
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
SYSTEMATIC REVIEW article
Front. Physiol. , 02 November 2017
Sec. Integrative Physiology
Volume 8 - 2017 | https://doi.org/10.3389/fphys.2017.00814
This article is part of the Research Topic Integrative Approaches to the Molecular Physiology of Inflammation View all 26 articles
The growth arrest-specific transcript 5 (GAS5) is a long noncoding RNA with low expression in multiple cancers. This meta-analysis aims to explore the association between GAS5 expression levels and cancer patients' prognosis. We collected all the relevant literatures about GAS5 expression levels associated with overall survival (OS), lymph node metastasis (LNM) and high tumor stage (II/III/IV) (HTS) from the PubMed and Web of Science. The hazard ratio (HR) and the corresponding 95% confidence interval (CI) were calculated to evaluate the link strength between GAS5 and cancer prognosis. A total of 934 patients from 14 studies were included to the present meta-analysis, according to the inclusion and exclusion criteria. The results demonstrated that low expression of GAS5 could predict poor OS in cancer patients (HR = 1.955, 95% CI: 1.551–2.465, P < 0.001). Meanwhile we also analyzed the following cancers independently: hepatocellular carcinoma (HR = 1.893, 95% CI: 1.103–3.249, P = 0.021) and urothelial carcinoma (HR = 1.653, 95% CI: 1.185–2.306, P = 0.003). Compared to the high GAS5 expression group, additionally, patients with low GAS5 expression in tumor tissues were more prone to lymph node metastasis (OR = 0.234, 95%CI: 0.153–0.358, P < 0.001) and high tumor stage (OR = 0.185, 95% CI:0.102–0.333, P < 0.001). In conclusion, this meta-analysis showed that GAS5 might be served as a novel biomarker for predicting prognosis in various types of cancers.
Cancer has become a global health problem. In recent years, the incidence of cancer has been increased year by year. According to WHO estimates, 14.1 million new cancer patients and 8.2 million deaths from cancer occurred worldwide in 2012 and more than 20 million new cases of cancer will be expected as early as 2025 (Ferlay et al., 2015). At present, cancer treatment includes surgery, radiotherapy, chemotherapy andetc., but the 5 years survival rate is still not ideal, especially some patients with lymph node metastasis or high stage tumor (Saika and Sobue, 2013). Therefore, it is important to find a new biological target that plays a guiding role in the carcinogenesis to detect cancer. It is also more conducive to early detection, early diagnosis and early treatment of tumor patients.
Long non-coding RNAs (lncRNAs) are noncoding RNAs with a length of more than 200 nucleotides that regulate gene expression (Mattick and Makunin, 2006). They were described as “noise,” and did not attract much attention in the past few decades (Ponjavic et al., 2007). With the application of whole genome sequencing and microarray, lncRNAs have attracted more and more attentions (Batista and Chang, 2013; Tang et al., 2013). The increasing evidence show that lncRNAs play a pivotal role in the development and progression of tumors, which means that they can be used as biomarkers for some tumors (Fang et al., 2017; Liu et al., 2017; Sun et al., 2017; Yang et al., 2017). However, only a few number of lncRNAs have corresponding functional features, and most of the functions of lncRNAs remain unclear.
The growth arrest-specific transcript 5 (GAS5) is a rising star among tumor-suppressive lncRNAs among all the kinds of lncRNAs (Ma et al., 2016). Recent studies have shown that GAS5 plays a key role in a variety of human diseases and participates a variety of biological processes, such as cell proliferation, cell apoptosis, epithelial-mesenchymal transition and etc. (Tan et al., 2017; Tao et al., 2017; Wen et al., 2017; Yang et al., 2017). Meanwhile, GAS5 is also involved in the progression of many types of cancer, such as bladder cancer (BC) (Zhang et al., 2017), colorectal cancer (CRC) (Yin et al., 2014; Li et al., 2017), non-small cell lung cancer (NSCLC) (Shi et al., 2015; Wu et al., 2016), breast cancer (BRC) (Li W. et al., 2016), hepatocellular carcinoma (HCC) (Tu et al., 2014; Chang et al., 2016; Hu et al., 2016), epithelial ovarian cancer (EOC) (Gao et al., 2015), gastric cancer (GC) (Sun et al., 2014), cervical cancer (CEC) (Cao et al., 2014), and head and neck squamous cell carcinoma (HNSCC) (Gee et al., 2011). The clinic pathological features of the patients, such as overall survival (OS), lymph node metastasis (LNM) and high tumor stage (II/III/IV) (HTS), are also highly correlated with the level of GAS5 expression in these cancers (Gee et al., 2011; Cao et al., 2014; Sun et al., 2014; Tu et al., 2014; Yin et al., 2014; Gao et al., 2015; Shi et al., 2015; Chang et al., 2016; Hu et al., 2016; Li J. et al., 2016; Wu et al., 2016; Droop et al., 2017; Li et al., 2017; Zhang et al., 2017). All these indicate that GAS5 can be a novel prognostic biomarker in unique cancer. To shed light on the relationship between GAS5 and cancer prognosis, the meta-analysis on the association between the expression of GAS5 and the prognosis of cancer is required. Although a meta-analysis has reported that the expression of GAS5 predicts poorer survival outcomes, only 4 literatures have been included in that work and the results may be incidental (Song et al., 2016). To verify the accuracy of the previous results, the present meta-analysis with 14 studies may provide a more accurate conclusion.
We searched potentially eligible literatures through PubMed, Web of Science to locate articles (published during March 2011 to April 2017), including articles referenced in the publications. We used “GAS5 or growth arrest specific 5” AND “cancer or tumor or carcinomas or neoplasm” as the keywords, in order to identify potentially relevant studies. Citation lists of retrieved articles were searched manually to ensure sensitivity of the search strategy.
All the eligible study data elements were independently assessed and extracted by two investigators. For inclusion in this meta-analysis, the studies met the following criteria: the association between GAS5 and cancer prognosis (OS) was investigated; patients were grouped according to the expression levels of GAS5;related clinic pathologic parameters were described, such as LNM, TNM and sufficient original data for calculating a hazard ratio (HR) with its 95% confidence interval (CI). Exclusion criteria are as the following: Duplicate publications; irrelevant to cancer, GAS5, or cancer prognosis; animal studies, letters, editorials, expert opinions, abstracts, case reports and reviews; studies without usable data.
According to the inclusion and exclusion criteria, two investigators extracted and reviewed the data independently (GQJ, XHB), and disagreements were discussed with two investigators (ZHJ, LYC) in conference. The following data were extracted: first author, publication date, country of origin, tumor type, total number of patients, number of high GAS5 expression group and low GAS5 expression group, number of patients with LNM, number of patients with HTS, detection method of GAS5 expression levels, follow-up month and cut-off values, multivariate analysis, hazard ratios (HRs), and corresponding 95% CI for OS.
Meta-analysis was performed using Stata12.0 software. Pooled hazard ratios (HRs) were extracted from the included studies; the log HR and standard error (SE) were used for aggregation of the survival results (Tierney et al., 2007). To determine the heterogeneity among the included studies, chi-square-based Q test and I2 statistics were used (Higgins et al., 2003). If the P < 0.1 or I2 > 50%, it means that significant heterogeneity existed among the included studies, thus the random-effects model was adopted to analyze the results. The fixed-effects model was applied when between-study heterogeneity was absent (P > 0.1 and I2 < 50%). The potential publication bias was assessed using the Eegg'stest and P < 0.05 was considered representative of statistically significant publication bias. Sensitivity analysis was performed by sequential omission of each individual study in order to validate the stability of outcomes in the present meta-analysis.
Two investigators (GQJ, XHB) performed the quality assessment of primary studies independently. We evaluated all eligible studies' quality by using the Newcastle-Ottawa Scale (NOS) for assessing the quality of studies in meta-analyses (Zeng et al., 2015). The higher scores indicated better methodological quality.
The initial search of the electronic database retrieved 137 literatures. After removing the duplicates, 104 articles were remained. Then we carefully screened the title and abstract, 25 literatures were excluded because the studies were irrelevant. Upon further review of the full articles, the articles with no survival outcomes, lymph node metastasis, TNM stage, animal testing and other factors were excluded. 14 articles were eventually selected for the present meta-analysis (Figure 1). A total of 934 patients were included among these studies, with a maximum sample size of 106 and a minimum sample size of 24 patients (Mean 67). The publication years of the included studies were between 2011 and 2017. In these studies, one was from UK, one was from Germany and 8 were from China. A total of 9 different types of cancer were evaluated in studies of this meta-analysis (3 hepatocellular carcinoma, 2 colorectal cancer, 2 non-small cell lung cancer, 2 urothelial carcinoma, 1 breast cancer, 1 epithelial ovarian cancer, 1 gastric cancer, 1 cervical cancer and 1 head and neck squamous cell carcinoma). The expression of GAS5 was detected by qRT-PCR and the cut-off values included in the studies were inconsistent. All diagnoses of LNM and TNM were based on pathology. Hazard ratios with the corresponding 95% CIs were extracted from the graphical survival plots and the articles. The main characteristics of the eligible studies were summarized in Table 1. The Newcastle-Ottawa Scale (NOS) confirmed that all the studies were of high quality (Table 2).
Among the included studies, 10 reported the overall survival (OS) of 730 patients according to GAS5 expression levels. In order to study the relationship between GAS5 expression level and prognosis, the fixed-effect model was used to calculate the pooled HR with corresponding 95% CI because heterogeneity analysis showed that low between-study heterogeneity among those nine studies for GAS5 expression was found (I2 = 0.0%, P(H) = 0.728). We found an inverse relationship that low expression of GAS5 might be associated with poor overall survival outcome (HR = 1.955, 95% CI:1.551–2.465, P < 0.001, fixed-effect model) (Figure 2). In a subgroup analysis of cancer sites, we also found the similar significant adverse association between levels of GAS5 and OS in the following cancers (low/high): HCC (HR = 1.893, 95% CI: 1.103–3.249, P = 0.021, P(H) = 0.902), UC (HR = 1.653, 95% CI: 1.185–2.306, P = 0.003, P(H) = 0.268) and HR for the subgroup of other cancers was 2.641 (95%CI: 1.625–4.204, P < 0.001, P(H) = 0.730). We didn't perform subgroup analyses for CRC, BRC, EOC, GC, CEC, and HNSCC, because there is only one paper investigating these associations between GAS5 and OS (Figure 2) in each cancer type. Compared with the high expression group, the low GAS5 expression group indicates a poorer OS which was confirmed statistically significant.
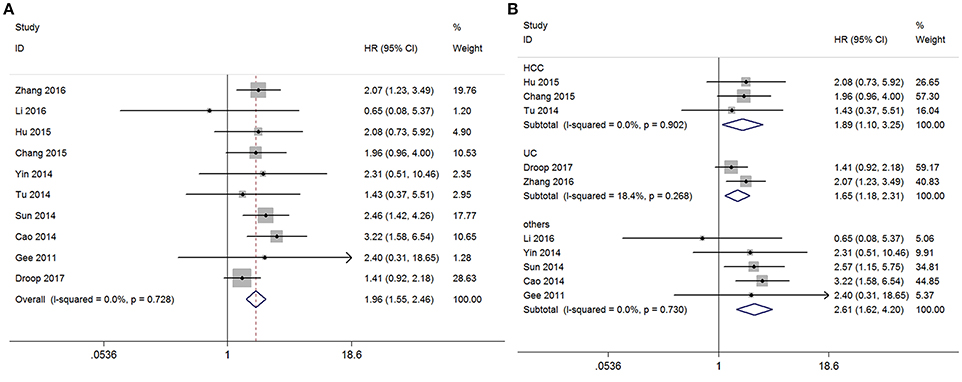
Figure 2. Meta-analysis of the pooled HRs of OS of different types of cancer with the level of GAS5 expression. (A) Forest plot for the correlation between GAS5 expression levels and OS in different cancer patients. (B) Subgroup analysis of HRs of OS by factor of different types of cancer.
Based on the differential expression levels of GAS5, seven studies reported 443 patients with lymph node metastasis. Because of the significant between-study heterogeneity (I2 = 59.6%, p = 0.021), the random-effects model was adopted to calculate the odds ratio (high GAS5 expression group vs. low GAS5 expression group; OR = 0.234, 95% CI: 0.153–0.358, P < 0.001). It demonstrated that patients with low GAS5 expression in tumor tissues were more prone to lymph node metastasis (Figure 3). In a subgroup analysis of cancer sites, we found the similar outcomes in CRC (OR = 0.353, 95% CI: 0.151–0.831, P = 0.017). OR for the subgroup of other cancers was 0.115 (95% CI: 0.06–0.221, P < 0.001). But the expression of GAS5 in NSCLC tumor tissues might not be a direct evidence of LNM (OR = 0.516, 95% CI: 0.229–1.164, P = 0.111). We didn't perform subgroup analyses for UC, BRC, HCC, EOC, GC, CEC, and HNSCC, there is only one paper investigating these associations between GAS5 and LNM (Figure 3) in each cancer type.
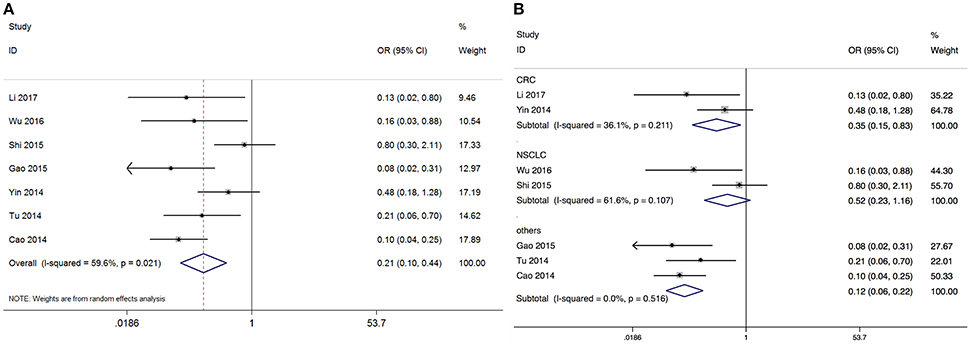
Figure 3. Meta-analysis of the LNM of different types of cancer with the level of GAS5 expression. (A) Forest plot for the correlation between GAS5 expression levels and LNM in different cancer patients. (B) Subgroup analysis of lymph node metastasis by factor of different types of cancer.
Five studies reported the HTS of 335 patients based on variousGAS5 expression levels. The fixed-effect model was adopted because there was no heterogeneity (I2 = 0.0%, p = 0.691). The odds ratio, expressed as high GAS5 expression group vs. low GAS5 expression group, was 0.185 (95% CI: 0.102–0.333, P < 0.001). The result showed that patients with low GAS5 expression in cancerous tissues were more prone to high tumor stage (Figure 4). All the results were listed in the Table 3.
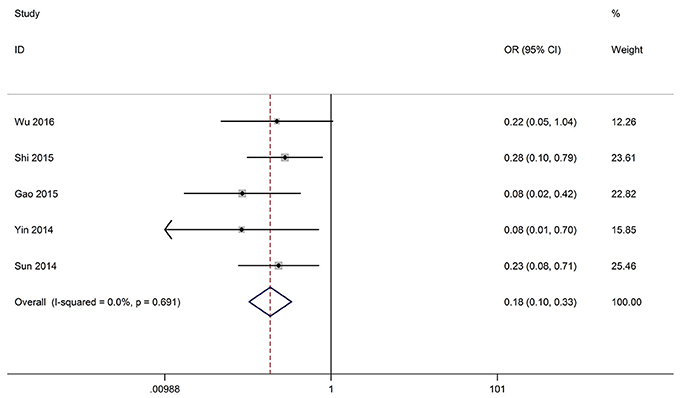
Figure 4. Forest plot for the correlation between GAS5 expression levels and HTS in different cancer patients.
To test the stability of the results of GAS5 and OS, we performed sensitivity analyses by sequentially removing each eligible study and the result was not significantly affected (Figure 5). We also performed a sensitivity analysis of lymph node metastasis and GAS5, and got similar results (Figure 5). We used Eegg's test to evaluate potential publication biases of the GAS5 and OS, and the result did not display obvious publication bias for the HR evaluations of OS (p = 0.996) (Figure 6).
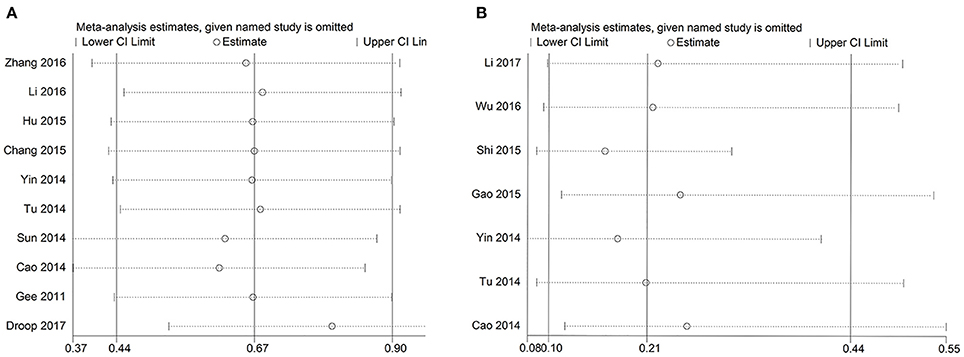
Figure 5. Sensitivity analysis of OS and LNM. (A) Sensitivity analysis of effect of individual studies on the pooled HRs for GAS5 and overall survival of patients. (B) Sensitivity analysis of effect of individual studies on ORs for GAS5 and lymph node metastasis of patients.
GAS5 has been reported to be down-regulated in multiple cancers, leading to changes in tumor cell production, proliferation, apoptosis, metastasis, and survival time (Chang et al., 2016; Hu et al., 2016; Zhang et al., 2017). In our meta-analysis, we explored the relationship between the level of GAS5 expression and cancer prognostic parameters. The results demonstrated that low expression levels of GAS5 predicted poor OS in various cancers and patients with low GAS5 expression in tumor tissues were more prone to lymph node metastasis. Meanwhile, we found patients with low GAS5 expression in cancerous tissues were more prone to high tumor stage. Our results showed that low expression levels of GAS5 could be a molecular biomarker of poor prognosis in cancer patients.
As shown in Figure 2, GAS5 and OS are positively related in OS analysis without heterogeneity and publication bias: the low expression of GAS5 predicts poorer survival outcomes. To investigate whether the above analysis was applicable in separate cancers, we made a subgroup analysis. The results was HCC (HR = 1.893, 95% CI: 1.103–3.249, P = 0.021, P(H) = 0.902), UC (HR = 1.653, 95% CI: 1.185–2.306, P = 0.003, P(H) = 0.268) which meant that the above conclusions applied equally in HCC and UC. Meanwhile, we investigated the association between the GAS5 expression levels and LNM and HTS, and we found that low GAS5 expression in cancerous tissues were more prone to LNM and HTS (Figures 3, 4). However, in LNM analysis we found that the included studies existed significantly heterogeneity. So we performed a subgroup analysis according to tumor type, and the results showed that the heterogeneity disappeared obviously in CRC(P for heterogeneity = 0.211, I2 = 36.1%, random-effects model), and other types of cancer(P for heterogeneity = 0.516, I2 = 0.0%, random-effects model), while the heterogeneity still existed in NSCLC(P for heterogeneity = 0.107, I2 = 61.6%, random-effects model) which might be caused by the different cut-off value methods which were adopted to define the high GAS5 expression group or low GAS5 expression group. In conclusion, all these results provided strong evidence for GAS5 as a potential biomarker for the prognosis of various cancers.
Nowadays, many lncRNAs have been found to be abnormally expressed in cancer. Therefore, many meta-analysis articles, like our study, have been used to reveal the correlation of lncRNAs and cancer prognosis. Several lines of studies, meanwhile, have revealed that a lot of lncRNAs play a important role in cancer prognosis, such as TUG1, SPRY4, MALAT1 (Wang et al., 2015, 2017; Yu et al., 2017). For instance, Wang et al. found that SPRY4 is remarkably upregulated in various cancer. Thus, they performed the meta-analysis to examine the association between the SPRY4-IT1 expression level and prognosis in cancer patients. Finally, they suggested the prognostic role of SPRY4-IT1 in human cancers, and increased SPRY4-IT1 expression was closely associated with advanced features of human cancers (Wang et al., 2017). Likewise, NEAT1, as a novel lncRNA, has been recently found to be up-regulated in several cancers, contributing to tumor proliferation, apoptosis, metastasis and survival. Chen et al. conduct a meta-analysis to clarify the association between high NEAT1 expression and poor prognosis. Eventually, they concluded that NEAT1 may serve as a molecular marker and a prognostic factor for patients with various cancers (Chen et al., 2017). Additionally, among these studies, it can be found that different lncRNA has specific signaling pathways in cancers. They move the extracellular signaling molecules into the cell and then, in some way, further affect cell phenotypic changes, such as cell metabolism, proliferation, invasion, apoptosis, and so on (Wang et al., 2015, 2017; Yu et al., 2017). To further investigate the value of GAS5, we analyzed and screened the signaling pathways and mechanisms of action from all GAS5 related literatures, which will be useful for future studies on tumorigenesis (Table 4).
There are several limitations in our study that should be acknowledged. Firstly, the present study used the summary data rather than a specific patient data. Secondly, the methods for distinguishing the cut-off value of GAS5 in high and low expression groups were inconsonant which inevitably could cause heterogeneity. Thirdly, most of the HR values were not directly reported in these included studies. We extracted and calculated them according to the survival curves, so inevitably there might be errors. Fourthly, different treatment methods for different types of cancer patients after surgery might have great influence on the survival time, which leaded to the heterogeneity of the researches. Fifthly, we only included English related literatures that could not be so comprehensive. Sixthly, most of the studies were from China, so the conclusion might not necessarily apply in other areas. Seventhly, we only included related studies reporting OS, LNM and HTS, and the articles on other prognostic indicators were thus excluded. In the light of the above deficiencies, a more comprehensive study covering larger samples, more regions, and more indicators will be needed to confirm our results.
In conclusion, our meta-analysis found that lncRNA GAS5 could sever as a molecular biomarker to predict the prognosis of various cancers and the low GAS5 expression could indicate the poor prognosis.
QG and HX performed Data extraction, HZ and JL did the data analysis. YL and WH designed the project and QG wrote the paper. YL supervised the project. WH provided financial support for the project.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
We are very grateful to all the colleagues who had contributed to the work and were not listed on the author list. This work was supported by the National Key Basic Research Program of China (973 Program) (2014CB745201), the Chinese High-Tech (863) Program (2014AA020607), National Natural Science Foundation of China (81402103), International S&T Cooperation program of China (ISTCP) (2014DFA31050), The National Science Foundation Projects of Guangdong Province (2014A030313717), the Shenzhen Municipal Government of China (ZDSYS201504301722174, JCYJ20150330102720130, GJHZ20150316154912494), and Special Support Funds of Shenzhen for Introduced High-Level Medical Team.
Batista, P. J., and Chang, H. Y. (2013). Long noncoding RNAs: cellular address codes in development and disease. Cell 152, 1298–1307. doi: 10.1016/j.cell.2013.02.012
Bian, D., Shi, W., Shao, Y., Li, P., and Song, G. (2017). Long non-coding RNA GAS5 inhibits tumorigenesis via miR-137 in melanoma. Am. J. Transl. Res. 9, 1509–1520.
Cao, Q., Wang, N., Qi, J., Gu, Z., and Shen, H. (2016). Long noncoding RNAGAS5 acts as a tumor suppressor in bladder transitional cell carcinoma via regulation of chemokine (CC motif) ligand 1 expression. Mol. Med. Reports 13, 27–34. doi: 10.3892/mmr.2015.4503
Cao, S., Liu, W., Li, F., Zhao, W., and Qin, C. (2014). Decreased expression of lncRNA GAS5 predicts a poor prognosis in cervical cancer. Int. J. Clin. Exp. Pathol. 7, 6776–6783.
Chang, L., Li, C., Lan, T., Wu, L., Yuan, Y., Liu, Q., et al. (2016). Decreased expression of long non-coding RNA GAS5 indicates a poor prognosis and promotes cell proliferation and invasion in hepatocellular carcinoma by regulating vimentin. Mol. Med. Rep. 13, 1541–1550. doi: 10.3892/mmr.2015.4716
Chen, Z., Xie, H., Zhong, Y., Liu, Y., and Huang, W. (2017). Role of nuclear paraspeckle assembly transcript 1 as a common molecular marker for prognosis in various cancers. Minerva. Med. 108, 477–479. doi: 10.23736/S0026-4806.17.05100-X
Dong, S., Qu, X., Li, W., Zhong, X., Li, P., Yang, S., et al. (2015). The long non-coding RNA, GAS5, enhances gefitinib-induced cell death in innate EGFR tyrosine kinase inhibitor-resistant lung adenocarcinoma cells with wide-type EGFR via downregulation of the IGF-1R expression. J. Hematol. Oncol. 8:43. doi: 10.1186/s13045-015-0140-6
Droop, J., Szarvas, T., Schulz, W. A., Niedworok, C., Niegisch, G., Scheckenbach, K., et al. (2017). Diagnostic and prognostic value of long noncoding RNAs as biomarkers in urothelial carcinoma. PLoS ONE 12:e0176287. doi: 10.1371/journal.pone.0176287
Fang, L., Sun, J., Pan, Z., Song, Y., Zhong, L., Zhang, Y., et al. (2017). Long non-coding RNA NEAT1 promotes hepatocellular carcinoma cell proliferation through the regulation of miR-129-5p-VCP-IkappaB. Am. J. Physiol. Gastrointest. Liver Physiol. 313:ajpgi.00426.02016. doi: 10.1152/ajpgi.00426.2016
Ferlay, J., Soerjomataram, I., Dikshit, R., Eser, S., Mathers, C., Rebelo, M., et al. (2015). Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int. J. Cancer 136, E359–E386. doi: 10.1002/ijc.29210
Gao, J., Liu, M., Zou, Y., Mao, M., Shen, T., Zhang, C., et al. (2015). Long non-coding RNA growth arrest-specific transcript 5 is involved in ovarian cancer cell apoptosis through the mitochondria-mediated apoptosis pathway. Oncol. Reports 34, 3212–3221. doi: 10.3892/or.2015.4318
Gee, H. E., Buffa, F. M., Camps, C., Ramachandran, A., Leek, R., Taylor, M., et al. (2011). The small-nucleolar RNAs commonly used for microRNA normalisation correlate with tumour pathology and prognosis. Br. J. Cancer 104, 1168–1177. doi: 10.1038/sj.bjc.6606076
Guo, C., Song, W. Q., Sun, P., Jin, L., and Dai, H. Y. (2015). LncRNA-GAS5 induces PTEN expression through inhibiting miR-103 in endometrial cancer cells. J. Biomed. Sci. 22:100. doi: 10.1186/s12929-015-0213-4
Higgins, J. P., Thompson, S. G., Deeks, J. J., and Altman, D. G. (2003). Measuring inconsistency in meta-analyses. Bmj 327, 557–560. doi: 10.1136/bmj.327.7414.557
Hu, L., Ye, H., Huang, G., Luo, F., Liu, Y., Liu, Y., et al. (2016). Long noncoding RNA GAS5 suppresses the migration and invasion of hepatocellular carcinoma cells via miR-21. Tum. Biol. 37, 2691–2702. doi: 10.1007/s13277-015-4111-x
Li, J., Huang, H., Li, Y., Li, L., Hou, W., and You, Z. (2016). Decreased expression of long non-coding RNA GAS5 promotes cell proliferation, migration and invasion, and indicates a poor prognosis in ovarian cancer. Oncol. Reports 36, 3241–3250. doi: 10.3892/or.2016.5200
Li, T., Mo, X., Fu, L., Xiao, B., and Guo, J. (2016). Molecular mechanisms of long noncoding RNAs on gastric cancer. Oncotarget 7, 8601–8612. doi: 10.18632/oncotarget.6926
Li, W., Zhai, L., Wang, H., Liu, C., Zhang, J., Chen, W., et al. (2016). Downregulation of LncRNA GAS5 causes trastuzumab resistance in breast cancer. Oncotarget 7, 27778–27786. doi: 10.18632/oncotarget.8413
Li, Y., Li, Y., Huang, S., He, K., Zhao, M., Lin, H., et al. (2017). Long non-coding RNA growth arrest specific transcript 5 acts as a tumour suppressor in colorectal cancer by inhibiting interleukin-10 and vascular endothelial growth factor expression. Oncotarget 8, 13690–13702 doi: 10.18632/oncotarget.14625
Liu, Q., Liu, H., Cheng, H., Li, Y., Li, X., and Zhu, C. (2017). Downregulation of long noncoding RNA TUG1 inhibits proliferation and induces apoptosis through the TUG1/miR-142/ZEB2 axis in bladder cancer cells. Onco Targets Ther. 10, 2461–2471. doi: 10.2147/OTT.S124595
Liu, X., Jiao, T., Wang, Y., Su, W., Tang, Z., and Han, C. (2016). Long non-coding RNA GAS5 acts as a molecular sponge to regulate miR-23a in gastric cancer. Minerva Med.. [Epub ahead of print].
Liu, Y., Zhao, J., Zhang, W., Gan, J., Hu, C., Huang, G., et al. (2015). lncRNA GAS5 enhances G1 cell cycle arrest via binding to YBX1 to regulate p21 expression in stomach cancer. Sci. Reports 5:10159. doi: 10.1038/srep10159
Liu, Z., Wang, W., Jiang, J., Bao, E., Xu, D., Zeng, Y., et al. (2013). Downregulation of GAS5 promotes bladder cancer cell proliferation, partly by regulating CDK6. PLoS ONE 8:e73991. doi: 10.1371/journal.pone.0073991
Luo, G., Liu, D., Huang, C., Wang, M., Xiao, X., Zeng, F., et al. (2017). LncRNA GAS5 Inhibits Cellular Proliferation by Targeting P27Kip1. Wuhan: Molecular Cancer Research (MCR).
Ma, C., Shi, X., Zhu, Q., Li, Q., Liu, Y., Yao, Y., et al. (2016). The growth arrest-specific transcript 5 (GAS5): a pivotal tumor suppressor long noncoding RNA in human cancers. Tum. Biol. 37, 1437–1444. doi: 10.1007/s13277-015-4521-9
Mattick, J. S., and Makunin, I. V. (2006). Non-coding RNA. Hum. Mol. Genet. 15, R17–R29. doi: 10.1093/hmg/ddl046
Mazar, J., Rosado, A., Shelley, J., Marchica, J., and Westmoreland, T. J. (2017). The long non-coding RNA GAS5 differentially regulates cell cycle arrest and apoptosis through activation of BRCA1 and p53 in human neuroblastoma. Oncotarget 8, 6589–6607. doi: 10.18632/oncotarget.14244
Mei, Y., Si, J., Wang, Y., Huang, Z., Zhu, H., Feng, S., et al. (2017). Long noncoding RNA GAS5 suppresses tumorigenesis by inhibiting miR-23a 5 expression in non-small cell lung cancer. Oncol. Res. 25, 1027–1037. doi: 10.3727/096504016X14822800040451
Pickard, M. R., and Williams, G. T. (2014). Regulation of apoptosis by long non-coding RNA GAS5 in breast cancer cells: implications for chemotherapy. Breast Cancer Res. Treat. 145, 359–370. doi: 10.1007/s10549-014-2974-y
Ponjavic, J., Ponting, C. P., and Lunter, G. (2007). Functionality or transcriptional noise? Evidence for selection within long noncoding RNAs. Genome Res. 17, 556–565. doi: 10.1101/gr.6036807
Renganathan, A., Kresoja-Rakic, J., Echeverry, N., Ziltener, G., Vrugt, B., Opitz, I., et al. (2014). GAS5 long non-coding RNA in malignant pleural mesothelioma. Mol. Cancer 13:119. doi: 10.1186/1476-4598-13-119
Saika, K., and Sobue, T. (2013). [Cancer statistics in the world]. Gan to kagaku ryoho. Cancer Chemother. 40, 2475–2480.
Shi, X., Sun, M., Liu, H., Yao, Y., Kong, R., Chen, F., et al. (2015). A critical role for the long non-coding RNA GAS5 in proliferation and apoptosis in non-small-cell lung cancer. Mol. Carcinogenesis 54 (Suppl. 1), E1–E12. doi: 10.1002/mc.22120
Song, W., Wang, K., Zhang, R. J., Dai, Q. X., and Zou, S. B. (2016). Long noncoding RNA GAS5 can predict metastasis and poor prognosis: a meta-analysis. Minerva Med. 107, 70–76.
Sun, M., Jin, F. Y., Xia, R., Kong, R., Li, J. H., Xu, T. P., et al. (2014). Decreased expression of long noncoding RNA GAS5 indicates a poor prognosis and promotes cell proliferation in gastric cancer. BMC Cancer 14:319. doi: 10.1186/1471-2407-14-319
Sun, Y., Zou, X., He, J., and Mao, Y. (2017). Identification of long non-coding RNAs biomarkers associated with progression of endometrial carcinoma and patient outcomes. Oncotarget. 8, 52604–52613. doi: 10.18632/oncotarget.17537
Tan, Q., Zuo, J., Qiu, S., Yu, Y., Zhou, H., Li, N., et al. (2017). Identification of circulating long non-coding RNA GAS5 as a potential biomarker for non-small cell lung cancer diagnosisnon-small cell lung cancer, long non-coding RNA, plasma, GAS5, biomarker. Int. J. Oncol. 50, 1729–1738. doi: 10.3892/ijo.2017.3925
Tang, J. Y., Lee, J. C., Chang, Y. T., Hou, M. F., Huang, H. W., Liaw, C. C., et al. (2013). Long noncoding RNAs-related diseases, cancers, and drugs. ScientificWorldJournal 2013:943539. doi: 10.1155/2013/943539
Tao, H., Zhang, J. G., Qin, R. H., Dai, C., Shi, P., Yang, J. J., et al. (2017). LncRNA GAS5 controls cardiac fibroblast activation and fibrosis by targeting miR-21 via PTEN/MMP-2 signaling pathway. Toxicology. 386, 11–18. doi: 10.1016/j.tox.2017.05.007
Tierney, J. F., Stewart, L. A., Ghersi, D., Burdett, S., and Sydes, M. R. (2007). Practical methods for incorporating summary time-to-event data into meta-analysis. Trials 8:16. doi: 10.1186/1745-6215-8-16
Tu, Z. Q., Li, R. J., Mei, J. Z., and Li, X. H. (2014). Down-regulation of long non-coding RNA GAS5 is associated with the prognosis of hepatocellular carcinoma. Int. J. Clin. Exp. Pathol. 7, 4303–4309.
Wang, D., Ding, L., Wang, L., Zhao, Y., Sun, Z., Karnes, R. J., et al. (2015). LncRNA MALAT1 enhances oncogenic activities of EZH2 in castration-resistant prostate cancer. Oncotarget 6, 41045–41055. doi: 10.18632/oncotarget.5728
Wang, M., Dong, X., Feng, Y., Sun, H., Shan, N., and Lu, T. (2017). Prognostic role of the long non-coding RNA, SPRY4 Intronic Transcript 1, in patients with cancer: a meta-analysis. Oncotarget 8, 33713–33724. doi: 10.18632/oncotarget.16735
Wen, Q., Liu, Y., Lyu, H., Xu, X., Wu, Q., Liu, N., et al. (2017). Long noncoding RNA GAS5, which acts as a tumor suppressor via microRNA 21, regulates cisplatin resistance expression in cervical cancer. Int. J. Gynecol. Cancer. 27, 1096–1108. doi: 10.1097/IGC.0000000000001028
Wu, Y., Lyu, H., Liu, H., Shi, X., Song, Y., and Liu, B. (2016). Downregulation of the long noncoding RNA GAS5-AS1 contributes to tumor metastasis in non-small cell lung cancer. Sci. Reports 6:31093. doi: 10.1038/srep31093
Xue, D., Zhou, C., Lu, H., Xu, R., Xu, X., and He, X. (2016). LncRNA GAS5 inhibits proliferation and progression of prostate cancer by targeting miR-103 through AKT/mTOR signaling pathway. Tum. Biol. doi: 10.1007/s13277-016-5429-8
Xue, Y., Ni, T., Jiang, Y., and Li, Y. (2017). LncRNA GAS5 Inhibits Tumorigenesis and Enhances Radiosensitivity By Suppressing miR-135b Expression in Non-Small Cell Lung Cancer. Oncol. Res. 25, 1305–1316. doi: 10.3727/096504017X14850182723737
Yang, Y., Shen, Z., Yan, Y., Wang, B., Zhang, J., Shen, C., et al. (2017). Long non-coding RNA GAS5 inhibits cell proliferation, induces G0/G1 arrest and apoptosis, and functions as a prognostic marker in colorectal cancer. Oncol. Lett. 13, 3151–3158 doi: 10.3892/ol.2017.5841
Yin, D., He, X., Zhang, E., Kong, R., De, W., and Zhang, Z. (2014). Long noncoding RNA GAS5 affects cell proliferation and predicts a poor prognosis in patients with colorectal cancer. Med. Oncol. 31:253. doi: 10.1007/s12032-014-0253-8
Yu, X. H., Guo, W., Zhang, J., Ma, C., Chu, A. J., Wen, B. L., et al. (2017). Long non-codingRNA (lncRNA) TUG1 and the prognosis of cancer: a meta-analysis. Cell. Mol. Biol. 63, 36–39. doi: 10.14715/cmb/2017.63.3.7
Zeng, X., Zhang, Y., Kwong, J. S., Zhang, C., Li, S., Sun, F., et al. (2015). The methodological quality assessment tools for preclinical and clinical studies, systematic review and meta-analysis, and clinical practice guideline: a systematic review. J. Evid. Based Med. 8, 2–10. doi: 10.1111/jebm.12141
Zhang, H., Guo, Y., Song, Y., and Shang, C. (2017). Long noncoding RNA GAS5 inhibits malignant proliferation and chemotherapy resistance to doxorubicin in bladder transitional cell carcinoma. Cancer Chemother. Pharmacol. 79, 49–55. doi: 10.1007/s00280-016-3194-4
Keywords: lncRNA, GAS5, cancer, prognosis, lymph node metastasis, high tumor stage, meta-analysis
Citation: Gao Q, Xie H, Zhan H, Li J, Liu Y and Huang W (2017) Prognostic Values of Long Noncoding RNA GAS5 in Various Carcinomas: An Updated Systematic Review and Meta-Analysis. Front. Physiol. 8:814. doi: 10.3389/fphys.2017.00814
Received: 19 July 2017; Accepted: 03 October 2017;
Published: 02 November 2017.
Edited by:
Carlos Rosales, National Autonomous University of Mexico, MexicoReviewed by:
Antonio Longo, Università degli Studi di Catania, ItalyCopyright © 2017 Gao, Xie, Zhan, Li, Liu and Huang. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Yuchen Liu, bGl1eXVjaGVubWRjZ0AxNjMuY29t
Weiren Huang, cG9ueTg5ODBAMTYzLmNvbQ==
†These authors have contributed equally to this work.
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.