
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Microbiol. , 25 March 2025
Sec. Antimicrobials, Resistance and Chemotherapy
Volume 16 - 2025 | https://doi.org/10.3389/fmicb.2025.1550894
Background: Invasive candidiasis (IC) is an increasingly common, expensive, and potentially fatal infection. However, IC caused by multiple Candida species is rarely reported in China. Herein, we revealed a complex IC caused by multiple Candida species, comprising the rare C. norvegensis, C. albicans, C. glabrata, and C. tropicalis. The resistance mechanism of azole and echinocandin resistance were explored further.
Methods: The isolates were confirmed using internal transcribed spacer (ITS) sequencing. The resistance mechanisms were investigated using PCR-based sequencing, quantitative real-time reverse transcription PCR, and rhodamine 6G efflux quantification.
Results: Antifungal susceptibility testing showed this complex infection was associated with cross-resistance to azole and echinocandin drugs. For C. glabrata, the acquired echinocandin resistance was likely caused by a novel mutational pattern (1,3-beta-D-glucan synthase subunits FKS1-S629P and FKS2-W1497stop) while the acquired azole resistance in C. glabrata RJ05 was related to complex mechanisms including enhanced efflux activity, pleiotropic drug resistance 1 (PDR1) mutation, and increased expression of Candida drug resistance 1 (CDR1) and CDR2. Additionally, the azole resistance of C. tropicalis was caused by two lanosterol 14-alpha demethylase (ERG11) mutations: Y132F and S154F.
Conclusion: Our study revealed a case of clinically complex, multiple Candida invasive infections, further uncovering the resistance mechanisms to azoles and echinocandins. These findings provide valuable references for the diagnosis and treatment of invasive candidiasis (IC) in clinical practice.
Candida species can colonize multiple parts of the human body, causing diseases when the body’s immune system is impaired or autoimmune barriers are disrupted (Giannella et al., 2024; Iliev et al., 2024). Invasive fungal diseases (IFDs) have become a major threat to public health, with high incidence and fatality rates, resulting in approximately 3.8 million patient deaths each year (Bassetti et al., 2017; Lass-Flörl et al., 2024; Sedik et al., 2024). Among them, invasive candidiasis (IC) is an increasingly common, expensive, and potentially fatal infection. Previously, IC was mainly caused by Candida albicans. However, in the past decade, the contribution of non-C. albicans Candida species (NCACs) to IC has increased (Bays et al., 2024; Salimi et al., 2024). More importantly, infections by rare Candida species, such as C. norvegensis, have displayed a rising trend over recent years (Pinho et al., 2024).
Antifungal resistance constitutes an important issue in the treatment of Candida infections. Triazoles like fluconazole (FLU), voriconazole (VRC), itraconazole (ITR), and posaconazole (POS) are the most commonly used antifungals. These drugs via bind to and inhibit the key ergosterol biosynthesis pathway enzyme, 14-α-demethylase, which is encoded by ERG11. However, resistance to azoles has become very common (Meneghello et al., 2024). To combat this, it is recommended to use echinocandins, including caspofungin (CAS), micafungin (MIF), and anidulafungin (ANF), as first-line IC treatments, because they target the catalytic subunit of β-1,3-glucan synthase (encoded by Candida gene FKS1, except in C. glabrata, in which it is encoded by FKS1 and FKS2)(Pappas et al., 2016). This enzyme is crucial for the biosynthesis of β-1,3-glucan, a component of the cell wall of fungi. However, the emergence of echinocandin resistance is a concerning trend, with C. glabrata isolates showing resistance rates of 10.3% in a cancer center in United States (Farmakiotis et al., 2014). However, in China, the resistance rate is below 1% (Hou et al., 2017; Song et al., 2020).
Azole resistance among Candida spp. results from ATP-binding cassette (ABC) family multidrug transporter overexpression, including that of CDR1, CDR2, and SNQ2 (Marie and White, 2009; Abbes et al., 2013). These transporters are regulated by the zinc finger transcription factor PDR1. Therefore, functional mutations in PDR1 can lead to reduced susceptibility to azoles (Ferrari et al., 2009). Meanwhile, the overexpression of drug targets (lanosterol 14α-demethylase, encoded by ERG11) and mutations are also common (Hull et al., 2012; Nguyen et al., 2024; Tanwar et al., 2024). What’s more, alternative mechanisms, like biofilm formation and mitochondrial defects, have also been reported recently (Ferrari et al., 2011; Nguyen et al., 2024). In contrast, the mechanisms underlying echinocandin resistance are relatively straightforward, primarily involving mutations in the hotspot regions of FKS1 and FKS2 encoding subunits of the 1,3-β-D-glucan synthase complex (Zimbeck et al., 2010; Arastehfar et al., 2020). However, mutations outside the hotspot regions also have been reported (Hou et al., 2019).
Recently, IC with multiple Candida species infection has gained increased attention, particularly in patients whose immune systems are compromised, e.g., those living with HIV/AIDS or receiving chemotherapy (García-Salazar et al., 2024; Sadeghi et al., 2024). A comprehensive analysis over 10 years showed that 2.8% of patients had mixed fungemia, indicating the prevalence still remained low (Ferngren et al., 2024). However, mixed candidiasis could complicate diagnosis and treatment, because different Candida species might exhibit varying degrees of virulence and resistance to antifungal therapies. According to a previous study (Soriano-Abarca et al., 2024), the overexpression of HWP1 (encoding virulence factor hyphal wall protein 1) and ASL3 (encoding virulence factor agglutinin-like sequence 3) under mixed growth conditions of C. albicans and non-C. albicans indicated a synergistic relationship and increased mycelial growth and adhesion. The interplay between multiple species might lead to more severe infections. Therefore, comprehending the antifungal susceptibility profiles and resistance mechanisms of each type of Candida species is of paramount importance for clinical therapy and prognosis of IC.
However, IC caused by multiple Candida species is not reported frequently in China. Moreover, the antifungal resistance mechanisms in these pathogens have not been explored in depth and are thus mostly unknown. Herein, we present eight clinical isolates from four different Candida species, including the rare C. norvegensis, which were isolated from one patient. This complex infection was associated with cross-resistance to azole and echinocandin drugs. Moreover, the molecular mechanisms of the multidrug resistance phenotypes were clarified.
The eight strains were isolated from the infecting mixed fungemia population using single-colony isolation techniques. Briefly, clinical samples were cultured on chromogenic Candida agar plates (CHROMagar™, Aubervilliers, France) and incubated for 48 h at 37°C. The resultant colonies were preliminarily examined for their colony color following the supplier’s guidelines to identify the Candida spp. Then the individual colonies with distinct morphological characteristics were picked and subcultured for purification.
Species identification was further confirmed utilizing internal transcribed spacer (ITS) sequencing and Matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS) (Vitek MS, bioMérieux, Marcy l’Etoile, France). Primers ITS1 and ITS4 (ITS1:5′-TCCGTAGGTGAACCTGCG-3′; ITS4:5′-TCCTCCGCTTATTGATATGC-3′) were employed to amplify and sequence the ITS regions (Zhang et al., 2014). BLAST searches1 were employed for species identification (>99% similarity).
The Ethics Committee of Ruijin Hospital, Shanghai Jiaotong University School of Medicine provided approval for this study (approval number: MX-B4621R).
A colorimetric microdilution panel, DL-96Fungus (Dier Biotech, Guangzhou, China) was employed to test the in vitro susceptibility of the isolates to eight drugs (amphotericin B, 5-flucytosine, micafungin, caspofungin, fluconazole, voriconazole, posaconazole and itraconazole) according to the supplier’s guidelines. The isolates were grown at 37°C for 24 h to ascertain the minimum inhibitory concentrations (MICs). The quality control species comprised Candida parapsilosis ATCC 22019 and Candida krusei ATCC 6258. C. parapsilosis ATCC 22019 is a well-established QC strain with known susceptibility to azoles and echinocandins, making it suitable for verifying the performance of these antifungals. C. krusei ATCC 6258 is intrinsically resistant to fluconazole, serving as a control for azole resistance testing. Resistance breakpoints for Candida species (C. albicans, C. glabrata, and C. tropicalis) in this study were defined according to the guidelines of the Clinical and Laboratory Standards Institute (CLSI; M27M44S) (Clinical and Laboratory Standards Institute, 2022b). The MIC values of the two control strains were consistently within the CLSI-defined acceptable ranges for each antifungal agent tested, confirming the validity of our susceptibility results. For those isolates without CLSI-determined clinical breakpoints, the epidemiological cut-off values (ECVs) were used to define the isolates as wild-type (WT) or non-wild-type (non-WT) (Clinical and Laboratory Standards Institute, 2022a). However, because of the scarcity of C. norvegensis, there are no clinical breakpoints or ECVs for this species.
DNA extraction and purification from overnight suspension were performed employing a TIANamp Yeast DNA Kit (Tiangen Biotech, Beijing, China) in accordance with the supplier’s manual. To identify mutations associated with antifungal resistance in the resistant isolates, the open reading frames (ORFs) of CgERG11, CgPDR1, CgFKS1, and CgFKS2 in C. glabrata and CtERG11 in C. tropicalis were amplified using PrimeSTAR® HS DNA Polymerase (Takara, Shiga, Japan) and specific primers (Supplementary Table S1) (Jiang et al., 2013; Wang et al., 2021). Genewiz (Beijing, China) determined the sequences of all the amplicons in both directions. The sequences were compared with those of reference strains and Snapgene 7.2 software (GSL Biotech LLC, Boston, MA, United States) was employed to identify single nucleotide polymorphisms (SNPs). The obtained nucleotide sequences were compared with those deposited in GenBank (CgERG11, accession no. L40389.1; CgPDR1, accession no. NC005967; CgFKS1, accession no. HM366440.1; CgFKS2, accession no. HM366442.1; and CtERG11. accession no. M23673).
The fluorescent dye rhodamine 6G resembles the membrane transporters of azoles in certain Candida spp. (Yao et al., 2019). Therefore, rhodamine 6G efflux pump activity of Candida spp. was determined utilizing an ultraviolet-visual (UV–Vis) spectrophotometer (Thermo Scientific, Waltham, MA, United States) (Fattouh et al., 2024). Yeast peptone dextrose (YPD) medium was used to adjust overnight-cultured isolates to 5 × 107 cells/mL. The Candida cells were centrifuged and rinsed thrice using glucose-free phosphate-buffered saline (PBS). Subsequently, the cells were depleted of energy by incubation for 4 h at 30°C with continuous shaking. Next, we added rhodamine 6G (Sigma-Aldrich, St Louis, MO, United States; final concentration = 10 μM) to the cells and shook them at for 2 h at 30°C. Following three rinses in cold sterile PBS, the Candida cells were reconstituted in 10 mL of PBS, followed by incubation at 30°C with gentle shaking. At 15, 30, 45, and 60 min, we took 1 mL of supernatant, which was centrifuged at 9,000 × g for 2 min to remove the Candida cells. The optical density (OD) values at 527 nm were determined for the supernatant employing a UV–Vis spectrophotometer. We generated a standard curve to determine the concentration of rhodamine 6G in the supernatant. Average values were determined from triplicated experiments. The negative control was treated with sodium azide (NaN₃), an ATPase inhibitor, to block energy-dependent efflux pump activity. A known high-efflux Candida strain, which was previously stored in the lab, was used as the positive control.
The isolates were grown in YPD medium to the mid exponential phase [OD at 600 nm (OD600) = 0.4–0.5] and then a Yeast RNAiso Reagent Kit (Takara, Shiga, Japan) was used to extract total RNA according to the supplier’s guidelines. Then, a PrimeScript RT reagent kit (Takara) was employed to reverse transcribe RNA to cDNA. Each reaction can reverse transcribe 1 μg RNA. The expression levels of CgCDR1, CgCDR2, CgSNQ2, CgPDR1, CgERG11, and CtERG11 were determined utilizing the cDNA as templates and a SYBR Premix Ex Taq Kit (Takara, Shiga, Japan) on a 7,300 real-time PCR System (Applied Biosystems, Shanghai, China), as detailed in a previous publication. The cDNA content in per RT-qPCR reaction is 75 ng/20 μL. The qPCR reactions were carried out thus: 95°C denaturation for 45 s, then 40 cycles of 5 s at 95°C and 45 s at 60°C. Table 1 shows the primers used for qPCR (Jiang et al., 2013; Yao et al., 2019). Target gene quantification employed the 2−△△CT method (Livak and Schmittgen, 2001). The expression of ACT1 gene (encoding actin1) was detected as an internal control. The RT-qPCR assays were carried out in triplicate. No-template control reactions were performed for each primer set by replacing the cDNA template with nuclease-free water. To rule out genomic DNA contamination, we conducted no-reverse transcription control reactions by omitting the reverse transcriptase enzyme during cDNA synthesis. To confirm the specificity and efficiency of our RT-qPCR assay, positive controls were included for each target gene using corresponding clinical strains with confirmed highly expression of the respective gene.
Whole genome sequencing was performed for two C. glabrata strains RJ04 and RJ05. The genome DNA was extracted as described previously in section 2.3. Library preparation and next-generation sequencing was conducted by Illumina PE150. Raw sequencing data was assessed for quality using FastQC. Then the short sequencing reads were de novo assembled into contiguous sequences (contigs). FastANI v1.342 was applied for fast alignment-free computation of whole-genome Average Nucleotide Identity (ANI). Snippy v4.6.0 was used to detect the Single Nucleotide Polymorphisms (SNPs) based on the whole genome. C. glabrata RJ04 was used as reference strain.
Data were analyzed and visualized using GraphPad Prism 10 software (GraphPad Inc., La Jolla, CA, United States). Two-way ANOVA was employed to assess statistical difference of extracellular concentrations of rhodamine 6G relative to that in the control. Multiple unpaired t-tests were used to test the significance of expression level change. p < 0.05 is considered as a significant difference.
Figure 1 shows all the Candida species isolated from an 86-year-old female patient who was treated at Ruijin Hospital, Shanghai Jiao Tong University School of Medicine. She was admitted to the emergency room suffering from abdominal pain, emesis, and unconsciousness. An abdominal computed tomography scan showed choledocholithiasis with obstructive dilation and pneumatosis. She was then diagnosed with obstructive suppurative cholangitis and transferred to the ICU for further treatment.
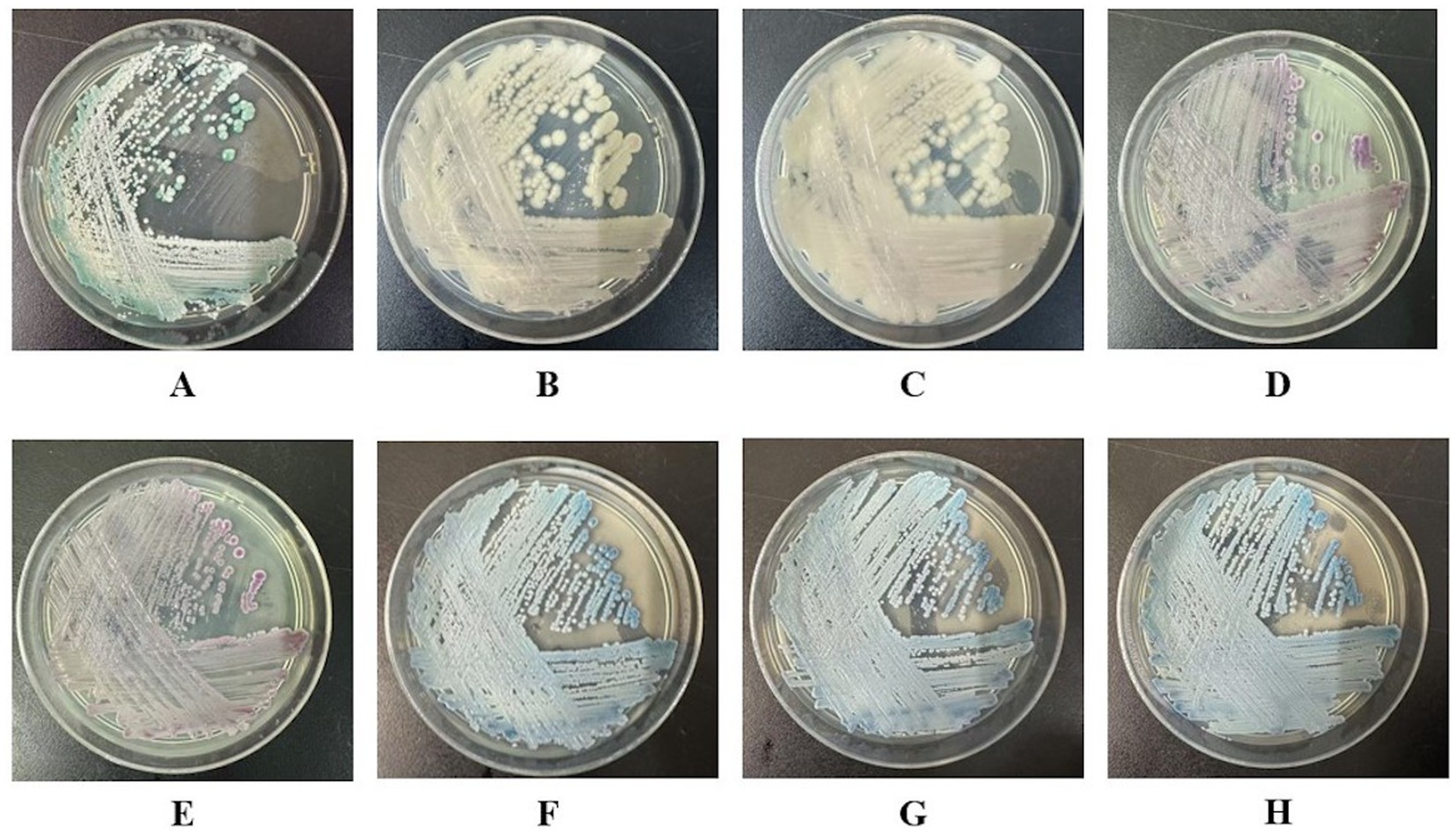
Figure 1. Candida colony morphology on the chromogenic Candida agar plates (CHROMagar™, Aubervilliers, France) of all the eight strains RJ01 ~ RJ08 collected from the same patient. (A) C. albicans RJ01; (B,C) C. norvegensis RJ02, RJ03; (D,E) C. glabrata RJ04, RJ05; (F–H) C. tropicalis RJ06, RJ07, RJ08.
As can be seen from Figure 2, at the beginning of treatment, C. albicans (RJ01) and C. norvegensis (RJ02, RJ03) were isolated from bile specimens. The patient was treated with CAS (50 mg qd) and FLU (200 mg qd) successively. After treatment, C. albicans and C. norvegensis were no longer detected. Instead, C. glabrata (RJ04, RJ05) isolates were detected in bile specimens. Then, the treatment switched to VRC (150 mg q12h). During this treatment period, C. tropicalis (RJ06, RJ07, RJ08) isolates were consistently detected in bile specimens. The detection of multiple Candida species from sterile fluid bile suggested that the patient might have IC. Meanwhile, Acinetobacter baumannii was repeatedly detected in her sputum and bile specimens, and later, blood infection with Staphylococcus epidermidis also appeared. The patient died after treatment failed because of septic shock caused by a severe infection throughout her body.
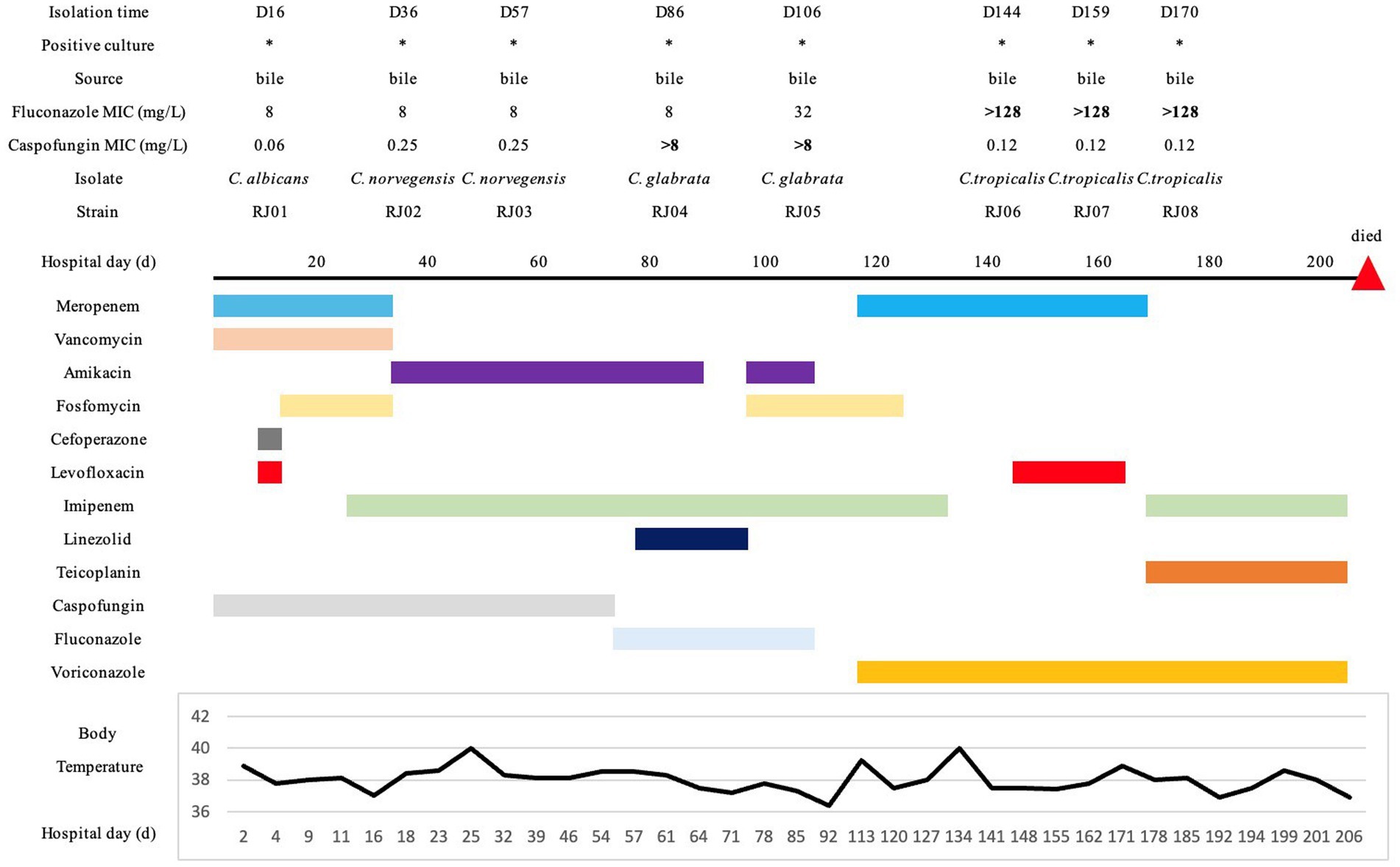
Figure 2. Medical history chart. The specimen sources, drug susceptibility, and isolation times of all the isolates were marked on the chart. All the medication histories and treatment durations were indicated in different colors. Additionally, the temperature changes during hospitalization were displayed at the bottom of the chart.
Table 2 shows the result of antifungal susceptibility testing of all eight isolates in this study toward the eight tested antifungal drugs. C. norvegensis (RJ02, RJ03) showed relatively low MIC values toward most of the antifungal drugs. However, there were no clinical breakpoints or epidemiological cut-off values for C. norvegensis because of its rarity. Additionally, C. albicans (RJ01) showed susceptibility or non-WT to all the tested drugs. For C. glabrata, isolate RJ04 displayed high-level resistance to MIF and CAS (MIC > 8 μg/mL). However, C. glabrata RJ05 showed an additional non-WT to azoles (POS and ITR) after treatment (MIC > 16 μg/mL). Moreover, all C. tropicalis isolates (RJ06, RJ07, and RJ08) demonstrated significant resistance to FLU (MIC > 128 μg/mL).
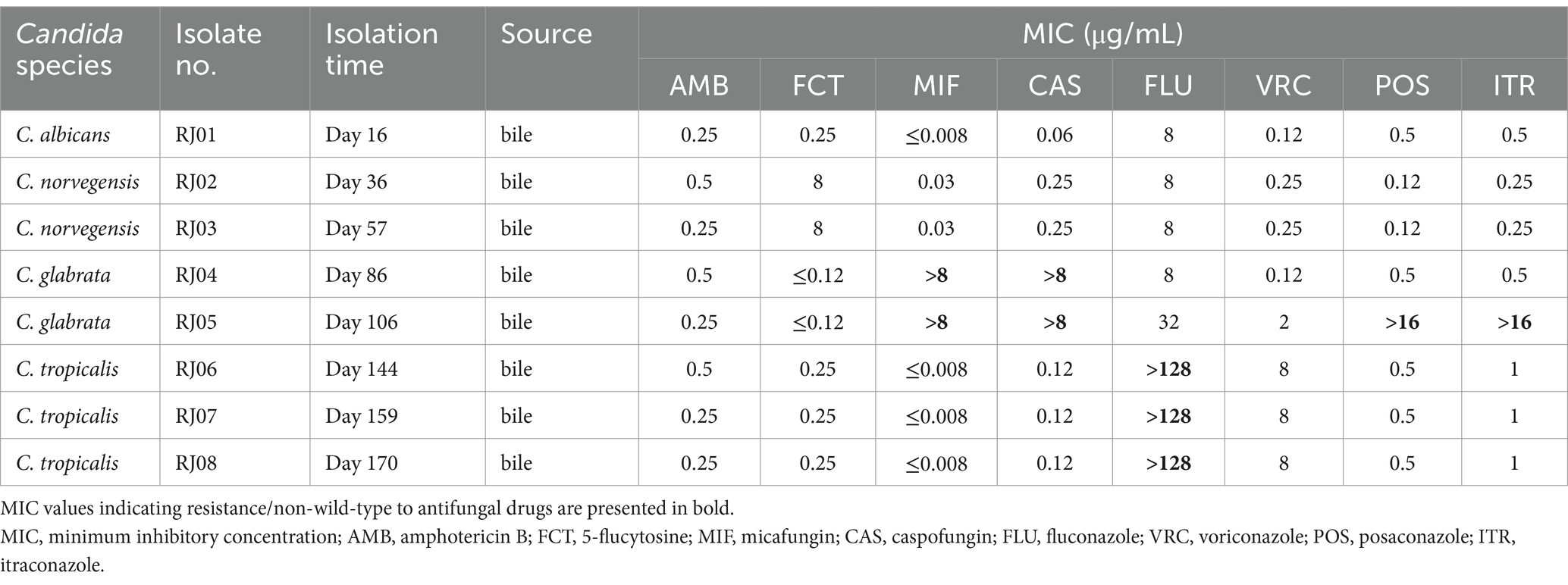
Table 2. Antifungal susceptibility results of all the eight clinical strains isolated from the patient sequentially.
The two C. glabrata strains RJ04 and RJ05 exhibited significant differences in their susceptibility to azoles. Based on whole-genome data, we performed a homology alignment between the two strains. The assembled genomes were subjected to cross-comparison analysis, revealing an overall similarity of 99.9973%. RJ04 and RJ05 were proven to be highly homologous, and might be two mutant strains derived from the same maternal strain. Further, using RJ04 as the reference genome, we identified 14 SNP variants, 3 complex variants, 1 deletion variant, and 1 MNP variant between the two samples at the read level. The detailed mutation sites are listed in Table 3.
A UV–Vis spectrophotometer was used to determine rhodamine 6G efflux. C. glabrata and C. tropicalis acquire resistance to azole antifungals via efflux pump overexpression (Jiang et al., 2013; Yao et al., 2019). Rhodamine 6G was introduced into the cells of C. glabrata and C. tropicalis, and then its efflux into the extracellular fluid of isolates RJ05, RJ06, RJ07, RJ08, and the control strains was quantified in three biological replicates. The controls comprised C. glabrata RJC1 and C. tropicalis RJC2. They were preserved in our laboratory and were both susceptible and wild-type to all the tested drugs, including azoles.
As shown in Figure 3A, at 15, 30, 45, and 60 min, C. tropicalis RJ05 exhibited significantly higher extracellular concentrations of rhodamine 6G relative to that in the control. According to two-way ANOVA analysis, the p-value was 0.0166, revealing that RJ05 has significantly enhanced efflux activity, thereby contributing to resistance against azole drugs. However, at all timepoints, the concentrations of rhodamine 6G in C. tropicalis RJ06, RJ07, and RJ08 were comparable to that of the control strain, and the statistical analysis yielded a p-value of 0.3339, indicating that the efflux pump did not contribute to the azole resistance in these C. tropicalis isolates (Figure 3B).
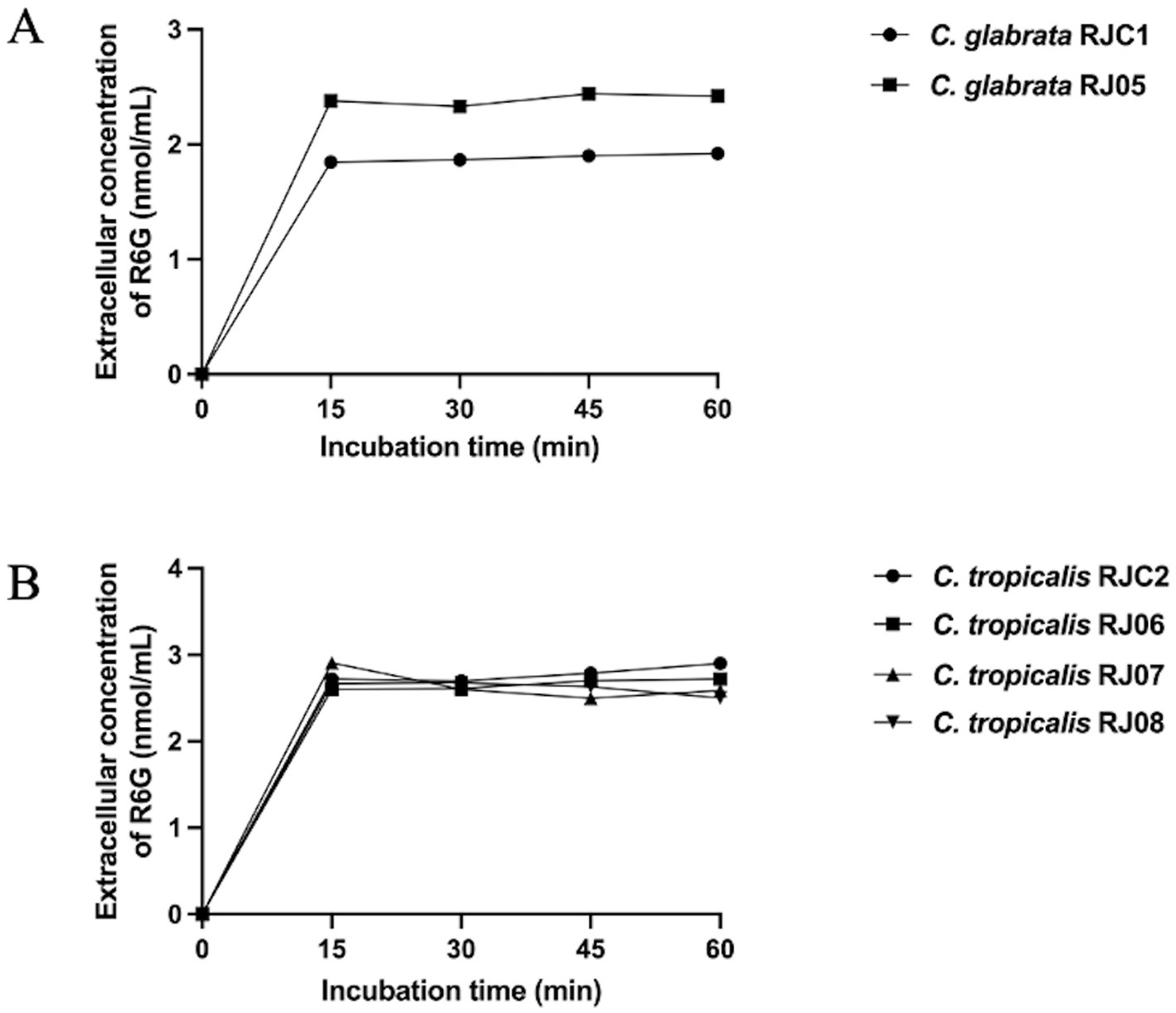
Figure 3. Quantification of extracellular concentration of R6G (nmol/mL) during 60 min. (A) Each data point for the control strain C. glabrata RJC1 (black circles) and C. glabrata RJ05 (black squares). A significant increase in extracellular R6G (p = 0.0166) was detected in C. glabrata RJ05 compared with the control strain. (B) Each data point for the control strain C. tropicalis RJC2 (black circles) and test strains C. tropicalis RJ06 (black squares), C. tropicalis RJ07 (black triangle) and C. tropicalis RJ08 (black inverted triangle). No significant increase in extracellular R6G (p = 0.3339) was detected in test strains compared with the control strain.
Analysis of the DNA sequence of the CgERG11 ORFs in C. glabrata showed that there were no mutations in this gene in isolates RJ04 and RJ05 in comparison with the CgERG11 sequence deposited in GenBank (accession number: L40389.1). However, CgPDR1 in isolate RJ05 encoded a protein with four missense mutations (S76P, V91I, L98S, and T143P). Furthermore, in C. glabrata RJ04 and RJ05, a mutation in FKS1 resulting in an S629P substitution, together with a premature stop codon in FKS2 (W1497stop), were observed, which mediated high-level resistance to micafungin and caspofungin (MIC > 8 μg/mL). The FKS mutation pattern has not been reported previously (Supplementary Figures S1A–C).
Analysis of the CtERG11 ORF DNA sequences from C. tropicalis RJ06, RJ07, and RJ08 identified two missense mutations, Y132F and S154F (Supplementary Figure S1D).
According to the results of RT-qPCR in Figure 4A, CgCDR1 (21.75-fold vs. 1-fold, p = 0.000164) and CgCDR2 (8.507-fold vs. 1-fold, p = 0.001162) expression levels were significantly higher in C. glabrata RJ05 than in the control strain C. glabrata RJC1, while C. glabrata RJ04 showed no significant difference (p > 0.05). In addition, the expression levels of CgSNQ2, CgERG11, and CgPDR1 in C. glabrata RJ04 and RJ05 exhibited no significant difference to those in the control isolate (p > 0.05). Moreover, the CtERG11 expression levels in three C. tropicalis isolates resistant to fluconazole (RJ06, RJ07, RJ08) were not increased significantly compared with those in C. tropicalis RJC2 (p > 0.05) (Figure 4B).
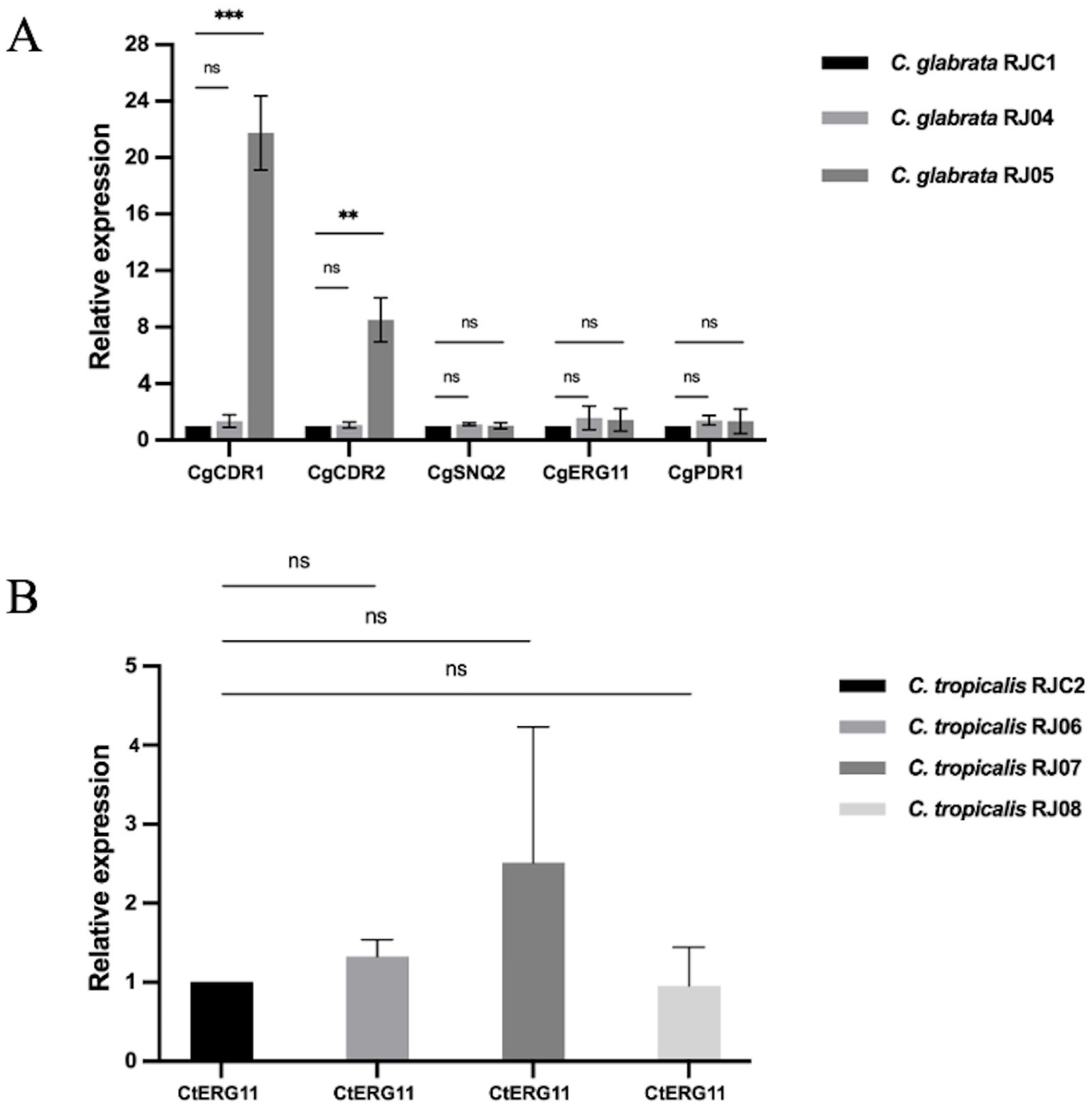
Figure 4. The relative expression level of resistance-related genes. Data presented as means ± SE (error bars) from biological triplicates with technical triplicates. Multiple unpaired t-tests were used to test the significance of expression level change. p < 0.05 is considered to indicate a significant difference. (A) For C. glabrata, the relative expression of each target gene (CgCDR1, CgCDR2, CgSNQ2, CgERG11, CgPDR1) was compared to that of ACT1 as internal control. C. glabrata RJC1 was used as external reference. A significant increase was observed in the CgCDR1 (21.75-fold vs. 1-fold, p = 0.000164) and CgCDR2 (8.507-fold vs. 1-fold, p = 0.001162) from C. glabrata RJ05. No significant increase in the two genes was found in RJ04 (1.346-fold vs. 1-fold, p = 0.241881; 1.058-fold vs. 1-fold, p = 0.661751). Additionally, no significant change was observed in the CgSNQ2 (1.118-fold vs. 1-fold, p = 0.131953; 1.016-fold vs. 1-fold, p = 0.899801), CgERG11 (1.575-fold vs. 1-fold, p = 0.304047; 1.435-fold vs. 1-fold, p = 0.391636), CgPDR1 (1.393-fold vs. 1-fold, p = 0.114514; 1.33-fold vs. 1-fold; p = 0.552051). (B) For C. tropicalis, the relative expression of target gene CtERG11 was comparable to that of ACT1 as internal control. C. tropicalis RJC2 was used as external reference. No significant increase was observed in C. tropicalis RJ06, RJ07 and RJ08 (1.33-fold vs. 1-fold, p = 0.1149; 2.52-fold vs. 1-fold, p = 0.2670; 0.95-fold vs. 1-fold, p = 0.8875). **p < 0.01, ***p < 0.001, “ns” denotes no significance.
IC caused by multiple Candida species has attracted the attention of epidemiologists in recent years due to its high mortality rate. Relevant reports remain scarce in China. Besides, clarifying the antifungal drug resistance mechanism of each Candida strain is of crucial significance for clinical treatment.
In this work, we revealed a rare clinical case of IC caused by multiple Candida species in a patient with obstructive suppurative cholangitis. In the ICU, patients with IC might present as deep-seated candidiasis with candidemia, deep-seated candidiasis or candidemia (Azim and Ahmed, 2024). For deep-seated candidiasis, the most frequently encountered form is intra-abdominal candidiasis (Vergidis et al., 2016). In this study, all the strains were isolated from bile samples from a sterile site, thus meeting the diagnostic criteria. The known risk factors for IC include ICU admission, placement of central venous catheters, treatment with broad-spectrum antibiotics, abdominal surgery, and immunosuppression (Lass-Flörl et al., 2024), which were consistent with the patient’s medical history.
Low susceptibility to certain antifungal drugs is a significant and pressing issue to be resolved for the treatment of IC with multiple Candida species. Notably, in mixed infections, different resistant phenotypes of Candida species could complement each other, ultimately leading to multidrug resistance. In C. glabrata, azole resistance is primarily driven by ATP-binding cassette (ABC) family multidrug transporter overexpression, including that of CDR1, CDR2, and SNQ2 (Bennett et al., 2004; Torelli et al., 2008), which is in accordance with the resistance mechanism in C. glabrata RJ05 in this study. The azole resistance was mediated by complex mechanisms comprising increased efflux ability, PDR1 mutation, and increased expression of CDR1 and CDR2. Notably, azole resistance was not likely to be mediated by the substitutions in PDR1 (S76P, V91I, L98S, and T143P), because the mutations were not gain-of-function mutations, as reported previously (Hou et al., 2018). Moreover, antifungal resistance can be acquired by C. glabrata during therapy (Borst et al., 2005). In this study, C. glabrata RJ05 exhibited non-WT to posaconazole and itraconazole after in vivo fluconazole exposure. Echinocandin resistance in C. glabrata has been associated with mutations in FKS1 and FKS2 (Pfaller et al., 2012). Mutations in FKS2 are more common than the ones in FKS1 (Zimbeck et al., 2010; Alexander et al., 2013). However, a novel mutational pattern mediating high level echinocandin-resistance (MIC > 8 μg/mL) was found in this study. The mutations were both identified in C. glabrata RJ04 and RJ05 strains: an S629P mutation in FKS1 HS1 region and a premature stop codon (W1497stop) in FKS2, which have not been reported previously. Similarly, Hou et al. (2019) has demonstrated that both mutations in FKS1 (W508stop) and FKS2 (E655K) might lead to higher MICs to echinocandins than single mutations. This revealed that more than one mutation within the same isolates may potentially mediate a high-level echinocandin-resistant phenotype in C. glabrata. However, more experiments are required to validate this hypothesis. As for C. tropicalis, the azole antifungal target cytochrome P450 lanosterol 14a-demethylase encoded by ERG11 play a rather vital role in azole resistance. Two mutations, Y132F and S154F in ERG11, were demonstrated to be responsible for fluconazole resistance in C. tropicalis RJ06, RJ07, and RJ08 in this study. This resistance mechanism has been experimentally proven in the previous study and interestingly, these two mutations always occurred together (Jiang et al., 2013).
In vivo evolution appears to be quite interesting. Homology analysis showed that RJ05 has 19 SNPs compared to RJ04, suggesting that these two strains are homologous. Previous literature (Misas et al., 2024) has reported that a threshold of 96 SNPs is considered indicative of homology between two strains. Therefore, RJ05 may have evolved from RJ04 under the sustained pressure of fluconazole. In fact, there have been previous reports of in vivo evolution of C. glabrata resistance to fluconazole, mainly involving the upregulation of CgCDR1, CgCDR2, and CgERG11 expression (Redding et al., 2003). However, in our study, RJ05 exhibits simultaneous resistance to both echinocandins and azoles, which complicates clinical treatment. It is noteworthy that RJ05 has multiple protein mutations compared to RJ04, including in proteins such as KES1, Negative Growth Regulatory Protein NGR1, and Flocculation Protein FLO9. Whether these proteins play a role in the development of azole resistance remains to be further investigated.
C. norvegensis is an uncommon Candida species. Recently, invasive infections by rare Candida species have been on the rise (Pinho et al., 2024). A recent report showed that C. norvegensis could display decreased anti-fungal susceptibility (Stavrou et al., 2020). In the clinic, the interpretation of symptoms and options for treatment for uncommon Candida spp. are a challenge because we lack clinical breakpoints to determine their susceptibility profile.
Thus, mixed candidiasis poses significant challenges for clinical treatment, especially regarding uncommon Candida species, because of clinicians’ lack of experience of these species. An observational cohort study in Europe showed that the decreased attributable mortality might be the result of improved outcomes for bloodstream infections involving C. parapsilosis and C. albicans, while bloodstream infections involving other Candida species still exhibit higher attributable mortality rates (Salmanton-García et al., 2024). Most significantly, mixed candidiasis mediates both azole and echinocandin resistance, resulting in a situation where no effective drugs are available. This also encourages the development of new antifungal drugs.
In summary, this study aids the clinical treatment of IC and deepens our knowledge regarding the mechanisms underlying Candida resistance to multiple drugs. Although multiple fungal infections are still relatively rare (Ferngren et al., 2024), the resulting multi-drug resistance and pathogenicity should be taken seriously. Future studies involving mixed populations from multiple patients will be necessary to validate our observations and provide a broader perspective on the diversity and clinical significance of mixed candidiasis. An in-depth comprehension of the dynamics of these co-infections, including their epidemiology, risk factors, and underlying mechanisms is essential to develop targeted treatment strategies and improve patient management in clinical settings.
The datasets presented in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found in the article/Supplementary material.
The studies involving humans were approved by the Ethics Committee of Ruijin Hospital, Shanghai Jiaotong University School of Medicine (approval number: MX-B4621R). The studies were conducted in accordance with the local legislation and institutional requirements. Written informed consent for participation was not required from the participants or the participants’ legal guardians/next of kin in accordance with the national legislation and institutional requirements.
S-JH: Conceptualization, Formal analysis, Investigation, Methodology, Writing – original draft, Writing – review & editing. Y-HS: Formal analysis, Investigation, Methodology, Writing – original draft, Writing – review & editing, Conceptualization. GL: Methodology, Writing – original draft. J-YL: Methodology, Writing – review & editing. J-TZ: Formal analysis, Investigation, Writing – review & editing. L-LW: Formal analysis, Investigation, Writing – review & editing. M-JX: Conceptualization, Funding acquisition, Resources, Supervision, Writing – review & editing.
The author(s) declare that financial support was received for the research and/or publication of this article. This work was supported by the National Natural Science Foundation of China [#81871706], Shanghai Municipal Health Commission [#202240205], [#201840227], and [#201740069], Natural Science Foundation of Shanghai [#22ZR1439800] and [#15ZR1426900], Shanghai Huangpu District Health Commission [#HLM202303], the Program of Shanghai Key Specialty [#ZK2012A21], and Excellent Youth of Huangpu District of Shanghai [#RCPY1407].
The authors would like to thank reviewers for helpful comments.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The authors declare that no Gen AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
The Supplementary material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fmicb.2025.1550894/full#supplementary-material
Abbes, S., Mary, C., Sellami, H., Michel-Nguyen, A., Ayadi, A., and Ranque, S. (2013). Interactions between copy number and expression level of genes involved in fluconazole resistance in Candida glabrata. Front. Cell. Infect. Microbiol. 3:74. doi: 10.3389/fcimb.2013.00074
Alexander, B. D., Johnson, M. D., Pfeiffer, C. D., Jiménez-Ortigosa, C., Catania, J., Booker, R., et al. (2013). Increasing echinocandin resistance in Candida glabrata: clinical failure correlates with presence of FKS mutations and elevated minimum inhibitory concentrations. Clin. Infect. Dis. 56, 1724–1732. doi: 10.1093/cid/cit136
Arastehfar, A., Daneshnia, F., Salehi, M., Yaşar, M., Hoşbul, T., Ilkit, M., et al. (2020). Low level of antifungal resistance of Candida glabrata blood isolates in Turkey: fluconazole minimum inhibitory concentration and FKS mutations can predict therapeutic failure. Mycoses 63, 911–920. doi: 10.1111/myc.13104
Azim, A., and Ahmed, A. (2024). Diagnosis and management of invasive fungal diseases in non-neutropenic ICU patients, with focus on candidiasis and aspergillosis: a comprehensive review. Front. Cell. Infect. Microbiol. 14:1256158. doi: 10.3389/fcimb.2024.1256158
Bassetti, M., Garnacho-Montero, J., Calandra, T., Kullberg, B., Dimopoulos, G., Azoulay, E., et al. (2017). Intensive care medicine research agenda on invasive fungal infection in critically ill patients. Intensive Care Med. 43, 1225–1238. doi: 10.1007/s00134-017-4731-2
Bays, D. J., Jenkins, E. N., Lyman, M., Chiller, T., Strong, N., Ostrosky-Zeichner, L., et al. (2024). Epidemiology of invasive candidiasis. Clin. Epidemiol. 16, 549–566. doi: 10.2147/clep.S459600
Bennett, J. E., Izumikawa, K., and Marr, K. A. (2004). Mechanism of increased fluconazole resistance in Candida glabrata during prophylaxis. Antimicrob. Agents Chemother. 48, 1773–1777. doi: 10.1128/aac.48.5.1773-1777.2004
Borst, A., Raimer, M. T., Warnock, D. W., Morrison, C. J., and Arthington-Skaggs, B. A. (2005). Rapid acquisition of stable azole resistance by Candida glabrata isolates obtained before the clinical introduction of fluconazole. Antimicrob. Agents Chemother. 49, 783–787. doi: 10.1128/aac.49.2.783-787.2005
Clinical and Laboratory Standards Institute (2022a). Epidemiological cutoff values for antifungal susceptibility testing M57S. 4th Edn. USA: Clinical and Laboratory Standards Institute.
Clinical and Laboratory Standards Institute (2022b). Performance standards for antifungal susceptibility testing of yeasts M27M44S. 3rd Edn. USA: Clinical and Laboratory Standards Institute.
Farmakiotis, D., Tarrand, J. J., and Kontoyiannis, D. P. (2014). Drug-resistant Candida glabrata infection in cancer patients. Emerg. Infect. Dis. 20, 1833–1840. doi: 10.3201/eid2011.140685
Fattouh, N., Khalaf, R. A., and Husni, R. (2024). Candida glabrata hospital isolate from Lebanon reveals micafungin resistance associated with increased chitin and resistance to a cell-surface-disrupting agent. J. Glob. Antimicrob. Resist. 37, 62–68. doi: 10.1016/j.jgar.2024.02.012
Ferngren, G., Yu, D., Unalan-Altintop, T., Dinnétz, P., and Özenci, V. (2024). Epidemiological patterns of candidaemia: a comprehensive analysis over a decade. Mycoses 67:e13729. doi: 10.1111/myc.13729
Ferrari, S., Ischer, F., Calabrese, D., Posteraro, B., Sanguinetti, M., Fadda, G., et al. (2009). Gain of function mutations in CgPDR1 of Candida glabrata not only mediate antifungal resistance but also enhance virulence. PLoS Pathog. 5:e1000268. doi: 10.1371/journal.ppat.1000268
Ferrari, S., Sanguinetti, M., De Bernardis, F., Torelli, R., Posteraro, B., Vandeputte, P., et al. (2011). Loss of mitochondrial functions associated with azole resistance in Candida glabrata results in enhanced virulence in mice. Antimicrob. Agents Chemother. 55, 1852–1860. doi: 10.1128/aac.01271-10
García-Salazar, E., Benavidez-López, S., Bonifaz, A., Hernández-Mendoza, E. A., Ramírez-Magaña, X., Reyes-Montes, M. D. R., et al. (2024). Fungal coinfection/superinfection in COVID-19 patients in a tertiary hospital in Mexico. Biomedica 44, 328–339. doi: 10.7705/biomedica.7251
Giannella, M., Lanternier, F., Dellière, S., Groll, A. H., Mueller, N. J., Alastruey-Izquierdo, A., et al. (2024). Invasive fungal disease in the immunocompromised host: changing epidemiology, new antifungal therapies, and management challenges. Clin. Microbiol. Infect. 31, 29–36. doi: 10.1016/j.cmi.2024.08.006
Hou, X., Healey, K. R., Shor, E., Kordalewska, M., Ortigosa, C. J., Paderu, P., et al. (2019). Novel FKS1 and FKS2 modifications in a high-level echinocandin resistant clinical isolate of Candida glabrata. Emerg. Microbes Infect. 8, 1619–1625. doi: 10.1080/22221751.2019.1684209
Hou, X., Xiao, M., Chen, S. C., Kong, F., Wang, H., Chu, Y. Z., et al. (2017). Molecular epidemiology and antifungal susceptibility of Candida glabrata in China (august 2009 to July 2014): a multi-center study. Front. Microbiol. 8:880. doi: 10.3389/fmicb.2017.00880
Hou, X., Xiao, M., Wang, H., Yu, S. Y., Zhang, G., Zhao, Y., et al. (2018). Profiling of PDR1 and MSH2 in Candida glabrata bloodstream isolates from a multicenter study in China. Antimicrob. Agents Chemother. 62:e00153-18. doi: 10.1128/aac.00153-18
Hull, C. M., Parker, J. E., Bader, O., Weig, M., Gross, U., Warrilow, A. G., et al. (2012). Facultative sterol uptake in an ergosterol-deficient clinical isolate of Candida glabrata harboring a missense mutation in ERG11 and exhibiting cross-resistance to azoles and amphotericin B. Antimicrob. Agents Chemother. 56, 4223–4232. doi: 10.1128/aac.06253-11
Iliev, I. D., Brown, G. D., Bacher, P., Gaffen, S. L., Heitman, J., Klein, B. S., et al. (2024). Focus on fungi. Cell 187, 5121–5127. doi: 10.1016/j.cell.2024.08.016
Jiang, C., Dong, D., Yu, B., Cai, G., Wang, X., Ji, Y., et al. (2013). Mechanisms of azole resistance in 52 clinical isolates of Candida tropicalis in China. J. Antimicrob. Chemother. 68, 778–785. doi: 10.1093/jac/dks481
Lass-Flörl, C., Kanj, S. S., Govender, N. P., Thompson, G. R. 3rd, Ostrosky-Zeichner, L., and Govrins, M. A. (2024). Invasive candidiasis. Nat. Rev. Dis. Primers 10:20. doi: 10.1038/s41572-024-00503-3
Livak, K. J., and Schmittgen, T. D. (2001). Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25, 402–408. doi: 10.1006/meth.2001.1262
Marie, C., and White, T. C. (2009). Genetic basis of antifungal drug resistance. Curr. Fungal Infect. Rep. 3, 163–169. doi: 10.1007/s12281-009-0021-y
Meneghello, S., Bernabè, G., Di Pietra, G., Di Sopra, S., Del Vecchio, C., Cattelan, A. M., et al. (2024). Prevalence, species distribution and resistance of Candidemia in pediatric and adult patients in a Northeast Italy university hospital. J. Fungi (Basel) 10:707. doi: 10.3390/jof10100707
Misas, E., Seagle, E., Jenkins, E. N., Rajeev, M., Hurst, S., Nunnally, N. S., et al. (2024). Genomic description of acquired fluconazole-and echinocandin-resistance in patients with serial Candida glabrata isolates. J. Clin. Microbiol. 62:e0114023. doi: 10.1128/jcm.01140-23
Nguyen, B. V. G., Tran, L. X. T., Vo, T. H., Nguyen-Ha, A. T., Le, M. T., and Nguyen, P. V. (2024). Biofilm formation and drug susceptibility of biofilm Candida spp. clinically isolated from nasopharyngeal cancer patients in Vietnam. Iran. J. Microbiol. 16, 698–707. doi: 10.18502/ijm.v16i5.16806
Pappas, P. G., Kauffman, C. A., Andes, D. R., Clancy, C. J., Marr, K. A., Ostrosky-Zeichner, L., et al. (2016). Clinical practice guideline for the Management of Candidiasis: 2016 update by the Infectious Diseases Society of America. Clin. Infect. Dis. 62, e1–e50. doi: 10.1093/cid/civ933
Pfaller, M. A., Castanheira, M., Lockhart, S. R., Ahlquist, A. M., Messer, S. A., and Jones, R. N. (2012). Frequency of decreased susceptibility and resistance to echinocandins among fluconazole-resistant bloodstream isolates of Candida glabrata. J. Clin. Microbiol. 50, 1199–1203. doi: 10.1128/jcm.06112-11
Pinho, S., Miranda, I. M., and Costa-de-Oliveira, S. (2024). Global epidemiology of invasive infections by uncommon Candida species: a systematic review. J. Fungi (Basel) 10:558. doi: 10.3390/jof10080558
Redding, S. W., Kirkpatrick, W. R., Saville, S., Coco, B. J., White, W., Fothergill, A., et al. (2003). Multiple patterns of resistance to fluconazole in Candida glabrata isolates from a patient with oropharyngeal candidiasis receiving head and neck radiation. J. Clin. Microbiol. 41, 619–622. doi: 10.1128/jcm.41.2.619-622.2003
Sadeghi, A., Houri, H., Lotfali, E., Ghadirzadeh, E., and Rajabnia, M. (2024). Biliary co-infection by multidrug-resistant Candida glabrata and Candida albicans in a case of pancreatic cancer with cholangitis: a case report and review of literature. Med. Mycol. Case Rep. 43:100625. doi: 10.1016/j.mmcr.2023.100625
Salimi, M., Javidnia, J., Abastabar, M., Mobayen, M. R., Moslemi, A., Rahimzadeh, G., et al. (2024). Multi-state evaluation of Candida infections in burn patients. Mycoses 67:e13788. doi: 10.1111/myc.13788
Salmanton-García, J., Cornely, O. A., Stemler, J., Barać, A., Steinmann, J., Siváková, A., et al. (2024). Attributable mortality of candidemia - results from the ECMM Candida III multinational European observational cohort study. J. Infect. 89:106229. doi: 10.1016/j.jinf.2024.106229
Sedik, S., Wolfgruber, S., Hoenigl, M., and Kriegl, L. (2024). Diagnosing fungal infections in clinical practice: a narrative review. Expert Rev. Anti-Infect. Ther. 22, 935–949. doi: 10.1080/14787210.2024.2403017
Song, Y., Chen, X., Yan, Y., Wan, Z., Liu, W., and Li, R. (2020). Prevalence and antifungal susceptibility of pathogenic yeasts in China: a 10-year retrospective study in a teaching hospital. Front. Microbiol. 11:1401. doi: 10.3389/fmicb.2020.01401
Soriano-Abarca, M., Tapia, J. C., Cáceres-Valdiviezo, M. J., Morey-León, G., Fernández-Cadena, J., Díaz-Cevallos, L., et al. (2024). Virulence-related genes expression in planktonic mixed cultures of Candida albicans and non-Albicans Candida species. Microb. Physiol. 34, 243–254. doi: 10.1159/000540991
Stavrou, A. A., Pérez-Hansen, A., Lackner, M., Lass-Flörl, C., and Boekhout, T. (2020). Elevated minimum inhibitory concentrations to antifungal drugs prevail in 14 rare species of candidemia-causing Saccharomycotina yeasts. Med. Mycol. 58, 987–995. doi: 10.1093/mmy/myaa005
Tanwar, S., Kalra, S., and Bari, V. K. (2024). Insights into the role of sterol metabolism in antifungal drug resistance: a mini-review. Front. Microbiol. 15:1409085. doi: 10.3389/fmicb.2024.1409085
Torelli, R., Posteraro, B., Ferrari, S., La Sorda, M., Fadda, G., Sanglard, D., et al. (2008). The ATP-binding cassette transporter-encoding gene CgSNQ2 is contributing to the CgPDR1-dependent azole resistance of Candida glabrata. Mol. Microbiol. 68, 186–201. doi: 10.1111/j.1365-2958.2008.06143.x
Vergidis, P., Clancy, C. J., Shields, R. K., Park, S. Y., Wildfeuer, B. N., Simmons, R. L., et al. (2016). Intra-abdominal candidiasis: the importance of early source control and antifungal treatment. PLoS One 11:e0153247. doi: 10.1371/journal.pone.0153247
Wang, Q., Li, Y., Cai, X., Li, R., Zheng, B., Yang, E., et al. (2021). Two sequential clinical isolates of Candida glabrata with multidrug-resistance to Posaconazole and Echinocandins. Antibiotics (Basel) 10:1217. doi: 10.3390/antibiotics10101217
Yao, D., Chen, J., Chen, W., Li, Z., and Hu, X. (2019). Mechanisms of azole resistance in clinical isolates of Candida glabrata from two hospitals in China. Infect. Drug Resist. 12, 771–781. doi: 10.2147/idr.S202058
Zhang, L., Xiao, M., Wang, H., Gao, R., Fan, X., Brown, M., et al. (2014). Yeast identification algorithm based on use of the Vitek MS system selectively supplemented with ribosomal DNA sequencing: proposal of a reference assay for invasive fungal surveillance programs in China. J. Clin. Microbiol. 52, 572–577. doi: 10.1128/jcm.02543-13
Keywords: invasive candidiasis, Candida glabrata , Candida norvegensis , Candida tropicalis , antifungal susceptibility testing, azole resistance, echinocandin resistance
Citation: Huang S-J, Song Y-H, Lv G, Liu J-Y, Zhao J-T, Wang L-L and Xiang M-J (2025) Emergence of invasive candidiasis with multiple Candida species exhibiting azole and echinocandin resistance. Front. Microbiol. 16:1550894. doi: 10.3389/fmicb.2025.1550894
Received: 24 December 2024; Accepted: 04 March 2025;
Published: 25 March 2025.
Edited by:
Renátó Kovács, University of Debrecen, HungaryReviewed by:
Mohsan Ullah Goraya, Huaqiao University, ChinaCopyright © 2025 Huang, Song, Lv, Liu, Zhao, Wang and Xiang. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Ming-Jie Xiang, bWp4aWFuZzEyMzQ1NkAxMjYuY29t
†These authors have contributed equally to this work and share first authorship
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.