
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Microbiol. , 26 March 2025
Sec. Virology
Volume 16 - 2025 | https://doi.org/10.3389/fmicb.2025.1518225
 Xizi Deng1†
Xizi Deng1† Jiaojiao Li1†
Jiaojiao Li1† Ruiying He1
Ruiying He1 Yingfen Wen1
Yingfen Wen1 Yaqing Lin1
Yaqing Lin1 Liya Li1
Liya Li1 Xuemei Ling2
Xuemei Ling2 Fengyu Hu1
Fengyu Hu1 Linghua Li2*
Linghua Li2* Yun Lan1*
Yun Lan1*Background: The development of human immunodeficiency virus (HIV) drug resistance significantly impairs patients’ quality of life. However, the HIV-1 drug resistance patterns in the central nervous system (CNS) have been poorly studied.
Objective: We aimed to compare HIV-1 genotypes and drug resistance mutations (DRMs) derived from the cerebrospinal fluid (CSF) and plasma of antiretroviral therapy (ART)-naive or -experienced patients.
Methods: The matched CSF and plasma samples from 59 patients with HIV were subjected to HIV proteinase (PR), reverse transcriptase (RT), and integrase (IN) gene sequencing. To determine the HIV-1 genotypes, sequences were assessed with the Context-based Modelling for Expeditious Typing (COMET) tool, and the neighbour-joining (NJ) phylogenetic tree was used to confirm the results. Quality control based on genotype and phylogenetic tree analysis was conducted to assess potential sequence contamination during the detection process. The HIV-1 drug resistance database of Stanford University was used to identify DRMs and sensitivity to four drug classes [protease inhibitors (PIs), nucleoside reverse transcriptase inhibitors (NRTIs), nonnucleoside reverse transcriptase inhibitors (NNRTIs), and integrase strand transfer inhibitors (INSTIs)].
Results: Of the 59 patients with HIV with matched CSF and plasma samples, samples from 37 were included in the study after excluding the samples that failed to be successfully amplified. CRF01_AE was the most frequently occurring genotype, with a frequency of 46.0% (17/37), followed by CRF07_BC (27.0%, 10/37) and CRF55_01B (10.8%, 4/37). Among the 37 patients, 37.8% (14/37) carried at least one DRM, and the mutation sites were consistent in both CSF and matched plasma, except one. NNRTI-related resistance mutations were the predominant DRMs, particularly V179D/E, present in 71.4% (10/14) of patients with DRM sites, primarily in ART-naive patients.
Conclusion: A high concordance of HIV-1 DRMs between CSF and plasma samples was observed. No unique mutations were identified in CSF other than those in plasma, indicating that the mutant variants in CSF were derived from blood.
The central nervous system (CNS) is an important site of human immunodeficiency virus (HIV) replication and persistence (Honeycutt et al., 2018). HIV-1 may invade the CNS through infected CD4+ T cells that cross the blood–brain barrier (BBB) soon after systemic infection (Sturdevant et al., 2015). HIV-1 entering the CNS can establish a viral neural reservoir by infecting and adapting to microglia (a brain-specific macrophage population) with low expression of CD4, leading to the evolution of compartmentalized HIV-1 variants in the CNS (Joseph et al., 2019; Chen et al., 2022; Ko et al., 2019). The ability of antiretroviral drugs to penetrate the BBB varies. Findings from several studies have suggested that concentrations of particular drugs may be insufficient to suppress viral replication in the CNS (Asahchop et al., 2017; Moreno-Gamez et al., 2015). The limited penetration of some antiretroviral drugs into the CNS through the BBB, combined with incomplete adherence to ART and ART interruption, may contribute to the compartmentalization of HIV-1 drug resistance between the CSF and plasma, which ultimately results in therapeutic failure (Gianella et al., 2016; Kincer et al., 2023; Vissers et al., 2010; Cysique and Brew, 2009). Findings from previous studies have indicated that drug resistance can develop differently in the two compartments and that the CSF can serve as a viral reservoir for drug resistance mutations (DRMs). In addition, significant differences in the relative frequencies of certain DRMs, such as M184I/V, between CSF and plasma have been reported (Bergroth et al., 2009; Stingele et al., 2001; Kelentse et al., 2020). DRMs in the CSF could evade routine blood diagnostics. Consequently, therapeutic success may be compromised if treatment is adjusted solely based on genotypic resistance patterns detected in one compartment. Currently, HIV-1 drug resistance patterns in the CNS have been poorly studied in China. Therefore, in this study, we aimed to compare HIV-1 genotypes and DRMs derived from the CSF and plasma of ART-naive and ART-experienced patients.
We retrospectively collected matched CSF and plasma samples from 59 ART-naive or ART-experienced patients with HIV who were hospitalized at Guangzhou Eighth People’s Hospital from February 2018 to February 2019. ART-naive patients were patients who have had no prior exposure to any antiretroviral drugs or treatment regimens; ART-experienced patients were patients previously treated with ART and currently taking ART who have experienced virological failure. CSF and plasma samples were obtained for HIV protease (PR), reverse transcriptase (RT), and integrase (IN) gene sequencing. Epidemiological data for the patients (including age, sex, transmission route, year at diagnosis, and treatment regimen) were downloaded from the National Free Antiretroviral Treatment Database for Disease Control and Prevention in China. The Clinical data (HIV-1 RNA viral load, total protein amount in CSF, CSF glucose level, CSF white blood cell count, CSF chloride level, and plasma CD4+/CD8+ T-cell counts) of these patients were collected and anonymously analyzed.
The study was approved by the Institutional Review Board of Guangzhou Eighth People’s Hospital, and all patients signed informed consent forms.
Viral RNA was extracted from 200 μL CSF and plasma samples with an automatic magnetic bead-based Virus RNA Extraction Kit (Daan, China) according to the manufacturer’s instructions. Amplification of the entire PR gene plus partial RT gene (PRRT region, HXB2 2,253–3,318, covering all 99 amino acids of PR and the first 240 amino acids of RT) and entire IN gene (IN region, HXB2 4,230–5,093, covering all 288 amino acids of IN) was performed with an in-house RT polymerase chain reaction (RT–PCR) procedure, as previously described (Lan et al., 2022; Lan et al., 2020). The amplified PCR products were separated by 1% agarose gel electrophoresis and the positive products were sent for Sanger sequencing (Tianyi Huiyuan Genomics Company, China). The obtained sequences were spliced with Sequencher analysis software (Version 5.4.6) and then aligned using BioEdit software (Version 7.2).
The sequences were aligned with 2013 HIV-1 subtyping reference strains downloaded from the Los Alamos HIV Sequence Database via BioEdit software. To determine the HIV-1 genotypes, sequences were assessed with the Context-based Modelling for Expeditious Typing (COMET) tool (Struck et al., 2014). The neighbour-joining (NJ) phylogenetic tree was used for confirmation. The phylogenetic tree was constructed via the NJ method with the Kimura two-parameter model with the MEGA program 6.0(Tamura et al., 2013), and the reliability of the tree topologies was assessed by bootstrapping with 1,000 replications. The pairwise genetic distance between CSF and plasma sequences was calculated using the MEGA program.
Quality control of the sequences derived from CSF and blood was conducted through genotyping and phylogenetic analysis. Genotype consistency between CSF and plasma was evaluated. Sequences were considered reliable for DRMs and drug resistance (DR) if the genotypes matched and CSF and plasma sequences clustered in the phylogenetic tree, thus excluding contamination. Inconsistent sequences were excluded from further analysis.
The HIV-1 drug resistance database of Stanford University (HIVdb version 9.1, https://hivdb.stanford.edu/hivdb/by-sequences/) was used to identify DRMs and sensitivity to four drug classes [protease inhibitors (PIs), nucleoside reverse transcriptase inhibitors (NRTIs), nonnucleoside reverse transcriptase inhibitors (NNRTIs) or integrase strand transfer inhibitors (INSTIs)]. Sequences associated with low-level, intermediate, or high-level resistance categories were defined as conferring drug resistance.
All the statistical analyses were performed using the IBM SPSS program, version 25.0. Qualitative variables are presented as percentages (%), and quantitative variables are presented as medians and interquartile ranges (IQRs) or ranges.
Among all the 59 patients with HIV with matched CSF and plasma samples, 4 CSF samples had low viral loads and could not be tested for drug resistance. Among the remaining 55 CSF samples, PRRT sequences were obtained for 49 (89.1%), and IN sequences were obtained for 48 (87.3%). Among all 59 plasma samples, PRRT sequences were obtained for 56 (94.9%), and IN sequences were obtained for 54 (91.5%). Finally, 37 patients with HIV with matched CSF and plasma PRRT and IN sequences were included in this study, including 27 ART-naive patients and 10 ART-experienced patients. The age of the patients ranged from 25 to 73 years, with a median (IQR) age of 45(30–53) years. In total, 81.1% (30/37) of the patients were male. Approximately two-thirds of the patients had confirmed infections in 2018. Heterosexual (HET) contact was the predominant risk group (51.4%, 19/37), followed by men who have sex with men (MSM) (21.6%, 8/37) and people who inject drugs (5.4%, 2/37). The median CSF and plasma HIV-1 RNA viral load at the time of the drug resistance test were 4.89 and 4.95 (log 10, copies/mL), respectively. All of the ART-experienced patients were treated with two NRTIs, with 60.0% of whom were treated with NNRTIs and 20.0% of whom were treated with PIs or INSTIs. The demographic characteristics of the patients are summarized in Table 1.
According to COMET HIV-1 analysis based on PRRT sequences, HIV genotypes identified in the 37 matched CSF and plasma samples were identical: CRF01_AE was the most frequently occurring genotype, with a frequency of 46.0% (17/37), followed by CRF07_BC (27.0%, 10/37) and CRF55_01B (10.8%, 4/37). A total of five genotypes or circulating recombinant forms (CRFs) were confirmed (94.6%, 35/37) according to the NJ phylogenetic tree based on 37 matched CSF and plasma PRRT sequences (Figure 1A), which was constructed to determine the evolutionary relationship of these sequences. In addition, 2 recombinant strains were observed (COMET tool ‘unassigned’ and not clustered with any reference sequences by the phylogenetic tree), which were classified as ‘other’ genotypes (Figure 1A). CRF07_BC, CRF08_BC, and genotype C cannot fall into clusters on the basis of IN sequences, as they lack the necessary breakpoints for genotyping (Figure 1B). Genetic distances were calculated for all patients with paired CSF-plasma PRRT or IN genes after removing the DRMs. The median genetic distance for the PRRT sequences was 0.0%, ranging from 0.0 to 0.9%, except for two patients whose distances were 0.1 and 0.9%, respectively. For IN sequences, the median genetic distance was also 0.0%, with a range of 0.0 to 0.9%, and all but five patients had distances less than or equal to 0.1%, with those five patients all having a distance of 0.1%.
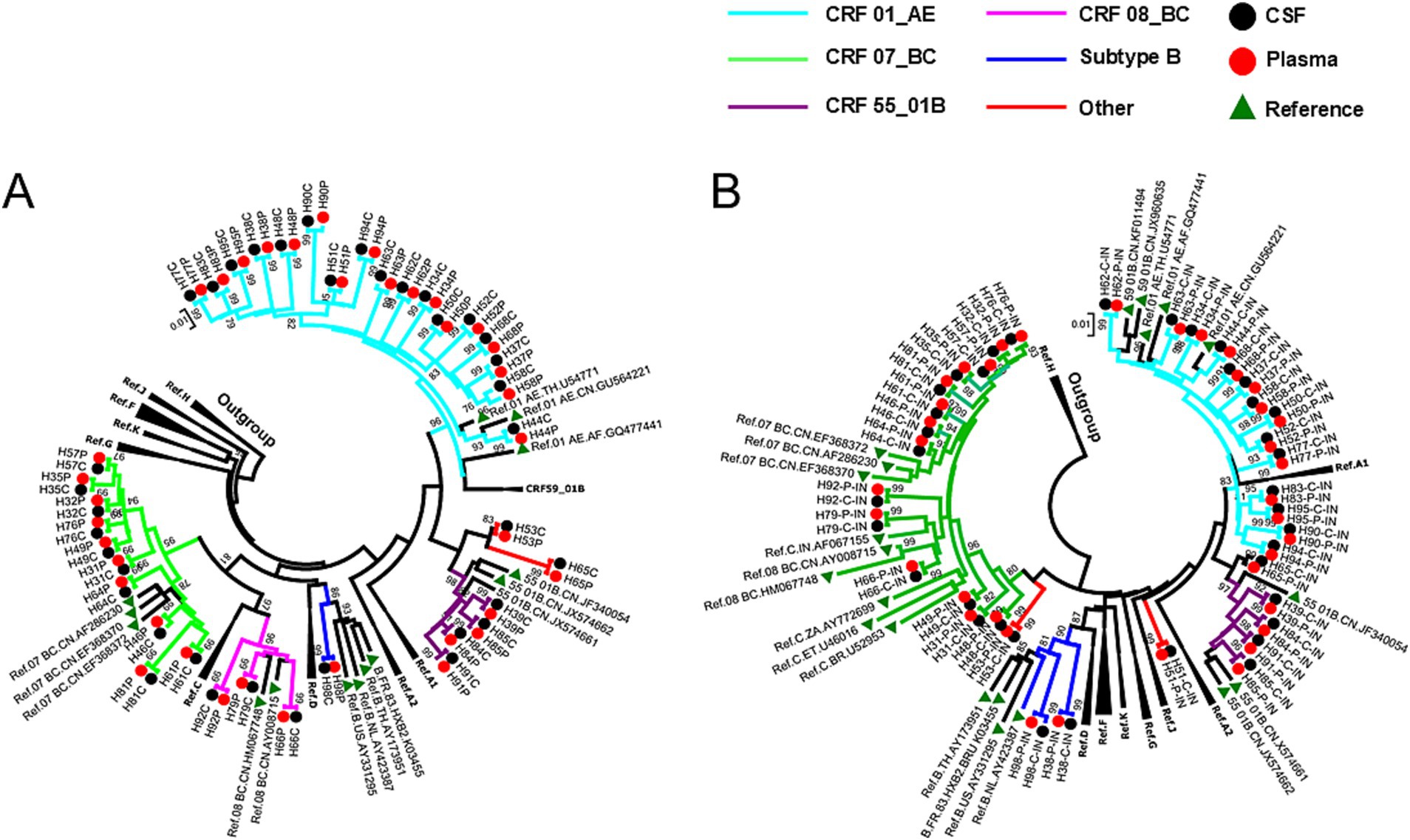
Figure 1. Neighbour-joining (NJ) phylogenetic tree based on protease (PR) gene plus partial reverse transcriptase (RT) gene (PRRT) sequences (A) or integrase (IN) sequences (B) obtained from 37 patients whose main genotypes were initially identified by Context-based modelling for expeditious typing (COMET) of human immunodeficiency virus (HIV). Reference sequences were downloaded from the Los Alamos HIV Sequence Database (https://www.hiv.lanl.gov/content/index); subtype H sequences were selected as the outgroup. Genetic distances were calculated for all patients whose drug-resistance mutation (DRM) sites were removed from paired cerebrospinal fluid (CSF)–plasma PRRT or IN genes. The median (range) genetic distance of the PRRT sequences was 0.0% (0.0–0.9%), which was less than 0.1% for all patients except for two (0.1 and 0.9%). The median (range) genetic distance of the IN sequences was 0.0% (0.0–0.9%) and was less than or equal to 0.1% for all patients except for five (all 0.1%).
Among the 37 patients, 14 (37.8%) carried at least one DRM. Among them, 71.4% (10/14) were ART-naive patients, whereas 28.6% (4/14) were ART-experienced patients (Table 2).
Among the 14 patients with DRM sites, the CSF and plasma genotypes were identical [CRF01_AE (35.7%, 5/14), CRF55_01B (28.6%, 4/14) and CRF07_BC (21.4%, 3/14)], the mutation sites were consistent except in one patient (the IN drug mutation G163R was present in the plasma only for one ART-naive patient), and approximately half of the patients (42.9%, 6/14) had more than one mutation site. The DRMs identified were mainly NNRTI, NRTI, and IN resistance mutations; no PI resistance mutations were found. The NRTI-related resistance mutation sites were K65R, D67N, S68G, K70E, M184V and T215S (all 7.1%, 1/14); the NNRTI-related resistance mutation sites were mainly V179D/E (71.4%, 10/14), A98G, L100I, K103N, V106I, E138G, and G190S (all 7.1%, 1/14); and the IN-related resistance mutation sites were L74M, A128T and G163R (all 7.1%, 1/14) (Table 2).
Five of the 14 patients with DRM sites exhibited varying levels of antiviral drug resistance, with four demonstrating resistances to antiviral agents currently used in China. Notably, patient H81 harbored a resistance mutation that confers resistance to D4T, a drug not currently in use within the country (Table 3).
In this study, we conducted a comparative analysis of PI, NRTI, NNRTI, and INSTI resistance profiles in the CSF and plasma of patients with HIV, irrespective of their ART status.
The COMET HIV-1 and the phylogenetic tree analysis of PRRT sequences revealed consistent findings regarding HIV-1 genotypes in CSF and plasma, thereby validating the reliability of the genotyping results. Specifically, we found that CRF01_AE, CRF07_BC, and CRF55_01B were the dominant genotypes, and these genotypes are consistent with the predominant genotypes currently circulating in Guangdong Province.(Zhou et al., 2019).
Our study reveals a high concordance of HIV-1 DRMs between CSF and plasma, with no mutations identified in CSF that were absent in plasma. The similar phenomena were also observed in a study with 62 patients conducted in Chongqing, China, where the major stain was CRF07_BC (64.5%) (Wang et al., 2023). The two studies conducted in Chongqing and Guangzhou consistently indicated that few patients carried additional DRMs in plasma, and no mutations were found in the CSF that were not already present in the plasma. We identified NNRTI-related resistance mutations as the predominant DRMs, particularly V179D/E, which were present in 71.4% (10/14) of patients with DRM sites, primarily in ART-naive patients. This observation underscores the importance of monitoring DRMs in HIV CRFs before initiating ART and highlights the urgent need for effective preventive measures to reduce transmission. Furthermore, we identified one patient who exhibited an additional HIV-1 DRM, G163R, in the plasma. G163R/K are nonpolymorphic in all subtypes except subtype F. These mutations have been predominantly identified in individuals who are undergoing treatment with raltegravir (RAL). These mutations seem to be accessory mutations since they commonly cooccur with other INSTI resistance mutations, particularly N155H. However, the associated phenotypic effects have not been thoroughly characterized (Hachiya et al., 2017; Margot et al., 2019; Sax et al., 2011). Retrospective analysis of the clinical data from patients harbouring the G163R mutation revealed that the patient had no prior exposure to any antiretroviral drugs or treatment regimens, suggesting the possibility of transmitted drug resistance. Additionally, it was hypothesized that the frequency of quasispecies harbouring the G163R mutation was relatively low and that the insufficient sequencing depth of Sanger sequencing may have contributed to the failure to detect this mutation in CSF samples.
Our research has certain limitations, primarily stemming from the limited sample size. The study population consisted of patients with detectable HIV viral loads in plasma and clinical indications for lumbar puncture during the study period, representing a highly specific subset of individuals living with HIV in China. Consequently, the generalizability of our findings to the broader population of individuals living with HIV in China may be limited. Moreover, the inadequate depth of Sanger sequencing may have resulted in the oversight of certain low-abundance DRMs. Therefore, further studies with larger sample sizes and next-generation sequencing need to be conducted to achieve a more comprehensive assessment of the drug resistance profiles in both CSF and plasma.
In conclusion, we revealed a high concordance of HIV-1 DRMs between CSF and plasma, with no mutations identified in CSF that were absent in plasma. NNRTI-related resistance mutations were predominantly detected, notably the V179D/E mutations, primarily in ART-naive patients, most likely attributable to polymorphisms. These results indicate that when a patient’s ART regimen is adjusted according to the drug resistance profile derived from plasma, the antiretroviral drug would remain effective in the CSF.
The datasets presented in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found at: https://www.ncbi.nlm.nih.gov/genbank/, PV240106-PV240179; PV242912-242985.
The studies involving humans were approved by the Institutional Review Board of Guangzhou Eighth People’s Hospital. The studies were conducted in accordance with the local legislation and institutional requirements. The participants provided their written informed consent to participate in this study.
XD: Data curation, Formal analysis, Writing – original draft. JL: Data curation, Formal analysis, Writing – original draft. RH: Methodology, Writing – original draft. YW: Methodology, Writing – original draft. YaL: Methodology, Writing – original draft. LyL: Investigation, Resources, Writing – original draft. XL: Project administration, Supervision, Validation, Writing – original draft. FH: Writing – review & editing. LnL: Project administration, Supervision, Validation, Writing – review & editing. YuL: Formal analysis, Investigation, Methodology, Project administration, Writing – original draft, Writing – review & editing.
The author(s) declare that financial support was received for the research and/or publication of this article. This study was supported by the Science and Technology Project of Guangzhou (2024A03J0883, 2025A03J3863), the Medical Key Discipline Program of Guangzhou-Viral Infectious Diseases (2021–2023).
We thank the BioBank of Guangzhou Eighth People’s Hospital for Biosamples and Services.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The authors declare that no Gen AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Asahchop, E. L., Meziane, O., Mamik, M. K., Chan, W. F., Branton, W. G., Resch, L., et al. (2017). Reduced antiretroviral drug efficacy and concentration in HIV-infected microglia contributes to viral persistence in brain. Retrovirology 14. Published 2017:47. doi: 10.1186/s12977-017-0370-5
Bergroth, T., Ekici, H., Gisslén, M., Hagberg, L., and Sönnerborg, A. (2009). Difference in drug resistance patterns between minor HIV-1 populations in cerebrospinal fluid and plasma. HIV Med. 10, 111–115. doi: 10.1111/j.1468-1293.2008.00659.x
Chen, J., Zhou, T., Zhang, Y., Luo, S., Chen, H., Chen, D., et al. (2022). The reservoir of latent HIV. Front. Cell. Infect. Microbiol. 12:945956. Published 2022. doi: 10.3389/fcimb.2022.945956
Cysique, L. A., and Brew, B. J. (2009). Neuropsychological functioning and antiretroviral treatment in HIV/AIDS: a review. Neuropsychol. Rev. 19, 169–185. doi: 10.1007/s11065-009-9092-3
Gianella, S., Kosakovsky Pond, S. L., Oliveira, M. F., Scheffler, K., Strain, M. C., de la Torre, A., et al. (2016). Compartmentalized HIV rebound in the central nervous system after interruption of antiretroviral therapy. Virus Evol. 2:vew020. Published 2016. doi: 10.1093/ve/vew020
Hachiya, A., Kirby, K. A., Ido, Y., Shigemi, U., Matsuda, M., Okazakiet, R., et al. (2017). Impact of HIV-1 Integrase L74F and V75I Mutations in a Clinical Isolate on Resistance to Second-Generation Integrase Strand Transfer Inhibitors. Antimicrob. Agents Chemother. 61:e00315–17. doi: 10.1128/AAC.00315-17
Honeycutt, J. B., Liao, B., Nixon, C. C., Cleary, R. A., Thayer, W. O., Birath, S. L., et al. (2018). T cells establish and maintain CNS viral infection in HIV-infected humanized mice. J. Clin. Invest. 128, 2862–2876. doi: 10.1172/JCI98968
Joseph, S. B., Kincer, L. P., Bowman, N. M., Evans, C., Vinikoor, M. J., Lippincott, C. K., et al. (2019). Human immunodeficiency virus type 1 RNA detected in the central nervous system (CNS) after years of suppressive antiretroviral therapy can originate from a replicating CNS reservoir or clonally expanded cells. Clin. Infect. Dis. 69, 1345–1352. doi: 10.1093/cid/ciy1066
Kelentse, N., Moyo, S., Mogwele, M., Lechiile, K., Moraka, N. O., Maruapula, D., et al. (2020). Differences in human immunodeficiency virus-1C viral load and drug resistance mutation between plasma and cerebrospinal fluid in patients with human immunodeficiency virus-associated cryptococcal meningitis in Botswana. Medicine (Baltimore) 99:e22606. doi: 10.1097/MD.0000000000022606
Kincer, L. P., Joseph, S. B., Gilleece, M. M., Hauser, B. M., Sizemore, S., Zhou, S., et al. (2023). Rebound HIV-1 in cerebrospinal fluid after antiviral therapy interruption is mainly clonally amplified R5 T cell-tropic virus. Nat. Microbiol. 8, 260–271. doi: 10.1038/s41564-022-01306-6
Ko, A., Kang, G., Hattler, J. B., Galadima, H. I., Zhang, J., Li, Q., et al. (2019). Macrophages but not Astrocytes Harbor HIV DNA in the brains of HIV-1-infected Aviremic individuals on suppressive antiretroviral therapy. J. Neuroimmune Pharmacol. 14, 110–119. doi: 10.1007/s11481-018-9809-2
Lan, Y., Li, L., Chen, W., Deng, X., Li, J., Fan, Q., et al. (2020). Absence of integrase inhibitor-associated resistance among antiretroviral therapy-Naïve HIV-1-infected adults in Guangdong Province, China, in 2018. Infect. Drug Resist. 13, 4389–4394. Published 2020. doi: 10.2147/IDR.S284917
Lan, Y., Li, F., Li, L., Deng, X., Li, L., Li, J., et al. (2022). Drug resistance and genetic transmission characteristics of HIV-1 CRF59_01B in infected patients in Guangdong Province, China. J. Antimicrob. Chemother. 77, 1960–1968. doi: 10.1093/jac/dkac132
Margot, N. A., Ram, R. R., White, K. L., Abram, M. E., and Callebaut, C. (2019). Antiviral activity of HIV-1 integrase strand-transfer inhibitors against mutants with integrase resistance-associated mutations and their frequency in treatment-naïve individuals. J Med Virol. 91, 2188–2194. doi: 10.1002/jmv.25564
Moreno-Gamez, S., Hill, A. L., Rosenbloom, D. I., Petrov, D. A., Nowak, M. A., and Pennings, P. S. (2015). Imperfect drug penetration leads to spatial monotherapy and rapid evolution of multidrug resistance. Proc. Natl. Acad. Sci. USA 112, E2874–E2883. doi: 10.1073/pnas.1424184112
Sax, P. E., Tierney, C., Collier, A. C., Daar, E. S., Mollan, K., Budhathoki, C., et al. (2011). Abacavir/lamivudine versus tenofovir DF/emtricitabine as part of combination regimens for initial treatment of HIV: final results. J Infect Dis. 204, 1191–1201. doi: 10.1093/infdis/jir505
Stingele, K., Haas, J., Zimmermann, T., Stingele, R., Hübsch–Müller, C., Freitag, M., et al. (2001). Independent HIV replication in paired CSF and blood viral isolates during antiretroviral therapy. Neurology 56, 355–361. doi: 10.1212/wnl.56.3.355
Struck, D., Lawyer, G., Ternes, A. M., Schmit, J. C., and Bercoff, D. P. (2014). COMET: adaptive context-based modeling for ultrafast HIV-1 subtype identification. Nucleic Acids Res. 42:e144. doi: 10.1093/nar/gku739
Sturdevant, C. B., Joseph, S. B., Schnell, G., Price, R. W., Swanstrom, R., and Spudich, S. (2015). Compartmentalized replication of R5 T cell-tropic HIV-1 in the central nervous system early in the course of infection. PLoS Pathog. 11:e1004720. doi: 10.1371/journal.ppat.1004720
Tamura, K., Stecher, G., Peterson, D., Filipski, A., and Kumar, S. (2013). MEGA6: molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 30, 2725–2729. doi: 10.1093/molbev/mst197
Vissers, M., Stelma, F. F., and Koopmans, P. P. (2010). Could differential virological characteristics account for ongoing viral replication and insidious damage of the brain during HIV 1 infection of the central nervous system? J. Clin. Virol. 49, 231–238. doi: 10.1016/j.jcv.2010.08.001
Wang, J., Li, M., Li, J., and Deng, R. (2023). Differences in drug resistance of HIV-1 genotypes in CSF and plasma and analysis of related factors. Virulence 14:2171632. doi: 10.1080/21505594.2023.2171632
Keywords: HIV-1, CSF, plasma, drug resistance, genotype
Citation: Deng X, Li J, He R, Wen Y, Lin Y, Li L, Ling X, Hu F, Li L and Lan Y (2025) High concordance of human immunodeficiency virus-1 genotypic drug resistance generated from paired cerebrospinal fluid and plasma in antiretroviral therapy -naive or -experienced patients. Front. Microbiol. 16:1518225. doi: 10.3389/fmicb.2025.1518225
Received: 29 October 2024; Accepted: 25 February 2025;
Published: 26 March 2025.
Edited by:
Bin Su, Capital Medical University, ChinaReviewed by:
Ting Pan, Sun Yat-sen University, ChinaCopyright © 2025 Deng, Li, He, Wen, Lin, Li, Ling, Hu, Li and Lan. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Linghua Li, bGxoZWxpemFAMTI2LmNvbQ==; Yun Lan, bGFueXVuLnZpcEAxMjYuY29t
†These authors have contributed equally to this work
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.