
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Microbiol. , 17 March 2025
Sec. Antimicrobials, Resistance and Chemotherapy
Volume 16 - 2025 | https://doi.org/10.3389/fmicb.2025.1553148
This article is part of the Research Topic Emerging Antimicrobials: Sources, Mechanisms of Action, Spectrum of Activity, Combination Antimicrobial Therapy, and Resistance Mechanisms View all 21 articles
 Jiming Wu1†
Jiming Wu1† Jisheng Zhang1†
Jisheng Zhang1† Jianmin Wang1†
Jianmin Wang1† Jin Wang1
Jin Wang1 Xushan Liang1
Xushan Liang1 Chunli Wei1
Chunli Wei1 Wenzhang Long1
Wenzhang Long1 Yang Yang1
Yang Yang1 Yuhui Chen1
Yuhui Chen1 Mingjing Liao1
Mingjing Liao1 Youtao Liang1
Youtao Liang1 Kaixin Yu2
Kaixin Yu2 Xiaoli Zhang1*
Xiaoli Zhang1*Due to the complexity of identifying the Enterobacter cloacae complex (ECC) at the species level, little is known about the distribution of carbapenem-resistant ECC (CRECC). Plasmid-mediated mcr family genes are significant contributors to polymyxin resistance. The emergence of the mcr-9 gene has further complicated the landscape of polymyxin resistance in CRECC. Our study aimed to ascertain the prevalence of CRECC and the mcr-9 gene, and to elucidate the mechanisms underlying high-level resistance to polymyxin B (PB). In this study, we collected 212 non-replicating ECC strains, identifying 38 CRECC strains (17.9%, 38/212) and Enterobacter hormaechei (71.1%, 27/38) as the predominant endemic strains. Among these, 10 CRECC strains (36.3%, 10/38) were found to harbor the mcr-9 gene. Interestingly, the presence of mcr-9 did not significantly impact PB resistance or impose a fitness cost. While overexpression of mcr-9 can enhance PB resistance within a certain range and may incur fitness costs, it does not result in high-level PB resistance. The PB resistance of 17 CRECC strains was notably increased (from 16 to 128 mg/L), accompanied by mutations in the phoP/Q and mgrB genes. Notably, two novel insertion sequences, IS5D and IS1X2, were discovered within the mgrB gene. The inactivation of mgrB results in the loss of its negative regulatory effect on the two-component system. Protein structure predictions indicated that mutations in phoQ primarily affect the phosphatase (HAMP) and histidine kinase domains. This research significantly expands our comprehension of the complexities of PB resistance, highlighting the multifactorial nature of antibiotic resistance mechanisms.
Carbapenem antibiotics are a key treatment strategy for Gram-negative enterobacterial infections, and the prevalence of carbapenem-resistant Enterobacterales (CRE) has been progressively increasing due to the extensive and long-term use of antibiotics (Yoon et al., 2019; Lee et al., 2022; Puljko et al., 2024). Among them, carbapenem-resistant Enterobacter cloacae complex (CRECC) is the third most common pathogen, often causing serious infections (Zhang et al., 2018; Girlich et al., 2021). CRECC frequently exhibits multidrug resistance, forcing us to reintroduce polymyxins to combat these difficult-to-treat bacteria (Li et al., 2006). Polymyxin B (PB), belonging to the polymyxin family along with polymyxin E, is considered as a last resort against multi-drug resistant Enterobacterales (Nang et al., 2021). However, there has been an alarming rise in global reports of PB resistance, posing a serious public health concern (Zhang et al., 2021; Liao et al., 2022). Enterobacter cloacae complex (ECC) is a complex community composed of multiple species, including E. hormaechei, E. asburiae, E. kobei and E. cloacae etc. The members of ECC are difficult to distinguish using the conventional phenotypic/biochemical or mass spectrometry identification methods available in clinical laboratories (Garinet et al., 2018; Girlich et al., 2021). Therefore, the prevalence trend of CRECC at species level still needs to be investigated.
Furthermore, previous studies have demonstrated that the pressure of antibiotics creates resistance to PB, leading to an increased risk of clinical treatment failure (Huang et al., 2020; Li et al., 2021). While most epidemiological reports primarily focus on the inherent resistance of carbapenem-resistant Enterobacterales to PB prior to exposure (Zhang et al., 2021), it is crucial to investigate cases where high-level resistance to polymyxins emerges in strains after exposure (Zhu et al., 2019; Wu et al., 2023). The resistance mechanisms in Enterobacter for PB are categorized into chromosomally mediated intrinsic resistance and plasmid-mediated acquired resistance (Poirel et al., 2017; Choi et al., 2020). Intrinsic drug resistance is primarily attributed to the PhoPQ two-component system and its negative regulator MgrB, which modulate downstream arnBCADTEF expression and subsequently mediate L-Ara4N modification of lipid A (Nang et al., 2021). While the PmrAB two-component system has been implicated in regulating polymyxin resistance in Klebsiella pneumoniae, studies have reported no impact of pmrAB on PB resistance within ECC (Doijad et al., 2023). These findings highlight distinct mechanisms of PB resistance across different bacterial species. Plasmid-mediated polymyxin resistance predominantly involves mobile colistin-resistant genes, including mcr-9 gene with three subtypes (Li et al., 2020; Zelendova et al., 2023). Previous investigations suggest that mcr-9 is more likely to be acquired and stably inherited by CRE strains; furthermore, there have been reports linking epidemic highly virulent clones carrying both mcr-9 and carbapenemase genes with E. hormaechei infections (Xu et al., 2022b; Zhou et al., 2022). However, further studies are needed to elucidate the specific effects of mcr-9 on PB resistance and its association with patient clinical outcomes or potential outbreaks posing public health concerns—particularly within widespread mcr-9-carrying ECC isolates (Liao et al., 2022; Xu et al., 2022a).
Therefore, a comprehensive analysis was conducted on the prevalence of CRECC and mcr-9 in clinical ECC isolates. The impact of mcr-9 and its overexpression on both PB resistance and growth of ECC strains were investigated. Moreover, an in vitro induction approach was employed to elucidate the underlying mechanism behind the development of high-level PB resistance in CRECC.
From December 2018 to April 2023, a total of 212 non-repetitive isolates of ECC were collected from clinical specimens at a teaching hospital (Supplementary Table S2). Identification was conducted using the VITEK-2 automated microbiological analyzer and MALDI-TOF MS (Bruker, Billerica, MA, United States). Based on the results of clinical drug sensitivity testing, 38 CRECC isolates were identified. The minimum inhibitory concentration (MIC) of PB was determined using the broth microdilution method with strain ATCC25922 as the quality control strain. Each MIC determination was repeated three times following CLSI (Clinical and Laboratory Standards Institute) guidelines (Satlin et al., 2020).
WGS was performed on 212 ECC isolates using Illumina HiSeq PE150 (Illumina, San Diego, CA, United States), and 10 CRECC isolates carrying mcr-9 for further long-read sequencing (Oxford Nanopore, Oxford, United Kingdom). Long-read sequencing assembly was performed using unicycler v0.4.8 and was corrected using pilon v1.22 (Walker et al., 2014; Wick et al., 2017). All strains contained 3,048 core genomes according to the reference genome (GenBank accession no. CP010377). The visualization of cg-MLST results is based on the previous methodology (Stamatakis, 2014). The results of core genome single nucleotide polymorphism (cg-SNP) were visualized using ggtree after building the evolutionary tree with FastMe (Lefort et al., 2015; Xie et al., 2023). FastANI was employed to rapidly calculate ANI and classify ECC species (Jain et al., 2018).
In order to obtain CRECC strains with high-level resistance to PB through induction, we made certain modifications based on previous studies and conducted a 7-day in vitro induction experiment (Wu et al., 2023). Initially, overnight cultures of 36 isolates of CRECC were incubated in 1 mg/L PB for 24 h. Subsequently, 10 μL aliquot of the overnight culture was inoculated onto blood plates and incubated at 37°C for 18 h to assess growth continuation. Simultaneously, a volume of 20 μL from the overnight culture was inoculated into fresh LB broth (4 mL) with doubled PB concentration. Finally, the MIC of induced strains toward PB was determined using the broth microdilution method.
All successfully induced CRECC isolates with high levels of PB resistance were detected by PCR for mutations in related genes. Genomic DNA was extracted using TIANamp bacterial DNA kit (Tiangen Biotechnology, Beijing, China) according to the manufacturer’s recommendations. The collected DNA was detected by gel electrophoresis using 1.2% agarose (Invitrogen) in 0.5 × Tris Borate EDTA running buffer (BIO-RAD) and quantified by ND 2000 (NanoDrop Technologies). Sanger sequencing of the positive amplification products was performed at Shanghai Shenggong (Shangai, China).
According to the method described previously, the bonding experiment was conducted using the membrane bonding method (Gong et al., 2018). The donor (CRECC68) and recipient culture (E. coli EC600) were mixed in a 1:3 ratio in Luria-Bertani (LB) broth. The mixture was inoculated in a filter membrane and incubated at 37 ° C for 24 h. Meropenem (1 mg/L) and rifampicin (600 mg/L) were added to the Muller-Hinton agar plate to screen the transconjugants. The presence of transconjugants were confirmed by VITEK-2 compact system and PCR.
The upper and downstream homologous recombination arm of the mcr-9.1 gene was amplified from the CRECC68 genome by superfidelity DNA polymerase; the apramycin resistance gene (apr) was amplified from the pUC57-apr plasmid; the upper and downstream homologous recombination arm of the mcr-9.1 gene and apr by fusion PCR to generate the target fragment Δmcr-9.1, which was cloned into the suicide plasmid pCVD442 to obtain the target plasmid pCVD442-Δmcr-9.1. By electroporation, pCVD442-Δmcr-9.1 was transferred to E. coli β2155 to obtain the donor bacteria β2155/pCVD442-Δmcr-9.1. β2155/pCVD442-Δmcr-9.1 was used as donor to conjugate with the receptor of E. cloacae. Several Ecl/pCVD442-Δmcr-9.1 clone solutions were spread on LB plates containing 10% sucrose and grown until the single clones were formed. The clones with the mcr-9.1 gene knockout were selected by PCR screening and named CRECC68-Δmcr-9.1. PB susceptibility testing was subsequently performed on CRECC68-Δmcr-9.1 to verify the function of the mcr-9.1 gene. The primers used in this experiment are shown in the Supplementary Table S1.
The full length of the mcr-9.2 gene was amplified from the genomic DNA of CRECC414 isolate using PCR. The PCR products and pET-28a plasmid were enzymatically digested at 37°C for 3 h. The purified target gene and pET-28a plasmid were ligated using T4 DNA ligase, followed by transformation into DH5α recipient cells and spreading on LB plates containing kanamycin (100 μg/mL). Monoclonal strains were selected the next day and verified through PCR and agarose gel electrophoresis. After successful verification, the PB MIC value of recombinant bacterium DH5α + pET-28a + mcr-9.2 was determined.
The growth curve of the strain was determined by measuring the absorbance at 600 nm using an Multifunctional Microplate Reader (Thermo Scientific) within a 24-h period. The strain culture was inoculated in LB medium broth at a ratio of 1:100 and incubated at 37°C with agitation at 180 rpm. At the third hour, IPTG (1 μM) was added to induce the expression of mcr-9 in the recombinant strain. The experiment was repeated three times.
The N-phenyl-1-naphthylamine (NPN) uptake test was employed to assess the permeability of the outer membrane, following previously established protocols (Burow et al., 2009; Lu et al., 2022). In brief, bacterial samples were collected during the logarithmic growth phase. Cultures of E. coli harboring the recombinant plasmid pET-28a + mcr-9.2 and control strains were centrifuged at 1,000 g for 5 min. Subsequently, cells were washed thrice with PBS and resuspended to an optical density (OD600) of 0.5. The bacterial suspension was thoroughly mixed with NPN at a final concentration of 10 μM. After incubation at room temperature for 10 min, cell suspensions were analyzed using Multifunctional Microplate Reader (Thermo Scientific). Fluorescence emission was measured at an excitation wavelength of 350 nm and an emission wavelength of 450 nm.
We collected 212 non-duplicate clinical strains of the ECC, including 38 (17.9%) CRECC and 174 (82.1%) carbapenem-sensitive ECC (CSECC) (Figure 1A; Supplementary Table S2). The prevalence of carbapenem resistance among the ECC strains was as follows: E. hormaechei (20.9%, 27/129), E. asburiae (23.5%, 4/17), E. kobei (17.6%, 3/17), E. roggenkampii (5.9%, 1/17), E. cloacae (13.3%, 2/15), and E. mori (16.7%, 1/6). All isolates of E. ludwigii and E. bugandensis exhibited susceptibility to carbapenems. The mcr-9 carriage rate among ECC strains was 7.5% (16/212), with the following distribution: E. hormaechei (n = 8), E. kobei (n = 3), E. asburiae (n = 3), E. cloacae (n = 1), and E. ludwigii (n = 1) (Figure 1B). The metallo-carbapenemase gene blaNDM-1 was found in 76.3% (29/38) of CRECC strains (Figure 1B). Additionally, Figure 1B depicts the specific carriage patterns of AmpC β-lactamase and ESBL genes. Notably, mcr-1, a gene associated with polymyxin resistance, was not detected.
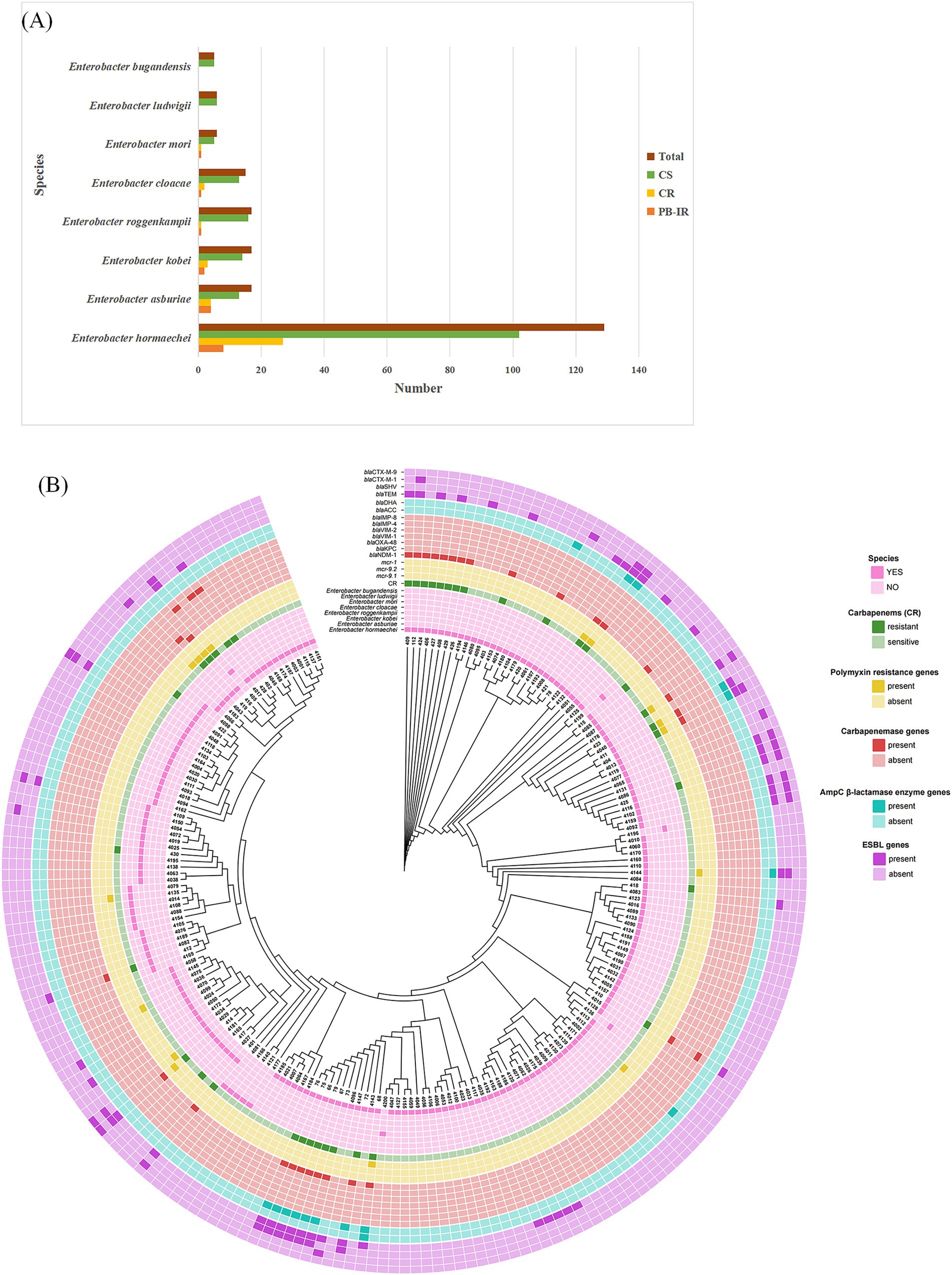
Figure 1. Epidemiological background of isolates. (A) Distribution characteristics of 212 isolates of Enterobacter cloacae complex at species level. CS, carbapenem sensitive; CR, carbapenem resistant; PB-IR, Polymyxin B (PB) induced resistance. (B) Phylogenetic analysis and characterization of drug resistance gene carriage were conducted in a cohort of 212 ECC strains.
A total of 38 non-duplicate clinical strains of CRECC were collected. Of these, 10 carried the mcr-9 gene, with 9 containing mcr-9.1 and 1 containing mcr-9.2 (Table 1; Figure 1B). The 38 CRECC isolates were classified into six species, with E. hormaechei being the predominant strain (71.1%, 27/38). Other species included E. asburiae (10.5%, 4/38), E. kobei (7.9%, 3/38), E. roggenkampii (2.6%, 1/38), E. cloacae (5.3%, 2/38), and E. mori (2.6%, 1/38). The main clinical sources of CRECC strains were blood (44.7%, 17/38) and urine (18.4%, 7/38). The mcr-9 gene was present in 26.3% (10/38) of CRECC strains, compared to 3.4% (6/174) in CSECC. CRECC strains are often accompanied by mcr-9 gene, which is consistent with relevant studies (Xu et al., 2022a; Zhou et al., 2022; Zeng et al., 2023). The high prevalence of the mcr-9 gene in CRECC strains had been concerning due to its association with high levels of PB resistance, presenting a significant public health issue.
To investigate the impact of mcr-9 on PB resistance in clinical strains, we performed gene knockout using homologous recombination and successfully generated the CRECC68-Δmcr-9.1 deletion mutant. The PB MIC for wild-type CRECC68 strain was 4 mg/L, whereas the MIC (PB) for CRECC68-Δmcr-9.1 decreased by half compared to the wild-type strain (Table 2). Considering the complexity of clinical strain backgrounds, we conducted conjugation and cloning experiments separately. The transconjugant J68 containing mcr-9.1 gene was obtained by conjugating the plasmid carrying mcr-9.1 to EC600. Additionally, we cloned the mcr-9.2 gene into pET-28a vector to construct recombinant strain DH5α + pET-28a + mcr-9.2. PB MICs were determined using microbroth dilution method for EC600, J68, DH5α, DH5α + pET-28a, and DH5α + pET-28a + mcr-9.2 strains. Our results showed that all strains were sensitive to PB, with no significant differences in MIC between EC600 and J68 in the conjugation experiment (Table 2). Similarly, no significant differences in PB MICs were observed among DH5α, DH5α + pET-28a, and DH5α + pET-28a + mcr-9.2 strains. However, upon IPTG induction for high-level expression of mcr-9.2 in BL21 (DE3)pLysS+pET-28a + mcr-9.2 strains, we observed a doubling of the MIC compared to control strains without IPTG. Overall, our findings suggest that both mcr-9.1 and mcr-9.2 contribute minimally to PB resistance, indicating that mcr-9 does not confer high levels of resistance.
To assess the fitness cost associated with the mcr-9 gene, we conducted a growth curve analysis. The growth rate of J68 was comparable to that of EC600, and no significant difference was observed between CRECC68 and CRECC68-Δmcr-9.1 (Figure 2A). Similarly, the growth of BL21 (DE3)pLysS+pET-28a + mcr-9.2 was consistent with that of BL21 (DE3)pLysS+pET-28a (Figure 2B). In addition, the high expression of mcr-9.2 was induced by IPTG, and the bacterial growth was monitored. The growth of BL21(DE3)pLysS+pET-28a + mcr-9.2(IPTG) with high expression of mcr-9.2 gene was slower compared to BL21(DE3)pLysS+pET-28a(IPTG) (Figure 2B). These results indicate that overexpression of mcr-9.2 incurs a fitness cost.
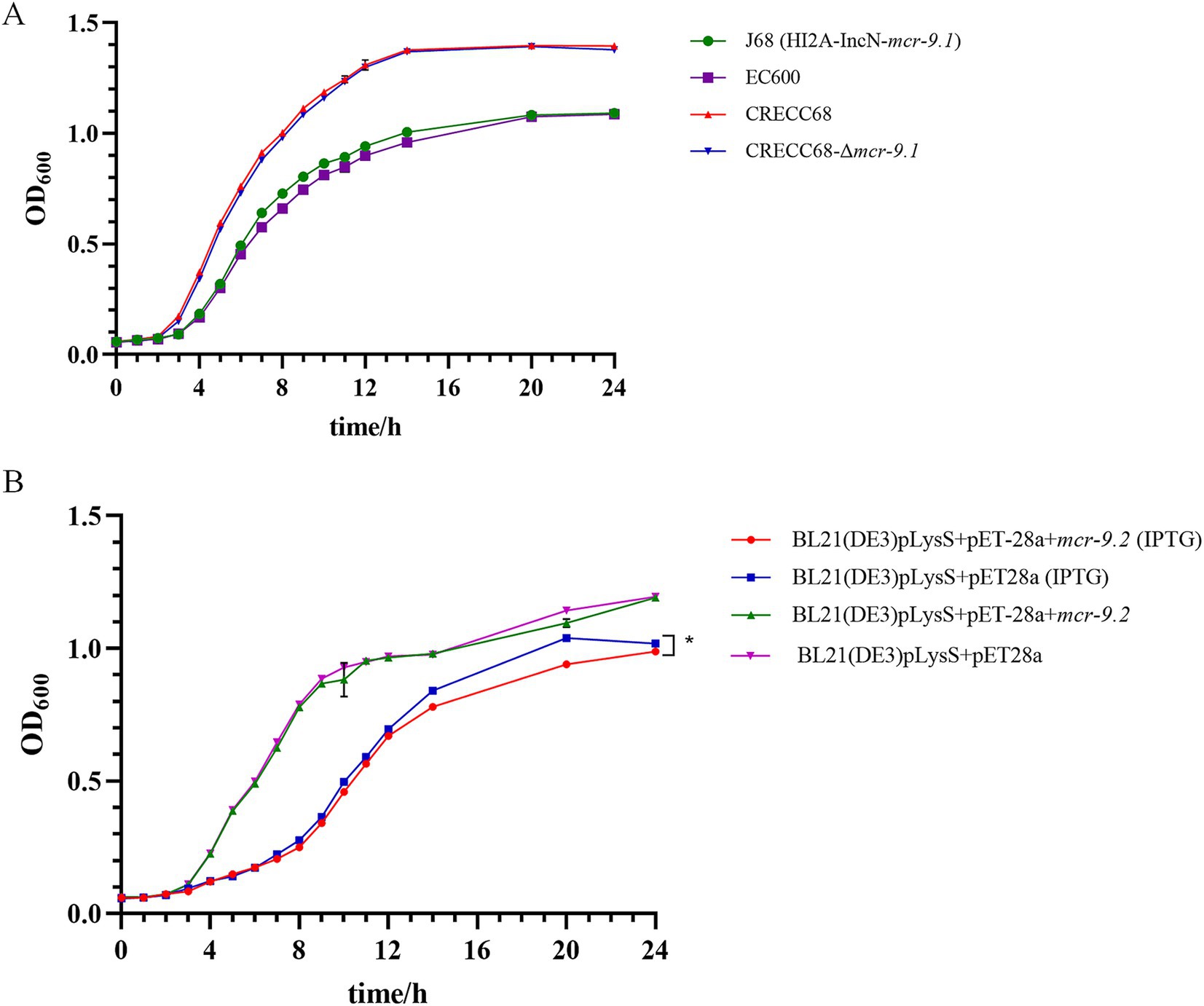
Figure 2. The influence of carrying mcr-9 and overexpression of mcr-9 on the growth of ECC. (A) Growth curves of transconjugant J68, receptor EC600, wild-type CRECC68 and mcr-9.1 deletion mutant strain CREC68-Δmcr-9.1. (B) Growth curves of Escherichia coli BL21(DE3) pLysS harboring empty vector pET-28a or pET-28a + mcr-9.2 with or without 1 mM IPTG induction. The data shows the mean ± SD of three independent replicates. Statistical significance was determined via a two-way ANOVA; *p ≤ 0.05.
NPN is a hydrophobic fluorescent probe that can integrate into cell membranes (Lee and Je, 2013). Its fluorescence intensity increases in hydrophobic environments, making it a useful indicator for assessing membrane permeability (Lu et al., 2022). Therefore, we determined the permeability of the outer membrane by NPN uptake test. The fluorescence intensity of BL21(DE3)pLysS+pET-28a + mcr-9.2 significantly increased with 1 mM IPTG compared to 0 mM IPTG (Figure 3). In contrast, there was no significant difference in the fluorescence intensity of the control group BL21(DE3)pLysS+pET-28a at 1 mM IPTG versus 0 mM IPTG (Figure 3). Thus, overexpression of mcr-9.2 in the presence of IPTG leads to a significant increase in outer membrane permeability (Figure 3).
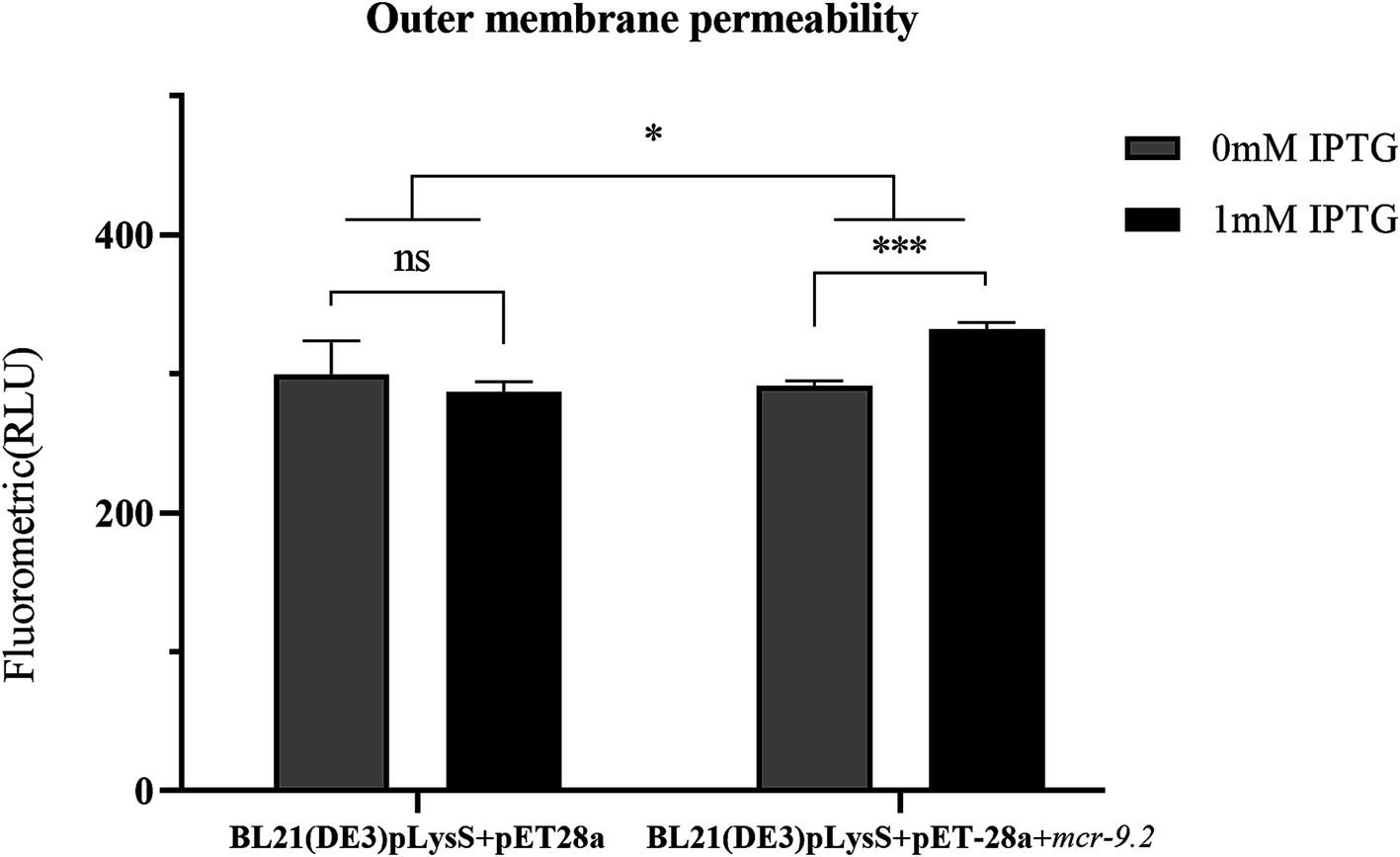
Figure 3. Effect of mcr-9.2 expression on membrane permeability. Outer membrane permeability of Escherichia coli BL21(DE3) pLysS harboring empty vector pET-28a or pET-28a + mcr-9.2 with or without 1 mM IPTG induction. The asterisks indicate statistical significance at different levels by paired t-test: *p ≤ 0.05; ***p ≤ 0.001; ns, not significant.
In vitro induction of PB was performed on 36 strains of low-level resistant CRECC (excluding CRECC405 and CRECC417) (Figure 4A; Table 3). After 7 days of continuous induction, the drug-resistant phenotype of 17 CRECC strains showed significant changes. These strains included E. hormaechei (n = 8), E. asburiae (n = 4), E. kobei (n = 2), E. roggenkampii (n = 1), E. cloacae (n = 1), and E. mori (n = 1). The MIC of PB was determined for these 17 induced CRECC strains using the microbroth dilution method. A significant increase in MIC values, ranging from 16 mg/L to >128 mg/L, was observed upon exposure to PB, as shown in Table 3.
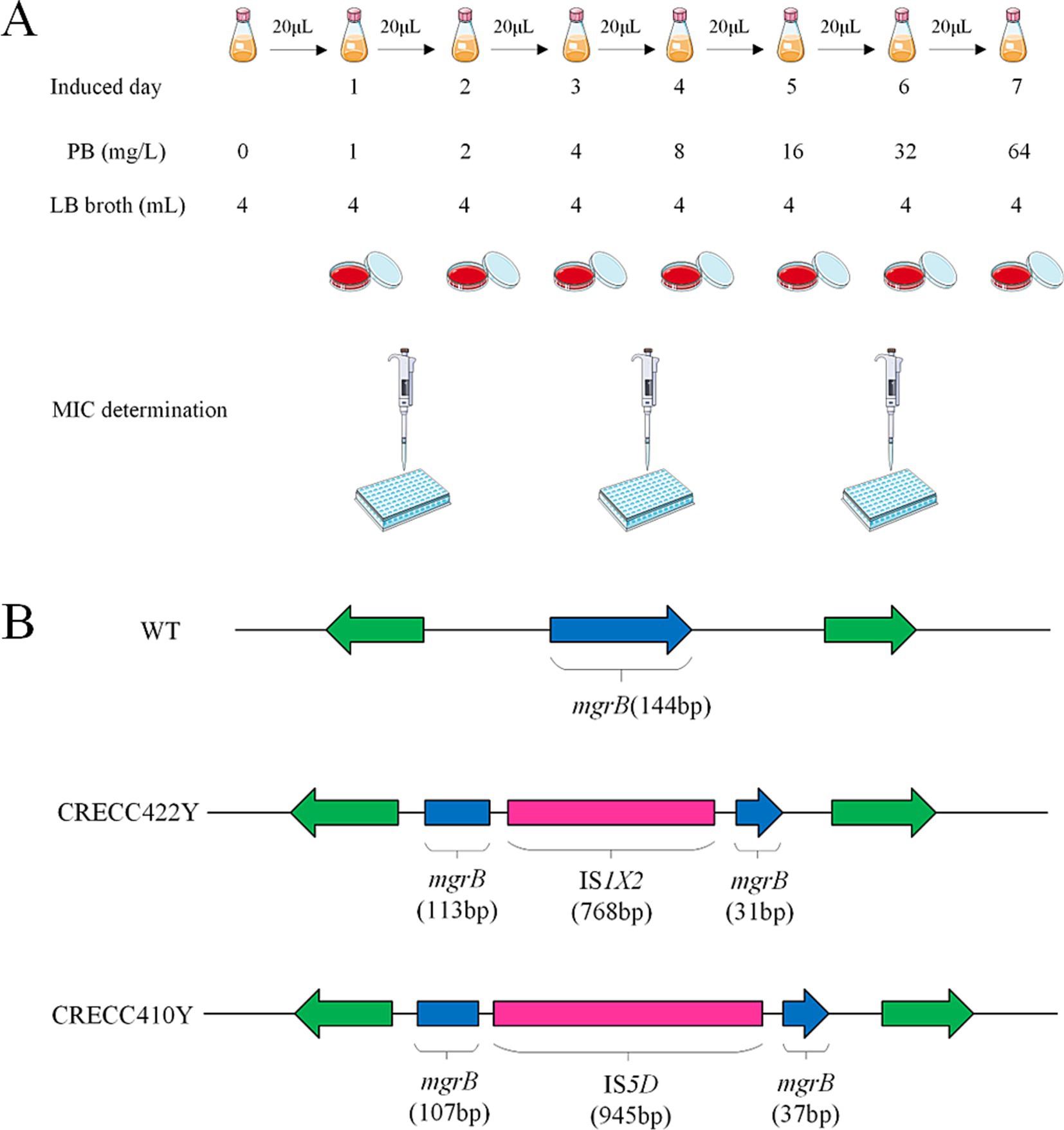
Figure 4. Experimental flow graph and analysis on insertional inactivated mgrB gene of in vitro polymyxin B inducted resistant variants. (A) Schematic diagram of polymyxin B induction experiment in vitro. (B) Insertional inactivation mgrB gene of carbapenem-resistant Enterobacter cloacae complex (CRECC) strains in vitro. Schematic representation of the mgrB gene in vitro-induced variants, CRECC422Y and CRECC410Y, derived from CRECC422 and CRECC410.
To investigate the molecular mechanisms underlying high-level drug resistance, we screened genes associated with the regulation of polymyxin resistance. Previous studies have identified the two-component system genes phoP and phoQ, as well as the negative regulatory gene mgrB, as crucial factors in polymyxin resistance (Doijad et al., 2023). Prior to exposure to PB, only CRECC430 and CRECC412 exhibited mutations (Table 3). In CRECC430, the phoP gene was truncated, and multiple amino acid substitutions were observed in phoQ. In CRECC412, the phoQ gene was truncated. The MICs of PB for pre-induction CRECC isolates ranged from 0.5 to 4 mg/L. After PB induction, the MICs of PB for 17 CRECC strains increased significantly, ranging from 16 to >128 mg/L (Table 3). The most frequently mutated gene was phoQ (58.8%, 10/17), with various mutations including amino acid substitutions (n = 5), frameshift mutations (n = 3), a nonsense mutation (n = 1), and gene truncations (n = 1) (Table 3). In contrast, no new mutations were found in the phoP gene after exposure to PB. The negative regulatory gene mgrB also exhibited multiple mutations, including deletions (n = 3), insertion sequences (n = 2), amino acid substitutions (n = 2), and a frameshift mutation (Table 3).
Insertion sequence disruption of mgrB is a key mechanism for polymyxin resistance. We found two cases with insertion sequences in polymyxin-exposed strains, identified by IS Finder analysis as IS1X2 (CRECC422Y) and IS5D (CRECC410Y) (Figure 4B). IS1X2, belonging to the IS1 family, fragments the mgrB gene into 113 bp and 31 bp segments. IS5D, from the IS5 family, is inserted after 107 bp of the mgrB gene. Notably, these insertion sequences are reported here for the first time in the mgrB gene of CRECC.
We identified several novel non-synonymous mutations in phoQ among polymyxin-resistant mutant strains (Figure 5; Table 3). To pinpoint these mutations, we used the AlphaFold sequencing scheme to model the structures of wild-type and mutant PhoQ dimers. The 3D structures of both wild-type and mutant PhoQ showed similar arrangements, covering the transmembrane region, periplasmic sensor domain, HAMP domain and histidine kinase region (Figure 5). The observed mutation sites in resistant mutants CRECC68Y (A351T), CRECC410Y (C423R), CRECC67Y (E397A), and CRECC75Y (E304V) were within the histidine kinase domain (Figures 5A–C,E). Secondary structure analysis revealed that all mutation sites, including CRECC68Y (A351T), CRECC410Y (C423R), CRECC72Y (E232K), and CRECC75Y (E304V), were within α helices (Figures 5A,B,D,E). Single point mutations in this region can affect domain energetics and their interactions, impacting polymyxin sensitivity (Mensa et al., 2021).
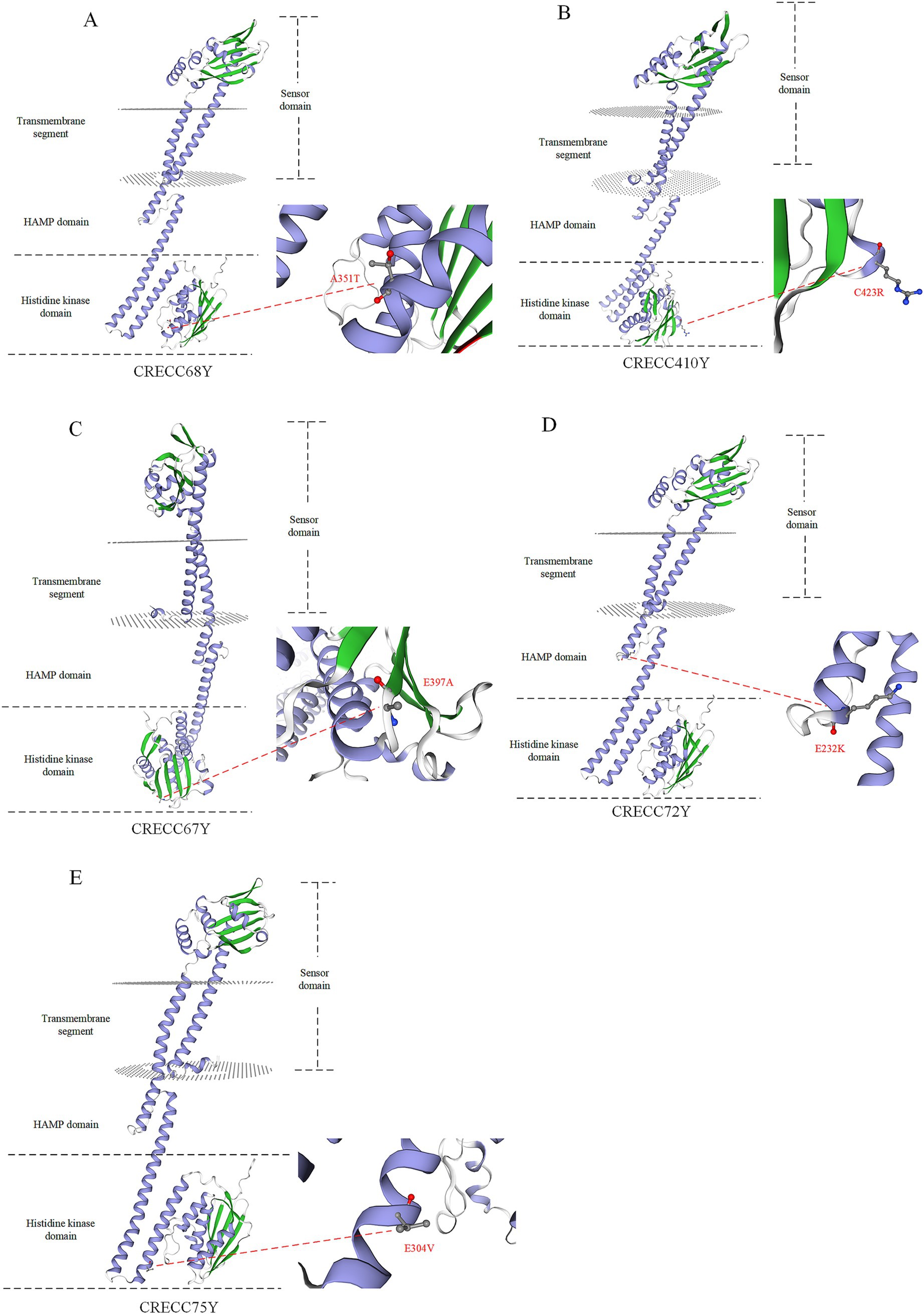
Figure 5. 3D simulation snapshots of PhoQ protein in polymyxin B resistant mutant strains. (A) Resistant mutant strain CRECC68Y (A351T). (B) Resistant mutant strain CRECC410 (C423R). (C) Resistant mutant strain CRECC67Y (E397A). (D) Resistant mutant strain CRECC72Y (E232K). (E) Resistant mutant strain CRECC75Y (E304V). The alpha helix of the protein is represented by the blue band while beta type structure is depicted by the green band.
Clinical laboratories should focus on accurately identifying ECC species and providing improved guidance on carbapenem use. Additionally, polymyxin resistance in CRECC strains is increasingly common, with many resistant strains harboring the mcr-9 gene (Kim et al., 2021; Wu et al., 2022; Yan et al., 2024). This paper discusses the specific effects of mcr-9 carriage and overexpression.
This study found a significantly higher prevalence of the mcr-9 gene in CRECC strains (26.3%) compared to other previous reports (Huang et al., 2020; Wang et al., 2020). Specifically, 9 strains were identified as mcr-9.1 and only one as mcr-9.2, with mcr-9.1 showing a more pronounced epidemic trend (Table 1). The prevalence of mcr-9 in CRECC was notably higher than in CSECC, which is consistent with previous studies (Zhou et al., 2022). These findings highlight the need to be cautious about the potential impact of mcr-9. Despite this, the results indicated that both mcr-9.1 and mcr-9.2 had a very limited effect on PB resistance. This research represents the first exploration of how different subtypes of the mcr-9 gene contribute to resistance. The mcr-9-mediated drug resistance may be linked to its expression level, as high expression of mcr-9 is observed in polymyxin-resistant strains (Kieffer et al., 2019; Jiang et al., 2022). We introduced the plasmid pET-28a + mcr-9.2 into the BL21(DE3) pLysS expression strain and induced high expression of mcr-9.2 with IPTG. However, results showed that even with high mcr-9.2 expression in BL21(DE3)pLysS+pET-28a + mcr-9.2(IPTG), there was only a two-fold increase in the MIC of PB compared to control strains (Table 2). Previous studies have demonstrated that the overexpression of mcr-1 imposes a fitness cost on E. coli (Lu et al., 2022). To investigate whether mcr-9 has similar effects in E. cloacae, we compared the growth of strains with high expression of mcr-9 to control strains. No significant differences were observed between E. coli J68 and its control strain EC600, or E. cloacae CRECC68 and CRECC68-Δmcr-9.1, suggesting that mcr-9 alone does not significantly impact bacterial resistance or incur a fitness cost (Figure 2A). This lack of a fitness cost in mcr-9-positive strains might contribute to the widespread prevalence of the mcr-9 gene in CRECC. However, IPTG-induced overexpression of mcr-9 led to slower growth in strain BL21(DE3)pLysS+pET-28a + mcr-9.2(IPTG) compared to the control strain (Figure 2B). This suggests that elevated mcr-9 expression imposes a fitness cost on BL21(DE3)pLysS+pET-28a + mcr-9.2(IPTG), though it does not confer high-level resistance to PB. Moreover, strain BL21(DE3)pLysS+pET-28a + mcr-9.2 exhibited significantly increased membrane permeability in the presence of IPTG (Figure 3). We speculate that mcr-9 overexpression may disrupt the bacterial outer membrane structure, increasing permeability and causing leakage of cell contents.
Exposure to PB has been shown to induce drug resistance, as demonstrated by reports of Enterobacter strains initially sensitive to PB evolving resistance during treatment and displaying a pronounced drug-resistant phenotype (Wu et al., 2023). In our in vitro studies, clinical CRECC strains with either PB sensitivity or low-level resistance were transformed to high-level resistance (PB MIC ≥16 mg/L) (Table 3). Sequence analysis of these genes after PB exposure revealed several CRECC strains with mutations including amino acid substitutions, frameshift mutations, premature terminations and partial deletions in the phoQ gene (Table 3). However, only one truncation was observed in the phoP gene, and it was not related to PB resistance (Table 3). The phoQ gene appears to play a more significant role than the phoP gene in the development of PB resistance in ECC. The mgrB gene plays a key role in acquired drug resistance (Mhaya et al., 2020; Yang et al., 2020). Among the CRECC strains, eight (22.2%, 8/36) showed alterations in the mgrB gene, including deletions (n = 3), insertion-induced inactivation (n = 2), amino acid substitutions (n = 2), and frameshift mutations (n = 1) (Table 3). We identified two novel mutations in the mgrB gene: a tryptophan substitution at position 47 in CRECC66 and a proline substitution at position 40 in CRECC414 (Table 3). Notably, insertion sequences were exclusively found in the mgrB gene, with IS1X2 from the IS1-family and IS5D from the IS5-family disrupting the mgrB gene in CRECC422Y and CRECC410Y, respectively (Figure 4B). This represents the first report of two insertion sequences within the mgrB gene of ECC, highlighting the need for further surveillance to evaluate their potential widespread distribution. It is intriguing that strains CRECC402 and CRECC416 have no resistance mutations, and this observation requires further exploration of potential resistance mechanisms. Resistance could arise from regulatory changes rather than mutations directly associated with polymyxin resistance. For instance, alterations in the regulation of genes associated with the stress response, quorum sensing, or biofilm formation could indirectly confer resistance to polymyxins (Harms et al., 2016; Hwang and Kim, 2024).
Bacterial cells must continuously adapt to survive changing external environments. In bacteria and some lower eukaryotes, two-component systems connect extracellular stimuli with intracellular adaptive responses (Cheung et al., 2008). PhoQ, a transmembrane histidine kinase, is essential for sensing various environmental signals and is involved in the metabolic homeostasis and pathogenesis of many Gram-negative bacteria (Doijad et al., 2023). This study identified several novel mutations in the phoQ gene, prompting an evaluation of their potential impact on protein function. Mensa et al. demonstrated that E232A mutations exhibit diverse effects, revealing how allosteric coupling of domain equilibria is influenced by point substitutions (Mensa et al., 2021). Similarly, we identified the E232K mutation (CRECC72Y) in the HAMP domain which disrupts the balance between sensor and catalytic domains, significantly affecting ligand sensitivity of PhoQ as well as signal magnitude and direction (Figure 5D). This study highlights the prevalence of CRECC at the species level and elucidates the molecular mechanisms underlying high levels of PB resistance. However, while multiple mutations associated with high PB resistance were identified, further research is needed to fully understand their specific regulatory mechanisms.
In conclusion, our study reveals that E. hormaechei is the predominant species, and the blaNDM-1 gene is a major contributor to carbapenem resistance in the clinical CRECC outbreak within this region. The prevalence of mcr-9 in CRECC is higher than that in CSECC. However, both mcr-9.1 and mcr-9.2 genes are generally present in a silent carriage state in ECC.
Overexpression of mcr-9 leads to a little increase in the MIC of PB, while also increasing membrane permeability and imposing fitness costs. It is worth emphasizing that mcr-9 is not responsible for high levels of resistance in PB, but does not exclude possible resistance mutations.
Mutations in the two-component system genes phoP/Q and the negatively regulated gene mgrB are key mechanisms mediating high-level PB resistance, with phoQ showing a higher likelihood than phoP in conferring this resistance. The horizontal transfer potential of insertion sequences significantly contributes to polymyxin resistance, highlighting the need for prompt clinical intervention. Therefore, monitoring IS elements could be crucial in preventing the spread of PB resistance and reducing the risk of treatment failure.
The genome sequence has been submitted to the National Center for Biotechnology Information, and the accession number is provided in the Supplementary Table S2. All primers used in this study are shown in the Supplementary Table S1.
The studies involving humans were approved by the Ethics Committee of Yongchuan Hospital of Chongqing Medical University. The studies were conducted in accordance with the local legislation and institutional requirements. Written informed consent for participation was not required from the participants or the participants’ legal guardians/next of kin in accordance with the national legislation and institutional requirements.
JimW: Formal analysis, Investigation, Methodology, Project administration, Visualization, Writing – original draft, Writing – review & editing. JZ: Conceptualization, Investigation, Methodology, Resources, Writing – review & editing. JiaW: Investigation, Methodology, Resources, Supervision, Writing – review & editing. JinW: Data curation, Methodology, Resources, Writing – review & editing. XL: Investigation, Project administration, Writing – review & editing. CW: Data curation, Investigation, Methodology, Writing – review & editing. WL: Investigation, Visualization, Writing – review & editing. YY: Data curation, Resources, Writing – review & editing. YC: Data curation, Investigation, Writing – review & editing. ML: Data curation, Resources, Writing – review & editing. YL: Formal analysis, Methodology, Writing – review & editing. KY: Formal analysis, Investigation, Writing – review & editing. XZ: Conceptualization, Funding acquisition, Writing – review & editing, Writing – original draft.
The author(s) declare that financial support was received for the research and/or publication of this article. This work was supported by Joint project of Chongqing Health Commission and Science and Technology Bureau (2023MSXM018), Yongchuan Natural Science Foundation (2021yc-jckx20053), Talent introduction project of Yongchuan Hospital of Chongqing Medical University (YJYJ202004 and YJYJ202005), Chongqing public health key specialty project, Program for Youth Innovation in Future Medicine, Chongqing Medical University (W0113), and Graduate Innovation Fund project of the Fifth Clinical College of Chongqing Medical University (YJSCX202303).
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Gen AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
The Supplementary material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fmicb.2025.1553148/full#supplementary-material
Burow, L. C., Mabbett, A. N., Borrás, L., and Blackall, L. L. (2009). Induction of membrane permeability in Escherichia coli mediated by lysis protein of the ColE7 operon. FEMS Microbiol. Lett. 298, 85–92. doi: 10.1111/j.1574-6968.2009.01705.x
Cheung, J., Bingman, C. A., Reyngold, M., Hendrickson, W. A., and Waldburger, C. D. (2008). Crystal structure of a functional dimer of the PhoQ sensor domain. J. Biol. Chem. 283, 13762–13770. doi: 10.1074/jbc.M710592200
Choi, Y., Lee, J.-Y., Lee, H., Park, M., Kang, K., Lim, S.-K., et al. (2020). Comparison of fitness cost and virulence in chromosome-and plasmid-mediated Colistin-resistant Escherichia coli. Front. Microbiol. 11:798. doi: 10.3389/fmicb.2020.00798
Doijad, S. P., Gisch, N., Frantz, R., Kumbhar, B. V., Falgenhauer, J., Imirzalioglu, C., et al. (2023). Resolving colistin resistance and heteroresistance in Enterobacter species. Nat. Commun. 14:140. doi: 10.1038/s41467-022-35717-0
Garinet, S., Fihman, V., Jacquier, H., Corvec, S., Le Monnier, A., Guillard, T., et al. (2018). Elective distribution of resistance to beta-lactams among Enterobacter cloacae genetic clusters. J. Infect. 77, 178–182. doi: 10.1016/j.jinf.2018.05.005
Girlich, D., Ouzani, S., Emeraud, C., Gauthier, L., Bonnin, R. A., Le Sache, N., et al. (2021). Uncovering the novel Enterobacter cloacae complex species responsible for septic shock deaths in newborns: a cohort study. Lancet Microbe 2, e536–e544. doi: 10.1016/S2666-5247(21)00098-7
Gong, X., Zhang, J., Su, S., Fu, Y., Bao, M., Wang, Y., et al. (2018). Molecular characterization and epidemiology of carbapenem non-susceptible Enterobacteriaceae isolated from the eastern region of Heilongjiang Province, China. BMC Infect. Dis. 18:417. doi: 10.1186/s12879-018-3294-3
Harms, A., Maisonneuve, E., and Gerdes, K. (2016). Mechanisms of bacterial persistence during stress and antibiotic exposure. Science (New York, N.Y.) 354:aaf4268. doi: 10.1126/science.aaf4268
Huang, H., Dong, N., Shu, L., Lu, J., Sun, Q., Chan, E. W.-C., et al. (2020). Colistin-resistance gene mcr in clinical carbapenem-resistant Enterobacteriaceae strains in China, 2014-2019. Emerg. Microb. Infect. 9, 237–245. doi: 10.1080/22221751.2020.1717380
Hwang, D., and Kim, H. J. (2024). Mechanisms of Polymyxin resistance in acid-adapted Enteroinvasive Escherichia coli NCCP 13719 revealed by Transcriptomics. Microorganisms 12:2549. doi: 10.3390/microorganisms12122549
Jain, C., Rodriguez-R, L. M., Phillippy, A. M., Konstantinidis, K. T., and Aluru, S. (2018). High throughput ANI analysis of 90K prokaryotic genomes reveals clear species boundaries. Nat. Commun. 9:5114. doi: 10.1038/s41467-018-07641-9
Jiang, S., Wang, X., Yu, H., Zhang, J., Wang, J., Li, J., et al. (2022). Molecular antibiotic resistance mechanisms and co-transmission of the mcr-9 and metallo-β-lactamase genes in carbapenem-resistant Enterobacter cloacae complex. Front. Microbiol. 13:1032833. doi: 10.3389/fmicb.2022.1032833
Kieffer, N., Royer, G., Decousser, J.-W., Bourrel, A.-S., Palmieri, M., Ortiz De La Rosa, J.-M., et al. (2019). Mcr-9, an inducible gene encoding an acquired Phosphoethanolamine transferase in Escherichia coli, and its origin. Antimicrob. Agents Chemother. 63, e00965–e00919. doi: 10.1128/AAC.00965-19
Kim, J. S., Yu, J. K., Jeon, S. J., Park, S. H., Han, S. H. E., Park, S. H., et al. (2021). Distribution of mcr genes among carbapenem-resistant Enterobacterales clinical isolates: high prevalence of mcr-positive Enterobacter cloacae complex in Seoul, Republic of Korea. Int. J. Antimicrob. Agents 58:106418. doi: 10.1016/j.ijantimicag.2021.106418
Lee, Y. L., Chen, H. M., Hii, I. M., and Hsueh, P. R. (2022). Carbapenemase-producing Enterobacterales infections: recent advances in diagnosis and treatment. Int. J. Antimicrob. Agents 59:106528. doi: 10.1016/j.ijantimicag.2022.106528
Lee, D.-S., and Je, J.-Y. (2013). Gallic acid-grafted-chitosan inhibits foodborne pathogens by a membrane damage mechanism. J. Agric. Food Chem. 61, 6574–6579. doi: 10.1021/jf401254g
Lefort, V., Desper, R., and Gascuel, O. (2015). FastME 2.0: a comprehensive, accurate, and fast distance-based phylogeny inference program. Mol. Biol. Evol. 32, 2798–2800. doi: 10.1093/molbev/msv150
Li, Y., Dai, X., Zeng, J., Gao, Y., Zhang, Z., and Zhang, L. (2020). Characterization of the global distribution and diversified plasmid reservoirs of the colistin resistance gene mcr-9. Sci. Rep. 10:8113. doi: 10.1038/s41598-020-65106-w
Li, L., Li, R., Qi, C., Gao, H., Wei, Q., Tan, L., et al. (2021). Mechanisms of polymyxin resistance induced by Salmonella typhimurium in vitro. Vet. Microbiol. 257:109063. doi: 10.1016/j.vetmic.2021.109063
Li, J., Nation, R. L., Turnidge, J. D., Milne, R. W., Coulthard, K., Rayner, C. R., et al. (2006). Colistin: the re-emerging antibiotic for multidrug-resistant gram-negative bacterial infections. Lancet Infect. Dis. 6, 589–601. doi: 10.1016/S1473-3099(06)70580-1
Liao, W., Cui, Y., Quan, J., Zhao, D., Han, X., Shi, Q., et al. (2022). High prevalence of colistin resistance and mcr-9/10 genes in Enterobacter spp. in a tertiary hospital over a decade. Int. J. Antimicrob. Agents 59:106573. doi: 10.1016/j.ijantimicag.2022.106573
Lu, Y., Liu, J.-H., Yue, C., Bergen, P. J., Wu, R., Li, J., et al. (2022). Overexpression of mcr-1 disrupts cell envelope synthesis and causes the dysregulation of carbon metabolism, redox balance and nucleic acids. Int. J. Antimicrob. Agents 60:106643. doi: 10.1016/j.ijantimicag.2022.106643
Mensa, B., Polizzi, N. F., Molnar, K. S., Natale, A. M., Lemmin, T., and DeGrado, W. F. (2021). Allosteric mechanism of signal transduction in the two-component system histidine kinase PhoQ. eLife 10:e73336. doi: 10.7554/eLife.73336
Mhaya, A., Bégu, D., Tounsi, S., and Arpin, C. (2020). MgrB inactivation is responsible for acquired resistance to Colistin in Enterobacter hormaechei subsp. steigerwaltii. Antimicrob. Agents Chemother. 64, e00128–e00120. doi: 10.1128/AAC.00128-20
Nang, S. C., Azad, M. A. K., Velkov, T., Zhou, Q., and Li, J. (2021). Rescuing the last-line Polymyxins: achievements and challenges. Pharmacol. Rev. 73, 679–728. doi: 10.1124/pharmrev.120.000020
Poirel, L., Jayol, A., and Nordmann, P. (2017). Polymyxins: antibacterial activity, susceptibility testing, and resistance mechanisms encoded by plasmids or chromosomes. Clin. Microbiol. Rev. 30, 557–596. doi: 10.1128/CMR.00064-16
Puljko, A., Barišić, I., Dekić Rozman, S., Križanović, S., Babić, I., Jelić, M., et al. (2024). Molecular epidemiology and mechanisms of carbapenem and colistin resistance in Klebsiella and other Enterobacterales from treated wastewater in Croatia. Environ. Int. 185:108554. doi: 10.1016/j.envint.2024.108554
Satlin, M. J., Lewis, J. S., Weinstein, M. P., Patel, J., Humphries, R. M., Kahlmeter, G., et al. (2020). Clinical and laboratory standards institute and European committee on antimicrobial susceptibility testing position statements on Polymyxin B and Colistin clinical breakpoints. Clin. Infect. Dis. 71, e523–e529. doi: 10.1093/cid/ciaa121
Stamatakis, A. (2014). RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics (Oxford, England) 30, 1312–1313. doi: 10.1093/bioinformatics/btu033
Walker, B. J., Abeel, T., Shea, T., Priest, M., Abouelliel, A., Sakthikumar, S., et al. (2014). Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS One 9:e112963. doi: 10.1371/journal.pone.0112963
Wang, Y., Liu, F., Hu, Y., Zhang, G., Zhu, B., and Gao, G. F. (2020). Detection of mobile colistin resistance gene mcr-9 in carbapenem-resistant Klebsiella pneumoniae strains of human origin in Europe. J. Infect. 80, 578–606. doi: 10.1016/j.jinf.2019.12.016
Wick, R. R., Judd, L. M., Gorrie, C. L., and Holt, K. E. (2017). Unicycler: resolving bacterial genome assemblies from short and long sequencing reads. PLoS Comput. Biol. 13:e1005595. doi: 10.1371/journal.pcbi.1005595
Wu, L., Xu, T., Ji, Y., Song, J., Wang, F., Huang, J., et al. (2022). Occurrence and characteristics of Mcrs among gram-negative Bacteria causing bloodstream infections of infant inpatients between 2006 and 2019 in China. Microbiol. Spectr. 10:e0193821. doi: 10.1128/spectrum.01938-21
Wu, Y., Zhao, J., Li, Z., Liu, X., Hu, Y., Zhang, F., et al. (2023). Within-host acquisition of colistin-resistance of an NDM-producing Klebsiella quasipneumoniae subsp. similipneumoniae strain through the insertion sequence-903B-mediated inactivation of mgrB gene in a lung transplant child in China. Front. Cell. Infect. Microbiol. 13:1153387. doi: 10.3389/fcimb.2023.1153387
Xie, J., Chen, Y., Cai, G., Cai, R., Hu, Z., and Wang, H. (2023). Tree visualization by one table (tvBOT): a web application for visualizing, modifying and annotating phylogenetic trees. Nucleic Acids Res. 51, W587–W592. doi: 10.1093/nar/gkad359
Xu, T., Xue, C.-X., Chen, Y., Huang, J., Wu, W., Lu, Y., et al. (2022a). Frequent convergence of mcr-9 and carbapenemase genes in Enterobacter cloacae complex driven by epidemic plasmids and host incompatibility. Emerg. Microb. Infect. 11, 1959–1972. doi: 10.1080/22221751.2022.2103456
Xu, T., Xue, C.-X., Huang, J., Wu, J., Chen, R., and Zhou, K. (2022b). Emergence of an epidemic hypervirulent clone of Enterobacter hormaechei coproducing mcr-9 and carbapenemases. Lancet Microbe 3, e474–e475. doi: 10.1016/S2666-5247(22)00122-7
Yan, Z., Ju, X., Zhang, Y., Wu, Y., Sun, Y., Xiong, P., et al. (2024). Analysis of the transmission chain of carbapenem-resistant Enterobacter cloacae complex infections in clinical, intestinal and healthcare settings in Zhejiang province, China (2022-2023). Sci. Total Environ. 920:170635. doi: 10.1016/j.scitotenv.2024.170635
Yang, T.-Y., Wang, S.-F., Lin, J.-E., Griffith, B. T. S., Lian, S.-H., Hong, Z.-D., et al. (2020). Contributions of insertion sequences conferring colistin resistance in Klebsiella pneumoniae. Int. J. Antimicrob. Agents 55:105894. doi: 10.1016/j.ijantimicag.2020.105894
Yoon, E.-J., Kim, D., and Jeong, S. H. (2019). Bloodstream infections and carbapenem-resistant Enterobacteriaceae in South Korea. Lancet Infect. Dis. 19, 931–932. doi: 10.1016/S1473-3099(19)30431-1
Zelendova, M., Papagiannitsis, C. C., Sismova, P., Medvecky, M., Pomorska, K., Palkovicova, J., et al. (2023). Plasmid-mediated colistin resistance among human clinical Enterobacterales isolates: national surveillance in the Czech Republic. Front. Microbiol. 14:1147846. doi: 10.3389/fmicb.2023.1147846
Zeng, H., Tan, Y., Su, J., Gao, F., Lei, T., and Liang, B. (2023). Co-occurrence of blaNDM-1, rmt C, and mcr-9 in multidrug-resistant Enterobacter kobei strain isolated from an infant with urinary tract infection. J. Glob. Antimicrob. Resist. 33, 221–226. doi: 10.1016/j.jgar.2023.03.011
Zhang, X., Qu, F., Jia, W., Huang, B., Shan, B., Yu, H., et al. (2021). Polymyxin resistance in carbapenem-resistant Enterobacteriaceae isolates from patients without polymyxin exposure: a multicentre study in China. Int. J. Antimicrob. Agents 57:106262. doi: 10.1016/j.ijantimicag.2020.106262
Zhang, Y. W., Wang, Q., Yin, Y. Y., Chen, H. B., Jin, L. Y., Gu, B., et al. (2018). Epidemiology of Carbapenem-resistant Enterobacteriaceae infections: report from the China CRE network. Antimicrob. Agents Chemother. 62, e01882–e01817. doi: 10.1128/AAC.01882-17
Zhou, H., Wang, S., Wu, Y., Dong, N., Ju, X., Cai, C., et al. (2022). Carriage of the mcr-9 and mcr-10 genes in clinical strains of the Enterobacter cloacae complex in China: a prevalence and molecular epidemiology study. Int. J. Antimicrob. Agents 60:106645. doi: 10.1016/j.ijantimicag.2022.106645
Zhu, Y., Galani, I., Karaiskos, I., Lu, J., Aye, S. M., Huang, J., et al. (2019). Multifaceted mechanisms of colistin resistance revealed by genomic analysis of multidrug-resistant Klebsiella pneumoniae isolates from individual patients before and after colistin treatment. J. Infect. 79, 312–321. doi: 10.1016/j.jinf.2019.07.009
Keywords: Enterobacter cloacae complex, mcr-9 , polymyxin, CRECC, epidemiology, antibiotic resistance
Citation: Wu J, Zhang J, Wang J, Wang J, Liang X, Wei C, Long W, Yang Y, Chen Y, Liao M, Liang Y, Yu K and Zhang X (2025) Insertion sequences in mgrB and mutations in two-component system genes confer high polymyxin resistance to carbapenem-resistant Enterobacter cloacae complex strains. Front. Microbiol. 16:1553148. doi: 10.3389/fmicb.2025.1553148
Received: 30 December 2024; Accepted: 03 March 2025;
Published: 17 March 2025.
Edited by:
John Osei Sekyere, University of Pretoria, South AfricaReviewed by:
Shin-ichi Yokota, Sapporo Medical University, JapanCopyright © 2025 Wu, Zhang, Wang, Wang, Liang, Wei, Long, Yang, Chen, Liao, Liang, Yu and Zhang. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Xiaoli Zhang, am1zenhsMTIzQDE2My5jb20=
†These authors have contributed equally to this work
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.