
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Microbiol. , 03 April 2025
Sec. Microbial Physiology and Metabolism
Volume 16 - 2025 | https://doi.org/10.3389/fmicb.2025.1579813
This article is part of the Research Topic Structure and Mechanism of Microbial Membrane Active Transporters View all 4 articles
 Dmitrii M. Bubnov1*†
Dmitrii M. Bubnov1*† Andrey A. Khozov1,2†
Andrey A. Khozov1,2† Tatiana V. Vybornaya1
Tatiana V. Vybornaya1 Agnessa A. Stepanova1,2
Agnessa A. Stepanova1,2 Sergey V. Molev1,2
Sergey V. Molev1,2 Olga E. Melkina1
Olga E. Melkina1 Gennadii A. Badun3
Gennadii A. Badun3 Maria G. Chernysheva3
Maria G. Chernysheva3 Ilia A. Skob4
Ilia A. Skob4 Alexander I. Netrusov2
Alexander I. Netrusov2 Sergey P. Sineoky1
Sergey P. Sineoky1In this study, we identified eight multicopy suppressors (yhjE, sdaC, ydgI, alaE, ychE, yqeG, proP, and yjeM) and three distinct classes of chromosomal mutations (lrp, marC, and cycA) capable of complementing the growth defect caused by threonine uptake deficiency in the sstT tdcC livKHMGF brnQ thrP strain. YhjE, SdaC, YdgI, AlaE, mutant MarC, and CycA exhibited measurable threonine-specific uptake activity in the in vitro assay. Phenotypic assays revealed that YhjE and SdaC were the main entry points for threonine in a strain lacking major threonine-specific permeases. A derivative of the threonine-auxotrophic sstT tdcC livKHMGF brnQ thrP mutant, harboring deletions of eight multicopy suppressors, exhibited significantly reduced fitness at subsaturating threonine concentrations and improved fitness at toxic threonine concentrations, indicating a defect in membrane permeability. These results may help guide the effective construction of threonine-producing strains, extend knowledge on the substrate preferences of SdaC, AlaE, and ProP, and provide clues for further studies on the exact substrate range of YhjE, YdgI, YjeM, YchE, MarC, and YqeG whose physiologically relevant functions have not yet been established.
The transport of metabolites across the cytoplasmic membrane is a process that plays a pivotal role in the assimilation of carbon and energy sources, as well as other compounds required for biosynthetic reactions. This cellular subsystem mediates the interactions between cells within a population and provides an adequate response to variations in the composition of the surrounding environment. The mechanisms of solute uptake are of great interest in clinical practice, with recent studies showing that functional amino acid transport systems are essential virulence factors for the severe human pathogens Yersinia pestis and Bacillus anthracis (Palace et al., 2014; Dutta et al., 2022). Hence, uptake systems are promising targets for the production of live-attenuated vaccines and new antibacterial drugs. Membrane transport systems are primary targets for the development of industrial strains capable of producing valuable chemicals, especially amino acids. Inactivation of membrane proteins that facilitate the uptake of a desired compound and overexpression of appropriate exporters promote their excretion into the medium, thereby preventing feedback inhibition of key biosynthetic enzymes and toxicity to producer cells (Okamoto et al., 1997; Kruse et al., 2001, 2002; Doroshenko et al., 2007; Park et al., 2007; Mundhada et al., 2016). Therefore, the mechanisms of transmembrane transport have been extensively studied over the past few decades. However, the relevant functions of the various membrane proteins remain unclear. For instance, the “y-om” of Escherichia coli, which is a set of genes that lack experimental evidence of function, contains 295 genes encoding putative membrane transporters (Ghatak et al., 2019), whose substrate specificity and physiologically relevant functions remain unknown.
In our previous study, we identified and characterized a high-affinity permease, ThrP, which is capable of translocating L-threonine and L-serine across the cytoplasmic membrane of E. coli K-12 (Khozov et al., 2023). The results suggest that the LIV-I system also participates in L-threonine transport while having KM and Vmax similar to those of ThrP. Additionally, we found that BrnQ, a dedicated transporter for L-isoleucine, L-valine, and L-leucine, is active toward L-threonine under normal conditions and is a primary L-threonine permease at unphysiologically high substrate concentrations. Together with the previously described L-threonine transporters SstT and TdcC (Hama et al., 1987; Sumantran et al., 1990; Ogawa et al., 1997), these proteins confer a major fraction of the L-threonine uptake activity detectable at substrate concentrations of 50–100 μM. Meanwhile, data on the phenotype of a mutant strain auxotrophic for L-threonine and lacking all the listed transport systems suggested that the cell could still consume L-threonine from the medium when its concentration was increased to 1.0–2.0 mM. This observation led us to presume that the cell possessed unknown transport systems that exhibited low-affinity L-threonine transport activity.
In the present study, we identified several permeases via screening of multicopy suppressors and genomic suppressor mutations that complement the growth defect of a strain lacking the major L-threonine transport systems. The results shed light on the physiologically relevant function of several “y”-genes and provide an approach for further elucidation of their role in E. coli cell biology. This study also extends the substrate range of well-established amino acid carriers.
The bacterial strains used in this study were derived from E. coli K-12 MG1655. The genotypes and relevant characteristics of all strains and plasmids used are listed in Tables 1, 2, respectively. The strains were constructed using a combination of λRed-mediated recombineering (Datsenko and Wanner, 2000) and P1 transduction (Thomason et al., 2007). The pBR-yhjE, pBR-yjeM, pBR-sdaC, pBR-ydgI, pBR-alaE, pBR-ychE, pBR-yqeG, and pBR-proP plasmids were recovered from a genomic library of the B1426 strain, based on their ability to enable the growth of the sstT tdcC thrP livKHMGF brnQ mutant B1895 strain on M9 agar supplemented with 0.2 g/L glucose and 1 mM threonine. The exact coordinates (according to the genome of the MG1655 strain; GenBank accession number U00096.3) of the genomic inserts found within the plasmids upon sequencing are listed in Table 2. The pDEW_yhjE reporter plasmid was constructed by ligating two DNA fragments. The insert was amplified from the chromosome of MG1655 using primers yhjE_p_F (5′-ATCGATGAATTCAATAGGCCGGATGCGGCG-3′) and yhjE_p_R (5- GAGCTCGGTACCCGGTTATTTTTTGGCTAACGAATAGC-3′). The resulting fragment was digested using KpnI and EcoRI and ligated into the pDEW201 vector (Van Dyk and Rosson, 1998) that was linearized using the same endonucleases.
Bacteria were routinely grown in lysogeny broth (LB) (10 g/L tryptone, 5 g/L yeast extract, and 10 g/L sodium chloride) at 37°C with shaking at 220 rpm. A solid LB medium was prepared by adding 20 g/L agar to the LB medium. Ampicillin (200 μg/mL), kanamycin (100 μg/mL), spectinomycin (50 μg/mL), and chloramphenicol (20 μg/mL) were added to the medium as needed. Either solid or liquid M9 minimal medium (Sambrook et al., 1989) with 0.2% glucose was used for the phenotype and transport assays. The medium was supplemented with L-threonine, as indicated in the Results section and figure legends. Phenotypic assays were performed as previously described (Khozov et al., 2023).
Standard methods were used for chromosomal DNA isolation, restriction enzyme digestion, agarose gel electrophoresis, ligation, and transformation (Sambrook et al., 1989). PCR amplification was performed using DreamTaq (Thermo Fisher Scientific, Vilnius, Lithuania) or KAPA HIFI (Kapa Biosystems, Wilmington, MA, United States) polymerase. Plasmids were isolated and DNA fragments were extracted using GeneJET Plasmid Miniprep and GeneJET Gel Extraction kits (Thermo Fisher Scientific).
A genomic library of the sstT tdcC thrBC mutant B1426 strain was prepared using the pBR322 vector, as described previously (Khozov et al., 2023). The library was electroporated into the B1895 strain, and transformants were selected on M9 agar plates supplemented with 200 mg/L ampicillin and 1 mM threonine and incubated for 3 days. The resulting colonies were repurified on the same medium to be used for plasmid isolation, followed by sequencing of the genomic inserts using the following primers: 5′- GGTTGAGGCCGTTGAGCAC-3′ and 5′- ACATTAACCTATAAAAATAGGCG-3′.
First, several independent cultures of the B1950 strain were inoculated by picking single colonies into 50 mL test tubes with 20 mL of LB medium, which were incubated overnight with shaking at 37°C and 220 rpm. The next day, each culture was washed twice with 50 mL of 0.9% NaCl solution. The cells were concentrated 10 times relative to the original volume, after which 100 μL aliquots were plated onto M9 agar plates supplemented with 0.2 g/L glucose and 400 μM threonine and incubated at 37°C for 2 days. The resulting mutants were picked, restreaked on the same medium, and subjected to genome resequencing on an Illumina MiSeq (Illumina, CA, United States). Paired-end 2 × 250 bp reads were aligned against the reference genome of the MG1655 strain (U00096.3 GenBank entry) using Breseq 0.36.1 (Barrick et al., 2014) in consensus mode. A set of common mutations detected in the genomes of all the analyzed strains was considered to be inherited from the parental strain and excluded from further analysis.
Uniformly labeled L-[U-14C]threonine was obtained from Moravek Biochemicals (United States). To measure L-[U-14C]threonine uptake, cells were grown overnight in 5 mL of M9 medium supplemented with 0.2% glucose at 37°C with shaking at 220 rpm. Plasmid-carrying strains were grown in the presence of 100 mg/L ampicillin, whereas the other strains were incubated in antibiotic-free M9 medium. For threonine-auxotrophic strains, the medium was supplemented with 50 mM threonine. The overnight culture was diluted in 20 mL of the same medium to obtain an OD600 of 0.0625 and was grown until the OD600 reached 0.5. The cells were harvested in 50 mL polypropylene tubes via centrifugation at 5,000 × g at 4°C for 5 min. The supernatant was discarded, and the cells were washed once with 35 mL of M9 medium. The cells were resuspended in 10 mL of M9 medium with 0.2% glucose and incubated for 2 h at 37°C with shaking at 220 rpm to ensure intracellular threonine exhaustion. All subsequent steps were performed on ice. The cells were harvested, and the pellet was resuspended in 1 mL of M9 medium and transferred to a precooled 1.5 mL tube. The tube was then centrifuged for 45 s in a cooled microcentrifuge rotor at 12,000 × g, and the supernatant was thoroughly aspirated. The pellet was resuspended in 200 μL of M9 medium with 0.2% glucose. The suspension was diluted with the same medium to obtain an OD600 of 32, and chloramphenicol was added at a final concentration of 50 μg/mL to stop protein synthesis. Both the cell suspension and labeled substrate dissolved in M9 medium with 0.2% glucose were separately preincubated for 20 min at 37°C. Uptake was initiated by adding the cell suspension to the substrate solution to obtain an OD600 of 10. The reaction mixture was incubated at 37°C. Subsequently, 12.5 μL sample fractions were periodically collected and immediately filtered through 0.45-μm GVS North America 13-mm nitrocellulose membranes presoaked in M9 medium on a vacuum manifold, followed by two washes with 1 mL of the same medium. Membranes were air-dried at 37°C for 18–20 h, and radioactivity was measured using 5 mL of GC-106 scintillation liquid (4 g 2,5-diphenyloxazole and 0.1 g 2,2′-(1,4-phenylene)bis(5-phenyl-1,3-oxazole) dissolved in 1 L of toluene) on a RackBeta1215 liquid scintillation spectrometer (LKB, Finland). The amount of radioactivity absorbed by the membrane was used as control. For each substrate concentration, the cell-free reaction mixture was incubated, filtered, washed, and counted as being identical to the experimental reactions. The measured values were then subtracted from those obtained in the appropriate experiment. Transport activity was expressed as nanomoles of substrate taken up by 1 mg dry cellular weight (DCW) in 1 min. The DCW value was calculated based on the OD600 of the cell suspension, as described previously (Khozov et al., 2023). All comparisons were performed using two-tailed Student’s t-test with unequal variances at a significance level of 5%.
A single colony of the assayed strain was inoculated into 5 mL of LB medium supplemented with 100 mg/L ampicillin to maintain the pDEW_yhjE plasmid. The overnight culture was diluted to an initial OD600 of 0.004 with fresh M9 medium supplemented with 2 g/L glucose and 40 mM L-threonine. Two hundred microliters of the culture were added to each well of a 96-well plate (black-walled, transparent flat bottom; cat. #665096 Greiner Bio-One, Frickenhausen, Germany). The outer wells were not used to avoid edge effects. A well containing a sterile medium was used as a blank. The plates were incubated at 37°C with double-orbital shaking at 600 rpm using a CLARIOstar Plus luminometer (BMG Labtech, Ortenberg, Germany), and OD600 and luminescence were measured every 15 min. No luminescence emission filter was used. The photomultiplier gain was automatically controlled using an enhanced dynamic range function. The measured values were normalized to a 1 s accumulation time. The acquired data were analyzed using the MARS software. Blank values were subtracted from the raw OD600 and relative luminescence units (RLU) values. The corrected RLU reads at each time point were divided by the corresponding OD600 values to normalize the RLU per cell mass for each well. The average RLU/OD600 values and standard deviations were calculated and plotted against the OD600.
The protein structures predicted using AlphaFold 2 (Jumper et al., 2021) were obtained from the AlphaFold Protein Structure Database (Varadi et al., 2022). Structural visualization was performed using UCSF ChimeraX (Pettersen et al., 2021).
Evolutionary history was inferred using the maximum likelihood method, and the Whelan and Goldman + Freq. Model (Whelan and Goldman, 2001). The initial tree(s) for the heuristic search were automatically obtained by applying the Neighbor-Join and BioNJ algorithms to a matrix of pairwise distances estimated using the JTT model and then selecting the topology with a superior log likelihood value. A discrete gamma distribution was used to model the evolutionary rate differences among sites [five categories (+G, parameter = 8.5853)]. The rate variation model allowed some sites to be evolutionarily invariable ([+I], 0.00% sites). Evolutionary analyses were conducted using MEGA11 (Tamura et al., 2021). The phylogenetic affiliations of the transporters were determined according to the Transporter Classification Database (Saier et al., 2014).
To identify permeases exhibiting uptake activity toward threonine on the ∆sstT ∆tdcC ∆brnQ ∆livKHMGF ∆thrP genetic background, we exploited the inability of the threonine-auxotrophic B1895 strain carrying these mutations to grow on a minimal medium with threonine unless its concentration is increased to 2.5 mM. After electroporation of this strain with a genomic library prepared using the chromosome of the thrBC sstT tdcC strain B1426 and the pBR322 vector, we selected transformants that regained the ability to grow on M9 agar supplemented with 1 mM threonine. Sequencing of chromosomal regions within plasmids isolated from the transformants revealed eight candidate genes encoding known or putative membrane proteins. Among these, three encode previously characterized amino acid carriers. The product of sdaC is an H+/L-serine symporter (Hama et al., 1988; Shao et al., 1994). ProP is an H+-dependent transport system specific toward zwitterionic osmolytes such as proline and glycine betaine, which protect cells from osmotic stress (MacMillan et al., 1999). In addition, it translocates a wide range of compounds including taurine, proline betaine, pipecolate, azetidine-2-carboxylate, 3,4-dehydroproline, carnitine, and ectoine (Csonka, 1981; Stalmach et al., 1983; McLaggan and Epstein, 1991; Jebbar et al., 1992; Verheul et al., 1998; MacMillan et al., 1999). AlaE is an H+-dependent antiporter that translocates L-alanine from the cytoplasm to the periplasmic space (Kim et al., 2015; Katsube et al., 2019; Ihara et al., 2022). The functions of the remaining five genes, yqeG, yhjE, ychE, ydgI, and yjeM, are unknown or are controversial, as inferred from literature analysis and the EcoCyc database (Keseler et al., 2017).
We verified that the amplification of the eight identified genes, rather than other mutations elsewhere in the genome, suppressed the growth defects of the B1895 strain at non-permissive threonine concentrations. We retransformed B1895 cells with appropriate plasmids and tested the growth of the transformants on minimal medium. All plasmid-carrying strains exhibited significantly improved fitness compared with the parental plasmid-free strain (Figure 1A). Among the multicopy suppressors, sdaC, proP, alaE, yhjE, ychE, ydgI, and yjeM had similar effects whereas yqeG stimulated growth in the medium containing the lowest threonine concentration to a lesser extent. Furthermore, we examined the phenotypes of constructed mutant derivatives of B1895 lacking incremental combinations of suppressors. Inactivation of yhjE and sdaC caused a prominent growth defect, whereas deletion of other genes resulted in no distinguishable phenotype under the conditions used (Figure 1B). This may be explained by the relatively low contribution of proP, alaE, ychE, ydgI, yjeM, and yqeG to the total threonine uptake, which was negligible in the case of wild-type alleles but became significant when these genes were amplified on the multicopy vector. The B2818 strain, which lacked all known threonine permeases and the eight identified suppressors, grew well on minimal plates with 6.4 mM threonine, indicating that it still expresses unidentified transport system(s). The activity of this carrier may also exceed that of the other carriers whose deletions result in no visible growth defects, thereby hiding their phenotype. This may represent a multi-subunit complex whose dedicated genes cannot be retrieved from the genome en bloc. Alternatively, the amplification of this system using the pBR322 vector may be deleterious to cell viability. Therefore, other approaches should be used to identify the remaining permease(s).
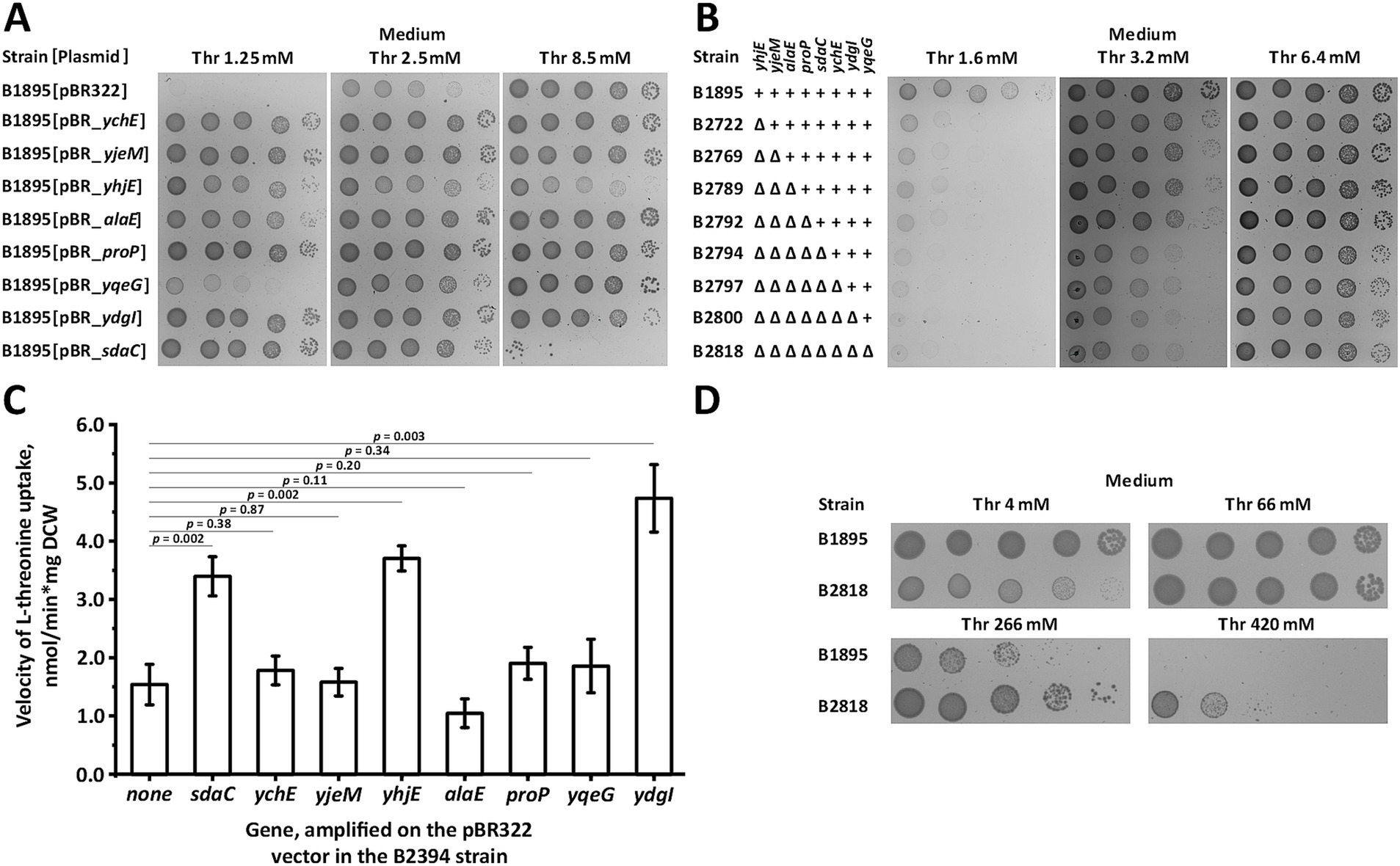
Figure 1. Screening and analysis of the multicopy suppressors of threonine uptake defect. Threonine transport activity was measured as described under “Threonine uptake assay” in the Materials and Methods. “Thr” indicates L-threonine. (A) Phenotypic assay of the B1895 threonine-auxotrophic strain and its derivatives carrying pBR-ychE, pBR-yjeM, pBR-yhjE, pBR-alaE, pBR-proP, pBR-yqeG, pBR-ydgI, and pBR-sdaC plasmids, and the empty pBR322 vector on minimal plates with varying threonine concentrations. (B) Comparison of the phenotypes of B1895 and its derivatives lacking incremental combinations of the multicopy suppressors on minimal plates with varying threonine concentrations. (C) Comparison of threonine transport activity in the B2394 strain lacking the SstT, TdcC, ThrP, BrnQ, and LIV-I carriers with its derivatives overexpressing SdaC, YchE, YjeM, YhjE, AlaE, ProP, YqeG, and YdgI due to the presence of the corresponding plasmids. Measurements were performed using 800 μM L-threonine, and the reaction time was 1 min. The values shown are the average of three independent biological replicates. Error bars indicate standard deviation. p-values were calculated using two-tailed Student’s t-test with unequal variances. (D) Comparison of the growth fitness of B1895 and its derivative, B2818, lacking eight multicopy suppressors on minimal plates with toxic threonine concentrations.
Unlike the other genes, sdaC rendered the B1895 strain sensitive to threonine in the medium at a concentration of 8.5 mM while being overexpressed on the multicopy plasmid (Figure 1A). In this manner, it resembles BrnQ, a high-capacity threonine carrier whose inactivation, based on our data, renders cells threonine-resistant (Khozov et al., 2023). Hence, we presumed that the inactivation of the identified suppressors exerts a similar effect. Indeed, a comparison of the phenotype of the parental B1895 and the B2818 strains lacking all eight suppressors revealed that the latter had significantly improved fitness under toxic threonine concentrations, thereby confirming the reduction of membrane permeability for threonine due to the introduced mutations (Figure 1D).
The observed phenotype of cells carrying the suppressors in the multicopy vector strongly suggests that the products sdaC, proP, alaE, yqeG, yhjE, ychE, ydgI, and yjeM can facilitate threonine translocation under the tested conditions. In addition, the reduced fitness compared with that of the corresponding parent of the strains lacking a chromosomal copy of the suppressor supports the hypothesis that yhjE and sdaC are involved in threonine uptake. Finally, the sensitivity of the sdaC-overexpressing strain to threonine indicates that SdaC can serve as a high-capacity threonine-specific permease. To verify this, we evaluated the threonine uptake in the threonine-prototrophic ∆sstT ∆tdcBCDE::neo ∆thrP ΔbrnQ ∆livKHMGF mutant B2394 overexpressing the suppressors due to the presence of the corresponding plasmids. In agreement with the phenotypic tests, the strains overexpressing sdaC, yhjE, and ydgI exhibited an approximately two-fold increase in activity (Figure 1C). Amplification of ychE, yjeM, alaE, proP, and yqeG did not noticeably affect activity, indicating that their effect on the growth of the threonine uptake-defective strain may occur via an indirect mechanism. However, this phenomenon is most likely due to the requirement of only a slight increase in transport activity to restore growth of the B1895 strain, which is barely detectable via direct activity measurements.
Next, we determined whether chromosomal suppressor mutations capable of restoring the growth defects caused by the inactivation of transport systems can be selected. We plated the B1950 strain lacking sstT, tdcC, thrP, brnQ, livKHMGF, yjeM, sdaC, ychE, and ydgI on M9 plates with 0.4 mM threonine. Under these conditions, B1950 showed no detectable growth, thereby allowing the selection of mutants that regained the ability to utilize exogenous threonine. Genome resequencing of seven independent mutants revealed three distinct sites of the causative mutations (Table 3). The first site is the gene encoding MarC, a protein with unknown function but is associated with isobutanol- and isoprenol-tolerant phenotypes (Minty et al., 2011; Babel and Krömer, 2020). Interestingly, MarC shared some homology with YchE (Figure 2C). We identified four amino acid substitutions, S140SS, V145E, L10Q, and G11E, which improved the growth of the B1950 strain. The second site was cycA, in which V226A and C110S substitutions were detected. CycA is a well-known H+-dependent symporter that translocates glycine and L-alanine as well as D-alanine, D-serine, β-alanine, and an antibiotic D-cycloserine (Robbins and Oxender, 1973; Schneider et al., 2004; Baisa et al., 2013). Recent data indicate that L-valine and α-aminobutyrate are also substrates of CycA (Hook et al., 2022). CycA shares homology with the known threonine/serine carrier ThrP (Figure 2D). The last site was lrp, which encodes a global transcription regulator that controls more than 10% of E. coli genes, including those involved in amino acid uptake (Tani et al., 2002).
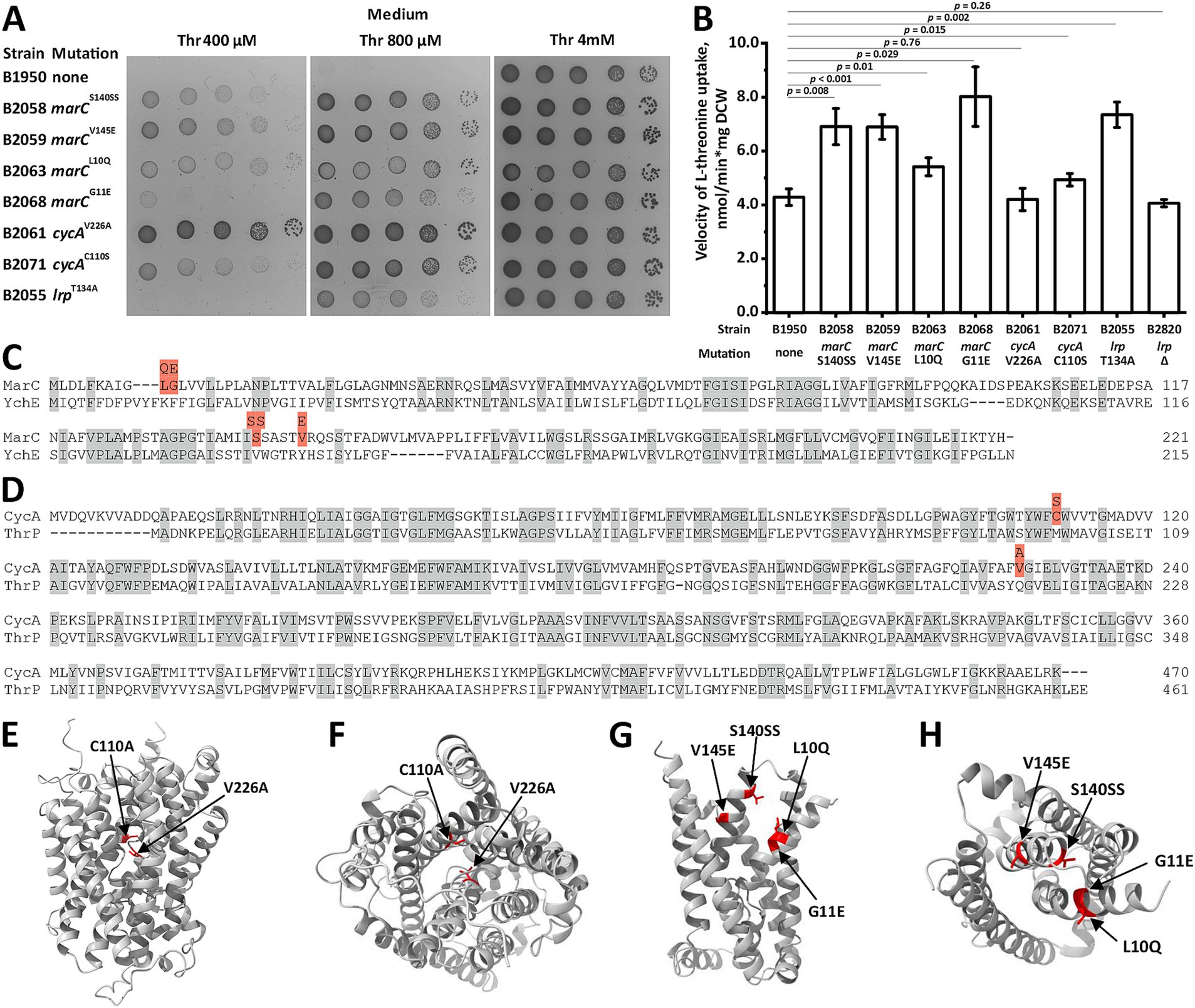
Figure 2. Screening and analysis of chromosomal mutations that suppress threonine uptake defect. Threonine transport activity was measured as described under “Threonine uptake assay” in the Materials and Methods. “Thr” indicated L-threonine. (A) Phenotypic assay for the B1950 threonine-auxotrophic strain and its derivatives harboring mutations in marC, cycA, and lrp loci on minimal plates with varying threonine concentrations. (B) Comparison of threonine transport activity in the B1950 strain its derivatives harboring mutations in marC, cycA, and lrp loci. Measurements were performed using 2.5 mM L-threonine, and the reaction time was 1 min. The values shown are the average of three independent biological replicates. Error bars indicate standard deviation. p-values were calculated using two-tailed Student’s t-test with unequal variances. (C,D) MarC-YchE and CycA-ThrP alignments performed using the Clustal Omega algorithm (Sievers and Higgins, 2014) and UniProt web service (Bateman et al., 2025). The gray and red fillings indicate identical amino acid residues and the amino acid substitutions detected in the mutant derivatives of B1950 that can grow at a non-permissive threonine concentration, respectively. (E–H) Ternary structures of CycA and MarC predicted by AlphaFold2 (Jumper et al., 2021). The red filling indicates mutated residues. (E) CycA, side view. (F) CycA, top view. (G) MarC, side view. (H) MarC, top view.
Expectedly, all mutants grew faster at restrictive threonine concentrations (0.4 and 0.8 mM) compared with the parental B1950 strain (Figure 2A). The mutations marCV145E, marCS140SS, marCL10Q, and cycAC110S provided similar fitness advantages, whereas marCG11E and lrpT134A stimulated growth to a lesser extent. The B2061 strain harboring cycAV226A grew significantly faster than the other mutant strains. When the threonine content in the medium was increased to 4 mM, all mutants and the parental B1950 strain exhibited similar fitness. Direct measurement of threonine uptake revealed that all mutant strains, except B2061, which carried a cycAV226A substitution, had significantly increased threonine uptake activity at 2.5 mM (Figure 2B). This inconsistency may be due to the effect of the V226A substitution on the KM of CycA toward threonine rather than the Vmax of translocation. In this case, at a substrate concentration close to saturation, no alteration in activity should be observed.
Analysis of the MarC and CycA crystal structures predicted by AlphaFold 2 (Jumper et al., 2021) revealed that the mutations occupied a compact region within both proteins and were located on the surface of the transmembrane helices that constitute the inner interface of the molecules (Figures 2E–H). Hence, the selected amino acid substitutions may affect the substrate binding and specificity of MarC and CycA, thereby allowing them to translocate threonine at a velocity sufficient to support cell growth at threonine concentrations that are non-permissive for the parental strain.
Next, we investigated the mechanism by which the T134A substitution in Lrp confers a fitness advantage to threonine uptake-defective strains. First, we compared the transport activity of the B2055 strain (B1950 lrpT134A) with that of its isogenic derivative, B2820, which carries the ∆lrp allele. The strain lacking Lrp did not exhibit increased transport activity, whereas B2055 did (Figure 2B). Thus, T134A is not a loss-of-function mutation. Next, we directly assessed the function of LrpT134A. In our preliminary study, we found that the activity of the yhjE promoter was strictly dependent on Lrp. This allowed us to estimate Lrp activity using a transcriptional fusion of the yhjE promoter and luxCDABE operon of Photorhabdus luminescens in the stains carrying the lrpwt, lrpT134A, or ∆lrp alleles (Figure 3B). The results indicate that LrpT134A activates yhjE transcription to a slightly lower extent than wild-type Lrp. At the same time, lrpT134A confers a phenotype distinct from that of the ∆lrp mutation, whose introduction led to a two-fold reduction in yhjE promoter activity compared with that in the B1950 strain carrying the wild-type lrp allele. Hence, this experiment indicates that consistent with the activity measurement results, lrpT134A confers the synthesis of active but altered Lrp. Interestingly, the same mutation has been reported to cause a defect in the activation of the papBA operon encoding pyelonephritis-associated pili and shows normal activation of the ilvIH operon (Kaltenbach et al., 1998).

Figure 3. Analysis of the lrpT134A mutation phenotype. The threonine uptake measurement and in vivo luminescence assays were performed as described under “Threonine uptake assay” and “Measurement of in vivo luminescence and bacterial growth,” respectively, in the Materials and Method. “Thr” indicated L-threonine. (A) Phenotypic assay for the B2055 threonine-auxotrophic strain carrying the lrpT134A allele and its derivatives harboring yhjE, proP, alaE, and yqeG deletions on minimal plates with varying threonine concentrations. (B) Quantification of yhjE promoter activity via bioluminescence measurement in strains carrying PyhjE-luxCDABE transcriptional fusion and either the lrpwt, lrpT134A, or ∆lrp allele. The values shown are the average of three independent biological replicates. Horizontal error bars and filled area indicate standard deviation for OD600 and RLU/OD600, respectively. (C) Evaluation of AlaE activity in the B2055 strain carrying the lrpT134A allele. Measurements were performed using 800 μM L-threonine, and the reaction time was 1 min. The values shown are the average of three independent biological replicates. Error bars indicate standard deviation. p-values were calculated using two-tailed Student’s t-test with unequal variances. (D) Comparison of the growth fitness of B1895 and its derivatives overexpressing either AlaE or SdaC on minimal plates supplemented with toxic threonine concentrations.
Based on this reasoning and the fact that Lrp controls the expression of various proteins involved in amino acid metabolism and transport processes, we presumed that the mutation led to increased synthesis of a transport system specific to threonine. To identify the permease, we inactivated yhjE, proP, alaE, and yqeG in B2055 cells (Figure 3A). The disruption of yhjE and proP had no additional defect on the growth of B2055. Meanwhile, YhjE exhibited a prominent phenotype in the genetic background of the B1895 strain carrying the wild-type lrp allele. Consistent with our assumption, the results indicate that unlike B1895, B2055 possesses a primary threonine permease other than YhjE while YhjE makes a minor contribution to total threonine uptake. Indeed, the subsequent disruption of alaE led to a drastic reduction in the fitness at limiting threonine concentrations (0.8–1.6 mM). Moreover, unlike B2055 which exhibited sensitivity to threonine at 66 mM, B2873 lacking alaE exhibited only slightly reduced fitness under such conditions compared with the parental B1950 strain. We also found that similar to sdaC, alaE amplified on the multicopy vector conferred the B1895 strain carrying the wild-type lrp allele with sensitivity to threonine (Figure 3D), further confirming that alaE exhibits transport activity specific toward threonine and is expressed, to some extent, even in the background of the lrpwt allele. Finally, we directly measured transport activity using cells of the B2055 strain carrying the pBR_alaE plasmid and found that it had significantly increased activity compared with those of the same strain with the empty vector (Figure 3C), whereas a similar experiment with the B2394 strain carrying lrpwt allele did not reveal AlaE activity (Figure 1C). Taken together, these results prove that lrpT134A leads to impaired or altered Lrp function, which in turn causes increased alaE synthesis and suppression of threonine uptake defects in the B1950 strain.
The main purpose of this study was to identify the permeases that mediate threonine uptake in strains lacking the main threonine-specific transport systems SstT, TdcC, ThrP, LIV-I, and BrnQ. Screening for multicopy and chromosomal suppressors identified 10 membrane protein-encoding genes, namely yhjE, yjeM, ydgI, ychE, marC, yqeG, sdaC, alaE, cycA, and proP, whose amplification on the multicopy vector or specific mutation restored the growth of the threonine uptake-defective strain at non-permissive threonine concentrations. We provided additional evidence for threonine-specific uptake activity for six of the 10 genes. Specifically, the inactivation of yhjE and sdaC confers further impairment to the fitness of the corresponding mutants compared with the parental strains at restrictive threonine concentrations. This indicates that YhjE and SdaC serve as major threonine-specific carriers in the sstT tdcC thrP livKHMGF brnQ strain. Next, the strains overexpressing yhjE, sdaC, ydgI, and alaE (the last in combination with the lrpT134A allele) or carrying the mutant marC and cycA alleles exhibited significantly increased threonine uptake activity, as revealed by in vitro experiments. Finally, sdaC and alaE overexpression rendered cells threonine-sensitive, suggesting a substantial increase in total threonine intake from the medium. We did not provide such arguments for yjeM, ychE, yqeG and proP. Thus, these genes may affect the phenotype of the threonine uptake-deficient strain via an indirect mechanism. However, taking into account the observed phenotype of these genes, membrane localization of corresponding proteins, and the phylogenetic relationship between YjeM and YdgI, YchE and MarC, YqeG and SdaC/TdcC, and ProP and YhjE (Figure 4), as well as the intrinsic substrate promiscuity of ProP, we tended to state the threonine-specific transport activity of these proteins, which, nevertheless, are undetectable under our measurement conditions.

Figure 4. Phylogenic tree of the transport systems involved in threonine uptake. The tree with the highest log likelihood (−13131.09) is shown. The tree is drawn to scale, with branch lengths measured in the number of substitutions per site. All positions with less than 50% site coverage were eliminated. The indicated phylogenetic affiliations of transporters were determined according to the Transporter Classification Database (Saier et al., 2014).
All the identified membrane proteins had a minor contribution, if any, to the threonine-specific transport activity in wild-type cells that express the dedicated transport systems SstT, TdcC, ThrP and LIV-I. Therefore, threonine uptake is unlikely to be a function of the carriers relevant to normal E. coli physiology. What knowledge can be gained from the results? Notably, the functions of the yhjE, yjeM, ydgI, ychE, yqeG, and marC genes are unclear. Thus, the obtained data can provide insight into the roles of the listed genes and their products. At this point, we can approach the understanding of YhjE closely. Recently, the ΔyhjE mutant has been reported to be defective in the formation of the bo3 terminal oxidase (Khalfaoui-Hassani et al., 2023). Based on this observation, the authors considered the involvement of YhjE in Cu2+ and Fe2+ ion transport. Experimental testing of this hypothesis showed that neither of these processes was impaired by yhjE inactivation. Therefore, the mechanisms underlying this phenotype remain elusive. In a distinct study, mutation of yhjE was reported to render cells resistant to D-valine (Maeda et al., 2020), suggesting that YhjE serves as the main entry point for this compound. It is worth noting, BrnQ and LIV-I, two carriers specific for branched-chain amino acids, can translocate threonine (Khozov et al., 2023). Together with our results, these indicate that the most probable substrates of YhjE are L-isoleucine, L-valine, or L-leucine. Moreover, previous studies have shown that E. coli possesses one or more transport systems unrelated to BrnQ and LIV-I (Iaccarino et al., 1978). YhjE may represent one of them; however, further studies are required to confirm this hypothesis.
In our experiments, we identified two genes, ychE and marC, whose products belong to the 6 TMS Neutral Amino Acid Transporter (NAAT) family (according to the Transporter Classification Database, #2.A.95). Four specific marC mutations and ychE amplification were found to be capable of suppressing threonine uptake deficiency. Previously, MarC was assumed to be involved in multiple drug resistance (Hächler et al., 1991; Cohen et al., 1993; White et al., 1997) based on its proximity to the marRAB operon, which controls the expression of several genes involved in resistance to antibiotics (Keeney et al., 2008; Warner and Levy, 2010), organic solvents (White et al., 1997), oxidative stress, and heavy metals (Alekshun and Levy, 1999). However, a later study reported that inactivation of neither MarC nor YchE affected susceptibility to various antibiotics (McDermott et al., 2008). Different studies have found that non-synonymous single nucleotide polymorphisms and loss-of-function mutations in marC increased the tolerance of E. coli cells to isobutanol (Minty et al., 2011) and isoprenol (Babel and Krömer, 2020). However, considering the regulatory role of the adjacent marRAB operon, whether these phenotypes are related to the function of MarC itself or whether the selected mutations in marC alter marRAB expression remains unclear. SnatA, another member of the NAAT Family from the hyperthermophilic archaeon Thermococcus sp. KS-1 which, when overexpressed in E. coli, catalyzes glycine translocation (Akahane et al., 2003), and its activity is inhibited by L-threonine, L-serine, L-alanine, and L-cysteine. Although we cannot exclude the possibility that MarC translocates isobutanol and isoprenol, this fact, together with our results, may suggest that the physiologically relevant substrate range of MarC and YchE includes some of these amino acids or structurally similar compounds. Strains whose growth on minimal medium depends on the function of YchE and mutant MarC can further be used for competitive inhibition assays to identify the exact substrates of these transport systems. The same approach could be applied to yjeM, ydgI, and yqeG. A literature search and EcoCyc database examination revealed no clues regarding the function of the corresponding proteins, except that YdgI is similar to the arginine:ornithine antiporter ArcD of Pseudomonas aeruginosa (Bourdineaud et al., 1993; Herzberg et al., 2006). The ability of these genes, when amplified on a plasmid, to mitigate threonine uptake deficiency and the directly measurable in vitro activity of YdgI toward threonine can be used to decipher their physiologically relevant substrates and roles in cellular biology.
The present work extends the substrate range of the SdaC and ProP transport systems. ProP is known to exhibit intrinsic substrate promiscuity and translocates to a wide range of compounds whereas SdaC has been reported to be strictly specific to L-serine and inactive toward threonine (Shao et al., 1994). In contrast to the conclusions of this study, we found that SdaC supports the growth of strains defective in threonine transport at concentrations of 1–3 mM (Figure 4). Moreover, the SdaC-overexpressing strain exhibited threonine sensitivity similar to that of strains expressing the active BrnQ carrier. Thus, SdaC may be a low-affinity and high-capacity threonine-specific permease.
One of the most interesting results was the threonine uptake activity of AlaE. Indeed, we found that alaE is a multicopy suppressor of the threonine growth defects in the B1895 strain. Increased alaE expression was the exact mechanism by which the B2055 strain carrying the lrpT134A allele overcame deficiency in threonine uptake. Next, the wild-type allele on the lrpT134A background and amplification of alaE on the lrpwt background conferred higher sensitivity to threonine in the corresponding strains compared with that of their counterparts lacking the chromosomal alaE copy and pBR-alaE plasmid, respectively. Finally, the threonine-specific activity of AlaE was directly measured in the strain carrying the pBR_alaE plasmid and lrpT134A. Taken together, these results indicate that AlaE can translocate threonine from the external environment to the cytosol. At the same time, there is strong evidence that AlaE is an H+/L-alanine antiporter that operates in the reverse direction and protects cells from toxic L-alanine accumulation (Hori et al., 2011a, 2011b; Kim et al., 2015). These functions are unlikely to be two independent activities in a single transport system. We presume that AlaE is specific to both substrates and can translocate them in both directions, depending on the ratio of their concentrations in the cytosol and external environment, and the magnitude of the electrochemical potential of H+ ions on the cytoplasmic membrane.
Finally, the obtained results can be used in metabolic engineering to construct effective threonine-producing strains. We have shown that the strain lacking yhjE, yjeM, ydgI, ychE, yqeG, sdaC, alaE, and proP as well as sstT, tdcC, thrP, and livKHMGF exhibit significantly increased fitness under toxic threonine concentrations up to 420 mM compared with its counterpart harboring wild-type alleles of the eight multicopy suppressors. Notably, this phenotype resembles that of strains that overexpress specific threonine exporters, such as RhtA or RhtC (Zakataeva et al., 1999; Livshits et al., 2003), which are commonly used to confer threonine resistance and enhance the ability of producing strains to accumulate threonine (Debabov, 2003; Lee et al., 2007; Dong et al., 2011). Therefore, improving fitness by reducing membrane permeability to threonine could serve as a complementary strategy for constructing industrial strains alongside the overexpression of exporters. Furthermore, it is well established that the ability to accumulate threonine inversely correlates with the threonine-specific uptake activity of cells (Kruse et al., 2001, 2002). The underlying mechanism of this phenomenon may involve the prevention of intracellular threonine accumulation and the subsequent inhibition of key biosynthetic enzymes. Additionally, a producer cell that simultaneously expresses both RhtA/RhtC and threonine-specific uptake systems inevitably translocates threonine in both directions via proton motive force-dependent pathways. This creates a futile cycle that dissipates energy and reduces product yield, which can be mitigated by inactivating the uptake transport systems.
The original contributions presented in the study are included in the article/supplementary material, further inquiries can be directed to the corresponding author.
DB: Writing – original draft, Writing – review & editing, Conceptualization, Project administration, Methodology. AK: Writing – original draft, Writing – review & editing, Investigation, Conceptualization. TV: Writing – review & editing, Conceptualization, Investigation. AS: Writing – review & editing, Investigation. SM: Writing – review & editing, Investigation. OM: Writing – review & editing, Investigation. GB: Writing – review & editing, Methodology. MC: Writing – review & editing, Methodology. IS: Investigation, Writing – original draft. AN: Project administration, Writing – review & editing, Methodology. SS: Writing – review & editing, Funding acquisition, Supervision.
The author(s) declare that financial support was received for the research and/or publication of this article. The experiments on phenotypic characterization and construction of bacterial strains were supported by the State Assignment of National Research Center “Kurchatov Institute.” The evaluation of the promoter activity was supported by the grant from the Ministry of Science and Higher Education of the Russian Federation (no. 075–15–2019-1659). In vitro measurement of transport activity was supported by the grant from the Ministry of Science and Higher Education of the Russian Federation (no. 122030200324–1).
This study was conducted using scientific equipment from the Bioresource Center Russian Collection of Industrial Microorganisms (BRC VKPM) and the Department of Radiochemistry (Faculty of Chemistry, Lomonosov Moscow State University).
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The authors declare that no Gen AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Akahane, S., Kamata, H., Yagisawa, H., and Hirata, H. (2003). A novel neutral amino acid transporter from the hyperthermophilic archaeon Thermococcus sp. KS-1. J. Biochem. 133, 173–180. doi: 10.1093/jb/mvg021
Alekshun, M. N., and Levy, S. B. (1999). The mar regulon: multiple resistance to antibiotics and other toxic chemicals. Trends Microbiol. 7, 410–413. doi: 10.1016/S0966-842X(99)01589-9
Babel, H., and Krömer, J. O. (2020). Evolutionary engineering of E. coli MG1655 for tolerance against isoprenol. Biotechnol. Biofuels 13:183. doi: 10.1186/s13068-020-01825-6
Baisa, G., Stabo, N. J., and Welch, R. A. (2013). Characterization of Escherichia coli D-cycloserine transport and resistant mutants. J. Bacteriol. 195, 1389–1399. doi: 10.1128/JB.01598-12
Barrick, J. E., Colburn, G., Deatherage, D. E., Traverse, C. C., Strand, M. D., Borges, J. J., et al. (2014). Identifying structural variation in haploid microbial genomes from short-read resequencing data using breseq. BMC Genomics 15:1039. doi: 10.1186/1471-2164-15-1039
Bateman, A., Martin, M.-J., Orchard, S., Magrane, M., Adesina, A., Ahmad, S., et al. (2025). UniProt: the universal protein knowledgebase in 2025. Nucleic Acids Res. 53, D609–D617. doi: 10.1093/nar/gkae1010
Bolivar, F., Rodriguez, R. L., Greene, P. J., Betlach, M. C., Heyneker, H. L., Boyer, H. W., et al. (1977). Construction and characterization of new cloning vehicles. II. A multipurpose cloning system. Gene 2, 95–113.
Bourdineaud, J. P., Heierli, D., Gamper, M., Verhoogt, H. J., Driessen, A. J., Konings, W. N., et al. (1993). Characterization of the arcD arginine:ornithine exchanger of Pseudomonas aeruginosa. Localization in the cytoplasmic membrane and a topological model. J. Biol. Chem. 268, 5417–5424. doi: 10.1016/S0021-9258(18)53337-8
Cohen, S. P., Hächler, H., and Levy, S. B. (1993). Genetic and functional analysis of the multiple antibiotic resistance (mar) locus in Escherichia coli. J. Bacteriol. 175, 1484–1492. doi: 10.1128/jb.175.5.1484-1492.1993
Csonka, L. N. (1981). The role of proline in osmoregulation in salmonella typhimurium and Escherichia coli. Basic Life Sci. 18, 533–542. doi: 10.1007/978-1-4684-3980-9_32
Datsenko, K. A., and Wanner, B. L. (2000). One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl. Acad. Sci. USA 97, 6640–6645. doi: 10.1073/pnas.120163297
Debabov, V. G. (2003). “The threonine story” in Microbial production of l-amino acids. eds. R. Faurie, J. Thommel, B. Bathe, V. G. Debabov, S. Huebner, and M. Ikeda, et al. (Berlin, Heidelberg: Springer), 113–136.
Dong, X., Quinn, P. J., and Wang, X. (2011). Metabolic engineering of Escherichia coli and Corynebacterium glutamicum for the production of L-threonine. Biotechnol. Adv. 29, 11–23. doi: 10.1016/j.biotechadv.2010.07.009
Doroshenko, V., Airich, L., Vitushkina, M., Kolokolova, A., Livshits, V., and Mashko, S. (2007). YddG from Escherichia coli promotes export of aromatic amino acids. FEMS Microbiol. Lett. 275, 312–318. doi: 10.1111/j.1574-6968.2007.00894.x
Dutta, S., Corsi, I. D., Bier, N., and Koehler, T. M. (2022). BrnQ-type branched-chain amino acid transporters influence Bacillus anthracis growth and virulence. MBio 13:e0364021. doi: 10.1128/mbio.03640-21
Ghatak, S., King, Z. A., Sastry, A., and Palsson, B. O. (2019). The y-ome defines the 35% of Escherichia coli genes that lack experimental evidence of function. Nucleic Acids Res. 47, 2446–2454. doi: 10.1093/nar/gkz030
Hächler, H., Cohen, S. P., and Levy, S. B. (1991). marA, a regulated locus which controls expression of chromosomal multiple antibiotic resistance in Escherichia coli. J. Bacteriol. 173, 5532–5538. doi: 10.1128/jb.173.17.5532-5538.1991
Hama, H., Shimamoto, T., Tsuda, M., and Tsuchiya, T. (1987). Properties of a Na+-coupled serine-threonine transport system in Escherichia coli. Biochim. Biophys Acta 905, 231–239. doi: 10.1016/0005-2736(87)90451-2
Hama, H., Shimamoto, T., Tsuda, M., and Tsuchiya, T. (1988). Characterization of a novel L-serine transport system in Escherichia coli. J. Bacteriol. 170, 2236–2239. doi: 10.1128/jb.170.5.2236-2239.1988
Herzberg, M., Kaye, I. K., Peti, W., and Wood, T. K. (2006). YdgG (TqsA) controls biofilm formation in Escherichia coli K-12 through autoinducer 2 transport. J. Bacteriol. 188, 587–598. doi: 10.1128/JB.188.2.587-598.2006
Hook, C., Eremina, N., Zaytsev, P., Varlamova, D., and Stoynova, N. (2022). The Escherichia coli amino acid uptake protein CycA: regulation of its synthesis and practical application in L-isoleucine production. Microorganisms 10:647. doi: 10.3390/microorganisms10030647
Hori, H., Ando, T., Isogai, E., Yoneyama, H., and Katsumata, R. (2011a). Identification of an L-alanine export system in Escherichia coli and isolation and characterization of export-deficient mutants. FEMS Microbiol. Lett. 316, 83–89. doi: 10.1111/j.1574-6968.2010.02196.x
Hori, H., Yoneyama, H., Tobe, R., Ando, T., Isogai, E., and Katsumata, R. (2011b). Inducible L-alanine exporter encoded by the novel gene ygaW (alaE) in Escherichia coli. Appl. Environ. Microbiol. 77, 4027–4034. doi: 10.1128/AEM.00003-11
Iaccarino, M., Guardiola, J., and De Felice, M. (1978). On the permeability of biological membranes. J. Membr. Sci. 3, 287–302. doi: 10.1016/S0376-7388(00)83028-8
Ihara, K., Kim, S., Ando, T., and Yoneyama, H. (2022). Importance of transmembrane helix 4 of L-alanine exporter AlaE in oligomer formation and substrate export activity in Escherichia coli. Microbiology 168:1147. doi: 10.1099/mic.0.001147
Jebbar, M., Talibart, R., Gloux, K., Bernard, T., and Blanco, C. (1992). Osmoprotection of Escherichia coli by ectoine: uptake and accumulation characteristics. J. Bacteriol. 174, 5027–5035. doi: 10.1128/jb.174.15.5027-5035.1992
Jumper, J., Evans, R., Pritzel, A., Green, T., Figurnov, M., Ronneberger, O., et al. (2021). Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589. doi: 10.1038/s41586-021-03819-2
Kaltenbach, L., Braaten, B., Tucker, J., Krabbe, M., and Low, D. (1998). Use of a two-color genetic screen to identify a domain of the global regulator Lrp that is specifically required for pap phase variation. J. Bacteriol. 180, 1224–1231. doi: 10.1128/JB.180.5.1224-1231.1998
Katsube, S., Ando, T., and Yoneyama, H. (2019). L-alanine exporter, AlaE, of Escherichia coli functions as a safety valve to enhance survival under feast conditions. Int. J. Mol. Sci. 20:4942. doi: 10.3390/ijms20194942
Keeney, D., Ruzin, A., McAleese, F., Murphy, E., and Bradford, P. A. (2008). MarA-mediated overexpression of the AcrAB efflux pump results in decreased susceptibility to tigecycline in Escherichia coli. J. Antimicrob. Chemother. 61, 46–53. doi: 10.1093/jac/dkm397
Keseler, I. M., Mackie, A., Santos-Zavaleta, A., Billington, R., Bonavides-Martinez, C., Caspi, R., et al. (2017). The EcoCyc database: reflecting new knowledge about Escherichia coli K-12. Nucleic Acids Res. 45, D543–D550. doi: 10.1093/nar/gkw1003
Khalfaoui-Hassani, B., Blaby-Haas, C. E., Verissimo, A., and Daldal, F. (2023). The Escherichia coli MFS-type transporter genes yhjE, ydiM, and yfcJ are required to produce an active bo3 quinol oxidase. PLoS One 18:e0293015. doi: 10.1371/journal.pone.0293015
Khozov, A. A., Bubnov, D. M., Plisov, E. D., Vybornaya, T. V., Yuzbashev, T. V., Agrimi, G., et al. (2023). A study on L-threonine and L-serine uptake in Escherichia coli K-12. Front. Microbiol. 14:1151716. doi: 10.3389/fmicb.2023.1151716
Kim, S., Ihara, K., Katsube, S., Hori, H., Ando, T., Isogai, E., et al. (2015). Characterization of the L-alanine exporter AlaE of Escherichia coli and its potential role in protecting cells from a toxic-level accumulation of L-alanine and its derivatives. Microbiology 4, 632–643. doi: 10.1002/mbo3.269
Kruse, D., Krämer, R., Eggeling, L., Rieping, M., Pfefferle, W., Tchieu, J. H., et al. (2002). Influence of threonine exporters on threonine production in Escherichia coli. Appl. Microbiol. Biotechnol. 59, 205–210. doi: 10.1007/s00253-002-0987-7
Kruse, D., Six, S., Krämer, R., and Burkovski, A. (2001). Analysis of threonine uptake in Escherichia coli threonine production strains. Biotechnol. Lett. 23, 401–404. doi: 10.1023/A:1005652211609
Lee, K. H., Park, J. H., Kim, T. Y., Kim, H. U., and Lee, S. Y. (2007). Systems metabolic engineering of Escherichia coli for L-threonine production. Mol. Syst. Biol. 3:149. doi: 10.1038/msb4100196
Livshits, V. A., Zakataeva, N. P., Aleshin, V. V., and Vitushkina, M. V. (2003). Identification and characterization of the new gene rhtA involved in threonine and homoserine efflux in Escherichia coli. Res. Microbiol. 154, 123–135. doi: 10.1016/S0923-2508(03)00036-6
MacMillan, S. V., Alexander, D. A., Culham, D. E., Kunte, H. J., Marshall, E. V., Rochon, D., et al. (1999). The ion coupling and organic substrate specificities of osmoregulatory transporter ProP in Escherichia coli. Biochim. Biophys. Acta 1420, 30–44. doi: 10.1016/S0005-2736(99)00085-1
Maeda, T., Iwasawa, J., Kotani, H., Sakata, N., Kawada, M., Horinouchi, T., et al. (2020). High-throughput laboratory evolution reveals evolutionary constraints in Escherichia coli. Nat. Commun. 11:5970. doi: 10.1038/s41467-020-19713-w
McDermott, P. F., McMurry, L. M., Podglajen, I., Dzink-Fox, J. L., Schneiders, T., Draper, M. P., et al. (2008). The marC gene of Escherichia coli is not involved in multiple antibiotic resistance. Antimicrob. Agents Chemother. 52, 382–383. doi: 10.1128/AAC.00930-07
McLaggan, D., and Epstein, W. (1991). Escherichia coli accumulates the eukaryotic osmolyte taurine at high osmolarity. FEMS Microbiol. Lett. 65, 209–213. doi: 10.1016/0378-1097(91)90304-S
Minty, J. J., Lesnefsky, A. A., Lin, F., Chen, Y., Zaroff, T. A., Veloso, A. B., et al. (2011). Evolution combined with genomic study elucidates genetic bases of isobutanol tolerance in Escherichia coli. Microb. Cell Factories 10:18. doi: 10.1186/1475-2859-10-18
Mundhada, H., Schneider, K., Christensen, H. B., and Nielsen, A. T. (2016). Engineering of high yield production of L-serine in Escherichia coli. Biotechnol. Bioeng. 113, 807–816. doi: 10.1002/bit.25844
Ogawa, W., Kayahara, T., Tsuda, M., Mizushima, T., and Tsuchiya, T. (1997). Isolation and characterization of an Escherichia coli mutant lacking the major serine transporter, and cloning of a serine transporter gene. J. Biochem. 122, 1241–1245. doi: 10.1093/oxfordjournals.jbchem.a021887
Okamoto, K., Kino, K., and Ikeda, M. (1997). Hyperproduction of L-threonine by an Escherichia coli mutant with impaired L-threonine uptake. Biosci. Biotechnol. Biochem. 61, 1877–1882. doi: 10.1271/bbb.61.1877
Palace, S. G., Proulx, M. K., Lu, S., Baker, R. E., and Goguen, J. D. (2014). Genome-wide mutant fitness profiling identifies nutritional requirements for optimal growth of Yersinia pestis in deep tissue. MBio 5:e01385-14. doi: 10.1128/mBio.01385-14
Park, J. H., Lee, K. H., Kim, T. Y., and Lee, S. Y. (2007). Metabolic engineering of Escherichia coli for the production of L-valine based on transcriptome analysis and in silico gene knockout simulation. Proc. Natl. Acad. Sci. USA 104, 7797–7802. doi: 10.1073/pnas.0702609104
Pettersen, E. F., Goddard, T. D., Huang, C. C., Meng, E. C., Couch, G. S., Croll, T. I., et al. (2021). UCSF CHIMERAX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82. doi: 10.1002/pro.3943
Robbins, J. C., and Oxender, D. L. (1973). Transport systems for alanine, serine, and glycine in Escherichia coli K-12. J. Bacteriol. 116, 12–18. doi: 10.1128/jb.116.1.12-18.1973
Saier, M. H., Reddy, V. S., Tamang, D. G., and Västermark, A. (2014). The transporter classification database. Nucleic Acids Res. 42, D251–D258. doi: 10.1093/nar/gkt1097
Sambrook, J., Fritsch, E. F., and Maniatis, T. (1989). Molecular cloning: a laboratory manual. Cold Spring Harbor, New York: Cold Spring Harbor Laboratory Press.
Schneider, F., Krämer, R., and Burkovski, A. (2004). Identification and characterization of the main beta-alanine uptake system in Escherichia coli. Appl. Microbiol. Biotechnol. 65, 576–582. doi: 10.1007/s00253-004-1636-0
Shao, Z., Lin, R. T., and Newman, E. B. (1994). Sequencing and characterization of the sdaC gene and identification of the sdaCB operon in Escherichia coli K-12. Eur. J. Biochem. 222, 901–907. doi: 10.1111/j.1432-1033.1994.tb18938.x
Sievers, F., and Higgins, D. G. (2014). Clustal omega, accurate alignment of very large numbers of sequences. Methods Mol. Biol. 1079, 105–116. doi: 10.1007/978-1-62703-646-7_6
Stalmach, M. E., Grothe, S., and Wood, J. M. (1983). Two proline porters in Escherichia coli K-12. J. Bacteriol. 156, 481–486. doi: 10.1128/jb.156.2.481-486.1983
Sumantran, V. N., Schweizer, H. P., and Datta, P. (1990). A novel membrane-associated threonine permease encoded by the tdcC gene of Escherichia coli. J. Bacteriol. 172, 4288–4294. doi: 10.1128/jb.172.8.4288-4294.1990
Tamura, K., Stecher, G., and Kumar, S. (2021). MEGA11: molecular evolutionary genetics analysis version 11. Mol. Biol. Evol. 38, 3022–3027. doi: 10.1093/molbev/msab120
Tani, T. H., Khodursky, A., Blumenthal, R. M., Brown, P. O., and Matthews, R. G. (2002). Adaptation to famine: a family of stationary-phase genes revealed by microarray analysis. Proc. Natl. Acad. Sci. 99, 13471–13476. doi: 10.1073/pnas.212510999
Thomason, L. C., Costantino, N., and Court, D. L. (2007). E. coli genome manipulation by P1 transduction. Curr. Protoc. Mol. Biol. 1, 1.17.1–1.17.8. doi: 10.1002/0471142727.mb0117s79
Van Dyk, T. K., and Rosson, R. A. (1998). “Photorhabdus luminescens luxCDABE promoter probe vectors” in Bioluminescence methods and protocols. ed. R. A. LaRossa (Totowa, NJ: Humana Press), 85–95.
Varadi, M., Anyango, S., Deshpande, M., Nair, S., Natassia, C., Yordanova, G., et al. (2022). AlphaFold protein structure database: massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Res. 50, D439–D444. doi: 10.1093/nar/gkab1061
Verheul, A., Wouters, J. A., Rombouts, F. M., and Abee, T. (1998). A possible role of ProP, ProU and CaiT in osmoprotection of Escherichia coli by carnitine. J. Appl. Microbiol. 85, 1036–1046. doi: 10.1111/j.1365-2672.1998.tb05269.x
Warner, D. M., and Levy, S. B. (2010). Different effects of transcriptional regulators MarA, SoxS and rob on susceptibility of Escherichia coli to cationic antimicrobial peptides (CAMPs): rob-dependent CAMP induction of the marRAB operon. Microbiology 156, 570–578. doi: 10.1099/mic.0.033415-0
Whelan, S., and Goldman, N. (2001). A general empirical model of protein evolution derived from multiple protein families using a maximum-likelihood approach. Mol. Biol. Evol. 18, 691–699. doi: 10.1093/oxfordjournals.molbev.a003851
White, D. G., Goldman, J. D., Demple, B., and Levy, S. B. (1997). Role of the acrAB locus in organic solvent tolerance mediated by expression of marA, soxS, or robA in Escherichia coli. J. Bacteriol. 179, 6122–6126. doi: 10.1128/jb.179.19.6122-6126.1997
Keywords: Escherichia coli, L-threonine uptake, transmembrane transport, amino acid transporter, membrane proteins
Citation: Bubnov DM, Khozov AA, Vybornaya TV, Stepanova AA, Molev SV, Melkina OE, Badun GA, Chernysheva MG, Skob IA, Netrusov AI and Sineoky SP (2025) Multiple routes for non-physiological L-threonine uptake in Escherichia coli K-12. Front. Microbiol. 16:1579813. doi: 10.3389/fmicb.2025.1579813
Received: 20 February 2025; Accepted: 18 March 2025;
Published: 03 April 2025.
Edited by:
Phillip E. Klebba, Kansas State University, United StatesReviewed by:
Michael Benedik, Texas A&M University, United StatesCopyright © 2025 Bubnov, Khozov, Vybornaya, Stepanova, Molev, Melkina, Badun, Chernysheva, Skob, Netrusov and Sineoky. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Dmitrii M. Bubnov, YnVibm92LmRtaXRyaWlAbWFpbC5ydQ==
†These authors have contributed equally to this work and share first authorship
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.