
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Microbiol. , 05 March 2025
Sec. Food Microbiology
Volume 16 - 2025 | https://doi.org/10.3389/fmicb.2025.1541084
Salmonella spp., as a major foodborne pathogen, pose significant threats to public health globally and has been an important zoonotic contamination for poultry industry that should receive increasing attentions. This study aimed to comprehensively investigate the prevalence, antimicrobial resistances, virulence factors, and plasmid types of Salmonella isolates collected from chickens, ducks, and geese across eight cities in Shandong between 2019 and 2022. Out of 300 samples, 53 Salmonella strains (17.67%) were isolated, with varied prevalence from 8.33% to 25.00% in different cities of Shandong. A total of seven serotypes were identified among the 53 Salmonella isolates, wherein the S. Enteritidis (45.28%), S. Pullorum (22.64%) and S. Typhimurium (16.98%) were identified as the most prevalent. Whole-genome sequencing analysis revealed that ST11, ST92, and ST19 were the predominant sequence types for S. Enteritidis, S. Pullorum, and S. Typhimurium, respectively. Phylogenetic analysis indicated that potential clonal spread of S. Enteritidis, S. Pullorum, and S. Typhimurium occurred across different regions, particularly the evidences supported that the S. Typhimurium isolates were dispersed in a cross-species manner. Finally, the phenotypic and genotypic profiling of antibiotic resistance among the isolates revealed that these isolates were multidrug resistant with corresponding antibiotic resistance genes (ARGs) including blaTEM, aac, aph, tet(A), and tet(B) to confer them with resistances to commonly-used veterinary drugs such as β-lactams, quinolones, macrolides. To sum, this study provides valuable insights into the current epidemiology of Salmonella in poultry industry in one of the biggest provinces in China, and shedding the light on the urgent necessity for further approaches to prevent and decontaminate such MDR Salmonella in livestock under One Health concept.
Salmonella spp., as one of the most prevalent foodborne pathogen, are reportedly to cause severe and acute intestinal diseases called salmonellosis via contaminated food chain (Chu et al., 2024). The previous studies indicated that Salmonella spp. are versatile pathogens capable of infecting a wide range of host animals, including chickens, cattle, and pigs, which are closely related to anthropogenic environments and humans. The authorities in the United States estimated that Salmonella causes approximately 1.35 million cases of infections with 420 deaths reports annually (Centers for Disease Control and Prevention | CDC). It is noted that the poultry has been a major source and an important reservoir of Salmonella spp. (Andoh et al., 2017; Feasey et al., 2012; Pui et al., 2011). Contaminated poultry products such as meat and eggs, are common cues for Salmonella accumulation and subsequent salmonellosis in humans (Lamas et al., 2016; Tedersoo et al., 2022). A directly evidence is the large foodborne infection outbreak across the U.S. in 2018 had been linked to raw chicken meat, highlighting the widespread prevalence of Salmonella in the broiler industry serves as a health threat that should not be neglected (Chu et al., 2023). Additionally, Salmonella spp. demonstrated high adaptive tolerance in different harsh conditions to transmit from animals and environments to related products through fecal contamination (Smith et al., 2023). A prior study on free-ranged poultry revealed that Salmonella was able to transmit from poultry to workers within in the same ecology via daily contacts. Therefore, longitudinal epidemiological analysis of Salmonella in poultry is crucial for in-depth understanding toward prevalence and genomic insights, which helps to better control on their potential outbreak.
Currently, antibiotics like fluoroquinolones and cephalosporin are frequently used to eradicate Salmonella infections (Chiu et al., 2002). However, the misuse and overuse of such antibiotics in animals have led to a global increase of antibiotic resistance in Salmonella spp. dampening the efficacies of such agents for clinical treatment against Salmonella (Wang et al., 2021; Xu et al., 2020). Previous studies have shown that infections caused by antibiotic-resistant bacteria (ARB) greatly challenge the current treatment paradigm and are associated with higher mortality rates in clinical settings (Crump et al., 2008). Moreover, it has been confirmed that both farm animals and wildlife are able to carry the identical Salmonella serotypes, thereby leading to high level dispersion of such ARB among the different host in the same niches (Cui et al., 2019).
Innovations in genomic analysis contribute accumulative genomic data of Salmonella in the public database, which generated from cost-effective, high-throughput whole-genome sequencing (WGS) (Wang et al., 2023). WGS is progressively becoming the standard approach for in depth understanding of pathogens. As a foundational tool in current microbiology studies, WGS data also facilitate the identification of virulence factors and antimicrobial resistance (AMR) genes (Chiou et al., 2023; Chiou et al., 2022; Collineau et al., 2019; Sohail et al., 2021). Notably, Salmonella Pathogenic Islands (SPIs) that house Salmonella virulence genes play a significant role in the pathogenesis of Salmonella in divergent hosts (Hensel, 2004; Rychlik et al., 2009). The previous investigations highlighted that SPIs vary among the serotypes and may explain differences in virulence across Salmonella serotypes (Zhao et al., 2020). Therefore, the surveillance of these ARGs and virulence factors are crucial for comprehending the evolution and pathogenicity of Salmonella.
In this study, we conducted a longitudinal study on the prevalence of Salmonella spp. in chicken, duck, and goose populations across eight cities in Shandong, China from 2019 to 2022. Subsequently, we phenotypically and genotypically identified the antimicrobial resistance of collected Salmonella strains by analyzing MLST, phylogenetic relationships, antibiotic resistance genes (ARGs), and virulence factors. This study provides vital information into the molecular epidemiology and potential pathogenicity of poultry-derived Salmonella, and contributes to development of timely control before the spread of such concerning pathogens.
A total of 300 fecal samples were collected from the chickens (n = 225), ducks (n = 67), and geese (n = 8) from eight regions in Shandong, China, including Liaocheng, Weifang, Yantai, Qingdao, Zibo, Heze, Jinan, and Dongying, during 2019 to 2022. The sampling was conducted with the informed consent of the poultry farms. The samples were labeled, placed in sterile plastic sample bags, transported to the laboratory on ice, and processed promptly.
The collected samples were inoculated onto Salmonella-Shigella (SS) selective agar, where Salmonella colonies manifest as black, pale yellow, or colorless on the SS agar medium (Maddocks et al., 2002). Presumptive Salmonella isolates underwent PCR examination targeting the invA and ompC genes, with the Salmonella Abony NCTC 6017 as a positive control (Chu et al., 2023). The serotype of the Salmonella isolates was determined using the Kauffmann-White scheme via slide agglutination with commercial O and H antisera (DK-2300 CPH.S Denmark). All Salmonella isolates were preserved as 50% glycerol stocks in a − 80°C freezer. Confirmed Salmonella isolates were serotyped according to the Kauffmann-White-Le Minor’s scheme (Guibourdenche et al., 2010).
All Salmonella isolates were subjected to the test for 15 antibiotics and the results were interpreted using the Kirby-Bauer disk diffusion method for antimicrobial susceptibility (https://www.addl.purdue.edu/newsletters/1997/spring/dds.shtml) on commercial antimicrobial disks (Hangzhou Microbial Reagent Co., Ltd. Hangzhou, Zhejiang, China) following the guidelines established by the Clinical Laboratory Standards Institute (CLSI: https://clsi.org/) (Humphries et al., 2021). Based on CLSI zone diameter interpretation criteria each strain was categorized as resistant, intermediate, or susceptible. Strains displaying resistance to a minimum of three different antibiotic classes, excluding cross-resistance mechanisms, were classified as multidrug-resistant (MDR) (Wang et al., 2023). The tested antibiotics included: amikacin, tobramycin, neomycin, doxycycline, azithromycin, lincomycin, polymyxin, amoxicillin, clindamycin, imipenem, spectinomycin, levofloxacin, enrofloxacin, cefotaxime, and florfenicol. The AST was quality controlled by using Escherichia coli ATCC 25922 as the reference strain.
All Salmonella isolates were subjected to the WGS after extraction of genomic DNA using the commercial kit (TIANGEN, China) following the manufacturer’s instructions. The purity and concentration of DNA were assessed with a Nanodrop 2000c spectrophotometer. WGS was carried out using the Illumina Hi Seq 2,500 system (Novo gene, Guangzhou, China) with a paired-end 2 × 150 bp sequencing protocol. All data were filtered to remove adaptors and low-quality reads using fastp v0.23.4 (fastp -i in.R1.fq.gz -I in.R2.fq.gz -o out.R1.fq.gz -O out.R2.fq.gz) (Chen et al., 2018b) and Fastqc v0.12.1 (fastqc *.fq.gz) (https://ifbic05c871deb3994101hwc60ck5ncovf6xf9fiac.eds.tju.edu.cn). Draft genomes were de novo assembled using SPAdes version 3.13.1. Multi-locus sequence typing (MLST) analysis was carried out using the online tool available at https://github.com/tseemann/mlst. Virulence genes, plasmid incompatibility (Inc) groups, and antibiotic resistance genes (ARGs) were analyzed using the online toolkits at https://github.com/tseemann/abricate. Phylogenetic trees for the Salmonella isolates were constructed using CSI Phylogeny version from https://www.genomicepidemiology.org/. When utilizing all the aforementioned online tools, default parameters were applied (Figure 1).
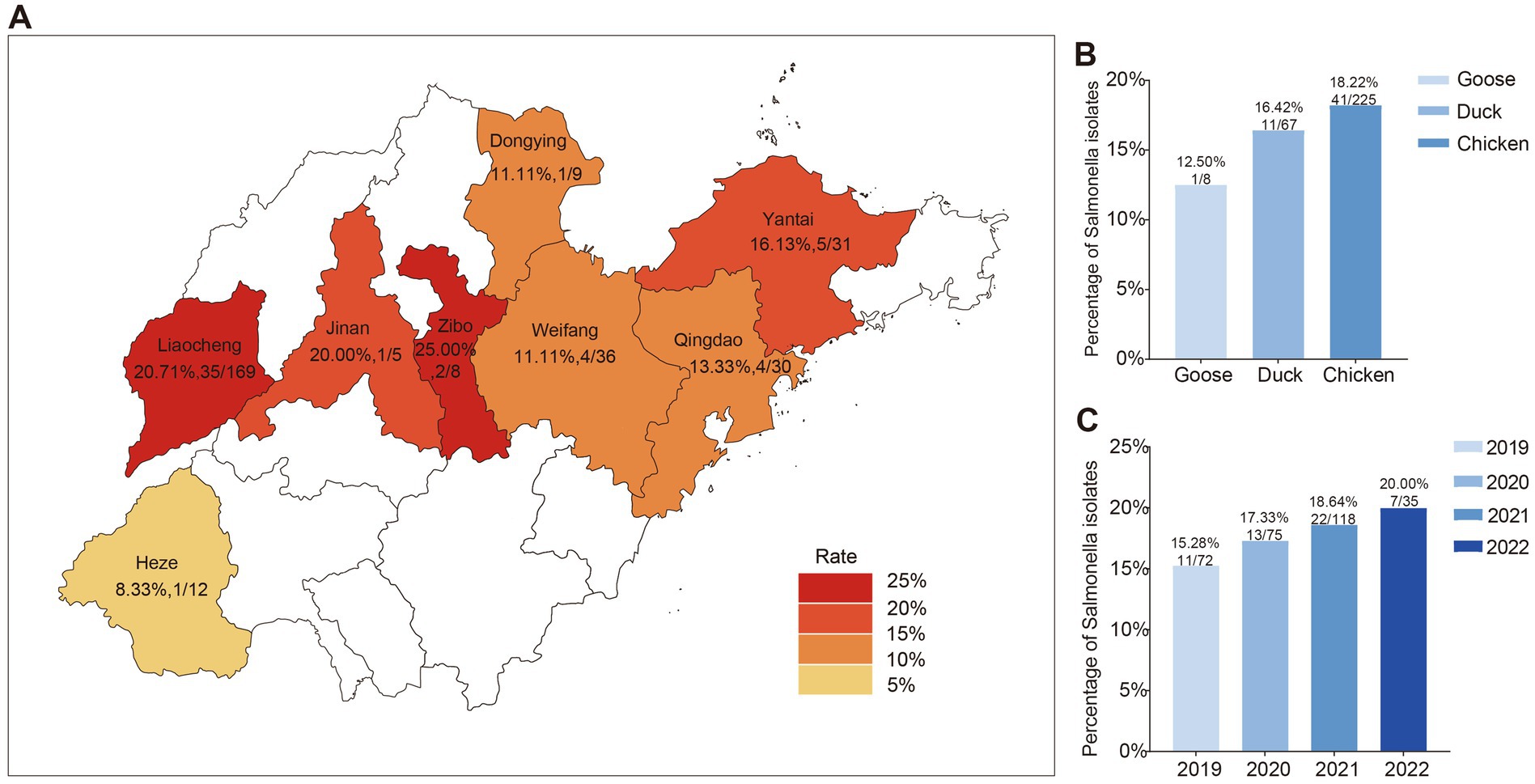
Figure 1. Geographic distribution and epidemiological overview of Salmonella in Shandong province. (A) Map of Salmonella sampling areas in Shandong province. (B) Contamination rates of Salmonella Isolates across various hosts. (C) Isolation rates of Salmonella Isolates over different years.
In this study, a total of 300 samples encompassing 225 from chickens, 67 from ducks, and 8 from geese, where 53 Salmonella isolates (17.67%, 53/300) were collected across eight cities in Shandong province between 2019 and 2022. The regional distribution of Salmonella isolates was shown as follows: Liaocheng (20.71%, 35/169); Weifang (11.11%, 4/35); Yantai (16.13%, 5/31); Qingdao (13.33%, 4/30); Zibo (25.00%, 2/8); Heze (8.33%, 1/12); Jinan (20.00%, 1/5); Dongying (11.11%, 1/9) (Figure 1A). As depicted in Figure 1B, the isolation rate in chickens was 18.22% (41/225), which was the highest among poultry species, including chickens, ducks, and geese. The temporal dynamic of the Salmonella prevalence numerically increased in first 3 years (11, 13, 22 respectively) then followed a slight decrease in 2022. However, the prevalence rate of Salmonella isolates in 2022 (20.00%, 7/35) was significantly higher compared to 2019 (15.28%, 11/72), suggesting a putative increase in salmonellosis incidence in poultry farms in Shandong during 2019–2022 (Figure 1C). A total of 7 serotypes were identified among the 53 Salmonella isolates, yet 2 isolates were not successfully determined (Figure 2A). The most prevalent serotype was S. Enteritidis (45.28%, 24/53), followed by S. Pullorum (22.64%, 12/53), S. Typhimurium (16.98%, 9/53), S. Kentucky (3.77%, 2/53), S. Saintpaul (1.89%, 1/53), and others. It was of note that the chicken samples from Liaocheng predominantly carried the isolates belonging to S. Enteritidis and S. Pullorum (Figure 2B).
In this study, all 53 Salmonella isolates were subjected to the AST with 15 commonly-used antibiotics. As depicted in Figure 2C, the all of isolates demonstrated resistance to clindamycin (100.00%, 53/53), and majority of them were found to resistant to lincomycin (79.25%, 42/53), enrofloxacin (75.47%, 40/53), and azithromycin (71.70%, 38/53). In addition, certain strains exhibited moderate resistance to azithromycin (64.15%, 34/53). Conversely, certain isolates showed low-level resistance to imipenem (100.00%, 53/53), florfenicol (90.57%, 48/53), spectinomycin (84.91%, 45/53), amikacin (83.02%, 44/53), tobramycin (81.13%, 43/53). Notably, the enrofloxacin resistance rate (100.00%) of Salmonella isolates in 2022 was significantly higher compared to the samples collected in years before. Moreover, a large portion of Salmonella isolates exhibited a multi-resistant profile, showcasing resistance to multiple antibiotics. Specifically, 88.68% (47/53) of the strains demonstrated resistance to three or more antibiotics, with an alarming strain S9 displaying resistance to 13 antibiotics (Figure 2D).

Figure 2. The serotypes and resistance phenotype of 53 Salmonella isolates. (A) The serotype identification of Salmonella. (B) Serotype distribution of Salmonella in different regions. (C) Susceptibility of Salmonella to various antimicrobial agents. (D) Numbers of resistant Salmonella isolates.
All Salmonella isolates were subjected to the WGS for harvesting the genomic insights, based on which the subsequent MLST analysis confirmed that S. indiana, S. Kentucky, S. Enteritidis, and S. Typhimurium were assigned to sequence types ST17, ST198, ST11, and ST19, respectively. Moreover, S. Pullorum isolates were categorized into two distinct STs, namely the ST92 (n = 10) and ST3717 (n = 2) (Figure 3). The core-genome sequences of S. Typhimurium isolates from chicken farms in Yantai (S14) were intriguingly found to be identical to the strains from duck farms in Liaocheng (S38), and a S. Kentucky isolate from Yantai (S9) demonstrated high similarity with an isolate from Liaocheng (S51) by showing only 20 single nucleotide polymorphisms (SNPs) in the genome. Likewise, a S. Enteritidis isolate from chickens in Liaocheng (S46) was phylogenetically similar to an isolate in Jinan (S173) with only SNP count of 30 (Figure 3). These findings collectively suggested that clonal spread at strain level might be a significant driving force for the dissemination of S. Typhimurium and S. Enteritidis across regions in Shandong province. Furthermore, one S. Enteritidis isolate (S50) exhibited more than 30,000 SNPs with many other isolates (S45, S46, S47, S5, S52, S53, S54, S83, S84, S86) from the same sampling site in Liaocheng, indicating the high genetic diversity among S. Enteritidis genogroups presented within the same geographical area.
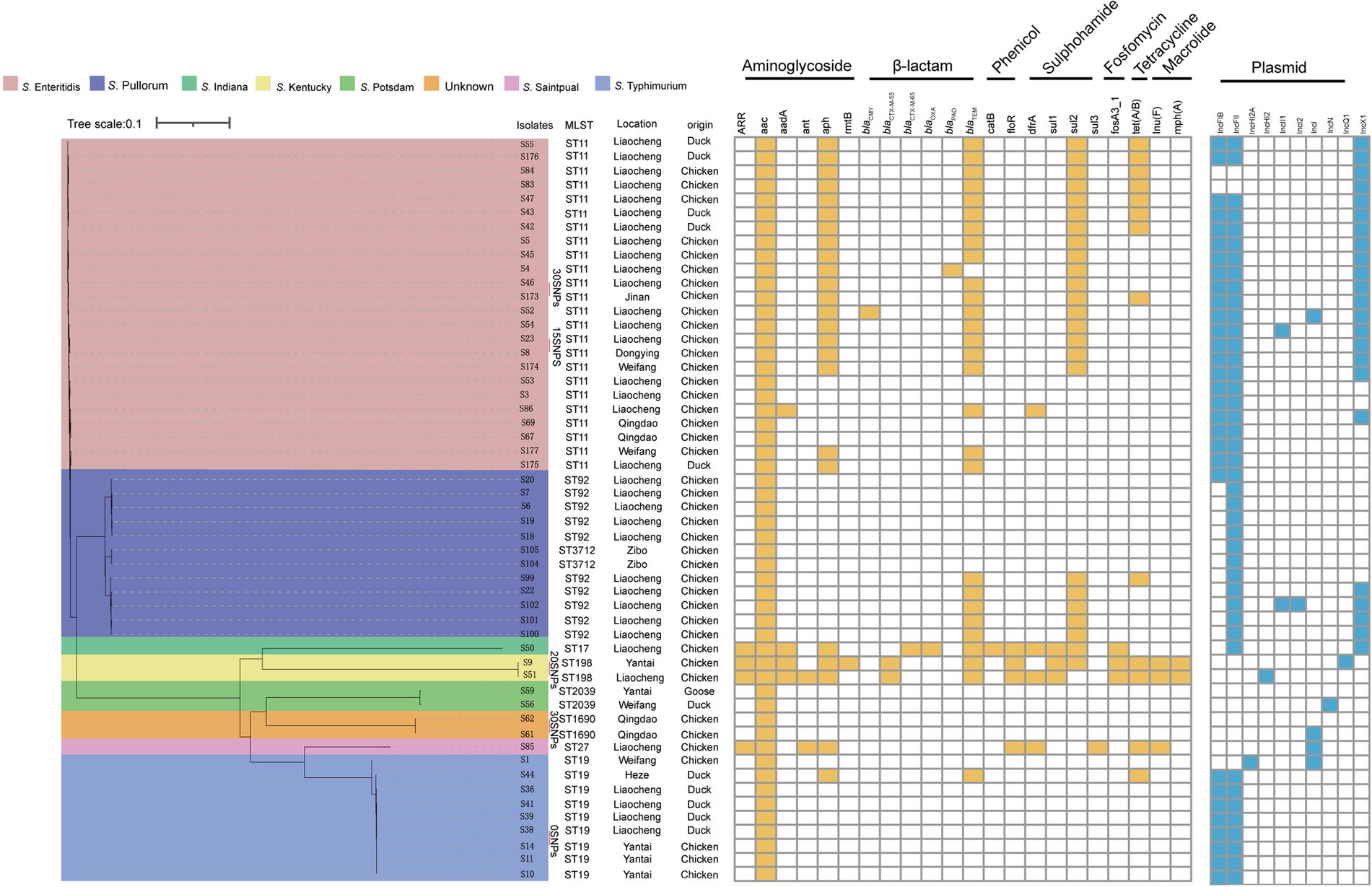
Figure 3. Phylogenetic structures, MLST and antibiotic resistance genotypes of 53 Salmonella isolates.
As the AST suggested the presence of phenotypical resistances against commonly-used antibiotic, we next sought to investigate the genotypic features of the antibiotic resistances. The genomic analysis revealed that 22 different types of ARGs were detected to confer resistances to 7 classes of antibiotics (Figure 3). As one of the most clinically concerning resistance genotype, the ESBL-producing genes were frequently detected among the Salmonella isolates, where blaCTX-M and blaTEM were identified as most dominant subtypes with prevalence rate of 54.72% (29/53) and 52.83% (28/53) respectively, followed by with other β-lactam resistance genes such as, blaCMY (1.89%, 1/53), blaPAO (1.89%, 1/53), and blaOXA (1.89%, 1/53). Additionally, a plenty of other clinically significant ARGs were detected, including aminoglycoside resistance genes ARR (7.55%), aac (100.00%), aad (7.55%), ant (3.77%), aph (45.28%), and rmtB (1.89%); phenicol resistance genes floR (7.55%) and catB (1.89%); macrolide resistance genes lnu (5.66%) and mph (3.77%); sulphonamide resistance genes sul1 (5.66%), sul2 (45.28%), and sul3 (1.89%); trimethoprim resistance genes dfrA (9.43%); fosfomycin resistance gene fosA3 (5.66%); and tetracycline resistance genes tet(A/B) (24.53%).
In the prokaryotes, the plasmids are deemed as the most imported carrier to facilitate the ARGs transmission. Thus, we have profiled the genetic characteristics of plasmids among the isolates. The results showed that a total of 9 plasmid replicon types were identified among the Salmonella isolates, including IncFIB (58.49%, 31/53), IncFII (81.13%, 43/53), IncHI2 (1.89%, 1/53), IncI1 (3.77%, 2/53), IncI2 (1.89%, 1/53), IncI (7.55%,4/53), IncN (1.89%, 1/53), IncQ (1.89%, 1/53), and IncX1 (43.40%, 23/53). Of note, no plasmid replicon was detected in S. Kentucky isolates. Among the plasmids, IncFIB and IncFII, which are the most prevalent, are primarily concentrated in S. Enteritidis and S. Typhimurium serotypes of Salmonella. This suggests that the presence of these plasmids may be associated with the serotype of the Salmonella strains. Furthermore, IncFIB was found in seven cities other than Zibo, while IncFII was detected in all of the surveyed cities. This indicates that the types of plasmids vary across different regions (Figure 4).
As to the harbored virulence factors, a total of 57 types of genes that responsible for Salmonella virulence were identified in all Salmonella isolates, most of which (31/57) were found to be conserved across all isolates. These virulence factors included handful genes belong to gene clusters encoding type III secretion systems (T3SS), SPI-2 effectors, and the Salmonella virulence plasmid (pSV). Notably, genes such as spv, lpfABCDE, and pefABCD were present in all isolates. Additionally, cdtB and pltAB genes were exclusively determined in the S. indiana isolates, in which genes like cdtA, cdtC, mig-5/−14, and rck were also found to be co-existed.
Salmonella spp. is considered one of the most significant foodborne pathogens and rank as the third leading cause of human mortality related to diarrheal illnesses. Animals serve as the primary reservoir for Salmonella, with animal-derived foods being the essential route for the transmission to humans via food chain (Ferrari et al., 2019). This study aimed to assess the prevalence and distribution of Salmonella isolates from poultry species in the Shandong province, which is one of the major bases of agricultural and livestock industry in China.
From the samples collected in eight major cities in Shandong during 2019 to 2022, a total of 53 Salmonella isolates were identified. This isolation rate was higher than that previously reported in Xinjiang (11.0%, 8/73) and Guangdong Province (3.58%, 301/8405), yet much lower than that in Tibet (30.4%, 70/230) and Henan (48.7%, 131/269) (Perry et al., 2024). However, the exact prevalence rates of Salmonella were observed heterogeneous in the different sampling sites of the eight cities, ranging from 8.3 to 25% accordingly. A previous study reported Salmonella detection rates in 12 provinces were ranging from 3.6 to 12.9% (Gong et al., 2014), indicating the Salmonella prevalence might be under certain geographic influences. The study also showed a notable presence of Salmonella contamination in poultry in Shandong Province, particularly in chickens which was with an alarming rate of 18.2% (41/225) (Liyuan et al., 2023). These results implied a wide presence of Salmonella contaminations in varied poultry breeds. Moreover, the isolation rate of Salmonella was also reported to be varied across the different environments, as some studies evidencing that the detection of Salmonella increased from poultry farms (9.1%) to slaughterhouses (9.5%) and finally enriched in the retail markets (41.8%) (Dargatz et al., 2016; Papadopoulos et al., 2016; Shang et al., 2019). To conclude, these differences were under impact of many factors such as geographical factors, environmental conditions and poultry species/breeds (Cao et al., 2017).
The prior study highlighted the increasing prevalence of Salmonella were potentially influenced by climate changing and dietary supplementations. For example, investigation by Sonora showed that Salmonella isolation rates were found to elevate in concert with the increment of temperature (Davies and Breslin, 2002). This suggests that global warming and climate change may promote Salmonella dispersion in nature especially in the anthropogenic environments. Factors such as antibiotic use, poultry density, and environmental pollution are as well believed to contribute to the global rise of Salmonella prevalence (Davies and Breslin, 2002; Sohail et al., 2021). It underscores the urgent necessity to monitor the Salmonella spp. and the preventive measures by relevant authorities.
This study identified S. Enteritidis as the predominant serotype among the collected isolates, followed by S. Pullorum and S. Typhimurium. In recent years, the contamination of S. Enteritidis has been a critical health concern in many provinces of China (Gong et al., 2014; Yang et al., 2019). This indicates that the widespread of S. Enteritidis is plausibly linked to the extensive contamination of poultry and poultry products. Cases of salmonellosis due to the consumption of contaminated poultry products, including meat, eggs and fresh products, have been reported as early in 2011 (Collineau et al., 2020; Middleton et al., 2014; Varga et al., 2012). S. Typhimurium, one of the most common serotypes in humans, have been frequently associated with severe infections on pigs, cattle, and other livestock in the European Union and the United States (Kuus et al., 2021). However, currently, this species has spread extensively worldwide, affecting not only fresh pork and beef but also poultry meat and poultry-related products. The consumption of contaminated livestock and poultry products has been proposed as a significant driver for bacterial infections in human (Perry et al., 2019; Ploton et al., 2017; Shen et al., 2022).
The tremendous use of antibiotics in animal production to promote growth and combat infection contributes to the rapid development of antibiotic resistance among pathogens (Liyuan et al., 2023). Our study revealed high levels of resistance in Salmonella isolates to clindamycin, lincomycin, ampicillin, enrofloxacin, azithromycin, and doxycycline, possibly owing to large-scale antibiotic usage for poultry farming. The S9 Salmonella strain exhibits resistance to 15 different antibiotics, making it a typical example of multidrug-resistant (MDR). Salmonella strains in the late 1980s challenged treatment choices amoxicillin. The MDR strains of Salmonella exhibit various resistance pattern, including ACSSuT (ampicillin, chloramphenicol, streptomycin, sulfamethoxazole, tetracycline) and AKSSuT (ampicillin, kanamycin, streptomycin, sulfamethoxazole, tetracycline) resistance pattern has been commonly reported. And this often includes first-line antibiotics (β-lactamases, macrolides), and the rise of MDR strains has compelled clinicians to resort to second-line agents (quinolones, tetracyclines and third-generation cephalosporins) (Rahman et al., 2023; Ramatla et al., 2021; Rowe et al., 1997; Samia et al., 2021). It is of particular concern that β-lactams and fluoroquinolones, which are typically the primary options for treating salmonellosis, were found to be lack of potency to the strains collected in our study due to carriage of corresponding ARGs. As observed with the emergence of Salmonella Infantis (ESI) clone in 2014, which was first detected in retail meats in Tennessee, by 2019, it had spread across the United States. The pESI plasmid underwent significant recombination, carrying not only the extended-spectrum β-lactamase gene blaCTX-M-65, but also the gyrA mutation, which confers resistance to fluoroquinolones, further limiting treatment options (Perry et al., 2024). Additionally, Salmonella strains are often transmitted through contaminated food, wastewater, animals, and other sources (Cosby et al., 2019; Gebreyes et al., 2008). This suggests that MDR strains have the potential for spread, which undoubtedly poses a significant threat to public health, exacerbates the burden on clinical medicine, and imposes a tremendous economic strain on society. However, the resistances to cephalosporins and fluoroquinolones have been observed to increase in many countries, possibly due to incorrect antibiotic usage. Additionally, Shea (2004) highlighted that prolonged antibiotic therapy contributed to development of antimicrobial resistance. Ciprofloxacin and enrofloxacin-resistant Salmonella isolates have been widely found in aquaculture, poultry, poultry meat, and pig manure (Ubeyratne et al., 2023; Wang et al., 2022), which threaten both animal wellbeing and human health under One Health concept.
MLST analysis revealed that all S. Enteritidis, S. Typhimurium, S. Kentucky, and S. indiana isolates belonging to ST11, ST19, ST198, and ST17, respectively. ST11 emerged as a predominant serotype, particularly prevalent chicken samples in our cases. Notably, 95% of S. Enteritidis isolates in England and Wales from April 2014 to March 2015 were classified as ST11 (Aung et al., 2022). The European Food Safety Authority (EFSA) indicated that the outbreak of infection of Salmonella ST11 in 2022 in Europe was associated with eggs and egg products, underscoring the urgent need for controlling the Salmonella contamination in poultry and its supply chain. The ST19 is the predominant genotype of Salmonella isolate with broad geographic distribution (Achtman et al., 2012; Gómez-Baltazar et al., 2023). The ST19 Salmonella is commonly found in patients with diarrhea and is always detected in various origins including animals, food, and humans (Sun et al., 2014; Wong et al., 2013). S. Kentucky, an emerging pathogen, has been increasingly responsible for human non-typhoidal Salmonella infections since 2005 (Mahindroo et al., 2019). While S. Kentucky isolates are frequently found in poultry and livestock yet rarely associated with human infection. Furthermore, the ST198 has been observed to spread independently among poultry and other livestock, occasionally leading to human infections (Samper-Cativiela et al., 2022). In this study, S. Pullorum was found with ST92 and ST3717.
Phylogenetic analyses are crucial for understanding microbial evolution and infectious disease transmission. Bacterial phylogenies are often inferred from SNP alignments, with SNPs as the method in the genetic relationship between isolates, allowing for comparison of genetic information at the genome level for each isolate (Dallman et al., 2018; Mona et al., 2025). In this study, the core-genome sequences of S. Kentucky isolate from Yantai (S9) demonstrated high similarity with an isolate from Liaocheng (S51) by showing only 20 single nucleotide polymorphisms (SNPs) in the genome. The genetic distance between these isolates is small, suggesting that they may originate from the same clonal group, which is consistent with previous Salmonella studies. Specifically, isolates with a small SNP distance are highly likely to come from the same clonal lineage (Wang et al., 2018). Additionally, SNP analysis revealed significant genetic diversity among these isolates. For example, isolate S50 showed over 30,000 SNP differences compared to several other isolates from the same sampling site (Liaocheng), including S45, S46, S47, S5, S52, S53, S54, S83, S84, and S86. This finding suggests that the genetic variation may be related to the source and time of isolation, with the primary factor being the origin of the Salmonella isolates (Levent et al., 2021). With this estimate of genetic similarity, bioinformaticians can better identify isolates that likely have a recent common source, and provides essential data for public health management.
The phenotypic resistances are closely associated with ARGs. Genotypic analysis revealed that the most prevalent ARG gene was aac, followed by aph, sul2, and tet (A/B). These genes conferred the resistances to aminoglycosides, sulfonamides, and tetracyclines (Ubeyratne et al., 2023). Further analyses also uncovered the presence of resistance genes against β-lactamase, including blaTEM, blaOXA, blaCTX-M-65, blaCTX-M-55, and blaCMY. Among them, blaTEM was the most prevalent (Olesen et al., 2004). These β-lactamases are active to hydrolyze the cephalosporins, dampening the efficacies of such antibiotics. Notably, genes like aadA rendered the isolates resistant to streptomycin, gentamicin, and tobramycin (Chu et al., 2023; Samia et al., 2021). Previous studies have also identified these genes and their impact on antimicrobial resistance in Salmonella spp.
Plasmids have been reported as a key player in the dissemination of ARGs, virulence genes, and other traits that provide a fitness advantage to host bacteria. Analysis on plasmid in the current study revealed the presence of 9 different plasmid replicons existing in the Salmonella isolates, with the IncFII as the dominant replicons. These plasmids were found in various host bacteria, indicating their ability to spread the harbored genes with their transmissions. IncFII and IncFIB are the most common variants within the IncF plasmid family. They are important vectors for the carriage and transmission of antibiotic resistance and virulence genes (Kline, 1985; Ravi et al., 2017; Villa et al., 2010). They are important vectors for the transmission of antibiotic resistance and virulence genes. In particular, IncFII plasmids are also the main vector of mcr-1 in E. coli isolates (Sugawara et al., 2019; Xavier et al., 2016). The IncF plasmids in Enterobacteriaceae are of particular interest since they contribute to the carriage and spread of ARGs and virulence genes (Carattoli, 2011). IncF plasmids have been isolated that contain ARGs genes. IncF plasmids in Salmonella isolated in China carried fluoroquinolone resistance genes (Chen et al., 2018a). In Salmonella isolated in the United States, IncF plasmids have been associated with strAB, tetA, tetC, tetD, aphA (aminoglycosides), and sul2 (Han et al., 2012; McMillan et al., 2019; McMillan et al., 2020), and they were most frequently described carrying ESBL genes, encoding carbapenemases genes, and plasmid-mediated quinolone resistance (PMQR) genes (Rozwandowicz et al., 2018). In addition, they also carry specific virulence traits such as cytotoxins and adhesion factors as accessory genes (Johnson and Nolan, 2009). In this study, IncF family plasmids were isolated from multiple serotypes of Salmonella, primarily S. Enteritidis and S. Typhimurium, across different regions. This suggests that IncF family plasmids may be associated with specific regions and serotypes. These findings are in accordance with previous studies, where the IncF family was detected in multiple serotypes of Salmonella across different regions and farms (NCF3, −5, and − 6) (Pornsukarom and Thakur, 2017; Wang et al., 2013). The association between plasmid types and specific serotypes or geographical regions may lead to the spread of antibiotic resistance and treatment limitations, exacerbating the antibiotic resistance crisis and posing a significant threat to global public health. Furthermore, IncHI2, as part of the backbone element of the pSE380T plasmid, in conjunction with the IncFIA plasmid, produces a rare fusion product encoding both virulence and resistance genes with high transmissibility (Villa et al., 2010; Wong et al., 2017). Therefore, monitoring the prevalence of ARGs together with plasmids profiling is crucial for the prevention and control of antibiotic-resistant Salmonella.
Analysis of WGS data revealed that the Salmonella isolates were found with multiple virulence factors including invABCEFGHIJ, prgHIJK, hilACD, sicAP, sipABCD, sopABD/D2E2, spaOPQRS, sptP, and mgtBC. Within these, virulence genes like invA, mgtC, and sopB are clustered in the Salmonella Pathogenicity Island (SPI) on the chromosomes, help Salmonella to establish systematic infection by mediating cell invasion, intracellular survival, and inflammatory responses (Marcus et al., 2000). For instance, hilACD are able to activate transcription of key virulence genes on SPI-1, facilitating the release of essential effector proteins for invading the host intestine (Bosire et al., 2020). Furthermore, the spv genes located on the Salmonella virulence plasmid (IncF), which is involved in survival and systemic infection in host cells, were detected in S. Enteritidis and S. Typhimurium. The spvABCD genes are located upstream of the genes pefA (plasmid-encoded fimbriae) and rck (resistance to complement killing) in a virulence plasmid by the upstream spvR gene (Monte et al., 2020; Silva et al., 2017). However, spvR, which is needed for the regulation of the spv locus (Guiney and Fierer, 2011), was absent in our strains. Previous studies have indicated that the spv operon, which plays a role Salmonella survival within the host cell during systemic infection, affects the formation ofautophagosomes, as well as highlight its association in killing of macrophages and neutrophils (Wu et al., 2016), being crucially required for virulence in vivo (Passaris et al., 2018), including aggravated damage in zebrafish infection model (Cao et al., 2023; Hsu et al., 2019; Passaris et al., 2018). Moreover, other virulence factors like pefABCD, lpfABCDE, cdtB, pltAB and rck were detected. The lpfABCDE and pefABCD gene clusters on pSV are reported to regulate the formation of long polar fimbriae fimbriae. The cdtB is a common virulence gene found in certain Gram-negative bacteria, and the expression of cdtB promotes toxin production in non-typhoidal Salmonella (Wójcicki et al., 2021). In addition, cdtB was characterized in S. typhi as one component of the cytolethal distending toxin (CDT) (Haghjoo and Galán, 2004). Several Gram-negative bacterial pathogens, including S. typhi, produce CDT, which arrests growth, induces apoptosis of infected host cells, and enhances the persistence of pathogenic bacteria in the host. The cdtB genes in S. typhi is located on a pathogenicity island upstream of the pltA and pltB genes, which encode pertussis-like toxins A and B (PltA and PltB). These toxins form a complex with cdtB, with the pltB gene having no apparent effect on cellular distension. However, when cdtB is combined with pltA, it induces noticeable distension in both the cytoplasm and the nucleus (Mezal et al., 2014). In S. Javiana, cdtB may require PltA for the typical signs of cytolethal distension, such as cytoplasmic and nuclear swelling. The cdtB gene, in conjunction with pltA (pertussis-like toxin A) and pltB (pertussis-like toxin B), is necessary to induce intoxication signs in eukaryotic cells, such as cellular distension and cell cycle arrest (Mezal et al., 2014; Spanò et al., 2008). In the other hand, cdtB, with the assistance of cdtAC, is translocated into the nucleus of target cells, where it induces double-strand breaks in DNA, resulting in cell apoptosis (Ohara, 2004; Thakur et al., 2022).
This study has identified a total of 53 Salmonella isolates from 300 samples collected during 2019 to 2022 in Shandong, China. Among the 53 Salmonella isolates, seven serotypes were identified wherein the S. Enteritidis was found to be most prevalent, followed by S. Pullorum and S. Typhimurium. WGS analysis revealed that ST11, ST92, and ST19 were the dominant sequence types in Salmonella in this study. The findings underscore the importance of epidemiological surveillance in the major agricultural area like Shandong Province. Furthermore, the majority of Salmonella isolates exhibited features of multi-drug resistance, highlighting the need for precision controlling strategies against such resistant Salmonella.
The majority of Salmonella isolates in this study exhibited multidrug resistance, highlighting the need for precise control strategies to address these resistant strains. This underscores the critical role of the One Health concept, which recognizes the interconnectedness of human, animal, and environmental health. The emergence and spread of multidrug-resistant Salmonella are influenced by various factors, including human, animal, and environmental elements. Therefore, an integrated approach should not only monitor human health but also coordinate animal health management and environmental surveillance. Such a strategy is essential for curbing the spread of resistant bacteria and mitigating the public health risks posed by multidrug-resistant Salmonella.
However, several limitations exist in this study. Firstly, it was region-specific, focusing solely on Shandong Province, which limits the generalizability of the findings. Additionally, the sample size was relatively small, and the study did not include Salmonella strains from other animals, such as cattle, sheep, and pigs, which may provide insights into cross-species transmission. Moreover, the impact of the rearing environment on Salmonella was not explored. To address these gaps, future research should expand the study to broader regions, investigate cross-species transmission, and examine the influence of environmental factors and farming practices. Incorporating the One Health approach into such studies will be crucial for developing more effective strategies to explore and combat antimicrobial resistance.
The datasets presented in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found at: https://submit.ncbi.nlm.nih.gov/subs/, Bioproject ID PRJNA1182193.
The manuscript presents research on animals that do not require ethical approval for their study.
LC: Data curation, Formal analysis, Investigation, Writing – original draft. YS: Data curation, Investigation, Software, Visualization, Writing – original draft. MW: Data curation, Funding acquisition, Supervision, Visualization, Writing – review & editing. YL: Funding acquisition, Project administration, Resources, Supervision, Writing – review & editing. ZS: Resources, Supervision, Validation, Writing – review & editing.
The author(s) declare that financial support was received for the research, authorship, and/or publication of this article. This work was in part supported by the National Natural Science Foundation of China (32372957), Foundation for Innovative Research Groups of the National Natural Science Foundation of China (32202859).
We thank all the contributors of this research topic and we wish you all a good reading.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The authors declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Achtman, M., Wain, J., Weill, F. X., Nair, S., Zhou, Z., Sangal, V., et al. (2012). Multilocus sequence typing as a replacement for serotyping in Salmonella enterica. PLoS Pathog. 8:e1002776. doi: 10.1371/journal.ppat.1002776
Andoh, L. A., Ahmed, S., Olsen, J. E., Obiri-Danso, K., Newman, M. J., Opintan, J. A., et al. (2017). Prevalence and characterization of Salmonella among humans in Ghana. Trop. Med. Health. 45:3. doi: 10.1186/s41182-017-0043-z
Aung, K. T., Khor, W. C., Ong, K. H., Tan, W. L., Wong, Z. N., Oh, J. Q., et al. (2022). Characterisation of Salmonella enteritidis ST11 and ST1925 associated with human intestinal and extra-intestinal infections in Singapore. Int. J. Environ. Res. Public Health 19:5671. doi: 10.3390/ijerph19095671
Bosire, E. M., Eade, C. R., Schiltz, C. J., Reid, A. J., Troutman, J., Chappie, J. S., et al. (2020). Diffusible signal factors act through AraC-type transcriptional regulators as chemical cues to repress virulence of enteric pathogens. Infect. Immun. 88, e00226–e00220. doi: 10.1128/iai.00226-20
Cao, T. T., Deng, G. H., Fang, L. X., Yang, R. S., Sun, J., Liu, Y. H., et al. (2017). Characterization of quinolone resistance in Salmonella enterica from farm animals in China. J. Food Prot. 80, 1742–1748. doi: 10.4315/0362-028x.Jfp-17-068
Cao, G., Zhao, S., Kuang, D., Hsu, C. H., Yin, L., Luo, Y., et al. (2023). Geography shapes the genomics and antimicrobial resistance of Salmonella enterica Serovar Enteritidis isolated from humans. Sci. Rep. 13:1331. doi: 10.1038/s41598-022-24150-4
Carattoli, A. (2011). Plasmids in gram negatives: molecular typing of resistance plasmids. Int. J. Med. Microbiol. 301, 654–658. doi: 10.1016/j.ijmm.2011.09.003
Chen, K., Dong, N., Zhao, S., Liu, L., Li, R., Xie, M., et al. (2018a). Identification and characterization of conjugative plasmids that encode ciprofloxacin resistance in Salmonella. Antimicrob. Agents Chemother. 62:e00575-18. doi: 10.1128/aac.00575-18
Chen, S., Zhou, Y., Chen, Y., and Gu, J. (2018b). Fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890. doi: 10.1093/bioinformatics/bty560
Chiou, C. S., Hong, Y. P., Wang, Y. W., Chen, B. H., Teng, R. H., Song, H. Y., et al. (2023). Antimicrobial resistance and mechanisms of azithromycin resistance in Nontyphoidal Salmonella isolates in Taiwan, 2017 to 2018. Microbiol Spectr. 11:e0336422. doi: 10.1128/spectrum.03364-22
Chiou, C. S., Liao, Y. S., Chen, B. H., Lu, M. C., Hong, Y. P., Wang, Y. W., et al. (2022). Demographic features of invasive meningococcal disease in Taiwan, 1993 to 2020, and genetic characteristics of Neisseria meningitidis isolates, 2003 to 2020. Microbiol Spectr. 10:e0088222. doi: 10.1128/spectrum.00882-22
Chiu, C. H., Wu, T. L., Su, L. H., Chu, C., Chia, J. H., Kuo, A. J., et al. (2002). The emergence in Taiwan of fluoroquinolone resistance in Salmonella enterica serotype choleraesuis. N. Engl. J. Med. 346, 413–419. doi: 10.1056/NEJMoa012261
Chu, Y., Ruan, Y.-X., Liu, J.-Q., Zhang, Y., Wang, M.-G., Liao, X.-P., et al. (2024). Population genomics of emerging multidrug-resistant Salmonella derby from pork and human in Guangdong, China. LWT Food Sci. Technol. 9:e0116423. doi: 10.1016/j.lwt.2024.116535
Chu, Y., Wang, D., Hao, W., Sun, R., Sun, J., Liu, Y., et al. (2023). Prevalence, antibiotic resistance, virulence genes and molecular characteristics of Salmonella isolated from ducks and wild geese in China. Food Microbiol. 118:104423. doi: 10.1016/j.fm.2023.104423
Collineau, L., Boerlin, P., Carson, C. A., Chapman, B., Fazil, A., Hetman, B., et al. (2019). Integrating whole-genome sequencing data into quantitative risk assessment of foodborne antimicrobial resistance: a review of opportunities and challenges. Front. Microbiol. 10:1107. doi: 10.3389/fmicb.2019.01107
Collineau, L., Phillips, C., Chapman, B., Agunos, A., Carson, C., Fazil, A., et al. (2020). A within-flock model of Salmonella Heidelberg transmission in broiler chickens. Prev. Vet. Med. 174:104823. doi: 10.1016/j.prevetmed.2019.104823
Cosby, D. E., Cox, N. A., Harrison, M. A., Wilson, J. L., Buhr, R. J., and Fedorka-Cray, P. J. (2019). Salmonella and antimicrobial resistance in broilers: A review. J. Appl. Poult. Res. 24, 408–426. doi: 10.3382/japr/pfv038
Crump, J. A., Kretsinger, K., Gay, K., Hoekstra, R. M., Vugia, D. J., Hurd, S., et al. (2008). Clinical response and outcome of infection with Salmonella enterica serotype Typhi with decreased susceptibility to fluoroquinolones: a United States foodnet multicenter retrospective cohort study. Antimicrob. Agents Chemother. 52, 1278–1284. doi: 10.1128/aac.01509-07
Cui, M., Zhang, P., Li, J., Sun, C., Song, L., Zhang, C., et al. (2019). Prevalence and characterization of fluoroquinolone resistant Salmonella isolated from an integrated broiler chicken supply chain. Front. Microbiol. 10:1865. doi: 10.3389/fmicb.2019.01865
Dallman, T., Ashton, P., Schafer, U., Jironkin, A., Painset, A., Shaaban, S., et al. (2018). SnapperDB: a database solution for routine sequencing analysis of bacterial isolates. Bioinformatics 34, 3028–3029. doi: 10.1093/bioinformatics/bty212
Dargatz, D. A., Kopral, C. A., Erdman, M. M., and Fedorka-Cray, P. J. (2016). Prevalence and antimicrobial resistance of Salmonella isolated from cattle feces in United States feedlots in 2011. Foodborne Pathog. Dis. 13, 483–489. doi: 10.1089/fpd.2016.2128
Davies, R., and Breslin, M. (2002). Environmental contamination and detection of Salmonella enterica serovar enteritidis in laying flocks. Vet. Rec. 149, 699–704. doi: 10.1136/vr.149.23.699
Feasey, N. A., Dougan, G., Kingsley, R. A., Heyderman, R. S., and Gordon, M. A. (2012). Invasive non-typhoidal salmonella disease: an emerging and neglected tropical disease in Africa. Lancet 379, 2489–2499. doi: 10.1016/s0140-6736(11)61752-2
Ferrari, R. G., Rosario, D. K. A., Cunha-Neto, A., Mano, S. B., Figueiredo, E. E. S., and Conte-Junior, C. A. (2019). Worldwide epidemiology of Salmonella Serovars in animal-based foods: a Meta-analysis. Appl. Environ. Microbiol. 85, e00591–e00519. doi: 10.1128/aem.00591-19
Gebreyes, W. A., Thakur, S., Dorr, P., Tadesse, D. A., Post, K., and Wolf, L. (2008). Occurrence of spvA virulence gene and clinical significance for multidrug-resistant Salmonella strains. J. Clin. Microbiol. 47, 777–780. doi: 10.1128/jcm.01660-08
Gómez-Baltazar, A., Vázquez-Marrufo, G., Astiazaran-Garcia, H., Ochoa-Zarzosa, A., Canett-Romero, R., García-Galaz, A., et al. (2023). Comparative virulence of the worldwide ST19 and emergent ST213 genotypes of Salmonella enterica serotype Typhimurium strains isolated from food. Microbes Infect. 25:105019. doi: 10.1016/j.micinf.2022.105019
Gong, J., Zhang, J., Xu, M., Zhu, C., Yu, Y., Liu, X., et al. (2014). Prevalence and fimbrial genotype distribution of poultry Salmonella isolates in China (2006 to 2012). Appl. Environ. Microbiol. 80, 687–693. doi: 10.1128/aem.03223-13
Guibourdenche, M., Roggentin, P., Mikoleit, M., Fields, P. I., Bockemühl, J., Grimont, P. A., et al. (2010). Supplement 2003-2007 (No. 47) to the white-Kauffmann-Le minor scheme. Res. Microbiol. 161, 26–29. doi: 10.1016/j.resmic.2009.10.002
Guiney, D. G., and Fierer, J. (2011). The role of the spv genes in Salmonella pathogenesis. Front. Microbiol. 2:129. doi: 10.3389/fmicb.2011.00129
Haghjoo, E., and Galán, J. E. (2004). Salmonella typhi encodes a functional cytolethal distending toxin that is delivered into host cells by a bacterial-internalization pathway. Proc. Natl. Acad. Sci. USA 101, 4614–4619. doi: 10.1073/pnas.0400932101
Han, J., Lynne, A. M., David, D. E., Tang, H., Xu, J., Nayak, R., et al. (2012). DNA sequence analysis of plasmids from multidrug resistant Salmonella enterica serotype Heidelberg isolates. PLoS One 7:e51160. doi: 10.1371/journal.pone.0051160
Hensel, M. (2004). Evolution of pathogenicity islands of Salmonella enterica. Int. J. Med. Microbiol. 294, 95–102. doi: 10.1016/j.ijmm.2004.06.025
Hsu, C. H., Li, C., Hoffmann, M., McDermott, P., Abbott, J., Ayers, S., et al. (2019). Comparative genomic analysis of virulence, antimicrobial resistance, and plasmid profiles of Salmonella Dublin isolated from sick cattle, retail beef, and humans in the United States. Microb. Drug Resist. 25, 1238–1249. doi: 10.1089/mdr.2019.0045
Humphries, R., Bobenchik, A. M., Hindler, J. A., and Schuetz, A. N. (2021). Overview of changes to the clinical and laboratory standards institute performance standards for antimicrobial susceptibility testing, M100. J. Clin. Microbiol. 59:e0021321. doi: 10.1128/jcm.00213-21
Johnson, T. J., and Nolan, L. K. (2009). Pathogenomics of the virulence plasmids of Escherichia coli. Microbiol. Mol. Biol. Rev. 73, 750–774. doi: 10.1128/mmbr.00015-09
Kline, B. C. (1985). A review of mini-F plasmid maintenance. Plasmid 14, 1–16. doi: 10.1016/0147-619x(85)90027-7
Kuus, K., Kramarenko, T., Sõgel, J., Mäesaar, M., Fredriksson-Ahomaa, M., and Roasto, M. (2021). Prevalence and serotype diversity of Salmonella enterica in the Estonian meat production chain in 2016-2020. Pathogens 10:1622. doi: 10.3390/pathogens10121622
Lamas, A., Fernandez-No, I. C., Miranda, J. M., Vázquez, B., Cepeda, A., and Franco, C. M. (2016). Prevalence, molecular characterization and antimicrobial resistance of Salmonella serovars isolated from northwestern Spanish broiler flocks (2011-2015). Poult. Sci. 95, 2097–2105. doi: 10.3382/ps/pew150
Levent, G., Schlochtermeier, A., Ives, S. E., Norman, K. N., Lawhon, S. D., Loneragan, G. H., et al. (2021). High-resolution genomic comparisons within Salmonella enterica serotypes derived from beef feedlot cattle: parsing the roles of cattle source, pen, animal, sample type and production period. Appl. Environ. Microbiol. 87:e0048521. doi: 10.1128/aem.00485-21
Liyuan, Z., Gang, L., Wenli, T., Xiangbin, S., Xiaoyu, Z., Chu, W., et al. (2023). Antimicrobial resistance and genomic characteristics of Salmonella from broilers in Shandong Province. Front. Vet. Sci. 10:1292401. doi: 10.3389/fvets.2023.1292401
Maddocks, S., Olma, T., and Chen, S. (2002). Comparison of CHROMagar Salmonella medium and xylose-lysine-desoxycholate and Salmonella-Shigella agars for isolation of Salmonella strains from stool samples. J. Clin. Microbiol. 40, 2999–3003. doi: 10.1128/jcm.40.8.2999-3003.2002
Mahindroo, J., Thanh, D. P., Nguyen, T. N. T., Mohan, B., Thakur, S., Baker, S., et al. (2019). Endemic fluoroquinolone-resistant Salmonellaenterica serovar Kentucky ST198 in northern India. Microb Genom. 5:e000275. doi: 10.1099/mgen.0.000275
Marcus, S. L., Brumell, J. H., Pfeifer, C. G., and Finlay, B. B. (2000). Salmonella pathogenicity islands: big virulence in small packages. Microbes Infect. 2, 145–156. doi: 10.1016/s1286-4579(00)00273-2
McMillan, E. A., Gupta, S. K., Williams, L. E., Jové, T., Hiott, L. M., Woodley, T. A., et al. (2019). Antimicrobial resistance genes, cassettes, and plasmids present in Salmonella enterica associated with United States food animals. Front. Microbiol. 10:832. doi: 10.3389/fmicb.2019.00832
McMillan, E. A., Jackson, C. R., and Frye, J. G. (2020). Transferable plasmids of Salmonella enterica associated with antibiotic resistance genes. Front. Microbiol. 11:562181. doi: 10.3389/fmicb.2020.562181
Mezal, E. H., Bae, D., and Khan, A. A. (2014). Detection and functionality of the CdtB, PltA, and PltB from Salmonella enterica serovar Javiana. Pathogens Disease. 72, 95–103. doi: 10.1111/2049-632x.12191
Middleton, D., Savage, R., Tighe, M. K., Vrbova, L., Walton, R., Whitfield, Y., et al. (2014). Risk factors for sporadic domestically acquired Salmonella serovar Enteritidis infections: a case-control study in Ontario, Canada, 2011. Epidemiol. Infect. 142, 1411–1421. doi: 10.1017/s0950268813001945
Mona, L. T., Leo, A. F., George, T., Torsten, S., Danielle, J. I., Timothy, P. S., et al. (2025). Exploring SNP filtering strategies: the influence of strict vs soft core. Microbial Genom. 11:001346. doi: 10.1099/mgen.0.001346
Monte, D. F. M., Sellera, F. P., Lopes, R., Keelara, S., Landgraf, M., Greene, S., et al. (2020). Class 1 integron-borne cassettes harboring blaCARB-2 gene in multidrug-resistant and virulent Salmonella typhimurium ST19 strains recovered from clinical human stool samples, United States. PLoS One 15:e0240978. doi: 10.1371/journal.pone.0240978
Ohara, M. (2004). Cytolethal distending toxin: a bacterial bullet targeted to nucleus. J. Biochem. 136, 409–413. doi: 10.1093/jb/mvh154
Olesen, I., Hasman, H., and Aarestrup, F. M. (2004). Prevalence of beta-lactamases among ampicillin-resistant Escherichia coli and Salmonella isolated from food animals in Denmark. Microb. Drug Resist. 10, 334–340. doi: 10.1089/mdr.2004.10.334
Papadopoulos, T., Petridou, E., Zdragas, A., Mandilara, G., Nair, S., Peters, T., et al. (2016). Comparative study of all Salmonella enterica serovar Enteritidis strains isolated from food and food animals in Greece from 2008 to 2010 with clinical isolates. Eur. J. Clin. Microbiol. Infect. Dis. 35, 741–746. doi: 10.1007/s10096-016-2591-2
Passaris, I., Cambré, A., Govers, S. K., and Aertsen, A. (2018). Bimodal expression of the Salmonella typhimurium spv operon. Genetics 210, 621–635. doi: 10.1534/genetics.118.300822
Perry, J., Arnold, K., Satuchne, C., Koren, O., Kenigswald, G., and Elnekave, E. (2024). Accumulation of resistance genes in Salmonella typhimurium transmitted between poultry and dairy farms increases the risk to public health. Appl. Environ. Microbiol. 90:e0229723. doi: 10.1128/aem.02297-23
Perry, I. D., Nguyen, T., Sherina, V., Love, T. M. T., Miller, R. K., Krishnan, L., et al. (2019). Analysis of the capacity of Salmonella enterica Typhimurium to infect the human placenta. Placenta 83, 43–52. doi: 10.1016/j.placenta.2019.06.386
Ploton, M. C., Gaschignard, J., Lemaitre, C., Cadennes, A., Germanaud, D., Poncelet, G., et al. (2017). Salmonella typhimurium bacteraemia complicated by meningitis and brain abscess in a 3-month-old boy. J. Paediatr. Child Health 53, 204–205. doi: 10.1111/jpc.13433
Pornsukarom, S., and Thakur, S. (2017). Horizontal dissemination of antimicrobial resistance determinants in multiple Salmonella serotypes following isolation from the commercial swine operation environment after manure application. Appl. Environ. Microbiol. 83:e01503-17. doi: 10.1128/aem.01503-17
Pui, C. F., Wong, W. C., Chai, L. C., Lee, H. Y., Noorlis, A., Zainazor, T. C., et al. (2011). Multiplex PCR for the concurrent detection and differentiation of Salmonella spp., Salmonella typhi and Salmonella typhimurium. Trop. Med. Health. 39, 9–15. doi: 10.2149/tmh.2010-20
Rahman, M. K., Hassan, M. M., Islam, S., Rostal, M. K., Uddin, M. H., Hagan, E., et al. (2023). Characterization and epidemiology of antimicrobial resistance patterns of Salmonella spp. and Staphylococcus spp. in free-ranging rhesus macaque (Macaca mulatta) at high-risk interfaces with people and livestock in Bangladesh. Front. Vet Sci. 10:1103922. doi: 10.3389/fvets.2023.1103922
Ramatla, T., Tawana, M., Onyiche, T. E., Lekota, K. E., and Thekisoe, O. (2021). Prevalence of antibiotic resistance in Salmonella serotypes concurrently isolated from the environment, animals, and humans in South Africa: a systematic review and Meta-analysis. Antibiotics 10:1435. doi: 10.3390/antibiotics10121435
Ravi, A., Valdés-Varela, L., Gueimonde, M., and Rudi, K. (2017). Transmission and persistence of IncF conjugative plasmids in the gut microbiota of full-term infants. FEMS Microbiol. Ecol. 94:158. doi: 10.1093/femsec/fix158
Rowe, B., Ward, L. R., and Threlfall, E. J. (1997). Multidrug-resistant Salmonella typhi: a worldwide epidemic. Clin. Infect. Dis. 24, S106–S109. doi: 10.1093/clinids/24.supplement_1.s106
Rozwandowicz, M., Brouwer, M. S. M., Fischer, J., Wagenaar, J. A., Gonzalez-Zorn, B., Guerra, B., et al. (2018). Plasmids carrying antimicrobial resistance genes in Enterobacteriaceae. J. Antimicrob. Chemother. 73, 1121–1137. doi: 10.1093/jac/dkx488
Rychlik, I., Karasova, D., Sebkova, A., Volf, J., Sisak, F., Havlickova, H., et al. (2009). Virulence potential of five major pathogenicity islands (SPI-1 to SPI-5) of Salmonella enterica serovar Enteritidis for chickens. BMC Microbiol. 9:268. doi: 10.1186/1471-2180-9-268
Samia, D., Bakir, M., Rachid, E., Chaffia, B., Omar, B., Rolain, J. M., et al. (2021). Prevalence and genotypic characterization of Salmonella spp. from chicken meats marketed in the province of Skikda, Algeria. J. Infect. Dev. Ctries. 15, 523–529. doi: 10.3855/jidc.13986
Samper-Cativiela, C., Diéguez-Roda, B., Trigo da Roza, F., Ugarte-Ruiz, M., Elnekave, E., Lim, S., et al. (2022). Genomic characterization of multidrug-resistant Salmonella serovar kentucky ST198 isolated in poultry flocks in Spain (2011-2017). Microb. Genom. 8:000773. doi: 10.1099/mgen.0.000773
Shang, K., Wei, B., Jang, H.-K., and Kang, M. (2019). Phenotypic characteristics and genotypic correlation of antimicrobial resistant (AMR) Salmonella isolates from a poultry slaughterhouse and its downstream retail markets. Food Control 100, 35–45. doi: 10.1016/j.foodcont.2018.12.046
Shea, K. M. (2004). Nontherapeutic use of antimicrobial agents in animal agriculture: implications for pediatrics. Pediatrics 114, 862–868. doi: 10.1542/peds.2004-1233
Shen, W., Chen, H., Geng, J., Wu, R. A., Wang, X., and Ding, T. (2022). Prevalence, serovar distribution, and antibiotic resistance of Salmonella spp. isolated from pork in China: a systematic review and meta-analysis. Int. J. Food Microbiol. 361:109473. doi: 10.1016/j.ijfoodmicro.2021.109473
Silva, C., Puente, J. L., and Calva, E. (2017). Salmonella virulence plasmid: pathogenesis and ecology. Pathogens Disease 75:070. doi: 10.1093/femspd/ftx070
Smith, J. C., Varriano, S., Roach, K., Snipes, Z., Dawson, J. L., Shealy, J., et al. (2023). Prevalence and molecular characterization of Salmonella isolated from wild birds in fresh produce environments. Front. Microbiol. 14:1272916. doi: 10.3389/fmicb.2023.1272916
Sohail, M. N., Rathnamma, D., Priya, S. C., Isloor, S., Naryanaswamy, H. D., Ruban, S. W., et al. (2021). Salmonella from farm to table: isolation, characterization, and antimicrobial resistance of Salmonella from commercial broiler supply chain and its environment. Biomed. Res. Int. 2021:3987111. doi: 10.1155/2021/3987111
Spanò, S., Ugalde, J. E., and Galán, J. E. (2008). Delivery of a Salmonella typhi exotoxin from a host intracellular compartment. Cell Host Microbe 3, 30–38. doi: 10.1016/j.chom.2007.11.001
Sugawara, Y., Akeda, Y., Hagiya, H., Sakamoto, N., Takeuchi, D., Shanmugakani, R. K., et al. (2019). Spreading patterns of NDM-producing Enterobacteriaceae in clinical and environmental settings in Yangon, Myanmar. Antimicrob. Agents Chemother. 63:e01924-18. doi: 10.1128/aac.01924-18
Sun, J., Ke, B., Huang, Y., He, D., Li, X., Liang, Z., et al. (2014). The molecular epidemiological characteristics and genetic diversity of Salmonella typhimurium in Guangdong, China, 2007-2011. PLoS One 9:e113145. doi: 10.1371/journal.pone.0113145
Tedersoo, T., Roasto, M., Mäesaar, M., Häkkinen, L., Kisand, V., Ivanova, M., et al. (2022). Antibiotic resistance in Campylobacter spp. isolated from broiler chicken meat and human patients in Estonia. Microorganisms 10:1067. doi: 10.3390/microorganisms10051067
Thakur, R., Suri, C. R., and Rishi, P. (2022). Contribution of typhoid toxin in the pathogenesis of Salmonella typhi. Microb. Pathog. 164:105444. doi: 10.1016/j.micpath.2022.105444
Ubeyratne, K. H., Madalagama, R. P., Liu, X., Pathirage, S., Ariyawansa, S., Wong, M. K. L., et al. (2023). Phenotypic and genotypic characterization of antibiotic-resistant Salmonella isolated from humans, aquaculture, and poultry in Sri Lanka: a retrospective study. J. Infect. Public Health 16, 203–209. doi: 10.1016/j.jiph.2023.10.028
Varga, C., Middleton, D., Walton, R., Savage, R., Tighe, M. K., Allen, V., et al. (2012). Evaluating risk factors for endemic human Salmonella enteritidis infections with different phage types in Ontario, Canada using multinomial logistic regression and a case-case study approach. BMC Public Health 12:866. doi: 10.1186/1471-2458-12-866
Villa, L., García-Fernández, A., Fortini, D., and Carattoli, A. (2010). Replicon sequence typing of IncF plasmids carrying virulence and resistance determinants. J. Antimicrob. Chemother. 65, 2518–2529. doi: 10.1093/jac/dkq347
Wang, Y. U., Pettengill, J. B., Pightling, A., Timme, R., Allard, M., Strain, E., et al. (2018). Genetic diversity of Salmonella and Listeria isolates from food facilities. J. Food Prot. 81, 2082–2089. doi: 10.4315/0362-028x.jfp-18-093
Wang, J., Stephan, R., Karczmarczyk, M., Yan, Q., Hächler, H., and Fanning, S. (2013). Molecular characterization of Bla ESBL-harboring conjugative plasmids identified in multi-drug resistant Escherichia coli isolated from food-producing animals and healthy humans. Front. Microbiol. 4:188. doi: 10.3389/fmicb.2013.00188
Wang, Z., Wang, H., Chen, D., and Li, Y. (2023). Genomic characterization and comparative genomic analysis of pathogenic Vibrio isolated from aquaculture-grown white-leg shrimp (Penaeus vannamei) in Guangdong and Jiangsu, China. Aquaculture. 580:740302. doi: 10.1016/j.aquaculture.2023.740302
Wang, X., Xie, Y., Cai, H., Duan, S., Song, X., Wu, Y., et al. (2021). Growth and survival characteristics of Salmonella enterica regarding antibiotic resistance phenotypes. LWT Food Sci. Technol. 149:111872. doi: 10.1016/j.lwt.2021.111872
Wang, Z., Zhang, J., Liu, S., Zhang, Y., Chen, C., Xu, M., et al. (2022). Prevalence, antimicrobial resistance, and genotype diversity of Salmonella isolates recovered from retail meat in Hebei Province, China. Int. J. Food Microbiol. 364:109515. doi: 10.1016/j.ijfoodmicro.2021.109515
Wójcicki, M., Świder, O., Daniluk, K. J., Średnicka, P., Akimowicz, M., Roszko, M., et al. (2021). Transcriptional regulation of the multiple resistance mechanisms in Salmonella-a review. Pathogens 10:801. doi: 10.3390/pathogens10070801
Wong, M. H., Chan, E. W., and Chen, S. (2017). IS26-mediated formation of a virulence and resistance plasmid in Salmonella enteritidis. J. Antimicrob. Chemother. 72, 2750–2754. doi: 10.1093/jac/dkx238
Wong, M. H., Yan, M., Chan, E. W., Liu, L. Z., Kan, B., and Chen, S. (2013). Expansion of Salmonella enterica Serovar Typhimurium ST34 clone carrying multiple resistance determinants in China. Antimicrob. Agents Chemother. 57, 4599–4601. doi: 10.1128/aac.01174-13
Wu, S.-Y., Wang, L.-D., Li, J.-L., Xu, G.-M., He, M.-L., Li, Y.-Y., et al. (2016). Salmonella spv locus suppresses host innate immune responses to bacterial infection. Fish Shellfish Immunol. 58, 387–396. doi: 10.1016/j.fsi.2016.09.042
Xavier, B. B., Lammens, C., Butaye, P., Goossens, H., and Malhotra-Kumar, S. (2016). Complete sequence of an IncFII plasmid harbouring the colistin resistance gene mcr-1 isolated from Belgian pig farms. J. Antimicrob. Chemother. 71, 2342–2344. doi: 10.1093/jac/dkw191
Xu, Z., Wang, M., Zhou, C., Gu, G., Liang, J., Hou, X., et al. (2020). Prevalence and antimicrobial resistance of retail-meat-borne Salmonella in southern China during the years 2009-2016: the diversity of contamination and the resistance evolution of multidrug-resistant isolates. Int. J. Food Microbiol. 333:108790. doi: 10.1016/j.ijfoodmicro.2020.108790
Yang, J., Ju, Z., Yang, Y., Zhao, X., Jiang, Z., and Sun, S. (2019). Serotype, antimicrobial susceptibility and genotype profiles of Salmonella isolated from duck farms and a slaughterhouse in Shandong province, China. BMC Microbiol. 19:202. doi: 10.1186/s12866-019-1570-z
Keywords: Salmonella, serotype, antimicrobial resistance genes, virulence genes, poultry
Citation: Chen L, Shi Y, Wang M, Li Y and Si Z (2025) Comprehensive epidemiological profiling of poultry-derived Salmonella spp. in Shandong, China, 2019–2022: a longitudinal study of prevalence, antibiotic resistances, virulence factors and molecular characteristics. Front. Microbiol. 16:1541084. doi: 10.3389/fmicb.2025.1541084
Received: 07 December 2024; Accepted: 19 February 2025;
Published: 05 March 2025.
Edited by:
Arun K. Bhunia, Purdue University, United StatesReviewed by:
Adriana Morar, Banat University of Agricultural Sciences and Veterinary Medicine, RomaniaCopyright © 2025 Chen, Shi, Wang, Li and Si. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Zhenshu Si, c2l6aGVuc2h1QGxjdS5lZHUuY24=
†These authors have contributed equally to this work
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.