- 1Department of Basic Sciences, Shoushtar Faculty of Medical Sciences, Shoushtar, Iran
- 2Department of Microbiology, Shahid Beheshti University, Tehran, Iran
- 3Quality Control Department of Temad Mfg, Co., Tehran, Iran
- 4Department of Microbiology, Science and Research Branch, Islamic Azad University, Tehran, Iran
- 5School of Medicine, Hamadan University of Medical Sciences, Hamadan, Iran
- 6Department of Microbiology, School of Medicine, Tehran University of Medical Sciences, Tehran, Iran
- 7Department of Bacteriology, Pasteur Institute of Iran, Tehran, Iran
Background: Staphylococcus is a genus of bacteria responsible for various infections ranging from mild skin to severe systemic diseases. Methicillin-resistant Staphylococcus aureus (MRSA) and coagulase-negative staphylococci (CoNS) are significant challenges owing to their resistance to multiple antibiotics, including macrolides, such as erythromycin, clarithromycin, and azithromycin.
Objective: This study aimed to systematically review and synthesize data on the prevalence of macrolide resistance in Staphylococcus spp., identify trends and changes in resistance patterns over time, and assess how testing methods and guidelines affect reported resistance rates.
Methods: The study conducted a systematic search of the Scopus, PubMed, Web of Science, and EMBASE databases. Studies have reported the proportion of macrolide-resistant Staphylococcus spp. Two authors independently extracted and analyzed the data using a random-effects model. Heterogeneity was assessed, and subgroup analyses were performed based on country, continent, species, AST guidelines, methods, and period.
Results: In total, 223 studies from 76 countries were included. The pooled prevalence of resistance to erythromycin, clarithromycin, and azithromycin were 57.3, 52.6, and 57.9%, respectively. Significant heterogeneity was observed across studies (I2 > 95%, p < 0.001). Oceania (72%) had the highest erythromycin resistance, whereas Europe had the lowest (40.7%). Subgroup analyses revealed variations in resistance based on the species, with higher resistance in MRSA than in MSSA and CoNS than in other species. Over time, a slight decrease in erythromycin resistance has been observed (59.6% from 2015–2019 to 55% from 2020–2023).
Conclusion: This study emphasizes the high prevalence of macrolide resistance in Staphylococcus spp. and its notable regional variation. These findings highlight the necessity for standardized methodologies and global surveillance to manage macrolide resistance effectively. Controlling antibiotic resistance should prioritize enhancing public health measures and updating treatment guidelines.
Systematic review registration: https://www.crd.york.ac.uk/prospero/display_record.php?RecordID=557756, CRD42024557756.
1 Introduction
Staphylococcus is a genus of bacteria that can cause many infections, from mild skin infections to serious systemic diseases. These infections can affect the skin, lungs, bloodstream, and medical devices and have become a significant treatment challenge, particularly for methicillin-resistant Staphylococcus aureus (MRSA) (Tong et al., 2015; Cheung et al., 2021). It is estimated that approximately 30% of people carry S. aureus on their bodies without any symptoms. In 2019, S. aureus was associated with more than 1 million deaths, with an estimated range of 816,000 to 1,470,000 deaths (Ikuta et al., 2022). In the United States, the rate of invasive MRSA infections in the black population (66.5 cases per 100,000 person-years) is more than twice that of the white population (27.7 cases per 100,000 person-years). In Australia, the incidence of Staphylococcus aureus bacteremia (SAB) is 5.8 to 20 times higher among Indigenous Australians than among non-Indigenous Australians. Similarly, in New Zealand, Māori and Pacific Island communities have significantly higher rates of SAB than those of European descent (Tong et al., 2015). In recent years, there has been a significant increase in the rate of MRSA colonization in healthy individuals, potentially contributing to the spread of MRSA in both community and hospital settings (Barcudi et al., 2020). In addition, MRSA is a pathogen resistant to multiple antibiotics, complicating infection management and leading to increased healthcare costs and adverse outcomes (Abebe and Birhanu, 2023; Lan et al., 2024; Saleem et al., 2025). Globally, the pathogen-drug combination with the most significant increase in attributable burden was MRSA. Its attributable deaths have doubled from 57,200 (range 34,100-80,300) in 1990 to 130,000 (range 113,000-146,000) in 2021(Naghavi et al., 2024).
Antibiotic resistance is a global health crisis that threatens the effectiveness of treatments for bacterial infections. Misuse and overuse of antibiotics have accelerated the development of resistance, rendering many therapies ineffective (Yadav and Kapley, 2021; Estany-Gestal et al., 2024). Macrolides, such as erythromycin, clarithromycin, and azithromycin, are widely used to treat various staphylococcal infections. However, the increasing emergence of macrolide resistance in Staphylococcus spp. has become a critical challenge in treating infections caused by these bacteria. Resistance to macrolides has been attributed to the methylation of specific targets in the 23S rRNA by methylases encoded by erm genes, particularly erm(C) and erm(A), which can be constitutive or inducible. In addition, efflux pumps, such as ABC-F proteins encoded by msr genes and major facilitator superfamily transporters encoded by mef genes, drug inactivation by phosphotransferases encoded by mph genes, and esterase encoded by ere genes, confer macrolide resistance (Leclercq, 2002; Miklasinska-Majdanik, 2021; El Mammery et al., 2023; Mahfouz et al., 2023). These mechanisms show regional variation, reflecting differences in the prevalence of resistance genes and differences in antibiotic use practices (Miklasinska-Majdanik, 2021).
Overall, antibiotic resistance reduces the effectiveness of these antibiotics and complicates the treatment of common staphylococcal infections such as skin infections, pneumonia, and bacteremia.
The global burden of macrolide-resistant staphylococci affects both public health and healthcare systems. Data indicate increasing infection rates and resistance patterns, particularly in healthcare-associated infections where S. aureus is a leading cause of morbidity and mortality (An et al., 2024). The economic impact is also profound, with resistant infections leading to longer hospital stays, more complex treatment regimens, and increased healthcare costs (Lodise and McKinnon, 2007). However, the limited number of effective treatment options for resistant infections increases the risk of adverse outcomes. This underscores the importance of developing novel therapeutic approaches and implementing stringent infection control measures (Guo et al., 2020).
Previous research on macrolide resistance in staphylococci has been limited by study design and reporting inconsistencies, making it difficult to draw robust conclusions and identify consistent trends. In addition, many studies require extensive regional analyses, limiting the generalizability of findings and their impact on global health. Furthermore, gaps in understanding the temporal trends and dynamics of resistance highlight the need for longitudinal studies and broader surveillance efforts (Leclercq, 2002; Khader et al., 2019). Hence, standardized methodologies and collaborative efforts across regions are essential to improving our understanding and managing macrolide resistance in staphylococci.
The primary objective of this study was to systematically review and analyze the available data on the prevalence of macrolide resistance in Staphylococcus spp.
The secondary objectives were to identify trends and changes in resistance patterns over time, explore heterogeneity in resistance rates across regions and populations, and assess the impact of testing methods and guidelines on reported resistance rates. By addressing these objectives, this study aimed to fill the existing knowledge gaps and provide comprehensive insights into the dynamics of macrolide resistance in Staphylococcus spp. to guide future research and clinical practice.
2 Methods
This study was conducted according to PRISMA guidelines and included a meta-analysis to increase the robustness of the results. The study was registered in the PROSPERO registry under the code CRD42024557756.
2.1 Eligibility criteria
The inclusion criteria for this meta-analysis stipulated that studies must investigate Staphylococcus spp. macrolide resistance, report resistance rates, specify sample size determination and have complete English-language articles available. Only cross-sectional studies providing antimicrobial resistance (AMR) data, mainly those reporting baseline resistance levels before any interventions, were included. Such studies offer a population-based overview of resistance rates at a specific time and are, therefore, suitable for estimating the prevalence of macrolide resistance. Studies were excluded if published in languages other than English and were review articles, case reports, and case series studies.
2.2 Information sources
A comprehensive search was conducted in several major online databases, including Scopus, PubMed, Web of Science, and EMBASE, focusing on studies published through December 2023. These databases were selected for their extensive coverage of biomedical literature, ensuring a broad scope for the systematic review.
2.3 Search strategy
The search syntax was tailored to each database according to their respective guidelines (“Staphylococcus*” OR “S. aureus” OR “S. epidermidis” OR “S. saprophyticus” OR “S. lugdunensis” OR “S. hominis” OR “S. capitis” OR “S. haemolyticus” OR “CoNS” OR “MRCoNS” OR “MRSA” OR “MSCoNS” OR “VISA” OR “VSSA”) AND (macrolide* OR azithromycin OR clarithromycin OR erythromycin OR roxithromycin OR telithromycin OR spiramycin OR fidaxomicin) AND (resistant* OR susceptible*). This rigorous methodological approach ensured comprehensive coverage of relevant research topics.
2.4 Selection process
The systematic online database search results were imported into EndNote (version 20), removing duplicate entries. Two authors (NG and EP) independently screened and analyzed the relevant publications to minimize bias. Disagreements were resolved by a third author (TN).
2.5 Data collection process
Data extracted included first author(s), publication year, country, diagnostic method, sample source, number of positive tests, and total sample size. To ensure accuracy, two authors (MM and MB) extracted the data independently, and any disagreements were resolved by consensus.
2.6 Study risk of bias assessment
The quality of the included studies was assessed using the JBI tool. Two authors (MB and TN) independently evaluated the quality, and a third author (MSH) resolved disagreements.
2.7 Synthesis methods
This comprehensive systematic review and meta-analysis aimed to determine the global prevalence of macrolide-resistant Staphylococcus species. The analysis used proportions as the primary outcome measure. The main objective was to assess the prevalence of macrolide-resistant Staphylococcus strains, while the secondary objective sought to identify sources of heterogeneity between studies. Subgroup analyses investigated potential variability in resistance rates across different demographic and methodological factors. Additionally, trends in macrolide resistance over time were examined.
A random effects model was employed to analyze the data, allowing for considering variability within and between studies. The degree of heterogeneity was estimated using the DerSimonian-Laird method for τ2. Along with τ2, the Q-test for heterogeneity and the I2 statistic (Higgins and Thompson, 2002) were also calculated. Heterogeneity was considered present if τ2 > 0, regardless of the Q-test results.
Subgroup analyses were performed across various factors to explore sources of heterogeneity, including countries, continents, antibiotic susceptibility testing (AST) guidelines, AST methods, Staphylococcus species, coagulase status, and year groups. This stratification helped identify macrolide resistance patterns and potential drivers across regions and testing protocols.
A Logit Transformation was applied to the proportions of macrolide-resistant Staphylococcus species to account for variations in the proportion data and stabilize the variance. The logit transformation—also known as the log-odds transformation—was used to ensure that the outcome variable remained within the 0 to 1 range, mainly when dealing with extreme proportions of resistance. This transformation also normalized the distribution of proportions, facilitating more accurate meta-regression modeling.
Meta-regression analysis was conducted to explore temporal trends in macrolide resistance over time. Moderator variables included country, continent, AST guidelines, and year group. This analysis aimed to identify how macrolide resistance in Staphylococcus species has evolved across different geographical regions and under varying testing conditions.
Outliers and influential studies were identified using studentized residuals and Cook’s distances. Studies with studentized residuals exceeding the 100 × (1–0.05 / (2 × k)) th percentile of a standard normal distribution were flagged as potential outliers (after applying a Bonferroni correction for α = 0.05 and for k studies in the meta-analysis). Studies with Cook’s distances greater than the median plus six times the interquartile range of Cook’s distances were considered influential and examined for their impact on the overall estimates.
Funnel plot asymmetry was assessed using rank correlation and regression tests, with the standard error of the observed results serving as the predictor. This approach was used to evaluate potential publication bias. All statistical analyses were performed using R (version 4.2.1) and the metafor package (version 3.8.1) (Cochran, 1954; Begg and Mazumdar, 1994; Higgins and Thompson, 2002; Sterne and Egger, 2005; Viechtbauer, 2010; Viechtbauer and Cheung, 2010; Kuhn et al., 2015).
3 Results
3.1 Descriptive statistics
A total of 21,273 records as results of the systematic search were collected in reference manager software (EndNote version 20), and 14,285 duplicated articles were removed. Thousand eighty-eight articles were assessed in the title abstract for this section; 990 full-text articles were evaluated and excluded. Eventually, this systematic review and meta-analysis included 207 eligible studies. The reports came from 76 countries and six continents. The reports cover the years 2015 to 2023. The screening and selection of presages are summarized in the PRISMA flowchart (Figure 1). Characteristics and references of included studies are presented in Table 1.
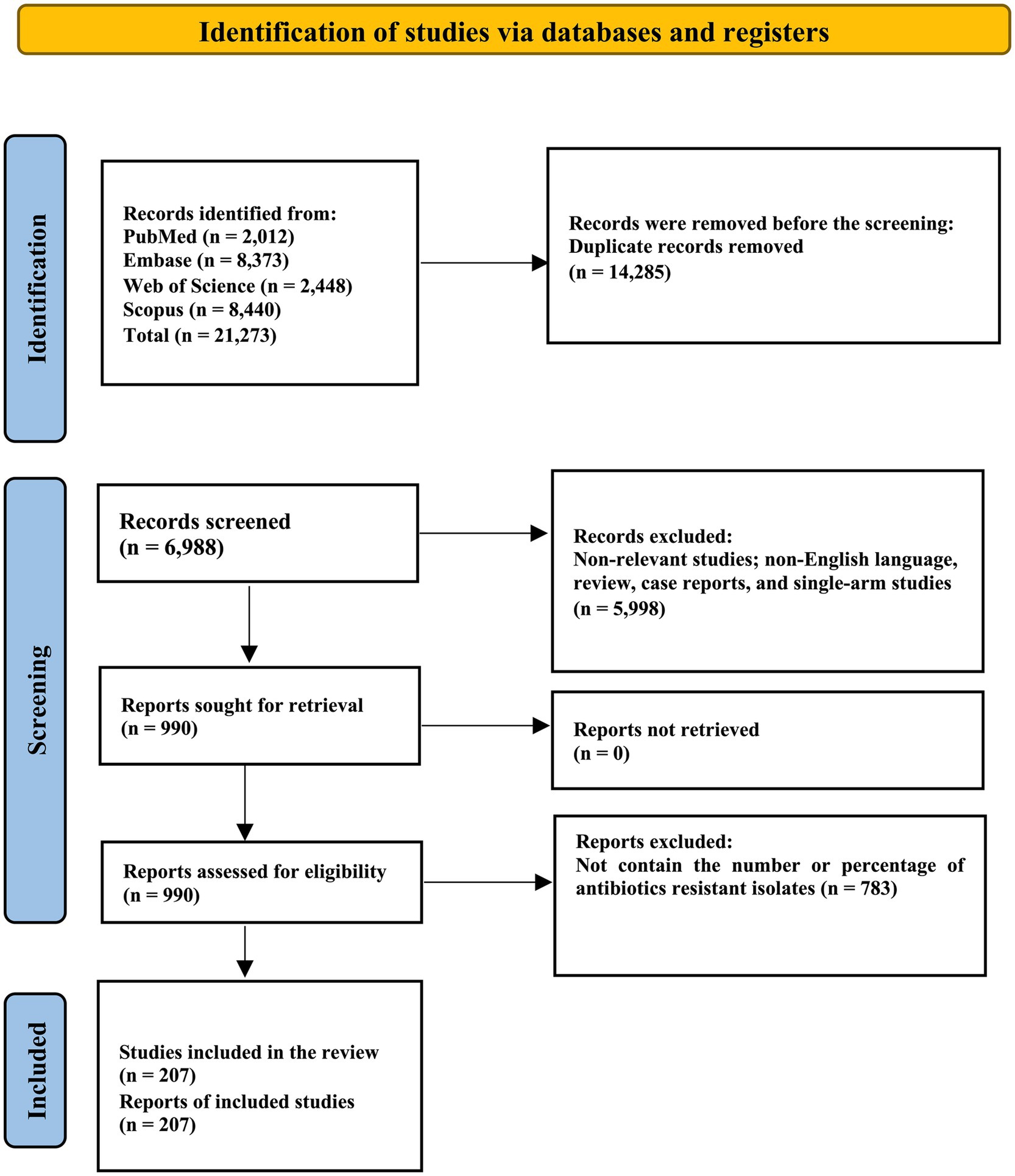
Figure 1. PRISMA flow diagram of study selection: this diagram illustrates the process of study identification, screening, eligibility assessment, and inclusion for the review. From a total of 21,273 records identified through databases, 207 studies were included in the final review after exclusion based on criteria such as duplication, irrelevance, and lack of data on antibiotic-resistant isolates.
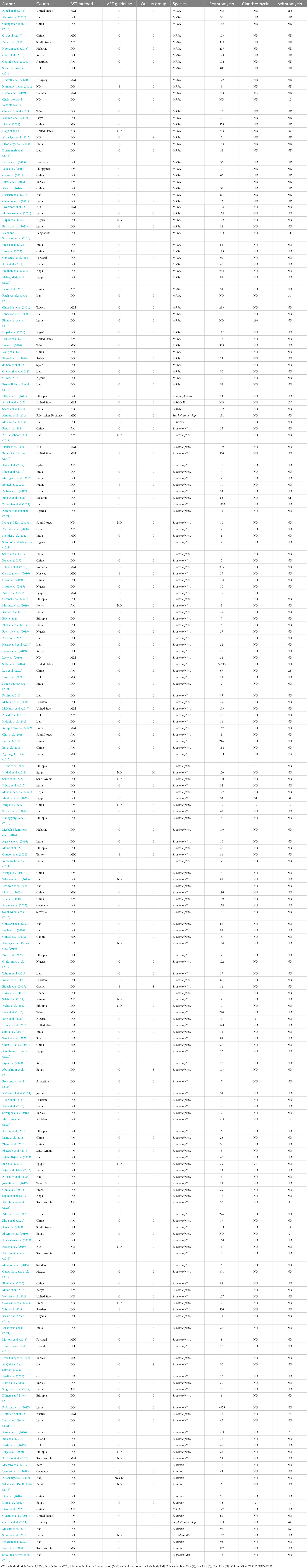
Table 1. A summary of the included studies in the meta-analysis is provided below, highlighting the characteristics employed.
3.2 Comprehensive overview of antibiotic resistance prevalence
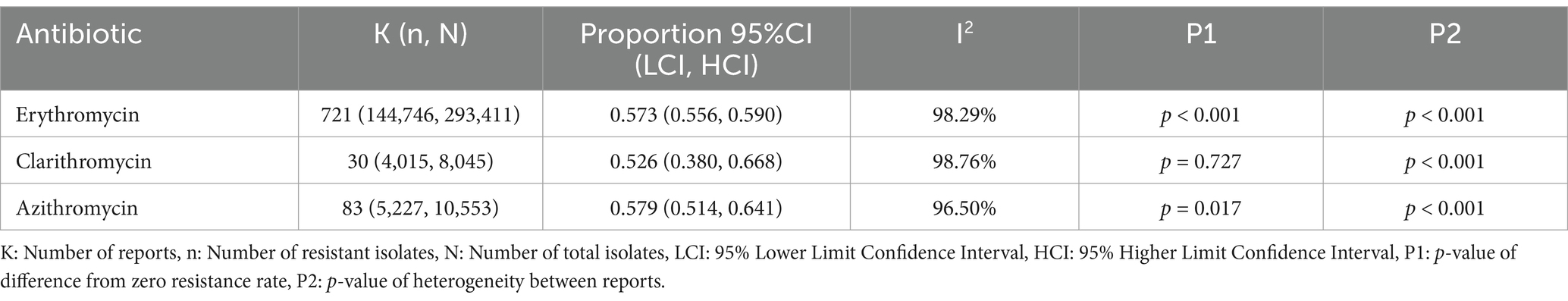
Among 360 reports, the proportion of erythromycin-resistant isolates was 0.573 (95% CI: 0.556–0.590), based on 144,746 resistant isolates out of 293,411 isolates tested. The heterogeneity among reports was significant (I2 = 96.09%, p = 0.001). Similarly, the proportion of clarithromycin resistance, as assessed by 30 reports involving 4,015 resistant isolates out of 8,045 tested isolates, was 0.526 (95% CI: 0.380–0.668), with significant heterogeneity between reports (I2 = 98.76%, p = 0.001). In addition, the proportion of azithromycin-resistant isolates, derived from 83 reports containing 5,227 resistant isolates out of 10,553 isolates tested, was 0.579 (95% CI: 0.514–0.641), again with significant heterogeneity between reports (I2 = 96.50%, p = 0.001).
3.2.1 Prevalence of erythromycin resistance
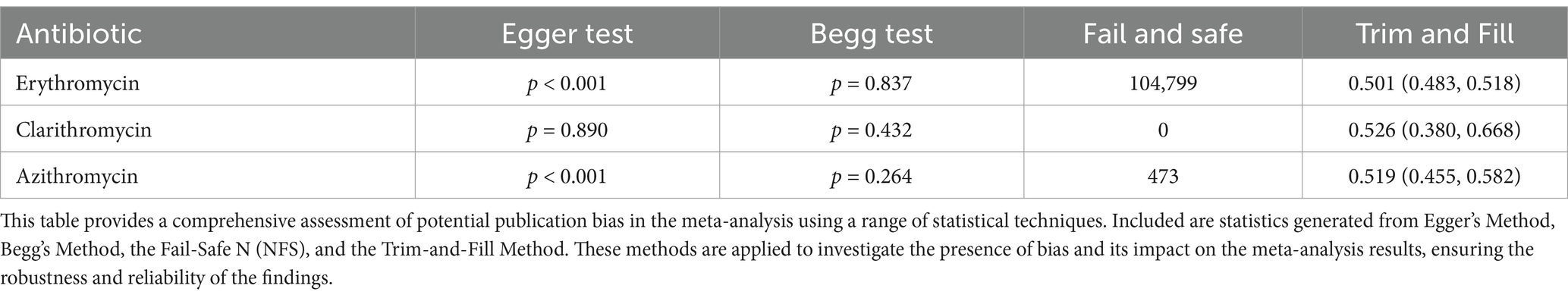
A total of 293,411 isolates from 721 studies were included in the erythromycin resistance analysis. The estimated mean proportion based on the random effects model was 0.573 (95% CI: 0.556–0.590). This result indicates that the mean proportion differed significantly from zero (z = 8.400, p < 0.001). The heterogeneity between studies was significant, as noted in the Q-test (Q(720) = 42,007.095, I2 = 98.29%, p < 0.001) (Table 2). A forest plot illustrating the observed results and the random effects model estimate is shown in Figure 2. Using the trim-and-fill method, the adjusted proportion was 0.501 (95% CI: 0.483–0.518). Analysis of the studentized residuals identified several studies with values greater than 3.979, suggesting potential outliers within the model. After excluding these potential outliers, the proportion was 0.501 (95% CI: 0.483–0.518). Cook’s distance analysis also indicated that several studies were overly influential. After removing these influential studies, the proportion remained unchanged at 0.501 (95% CI: 0.483–0.518). Both the rank correlation test and the regression test suggested a potential funnel plot asymmetry (p < 0.001 and p = 0.018, respectively) (Table 3).
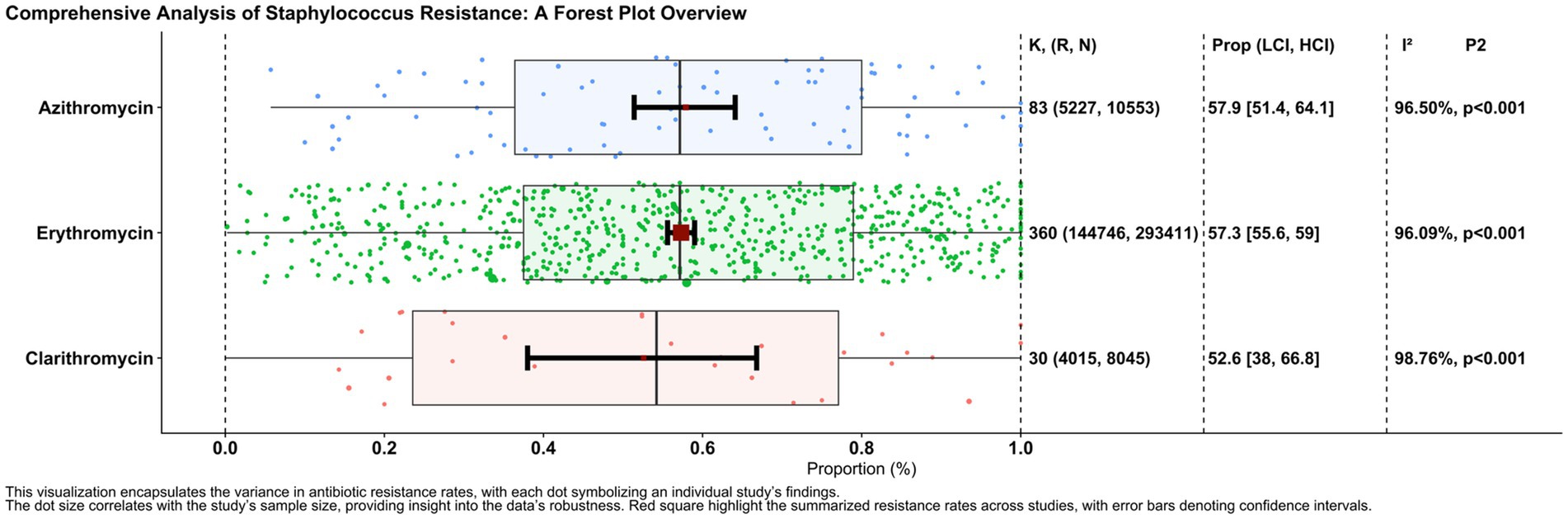
Figure 2. Forest plot of resistance rates for macrolide antibiotics against Staphylococcus: the forest plot summarizes the resistance rates of Staphylococcus species to Azithromycin, Erythromycin, and Clarithromycin across various studies. Each dot represents an individual study’s data point, with red squares indicating pooled resistance estimates and black bars showing confidence intervals.
3.2.2 Prevalence of clarithromycin resistance
The clarithromycin resistance analysis included Eight forty-five isolates from 30 studies. The estimated average proportion based on the random-effects model was 0.526 (95%CI, 0.380, 0.668). Therefore, the average outcome was not significantly different from zero (z = 0.349, p = 0.727). According to the Q test, the outcomes were heterogeneous (Q (29) = 2347.241, I 2 = 98.76%, p < 0.001). A forest plot showing the observed outcomes and the estimate based on the random effects model is shown in Figure 2. With the fill and trim method implementation, the proportion changed to 0.526 (95%CI, 0.380, 0.668). Examination of the studentized residuals revealed that none of the studies had values greater than 3.144. Hence, there was no indication of outliers in the context of this model. According to Cook’s distance, none of the studies could be considered overly influential. Neither the rank correlation nor the regression test indicated funnel plot asymmetry (p = 0.432 and p = 0.890, respectively) (Figure 3).
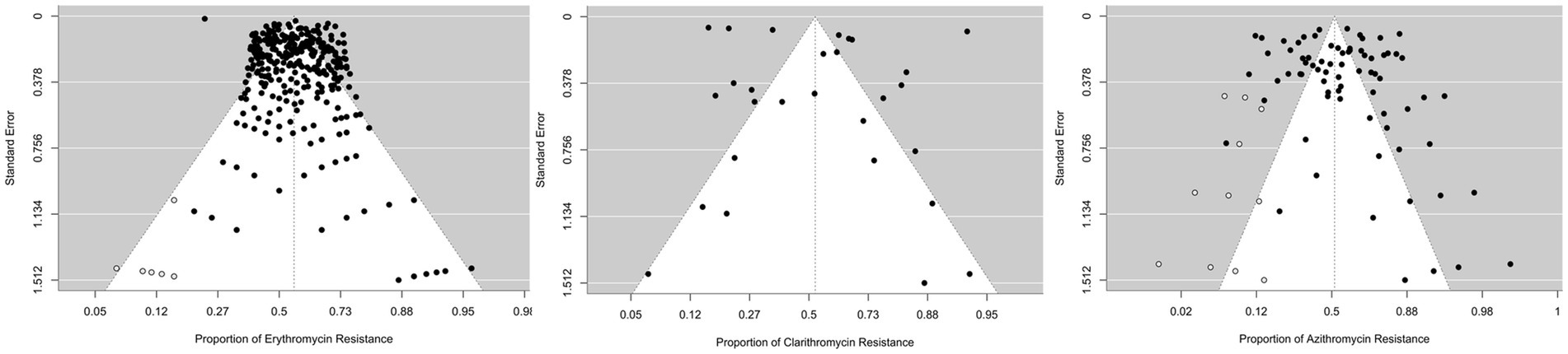
Figure 3. Funnel plots for publication bias analysis: funnel plots assessing the presence of publication bias in resistance studies for Erythromycin (left), Clarithromycin (middle), and Azithromycin (right). Symmetrical distributions indicate minimal bias, whereas asymmetries may suggest potential bias.
3.2.3 Prevalence of azithromycin resistance
The analysis of azithromycin resistance included data from 83 studies with 10,553 isolates. Using a random effects model, the estimated mean proportion was 0.579 (95% CI: 0.514, 0.641), indicating that the mean outcome differed significantly from zero (z = 2.385, p = 0.017). The heterogeneity of the outcomes was confirmed by the Q-test (Q(82) = 2342.061, I2 = 96.50%, p < 0.001). After using the fill-and-trim method, the proportion was adjusted to 0.519 (95% CI: 0.455, 0.582). Analysis of the studentized residuals showed no study exceeded a value of 3.431, indicating no outliers in the model. Furthermore, Cook’s distance analysis indicated that no single study had an undue influence on the results. While the regression test revealed funnel plot asymmetry (p < 0.001), the rank correlation test did not reveal significant asymmetry (p = 0.264).
3.3 Subgroup analysis
This section provides a detailed summary of the subgroup analyses performed on antimicrobial resistance. The full dataset is available in Table 4. The analyses examined variations in resistance rates across geographic regions, antimicrobial susceptibility testing (AST) methods, time trends, and study quality.
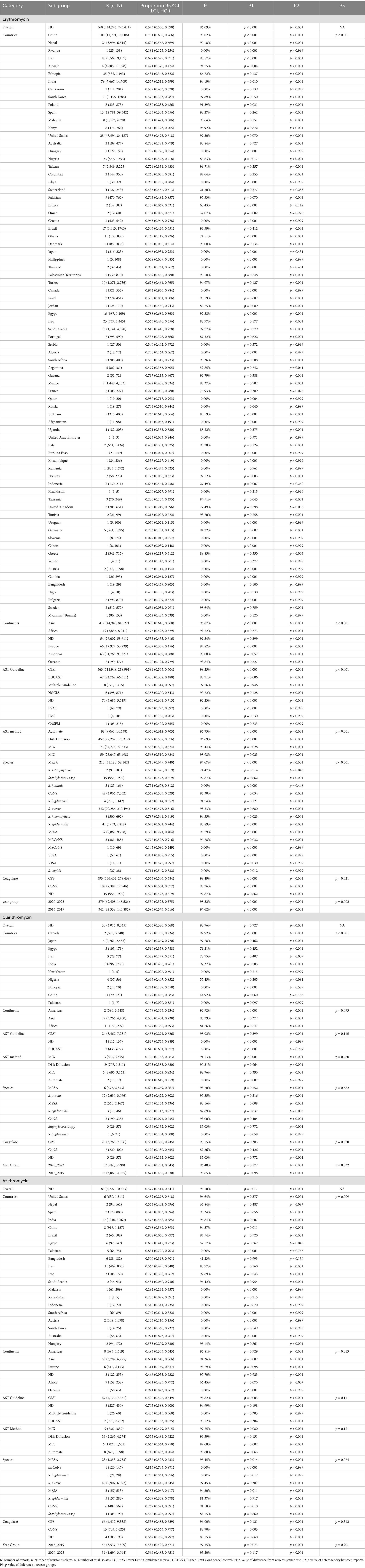
Table 4. Meta-analysis statistics of worldwide antibiotic resistance in staphylococcus spp. and subgroup analysis results.
3.3.1 Subgroup analysis based on countries
Subgroup analysis revealed statistically significant differences in antimicrobial resistance prevalence between countries for azithromycin, clarithromycin, and erythromycin. Austria had the lowest resistance rate for azithromycin, with a prevalence of 13.5%, while Australia had the highest resistance rate at 92.1%. Pakistan had the lowest resistance rate (14.3%) for clarithromycin, while China had the highest (72.9%). The Philippines had the lowest resistance rate of 2.8% for erythromycin, while Canada had the highest resistance rate of 97.4% (Figure 4).
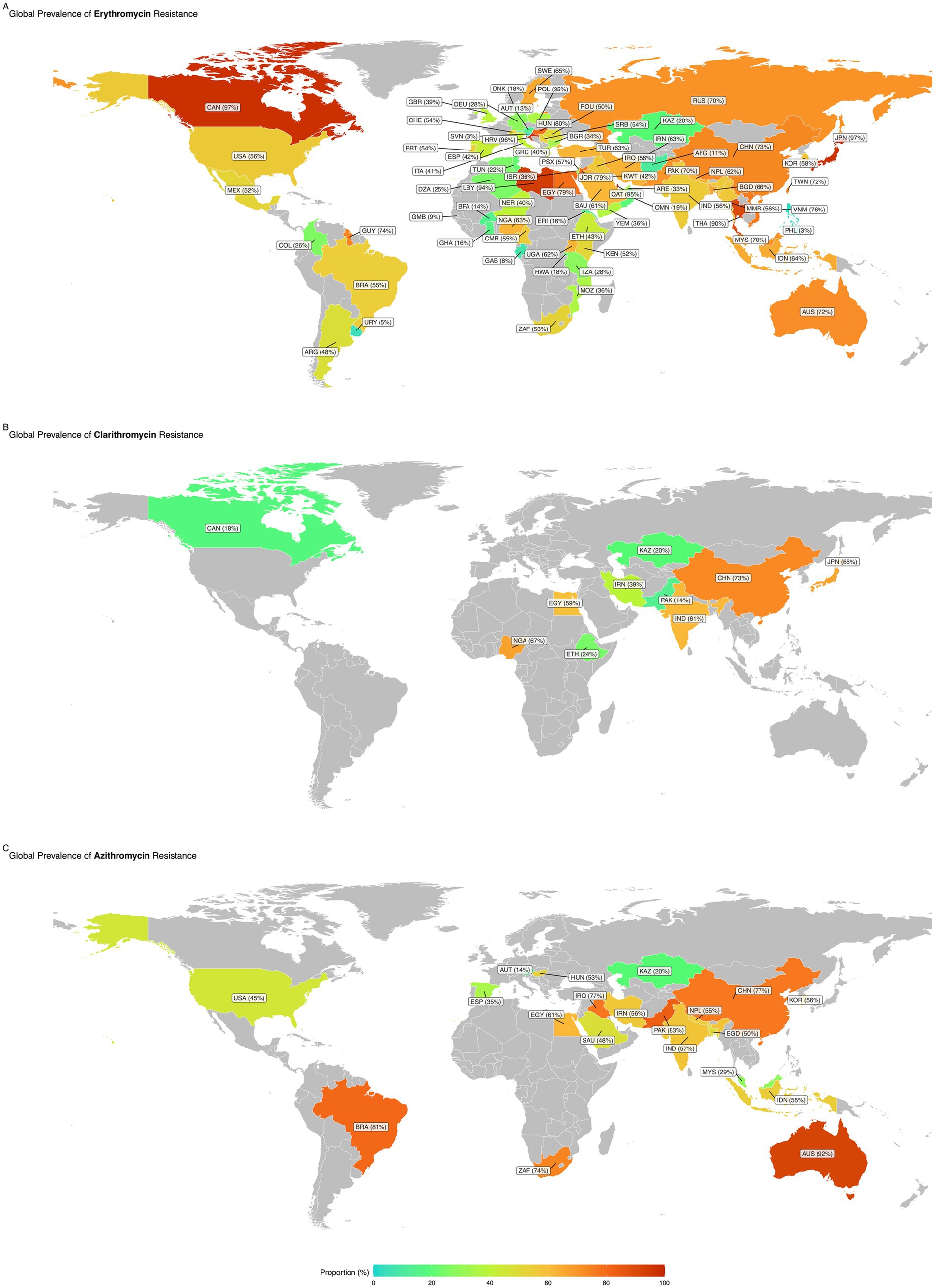
Figure 4. Global prevalence of antibiotic resistance in Staphylococcus: maps showing the worldwide prevalence of resistance to Erythromycin (A), Clarithromycin (B), and Azithromycin (C). Regions with higher resistance proportions are highlighted in warmer colors (e.g., red), while areas with lower resistance rates are shown in cooler tones (e.g., green).
3.3.2 Subgroup analysis based on continents
Subgroup analysis revealed statistically significant differences in antimicrobial resistance prevalence between continents, particularly for azithromycin and erythromycin. Europe had the lowest resistance rate for azithromycin, with a prevalence of 31.1%, while Oceania had the highest resistance rate of 92.1%. Similarly, Europe had the lowest resistance rate for erythromycin, with a prevalence of 40.7%, while Oceania had the highest resistance rate at 72% (Figure 5A).
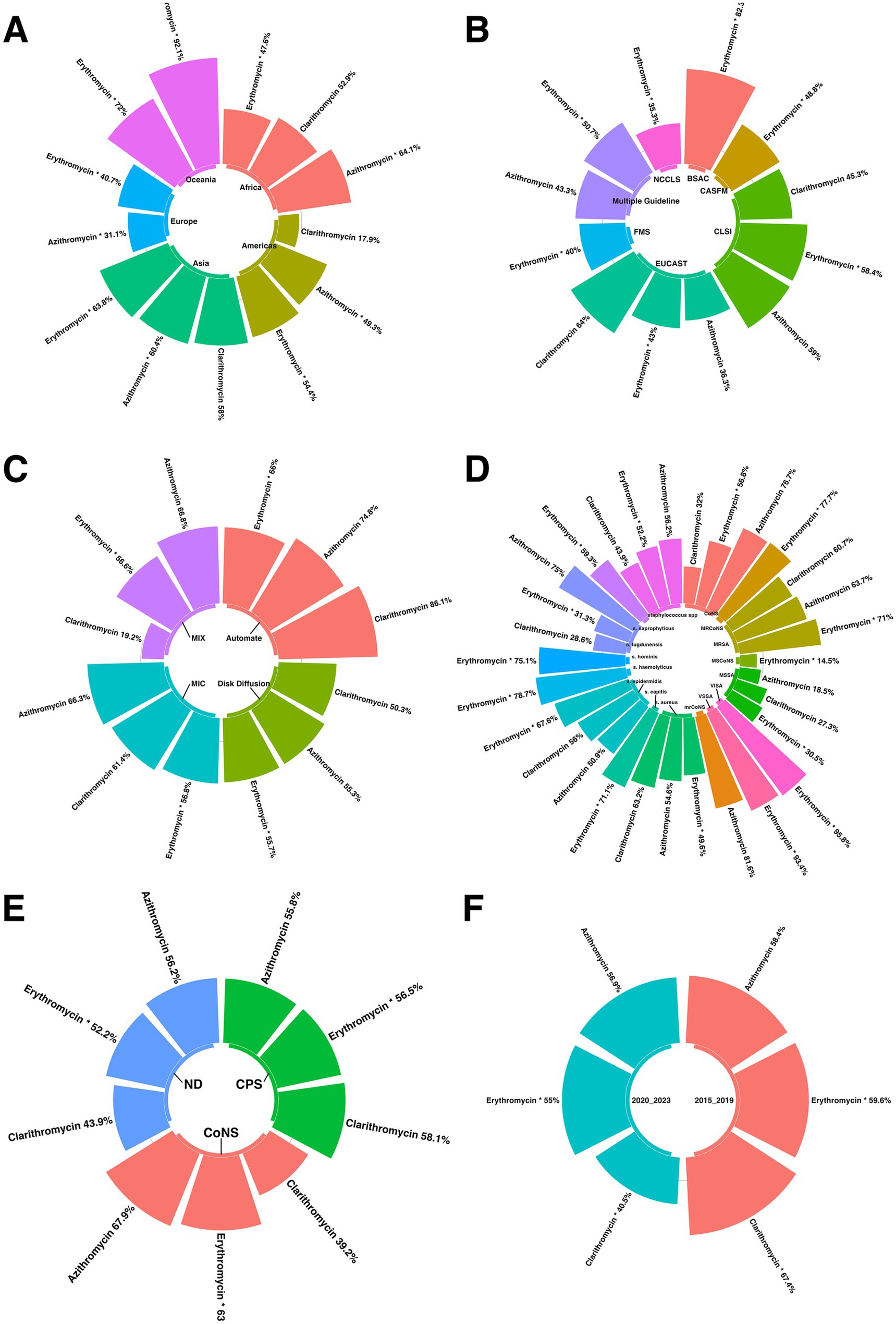
Figure 5. subgroup analysis results were illustrated in figures (A) compression of the prevalence of antibiotic-resistant staphylococcus isolates between continents; (B) Compression of the prevalence of antibiotic-resistant staphylococcus isolates between AST guideline; (C) Compression of the prevalence of staphylococcus isolates AST method; (D) Compression of the prevalence of antibiotic-resistant staphylococcus isolates based on species (E) Compression of the prevalence of antibiotic-resistant staphylococcus isolates based on coagulase; (F) Compression of the prevalence of staphylococcus isolates before and after 2020.
3.3.3 Subgroup analysis based on AST guideline
The subgroup analysis identified statistically significant differences in antibiotic resistance prevalence, including erythromycin, between different antimicrobial susceptibility testing (AST) guidelines. For erythromycin, the NCCLS guideline showed the lowest resistance rate with a prevalence of 35.3%, while the BSAC guideline showed the highest resistance rate at 82.3% (Figure 5B).
3.3.4 Subgroup analysis based on the AST method
Subgroup analysis revealed a statistically significant disparity in the prevalence of antibiotic resistance, including erythromycin, among the various AST methods. For erythromycin, the AST method with the lowest resistance rate was Disk Diffusion, with a prevalence of 55.7%. Conversely, the AST method, with the highest resistance rate, was automated, with a prevalence rate of 66% (Figure 5C).
3.3.5 Subgroup analysis based on species
Subgroup analysis revealed statistically significant differences in antibiotic resistance prevalence among different species, including erythromycin. For erythromycin, MSCoNS had the lowest resistance rate with a prevalence of 14.5%, while VISA had the highest resistance rate with a prevalence of 95.8% (Figure 5D).
3.3.6 Subgroup analysis based on coagulase
Subgroup analysis revealed statistically significant differences in the prevalence of antibiotic resistance, including erythromycin, among different coagulase types. For erythromycin, the coagulase type with the lowest resistance rate was ND, with a prevalence of 52.2%. In contrast, the highest resistance rate was observed for CoNS, with a prevalence of 63.2% (Figure 5E).
3.3.7 Subgroup analysis based on year-group
The subgroup analysis identified statistically significant differences in antibiotic resistance prevalence among different groups, including clarithromycin and erythromycin. For clarithromycin, the period with the lowest resistance rate was 2020–2023, with a prevalence of 40.5%, while the highest resistance rate was observed in 2015–2019, with a prevalence of 67.4%. Similarly, for erythromycin, the lowest resistance rate occurred during 2020–2023, with a prevalence of 55%, while the highest resistance rate was observed during 2015–2019, with a prevalence of 59.6% (Figure 5F).
3.4 Meta-regression
Meta-regression analysis was performed to examine the relationship between antimicrobial resistance rates and year of reporting. No statistically significant correlation was observed for erythromycin (r = −0.041, p-value = 0.007, 95% CI [−0.071, −0.011]) (Figure 6A). Similarly, the correlation was not statistically significant for clarithromycin (r = −0.123, p-value = 0.263, 95% CI [−0.339, 0.093]) (Figure 6B). These results suggest that resistance rates for azithromycin and clarithromycin remained relatively stable over the study period. In contrast, a statistically significant positive correlation was observed for azithromycin (r = 0.005, p-value = 0.929, 95% CI [−0.1, 0.11]) (Figure 6C), indicating an upward trend in erythromycin resistance rates over time.
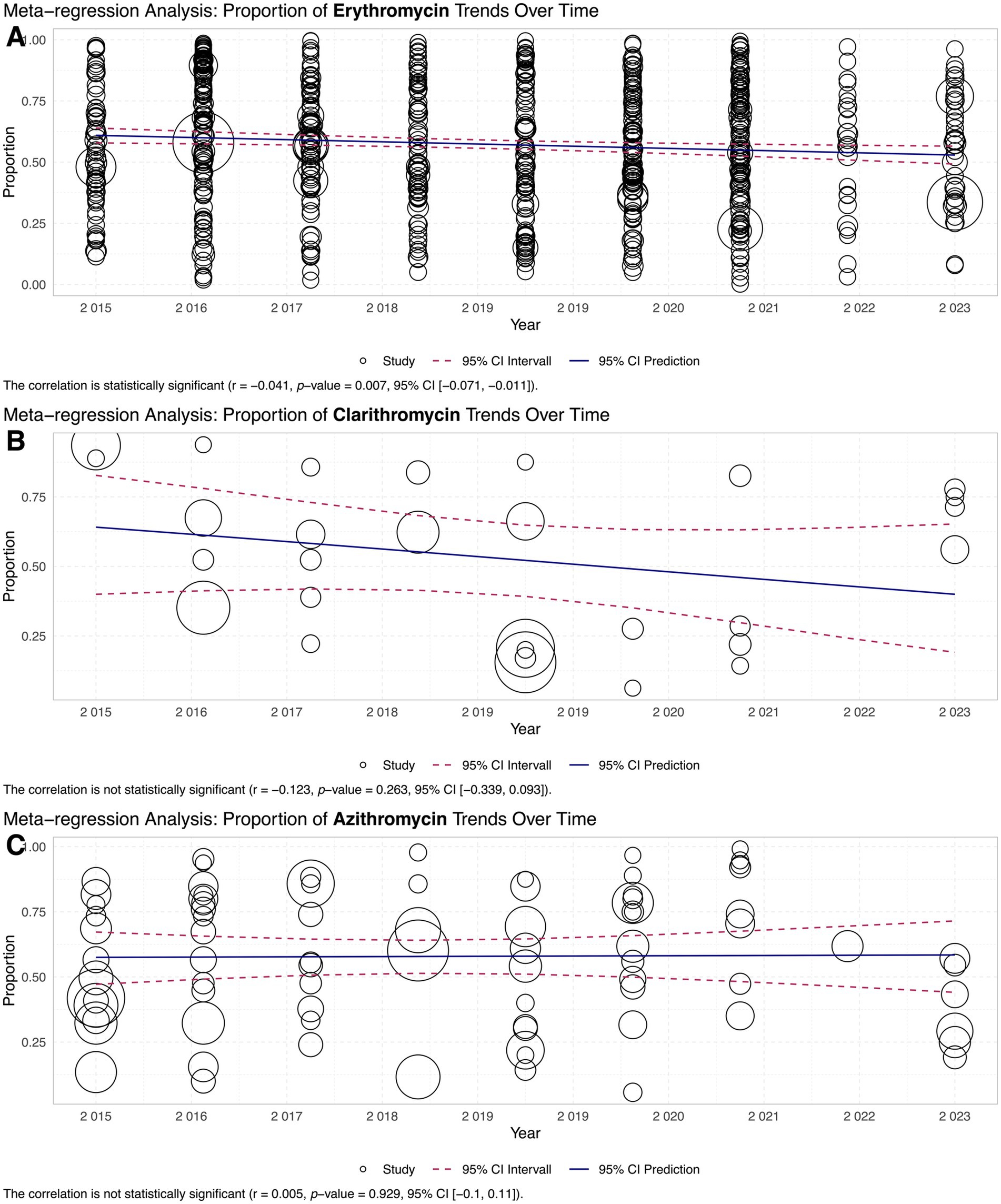
Figure 6. Trends in antibiotic resistance over time (2015–2023): meta-regression analysis plots the trends of resistance proportions for: (A) Erythromycin, showing a slight but statistically significant decline. (B) Clarithromycin, demonstrating a non-significant downward trend. (C) Azithromycin, with no significant trend observed. Data points represent study-specific resistance proportions over time, with bubble sizes reflecting sample size.
4 Discussion
This systematic review and meta-analysis thoroughly evaluated the prevalence and trends of macrolide resistance in Staphylococcus species, explicitly focusing on resistance to erythromycin, clarithromycin, and azithromycin. By analyzing data from 207 studies conducted in 76 countries between 2015 and 2023, our findings provide valuable insights into global patterns of macrolide resistance in Staphylococcus species. Erythromycin, the first macrolide antibiotic discovered, remains effective in treating minor skin infections caused by penicillin-resistant S. aureus strains (Washington and Wilson, 1985). This meta-analysis revealed that erythromycin was the most commonly tested macrolide in antibiotic susceptibility studies, with data from 207 studies in 76 countries. The pooled prevalence of resistance was 57.3%, with significant heterogeneity between studies (I2 = 96.09%, p < 0.001). Evidence of publication bias was also detected using Egger’s test (p < 0.001), resulting in an adjusted pooled prevalence of 50.1% after Fill and Trim analysis. These variations may be due to differences in study populations, periods, sampling methods, or clinical specimen types.
Subgroup analyses revealed significant regional differences in erythromycin resistance rates. Oceania had the highest resistance rate (72%, based on two reports), while Asia contributed the most studies (417 reports) with a pooled prevalence of 63.8%. In particular, China, Iran, and India reported resistance rates of 73.1, 62.7, and 55.7%, respectively, based on 105, 85, and 79 reports. In contrast, Europe had the lowest pooled prevalence of erythromycin-resistant isolates (40.7%, 44 reports), with Spain (13 reports) and Poland (8 reports) reporting prevalence rates of 42.5 and 35%, respectively. The lower resistance rates in Europe reflect increased public awareness and effective public health interventions to curb antimicrobial resistance.
On the other hand, prevalence rates of over 90% for erythromycin-resistant isolates in countries such as Qatar, Canada, Libya, Japan, and Croatia raise significant concerns. However, because these findings are based on AST performed at a single clinical center in each country, the results cannot be generalized to the entire population in these regions. This underscores the need for comprehensive national surveillance systems to monitor antimicrobial resistance in these areas.
Subgroup analysis by species revealed a pooled prevalence of erythromycin resistance in 49.6% of S. aureus isolates (342 reports). In addition, some studies included in this meta-analysis reported erythromycin resistance rates for S. aureus in two subgroups: MSSA (methicillin-susceptible S. aureus) and MRSA. The prevalence of resistance was significantly higher in MRSA than in MSSA (71% vs. 30.5%). However, more studies have focused on MRSA than MSSA (212 vs. 37). These findings are consistent with other meta-analyses that have reported pooled prevalence rates of erythromycin-resistant S. aureus isolates (Eshetie et al., 2016; Khanal et al., 2021; Chelkeba et al., 2022; Chelkeba and Melaku, 2022; Ezeh et al., 2023; Xu et al., 2024). However, most of these studies were based on data from one African country and had fewer studies than ours. Moreover, Chelkeba et al. (2022) and Chelkeba and Melaku (2022), during two separate meta-analyses conducted in Ethiopia, reported 50 and 45% prevalence rates for erythromycin-resistant S. aureus isolates in women with bacteriuria and patients with wound infections, respectively. In a meta-analysis review, Ezeh et al. (2023) reported a prevalence rate of 47% for erythromycin-resistant S. aureus isolates in Nigeria (66 reports) up to 2022. However, data from our meta-analysis highlighted a higher prevalence of erythromycin resistance in Nigeria (23 reports) than in Ezeh et al. (2023) (62.6% vs. 47%). The observed discrepancy in prevalence rates may be due to differences in the periods and number of studies included in these two meta-analyses. Subgroup analysis by species revealed a high pooled prevalence of erythromycin resistance among CoNS isolates at 56.8% (based on 42 reports). In addition, some studies independently reported the frequency of specific CoNS species, allowing pooled prevalence rates to be calculated for each species. Among these, S. epidermidis was the most commonly studied CoNS species (41 reports), with a pooled erythromycin resistance prevalence of 67.7%. Similar to our findings, Deyno et al. (2018) also reviewed the prevalence of antimicrobial resistance among clinical isolates of CoNS in Ethiopia through 2016, reporting a 30% prevalence of erythromycin-resistant CoNS. The discrepancy between our findings and Deyno et al. (2018) may be due to differences in the periods and geographic regions covered by these two meta-analyses. Specifically, our meta-analysis included data collected between 2015 and 2023, whereas Deyno et al. (2018) focused on data up to 2016. Furthermore, our study provided a global overview of antimicrobial resistance prevalence, whereas Deyno et al. (2018) limited their analysis to Ethiopia.
In addition, five studies reported a resistance prevalence of 77.7% among [methicillin-resistant Coagulase-Negative Staphylococci (MRCoNS)], which was significantly higher than the 14.5% reported in a single survey of MSCoNS. However, due to the unequal number of studies, this comparison lacks balance, and further research is needed to make a comprehensive and accurate comparison.
Overall, the prevalence of MRCoNS was significantly lower than that of MRSA. This difference may be attributed to the lower frequency of CoNS infections than S. aureus infections, reducing antimicrobial exposure. However, CoNS have transitioned from being non-pathogenic to emerging as pathogenic strains, potentially acquiring resistance genes from S. aureus (Yu et al., 2017).
In contrast, the prevalence of erythromycin resistance decreased slightly over time, from 59.6% in 2015–2019 to 55% in 2020–2023. This decline may reflect increased national efforts to combat antimicrobial resistance and the implementation of updated treatment guidelines and surveillance systems in developed countries. Similarly, a meta-analysis by Xu et al. (2024), found no significant change in erythromycin-resistant S. aureus isolates from Cystic fibrosis patients when comparing the periods 2008–2015 and 2015–2021.
Based on AST guidelines, the subgroup analysis showed higher resistance levels in the CLSI group compared to the EUCAST group (58.4% vs. 43%). However, this finding may be influenced by more studies using CLSI guidelines (563) compared to EUCAST guidelines (67 studies). Both guidelines are widely used but differ in their breakpoints for determining resistance. For example, EUCAST defines resistance as MIC >1, whereas CLSI uses MIC ≥8. Similarly, EUCAST considers a zone diameter of <21 mm resistant, while CLSI uses a zone diameter of ≤13 mm. These differences and variations in the number of studies likely contributed to the observed differences in erythromycin resistance prevalence.
This meta-analysis found fewer studies evaluated susceptibility testing for azithromycin and clarithromycin than erythromycin. It may be due to the limited clinical use of azithromycin and clarithromycin for treating staphylococcal infections compared to erythromycin. The pooled prevalence of azithromycin resistance was similar to that of erythromycin (57.3% vs. 57.9%). However, significant heterogeneity between studies was observed (I2 = 96.5%, p < 0.001), and Egger’s test indicated potential publication bias (p < 0.001). After applying fill and trim analysis, the pooled prevalence of azithromycin resistance was adjusted to 51.9%.
The highest resistance rates were reported in Oceania (92.1%, based on one report), while most studies (58 reports) were conducted in Asia, with a pooled prevalence of 60.4%. Specifically, India and Iran contributed 17 and 11 reports, respectively, with 57.5 and 56.3% resistance prevalence rates. Like erythromycin, Europe had the lowest prevalence of azithromycin resistance (31.1%, based on six studies). This low prevalence may be due to the limited number of European studies and the infrequent use of azithromycin to treat staphylococcal infections in this region. Alarmingly, high levels of azithromycin-resistant isolates were identified in Pakistan, Brazil, and China.
Subgroup analysis by species showed that S. aureus was the most commonly studied species, with a pooled resistance prevalence of 54.6% (40 reports). In addition, 23 studies reported a high prevalence of azithromycin resistance among MRSA isolates (63.7%), compared with only three studies evaluating MSSA isolates, which showed a much lower resistance prevalence of 18.5%. However, this comparison was biased due to the unequal number of studies. Subgroup analysis by the AST method showed that disc diffusion was the most commonly used method for antibiotic susceptibility testing, probably because of its accessibility and widespread acceptance. However, the highest prevalence of azithromycin resistance was associated with the automated method (74.8%, based on eight reports). Like erythromycin, the prevalence of azithromycin resistance decreased slightly over time, from 58.4% in 2015–2019 to 56.9% in 2020–2023.
Clarithromycin was the third macrolide antibiotic studied in this meta-analysis, with a pooled resistance prevalence of 52.6%; however, there was considerable heterogeneity between studies (I2 = 98.76%, p < 0.001). Most of the reports (17) were from Asia, with a pooled prevalence of 58%. S. aureus was the dominant species, with a resistance prevalence of 63.2%; six studies showed a prevalence rate of 60.7% among MRSA isolates and 27.3% among MSSA isolates (two reports). In contrast to erythromycin and azithromycin, the prevalence of resistance to clarithromycin decreased significantly over different periods (67.4% from 2015 to 2019 and 40.5% from 2020 to 2023).
Clarithromycin, the third macrolide antibiotic examined in this meta-analysis, had a pooled resistance prevalence of 52.6%, although significant heterogeneity between studies was observed (I2 = 98.76%, p < 0.001). Most reports (17 studies) were from Asia, with a pooled resistance prevalence of 58%. S. aureus was the predominant species, with a resistance prevalence of 63.2%. Among MRSA isolates, six studies reported a resistance prevalence of 60.7%, while MSSA isolates had a lower prevalence of 27.3% (based on two reports). In contrast to erythromycin and azithromycin, clarithromycin resistance decreased significantly over time, from 67.4% in 2015–2019 to 40.5% in 2020–2023.
This meta-analysis is the first to compare the prevalence of resistance to azithromycin and clarithromycin in Staphylococcus species. As a result, no previous meta-analyses have provided comparable global results.
A significant limitation of this study is the lack of differentiation between Staphylococcus species isolated from healthcare and community settings. This distinction is critical, as antibiotic resistance rates in healthcare settings are typically higher than in the community. Another limitation is the lack of data on resistance to newer macrolides, primarily due to the limited number of studies investigating them. This gap highlights the need for further research to provide accurate and comprehensive evidence.
5 Conclusion
This meta-analysis highlights a relatively high prevalence of macrolide resistance in S. aureus and CoNS isolates worldwide. These elevated resistance rates underscore the importance of regular epidemiologic surveillance of antimicrobial resistance and the implementation of stewardship programs. Most of the studies included in this analysis were conducted in Asia, while Europe had the lowest macrolide resistance rate. In addition, resistance to erythromycin and azithromycin remained relatively stable between 2015–2019 and 2020–2023. Nevertheless, antimicrobial susceptibility testing before treatment is recommended, and further research into the molecular and genetic mechanisms of macrolide resistance is strongly encouraged.
Data availability statement
The original contributions presented in the study are included in the article/supplementary material, further inquiries can be directed to the corresponding authors.
Author contributions
TN: Data curation, Writing – original draft, Writing – review & editing. AZ: Supervision, Writing – original draft, Writing – review & editing. EP: Investigation, Project administration, Writing – original draft. MM: Investigation, Resources, Visualization, Writing – review & editing. NG: Project administration, Validation, Writing – original draft. MB: Investigation, Methodology, Project administration, Writing – original draft. MS: Formal analysis, Software, Supervision, Writing – original draft.
Funding
The author(s) declare that no financial support was received for the research and/or publication of this article.
Conflict of interest
AZ was employed by Quality Control Department of Temad Mfg, Co.
The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Generative AI statement
The author(s) declare that no Generative AI was used in the creation of this manuscript.
Publisher’s note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
References
Abbasi, M., BaseriSalehi, M., Bahador, N., and Taherikalani, M. (2017). Antibiotic resistance patterns and virulence determinants of different SCCmec and Pulsotypes of Staphylococcus Aureus isolated from a major Hospital in Ilam, Iran. Open Microbiol. J. 11, 211–223. doi: 10.2174/1874285801711010211
Abdulmanea, A. A., Alharbi, N. S., Somily, A. M., Khaled, J. M., and Algahtani, F. H. (2023). The prevalence of the virulence genes of Staphylococcus aureus in sickle cell disease patients at KSUMC, Riyadh, Saudi Arabia. Antibiotics (Basel) 12:1221. doi: 10.3390/antibiotics12071221
Abebe, A. A., and Birhanu, A. G. (2023). Methicillin resistant Staphylococcus aureus: molecular mechanisms underlying drug resistance development and novel strategies to combat. Infect Drug. Resist. 16, 7641–7662. doi: 10.2147/IDR.S428103
Abouelnour, A., Zaki, M., Hassan, R., and Elkannishy, S. (2019). Phenotypic and genotypic identification of Staphylococcus aureus resistant to clindamycin in Mansoura university children hospital, Egypt. Afr. J. Clin. Exp. Microbiol. 21, 30–35. doi: 10.4314/ajcem.v21i1.4
Ackers-Johnson, G., Kibombo, D., Kusiima, B., Nsubuga, M. L., Kigozi, E., Kajumbula, H. M., et al. (2021). Antibiotic resistance profiles and population structure of disease-associated Staphylococcus aureus infecting patients in Fort Portal regional referral hospital, Western Uganda. Microbiology (Reading) 167:001000. doi: 10.1099/mic.0.001000
Adhikari, P., Basyal, D., Rai, J. R., Bharati, L., Budthapa, A., Gharti, K. P., et al. (2023). Prevalence, antimicrobial susceptibility pattern and multidrug resistance of methicillin-resistant Staphylococcus aureus isolated from clinical samples at a tertiary care teaching hospital: an observational, cross-sectional study from the Himalayan country, Nepal. BMJ Open 13:e067384. doi: 10.1136/bmjopen-2022-067384
Agarwal, L., Singh, A. K., Agarwal, A., and Agarwal, A. (2016). Methicillin and mupirocin resistance in nasal colonizers coagulase-negative Staphylococcus among health care workers. Med. J. Dr. DY Patil Univ. 9, 479–483. doi: 10.4103/0975-2870.186070
Aguinagalde, L., Diez-Martinez, R., Yuste, J., Royo, I., Gil, C., Lasa, I., et al. (2015). Auranofin efficacy against MDR Streptococcus pneumoniae and Staphylococcus aureus infections. J. Antimicrob. Chemother. 70, 2608–2617. doi: 10.1093/jac/dkv163
Ahangarzadeh Rezaee, M., Mirkarimi, S. F., Hasani, A., Sheikhalizadeh, V., Soroush, M. H., and Abdinia, B. (2016). Molecular typing of Staphylococcus aureus isolated from clinical specimens during an eight-year period (2005–2012) in Tabriz, Iran. Arch. Pediatr. Infect. Dis. 4:e35563. doi: 10.5812/pedinfect.35563
Ahmad, M., Kumar, P., Sultan, A., Akhtar, A., Chaudhary, B., and Khan, F. (2020). Prevalence of community acquired Uropathogens and their antimicrobial susceptibility in patients from the urology unit of a tertiary care medical center. J. Pure Appl. Microbiol. 14, 2009–2015. doi: 10.22207/JPAM.14.3.40
Akbariyeh, H., Nahaei, M. R., Hasani, A., and Pormohammad, A. (2017). Intrinsic and acquired methicillin-resistance detection in Staphylococcus aureus and its relevance in therapeutics. Arch. Pediatr. Infect. Dis. 5:e39185. doi: 10.5812/pedinfect.39185
Akpaka, P. E., Roberts, R., and Monecke, S. (2017). Molecular characterization of antimicrobial resistance genes against Staphylococcus aureus isolates from Trinidad and Tobago. J. Infect. Public Health 10, 316–323. doi: 10.1016/j.jiph.2016.05.010
Al Zebary, M. K., Yousif, S. Y., and Assafi, M. S. (2017). The prevalence, molecular characterization and antimicrobial susceptibility of isolated from impetigo cases in Duhok, Iraq. Open Dermatol. J. 11, 22–29. doi: 10.2174/1874372201711010022
Al-Habsi, T. H. A., Al-Lamki, R. N. A., and Mabruk, M. (2020). Antibiotic susceptibility pattern of bacterial isolates from wound infections among patients attending a tertiary care hospital in Oman. Biomed. Pharma. J. 13, 2069–2080. doi: 10.13005/bpj/2087
Al-Humaidan, O. S., El-Kersh, T. A., and Al-Akeel, R. A. (2015). Risk factors of nasal carriage of Staphylococcus aureus and methicillin-resistant Staphylococcus aureus among health care staff in a teaching hospital in Central Saudi Arabia. Saudi Med. J. 36, 1084–1090. doi: 10.15537/smj.2015.9.12460
Almasri, M., Abu Hasan, N., and Sabbah, N. (2016). Macrolide and lincosamide resistance in staphylococcal clinical isolates in Nablus, Palestine. Turk. J. Med. Sci. 46, 1064–1070. doi: 10.3906/sag-1503-121
Almohammady, M. N., Eltahlawy, E. M., and Reda, N. M. (2020). Pattern of bacterial profile and antibiotic susceptibility among neonatal sepsis cases at Cairo University children hospital. J. Taibah Univ. Med. Sci. 15, 39–47. doi: 10.1016/j.jtumed.2019.12.005
Al-Naqshbandi, A. A., Chawsheen, M. A., and Abdulqader, H. H. (2019). Prevalence and antimicrobial susceptibility of bacterial pathogens isolated from urine specimens received in rizgary hospital—Erbil. J. Infect. Public Health 12, 330–336. doi: 10.1016/j.jiph.2018.11.005
Al-Qaisi, M. M., and Al-Salmani, T. S. (2020). Phenotypic detection of macrolide-Lincosamide-Streptogramin resistance among Staphylococcus aureus and Staphylococcus epidermidis in Baghdad, Iraq. Int. J. Drug Deliv. Technol. 10, 431–436. doi: 10.25258/ijddt.10.3.22
AL-Salihi, S. S., Karim, G. F., Al-Bayati, A., and Obaid, H. M. (2023). Prevalence of methicillin-resistant and methicillin sensitive Staphylococcus aureus nasal carriage and their antibiotic resistant patterns in Kirkuk City, Iraq. J. Pure Appl. Microbiol. 17, 329–337. doi: 10.22207/JPAM.17.1.22
Al-Tamimi, M., Himsawi, N., Abu-Raideh, J., Khasawneh, A. I., Jazar, D. A., Al-Jawaldeh, H., et al. (2021). Phenotypic and molecular screening of Nasal S. aureus from adult hospitalized patients for methicillin- and vancomycin-resistance. Infect. Disord. Drug Targets 21, 68–77. doi: 10.2174/1871526520666200109143158
An, N. V., Hai, L. H. L., Luong, V. H., Vinh, N. T. H., Hoa, P. Q., Hung, L. V., et al. (2024). Antimicrobial resistance patterns of Staphylococcus Aureus isolated at a general Hospital in Vietnam between 2014 and 2021. Infect. Drug Resist. 17, 259–273. doi: 10.2147/IDR.S437920
Arabestani, M. R., Rastiyani, S., Alikhani, M. Y., and Mousavi, S. F. (2018). The relationship between prevalence of antibiotics resistance and virulence factors genes of MRSA and MSSA strains isolated from clinical samples, West Iran. Oman Med. J. 33, 134–140. doi: 10.5001/omj.2018.25
Asaad, A. M., Ansar Qureshi, M., and Mujeeb Hasan, S. (2016). Clinical significance of coagulase-negative staphylococci isolates from nosocomial bloodstream infections. Infect Dis. (Lond) 48, 356–360. doi: 10.3109/23744235.2015.1122833
Asbell, P. A., Sanfilippo, C. M., Pillar, C. M., DeCory, H. H., Sahm, D. F., and Morris, T. W. (2015). Antibiotic resistance among ocular pathogens in the United States five-year results from the antibiotic resistance monitoring in ocular microorganisms (ARMOR) surveillance study. JAMA Ophthalmol. 133, 1445–1454. doi: 10.1001/jamaophthalmol.2015.3888
Baek, Y. S., Jeon, J., Ahn, J. W., and Song, H. J. (2016). Antimicrobial resistance of Staphylococcus aureus isolated from skin infections and its implications in various clinical conditions in Korea. Int. J. Dermatol. 55, e191–e197. doi: 10.1111/ijd.13046
Bai, B., Lin, Z., Pu, Z., Xu, G., Zhang, F., Chen, Z., et al. (2019). In vitro activity and Heteroresistance of Omadacycline against clinical Staphylococcus aureus isolates from China reveal the impact of Omadacycline susceptibility by branched-chain amino acid transport system II carrier protein, Na/pi cotransporter family protein, and fibronectin-binding protein. Front. Microbiol. 10:2546. doi: 10.3389/fmicb.2019.02546
Banawas, S. S., Alobaidi, A. S., Dawoud, T. M., AlDehaimi, A., Alsubaie, F. M., Abdel-Hadi, A., et al. (2023). Prevalence of multidrug-resistant Bacteria in healthcare-associated bloodstream infections at hospitals in Riyadh, Saudi Arabia. Pathogens 12:1075. doi: 10.3390/pathogens12091075
Barcudi, D., Sosa, E. J., Lamberghini, R., Garnero, A., Tosoroni, D., Decca, L., et al. (2020). MRSA dynamic circulation between the community and the hospital setting: new insights from a cohort study. J. Infect. 80, 24–37. doi: 10.1016/j.jinf.2019.10.001
Baz, A. A., Bakhiet, E. K., Abdul-Raouf, U., and Abdelkhalek, A. (2021). Prevalence of enterotoxin genes (SEA to SEE) and antibacterial resistant pattern of Staphylococcus aureus isolated from clinical specimens in Assiut city of Egypt. Egyptian J. Med. Hum. Genet. 22, 1–12. doi: 10.1186/s43042-021-00199-0
Begg, C. B., and Mazumdar, M. (1994). Operating characteristics of a rank correlation test for publication bias. Biometrics 50, 1088–1101. doi: 10.2307/2533446
Belbase, A., Pant, N. D., Nepal, K., Neupane, B., Baidhya, R., Baidya, R., et al. (2017). Antibiotic resistance and biofilm production among the strains of Staphylococcus aureus isolated from pus/wound swab samples in a tertiary care hospital in Nepal. Ann. Clin. Microbiol. Antimicrob. 16:15. doi: 10.1186/s12941-017-0194-0
Belete, M. A. (2020). Bacterial profile and ESBL screening of urinary tract infection among asymptomatic and symptomatic pregnant women attending antenatal Care of Northeastern Ethiopia Region. Infect Drug Resist. 13, 2579–2592. doi: 10.2147/IDR.S258379
Bensaci, M., and Sahm, D. (2017). Surveillance of tedizolid activity and resistance: in vitro susceptibility of gram-positive pathogens collected over 5 years from the United States and Europe. Diagn. Microbiol. Infect. Dis. 87, 133–138. doi: 10.1016/j.diagmicrobio.2016.10.009
Bhatt, P., Tandel, K., Singh, A., Mugunthan, M., Grover, N., and Sahni, A. K. (2016). Species distribution and antimicrobial resistance pattern of coagulase-negative staphylococci at a tertiary care Centre. Med. J. Armed Forces India 72, 71–74. doi: 10.1016/j.mjafi.2014.12.007
Bhattacharya, S., Pal, K., Jain, S., Chatterjee, S. S., and Konar, J. (2016). Surgical site infection by methicillin resistant Staphylococcus aureus- on decline? J. Clin. Diagn. Res. 10, DC32–DC36. doi: 10.7860/JCDR/2016/21664.8587
Bhavana, A. M., Kumari, P. H. P., Mohan, N., Chandrasekhar, V., Vijayalakshmi, P., and Manasa, R. V. (2019). Bacterial vaginosis and antibacterial susceptibility pattern of asymptomatic urinary tract infection in pregnant women at a tertiary care hospital, Visakhaptn, India. Iran J. Microbiol. 11, 488–495
Biset, S., Moges, F., Endalamaw, D., and Eshetie, S. (2020). Multi-drug resistant and extended-spectrum beta-lactamases producing bacterial uropathogens among pregnant women in Northwest Ethiopia. Ann. Clin. Microbiol. Antimicrob. 19:25. doi: 10.1186/s12941-020-00365-z
Bishr, A. S., Abdelaziz, S. M., Yahia, I. S., Yassien, M. A., Hassouna, N. A., and Aboshanab, K. M. (2021). Association of Macrolide Resistance Genotypes and Synergistic Antibiotic Combinations for combating macrolide-resistant MRSA recovered from hospitalized patients. Biology (Basel) 10. doi: 10.3390/biology10070624
Bolatchiev, A. (2020). Antibacterial activity of human defensins against Staphylococcus aureus and Escherichia coli. PeerJ 8:e10455. doi: 10.7717/peerj.10455
Boncompain, C. A., Suarez, C. A., Squeff, M., Belluzo, V., Piccirilli, G., Molteni, A., et al. (2023). Phenotypic and molecular characterization of Staphylococcus aureus isolates conducted in nares of psoriatic patients attending a public hospital in Argentina. Rev. Argent. Microbiol. 55, 3–11. doi: 10.1016/j.ram.2022.02.008
Cavalcante, F. S., Alvarenga, C., Saintive, S., Dios, E., Carvalho, D., and Netto, K. (2020). Staphylococcus aureus nasal isolates may have the same genetic profile in atopic dermatitis paediatric patients and their close contacts. J. Med. Microbiol. 69, 850–853. doi: 10.1099/jmm.0.001197
Cavanagh, J. P., Wolden, R., Heise, P., Esaiassen, E., Klingenberg, C., and Aarag Fredheim, E. G. (2016). Antimicrobial susceptibility and body site distribution of community isolates of coagulase-negative staphylococci. APMIS 124, 973–978. doi: 10.1111/apm.12591
Chaleshtori, S. H., and Kachoie, M. A. (2016). Chemical composition and antimicrobial effects of calendula officinalis grown under chemical and biological conditions on the methicillin-resistant staphylococcus aureus isolated from hospital infections. Biosci. Biotechnol. Res. Asia 13, 1787–1796. doi: 10.13005/bbra/2331
Changchien, C. H., Chen, S. W., Chen, Y. Y., and Chu, C. (2016). Antibiotic susceptibility and genomic variations in Staphylococcus aureus associated with skin and soft tissue infection (SSTI) disease groups. BMC Infect. Dis. 16:276. doi: 10.1186/s12879-016-1630-z
Chauhan, S., Surender,, and Rappai, T. (2021). Mupirocin resistance in staphylococcus aureus isolated from nasal swabs of ICU and OT staff- a study from a tertiary care hospital. J Pure Appl Microbiol 15, 2059–2064. doi: 10.22207/JPAM.15.4.28
Chelkeba, L., Fanta, K., Mulugeta, T., and Melaku, T. (2022). Bacterial profile and antimicrobial resistance patterns of common bacteria among pregnant women with bacteriuria in Ethiopia: a systematic review and meta-analysis. Arch. Gynecol. Obstet. 306, 663–686. doi: 10.1007/s00404-021-06365-4
Chelkeba, L., and Melaku, T. (2022). Epidemiology of staphylococci species and their antimicrobial-resistance among patients with wound infection in Ethiopia: a systematic review and meta-analysis. J. Glob. Antimicrobial Resist. 29, 483–498. doi: 10.1016/j.jgar.2021.10.025
Chen, P. Y., Chuang, Y. C., Wang, J. T., Sheng, W. H., Chen, Y. C., and Chang, S. C. (2021). Sequence type 8 as an emerging clone of methicillin-resistant Staphylococcus aureus causing bloodstream infections in Taiwan. Emerg Microbes Infect 10, 1908–1918. doi: 10.1080/22221751.2021.1981158
Chen, Y. L., Kang, E. Y., Yeh, L. K., Ma, D. H. K., Tan, H. Y., Chen, H. C., et al. (2021). Clinical features and molecular characteristics of methicillin-susceptible Staphylococcus aureus ocular infection in Taiwan. Antibiotics (Basel) 10. doi: 10.3390/antibiotics10121445
Chen, P., Sun, F., Feng, W., Hong, H., Li, B., and Song, J. (2021). Pathogenic characteristics Ofstaphylococcus aureusisolates from arthroplasty infections. Int. J. Artif. Organs 44, 208–214. doi: 10.1177/0391398820948877
Cheung, G. Y. C., Bae, J. S., and Otto, M. (2021). Pathogenicity and virulence of Staphylococcus aureus. Virulence 12, 547–569. doi: 10.1080/21505594.2021.1878688
Choi, E. Y., Han, J. Y., Lee, H., Lee, S. C., Koh, H. J., Kim, S. S., et al. (2019). Impact of antibiotic resistance of pathogens and early vitrectomy on the prognosis of infectious endophthalmitis: a 10-year retrospective study. Graefes Arch. Clin. Exp. Ophthalmol. 257, 805–813. doi: 10.1007/s00417-019-04261-x
Cochran, W. G. (1954). The combination of estimates from different experiments. Biometrics 10, 101–129. doi: 10.2307/3001666
Conceicao, T., de Lencastre, H., and Aires-de-Sousa, M. (2021). Prevalence of biocide resistance genes and chlorhexidine and mupirocin non-susceptibility in Portuguese hospitals during a 31-year period (1985-2016). J. Glob. Antimicrob. Resist. 24, 169–174. doi: 10.1016/j.jgar.2020.12.010
Coombs, G. W., Daley, D. A., Mowlaboccus, S., Lee, Y. T., and Pang, S.R. Australian Group on Antimicrobial (2020). Australian group on antimicrobial resistance (AGAR) Australian Staphylococcus aureus Sepsis outcome Programme (ASSOP) annual report 2018. Commun. Dis. Intell. 44:44. doi: 10.33321/cdi.2020.44.18
Dayie, N., Osei, M. M., Opintan, J. A., Tetteh-Quarcoo, P. B., Kotey, F. C. N., Ahenkorah, J., et al. (2021). Nasopharyngeal carriage and antimicrobial susceptibility profile of Staphylococcus aureus among children under five years in Accra. Pathogens 10. doi: 10.3390/pathogens10020136
de Benito, S., Alou, L., Becerro-de-Bengoa-Vallejo, R., Losa-Iglesias, M. E., Gomez-Lus, M. L., Collado, L., et al. (2018). "prevalence of Staphylococcus spp. nasal colonization among doctors of podiatric medicine and associated risk factors in Spain." Antimicrob resist. Infect. Control. 7:24. doi: 10.1186/s13756-018-0318-0
Demir, C., Demirci, M., Yigin, A., Tokman, H. B., and Cetik Yildiz, S. (2020). Presence of biofilm and adhesin genes in Staphylococcus aureus strains taken from chronic wound infections and their genotypic and phenotypic antimicrobial sensitivity patterns. Photodiagn. Photodyn. Ther. 29:101584. doi: 10.1016/j.pdpdt.2019.101584
Deyno, S., Fekadu, S., and Seyfe, S. (2018). Prevalence and antimicrobial resistance of coagulase negative staphylococci clinical isolates from Ethiopia: a meta-analysis. BMC Microbiol. 18:43. doi: 10.1186/s12866-018-1188-6
Dilnessa, T., and Bitew, A. (2016). Prevalence and antimicrobial susceptibility pattern of methicillin resistant Staphylococcus aureus isolated from clinical samples at Yekatit 12 hospital medical college, Addis Ababa, Ethiopia. BMC Infect. Dis. 16:398. doi: 10.1186/s12879-016-1742-5
Diriba, K., Kassa, T., Alemu, Y., and Bekele, S. (2020). In vitro biofilm formation and antibiotic susceptibility patterns of Bacteria from suspected external eye infected patients attending ophthalmology clinic, Southwest Ethiopia. Int J Microbiol 2020:8472395. doi: 10.1155/2020/8472395
Dormanesh, B., Siroosbakhat, S., Khodaverdi Darian, E., and Afsharkhas, L. (2015). Methicillin-resistant Staphylococcus aureus isolated from various types of hospital infections in pediatrics: Panton-valentine Leukocidin, staphylococcal chromosomal cassette mec SCCmec phenotypes and antibiotic resistance properties. Jundishapur J. Microbiol. 8:e11341. doi: 10.5812/jjm.11341
Doss, R. W., Mostafa, A. M. A., Arafa, A. E. E.-D., and Radi, N. A. E.-M. (2017). Relationship between lipase enzyme and antimicrobial susceptibility of Staphylococcus aureus-positive and Staphylococcus epidermidis-positive isolates from acne vulgaris. J. Egypt. Women Dermatol. Soc. 14, 167–172. doi: 10.1097/01.EWX.0000516051.01553.99
Duncan, L. R., Sader, H. S., Flamm, R. K., Jones, R. N., and Mendes, R. E. (2016). Oritavancin in vitro activity against contemporary Staphylococcus aureus isolates responsible for invasive community- and healthcare-associated infections among patients in the United States (2013-2014). Diagn. Microbiol. Infect. Dis. 86, 303–306. doi: 10.1016/j.diagmicrobio.2016.07.025
Eibach, D., Nagel, M., Hogan, B., Azuure, C., Krumkamp, R., Dekker, D., et al. (2017). Nasal carriage of Staphylococcus aureus among children in the Ashanti region of Ghana. PLoS One 12:e0170320. doi: 10.1371/journal.pone.0170320
El Mammery, A., Ramirez de Arellano, E., Canada-Garcia, J. E., Cercenado, E., Villar-Gomara, L., Casquero-Garcia, V., et al. (2023). An increase in erythromycin resistance in methicillin-susceptible Staphylococcus aureus from blood correlates with the use of macrolide/lincosamide/streptogramin antibiotics. EARS-net Spain (2004-2020). Front. Microbiol. 14:1220286. doi: 10.3389/fmicb.2023.1220286
El-Amir, M. I., El-Feky, M. A., Abo Elwafa, D. A., and Abd-Elmawgood, E. A. (2019). Rapid diagnosis of neonatal sepsis by PCR for detection of 16S rRNA gene, while blood culture and PCR results were similar in E.Coli-predominant EOS cases. Infect Drug Resist 12, 2703–2710. doi: 10.2147/IDR.S213958
El-Baghdady, K. Z., El-Borhamy, M. I., and Abd El-Ghafar, H. A. (2020). Prevalence of resistance and toxin genes in community-acquired and hospital-acquired methicillin-resistant Staphylococcus aureus clinical isolates. Iran. J. Basic Med. Sci. 23, 1251–1260. doi: 10.22038/ijbms.2020.40260.9534
El-Kersh, T. A., Marie, M. A., Al-Sheikh, Y. A., Al-Agamy, M. H., and Al Bloushy, A. A. (2016). Prevalence and risk factors of early fecal carriage of enterococcus faecalis and Staphylococcus spp and their antimicrobial resistant patterns among healthy neonates born in a hospital setting in Central Saudi Arabia. Saudi Med. J. 37, 280–287. doi: 10.15537/smj.2016.3.13871
Elzorkany, K. M. A., Elbrolosy, A. M., and Salem, E. H. (2019). Methicillin-resistant Staphylococcus aureus carriage in hemodialysis vicinity: prevalence and decolonization approach. Indian J Nephrol 29, 282–287. doi: 10.4103/ijn.IJN_56_18
Eshetie, S., Tarekegn, F., Moges, F., Amsalu, A., Birhan, W., and Huruy, K. (2016). Methicillin resistant Staphylococcus aureus in Ethiopia: a meta-analysis. BMC Infect. Dis. 16:689. doi: 10.1186/s12879-016-2014-0
Esmaeili Benvidi, M., Houri, H., Ghalavand, Z., Nikmanesh, B., Azimi, H., Samadi, R., et al. (2017). Toxin production and drug resistance profiles of pediatric methicillin-resistant Staphylococcus aureus isolates in Tehran. J. Infect. Dev. Ctries. 11, 759–765. doi: 10.3855/jidc.9360
Estany-Gestal, A., Salgado-Barreira, A., and Vazquez-Lago, J. M. (2024). Antibiotic use and antimicrobial resistance: a global public health crisis. Antibiotics (Basel) 13. doi: 10.3390/antibiotics13090900
Ezeh, C. K., Eze, C. N., Dibua, M. E. U., and Emencheta, S. C. (2023). A meta-analysis on the prevalence of resistance of Staphylococcus aureus to different antibiotics in Nigeria. Antimicrob. Resist. Infect. Control 12:40. doi: 10.1186/s13756-023-01243-x
Farah, S. M., Alshehri, M. A., Alfawaz, T. S., Alasmeri, F. A., Alageel, A. A., and Alshahrani, D. A. (2019). Trends in antimicrobial susceptibility patterns in king Fahad Medical City, Riyadh, Saudi Arabia. Saudi Med. J. 40, 252–259. doi: 10.15537/smj.2019.3.23947
Fasihi, Y., Saffari, F., Kandehkar Ghahraman, M. R., and Kalantar-Neyestanaki, D. (2016). Molecular detection of macrolide and Lincosamide-resistance genes in clinical methicillin-resistant Staphylococcus aureus isolates from Kerman, Iran. Archiv. Pediatr. Infect. Dis. 5:e37761. doi: 10.5812/pedinfect.37761
Fateh Amirkhiz, M., Ahangarzadeh Rezaee, M., Hasani, A., Aghazadeh, M., and Naghili, B. (2015). SCCmec typing of methicillin-resistant Staphylococcus aureus: An eight year experience. Archiv. Pediatr. Infect. Dis. 3:e30632. doi: 10.5812/pedinfect.30632
Fateh Dizji, P., Khosravy, M., Saeedi, A. A., Asli, M., Sepahvand, S., and Darvishi, M. (2023). Prevalence of clindamycin-resistant Staphylococcus aureus induced by macrolide resistance, Iran, 2019-2021. Iran. J. Med. Microbiol. 17, 256–261. doi: 10.30699/ijmm.17.2.256
Firoozeh, F., Omidi, M., Saffari, M., Sedaghat, H., and Zibaei, M. (2020). Molecular analysis of methicillin-resistant Staphylococcus aureus isolates from four teaching hospitals in Iran: the emergence of novel MRSA clones. Antimicrob. Resist. Infect. Control 9:112. doi: 10.1186/s13756-020-00777-8
Fu, Y., Xiong, M., Li, X., Zhou, J., Xiao, X., Fang, F., et al. (2020). Molecular characteristics, antimicrobial resistance and virulence gene profiles of Staphylococcus aureus isolates from Wuhan, Central China. Infect Drug Resist 13, 2063–2072. doi: 10.2147/IDR.S249988
Gajdacs, M., Abrok, M., Lazar, A., and Burian, K. (2021). Urinary tract infections in elderly patients: a 10-year study on their epidemiology and antibiotic resistance based on the WHO access, watch, reserve (AWaRe) classification. Antibiotics (Basel) 10:1098. doi: 10.3390/antibiotics10091098
Garza-Gonzalez, E., Morfin-Otero, R., Mendoza-Olazaran, S., Bocanegra-Ibarias, P., Flores-Trevino, S., Rodriguez-Noriega, E., et al. (2019). A snapshot of antimicrobial resistance in Mexico. Results from 47 centers from 20 states during a six-month period. PLoS One 14:e0209865. doi: 10.1371/journal.pone.0209865
Getaneh, A., Ayalew, G., Belete, D., Jemal, M., and Biset, S. (2021). Bacterial etiologies of ear infection and their antimicrobial susceptibility pattern at the University of Gondar Comprehensive Specialized Hospital, Gondar, Northwest Ethiopia: a six-year retrospective study. Infect Drug Resist 14, 4313–4322. doi: 10.2147/IDR.S332348
Gitau, W., Masika, M., Musyoki, M., Museve, B., and Mutwiri, T. (2018). Antimicrobial susceptibility pattern of Staphylococcus aureus isolates from clinical specimens at Kenyatta National Hospital. BMC. Res. Notes 11:226. doi: 10.1186/s13104-018-3337-2
Goes, I., Romero, L. C., Turra, A. J., Gotardi, M. A., Rodrigues, T., Santos, L. O., et al. (2021). Prevalence of nasal carriers of methicillin-resistant Staphylococcus aureus in primary health care units in Brazil. Rev. Inst. Med. Trop. Sao Paulo 63:e14. doi: 10.1590/s1678-9946202163014
Goudarzi, M., Abiri, P., Nasirian, S., and Afshari, S. G. (2018). SCCmec and spa typing of Staphylococcus aureus strains isolated from patients with urinary tract infection: emergence of spa types t426 and t021 in Iran. Jundishapur J. Microbiol. 11:e62169. doi: 10.5812/jjm.62169
Goudarzi, M., Tayebi, Z., Fazeli, M., Miri, M., and Nasiri, M. J. (2020). Molecular characterization, drug resistance and virulence analysis of constitutive and inducible clindamycin resistance Staphylococcus aureus strains recovered from clinical samples, Tehran—Iran. Infect Drug Resist 13, 1155–1162. doi: 10.2147/IDR.S251450
Gungor, S., Karagoz, A., Kocak, N., and Arslantas, T. (2021). Methicillin-resistant Staphylococcus aureus in a Turkish hospital: characterization of clonal types and antibiotic susceptibility. J. Infect. Dev. Ctries. 15, 1854–1860. doi: 10.3855/jidc.14963
Guo, Y., Ding, Y., Liu, L., Shen, X., Hao, Z., Duan, J., et al. (2019). Antimicrobial susceptibility, virulence determinants profiles and molecular characteristics of Staphylococcus epidermidis isolates in Wenzhou, eastern China. BMC Microbiol. 19:157. doi: 10.1186/s12866-019-1523-6
Guo, Y., Song, G., Sun, M., Wang, J., and Wang, Y. (2020). Prevalence and therapies of antibiotic-resistance in Staphylococcus aureus. Front. Cell. Infect. Microbiol. 10:107. doi: 10.3389/fcimb.2020.00107
Guo, Y., Wang, B., Rao, L., Wang, X., Zhao, H., Li, M., et al. (2021). Molecular characteristics of rifampin-sensitive and -resistant isolates and characteristics of rpoB gene mutations in methicillin-resistant Staphylococcus aureus. Infect Drug Resist 14, 4591–4600. doi: 10.2147/IDR.S336200
Hailegiyorgis, T. T., Sarhie, W. D., and Workie, H. M. (2018). Isolation and antimicrobial drug susceptibility pattern of bacterial pathogens from pediatric patients with otitis media in selected health institutions, Addis Ababa, Ethiopia: a prospective cross-sectional study. BMC Ear Nose Throat Disord 18:8. doi: 10.1186/s12901-018-0056-1
Hasanvand, H., Teymouri, F., Ohadi, E., Azadegan, A., and Sadeghi Kalani, B. (2019). Biofilm formation in Staphylococcus epidermidis isolated from hospitalized patients. Archiv. Clin. Infect. Dis. 14:e64496. doi: 10.5812/archcid.64496
Higgins, J. P., and Thompson, S. G. (2002). Quantifying heterogeneity in a meta-analysis. Stat. Med. 21, 1539–1558. doi: 10.1002/sim.1186
Hoffmann, K., den Heijer, C. D., George, A., Apfalter, P., and Maier, M. (2015). Prevalence and resistance patterns of commensal S. aureus in community-dwelling GP patients and socio-demographic associations. A cross-sectional study in the framework of the APRES-project in Austria. BMC Infect. Dis. 15:213. doi: 10.1186/s12879-015-0949-1
Horvath, A., Dobay, O., Sahin-Toth, J., Juhasz, E., Pongracz, J., Ivan, M., et al. (2020). Characterisation of antibiotic resistance, virulence, clonality and mortality in MRSA and MSSA bloodstream infections at a tertiary-level hospital in Hungary: a 6-year retrospective study. Ann. Clin. Microbiol. Antimicrob. 19:17. doi: 10.1186/s12941-020-00357-z
Ibadin, E. E., Enabulele, I. O., and Muinah, F. (2017). Prevalence of mecA gene among staphylococci from clinical samples of a tertiary hospital in Benin City, Nigeria. Afr. Health Sci. 17, 1000–1010. doi: 10.4314/ahs.v17i4.7
Ikuta, K., Swetschinski, L., Robles, G., Sharara, F., Mestrovic, T., Gray, A., et al. (2022). Global mortality associated with 33 bacterial pathogens in 2019: a systematic analysis for the global burden of disease study 2019. Lancet 400, 2221–2248. doi: 10.1016/S0140-6736(22)02185-7
Iliya, S., Mwangi, J., Maathai, R., and Muriuki, M. (2020). Phenotypic analysis and antibiotic susceptibility of methicillin-resistant Staphylococcus aureus in Kiambu County, Kenya. J. Infect. Dev. Ctries. 14, 597–605. doi: 10.3855/jidc.12174
Islam, T. A. B., and Shamsuzzaman, S. (2015). Prevalence and antimicrobial susceptibility pattern of methicillin-resistant, vancomycin-resistant, and Panton-valentine leukocidin positive Staphylococcus aureus in a tertiary care hospital Dhaka, Bangladesh. Tzu Chi Med. J. 27, 10–14. doi: 10.1016/j.tcmj.2014.12.001
Javidnia, S., Talebi, M., Katouli, M., Shojaie, A., Lari, A. R., and Pourshafie, M. R. (2015). Clonal diversity of meticillin-resistant staphylococcus aureus isolated from intensive care unit. Infect. Dis. Clin. Pract. 23, 128–130. doi: 10.1097/IPC.0000000000000230
Joachim, A., Moyo, S. J., Nkinda, L., Majigo, M., Mmbaga, E., Mbembati, N., et al. (2017). Prevalence of methicillin-resistant Staphylococcus aureus carriage on admission among patients attending regional hospitals in Dar Es Salaam, Tanzania. BMC. Res. Notes 10:417. doi: 10.1186/s13104-017-2668-8
John, B., Mabekoje Oladele, O., Aminat, H., Saba, M. A., Danasabe, D., Legbo, M. I., et al. (2023). Occurrence of Staphylococcus associated with urinary tract infections among women attending Ibrahim Badamasi Babangida (IBB) specialist hospital, Minna, Nigeria. Tanzan. J. Health Res. 24, 17–30. doi: 10.4314/thrb.v24i2
Juda, M., Chudzik-Rzad, B., and Malm, A. (2016). The prevalence of genotypes that determine resistance to macrolides, lincosamides, and streptogramins B compared with spiramycin susceptibility among erythromycin-resistant Staphylococcus epidermidis. Mem. Inst. Oswaldo Cruz 111, 155–160. doi: 10.1590/0074-02760150356
Junaidi, N. S. S. A., Shakrin, N. N. S. M., Huri, M. F. D., Kamarudin, A. Z., Desa, M. N. M., and Yunus, W. M. Z. W. (2023). Antibiotic resistance and molecular typing of clinical Staphylococcus aureus isolates from Malaysian military hospital. Asian Pac J Trop Med 16, 220–231. doi: 10.4103/1995-7645.377743
Kahsay, A. G., Hagos, D. G., Abay, G. K., and Mezgebo, T. A. (2018). Prevalence and antimicrobial susceptibility patterns of methicillin-resistant Staphylococcus aureus among janitors of Mekelle university, North Ethiopia. BMC. Res. Notes 11:294. doi: 10.1186/s13104-018-3399-1
Kang, S. H., and Kim, M. K. (2019). Antibiotic sensitivity and resistance of bacteria from odontogenic maxillofacial abscesses. J. Korean Assoc. Oral Maxillofac. Surg. 45, 324–331. doi: 10.5125/jkaoms.2019.45.6.324
Khader, K., Thomas, A., Jones, M., Toth, D., Stevens, V., Samore, M. H., et al. (2019). Variation and trends in transmission dynamics of methicillin-resistant Staphylococcus aureus in veterans affairs hospitals and nursing homes. Epidemics 28:100347. doi: 10.1016/j.epidem.2019.100347
Khan, F. Y., Abu-Khattab, M., Almaslamani, E. A., Hassan, A. A., Mohamed, S. F., Elbuzdi, A. A., et al. (2017). Acute bacterial meningitis in Qatar: a hospital-based study from 2009 to 2013. Biomed. Res. Int. 2017:2975610. doi: 10.1155/2017/2975610
Khan, S., Singh, P., Siddiqui, Z., and Ansari, M. (2015). Pregnancy-associated asymptomatic bacteriuria and drug resistance. J. Taibah Univ. Med. Sci. 10, 340–345. doi: 10.1016/j.jtumed.2015.01.011
Khanal, A., Sulochan, G. C., Gaire, A., Khanal, A., Estrada, R., Ghimire, R., et al. (2021). Methicillin-resistant Staphylococcus aureus in Nepal: a systematic review and meta-analysis. Int. J. Infect. Dis. 103, 48–55. doi: 10.1016/j.ijid.2020.11.152
Khemiri, M., Akrout Alhusain, A., Abbassi, M. S., El Ghaieb, H., Santos Costa, S., Belas, A., et al. (2017). Clonal spread of methicillin-resistant Staphylococcus aureus-t6065-CC5-SCCmecV-agrII in a Libyan hospital. J. Glob. Antimicrob. Resist. 10, 101–105. doi: 10.1016/j.jgar.2017.04.014
Kim, H. J., Choi, Q., Kwon, G. C., and Koo, S. H. (2020). Molecular epidemiology and virulence factors of methicillin-resistant Staphylococcus aureus isolated from patients with bacteremia. J. Clin. Lab. Anal. 34:e23077. doi: 10.1002/jcla.23077
Kong, Y., Ye, J., Zhou, W., Jiang, Y., Lin, H., Zhang, X., et al. (2018). Prevalence of methicillin-resistant Staphylococcus aureus colonisation among healthcare workers at a tertiary care hospital in southeastern China. J. Glob. Antimicrob. Resist. 15, 256–261. doi: 10.1016/j.jgar.2018.08.013
Kpeli, G., Darko Otchere, I., Lamelas, A., Buultjens, A. L., Bulach, D., Baines, S. L., et al. (2016). Possible healthcare-associated transmission as a cause of secondary infection and population structure of Staphylococcus aureus isolates from two wound treatment centres in Ghana. New Microb. New Infect. 13, 92–101. doi: 10.1016/j.nmni.2016.07.001
Kuhn, M., Wing, J., Weston, S., Williams, A., Keefer, C., Engelhardt, A., et al. (2015). Caret: Classification and regression training. (Version 7.0-1) [Computer software]. Available at: https://github.com/topepo/caret/
Kulshrestha, N., Ghatak, T., Gupta, P., Singh, M., Agarwal, J., and Mishra, P. (2021). Surveillance of health-care workers hand to detect carriage of multidrug-resistant Staphylococcus spp. in a tertiary care center: An observational study. Med. J. Dr. D.Y. Patil Vidyapeeth 14, 403–408. doi: 10.4103/mjdrdypu.mjdrdypu_372_20
Kumar, S., and Shetty, V. A. (2021). Prevalence and susceptibility profiles of methicillin sensitive Staphylococcus aureus from community and hospital associated infections. J. Clin. Diagn. Res. 15:5. doi: 10.7860/JCDR/2021/48115.14622
Kumar, R. A., Thirugnanamani, R., Dodeja, S., and Satish, H. S. (2018). Bacterial profile and antibiotic sensitivity in patients with chronic rhinosinusitis undergoing functional endoscopic sinus surgery: a prospective study. Int. J. Clin. Rhinol. 10, 137–141. doi: 10.5005/jp-journals-10013-1325
Kurup, R., and Ansari, A. A. (2019). A study to identify bacteriological profile and other risk factors among diabetic and non-diabetic foot ulcer patients in a Guyanese hospital setting. Diabetes Metab. Syndr. 13, 1871–1876. doi: 10.1016/j.dsx.2019.04.024
Lan, T., Zhang, B., Liu, J. L., Jia, Q., Gao, J., Cao, L., et al. (2024). Prevalence and antibiotic resistance patterns of methicillin-resistant Staphylococcus aureus (MRSA) in a hospital setting: a retrospective study from 2018 to 2022. Indian J. Microbiol. 64, 1035–1043. doi: 10.1007/s12088-024-01228-3
Larsen, J., Petersen, A., Sorum, M., Stegger, M., van Alphen, L., Valentiner-Branth, P., et al. (2015). Meticillin-resistant Staphylococcus aureus CC398 is an increasing cause of disease in people with no livestock contact in Denmark, 1999 to 2011. Euro Surveill. 20. doi: 10.2807/1560-7917.ES.2015.20.37.30021
Leclercq, R. (2002). Mechanisms of resistance to macrolides and lincosamides: nature of the resistance elements and their clinical implications. Clin. Infect. Dis. 34, 482–492. doi: 10.1086/324626
Lee, Y. C., Chen, P. Y., Wang, J. T., and Chang, S. C. (2020). Prevalence of fosfomycin resistance and gene mutations in clinical isolates of methicillin-resistant Staphylococcus aureus. Antimicrob. Resist. Infect. Control 9:135. doi: 10.1186/s13756-020-00790-x
Lee, S. O., Lee, S., Park, S., Lee, J. E., and Lee, S. H. (2019). The cefazolin inoculum effect and the presence of type a blaZ gene according to agr genotype in methicillin-susceptible Staphylococcus aureus bacteremia. Infect Chemother. 51, 376–385. doi: 10.3947/ic.2019.51.4.376
Leibler, J. H., Leon, C., Cardoso, L. J. P., Morris, J. C., Miller, N. S., Nguyen, D. D., et al. (2017). Prevalence and risk factors for MRSA nasal colonization among persons experiencing homelessness in Boston, MA. J. Med. Microbiol. 66, 1183–1188. doi: 10.1099/jmm.0.000552
Lenart-Boron, A., Wolny-Koladka, K., Stec, J., and Kasprowic, A. (2016). Phenotypic and molecular antibiotic resistance determination of airborne coagulase negative Staphylococcus spp. strains from healthcare facilities in southern Poland. Microb. Drug Resist. 22, 515–522. doi: 10.1089/mdr.2015.0271
Lennartz, F. E., Schwartbeck, B., Dubbers, A., Grosse-Onnebrink, J., Kessler, C., Kuster, P., et al. (2019). The prevalence of Staphylococcus aureus with mucoid phenotype in the airways of patients with cystic fibrosis-a prospective study. Int. J. Med. Microbiol. 309, 283–287. doi: 10.1016/j.ijmm.2019.05.002
Li, S., Guo, Y., Zhao, C., Chen, H., Hu, B., Chu, Y., et al. (2016). In vitro activities of tedizolid compared with other antibiotics against gram-positive pathogens associated with hospital-acquired pneumonia, skin and soft tissue infection and bloodstream infection collected from 26 hospitals in China. J. Med. Microbiol. 65, 1215–1224. doi: 10.1099/jmm.0.000347
Li, S., Han, Z., He, J., Gao, S., Liu, D., Liu, L., et al. (2018). Society for Translational Medicine expert consensus on the use of antibacterial drugs in thoracic surgery. J. Thorac. Dis. 10, 6356–6374. doi: 10.21037/jtd.2018.10.108
Liang, J., Hu, Y., Fu, M., Li, N., Wang, F., Yu, X., et al. (2023). Resistance and molecular characteristics of methicillin-resistant Staphylococcus aureus and heterogeneous vancomycin-intermediate Staphylococcus aureus. Infect Drug Resist 16, 379–388. doi: 10.2147/IDR.S392908
Liang, B., Mai, J., Liu, Y., Huang, Y., Zhong, H., Xie, Y., et al. (2018). Prevalence and characterization of Staphylococcus aureus isolated from women and children in Guangzhou, China. Front. Microbiol. 9:2790. doi: 10.3389/fmicb.2018.02790
Lin, J., Wu, C., Yan, C., Ou, Q., Lin, D., Zhou, J., et al. (2018). A prospective cohort study of Staphylococcus aureus and methicillin-resistant Staphylococcus aureus carriage in neonates: the role of maternal carriage and phenotypic and molecular characteristics. Infect Drug Resist 11, 555–565. doi: 10.2147/IDR.S157522
Liu, C., Chen, Z. J., Sun, Z., Feng, X., Zou, M., Cao, W., et al. (2015). Molecular characteristics and virulence factors in methicillin-susceptible, resistant, and heterogeneous vancomycin-intermediate Staphylococcus aureus from Central-Southern China. J. Microbiol. Immunol. Infect. 48, 490–496. doi: 10.1016/j.jmii.2014.03.003
Livermore, D. M., Mushtaq, S., Warner, M., James, D., Kearns, A., and Woodford, N. (2015). Pathogens of skin and skin-structure infections in the UK and their susceptibility to antibiotics, including ceftaroline. J. Antimicrob. Chemother. 70, 2844–2853. doi: 10.1093/jac/dkv179
Lodise, T. P. Jr., and McKinnon, P. S. (2007). Burden of methicillin-resistant Staphylococcus aureus: focus on clinical and economic outcomes. Pharmacotherapy 27, 1001–1012. doi: 10.1592/phco.27.7.1001
Luo, Z. G., Ying, X. R., Shen, C., Ren, Y., Wang, S. B., and Wu, G. F. (2020). Characteristics and drug resistance of pathogens in urinary tract infection patients complicated with urinary calculi. Indian J. Pharm. Sci. 82, 922–927.
Mahfouz, A. A., Said, H. S., Elfeky, S. M., and Shaaban, M. I. (2023). Inhibition of erythromycin and erythromycin-induced resistance among Staphylococcus aureus clinical isolates. Antibiotics (Basel) 12. doi: 10.3390/antibiotics12030503
Maina, D., Omuse, G., Revathi, G., and Adam, R. D. (2016). Spectrum of microbial diseases and resistance patterns at a private teaching Hospital in Kenya: implications for clinical practice. PLoS One 11:e0147659. doi: 10.1371/journal.pone.0147659
Maleki, D. T., Ghalavand, Z., Laabei, M., Nikmanesh, B., Houri, H., Kodori, M., et al. (2019). Molecular analysis of accessory gene regulator functionality and virulence genes in Staphylococcus aureus derived from pediatric wound infections. Infect. Genet. Evol. 73, 255–260. doi: 10.1016/j.meegid.2019.05.013
Mama, M., Aklilu, A., Misgna, K., Tadesse, M., and Alemayehu, E. (2019). Methicillin- and inducible clindamycin-resistant Staphylococcus aureus among patients with wound infection attending Arba Minch hospital, South Ethiopia. Int J Microbiol 2019:2965490. doi: 10.1155/2019/2965490
Manandhar, S., Shrestha, R., Tuladhar, R. S., and Lekhak, S. (2021). Inducible clindamycin resistance and biofilm production among staphylococci isolated from tertiary care hospitals in Nepal. Infect Dis. Rep. 13, 1043–1052. doi: 10.3390/idr13040095
Mansson, E., Hellmark, B., Sundqvist, M., and Soderquist, B. (2015). Sequence types of Staphylococcus epidermidis associated with prosthetic joint infections are not present in the laminar airflow during prosthetic joint surgery. APMIS 123, 589–595. doi: 10.1111/apm.12392
Mascaro, V., Capano, M. S., Iona, T., Nobile, C. G. A., Ammendolia, A., and Pavia, M. (2019). Prevalence of Staphylococcus aureus carriage and pattern of antibiotic resistance, including methicillin resistance, among contact sport athletes in Italy. Infect Drug Resist. 12, 1161–1170. doi: 10.2147/IDR.S195749
McHardy, I. H., Veltman, J., Hindler, J., Bruxvoort, K., Carvalho, M. M., and Humphries, R. M. (2017). Clinical and microbiological aspects of beta-lactam resistance in Staphylococcus lugdunensis. J. Clin. Microbiol. 55, 585–595. doi: 10.1128/JCM.02092-16
Mehreen, A., Liaqat, I., Arshad, M., Waheed, M., and Arshad, N. (2018). Characterization of Staphylococcus aureus from sore throat patients: association among host immune evasion and toxin genes. Pak. J. Zool. 50. doi: 10.17582/journal.pjz/2018.50.6.2261.2272
Mesbah Elkammoshi, A., Ghasemzadeh-Moghaddam, H., Amin Nordin, S., Mohd Taib, N., Kumar Subbiah, S., Neela, V., et al. (2016). A low prevalence of inducible macrolide, Lincosamide, and Streptogramin B resistance phenotype among methicillin-susceptible Staphylococcus aureus isolated from Malaysian patients and healthy individuals. Jundishapur J. Microbiol. 9:e37148
Miklasinska-Majdanik, M. (2021). Mechanisms of resistance to macrolide antibiotics among Staphylococcus aureus. Antibiotics (Basel) 10. doi: 10.3390/antibiotics10111406
Modukuru, G. K., Surya, P. M. S., Kakumanu, V. R., and Yarava, S. (2021). Phenotypic characterization of macrolide-Lincosamide-Streptogramin B resistance in Staphylococcus aureus. J. Pure Appl. Microbiol. 15, 689–694. doi: 10.22207/JPAM.15.2.18
Mostafa, M., Siadat, S. D., Shahcheraghi, F., Vaziri, F., Japoni-Nejad, A., Vand Yousefi, J., et al. (2015). Variability in gene cassette patterns of class 1 and 2 integrons associated with multi drug resistance patterns in Staphylococcus aureus clinical isolates in Tehran-Iran. BMC Microbiol. 15:152. doi: 10.1186/s12866-015-0488-3
Mottola, C., Matias, C. S., Mendes, J. J., Melo-Cristino, J., Tavares, L., Cavaco-Silva, P., et al. (2016). Susceptibility patterns of Staphylococcus aureus biofilms in diabetic foot infections. BMC Microbiol. 16:119. doi: 10.1186/s12866-016-0737-0
Muhammad, A., Khan, S. N., Ali, N., Rehman, M. U., and Ali, I. (2020). Prevalence and antibiotic susceptibility pattern of uropathogens in outpatients at a tertiary care hospital. New Microbes New Infect 36:100716. doi: 10.1016/j.nmni.2020.100716
Murugesan, S., Perumal, N., Mahalingam, S. P., Dilliappan, S. K., and Krishnan, P. (2015). Analysis of antibiotic resistance genes and its associated SCCmec types among nasal carriage of methicillin resistant coagulase negative staphylococci from community settings, Chennai, southern India. J. Clin. Diagn. Res. 9:DC01-05. doi: 10.7860/JCDR/2015/11733.6307
Mutonga, D. M., Mureithi, M. W., Ngugi, N. N., and Otieno, F. C. F. (2019). Bacterial isolation and antibiotic susceptibility from diabetic foot ulcers in Kenya using microbiological tests and comparison with RT-PCR in detection of S. Aureus and MRSA. BMC. Res. Notes 12:244. doi: 10.1186/s13104-019-4278-0
Naghavi, M., Vollset, S., Ikuta, K., Swetschinski, L., Gray, A., Wool, E., et al. (2024). Global burden of bacterial antimicrobial resistance 1990-2021: a systematic analysis with forecasts to 2050. Lancet 404, 1199–1226. doi: 10.1016/S0140-6736(24)01867-1
Napp, M., Daeschlein, G., von Podewils, S., Hinz, P., Emmert, S., Haase, H., et al. (2016). In vitro susceptibility of methicillin-resistant and methicillin-susceptible strains of Staphylococcus aureus to two different cold at plasma sources. Infection 44, 531–537. doi: 10.1007/s15010-016-0888-9
Nasirian, S., Saadatmand, S., Goudarzi, H., Goudarzi, M., and Azimi, H. (2018). Molecular investigation of methicillin-resistant Staphylococcus aureus strains recovered from the intensive care unit (ICU) based on toxin, adhesion genes and agr locus type analysis. Arch. Clin. Infect. Dis. 13:e14495.
Nichol, K. A., Adam, H. J., Golding, G. R., Lagace-Wiens, P. R. S., Karlowsky, J. A., Hoban, D. J., et al. (2019). Characterization of MRSA in Canada from 2007 to 2016. J. Antimicrob. Chemother. 74, iv55–iv63. doi: 10.1093/jac/dkz288
Noordin, A., Sapri, H. F., Mohamad Sani, N. A., Leong, S. K., Tan, X. E., Tan, T. L., et al. (2016). Antimicrobial resistance profiling and molecular typing of methicillin-resistant Staphylococcus aureus isolated from a Malaysian teaching hospital. J. Med. Microbiol. 65, 1476–1481. doi: 10.1099/jmm.0.000387
Numanovic, F., Dermota, U., Smajlovic, J., Janezic, S., Tihic, N., Delibegovic, Z., et al. (2021). Characterization and clonal representation of MRSA strains in Tuzla Canton, Bosnia and Herzegovina, from 2009 to 2017. Med. Glas. (Zenica) 18, 38–46. doi: 10.17392/1265-21
Okuda, K. V., Toepfner, N., Alabi, A. S., Arnold, B., Belard, S., Falke, U., et al. (2016). Molecular epidemiology of Staphylococcus aureus from Lambarene, Gabon. Eur. J. Clin. Microbiol. Infect. Dis. 35, 1963–1973. doi: 10.1007/s10096-016-2748-z
Olufunmiso, O., Tolulope, I., and Roger, C. (2017). Multidrug and vancomycin resistance among clinical isolates of Staphylococcus aureus from different teaching hospitals in Nigeria. Afr. Health Sci. 17, 797–807. doi: 10.4314/ahs.v17i3.23
Ouidri, M. A. (2018). Screening of nasal carriage of methicillin-resistant Staphylococcus aureus during admission of patients to Frantz fanon hospital, Blida, Algeria. New Microbes New Infect 23, 52–60. doi: 10.1016/j.nmni.2018.02.006
Oydanich, M., Dingle, T. C., Hamula, C. L., Ghisa, C., and Asbell, P. (2017). Retrospective report of antimicrobial susceptibility observed in bacterial pathogens isolated from ocular samples at Mount Sinai hospital, 2010 to 2015. Antimicrob. Resist. Infect. Control 6:29. doi: 10.1186/s13756-017-0185-0
Parastan, R., Kargar, M., Solhjoo, K., and Kafilzadeh, F. (2020). A synergistic association between adhesion-related genes and multidrug resistance patterns of Staphylococcus aureus isolates from different patients and healthy individuals. J. Glob. Antimicrob. Resist. 22, 379–385. doi: 10.1016/j.jgar.2020.02.025
Peng, X., Zhu, Q., Liu, J., Zeng, M., Qiu, Y., Zhu, C., et al. (2021). Prevalence and antimicrobial resistance patterns of bacteria isolated from cerebrospinal fluid among children with bacterial meningitis in China from 2016 to 2018: a multicenter retrospective study. Antimicrob. Resist. Infect. Control 10:24. doi: 10.1186/s13756-021-00895-x
Peterside, O., Pondei, K., and Akinbami, F. O. (2015). Bacteriological profile and antibiotic susceptibility pattern of neonatal Sepsis at a teaching Hospital in Bayelsa State, Nigeria. Trop. Med. Health 43, 183–190. doi: 10.2149/tmh.2015-03
Petrović, J. L., Kuljić, K. N., Ristanović, E., Jošić, D., and Lepšanović, Z. (2016). Prevalence of Panton-valentine leukocidin genes in community-associated methicillin-resistant Staphylococcus aureus in the district of Pomoravlje. Vojnosanit. Pregl. 73, 256–260. doi: 10.2298/VSP140715003P
Pfaller, M. A., Huband, M. D., Shortridge, D., and Flamm, R. K. (2020). Surveillance of Omadacycline activity tested against clinical isolates from the United States and Europe: report from the SENTRY antimicrobial surveillance program, 2016 to 2018. Antimicrob. Agents Chemother. 64:e02488–19. doi: 10.1128/AAC.02488-19
Pradhan, P., Rajbhandari, P., Nagaraja, S. B., Shrestha, P., Grigoryan, R., Satyanarayana, S., et al. (2021). Prevalence of methicillin-resistant Staphylococcus aureus in a tertiary hospital in Nepal. Public Health Action 11, 46–51. doi: 10.5588/pha.21.0042
Preeja, P. P., Kumar, S. H., and Shetty, V. (2021). Prevalence and characterization of methicillin-resistant Staphylococcus aureus from community- and hospital-associated infections: a tertiary care center study. Antibiotics (Basel) 10:197. doi: 10.3390/antibiotics10020197
Pushkar, Aashana, Sharma, M., and Yadav, A. (2022). "Prevalence and antimicrobial resistance of methicillin resistant Staphylococcus aureus (MRSA) isolated from blood culture in tertiary care hospital in Haryana." Pravara. Med. Rev. 14, 64–68. doi: 10.36848/PMR/2022/99100.51095
Qin, Y., Wen, F., Zheng, Y., Zhao, R., Hu, Q., and Zhang, R. (2017). Antimicrobial resistance and molecular characteristics of methicillin-resistant Staphylococcus aureus isolates from child patients of high-risk wards in Shenzhen, China. Jpn. J. Infect. Dis. 70, 479–484. doi: 10.7883/yoken.JJID.2016.328
Rahimi, F. (2016). Characterization of resistance to aminoglycosides in methicillin-resistant Staphylococcus aureus strains isolated from a tertiary Care Hospital in Tehran, Iran. Jundishapur J. Microbiol. 9:e29237. doi: 10.5812/jjm.29237
Rajkumar, S., Sistla, S., Manoharan, M., Sugumar, M., Nagasundaram, N., Parija, S. C., et al. (2017). Prevalence and genetic mechanisms of antimicrobial resistance in Staphylococcus species: a multicentre report of the indian council of medical research antimicrobial resistance surveillance network. Indian J. Med. Microbiol. 35, 53–60. doi: 10.4103/ijmm.IJMM_16_427
Ramakrishna, M. S., Jeyamani, L., Abimannan, G. C., and Vajravelu, L. K. (2021). Microbial profile and Antibiogram pattern analysis of skin and soft tissue infections at a tertiary Care Center in South India. J. Pure Appl. Microbiol. 15, 915–925. doi: 10.22207/JPAM.15.2.50
Rampelotto, R. F., Coelho, S. S., Franco, L. N., Mota, A. D. D., Calegari, L. F., Jacobi, L. F., et al. (2022). Coagulase-negative staphylococci isolates from blood cultures of newborns in a tertiary hospital in southern Brazil. Braz. J. Pharm. Sci. 58:e19664. doi: 10.1590/s2175-97902022e19664
Raut, S., Bajracharya, K., Adhikari, J., Pant, S. S., and Adhikari, B. (2017). Prevalence of methicillin resistant Staphylococcus aureus in Lumbini medical college and teaching hospital, Palpa, Western Nepal. BMC. Res. Notes 10:187. doi: 10.1186/s13104-017-2515-y
Roden, L., Gorlich, D., Omran, H., Peters, G., Grosse-Onnebrink, J., and Kahl, B. C. (2019). A retrospective analysis of the pathogens in the airways of patients with primary ciliary dyskinesia. Respir. Med. 156, 69–77. doi: 10.1016/j.rmed.2019.08.009
Rukan, M., Jamil, H., Bokhari, H. A., Khattak, A. A., Khan, A. N., Ullah, Z., et al. (2021). Nasal carriage of highly resistant methicillin resistant Staphylococcus aureus (MRSA) strains by hospital staff in Hazara region of Pakistan. J. Pak. Med. Assoc. 71, 47–50. doi: 10.47391/JPMA.177
Saini, V., Jain, C., Singh, N. P., Alsulimani, A., Gupta, C., Dar, S. A., et al. (2021). Paradigm shift in antimicrobial resistance pattern of bacterial isolates during the COVID-19 pandemic. Antibiotics (Basel) 10:954. doi: 10.3390/antibiotics10080954
Sakabe, D., and Del Fiol Fde, S. (2016). Profile of infections and antimicrobial treatment among burn-injury patients. Am. J. Infect. Control 44, 950–952. doi: 10.1016/j.ajic.2016.03.063
Salah, A., Al-Subol, I., Hudna, A., Alhaj, A., Alqubaty, A. R., Farie, W., et al. (2021). Neonatal sepsis in Sana'a city, Yemen: a predominance of Burkholderia cepacia. BMC Infect. Dis. 21:1108. doi: 10.1186/s12879-021-06808-y
Salarvand, S., Abdollahi, A., Doraghi, M., Miratashi Yazdi, S. A., Panahi, Z., Mortazavi, S. M. J., et al. (2023). Microbiological profile and drug resistance in bone and joint infections: a survey in orthopedic wards of a great referral Hospital in Tehran, Iran. Jundishapur J. Microbiol. 16:e137125. doi: 10.5812/jjm-137125
Saleem, M., Ahmad, I., Salem, A. M., Almarshedy, S. M., Moursi, S. A., Syed Khaja, A. S., et al. (2025). Molecular and genetic analysis of methicillin-resistant Staphylococcus aureus (MRSA) in a tertiary care hospital in Saudi Arabia. Naunyn. Schmiedebergs Arch. Pharmacol. doi: 10.1007/s00210-024-03771-8
Sanchez, A., Benito, N., Rivera, A., Garcia, L., Miro, E., Mur, I., et al. (2020). Pathogenesis of Staphylococcus epidermidis in prosthetic joint infections: can identification of virulence genes differentiate between infecting and commensal strains? J. Hosp. Infect. 105, 561–568. doi: 10.1016/j.jhin.2020.04.026
Sapkota, J., Sharma, M., Jha, B., and Bhatt, C. P. (2019). Prevalence of Staphylococcus aureus isolated from clinical samples in a tertiary care hospital: a descriptive cross-sectional study. JNMA J. Nepal Med. Assoc. 57, 398–402. doi: 10.31729/jnma.4673
Saxena, S., Priyadarshi, M., Saxena, A., and Singh, R. (2019). Antimicrobial consumption and bacterial resistance pattern in patients admitted in I.C.U at a tertiary care center. J. Infect. Public Health 12, 695–699. doi: 10.1016/j.jiph.2019.03.014
Selim, S., Faried, O. A., Almuhayawi, M. S., Saleh, F. M., Sharaf, M., El Nahhas, N., et al. (2022). Incidence of vancomycin-resistant Staphylococcus aureus strains among patients with urinary tract infections. Antibiotics (Basel) 11:408. doi: 10.3390/antibiotics11030408
Shashindran, N., Nagasundaram, N., Thappa, D. M., and Sistla, S. (2016). Can Panton valentine Leukocidin gene and clindamycin susceptibility serve as predictors of community origin of MRSA from skin and soft tissue infections? J. Clin. Diagn. Res. 10, DC01–DC04. doi: 10.7860/JCDR/2016/14531.7036
Sheeba, V., Vedachalam, D., and Affan, T. F. (2021). An increasing trend in the antimicrobial resistance of bacterial isolates from skin and soft tissue infections in a tertiary care hospital. J. Pure Appl. Microbiol. 15, 803–812. doi: 10.22207/JPAM.15.2.34
Shidiki, A., Pandit, B., and Vyas, A. (2018). Incidence and antibiotic profile of bacterial isolates from neonatal septicemia in national medical college and teaching hospital, Birgunj, Nepal. Res. J. Pharm. Technol. 11, 2238–2242. doi: 10.5958/0974-360X.2018.00414.6
Shittu, A. O., Oyedara, O., Okon, K., Raji, A., Peters, G., von Muller, L., et al. (2015). An assessment on DNA microarray and sequence-based methods for the characterization of methicillin-susceptible Staphylococcus aureus from Nigeria. Front. Microbiol. 6:1160. doi: 10.3389/fmicb.2015.01160
Shivappa, S. G., Morubagal, R. R., Mahale, R. P., and Gowda, R. S. (2018). Prevalence and Antibiogram of methicillin sensitive and methicillin resistant Staphylococcus aureus isolated from pus samples in a tertiary care teaching hospital. J. Pure Appl. Microbiol. 12, 2297–2303. doi: 10.22207/JPAM.12.4.71
Singh, N., and Hota, S., S. snigdha Panda, D. Pattnaik, A. Praharaj and J. Jena (2019). "Prevalence and antibiotic resistance profile of coagulase negative staphylococci causing true Bacteraemia in a tertiary care hospital." Indian J. Public Health 10:479. doi: 10.5958/0976-5506.2019.02474.4
Skender, K., Machowska, A., Singh, V., Goel, V., Marothi, Y., Lundborg, C. S., et al. (2022). Antibiotic use, incidence and risk factors for orthopedic surgical site infections in a teaching Hospital in Madhya Pradesh, India. Antibiotics (Basel) 11:748. doi: 10.3390/antibiotics11060748
Solomon, J. G., and Salaudeen, A. G. (2021). Antibiotics resistance, sensitivity pattern and development of antibiogram to support empirical prescription in health facilities in south senatorial district of Kwara state, Nigeria 9, 35–45. doi: 10.21522/TIJPH.2013.09.03.Art004
Soroush, S., Jabalameli, F., Taherikalani, M., Amirmozafari, N., Fooladi, A. A., Asadollahi, K., et al. (2016). Investigation of biofilm formation ability, antimicrobial resistance and the staphylococcal cassette chromosome mec patterns of methicillin resistant Staphylococcus epidermidis with different sequence types isolated from children. Microb. Pathog. 93, 126–130. doi: 10.1016/j.micpath.2016.01.018
Sotoudeh Anvari, M., Kianinejad, R., Boroumand, M. A., Arzhan, S., and Jalali, A. (2015). Bacterial pericarditis and antimicrobial resistance at the Tehran Heart Center, Iran. J. Infect. Dev. Ctries. 9, 780–784. doi: 10.3855/jidc.6027
Soumya, K. R., Philip, S., Sugathan, S., Mathew, J., and Radhakrishnan, E. K. (2017). Virulence factors associated with Coagulase Negative Staphylococci isolated from human infections. 3 Biotech 7:140. doi: 10.1007/s13205-017-0753-2
Sterne, J. A. C., and Egger, M. (2005). Regression methods to detect publication and other Bias in Meta-analysis. Public. Bias Meta Analysis, 99–110. doi: 10.1002/0470870168.ch6
Sultan, A., Rizvi, M., Khan, F., Sami, H., Shukla, I., and Khan, H. M. (2015). Increasing antimicrobial resistance among uropathogens: is fosfomycin the answer? Urol. Ann. 7, 26–30. doi: 10.4103/0974-7796.148585
Suneel Kumar, A., Smiline Girija, A. S., and Naga Srilatha, B. (2021). Characterization of biofilm producing methicillin resistant coagulase negative staphylococci from India. Acta Microbiol. Immunol. Hung. 69, 35–40. doi: 10.1556/030.2021.01538
Sutter, D. E., Milburn, E., Chukwuma, U., Dzialowy, N., Maranich, A. M., and Hospenthal, D. R. (2016). Changing susceptibility of Staphylococcus aureus in a US pediatric population. Pediatrics 137: e20153099. doi: 10.1542/peds.2015-3099
Svent-Kucina, N., Pirs, M., Kofol, R., Blagus, R., Smrke, D. M., Bilban, M., et al. (2016). Molecular characterization of Staphylococcus aureus isolates from skin and soft tissue infections samples and healthy carriers in the Central Slovenia region. APMIS 124, 309–318. doi: 10.1111/apm.12509
Taha, L., Stegger, M., and Soderquist, B. (2019). Staphylococcus lugdunensis: antimicrobial susceptibility and optimal treatment options. Eur. J. Clin. Microbiol. Infect. Dis. 38, 1449–1455. doi: 10.1007/s10096-019-03571-6
Tahbaz, S. V., Azimi, L., Nowroozi, J., Armin, S., and Fallah, F. (2019). Multilocus sequence typing and antibiotic resistant patterns of the meticillin-resistant Staphylococcus aureus isolates from different clinical specimens. Rev. Res. Med. Microbiol. 30, 77–82. doi: 10.1097/MRM.0000000000000176
Taherirad, A., Jahanbakhsh, R., Shakeri, F., Anvary, S., and Ghaemi, E. A. (2016). Staphylococcal cassette chromosome mec types among methicillin-resistant Staphylococcus aureus in northern Iran. Jundishapur J. Microbiol. 9:e33933
Talapan, D., Sandu, A. M., and Rafila, A. (2023). Antimicrobial resistance of Staphylococcus aureus isolated between 2017 and 2022 from infections at a tertiary Care Hospital in Romania. Antibiotics (Basel) 12:974. doi: 10.3390/antibiotics12060974
Tang, B., Gong, T., Cui, Y., Wang, L., He, C., Lu, M., et al. (2020). Characteristics of oral methicillin-resistant Staphylococcus epidermidis isolated from dental plaque. Int. J. Oral Sci. 12:15. doi: 10.1038/s41368-020-0079-5
Tekeli, A., Ocal, D. N., Ozmen, B. B., Karahan, Z. C., and Dolapci, I. (2016). Molecular characterization of methicillin-resistant Staphylococcus aureus bloodstream isolates in a Turkish university hospital between 2002 and 2012. Microb. Drug Resist. 22, 564–569. doi: 10.1089/mdr.2015.0116
Tong, S. Y., Davis, J. S., Eichenberger, E., Holland, T. L., and Fowler, V. G. Jr. (2015). Staphylococcus aureus infections: epidemiology, pathophysiology, clinical manifestations, and management. Clin. Microbiol. Rev. 28, 603–661. doi: 10.1128/CMR.00134-14
Tsige, Y., Tadesse, S., Eyesus, T., Tefera, M. M., Amsalu, A., and Menberu, M. A. (2020). Prevalence of methicillin-resistant Staphylococcus aureus and associated risk factors among patients with wound infection at referral hospital, Northeast Ethiopia. J Pathog 2020:3168325.
Ukpai, E. G., Chukwura, E. I., Moses, I. B., Ugbo, E. N., Agumah, N. B., Okata-Nwali, O. D., et al. (2021). Prevalence and Antibiogram of healthcare-associated methicillin-resistant Staphylococcus aureus (HA-MRSA) in Ebonyi state, Nigeria. Int. J. Pharma. Sci. Rev. Res. 69, 104–111. doi: 10.47583/ijpsrr.2021.v69i01.016
Ullah, H., Bashir, K., Idrees, M., Ullah, A., Hassan, N., Khan, S., et al. (2022). Phylogenetic analysis and antimicrobial susceptibility profile of uropathogens. PLoS One 17:e0262952. doi: 10.1371/journal.pone.0262952
Uyar Güleç, G., Öncü, S., Bozdoğan, B., Öztürk, B., Ertuğrul, B., and Sakarya, S. (2020). Phenotypic and molecular detection of macrolide lincosamide streptogramin B resistance in clinical isolates of staphylococci. FLORA 25, 190–196. doi: 10.5578/flora.68683
Valle, D. L., Paclibare, P. A., Cabrera, E. C., and Rivera, W. L. (2016). Molecular and phenotypic characterization of methicillin-resistant Staphylococcus aureus isolates from a tertiary hospital in the Philippines. Trop Med Health 44:3. doi: 10.1186/s41182-016-0003-z
Viechtbauer, W. (2010). Conducting Meta-analyses in R with the meta for Package. J. Stat. Softw. 36, 1–48. doi: 10.18637/jss.v036.i03
Viechtbauer, W., and Cheung, M. W. (2010). Outlier and influence diagnostics for meta-analysis. Res. Synth. Methods 1, 112–125. doi: 10.1002/jrsm.11
Vijay, S., and Dalela, G. (2016). Prevalence of LRTI in patients presenting with productive cough and their antibiotic resistance pattern. J. Clin. Diagn. Res. 10, DC09–DC12. doi: 10.7860/JCDR/2016/17855.7082
Wan, T. W., Hung, W. C., Tsai, J. C., Lin, Y. T., Lee, H., Hsueh, P. R., et al. (2016). Novel structure of Enterococcus faecium-originated ermB-positive Tn1546-like element in Staphylococcus aureus. Antimicrob. Agents Chemother. 60, 6108–6114. doi: 10.1128/AAC.01096-16
Wang, R., Li, X., Wang, Q., Zhang, Y., and Wang, H. (2017). Microbiological characteristics and clinical features of cardiac implantable electronic device infections at a tertiary Hospital in China. Front. Microbiol. 8:360. doi: 10.3389/fmicb.2017.00360
Wangai, F. K., Masika, M. M., Lule, G. N., Karari, E. M., Maritim, M. C., Jaoko, W. G., et al. (2019). Bridging antimicrobial resistance knowledge gaps: the east African perspective on a global problem. PLoS One 14:e0212131. doi: 10.1371/journal.pone.0212131
Washington, J. A., and Wilson, W. R. (1985). Erythromycin: a microbial and clinical perspective after 30 years of clinical use (1). Mayo Clin Proc. 1985 60:189–203. doi: 10.1016/s0025-6196(12)60219-5
Weldu, Y., Naizgi, M., Hadgu, A., Desta, A. A., Kahsay, A., Negash, L., et al. (2020). Neonatal septicemia at intensive care unit, Ayder comprehensive specialized hospital, Tigray, North Ethiopia: bacteriological profile, drug susceptibility pattern, and associated factors. PLoS One 15:e0235391. doi: 10.1371/journal.pone.0235391
Wurster, J. I., Bispo, P. J. M., Van Tyne, D., Cadorette, J. J., Boody, R., and Gilmore, M. S. (2018). Staphylococcus aureus from ocular and otolaryngology infections are frequently resistant to clinically important antibiotics and are associated with lineages of community and hospital origins. PLoS One 13:e0208518. doi: 10.1371/journal.pone.0208518
Xie, X., Bao, Y., Ouyang, N., Dai, X., Pan, K., Chen, B., et al. (2016). Molecular epidemiology and characteristic of virulence gene of community-acquired and hospital-acquired methicillin-resistant Staphylococcus aureus isolates in Sun Yat-sen memorial hospital, Guangzhou, southern China. BMC Infect. Dis. 16:339. doi: 10.1186/s12879-016-1684-y
Xu, Z., Liu, S., Chen, L., Liu, Y., Tan, L., Shen, J., et al. (2019). Antimicrobial resistance and molecular characterization of methicillin-resistant coagulase-negative staphylococci from public shared bicycles in Tianjin, China. J. Glob. Antimicrob. Resist 19, 231–235. doi: 10.1016/j.jgar.2019.03.008
Xu, X., Zhang, X., Zhang, G., and Abbasi Tadi, D. (2024). Prevalence of antibiotic resistance of Staphylococcus aureus in cystic fibrosis infection: a systematic review and meta-analysis. J. Glob. Antimicrob. Resist. 36, 419–425. doi: 10.1016/j.jgar.2023.05.006
Yadav, S., and Kapley, A. (2021). Antibiotic resistance: global health crisis and metagenomics. Biotechnol. Rep. (Amst.) 29:e00604. doi: 10.1016/j.btre.2021.e00604
Yang, H., Wang, W. S., Tan, Y., Zhang, D. J., Wu, J. J., and Lei, X. (2017). Investigation and analysis of the characteristics and drug sensitivity of bacteria in skin ulcer infections. Chin. J. Traumatol. 20, 194–197. doi: 10.1016/j.cjtee.2016.09.005
Yao, Z., Wu, Y., Xu, H., Lei, Y., Long, W., Li, M., et al. (2023). Prevalence and clinical characteristics of methicillin-resistant Staphylococcus aureus infections among dermatology inpatients: a 7-year retrospective study at a tertiary care center in Southwest China. Front. Public Health 11:1124930. doi: 10.3389/fpubh.2023.1124930
Yitayeh, L., Gize, A., Kassa, M., Neway, M., Afework, A., Kibret, M., et al. (2021). Antibiogram profiles of Bacteria isolated from different body site infections among patients admitted to GAMBY teaching general hospital, Northwest Ethiopia. Infect Drug Resist 14, 2225–2232. doi: 10.2147/IDR.S307267
Yu, W., Kim, H. K., Rauch, S., Schneewind, O., and Missiakas, D. (2017). Pathogenic conversion of coagulase-negative staphylococci. Microbes Infect. 19, 101–109. doi: 10.1016/j.micinf.2016.12.002
Zamanian, M. H., Shirvani, M., Janbakhsh, A., Sayad, B., Vaziri, S., Mohseni Afshar, Z., et al. (2021). Antibiotic Resistance in Staphylococcus aureus in Patients Hospitalized in Imam Reza Hospital of Kermanshah, Iran (2016–2018). J. Kermanshah Univ. Med. Sci. 25:e118807.
Zhang, J., Gu, F. F., Zhao, S. Y., Xiao, S. Z., Wang, Y. C., Guo, X. K., et al. (2015). Prevalence and molecular epidemiology of Staphylococcus aureus among residents of seven nursing homes in Shanghai. PLoS One 10:e0137593. doi: 10.1371/journal.pone.0137593
Keywords: Staphylococcus , macrolide, meta-analysis, methicillin-resistant Staphylococcus aureus, coagulase-negative staphylococci
Citation: Navidifar T, Zare Banadkouki A, Parvizi E, Mofid M, Golab N, Beig M and Sholeh M (2025) Global prevalence of macrolide-resistant Staphylococcus spp.: a comprehensive systematic review and meta-analysis. Front. Microbiol. 16:1524452. doi: 10.3389/fmicb.2025.1524452
Edited by:
Takashi Azuma, Osaka Medical College, JapanReviewed by:
Okon Okwong Kenneth, Federal University Wukari, NigeriaGuo Yinjuan, Tongji University, China
Copyright © 2025 Navidifar, Zare Banadkouki, Parvizi, Mofid, Golab, Beig and Sholeh. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Mohammad Sholeh, bW9oYW1tYWQuc2hvbGVoLm1haWxAZ21haWwuY29t; Masoumeh Beig, YmVpZ21hc291bWVoQGdtYWlsLmNvbQ==
†These authors have contributed equally to this work
 Tahereh Navidifar
Tahereh Navidifar Abbas Zare Banadkouki2,3†
Abbas Zare Banadkouki2,3† Elnaz Parvizi
Elnaz Parvizi Maryam Mofid
Maryam Mofid Narges Golab
Narges Golab Masoumeh Beig
Masoumeh Beig Mohammad Sholeh
Mohammad Sholeh