- 1State Key Laboratory for Biology of Plant Diseases and Insect Pests, Institute of Plant Protection, Chinese Academy of Agricultural Sciences, Beijing, China
- 2Western Agricultural Research Center, Chinese Academy of Agricultural Sciences, Changji, China
Introduction: Aphids are often infected with diverse bacterial symbionts that enhance their ecological adaptation. While geographic factors significantly influence aphid bacterial communities, research on environmental effects on the cotton aphid Aphis gossypii Glover feeding on cotton plants across China's major cotton-growing regions is limited.
Methods: This study examined the influence of geographic factors on the endosymbiotic bacterial community and diversity of A. gossypii by analyzing 58 field samples from 24 locations across China's major cotton-growing regions (2021–2022) using 16S rRNA (V3–V4) high-throughput sequencing.
Results and discussion: Our results demonstrate that geography is an important factor in shaping the endosymbiotic bacterial composition and diversity of A. gossypii. Among China's three major cotton-growing regions, the Yangtze River Basin exhibited the highest bacterial diversity, followed by the Northwestern Inland Region, and then the Yellow River Basin. Acinetobacter, Lactobacillus, Serratia, and Aeromonas were more abundant in the Yangtze River Basin, with positive correlations observed for Acinetobacter, Serratia, and Aeromonas in relation to annual precipitation. In contrast, Candidatus Uzinura, dominant in southern Xinjiang, displayed negative correlations with precipitation and longitude but a positive correlation with altitude, and this report is the first detection of it in A. gossypii. Buchnera was ubiquitous and negatively associated with both precipitation and temperature, while Arsenophonus showed no significant environmental correlations. These findings highlight the distinct influences of geographic factors on A. gossypii endosymbiotic communities across China's major cotton-growing regions, broadening our understanding of aphid–endosymbiont–environment interactions and offering potential avenues for biocontrol strategies.
1 Introduction
The cotton aphid (Aphis gossypii Glover; Hemiptera: Aphididae) is a major global agricultural pest, infesting over 600 plant species and causing substantial economic losses in cotton production through direct feeding and virus transmission (Carletto et al., 2009; Liu et al., 2017; Wu and Guo, 2005). In China, cotton remains one of the most economically important crops, primarily grown in three major regions: the Northwest Inland, the Yellow River Basin, and the Yangtze River Basin. Among these, Xinjiang has emerged as a leading cotton-producing region in China (Feng et al., 2022; Wu and Guo, 2005).
Aphids commonly harbor diverse symbiotic bacteria that play a critical role in their survival, fitness, and ecological adaptability. While, the composition and diversity of symbiotic bacterial communities in aphids are influenced by multiple factors, including genetics, geography, and host plant species (Baumann, 2005; Łukasik et al., 2013; Oliver et al., 2003; Russell and Moran, 2006; Guo et al., 2017; Oliver et al., 2010; Tougeron and Iltis, 2022; Li et al., 2021; Oliver et al., 2005). The symbiotic bacterial communities are quite different among the different aphid species; however, most studies on aphid–bacteria interactions have focused on the pea aphid (Acyrthosiphon pisum). In A. pisum, host plants play a critical role in shaping bacterial communities, and several previous studies have highlighted significant differences in bacterial community composition and prevalence among populations on different host plants (Ferrari et al., 2012; Gauthier et al., 2015). Geography also exerts a strong influence on these symbiotic associations. For instance, the bacterial community structure of A.pisum was strongly influenced by geography (Zhang et al., 2021). Distinct distribution patterns of secondary symbionts—particularly Regiella insecticola—were observed in Japanese A. pisum populations (Tsuchida et al., 2002). Beyond pea aphids, the grain aphid Sitobion avenae harbors symbiotic bacteria with clear geographic distribution patterns, including a strong positive correlation between Regiella insecticola infection and environmental parameters such as mean annual temperature and precipitation (Hu et al., 2020). Another study showed that symbiotic bacteria in four cereal aphid species and A. pisum in two agroclimatic zones of Chile varied across aphid species, host plants, and geographic regions (Sepúlveda et al., 2017). Moreover, the bacterial communities associated with wheat aphid, Sitobion miscanthi, in China's major wheat-growing region affected pesticide resistance and were modulated by environmental factors (Wang et al., 2022).
Aphis gossypii, like other aphids, harbors diverse endosymbiotic bacteria that enhance its ecological adaptation. Its bacterial communities are mostly influenced by host plants and geography. For instance, the symbiont diversity in Japanese and Australian populations of A. gossypii was primarily shaped by geography rather than host plant (Najar-Rodríguez et al., 2009). Moreover, another study revealed that consistent bacterial community structures within provinces, but found significant inter-province variation in northern China, indicating strong geographical effect (Zhao et al., 2016). In contrast, host plant was a more significant determinant of symbiont composition than geography in several A. gossypii populations (Xu et al., 2020). These studies emphasize the complex interplay of factors driving microbial symbiont associations in A. gossypii. Despite evidence indicating that geographic factors can shape the bacterial communities of A. gossypii, previous research has been based on limited sampling across diverse host plants. Consequently, there has been a comprehensive study of A. gossypii populations exclusively associated with cotton in China's three main cotton-producing regions. Here, we performed 16S rRNA (V3–V4) high-throughput sequencing on 58 cotton aphid field samples from 24 cotton fields distributed over these three regions, demonstrating distinct geographic influences on A. gossypii symbiotic bacterial communities. These findings expand our understanding of aphid–symbiont–environment interactions and offer new prospects for developing biocontrol strategies.
2 Materials and methods
2.1 Sampling of Aphis gossypii from different geographical regions
To investigate the diversity of symbiotic microbial communities in different geographical populations of A. gossypii, a total of 58 field samples were collected in 2021 and 2022 from 24 different geographic locations across three major cotton-growing regions in China, including the Northwest Inland Cotton Region (Northwest), the Yellow River Basin Cotton Region (Yellow) and the Yangtze River Basin Cotton Region (Yangtze) (Figure 1 and Table 1 for details).
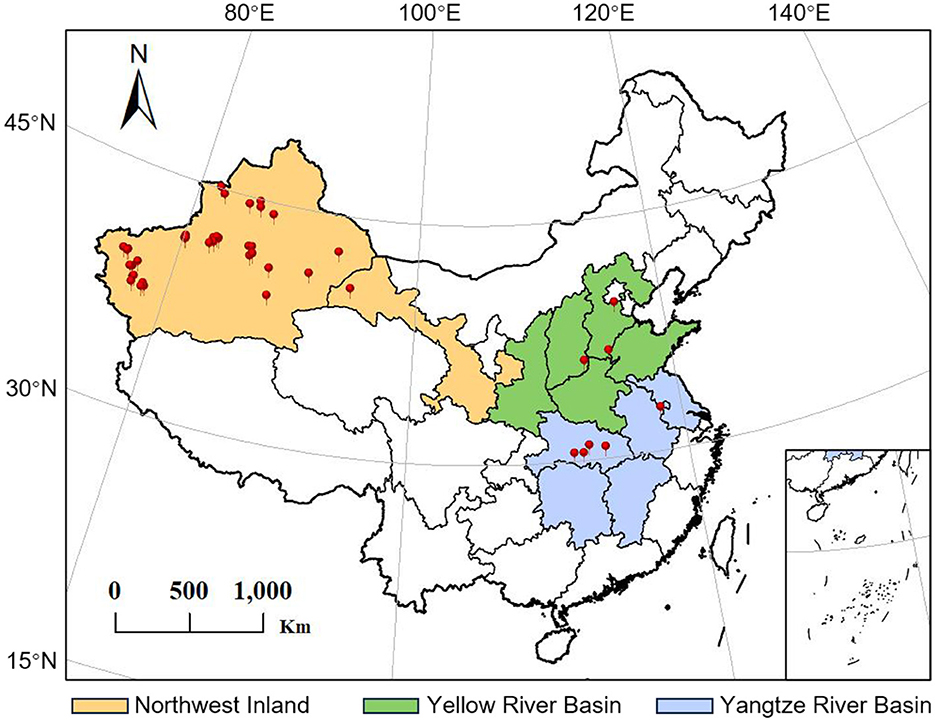
Figure 1. Collections sites of A. gossypii from three major cotton-growing regions in China. Detailed information on the specific locations and collection dates is provided in Table 1.
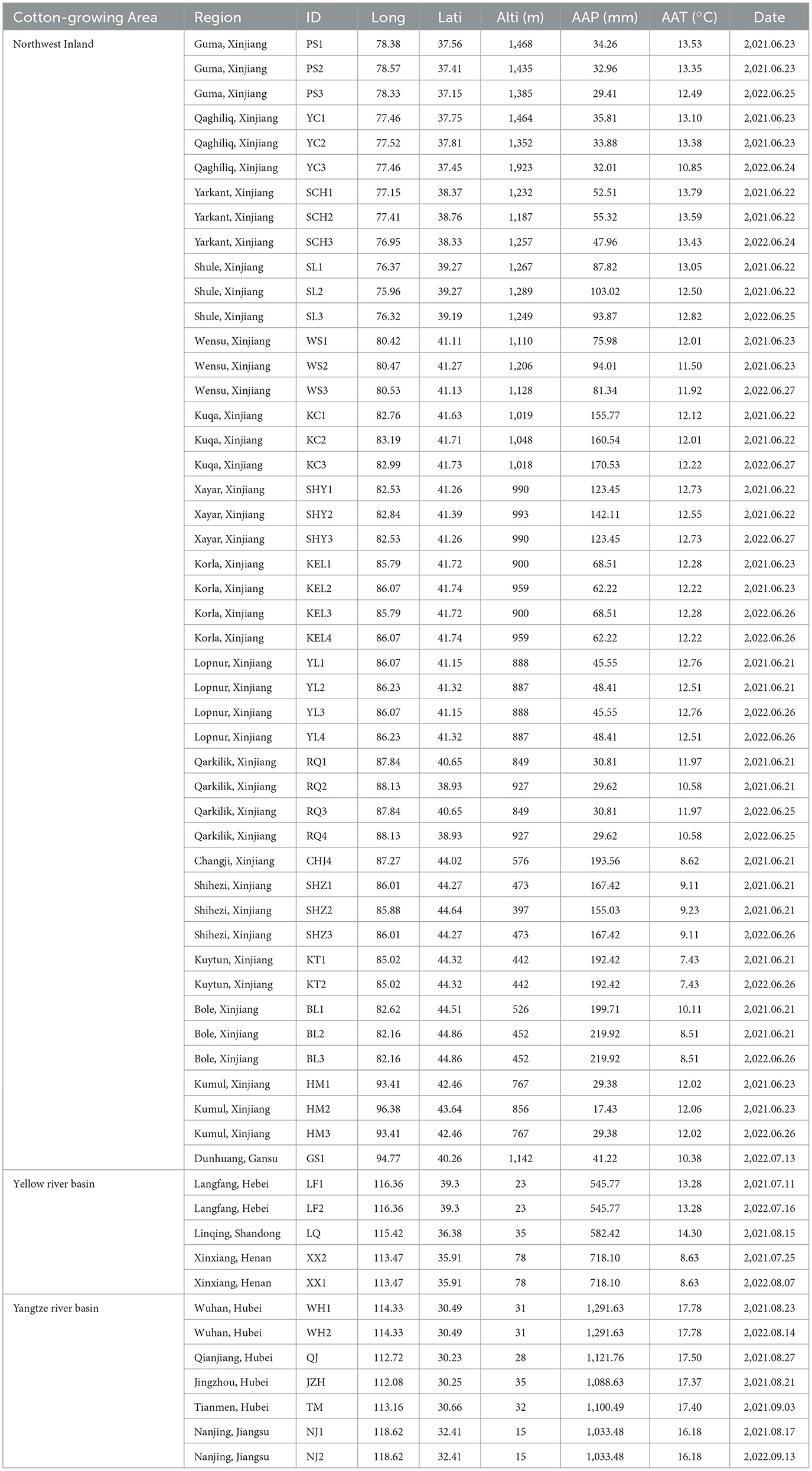
Table 1. Sampling information for A. gossypii collected from three major cotton-growing regions in China.
In each location, at least one cotton field that had been continuously cultivated with cotton for several years was selected (if a second field was included in the same location, there were > 10 km between fields). Samples of A. gossypii were collected from each cotton field using a five-point sampling method, ensuring that the distance between each sampling point was >10 m, and only one cotton plant was selected from each sampling point. We collected 30–50 wingless adult aphids from each sampling point. Samples from the same field were considered as one sample, from which 10 individual aphids were randomly selected from each sampling point, resulting in 50 wingless aphids as the group to be examined. These 50 aphids were placed in a 1.5 mL EP tube containing 1 mL of absolute ethanol and stored at −20°C until examined.
2.2 DNA isolation, 16s rRNA gene PCR amplification and high-throughput sequencing
Total genomic DNA was extracted from 58 samples of A. gossypii samples (each of 50 aphids) collected from 24 geographical locations. Before DNA extraction, we discarded the ethanol from the sample tubes, and triple-rinsed the aphids with sterile ultrapure water (3 min per rinse). Genomic DNA was extracted from the group of 50 aphids to using the NucleoSpin 96 Tissue DNA kit (Genetic Biotechnology International Trading Co., Shanghai, China) following the manufacturer's protocol. The quality of extracted DNA was assessed by 1.5% agarose gel electrophoresis, while DNA concentration and purity were quantified using a NanoDrop 2000 UV spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA). DNA samples were subsequently diluted to 100 ng/μL with TE buffer and stored at −20°C.
The V3-V4 hypervariable region of the 16S rRNA gene was amplified from the A. gossypii genomic DNA using the universal bacterial primers 338F (5′-ACTCCTACGGGAGGCAGCA-3′) and 806R (5′-GGACTACHVGGGTWTCTAAT-3′) (Yu et al., 2005). Each PCR reaction (20 μL total volume) contained 40–50 ng of template DNA, 0.3 μL of each primer (10 μM), 5 μL of KOD FX Neo buffer, 2 μL of dNTPs (2 mM each), 0.2 μL of KOD FX Neo polymerase, and nuclease-free water to make up the volume to 20 μL. The thermal cycling conditions were as follows: initial denaturation at 95°C for 5 min, 30 cycles of denaturation at 95°C for 30 s, annealing at 50°C for 30 s, and extension at 72°C for 40 s, followed by a final extension at 72°C for 7 min. The completed reaction products were held at 4°C. The PCR products were purified using the Omega DNA Purification Kit (Omega Bio-tek, Norcross, GA, USA) and quantified using a Qsep-400 DNA analyzer (BiOptic, New Taipei City). The purified amplicons were then used for small-fragment library construction. Paired-end sequencing (2 × 250 bp) was performed on an Illumina NovaSeq 6000 platform (Beijing Biomics Technology Co., Ltd., Beijing, China).
2.3 Bioinformatic analysis
The paired-end reads were assembled using FLASH (version 1.2.11) (Magoč and Salzberg, 2011) with a minimum overlap of 10 bp and a maximum mismatch ratio of 20%, producing raw tags. These raw tags were then utilized for operational taxonomic unit (OTU) clustering. Tags with an average quality score below 20 or a length shorter than 75% of their original size were filtered using Trimmomatic (version 0.33) (Bolger et al., 2014). Primer sequences were identified and removed with Cutadapt (version 1.8.3) (Kechin et al., 2017). Chimeric sequences were removed using UCHIME (Edgar et al., 2011) by comparing query fragments to a reference database, producing clean tags. The clean tags were clustered into OTUs at a 97% similarity threshold using USEARCH (version 10.0) (Edgar, 2013) and OTUs representing < 0.005% of the total sequence count were filtered out (Bokulich et al., 2013). Taxonomic annotation of the filtered OTUs was carried out using QIIME2 (Bolyen et al., 2019), applying a Naive Bayes classifier trained on the SILVA database (version 138.1) (Quast et al., 2013) at a confidence threshold of 70%. Both alpha diversity and beta diversity metrics were calculated using QIIME2, and visualized using the ggplot2 package (Ginestet, 2011) Redundancy analysis (RDA) was performed in R using vegan package (Oksanen et al., 2018) to evaluate environmental factors influencing microbial community composition. All analyses were conducted on the BMKCloud platform.
2.4 Environmental factor data collection
Th0065 sampling locations varied in altitude, precipitation, temperature, and latitude/longitude. In this study, the historical monthly precipitation and temperature data (2010–2021) were downloaded from the WorldClim website (https://worldclim.org/data/monthlywth.html). Latitude and altitude were recorded during sampling using a mobile phone's built-in compass app. Precipitation and temperature data were extracted from the WorldClim dataset (Harris et al., 2020) in ArcMap 10.8 based on the recorded coordinates.
2.5 Statistical analysis
For the downstream statistical analysis, samples were categorized into three geographical groups: Northwest (46 samples from 16 locations), Yellow (5 samples from 3 locations), and Yangtze (7 samples from 5 locations).
Alpha diversity indices (Simpson, Shannon, Chao1, and ACE) were calculated to assess bacterial community diversity and richness across the three geographical groups using the R package vegan. Pairwise inter-group significance was calculated using the non-parametric Wilcoxon test (P < 0.05), and the results were visualized using the R package ggplot2. Principal coordinate analysis (PCoA) based on the Weighted UniFrac distances matrix at the OTU level was conducted using the R package GuniFrac (Chen et al., 2012), and the results were visualized with the R package ggplot2. To further assess the significance of differences in bacterial communities across the geographical regions, PERMANOVA and ANOSIM analysis values at a 95% confidence level were calculated using the anoism and adonis functions of in the vegan package.
The influence of environmental factors (precipitation, temperature, altitude, and geographic coordinates) on microbial community composition were assessed using redundancy analysis (RDA) and Mantel tests at the genus level using the vegan package. Pearson correlation analysis was conducted to explore the relationships between environmental factors and the relative abundances of the 11 common endosymbionts. Significant correlations were identified using a correlation coefficient threshold of 0.5 and a significance level of P < 0.05. Correlation heatmaps were generated using the pheatmap package in R.
3 Results
3.1 Bacterial communities in Aphis gossypii from three different geographical regions
High-throughput sequencing of 58 samples generated a total of 4,398,796 paired-end reads. After quality control and assembly, 4,335,095 clean reads were obtained, with a minimum of 42,118 clean reads and an average of 74,743 clean reads per sample. These reads were clustered into 42,757 OTUs and classified into 51 phyla, 130 classes, 403 orders, 886 families, 2,168 genera, and 3,084 species.
At the phylum level, Proteobacteria was the dominant one across all regions, though its relative abundance varied substantially: 89.7% in the Northwest Inland, 95.7% in the Yellow River Basin, and 42.8% in the Yangtze River Basin. Firmicutes was the second most abundant phylum in the Yangtze region (14.2%), while its abundance was notably lower in the Northwest Inland (3.8%) and Yellow River Basin (2.1%) regions. In addition, the phyla Bacteroidota and Actinobacteriota exhibited substantially higher abundance in the Yangtze region, at 8.5 and 6.6%, respectively, compared to 3.5% and 0.8% in the Northwest, and 0.6% and 0.4%. in the Yellow River Basin (Supplementary Table 1).
At the genus level, the primary symbiont Buchnera was detected alongside other commonly encountered endosymbiotic genera, including Arsenophonus, Candidatus Hamiltonella, Wolbachia, Rickettsia, Serratia, Spiroplasma, Candidatus Uzinura, Acinetobacter, Aeromonas, and Lactobacillus (Figure 2E; Table 2). Buchnera predominated in all three regions, reaching its highest relative abundance in the Yellow River Basin (93.4%), followed by the Northwest Inland (89.0%) and the Yangtze River Basin (59.7%). Arsenophonus showed moderate levels in the Northwest Inland (6.3%) and the Yellow River Basin (6.1%), but declined to 2.7% in the Yangtze River Basin. Candidatus Uzinura was generally low, detected at 2.4% in the Northwest Inland but almost absent in the Yellow River Basin (0.01%) and the Yangtze River Basin (0.06%). By contrast, Aeromonas, Lactobacillus, and Acinetobacter reached higher abundances in the Yangtze River Basin (17.6, 12.0, and 6.4%, respectively), while remaining markedly lower in the Northwest Inland (0.06, 0.65, and 1.56%), and effectively undetected in the Yellow River Basin except for Lactobacillus (0.39%) and Acinetobacter (0.06%). Serratia and Rickettsia occurred at low levels across all regions, with Serratia slightly more abundant in the Yangtze River Basin (0.84%) compared to the Northwest Inland (0.01%) and the Yellow River Basin (0.004%). Rickettsia similarly peaked in the Yangtze River Basin (0.35%) relative to the Northwest Inland (0.01%) and the Yellow River Basin (0.07%). Other genera, including Candidatus Hamiltonella, Wolbachia, and Spiroplasma, were consistently present at very low abundances. Candidatus Hamiltonella remained at 0.01% in both the Northwest Inland and Yellow River Basin, increasing to 0.14% in the Yangtze River Basin. Wolbachia followed a similar trend, peaking at 0.13% in the Yangtze River Basin compared to 0.02% in the Northwest Inland and 0.01% in the Yellow River Basin. Spiroplasma appeared only in trace amounts (0.03% in the Northwest Inland and 0.02% in the Yangtze River Basin) and was absent in the Yellow River Basin.
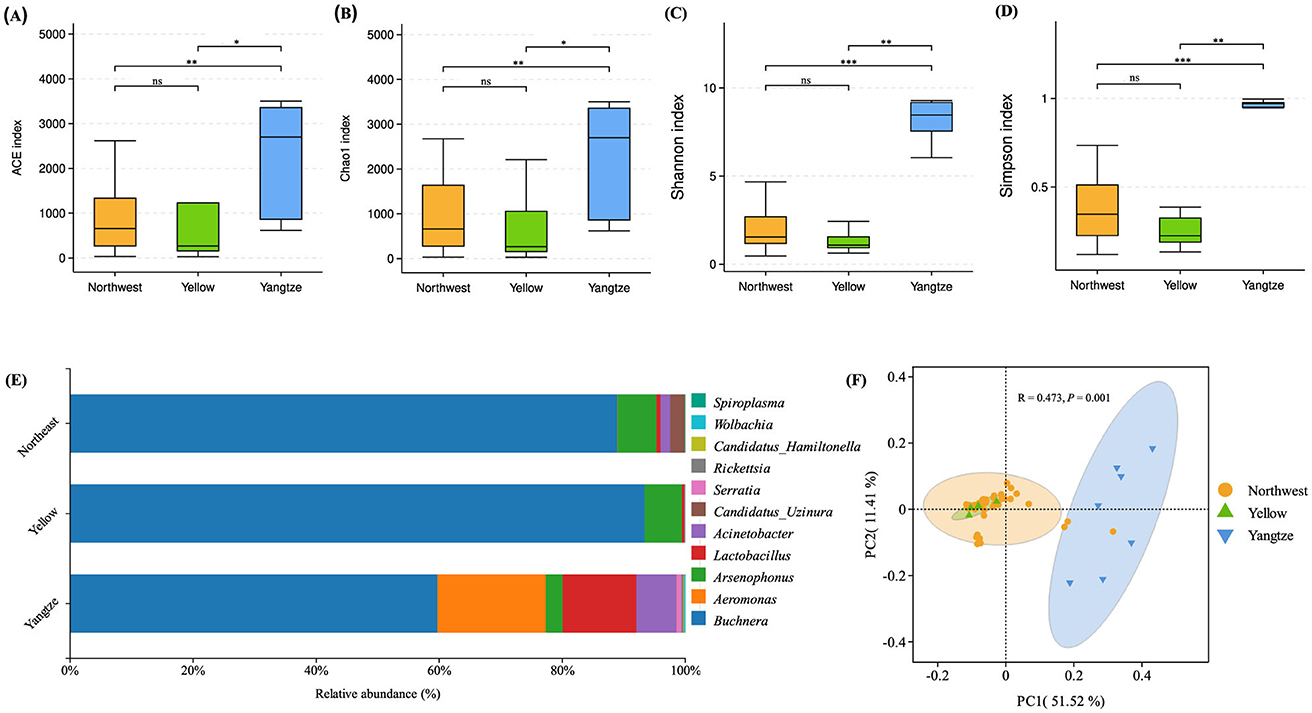
Figure 2. Bacterial diversity in A. gossypii across cotton-growing regions (Northwest Inland, Yellow River Basin, and Yangtze River Basin) analyzed via 16S rRNA sequencing. Bacterial community richness and diversity were evaluated using the ACE (A), Chao1 (B), Shannon (C), and Simpson (D) indices. Significant differences among groups were determined using the Wilcoxon test at the OTU level. The relative abundances of symbiotic bacteria at the genus level are depicted in panel (E), while principal component analysis (PCoA) based on Weighted UniFrac distances at the OTU level is presented in panel (F). Significance levels: *P < 0.05, **0.001 < P < 0.05, ***P < 0.001. Not significant (ns): P ≥ 0.05
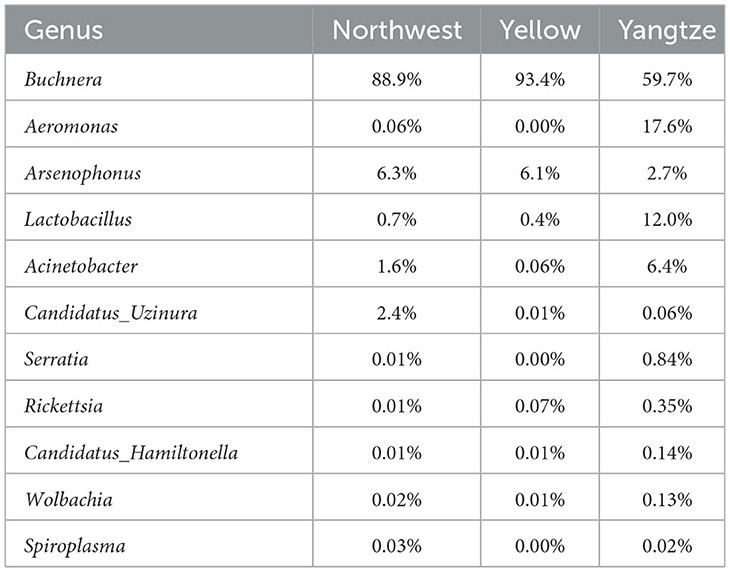
Table 2. Relative abundances of symbionts in A. gossypii from three major cotton-growing regions in China.
3.2 Bacteria diversity in Aphis gossypii from three different geographical regions
When we assessed alpha diversity indices (ACE, Chao1, Simpson, and Shannon) and phylogenetic diversity (PD) across the three regions, we found significant variation (Wilcoxon test, P < 0.05) (Figures 2A–D; Table 3). Samples from the Yangtze region exhibited notably higher diversity in all metrics (PD: mean = 217.77, SE = 49.91, Shannon: mean = 8.26, SE = 0.43, Simpson: mean = 0.97, SE = 0.01, ACE: mean = 2,167.72, SE = 490.47, Chao1: mean = 2,166.14, SE = 489.91). In contrast, the Northwest Inland region showed moderate diversity (PD: mean = 109.47, SE = 9.39, Shannon: mean = 2.60, SE = 0.27, Simpson: mean = 0.39, SE = 0.03, ACE: mean = 808.53, SE = 95.48, Chao1: mean = 898.05, SE = 111.18), while the Yellow River Basin exhibited the lowest overall values (PD: mean = 74.76, SE = 28.88, Shannon: mean = 1.29, SE = 0.30, Simpson: mean = 0.25, SE = 0.04, ACE: mean = 813.68, SE = 531.56, Chao1: mean = 674.92, SE = 397.78). Statistical analysis using the Wilcoxon test confirmed significant differences in alpha diversity indices among the regions, with the Yangtze region showing highly significant differences in indices compared to both the Northwest Inland (Simpson: P < 0.001; Shannon: P < 0.001; ACE: P < 0.01; Chao1: P < 0.01) and Yellow River Basin (Simpson: P < 0.01; Shannon: P < 0.01; ACE: P < 0.05; Chao1: P < 0.05). In contrast, no significant differences were found in the four alpha diversity indices between the Northwest and Yellow River regions (P > 0.05 for all indices).
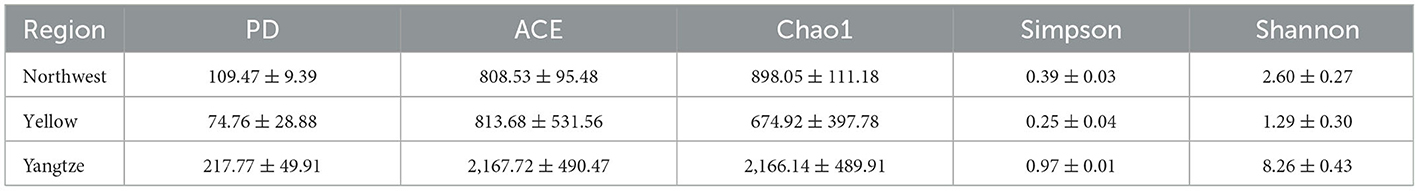
Table 3. The alpha diversity indices (mean, SE) (ACE, Chao1, Simpson, and Shannon) and phylogenetic diversity (PD) of microbial symbionts of A. gossypii from three major cotton-growing regions in China.
Beta diversity analysis using Weighted UniFrac-based principal coordinate analysis (PCoA) revealed distinct clustering of bacterial communities among the three geographic regions (Figure 2F). Notably, samples from the Yangtze region formed a clearly separate cluster, reflecting significant differences in community structure relative to those from the Yellow River Basin or Northwest Inland (PERMANOVA: R2 = 0.166, P = 0.001; ANOSIM: R = 0.321, P = 0.009).
These results indicate that the Yangtze region has greater richness and abundance in the symbiotic bacterial community of A. gossypii than the other two regions.
3.3 The effect of different environmental conditions on structuring of symbiont communities in Aphis gossypii
Environmental factors significantly influenced the distribution and composition of symbiotic bacterial communities in A. gossypii across from the three geographical regions (Northwest Inland, Yangtze River Basin, and Yellow River Basin). Mantel test results (Table 4) identified annual average precipitation (AAP, r = 0.7113, P = 0.001) as the most significant environmental factor, followed by latitude (Lati, r = 0.6733, P = 0.001), annual average temperature (AAT, r = 0.5821, P = 0.001), longitude (Long, r = 0.4393, P = 0.001), and altitude (Alti, r = 0.3164, P = 0.001).
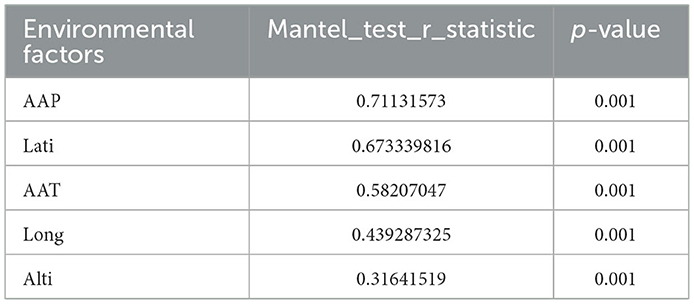
Table 4. Mantel test results between environmental factors and bacterial communities in A. gossypii from three major cotton-growing regions in China.
These findings indicate that the bacterial communities associated with A. gossypii are significantly influenced by geographic factors. The redundancy analysis (RDA) results (Figure 3) further confirmed that environmental conditions play a important role in shaping cotton aphid‘s symbiotic bacterial communities, emphasizing the complex interplay between ecological variables and microbial diversity across different geographic contexts.
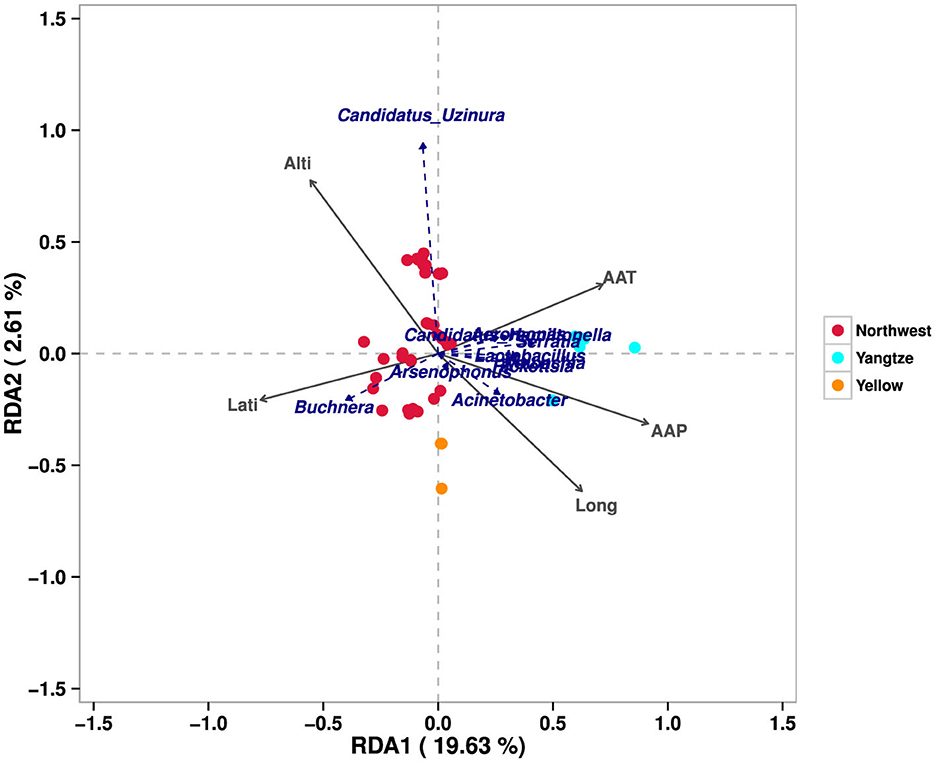
Figure 3. Redundancy analysis (RDA) bacterial communities and environmental factors. This figure illustrates the relationship between symbiotic bacterial communities in A. gossypii and various environmental factors across three major cotton-growing regions in China. Each colored point represents populations of A. gossypii from distinct geographical regions: red represents the Northwest Inland, cyan denotes the Yangtze River Basin, and orange corresponds to the Yellow River Basin. The blue text highlights the different symbiotic bacterial taxa identified within these populations. Additionally, the arrows signify the environmental factors; an angle of <90° between two arrows indicates a positive correlation between the corresponding factors.
Additionally, the heatmap based on Pearson correlation coefficients (threshold = 0.1, significance P < 0.05) corroborated the associations identified in the Mantel test and RDA. Notably, Candidatus Uzinura exhibited significant negative correlations with longitude (r < −0.6, P < 0.001) and AAP (r < −0.5, P < 0.01), while showing a strong positive correlation with altitude (r > 0.6, P < 0.001). Conversely, Serratia displayed a significant negative correlation with AAP (r < −0.6, P < 0.01), suggesting its lower occurrence in regions with high precipitation. Furthermore, Acinetobacter demonstrated positive correlations with both AAP (r > 0.4, P < 0.01) and AAT (r > 0.4, P < 0.01) (Figure 4). These correlations enhance our understanding of how specific environmental factors shape the composition of symbiotic bacterial communities in A. gossypii.
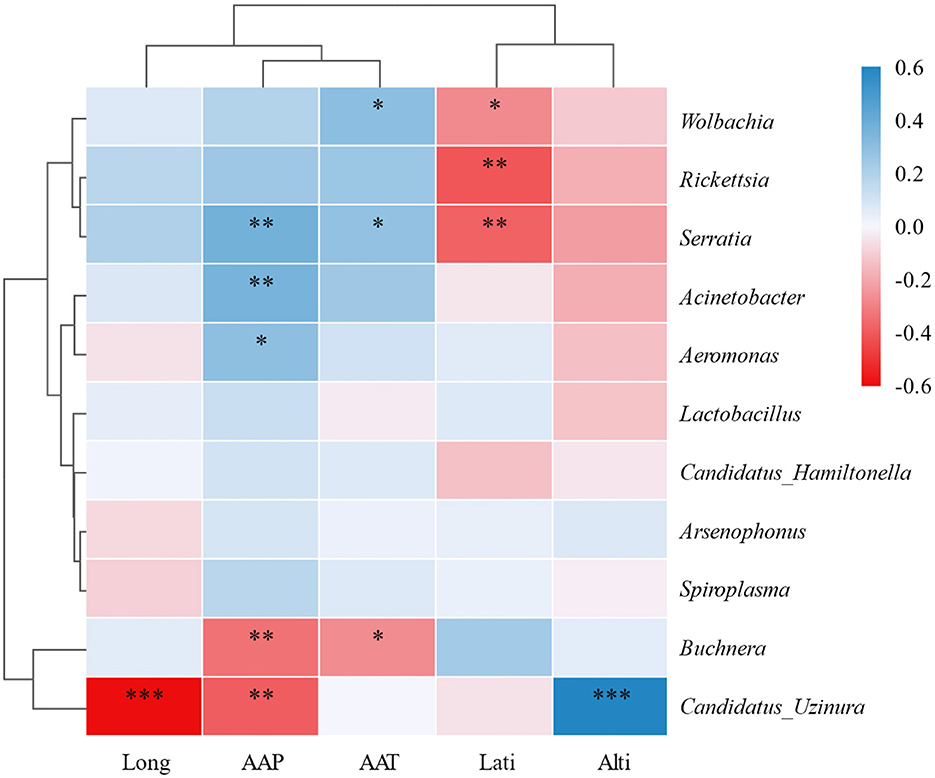
Figure 4. Heatmap of correlations between symbiotic bacteria in Aphis gossypii and environmental factors. Rows represent environmental factors, and columns represent symbiotic bacteria. Color intensity reflects correlation strength and direction: darker blue indicates stronger negative correlations, darker red indicates stronger positive correlations. Significance levels: *P < 0.05, **0.001 < P < 0.05, ***P < 0.001. Pearson correlation was used with a coefficient threshold of 0.1 and significance set at P < 0.05.
4 Discussion
We analyzed 58 field samples from 24 locations across China‘s three major cotton-growing regions during 2021–2022 using 16S rRNA (V3–V4) sequencing. Our results indicate that geographic factors, such as precipitation, temperature, altitude, longitude, and latitude, significantly influenced the diversity and composition of bacterial communities in A. gossypii. Previous studies have similarly highlighted the important role of environmental factors in shaping aphid endosymbiont communities in various geographic regions (Najar-Rodríguez et al., 2009; Zhao et al., 2016; Martins et al., 2023; Sepúlveda et al., 2017; Tsuchida et al., 2002). In contrast, some research suggests that host plants may have a stronger influence than geography on symbiotic bacteria composition in A. gossypii populations (Xu et al., 2020). However, because all A. gossypii samples in our study were collected exclusively from cotton plants, host plant variability likely had minimal impact on our results. Their (Xu et al., 2020) study sampled A. gossypii from 25 different host plant families, which may account for the differing conclusions regarding host plant influence on symbiont composition. Moreover, another study indicated that geographic factors had a greater effect on the diversity of symbiotic bacteria in A. gossypii than host plants (Najar-Rodríguez et al., 2009). These studies highlight the complex interplay among geography, host plants, and the community structure of symbiotic bacteria in A. gossypii, warranting further investigation. Our results contribute to this discussion and highlight the importance of understanding these dynamics to elucidate the factors driving microbial community structures in significant agricultural pests.
In this study, we detected Buchnera, an obligate endosymbiont of aphids, in all samples (Supplementary Figure 1), confirming its crucial role in aphid physiology, particularly in amino acid synthesis and nutrient provision (Brinza et al., 2009; Douglas, 1998). Heatmap analysis revealed a negative correlation between Buchnera abundance and precipitation and temperature (P < 0.05). Despite its lower relative abundance in the Yangtze River Basin (59.7%) compared to the Northwest inland (88.9%) and Yellow River Basin (93.4%), it significantly surpassed other symbionts (Figure 3; Table 2). Arsenophonus, which was consistently identified as the predominant facultative endosymbiont in A. gossypii, showed the highest infection prevalence and relative abundance among the symbiotic bacteria analyzed, aligning with previous studies (Xu et al., 2020; Zhang et al., 2021; Zhao et al., 2016), while Arsenophonus exhibited no significant correlation with environmental factors. Furthermore, a recent study indicated a significant correlation between gossypol concentrations and Arsenophonus abundance, suggesting that this symbiont confers adaptive advantages to A. gossypii in gossypol-rich environments (Chang et al., 2022). Given that all A. gossypii samples in our research were sourced from cotton plants, known for their high gossypol content, this likely accounts for the observed high prevalence and relative abundance of Arsenophonus in our results.
Moreover, Wolbachia was positively correlated with temperature and precipitation but was negatively associated with latitude. A recent study reported positive correlations between Wolbachia and both precipitation and temperature (Yang et al., 2023). Rickettsia also showed a negative correlation with latitude, while Serratia was negatively correlated with latitude and had a weak but positive association with temperature and precipitation. In contrast, Acinetobacter and Aeromonas were positively correlated with precipitation and weakly but negatively correlated with latitude (Figure 3). In contrast, Aeromonas, a facultative symbiont commonly associated with aquatic or semi-aquatic insects (Dubey et al., 2022), was predominantly enriched in the Yangtze River Basin, a humid region characterized by high rainfall. In contrast, Candidatus Uzinura exhibited a significant positive correlation with altitude and negative correlations with both precipitation and longitude. It was predominantly detected in cotton aphids from southern Xinjiang (Guma, Qaghiliq, Yarkant, Shule, Wensu, Kuqa, and Qarkilik), where its relative abundance consistently exceeded 2% (Supplementary Figure 2). This region features fruit-cotton intercropping systems, such as jujube-cotton cultivation, which create favorable microhabitats for armored scale insects, significant agricultural pests of fruit and nut trees (Wang et al., 2021; Wei et al., 2016). As a primary endosymbiont of armored scales, Candidatus Uzinura plays a critical role in nutritional supplementation (Gruwell et al., 2012, 2007). These systems likely facilitate the symbiont's distribution and proliferation, leading to selective pressures that promote horizontal transmission to other arthropods, including cotton aphids. While direct evidence of Candidatus Uzinura in A. gossypii, or its horizontal transfer from armored scale insects, is lacking, horizontal symbiont transmission among arthropods has been documented (Chrostek et al., 2017; Gonella et al., 2015). Furthermore, this microbe‘s detection in southern Xinjiang, near the arid Taklamakan Desert, aligns with findings of this endosymbiont in insects from extremely dry environments (Hakobyan et al., 2023). These observations suggest that Candidatus Uzinura may confer drought resilience to its host insects. Further research is essential to elucidate its role in host survival, ecological significance, and mechanisms of horizontal transmission across arthropod species.
We investigated the endosymbiotic bacterial communities and diversity of Aphis gossypii across China's three major cotton-growing regions. Our findings indicate that geographic factors exert the most significant influence on the composition and diversity of the symbiont community in A. gossypii. Additionally, this study highlights the critical relationships between symbiont communities and environmental factors—including precipitation, temperature, altitude, longitude, and latitude—that may have been underestimated in prior research. Furthermore, this study broaden our understanding of aphid–endosymbiont–environment interactions and offering potential avenues for biocontrol strategies.
Data availability statement
The raw sequence data reported in this paper have been deposited in the Genome Sequence Archive (Chen et al., 2021) at the National Genomics Data Center (CNCB-NGDC Members and Partners., 2022), China National Center for Bioinformation / Beijing Institute of Genomics, Chinese Academy of Sciences (GSA: CRA024049) and are publicly accessible at https://ngdc.cncb.ac.cn/gsa/, project number PRJCA037416.
Author contributions
AA: Formal analysis, Investigation, Writing – original draft. YG: Investigation, Writing – original draft. JL: Investigation, Writing – original draft. YL: Conceptualization, Data curation, Funding acquisition, Methodology, Project administration, Resources, Software, Supervision, Validation, Visualization, Writing – review & editing.
Funding
The author(s) declare that financial support was received for the research and/or publication of this article. This study was financially supported by the National Key R&D Program of China (2022YFD1400300) and China Agriculture Research System (CARS-15-19).
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Generative AI statement
The author(s) declare that no Gen AI was used in the creation of this manuscript.
Publisher's note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Supplementary material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fmicb.2025.1569543/full#supplementary-material
References
Baumann, P. (2005). Biology bacteriocyte-associated endosymbionts of plant sap-sucking insects. Annu. Rev. Microbiol. 59, 155–189. doi: 10.1146/annurev.micro.59.030804.121041
Bokulich, N. A., Subramanian, S., Faith, J. J., Gevers, D., Gordon, J. I., Knight, R., et al. (2013). Quality-filtering vastly improves diversity estimates from Illumina amplicon sequencing. Nat. Methods 10, 57–59. doi: 10.1038/nmeth.2276
Bolger, A. M., Lohse, M., and Usadel, B. (2014). Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120. doi: 10.1093/bioinformatics/btu170
Bolyen, E., Rideout, J. R., Dillon, M. R., Bokulich, N. A., Abnet, C. C., Al-Ghalith, G. A., et al. (2019). Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857. doi: 10.1038/s41587-019-0209-9
Brinza, L., Viñuelas, J., Cottret, L., Calevro, F., Rahb,é, Y., Febvay, G., et al. (2009). Systemic analysis of the symbiotic function of Buchnera aphidicola, the primary endosymbiont of the pea aphid Acyrthosiphon pisum. C. R. Biol. 332, 1034–1049. doi: 10.1016/j.crvi.2009.09.007
Carletto, J., Lombaert, E., Chavigny, P., Brévault, T., Lapchin, L., and Vanlerberghe-Masutti, F. (2009). Ecological specialization of the aphid Aphis gossypii Glover on cultivated host plants. Mol. Ecol. 18, 2198–2212. doi: 10.1111/j.1365-294X.2009.04190.x
Chang, C. Y., Sun, X. W., Tian, P. P., Miao, N. H., Zhang, Y. L., and Liu, X. D. (2022). Plant secondary metabolite and temperature determine the prevalence of Arsenophonus endosymbionts in aphid populations. Environ. Microbiol. 24, 3764–3776. doi: 10.1111/1462-2920.15929
Chen, J., Bittinger, K., Charlson, E. S., Hoffmann, C., Lewis, J., Wu, G. D., et al. (2012). Associating microbiome composition with environmental covariates using generalized UniFrac distances. Bioinformatics 28, 2106–2113. doi: 10.1093/bioinformatics/bts342
Chrostek, E., Pelz-Stelinski, K., Hurst, G. D. D., and Hughes, G. L. (2017). Horizontal transmission of intracellular insect symbionts via plants. Front. Microbiol. 8:2237. doi: 10.3389/fmicb.2017.02237
Douglas, A. E. (1998). Nutritional interactions in insect-microbial symbioses: aphids and their symbiotic bacteria Buchnera. Annu. Rev. Entomol. 43, 17–37. doi: 10.1146/annurev.ento.43.1.17
Dubey, S., Ager-Wick, E., Kumar, J., Karunasagar, I., Karunasagar, I., Peng, B., et al. (2022). Aeromonas species isolated from aquatic organisms, insects, chicken, and humans in India show similar antimicrobial resistance profiles. Front. Microbiol. 13:1008870. doi: 10.3389/fmicb.2022.1008870
Edgar, R. C. (2013). UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat. Methods 10, 996–998. doi: 10.1038/nmeth.2604
Edgar, R. C., Haas, B. J., Clemente, J. C., Quince, C., and Knight, R. (2011). UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27, 2194–2200. doi: 10.1093/bioinformatics/btr381
Feng, L., Chi, B.-J., and Dong, H.-Z. (2022). Cotton cultivation technology with Chinese characteristics has driven the 70-year development of cotton production in China. J. Integr. Agric. 21, 597–609. doi: 10.1016/S2095-3119(20)63457-8
Ferrari, J., West, J. A., Via, S., and Godfray, H. C. (2012). Population genetic structure and secondary symbionts in host-associated populations of the pea aphid complex. Evolution 66, 375–390. doi: 10.1111/j.1558-5646.2011.01436.x
Gauthier, J. P., Outreman, Y., Mieuzet, L., and Simon, J. C. (2015). Bacterial communities associated with host-adapted populations of pea aphids revealed by deep sequencing of 16S ribosomal DNA. PLoS ONE 10:e0120664. doi: 10.1371/journal.pone.0120664
Ginestet, C. (2011). ggplot2: elegant graphics for data analysis. J. R. Stat. Soc. Ser. A. Stat. Soc. 174, 245–246. doi: 10.1111/j.1467-985X.2010.00676_9.x
Gonella, E., Pajoro, M., Marzorati, M., Crotti, E., Mandrioli, M., Pontini, M., et al. (2015). Plant-mediated interspecific horizontal transmission of an intracellular symbiont in insects. Sci. Rep. 5:15811. doi: 10.1038/srep15811
Gruwell, M. E., Flarhety, M., and Dittmar, K. (2012). Distribution of the primary endosymbiont (candidatus Uzinura diaspidicola) within host insects from the scale insect family Diaspididae. Insects 3, 262–269. doi: 10.3390/insects3010262
Gruwell, M. E., Morse, G. E., and Normark, B. B. (2007). Phylogenetic congruence of armored scale insects (Hemiptera: Diaspididae) and their primary endosymbionts from the phylum bacteroidetes. Mol. Phylogenet. Evol. 44, 267–280. doi: 10.1016/j.ympev.2007.01.014
Guo, J., Hatt, S., He, K., Chen, J., Francis, F., and Wang, Z. (2017). Nine facultative endosymbionts in aphids. A review. J. Asia-Pac. Entomol. 20, 794–801. doi: 10.1016/j.aspen.2017.03.025
Hakobyan, A., Velte, S., Sickel, W., Quandt, D., Stoll, A., and Knief, C. (2023). Tillandsia landbeckii phyllosphere and laimosphere as refugia for bacterial life in a hyperarid desert environment. Microbiome 11:246. doi: 10.1186/s40168-023-01684-x
Harris, I., Osborn, T. J., Jones, P., and Lister, D. (2020). Version 4 of the CRU TS monthly high-resolution gridded multivariate climate dataset. Sci. Data 7:109. doi: 10.1038/s41597-020-0453-3
Hu, Z., Su, D., Li, D., Tong, Z., Zhang, C., Zhang, G., et al. (2020). Diversity of secondary endosymbionts among different geographical populations of the grain aphid, Sitobion avenae (Fabricius) (Hemiptera: Aphididae) in China. Entomol. Gen. 40, 253–262. doi: 10.1127/entomologia/2020/0875
Kechin, A., Boyarskikh, U., Kel, A., and Filipenko, M. (2017). cutPrimers: a new tool for accurate cutting of primers from reads of targeted next generation sequencing. J. Comput. Biol. 24, 1138–1143. doi: 10.1089/cmb.2017.0096
Li, Q., Sun, J., Qin, Y., Fan, J., Zhang, Y., Tan, X., et al. (2021). Reduced insecticide susceptibility of the wheat aphid Sitobion miscanthi after infection by the secondary bacterial symbiont Hamiltonella defensa. Pest Manag. Sci. 77, 1936–1944. doi: 10.1002/ps.6221
Liu, X. D., Xu, T. T., and Lei, H. X. (2017). Refuges and host shift pathways of host-specialized aphids Aphis gossypii. Sci. Rep. 7:2008. doi: 10.1038/s41598-017-02248-4
Łukasik, P., Guo, H., van Asch, M., Ferrari, J., and Godfray, H. C. (2013). Protection against a fungal pathogen conferred by the aphid facultative endosymbionts Rickettsia and Spiroplasma is expressed in multiple host genotypes and species and is not influenced by co-infection with another symbiont. J. Evol. Biol. 26, 2654–2661. doi: 10.1111/jeb.12260
Magoč, T., and Salzberg, S. L. (2011). FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963. doi: 10.1093/bioinformatics/btr507
Martins, M., César, C. S., and Cogni, R. (2023). The effects of temperature on prevalence of facultative insect heritable symbionts across spatial and seasonal scales. Front. Microbiol. 14:1321341. doi: 10.3389/fmicb.2023.1321341
Najar-Rodríguez, A. J., McGraw, E. A., Mensah, R. K., Pittman, G. W., and Walter, G. H. (2009). The microbial flora of Aphis gossypii: patterns across host plants and geographical space. J. Invertebr. Pathol. 100, 123–126. doi: 10.1016/j.jip.2008.10.005
Oksanen, J., Blanchet, F. G., Friendly, M., Kindt, R., Legendre, P., McGlinn, D., et al. (2018). Package ‘vegan', community ecology package. Version 2.5-2. Available online at: https://github.com/vegandevs/vegan
Oliver, K. M., Degnan, P. H., Burke, G. R., and Moran, N. A. (2010). Facultative symbionts in aphids and the horizontal transfer of ecologically important traits. Ann. Rev. Entomol. 55, 247–266. doi: 10.1146/annurev-ento-112408-085305
Oliver, K. M., Moran, N. A., and Hunter, M. S. (2005). Variation in resistance to parasitism in aphids is due to symbionts not host genotype. Proc. Natl. Acad. Sci. U. S. A. 102, 12795–12800. doi: 10.1073/pnas.0506131102
Oliver, K. M., Russell, J. A., Moran, N. A., and Hunter, M. S. (2003). Facultative bacterial symbionts in aphids confer resistance to parasitic wasps. Proc. Natl. Acad. Sci. U. S. A. 100, 1803–1807. doi: 10.1073/pnas.0335320100
Quast, C., Pruesse, E., Yilmaz, P., Gerken, J., Schweer, T., Yarza, P., et al. (2013). The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic. Acids Res. 41, D590–596. doi: 10.1093/nar/gks1219
Russell, J. A., and Moran, N. A. (2006). Costs and benefits of symbiont infection in aphids: variation among symbionts and across temperatures. Proc. Biol. Sci. 273, 603–610. doi: 10.1098/rspb.2005.3348
Sepúlveda, D. A., Zepeda-Paulo, F., Ramírez, C. C., Lavandero, B., and Figueroa, C. C. (2017). Diversity, frequency, and geographic distribution of facultative bacterial endosymbionts in introduced aphid pests. Insect. Sci. 24, 511–521. doi: 10.1111/1744-7917.12313
Tougeron, K., and Iltis, C. (2022). Impact of heat stress on the fitness outcomes of symbiotic infection in aphids: a meta-analysis. Proc. R. Soc. B Biol. Sci. 289:20212660. doi: 10.1098/rspb.2021.2660
Tsuchida, T., Koga, R., Shibao, H., Matsumoto, T., and Fukatsu, T. (2002). Diversity and geographic distribution of secondary endosymbiotic bacteria in natural populations of the pea aphid, Acyrthosiphon pisum. Mol. Ecol. 11, 2123–2135. doi: 10.1046/j.1365-294X.2002.01606.x
Wang, C., Li, X., Jin, D., Gong, P., Li, Q., Zhang, Y., et al. (2022). Implications of environmentally shaped microbial communities for insecticide resistance in Sitobion miscanthi. Environ. Res. 215:114409. doi: 10.1016/j.envres.2022.114409
Wang, X. Y., Yang, T., Shen, L., Zhang, W. L., and Li, L. H. (2021). Formation of factors influencing cotton yield in jujube-cotton intercropping systems in Xinjiang, China. Agrofor. Syst. 95, 1–13. doi: 10.1007/s10457-020-00571-w
Wei, J., Niu, M., and Feng, J. (2016). Diversity and distribution patterns of scale insects in China. Ann. ESA 2016:109. doi: 10.1093/aesa/sav159
Wu, K. M., and Guo, Y. Y. (2005). The evolution of cotton pest management practices in China. Ann. Rev. Entomol. 50, 31–52. doi: 10.1146/annurev.ento.50.071803.130349
Xu, S., Jiang, L., Qiao, G., and Chen, J. (2020). The bacterial flora associated with the polyphagous Aphid Aphis gossypii Glover (Hemiptera: Aphididae) is strongly affected by host plants. Microb. Ecol. 79, 971–984. doi: 10.1007/s00248-019-01435-2
Yang, K., Zhang, H. Y., Wang, P., Jin, G. X., and Chu, D. (2023). Both symbionts and environmental factors contribute to shape the microbiota in a pest insect, Sogatella furcifera. Front. Microbiol. 14:1336345. doi: 10.3389/fmicb.2023.1336345
Yu, Y., Lee, C., Kim, J., and Hwang, S. (2005). Group-specific primer and probe sets to detect methanogenic communities using quantitative real-time polymerase chain reaction. Biotechnol. Bioeng. 89, 670–679. doi: 10.1002/bit.20347
Zhang, S., Su, H., Jiang, W., Hu, D., Ali, I., Jin, T., et al. (2021). Symbiotic microbial studies in diverse populations of Aphis gossypii, existing on altered host plants in different localities during different times. Ecol. Evol. 11, 13948–13960. doi: 10.1002/ece3.8100
Keywords: Aphis gossypii, bacterial communities, 16S rRNA gene sequencing, environmental factors, cotton
Citation: Alimu A, Gao Y, Liu J and Lu Y (2025) Geographic factors influence communities of symbiotic bacterial communities in Aphis gossypii across China's major cotton regions. Front. Microbiol. 16:1569543. doi: 10.3389/fmicb.2025.1569543
Received: 01 February 2025; Accepted: 10 March 2025;
Published: 01 April 2025.
Edited by:
Decai Jin, Chinese Academy of Sciences, ChinaReviewed by:
Yong Zhang, Anhui Agricultural University, ChinaXingya Wang, Shenyang Agricultural University, China
Copyright © 2025 Alimu, Gao, Liu and Lu. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Yanhui Lu, bHV5YW5odWlAY2Fhcy5jbg==
 Abulaiti Alimu
Abulaiti Alimu Yu Gao
Yu Gao Jinping Liu1
Jinping Liu1 Yanhui Lu
Yanhui Lu