
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Pharmacol. , 04 April 2025
Sec. Gastrointestinal and Hepatic Pharmacology
Volume 16 - 2025 | https://doi.org/10.3389/fphar.2025.1492694
 Qiang Li1
Qiang Li1 Pan Pan2
Pan Pan2 Qingqing Xian1
Qingqing Xian1 Jingtan Li1
Jingtan Li1 Jingting Wang1
Jingting Wang1 Jiaying Cai1
Jiaying Cai1 Jing Wang1
Jing Wang1 Yanfei Jia1
Yanfei Jia1 Haiji Sun3
Haiji Sun3 Lulu Zhang1*
Lulu Zhang1* Xiaoli Ma1*
Xiaoli Ma1*Background: The transcription factor GLI1, aberrantly activated in gastric cancer, drives tumor progression, yet no approved inhibitors currently target this molecule. G-quadruplex (G4) motifs in promoter regions have emerged as promising therapeutic targets. This study explores G4 stabilization in the GLI1 promoter as a novel strategy to suppress gastric cancer progression.
Methods: G4 formation in the GLI1 promoter was validated using circular dichroism. A dual-luciferase assay screened FDA-approved drugs for G4-stabilizing activity, identifying alpha-estradiol and (R)-(-)-ibuprofen as candidates. These compounds were evaluated for anti-tumor effects through in vitro assays (proliferation, migration, invasion) and in vivo xenograft models. Mechanistic insights into GLI1/PRKACB signaling were obtained via chromatin immunoprecipitation and pathway analysis.
Results: Stable G4 structures were confirmed in the GLI1 promoter. Alpha-estradiol and (R)-(-)-ibuprofen suppressed GLI1 transcription and protein levels, significantly inhibiting gastric cancer cell proliferation, migration, invasion, and stemness. In vivo, both compounds reduced tumor growth and metastasis, with (R)-(-)-ibuprofen synergizing with cisplatin to enhance efficacy. Mechanistically, GLI1 directly regulated PRKACB expression, and G4 stabilization downregulated PRKACB, impairing epithelial-mesenchymal transition and cancer stemness.
Conclusion: Targeting GLI1 G4 structures with alpha-estradiol and (R)-(-)-ibuprofen effectively inhibits gastric cancer progression by blocking GLI1/PRKACB signaling. This study highlights G4-targeted therapy as a novel and clinically translatable strategy for gastric cancer treatment.
Gastric cancer is the fifth most commonly diagnosed cancer and the fourth leading cause of cancer-related death in the world (Sung et al., 2021). Due to dramatic advancements in Helicobacter pylori eradication, nutritional improvement, earlier diagnosis, and therapeutic breakthroughs, the incidence of gastric cancer has declined over the past few decades (Park and Herrero, 2021). However, there is an unmet clinical demand for targeted therapies for advanced gastric cancer (Smyth et al., 2020). Therefore, the identification of novel therapeutic targets and elucidation of the underlying mechanisms are urgently needed in gastric cancer.
Glioma-associated oncogene homolog 1 (GLI1) is an important transcription factor downstream of the Hedgehog (Hh) signaling pathway and can be used as a marker of Hh signaling pathway activation (Wang et al., 2019; Jagani et al., 2010). The Hh/GLI signaling pathway is aberrantly activated in various tumors, including gastric cancer (Qi et al., 2020), lung adenocarcinoma (Li et al., 2016), oral squamous carcinoma (Shimo et al., 2016), breast carcinoma (Hegde et al., 2008), and hepatocellular carcinoma (Yan et al., 2016), and is closely associated with tumorigenesis, progression, invasion, and metastasis. In addition, studies (Aberger et al., 2012; Kern et al., 2015) have found that the Hh/GLI signaling pathway can promote tumor progression through other signaling pathways through a cross-regulatory phenomenon. Various small-molecule inhibitors of the Hh pathway are currently being developed; however, there are two major problems. Most inhibitors have been developed to target PTCH1 and SMO (Xu et al., 2019). Previous antitumor studies have focused on SMO targets upstream of the Hh pathway; however, in recent years, GLI has been found to play a more important role than SMO in cancer development (Stecca and Ruiz i Altaba, 2010). Second, only two drugs (GDC-0449 and LDE225) have been approved by the FDA for the treatment of basal cell carcinoma and medulloblastoma, and no molecular inhibitors have been approved for the treatment of gastric cancer (Rubin and de Sauvage, 2006; Bariwal et al., 2019). Therefore, blocking or silencing the expression of GLI1 will significantly help suppress gastric cancer progression.
G-quadruplexes are a class of guanine-rich DNA or RNA secondary structures stabilized by Hoogsteen hydrogen bonds (Nishio et al., 2021). Many potential G-quadruplex formation motifs are present in the human genome. G-quadruplex structures regulate genomic stability, DNA replication, and gene expression, many of which are involved in oncogenesis (Awadasseid et al., 2021). Therefore, searching for ligands that can induce G-quadruplex formation in guanine-rich regions of DNA and screening for potential compounds as anticancer drugs represent research hotspots in the field of cancer therapy (Tassinari et al., 2021). Through in vitro experiments, we verified sequences that could form a G-quadruplex structure and thus could be used as molecular targets for inhibitor development.
cAMP-dependent protein kinase catalytic subunit β (PRKACB) mediates the cAMP-dependent pathway associated with various cellular processes such as cell proliferation, cell cycle, chromatin condensation, nuclear membrane reorganization, and intracellular transport mechanisms (Wang et al., 2020). Recently, it was found that PRKACB is likely associated with the epithelial mesenchymal transition (Falch et al., 2018). PRKACB has two isoforms, Cβ1 and Cβ2, of which the Cβ1 isoform is abnormally expressed in gastric cancer tissues (Furuta et al., 2012). PRKACB regulates p75, PKA, and other key molecules involved in cell survival, differentiation, metabolism and apoptosis (Kamilaris et al., 2020). Excessive activity can cause diseases such as adrenal tumors (Espiard et al., 2018). Therefore, PRKACB has an important physiological function; however, in gastric cancer, the function of PRKACB is not clear, and its upstream and downstream regulatory molecular pathways are rarely reported.
Here, we studied the effects of G-quadruplex regulators of the GLI1 promoter on gastric cancer progression. Furthermore, this effect was confirmed by inhibition of the GLI/PRKACB pathway in gastric cancer.
The expression of GLI1 and its correlation with clinical stage and lymph node metastasis of gastric cancer were analyzed using the online databases GEPIA (http://gepia.cancer-pku.cn) and UALCAN (http://ualcan.path.uab.edu/index.html), respectively. The correlation between GLI1 and PRKACB and their clinical significance with overall survival (OS) was analyzed using TIMER (https://www.timeanddate.com/timer) and The Human Protein Atlas (https://www.proteinatlas.org/).
The tissue microarray (HStm-Muc060CS-01) consists of 30 adenocarcinoma and 30 tumor-adjacent tissue samples. Each group of paired carcinoma and tumor-adjacent tissues was collected and classified according to the clinical information. Twenty of the 30 specimens were from males and 10 were from females, ranging from to 45–82 years old (mean age, 61 years). The tissues of the patient specimens were obtained from the Department of Pathology, Central Hospital, affiliated with Shandong First Medical University.
The human gastric epithelial cell line GES-1, gastric adenocarcinoma cell line HGC-27, and AGS were purchased from IBCB (Shanghai Institute of Biochemistry and Cell Biology, Chinese Academy of Science). Cells were grown in RPMI 1640 medium (HyClone) and Ham’s F-12 nutrient medium supplemented with 10% fetal bovine serum (FBS) (Gibco) and 1% penicillin and streptomycin (Macgene). Cells were cultured at 37°C in a humidified atmosphere containing 5% CO2. Human gastric cancer cell lines were treated with 20 µM GANT61 (an inhibitor of GLI1), 1 µM alpha-estradiol or (R)-(−)-Ibuprofen for 24 h.
AGS and HGC-27 cells were seeded in 6-well plates at a density of 5 × 105 cells per well and cultured overnight to reach 70%–80% confluence. Then the cells were transfected with the PRKACB plasmid in opti-MED medium using Lipofectamine 2000 Transfection Reagent at room temperature, according to the manufacturer’s protocol. 24 h after transfection, PRKACB expression in AGS and HGC-27 cells was assessed by Western blot analysis.
Immunohistochemical staining was performed using a streptavidin-peroxidase assay. Tissue sections were cut from the formalin-fixed paraffin-embedded tissue blocks at a thickness of 4 µm and mounted on glass slides.The slides were then dewaxed in xylene for 10 min twice and rehydrated through a series of graded ethanol solutions (100%, 95%, 80%, 70%) for 5 min each. Antigen retrieval was carried out by boiling the slides in a citrate buffer (pH 6.0) in a microwave oven for 10 min. Then, the slides were incubated with primary antibodies: anti-GLI1 MAb (1:200; Proteintech, cat no. 66905-1-Ig), anti-vimentin RAb (1:2500; Proteintech, cat no. 10366-1-AP), anti-Zeb1 RAb (1:250; Proteintech, cat no. 21544-1-AP), anti-PRKACB RAb (1:200; Proteintech, cat no. 12232-1-AP) and anti-Ki67 MAb (1:50; Santa Cruz, sc-23900) overnight at 4°C. The slides were then incubated with a biotinylated anti-mouse secondary antibody (OriGene cat no. PV-9002) and biotinylated anti-rabbit antibody (OriGene, cat no. PV-9001) for 20 min at 37°C. The immunohistochemical analyses were performed simultaneously by two independent researchers.
Cell monolayers were lysed in RIPA buffer to obtain total cell protein extracts. Total protein was quantified using a BCA proteometric assay. Proteins were separated by 10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and transferred to a PVDF membrane. The membrane was incubated with the primary antibody at 4°C overnight. Then, the secondary antibody was added at room temperature for 45 min and the proteins were visualized using the FluorChenE Gel Imaging System (Protein Simple). For Western blot analysis, the primary antibodies anti-GLI1 MAb (1:2000; Proteintech, cat no. 66905-1-Ig), anti-PRKACB RAb (1:1000; Proteintech, cat no. 12232-1-AP), anti-Zeb1 (1:1000, Proteintech, cat no. 21544-1-AP), anti-vimentin (1:5000; Proteintech, cat no. 10366-1-AP), anti-GAPDH (1:5000; Proteintech, cat no. 10494-1-AP), anti-CD44 RAb (1:3000; Proteintech, cat no. 15675-1-AP) and anti-Sox2 RAb (1:2000; Cell Signaling, 23064S) were used. GAPDH was used as an internal standard.
Chromatin immunoprecipitation (ChIP) assays were performed using a ChIP Assay Kit (CST, USA). The manufacturer’s instructions were followed. Briefly, cells were cultured to approximately 1 × 107 cells, cross-linked the proteins and DNA for 10 min at room temperature with 1% formaldehyde. The cells were harvested, washed twice with cold PBS, and lysed in SDS lysis buffer (50 mM Tris-HCl pH 8.0, 10 mM EDTA, 1% SDS) containing protease inhibitors. The lysates were sonicated using a sonicator to shear the DNA into fragments of 200–500 bp. The sonicated lysates were centrifuged at 14,000 rpm for 10 min at 4°C, and the supernatants were collected. The supernatants were pre-cleared by incubating with protein A/G agarose beads (CST) for 1 h at 4°C. After pre-clearing, the supernatants were divided into two aliquots and incubated with 2 µg of GLI (PTG) or IgG antibody per reaction overnight at 4°C with gentle rotation. Protein A/G agarose beads were added to the reactions and incubated for 2 h at 4°C to capture the antibody-DNA complexes. DNA was eluted from the beads by incubating with elution buffer (1% SDS, 0.1 M NaHCO3) for 15 min at room temperature twice. Real-time fluorescence quantitative PCR was performed to amplify the precipitated DNA using primers specific for the PRKACB promoter. The primer sequences were as follows: forward 5′-TTCAGATCTACACAGAAGTTGC-3′ and reverse 5′-CTCGCACATTGAGGTATTCG-3’. Non-immunoprecipitated chromatin fragments were used as the input controls, and the relative enrichment of the target DNA was calculated using the comparative Ct method.
Cell migration was evaluated using a wound-healing assay. Briefly, 2 × 105 AGS or HGC cells per well were seeded in six-well plates. After the cells reached approximately 90% confluence, the cell monolayers were wounded vertically with a 20 µL or 200 µL sterile pipette tip. The wells were washed with phosphate buffered saline (PBS) and fresh medium was added. The plates were placed in a 37°C incubator with 5% CO2, and the migrated cells were observed and photographed at 0 h and 24 h after scratching using an Olympus CKX41 microscope. The width of the wound was measured using image analysis software (ImageJ), and the migration rate was calculated as the ratio of the decrease in wound width to the initial wound width.
The cell migration assay was conducted using 24-well plates with 8-μm pore Transwell chambers (Corning, United States). A total of 1 × 105 AGS or HGC cells were placed in the upper chamber in serum-free medium. The lower chamber was filled with culture medium containing 20% FBS. After 24 h, the non-migrated cells were removed with clean swabs and the cells on the bottom surface were stained with 0.1% crystal violet dye. For the cell invasion assay, AGS or HGC cells were seeded at a density of 1 × 105 cells/well in the upper chamber filled with Matrigel (Corning, United States). After 24 h, non-invading cells were removed with swabs and the cells on the underside were stained with 0.1% crystal violet dye. The migration and invasion abilities of AGS and HGC cells were quantified by inverted contrast microscopy.
The cells were digested with trypsin and resuspended for cell counting, and 5 × 103 cells were spread in a low-adhesion 6-well plate with three replicate wells. The plate was placed in a 37°C incubator with 5% CO2 and cultured for 7 days. During the culture process, the medium was not changed. After 7 days of culture, photographs were taken under a microscope, and five fields were randomly selected to count the number of sphere-forming cells using image analysis software (ImageJ). The sphere formation efficiency was calculated as the ratio of the number of sphere-forming cells to the total number of seeded cells.
The assay was carried out according to the instructions of the Cell-Light EdU Apollo 567 In Vitro kit (Ribobio Ltd, China). In 96-well plates, 5 × 103 cells were added to each well and cultured overnight. 100 μL culture medium (containing 0.1% EdU) was added to each well for 2 h before fixation in 4% paraformaldehyde. After permeabilization with 0.5% Triton X-100, 100 μL 1×Apollo-staining reaction liquid was added to cells at 37°C for 30 min, then the cells were counterstained with 4′,6-diamidino-2-phenylindole (DAPI) and imaged using a fluorescence microscope (Olympus, Tokyo, Japan). The images were analyzed using image analysis software (ImageJ), and the percentage of proliferating cells (EdU-positive cells) was calculated as the ratio of the number of EdU-positive cells to the total number of DAPI-stained cells.
In a 24-well plate, 1000 cells were added to each well. The cells were cultured in the complete culture medium described above for 10 days. During the culture process, the medium was changed every 3 days. After 10 days of incubation, 500 μL methanol was added to each well for 15 min. After washing twice with PBS buffer, each well was stained with 0.1% crystal violet dye for 30 min.The colonies were counted under a light microscope, and the clone formation efficiency was calculated as the ratio of the number of colonies to the number of seeded cells.
The purified protein was subjected to SDS-PAGE, and the stained and decolorized gels were used to determine the purity of the target protein. The gel was stained with Coomassie Brilliant Blue R-250 and then decolorized using a destaining solution (a mixture of methanol, acetic acid, and water). The purified protein can be used for CD spectroscopy when the content of target protein is 95% or more. The spectra were recorded in the far-ultraviolet, 190–240 nm, at room temperature using a circular dichroic chromatograph. The concentration of the sample was 100 μg/mL and the optical diameter of the sample cup was 0.2 cm. The resolution was 0.5 nm, bandwidth of 0.5 nm, sensitivity of 50 mdeg, and speed of 0.8 nm/min. Secondary structures of the peptides were calculated using Yang’s algorithm (Yang. Jsr). The algorithm analyzes the CD spectra data and estimates the proportions of different secondary structures.
Human gastric cancer cells were plated in 6-well plates at a density of 2 × 105 cells per well and cultured in the complete culture medium described above for 24 h. Then, the cells in each well were transfected with 1 μM plasmid (including the wild-type or mutant GLI1 promoter reporter plasmid and the control Renilla luciferase plasmid). Briefly, the plasmid DNA was diluted in Opti - MEM medium (Gibco), and Lipofectamine 2000 was added at a ratio of 1:2 (μg of DNA to μL of Lipofectamine 2000). The mixture was incubated at room temperature for 20 min to form transfection complexes. Subsequently, the transfection complexes were added to the cells dropwise, and the cells were incubated in a 37°C incubator with 5% CO2 for 6 h. After that, the medium was replaced with fresh complete culture medium. After transfection, the cells were harvested 24 h after drug treatment and manipulated using a Dual-Glo kit (Promega, 0000551855). Assays were performed using the Dual-Luciferase Assay Luciferase Assay System (Promega) according to the manufacturer’s protocol. The firefly/renal luciferase ratio was calculated as the normalized reporter gene activity, with the control value set at 1.0. Each experiment was repeated at least three times to ensure the reliability of the data.
The animal assays were approved by the Institutional Animal Care and Use Committee of the Central Hospital affiliated with Shandong First Medical University. BALB/c nude mice (4 weeks old, male, 18–20 g) were purchased and acclimatized to the experimental conditions for 1 week. The mice were randomly divided into control and experimental groups. The experimental groups were alpha-estradiol (n = 6), (R)-(−)-ibuprofen (n = 6), GANT61 (n = 6), cisplatin (n = 6), and cisplatin in combination with (R)-(−)-ibuprofen (n = 6). A total of 2 × 106 HGC cells suspended in 100 μL of medium were injected intravenously (IV), or 1 × 107 HGC cells suspended in 50 μL of medium mixed with 50 μL Matrigel Basement Membrane Matrix High Concentration (Corning 354248) were injected subcutaneously (SC) into the BALB/c nude mice. The drug was injected intraperitoneally daily for 1 week. On days 5, 7, 9, 11, 13, and 15, the tumor size in each mouse was measured. The mice were subsequently sacrificed, and the tumor tissue was collected for analysis.
All data were analyzed and graphed using GraphPad Prism 7.0. Data were expressed as (mean ± SD). Group t-tests were used for two independent samples. The relationship between GLI1 and PRKACB expression and pathological parameters was examined using the Fisher’s exact test. Survival analysis was performed by Kaplan–Meier curves. Statistical significance was set at P < 0.05.
A TIMER database was used to analysis the expression of GLI1 in various cancer. GLI1 expression significantly increased in gastric cancer, kidney renal clear cell carcinoma and hepatocellular carcinoma and we selected GC for our study (Figure 1A). For gastric cancer in the TCGA database, high GLI1 expression was significantly associated with clinical stage and lymph node status (Figures 1B, C). A survival analysis of patients with high GLI1 expression, using the UALCAN online database showed that GLI1 expression was associated with poor prognosis in patients with GC (Figure 1D). In addition, GLI1 protein level in gastric cancer tissues was significantly higher than that in tumor-adjacent tissues, which is consistent with the results of previous studies (Figure 1E).
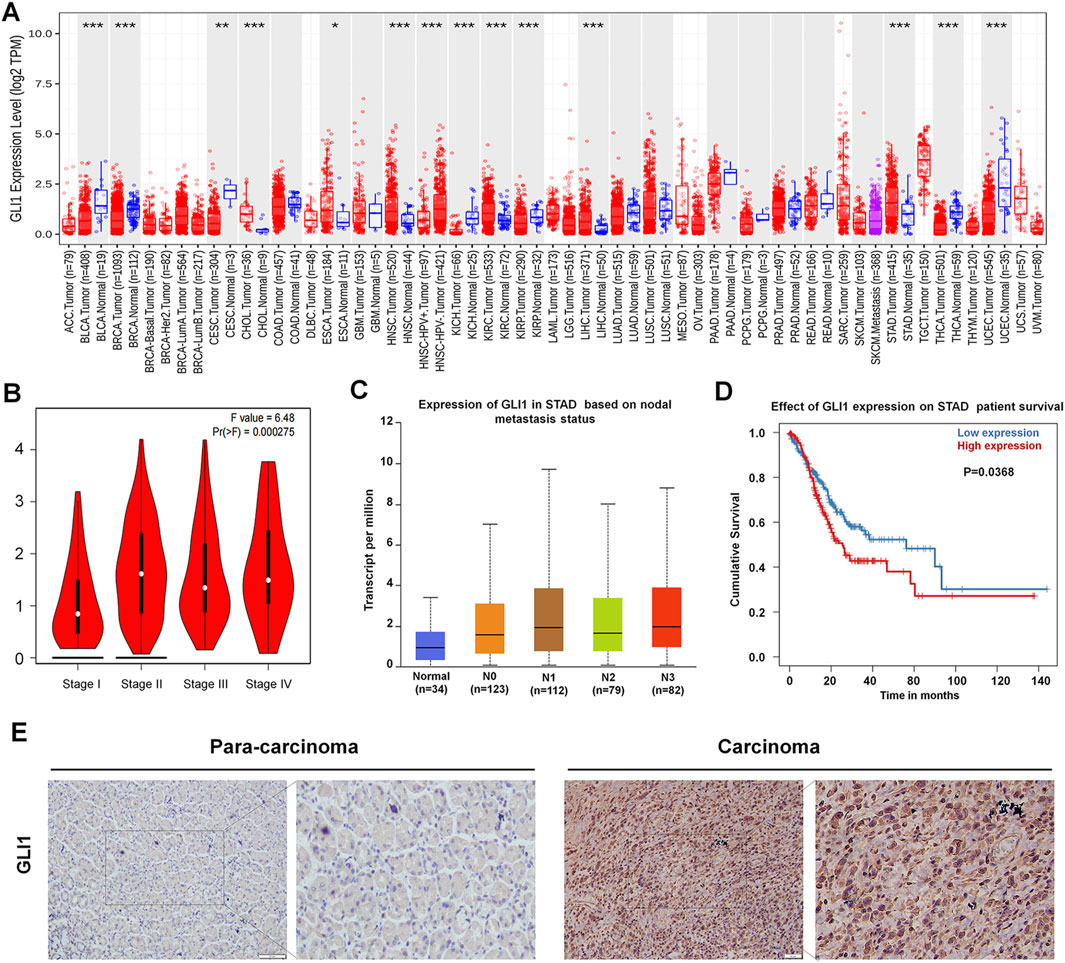
Figure 1. GLI1 expression in gastric cancer. (A) GLI1 expression in various cancers, from the TIMER database. (B) GLI1 expression is correlated with clinical stages. (C) GLI1 expression is correlated with lymph node metastasis status. (D) GLI1 expression is correlated with poor prognosis. (E) GLI1 expression in patients with gastric cancer.
Circular dichroism (CD) analysis revealed that the GLI1 promoter region can form G-quadruplex structures (Figure 2A). We mutated the region of the GLI1 promoter that readily formed G-quadruplexes and screened the best mutated fragments using a dual-luciferase reporter assay (Figures 2B, C). Using the FDA drug library (Selleck, Cherry pick libarary, Z590771), we initially screened six drugs that only targeted wild-type gastric cancer cells but had no effect on mutated-type gastric cancer cells (Figure 2D). To further demonstrate the effects of these drugs, we validated them by using a dual-luciferase reporter assay. The results showed that alpha-estradiol (α-Estradiol) and (R)-(−)-ibuprofen significantly reduced the expression of GLI1 transcripts (Figure 2E). In addition, Western blotting results showed that the protein level of GLI1 was significantly reduced with the addition of alpha-estradiol and (R)-(−)-ibuprofen compared to that in the control group (Figure 2F). These results suggest that α-estradiol and (R)-(−)-ibuprofen can act as G-quadruplexes stabilizers on the GLI1 promoter to influence the state of gastric cancer cells.
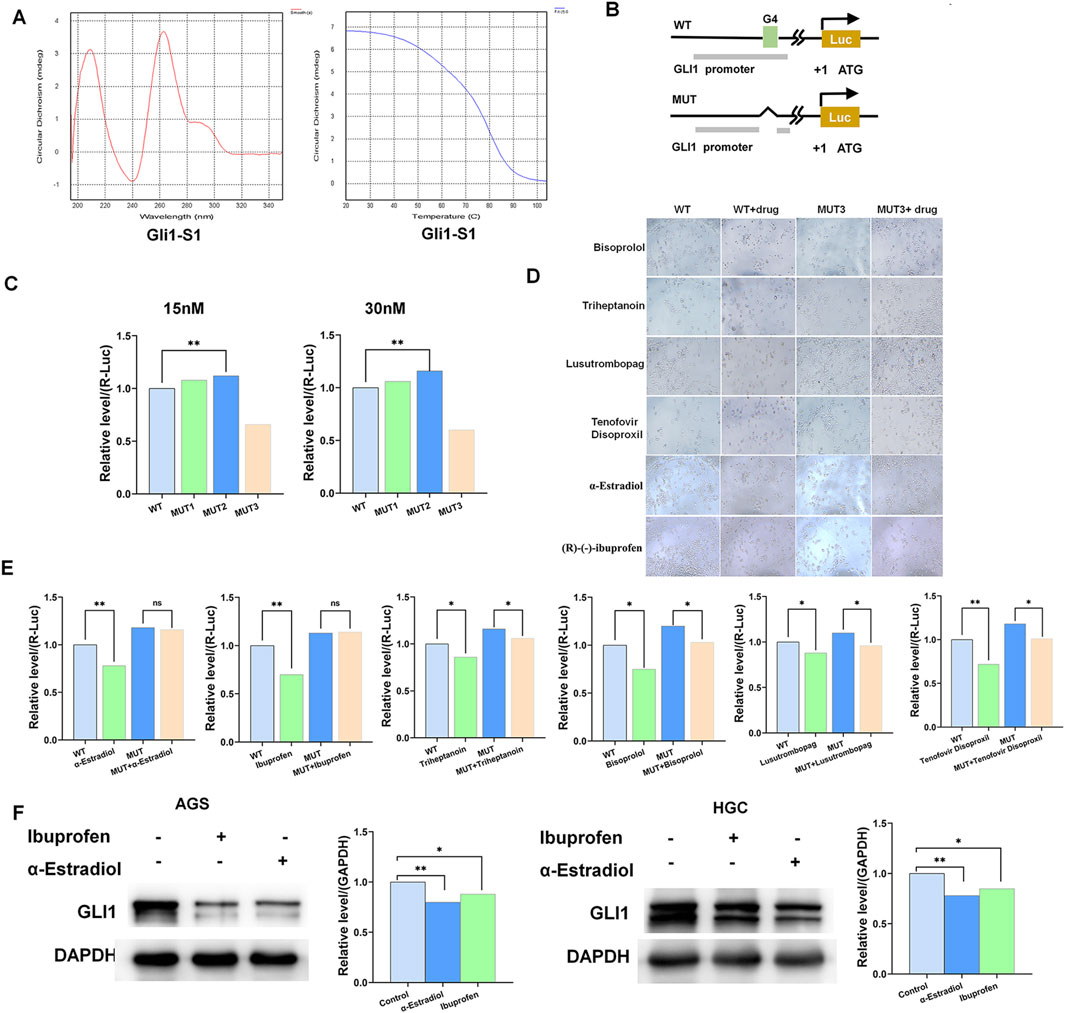
Figure 2. α-Estradiol and ibuprofen as the G-quadruplex stabilizers of the GLI1 promoter mediate cell migration and invasion in gastric cancer. (A) Circular dichroism spectrum of GLI1 G-quadruplex. (B, C) The activity of the G-quadruplex in HGC cells determined by luciferase reporter assays. (D, E) Alpha-estradiol and ibuprofen stabilize the G-quadruplex, to inhibiting gastric cancer cell proliferation. (F) GLI1 expression is inhibited by alpha-estradiol and ibuprofen treatment.
To validate the effects of alpha-estradiol and (R)-(−)-ibuprofen, a series of in vitro experiments were conducted. The results showed that the cell proliferation rate was significantly reduced in the groups treated with alpha-estradiol or (R)-(−)-ibuprofen (Figure 3A). Wound healing was significantly slower than that in the group without the addition of estradiol (Figure 3B). Similarly, the migration and invasion of AGS and HGC cells were significantly reduced by the addition of alpha-estradiol and (R)-(−)-ibuprofen compared to those in the control group (Figure 3C). These results show that alpha-estradiol and (R)-(−)-ibuprofen inhibit the migration and invasion of gastric cancer cells by regulating the stability of G-quadruplexes in the GLI1 promoter.
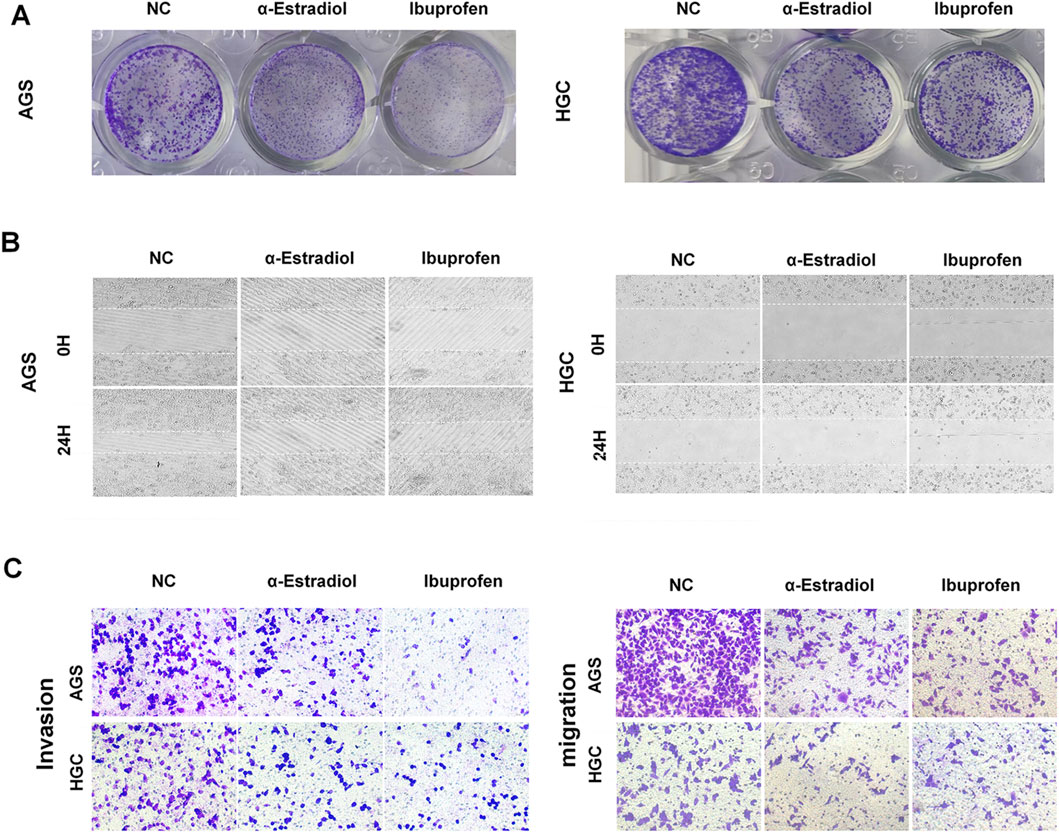
Figure 3. Alpha-estradiol and ibuprofen mediated gastric cancer cell migration and invasion. (A) Alpha-estradiol and ibuprofen inhibit cell proliferation in the colony-formation assay. (B) Alpha-estradiol and ibuprofen inhibit cell migration in the wound healing assay. (C) Alpha-estradiol and ibuprofen inhibit cell invasion in the transwell assay.
To further validate the regulatory functions of alpha-estradiol and (R)-(−)-ibuprofen in gastric cancer growth, we implanted human HGC cells into BALB/c nude mice. In vivo, the results showed that alpha-estradiol and (R)-(−)-ibuprofen significantly inhibited tumor growth. This was consistent with the results obtained using GANT61, a laboratory-recognized GLI1 inhibitor (Figure 4A). In addition, there was a corresponding improvement in tumor volume and weight after treatment with alpha-estradiol and (R)-(−)-ibuprofen (Figures 4B, C). The effects of alpha-estradiol and (R)-(−)-ibuprofen on invasion and metastasis of gastric cancer cells were investigated by tail vein injection experiments. The results showed that alpha-estradiol and (R)-(−)-ibuprofen inhibited the lung metastasis of gastric cancer cells (Figure 4D). In addition, immunohistochemical analysis of tumor tissues showed that alpha-estradiol and (R)-(−)-ibuprofen inhibited the expression of GLI1, Zeb1 and Vimentin (Figure 4E). These data confirmed that GLI1-mediated tumor growth and metastasis in gastric cancer can be inhibited by alpha-estradiol and (R)-(−)-ibuprofen.
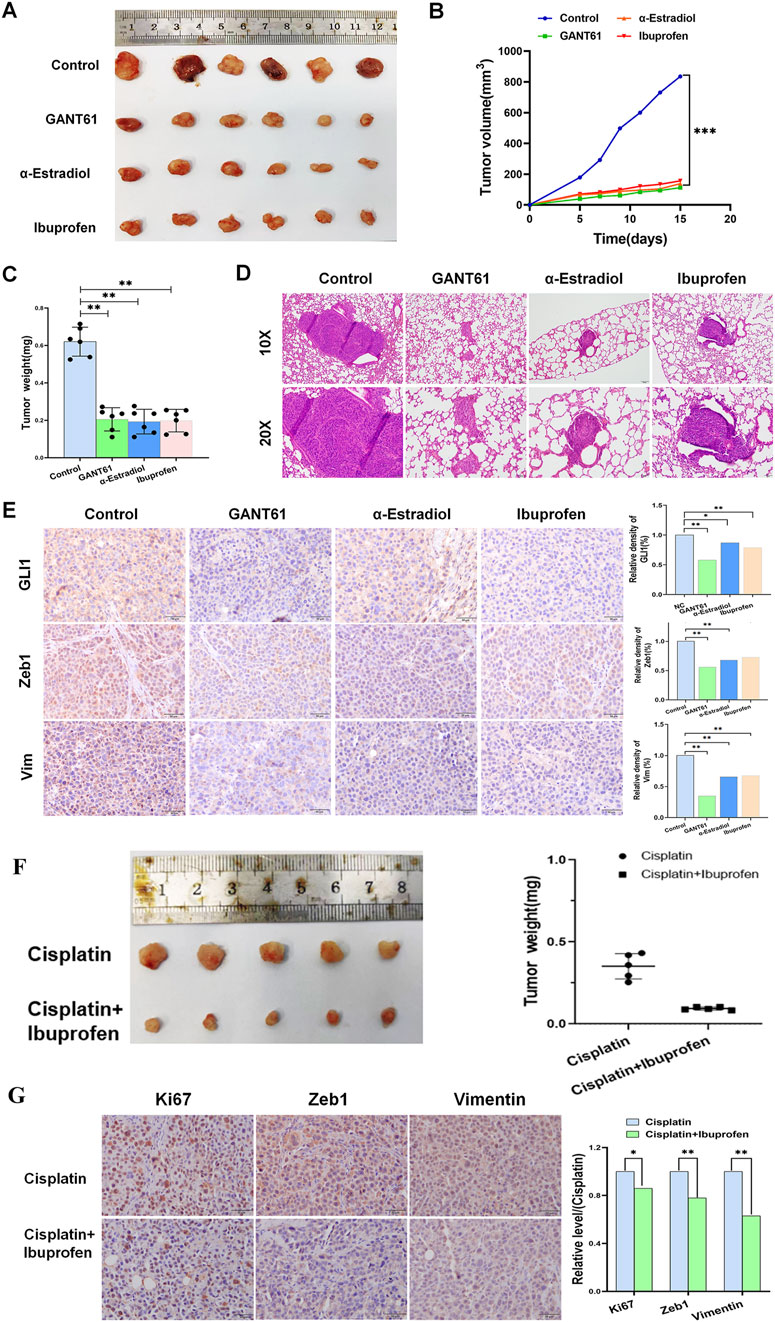
Figure 4. Alpha-estradiol and ibuprofen suppress gastric cancer cell growth in vivo. (A) Representative tumor size of isolated tumors from mice injected with HGC cells. (B) HGC cell-derived xenograft tumor volume measurements. (C) HGC cell-derived xenograft tumor weight from mice. (D) Representative hematoxylin and eosin staining of lung from HGC cell-derived xenograft mice with GANT61, alpha-estradiol and ibuprofen treatment. (E) GLI1, Zeb1, and Vimentin expression in GANT61, alpha-estradiol and ibuprofen treatment groups. (F) Ibuprofen in combination with Cisplatin decreased tumor size and weight compared with Cisplatin alone. (G) The expression of Ki67, Zeb1, Vimentin decreased in combination treatment. *P < 0.05, **P < 0.01.
Given the clinical roles of these two drugs, we speculated that they could be used as adjuvants to enhance the efficacy of chemotherapy drugs. The results showed that cisplatin treatment in combination with (R)-(−)-ibuprofen was more effective than cisplatin treatment alone (Figure 4F). In addition, immunohistochemistry results showed that the combination therapy further reduced the expression of Zeb1 and Ki67, which was consistent with previous results (Figure 4G).
It was reported that PRKACB expression was elevated in gastric cancer tissues and was associated with tumor development. To verify whether there was an association between GLI1 and PRKACB, we analyzed the data using the TIMER database and found that the correlation coefficient between PRKACB and GLI1 expression in gastric cancer was the highest amongst the analyzed tumors (Figures 5A, B). Patients with high PRKACB expression showed poorer survival rates (Figure 5C) and significantly correlated with age, Lauren stage, PNI, TNM stage, pathological stage, and Borrmann stage (Table 1). In addition, the expression levels of GLI1 and PRKACB were reduced after treatment with alpha-estradiol and (R)-(−)-ibuprofen in gastric xenograft tissues (Figure 5D). These results suggested that GLI1 may be related to PRKACB expression in gastric cancer.
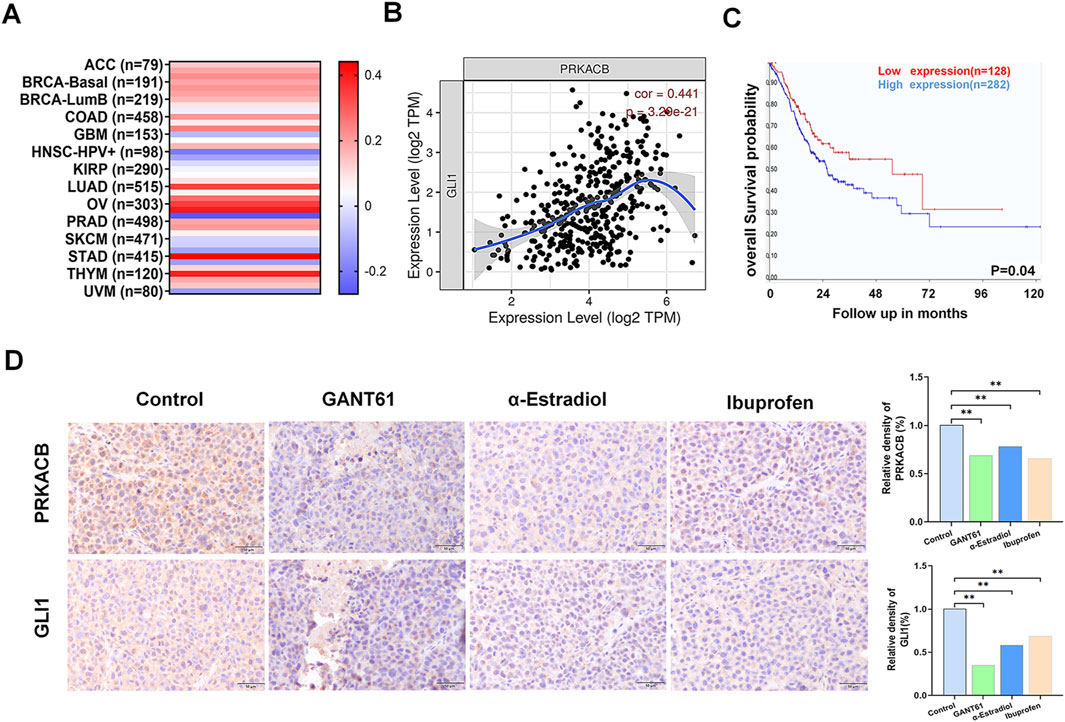
Figure 5. GLI1 and PRKACB expressions are correlated in gastric cancer. (A) Correlation between GLI1 and PRKACB expression in various cancers; (B) The association of GLI1 expression with PRKACB. (C) PRKACB expression correlates with poor prognosis. (D) GLI1 and PRKACB expression in GANT61, alpha-estradiol and ibuprofen treatment groups in gastric cancer and tumor-adjacent tissues. Scale bars 100 μm *P < 0.05.
To verify the relationship between GLI1 and PRKACB, we selected AGS and HGC cell lines with low PRKACB expression (Figure 6A). After adding GANT61, we found that PRKACB expression decreased, along with a reduction in GLI1 expression (Figure 6B). GLI1 acts as a transcription factor that regulates the expression of several molecules. We speculate that GLI1 directly binds to the promoter region of PRKACB and regulates its expression. Transcriptional element analysis (http://jaspar.genereg.net) revealed that the transcription factor GLI1 directly bound to the PRKACB promoter at −418 to −408 (Figure 6C). Next, we conducted ChIP. The results showed that GLI1 could bind to the promoter of PRKACB, and that this binding was reduced upon the addition of GANT61 (Figure 6D). The addition of GANT61 to the control and PRKACB overexpression groups significantly suppressed epithelial mesenchymal transition markers compared with the control and PRKACB overexpression groups (Figures 6E, F). Taken together, these findings suggest that GLI1 mediates PRKACB expression in gastric cells in vitro.
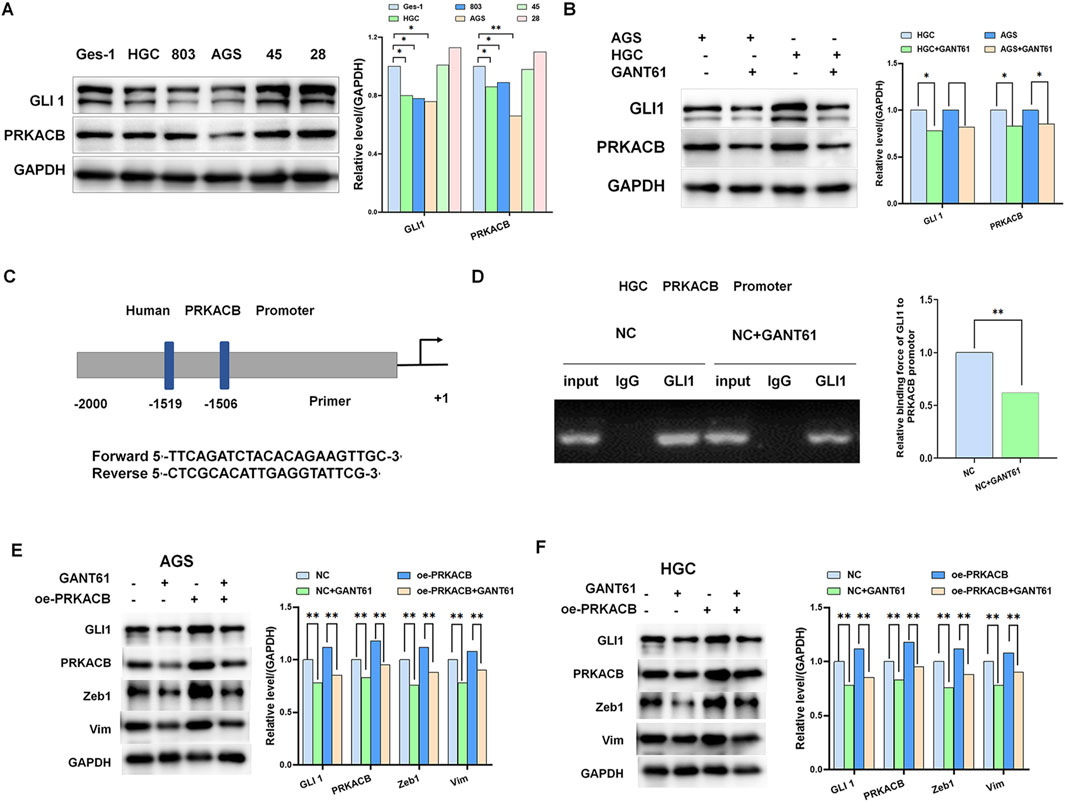
Figure 6. GLI1 mediates PRKACB expression in gastric cancer in vitro. (A) Expression of GLI1 and PRKACB in gastric cancer cell lines. (B) GANT61 inhibits PRKACB expression in HGC and AGS gastric cancer cell lines. (C, D) Chromatin immunoprecipitation assay showing GLI1 binding to the PRKACB promoter in gastric cancer. (E, F) GANT61 downregulated the expression of PRKACB and Zeb1 in gastric cancer. *P < 0.05, **P < 0.01.
Both GLI1and PRKACB have been reported to significantly promote the proliferation, migration, and invasion of cancer cells. The EdU assay showed that the proliferation of the GANT61 and overexpressing (oe)-PRKACB + GANT61 groups was significantly lower than that of the control and oe-PRKACB groups (Figure 7A). Wound healing in the GANT61 and oe-PRKACB + GANT61 groups was significantly slower than in the control and oe-PRKACB groups (Figure 7B). The GANT61 and oe-PRKACB + GANT61 groups showed reduced migration and invasion of AGS and HGC cells compared to the control and oe-PRKACB groups (Figure 7C).
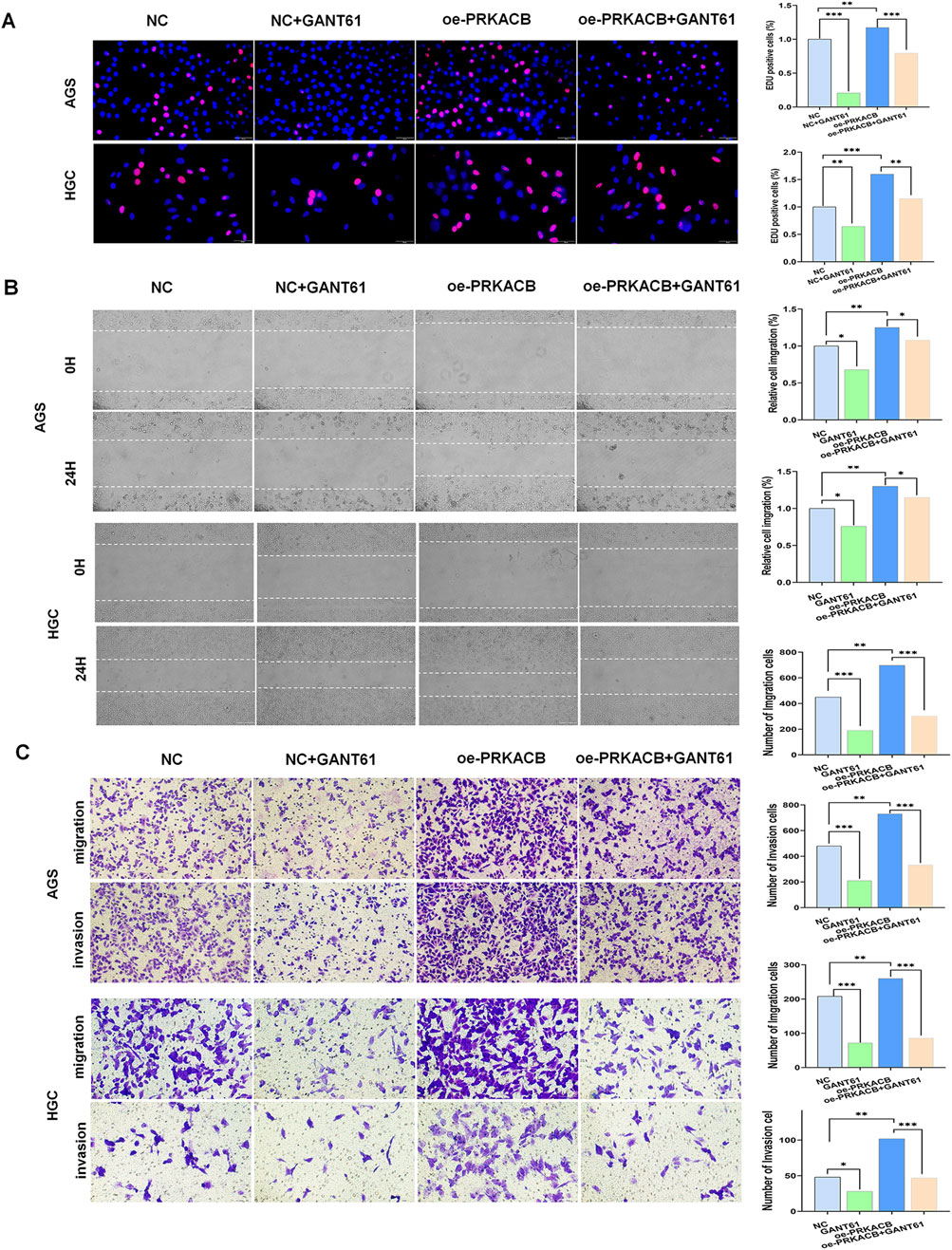
Figure 7. The expression of GLI1 and PRKACB synergistically affects cell proliferation, migration, invasion and stem cell-like properties in HGC and AGS cells. (A) EdU incorporation assay shows that silencing GLI1 and/or PRKACB reduces cell proliferation. Red denotes positive spots, while blue (DAPI) indicates nuclei (n = 3). Scale bar, 100 µm. (B) Wound healing assay shows that GLI1 and/or PRKACB downregulation inhibit the migration of HGC and AGS cells. Scale bar, 100 µm (n = 3). *P < 0.05, **P < 0.01. (C) Transwell assays show that GLI1 and/or PRKACB downregulation inhibit the invasion of HGC and AGS cells. Scale bar, 100 µm (n = 3). *P < 0.05, **P < 0.01.
GLI1 and PRKACB are highly correlated with cell stemness, so we further explored the correlation with cancer cell stemness. SOX2 and CD44 are markers for cancer stem cells. The levels of CD44 and SOX2 decreased after treatment with GANT61 in AGS and HGC cell lines (Figures 8A, B). The sphere formation assay demonstrated that sphere formation efficiency was significantly attenuated in the GANT61 and oe-PRKACB + GANT61 groups compared to that in the control and oe-PRKACB groups in AGS and HGC cells (Figure 8C). These results suggested that GLI1 and PRKACB are involved in gastric cancer cell stemness.
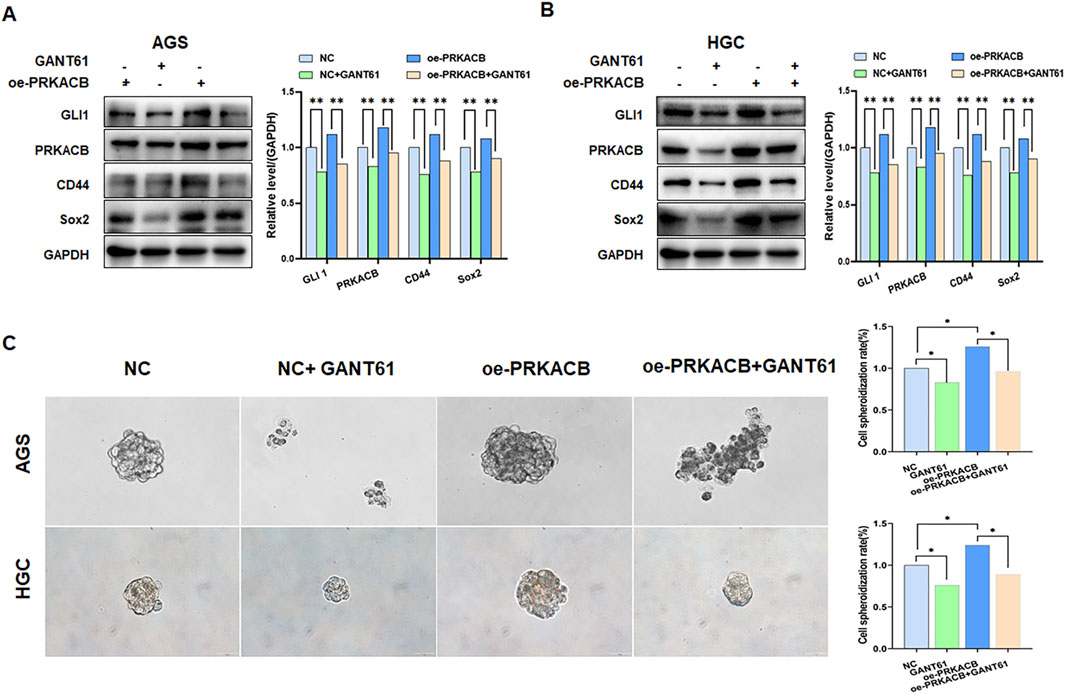
Figure 8. Expression of GLI1and PRKACB affects stem cell-like properties in HGC and AGS cells. (A, B) Inhibiting GLI1 or/and overexpressing PRKACB decreases CD44 in AGS and HGC cells. **P < 0.01. (C) Representative spheroid images formed by single cells with inhibition of GLI1 expression and/or overexpressing PRKACB.
GLI1 and PRKACB expression was examined using immunohistochemistry in tissue microarrays containing 30 gastric cancer and 30 tumor-adjacent tissue samples. GLI1 protein levels were significantly higher in gastric cancer tissues than in tumor-adjacent tissue samples, consistent with PRKACB protein levels in gastric cancer tissues (Figure 9A). Spearman’s correlation analysis revealed a correlation between GLI1 and PRKACB expression levels (Figure 9B) (P < 0.05). The data from the tissue microarrays suggest that GLI1 is strongly associated with PRKACB in patient samples.
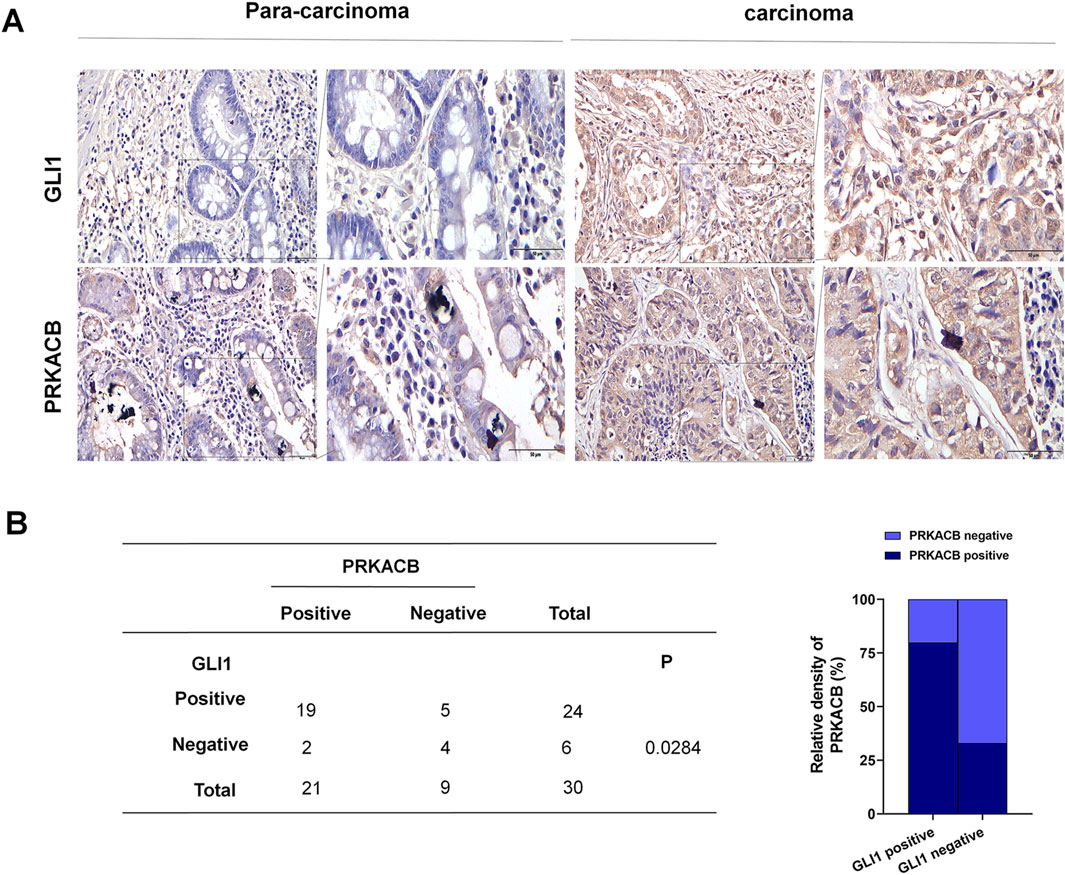
Figure 9. PRKACB and GLI1 expression in gastric cancer tissue microarray. (A) GLI1 and PRKACB expression in gastric cancer carcinoma and tumor-adjacent tissues. (B) Correlation between GLI1 and PRKACB expression in gastric cancer tissues. Scale bars are 100 µm.
Gastric cancer is a serious threat to human health. According to the 2021 statistics compiled by the American Cancer Society, incidence and mortality rates remain high (Siegel et al., 2021). The treatment of gastric cancer is multidisciplinary and comprehensive, including endoscopic treatment, surgery, radiotherapy, and systemic chemotherapy. However, the efficiency and side effects of these methods limit their clinical application, and targeted therapy, as a widely recognized direction of research on the disease, has been gradually applied to clinical research (Alsina et al., 2023). Drugs that specifically target nucleic acid structures through binding, intercalation, or covalent modification have been developed to treat various diseases (Tateishi-Karimata and Sugimoto, 2020). The impact of G4 structures on DNA and RNA functions presents an enticing opportunity to modulate these folded structures and exert control over their diverse functions. The distinctive structural features of G4 have facilitated the development of diverse structure-selective small molecule ligands (Carvalho et al., 2020; Berner et al., 2024). In this project, we combined previous studies with the current research status and used the GLI1 molecule in the Hh pathway as a regulatory molecule to develop inhibitors for G-quadruplexes, providing a new option for the treatment of gastric cancer. Furthermore, we confirmed that G-quadruplex regulators of the GLI1 promoter inhibited gastric cancer progression by downregulating PRKACB, an effector molecule downstream of GLI1.
Aberrant activation of Hh signaling is observed in many tumors and accounts for approximately 25% of human cancer deaths (Lum and Beachy, 2004). The Hh pathway is active in stem cells and largely inactive in adult organisms. Aberrant activation of this pathway in adult cells affects the downstream cellular processes and leads to the development of multiple cancers, including gastric cancer, tissue infiltration, and drug resistance (Karhadkar et al., 2004; Zhu et al., 2019; Huang et al., 2006). GLI is a transcription factor belonging to the zinc finger protein family and a major effector of Hh signaling (Doheny et al., 2020). In vertebrates, there are three members of the GLI gene family: GLI1, GLI2 and GLI3 (Niewiadomski et al., 2019). GLI1 can act as a transcriptional activator (Hynes et al., 1997) and Hh ligands can induce GLI1 expression, which provides a positive feedback loop for Hh signaling (Regl et al., 2002). In human gliomas the Hh/GLI1 pathway plays an important role in the self-renewal and tumorigenicity of cancer stem cell-like cells (Clement et al., 2007). Our study showed that GLI1 was overexpressed in gastric cancer tissue samples and that its expression was correlated with adverse clinicopathological parameters, suggesting a role for GLI1 in gastric carcinogenesis, progression, and metastasis. By far, no molecular inhibitor of GLI1 has been approved for the treatment of gastric cancer. Attempting to block or silence the expression of GLI1 will significantly help suppress gastric cancer progression.
In recent years, the exploration of nucleic acid secondary structures has drawn increasing attention. A G-quadruplex is a nucleic acid secondary structure usually found in telomeres, or the promoter region of a gene, and is formed by the folding of a sequence rich in guanine bases (Zhang et al., 2022) (Kosiol et al., 2021). The formation of G-quadruplexes is closely related to biological processes such as gene expression regulation, genome replication, and telomere elongation, and influences cellular senescence and apoptosis, which are involved in the onset and progression of cancer and other diseases (Tan et al., 2018; Hirashima and Seimiya, 2015; Murat and Balasubramanian, 2014; Huppert and Balasubramanian, 2005). G-quadruplexes located in the promoter and 5′-untranslated regions play a regulatory role in gene expression, and the formation of G-quadruplexes tends to repress the expression of oncogenes such as MYC, FGR, hTERT, and BCL2 (Hänsel-Hertsch et al., 2017; Hänsel-Hertsch et al., 2018; Lin et al., 2024). Therefore, these G-quadruplex structures may serve as potential targets for anticancer drugs. The guanine-rich sequence in the promoter region of GLI1 forms a G-quadruplex structure, which is a good target for inhibitor development. In this study, through meticulous molecular ligand screening, two inhibitors, namely, alpha-estradiol and (R)-(−)-ibuprofen, were identified. Our investigations revealed that these inhibitors could effectively modulate GLI1 expression and consequently intervene in the progression of gastric cancer, opening up new avenues for future therapeutic strategies.
Our results have showed that G-quadruplex regulators on the GLI1 promoter inhibited gastric cancer progression by downregulating PRKACB, an effector molecule downstream of GLI1. PRKACB is located at chromosome 1p31.1, has a total gene length of 160 kb, and is a member of the serine/threonine protein kinase family (Zhu et al., 2005). As an important molecule in the cellular cascade signaling pathway, the activated catalytic subunit can transmit extracellular stimulatory signals and participate in the regulation of cell proliferation, differentiation and apoptosis (Skalhegg and Tasken, 2000). It is a key link in the regulation of the cell cycle and is closely related to cell growth, differentiation, gene expression, tumor growth, metastasis, and other physiological and pathological processes (Yao et al., 2020; Higuchi et al., 2003; Kvissel et al., 2007). Our study showed that PRKACB is highly expressed in gastric cancer and acts as a downstream regulator of GLI1 to promote gastric cancer progression.
There are some limitations of the present study that could be addressed in our following study. G-quadruplex has potential as a drug target, but due to its wide distribution and diversity in the genome, developing GLI1 G-quadruplex targeted drugs with high specificity and minimal side effects still faces challenges. The clinical value should be further confirmed in a larger cohort including preclinical intervention studies. To further explore the apoptosis and the changes in the phases of the cell cycle via the GLI1/PRKACB pathway following treatment with these inhibitors would provide a more comprehensive understanding of the mechanisms.
In this study, a G-quadruplex was used as a target for inhibitor exploration, and a GLI1 molecule inhibitor was used as an entry point to verify the relationship between its aberrant activation and gastric cancer progression. In addition, G-quadruplex regulators of the GLI1 promoter inhibited gastric cancer cell proliferation, migration, and invasion by downregulating PRKACB, which provides more options for the clinical treatment of gastric cancer (Figure 10).
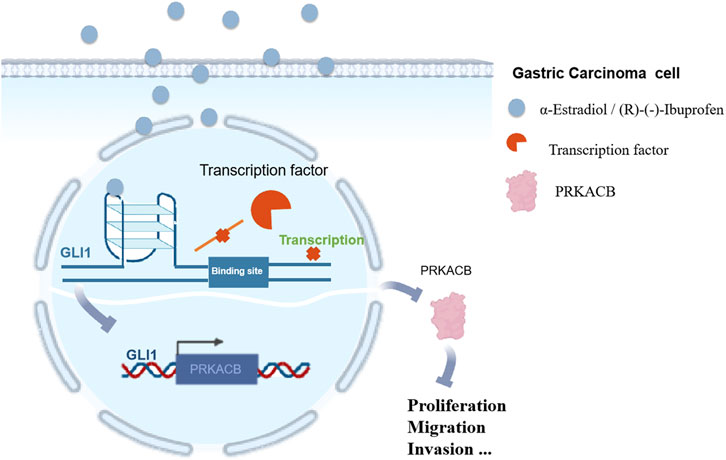
Figure 10. GLI1 G-quadruplex regulators inhibit gastric cancer progression via down-regulating PRKACB.
The data supporting the findings of this study are available from the corresponding author upon reasonable request.
The study was approved by the Ethics committee of the Institutional Animal Care and Use Committee of Central Hospital Affiliated to Shandong First Medical University and conformed to the Declaration of Helsinki (approval no. JNCH2021-77). The study involving the usage of human tissues was performed in accordance with the Declaration of Helsinki and with the approval of the ethics committee of Shanghai Outdo Biotech (approval no. SHXC2021YF01).
QL: Formal Analysis, Project administration, Writing–original draft, Writing–review and editing. PP: Methodology, Writing–original draft, Writing–review and editing. QX: Resources, Writing–original draft, Writing–review and editing. JL: Software, Writing–original draft, Writing–review and editing. JtW: Data curation, Writing–original draft, Writing–review and editing. JC: Investigation, Writing–original draft, Writing–review and editing. JW: Conceptualization, Investigation, Project administration, Validation, Writing–original draft, Writing–review and editing. YJ: Formal Analysis, Validation, Writing–original draft, Writing–review and editing. HS: Conceptualization, Methodology, Writing–original draft, Writing–review and editing. LZ: Conceptualization, Funding acquisition, Supervision, Validation, Writing–original draft, Writing–review and editing. XM: Conceptualization, Investigation, Supervision, Validation, Writing–original draft, Writing–review and editing.
The author(s) declare that financial support was received for the research and/or publication of this article. This work was supported by grants from the Natural Science Foundation of Shandong Province (ZR2022QC156), the National Natural Science Foundation of China (32170496 and 31970728).
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Aberger, F., Kern, D., Greil, R., and Hartmann, T. N. (2012). Canonical and noncanonical Hedgehog/GLI signaling in hematological malignancies. Vitam. Horm. 88, 25–54. doi:10.1016/B978-0-12-394622-5.00002-X
Alsina, M., Arrazubi, V., Diez, M., and Tabernero, J. (2023). Current developments in gastric cancer: from molecular profiling to treatment strategy. Nat. Rev. Gastroenterol. Hepatol. 20, 155–170. doi:10.1038/s41575-022-00703-w
Awadasseid, A., Ma, X., Wu, Y., and Zhang, W. (2021). G-quadruplex stabilization via small-molecules as a potential anti-cancer strategy. Biomed. Pharmacother. 139, 111550. doi:10.1016/j.biopha.2021.111550
Bariwal, J., Kumar, V., Dong, Y., and Mahato, R. I. (2019). Design of Hedgehog pathway inhibitors for cancer treatment. Med. Res. Rev. 39, 1137–1204. doi:10.1002/med.21555
Berner, A., Das, R. N., Bhuma, N., Golebiewska, J., Abrahamsson, A., Andreasson, M., et al. (2024). G4-Ligand-Conjugated oligonucleotides mediate selective binding and stabilization of individual G4 DNA structures. J. Am. Chem. Soc. 146, 6926–6935. doi:10.1021/jacs.3c14408
Carvalho, J., Mergny, J. L., Salgado, G. F., Queiroz, J. A., and Cruz, C. (2020). G-Quadruplex, friend or foe: the role of the G-quartet in anticancer strategies. Trends Mol. Med. 26, 848–861. doi:10.1016/j.molmed.2020.05.002
Clement, V., Sanchez, P., de Tribolet, N., Radovanovic, I., and Ruiz i Altaba, A. (2007). HEDGEHOG-GLI1 signaling regulates human glioma growth, cancer stem cell self-renewal, and tumorigenicity. Curr. Biol. 17, 165–172. doi:10.1016/j.cub.2006.11.033
Doheny, D., Manore, S. G., Wong, G. L., and Lo, H. W. (2020). Hedgehog signaling and truncated GLI1 in cancer. Cells 9, 2114. doi:10.3390/cells9092114
Espiard, S., Knape, M. J., Bathon, K., Assié, G., Rizk-Rabin, M., Faillot, S., et al. (2018). Activating PRKACB somatic mutation in cortisol-producing adenomas. JCI Insight 3, e98296. doi:10.1172/jci.insight.98296
Falch, C. M., Sundaram, A. Y. M., Øystese, K. A., Normann, K. R., Lekva, T., Silamikelis, I., et al. (2018). Gene expression profiling of fast- and slow-growing non-functioning gonadotroph pituitary adenomas. Eur. J. Endocrinol. 178, 295–307. doi:10.1530/EJE-17-0702
Furuta, K., Arao, T., Sakai, K., Kimura, H., Nagai, T., Tamura, D., et al. (2012). Integrated analysis of whole genome exon array and array-comparative genomic hybridization in gastric and colorectal cancer cells. Cancer Sci. 103, 221–227. doi:10.1111/j.1349-7006.2011.02132.x
Hänsel-Hertsch, R., Di Antonio, M., and Balasubramanian, S. (2017). DNA G-quadruplexes in the human genome: detection, functions and therapeutic potential. Nat. Rev. Mol. Cell. Biol. 18, 279–284. doi:10.1038/nrm.2017.3
Hänsel-Hertsch, R., Spiegel, J., Marsico, G., Tannahill, D., and Balasubramanian, S. (2018). Genome-wide mapping of endogenous G-quadruplex DNA structures by chromatin immunoprecipitation and high-throughput sequencing. Nat. Protoc. 13, 551–564. doi:10.1038/nprot.2017.150
Hegde, G. V., Munger, C. M., Emanuel, K., Joshi, A. D., Greiner, T. C., Weisenburger, D. D., et al. (2008). Targeting of sonic hedgehog-GLI signaling: a potential strategy to improve therapy for mantle cell lymphoma. Mol. Cancer Ther. 7, 1450–1460. doi:10.1158/1535-7163.MCT-07-2118
Higuchi, H., Yamashita, T., Yoshikawa, H., and Tohyama, M. (2003). PKA phosphorylates the p75 receptor and regulates its localization to lipid rafts. Embo J. 22, 1790–1800. doi:10.1093/emboj/cdg177
Hirashima, K., and Seimiya, H. (2015). Telomeric repeat-containing RNA/G-quadruplex-forming sequences cause genome-wide alteration of gene expression in human cancer cells in vivo. Nucleic Acids Res. 43, 2022–2032. doi:10.1093/nar/gkv063
Huang, S., He, J., Zhang, X., Bian, Y., Yang, L., Xie, G., et al. (2006). Activation of the hedgehog pathway in human hepatocellular carcinomas. Carcinogenesis 27, 1334–1340. doi:10.1093/carcin/bgi378
Huppert, J. L., and Balasubramanian, S. (2005). Prevalence of quadruplexes in the human genome. Nucleic Acids Res. 33, 2908–2916. doi:10.1093/nar/gki609
Hynes, M., Stone, D. M., Dowd, M., Pitts-Meek, S., Goddard, A., Gurney, A., et al. (1997). Control of cell pattern in the neural tube by the zinc finger transcription factor and oncogene Gli-1. Neuron 19, 15–26. doi:10.1016/s0896-6273(00)80344-x
Jagani, Z., Mora-Blanco, E. L., Sansam, C. G., McKenna, E. S., Wilson, B., Chen, D., et al. (2010). Loss of the tumor suppressor Snf5 leads to aberrant activation of the Hedgehog-Gli pathway. Nat. Med. 16, 1429–1433. doi:10.1038/nm.2251
Kamilaris, C. D. C., Hannah-Shmouni, F., and Stratakis, C. A. (2020). Adrenocortical tumorigenesis: lessons from genetics. Best. Pract. Res. Clin. Endocrinol. Metab. 34, 101428. doi:10.1016/j.beem.2020.101428
Karhadkar, S. S., Bova, G. S., Abdallah, N., Dhara, S., Gardner, D., Maitra, A., et al. (2004). Hedgehog signalling in prostate regeneration, neoplasia and metastasis. Nature 431, 707–712. doi:10.1038/nature02962
Kern, D., Regl, G., Hofbauer, S. W., Altenhofer, P., Achatz, G., Dlugosz, A., et al. (2015). Hedgehog/GLI and PI3K signaling in the initiation and maintenance of chronic lymphocytic leukemia. Oncogene 34, 5341–5351. doi:10.1038/onc.2014.450
Kosiol, N., Juranek, S., Brossart, P., Heine, A., and Paeschke, K. (2021). G-quadruplexes: a promising target for cancer therapy. Mol. Cancer 20, 40. doi:10.1186/s12943-021-01328-4
Kvissel, A. K., Ramberg, H., Eide, T., Svindland, A., Skålhegg, B. S., and Taskén, K. A. (2007). Androgen dependent regulation of protein kinase A subunits in prostate cancer cells. Cell. Signal 19, 401–409. doi:10.1016/j.cellsig.2006.07.011
Li, H., Yue, D., Jin, J. Q., Woodard, G. A., Tolani, B., Luh, T. M., et al. (2016). Gli promotes epithelial-mesenchymal transition in human lung adenocarcinomas. Oncotarget 7, 80415–80425. doi:10.18632/oncotarget.11246
Lin, H., Hassan Safdar, M., Washburn, S., S, S. A., Dickerhoff, J., Ayers, M., et al. (2024). Fibroblast growth receptor 1 is regulated by G-quadruplex in metastatic breast cancer. Commun. Biol. 7, 963. doi:10.1038/s42003-024-06602-x
Lum, L., and Beachy, P. A. (2004). The Hedgehog response network: sensors, switches, and routers. Science 304, 1755–1759. doi:10.1126/science.1098020
Murat, P., and Balasubramanian, S. (2014). Existence and consequences of G-quadruplex structures in DNA. Curr. Opin. Genet. Dev. 25, 22–29. doi:10.1016/j.gde.2013.10.012
Niewiadomski, P., Niedziółka, S. M., Markiewicz, Ł., Uśpieński, T., Baran, B., and Chojnowska, K. (2019). Gli proteins: regulation in development and cancer. Cells 8, 147. doi:10.3390/cells8020147
Nishio, M., Tsukakoshi, K., and Ikebukuro, K. (2021). G-quadruplex: flexible conformational changes by cations, pH, crowding and its applications to biosensing. Biosens. Bioelectron. 178, 113030. doi:10.1016/j.bios.2021.113030
Park, J. Y., and Herrero, R. (2021). Recent progress in gastric cancer prevention. Best. Pract. Res. Clin. Gastroenterol. 50-51, 101733. doi:10.1016/j.bpg.2021.101733
Qi, W., Yang, Z., Feng, Y., Li, H., Che, N., Liu, L., et al. (2020). Gli1 regulates stemness characteristics in gastric adenocarcinoma. Diagn Pathol. 15, 60. doi:10.1186/s13000-020-00949-5
Regl, G., Neill, G. W., Eichberger, T., Kasper, M., Ikram, M. S., Koller, J., et al. (2002). Human GLI2 and GLI1 are part of a positive feedback mechanism in Basal Cell Carcinoma. Oncogene 21, 5529–5539. doi:10.1038/sj.onc.1205748
Rubin, L. L., and de Sauvage, F. J. (2006). Targeting the Hedgehog pathway in cancer. Nat. Rev. Drug Discov. 5, 1026–1033. doi:10.1038/nrd2086
Shimo, T., Matsumoto, K., Takabatake, K., Aoyama, E., Takebe, Y., Ibaragi, S., et al. (2016). The role of sonic hedgehog signaling in osteoclastogenesis and jaw bone destruction. PLoS One 11, e0151731. doi:10.1371/journal.pone.0151731
Siegel, R. L., Miller, K. D., Fuchs, H. E., and Jemal, A. (2021). Cancer statistics, 2021. CA Cancer J. Clin. 71, 7–33. doi:10.3322/caac.21654
Skalhegg, B. S., and Tasken, K. (2000). Specificity in the cAMP/PKA signaling pathway. Differential expression,regulation, and subcellular localization of subunits of PKA. Front. Biosci. 5, D678–D693. doi:10.2741/skalhegg
Smyth, E. C., Nilsson, M., Grabsch, H. I., van Grieken, N. C., and Lordick, F. (2020). Gastric cancer. Lancet 396, 635–648. doi:10.1016/S0140-6736(20)31288-5
Stecca, B., and Ruiz i Altaba, A. (2010). Context-dependent regulation of the GLI code in cancer by HEDGEHOG and non-HEDGEHOG signals. J. Mol. Cell. Biol. 2, 84–95. doi:10.1093/jmcb/mjp052
Sung, H., Ferlay, J., Siegel, R. L., Laversanne, M., Soerjomataram, I., Jemal, A., et al. (2021). Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71, 209–249. doi:10.3322/caac.21660
Tan, W., Yi, L., Zhu, Z., Zhang, L., Zhou, J., and Yuan, G. (2018). Hsa-miR-1587 G-quadruplex formation and dimerization induced by NH(4)(+), molecular crowding environment and jatrorrhizine derivatives. Talanta 179, 337–343. doi:10.1016/j.talanta.2017.11.041
Tassinari, M., Richter, S. N., and Gandellini, P. (2021). Biological relevance and therapeutic potential of G-quadruplex structures in the human noncoding transcriptome. Nucleic Acids Res. 49, 3617–3633. doi:10.1093/nar/gkab127
Tateishi-Karimata, H., and Sugimoto, N. (2020). Chemical biology of non-canonical structures of nucleic acids for therapeutic applications. Chem. Commun. (Camb) 56, 2379–2390. doi:10.1039/c9cc09771f
Wang, F., Stappenbeck, F., and Parhami, F. (2019). Inhibition of hedgehog signaling in fibroblasts, pancreatic, and lung tumor cells by Oxy186, an oxysterol analogue with drug-like properties. Cells 8, 509. doi:10.3390/cells8050509
Wang, Y., Guo, T., Liu, Q., and Xie, X. (2020). CircRAD18 accelerates the progression of acute myeloid leukemia by modulation of miR-206/PRKACB Axis. Cancer Manag. Res. 12, 10887–10896. doi:10.2147/CMAR.S277432
Xu, Y., Song, S., Wang, Z., and Ajani, J. A. (2019). The role of hedgehog signaling in gastric cancer: molecular mechanisms, clinical potential, and perspective. Cell. Commun. Signal 17, 157. doi:10.1186/s12964-019-0479-3
Yan, Z., Xu, L., Zhang, J., Lu, Q., Luo, S., and Xu, L. (2016). Aldehyde dehydrogenase 1A1 stabilizes transcription factor Gli2 and enhances the activity of Hedgehog signaling in hepatocellular cancer. Biochem. Biophys. Res. Commun. 471, 466–473. doi:10.1016/j.bbrc.2016.02.052
Yao, X., Hu, W., Zhang, J., Huang, C., Zhao, H., and Yao, X. (2020). Application of cAMP-dependent catalytic subunit β (PRKACB) low expression in predicting worse overall survival: a potential therapeutic target for colorectal carcinoma. J. Cancer 11, 4841–4850. doi:10.7150/jca.46156
Zhang, C., Fu, S., Zhang, F., Han, M., Wang, X., Du, J., et al. (2022). Affibody modified G-quadruplex DNA micelles incorporating polymeric 5-fluorodeoxyuridine for targeted delivery of curcumin to enhance synergetic therapy of HER2 positive gastric cancer. Nanomater. (Basel) 12, 696. doi:10.3390/nano12040696
Zhu, H., Lu, W., Laurent, C., Shaw, G. M., Lammer, E. J., and Finnell, R. H. (2005). Genes encoding catalytic subunits of protein kinase A and risk of spina bifida. Birth Defects Res. A Clin. Mol. Teratol. 73, 591–596. doi:10.1002/bdra.20175
Keywords: G-quadruplex, GLI1, alpha-estradiol, (R)-(−)-ibuprofen, PRKACB, gastric cancer
Citation: Li Q, Pan P, Xian Q, Li J, Wang J, Cai J, Wang J, Jia Y, Sun H, Zhang L and Ma X (2025) Alpha-estradiol and (R)-(−)-ibuprofen inhibit gastric cancer progression via GLI1 G-quadruplex. Front. Pharmacol. 16:1492694. doi: 10.3389/fphar.2025.1492694
Received: 07 September 2024; Accepted: 17 March 2025;
Published: 04 April 2025.
Edited by:
Angelo A. Izzo, University of Naples Federico II, ItalyReviewed by:
Azna Zuberi, Northwestern University, United StatesCopyright © 2025 Li, Pan, Xian, Li, Wang, Cai, Wang, Jia, Sun, Zhang and Ma. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Lulu Zhang, emhhbmdfbHVsdV8yMDE2QDE2My5jb20=; Xiaoli Ma, TWF4aWFvbGlfam5Ac2R1LmVkdS5jbg==
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.