
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Microbiol. , 27 February 2025
Sec. Infectious Agents and Disease
Volume 16 - 2025 | https://doi.org/10.3389/fmicb.2025.1514643
Introduction: Klebsiella pneumoniae and K. oxytoca are members of Enterobacteriaceae. They are Gram-negative, non-motile rods that are ubiquitous in the environment and part of the human intestinal microbiota. These opportunistic pathogens may cause pneumonia and urinary tract infections. Klebsiella species are genetically and biochemically similar; therefore, it is important to find reliable methods for their differentiation.
Methods: This study presents the results of biochemical assays, PCR, and MALDI-TOF mass spectrometry (MS) performed on 35 Klebsiella isolates obtained from the urine of patients from central Poland.
Results: Among biochemical methods, the indole test demonstrated the highest discriminatory power, whereas the determination of growth at 10°C was the least effective. For all strains biochemically identified as K. pneumoniae, a 108-bp amplicon was detected, indicating the presence of the rpoB gene in their genome. Only 12 K. oxytoca isolates produced a product of the pehX gene. All tested strains were analyzed using the MALDI-TOF Biotyper, which confirmed, with high-quality scores, their identification based on api 20E and indole tests. Strain 0.011 was identified as Raoultella ornithinolytica.
Conclusion: MALDI-TOF MS analysis proved to be the most reliable method for identifying K. oxytoca and K. pneumoniae, with the potential for phylogroup differentiation.
Klebsiella oxytoca and K. pneumoniae are non-motile, Gram-negative bacilli belonging to the Enterobacteriaceae. These species are ubiquitous in the environment where they play the role of diazotrophs (Anbuselvi et al., 2017; Brisse et al., 2006; Kovtunovych et al., 2003). Both species also colonize the skin, oropharynx, and intestines of healthy humans and animals (Yang et al., 2022). These bacteria are regarded as opportunistic pathogens that affect mainly immunocompromised patients and cause infections of the respiratory and urinary tracts (Martin and Bachman, 2018). Among clinical Klebsiella spp. isolates, K. pneumoniae predominates over K. oxytoca strains; however, the increasing K. oxytoca isolation from the samples indicates the species is a clinically significant pathogen (Gómez et al., 2021; Singh et al., 2016). K. pneumoniae was first isolated from a patient by Friedländer in 1982, but its present name was introduced 5 years later by Trevisan, whereas K. oxytoca was first described in 1886 by Flügge as Bacillus oxytocus perniciosus, and the current name, K. oxytoca, was introduced in 1956 by Lautrop (Brisse et al., 2006; Yang et al., 2022). The existence of this species was questionable for many years, and K. oxytoca was even regarded as a biogroup of K. pneumonia. Since then, the Klebsiella taxonomy has been significantly reorganized, e.g., some Klebsiella species such as Klebsiella terrigena, Klebsiella planticola, and Klebsiella ornithinolytica have been reassigned to the newly established genus Raoultella (Brisse et al., 2006).
The K. oxytoca group has been found as a cause of urinary tract infections or pneumonia acquired in hospitals and antibiotic-associated hemorrhagic colitis in neonates (Cuénod et al., 2021). Isolation of K. oxytoca from severe medical cases and its increasingly occurring multidrug resistance phenotype requires the development of a diagnostic method for quick and proper identification of the species. Klebsiella species are genetically similar (Bridel et al., 2021). They also possess similar phenotypic and biochemical features. Thus, the discriminatory power of 16S rRNA gene sequencing and metabolic reaction profiles may show different effectiveness depending on the species (Cuénod et al., 2021). Specific PCR conditions with species-specific primers targeting (a) gene pehX encoding the polygalacturonase, which cleaves a polygalacturonic chain of demethoxylated pectin, and (b) gene rpoB coding the beta subunit of RNA polymerase have been developed to K. oxytoca and K. pneumoniae identification. The method allowed the identification of amplicons of 343 bp for K. oxytoca strains and 108 bp for K. pneumoniae (Chander et al., 2011; Kovtunovych et al., 2003).
The results of biochemical reactions that are indicated for K. pneumoniae and K. oxytoca identification may also vary depending on the tested strains, which may result in the incorrect identification of the species (Hansen et al., 2004). Among the biochemical reactions commonly used by reference laboratories, two features can be highlighted: indole production from tryptophan and pectate degradation. These traits differentiate the K. oxytoca complex (positive results) not only from K. pneumoniae but also from the majority of Raoultella species. Only Raoultella ornithinolytica is also capable of indole production. Gas production from lactose at 44.5°C is a feature unique to Klebsiella pneumoniae subsp. pneumoniae and is not observed in the K. oxytoca complex or in Raoultella species (Hansen et al., 2004; Brisse et al., 2006). The majority of K. oxytoca strains are capable of growth at 10°C, whereas 93–100% of tested Raoultella species also demonstrate this ability. In contrast, the growth of K. pneumoniae at 10°C has not been commonly observed; however, this result may be method-dependent (Hansen et al., 2004).
However, it should be remembered that phylogenetic analyses have revealed that K. pneumoniae and K. oxytoca form two complexes, and each of them includes phylogenetic groups: Kp1–Kp7 and Ko1–Ko4, Ko6–Ko9, respectively (Gómez, et al., 2021; Rodrigues et al., 2019). Kp1 contains K. pneumoniae sensu stricto with three subspecies: pneumoniae, ozaenae, and rhinoscleromatis, whereas the Ko2 phylogroup includes K. oxytoca sensu stricto. The remaining species within each complex are as follows: Klebsiella michiganensis (Ko1), K. spallanzanii (Ko3), K. pasteurii (Ko4), K. grimontii (Ko6), K. huaxiensis (Ko8), K. quasipneumoniae subsp. quasipneumoniae (Kp2), K. quasipneumoniae subsp. similipneumoniae (Kp3), K. variicola (Kp4), K. variicola susp. tropicalensis (Kp5), K. quasivariicola (Kp6), and K. africanensis (Kp7) (Bridel et al., 2021; Brisse et al., 2006; Martínez et al., 2004). Ko5 is recognized as a sub-phylogroup of Ko1, and Ko9 as a sub-phylogroup of Ko3, whereas the taxonomic status of Ko7 remains unresolved (Rodrigues et al., 2019; Yang et al., 2022). Despite the fact that each species within the K. oxytoca complex shows a unique phenotypic profile, such identification has usually been done only for a few strains; moreover, the results of some phenotypic tests were variable even within one phylogroup (Yang et al., 2022). Variable results were observed for Klebsiella spallanzanii (Ko3) and Klebsiella grimontii (Ko6) when dl-α-glycerol-phosphate utilization was tested, a characteristic that differentiates Klebsiella huaxiensis (Ko8 – negative result) from the K. oxytoca complex (Merla et al., 2019). As for the K. pneumoniae complex, the situation was clear only regarding the utilization of N-acetyl-neuraminic acid, a feature characteristic exclusively of Kp4, and the utilization of 4-hydroxy-l-proline, which was negative only for Kp1 within the K. pneumoniae complex (Rodrigues et al., 2019).
To distinguish species within the Klebsiella oxytoca complex, species-specific matrix-assisted laser desorption ionization-time-of-flight mass spectrometry (MALDI-TOF MS) markers were proposed by Merla et al. (2019), who identified 31 biomarkers, and by Cuénod et al. (2021), who proposed 25 ribosomal subunit proteins for the identification of different Klebsiella species. However, it should be noted that successful identification and reproducibility depend on the validity of the MALDI Biotyper Database. Currently, nine phylogroups of the K. oxytoca complex are assigned to respective β-lactamase gene blaOXY variants (Cosic et al., 2021; Yang et al., 2022). Cosic et al. (2021) have recently identified species-specific gene loci and proposed their combinations (blaOXY-2, blaOXY-4, blaOXY-1&5, blaOXY-6, leupAB, orfABC, orfA’) as useful in differentiating biochemically identical species: K. oxytoca, K. michiganensis, K. pasteurii, and K. grimontii (Supplementary Figure S1).
Another method applied for species identification is the Kleborate framework (Lam et al., 2021). This method allowed Stewart et al. (2022) to demonstrate species diversity among 92 isolates previously identified by a clinical laboratory as K. oxytoca. However, Kleborate was designed to screen genome assemblies of Klebsiella pneumoniae and the Klebsiella pneumoniae species complex (KpSC), mainly to detect and track clinically relevant AMR (acquired antimicrobial resistance) and virulence determinants (e.g., the rmp hypermucoidy locus and the rmpA2 gene) from the genome database. This tool can also be used for K or O locus identification and prediction of K and O serotypes when switched to Kaptive (Wyres et al., 2016).
To gain a better understanding of the emerging pathogen K. oxytoca and the more commonly isolated K. pneumoniae, it is extremely important to use reliable methods for their identification. These studies present the results of different kinds of methods, phenotypic, molecular, and matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS), to indicate which is the most appropriate for the species classification to the K. pneumoniae or K. oxytoca group.
A total of 35 Klebsiella spp. strains (0.04, 0.05, 0.09–0.011, 0.013, 0.019, 0.021–0.025, 0.029, 0.030, 0.033, 0.034, 0.038, 0.040, 0.042, 0.045, 0.046, 0.050, 0.054, 0.055, 0.057, 0.060–0.063, 0.065, 0.067–0.069, 0.071, 0.079) were selected for the study. The species affiliation of the strains was initially estimated (by rapid identification assay) in the Synevo laboratory, from which the strains came. All 35 strains are isolated from the urine of patients from the Łódź area, Poland. The type strains of K. oxytoca (8724) and K. pneumoniae species (700603; 10031) were obtained from the American Type Culture Collection (ATCC).
All 35 tested strains were biochemically identified by a standardized system api 20E (BioMerieux) according to the producer’s instructions.
The indole test was performed on a bacterial culture (after incubation at 37°C for 24 h) on a medium with DL-tryptophan according to the method developed by Ewing (1986). A positive result was shown after the addition of a few drops of Kovac’s reagent and the appearance of a red-violet ring on the culture broth surface. Klebsiella oxytoca can produce indole from DL-tryptophan, whereas Klebsiella pneumoniae strains do not exhibit this ability.
Growth determination at 10°C was conducted as Naemura and Seidler described (1978) by inoculating nutrient broth (NB) with an overnight culture in NB. The tubes were incubated at 10°C for 72 h. This ability is considered characteristic of Klebsiella oxytoca but not of Klebsiella pneumoniae strains. The optical density was measured at a wavelength of 600 nm every 24 h. OD600 > 0.02 was defined as a positive result.
The ability of the strains to produce gas from lactose at 44.5°C was tested by using two kinds of lactose broths: EC (0.4% lactose, bile salts, casein digest, dipotassium phosphate, potassium phosphate, and sodium chloride) and 1% lactose broth (1% lactose and peptone). In each broth, a Durham tube was inserted to observe CO2 production. Gas production from lactose at 44.5°C is the characteristic of Klebsiella pneumoniae subsp. pneumoniae and is not observed in the Klebsiella oxytoca complex.
The procedures were repeated to obtain reproducible results.
Genomic DNA was obtained from the tested strains cultured for 18 h in L-broth by the use of a commercial Genomic Mini Kit (A&A Biotechnology) according to the manufacturer’s instructions. DNA suspended in Tris buffer (10 mM Tris, pH = 8) was stored at −20°C.
Species (KP – K. pneumoniae and KO – K. oxytoca) specific pairs of primers targeting the rpoB or pehX gene (KP – forward [F] and reverse [R]; KO – F and R) were used for amplification:
KP (F) 5’-CAA CGG TGT GGT TAC TGA CG-3′; 5’-TCT ACG AAG TGG CCG TTT TC-3′ (R); KO (F) 5′-GAT ACG GAG TAT GCC TTT ACG GTG-3′; 5′-TAG CCT TTA TCA AGC GGA TAC TGG-3′ (R) (Chander et al., 2011). The oligonucleotides were synthesized by Bionovo. PCR amplification was performed at a final volume of 50 μL, containing 1 μL of template DNA, 1 μL of each primer, and 25 μL of HotStarTaq™ DNA polymerase Master Mix Kit (Qiagen). PCR was performed in a thermocycler (Eppendorf) according to the following conditions (Table 1).
PCR products were submitted to 1% agarose gel electrophoresis in TAE buffer (40 mM Tris, pH 7.6; 20 mM C2H4O2, 1 mM EDTA) at 70 V. The gel was stained with ethidium bromide and visualized under an ultraviolet transilluminator (GelDoc 2000). The 108-bp amplicons are expected to be generated when a rpoB-specific pair of primers is used. These primers have been found to be Klebsiella pneumoniae-specific. The 343-bp amplicons were detected when a pehX-specific pair of primers was applied, and these products are typical for Klebsiella oxytoca strains. The band position of the 108-bp and the 343-bp amplicons was established in reference to a 100–3,000-bp or 100–2000-bp DNA ladder (BLIRT DNA-GDAŃSK) used as a molecular weight standard. The procedure was repeated to obtain reproducible results and ensure good-quality product visualization.
Strains were also identified by using MALDI Biotyper (Bruker) at the Jagiellonian Center of Innovation (Krakow, Poland) (strains 0.010, 0.011, 0.013, 0.021, 0.050, 0.055, 0.071, 0.061, and 0.067) and Laboratory of Microbial Immunochemistry and Vaccines, Hirszfeld Institute of Immunology and Experimental Therapy, Polish Academy of Sciences (Wrocław, Poland) (all tested strains). The 18-h Klebsiella isolates were spotted onto a plate according to the direct transfer method with the α-cyano-4-hydroxycinnamic acid as a matrix (10 mg/mL in acetonitrile 50%, water 47.5%, and trifluoroacetic acid 2.5%). MALDI-TOF MS spectra were calibrated with the Bruker Bacterial Test Standard (Bruker Daltonics, Germany) with the reference masses: 3637.8, 5096.8, 5381.4, 6255.4, 7274.5, 10300.1, 13683.2, and 16952.3 Da. The MALDI-TOF mass spectra were obtained in a positive linear mode within a mass range of 2–20 kDa by using an ultrafleXtreme (Bruker, Bremen, Germany) mass spectrometer and analyzed by Biotyper software (version 3.0). The score values were interpreted as follows: ≥2.00 – high-confidence identification; 1.70–1.99 – probably genus identification. The m/z data obtained were analyzed in relation to the 31 markers (ions) described previously for K. oxytoca species complexes (Merla et al., 2019) using the Bruker Flex Analyses software (v.3.4).
The species identification was confirmed by api 20E system. For most strains identified by the api system as K. oxytoca, low discrimination was obtained according to the Analytical Profile Index (Tables 2, 3). However, %id ≥80 indicated an acceptable identification. The doubtful results concerned the urease, indole, lysine, rhamnose, or Voges–Proskauer tests. As for the urease test, an extension of the incubation time to 48 h resulted in the occurrence of a positive reaction. In the case of one strain, 0.069, species identification by api 20 E was impossible (Tables 2, 3). For the reference strains, K. oxytoca ATCC 8724 and K. pneumoniae ATCC 700603, the api 20E results were accurate and did not raise any doubts.
For further biochemical identification of the tested strains, three biochemical tests (indole test, growth at 10°C, and gas from lactose at 44.5°C) were chosen from the test panel for the differentiation of Klebsiella species. Among the strains identified as K. oxytoca by Synevo, only one, 0.050, did not produce indole from tryptophan, which is not a feature typical for this species (Table 2). The reference strain K. oxytoca ATCC 8724 produced indole, which was indicated by the presence of a red-violet ring on the surface of the culture broth, whereas K. pneumoniae ATCC 700603 gave a negative result in the indole assay.
As can be seen in Table 2, the ability to produce gas from lactose at 44.5°C appeared to be medium-dependent. More strains classified as K. pneumoniae (62.5%) produced CO2 when grown in EC broth than when cultured in 1% lactose broth (50%) (Table 2). None of the K. oxytoca strains tested in the studies occurred to be gas producer, which is consistent with the results for the K. oxytoca reference strain, 8724. K. pneumoniae ATCC 700603 produced gas in both types of media (Table 2).
When the growth response in nutrient broth at 10°C was tested after 24 h of incubation, most K. oxytoca strains (16 tested strains and K. oxytoca ATCC 8724) and 5 K. pneumoniae isolates were able to grow–OD values of the cultures were higher than the OD of 0.02 (Table 2). K. pneumoniae ATCC 700603 strain was unable to grow at 10°C. After 48 h of incubation, growth was observed for most of the tested strains except K. oxytoca (0.022 and 0.063). When the time of incubation was extended to 72 h, only one strain, K. oxytoca 0.063, was unable to grow.
All strains identified by api 20E as K. pneumoniae and the species type strain were screened for the presence of the rpoB gene with primer pairs (KP(F) and KP(R)). The 108-bp fragment regarded as characteristic of the K. pneumoniae species was obtained from the DNA of all tested strains, which stays in agreement with biochemical identification [api 20E, indole assay, and gas production from lactose] (Figure 1A and Table 2). For K. oxytoca ATCC 8724 and E. coli ATCC 25966 (negative controls), slightly visible bands were obtained in gel (Figure 1A). For one strain, 0.034, identified by the laboratory as K. oxytoca, the results of both molecular and biochemical studies (including api 20E) appeared to be typical for K. pneumoniae.
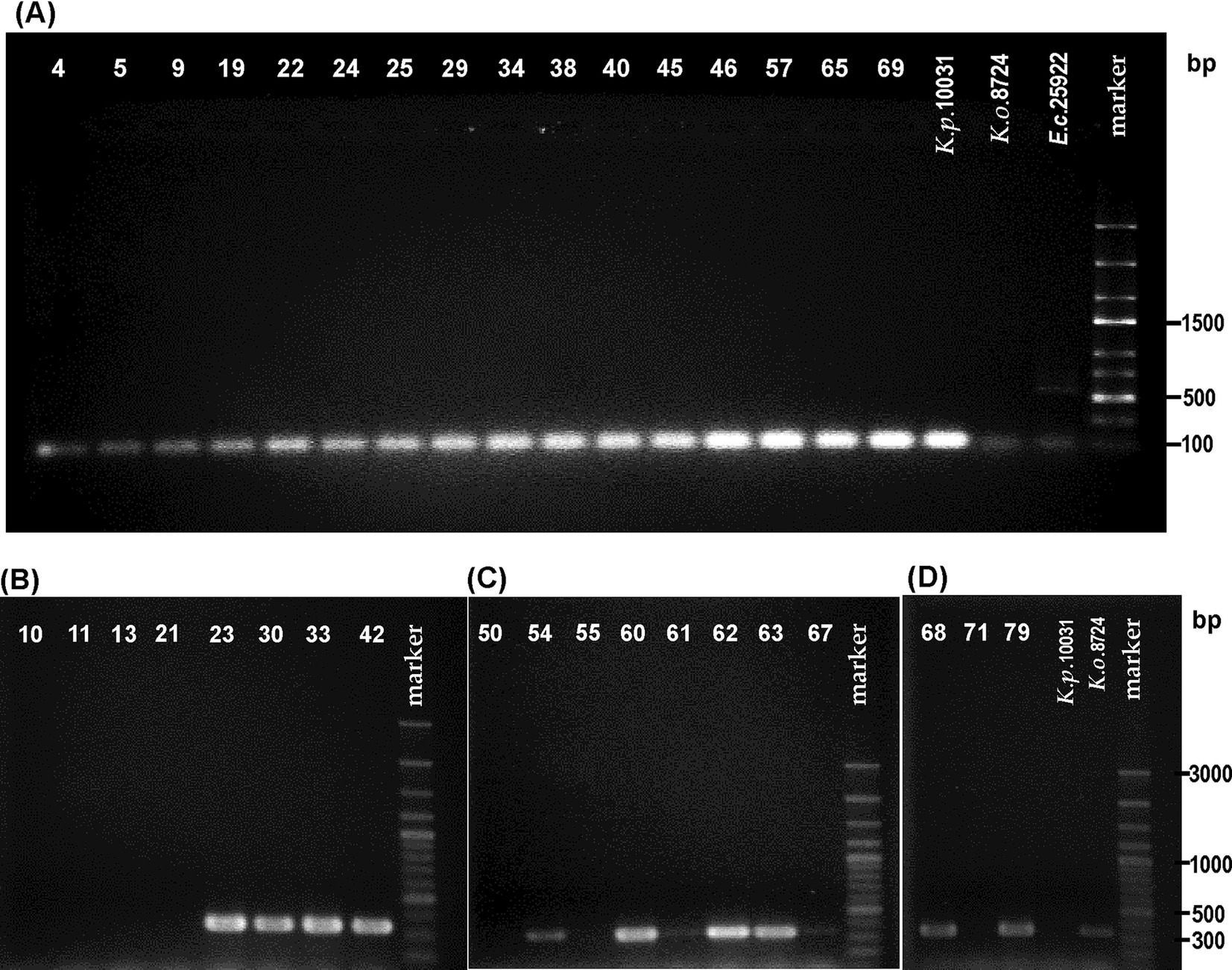
Figure 1. Visualization of electrophoretically separated PCR products of: (A) rpoB gene (expected size: 108 bp) for the strains biochemically identified as Klebsiella pneumoniae, (B–D) pehX gene (expected size: 343 bp) for the strains biochemically identified as Klebsiella oxytoca. The strain numbers are simplified (e.g., 0.010–10) to make the figure clearer. The band position of the amplicon was established in reference to 100–3,000-bp DNA or 100–2000-bp ladders used as a molecular weight standard. The results of the PCR control targeting the pehX and rpoB genes are presented in Supplementary Figure S1.
Subsequently, 18 strains biochemically classified as K. oxytoca (also the ones for which the api results were doubtful) and strain 0.050 (identified as K. oxytoca) were screened for the presence of the pehX gene. As Figures 1B–D show that 12 from the 18 tested strains generated 343-bp amplicons with primer pair KO(F) and KO(R). For two strains, 0.061 and 0.067, the bands were barely visible. Thus, DNA from the 0.061 and 0.067 strains, together with those that did not generate the 343-bp fragment (0.010, 0.011, 0.013, 0.021, 0.050, 0.055, and 0.071), was checked for the presence of gene rpoB. Surprisingly, all tested strains were positive upon PCR with primer pairs (KP(F) and KP(R)), and 108-bp amplicons were obtained, which presumably indicated these isolates as K. pneumoniae (Figure 2).
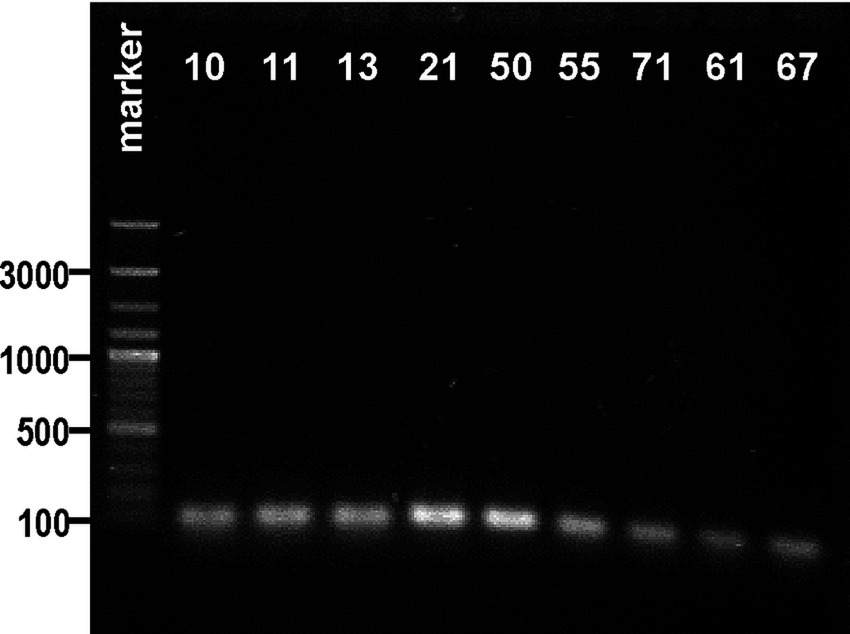
Figure 2. Visualization of electrophoretic separated PCR products of the rpoB gene (expected size: 108 bp) for the strains biochemically identified as Klebsiella oxytoca in agarose gel. The strain numbers are simplified (e.g., 0.010–10) to make the figure clearer. The band position of the amplicon was established in reference to a 100–3,000-bp DNA ladder used as a molecular weight standard. The results of the PCR control targeting the pehX and rpoB genes are presented in Supplementary Figure S1.
The final part of the study includes the species classification of the tested Klebsiella isolates using MALDI-TOF MS Biotyper (Bruker). 17 strains were classified as K. oxytoca, 17 – as K. pneumoniae and one strain, 0.011, was identified as Raoultella ornithinolytica. For all strains, score values ≥2 were obtained, which proved high-quality identification (Table 3).
The m/z data obtained in the present study were analyzed in relation to the identified markers (Merla et al., 2019). Many peaks appeared to be common for the majority of strains tested (e.g., m/z at 2067, 2704, 2837, 3134, 3578, 4731, 5410, and 6254/55). As shown in Table 4, the tested strains could not be classified to determine the phylogroup. One m/z value, 5052, which is typical only for two phylogroups, was found for four strains tested (0.013, 0.054, 0.067, and 0.071), suggesting their affiliation with Ko1 or Ko6 (Table 4). For K. oxytoca 0.071 and 0.013 strains, only one specific peak was observed, whereas other strains from Table 4 shared peaks with the other isolates. For example, three other specific peaks at m/z 2591, 3132, and 2702 were observed for the K. oxytoca 0.055 strain, and they were described by Merla et al. (2019) as those associated with the Ko1, Ko4, Ko6, Ko2 (m/z 2591, 3132) and Ko1, Ko4, Ko6, Ko3, Ko2 (m/z 2702) phylogroups (Table 4).
For many years, the diagnostic methods of species identification were focused on phenotypic characterization (Hansen et al., 2004). However, commercial biochemical tests for Enterobacteriaceae often fail to differentiate Klebsiella species (Chander et al., 2011; Hasan and Mohammed Bakr, 2022). The results by api 20E did not fully confirm the species affiliation indicated by the laboratory from which the strains came (Table 3). Strain 0.021 previously classified as K. pneumoniae, appeared to be K. oxytoca according to the api 20E numerical profile, whereas strain 0.034 previously designated as K. oxytoca was identified by api 20E as K. pneumoniae. However, the api 20E system also failed to identify one strain, 0.069, for which no appropriate numerical profile could be found in the Analytical Profile Index (Table 3).
In the biochemical studies, we aimed to check if the api 20E-based identification correlates with that made by using the three methods: indole test, growth determination at 10°C, and gas production from lactose at 44.5°C. As shown in Tables 2, 3, only the indole test proved to be fully effective in the differentiation between K. oxytoca (positive results) and K. pneumoniae (negative results), and its outcomes were consistent with those obtained by api 20E and MALDI Biotyper (Tables 2, 3). Such results indicate the indole assay as a reliable tool in the differentiation between K. oxytoca and K. pneumoniae. The data presented by Hansen et al. (2004) also showed this test to be effective for K. oxytoca identification (90–100% of the tested samples were positive for K. oxytoca).
The results obtained by the method of detecting gas production from lactose were not as evident as those of the indole test. This ability should be expressed by the majority of K. pneumoniae strains, which was proved by Hansen et al. (2004), who observed the positive results of the method for 89.3% of K. pneumoniae isolates. From the strains classified by api 20E (in the present study) as K. pneumoniae, 62.5% produced gas in EC broth and 50% in 1% lactose broth. Only 25% of K. pneumoniae strains showed gas production in both media. According to the technical data of the HiMedia Laboratories, the observed differences may result from the presence of the EC broth of phosphate salts controlling the pH during the fermentation process. False-negative reactions may occur due to a low pH. Obtaining more positive results in EC broth indicates that this medium is more appropriate for the detection of gas production from lactose at 44.5°C than 1% lactose broth. In the studies by Hansen et al. (2004), 3.3–6.7% of K. oxytoca strains produced gas from lactose. In this work, the K. oxytoca strains were unable to do that (Table 2), which is typical for this species. The negative results achieved for some K. pneumoniae strains may have resulted from the fact that they represent subspecies other than pneumoniae. Most K. pneumoniae ssp. rhinoscleromatis strains did not produce gas from lactose at 44.5°C, and only for 3.7–11.1% of K. pneumoniae spp. ozaenae gas was detected (Hansen et al., 2004). Interspecies differences in the ability to ferment lactose at 44.5°C among K. pneumoniae suggested this method as a questionable tool for Klebsiella species differentiation. However, this procedure is regarded as an effective method for detecting fecal coliform-positive Klebsiella in the environment and monitoring fecal contamination (Naemura and Seidler, 1978).
From all biochemical methods applied in the present study, a test for growing at 10°C showed the worst differentiating effect. This ability should be typical for K. oxytoca and not expressed by K. pneumoniae. However, when Hansen et al. (2004) analyzed the results obtained by one of the three laboratories, it was revealed that 57% of K. pneumoniae isolates were also capable of growing at 10°C. Naemura and Seidler indicated 72 h of incubation as a time adequate for noting a growth response at 10°C. In the present study, almost all strains tested, except K. oxytoca 0.063, grew after 72 h of incubation. In this case, the better discrimination power of the method was observed after 24 h of incubation when most of the tested K. pneumoniae strains did not grow, whereas most of the K. oxytoca isolates were capable of growing (OD600 > 0.02). The increase in OD values observed for 48 h-cultured and 72 h-cultured strains, independently of the species they represent, suggests that time extension is unsuitable for Klebsiella species differentiation.
Ambiguous results and interspecies differences observed for some biochemical reactions could result from strains’ phenotypic variations modified by environmental conditions. Thus, the standardization of biochemical tests for distinguishing K. oxytoca from K. pneumoniae is not an easy task. Moreover, using phenotypic markers for species distinguishing is not very reliable; it is time-consuming and often does not allow correcting the identification since related species frequently present similar biochemical patterns (Chander et al., 2011, Hasan and Mohammed Bakr, 2022). In contrast, PCR-based methods using different specific primers and amplification conditions have been widely suggested as reliable diagnostic tools for the correct identification of K. pneumoniae and K. oxytoca (Chander et al., 2011; Hasan and Mohammed Bakr, 2022; Kovtunovych et al., 2003). Two pairs of specific primers were proposed to be used in distinguishing these two groups of species: targeting the polygalacturonase gene and gene rpoB. These specific primers were also used in this study for PCR performed under the conditions described by Chander et al. with a few modifications (Table 1). For all strains biochemically identified as K. pneumoniae, 108-bp amplicons were obtained, which indicates the presence of the rpoB gene in the genome of the isolates tested. However, the K. oxytoca and E. coli reference strains also demonstrated a PCR product with 108 bp, which was only slightly visible in gel compared to those obtained for the K. pneumoniae isolates (Figure 1A). The rpoB is one of the genetic markers tested during the 16S rRNA gene sequence analysis. Recent phylogenetic studies conducted on 37 K. pneumoniae strains revealed that the rrs gene sequences coding for 16S rRNA were > 98.8% similar (Rodrigues et al., 2019). Thus, it can be seen that 16S rRNA is not always a proper target for species identification within Enterobacteriaceae (Cuénod et al., 2021).
The strains designed by API as K. oxytoca were also identified using specific primers for pehX gene detection. A PCR amplicon of 343 bp was obtained for twelve isolates, but for two of them (0.061 and 0.067) the product was obtained in small amounts (the slightly visible bands in Figures 1C, D). It is worth mentioning that K. michiganensis has been reported to be negative for pehX by PCR (Yang et al., 2022). Using PCR with the same set of primer pairs and similar PCR conditions, Chander et al. (2011) also did not confirm the biochemical identification of 20% of the tested K. oxytoca strains of human origin. The BLAST analysis indicated that pehX was truncated between nucleotides 1,977 and 1,983 in a few K. oxytoca genomes, including K. michiganensis strains A10 and A11, K. oxytoca strain 112, and K. pasteurii strain (Yang et al., 2022). For such strains, the detection of the pehX product by PCR could be hampered. The results of biochemical assays and PCR obtained for strains 0.050 and 0.034, previously identified as K. oxytoca, appeared to be typical for K. pneumoniae species. It is an unusual situation since in most cases strains previously attributed to K. pneumoniae appeared to be K. oxytoca, like in the studies of Kovtunovych et al. (2003), who developed a PCR specific for K. oxytoca. They succeeded in identifying six K. oxytoca strains previously deposited as K. pneumoniae in international collections.
The last part of the studies concerned species identification by the use of MALDI-TOF mass spectrometry analysis, the method that is increasingly applied for Klebsiella species identification (Figure 3) (Bridel et al., 2021; Cuénod et al., 2021; Merla et al., 2019; Rodrigues et al., 2019). The species identification by the MALDI Biotyper is consistent with the one obtained by api 20E; however, the latter appeared to have a lower ability to discriminate than the MALDI Biotyper (Tables 2, 3). Moreover, one strain, 0.069, which could not be identified by api 20E, as defined by the MALDI Biotyper as K. pneumoniae with a high probability of identification (Tables 2, 3).
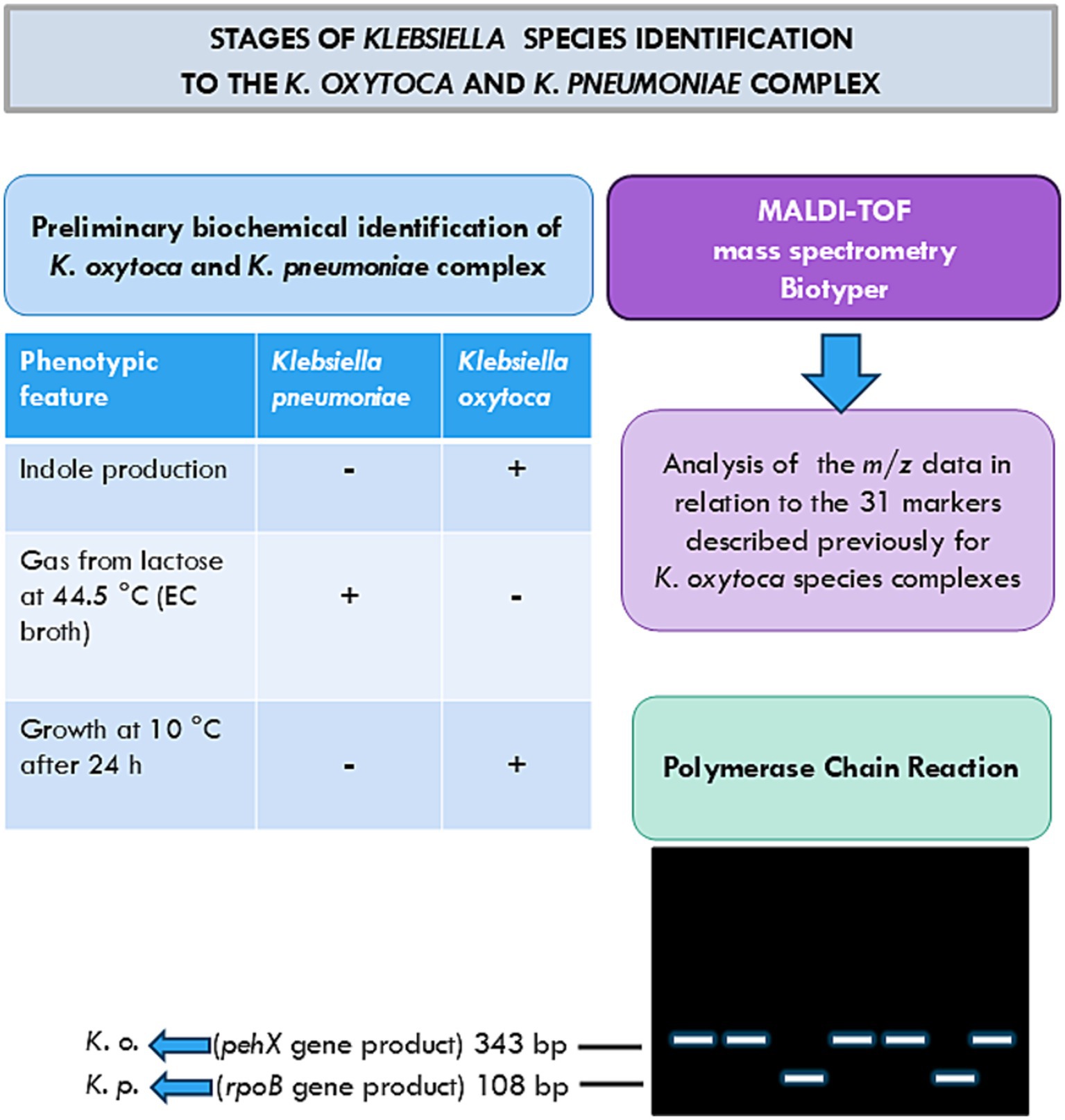
Figure 3. Stages of Klebsiella species identification by the K. oxytoca and K. pneumoniae complex used in these studies (Chander et al., 2011; Hansen et al., 2004; Kovtunovych et al., 2003; Merla et al., 2019).
One strain (0.011), biochemically identified as K. oxytoca, was designed by the MALDI Biotyper as Raoultella ornithinolytica with an identification rate equal to 2.27 (Table 3). Furthermore, Merla et al. identified seven strains of Ko3 and Ko8 as Raoultella ornithinolytica, which they explained by the lack of reference spectra of most phylogroups in the reference database. Brisse et al. (2006) noticed that the phylogenetic analysis of the rpoB gene sequence did not demonstrate the monophyly of Raoultella. Moreover, MALDI-TOF MS spectra and many biochemical features, including indole production, growth at 10°C, and gas production from lactose at 44.5°C are very similar for K. oxytoca and Raoultella ornithinolytica, which significantly hampers these species’ differentiation (Hansen et al., 2004; Merla et al., 2019).
Merla et al. (2019) also provided 31 identification biomarkers for all members of the K. oxytoca species complex based on a MALDI-TOF mass spectrometry analysis obtained for 30 strains from all phylogroups described so far: Ko1 (7), Ko2 (5), Ko3 (4), Ko4 (5), Ko6 (6), and Ko8 (3) (Figure 3). Analysis of the raw data obtained in the present study enabled the identification of a few m/z values assigned by Merla et al. (2019) to the Klebsiella oxytoca phylogroups. As Merla et al. observed, some peaks are shared by a few phylogroups; e.g., Ko4 shared most of its spectral peaks with Ko1 and Ko6, which can also be seen in Table 4 (Merla et al., 2019). To the Ko1, Ko4, Ko6, and Ko2 phylogroups, nine different m/z values were assigned: 2591, 5187, 3132, 6266, 3949, 7898, 6135, 7109, and 6767 (Merla et al., 2019). Thus, detecting these values in a strain may confirm its belonging to one of the mentioned phylogroups. For example, for K. oxytoca strains 0.021 and 0.061, four and three of these values were, respectively, detected (Table 4). Only for the Ko2, Ko3, and Ko8 phylogroups, specific peak combinations were proposed as biomarkers for discriminating each of these groups (Merla et al., 2019), and these peak positions could not be found in the present studies. For the K. oxytoca 0.013 and 0.071 strains, one specific peak was found (5052 m/z) reported by Merla et al. as that specific to Ko1 and Ko6 phylogroups.
The present studies indicate the MALDI-TOF mass spectrometry analysis as the most reliable method of species differentiation between K. oxytoca and K. pneumoniae groups, which also enables indicating a phylogroup within these groups. This method has been increasingly used for bacterial identification by clinical laboratories, as it is less costly than genetic identification through whole-genome sequencing, which requires DNA isolation and multi-stage bioinformatics analysis with specialized software. However, the assignment of the strains within the K. pneumoniae or K. oxytoca complex to an appropriate phylogroup is more difficult and requires reproducing similar parameters such as the standards, database, and software, since small differences in these conditions may result in slightly different m/z values. When K. pneumoniae (n = 10), K. variicola (n = 10), K. oxytoca (n = 8), K. michiganensis (n = 12), and K. grimontii (n = 10) were analyzed on four different MALDI-TOF MS systems in various laboratories, the reproducibility of detected marker masses and the sensitivity at the species level varied depending on the system, which was explained by differences in the quality of mass spectra. Mass spectral quality is crucial in distinguishing species within the K. oxytoca group, as the ribosomal marker masses are found in a high mass range (Cuénod et al., 2021). MALDI Biotyper library versions are very important, as differences between versions may lead to species misidentification. For example, Ohama et al. reported the misidentification of K. quasipneumoniae as K. pneumoniae when using MALDI Biotyper version 9, whereas version 57 provided the correct identification (Ohama et al., 2022). It is worth remembering that the phylogenetic division of the K. oxytoca and K. pneumoniae complexes into phylogroups has been made quite recently, and MALDI-TOF MS analyses using more strains are desirable. The Kleborate framework is also being increasingly applied, primarily for tracking pathogens and AMR determinants (Lam et al., 2021). Using this tool solely for taxonomic assignment to species and subspecies still requires whole-genome sequencing (WGS), which is not a fully accessible method. Nevertheless, WGS data provide the information important for both methods of species identification, Kleborate and MALDI-TOF MS, and enhance the database with species-specific phylogenetic markers.
In future studies, the authors will consider sequencing the whole genomes of the tested strains or sequencing the amplicons in the search for the phosphoenolpyruvate mutase gene, tctA (tripartite tricarboxylate transporter gene), diguanylate phosphodiesterase gene (for the identification of K. pneumoniae complex species), and blaOXY-(1–9) genes (for the identification of K. oxytoca complex species) (Cosic et al., 2021; Barrios-Camacho et al., 2021; Yang et al., 2022) (Supplementary Figure S2).
The original contributions presented in the study are included in the article/Supplementary material, further inquiries can be directed to the corresponding author/s.
AP: Conceptualization, Data curation, Investigation, Methodology, Validation, Visualization, Writing – original draft, Writing – review & editing. AM: Data curation, Methodology, Validation, Writing – original draft. JŁ: Data curation, Formal analysis, Supervision, Validation, Writing – review & editing.
The author(s) declare that financial support was received for the research, authorship, and/or publication of this article. This research was financially supported partially by the University of Lodz, Poland.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The authors declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
The Supplementary material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fmicb.2025.1514643/full#supplementary-material
Anbuselvi, S., Harinee, C., and Kalaivani,. (2017). Isolation of Klebsiella sp. from medical garbage soil and its effect in growth of chickpea plant. Int. J. Chem. Tech. Res. 10, 232–235.
Barrios-Camacho, H., Silva-Sánchez, J., Cercas-Ayala, E., Lozano-Aguirre, L., Morfín-Otero, R., Hernández-Castro, R., et al. (2021). PCR system for the correct differentiation of the main bacterial species of the Klebsiella pneumoniae complex arch. Microbiol 204:73. doi: 10.1007/s00203-021-02668-x
Bridel, S., Watts, S. C., Judd, L. M., Harshegyi, T., Passet, V., Rodrigues,, et al. (2021). Klebsiella MALDI TypeR: a web-based tool for Klebsiella identification based on MALDI-TOF mass spectrometry. Res. Microbiol. 172:103835. doi: 10.1016/j.resmic.2021.103835
Brisse, S., Grimont, F., and Grimont, P. A. D. (2006). “The genus Klebsiella” in The prokaryotes: A handbook on the biology of Bacteria volume 6: Proteobacteria: Gamma subclass. eds. M. Dworkin, S. Falkow, E. Rosenberg, K.-H. Schleifer, and E. Stackebrandt (New York, NY: Springer New York), 159–196.
Chander, Y., Ramakrishnan, M. A., Jindal, N., Hanson, K., and Goyal, S. M. (2011). Differentiation of Klebsiella pneumoniae and K. oxytoca by multiplex polymerase chain reaction. Intern. J. Appl. Res. Vet. Med. 9, 138–142.
Cosic, A., Leitner, E., Petternel, C., Galler, H., Reinthaler, F. F., Herzog-Obereder, K. A., et al. (2021). Variation in accessory genes within the Klebsiella oxytoca species complex delineates monophyletic members and simplifies coherent genotyping. Front. Microbiol. 12:692453. doi: 10.3389/fmicb.2021.692453
Cuénod, A., Wüthrich, D., Seth-Smith, H. M. B., Ott, C., Gehringer, C., Foucault, F., et al. (2021). Whole-genome sequence-informed MALDITOF MS diagnostics reveal importance of Klebsiella oxytoca group in invasive infections: a retrospective clinical study. Genome Med. 13:150. doi: 10.1186/s13073-021-00960-5
Ewing, W. H. (1986). Edwards and Ewing’s identification of Enterobacteriaceae. 4th Edn. New York: Elsevier 536.
Gómez, M., Valverde, A., del Campo, R., Rodríguez, J. M., and Maldonado-Barragán, A. (2021). Phenotypic and molecular characterization of commensal, community-acquired and nosocomial Klebsiella spp. Microorganisms 9:2344. doi: 10.3390/microorganisms9112344
Hansen, D. S., Aucken, H. M., Abiola, T., and Podschun, R. (2004). Recommended test panel for differentiation of Klebsiella species on the basis of a trilateral interlaboratory evaluation of 18 biochemical tests. J. Clin. Microbiol. 42, 3665–3669. doi: 10.1128/JCM.42.8.3665-3669.2004
Hasan, S. A., and Mohammed Bakr, M. (2022). Bacteriological and molecular detection of Klebsiella oxytoca and its resistance to antibiotics among clinical specimens from Kirkuk, Iraq. Archiv. Razi Institute. 77, 1521–1525. doi: 10.22092/ari.2022.357753.2095
Kovtunovych, G., Lytvynenko, T., Negrutska, V., Lar, O., Brisse, S., and Kozyrovska, N. (2003). Identification of Klebsiella oxytoca using a specific PCR assay targeting the polygalacturonase pehX gene. Res. Microbiol. 154, 587–592. doi: 10.1016/S0923-2508(03)00148-7
Lam, M. M. C., Wick, R. R., Watts, S. C., Cerdeira, L. T., Wyres, K. L., and Holt, K. E. (2021). A genomic surveillance framework and genotyping tool for Klebsiella pneumoniae and its related species complex. Nat. Commun. 12:4188. doi: 10.1038/s41467-021-24448-3
Martin, R. M., and Bachman, M. A. (2018). Colonization, infection, and the accessory genome of Klebsiella pneumoniae. Front. Cell. Infect. Microbiol. 8:4. doi: 10.3389/fcimb.2018.00004
Martínez, J., Martínez, L., Rosenblueth, M., Silva, J., and Martínez-Romero, E. (2004). How are gene sequence analyses modifying bacterial taxonomy? The case of Klebsiella. Int. Microbiol. 7, 261–268
Merla, C., Rodrigues, C., Passet, V., Corbella, M., Thorpe, H. A., Kallonen, T. V. S., et al. (2019). Description of Klebsiella spallanzanii sp. nov. and of Klebsiella pasteurii sp. nov. Front. Microbiol. 10:2360. doi: 10.3389/fmicb.2019.02360
Naemura, L. G., and Seidler, R. J. (1978). Significance of low-temperature growth associated with the fecal coliform response, indole production, and pectin liquefaction in Klebsiella. Appl. Environ. Microbiol. 35, 392–396. doi: 10.1128/aem.35.2.392-396.1978
Ohama, Y., Nomura, Y., Mizoguchi, M., Higurashi, Y., Okamoto, K., and Harada, S. (2022). Accurate identification of Klebsiella variicola by MALDI-TOF mass spectrometry in clinical microbiology laboratories. Microbiol. Spectr. 10, e02844–e02822. doi: 10.1128/spectrum.02844-22
Rodrigues, C., Passet, V., Rakotondrasoa, A., Diallo, T. A., Criscuolo, A., and Brisse, S. (2019). Description of Klebsiella africanensis sp. nov., Klebsiella variicola subsp. tropicalensis subsp. nov. and Klebsiella variicola subsp. variicola subsp. nov. Res. Microbiol. 170, 165–170. doi: 10.1016/j.resmic.2019.02.003
Singh, L., Cariappa, M. P., and Kaur, M. (2016). Klebsiella oxytoca: an emerging pathogen? Med. J. Armed Forces India. 72, 59–61. doi: 10.1016/j.mjafi.2016.05.002
Stewart, J., Judd, L. M., Jenney, A., Holt, K. E., Wyres, K. L., and Hawkey, J. (2022). Epidemiology and genomic analysis of Klebsiella oxytoca from a single hospital network in Australia. BMC Infect. Dis. 22:704. doi: 10.1186/s12879-022-07687-7
Wyres, K. L., Wick, R. R., Gorrie, C., Jenney, A., Follador, R., Thomson, N. R., et al. (2016). Identification of Klebsiella capsule synthesis loci from whole genome data. Microbial Genomics 2:e000102. doi: 10.1099/mgen.0.000102
Keywords: biochemical methods, Klebsiella oxytoca, Klebsiella pneumoniae, MALDI-TOF mass spectrometry, PCR, species identification
Citation: Palusiak A, Maciejewska A and Łukasiewicz J (2025) The results of polymerase chain reaction and MALDI-TOF mass spectrometry versus phenotypic distinction between Klebsiella pneumoniae and Klebsiella oxytoca. Front. Microbiol. 16:1514643. doi: 10.3389/fmicb.2025.1514643
Received: 21 October 2024; Accepted: 10 February 2025;
Published: 27 February 2025.
Edited by:
Jens Andre Hammerl, Bundesinstitut für Risikobewertung, GermanyReviewed by:
Sudhir Kumar, Iowa State University, United StatesCopyright © 2025 Palusiak, Maciejewska and Łukasiewicz. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Agata Palusiak, YWdhdGEucGFsdXNpYWtAYmlvbC51bmkubG9kei5wbA==
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.