
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
REVIEW article
Front. Microbiol. , 18 March 2025
Sec. Microbial Symbioses
Volume 16 - 2025 | https://doi.org/10.3389/fmicb.2025.1550749
Microbiomes combine the species and activities of all microorganisms living together in a specific habitat. They comprise unique ecological niches with influences that scale from local to global ecosystems. Understanding the connectivity of microbiomes across academic disciplines is important to help mitigate global climate change, reduce food insecurity, control harmful diseases, and ensure environmental sustainability. However, most publications refer to individual microbiomes, and those integrating two or more related disciplines are rare. This review examines the multifaceted benefits of microbiomes across agriculture, food manufacturing and preservation, the natural environment, human health, and biocatalyst processes. Plant microbiomes, by improving plant nutrient cycling and increasing plant abiotic and biotic stress resilience, have increased crop yields by over 20%. Food microbiomes generate approximately USD 30 billion to the global economy through the fermented food industry alone. Environmental microbiomes help detoxify pollutants, absorb more than 90% of heavy metals, and facilitate carbon sequestration. For human microbiomes, an adult person can carry up to 38 trillion microbes which regulate well being, immune functionality, reproductive function, and disease prevention. Microbiomes are used to optimize biocatalyst processes which produce bioenergy and biochemicals; bioethanol production alone is valued at over USD 83 billion p.a. However, challenges, including knowledge gaps, engaging indigenous communities, technical limitations, regulatory considerations, the need for interdisciplinary collaboration, and ethical issues, must be overcome before the potential for microbiomes can be more effectively realized.
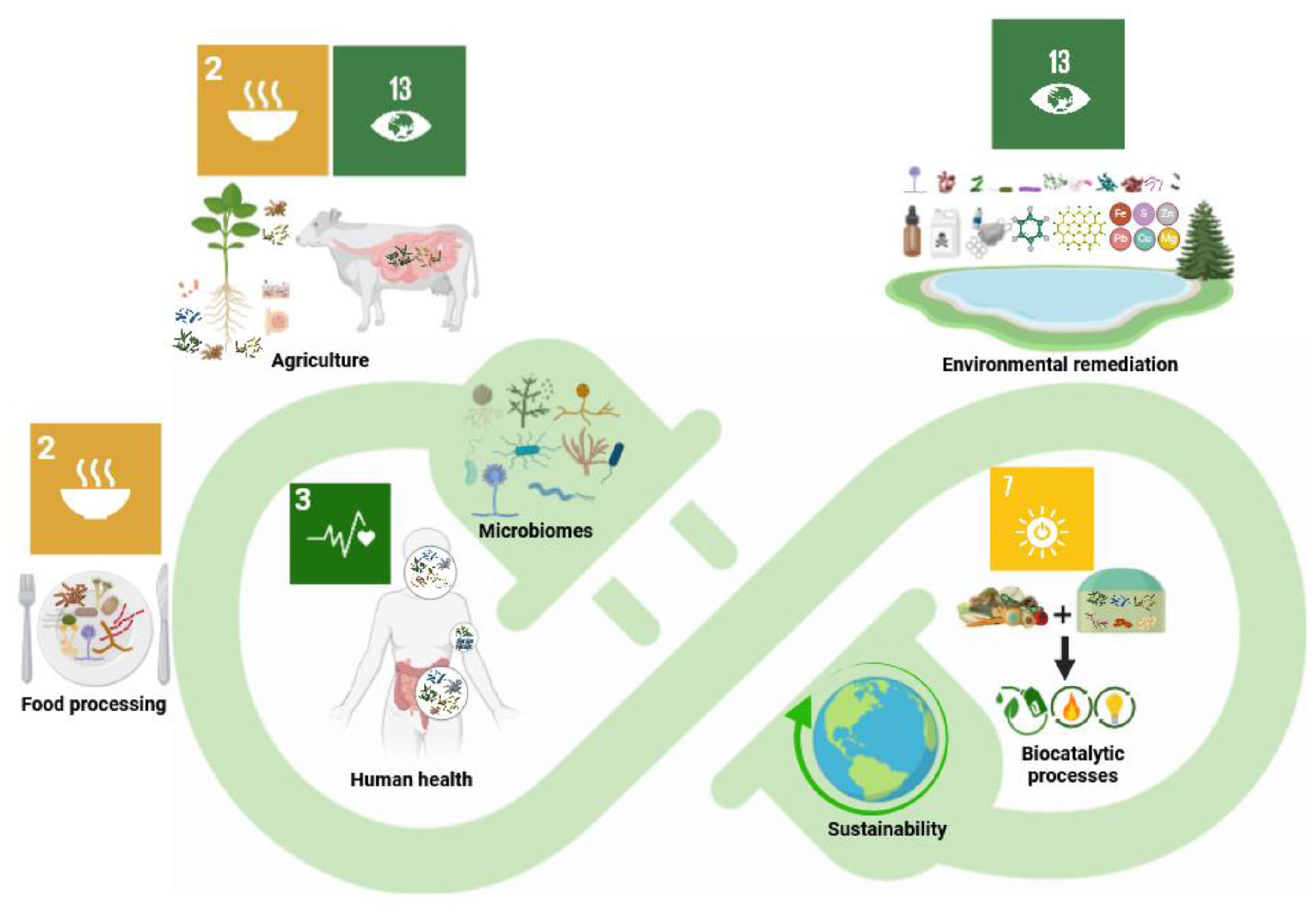
Graphical Abstract. Multifaceted benefits of microbiomes across academic disciplines for a sustainable world. Unlocking the full potential of microbiomes will contribute to achieving sustainable development goals such as zero hunger, good health and well being, affordable and clean energy, and mitigation of climate change.
A microbiome is the combination of all the microorganisms living together in a particular habitat. The expression of their collective activities and interactions creates unique ecological niches that impact surrounding ecosystems (Berg et al., 2020). All plants and animals establish symbiotic associations with an extensive diversity of microorganisms, forming functional integrated super-organismal units termed holobionts (the combination of an eukaryotic organism and its microbiome comprising a single ecological unit) (Berg et al., 2017; Poole, 2017; Zilber-Rosenberg and Rosenberg, 2008; Baedke et al., 2020; Gilbert, 2014; Brinker et al., 2019). The concept of symbiosis is now considered the rule rather than the exception and is emerging as one of the central principles of contemporary biology (Gilbert, 2014).
In both natural and productive (e.g., agricultural) environments, plant microbiomes improve plant nutrient acquisition and protect plants against biotic and abiotic stresses. They enhance growth, productivity, and biodiversity, delivering a wide range of ecosystem functions (Salas-Gonzalez et al., 2021; Mendoza-Mendoza et al., 2024; Chieb and Gachomo, 2023; Gibert et al., 2019). Microbiomes are key players in many aspects of food manufacturing and preservation, contributing to the development of various fermented foods and extending the shelf life of many food products. People consume food shaped by microbiomes daily (Siddiqui et al., 2023), deriving value from distinct flavor profiles, prolonged preservation, and nutritional advantages conferred by microorganisms in the context of fermented foods (e.g., cheese, pickled vegetables and fruit, tempeh, and yogurt) and beverages (e.g., beer, cider, whiskey, and wine) (Louw et al., 2023; Piao et al., 2015; Pinto et al., 2015). Humans are profoundly influenced by their microbiomes. An adult human weighing around 70 kilograms may have up to 38 trillion microbes from 3,000 species (Sender et al., 2016), all influencing health and wellbeing. For example, the gut, skin, oral, and vaginal microbiomes influence digestion, nutrient absorption, immune system regulation, and reproductive functions (Zheng et al., 2020; Coelho et al., 2021; Ma et al., 2023). Current findings suggest that the gut microbiota influence mood and mental health, including anxiety, depression, bipolar disorder, schizophrenia, and obsessive-compulsive disorder, among others (Gupt et al., 2024). Microbiomes help create toxic-free ecosystems, such as in bioremediation for environmental cleanup and wastewater treatment (Rani et al., 2019; Ayilara and Babalola, 2023; Abatenh et al., 2017; Xu et al., 2014; Hu et al., 2021; Cheng et al., 2022; Ruan et al., 2024). A summary of key microbiome taxa and their roles across academic disciplines is presented in Table 1.
Soil and environmental microbiomes improve soil health, degrade phytotoxicity, and enhance plant nutrient availability (Table 1). These collectively contribute to sustainable food production. This toxin-free environment nurtures plant microbiomes that support plant growth, builds stress resilience, and increases plant yield. The plant microbiome transforms it into highly nutritious and healthy diets for humans and animals. When humans consume plants, they consume plant holobionts, impacting their gut microbiomes and functioning, playing a key role in maintaining well being and immunity. Waste generated by this process and other agricultural byproducts is transformed into bioenergy or biofuel by biocatalyst microbiomes. Finally, the final waste turns into biofertilizer, which increases plant productivity and improves microbiome diversity in the environment and soil. This interconnected process is indefinitely cyclical. Recent One Health research has recognized the important of this interconnected nature of microbiomes for soil, plants, animal, and human health (van Bruggen et al., 2019; Law et al., 2024; Ma et al., 2023). However, these studies tended to emphasize the role of microbiomes for health-related outcomes rather than explicit conceptualization of their cyclical nature in sustaining ecosystem functions. Therefore, this article introduces the “circular microbiome system” concept, which refers to the interconnectedness of microbiomes across life science disciplines in a virtuous and interdependent cycle. For example, the inclusion of biocatalyst microbiomes that transform waste into bioenergy or biofertilizer and the use of indigenous knowledge to complete the loop. Despite this inherent interconnection, microbiome systems are often studied in isolation. To fully leverage the potential of microbiomes, we need a holistic approach that formally integrates interconnected domains, recognizing the microbiome as a unified, interconnected system that sustains life on Earth. As microbiomes play a fundamental role in optimizing sustainability, the understanding and utilization of microbiomes offer tremendous opportunities for improving food production, environmental remediation, human health, and biocatalyst processes. This review examines the multifaceted application of microbiomes involved in the sustainability of diverse ecosystems and identifies challenges that may delay the potential use of microbiomes.
Microbiomes play a significant role in enhancing nutrient cycling, suppressing diseases and insect pests, promoting stress tolerance, regulating phytohormones (Figure 1), and facilitating food processing (Sessitsch et al., 2023; Ayilara and Babalola, 2023; Munir et al., 2022; Kulkarni et al., 2024; Bhat et al., 2022; Solomon et al., 2023; Molefe et al., 2023; Suman et al., 2022; Figure 2). For instance, plant microbiomes help increase staple crop yield by up to 20% under field conditions (Trivedi et al., 2017).
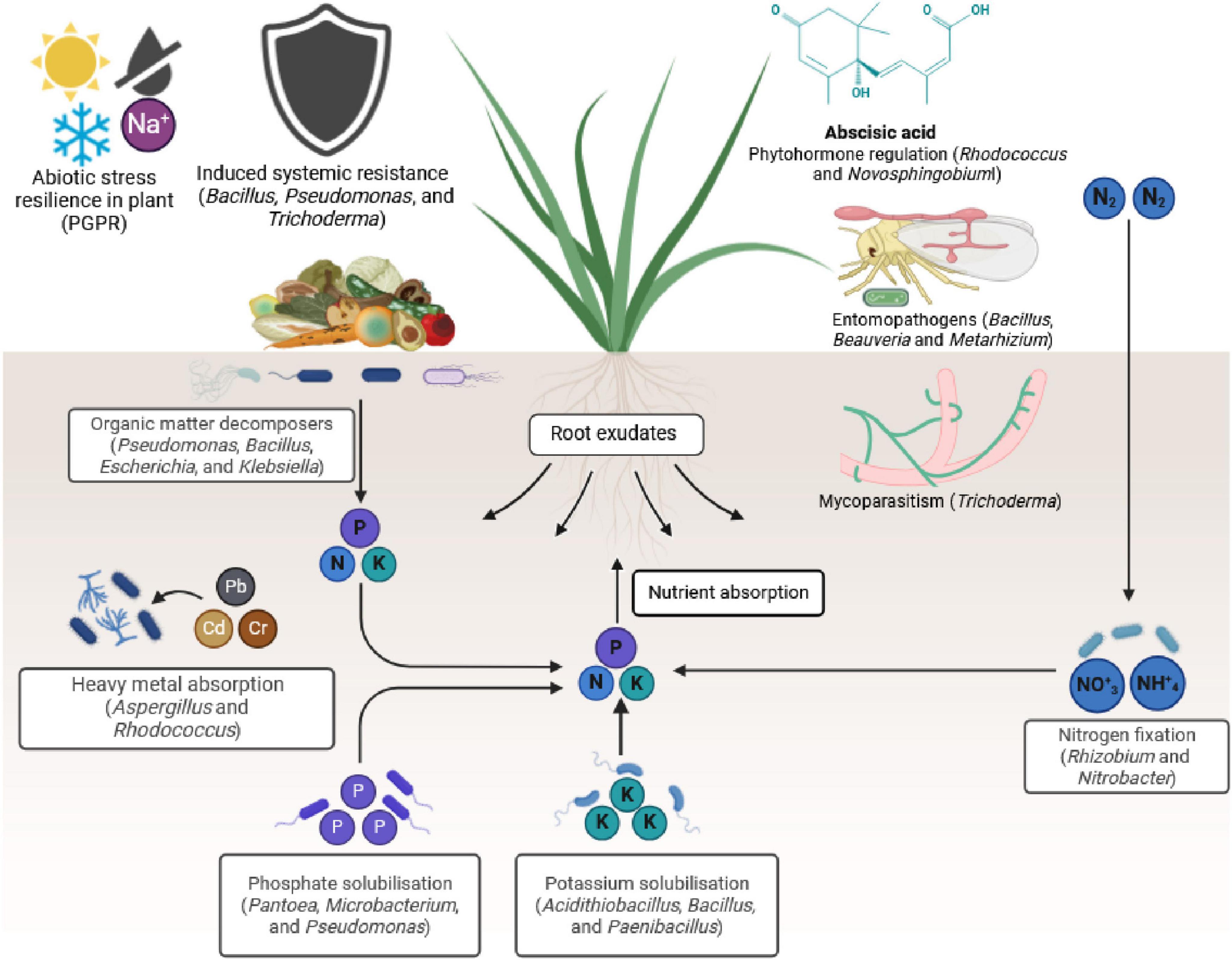
Figure 1. Mechanisms by which plant microbiomes can improve plant health for sustainable agricultural systems. They: (1) play a role in nutrient cycling, (2) are involved in pest and disease suppression, (3) induce plant resistance to abiotic stresses, (4) regulate phytohormones to respond to stresses, and (5) form beneficial biofilms (Figure was created using Biorender at https://app.biorender.com/).
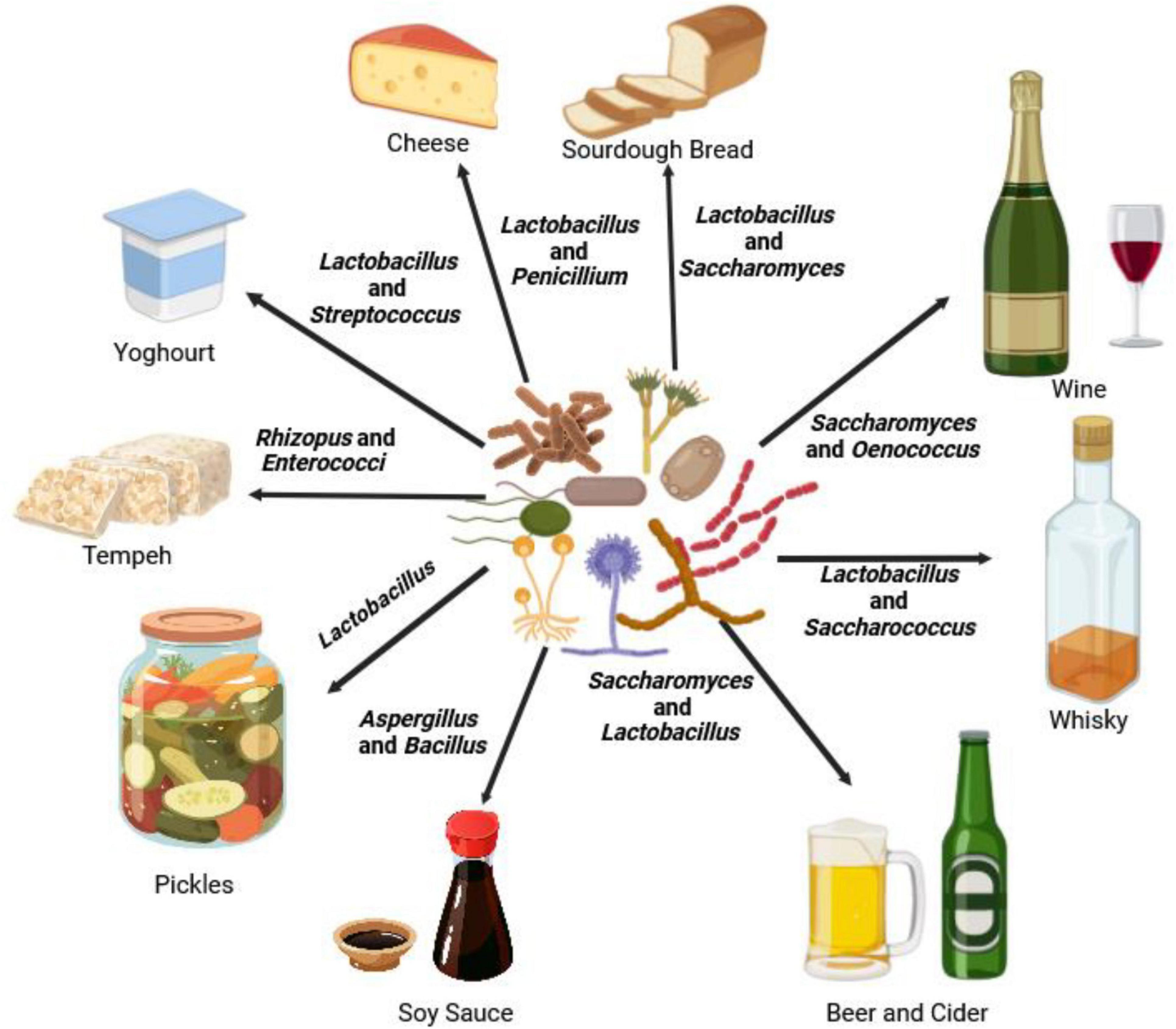
Figure 2. An illustration of how microbiomes such as Aspergillus, Bacillus (Yee et al., 2021), Enterococci (Moreno et al., 2002), Lactobacillus (Giraffa et al., 2010), Oenococcus (du Toit et al., 2011), Penicillium (Ropars et al., 2020), Rhizopus (Nout and Aidoo, 2011), Saccharomyces (Walker and Stewart, 2016), and Streptococcus (Mena and Aryana, 2012) are used for food (bread, cheese, pickles, soy sauce, tempeh, and yogurt) and beverages (beer, cider, wine, and whiskey) processing.
Microbiomes are integral to ecosystem nutrient transport, mineralization, and solubilization. For example, mycorrhizal fungi exchange resources with plants by facilitating the solubility and transport of insoluble soil nutrients (Aislabie and Deslippe, 2013), particularly phosphorus, to the plant root system. The mycorrhizal networks associated with roots can increase the effective volume of soil the root can access by orders of magnitude (Li et al., 1991). Similarly, the interaction of nutrients in the soil, such as nitrogen, phosphorus, or sulfur, dramatically alters their solubility and chemical reactivity, directly influencing their transport in the environment and/or movement and exchange between the biosphere, atmosphere, soil, and water.
Microbiomes support nutrient cycling by breaking down and transforming complex organic compounds into forms that plants can utilize as they have a high-to-low carbon-to-nitrogen (C:N) ratio of 12:1 or 8:1 and a short shelf life (Raza et al., 2023). Additionally, microbiomes can transform insoluble nutrients into plant-available soluble ones. Plant root bacterial families such as Xanthobacteraceae and Bryobacteraceae have genes related to organic compound intake and phosphorus, and nitrogen turnover, which function as phosphorus transporters, mineralizers, and solubilizers (Camargo et al., 2023). Besides enhancing the availability of macronutrients like nitrogen (Andrews and Andrews, 2017), phosphorus (Paries and Gutjahr, 2023; Finkel et al., 2019), potassium (Masood and Bano, 2016), sulfur (Aziz et al., 2016), and magnesium (Prasad et al., 2018), plant microbiomes also improve the availability of micronutrients such as iron (Rajkumar et al., 2010), copper (Ferrol et al., 2016), manganese (Nogueira and Cardoso, 2003), zinc (Kamran et al., 2017), molybdenum (Shi et al., 2020), boron (Quiroga et al., 2020), and nickel (Singh et al., 2022). For example, inoculation of plants with strains of Pseudomonas, Arthrobacter, and Bacillus spp. increased nitrogen, phosphorus, and potassium content in Astragalus mongholicus seedlings by 8% to 118% (Shi et al., 2023). Under field conditions in a tropical savanna climate, inoculating soybeans with N-fixation bacteria increased yield by 22% (439 kg per hectare) over that of uninoculated plants (Cordeiro and Echer, 2019). In an analysis of 97 peer-reviewed papers about the use of microbes to improve crop yield, Li et al. (2022) reported that around 22% of them focused on microbes which improved plant nutrient availability, for which key players were Enterobacter (around 27%) and Bacillus (around 26%). Therefore, plant microbiomes contribute substantially to improving plant nutrient availability and could contribute to reducing the reliance on fertilizers to mitigate nutrient deficiencies in the face of global climate change.
Plant microbiomes host a range of mechanisms to combat pests and maintain plant health and resilience. Some microbes produce antimicrobial compounds which can inhibit the growth of plant pathogens (Stracquadanio et al., 2020; Kumar et al., 2021; Zehra et al., 2017; Mendoza-Mendoza et al., 2024). For example, 13 Trichoderma species have been reported to produce 6-pentyl-α-pyrone, an antifungal compound which can control an array of plant pathogens (Mendoza-Mendoza et al., 2024). Beneficial microbes can induce systemic resistance in plants, activating defense mechanisms that enhance plant resilience against pests (Moran-Diez et al., 2021; Kulkarni et al., 2024). The inheritance effect of plant mitogen-activated protein kinases (MAPK) primed by Trichoderma spp. enables next-generation seedlings to better withstand stresses (Moran-Diez et al., 2021). Competition for resources is another mechanism whereby microbes outcompete pathogenic organisms for nutrients and space around plant roots (Chepsergon and Moleleki, 2023). For example, the incidence of the pathogen Ralstonia solanacearum was reduced by 47% when eggplant was inoculated with rhizosphere microbiomes (Jiang et al., 2022). In the rhizosphere, Bacillus spp. and Pseudomonas spp. colonized plant roots faster than pathogenic Fusarium spp. and Pythium spp., which reduced disease pressure (Chen et al., 2024).
Microbes can also control invertebrate pests (Santoyo, 2022; Chen et al., 2023; Shi et al., 2021). Entomopathogens such as Beauveria, Bacillus, and Metarhizium spp. are exploited from nature to be used as biopesticides to control an array of insect pests (Glare et al., 2012). For instance, a combination of Beauveria strains was found to increase efficacy for control of diamondback moth (Soth et al., 2022a). The production of secondary metabolites with anti-insect properties is a common trait. These metabolites deter or inhibit pest feeding and/or reproduction (Soth et al., 2022b; Mascarin and Jaronski, 2016). For example, insect-resistant sugarcane had a higher microbiome diversity in stems than in soil, with fungi (22%) dominating over bacteria (10%), while microbiomes of susceptible sugarcane and the surrounding soil shifted toward those of resistant plants after insect damage (Li et al., 2023). This reveals a protection function of microbiomes against sugarcane insects. In other systems, nematophagous Ascomycota and Basidiomycota fungi can control pathogenic nematodes directly via parasitism and indirectly through secondary toxic metabolites (Al-Ani et al., 2022). Microbes can also modulate plant production of volatiles, influencing the emission of compounds that attract natural enemies of pests, such as predatory insects (Francis et al., 2020) and parasitoids (Contreras-Cornejo et al., 2018). Moreover, endophytic Epichloë fungi in grasses, which are vertically transmitted through seeds, can protect grasses against various root-feeding invertebrate pests. This protection generates approximately US$200 million annually to the New Zealand economy (Johnson et al., 2013).
Microbes’ establishment of biofilms on plant surfaces indirectly creates a physical barrier that hinders pest attachment and colonization (Velmourougane et al., 2017). Nutrient cycling and improving plant health are facilitated by beneficial microbes, making plants less susceptible to pest attacks (Iqbal et al., 2023). Furthermore, microbes trigger plant’s defense mechanisms, resulting in the synthesis of defensive proteins and phytochemicals that deter pests or reduce susceptibility to the attack (War et al., 2012).
Plant microbiomes can foster adaptive resilience mechanisms to abiotic stresses like drought, salinity, and heat. In particular, the symbiosis of plant roots with diverse strains of plant growth-promoting rhizobacteria (PGPR) species is a key aspect of increasing drought tolerance by enhancing root proliferation, water and nutrient absorbance, osmotic regulation, antioxidant enzyme accumulation, photosynthetic activity, pigment synthesis, phytohormone adjustment, and upregulation of stress-related genes (Chieb and Gachomo, 2023). Microbes contribute to enhancing plant water use efficiency by influencing biosynthesis of the phytohormone ethylene (Suganuma et al., 1995). Ethylene regulates stomatal conductance and decreases transpiration rates, allowing the plant to better cope with drought conditions (Khan et al., 2024). Microbes can also trigger plants to generate production of osmoprotectants and antioxidants, thereby ensuring the integrity of plant cells in stress situations by regulating phytohormonal pathways and building root systems to absorb the available water (Carter et al., 2023). Like induced systemic resistance (ISR) to biotic diseases, microbes facilitate induced systemic tolerance (IST) by the establishment of resistance mechanisms against abiotic stresses (Yu et al., 2022). Microbiomes also mediate plant-microbe signaling, and they induce the upregulation of stress-responsive genes such as pathogenesis-related (PR) and pattern-recognition receptor (PRR) genes in plants (Liu et al., 2020).
Plant microorganisms also indirectly assist in the alleviation of abiotic stress. This can occur, for example, via promotion of the formation and expansion of the root system which provides a greater ability to absorb water and nutrients from the soil (Liu et al., 2021). Similarly, improved soil structure as a result of microbial activity can also conserve water and reduce the impact of drought on plant growth (Bogati and Walczak, 2022). Plant metabolites, including phytohormones and other secondary metabolites (coumarins, terpenoids, and benzoxazinoids) can stimulate microbial communities to help plants alleviate stress by improving nutrient cycling, suppressing biotic stresses, detoxifying phytotoxins, and balancing phytohormones (Su et al., 2023). The detoxification or sequestration of harmful substances by plant microbiomes (Sharma et al., 2021) also improves tolerance to stress (Montreemuk et al., 2024). Therefore, plant microbiomes help plants avoid abiotic stresses through (1) changing plant physical structure, (2) producing stress-protective substances, (3) manipulating plant resource uptake, (4) activating plant defense mechanisms, and (5) upregulating plant genes linked to response to stress.
Plant microbiomes regulate phytohormones, including auxins, cytokinins, gibberellins, abscisic acid, jasmonic acid, salicylic acid, and ethylene (Stec et al., 2016; Van der Meij et al., 2018; Ding et al., 2018; Li et al., 2019; Chen X. et al., 2019; Carvalhais et al., 2013; Reusche et al., 2013; McGuiness et al., 2019; Conway et al., 2022; Gonin et al., 2023). Auxin controls several facets of plant growth and development, such as cell elongation, maintaining apical dominance, and triggering root formation (Shani et al., 2017). Bacterial species such as Variovorax sp., Pseudomonas sp. and many other bacterial species (Gonin et al., 2023) interact with auxins in plants, with widespread impacts on growth and physiology (Conway et al., 2022; Leveau and Lindow, 2005).
Cytokinin primarily impacts plant physiological development and growth through cell division and shoot development regulation (Zurcher et al., 2013). Bacterial species like Azospirillum spp., Bacillus spp., and Pseudomonas spp. play a pivotal role in cytokinin signaling (Gupta et al., 2022; Hussain and Hasnain, 2011). For example, after inoculating Platycladus orientalis (an evergreen conifer) with Bacillus subtilis, Liu et al. (2013) reported a 97% and 48% increase in cytokinin contents of irrigated and drought-stressed plant shoots, respectively. As cytokinin also influences stomatal opening and closure (Hussain and Hasnain, 2011), increasing cytokinin influences plants’ resistance to biotic and abiotic stresses (Gupta et al., 2022; Todero Ritter et al., 2013). Gibberellin, another crucial plant hormone, plays a significant role in plant growth, including stem elongation, regulating seed germination, and inducing flowering (Castro-Camba et al., 2022). As with other hormones, a wide diversity of microorganisms can influence plant gibberellin levels. These include Bacillus spp. (Keswani et al., 2022), Azospirillum spp. (Cohen et al., 2009), Aspergillus fumigatus (Khan et al., 2011), Penicillium spp. (Khan et al., 2013), Pseudomonas spp. (Kapoor et al., 2016; Pandya and Desai, 2014), and Rhizobium spp. (Yanni et al., 2001). For example, inoculating gibberellin-deficient melon with Bacillus sp. LKE15 increased shoot length (by 33%), root length (by 9%), and plant fresh weight (by 65%) compared to the control plants (Kang et al., 2015).
Abscisic acid (ABA) has a distinct role in how plants react to environmental stress responses such as drought and salinity (Tuteja, 2007). It promotes dormancy, regulates stomatal closure, and metabolic regulation. When Phaseolus vulgaris roots were colonized by an entomopathogenic fungus (Metarhizium robertsii) and a plant pathogen (Fusarium solani) Hu and Bidochka (2021) reported that the plants downregulated and upregulated ABA, respectively, demonstrating that these fungi can influence in planta ABA. In rice and tomato, Rhodococcus sp. and Novosphingobium sp. upregulated ABA which resulted in increased plant biomass and biosynthesis (Belimov et al., 2014). Similarly, Bacillus licheniformis regulated ABA content, enabling Chrysanthemum to better withstand saline and alkaline conditions (Zhou et al., 2017).
Jasmonic acid (JA) is a phytohormone involved in various physiological processes in plants, especially stress response, defense against herbivores, and regulation of growth and development (Hewedy et al., 2023; Kulkarni et al., 2024). JA also plays another significant role in facilitating the interaction between the plant and its microbiomes (Pieterse et al., 2009). For instance, JA can stimulate beneficial bacterial and archaeal communities, including Streptomyces, Bacillus, and Lysinibacillus (Carvalhais et al., 2015). Similarly, salicylic acid (SA) is a key signaling molecule in plants, playing a central role in the regulation of many physiological processes, particularly in response to abiotic (Yang et al., 2023) and biotic stress (Yang et al., 2015). SA also stimulates beneficial microbes, including Rhodanobacter, Sphingomonas, and Micromonospora, which improved tolerance of watermelon to a Fusarium wilt disease (Hou et al., 2022).
Ethylene is another phytohormone that serves as a key regulator of plant growth, development, and stress responses, influencing processes like fruit ripening, leaf senescence, and abscission, while also interacting with other hormones to orchestrate adaptive plant responses (Iqbal et al., 2017; Khan et al., 2024). Besides plant genotype, microbiomes play a significant role in regulating ethylene levels in plants (Ravanbakhsh et al., 2018). Microbes such as Azospirillum spp., Rhizobium spp., Agrobacterium spp., Achromobacter spp., Burkholderia spp., Ralstonia spp., Pseudomonas spp., Enterobacter spp., and Kluyvera ascorbata are key species known to regulate ethylene (Blaha et al., 2006). Thus, microbiomes produce ethylene that can affect food quality and storage, as well as shelf life of ornamental flowers.
Microbiomes are important for food processing and preservation, contributing to the development of various fermented and other food products, and influencing the shelf life of many more. Every day, people consume food and drinks produced through the activities of microbiomes (Siddiqui et al., 2023). The current global market for fermented food alone is valued at around USD 30 billion (Voidarou et al., 2020). For thousands of years, people have been using microbial communities for processing food, including adding distinct flavors, prolonging preservation, and improving nutritional values. Examples include cheese, soy sauce, pickled vegetables and fruit, sourdough and other breads, tempeh, yogurt, beer, cider, kombucha, and wine (Louw et al., 2023; Tamang et al., 2020; Figure 2). The following are examples of traditional foods and drinks from different cultures which use microbial processes: Japan (miso, soy sauce, and sake) (Murooka and Yamshita, 2008), South Korea (kimchi and makgeolli) (Shin et al., 2016), China (doubanjiang and baijiu) (Guan et al., 2020; Niu et al., 2023), India (dosa and idli) (Tamang, 2022), Germany (sauerkraut and beer) (Ngcamu, 2023), France (Camembert cheese and wine) (Boisard, 2003; Phillips, 2020), Italy (parmesan cheese and balsamic vinegar) (Dalby, 2009; Giudici et al., 2015), Ethiopia (injera) (Neela and Fanta, 2020), Nigeria (ogi and burukutu) (Uzogara et al., 1990), Mexico (tequila, mezcal, tepache and pulque) (Ojeda-Linares et al., 2021; Becerra-Lucio et al., 2022; Aldrete-Tapia et al., 2020), Brazil (cachaça and caiçuma) (Lima et al., 2022), Polynesia (poi) (Brown et al., 2016), Turkey (ayran and boza) (S̨anlibaba and Tezel, 2023), Iran (doogh and traditional pickles) (Meybodi et al., 2016), and Cambodia (prohok) (LeGrand et al., 2020). In food processing, certain microbes such as lactic acid bacteria (the genus Lactobacillus) (Giraffa et al., 2010) and fungi (Rhizopus, Monascus, and Penicillium spp.) (Nout and Aidoo, 2011) are intentionally introduced to transform raw ingredients into flavorful and nutritious products. During fermentation, microbes transform complex carbohydrates, proteins, and fats into new compounds, enhancing the final product’s taste, texture, and nutritional profile (Hutkins, 2008). Additionally, organic acids and antimicrobial compounds produced by these microbes help inhibit the growth of spoilage microbes, contributing to the preservation of processed foods (Sharma et al., 2020) and preventing the growth of harmful bacteria and fungi that can lead foodborne illnesses and waste of food. Techniques such as pickling, canning, and salting rely on the ability of microbes to create an environment hostile to spoilage microbes (Amit et al., 2017). For example, beneficial microbes like Lactobacillus bacteria create an acidic environment in fermented pickles that preserves the vegetables and enhances their flavor (Behera et al., 2020).
For future food, algae (Spirulina spp., Chlorella spp., and Scenedesmus spp.), bacteria (Cellulomonas spp. and Alcaligenes spp.), fungi (Aspergillus spp., Penicillium spp., and Trichoderma spp.), and yeast (Saccharomyces spp. and Pichia sp.) are all potential sources of protein, fiber (cellulose and poly/oligosaccharides), and vitamins (A, B, C, D, and E) (Ravindra, 2000; Graham and Ledesma-Amaro, 2023). Around 470 microorganisms were found in plant-based protein alternates, of which nearly 95% are bacteria (Roch et al., 2024). Therefore, the utilization of microbes in the food industry not only adds diversity and richness to our culinary experiences but also may provide an alternate source of future food.
Microbiomes have a key role in the restoration and remediation of contaminated environments (Maqsood et al., 2023; Rafeeq et al., 2023). This bioremediation, which utilizes the metabolic abilities of microbiomes to enhance the degradation of pollutants, can reduce their harmful impacts on the environment (Ayilara and Babalola, 2023). For example, Rhodococcus rhodochrous (bacteria) and Aspergillus fumigatus (fungus) have the capacity for biomethylation, a process to detoxify heavy metals and metalloids (Hartmann and Six, 2023). Additionally, the bacterium Ideonella sakaiensis can break down plastic wastes (Yoshida et al., 2016). Further examples include bacteria in the genera Bacillus, Mycobacterium, Pseudomonas, and Nocardia that can degrade pesticides in water and soil (Bala et al., 2022; Figure 3).
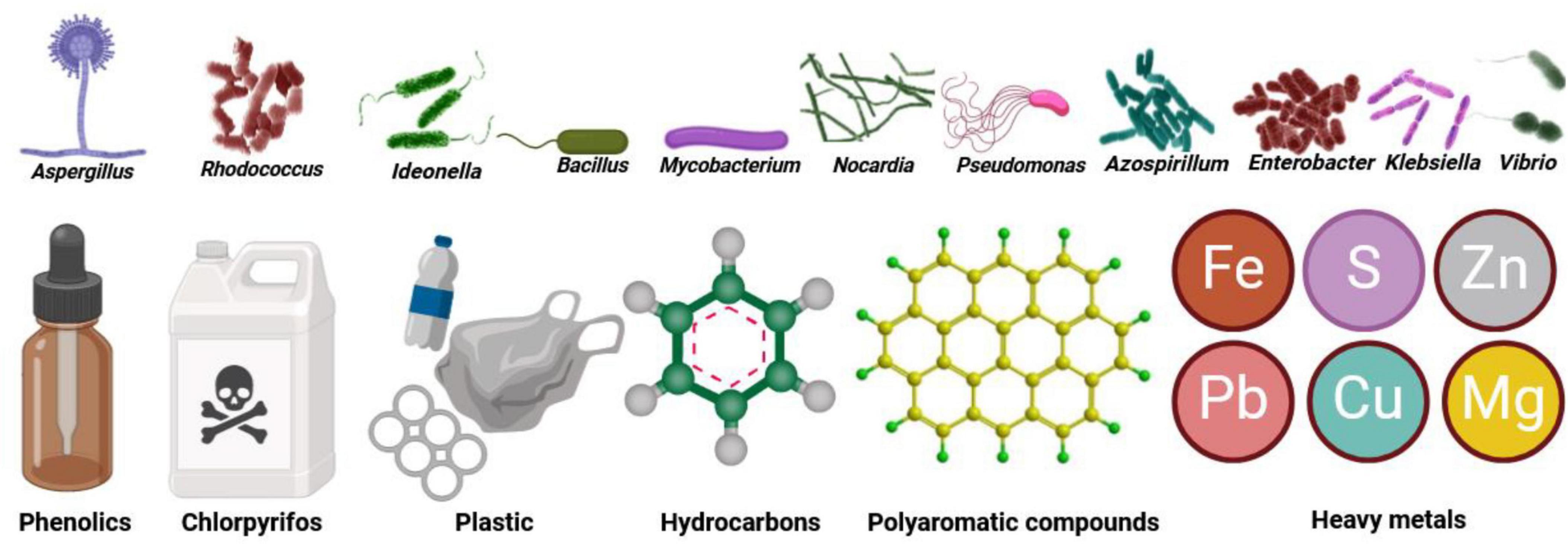
Figure 3. Environmental microbiomes such as Aspergillus, Bacillus, Ideonella, Mycobacterium, Nocardia, Pseudomonas, and Rhodococcus can break down chemicals and hydrocarbons as well as pesticidal pollutants. Other microbiomes, including Azospirillum, Enterobacter, Klebsiella, Pseudomonas, and Vibrio are responsible for breaking down heavy metals that are environmental pollutants.
Microbiomes are essential in the process of phytoremediation whereby plants (and their microbial symbionts) filter, immobilize or transform pollutants (Sessitsch et al., 2023; Wang et al., 2024; Nathan and Ammini, 2019; Shahid et al., 2020). A microorganism able to degrade pollutants in the soil is attracted to root exudates secreted from plants. Examples of plant-microbiome heavy metal remediation systems include Bacillus sp. in Arundo donax (a perennial cane) (Sarathambal et al., 2017), Enterobacter and Klebsiella sp. in Brassica napus (Jing et al., 2014), Pseudomonas in the fern Pteris vittate (Manzoor et al., 2019), and Phyllobacterium myrsinacearum in the succulent Sedum plumbizincicola (Ma et al., 2013). Plants allocated up to 22% of their carbon source to their symbiont ectomycorrhizal fungi to improve nutrients uptake under nutrient inefficiency conditions (Hobbie, 2006). Overall, the synergism between microbiomes and plants improves the ability to eradicate toxins from the environment, making it an effective and sustainable environmental restoration method (Aislabie and Deslippe, 2013). Hence, using microorganisms as the driving force in establishing environmental remediation, including phytoremediation, is not only low cost and sustainable, but also is in line with the biodiversity balance and preservation.
Microbiomes have been widely used in industrial wastewater treatment. For instance, microbial communities can decompose organic matter (Fenchel et al., 2012) and remove waterborne pollutants (e.g., nitrogen fixation by bacteria such as Enterobacter spp., Azospirillum spp., Pseudomonas spp., Klebsiella spp., and Vibrio spp.) (Haque et al., 2020; Lamers et al., 2012). This environmentally friendly method harnesses the natural abilities of microorganisms to degrade and detoxify pollutants present in industrial effluents (Nagda et al., 2022). In combating pollutants, various strains of bacteria and fungi have been used to target specific contaminants, facilitating the breakdown of organic compounds and removing heavy metals. For instance, Aspergillus tubingensis isolated from the mangrove rhizosphere can absorb more than 90% of heavy metals (98% of lead, 96% of nickel, 94% of zinc, and 92% of copper) (Mahanty et al., 2020). Additionally, when using green microalgae Chlorella vulgaris as a co-precipitate with other heavy metals, there was a 91% and 85% reduction of phosphoric and ammonium ions, respectively (Govarthanan et al., 2020). Another study showed the bioremediation ability of the mesophilic bacteria Lysinibacillus sp. in reducing nickel (by 70%), chromium (by 58%), and reactive black 5 dye (by 82%) (San Keskin et al., 2018). Overall, these examples show the potential for bioremediation to offer an effective solution for removing industrial pollutants from wastewater systems.
It has recently been shown that the human gut microbiome is not simply important for digestive health but plays a key role for the broader well being, physiology, inflammation, immunity, and disease status of people (Hou et al., 2022; Sender et al., 2016; Zhao et al., 2023; Li et al., 2023). The gut microbiome consists of approximately 38 trillion microbes from 3,000 species, including bacteria, viruses, and fungi. The main microbial groups of the human body are presented in Figure 4.
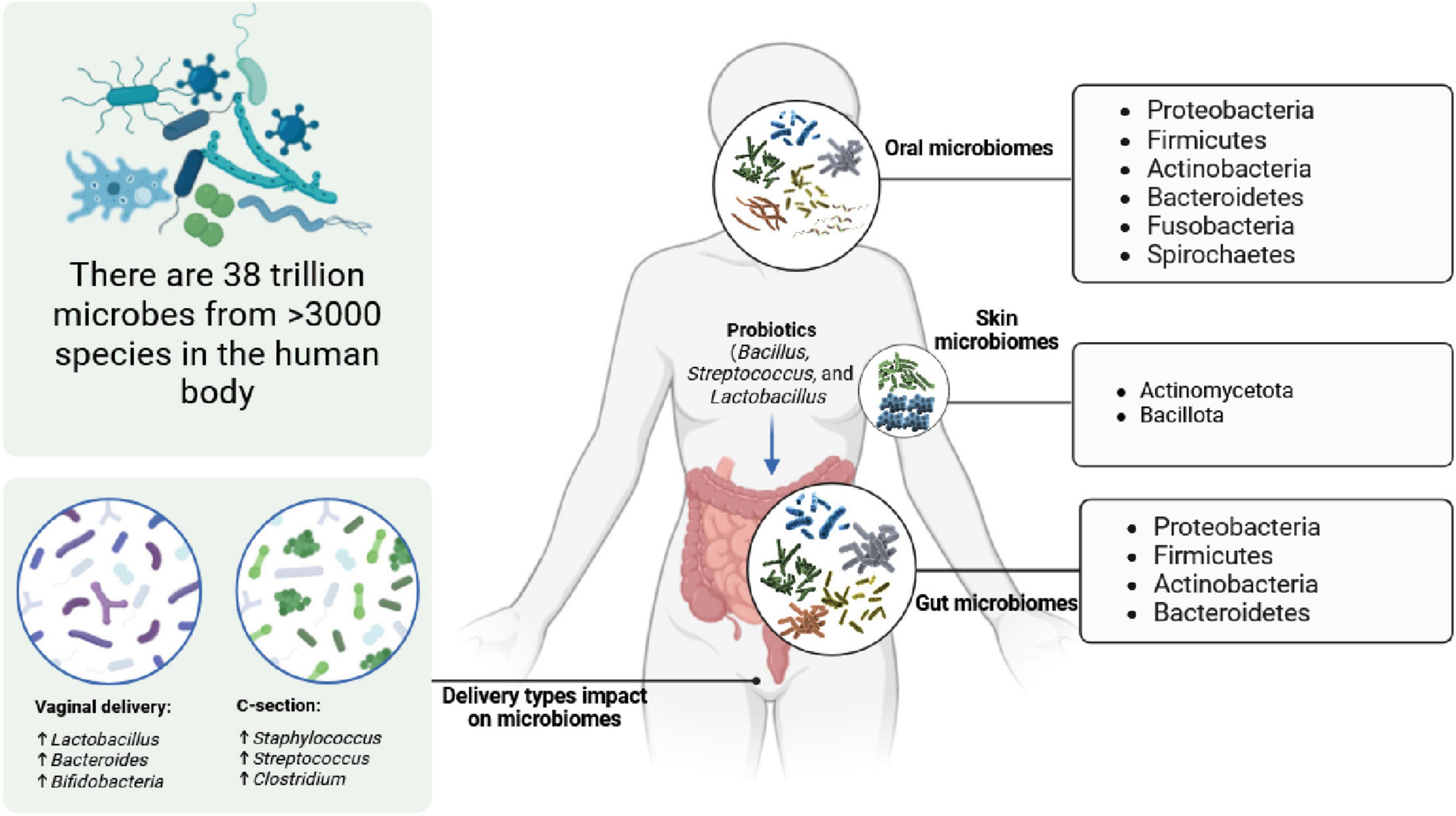
Figure 4. Human microbiome compositions in oral, skin, gut, and vaginal systems, and the impact of delivery method on a newborn baby’s microbiome community.
The gut microbiome is connected to the broader physiological state, affecting overall human health and wellbeing. For example, the gut-brain axis provides at least three well-characterized pathways of connection between the gut, brain, and broader body. As such, changes in the diversity, balance, or function of microbiomes in the gut can have profound effects on overall physical and mental health (Clapp et al., 2017; Rea et al., 2016; Campbell et al., 2023). For instance, adjusting microbial communities in the mice gut enhanced host resilience to chronic social defeat stress (Wang et al., 2020). Additionally, the human gut microbiome is known to have some roles in disease prevention, as disturbances in its composition are related to various health consequences, including autoimmune disorders, allergies, and metabolic diseases (Vijay and Valdes, 2022; Keswani et al., 2022).
Myriad foods which contain various nutrients can favorably influence the development of specific microorganisms in the gut; therefore, food choice affects microbial compositions. For instance, the intake of dietary fiber from plant-based products promotes the development of bacteria that promote digestive health and produce short-chain fatty acids, which play a role in gut health (Sun et al., 2022). However, changes in the gut microbiome related to adverse health outcomes can also be caused by food that has undergone processing and added sugar (Keswani et al., 2022). Therefore, a healthy microbiome in the gut is increasingly gaining importance as part of a holistic maintenance of health, immunity, and disease prevention through a good diet and a healthy lifestyle.
The possibility to employ microbiome-targeted therapies comprising probiotics and prebiotics is an exciting approach to promotion of health and treatment of numerous diseases (Yadav and Chauhan, 2022; Ji et al., 2023). Probiotics, which are live beneficial microorganisms, when used in adequate amounts, can positively influence the amount and functions of the gut microbiome (Hemarajata and Versalovic, 2013; Das et al., 2022). These microorganisms are involved in regaining a normal microbiota balance and improving immunological fitness. They may also be used for the prevention or treatment of conditions such as inflammatory bowel disease and even allergies (Yuan et al., 2023; Cristofori et al., 2021; Simon et al., 2021). For example, a mixture of probiotics (Medilac® containing Bacillus subtilis and Streptococcus faecium) in combination with Mosasal® containing 5 mg of mosapride (gastroprokinetic agent) was found to reduce abdominal pain within a month (Choi et al., 2015). Unlike prebiotics, pro- and post-biotics work by stimulating the growth of healthy bacteria and reducing the number of pathogens (Davani-Davari et al., 2019; Bedu-Ferrari et al., 2022). Synbiotics, the synergistic use of probiotics and prebiotics, can also be utilized to strengthen the effects of both probiotics and prebiotics (Simon et al., 2021; Kolida and Gibson, 2011). For example, a prophylactic treatment utilizing Lactobacillus plantarum decreased sepsis to an overwhelming extent in newborn babies (Panigrahi et al., 2017). Thus, microbiome-based therapeutics serve as the basis for personalized medicine approaches, creating new ways of staying healthy and fighting different diseases.
Emerging research on skin, oral, and vaginal microbiomes is shedding light on their significant implications for personalized medicine and disease management. These microbial communities play crucial roles in maintaining the health and function of their respective environments (O’Mahony and Comizzoli, 2023; Prescott et al., 2017). The skin microbiome (Propionibacterium acnes, Staphylococcus epidermidis, and Staphylococcus aureus) (Byrd et al., 2018), for instance, influences immune responses and skin conditions (Smythe and Wilkinson, 2023), while the oral microbiome (Firmicutes, Actinobacteria, Proteobacteria, Fusobacteriota, Bacteroidetes, and Spirochetes) (Prasad et al., 2018) is linked to oral health and systemic well being (Sedghi et al., 2021). Similarly, the vaginal microbiome (Lactobacillus spp. and Gardnerella vaginalis) plays a pivotal role in reproductive health (Chen et al., 2021). For example, vaginally born babies have a beneficial microbiome (higher Bacteroides, Bifidobacterium, and Lactobacillus), enhancing their immune systems. In contrast, Cesarean-born babies exhibit a less diverse microbiome (mostly Staphylococcus, Streptococcus, and Clostridium), resembling hospital settings, potentially impacting their immune development (Coelho et al., 2021; Korpela, 2021). Examining the complicated surroundings of these microbiomes gives a new path for personalized medicine, as the variations in microbial composition can determine the level of an individual’s susceptibility to specific diseases and affect treatment outcomes (Behrouzi et al., 2019; Cullen et al., 2020). Therefore, understanding the microbiome may make it possible to develop interventions aimed at a particular microbial disbalance, enhancing the effectiveness of the therapy and minimizing side effects.
Microbiomes have great potential to improve green processes and reduce environmental problems. Biocatalyst processes such as hydrogen production, carbon dioxide removal, and bioenergy can be significantly improved using molecular microbial engineering (Behera et al., 2022; Nagda et al., 2022). For instance, bacteria and archaea transform organic materials from animal and agricultural systems and sewage wastes into biogas. First, complex organic compounds are hydrolyzed by the bacterial phyla Firmicutes and Bacteroides into simpler substances (Mamindlapelli et al., 2021). Then, the bacterial genera Chloroflexi, Proteobacteria, and Atribacteria synthesize these substances into volatile fatty acids (VFAs). Acetogenic bacteria, including Acetobacterium, Clostridium, and Syntrophomonas, catabolize VFAs to form acetate, carbon dioxide, and hydrogen. Finally, the methanogenic archaeal genera Methanobacterium, Methanosarcina, and Methanococcus use acetate, carbon dioxide, and hydrogen as substrates for the final stage of the anaerobic process and produce methane and carbon dioxide (biogases) (Lim et al., 2020; Figure 5). Improvements are aimed at altering microbial ecosystem design to improve their efficiency, reliability, and productivity (Ramamurthy et al., 2021). Beneficial microorganisms can be introduced or environmental conditions optimized to allow microbiome-based interventions to achieve the goals of more environmentally friendly and sustainable industrial processes (Wu et al., 2021). This will allow the possibility of raising yields of recovered valuable materials such as bioproducts, biopolymers, and other value-added compounds, contributing to decreasing waste and reducing the environmental impact of industrial activities (Wiltschi et al., 2020).
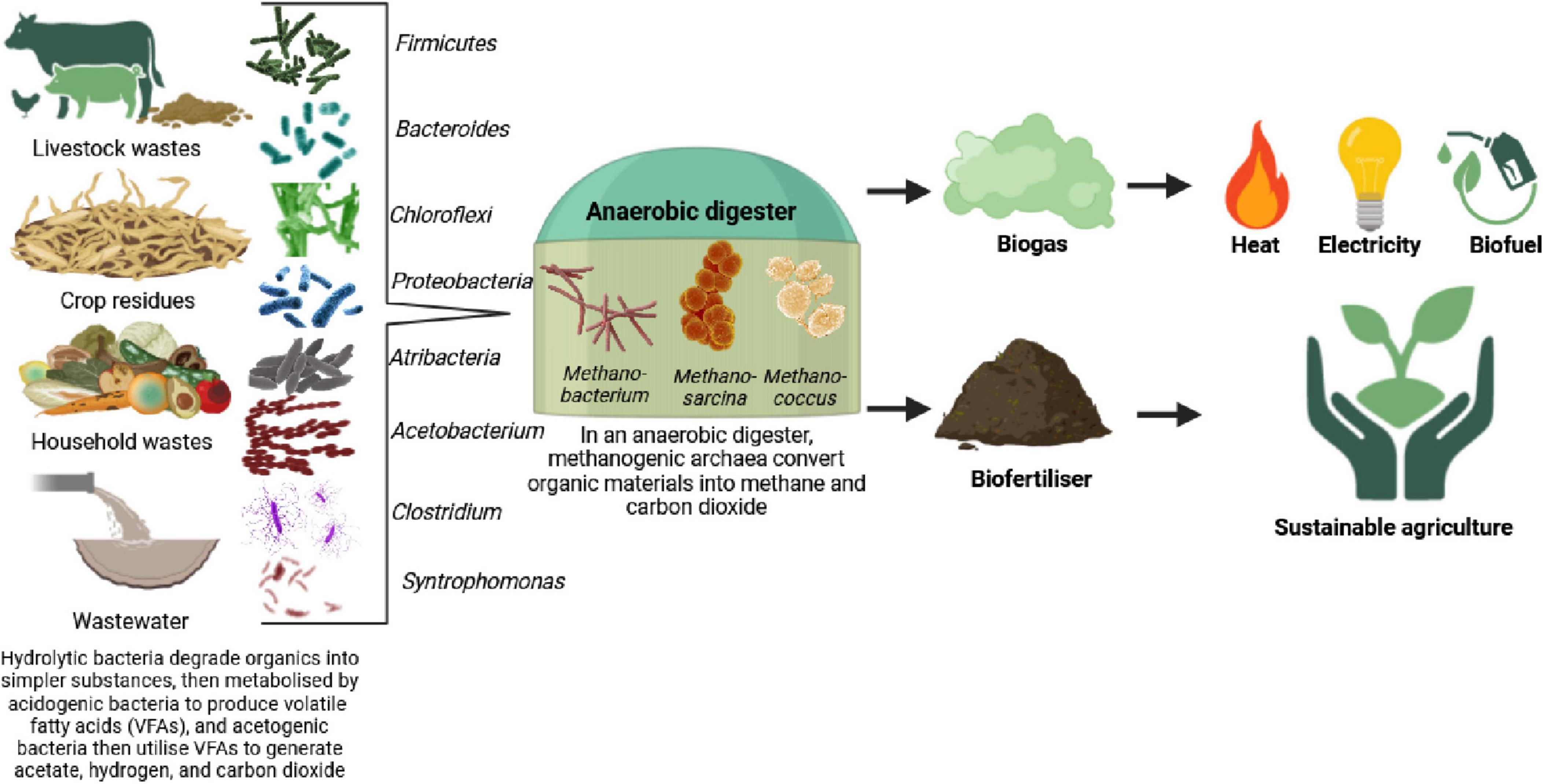
Figure 5. Graphical illustration of how microbiomes convert organic materials and wastewater into biogas, bioenergy, and biofertilizer.
Microbial biocatalysts are pivotal in advancing the sustainable production of biofuels, bioplastics, and biochemicals. Harnessing the metabolic capabilities of microorganisms allows efficient conversion of renewable resources into valuable bio-based products (Ramamurthy et al., 2021). Approximately 58 strains of bacteria, 24 species of fungi, and 17 species of yeast are known to have the ability to serve as crucial catalysts in the fermentation and transformation of biomass-derived feedstocks into ethanol, biodiesel, and other alternative fuels (Chen et al., 2014). For example, the bioethanol market is valued at USD 83.4 billion and is forecasted to increase to USD 114.7 billion by 2028 with an annual growth rate of 6.6% (MarketsandMarkets, 2024). The main microbes involved in biofuel production include Trichoderma reesei (which produces an enzyme that breaks down lignocellulosic biomass), Saccharomyces cerevisiae (utilizes hexose sugar), Scheffersomyces stipites (exploits pentose sugar) (Rastogi and Shrivastava, 2017), Penicillium echinulatum (produces cellulase and xylanase) (Todero Ritter et al., 2013), Anoxybacillus flavithermus (a pH and temperature stabilizer for xylanase synthesize) (Ellis and Magnuson, 2012), and Aspergillus spp. and Kluyveromyces spp. (which synthesize inulin) (Ricca et al., 2007). Combining T. reesei, S. cerevisiae, and S. stipites allowed a 67% yield of bioethanol from non-sterile wheat straw (Brethauer and Studer, 2014). Similarly, in the production of bioplastics, microbes contribute to the synthesis of polymers from renewable sources, offering eco-friendly alternatives to traditional petroleum-based plastics (Behera et al., 2022).
Bacterial species including Cupriavidus necator (Philip et al., 2007), Pseudomonas putida (Manoli et al., 2020), Aeromonas hydrophila and Pseudomonas aeruginosa (Chen, 2010), Pseudomonas mendocina (Zhang et al., 2020), Bacillus megaterium (Cal et al., 2021), and Alcaligenes eutrophus (Sathya et al., 2018), are reported to be involved in the biosynthesis of polyhydroxyalkanoates. Additionally, microbial biocatalysts are integral in the sustainable production of biochemicals, facilitating the transformation of organic materials into various high-value chemicals used in industries ranging from pharmaceuticals to agriculture (Wiltschi et al., 2020; Wu et al., 2021). Some microbes, like Bacillus sp., Aneurinibacillus sp., and Trichoderma harzianum, can produce lignin-degrading enzymes for sustainable processing of agro-byproducts (Sharma et al., 2022). For example, algae, bacteria, fungi, and yeasts play a significant role in transforming plant and food residues into microbial proteins, which are valuable as animal feeds (Rasool et al., 2023). This interdisciplinary approach, leveraging the capabilities of microbiomes, underscores the potential for environmentally friendly and economically viable solutions in the pursuit of a more sustainable future.
Microbiomes contribute sustainably to agriculture, human health, the environment, and biocatalyst processes. From flora and fauna sustainability to human and environmental health, microbiomes emerge as the key players in the complex regulation of ecological balance and resilience (Suman et al., 2022). Microbiomes could be pivotal in sustaining the world if their roles are correctly researched and standardized to guarantee their stability and resourcefulness in the era of climate change (Zheng et al., 2020). This illustrates the concept of a circular microbiome system, where microbiomes are interconnected across multiple dimensions of life, forming a continuous, interdependent, and virtuous cycle. The microbiome of the environment enhances soil fertility, creates a safe ecosystem, and supports sustainable food production. This, in turn, nurtures the plant’s microbiome, enabling healthy growth, resilience to diseases, and higher yields. The plant becomes food, its microbiome influencing the food microbiome, improving nutritional quality, preservation, and flavor. Once consumed, the food impacts the gut microbiome, which governs human health, digestion, and immunity. Waste generated by this process and other agricultural byproducts is transformed into bioenergy or biofuel by biocatalyst microbiomes. Finally, the final waste turns into biofertilizer, which increases plant productivity and improves microbiome diversity in the environment and soil. This interconnected process is aligned with the One Health concept, which is about interaction among humans, animals, and the ecosystem (Ma et al., 2023). Climate change has meant that around 58% of identified zoonotic diseases have emerged through pathogen spillover by altering wildlife distribution and behavior, vector and pathogen adaptability, and releasing ancient diseases from melting permafrost (Mora et al., 2022; Lim et al., 2023). As the circular microbiome system is to maintain biodiversity equilibrium and integrity, it may assist in the prevention of pathogen emergence. For example, the “Balance of Dynamic Factors” concept demonstrated the significant interactions among humans, animals, and the environment for disease prevention, health equity, and ecosystem sustainability (Zhou et al., 2024).
However, challenges need to be resolved to bring together an integrated whole-of-system approach. Addressing knowledge gaps and technical issues remains crucial to improving understanding of the microbiome and its broader applications. Attempts to understand the inner workings of microbial communities have been frustrated by their complexities, and an holistic view is required (Sergaki et al., 2018). Specifically, targeted research efforts must be carried out to close the knowledge gaps in diversity, structure, and interaction processes within microbiomes across different habitats. Moreover, technical obstacles, including developing new tools for high-throughput sequencing, bioinformatics tools, and standardized methodologies, will have to be available for use in the process (Barea, 2015). For instance, whole genome sequencing can be used to study microbiomes in different ecosystems (Aminu et al., 2024). Additionally, bioinformatic tools for data processing (FastQC, Cutadapt, DADA2, QIIME2, and MOTHUR), taxonomic profiling (Kraken2, MetaPhlAn, SILVA, and UNITE), functional profiling (HUMAnN, PICRUSt, KEGG, and EggNOG-mapper), diversity analysis (QIIME2, Phyloseq, Vegan, and MicrobiomeAnalyst), metagenomic assembly and annotation (MEGAHIT, MetaSPAdes, Prokka, and IMG/M), statistical analysis and visualization (LEfSe, ANCOM, STAMP, and ggplot2), microbial interaction and network analysis (SPIEC-EASI, CoNet, and Cytoscape), and metatranscriptomics and metabolomics (Kallisto, MetaCyc, and GNPS) can be complex and with steep learning curves for inexperienced researchers. The large amount of sequencing data generated can be computationally demanding and often requires high-performance computing infrastructure. Therefore, training and access to these tools, computing infrastructure, and dedicated bioinformatic support are all needed to help achieve the common goals.
Special consideration needs to be given to rights of Indigenous communities and peoples. As guardians of native spaces in many parts of the globe, the ecosystems they manage are often exploited (or “bio-prospected”) as a resource for microbiomes; essentially comprising biobanks of important and novel microbiomes (Warbrick et al., 2023). Māori people of Aotearoa (New Zealand), for example, practice kaitiakitanga (stewardship) to protect biodiversity resources, which provides insights for Western-trained researchers about holistic approaches for sustainable biological conservation (McAllister et al., 2023). For example, indigenous uses of the bark of the mānuka tree (Leptospermum scoparium) for its antimicrobial properties led to the discovery that mānuka honey has a wide range of health benefits (Zucchetta et al., 2022). Accompanying this is often a deep knowledge of the ecosystems themselves. This knowledge has been gained over generations of people living intimately as an integral part of these ecosystems - the first true “circular bioeconomies” - and incorporates knowledge and values of the connections between the soil and plants, the environment and communities. Recognition of their stewardship in protecting these places and the biodiversity resources they hold, which may benefit mankind, must be appropriately recognized and protected. Indeed, the interests of Indigenous communities are protected across the United Nations member countries within areas such as the Permanent Forum on Indigenous Issues, the World Intellectual Property Organization, and the Convention on Biological Diversity (Hudson et al., 2020). Article 31 of the UN Declaration on the Rights of Indigenous Peoples states that “Indigenous people have the right to maintain, control, protect… manifestations of their sciences, technologies and cultures, including human and genetic resources, seeds, medicines, knowledge of the properties of fauna and flora,…” (United Nations, 2007). This is supporting the Nagoya Protocol, a supplementary agreement to the Convention on Biological Diversity. The Nagoya Protocol includes the “equitable sharing of benefits from using genetic resources” (Buck and Hamilton, 2011). It is often the case that microbiome research – from the discovery of unique taxa, understanding function and ecology, through to taxonomic placement – is now supported by genetic and genomic information and, as such, underpins modern microbiology and microbiome sciences and, thereby, falls under the mandate of the Nagoya protocol.
Furthermore, Indigenous communities are often relatively impoverished and face a range of inequity issues spanning food security, access to medicine and health care, through to education, and access to financial and other support systems (Warbrick et al., 2023). Therefore, these communities may benefit significantly if a process of consultation and partnership is used, and ownership of their knowledge and microflora is appropriately protected and rewarded. There is an opportunity for science to help redress components of inequity in the systems and add value to all partners via participation, not exploitation.
Another big challenge is the establishment of solid regulatory frameworks able to control the use of microbiomes (Peixoto et al., 2022). As regulatory frameworks may vary globally, effective communication and documentation to define concepts and criteria of microbiome utilization with R&D investments to build stakeholder trust are bottlenecks to expanding the potential of microbiomes (Kostic et al., 2024). Maintaining an harmonious relationship between technology propulsion and environmental security is fundamental. Additionally, ethical considerations around this area of research become evident as the deliberate modification of microbiomes creates possibilities for unintended consequences and ecological impact (Lange et al., 2022). In the future, researchers, policymakers, and other relevant stakeholders must work together to design and develop regulatory guidelines which can be adopted as new knowledge emerges about the microbiome (Greenhough et al., 2020).
Furthermore, supporting explicit conversational spaces and active public participation, including with Indigenous communities, in developing ethical standards will be necessary to allow trustworthy practices and responsible use (Lange et al., 2022). In particular, indigenous cultural values, knowledge, and perspectives can bring and embody significant value in the life cycle of microbiome research, from discovery to development and utilization. That is, it has its own intrinsic and extrinsic values, as well as contributing to unique foods, practices, and customs of different cultures themselves. Furthermore, often, this information on the microorganisms and their relation to the environment cannot be obtained elsewhere; it sits with and is held by the people and is often passed across and down generations as oral knowledge. The Western approach of sharing knowledge in published format (print or online) cannot access this deep well of information.
Integrating different disciplines, such as microbiology, genetics, ecology, and computational sciences, is another challenge (Cullen et al., 2020). While some studies have explored a system approach to interdisciplinary and transdisciplinary collaboration (Meisner et al., 2022; Tomasulo et al., 2024), the practical aspects of such frameworks requires breaking technical and structural barriers. Sharing knowledge and cooperation are the keys in fully capturing the value of microbiome research. Moreover, retrieving multiple data sources from different methods and tools entails another challenge, thus necessitating the creation of common approaches. Since microbiome research is progressing rapidly, the future course should give precedence to the breaking down of silos and promoting collaboration of scientists from all disciplines (Greenhough et al., 2020). Tomasulo et al. (2024) proposed a model for a healthy ecosystem and encouraged innovative microbiome research via active collaboration among all relevant stakeholders to achieve this goal. Multidisciplinary research team creation, furthering cross-training, and establishing open data-sharing platforms will be key to potentially using microbiomes (Acosta et al., 2022; Amann et al., 2019; Tomasulo et al., 2024; Meisner et al., 2022). Adopting modern technologies and techniques, e.g., advanced sequencing methods and computational modeling, will lead to groundbreaking findings in basic and applied genomics (Bianconi et al., 2023). Therefore, overcoming these challenges will be instrumental in the implementation of interdisciplinary work in microbiome research, and this will mark the beginning of a new era wherein the complex interactions between microbiome function and adaptation to different ecosystems will be understood in totality.
The highly multifaceted nature of microbiomes plays a pivotal role in interdisciplinary ecosystems. From their critical involvement in agriculture and food production, through addressing environmental issues, impacting human health and finding novel and eco-friendly biocatalyst processes, microbiomes are a sustainable tool for a greener future. As a part of nutrient cycling, pest and disease management, and cleansing of environmental toxins, microbiomes offer a sustainable way to reduce environmental pollutants. They are also significant for human health, contributing largely to whole-body well being, immune system functioning, prevention of diseases, and personalized medicines. In biocatalyst processes, microbial community activity ranges from optimizing processes involved in wastewater treatment to fermentation of biofuels, bioplastics, and biochemicals. Such microbiome-focused innovations are important for sustainable ecological balance, biodiversity conservation, and assured eco-friendly industrial practices.
These opportunities are also accompanied by the challenges of knowledge gaps, integrating Indigenous knowledge, technical limitations, regulatory frameworks, interdisciplinary collaboration, and ethical considerations, all of which require attention. Breaking interdisciplinary barriers, creating a collaborative culture, and adopting innovative technologies will be key stepping stones in tackling these challenges. Constant assessment, ethical associations, and progressive points of view are essential for effective microbiome utilization. By achieving this, the true capabilities of microbiomes can be unveiled, allowing us to better understand microbial communities and how they interact with the ecosystem. The multifaceted benefits of microbiomes will provide both economically viable and environmentally friendly mechanisms for plant production, improve ecological equilibrium and biodiversity conservation, reduce industrial pollutants, and offer opportunities for improving human health and indigenous people’s values.
A clear roadmap to fully harness the multifaceted benefits of microbiomes is necessary. Future studies should focus on existing systems-based approaches to interdisciplinary and transdisciplinary collaboration (Meisner et al., 2022; Tomasulo et al., 2024) to address knowledge gaps, to include Indigenous knowledge, and to develop advanced tools like meta-omics and biological synthesis. Researchers and policymakers should work together to establish ethical regulatory frameworks, enrich international partnerships, and guarantee equitable access to microbiome innovation. Practical applications across academic disciplines mentioned in this review must be expanded with long-term planning and public intervention. In order to be able to add to the existing literature (Aminu et al., 2024; Buerger et al., 2023; Kostic et al., 2024), this roadmap envisages ethical awareness, Indigenous communities and practical applications, to provide greener, healthier, and more equitable sharing through microbiome innovations. Achieving this will reveal how microbiomes can be utilized as future tools for sustainable agricultural development, enhancing human health, conserving biodiversity, and harmonizing equity among researchers, policymakers, the public, and Indigenous peoples.
SS: Conceptualization, Data curation, Formal Analysis, Investigation, Methodology, Project administration, Resources, Validation, Visualization, Writing – original draft, Writing – review and editing. JH: Conceptualization, Data curation, Formal Analysis, Funding acquisition, Investigation, Methodology, Project administration, Resources, Supervision, Validation, Visualization, Writing – original draft, Writing – review and editing. HA: Data curation, Investigation, Supervision, Validation, Visualization, Writing – original draft, Writing – review and editing. SW: Conceptualization, Data curation, Investigation, Supervision, Validation, Visualization, Writing – original draft, Writing – review and editing. AM-M: Conceptualization, Data curation, Formal Analysis, Funding acquisition, Investigation, Methodology, Project administration, Resources, Supervision, Validation, Visualization, Writing – original draft, Writing – review and editing.
The author(s) declare that financial support was received for the research and/or publication of this article. This research was supported by the New Zealand Ministry of Business, Innovation, and Employment through LINX2201: fungal volatile organic compounds for sustainable agriculture in a changing environment.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The authors declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Abatenh, E., Gizaw, B., Tsegaye, Z., and Wassie, M. (2017). The role of microorganisms in bioremediation-A review. Open J. Environ. Biol. 2, 038–046.
Abranches, J., Zeng, L., Kajfasz, J. K., Palmer, S. R., Chakraborty, B., Wen, Z. T., et al. (2018). Biology of oral streptococci. Microbiol. Spectrum 6, 10–1128. doi: 10.1128/microbiolspec.gpp3-0042-2018
Acosta, J. N., Falcone, G. J., Rajpurkar, P., and Topol, E. J. (2022). Multimodal biomedical AI. Nat. Med. 28, 1773–1784.
Aislabie, J., and Deslippe, J. R. (2013). Soil microbes and their contribution to soil services. Ecosyst. Serv. New Zealand Conditions Trends 1, 143–161.
Al-Ani, L. K. T., Soares, F. E. D., Sharma, A., De Los Santos-Villalobos, S., Valdivia-Padilla, A. V., and Aguilar-Marcelino, L. (2022). Strategy of nematophagous fungi in determining the activity of plant parasitic nematodes and their prospective role in sustainable agriculture. Front. Fungal Biol. 3:863198. doi: 10.3389/ffunb.2022.863198
Aldrete-Tapia, J. A., Escalante-Minakata, P., Martínez-Peniche, R. A., Tamplin, M. L., and Hernández-Iturriaga, M. (2020). Yeast and bacterial diversity, dynamics and fermentative kinetics during small-scale tequila spontaneous fermentation. Food Microbiol. 86:103339. doi: 10.1016/j.fm.2019.103339
Amann, R. I., Baichoo, S., Blencowe, B. J., Bork, P., Borodovsky, M., Brooksbank, C., et al. (2019). Toward unrestricted use of public genomic data. Science 363, 350–352.
Aminu, S., Ascandari, A., Laamarti, M., Safdi, N. E. H., El Allali, A., and Daoud, R. (2024). Exploring microbial worlds: A review of whole genome sequencing and its application in characterizing the microbial communities. Crit. Rev. Microbiol. 50, 805–829. doi: 10.1080/1040841X.2023.2282447
Amit, S. K., Uddin, M. M., Rahman, R., Islam, S. R., and Khan, M. S. (2017). A review on mechanisms and commercial aspects of food preservation and processing. Agric. Food Sec. 6, 1–22.
Andrews, M., and Andrews, M. E. (2017). Specificity in legume-rhizobia symbioses. Int. J. Mol. Sci. 18:705.
Aplevicz, K. S., Ogliari, P. J., and Sant’anna, E. S. (2013). Influence of fermentation time on characteristics of sourdough bread. Braz. J. Pharm. Sci. 49, 233–239.
Asnicar, F., Berry, S. E., Valdes, A. M., Nguyen, L. H., Piccinno, G., Drew, D. A., et al. (2021). Microbiome connections with host metabolism and habitual diet from 1,098 deeply phenotyped individuals. Nat. Med. 27, 321–332. doi: 10.1038/s41591-020-01183-8
Ayilara, M. S., and Babalola, O. O. (2023). Bioremediation of environmental wastes: The role of microorganisms. Front. Agron. 5:1183691. doi: 10.3389/fagro.2023.1183691
Aziz, M., Nadipalli, R. K., Xie, X., Sun, Y., Surowiec, K., Zhang, J. L., et al. (2016). Augmenting sulfur metabolism and herbivore defense in Arabidopsis by bacterial volatile signaling. Front. Plant. Sci. 7:458. doi: 10.3389/fpls.2016.00458
Baedke, J., Fábregas-Tejeda, A., and Nieves Delgado, A. (2020). The holobiont concept before Margulis. J. Exp. Zool. B Mol. Dev. Evol. 334, 149–155.
Bala, S., Garg, D., Thirumalesh, B. V., Sharma, M., Sridhar, K., Inbaraj, B. S., et al. (2022). Recent strategies for bioremediation of emerging pollutants: A review for a green and sustainable environment. Toxics 10:484. doi: 10.3390/toxics10080484
Barea, J. M. (2015). Future challenges and perspectives for applying microbial biotechnology in sustainable agriculture based on a better understanding of plant-microbiome interactions. J. Soil Sci. Plant. Nutr. 15, 261–282.
Bautista-Gallego, J., Alessandria, V., Fontana, M., Bisotti, S., Taricco, S., Dolci, P., et al. (2014). Diversity and functional characterization of Lactobacillus spp. isolated throughout the ripening of a hard cheese. Int. J. Food Microbiol. 181, 60–66. doi: 10.1016/j.ijfoodmicro.2014.04.020
Becerra-Lucio, P. A., Diego-García, E., Guillén-Navarro, K., and Peña-Ramírez, Y. J. (2022). Unveiling the microbial ecology behind mezcal: A spirit drink with a growing global demand. Fermentation 8:662.
Bedu-Ferrari, C., Biscarrat, P., Langella, P., and Cherbuy, C. (2022). Prebiotics and the human gut microbiota: From breakdown mechanisms to the impact on metabolic health. Nutrients 14:2096. doi: 10.3390/nu14102096
Behera, S. S., El Sheikha, A. F., Hammami, R., and Kumar, A. (2020). Traditionally fermented pickles: How the microbial diversity associated with their nutritional and health benefits? J. Funct. Foods 70:103971.
Behera, S., Priyadarshanee, M., and Vandana Das, S. (2022). Polyhydroxyalkanoates, the bioplastics of microbial origin: Properties, biochemical synthesis, and their applications. Chemosphere 294:133723. doi: 10.1016/j.chemosphere.2022.133723
Behrouzi, A., Nafari, A. H., and Siadat, S. D. (2019). The significance of microbiome in personalized medicine. Clin. Transl. Med. 8:16.
Belimov, A. A., Dodd, I. C., Safronova, V. I., Dumova, V. A., Shaposhnikov, A. I., Ladatko, A. G., et al. (2014). Abscisic acid metabolizing rhizobacteria decrease ABA concentrations in planta and alter plant growth. Plant. Physiol. Biochem. 74, 84–91. doi: 10.1016/j.plaphy.2013.10.032
Berg, G., Köberl, M., Rybakova, D., Müller, H., Grosch, R., and Smalla, K. (2017). Plant microbial diversity is suggested as the key to future biocontrol and health trends. FEMS Microbiol. Ecol. 93:fix050. doi: 10.1093/femsec/fix050
Berg, G., Rybakova, D., Fischer, D., Cernava, T., Vergès, M.-C. C., Charles, T., et al. (2020). Microbiome definition re-visited: Old concepts and new challenges. Microbiome 8, 1–22.
Bhat, B. A., Tariq, L., Nissar, S., Islam, S. T., Islam, S. U., Mangral, Z., et al. (2022). The role of plant-associated rhizobacteria in plant growth, biocontrol and abiotic stress management. J. Appl. Microbiol. 133, 2717–2741. doi: 10.1111/jam.15796
Bianconi, I., Aschbacher, R., and Pagani, E. (2023). Current uses and future perspectives of genomic technologies in clinical microbiology. Antibiotics 12:1580. doi: 10.3390/antibiotics12111580
Blaha, D., Prigent-Combaret, C., Mirza, M. S., and Moenne-Loccoz, Y. (2006). Phylogeny of the 1-aminocyclopropane-1-carboxylic acid deaminase-encoding gene acdS in phytobeneficial and pathogenic Proteobacteria and relation with strain biogeography. FEMS Microbiol. Ecol. 56, 455–470. doi: 10.1111/j.1574-6941.2006.00082.x
Bogati, K., and Walczak, M. (2022). The impact of drought stress on soil microbial community, enzyme activities and plants. Agronomy 12:189.
Brethauer, S., and Studer, M. H. (2014). Consolidated bioprocessing of lignocellulose by a microbial consortium. Energy Environ. Sci. 7, 1446–1453.
Brinker, P., Fontaine, M. C., Beukeboom, L. W., and Salles, J. F. (2019). Host, symbionts, and the microbiome: The missing tripartite interaction. Trends Microbiol. 27, 480–488. doi: 10.1016/j.tim.2019.02.002
Brown, A. C., Ibrahim, S. A., and Song, D. (2016). Poi History, Uses, and Role in health. Fruits, Vegetables, and Herbs. Amsterdam: Elsevier.
Buck, M., and Hamilton, C. (2011). The nagoya protocol on access to genetic resources and the fair and equitable sharing of benefits arising from their utilization to the convention on biological diversity. Rev. Europ. Commun. Internat. Environ. Law 20, 47–61.
Buerger, A. N., Allen, H. L., Divis, H. R., Encina, E., Parker, J., and Boles, C. (2023). The worker microbiome and envirobiomes in the agriculture industry: A narrative review of current applications and understanding for worker health. J. Public Health Emerg. 7:21.
Byrd, A. L., Belkaid, Y., and Segre, J. A. (2018). The human skin microbiome. Nat. Rev. Microbiol. 16, 143–155.
Cal, A. J., Kibblewhite, R. E., Sikkema, W. D., Torres, L. F., Hart-Cooper, W. M., Orts, W. J., et al. (2021). Production of polyhydroxyalkanoate copolymers containing 4-hydroxybutyrate in engineered Bacillus megaterium. Int. J. Biol. Macromol. 168, 86–92. doi: 10.1016/j.ijbiomac.2020.12.015
Camargo, A. P., De Souza, R. S. C., Jose, J., Gerhardt, I. R., Dante, R. A., Mukherjee, S., et al. (2023). Plant microbiomes harbor potential to promote nutrient turnover in impoverished substrates of a Brazilian biodiversity hotspot. ISME J. 17, 354–370. doi: 10.1038/s41396-022-01345-1
Campbell, C., Kandalgaonkar, M. R., Golonka, R. M., Yeoh, B. S., Vijay-Kumar, M., and Saha, P. (2023). Crosstalk between gut microbiota and host immunity: Impact on inflammation and immunotherapy. Biomedicines 11:294.
Carrillo-Castañeda, G., Juárez Muñoz, J., Ramón Peralta-Videa, J., Gomez, E., and Gardea-Torresdey, J. L. (2002). Plant growth-promoting bacteria promote copper and iron translocation from root to shoot in alfalfa seedlings. J. Plant Nutr. 26, 1801–1814.
Carter, K. R., Nachtsheim, A. C., Dickman, L. T., Moore, E. R., Negi, S., Heneghan, J. P., et al. (2023). Drought conditioning of rhizosphere microbiome influences maize water use traits. Plant Soil 492, 587–604.
Carvalhais, L. C., Dennis, P. G., Badri, D. V., Kidd, B. N., Vivanco, J. M., and Schenk, P. M. (2015). Linking jasmonic acid signaling, root exudates, and rhizosphere microbiomes. Mol. Plant Microbe Interact 28, 1049–1058. doi: 10.1094/MPMI-01-15-0016-R
Carvalhais, L. C., Dennis, P. G., Badri, D. V., Tyson, G. W., Vivanco, J. M., and Schenk, P. M. (2013). Activation of the jasmonic acid plant defence pathway alters the composition of rhizosphere bacterial communities. PLoS One 8:e56457. doi: 10.1371/journal.pone.0056457
Castro-Camba, R., Sanchez, C., Vidal, N., and Vielba, J. M. (2022). Plant development and crop yield: The role of gibberellins. Plants 11:2650.
Chen, G.-Q. (2010). “Plastics completely synthesized by bacteria: Polyhydroxyalkanoates,” in Plastics From Bacteria: Natural Functions and Applications, ed. G. G.-Q. CHEN (Berlin: Springer).
Chen, J., Wang, X., Tang, D., and Wang, W. (2019). Oxidative stress adaptation improves the heat tolerance of Pseudomonas fluorescens SN15-2. Biol. Control 138:104070.
Chen, Q., Song, Y., An, Y., Lu, Y., and Zhong, G. (2024). Soil microorganisms: Their role in enhancing crop nutrition and health. Diversity 16:734. doi: 10.1002/elps.201300568
Chen, W., Modi, D., and Picot, A. (2023). Soil and phytomicrobiome for plant disease suppression and management under climate change: A review. Plants 12:2736. doi: 10.3390/plants12142736
Chen, X. H., Gu, Y., Zhou, X. F., and Zhang, Y. L. (2014). Asparagus stem as a new lignocellulosic biomass feedstock for anaerobic digestion: Increasing hydrolysis rate, methane production and biodegradability by alkaline pretreatment. Bioresour. Technol. 164, 78–85. doi: 10.1016/j.biortech.2014.04.070
Chen, X., Lu, Y., Chen, T., and Li, R. (2021). The female vaginal microbiome in health and bacterial vaginosis. Front. Cell Infect. Microbiol. 11:631972. doi: 10.3389/fcimb.2021.631972
Chen, X., Marszalkowska, M., and Reinhold-Hurek, B. (2019). Jasmonic acid, not salicyclic acid restricts endophytic root colonization of rice. Front. Plant Sci. 10:1758. doi: 10.3389/fpls.2019.01758
Cheng, M., Chen, D., Parales, R. E., and Jiang, J. (2022). Oxygenases as powerful weapons in the microbial degradation of pesticides. Annu. Rev. Microbiol. 76, 325–348. doi: 10.1146/annurev-micro-041320-091758
Chepsergon, J., and Moleleki, L. N. (2023). Rhizosphere bacterial interactions and impact on plant health. Curr. Opin. Microbiol. 73:102297.
Chieb, M., and Gachomo, E. W. (2023). The role of plant growth promoting rhizobacteria in plant drought stress responses. BMC Plant Biol. 23:407. doi: 10.1186/s12870-023-04403-8
Choi, C., Kwon, J., Kim, S., Myung, S. J., Park, K., Sohn, C. I., et al. (2015). Efficacy of combination therapy with probiotics and mosapride in patients with IBS without diarrhea: A randomized, double-blind, placebo-controlled, multicenter, phase II trial. Neurogastroenterol. Motil. 27, 705–716. doi: 10.1111/nmo.12544
Clapp, M., Aurora, N., Herrera, L., Bhatia, M., Wilen, E., and Wakefield, S. (2017). Gut microbiota’s effect on mental health: The gut-brain axis. Clin. Pract. 7:987.
Coelho, G. D. P., Ayres, L. F. A., Barreto, D. S., Henriques, B. D., Prado, M., and Passos, C. M. D. (2021). Acquisition of microbiota according to the type of birth: An integrative review. Rev. Lat. Am. Enfermagem 29:e3446. doi: 10.1590/1518.8345.4466.3446
Cohen, A. C., Travaglia, C. N., Bottini, R., and Piccoli, P. N. (2009). Participation of abscisic acid and gibberellins produced by endophytic in the alleviation of drought effects in maize. Botany 87, 455–462.
Contreras-Cornejo, H. A., Del-Val, E., Macías-Rodríguez, L., Alarcón, A., González-Esquivel, C. E., and Larsen, J. (2018). Trichoderma atroviride, a maize root associated fungus, increases the parasitism rate of the fall armyworm Spodoptera frugiperda by its natural enemy Campoletis sonorensis. Soil Biol. Biochem. 122, 196–202.
Conway, J. M., Walton, W. G., Salas-Gonzalez, I., Law, T. F., Lindberg, C. A., Crook, L. E., et al. (2022). Diverse MarR bacterial regulators of auxin catabolism in the plant microbiome. Nat. Microbiol. 7, 1817–1833. doi: 10.1038/s41564-022-01244-3
Cordeiro, C., and Echer, F. R. (2019). Interactive effects of nitrogen-fixing bacteria inoculation and nitrogen fertilization on soybean yield in unfavorable edaphoclimatic environments. Sci. Rep. 9:15606.
Cristofori, F., Dargenio, V. N., Dargenio, C., Miniello, V. L., Barone, M., and Francavilla, R. (2021). Anti-inflammatory and immunomodulatory effects of probiotics in gut inflammation: A door to the body. Front. Immunol. 12:578386. doi: 10.3389/fimmu.2021.578386
Cullen, C. M., Aneja, K. K., Beyhan, S., Cho, C. E., Woloszynek, S., Convertino, M., et al. (2020). Emerging priorities for microbiome research. Front. Microbiol. 11:136. doi: 10.3389/fmicb.2020.00136
Das, T. K., Pradhan, S., Chakrabarti, S., Mondal, K. C., and Ghosh, K. (2022). Current status of probiotic and related health benefits. Appl. Food Res. 2:100185.
Davani-Davari, D., Negahdaripour, M., Karimzadeh, I., Seifan, M., Mohkam, M., Masoumi, S. J., et al. (2019). Prebiotics: Definition, types, sources, mechanisms, and clinical applications. Foods 8:92. doi: 10.3390/foods8030092
Ding, Y., Sun, T., Ao, K., Peng, Y., Zhang, Y., Li, X., et al. (2018). Opposite roles of salicylic acid receptors NPR1 and NPR3/NPR4 in transcriptional regulation of plant immunity. Cell 173, 1454–1467 e15. doi: 10.1016/j.cell.2018.03.044
du Toit, M., Engelbrecht, L., Lerm, E., and Krieger-Weber, S. (2011). Lactobacillus: The next generation of malolactic fermentation starter cultures—an overview. Food Biopro. Technol. 4, 876–906.
Ellis, J. T., and Magnuson, T. S. (2012). Thermostable and alkalistable xylanases produced by the thermophilic bacterium Anoxybacillus flavithermus TWXYL3. ISRN Microbiol. 2012:517524. doi: 10.5402/2012/517524
Fenchel, T., Blackburn, H., King, G. M., and Blackburn, T. H. (2012). Bacterial Biogeochemistry: The Ecophysiology of Mineral Cycling. Cambridge, MA: Academic Press.
Ferrol, N., Tamayo, E., and Vargas, P. (2016). The heavy metal paradox in arbuscular mycorrhizas: From mechanisms to biotechnological applications. J. Exp. Bot. 67, 6253–6265. doi: 10.1093/jxb/erw403
Finkel, O. M., Salas-González, I., Castrillo, G., Spaepen, S., Law, T. F., Teixeira, P., et al. (2019). The effects of soil phosphorus content on plant microbiota are driven by the plant phosphate starvation response. PLoS Biol. 17:e3000534. doi: 10.1371/journal.pbio.3000534
Francis, F., Jacquemyn, H., Delvigne, F., and Lievens, B. (2020). From diverse origins to specific targets: Role of microorganisms in indirect pest biological control. Insects 11:533. doi: 10.3390/insects11080533
Giacomini, J. J., Torres-Morales, J., Dewhirst, F. E., Borisy, G. G., and Mark Welch, J. L. (2023). Site specialization of human oral Veillonella species. Microbiol. Spectrum 11:e04042–22. doi: 10.1128/spectrum.04042-22
Gibert, A., Tozer, W., and Westoby, M. (2019). Plant performance response to eight different types of symbiosis. New Phytol. 222, 526–542. doi: 10.1111/nph.15392
Gilbert, S. F. (2014). Symbiosis as the way of eukaryotic life: The dependent co-origination of the body. J. Biosci. 39, 201–209. doi: 10.1007/s12038-013-9343-6
Giraffa, G., Chanishvili, N., and Widyastuti, Y. (2010). Importance of lactobacilli in food and feed biotechnology. Res. Microbiol. 161, 480–487.
Giudici, P., Lemmetti, F., and Mazza, S. (2015). Balsamic Vinegars: Tradition, Technology, Trade. Cham: Springer.
Glare, T., Caradus, J., Gelernter, W., Jackson, T., Keyhani, N., Köhl, J., et al. (2012). Have biopesticides come of age? Trends Biotechnol. 30, 250–258.
Gonin, M., Salas-Gonzalez, I., Gopaulchan, D., Frene, J. P., Roden, S., Van De Poel, B., et al. (2023). Plant microbiota controls an alternative root branching regulatory mechanism in plants. Proc. Natl. Acad. Sci. 120:e2301054120. doi: 10.1073/pnas.2301054120
Govarthanan, M., Jeon, C.-H., Jeon, Y.-H., Kwon, J.-H., Bae, H., and Kim, W. (2020). Non-toxic nano approach for wastewater treatment using Chlorella vulgaris exopolysaccharides immobilized in iron-magnetic nanoparticles. Int. J. Biol. Macromol. 162, 1241–1249. doi: 10.1016/j.ijbiomac.2020.06.227
Greenhough, B., Read, C. J., Lorimer, J., Lezaun, J., Mcleod, C., Benezra, A., et al. (2020). Setting the agenda for social science research on the human microbiome. Palgrave Commun. 6, 1–11.
Guan, T., Lin, Y., Chen, K., Ou, M., and Zhang, J. (2020). Physicochemical factors affecting microbiota dynamics during traditional solid-state fermentation of Chinese strong-flavor Baijiu. Front. Microbiol. 11:2090. doi: 10.3389/fmicb.2020.02090
Guo, P., Zhang, K., Ma, X., and He, P. (2020). Clostridium species as probiotics: Potentials and challenges. J. Anim. Sci. Biotechnol. 11, 1–10. doi: 10.1186/s40104-019-0402-1
Gupt, A., Naudiyal, S., Rani, A., and Kumar, S. (2024). Mental health and the microbiome: A review of psychological impacts of gut microflora. Curr. Pharmacol. Rep. 10, 223–236.
Gupta, R., Elkabetz, D., Leibman-Markus, M., Jami, E., and Bar, M. (2022). Cytokinin-microbiome interactions regulate developmental functions. Environ. Microbiome 17:2. doi: 10.1186/s40793-022-00397-2
Haque, M. M., Mosharaf, M. K., Khatun, M., Haque, M. A., Biswas, M. S., Islam, M. S., et al. (2020). Biofilm producing rhizobacteria with multiple plant growth-promoting traits promote growth of tomato under water-deficit stress. Front. Microbiol. 11:542053. doi: 10.3389/fmicb.2020.542053
Hartmann, M., and Six, J. (2023). Soil structure and microbiome functions in agroecosystems. Nat. Rev. Earth Environ. 4, 4–18.
Hemarajata, P., and Versalovic, J. (2013). Effects of probiotics on gut microbiota: Mechanisms of intestinal immunomodulation and neuromodulation. Therap. Adv. Gastroenterol. 6, 39–51.
Hewedy, O. A., Elsheery, N. I., Karkour, A. M., Elhamouly, N., Arafa, R. A., Mahmoud, G. A., et al. (2023). Jasmonic acid regulates plant development and orchestrates stress response during tough times. Environ. Exp. Bot. 208:105260.
Hobbie, E. A. (2006). Carbon allocation to ectomycorrhizal fungi correlates with belowground allocation in culture studies. Ecology 87, 563–569. doi: 10.1890/05-0755
Hou, K., Wu, Z. X., Chen, X. Y., Wang, J. Q., Zhang, D., Xiao, C., et al. (2022). Microbiota in health and diseases. Signal Transduct. Target Ther. 7:135.
Hu, S., and Bidochka, M. J. (2021). Abscisic acid implicated in differential plant responses of Phaseolus vulgaris during endophytic colonization by Metarhizium and pathogenic colonization by Fusarium. Sci. Rep. 11:11327. doi: 10.1038/s41598-021-90232-4
Hu, S., Liu, G., Zhang, L., Gan, Y., Wang, B., Freilich, S., et al. (2021). A synergistic consortium involved in rac-dichlorprop degradation as revealed by DNA stable isotope probing and metagenomic analysis. Appl. Environ. Microbiol. 87:e0156221. doi: 10.1128/AEM.01562-21
Hudson, M., Garrison, N. A., Sterling, R., Caron, N. R., Fox, K., Yracheta, J., et al. (2020). Rights, interests and expectations: Indigenous perspectives on unrestricted access to genomic data. Nat. Rev. Genet. 21, 377–384. doi: 10.1038/s41576-020-0228-x
Hussain, A., and Hasnain, S. (2011). Interactions of bacterial cytokinins and IAA in the rhizosphere may alter phytostimulatory efficiency of rhizobacteria. World J. Microbiol. Biotechnol. 27, 2645–2654.
Hutkins, R. W. (2008). Microbiology and Technology of Fermented Foods. Hoboken, NJ: John Wiley & Sons.
Iqbal, B., Li, G. L., Alabbosh, K. F., Hussain, H., Khan, I., Tariq, M., et al. (2023). Advancing environmental sustainability through microbial reprogramming in growth improvement, stress alleviation, and phytoremediation. Plant Stress 10:100283.
Iqbal, N., Khan, N. A., Ferrante, A., Trivellini, A., Francini, A., and Khan, M. I. R. (2017). Ethylene role in plant growth, development and senescence: Interaction with other phytohormones. Front. Plant Sci. 8:475. doi: 10.3389/fpls.2017.00475
Ji, J., Jin, W., Liu, S. J., Jiao, Z., and Li, X. (2023). Probiotics, prebiotics, and postbiotics in health and disease. Med. Comm. 4:e420.
Jiang, G. F., Zhang, Y. L., Gan, G. Y., Li, W. L., Wan, W., Jiang, Y. Q., et al. (2022). Exploring rhizo-microbiome transplants as a tool for protective plant-microbiome manipulation. ISME Commun. 2:10. doi: 10.1038/s43705-022-00094-8
Jing, Y. X., Yan, J. L., He, H. D., Yang, D. J., Xiao, L., Zhong, T., et al. (2014). Characterization of bacteria in the rhizosphere soils of Polygonum pubescens and their potential in promoting growth and Cd, Pb, Zn uptake by Brassica napus. Int. J. Phytoremediation 16, 321–333. doi: 10.1080/15226514.2013.773283
Johnson, L. J., De Bonth, A. C., Briggs, L. R., Caradus, J. R., Finch, S. C., Fleetwood, D. J., et al. (2013). The exploitation of epichloae endophytes for agricultural benefit. Fungal Divers. 60, 171–188.
Kamran, S., Shahid, I., Baig, D. N., Rizwan, M., Malik, K. A., and Mehnaz, S. (2017). Contribution of zinc solubilizing bacteria in growth promotion and zinc content of wheat. Front. Microbiol. 8:2593. doi: 10.3389/fmicb.2017.02593
Kang, S. M., Radhakrishnan, R., Lee, K. E., You, Y. H., Ko, J. H., Kim, J. H., et al. (2015). Mechanism of plant growth promotion elicited by sp LKE15 in oriental melon. Acta Agric. Scand. B Soil Plant Sci. 65, 637–647.
Kapoor, R., Soni, R., and Kaur, M. (2016). Gibberellins production by fluorescent Pseudomonas isolated from Rhizospheric soil of Malus and Pyrus. Int. J. Agric. Environ. Biotechnol. 9, 193–199.
Karcher, N., Nigro, E., Punčochář, M., Blanco-Míguez, A., Ciciani, M., Manghi, P., et al. (2021). Genomic diversity and ecology of human-associated Akkermansia species in the gut microbiome revealed by extensive metagenomic assembly. Genome Biol. 22:209. doi: 10.1186/s13059-021-02427-7
Keswani, C., Singh, S. P., Garcia-Estrada, C., Mezaache-Aichour, S., Glare, T. R., Borriss, R., et al. (2022). Biosynthesis and beneficial effects of microbial gibberellins on crops for sustainable agriculture. J. Appl. Microbiol. 132, 1597–1615.
Khan, A. L., Hamayun, M., Kim, Y. H., Kang, S. M., Lee, J. H., and Lee, I. J. (2011). Gibberellins producing endophytic sp LH02 influenced endogenous phytohormonal levels, isoflavonoids production and plant growth in salinity stress. Process Biochem. 46, 440–447.
Khan, A. L., Waqas, M., Khan, A. R., Hussain, J., Kang, S. M., Gilani, S. A., et al. (2013). Fungal endophyte Penicillium janthinellum LK5 improves growth of ABA-deficient tomato under salinity. World J. Microbiol. Biotechnol. 29, 2133–2144. doi: 10.1007/s11274-013-1378-1
Khan, S., Alvi, A. F., and Khan, N. A. (2024). Role of ethylene in the regulation of plant developmental processes. Stresses 4, 28–53.
Kolida, S., and Gibson, G. R. (2011). Synbiotics in health and disease. Annu. Rev. Food Sci. Technol. 2, 373–393.
Kostic, T., Schloter, M., Arruda, P., Berg, G., Charles, T. C., Cotter, P. D., et al. (2024). Concepts and criteria defining emerging microbiome applications. Microbial Biotechnol. 17:e14550.
Kulkarni, O. S., Mazumder, M., Kini, S., Hill, E. D., Aow, J. S. B., Phua, S. M. L., et al. (2024). Volatile methyl jasmonate from roots triggers host-beneficial soil microbiome biofilms. Nat. Chem. Biol. 20, 473–483. doi: 10.1038/s41589-023-01462-8
Kumar, S., Shukla, V., Dubey, M. K., and Upadhyay, R. S. (2021). Activation of defense response in common bean against stem rot disease triggered by Trichoderma erinaceum and Trichoderma viride. J. Basic Microbiol. 61, 910–922. doi: 10.1002/jobm.202000749
Lamers, L. P., Van Diggelen, J. M., Op Den, Camp, H. J., Visser, E. J., Lucassen, E. C., et al. (2012). Microbial transformations of nitrogen, sulfur, and iron dictate vegetation composition in wetlands: A review. Front. Microbiol. 3:156. doi: 10.3389/fmicb.2012.00156
Lange, L., Berg, G., Cernava, T., Champomier-Verges, M. C., Charles, T., Cocolin, L., et al. (2022). Microbiome ethics, guiding principles for microbiome research, use and knowledge management. Environ. Microbiome 17:50.
Law, S. R., Mathes, F., Paten, A. M., Alexandre, P. A., Regmi, R., Reid, C., et al. (2024). Life at the borderlands: Microbiomes of interfaces critical to One Health. FEMS Microbiol. Rev. 48:fuae008. doi: 10.1093/femsre/fuae008
LeGrand, K., Borarin, B., and Young, G. M. (2020). Tradition and fermentation science of prohok, an ethnic fermented fish product of Cambodia. J. Ethnic Foods 7, 1–19.
Leveau, J. H., and Lindow, S. E. (2005). Utilization of the plant hormone indole-3-acetic acid for growth by Pseudomonas putida strain 1290. Appl. Environ. Microbiol. 71, 2365–2371. doi: 10.1128/AEM.71.5.2365-2371.2005
Li, G., Liu, P., Zhao, J., Su, L., Zhao, M., Jiang, Z., et al. (2023). Correlation of microbiomes in “plant-insect-soil” ecosystem. Front. Microbiol. 14:1088532. doi: 10.3389/fmicb.2023.1088532
Li, J., Wang, J., Liu, H., Macdonald, C. A., and Singh, B. K. (2022). Application of microbial inoculants significantly enhances crop productivity: A meta-analysis of studies from 2010 to 2020. J. Sustain. Agric. 1, 216–225.
Li, N., Han, X., Feng, D., Yuan, D., and Huang, L.-J. (2019). Signaling crosstalk between salicylic acid and ethylene/jasmonate in plant defense: Do we understand what they are whispering? Int. J. Mol. Sci. 20:671. doi: 10.3390/ijms20030671
Li, X.-L., George, E., and Marschner, H. (1991). Extension of the phosphorus depletion zone in VA-mycorrhizal white clover in a calcareous soil. Plant Soil 136, 41–48.
Lim, J. W., Park, T., Tong, Y. W., and Yu, Z. (2020). The microbiome driving anaerobic digestion and microbial analysis. Adv. Bioenergy 5, 1–61.
Lim, Y.-B., Parr, L., Popescu, S., Rezzonico, A., Van Beck, L., and Zarfos, M. R. (2023). Addressing the Threat of Disease Spillover: U.S.-Based Best Practices at the Nexus of Ecological and Biological Security. Washington, DC: The Council on Strategic Risks.
Lima, T. T. M., Hosken, B. D. O., Venturim, B. C., Lopes, I. L., and Martin, J. G. P. (2022). Traditional Brazilian fermented foods: Cultural and technological aspects. J. Ethnic Foods 9:35. doi: 10.1590/S1517-83822013000200001
Liu, F. C., Xing, S. J., Ma, H. L., Du, Z. Y., and Ma, B. Y. (2013). Cytokinin-producing, plant growth-promoting rhizobacteria that confer resistance to drought stress in container seedlings. Appl. Microbiol. Biotechnol. 97, 9155–9164. doi: 10.1007/s00253-013-5193-2
Liu, H. Q., Lu, X. B., Li, Z. H., Tian, C. Y., and Song, J. (2021). The role of root-associated microbes in growth stimulation of plants under saline conditions. Land. Degrad. Dev. 32, 3471–3486.
Liu, H., Brettell, L. E., Qiu, Z., and Singh, B. K. (2020). Microbiome-mediated stress resistance in plants. Trends Plant Sci. 25, 733–743.
Liu, S., Greenhut, I. V., Heist, E. P., Heist, M. R., and Moe, L. A. (2023). Bacterial community dynamics during distilled spirit fermentation: Influence of mash recipes and fermentation processes. Microbiol. Spectrum 11:e01624-23. doi: 10.1128/spectrum.01624-23
Louw, N. L., Lele, K., Ye, R., Edwards, C. B., and Wolfe, B. E. (2023). Microbiome assembly in fermented foods. Annu. Rev. Microbiol. 77, 381–402.
Ma, L.-C., Zhao, H.-Q., Wu, L. B., and Liu, C. (2023). Impacts of the microbiome on human, animal, and environmental health from a One Health perspective. Sci. One Health 2:100037.
Ma, Y., Rajkumar, M., and Freitas, H. (2009). Improvement of plant growth and nickel uptake by nickel resistant-plant-growth promoting bacteria. J. Hazardous Mater. 166, 1154–1161. doi: 10.1016/j.jhazmat.2008.12.018
Ma, Y., Rajkumar, M., Luo, Y., and Freitas, H. (2013). Phytoextraction of heavy metal polluted soils using Sedum plumbizincicola inoculated with metal mobilizing Phyllobacterium myrsinacearum RC6b. Chemosphere 93, 1386–1392. doi: 10.1016/j.chemosphere.2013.06.077
Mahanty, S., Chatterjee, S., Ghosh, S., Tudu, P., Gaine, T., Bakshi, M., et al. (2020). Synergistic approach towards the sustainable management of heavy metals in wastewater using mycosynthesized iron oxide nanoparticles: Biofabrication, adsorptive dynamics and chemometric modeling study. J. Water Process Eng. 37:101426.
Mamindlapelli, N. K., Arelli, V., Juntupally, S., Begum, S., Thenkrishnan, K., Maddala, R. K., et al. (2021). Understanding the substrate mediated microbial community shift within the anaerobic ecosystems via 16S metagenomic studies. Bioresour. Technol. Rep. 15:100793.
Manoli, M. T., Tarazona, N., Mato, A., Maestro, B., Sanz, J. M., Nogales, J., et al. (2020). “Molecular basis of medium-chain length-PHA metabolism of Pseudomonas putida,” in The Handbook of Polyhydroxyalkanoates, ed. M. Koller (Boca Raton, FL: CRC Press), 89–114.
Manzoor, M., Abid, R., Rathinasabapathi, B., De Oliveira, L. M., Da Silva, E., Deng, F., et al. (2019). Metal tolerance of arsenic-resistant bacteria and their ability to promote plant growth of Pteris vittata in Pb-contaminated soil. Sci. Total Environ. 660, 18–24. doi: 10.1016/j.scitotenv.2019.01.013
Maqsood, Q., Sumrin, A., Waseem, R., Hussain, M., Imtiaz, M., and Hussain, N. (2023). Bioengineered microbial strains for detoxification of toxic environmental pollutants. Environ. Res. 227:115665.
MarketsandMarkets. (2024). Bioethanol Market by Feedstock (Starch Based, Sugar Based, Cellulose-based), Fuel Blend (E5, E10, E15 to E70, E75& E85), end-use (Transportation, Pharmaceutical, Cosmetic, Alcoholic Beverages), Generation and Region Global Forecast to 2028. Pune: MarketsandMarkets Research Private Ltd.
Martín, R., Rios-Covian, D., Huillet, E., Auger, S., Khazaal, S., Bermúdez-Humarán, L. G., et al. (2023). Faecalibacterium: A bacterial genus with promising human health applications. FEMS Microbiol. Rev. 47:fuad039. doi: 10.1093/femsre/fuad039
Mascarin, G. M., and Jaronski, S. T. (2016). The production and uses of as a microbial insecticide. World J. Microbiol. Biotechnol. 32, 1–26.
Masood, S., and Bano, A. (2016). “Mechanism of potassium solubilization in the agricultural soils by the help of soil microorganisms,” in Potassium Solubilizing Microorganisms for Sustainable Agriculture, eds V. S. Meena, B. R. Maurya, J. P. Verma, and R. S. Meena (New Delhi: Springer).
McAllister, T., Hikuroa, D., and Macinnis-Ng, C. (2023). Connecting science to indigenous knowledge. New Zealand J. Ecol. 47, 1–13.
McGuiness, P. N., Reid, J. B., and Foo, E. (2019). The role of gibberellins and brassinosteroids in nodulation and arbuscular mycorrhizal associations. Front. Plant Sci. 10:269. doi: 10.3389/fpls.2019.00269
Meena, V. S., Bahadur, I., Maurya, B. R., Kumar, A., Meena, R. K., Meena, S. K., et al. (2016). Potassium-Solubilizing Microorganism in Evergreen Agriculture: An Overview. Berlin: Springer.
Meisner, A., Wepner, B., Kostic, T., Van Overbeek, L. S., Bunthof, C. J., De Souza, R. S. C., et al. (2022). Calling for a systems approach in microbiome research and innovation. Curr. Opin. Biotechnol. 73, 171–178.
Mena, B., and Aryana, K. J. (2012). Influence of ethanol on probiotic and culture bacteria Lactobacillus bulgaricus and Streptococcus thermophilus within a therapeutic product. Open J. Med. Microbiol. 2, 70–76.
Mendoza-Mendoza, A., Esquivel-Naranjo, E. U., Soth, S., Whelan, H., Alizadeh, H., Echaide-Aquino, J. F., et al. (2024). Uncovering the multifaceted properties of 6-pentyl-alpha-pyrone for control of plant pathogens. Front. Plant Sci. 15:1420068. doi: 10.3389/fpls.2024.1420068
Meybodi, N. M., Ebrahimi, M. T., and Mortazavian, A. M. (2016). “Ethnic fermented foods and beverage of Iran,” in Ethnic Fermented Foods and Alcoholic Beverages of Asia, ed. J. Tamang (Berlin: Springer), 309–322. doi: 10.1177/1082013211428212
Molefe, R. R., Amoo, A. E., and Babalola, O. O. (2023). Communication between plant roots and the soil microbiome; Involvement in plant growth and development. Symbiosis 90, 231–239.
Montreemuk, J., Stewart, T. N., and Prapagdee, B. (2024). Bacterial-assisted phytoremediation of heavy metals: Concepts, current knowledge, and future directions. Environ. Technol. Innov. 33:103488.
Mora, C., Mckenzie, T., Gaw, I. M., Dean, J. M., Von Hammerstein, H., Knudson, T. A., et al. (2022). Over half of known human pathogenic diseases can be aggravated by climate change. Nat. Climate Change 12, 869–875. doi: 10.1038/s41558-022-01426-1
Moran-Diez, M. E., Martinez, De Alba, A. E., Rubio, M. B., Hermosa, R., and Monte, E. (2021). Trichoderma and the plant heritable priming responses. J. Fungi 7:318. doi: 10.3390/jof7040318
Moreno, M. R., Leisner, J. J., Tee, L. K., Ley, C., Radu, S., Rusul, G., et al. (2002). Microbial analysis of Malaysian tempeh, and characterization of two bacteriocins produced by isolates of Enterococcus faecium. J. Appl. Microbiol. 92, 147–157. doi: 10.1046/j.1365-2672.2002.01509.x
Mukherjee, A., Lordan, C., Ross, R. P., and Cotter, P. D. (2020). Gut microbes from the phylogenetically diverse genus Eubacterium and their various contributions to gut health. Gut Microbes 12:1802866. doi: 10.1080/19490976.2020.1802866
Munir, N., Hanif, M., Abideen, Z., Sohail, M., El-Keblawy, A., Radicetti, E., et al. (2022). Mechanisms and strategies of plant microbiome interactions to mitigate abiotic stresses. Agronomy 12:2069.
Murooka, Y., and Yamshita, M. (2008). Traditional healthful fermented products of Japan. J. Industrial Microbiol. Biotechnol. 35:791.
Nagda, A., Meena, M., and Shah, M. P. (2022). Bioremediation of industrial effluents: A synergistic approach. J. Basic Microbiol. 62, 395–414.
Nathan, V. K., and Ammini, P. (2019). Carbon dioxide sequestering ability of bacterial carbonic anhydrase in a mangrove soil microcosm and its bio-mineralization properties. Water Air Soil Pollut. 230, 1–12.
Neela, S., and Fanta, S. W. (2020). Injera (An ethnic, traditional staple food of Ethiopia): A review on traditional practice to scientific developments. J. Ethnic Foods 7:32.
Ngcamu, S. P. (2023). The Effects of Sauerkraut on Human Health, Nutrition, and Food Security: A Review of the Literature. Stellenbosch: Stellenbosch University.
Niu, C., Xing, X., Wang, Y., Li, X., Zheng, F., Liu, C., et al. (2023). Characterization of color, metabolites and microbial community dynamics of doubanjiang during constant temperature fermentation. Food Res. Int. 174:113554. doi: 10.1016/j.foodres.2023.113554
Nogueira, M. A., and Cardoso, E. J. B. N. (2003). Mycorrhizal effectiveness and manganese toxicity in soybean as affected by soil type and endophyte. Sci. Agric. 60, 329–335.
Nout, M. R., and Aidoo, K. E. (2011). “Asian fungal fermented food,” in Industrial Applications, ed. M. Hofrichter (New York, NY: Springer-Verlag), 23–47.
O’Mahony, S., and Comizzoli, P. (2023). Special series on the role of the microbiome in reproduction and fertility. Reprod. Fertil. 4:e230080. doi: 10.1530/RAF-23-0080
O’Callaghan, A., and Van Sinderen, D. (2016). Bifidobacteria and their role as members of the human gut microbiota. Front. Microbiol. 7:925. doi: 10.3389/fmicb.2016.00925
Ojeda-Linares, C., Álvarez-Ríos, G. D., Figueredo-Urbina, C. J., Islas, L. A., Lappe-Oliveras, P., Nabhan, G. P., et al. (2021). Traditional fermented beverages of Mexico: A biocultural unseen foodscape. Foods 10:2390. doi: 10.3390/foods10102390
Pandya, N., and Desai, P. (2014). Screening and characterization of GA3 producing Pseudomonas monteilii and its impact on plant growth promotion. Int. J. Curr. Microbiol. Appl. Sci. 3, 110–115.
Panigrahi, P., Parida, S., Nanda, N. C., Satpathy, R., Pradhan, L., Chandel, D. S., et al. (2017). A randomized synbiotic trial to prevent sepsis among infants in rural India. Nature 548, 407–412.
Paries, M., and Gutjahr, C. (2023). The good, the bad, and the phosphate: Regulation of beneficial and detrimental plant-microbe interactions by the plant phosphate status. New Phytol. 239, 29–46. doi: 10.1111/nph.18933
Peixoto, R. S., Voolstra, C. R., Sweet, M., Duarte, C. M., Carvalho, S., Villela, H., et al. (2022). Harnessing the microbiome to prevent global biodiversity loss. Nat. Microbiol. 7, 1726–1735. doi: 10.1038/s41564-022-01173-1
Philip, S., Keshavarz, T., and Roy, I. (2007). Polyhydroxyalkanoates: Biodegradable polymers with a range of applications. J. Chem. Technol. Biotechnol. 82, 233–247.
Piao, H., Hawley, E., Kopf, S., Descenzo, R., Sealock, S., Henick-Kling, T., et al. (2015). Insights into the bacterial community and its temporal succession during the fermentation of wine grapes. Front. Microbiol. 6:809. doi: 10.3389/fmicb.2015.00809
Pieterse, C. M., Leon-Reyes, A., Van Der Ent, S., and Van Wees, S. C. (2009). Networking by small-molecule hormones in plant immunity. Nat. Chem. Biol. 5, 308–316. doi: 10.1038/nchembio.164
Pinto, C., Pinho, D., Cardoso, R., Custódio, V., Fernandes, J., Sousa, S., et al. (2015). Wine fermentation microbiome: A landscape from different Portuguese wine appellations. Front. Microbiol. 6:905. doi: 10.3389/fmicb.2015.00905
Poole, P. (2017). Shining a light on the dark world of plant root–microbe interactions. Proc. Natl. Acad. Sci. 114:4281. doi: 10.1073/pnas.1703800114
Prasad, D., Verma, N., Bakshi, M., Narayan, O. P., Singh, A. K., Dua, M., et al. (2018). Functional characterization of a magnesium transporter of root endophytic fungus Piriformospora indica. Front. Microbiol. 9:3231. doi: 10.3389/fmicb.2018.03231
Prescott, S. L., Larcombe, D.-L., Logan, A. C., West, C., Burks, W., Caraballo, L., et al. (2017). The skin microbiome: Impact of modern environments on skin ecology, barrier integrity, and systemic immune programming. World Allergy Organ. J. 10, 1–16. doi: 10.1186/s40413-017-0160-5
Quiroga, G., Erice, G., Aroca, R., and Ruiz-Lozano, J. M. (2020). Elucidating the possible involvement of maize aquaporins in the plant boron transport and homeostasis mediated by Rhizophagus irregularis under drought stress conditions. Int. J. Mol. Sci. 21:1748. doi: 10.3390/ijms21051748
Rafeeq, H., Afsheen, N., Rafique, S., Arshad, A., Intisar, M., Hussain, A., et al. (2023). Genetically engineered microorganisms for environmental remediation. Chemosphere 310:136751.
Rajkumar, M., Ae, N., Prasad, M. N., and Freitas, H. (2010). Potential of siderophore-producing bacteria for improving heavy metal phytoextraction. Trends Biotechnol. 28, 142–149.
Ramamurthy, P. C., Singh, S., Kapoor, D., Parihar, P., Samuel, J., Prasad, R., et al. (2021). Microbial biotechnological approaches: Renewable bioprocessing for the future energy systems. Microb. Cell Fact 20:55. doi: 10.1186/s12934-021-01547-w
Rani, N., Sangwan, P., Joshi, M., Sagar, A., and Bala, K. (2019). “Microbes: A key player in industrial wastewater treatment,” in Microbial Wastewater Treatment, eds M. P. Shah and S. Rodriguez-Couto (Amsterdam: Elsevier). doi: 10.1016/j.envres.2024.120216
Rasool, K., Hussain, S., Shahzad, A., Miran, W., Mahmoud, K. A., Ali, N., et al. (2023). Comprehensive insights into sustainable conversion of agricultural and food waste into microbial protein for animal feed production. Rev. Environ. Sci. Biotechnol. 22, 527–562.
Rastogi, M., and Shrivastava, S. (2017). Recent advances in second generation bioethanol production: An insight to pretreatment, saccharification and fermentation processes. Renew. Sustain. Energy Rev. 80, 330–340.
Ravanbakhsh, M., Sasidharan, R., Voesenek, L. A., Kowalchuk, G. A., and Jousset, A. (2018). Microbial modulation of plant ethylene signaling: Ecological and evolutionary consequences. Microbiome 6, 1–10. doi: 10.1186/s40168-018-0436-1
Raza, T., Qadir, M. F., Khan, K. S., Eash, N. S., Yousuf, M., Chatterjee, S., et al. (2023). Unrevealing the potential of microbes in decomposition of organic matter and release of carbon in the ecosystem. J. Environ. Manage 344:118529. doi: 10.1016/j.jenvman.2023.118529
Rea, K., Dinan, T. G., and Cryan, J. F. (2016). The microbiome: A key regulator of stress and neuroinflammation. Neurobiol. Stress 4, 23–33.
Reusche, M., Klaskova, J., Thole, K., Truskina, J., Novak, O., Janz, D., et al. (2013). Stabilization of cytokinin levels enhances Arabidopsis resistance against Verticillium longisporum. Mol. Plant Microbe Interact. 26, 850–860. doi: 10.1094/MPMI-12-12-0287-R
Ricca, E., Calabro, V., Curcio, S., and Iorio, G. (2007). The state of the art in the production of fructose from inulin enzymatic hydrolysis. Crit. Rev. Biotechnol. 27, 129–145. doi: 10.1080/07388550701503477
Roch, F.-F., Dzieciol, M., Quijada, N. M., Alteio, L. V., Mester, P.-J., and Selberherr, E. (2024). Microbial community structure of plant-based meat alternatives. npj Sci. Food 8:27.
Ropars, J., Caron, T., Lo, Y. C., Bennetot, B., and Giraud, T. (2020). The domestication of Penicillium cheese fungi. C R Biol. 343, 155–176.
Ruan, Z., Chen, K., Cao, W., Meng, L., Yang, B., Xu, M., et al. (2024). Engineering natural microbiomes toward enhanced bioremediation by microbiome modeling. Nat. Commun. 15:4694.
Saberi-Riseh, R., Fathi, F., and Moradzadeh-Eskandari, M. (2020). Effect of some Pseudomonas fluorescens and Bacillus subtilis strains on osmolytes and antioxidants of cucumber under salinity stress. J. Crop Protect. 9, 1–16.
Salas-Gonzalez, I., Reyt, G., Flis, P., Custodio, V., Gopaulchan, D., Bakhoum, N., et al. (2021). Coordination between microbiota and root endodermis supports plant mineral nutrient homeostasis. Science 371:eabd0695.
San Keskin, N. O., Celebioglu, A., Sarioglu, O. F., Uyar, T., and Tekinay, T. (2018). Encapsulation of living bacteria in electrospun cyclodextrin ultrathin fibers for bioremediation of heavy metals and reactive dye from wastewater. Colloids Surf B Biointerfaces 161, 169–176. doi: 10.1016/j.colsurfb.2017.10.047
S̨anlibaba, P., and Tezel, B. (2023). Traditional fermented foods in Anatolia. Acta Sci. Pol. Technol. Aliment. 22, 193–215.
Santoyo, G. (2022). How plants recruit their microbiome? New insights into beneficial interactions. J. Adv. Res. 40, 45–58.
Sarathambal, C., Khankhane, P. J., Gharde, Y., Kumar, B., Varun, M., and Arun, S. (2017). The effect of plant growth-promoting rhizobacteria on the growth, physiology, and Cd uptake of Arundo donax L. Int. J. Phytoremediation 19, 360–370. doi: 10.1080/15226514.2016.1225289
Sathya, A. B., Sivasubramanian, V., Santhiagu, A., Sebastian, C., and Sivashankar, R. (2018). Production of polyhydroxyalkanoates from renewable sources using bacteria. J. Polym. Environ. 26, 3995–4012.
Sedghi, L., Dimassa, V., Harrington, A., Lynch, S. V., and Kapila, Y. L. (2021). The oral microbiome: Role of key organisms and complex networks in oral health and disease. Periodontol 87, 107–131.
Sender, R., Fuchs, S., and Milo, R. (2016). Revised estimates for the number of human and bacteria cells in the body. PLoS Biol. 14:e1002533. doi: 10.1371/journal.pbio.1002533
Sergaki, C., Lagunas, B., Lidbury, I., Gifford, M. L., and Schafer, P. (2018). Challenges and approaches in microbiome research: From fundamental to applied. Front. Plant Sci. 9:1205. doi: 10.3389/fpls.2018.01205
Sessitsch, A., Wakelin, S., Schloter, M., Maguin, E., Cernava, T., Champomier-Verges, M. C., et al. (2023). Microbiome interconnectedness throughout environments with major consequences for healthy people and a healthy planet. Microbiol. Mol. Biol. Rev. 87:e0021222. doi: 10.1128/mmbr.00212-22
Shahid, M. J., Al-Surhanee, A. A., Kouadri, F., Ali, S., Nawaz, N., Afzal, M., et al. (2020). Role of microorganisms in the remediation of wastewater in floating treatment wetlands: A review. Sustainability 12:5559.
Shani, E., Salehin, M., Zhang, Y., Sanchez, S. E., Doherty, C., Wang, R., et al. (2017). Plant stress tolerance requires auxin-sensitive aux/iaa transcriptional repressors. Curr. Biol. 27, 437–444. doi: 10.1016/j.cub.2016.12.016
Sharma, P., Pandey, A. K., Udayan, A., and Kumar, S. (2021). Role of microbial community and metal-binding proteins in phytoremediation of heavy metals from industrial wastewater. Bioresour. Technol. 326:124750. doi: 10.1016/j.biortech.2021.124750
Sharma, R., Garg, P., Kumar, P., Bhatia, S. K., and Kulshrestha, S. (2020). Microbial fermentation and its role in quality improvement of fermented foods. Fermentation 6:106.
Sharma, V., Tsai, M. L., Nargotra, P., Chen, C. W., Kuo, C. H., Sun, P. P., et al. (2022). Agro-industrial food waste as a low-cost substrate for sustainable production of industrial enzymes: A critical review. Catalysts 12:1373.
Shi, S., Marshall, S., Schon, N., Dignam, B., Bell, N., and O’callaghan, M. (2021). Insights into the soil microbiome and prospects for its manipulation for improved pasture resilience. NZGA Res. Pract. Ser. 17, 163–178.
Shi, Z., Guo, X., Lei, Z., Wang, Y., Yang, Z., Niu, J., et al. (2023). Screening of high-efficiency nitrogen-fixing bacteria from the traditional Chinese medicine plant Astragalus mongolicus and its effect on plant growth promotion and bacterial communities in the rhizosphere. BMC Microbiol. 23:292. doi: 10.1186/s12866-023-03026-1
Shi, Z., Zhang, J., Lu, S., Li, Y., and Wang, F. (2020). Arbuscular mycorrhizal fungi improve the performance of sweet sorghum grown in a mo-contaminated soil. J. Fungi 6:44. doi: 10.3390/jof6020044
Shin, D.-H., Kim, Y.-M., Park, W.-S., and Kim, J.-H. (2016). “Ethnic fermented foods and beverages of Korea,” in Ethnic Fermented Foods and Alcoholic Beverages of Asia, ed. J. P. Tamang (Berlin: Springer), 263–308.
Siddiqui, S. A., Erol, Z., Rugji, J., Tasçi, F., Kahraman, H. A., Toppi, V., et al. (2023). An overview of fermentation in the food industry - looking back from a new perspective. Bioresour. Bioprocess. 10:85. doi: 10.1186/s40643-023-00702-y
Simon, E., Calinoiu, L. F., Mitrea, L., and Vodnar, D. C. (2021). Probiotics, prebiotics, and synbiotics: Implications and beneficial effects against irritable bowel syndrome. Nutrients 13:2112. doi: 10.3390/nu13062112
Singh, S. K., Wu, X., Shao, C., and Zhang, H. (2022). Microbial enhancement of plant nutrient acquisition. Stress Biol. 2:3.
Smythe, P., and Wilkinson, H. N. (2023). The skin microbiome: Current landscape and future opportunities. Int. J. Mol. Sci. 24:3950. doi: 10.3390/ijms24043950
Solomon, W., Mutum, L., Janda, T., and Molnár, Z. (2023). Potential benefit of microalgae and their interaction with bacteria to sustainable crop production. Plant Growth Regul. 101, 53–65.
Soth, S., Glare, T. R., Hampton, J. G., Card, S. D., and Brookes, J. J. (2022a). Biological control of diamondback moth—increased efficacy with mixtures of Beauveria fungi. Microorganisms 10:646. doi: 10.3390/microorganisms10030646
Soth, S., Glare, T. R., Hampton, J. G., Card, S. D., Brookes, J. J., and Narciso, J. O. (2022b). You are what you eat: Fungal metabolites and host plant affect the susceptibility of diamondback moth to entomopathogenic fungi. Peerj 10:e14491. doi: 10.7717/peerj.14491
Stec, N., Banasiak, J., and Jasinski, M. (2016). Abscisic acid - An overlooked player in plant-microbe symbioses formation? Acta Biochim. Pol. 63, 53–58. doi: 10.18388/abp.2015_1210
Stracquadanio, C., Quiles, J. M., Meca, G., and Cacciola, S. O. (2020). Antifungal activity of bioactive metabolites produced by Trichoderma asperellum and Trichoderma atroviride in liquid medium. J. Fungi 6:263. doi: 10.3390/jof6040263
Su, Y. W., Wang, J., Gao, W. Y., Wang, R. B., Yang, W. Q., Zhang, H. Y., et al. (2023). Dynamic metabolites: A bridge between plants and microbes. Sci. Total Environ. 899:165612. doi: 10.1016/j.scitotenv.2023.165612
Suganuma, N., Yamauchi, H., and Yamamoto, K. (1995). Enhanced production of ethylene by soybean roots after inoculation with Bradyrhizobium japonicum. Plant Sci. 111, 163–168.
Suiker, I. M., and Wösten, H. A. (2022). Spoilage yeasts in beer and beer products. Curr. Opin. Food Sci. 44:100815.
Suman, J., Rakshit, A., Ogireddy, S. D., Singh, S., Gupta, C., and Chandrakala, J. (2022). Microbiome as a key player in sustainable agriculture and human health. Front. Soil Sci. 2:821589. doi: 10.3389/fsoil.2022.821589
Sun, X., Xu, Z., Xie, J., Hesselberg-Thomsen, V., Tan, T., Zheng, D., et al. (2022). Bacillus velezensis stimulates resident rhizosphere Pseudomonas stutzeri for plant health through metabolic interactions. ISME J. 16, 774–787. doi: 10.1038/s41396-021-01125-3
Tamang, J. P. (2022). “Ethno-microbiology” of ethnic Indian fermented foods and alcoholic beverages. J. Appl. Microbiol. 133, 145–161. doi: 10.1111/jam.15382
Tamang, J. P., Cotter, P. D., Endo, A., Han, N. S., Kort, R., Liu, S. Q., et al. (2020). Fermented foods in a global age: East meets West. Compr. Rev. Food Sci. Food Saf. 19, 184–217. doi: 10.1111/1541-4337.12520
Todero Ritter, C. E., Camassola, M., Zampieri, D., Silveira, M. M., and Dillon, A. J. (2013). Cellulase and xylanase production by Penicillium echinulatum in submerged media containing cellulose amended with sorbitol. Enzyme Res. 2013:240219. doi: 10.1155/2013/240219
Tomasulo, A., Simionati, B., and Facchin, S. (2024). Microbiome One Health model for a healthy ecosystem. Science One Health 3:100065.
Trivedi, P., Schenk, P. M., Wallenstein, M. D., and Singh, B. K. (2017). Tiny microbes, big yields: Enhancing food crop production with biological solutions. Microbial Biotechnol. 10, 999–1003. doi: 10.1111/1751-7915.12804
United Nations. (2007). United Nations Declaration on the Rights of Indigenous Peoples. Washington, DC: United Nations.
Uzogara, S., Agu, L., and Uzogara, E. (1990). A review of traditional fermented foods, condiments and beverages in Nigeria: Their benefits and possible problems. Ecol. Food Nutr. 24, 267–288.
Valentino, V., De Filippis, F., Marotta, R., Pasolli, E., and Ercolini, D. (2024). Genomic features and prevalence of Ruminococcus species in humans are associated with age, lifestyle, and disease. Cell Rep. 43:115018. doi: 10.1016/j.celrep.2024.115018
van Bruggen, A. H., Goss, E. M., Havelaar, A., Van Diepeningen, A. D., Finckh, M. R., and Morris, J. G. (2019). One Health-Cycling of diverse microbial communities as a connecting force for soil, plant, animal, human and ecosystem health. Sci. Total Environ. 664, 927–937. doi: 10.1016/j.scitotenv.2019.02.091
Van der Meij, A., Willemse, J., Schneijderberg, M. A., Geurts, R., Raaijmakers, J. M., and Van Wezel, G. P. (2018). Inter-and intracellular colonization of Arabidopsis roots by endophytic actinobacteria and the impact of plant hormones on their antimicrobial activity. Antonie Van Leeuwenhoek 111, 679–690. doi: 10.1007/s10482-018-1014-z
Velmourougane, K., Prasanna, R., and Saxena, A. K. (2017). Agriculturally important microbial biofilms: Present status and future prospects. J. Basic Microbiol. 57, 548–573. doi: 10.1002/jobm.201700046
Vijay, A., and Valdes, A. M. (2022). Role of the gut microbiome in chronic diseases: A narrative review. Eur. J. Clin. Nutr. 76, 489–501.
Voidarou, C., Antoniadou, M., Rozos, G., Tzora, A., Skoufos, I., Varzakas, T., et al. (2020). Fermentative foods: Microbiology, biochemistry, potential human health benefits and public health issues. Foods 10:69. doi: 10.3390/foods10010069
Walker, G. M., and Hill, A. E. (2016). Saccharomyces cerevisiae in the production of whisk (e) y. Beverages 2:38.
Walker, G. M., and Stewart, G. G. (2016). Saccharomyces cerevisiae in the production of fermented beverages. Beverages 2:30.
Wang, C., Zhao, J., Zhang, H., Lee, Y.-K., Zhai, Q., and Chen, W. (2021). Roles of intestinal bacteroides in human health and diseases. Crit. Rev. Food Sci. Nutr. 61, 3518–3536. doi: 10.1080/10408398.2020.1802695
Wang, S., Qu, Y., Chang, L., Pu, Y., Zhang, K., and Hashimoto, K. (2020). Antibiotic-induced microbiome depletion is associated with resilience in mice after chronic social defeat stress. J. Affect. Disord. 260, 448–457. doi: 10.1016/j.jad.2019.09.064
Wang, W., Zhang, Z., Gao, J., and Wu, H. (2024). The impacts of microplastics on the cycling of carbon and nitrogen in terrestrial soil ecosystems: Progress and prospects. Sci. Total Environ. 915:169977. doi: 10.1016/j.scitotenv.2024.169977
War, A. R., Paulraj, M. G., Ahmad, T., Buhroo, A. A., Hussain, B., Ignacimuthu, S., et al. (2012). Mechanisms of plant defense against insect herbivores. Plant Signal Behav. 7, 1306–1320.
Warbrick, I., Heke, D., and Breed, M. (2023). Indigenous knowledge and the microbiome—bridging the disconnect between colonized places, peoples, and the unseen influences that shape our health and well-being. Msystems 8:e0875-22. doi: 10.1128/msystems.00875-22
Waweru, B., Turoop, L., Kahangi, E., Coyne, D., and Dubois, T. (2014). Non-pathogenic Fusarium oxysporum endophytes provide field control of nematodes, improving yield of banana (Musa sp.). Biol. Control 74, 82–88.
Wiltschi, B., Cernava, T., Dennig, A., Galindo Casas, M., Geier, M., Gruber, S., et al. (2020). Enzymes revolutionize the bioproduction of value-added compounds: From enzyme discovery to special applications. Biotechnol. Adv. 40:107520. doi: 10.1016/j.biotechadv.2020.107520
Wu, S., Snajdrova, R., Moore, J. C., Baldenius, K., and Bornscheuer, U. T. (2021). Biocatalysis: Enzymatic synthesis for industrial applications. Angew Chem. Int. Ed. Engl. 60, 88–119.
Xu, M., Zhang, Q., Xia, C., Zhong, Y., Sun, G., Guo, J., et al. (2014). Elevated nitrate enriches microbial functional genes for potential bioremediation of complexly contaminated sediments. Isme J. 8, 1932–1944. doi: 10.1038/ismej.2014.42
Yadav, M., and Chauhan, N. S. (2022). Microbiome therapeutics: Exploring the present scenario and challenges. Gastroenterol. Rep. 10:goab046.
Yang, H., Fang, R., Luo, L., Yang, W., Huang, Q., Yang, C., et al. (2023). Uncovering the mechanisms of salicylic acid-mediated abiotic stress tolerance in horticultural crops. Front. Plant Sci. 14:1226041. doi: 10.3389/fpls.2023.1226041
Yang, L., Li, B., Zheng, X.-Y., Li, J., Yang, M., Dong, X., et al. (2015). Salicylic acid biosynthesis is enhanced and contributes to increased biotrophic pathogen resistance in Arabidopsis hybrids. Nat. Commun. 6:7309. doi: 10.1038/ncomms8309
Yanni, Y. G., Rizk, R. Y., Abd El-Fattah, F. K., Squartini, A., Corich, V., Giacomini, A., et al. (2001). The beneficial plant growth-promoting association of Rhizobium leguminosarum bv. trifolii with rice roots. Aust. J. Plant Physiol. 28, 845–870.
Yee, C. S., Sohedein, M. N. A., Suan, O. P., Loen, A. W. W., Abd Rahim, M. H., Soumaya, S., et al. (2021). The production of functional γ-aminobutyric acid Malaysian soy sauce koji and moromi using the trio of Aspergillus oryzae NSK, Bacillus cereus KBC, and the newly identified Tetragenococcus halophilus KBC in liquid-state fermentation. Future Foods 4:100055.
Yoshida, S., Hiraga, K., Takehana, T., Taniguchi, I., Yamaji, H., Maeda, Y., et al. (2016). A bacterium that degrades and assimilates poly (ethylene terephthalate). Science 351, 1196–1199.
Yu, Y., Gui, Y., Li, Z., Jiang, C., Guo, J., and Niu, D. (2022). Induced systemic resistance for improving plant immunity by beneficial microbes. Plants 11:386.
Yuan, S., Wang, K. S., Meng, H., Hou, X. T., Xue, J. C., Liu, B. H., et al. (2023). The gut microbes in inflammatory bowel disease: Future novel target option for pharmacotherapy. Biomed. Pharmacother. 165:114893.
Zehra, A., Meena, M., Dubey, M. K., Aamir, M., and Upadhyay, R. S. (2017). Synergistic effects of plant defense elicitors and Trichoderma harzianum on enhanced induction of antioxidant defense system in tomato against Fusarium wilt disease. Bot. Stud. 58:44. doi: 10.1186/s40529-017-0198-2
Zhang, X., Lin, Y., Wu, Q., Wang, Y., and Chen, G. Q. (2020). Synthetic biology and genome-editing tools for improving PHA metabolic engineering. Trends Biotechnol. 38, 689–700.
Zhao, L. Y., Mei, J. X., Yu, G., Lei, L., Zhang, W. H., Liu, K., et al. (2023). Role of the gut microbiota in anticancer therapy: From molecular mechanisms to clinical applications. Signal. Transduct. Target Ther. 8:201.
Zheng, D. P., Liwinski, T., and Elinav, E. (2020). Interaction between microbiota and immunity in health and disease. Cell Res. 30, 492–506.
Zhou, C., Li, F., Xie, Y., Zhu, L., Xiao, X., Ma, Z., et al. (2017). Involvement of abscisic acid in microbe-induced saline-alkaline resistance in plants. Plant Signal. Behav. 12:e1367465. doi: 10.1080/15592324.2017.1367465
Zhou, Y., Xu, Z., Zhang, H., Liu, T., Zhou, J., Bi, Y., et al. (2024). When the microbiome meets One Health principle: Leading to the Holy Grail of biology and contributing to overall well-being and social sustainability. iMetaOmics 1:e30.
Zhu, F., Fang, Y., Wang, Z., Wang, P., Yang, K., Xiao, L., et al. (2022). Salicylic acid remodeling of the rhizosphere microbiome induces watermelon root resistance against Fusarium oxysporum f. sp. niveum infection. Front. Microbiol. 13:1015038. doi: 10.3389/fmicb.2022.1015038
Zilber-Rosenberg, I., and Rosenberg, E. (2008). Role of microorganisms in the evolution of animals and plants: The hologenome theory of evolution. FEMS Microbiol. Rev. 32, 723–735.
Zucchetta, C., Tangohau, W., Mccallion, A., Hardy, D. J., and Clavijo Mccormick, A. (2022). Exploring the chemical properties and biological activity of four New Zealand monofloral honeys to support the Māori vision and aspirations. Molecules 27:3282. doi: 10.3390/molecules27103282
Keywords: agriculture, biocatalyst processes, environmental bioremediation, food processing, human health, microbes, microbiomes
Citation: Soth S, Hampton JG, Alizadeh H, Wakelin SA and Mendoza-Mendoza A (2025) Microbiomes in action: multifaceted benefits and challenges across academic disciplines. Front. Microbiol. 16:1550749. doi: 10.3389/fmicb.2025.1550749
Received: 24 December 2024; Accepted: 24 February 2025;
Published: 18 March 2025.
Edited by:
Prem Lal Kashyap, Indian Institute of Wheat and Barley Research (ICAR), IndiaReviewed by:
Nitin Kamble, University of Cincinnati Medical Center, United StatesCopyright © 2025 Soth, Hampton, Alizadeh, Wakelin and Mendoza-Mendoza. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Sereyboth Soth, c2VyZXlib3RoLnNvdGhAbGluY29sbnVuaS5hYy5ueg==
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.