seedling development and up-regulated by cadmium in Arabidopsis thaliana. BMC Plant Biol. 8, 1–11. doi: 10.1186/1471-2229-8-22
Garcia, O., Bouige, P., Forestier, C., Dassa, E. (2004). Inventory and comparative analysis of rice and arabidopsis ATP-binding cassette (ABC) systems. J. Mol. Biol. 343, 249–265. doi: 10.1016/j.jmb.2004.07.093
Georgieva, M., Vassileva, V. (2023). Stress management in plants: examining provisional and unique dose-dependent responses. Int. J. Mol. Sci. 24. doi: 10.3390/ijms24065105
Guo, J., Zhou, R., Ren, X., Jia, H., Hua, L., Xu, H., et al. (2018). Effects of salicylic acid, Epi-brassinolide and calcium on stress alleviation and Cd accumulation in tomato plants. Ecotoxicology Environ. Saf. 157, 491–496. doi: 10.1016/j.ecoenv.2018.04.010
Guo, Z., Yuan, X., Li, L., Zeng, M., Yang, J., Tang, H., et al. (2022). Genome-wide analysis of the ATP-binding cassette (ABC) transporter family in zea mays L. and its response to heavy metal stresses. Int. J. Mol. Sci. 23, 2109. doi: 10.3390/ijms23042109
Gupta, O. P., Sharma, P., Gupta, R. K., Sharma, I. (2014). MicroRNA mediated regulation of metal toxicity in plants: present status and future perspectives. Plant Mol. Biol. 84, 1–18. doi: 10.1007/s11103-013-0120-6
Gupta, S., Seth, C. S. (2021). Salicylic acid alleviates chromium (VI) toxicity by restricting its uptake, improving photosynthesis and augmenting antioxidant defense in Solanum lycopersicum L. Physiol. Mol. Biol. Plants 27, 2651–2664. doi: 10.1007/s12298-021-01088-x
Iqbal, Z., Iqbal, M. S., Hashem, A., Abd_Allah, E. F., Ansari, M. I. (2021). Plant defense responses to biotic stress and its interplay with fluctuating dark/light conditions. Front. Plant Sci. 12. doi: 10.3389/fpls.2021.631810
Javaid, M. H., Khan, A. R., Salam, A., Neelam, A., Azhar, W., Ulhassan, Z., et al. (2022). Exploring the adaptive responses of plants to abiotic stresses using transcriptome data. Agriculture 12, 211. doi: 10.3390/agriculture12020211
Kamran, M., Danish, M., Saleem, M. H., Malik, Z., Parveen, A., Abbasi, G. H., et al. (2021). Application of abscisic acid and 6-benzylaminopurine modulated morpho-physiological and antioxidative defense responses of tomato (Solanum lycopersicum L.) by minimizing cobalt uptake. Chemosphere 263, 128169. doi: 10.1016/j.chemosphere.2020.128169
Kang, J., Park, J., Choi, H., Burla, B., Kretzschmar, T., Lee, Y., et al. (2011). Plant ABC transporters. The Arabidopsis book/American Society of Plant Biologists 9, e0153.
Kaur, R., Das, S., Bansal, S., Singh, G., Sardar, S., Dhar, H., et al. (2021). Heavy metal stress in rice: Uptake, transport, signaling, and tolerance mechanisms. Physiol. Plant 173, 430–448. doi: 10.1111/ppl.13491
Khan, I., Asaf, S., Jan, R., Bilal, S., Lubna Khan, A. L., Kim, K.-M., et al. (2023a). Genome-wide annotation and expression analysis of WRKY and bHLH transcriptional factor families reveal their involvement under cadmium stress in tomato (Solanum lycopersicum L.). Front. Plant Sci. 14. doi: 10.3389/fpls.2023.1100895
Khan, I., Lubna Asaf, S., Jan, R., Bilal, S., Khan, A. L., Kim, K.-M., et al. (2023b). Dynamic interplay of WRKY, GRAS, and ERF transcription factor families in tomato-endophytic fungal symbiosis: insights from transcriptome and genome-wide analysis. Front. Plant Sci. 14. doi: 10.3389/fpls.2023.1181227
Kim, D. Y., Bovet, L., Maeshima, M., Martinoia, E., Lee, Y. (2007). The ABC transporter AtPDR8 is a cadmium extrusion pump conferring heavy metal resistance. Plant J. 50, 207–218. doi: 10.1111/j.1365-313X.2007.03044.x
Kou, X., Zhao, Z., Xu, X., Li, C., Wu, J., Zhang, S. (2024). Identification and expression analysis of ATP-binding cassette (ABC) transporters revealed its role in regulating stress response in pear (Pyrus bretchneideri). BMC Genomics 25, 169. doi: 10.1186/s12864-024-10063-1
Kumar, M., Tomar, M., Bhuyan, D. J., Punia, S., Grasso, S., Sá, A. G. A., et al. (2021). Tomato (Solanum lycopersicum L.) seed: A review on bioactives and biomedical activities. BioMed. Pharmacother. 142, 112018. doi: 10.1016/j.biopha.2021.112018
Li, X., Liu, L., Sun, S., Li, Y., Jia, L., Ye, S., et al. (2023). Physiological and transcriptional mechanisms associated with cadmium stress tolerance in Hibiscus Syriacus L. BMC Plant Biol. 23, 286. doi: 10.1186/s12870-023-04268-x
Lu, Q., Chen, S., Li, Y., Zheng, F., He, B., Gu, M. (2020). Exogenous abscisic acid (ABA) promotes cadmium (Cd) accumulation in Sedum alfredii Hance by regulating the expression of Cd stress response genes. Environ. Sci. pollut. Res. 27, 8719–8731. doi: 10.1007/s11356-019-07512-w
Luo, T., Zou, T., Yuan, G., He, Z., Li, W., Tao, Y., et al. (2020). Less and shrunken pollen 1 (LSP1) encodes a member of the ABC transporter family required for pollen wall development in rice (Oryza sativa L.). Crop J. 8, 492–504. doi: 10.1016/j.cj.2019.09.001
Mallick, P. K. (2021). Medicinal values of tomato (Lycopersicon esculentum Mill.-Solanaceae). Int J. Appl Sci. Biotechnol. 9, 166–168.
Mareri, L., Parrotta, L., Cai, G. (2022). Environmental stress and plants. Int. J. Mol. Sci. 23, 5416. doi: 10.3390/ijms23105416
Mohan, A. G., Calenic, B., Ghiurau, N. A., Duncea-Borca, R. M., Constantinescu, A. E., Constantinescu, I. (2023). The golgi apparatus: A voyage through time, structure, function and implication in neurodegenerative disorders. Cells 12. doi: 10.3390/cells12151972
Naaz, S., Ahmad, N., Jameel, M. R., Al-Huqail, A. A., Khan, F., Qureshi, M. I. (2023). Impact of some toxic metals on important ABC transporters in soybean (Glycine max L.). ACS Omega 8, 27597–27611. doi: 10.1021/acsomega.3c03325
Nawaz, M., Sun, J., Shabbir, S., Khattak, W.A., Ren, G., Nie, X., et al. (2023). A review of plants strategies to resist biotic and abiotic environmental stressors. Sci. Total Environ. 900, 165832. doi: 10.1016/j.scitotenv.2023.165832
Nguyen, V. N. T., Moon, S., Jung, K.-H. (2014). Genome-wide expression analysis of rice ABC transporter family across spatio-temporal samples and in response to abiotic stresses. J. Plant Physiol. 171, 1276–1288. doi: 10.1016/j.jplph.2014.05.006
Ofori, P. A., Mizuno, A., Suzuki, M., Martinoia, E., Reuscher, S., Aoki, K., et al. (2018). Genome-wide analysis of ATP binding cassette (ABC) transporters in tomato. PloS One 13, e0200854. doi: 10.1371/journal.pone.0200854
Pang, K., Li, Y., Liu, M., Meng, Z., Yu, Y. (2013). Inventory and general analysis of the ATP-binding cassette (ABC) gene superfamily in maize (Zea mays L.). Gene 526, 411–428. doi: 10.1016/j.gene.2013.05.051
Park, S. Y., Fung, P., Nishimura, N., Jensen, D. R., Fujii, H., Zhao, Y., et al. (2009). Abscisic acid inhibits type 2C protein phosphatases via the PYR/PYL family of START proteins. Science 324, 1068–1071. doi: 10.1126/science.1173041
Park, J., Song, W. Y., Ko, D., Eom, Y., Hansen, T. H., Schiller, M., et al. (2012). The phytochelatin transporters AtABCC1 and AtABCC2 mediate tolerance to cadmium and mercury. Plant J. 69, 278–288. doi: 10.1111/j.1365-313X.2011.04789.x
Pompeu, G. B., Vilhena, M. B., Gratão, P. L., Carvalho, R. F., Rossi, M. L., Martinelli, A. P., et al. (2017). Abscisic acid-deficient sit tomato mutant responses to cadmium-induced stress. Protoplasma 254, 771–783. doi: 10.1007/s00709-016-0989-4
Pourrut, B., Lopareva-Pohu, A., Pruvot, C., Garçon, G., Verdin, A., Waterlot, C., et al. (2011). Assessment of fly ash-aided phytostabilisation of highly contaminated soils after an 8-year field trial: Part 2. Influence on plants. Sci. Total Environ. 409, 4504–4510. doi: 10.1016/j.scitotenv.2011.07.047
Qin, C., Lian, H., Alqahtani, F. M., Ahanger, M. A. (2024). Chromium mediated damaging effects on growth, nitrogen metabolism and chlorophyll synthesis in tomato can be alleviated by foliar application of melatonin and jasmonic acid priming. Scientia Hortic. 323, 112494. doi: 10.1016/j.scienta.2023.112494
Rekha, K., Usha, B., Keeran, N. S. (2021). “3 - Role of ABC transporters and other vacuolar transporters during heavy metal stress in plants,” in Metal and nutrient transporters in abiotic stress. Eds. Roychoudhury, A., Tripathi, D. K., Deshmukh, R. (Academic Press), 55–76.
Sánchez-Fernández, R., Davies, T. G., Coleman, J. O., Rea, P. A. (2001). The Arabidopsis thaliana ABC protein superfamily, a complete inventory. J. Biol. Chem. 276, 30231–30244. doi: 10.1074/jbc.M103104200
Shahid, M., Pourrut, B., Dumat, C., Nadeem, M., Aslam, M., Pinelli, E. (2014). Heavy-metal-induced reactive oxygen species: phytotoxicity and physicochemical changes in plants. Rev. Environ. Contam Toxicol. 232, 1–44. doi: 10.1007/978-3-319-06746-9_1
Song, W.-Y., Park, J., Mendoza-Cózatl, D. G., Suter-Grotemeyer, M., Shim, D., Hörtensteiner, S., et al. (2010). Arsenic tolerance in Arabidopsis is mediated by two ABCC-type phytochelatin transporters. Proc. Natl. Acad. Sci. 107, 21187–21192. doi: 10.1073/pnas.1013964107
Song, W.-Y., Park, J., Eisenach, C., Maeshima, M., Lee, Y., Martinoia, E. (2014). ABC transporters and heavy metals, in plant ABC transporters. Ed. Geisler, M. (Cham: Springer International Publishing), 1–17.
Sugiyama, A., Shitan, N., Sato, S., Nakamura, Y., Tabata, S., Yazaki, K. (2006). Genome-wide analysis of ATP-binding cassette (ABC) proteins in a model legume plant, Lotus japonicus: comparison with Arabidopsis ABC protein family. DNA Res. 13, 205–228. doi: 10.1093/dnares/dsl013
Suman, J., Uhlik, O., Viktorova, J., Macek, T. (2018). Phytoextraction of heavy metals: A promising tool for clean-up of polluted environment? Front. Plant Sci. 9. doi: 10.3389/fpls.2018.01476
Sun, X., Wang, S., Tian, J., Xiang, X., Zheng, H., Liu, H., et al. (2023). Synergistic interplay between ABA-generating bacteria and biochar in the reduction of heavy metal accumulation in radish, pakchoi, and tomato. Environ. Pollut. 333, 122084. doi: 10.1016/j.envpol.2023.122084
Wang, H., Liu, Y., Peng, Z., Li, J., Huang, W., Liu, Y., et al. (2019). Ectopic expression of poplar ABC transporter ptoABCG36 confers cd tolerance in arabidopsis thaliana. Int. J. Mol. Sci. 20, 3293. doi: 10.3390/ijms20133293
Wei, T., Li, X., Yashir, N., Li, H., Sun, Y., Hua, L., et al. (2021). Effect of exogenous silicon and methyl jasmonate on the alleviation of cadmium-induced phytotoxicity in tomato plants. Environ. Sci. pollut. Res. 28, 51854–51864. doi: 10.1007/s11356-021-14252-3
Wei, T., Sun, Y., Li, H., Hua, L., Ren, X., Jia, H., et al. (2022). Methyl jasmonate triggers cadmium detoxification in tomato plants via depressing cd2+ Influx, altering the subcellular distribution, and chemical forms of cadmium. Water Air Soil pollut. 233, 460. doi: 10.1007/s11270-022-05934-w
Wu, H., Fan, S., Gong, H., Guo, J. (2023). Roles of salicylic acid in selenium-enhanced salt tolerance in tomato plants. Plant Soil 484, 569–588. doi: 10.1007/s11104-022-05819-1
Xiong, T., Zhang, T., Dumat, C., Sobanska, S., Dappe, V., Shahid, M., et al. (2019). Airborne foliar transfer of particular metals in Lactuca sativa L.: translocation, phytotoxicity, and bioaccessibility. Environ. Sci. Pollut. Res. Int. 26, 20064–20078. doi: 10.1007/s11356-018-3084-x
Yang, H., Murphy, A. S. (2009). Functional expression and characterization of Arabidopsis ABCB, AUX 1 and PIN auxin transporters in Schizosaccharomyces pombe. Plant J. 59, 179–191. doi: 10.1111/j.1365-313X.2009.03856.x
Yu, C., Dong, W., Zhan, Y., Huang, Z.-A., Li, Z., Kim, I. S., et al. (2017). Genome-wide identification and expression analysis of ClLAX, ClPIN and ClABCB genes families in Citrullus lanatus under various abiotic stresses and grafting. BMC Genet. 18, 33. doi: 10.1186/s12863-017-0500-z
Zhang, Y., Xu, J., Li, R., Ge, Y., Li, Y., Li, R. (2023). Plants’ Response to abiotic stress: mechanisms and strategies. Int. J. Mol. Sci. 24. doi: 10.3390/ijms241310915
Zhang, J., Martinoia, E., Lee, Y. (2018). Vacuolar transporters for cadmium and arsenic in plants and their applications in phytoremediation and crop development. Plant Cell Physiol. 59, 1317–1325. doi: 10.1093/pcp/pcy006
Keywords: Solanum lycopersicum, heavy metals, ABC transporters, phylogenetic analysis, gene expression
Citation: Hashmi SS, Lubna, Bilal S, Jan R, Asif S, Abdelbacki AMM, Kim K-M, Al-Harrasi A and Asaf S (2025) Exploring the role of ATP-binding cassette transporters in tomato (Solanum lycopersicum) under cadmium stress through genome-wide and transcriptomic analysis. Front. Plant Sci. 16:1536178. doi: 10.3389/fpls.2025.1536178
Received: 28 November 2024; Accepted: 24 February 2025;
Published: 18 March 2025.
Edited by:
Wajid Zaman, Yeungnam University, Republic of KoreaReviewed by:
Yanping Jing, Beijing Forestry University, ChinaCopyright © 2025 Hashmi, Lubna, Bilal, Jan, Asif, Abdelbacki, Kim, Al-Harrasi and Asaf. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Sajjad Asaf, c2FqYWRhc2lmMjAwMEBnbWFpbC5jb20=; Kyung-Min Kim, a2ttQGtudS5hYy5rcg==
†These authors have contributed equally to this work
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.