
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Pharmacol. , 12 March 2025
Sec. Pharmacology of Infectious Diseases
Volume 16 - 2025 | https://doi.org/10.3389/fphar.2025.1511088
This article is part of the Research Topic Multidrug Resistant Bacteria: New Therapeutic Approaches for a Challenging Problem View all 3 articles
Aims: To establish a population pharmacokinetic (PopPK) model of polymyxin B (PMB) in critically ill patients based on steady-state trough (Ctrough,ss) and peak (Cpeak,ss) concentrations, optimize the dosing regimen, and evaluate the consistency of 24-hour steady-state area under the concentration-time curve (AUCss,24h) estimation between model-based and the two-point (Ctrough,ss and Cpeak,ss) methods.
Methods: PopPK modeling was performed using NONMEM, Monte Carlo simulations were used to optimize PMB dosing regimens. Bland-Altman analysis was used to evaluate the consistency between the two AUCss,24h estimation methods.
Results: A total of 95 patients, contributing 214 blood samples, were included and categorized into a modeling group (n = 80) and a validation group (n = 15). A one-compartment model was developed, with creatinine clearance (CrCL) and platelet count (PLT) identified as significant covariates influencing PK parameters. Simulation results indicated that when a Minimum Inhibitory Concentration (MIC) ≤ 0.5 mg·L-1, a probability of target attainment (PTA) ≥ 90% was achieved in all groups except for the 50 mg every 12 h (q12h) maintenance dose group. PTA decreased as CrCL increased, with slight variations observed across different PLT levels. The 75 mg and 100 mg q12h groups showed a higher proportion of AUCss,24h within the therapeutic window. Bland-Altman analysis revealed a mean bias of 12.98 mg·h·L-1 between the two AUCss,24h estimation methods. The Kappa test (κ = 0.51, P < 0.001) and McNemar’s test (P = 0.33) demonstrated moderate agreement, reflecting overall consistency with minor discrepancies in classification outcomes.
Conclusion: The PopPK model of PMB is well-suited for critically ill patients. The 75 mg q12h and 100 mg q12h regimens are appropriate for critically ill patients, with CrCL levels guiding individualized dosing. A two-point sampling strategy can be used for routine therapeutic drug monitoring (TDM) of PMB.
The global spread of carbapenem-resistant organisms (CRO) has become a major public health concern, leading to increased morbidity and mortality due to their widespread resistance to common antibiotics (Tacconelli et al., 2018; Brink, 2019). Due to the lack of effective alternatives, polymyxin B (PMB) — an antibiotic that was initially withdrawn in the 1950s due to concerns over nephrotoxicity and neurotoxicity (Poirel et al., 2017) — has been reintroduced as a last-line agent against CRO infections (Abdallah et al., 2015; Rabanal and Cajal, 2017; Zhang et al., 2020). In China, where new antimicrobial options remain scarce, PMB now serves as a cornerstone therapy for these CRO. However, uncertainties persist regarding its optimal dosing, particularly due to limited pharmacokinetic (PK) data in CRO-infected populations and the dosing regimen continues to be debated (Bergen et al., 2010; Onufrak et al., 2017; Manchandani et al., 2018). PMB was often used in critically ill patients who present with distinct physiological traits, such as higher Acute Physiology and Chronic Health Evaluation II (APACHE II) scores, lower serum albumin levels, hemodynamic instability, and significant variability in creatinine clearance (CrCL) (Roberts et al., 2014a). These factors can alter the drug’s PK parameters, like clearance and volume of distribution, potentially leading to suboptimal antibiotic concentrations at the infection site (Roberts et al., 2014a). This suboptimal drug exposure is associated with bacterial tolerance and poor outcomes (Huemer et al., 2020), underscoring the need to optimize PMB dosing in this patient population.
Several studies have utilized population pharmacokinetic (PopPK) modeling to optimize PMB dosing in critically ill patients (Sandri et al., 2013a; Luo et al., 2022; Ye et al., 2022; Liang et al., 2023; Tang et al., 2023), Sandri et al. and Liang et al. conducted studies with 24 and 22 critically ill patients, respectively, both of which had relatively small sample sizes. Luo et al. included critically ill patients with and without continuous renal replacement therapy (CRRT), but their study did not fully capture the PK characteristics of patients without CRRT. Ye et al. focused on optimizing PMB dosing in critically ill patients with varying renal function, enrolling 23 patients. Tang et al. studied critically ill patients with nosocomial pneumonia, which limited the applicability of their findings to other types of infections. International guidelines and a clinical study from the Chinese population on PMB usage recommend a concentration-time curve at steady state over 24 h (AUCss,24h) in the range of 50–100 mg·h·L-1 as the therapeutic window to ensure PMB safety and efficacy (Tsuji et al., 2019; Yang et al., 2022). The Chinese guidelines for therapeutic drug monitoring (TDM) of PMB (Liu et al., 2023) propose two methods for estimating the AUCss,24h. One approach uses steady-state trough (Ctrough,ss) and peak (Cpeak,ss) concentrations with a first-order elimination equation, while the other employs individual PK parameters from a PopPK model to estimate AUCss,24h. However, no studies have compared AUCss,24h estimates from these two methods, leaving the consistency of the results uncertain.
This study aims to: 1) to develop a PopPK model for PMB in critically ill patients with CRO infections to identify factors influencing PK variability in this population; 2) to select the optimal dosing regimen for this population based on Monte Carlo simulations of the final model; and 3) to assess the consistency between two AUCss,24h estimation methods, providing evidence for the TDM of PMB based on the two-point method using Ctrough,ss and Cpeak,ss.
A single-center prospective study was conducted in the intensive care unit (ICU) of the First Affiliated Hospital of Army Medical University from August 2021 to July 2024. Inclusion criteria: (1) patients aged ≥18 years; (2) patients receiving intravenous PMB for CRO infections confirmed by pathogen testing; (3) patients receiving at least four consecutive doses of intravenous PMB, or a loading dose followed by at least three consecutive doses. Exclusion criteria: (1) patients with missing clinical data; (2) patients hospitalized for fewer than 7 days; (3) patients receiving any form of renal replacement therapy during PMB treatment; (4) pregnant women. The research protocol was approved by the Ethics Committee of our hospital (No. (A) KY2021064).
Data collected from electronic medical records included: (1) demographic characteristics and main diseases; (2) PMB dose, administration route, and concurrent antibacterial agents; (3) routine blood test results; (4) liver and renal function indices, with CrCL calculated using the Cockcroft-Gault equation; (5) blood coagulation parameters; (6) other treatments such as extracorporeal membrane oxygenation (ECMO), mechanical ventilation, as well as relevant parameters such as the duration of these treatments.
After at least 48 h of treatment, two blood samples were collected: one immediately before the infusion and the other immediately after. The exact times of blood sampling and infusion were recorded for each patient. Plasma was separated by low-temperature, low-speed centrifugation and stored at −70°C until analysis.
PMB concentrations were analyzed using a validated ultra-performance liquid chromatography-tandem mass spectrometry (UPLC-MS/MS) method. Briefly, using polymyxin E2 as the internal standard, the assay was linear over 0.2–20.0 mg·L-1 and 0.05–5 mg·L-1 (r > 0.995) for PMB1 and PMB2 respectively. Intra-day and inter-day precision tests showed relative standard deviations (RSDs) ≤ 12.06%. The average extraction recovery ranged from 103.04% to 117.44%, and RSDs for the matrix effect and stability tests did not exceed 7.42%. PMB concentration was calculated as follows: total concentration of PMB = [PMB1 concentration/PMB1 molecular + PMB2 concentration/PMB2 molecular]*total PMB molecular (Liu et al., 2023).
The PopPK model was developed using nonlinear mixed-effects modeling software: NONMEM (version 7.5.1, ICON plc, United States), Pirana (version 23.1.2, Certara L.P., United States), and PsN (version 5.3.0, https://uupharmacometrics.github.io/PsN/), with the first-order conditional estimation method including interaction (FOCE-I). The base model was selected based on goodness-of-fit (GOF) diagnostic plots, relative standard error (RSE) of parameters, and the objective function value (OFV). Spearman correlation was used to evaluate the relationships between covariates and individual empirical Bayesian estimates (EBEs) of PK parameters before covariate selection. Covariates included age, weight, APACHE II score, the presence of sepsis, and all laboratory parameters listed in Table 1. A decrease in OFV >3.84 (P < 0.05, χ2, df = 1) for forward addition and an increase >6.63 (P < 0.01, χ2, df = 1) for backward elimination were the criteria for covariate inclusion. GOF plots were used to assess the model. To assess the stability of the final model and the precision of the PK parameters, a bootstrap method with 1,000 resampling iterations was performed. The final model was then subjected to 1,000 simulations for visual predictive checks (VPC) to evaluate the model’s predictive ability and accuracy. The accuracy and predictive performance of the model were further assessed using normalized prediction distribution errors (NPDE) plots. Statistical validation of the model was conducted using the t-test, Fisher’s test, Shapiro-Wilk test, and the Global test.
The patients were divided into a modeling group and a validation group based on the chronological order of enrollment and in accordance with the inclusion and exclusion criteria. External validation of the final model was performed using the validation group data. The mean prediction error (MPE), mean absolute prediction error (MAPE) (Equations 1, 2), F20 (20% of the absolute value of the prediction error), and F30 were calculated by comparing the predicted values with the observed values. Model performance was considered acceptable if MPE% ≤ ±20%, MAPE% ≤ 30%, F20 ≥ 35%, and F30 ≥ 50% (Mao et al., 2018).
predi denotes the i-th predicted value, and obsi its corresponding observed value.
Using the final PopPK model, 1,000 simulations were conducted for commonly used maintenance doses in ICU patients, specifically 50 mg, 75 mg, 100 mg, and 125 mg every 12 h (q12h) with a 1-hour infusion time. According to clinical research (Tang et al., 2023), PMB has demonstrated better clinical efficacy in treating CRO caused nosocomial pneumonia when AUCss,24h/minimum inhibitory concentration (MIC) ≥ 66.9. Given that the majority of patients in this study (65/80, 81.25%) had pulmonary infections, we selected AUCss,24h/MIC ≥ 66.9 as the PK/PD target, with a 90% target attainment probability (PTA) was considered to be effective. Furthermore, to mitigate the potential risk of nephrotoxicity associated with excessively high AUCss,24h, we defined the therapeutic window for AUCss,24h as 50–100 mg·h·L-1 (Tsuji et al., 2019; Yang et al., 2022). We analyzed the probability of the simulated population achieving this therapeutic window.
Data analysis was performed SPSS (version 26.0, IBM, United States) and GraphPad Prism (version 8.3.0, CA, United States). Variables with a normal distribution were expressed as mean ± standard deviation (SD), while non-normally distributed variables were reported as median (interquartile range, IQR). Categorical data were presented as percentages (%). Comparisons between two groups were conducted using the Mann-Whitney U test for non-normally distributed data and the independent t-test for normally distributed data. The estimation of AUCss,24h used the two-point method was presented in Equations 3–5 (Liu et al., 2023), while the estimation of AUCss,24h based on the PopPK model was obtained by dividing the total 24-hour drug dose by the individual clearance. Bland-Altman analysis was performed to evaluate the consistency of AUCss,24h estimates obtained by the two methods. AUCss,24h values were categorized as “within the therapeutic window” (50–100 mg·h·L-1), and McNemar’s test and Kappa test were applied to assess the consistency of AUCss,24h classifications.
Infusion time; τ: dosing interval; Csoi' is the exploratory concentration at the start of dosing based on the one-compartment linear elimination pharmacokinetic assumption; ke: elimination rate constant; n is the number of doses within 24 h.
The modeling group consisted of 80 patients with 184 PMB blood concentration samples, while the validation group included 15 patients with 30 samples. 11 patients in the modeling group underwent repeated sampling. Clinical characteristics and laboratory parameters are summarized in Table 1, showing no significant differences between the two groups. All patients were infected with CRO, in the modeling group, carbapenem-resistant Acinetobacter baumannii (CRAB) was the most common infection (70 cases), followed by carbapenem-resistant Enterobacterales (CRE; 56 cases, comprising 45 Klebsiella pneumoniae, 4 Enterobacter cloacae, 3 Serratia marcescens, 2 Escherichia coli, and 2 Citrobacter freundii), and carbapenem-resistant Pseudomonas aeruginosa (CRPA; 15 cases). In the validation group, CRE was more prevalent (11 cases) than CRAB (10 cases). In the modeling group, the MICs of PMB against CRO strains were: ≤0.5 mg·L-1 in 72.25% (57/80), 1 mg·L-1 in 13.75% (11/80), 2 mg L-1 in 10% (8/80), and 16 mg·L-1 in 3.75% (3/80). In the validation group, MIC values were ≤0.5 mg·L-1 in 60% (9/15), 1 mg·L-1 in 26.67% (4/15), and 2 mg·L-1 in 13.33% (2/15), with no statistically significant difference observed (P = 0.21).
A one-compartment model with first-order elimination best fit the population data of PMB in critically ill patients. Inter-individual variability was described using an exponential random effects model, while residual variability was described using both proportional and additive error models. Among the covariates evaluated, CrCL and platelet count (PLT) count were found to significantly influence CL, whereas no covariates had a significant effect on Vd (volume of distribution). The final PK model Equations 6, 7 is as follows:
Parameter estimates and GOF plots for the base model were presented in Supplementary Table S1 and Supplementary Figure S1 (Supplementary Material). The GOF plot (Figure 1) of the final model demonstrated strong agreement between observed and predicted values. The bootstrap median (Table 2) closely matched the population estimates of the final model, further confirming the robustness of the population PK model. The observed-versus-predicted plot showed a random scatter, indicating no systematic bias and suggesting that the model accurately describes the concentration data. The VPC plot (Figure 2) demonstrated that the median of the model predictions closely aligns with the median of the observed data, and most observed data points fall within the model’s 95% prediction interval. This indicated that the model has strong predictive ability and reasonable variability. Statistical tests for NPDE results included: t-test (P = 1), Fisher variance test (P = 1), Shapiro-Wilk normality test (P = 0.30), and Global test (P = 0.30). The NPDE histogram and Q-Q plot (Figure 3) showed that prediction errors were close to zero and symmetrically distributed, conforming to the normality assumption. NPDE showed no significant variation over time or across predicted concentrations, indicating that the model’s predictive performance is consistent and stable across various time points and concentration levels. External validation, conducted with 15 patients using the final model, showed an MPE% of 2.69%, MAPE% of 28.45%, F20 of 36.67%, and F30 of 73.33%, confirming acceptable predictive performance.
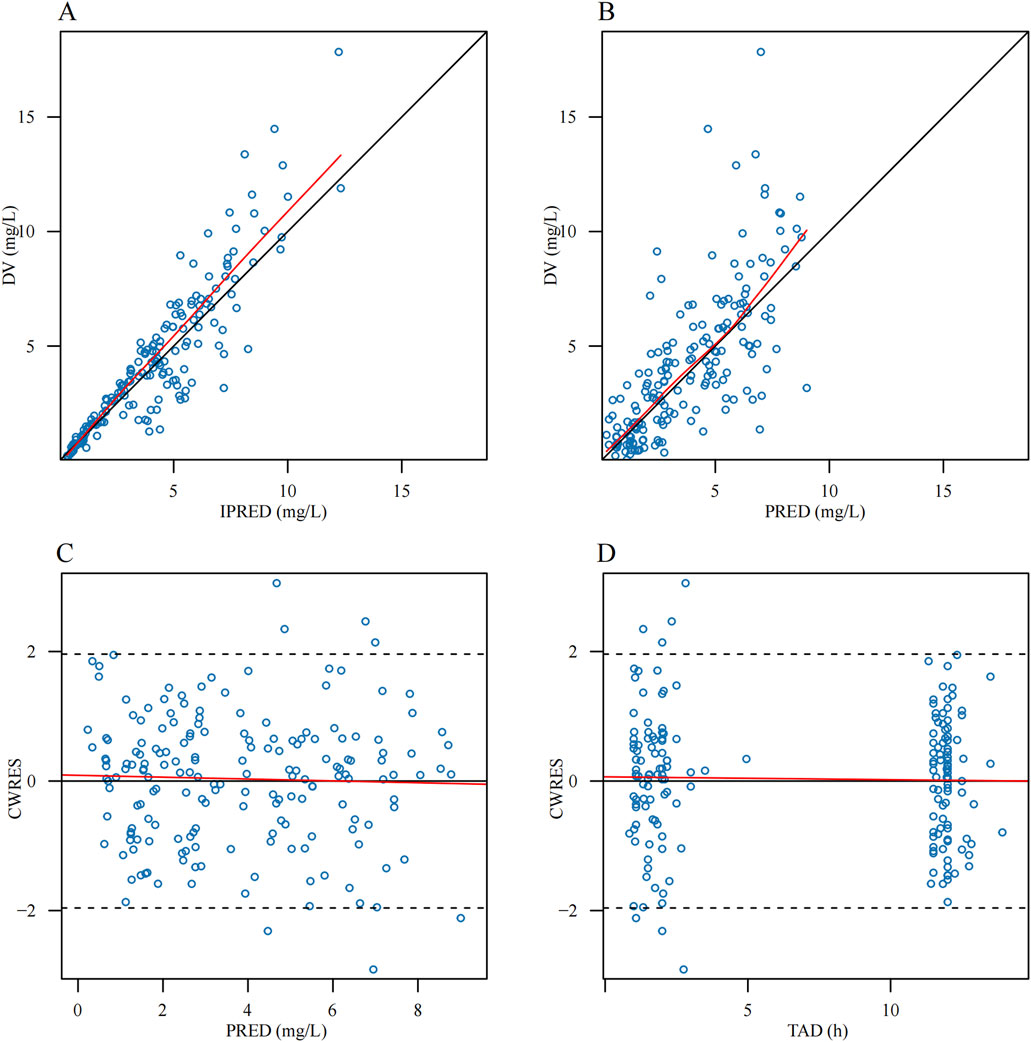
Figure 1. Goodness-of-fit plots of the final population pharmacokinetic model for polymyxin B. (A) Observed concentration (DV) versus individual prediction (IPRED) (B) DV versus population prediction (PRED) (C) Conditional weighted residuals (CWRES) versus PRED (D) CWRES versus time after dose (TAD). The red lines represent the locally weighted scatter plot smoothing (LOESS) curves.
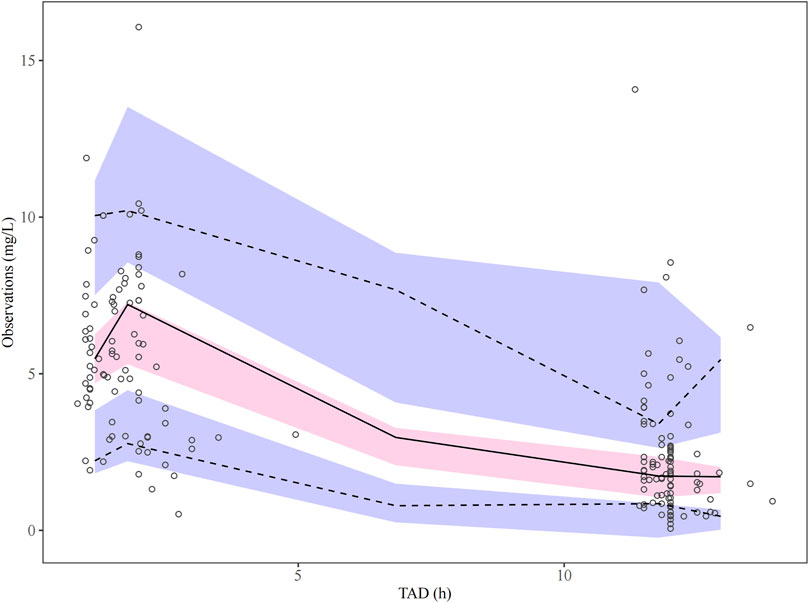
Figure 2. Visual predictive check plots for the PopPK final model. The solid black line indicates the median of the observed data, while the dashed lines show the 5th and 95th percentiles. The shaded regions represent the 95% confidence intervals for the 5th, 50th, and 95th percentiles derived from the simulations.
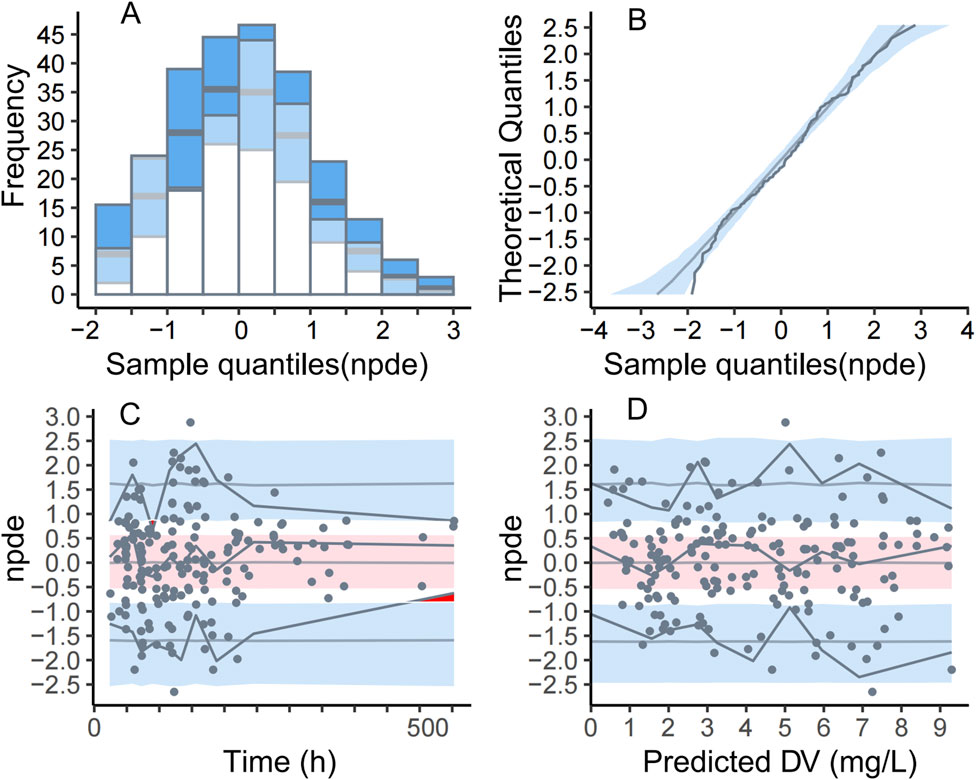
Figure 3. Normalized Prediction Distribution Errors (NPDE) plot for the final PopPK model. (A) Histogram of NPDE overlaid with the density plot of a standard normal distribution (B). Q-Q plot of NPDE against the standard normal distribution (C). Scatter plot of NPDE versus time after the first dose (D). Scatter plot of NPDE versus population predicted (PRED) The dots represent the NPDE calculated from the dataset. In panel B, the blue shaded area represents the 95% prediction interval for a standard normal random variable. In panels C and D, the red shaded area corresponds to the prediction interval for the median (50th percentile) of the NPDE, while the blue shaded area represents the prediction interval for the 5th and 95th percentiles. The solid lines depict the evolution of observed data across percentiles compared to model predictions.
A total of 106 AUCss,24h values were obtained from the modeling group (91 values) and the validation group (15 values). The AUCss,24h estimated by the PPK model within the range of 50–100 was 51.89% (55/106), while the AUCss,24h estimated using the two-point method was 46.23% (49/106). The Bland-Altman analysis results were shown in Figure 4. The mean difference of AUCss,24h between the two methods was 12.98 mg·h·L-1, indicated a small average difference between the methods. The majority of the sample differences fell within the 95% confidence interval, demonstrated consistency between the two estimation methods in most samples. The Kappa test revealed moderate agreement between the two methods (κ = 0.51, P < 0.001), while McNemar’s test showed no significant difference in classification outcomes (P = 0.33), indicating overall consistency with minor discrepancies.
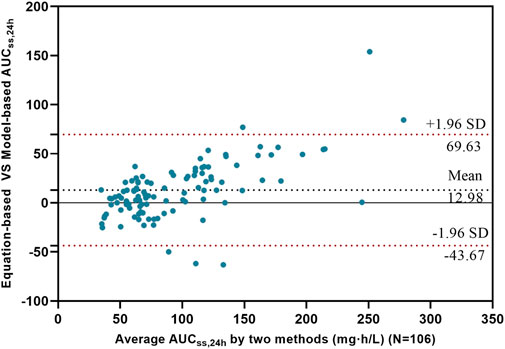
Figure 4. Bland-Altman analysis of AUCss,24h calculated by the two methods The black solid line represents the mean difference of AUCss,24h calculated by the two methods, while the red dashed lines represent the 95% confidence interval of the differences.
To evaluate the impact of varying renal function levels, CrCL values of 30, 60, 90, 120, and 150 mL. min-1 were used in the simulation. Since PLT levels in the modeling data are concentrated in the low and normal PLT ranges, the simulation used PLT at the quartiles of the modeling data: 85·109·L-1 and 266·109·L-1. Simulation results were presented in Figure 5 and Table 3. Forty different clinical scenarios were simulated. At MIC ≤ 0.5 mg·L-1, PTA ≥ 90% was achieved in all groups except the 50 mg q12h maintenance dose group. PTA decreased as CrCL increased for the same maintenance dose and PLT level. Compared with maintenance dose and CrCL levels, PLT levels seem to exert a relatively minor influence on the PTA and proportion of AUCss,24h within the therapeutic window. In the 50 mg q12h group, the proportion of AUCss,24h within the therapeutic window decreased as CrCL increased, whereas an opposite trend was noted in the 100 mg and 125 mg groups. Overall, the 75 mg and 100 mg q12h groups had a slightly higher (75 mg 58.13%; 100 mg 46.78%; 50 mg 42.40%; 125 mg 14.66%) proportion of AUCss,24h within the therapeutic window compared to other groups.
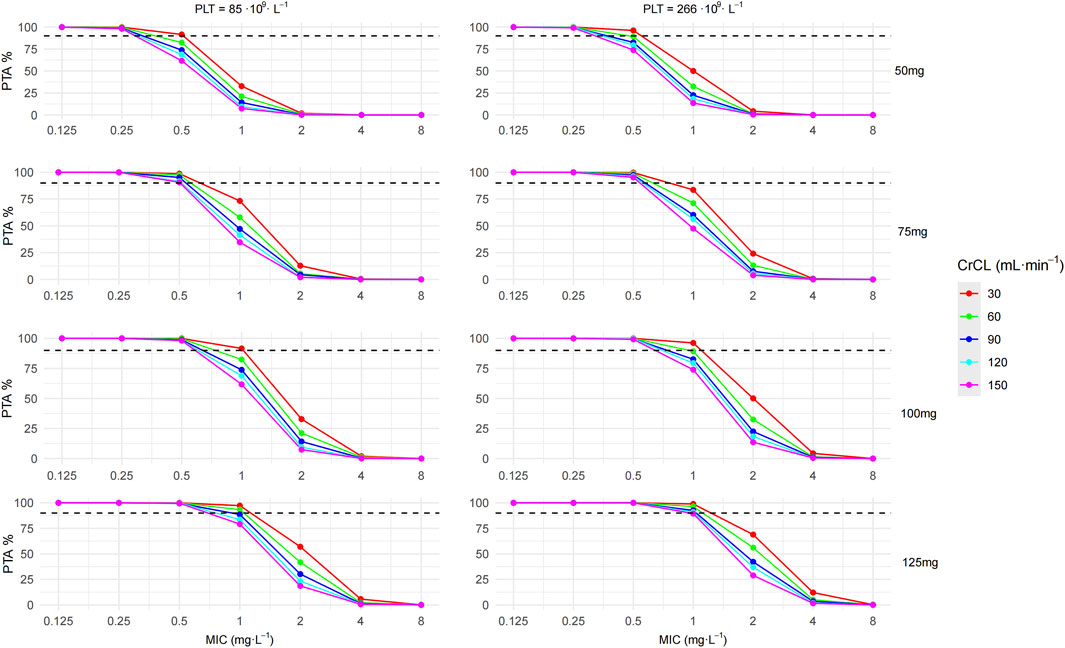
Figure 5. Probability of target attainment (PTA) for polymyxin B at a PK/PD target of AUCss,24h/MIC ≥ 66.9.
In this study, we develop a PopPK model for critically ill patients receiving PMB therapy using a two-point method based on Ctrough,ss and Cpeak,ss. Furthermore, it is the first to compare AUCss,24h estimates from the first-order elimination equation method based on Ctrough,ss and Cpeak,ss with those estimated using a PopPK model.
Our findings revealed that the one-compartment model with first-order elimination adequately described the population data, with CrCL and PLT identified as covariates influencing PMB PK. This model was found to be suitable for application in critically ill patients. Model-based simulations suggested that PLT levels had a minimal impact on the PTA. At MIC ≤ 0.5 mg·L-1, commonly used maintenance doses in critically ill patients (50, 75, 100, 125 mg q12h) generally achieved 90% PTA, except in the 50 mg maintenance dose group with high CrCL. The 75 mg and 100 mg q12h regimens showed a higher proportion of AUCss,24h values within the therapeutic window compared to other regimens, suggesting these dosing strategies may be optimal for this population. Additionally, CrCL levels should be considered for further optimization of individualized dosing. The comparison of AUCss,24h estimation methods showed good consistency, supporting the use of the Ctrough,ss and Cpeak,ss two-point sampling strategy for routine TDM.
Compared to other one-compartment PopPK studies of PMB (Table 4) (Kubin et al., 2018; Manchandani et al., 2018; Yu et al., 2020; Crass et al., 2021; Li et al., 2021), our results were similar to those of Manchandani et al. (CL = 2.5 L·h-1) and Kubin et al. (CL = 2.37 L·h-1) in CRO-infected patients, highlighting the significant influence of renal function on CL. In contrast, the lower CL observed in previous studies by Li et al. (1.18 L·h-1) and Yu et al. (1.59 L·h-1) may reflect differences in renal function and patient characteristics. Our study reported a value of 18 L, which is lower than the values reported in earlier studies by Manchandani et al. (34.3 L) and Kubin et al. (34.40 L), but is more closely aligned with the values from Li et al. (12.90 L) and Crass et al. (12.70 L). Variations in Vd may reflect differences in fluid status and organ function, as critically ill patients often experience tissue edema and hemodynamic instability, which affect drug distribution and explaining discrepancies across studies.
The association between PMB pharmacokinetics and CrCL is still debated. A study (Sandri et al., 2013b) reported that the urinary recovery of PMB in 17 patients ranged from 0.98% to 17.40%, while another study (Yu et al., 2020) reported a recovery rate of 23.56% in four patients. This suggested significant interindividual variability in PMB renal clearance. Our findings indicated that CrCL is a significant covariate influencing PMB pharmacokinetics, consistent with previous PopPK studies (Avedissian et al., 2018; Wang et al., 2020; Yu et al., 2020; Li et al., 2021; Wang et al., 2021; Luo et al., 2022; Ye et al., 2022). In this study, the similar proportions of patients with and without renal impairment represented a wide range of renal function levels. Simulations indicated higher PTA in patients with renal impairment. This may result from impaired glomerular filtration, leading to decreased PMB clearance and increased drug exposure. With increased maintenance doses, patients with renal impairment exhibited higher drug exposure, leading to more AUCss,24h values exceeding 100 mg·h·L-1.
In critically ill patients, conditions such as inflammation and organ dysfunction, especially in sepsis, can alter PLT levels, potentially affecting drug metabolism and clearance (van Dalen and Vree, 1990; van der Poll et al., 2013). Hypoalbuminemia, frequently observed in critically ill patients, may correlate with changes in PLT, influencing drug plasma protein binding and concentrations (Roberts et al., 2014b).Renal function changes, such as acute kidney injury, may also be associated with variations in PLT, thereby influenced drug clearance (Ulldemolins et al., 2011; Chawla et al., 2014). Our study identified PLT as a significant factor influencing the pharmacokinetics of PMB, and to our knowledge, this is the first report of such a finding. This finding opens new avenues for pharmacokinetic modeling and underscores the need for further investigation into the relationship between PLT and PMB pharmacokinetics.
Our study shows that although there is a small bias between the two AUCss,24h estimation methods, this difference is clinically acceptable and provides a simple yet reliable tool for clinical TDM without the need for complex modeling. However, the development of individualized dosing regimens still relies on the PopPK model, which integrates patient characteristics to enable precise predictions and adjustments, thereby optimizing the efficacy and safety of PMB therapy.
Despite the significant findings, our study has some limitations. First, we did not correlate PMB PK/PD parameters with patient outcomes, so the relationship between PMB exposure and clinical outcomes remains unclear. Second, most patients in our study had PLT counts within or below normal ranges, and to minimize simulation errors, we excluded populations with elevated PLT levels from modeling. Consequently, our findings may not apply to populations with elevated PLT counts. This novel observation should be interpreted with caution, especially as prior studies have not reported PLT’s influence on PMB pharmacokinetics. Lastly, we measured total plasma concentrations of PMB without evaluating free drug levels. If significant variability in albumin levels exists among patients, the applicability of our findings could be limited.
This study established a PopPK model for PMB in critically ill patients, identifying CrCL and PLT as covariates influencing PMB clearance. The 75 mg q12h and 100 mg q12h dosing regimens seem appropriate for critically ill patients; however, CrCL levels should be considered when selecting between these regimens to guide individualized dosing. The two-point sampling strategy based on Ctrough,ss and Cpeak,ss can be applied for routine TDM of PMB.
The raw data supporting the conclusions of this article will be made available by the authors, without undue reservation.
The studies involving humans were approved by Ethics Committee of First Affiliated Hospital of Army Medical University. The studies were conducted in accordance with the local legislation and institutional requirements. The participants provided their written informed consent to participate in this study.
JY: Formal Analysis, Software, Validation, Writing–original draft, Writing–review and editing. MY: Formal Analysis, Methodology, Supervision, Writing–review and editing. YG: Data curation, Investigation, Validation, Writing–review and editing. LC: Formal Analysis, Methodology, Writing–review and editing. GY: Data curation, Writing–review and editing. LX: Data curation, Writing–review and editing. FL: Formal Analysis, Project administration, Supervision, Writing–review and editing. YC: Conceptualization, Methodology, Project administration, Resources, Supervision, Writing–review and editing.
The author(s) declare that no financial support was received for the research, authorship, and/or publication of this article.
The authors acknowledge the valuable assistance of Riuxiang Liu and Yang Wang for their advice on model development.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fphar.2025.1511088/full#supplementary-material
ALB, Albumin; IL-6, Interleukin-6; ALT, Alanine Aminotransferase; INR, International normalized ratio; APACHE, Ⅱ Acute Physiology and Chronic Health Evaluation Ⅱ; IPRED, Individual prediction; APTT, Activated partial thromboplastin time; MIC, Minimum Inhibitory Concentration; AST, Aspartate Aminotransferase; NPDE, Normalized prediction distribution error; AUCss,24h, Concentration-time curve at steady state over 24 h; OFV, Objective Function Value; BUN, Blood urea nitrogen; PK/PD, pharmacokinetics/pharmacodynamics; CI, Confidence Interval; PLT, Platelet count; CL, Systemic Clearance; PMB, Polymyxin B; Cpeak,ss, Steady-state peak concentration; PopPK, Population Pharmacokinetics; CRAB, carbapenem-resistant Acinetobacter baumannii; PRDE, Population prediction; CrCL, Creatinine Clearance; PTA, Probability of Target Attainment; CRE, carbapenem-resistant Enterobacterales; TBIL, Total bilirubin; CRO, Carbapenem-resistant organisms; TBW, Total body weight; CRKP, Carbapenem-resistant Klebsiella pneumoniae; TDM, Therapeutic Drug Monitoring; Ctrough,ss, Steady-state trough concentration; TP, Total protein; CWRES, Conditional weighted residuals; UPLC-MS/MS, Ultra-performance liquid chromatography-tandem mass spectrometry; DV, Observed concentration; Vd, Volume of Distribution of the Central Compartment; eGFR, Estimated glomerular filtration rate; VPC, Visual predictive check; Fib, Fibrinogen; WBC, White Blood Cell; HGB, Hemoglobin.
Abdallah, M., Olafisoye, O., Cortes, C., Urban, C., Landman, D., and Quale, J. (2015). Activity of Eravacycline against Enterobacteriaceae and acinetobacter baumannii, Including multidrug-resistant isolates, from New York City. Antimicrob. Agents Chemother. 59, 1802–1805. doi:10.1128/AAC.04809-14
Avedissian, S. N., Miglis, C., Kubin, C. J., Rhodes, N. J., Yin, M. T., Cremers, S., et al. (2018). Polymyxin B pharmacokinetics in adult Cystic fibrosis patients. Pharmacother. J. Hum. Pharmacol. Drug Ther. 38, 730–738. doi:10.1002/phar.2129
Bergen, P. J., Bulitta, J. B., Forrest, A., Tsuji, B. T., Li, J., and Nation, R. L. (2010). Pharmacokinetic/pharmacodynamic investigation of colistin against Pseudomonas aeruginosa using an in vitro model. Antimicrob. Agents Chemother. 54, 3783–3789. doi:10.1128/AAC.00903-09
Brink, A. J. (2019). Epidemiology of carbapenem-resistant Gram-negative infections globally. Curr. Opin. Infect. Dis. 32, 609–616. doi:10.1097/QCO.0000000000000608
Chawla, L. S., Eggers, P. W., Star, R. A., and Kimmel, P. L. (2014). Acute kidney injury and chronic kidney disease as interconnected syndromes. N. Engl. J. Med. 371, 58–66. doi:10.1056/NEJMra1214243
Crass, R. L., Al Naimi, T., Wen, B., Souza, E., Murray, S., Pai, M. P., et al. (2021). Pharmacokinetics of polymyxin B in hospitalized adults with cystic fibrosis. Antimicrob. Agents Chemother. 65, e0079221. doi:10.1128/AAC.00792-21
Huemer, M., Mairpady Shambat, S., Brugger, S. D., and Zinkernagel, A. S. (2020). Antibiotic resistance and persistence-Implications for human health and treatment perspectives. EMBO Rep. 21, e51034. doi:10.15252/embr.202051034
Kubin, C. J., Nelson, B. C., Miglis, C., Scheetz, M. H., Rhodes, N. J., Avedissian, S. N., et al. (2018). Population pharmacokinetics of intravenous polymyxin B from clinical samples. Antimicrob. Agents Chemother. 62. doi:10.1128/AAC.01493-17
Li, Y., Deng, Y., Zhu, Z.-Y., Liu, Y.-P., Xu, P., Li, X., et al. (2021). Population pharmacokinetics of polymyxin B and dosage optimization in renal transplant patients. Front. Pharmacol. 12, 727170. doi:10.3389/fphar.2021.727170
Liang, D., Liang, Z., Deng, G., Cen, A., Luo, D., Zhang, C., et al. (2023). Population pharmacokinetic analysis and dosing optimization of polymyxin B in critically ill patients. Front. Pharmacol. 14, 1122310. doi:10.3389/fphar.2023.1122310
Liu, X., Huang, C., Bergen, P. J., Li, J., Zhang, J., Chen, Y., et al. (2023). Chinese consensus guidelines for therapeutic drug monitoring of polymyxin B, endorsed by the infection and Chemotherapy Committee of the Shanghai Medical Association and the Therapeutic Drug Monitoring Committee of the Chinese Pharmacological Society. J. Zhejiang University-SCIENCE B 24, 130–142. doi:10.1631/jzus.b2200466
Luo, X., Zhang, Y., Liang, P., Zhu, H., Li, M., Ding, X., et al. (2022). Population pharmacokinetics of polymyxin B and dosage strategy in critically ill patients with/without continuous renal replacement therapy. Eur. J. Pharm. Sci. 175, 106214. doi:10.1016/j.ejps.2022.106214
Manchandani, P., Thamlikitkul, V., Dubrovskaya, Y., Babic, J. T., Lye, D. C., Lee, L. S., et al. (2018). Population pharmacokinetics of polymyxin B. Clin. Pharmacol. & Ther. 104, 534–538. doi:10.1002/cpt.981
Mao, J. J., Jiao, Z., Yun, H. Y., Zhao, C. Y., Chen, H. C., Qiu, X. Y., et al. (2018). External evaluation of population pharmacokinetic models for ciclosporin in adult renal transplant recipients. Br. J. Clin. Pharmacol. 84, 153–171. doi:10.1111/bcp.13431
Onufrak, N. J., Rao, G. G., Forrest, A., Pogue, J. M., Scheetz, M. H., Nation, R. L., et al. (2017). Critical need for clarity in polymyxin B dosing. Antimicrob. Agents Chemother. 61. doi:10.1128/AAC.00208-17
Poirel, L., Jayol, A., and Nordmann, P. (2017). Polymyxins: antibacterial activity, susceptibility testing, and resistance mechanisms encoded by plasmids or chromosomes. Clin. Microbiol. Rev. 30, 557–596. doi:10.1128/CMR.00064-16
Rabanal, F., and Cajal, Y. (2017). Recent advances and perspectives in the design and development of polymyxins. Nat. Product. Rep. 34, 886–908. doi:10.1039/c7np00023e
Roberts, J. A., Abdul-Aziz, M. H., Lipman, J., Mouton, J. W., Vinks, A. A., Felton, T. W., et al. (2014a). Individualised antibiotic dosing for patients who are critically ill: challenges and potential solutions. Lancet Infect. Dis. 14, 498–509. doi:10.1016/S1473-3099(14)70036-2
Roberts, J. A., Paul, S. K., Akova, M., Bassetti, M., De Waele, J. J., Dimopoulos, G., et al. (2014b). DALI: Defining antibiotic levels in intensive care unit patients: are current -lactam antibiotic doses sufficient for critically ill patients? Clin. Infect. Dis. 58, 1072–1083. doi:10.1093/cid/ciu027
Sandri, A. M., Landersdorfer, C. B., Jacob, J., Boniatti, M. M., Dalarosa, M. G., Falci, D. R., et al. (2013a). Population pharmacokinetics of intravenous polymyxin B in critically ill patients: Implications for selection of dosage regimens. Clin. Infect. Dis. 57, 524–531. doi:10.1093/cid/cit334
Sandri, A. M., Landersdorfer, C. B., Jacob, J., Boniatti, M. M., Dalarosa, M. G., Falci, D. R., et al. (2013b). Pharmacokinetics of polymyxin B in patients on continuous venovenous haemodialysis. J. Antimicrob. Chemother. 68, 674–677. doi:10.1093/jac/dks437
Tacconelli, E., Carrara, E., Savoldi, A., Harbarth, S., Mendelson, M., Monnet, D. L., et al. (2018). Discovery, research, and development of new antibiotics: the WHO priority list of antibiotic-resistant bacteria and tuberculosis. Lancet Infect. Dis. 18, 318–327. doi:10.1016/S1473-3099(17)30753-3
Tang, T., Li, Y., Xu, P., Zhong, Y., Yang, M., Ma, W., et al. (2023). Optimization of polymyxin B regimens for the treatment of carbapenem-resistant organism nosocomial pneumonia: a real-world prospective study. Crit. Care 27, 164. doi:10.1186/s13054-023-04448-z
Tsuji, B. T., Pogue, J. M., Zavascki, A. P., Paul, M., Daikos, G. L., Forrest, A., et al. (2019). International consensus guidelines for the optimal use of the polymyxins: endorsed by the American College of Clinical Pharmacy (ACCP), European Society of Clinical Microbiology and Infectious Diseases (ESCMID), Infectious Diseases Society of America (IDSA), International Society for Anti-infective Pharmacology (ISAP), Society of Critical Care Medicine (SCCM), and Society of Infectious Diseases Pharmacists (SIDP). Pharmacother. J. Hum. Pharmacol. Drug Ther. 39, 10–39. doi:10.1002/phar.2209
Ulldemolins, M., Roberts, J. A., Rello, J., Paterson, D. L., and Lipman, J. (2011). The effects of hypoalbuminaemia on optimizing antibacterial dosing in critically ill patients. Clin. Pharmacokinet. 50, 99–110. doi:10.2165/11539220-000000000-00000
Van Dalen, R., and Vree, T. B. (1990). Pharmacokinetics of antibiotics in critically ill patients. Intensive Care Med. 16 (Suppl. 3), S235–S238. doi:10.1007/BF01709707
Van Der Poll, T., Schultz, M., and Levi, M. (2013). Sepsis and thrombosis. Seminars Thrombosis Hemostasis 39, 559–566. doi:10.1055/s-0033-1343894
Wang, P., Zhang, Q., Zhu, Z., Feng, M., Sun, T., Yang, J., et al. (2020). Population pharmacokinetics and limited sampling strategy for therapeutic drug monitoring of polymyxin B in Chinese patients with multidrug-resistant gram-negative bacterial infections. Front. Pharmacol. 11, 829. doi:10.3389/fphar.2020.00829
Wang, P., Zhang, Q., Zhu, Z., Pei, H., Feng, M., Sun, T., et al. (2021). Comparing the population pharmacokinetics of and acute kidney injury due to polymyxin B in Chinese patients with or without renal insufficiency. Antimicrob. Agents Chemother. 65. doi:10.1128/AAC.01900-20
Yang, J., Liu, S., Lu, J., Sun, T., Wang, P., and Zhang, X. (2022). An area under the concentration-time curve threshold as a predictor of efficacy and nephrotoxicity for individualizing polymyxin B dosing in patients with carbapenem-resistant gram-negative bacteria. Crit. Care 26, 320. doi:10.1186/s13054-022-04195-7
Ye, Q., Wang, Q., Chen, W., Zhang, R., Chen, Z., Li, P., et al. (2022). The population pharmacokinetics and dose optimization of polymyxin B in critically ill patients with or without extracorporeal membrane oxygenation. J. Clin. Pharm. Ther. 47, 1608–1618. doi:10.1111/jcpt.13711
Yu, X. B., Jiao, Z., Zhang, C. H., Dai, Y., Zhou, Z. Y., Han, L., et al. (2020). Population pharmacokinetic and optimization of polymyxin B dosing in adult patients with various renal functions. Br. J. Clin. Pharmacol. 87, 1869–1877. doi:10.1111/bcp.14576
Keywords: population pharmacokinetics, polymyxin B, critical illness, dosing optimization, consistency
Citation: Yang J, Yu M, Gan Y, Cheng L, Yang G, Xiong L, Liu F and Chen Y (2025) Population pharmacokinetics of polymyxin B in critically ill patients with carbapenem-resistant organisms infections: insights from steady-state trough and peak plasma concentration. Front. Pharmacol. 16:1511088. doi: 10.3389/fphar.2025.1511088
Received: 14 October 2024; Accepted: 20 February 2025;
Published: 12 March 2025.
Edited by:
Sonia Alejandra Gomez, National Scientific and Technical Research Council (CONICET), ArgentinaReviewed by:
Maximiliano Gabriel Castro, Jose Bernando Iturraspe Hospital, ArgentinaCopyright © 2025 Yang, Yu, Gan, Cheng, Yang, Xiong, Liu and Chen. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Fang Liu, bGl1ZmFuZzAyMDlAdG1tdS5lZHUuY24=; Yongchuan Chen, endtY3ljQHRtbXUuZWR1LmNu
†These authors have contributed equally to this work and share first authorship
‡These authors have contributed equally to this work
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.