
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
REVIEW article
Front. Oncol. , 17 March 2025
Sec. Cancer Molecular Targets and Therapeutics
Volume 15 - 2025 | https://doi.org/10.3389/fonc.2025.1521204
This article is part of the Research Topic New potential biomarkers and cellular strategies for the study of prostate cancer and testicular cancer cells View all 6 articles
 Fidelis Charles Bugoye1,2*
Fidelis Charles Bugoye1,2* Rispah Torrorey-Sawe2*
Rispah Torrorey-Sawe2* Richard Biegon2
Richard Biegon2 Nazima Dharsee3
Nazima Dharsee3 Fidelice Mafumiko1
Fidelice Mafumiko1 Herry Kibona4
Herry Kibona4 Said Aboud5
Said Aboud5 Kirtika Patel2
Kirtika Patel2 Simeon Mining2
Simeon Mining2Background: Prostate cancer is defined by the suppression of genes that suppress tumours and the activation of proto-oncogenes. These are the hallmarks of prostate cancer, and they have been linked to numerous genomic variations, which lead to unfavourable treatment outcomes. Prostate cancer can be categorised into various risk groups of tumour molecular subtypes grounded in the idea of genomic structural variations connected to TMPRSS2:ERG fusion and loss of PTEN. Research suggests that certain genomic alterations may be more prevalent or exhibit different patterns in prostate cancer tumours across populations. Studies have reported a higher frequency of PTEN loss and TMPRSS2:ERG fusion in prostate tumours of Black/African American men, which may contribute to the more aggressive nature of the disease in this population. Thus, therapeutically important information can be obtained from these structural variations, including correlations with poor prognosis and disease severity.
Methods: Peer-reviewed articles from 1998 to 2024 were sourced from PubMed and Google Scholar. During the review process, the following search terms were employed: “Tumour suppressor genes OR variations OR alterations OR oncogenes OR diagnostics OR ethnicity OR biomarkers OR prostate cancer genomics OR prostate cancer structural variations OR tumour and molecular subtypes OR therapeutic implications OR immunotherapy OR immunogenetics.”
Results: There was a total of 13,012 results for our search query: 5,903 publications from Google Scholar with the patent and citation unchecked filer options, and 7127 articles from PubMed with the abstract, free full text, and full-text options selected. Unpublished works were not involved. Except for four articles published between 1998 and 1999, all other selected articles published in 2000 and later were considered. However, papers with irrelevant information or redundant or duplicate content were not chosen for this review. Thus, 134 met the inclusion criteria and were ultimately retained for this review.
Conclusion: This review extracted 134 relevant articles about genomic structure variations in prostate cancer. Our findings demonstrate the importance of PTEN and TMPRSS2:ERG fusion and tumour molecular subtyping in prostate cancer precision medicine.
Prostate cancer (PCa) is one of the most prevalent cancers worldwide and a leading cause of cancer-related deaths among men (1, 2). Its diverse clinical presentation and unpredictable prognosis make it a significant public health concern. Developments in molecular oncology have drawn attention to how important genetic and molecular alterations are in promoting the advancement of PCa and resistance to treatment. These include deletion of the PTEN tumour-suppressor, TMPRSS2:ERG gene fusion, and other molecular subtypes that have become important biomarkers with significant effects on treatment and prognosis (3–5).
This review explores the therapeutic potential of targeting ERG/PTEN molecular subtypes emphasising their role in advancing personalised medicine for PCa treatment. The TMPRSS2:ERG fusion, one of the most frequent genetic alterations in PCa, occurs in 40%–50% of cases (3–5). This fusion produces a distinct molecular subtype linked to intermediate-to-aggressive disease symptoms, with the androgen-responsive TMPRSS2 promoter driving the overexpression of the ERG transcription factor (3–5). PCa progression is significantly influenced by ERG changes as well as the loss or inactivation of PTEN, a key regulator of the PI3K/AKT signalling pathway. PTEN deletions are found in approximately 40% of primary PCa cases and are significantly more common in castration-resistant and metastatic forms of the illness (3–5). The PI3K/AKT signalling pathway is uncheckedly activated when PTEN is lost, which promotes tumour development, survival, and resistance to treatment. The interplay between TMPRSS2:ERG fusion (often represented by ERG expression) and PTEN loss allows for the classification of PCa into molecular subtypes providing critical insights into tumour biology (4). It is noteworthy that tumours with both PTEN loss and ERG expression exhibit distinct clinical characteristics and therapeutic responses, indicating the potential of integrating these biomarkers to enhance personalised treatment strategies (6, 7). Understanding PCa heterogeneity has advanced significantly with the development of molecular subtyping based on ERG and PTEN expression (6, 7). These subtypes not only increase prognosis accuracy but also facilitate tailored therapies, like medicines that target ERG-mediated pathways or PI3K/AKT inhibitors for tumours losing PTEN (6, 7). Even though the roles of ERG and PTEN in PCa progression have been the focus of several studies, a thorough synthesis of current studies is required to completely clarify its clinical consequences (4, 6, 8).
This review aims to clarify the roles of ERG and PTEN molecular subtypes in PCa development and progression emphasising their potential as biomarkers for prognosis, detection, and targeted therapy. By integrating existing evidence, we seek to provide a groundwork for advancing personalised treatment approaches in PCa.
Peer-reviewed articles from 1999 to 2024 were sourced from PubMed and Google Scholar. During the review process, the following search terms were employed: “Tumour suppressor genes OR variations OR alterations OR oncogenes OR diagnostics OR ethnicity OR biomarkers OR prostate cancer genomics OR prostate cancer structural variations OR tumour and molecular subtypes OR therapeutic implications OR immunotherapy OR immunogenetics.”
There were 13,030 total results for our search query: 5,903 publications from Google Scholar with the patent and citation unchecked filer options and 7,127 articles from PubMed with the abstract, free full text, and full-text options selected.
The filtering method also yielded irrelevant results due to the overly broad key phrases in the search string. The following criteria for inclusion were applied during the manual screening of the obtained articles: 1) Articles discussing the phenotypic or genetic disorders associated with PCa, 2) publications detailing genetic changes or variants linked to PCa, 3) articles outlining changes in tumour-suppressor genes and oncogenes related to PCa, 4) articles exploring how genes associated with PCa may influence prognosis and diagnosis, 5) articles on the clinical applications of tumour subtyping and PCa risk classification, and 6) tumour-suppressor genes and oncogenes in the context of PCa treatments. Unpublished works were excluded. Except for four articles published between 1998 and 1999, all other selected articles published in 2000 and later were included. However, papers containing irrelevant information or redundant or duplicate content were not considered for this review. Thus, 134 articles were ultimately retained for this review.
The initiation, development, progression, and treatment resistance of PCa are driven by a complex interplay of genetic and molecular alterations reflecting the disease’s inherent heterogeneity (9, 10). Among the most well-reported and clinically relevant molecular alterations in PCa are the loss of PTEN and the TMPRSS2:ERG gene fusion. The TMPRSS2:ERG fusion leads to the overexpression of the ERG transcription factor, a key driver of tumour invasion and progression (9, 10). Similarly, PTEN, a tumour-suppressor gene that regulates the PI3K/AKT signalling pathway, is commonly lost or inactivated, particularly in advanced and metastatic tumours. The co-occurrence of these alterations is strongly associated with resistance to conventional therapies, aggressive tumour behaviour, and poor clinical outcomes. Advances in understanding the biology and clinical implications of ERG and PTEN alterations have illuminated the pathophysiology of PCa and paved the way for the development of targeted and personalised therapeutic strategies. These insights are crucial for improving patient outcomes and addressing the challenges posed by the disease’s heterogeneous nature.
Structurally, PTEN is located at the 10q23 locus on chromosome 10, with nine exons and eight introns. It is roughly 200-kb long and codes for a 403-amino acid multifunctional protein that has lipid phosphatase activity (9, 10). PTEN dephosphorylates phosphoinositide substrates to create a dual-specific protein phosphatase that is essential for controlling the PI3K/AKT signalling pathway (9, 10). This regulation affects essential cellular functions such as apoptosis, cell cycle regulation, cell invasion inhibition, and general tumour suppression (9, 10). Approximately 50% of castration-resistant prostate tumours frequently exhibit PTEN mutations, deletions, and inactivation, which contribute to dysregulated PI3K/AKT signalling (7). As a result, the onset, spread, and poor clinical outcomes of PCa are all strongly associated with the loss of PTEN function (9, 10).
PTEN, the most frequently altered tumour-suppressor gene (TSG) in PCa, plays a significant role in maintaining genomic stability through its nuclear activities (11). While PTEN is known for its ability to suppress tumours by preventing the oncogenic PI3K signalling pathway, the available findings suggest that its tumour-suppressive functions extend beyond its lipid phosphatase activity (12). Previous studies hypothesised that PTEN functions as a lipid phosphatase in the nucleus due to the presence of PI3K, AKT, and pyruvate dehydrogenase kinase 1 enzyme (PDK1) in this compartment (13). In addition, subsequent research has demonstrated that PTEN’s nuclear functions are not primarily regulated by its lipid phosphatase activity, as its main substrate, PIP3, is not significantly present in the nucleus (14). Given that several anti-PI3K medications are unable to completely suppress tumour growth in PTEN-deficient malignancies, available data suggest that PTEN has tumour-suppressive effects that extend beyond PI3K pathway inhibition (15, 16). Specifically, nuclear PTEN localises to heterochromatin, contributing to structural stability and reinforcing its phosphatase-independent tumour-suppressive roles (17). These findings highlight the multifaceted nature of PTEN’s functions and underscore its importance in both cytoplasmic and nuclear contexts.
Due to its dual-specificity phosphatase activity, PTEN operates in the cytoplasm to dephosphorylate phosphoinositol (3,4,5)-trisphosphate (PIP3), thereby preventing AKT activation and directly suppressing PI3K signalling (9). The PI3K pathway is typically activated when growth factors bind to receptor tyrosine kinases triggering the conversion of PIP2 to PIP3. This process promotes AKT activation through phosphorylation mediated by PDK1 (18). Once activated, AKT modulates several downstream targets, including mTOR, which enhance cell growth, proliferation, and survival (19).
This regulatory mechanism is disrupted when PTEN is lost, which results in uncontrolled PI3K/AKT signalling. In PCa, this dysregulation contributes to resistance to androgen deprivation therapy by promoting androgen-independent activation of the androgen receptor (AR) pathway. Furthermore, PTEN inactivation leads to a reduced control over vital cellular functions that are essential to the growth of tumours, such as energy metabolism, cell survival, proliferation, and structural integrity (18). In addition, as seen in Figure 1, PTEN is essential for the development and spread of cancer. These findings underscore the multifaceted contributions of PTEN to tumour biology and highlight its significance as a key regulator of cellular homeostasis and cancer progression.
The primary mechanism of PTEN loss in PCa is copy number variations, which differentiate PCa from other types of malignancies (20). Loss of PTEN function drives metabolic reprogramming influencing aggressive tumour growth and rapid cell proliferation (9, 21). This dysfunction affects chromatin structure leading to the loss of heterochromatic foci, reduced chromatin compaction, amplification of heterochromatic genes, disruption of heterochromatin protein 1, and, eventually, genomic instability and loss of PTEN function (22). Loss of PTEN function activates the PI3K–AKT signalling pathway, which is strongly connected with poor PCa outcomes, as illustrated in Figure 2. Available evidence suggests that PTEN could function as a genetic marker to differentiate aggressive PCa from indolent PCa, especially in clinically localised PCa (9, 23). Furthermore, PTEN facilitates tumour formation by modulating the tumour microenvironment (TME) and immune responses (9). The available research findings have shown that loss of PTEN function in PCa correlates with higher Gleason scores and advanced tumour stages. However, inconsistencies in reported findings, which are linked to methodological variations, participant selection, and population variability, underscore the need for further research (9, 24). Ethnic differences in PTEN loss have also been reported, with African-American men demonstrating lower rates of PTEN loss compared to European-American men (25, 26). Despite these insights, the relationship between racial background, PTEN loss, and poor prognosis is not well known and warrants further research (26).
PTEN loss is closely linked to the development of an immunosuppressive TME characterised by increased cytokine and chemokine signalling. This environment is enhanced with immunosuppressive cells, such as myeloid-derived suppressor cells (MDSCs), regulatory T cells (Tregs), and M2-polarised macrophages, which prevent antitumour immune responses (27). Additionally, PTEN-deficient tumours show increased expression of immune evasion markers, including programmed death-ligand 1 (PD-L1) and indoleamine 2,3-dioxygenase 1 (IDO1), which damage the cytotoxic activity of immune cells and facilitate immune escape (18). The loss of PTEN’s nuclear functions further exacerbates inflammation and cytokine signalling, contributing to an immunosuppressive TME (27). This milieu, dominated by M2 macrophages, MDSCs, and regulatory immune cells, suppresses antitumour immunity. Moreover, PTEN loss is associated with heightened genomic instability, which can generate neoantigens capable of activating CD8+ T cells and triggering an immune response (18). However, infiltrating tumour-associated macrophages may decrease the immune system’s reaction to counteract neoantigen-driven immunity in cancers with high levels of genomic instability (18).
The androgen-regulated gene TMPRSS2 is found on chromosome band 21q22 and is mainly expressed in the prostate (3). The ERG gene, also mapped to 21q22, is located approximately 3 Mb downstream of TMPRSS2. The TMPRSS2:ERG gene fusion, the most prevalent structural variation in PCa, has been reported in nearly half of all PCa cases (28, 29). This fusion is common in both early and advanced stages of the disease underlining its importance in PCa pathogenesis (5, 30, 31). While it is not frequently reported in normal prostate tissue, TMPRSS2:ERG fusions facilitate tumour progression by promoting angiogenesis, inflammation, and epithelial–mesenchymal transition ultimately leading to metastasis, advanced tumour stages, and increased mortality (32–34). Other ETS family members, such as ETV1, ETV4, and ETV5, as well as androgen receptor (AR) targets, like SLC45A3 and NDRG, are also implicated in gene fusions in PCa.
The TMPRSS2:ERG fusion occurs when androgen and prostate-specific regulatory regions of TMPRSS2 are juxtaposed with the first exon(s) of ERG leading to androgen-driven overexpression of the fusion transcript. In nearly 50%–60% of fusion-positive tumours, this results from an intronic deletion on chromosome 21, which deletes the genomic region between TMPRSS2 and ERG. Alternatively, complicated genomic rearrangements involving chromosome 21q22 and other chromosomes may also give this fusion (35–37). The recurring nature of this fusion is associated with a common deletion spot connecting ERG and TMPRSS2 (38). The deletion site is characterised by two types of breakpoints, with the 3′ end of ERG consistently fused to the 5′ end of TMPRSS2. This fusion leads to the overexpression of ERG, facilitated by the androgen-responsive promoter of TMPRSS2, resulting in elevated PCa cell invasion, proliferation, angiogenesis, and tumour aggressiveness (39, 40). Moreover, the TMPRSS2:ERG fusion triggers downstream oncogenes, further amplifying its carcinogenic effects (41).
Similarly, androgen receptor activation provides a key role in this process by enhancing TMPRSS2 promoter function leading to the production of an ERG mRNA fusion transcript. This transcript is translated into the ERG transcription factor, which consequently activates downstream oncogenic signalling pathways (42, 43). This molecular cascade provides two key therapeutic targets. First, androgen receptor inhibitors (e.g., enzalutamide and abiraterone) suppress TMPRSS2 activation reducing fusion transcript production and ERG overexpression (44). Second, bromodomain inhibitors (e.g., JQ1) disrupt the transcriptional machinery required for ERG-mediated gene activation, thereby preventing its oncogenic effects (42, 45). These targeted approaches, which interrupt critical nodes in the TMPRSS2:ERG fusion pathway, provide potential opportunities for personalised therapy in PCa (46, 47). The mechanisms regulating ERG overexpression in PCa cells are illustrated in Figure 2.
The transition from prostatic intraepithelial neoplasia (PIN) to PCa is characterised by higher Gleason scores, metastasis, advanced tumour stages, and reduced survival rates. This progression is facilitated by the persistent overexpression of the ERG oncogene (28, 48). In contrast to other ETS family members, such as SAM-pointed domain-containing ETS transcription factor (SPDEF) and ETS2 repressor factor (ERF), which are important for maintaining normal prostate epithelium, ERG is only moderately expressed in normal prostate cells (49), though, when overexpressed in prostate cells, ERG drives a range of oncogenic effects. For instance, ERG interacts with the PI3K oncogenic signalling pathway leading to tumourigenesis (48). Additionally, ERG enhances androgen receptor (AR) binding and enhances AR transcription, especially in PCa patients with loss of PTEN function (50). Notably, AR binding patterns are knowingly altered in cells with increased ERG expression (50, 51). Furthermore, ERG triggers the Wnt signalling pathway, elevating β-catenin activation and facilitating PCa development and progression (52–54). Despite these oncogenic functions, available findings suggest that ERG overexpression alone is insufficient to initiate cancer and is not reliably associated with disease progression (6, 48, 50, 55–57). Ethnic and population-based variations in ERG rearrangements and increased expression have also been reported (Table 1). In American populations, the overall frequency of ERG expression ranges between 50% and 55%, with rates of 28% observed among Caucasian Americans. Among African American populations, ERG expression rates vary considerably depending on the studied population. Findings from Sub-Saharan Africa remain inadequate. In Asian populations, ERG rearrangements appear in 50%–55% of cases, with rates of 28% in Indian patients and 49% in Chinese PCa patients (34). These differences highlight the impact of genetic and environmental factors on ERG-driven oncogenesis.
Significantly, ERG overexpression may function as an indicator of disease aggressiveness, and its interaction with other regulatory pathways, such as loss of PTEN, further highlights its role in PCa progression (75, 76). PCa individuals exhibiting ERG-positive high-grade prostatic intraepithelial neoplasia (HGPIN) are at a considerably elevated risk of developing PCa emphasising the clinical relevance of ERG as a potential biomarker for disease stratification and risk assessment (77).
The interaction between ERG and the androgen receptor (AR) plays an essential role in the aetiology and progression of PCa. This interaction is predominantly significant in the context of the TMPRSS2:ERG gene fusion, a common genetic alteration found in PCa (5). Available data suggest that ERG and AR collaborate to drive PCa development, with ERG modulating the AR transcriptional program and facilitating the expression of AR target genes that enhance tumour growth and survival (5). Moreover, ERG may indirectly influence AR signalling pathway by altering chromatin structure and accessibility, thereby promoting AR binding to DNA and transcriptional activation. In the lack of androgens, AR remains inactive in the cytoplasm, bound to the chaperone protein HSP90. Androgen binding triggers a conformational change in AR initiating it to dissociate from HSP90 and translocate into the nucleus (78). Once in the nucleus, AR binds to androgen response elements (AREs) in the promoter or enhancer regions of target genes to regulate their transcription. In normal prostate cells, this process mechanism upregulates genes, which are essential for prostate function (79). However, in PCa cells with the TMPRSS2:ERG fusion, androgen-bound AR aberrantly triggers the ERG oncogene leading to tumourigenesis (80). This dual role of androgen signalling pathway controlling both normal cellular functions and oncogenic signalling pathways underlines potential therapeutic targets (5). For instance, androgen deprivation therapies (ADTs), which reduce androgen levels or prevent AR role, have been reported to reduce ERG expression in TMPRSS2:ERG fusion-positive tumours, thus reducing their oncogenic potential (81). The association between ERG and AR also has significant effects for combination therapies in PCa. Combining AR-targeted therapies with agents that selectively disrupt ERG function or its downstream pathways provides potential synergistic options for treating TMPRSS2:ERG fusion-positive PCa (81). However, directly targeting ERG raises challenges due to its nature as a transcription factor. Despite this, ongoing research remains to explore strategies to prevent ERG–DNA binding or interrupt ERG–AR interactions (81). The mechanisms underlying androgen regulation of gene expression in PCa cells are illustrated in Figure 3.
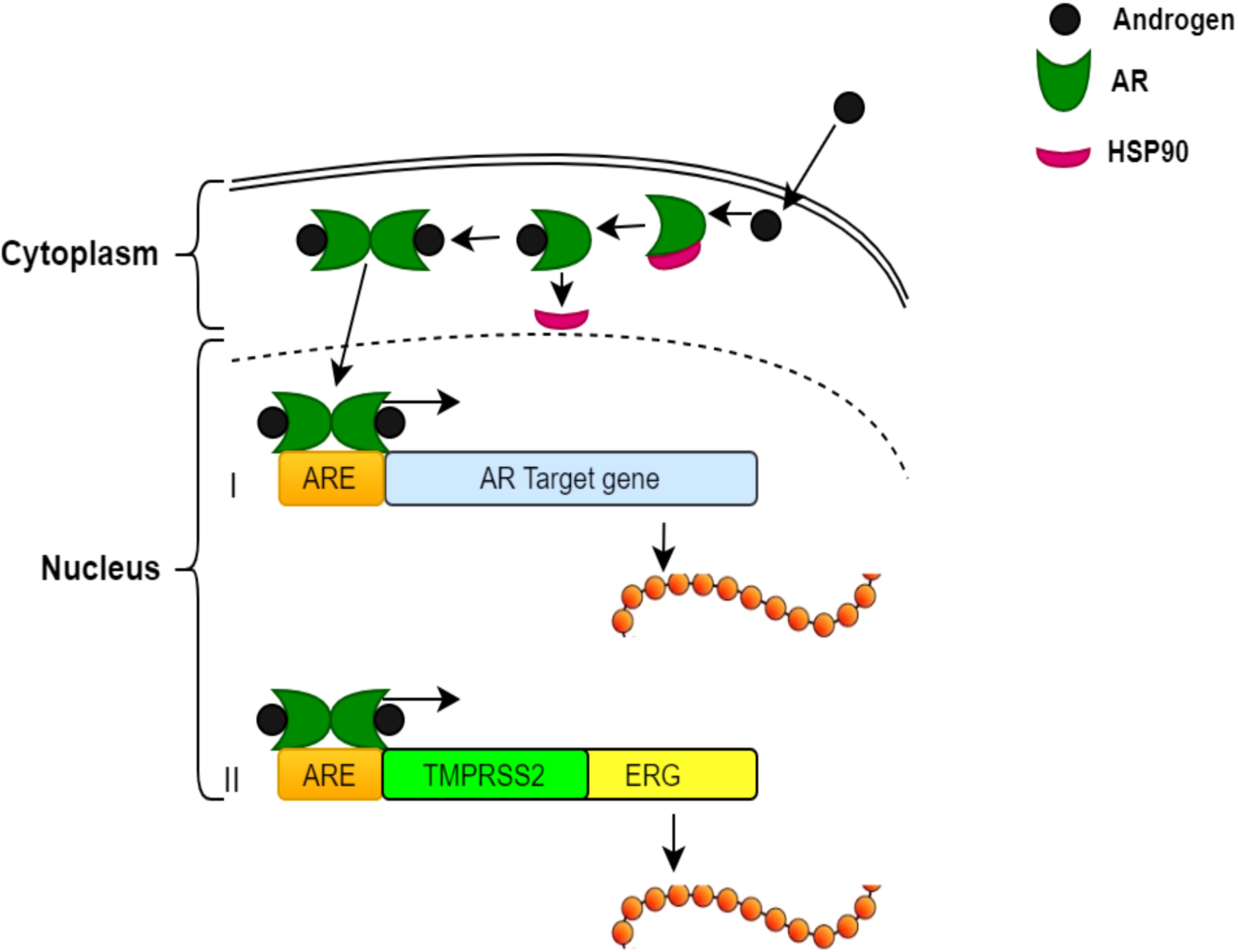
Figure 3. Mechanisms of androgen regulation of gene expression in PCa cells. Inactive AR in the cytoplasm bound to HSP90. The binding of androgens induces a conformational change and activates AR, releasing Hsp90. Activated AR moves to the nucleus. AR binds to androgen response elements (AREs) in the target genes’ promoter/enhancer regions. (I) Example of a typical androgen-regulated gene, the expression of which is induced when AR binds to the ARE at the promoter region. (II) TMPRSS2:ERG fusion—androgen binding to the ARE regulates the ERG oncogene.
Recent studies suggest that prostatic intraepithelial neoplasia (PIN) can advance to invasive cancer when ERG activation coincides with loss of PTEN function (82). While some studies have proposed a link between ERG expression and Gleason score, the current evidence remains inconclusive underscoring the need for further investigation. In contrast, there is stronger evidence associating reduced PTEN expression with higher Gleason scores and poor prognosis (74). Although the relationship between ERG overexpression and loss of PTEN function has not been extensively explored, studies investigating the therapeutic potential of targeting ERG and PTEN have yielded promising results (74). Furthermore, PTEN loss has been consistently associated with adverse clinical outcomes, including poor overall survival, unfavourable pathological tumour behaviour, and the development of castration-resistant and metastatic PCa (69, 83). These findings suggest that PTEN loss may represent a more clinically significant chromosomal alteration in PCa compared to TMPRSS2:ERG fusion emphasising its importance as a potential biomarker and therapeutic target.
Interleukin-6 (IL-6) cytokine signalling, PTEN loss, and activation of the PI3K–AKT pathway are among key contributors to the pathophysiology of PCa. Their complex relationship promotes tumour growth, boosts cell survival, and facilitates therapeutic resistance, underlining the significance of integrating molecular approaches in developing targeted therapies and improving prognostic assessment (84, 85). Chronic inflammation is closely linked to the genetic alterations and mechanisms facilitating PCa progression (86, 87). Among the key cytokine’s mediators of inflammation, IL-6 plays a key role in PCa pathogenesis by facilitating angiogenesis, tumour development, and disease progression. Pro-inflammatory cytokine, IL-6, is the most frequently associated inflammatory mediator in PCa (88). Beyond its inflammatory role, elevated IL-6 expression is strongly associated with poor clinical outcomes, treatment resistance, and advanced disease stages. The oncogenic effects of IL-6 are facilitated through IL-6 receptor (IL-6R) signalling, which promotes the PI3K–AKT pathway, thus stimulating PCa cell growth and survival rates (84, 85). This process introduces a positive feedback loop, where PTEN loss and consequent hyperactivation of the PI3K–AKT signalling pathway further upregulate IL-6 expression amplifying its oncogenic effects.
The therapeutic potential of targeting IL-6-mediated pathways is significant. Anti-IL-6 treatments, like monoclonal antibodies against IL-6 or IL-6R, may interfere with IL-6 signalling and slow the growth of PCa (84, 85). Additionally, the potential of combination therapy is highlighted by the dynamic interaction between PI3K–AKT signalling, PTEN loss, and IL-6. Strategies that target PTEN loss block the PI3K–AKT pathway and reduce IL-6 signalling together have the potential to improve PCa treatment outcomes. These combined approaches present an achievable means to improve patient outcomes and overcome treatment resistance (84, 85).
Prostate cancer (PCa) can be classified into four distinct molecular subtypes based on IHC staining analysis of ERG and PTEN expression. These molecular subtypes include the following: (1) normal ERG with loss of PTEN expression (ERG−/PTEN−), (2) rearranged ERG with normal PTEN expression (ERG+/PTEN+), (3) normal ERG with normal PTEN expression (ERG−/PTEN+), and (4) rearranged ERG with loss of PTEN expression (ERG+/PTEN−) (58, 89, 90). These tumour molecular subtypes have surfaced as valuable tools for patient stratification allowing personalised approaches to treatment based on individual molecular subtypes. The prevalence of these molecular subtypes varies across different populations, as summarised in Table 2. Additionally, these molecular subtypes present distinct therapeutic potentials, particularly in the setting of immunotherapy and personalised treatment, providing opportunities for more precise and efficient treatment strategies (92, 93).
Prostate cancer characterised by PTEN loss and absence of ERG expression (ERG−/PTEN−) signifies a distinct molecular subtype with profound biological and therapeutic implications (74). This subtype exhibits reduced dependence on androgen receptor (AR) signalling compared to ERG-positive tumours, contributing to resistance to AR-targeted therapies such as androgen deprivation therapy (ADT) (94). ERG−/PTEN− tumours are often more aggressive, driven by their reliance on PI3K/AKT signalling and heightened genomic instability, and are associated with poor prognosis (69, 95, 96). Notably, this subtype appears to be more prevalent in certain populations, such as African American men, who exhibit higher rates of PTEN loss and lower frequencies of TMPRSS2:ERG fusion highlighting potential ethnic and genetic variations in PCa subtypes (97). While ERG−/PTEN− tumours pose therapeutic challenges due to their reduced sensitivity to conventional AR-targeted therapies, they also present opportunities for novel treatment strategies (98). To address cross-pathway interactions, inhibitors that target the PI3K/AKT/mTOR pathway have shown potential, especially when combined with AR inhibitors (99). Additionally, immunotherapy, including immune checkpoint inhibitors, is a promising therapeutic approach for this subtype, as PTEN loss has been connected to immune evasion. The findings highlight how crucial it is to use both pathway-specific and immune-modulating treatment methods to treat ERG−/PTEN− tumours to enhance patient outcomes.
The ERG+/PTEN+ tumour molecular subtype of PCa is characterised by ERG rearrangements, most commonly the TMPRSS2:ERG fusion, and intact PTEN function (58). However, the occurrence of this molecular subtype varies due to the heterogeneousness of PTEN alterations, with PTEN loss reported in 20%–50% of cases. ERG rearrangements are found in nearly 40%–70% of prostate tumours (59, 60). Biologically, ERG rearrangements facilitate oncogenesis by promoting androgen receptor (AR)-regulated transcriptional pathways, which enhance tumour invasion and progression (100). The intact PTEN function in this molecular subtype helps regulate the PI3K/AKT signalling pathway differentiating it from the more aggressive ERG+/PTEN− molecular subtype (101, 102).
Clinically, ERG+/PTEN+ tumours are known to be sensitive to AR-targeted therapies, such as androgen deprivation therapy (ADT), abiraterone, and enzalutamide, which form the keystone of treatment for this tumour molecular subtype (103). Emerging therapeutic strategies, such as precision medicine approaches that target vulnerabilities in ERG driven pathways and DNA repair mechanisms, such as PARP inhibitors, may benefit a subset of these tumours, particularly those with additional mutations in DNA damage repair genes (104). Immune checkpoint inhibitors also represent a potential therapeutic avenue for ERG+/PTEN+ tumours, as this subtype may maintain a more immunologically active tumour microenvironment compared to PTEN-deficient subtypes, though the efficacy of these treatments remains under research (105, 106). Additionally, novel therapies targeting ERG-associated signalling pathways are being explored offering promising avenues for future treatment development (107). Despite these developments, several issues still need to be addressed, including the biology and clinical variation of this subtype in other populations and the need for biomarkers to direct tailored treatment. An improved understanding of the particular vulnerabilities of ERG+/PTEN+ tumours is necessary to improve treatment approaches and patient outcomes for patients with this genetic subtype of PCa (59, 60, 107).
The ERG−/PTEN+ tumour molecular subtype of PCa is described by the absence of ERG rearrangements, such as TMPRSS2:ERG fusions, and the retention of PTEN function, which is important for controlling the PI3K/AKT signalling pathway (58). This molecular subtype falls within the wide category of ERG-negative prostate tumours accounting for 30%–60% of PCa cases (58, 71). The occurrence of the ERG−/PTEN+ subtype varies throughout populations, with studies suggesting population-specific differences in disease biology. For instance, African American men demonstrate a higher frequency of PTEN-positive tumours and a reduced occurrence of ERG expression, stressing potential ethnic and genetic variations in PCa (97, 108).
The retention of PTEN function in ERG−/PTEN+ tumours distinguish them from more aggressive PTEN-deficient subtypes, as PTEN supports the regulation of cellular proliferation and survival (58, 61). Therapeutically, these molecular tumours continue to depend on androgen receptor (AR) signalling making them responsive to androgen deprivation therapy (ADT) and AR-targeted agents such as enzalutamide and abiraterone. Furthermore, the absence of PTEN loss may contribute to a more immunologically active tumour microenvironment fostering the possibility of exploring immunotherapies, including immune checkpoint inhibitors, potentially in combination with AR-targeted therapies (62, 103).
The lack of ERG expression in this molecular subtype indicates a dependency on alternative oncogenic signalling pathways emphasising the need for further research into precision medicine approaches and innovative therapeutic targets. Understanding the distinctive molecular and immunological characteristics of ERG−/PTEN+ tumour molecular subtypes is important for developing tailored treatment strategies and improving outcomes for patients with this subtype.
The presence of ERG rearrangements, most frequently TMPRSS2:ERG fusions, and the loss of PTEN function are characteristics of the ERG+/PTEN− molecular subtype of PCa (58, 109). With more invasive tumours, higher Gleason scores, and an increased likelihood to develop castration-resistant PCa, this subtype is associated with aggressive disease (109, 110). ERG alterations increase transcriptional activity mediated by the androgen receptor (AR), whereas PTEN loss interferes with the PI3K/AKT signalling pathway leading to uncontrolled cell proliferation and resistance to apoptosis (111). ERG rearrangements characterised by ERG expression, are more common in Caucasian men compared to African American and Asian men, and the occurrence of ERG+/PTEN− tumours varies by ethnic group. However, all ethnic groups exhibit PTEN loss, especially in advanced stages of the disease (60, 65). Therapeutically, AR-targeted treatments, such as abiraterone, enzalutamide, and androgen deprivation therapy (ADT), are efficient against ERG-driven tumours. However, PTEN loss often confers resistance by triggering alternative signalling pathways (62, 103). To address this, inhibitors targeting the PI3K/AKT/mTOR pathway are being researched, with promising results when used in combination with AR-targeted therapies (103, 112, 113). Additionally, ERG+/PTEN− tumours commonly exhibit immunosuppressive characteristics, such as reduced CD8+ T-cell infiltration and enhanced recruitment of myeloid-derived suppressor cells (18, 95). These characteristics reduce the efficacy of immune checkpoint inhibitors (ICIs), such as anti-PD-1/PD-L1 or anti-CTLA-4 antibodies (18), though combination therapies that simultaneously target the PI3K/AKT pathway and boost immune responses may improve outcomes in certain subtypes (18, 95).
Significant research gaps still exist despite these developments, especially in the areas of clarifying resistance mechanisms, discovering predictive biomarkers, and understanding population-specific variances. To create more individualised and successful treatment plans for patients with ERG+/PTEN− PCa, several issues must be resolved.
Due to the TMPRSS2:ERG fusion, the ERG gene is more expressed in PCa in both its early and late stages (5, 114, 115). It has been suggested that this fusion event is a diagnostic biomarker for PCa and is a useful tool for distinguishing tumour molecular subtypes (116). Research indicates that over 50% of PCa cases have the TMPRSS2:ERG fusion and consequent ERG overexpression (115, 117). The identification of ERG protein overexpression through immunostaining in PCa samples or the presence of TMPRSS2:ERG in prostate tissue can both be used as reliable diagnostic markers. However, the absence of ERG expression is not sufficient evidence that PCa is not present. TMPRSS2:ERG fusion may also be a potential urine-based biomarker for PCa detection, according to recent data (118, 119). The prognostic significance of the TMPRSS2:ERG fusion remains controversial, with studies reporting contradictory findings (28). While some research challenges its predictive utility, others suggest that it can function as a prognostic biomarker, with increased TMPRSS2:ERG frequency connected to worse clinical outcomes (6, 8, 120). These discrepancies may stem from variations in patient demographics, methodologies used to detect gene fusions, and the therapeutic effects on the tumour samples analysed. As a result, cautiousness is necessary when interpreting studies that emphasise the prognostic utility of the TMPRSS2:ERG fusion. Despite these challenges, the TMPRSS2:ERG fusion remains a potential therapeutic target due to its specificity to PCa and its overexpression throughout various stages of tumour progression (121). As the most common genetic alteration in PCa, targeting TMPRSS2:ERG at the molecular level has gained significant interest as a potential treatment strategy. Additionally, ongoing research continues to explore the TMPRSS2:ERG fusion gene for its potential as a different biomarker, therapeutic target, and diagnostic and prognostic indicator in PCa (121).
Evaluating the prognosis for individuals with PCa remains a key issue in disease management. Various potential molecular indicators and biomarkers are being introduced, with different institutions developing their standards for PCa risk assessment (122). Generally, blood PSA levels and Gleason scores are utilised to determine whether patients require active monitoring and treatment decisions (122). However, these markers are associated with several limitations when classifying PCa patients for adequate disease management. The Gleason grade is regarded as the most reliable predictive biopsy metric having been enhanced through modifications to the Gleason grading system (123, 124), though the amount of data obtainable through this approach is also limited, necessitating the development of cost-effective and straightforward predictive biomarkers to identify potentially aggressive prostate tumours and assist in categorising PCa patients into various predictive groups for treatment options (125, 126). According to previously published research, PTEN depletion has been linked to PCa progression through various methods such as IHC and FISH (125). PTEN loss or deficiency plays a significant role in PCa development, as evidenced by numerous studies. Several publications have also shown a direct correlation between PTEN loss and an increased risk of biochemical recurrence following prostatectomy proving useful for categorising patients with PCa into different prognosis groups for targeted treatments (68, 69, 125, 127).
Prostate cancer encounters significant challenges in patient classification and management selection due to its varied tumour molecular subtypes, variable tumour aggressiveness, and heterogeneous therapeutic responses (2). While tumour molecular subtyping presents a potential approach for stratifying patients and guiding personalised therapy (128), the intricacy of the disease complicates the accurate assessment of critical genetic alterations, such as ERG fusions and PTEN variants. Addressing these challenges requires the integration of multiple molecular biomarkers and comprehensive genomic profiling to better understand the underlying biology of PCa (129). Additionally, the varying susceptibility and resistance of diverse molecular subtypes to targeted therapies further complicate the clinical application of precision medicine. For example, tumours with ERG rearrangements or loss of PTEN may demonstrate different therapeutic sensitivities and resistance mechanisms necessitating tailored treatment strategies (129).
In line with this, there is great potential for enhancing risk classification, directing treatment choices, and developing tailored therapeutics for PCa by incorporating TMPRSS2:ERG fusion status and PTEN expression into clinical practice (130, 131). The goal of future research should be to clarify the molecular processes underlying PTEN and ERG dysregulation to aid in the development of tailored therapy strategies (130, 131). We anticipate that improvements in drug discovery and molecular profiling will strengthen our capacity for predicting clinical results using PTEN and ERG immunostaining and optimise therapeutic decisions. Precise detection of TMPRSS2:ERG fusions and loss of PTEN, particularly through reliable and cost-effective methods, such as immunohistochemistry (IHC), will be necessary for comprehensive genomic profiling (130, 131). Additionally, the development of advanced diagnostic and analytic tools to assess gene expression patterns will enable clinicians to select and monitor precise therapeutic options, eventually improving long-term patient outcomes (130, 131). These advancements underline the importance of incorporating molecular biomarkers into routine clinical practice to refine precision medicine approaches in PCa.
In conclusion, the application of PTEN and TMPRSS2:ERG fusion in PCa holds considerable promise for enhancing targeted treatment strategies and improving patient outcomes. The expression patterns of PTEN and ERG provide valuable insights into tumour characteristics, patient prognosis, and treatment response promising more tailored and effective therapeutic approaches. Loss-of-function affecting the phosphatase domain of PTEN, often associated with aggressive tumour phenotypes and poor prognosis, underscores the potential for targeted therapeutics aimed at restoring PTEN function or mitigating its downstream effects. Similarly, the prevalence of ERG overexpression in a substantial proportion of PCa presents opportunities for the development of ERG-targeted therapies and diagnostic tools. The interpretation and integration of tumour molecular subtyping analysis into clinical practice require further validation through robust clinical studies and the establishment of reliable, cost-effective testing methodologies. Additionally, the exploration of integrative techniques targeting multiple pathways affected by PTEN and ERG alterations holds promise for overcoming resistance mechanisms and enhancing treatment efficacy.
FB: Conceptualization, Data curation, Formal analysis, Methodology, Resources, Writing – original draft, Writing – review & editing. RS: Methodology, Project administration, Supervision, Validation, Writing – review & editing. RB: Conceptualization, Writing – review & editing. ND: Conceptualization, Writing – review & editing. FM: Resources, Supervision, Writing – review & editing. HK: Conceptualization, Writing – review & editing. SA: Conceptualization, Project administration, Supervision, Writing – review & editing. KP: Conceptualization, Supervision, Writing – review & editing. SM: Conceptualization, Resources, Supervision, Writing – review & editing.
The author(s) declare that financial support was received for the research and/or publication of this article. The preparation of this manuscript was facilitated by funding provided by the Government Chemist Laboratory Authority (GCLA), Tanzania, and Moi University, Kenya, through the P4HPTII project of the Intra-Africa Academic Mobility Scholarship scheme. The funders were not involved in the study design, data collection, analysis, interpretation, writing of this article, or the decision to submit it for publication.
We extend our gratitude to the staff members of the Muhimbili National Hospital in Tanzania, the Government Chemist Laboratory Authority, the National Institute for Medical Research, the Department of Pathology and Immunology Section at Moi University—Kenya, and the National Institute for Medical Research for their invaluable contributions and assistance with reviewing.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
DNA, deoxyribonucleic acid; TSGs, tumour-suppressor genes; PCa, prostate cancer.
1. Bugoye FC, Leyna GH, Moen K, Mmbaga EJ. Knowledge, perceived risk and utilization of prostate cancer screening services among men in Dar Es Salaam, Tanzania. Prostate Cancer. (2019) 2019. doi: 10.1155/2019/2463048
2. Bugoye FC, Torrorey-Sawe R, Biegon R, Dharsee N, Mafumiko FMS, Patel K, et al. Mutational spectrum of DNA damage and mismatch repair genes in prostate cancer. Front Genet. (2023) 14:1231536. doi: 10.3389/fgene.2023.1231536
3. Hermans KG, Boormans JL, Gasi D, Leenders GJv, Jenster G, Verhagen PC, et al. Overexpression of prostate-specific TMPRSS2(exon 0)–ERG fusion transcripts corresponds with favorable prognosis of prostate cancer. Clin Cancer Res. (2009) 15:6398–403. doi: 10.1158/1078-0432.CCR-09-1176
4. Kish EK, Choudhry M, Gamallat Y, Buharideen SM, Dhananjaya D, Bismar TA, et al. The expression of proto-oncogene ETS-related gene (ERG) plays a central role in the oncogenic mechanism involved in the development and progression of prostate cancer. Int J Mol Sci. (2022) 23. doi: 10.3390/ijms23094772
5. Tomlins SA, Rhodes DR, Perner S, Dhanasekaran SM, Mehra R, Sun X-W, et al. Recurrent fusion of TMPRSS2 and ETS transcription factor genes in prostate cancer. Sci (New York N.Y.). (2005) 310:644–8. doi: 10.1126/science.1117679
6. Pettersson A, Graff RE, Bauer SR, Pitt MJ, Lis RT, Stack EC, et al. The TMPRSS2:ERG rearrangement, ERG expression, and prostate cancer outcomes: A cohort study and meta-analysis. Cancer Epidemiology Biomarkers Prev. (2012) 21:1497–509. doi: 10.1158/1055-9965.EPI-12-0042
7. Wang Q, Wang J, Xiang H, Ding P, Wu T, Ji G. The biochemical and clinical implications of phosphatase and tensin homolog deleted on chromosome ten in different cancers. Am J Cancer Res. (2021) 11:5833–55.
8. Saramaki OR, Harjula AE, Martikainen PM, Vessella RL, Tammela TL, Visakorpi T. TMPRSS2–ERG fusion identified a subgroup of prostate cancers with a favorable prognosis. Clin Cancer Res. (2008) 14:3395–4000. doi: 10.1158/1078-0432.CCR-07-2051
9. Jamaspishvili T, Berman DM, Ross AE, Scher HI, De Marzo AM, Squire JA, et al. Clinical implications of PTEN loss in prostate cancer. Nat Rev Urol. (2018) 15:222–34. doi: 10.1038/nrurol.2018.9
10. Naderali E, Valipour B, Khaki AA, Soleymani Rad J, Alihemmati A, Rahmati M, et al. Positive effects of PI3K/akt signaling inhibition on PTEN and P53 in prevention of acute lymphoblastic leukemia tumor cells. Advanced Pharm Bull. (2019) 9:470–80. doi: 10.15171/apb.2019.056
11. Ho J, Cruise ES, Dowling RJO, Stambolic V. PTEN nuclear functions. Cold Spring Harbor Perspect Med. (2020) 10. doi: 10.1101/cshperspect.a036079
12. Maehama T, Dixon JE. PTEN: a tumour suppressor that functions as a phospholipid phosphatase. Trends Cell Biol. (1999) 9:125–8. doi: 10.1016/s0962-8924(99)01519-6
13. Déléris P, Gayral S, Breton-Douillon M. Nuclear Ptdlns(3,4,5)P3 signaling: an ongoing story. J Cell Biochem. (2006) 98:469–85. doi: 10.1002/jcb.20695
14. Lindsay Y, McCoull D, Davidson L, Leslie NR, Fairservice A, Gray A, et al. Localization of agonist-sensitive PtdIns(3,4,5)P3 reveals a nuclear pool that is insensitive to PTEN expression. J Cell Sci. (2006) 119:5160–8. doi: 10.1242/jcs.000133
15. Juric D, Castel P, Griffith M, Griffith OL, Won HH, Ellis H, et al. Convergent loss of PTEN leads to clinical resistance to a PI(3)Kα inhibitor. Nature. (2015) 518:240–4. doi: 10.1038/nature13948
16. Le X, Antony R, Razavi P, Treacy DJ, Luo F, Ghandi M, et al. Systematic functional characterization of resistance to PI3K inhibition in breast cancer. Cancer Discovery. (2016) 6:1134–47. doi: 10.1158/2159-8290.CD-16-0305
17. Amor DJ, Kalitsis P, Sumer H, Choo KHA. Building the centromere: from foundation proteins to 3D organization. Trends Cell Biol. (2004) 14:359–68. doi: 10.1016/j.tcb.2004.05.009
18. Vidotto T, Melo CM, Castelli E, Koti M, Dos Reis RB, Squire JA. Emerging role of PTEN loss in evasion of the immune response to tumours. Br J Cancer. (2020) 122:1732–43. doi: 10.1038/s41416-020-0834-6
19. Lee JO, Yang H, Georgescu MM, Di Cristofano A, Maehama T, Shi Y, et al. Crystal structure of the PTEN tumor suppressor: implications for its phosphoinositide phosphatase activity and membrane association. Cell. (1999) 99:323–34. doi: 10.1016/s0092-8674(00)81663-3
20. Poluri RTK, Audet-Walsh É. Genomic deletion at 10q23 in prostate cancer: more than PTEN loss? Front Oncol. (2018) 8:246. doi: 10.3389/fonc.2018.00246
21. Zhou X, Yang X, Sun X, Xu X, Li X, Guo Y, et al. Effect of PTEN loss on metabolic reprogramming in prostate cancer cells. Oncol Lett. (2019) 17:2856–66. doi: 10.3892/ol.2019.9932
22. Gong L, Govan JM, Evans EB, Dai H, Wang E, Lee S-W, et al. Nuclear PTEN tumor-suppressor functions through maintaining heterochromatin structure. Cell Cycle (Georgetown Tex.). (2015) 14:2323–32. doi: 10.1080/15384101.2015.1044174
23. Steelman LS, Bertrand FE, McCubrey JA. The complexity of PTEN: mutation, marker and potential target for therapeutic intervention. Expert Opin Ther Targets. (2004) 8:537–50. doi: 10.1517/14728222.8.6.537
24. Geybels MS, Fang M, Wright JL, Qu X, Bibikova M, Klotzle B, et al. PTEN loss is associated with prostate cancer recurrence and alterations in tumor DNA methylation profiles. Oncotarget. (2017) 8:84338–48. doi: 10.18632/oncotarget.20940
25. Huang FW, Mosquera JM, Garofalo A, Oh C, Baco M, Amin-Mansour A, et al. Exome sequencing of african-american prostate cancer reveals loss-of-function ERF mutations. Cancer Discovery. (2017) 7:973–83. doi: 10.1158/2159-8290.CD-16-0960
26. Tosoian JJ, Almutairi F, Morais CL, Glavaris S, Hicks J, Sundi D, et al. Prevalence and prognostic significance of PTEN loss in african-american and european-american men undergoing radical prostatectomy. Eur Urol. (2017) 71:697–700. doi: 10.1016/j.eururo.2016.07.026
27. Cheng F, Eng C. PTEN mutations trigger resistance to immunotherapy. In Trends Mol Med. (2019) 25:461–3. doi: 10.1016/j.molmed.2019.03.003
28. Hägglöf C, Hammarsten P, Strömvall K, Egevad L, Josefsson A, Stattin P, et al. TMPRSS2-ERG expression predicts prostate cancer survival and associates with stromal biomarkers. PloS One. (2014) 9:e86824. doi: 10.1371/journal.pone.0086824
29. Omar M, Xu Z, Rand SB, Salles DC, Schaeffer EM, Lotan TL, et al. Abstract 5369: Detection of ERG: TMPRSS2 gene fusion in prostate cancer from histopathology slides using attention-based deep learning. Cancer Res. (2023) 83:5369. doi: 10.1158/1538-7445.AM2023-5369
30. Barros-Silva JD, Paulo P, Bakken AC, Cerveira N, Løvf M, Henrique R, et al. Novel 5’ fusion partners of ETV1 and ETV4 in prostate cancer. Neoplasia (New York N.Y.). (2013) 15:720–6. doi: 10.1593/neo.13232
31. Cerveira N, Ribeiro FR, Peixoto A, Costa V, Henrique R, Jerönimo C, et al. TMPRSS2-ERG gene fusion causing ERG overexpression precedes chromosome copy number changes in prostate carcinomas, paired HGPIN lesions. Neoplasia. (2006) 8:826–32. doi: 10.1593/neo.06427
32. Liu F, Patient R. Genome-wide analysis of the zebrafish ETS family identifies three genes required for hemangioblast differentiation or angiogenesis. Circ Res. (2008) 103:1147–54. doi: 10.1161/CIRCRESAHA.108.179713
33. Testa U, Castelli G, Pelosi E. Cellular and molecular mechanisms underlying prostate cancer development: therapeutic implications. Medicines (Basel Switzerland). (2019) 6. doi: 10.3390/medicines6030082
34. Xiao Q, Sun Y, Dobi A, Srivastava S, Wang W, Srivastava S, et al. Systematic analysis reveals molecular characteristics of ERG-negative prostate cancer. Sci Rep. (2018) 8:12868. doi: 10.1038/s41598-018-30325-9
35. Hermans KG, Marion Rv, Dekken Hv, Jenster G, Weerden WMv, Trapman J. TMPRSS2:ERG fusion by translocation or interstitial deletion is highly relevant in androgen-dependent prostate cancer, but is bypassed in late-stage androgen receptor-negative prostate cancer. Cancer Res. (2006) 66:10658–63. doi: 10.1158/0008-5472.CAN-06-1871
36. Lorenzin F, Demichelis F. Past, current, and future strategies to target ERG fusion-positive prostate cancer. Cancers. (2022) 14. doi: 10.3390/cancers14051118
37. Perner S, Demichelis F, Beroukhim R, Schmidt FH, Mosquera J-M, Setlur S, et al. TMPRSS2:ERG fusion-associated deletions provide insight into the heterogeneity of prostate cancer. Cancer Res. (2006) 66:8337–41. doi: 10.1158/0008-5472.CAN-06-1482
38. Kohaar I, Li Q, Chen Y, Ravindranath L, Young D, Ali A, et al. Association of germline genetic variants with TMPRSS2-ERG fusion status in prostate cancer. Oncotarget. (2020) 11:1321–33. doi: 10.18632/oncotarget.27534
39. Giunchi F, Massari F, Altimari A, Gruppioni E, Nobili E, Fiorentino M, et al. Dual TMPRSS2:ERG fusion in a patient with lung and prostate cancers. Diagnostics. (2020) 10. doi: 10.3390/diagnostics10121109
40. Kobelyatskaya AA, Pudova EA, Snezhkina AV, Fedorova MS, Pavlov VS, Guvatova ZG, et al. Impact TMPRSS2–ERG molecular subtype on prostate cancer recurrence. Life. (2021) 11. doi: 10.3390/life11060588
41. Yamoah K, Lal P, Awasthi S, Naghavi AO, Rounbehler RJ, Gerke T, et al. TMPRSS2-ERG fusion impacts anterior tumor location in men with prostate cancer. Prostate. (2021) 81:109–17. doi: 10.1002/pros.24086
42. Manzar N, Ganguly P, Khan UK, Ateeq B. Transcription networks rewire gene repertoire to coordinate cellular reprograming in prostate cancer. Semin Cancer Biol. (2023) 89:76–91. doi: 10.1016/j.semcancer.2023.01.004
43. St John J, Powell K, Conley-LaComb MK, Chinni SR. TMPRSS2-ERG fusion gene expression in prostate tumor cells and its clinical and biological significance in prostate cancer progression. J Cancer Sci Ther. (2012) 4:94. doi: 10.4172/1948-5956.1000119
44. Semaan L, Mander N, Cher ML, Chinni SR. TMPRSS2-ERG fusions confer efficacy of enzalutamide in an in vivo bone tumor growth model. BMC Cancer. (2019) 19:1–10. doi: 10.1186/s12885-019-6185-0
45. Blee AM. Roles of TMPRSS2-erg gene fusions during prostate adenocarcinoma initiation and progression. Coll Medicine-Mayo Clinic. (2018). doi: 10.1593/neo.07822
46. Assadi M, Jokar N, Ghasemi M, Nabipour I, Gholamrezanezhad A, Ahmadzadehfar H. Precision medicine approach in prostate cancer. Curr Pharm Design. (2020) 26:3783–98. doi: 10.2174/1381612826666200218104921
47. Brenner JC. Therapeutic targeting of ETS rearranged cancers. Michingan USA: University of Michigan(2012).
48. Carver BS, Tran J, Chen Z, Carracedo-Perez A, Alimonti A, Nardella C, et al. ETS rearrangements and prostate cancer initiation. Nature. (2009) 457:E1; discussion E2–3. doi: 10.1038/nature07738
49. Consortium, T. Gte, Ardlie KG, Deluca DS, Segrè AV, Sullivan TJ, Young TR, et al. The Genotype-Tissue Expression (GTEx) pilot analysis: Multitissue gene regulation in humans. Science. (2015) 348:648–60. doi: 10.1126/science.1262110
50. Chen Y, Chi P, Rockowitz S, Iaquinta PJ, Shamu T, Shukla S, et al. ETS factors reprogram the androgen receptor cistrome and prime prostate tumorigenesis in response to PTEN loss. Nat Med. (2013) 19:1023–9. doi: 10.1038/nm.3216
51. Yu J, Yu J, Mani R-S, Cao Q, Brenner CJ, Cao X, et al. An integrated network of androgen receptor, polycomb, and TMPRSS2-ERG gene fusions in prostate cancer progression. Cancer Cell. (2010) 17:443–54. doi: 10.1016/j.ccr.2010.03.018
52. Iljin K, Wolf M, Edgren H, Gupta S, Kilpinen S, Skotheim RI, et al. TMPRSS2 fusions with oncogenic ETS factors in prostate cancer involve unbalanced genomic rearrangements and are associated with HDAC1 and epigenetic reprogramming. Cancer Res. (2006) 66:10242–6. doi: 10.1158/0008-5472.CAN-06-1986
53. Leshem O, Madar S, Kogan-Sakin I, Kamer I, Goldstein I, Brosh R, et al. TMPRSS2/ERG promotes epithelial to mesenchymal transition through the ZEB1/ZEB2 axis in a prostate cancer model. PloS One. (2011) 6:e21650. doi: 10.1371/journal.pone.0021650
54. Wu L, Zhao JC, Kim J, Jin H-J, Wang C-Y, Yu J. ERG is a critical regulator of Wnt/LEF1 signaling in prostate cancer. Cancer Res. (2013) 73:6068–79. doi: 10.1158/0008-5472.CAN-13-0882
55. Hoogland AM, Jenster G, Van Weerden WM, Trapman J, van der Kwast T, Roobol MJ, et al. ERG immunohistochemistry is not predictive for PSA recurrence, local recurrence or overall survival after radical prostatectomy for prostate cancer. Modern Pathol. (2012) 25:471–9. doi: 10.1038/modpathol.2011.176
56. Minner S, Enodien M, Sirma H, Luebke AM, Krohn A, Mayer PS, et al. ERG status is unrelated to PSA recurrence in radically operated prostate cancer in the absence of antihormonal therapy. Clin Cancer Res. (2011) 17:5878–88. doi: 10.1158/1078-0432.CCR-11-1251
57. Xu B, Chevarie-Davis M, Chevalier S, Scarlata E, Zeizafoun N, Dragomir A, et al. The prognostic role of ERG immunopositivity in prostatic acinar adenocarcinoma: a study including 454 cases and review of the literature. Hum Pathol. (2014) 45:488–97. doi: 10.1016/j.humpath.2013.10.012
58. Reid AHM, Attard G, Ambroisine L, Fisher G, Kovacs G, Brewer D, et al. Molecular characterisation of ERG, ETV1 and PTEN gene loci identifies patients at low and high risk of death from prostate cancer. Br J Cancer. (2010) 102:678–84. doi: 10.1038/sj.bjc.6605554
59. Shan L, Ji T, Su X, Shao Q, Du T, Zhang S. TMPRSS2-ERG fusion promotes recruitment of regulatory T cells and tumor growth in prostate cancer. Am J Med Sci. (2018) 356:72–8. doi: 10.1016/j.amjms.2018.03.023
60. Al Bashir S, Alorjani MS, Kheirallah K, Al Hamad M, Haddad HK, Al-Dwairy A, et al. PTEN, ERG, SPINK1, and TFF3 status and relationship in a prostate cancer cohort from Jordanian arab population. Medicina. (2024) 60:174. doi: 10.3390/medicina60010174
61. Ferraldeschi R, Rodrigues DN, Riisnaes R, Miranda S, Figueiredo I, Rescigno P, et al. PTEN protein loss and clinical outcome from castration-resistant prostate cancer treated with abiraterone acetate. Eur Urol. (2015) 67:795–802. doi: 10.1016/j.eururo.2014.10.027
62. Yoshimoto M, Cunha IW, Coudry RA, Fonseca FP, Torres CH, Soares FA, et al. FISH analysis of 107 prostate cancers shows that PTEN genomic deletion is associated with poor clinical outcome. Br J Cancer. (2007) 97:678–85. doi: 10.1038/sj.bjc.6603924
63. Yorioka MAW, Murta CB, Leite KRM, Cardili L, de Mello ES, de Carvalho Fazoli AJ, et al. ERG and PTEN role on active surveillance for low-risk prostate cancer in the multiparametric MRI era. Prostate. (2024) 85(4)364–73. doi: 10.1002/pros.24835
64. Abdelsalam RA, Khalifeh I, Box A, Kalantarian M, Ghosh S, Abou-Ouf H, et al. Molecular characterization of prostate cancer in Middle Eastern population highlights differences with Western populations with prognostic implication. J Cancer Res Clin Oncol. (2020) 146:1701–9. doi: 10.1007/s00432-020-03221-x
65. Bismar TA, Yoshimoto M, Duan Q, Liu S, Sircar K, Squire JA. Interactions and relationships of PTEN, ERG, SPINK1 and AR in castration-resistant prostate cancer. Histopathology. (2012) 60:645–52. doi: 10.1111/j.1365-2559.2011.04116.x
66. Rupp NJ, Freiberger SN, Ferraro DA, Laudicella R, Heimer J, Muehlematter UJ, et al. Immunohistochemical ERG positivity is associated with decreased PSMA expression and lower visibility in corresponding [68Ga]Ga-PSMA-11 PET scans of primary prostate cancer. Eur J Nucl Med Mol Imaging. (2024) 52:305–13. doi: 10.1007/s00259-024-06856-x
67. Blackburn J, Vecchiarelli S, Heyer EE, Patrick SM, Lyons RJ, Jaratlerdsiri W, et al. TMPRSS2-ERG fusions linked to prostate cancer racial health disparities: A focus on Africa. Prostate. (2019) 79:1191–6. doi: 10.1002/pros.v79.10
68. Troyer DA, Jamaspishvili T, Wei W, Feng Z, Good J, Hawley S, et al. A multicenter study shows PTEN deletion is strongly associated with seminal vesicle involvement and extracapsular extension in localized prostate cancer. Prostate. (2015) 75:1206–15. doi: 10.1002/pros.23003
69. Krohn A, Diedler T, Burkhardt L, Mayer P-S, De Silva C, Meyer-Kornblum M, et al. Genomic deletion of PTEN is associated with tumor progression and early PSA recurrence in ERG fusion-positive and fusion-negative prostate cancer. Am J Pathol. (2012) 181:401–12. doi: 10.1016/j.ajpath.2012.04.026
70. Yoshimoto M, Cutz J-C, Nuin PAS, Joshua AM, Bayani J, Evans AJ, et al. Interphase FISH analysis of PTEN in histologic sections shows genomic deletions in 68% of primary prostate cancer and 23% of high-grade prostatic intra-epithelial neoplasias. Cancer Genet Cytogenetics. (2006) 169:128–37. doi: 10.1016/j.cancergencyto.2006.04.003
71. Yoshimoto M, Joshua AM, Cunha IW, Coudry RA, Fonseca FP, Ludkovski O, et al. Absence of TMPRSS2: ERG fusions and PTEN losses in prostate cancer is associated with a favorable outcome. Modern Pathol. (2008) 21:1451–60. doi: 10.1038/modpathol.2008.96
72. Ahearn TU, Pettersson A, Ebot EM, Gerke T, Graff RE, Morais CL, et al. A prospective investigation of PTEN loss and ERG expression in lethal prostate cancer. J Natl Cancer Institute. (2016) 108:1–9. doi: 10.1093/jnci/djv346
73. Mitala Y, Ssenkumba B, Birungi A, Kiconco R, Mutakooha MM, Atwine R. A cross-sectional study of ERG expression and the relationship with clinicopathological features of Prostate cancer in Southwestern Uganda. Diagn Pathol. (2024) 19:67. doi: 10.1186/s13000-024-01494-1
74. Spinos T, Georgiou A, Voulgari O, Goutas D, Lazaris AC, Thymara I, et al. Association between ERG/PTEN genes and pathologic parameters of prostate cancer with an emphasis on gleason score: A literature review. Cancer Diagnosis Prognosis. (2023) 3:291–6. doi: 10.21873/cdp.10214
75. Carver BS, Tran J, Gopalan A, Chen Z, Shaikh S, Carracedo A, et al. Aberrant ERG expression cooperates with loss of PTEN to promote cancer progression in the prostate. Nat Genet. (2009) 41:619–24. doi: 10.1038/ng.370
76. Krohn A, Freudenthaler F, Harasimowicz S, Kluth M, Fuchs S, Burkhardt L, et al. Heterogeneity and chronology of PTEN deletion and ERG fusion in prostate cancer. Modern Pathol. (2014) 27:1612–20. doi: 10.1038/modpathol.2014.70
77. Park K, Dalton JT, Narayanan R, Barbieri CE, Hancock ML, Bostwick DG, et al. TMPRSS2:ERG gene fusion predicts subsequent detection of prostate cancer in patients with high-grade prostatic intraepithelial neoplasia. J Clin Oncology : Off J Am Soc Clin Oncol. (2014) 32:206–11. doi: 10.1200/JCO.2013.49.8386
78. Ni L, Llewellyn R, Kesler CT, Kelley JB, Spencer A, Snow CJ, et al. Androgen induces a switch from cytoplasmic retention to nuclear import of the androgen receptor. Mol Cell Biol. (2013). doi: 10.1128/MCB.00647-13
79. Wang Z. Androgen-responsive genes in prostate cancer: regulation, function and clinical applications. New York USA: Springer Science & Business Media(2013).
80. Makkonen HT. Androgen receptor-mediated gene activation in prostate cancer cells. Itä-Suomen yliopisto. (2010).
81. He Y, Xu W, Xiao Y-T, Huang H, Gu D, Ren S. Targeting signaling pathways in prostate cancer: mechanisms and clinical trials. Signal Transduction Targeted Ther. (2022) 7:198. doi: 10.1038/s41392-022-01042-7
82. Abou-Ouf H, Assem H, Ghosh S, Karnes RJ, Stoletov K, Palanisamy N, et al. High serine-arginine protein kinase 1 expression with PTEN loss defines aggressive phenotype of prostate cancer associated with lethal outcome and decreased overall survival. Eur Urol Open Sci. (2021) 23:1–8. doi: 10.1016/j.euros.2020.11.005
83. Mithal P, Allott E, Gerber L, Reid J, Welbourn W, Tikishvili E, et al. PTEN loss in biopsy tissue predicts poor clinical outcomes in prostate cancer. Int J Urol. (2014) 21:1209–14. doi: 10.1111/iju.12571
84. Culig Z, Puhr M. Interleukin-6: a multifunctional targetable cytokine in human prostate cancer. Mol Cell Endocrinol. (2012) 360:52–8. doi: 10.1016/j.mce.2011.05.033
85. Smith DA, Kiba A, Zong Y, Witte ON. Interleukin-6 and oncostatin-M synergize with the PI3K/AKT pathway to promote aggressive prostate Malignancy in mouse and human tissues. Mol Cancer Research : MCR. (2013) 11:1159–65. doi: 10.1158/1541-7786.MCR-13-0238
86. Georgescu MM, Kirsch KH, Kaloudis P, Yang H, Pavletich NP, Hanafusa H. Stabilization and productive positioning roles of the C2 domain of PTEN tumor suppressor. Cancer Res. (2000) 60:7033–8.
87. Haverkamp J, Charbonneau B, Ratliff TL. Prostate inflammation and its potential impact on prostate cancer: a current review. J Cell Biochem. (2008) 103:1344–53. doi: 10.1002/jcb.21536
88. Blum DL, Koyama T, M’Koma AE, Iturregui JM, Martinez-Ferrer M, Uwamariya C, et al. Chemokine markers predict biochemical recurrence of prostate cancer following prostatectomy. Clin Cancer Research : Off J Am Assoc Cancer Res. (2008) 14:7790–7. doi: 10.1158/1078-0432.CCR-08-1716
89. Brady L, Carlsson J, Baird A-M, Casey O, Vlajnic T, Murchan P, et al. Correlation of integrated ERG/PTEN assessment with biochemical recurrence in prostate cancer. Cancer Treat Res Commun. (2021) 29:100451. doi: 10.1016/j.ctarc.2021.100451
90. Lahdensuo K, Erickson A, Saarinen I, Seikkula H, Lundin J, Lundin M, et al. Loss of PTEN expression in ERG-negative prostate cancer predicts secondary therapies and leads to shorter disease-specific survival time after radical prostatectomy. Modern Pathol. (2016) 29:1565–74. doi: 10.1038/modpathol.2016.154
91. Morais CE, Gurgel DC, Teixeira AC, Arrudamattos TV, Alves da Silva AV, Tavora F. Prevalence of ERG expression and PTEN loss in a Brazilian prostate cancer cohort. Braz J Med Biol Res. (2019) 52:1–9. doi: 10.1590/1414-431x20198483
92. Lotan TL, Gupta NS, Wang W, Toubaji A, Haffner MC, Chaux A, et al. ERG gene rearrangements are common in prostatic small cell carcinomas. Modern Pathology : Off J United States Can Acad Pathology Inc. (2011) 24:820—828. doi: 10.1038/modpathol.2011.7
93. Yoshimoto M, Ludkovski O, DeGrace D, Williams JL, Evans A, Sircar K, et al. PTEN genomic deletions that characterize aggressive prostate cancer originate close to segmental duplications. Genes Chromosomes Cancer. (2012) 51:149–60. doi: 10.1002/gcc.20939
94. Cai C, Wang H, Xu Y, Chen S, Balk SP. Reactivation of androgen receptor–regulated TMPRSS2: ERG gene expression in castration-resistant prostate cancer. Cancer Res. (2009) 69:6027–32. doi: 10.1158/0008-5472.CAN-09-0395
95. Esteban-Villarrubia J, Ballesteros PA, Martín-Serrano M, Vico MR, Funes JM, de Velasco G, et al. Mechanisms of immune evasion in PTEN loss prostate cancer. Immuno. (2024) 4:444–60. doi: 10.3390/immuno4040028
96. Haney NM, Faisal FA, Lu J, Guedes LB, Reuter VE, Scher HI, et al. PTEN loss with ERG negative status is associated with lethal disease after radical prostatectomy. J Urol. (2020) 203:344–50. doi: 10.1097/JU.0000000000000533
97. Zhou CK, Young D, Yeboah ED, Coburn SB, Tettey Y, Biritwum RB, et al. TMPRSS2:ERG gene fusions in prostate cancer of west African men and aMeta-analysis of racial differences. Am J Epidemiol. (2017) 186:1352–61. doi: 10.1093/aje/kwx235
98. Ma F, Chen S, Calagua C, Arai S, Xie F, Voznesensky O, et al. Androgen deprivation-mediated activation of AKT is enhanced in prostate cancer with TMPRSS2:ERG fusion. BioRxiv. (2024). doi: 10.1101/2024.08.26.609679
99. Li J, Yadav SS, O’Connor J, Herzog B, Elaiho C, Yadav KK, et al. Combination effect of therapies targeting the pi3k-mtor and ar signaling pathways in prostate cancer: pd32-09. J Urol. (2016) 195:e764. doi: 10.1016/j.juro.2016.02.685
100. Silva KCS, Tambwe N, Mahfouz DH, Wium M, Cacciatore S, Paccez JD, et al. Transcription factors in prostate cancer: insights for disease development and diagnostic and therapeutic approaches. Genes. (2024) 15. doi: 10.3390/genes15040450
101. Glaviano A, Foo ASC, Lam HY, Yap KCH, Jacot W, Jones RH, et al. PI3K/AKT/mTOR signaling transduction pathway and targeted therapies in cancer. Mol Cancer. (2023) 22:138. doi: 10.1186/s12943-023-01827-6
102. Imada EL, Sanchez DF, Dinalankara W, Vidotto T, Ebot EM, Tyekucheva S, et al. Transcriptional landscape of PTEN loss in primary prostate cancer. BMC Cancer. (2021) 21:856. doi: 10.1186/s12885-021-08593-y
103. Graff RE, Pettersson A, Lis RT, DuPre N, Jordahl KM, Nuttall E, et al. The TMPRSS2: ERG fusion and response to androgen deprivation therapy for prostate cancer. Prostate. (2015) 75:897–906. doi: 10.1002/pros.22973
104. Brenner JC, Ateeq B, Li Y, Yocum AK, Cao Q, Asangani IA, et al. Mechanistic rationale for inhibition of poly(ADP-ribose) polymerase in ETS gene fusion-positive prostate cancer. Cancer Cell. (2011) 19:664–78. doi: 10.1016/j.ccr.2011.04.010
105. Shakfa N, Li D, Conseil G, Lightbody ED, Wilson-Sanchez J, Hamade A, et al. Cancer cell genotype associated tumor immune microenvironment exhibits differential response to therapeutic STING pathway activation in high-grade serous ovarian cancer. J ImmunoTherapy Cancer. (2023) 11. doi: 10.1136/jitc-2022-006170
106. Vidotto T, Melo CM, Lautert-Dutra W, Chaves LP, Reis RB, Squire JA. Pan-cancer genomic analysis shows hemizygous PTEN loss tumors are associated with immune evasion and poor outcome. Sci Rep. (2023) 13:5049. doi: 10.1038/s41598-023-31759-6
107. Shao L, Kahraman N, Yan G, Wang J, Ozpolat B, Ittmann M. Targeting the TMPRSS2/ERG fusion mRNA using liposomal nanovectors enhances docetaxel treatment in prostate cancer. Prostate. (2020) 80:65–73. doi: 10.1002/pros.23918
108. Sedarsky J, Degon M, Srivastava S, Dobi A. Ethnicity and ERG frequency in prostate cancer. Nat Rev Urol. (2017) 15:125-31. doi: 10.1038/nrurol.2017.140
109. Fallahabadi ZR, Noori Daloii MR, Mahdian R, Behjati F, Shokrgozar MA, Abolhasani M, et al. Frequency of PTEN alterations, TMPRSS2-ERG fusion and their association in prostate cancer. Gene. (2016) 575:755–60. doi: 10.1016/j.gene.2015.09.068
110. Repka MC, Sholklapper T, Zwart AL, Danner M, Ayoob M, Yung T, et al. Prognostic utility of biopsy-based PTEN and ERG status on biochemical progression and overall survival after SBRT for localized prostate cancer. Front Oncol. (2024) 14. doi: 10.3389/fonc.2024.1381134
111. Goel S, Bhatia V, Kundu S, Biswas T, Carskadon S, Gupta N, et al. Transcriptional network involving ERG and AR orchestrates Distal-less homeobox-1 mediated prostate cancer progression. Nat Commun. (2021) 12:5325. doi: 10.1038/s41467-021-25623-2
112. Cham J, Venkateswaran AR, Bhangoo M. Targeting the PI3K-AKT-mTOR pathway in castration resistant prostate cancer: A review article. Clin Genitourinary Cancer. (2021) 19:563.e1–7. doi: 10.1016/j.clgc.2021.07.014
113. Tortorella E, Giantulli S, Sciarra A, Silvestri I. AR and PI3K/AKT in prostate cancer: A tale of two interconnected pathways. Int J Mol Sci. (2023) 24. doi: 10.3390/ijms24032046
114. Kissick HT, On ST, Dunn LK, Sanda MG, Asara JM, Pellegrini KL, et al. The transcription factor ERG increases expression of neurotransmitter receptors on prostate cancer cells. BMC Cancer. (2015) 15:604. doi: 10.1186/s12885-015-1612-3
115. Tomlins SA, Palanisamy N, Siddiqui J, Chinnaiyan AM, Kunju LP. Antibody-based detection of ERG rearrangements in prostate core biopsies, including diagnostically challenging cases: ERG staining in prostate core biopsies. Arch Pathol Lab Med. (2012) 136:935–46. doi: 10.5858/arpa.2011-0424-OA
116. Scaravilli M, Koivukoski S, Latonen L. Androgen-driven fusion genes and chimeric transcripts in prostate cancer. Front Cell Dev Biol. (2021) 9:623809. doi: 10.3389/fcell.2021.623809
117. Merz C, von Mässenhausen A, Queisser A, Vogel W, Andrén O, Kirfel J, et al. IL-6 overexpression in ERG-positive prostate cancer is mediated by prostaglandin receptor EP2. Am J Pathol. (2016) 186:974–84. doi: 10.1016/j.ajpath.2015.12.009
118. Stephan C, Rittenhouse H, Hu X, Cammann H, Jung K. Prostate-specific antigen (PSA) screening and new biomarkers for prostate cancer (PCa). EJIFCC. (2014) 25:55–78.
119. Teoh JYC, Tsu JHL, Yuen SKK, Chan SYS, Chiu PKF, Lee W-M, et al. Prognostic significance of time to prostate-specific antigen (PSA) nadir and its relationship to survival beyond time to PSA nadir for prostate cancer patients with bone metastases after primary androgen deprivation therapy. Ann Surg Oncol. (2015) 22:1385–91. doi: 10.1245/s10434-014-4105-8
120. Demichelis F, Fall K, Perner S, Andrén O, Schmidt F, Setlur SR, et al. TMPRSS2:ERG gene fusion associated with lethal prostate cancer in a watchful waiting cohort. Oncogene. (2007) 26:4596–9. doi: 10.1038/sj.onc.1210237
121. Song C, Chen H. Predictive significance of TMRPSS2-ERG fusion in prostate cancer: a meta-analysis. Cancer Cell Int. (2018) 18:177. doi: 10.1186/s12935-018-0672-2
122. Bruinsma SM, Bangma CH, Carroll PR, Leapman MS, Rannikko A, Petrides N, et al. Active surveillance for prostate cancer: a narrative review of clinical guidelines. Nat Rev Urol. (2016) 13:151–67. doi: 10.1038/nrurol.2015.313
123. Barrett T, Haider MA. The emerging role of MRI in prostate cancer active surveillance and ongoing challenges. Am J Roentgenology. (2017) 208:131–9. doi: 10.2214/AJR.16.16355
124. Ma TM, Tosoian JJ, Schaeffer EM, Landis P, Wolf S, Macura KJ, et al. The role of multiparametric magnetic resonance imaging/ultrasound fusion biopsy in active surveillance. Eur Urol. (2017) 71:174–80. doi: 10.1016/j.eururo.2016.05.021
125. Bismar TA, Hegazy S, Feng Z, Yu D, Donnelly B, Palanisamy N, et al. Clinical utility of assessing PTEN and ERG protein expression in prostate cancer patients: a proposed method for risk stratification. J Cancer Res Clin Oncol. (2018) 144:2117–25. doi: 10.1007/s00432-018-2730-5
126. Epstein JI, Egevad L, Amin MB, Delahunt B, Srigley JR, Humphrey PA, et al. The 2014 international society of urological pathology (ISUP) consensus conference on gleason grading of prostatic carcinoma: definition of grading patterns and proposal for a new grading system. Am J Surg Pathol. (2016) 40. doi: 10.1097/PAS.0000000000000530
127. Lotan TL, Wei W, Ludkovski O, Morais CL, Guedes LB, Jamaspishvili T, et al. Analytic validation of a clinical-grade PTEN immunohistochemistry assay in prostate cancer by comparison with PTEN FISH. Modern Pathology : Off J United States Can Acad Pathology Inc. (2016) 29:904–14. doi: 10.1038/modpathol.2016.88
128. Abeshouse A, Ahn J, Akbani R, Ally A, Amin S, Andry CD, et al. The molecular taxonomy of primary prostate cancer. Cell. (2015) 163:1011–25. doi: 10.1016/j.cell.2015.10.025
129. Chung J-S, Morgan TM, Hong SK. Clinical implications of genomic evaluations for prostate cancer risk stratification, screening, and treatment: a narrative review. Prostate Int. (2020) 8:99–106. doi: 10.1016/j.prnil.2020.09.001
130. Mateo J, Carreira S, Sandhu S, Miranda S, Mossop H, Perez-Lopez R, et al. DNA-repair defects and olaparib in metastatic prostate cancer. New Engl J Med. (2015) 373:1697–708. doi: 10.1056/NEJMoa1506859
Keywords: tumour molecular subtype, PTEN, TMPRSS2:ERG, genome, immune response, prostate cancer, immunohistochemistry
Citation: Bugoye FC, Torrorey-Sawe R, Biegon R, Dharsee N, Mafumiko F, Kibona H, Aboud S, Patel K and Mining S (2025) Exploring therapeutic applications of PTEN, TMPRSS2:ERG fusion, and tumour molecular subtypes in prostate cancer management. Front. Oncol. 15:1521204. doi: 10.3389/fonc.2025.1521204
Received: 01 November 2024; Accepted: 21 February 2025;
Published: 17 March 2025.
Edited by:
Anna Grazia Recchia, Independent Researcher, Toronto, ON, CanadaReviewed by:
R.C. Koumar, Yenepoya University, IndiaCopyright © 2025 Bugoye, Torrorey-Sawe, Biegon, Dharsee, Mafumiko, Kibona, Aboud, Patel and Mining. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Fidelis Charles Bugoye, YnVnb3llODFAeWFob28uY28udWs=; Rispah Torrorey-Sawe, dG9ycm9yZXlAZ21haWwuY29t
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.