
94% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
REVIEW article
Front. Microbiol. , 22 January 2019
Sec. Microbial Physiology and Metabolism
Volume 9 - 2018 | https://doi.org/10.3389/fmicb.2018.03264
This article is part of the Research Topic Novel Metabolic Engineering Approaches for Producing Novel Chemicals View all 21 articles
Plant biomass is a promising carbon source for producing value-added chemicals, including transportation biofuels, polymer precursors, and various additives. Most engineered microbial hosts and a select group of wild-type species can metabolize mixed sugars including oligosaccharides, hexoses, and pentoses that are hydrolyzed from plant biomass. However, most of these microorganisms consume glucose preferentially to non-glucose sugars through mechanisms generally defined as carbon catabolite repression. The current lack of simultaneous mixed-sugar utilization limits achievable titers, yields, and productivities. Therefore, the development of microbial platforms capable of fermenting mixed sugars simultaneously from biomass hydrolysates is essential for economical industry-scale production, particularly for compounds with marginal profits. This review aims to summarize recent discoveries and breakthroughs in the engineering of yeast cell factories for improved mixed-sugar co-utilization based on various metabolic engineering approaches. Emphasis is placed on enhanced non-glucose utilization, discovery of novel sugar transporters free from glucose repression, native xylose-utilizing microbes, consolidated bioprocessing (CBP), improved cellulase secretion, and creation of microbial consortia for improving mixed-sugar utilization. Perspectives on the future development of biorenewables industry are provided in the end.
Legitimate concerns regarding the negative environmental impact and unsustainability of the petrochemical industry have resulted in extensive exploration in microbial production of fuels and chemicals (Chu and Majumdar, 2012). Currently, the majority of industrialized biochemical processes utilize crop sugar as substrate, which is non-ideal for a multitude of reasons, including minimal reductions in greenhouse gas emissions, food vs. fuel controversy, and uncompetitive margins compared to petrochemical counterparts (Carriquiry et al., 2011; Du et al., 2012; Latimer et al., 2014). Research foci have shifted toward utilizing lignocellulosic feedstock for second-generation fuel and chemical production because of its abundance, sustainability, and low price.
Although the prospect of second-generation fuel and chemical production offers multiple environmental and socioeconomical advantages, its current economic state renders the biorenewables industry incapable of competing with the petrochemical industry (Carriquiry et al., 2011). Multiple technical hurdles must be overcome to efficiently convert lignocellulosic biomass to biofuels/biochemicals.
Plant biomass is mainly comprised of cellulose, hemicellulose, and lignin with proportions of each component depending on the sources of plant biomass (Fengel and Wegener, 1983; Betts et al., 1991). After pretreatment and saccharification processing, the resulting biomass hydrolysates are mixtures of various hexoses and pentoses. Next to glucose, xylose and arabinose are the second and third most abundant sugars in most terrestrial plant biomass hydrolysates, respectively (Chandel et al., 2011; Kim et al., 2012). Efficient utilization of all types of sugars from plant biomass hydrolysates is essential for economic conversion of plant biomass to fuels and chemicals (Saha, 2003). Most microorganisms, including Saccharomyces cerevisiae and Escherichia coli, can efficiently utilize glucose as the optimal fermentation substrate, whereas non-glucose sugars such as xylose and arabinose are utilized at much lower efficiencies (Jojima et al., 2010).
Because of carbon catabolite repression, nearly all microorganisms utilize glucose preferentially over other sugars, resulting in extended fermentation periods and lower productivities (Zaldivar et al., 2001). Post-glucose depletion, other nutritional elements gradually become limited and certain fermentation end-products and/or side products that inhibit cell growth begin to accumulate in the culture media, eventually leading to the slow and incomplete fermentation of non-glucose sugars (Palmqvist et al., 1999; Graves et al., 2006). To address this issue, it is imperative to engineer microorganisms capable of simultaneous co-sugar utilization. Currently, microbial mixed-sugar conversion has been mostly studied in model microbial platforms such as E. coli and S. cerevisiae because of their well-understood physiology and genetic backgrounds, fast cell growth rates, and readily available genetic manipulation tools. Moreover, current commercial production of ethanol from sugarcane or cornstarch employs S. cerevisiae, and it was predicted that the integration of existing ethanol plants with engineered S. cerevisiae that produce biofuels from non-edible plant biomass can reduce the total cost by as much as 20% (Wooley et al., 1999; Peters et al., 2003).
This review describes recent advances in microbial conversion of mixed sugars from plant biomass, mainly focusing on S. cerevisiae. Various strategies used to ameliorate simultaneous mixed-sugar fermentation, including pathway introduction and optimization, discovery and engineering of novel specific transporters for special sugars, consolidated bioprocessing, and creation of microbial consortia, are discussed in detail. The vast majority of the studies discussed in this review analyzed bioethanol production. In these contexts, bioethanol is simply used as a reporter molecule for assessing mixed-sugar assimilation competence. In alignment with the broader scope laid out by the Department of Energy's Biomass Program (Aden et al., 2004), it is the hope that engineered strains can serve as microbial platforms to produce a wide array of biochemicals.
Xylose, the second most abundant sugar in plant biomass, is metabolized by microorganisms mainly via two distinct routes (Figure 1). In native xylose-utilizing bacteria, some fungi, and plants, xylose is converted to D-xylulose by xylose isomerase (XI or XylA) in a single step (Schellenberg et al., 1984; Wilhelm and Hollenberg, 1984; Banerjee et al., 1994; Kristo et al., 1996; Rawat et al., 1996; Maehara et al., 2013), whereas in most innate xylose-utilizing fungi, a more complex alternative route consisting of two redox reactions exists. Xylose is first reduced to xylitol by a NADPH-preferred xylose reductase (XR). The resulting xylitol is then oxidized to D-xylulose by NAD+-dependent xylose dehydrogenase (XDH) (Chakravorty et al., 1962; Bruinenberg et al., 1984). Subsequently, D-xylulose derived from either pathway is phosphorylated by a xylulokinase (XKS) into D-xylulose 5-phosphate (D-X5P), which is then channeled into the pentose phosphate pathway (PPP) (Xue and Ho, 1990; Rodriguez-Pena et al., 1998; Hahn-Hägerdal et al., 2007).
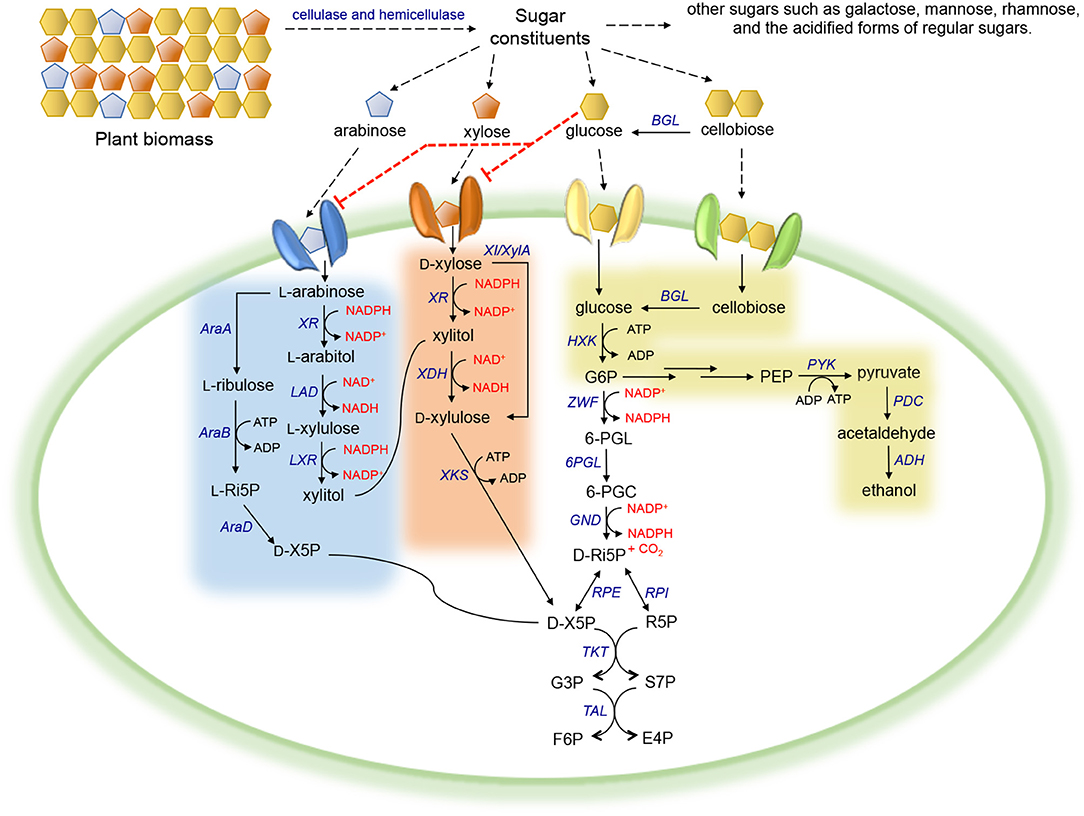
Figure 1. Carbohydrate metabolism in microorganisms. Red dotted line corresponds to inhibition. Abbreviation of metabolites—PEP, phosphoenolpyruvate; G6P, glucose-6-phosphate; 6-PGL, 6-phosphogluconolactone; 6-PGC, 6-phosphogluconate; D-Ri5P, D-ribulose-5-phosphate; D-X5P, D-xylulose-5-phosphate; R5P, ribose-5-phosphate; G3P, glyceraldehyde-3-phosphate; S7P, sedoheptulose-7-phosphate; F6P, fructose-6-phosphate; E4P, erythrose-4-phosphate; L-Ri5P, L-ribulose-5-phosphate. Abbreviation of enzymes—BGL, β-glucosidase; HXK, hexokinase; PYK, pyruvate kinase; PDC, pyruvate decarboxylase; ADH, alcohol dehydrogenase; ZWF, glucose-6-phosphate dehydrogenase; 6PGL, 6-phosphogluconolactonase; GND, 6-phosphogluconate dehydrogenase; RPI, ribose-5-phosphate isomerase; RPE, ribulose-5-phosphate epimerase; TKT, transketolase; TAL, transaldolase; XR, xylose reductase; XDH, xylose dehydrogenase; XKS, xylulokinase; XI/XylA, xylose isomerase; LAD, L-arabitol 4-dehydrogenase; LXR, L-xylulose reductase; AraA, L-arabinose isomerase; AraB, L-ribulokinase; AraD, L-ribulose-5-phosphate 4-epimerase.
Despite its broad industrial applications, S. cerevisiae cannot natively utilize xylose hydrolyzed from plant biomass, although gene homologs encoding XR, XDH, and XKS required for xylose metabolism are present in its genome (Hahn-Hägerdal et al., 2007). Overexpression of these native genes allowed for minimal cell growth on xylose (Toivari et al., 2004). Even after extensive evolution, S. cerevisiae strains with endogenous xylose metabolic pathways still could not metabolize xylose as efficiently as glucose (Attfield and Bell, 2006). This was mainly attributed to the imbalanced xylose-utilizing pathway, where the activities of XR and XDH were much lower compared to that of XKS.
To overcome this limitation, heterologous xylose-utilizing pathways were introduced into S. cerevisiae. First, despite its inability to ferment xylose, S. cerevisiae can grow on D-xylulose (Chiang et al., 1981), indicating that simply introducing a heterologous XI enables xylose utilization. The first highly functional Piromyces XI gene (Harhangi et al., 2003) that was introduced into S. cerevisiae conferred a specific growth rate of 0.005 h−1 on xylose under aerobic conditions (Kuyper et al., 2003, 2004). Continuous evolution in xylose media resulted in a mutant strain with improved growth rates of 0.18 h−1 under aerobic conditions and 0.03 h−1 under anaerobic conditions. The anaerobic ethanol yield from xylose was as high as 0.42 g g−1. Brat et al. identified the highly active Clostridium phytofermentans XI, a distant homolog of Piromyces XIs (Brat et al., 2009). Introducing a codon-optimized version into an industrial S. cerevisiae strain enabled an aerobic cell growth rate of 0.057 h−1 and anaerobic ethanol yield of 0.43 g g−1 when cultured in xylose. Subsequently, XIs from a series of species showing high similarities with Piromyces XI or C. phytofermentans XI were actively expressed in S. cerevisiae (Hahn-Hägerdal et al., 2007; Madhavan et al., 2009; Aeling et al., 2012; Hector et al., 2013; Peng et al., 2015). Particularly, through evolutionary engineering, XIs from Prevotella ruminicola (Hector et al., 2013), Bacteroides vulgatus, and Alistipes sp. HGB5 (Peng et al., 2015) displayed comparable enzyme activities to the best-reported XI from Piromyces. To date, many functionally expressed XIs in S. cerevisiae originated from mammal gut microflora, suggesting the evolutionary advantage of xylan-degrading bacteria present in mammalian guts. These findings have led to mammalian gut commensals serving as a repertoire for isolating novel genes involved in plant biomass degradation at standard temperatures (Peng et al., 2015).
In parallel to the XI pathway, the XR/XDH pathway has also been extensively studied in S. cerevisiae in the past few decades. Multitudinous efforts have been made to introduce genes encoding XR and XDH from natural xylose-assimilating species including Scheffersomyces stipitis, Neurospora crassa, and Candida tenuis into S. cerevisiae, leading to recombinant strains that grew aerobically on xylose media (Kotter et al., 1990; Kotter and Ciriacy, 1993; Tantirungkij et al., 1993; Jin et al., 2000; Petschacher and Nidetzky, 2008). However, xylose utilization was very slow in these recombinant strains and a large amount of xylitol accumulated, resulting in a low ethanol yield (Kotter and Ciriacy, 1993; Tantirungkij et al., 1993; Jin et al., 2000). Further investigation attributed this low xylose-utilizing efficiency to two main factors. The cofactor imbalance of XR and XDH was severe under oxygen-limited and anaerobic conditions, which resulted in excess formation of NADP+ and a shortage of NAD+. Because it lacked a transhydrogenase, S. cerevisiae cannot directly convert NADP+ to NAD+ (Hahn-Hägerdal et al., 2007). This imbalance was thought to result in xylitol by-product formation, pulling metabolic flux away from ethanol production (Karhumaa et al., 2007). Moreover, accumulation of PPP intermediates during xylose fermentation indicated that the capacity of the downstream PPP was constrained by PPP enzyme insufficiency (Kotter and Ciriacy, 1993). In the next two sections, strategies that have been implemented to improve xylose conversion in S. cerevisiae are discussed in detail.
To mitigate the cofactor imbalance issue, numerous efforts have focused on either isolating novel NADH-preferred XR or switching the cofactor specificities of existing XRs (Kang et al., 2003; Khoury et al., 2009) (Figure 2, strategy I). For example, the cofactor specificity of C. boidinii XR was altered through computational design of the cofactor binding pocket, and subsequent characterization validated that seven mutants were more specific to NADH (Khoury et al., 2009). Four of the seven mutants completely lost activity when NADPH was the only available cofactor, and more than 104-fold changes in substrate specificity from NADPH to NADH were observed for the mutant CbXR-K272R, S273E, N274G. Expression of the mutant XRs in S. cerevisiae containing the wild-type XDH resulted in increased xylose consumption rates, ethanol titers, and yields along with decreased xylitol levels (Jeppsson et al., 2006; Watanabe et al., 2007; Petschacher and Nidetzky, 2008; Bengtsson et al., 2009). Analogously, this strategy was applied to modify the specificity of XDH to NADP+ (Metzger and Hollenberg, 1995; Watanabe et al., 2005). Metzger et al. attempted to identify the NAD+-binding domain of S. stipitis XDH, and introduced a putative NADP+-recognition sequence (GSRPVC) of the alcohol dehydrogenase from Thermoanaerobium brockii (Metzger and Hollenberg, 1995). As a result, the mutant enzyme used both NAD+ and NADP+ as cofactors with equal apparent Km values. The most successful work in changing the preference of XDH from NAD+ to NADP+ was performed by introducing triple (D207A/I208R/F209S) or quadruple (D207A/I208R/F209S/N211R) mutations into S. stipitis XDH (Watanabe et al., 2005) (Figure 2, strategy II). The resulting XDH variants displayed a shifted specificity toward NADP+, which was more than 4,500-fold higher than that displayed by the wild-type enzyme in vitro. Functional expression of the mutants in S. cerevisiae resulted in a maximal 86% decrease in xylitol accumulation coupled with a 41% increase in ethanol production compared to the control strain containing wild-type XDH (Watanabe et al., 2007).
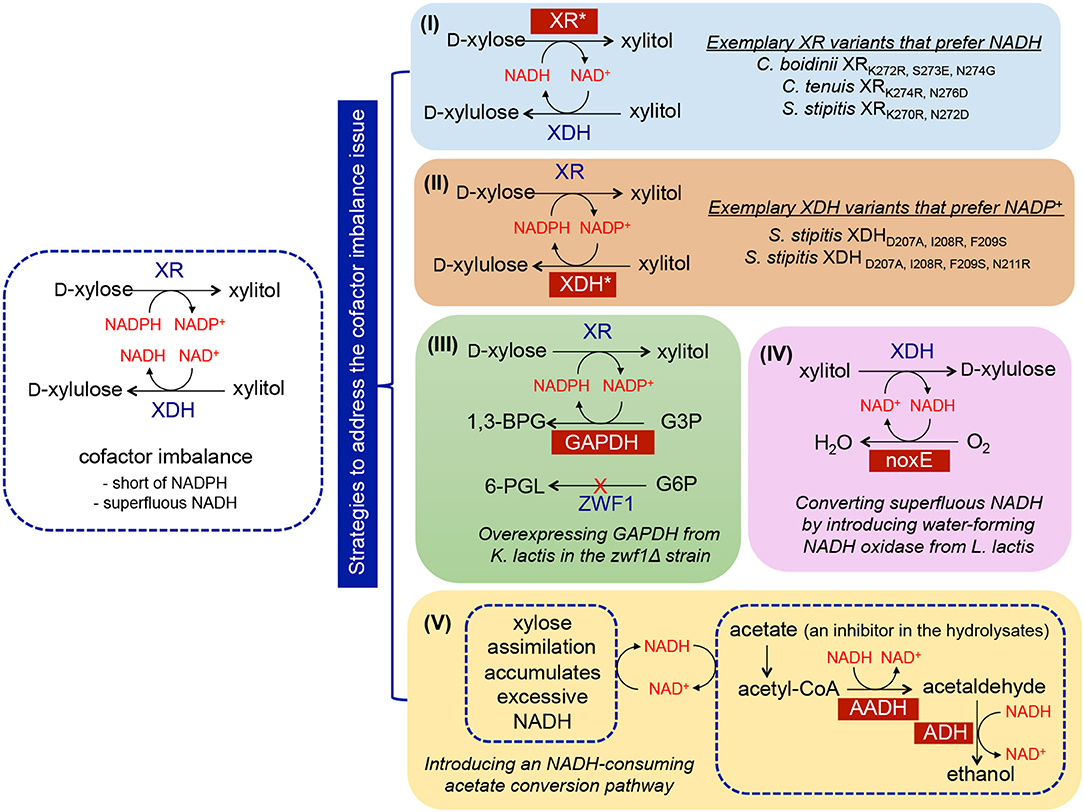
Figure 2. Implementation of metabolic engineering strategies in S. cerevisiae to enhance xylose utilization via ameliorating cofactor imbalance, including (I) discovery of NADH-preferred XR variants; (II) discovery of NADP+-preferred XDH variants; (III) enhancing NADPH regeneration through overexpressing NADP+-dependent GAPDH; (IV) enhancing NAD+ regeneration through overexpressing noxE; (V) improving NAD+ regeneration through the integration of acetate-consuming pathway. Abbreviation of metabolites−1,3-BPG, 1,3-bisphosphoglycerate; 6-PGL, 6-phosphogluconolactone; G6P, glucose-6-phosphate; G3P, glyceraldehyde-3-phosphate. Abbreviation of enzymes—XR, xylose reductase; XDH, xylose dehydrogenase; ZWF, glucose-6-phosphate dehydrogenase; GAPDH, glyceraldehyde-3-phosphate dehydrogenase; noxE, NADH oxidase; AADH, acetylating acetaldehyde dehydrogenase; ADH, alcohol dehydrogenase.
A viable alternative strategy for enabling cofactor balance in S. cerevisiae containing the XR/XDH pathway is to enhance the regeneration of NADPH or NAD+. In wild-type yeast, NADPH is mainly regenerated via the oxidative portion of the PPP, in which two of the three enzymes involved in converting glucose-6-phosphate (G6P) to D-ribulose-5-phosphate (D-Ri5P) utilize NADP+ as a cofactor (Figure 1). However, this conversion is coupled with CO2 formation, inevitably resulting in significant carbon loss and low product yield (Verho et al., 2003). To reduce the heavy reliance on the PPP to regenerate NADPH, a yeast GPD1 gene, encoding NADP+-dependent D-glyceraldehyde-3-phosphate dehydrogenase (GAPDH) originated from Kluyveromyces lactis, was introduced into S. cerevisiae containing the XR/XDH pathway (Verho et al., 2003) (Figure 2, strategy III). Compared to S. cerevisiae containing endogenous NAD+-dependent GAPDH, overexpression of K. lactis GAPDH converted D-glyceraldehyde-3-phosphate (G3P) to 1, 3-bisphosphoglycerate using NADP+ as a cofactor, which is beneficial for both regenerating NADPH and reducing the accumulation of excessive NADH produced by endogenous GAPDH. When the ZWF1 gene encoding glucose-6-phosphate dehydrogenase was simultaneously deleted, the K. lactis GAPDH became the major engine replenishing NADPH in recombinant S. cerevisiae, resulting in higher ethanol production rates and yields from xylose compared to those in the control strain. The same strategy was applied to improve NAD+ availability by heterologously expressing Lactococcus lactis NADH oxidase in S. cerevisiae containing the XR/XDH pathway, which mediated the conversion of NADH and oxygen to NAD+ and water (Heux et al., 2006; Vemuri et al., 2007; Zhang et al., 2012; Hou et al., 2014) (Figure 2, strategy IV). Introduction of the water-forming NADH oxidase mitigated this cofactor imbalance by converting excess intracellular NADH to NAD+ (Heux et al., 2006; Vemuri et al., 2007). During xylose fermentation, the recombinant strain overexpressing NADH oxidase from L. lactis demonstrated a 69.63% decrease in xylitol accumulation, a 53.85% reduction in glycerol production, and a 39.33% increase in ethanol yield (Zhang et al., 2012).
In addition to NADH oxidase, other NAD+-regenerating pathways were explored. For example, Wei et al. confirmed that in yeast, acetate can be converted to ethanol catalyzed by the endogenous acetyl-CoA synthetase and alcohol dehydrogenase (ADH) in conjunction with the heterologous acetylating acetaldehyde dehydrogenase (AADH) using NADH as a cofactor (Wei et al., 2013) (Figure 2, strategy V). A group of AADH homologs was tested, including bifunctional proteins isolated from E. coli, Piromyces Sp. E2, and Clostridium beijerinckii (catalyzing the conversion of acetyl-CoA to acetaldehyde followed by subsequent conversion to ethanol), and the proteins involved in ethanolamine or 4-hydroxy-2-ketovalerate catabolism from E. coli. Introduction of these homologs into S. cerevisiae harboring the XR/XDH pathway enabled simultaneous acetate and xylose consumption under anaerobic conditions. The highest improvement was achieved by overexpressing the bifunctional E. coli AdhE, which led to 17% and 21% increases in ethanol yield and productivity, respectively. This improvement was attributed to the increased regeneration of NAD+ (by 50%) and decreased levels of glycerol and xylitol byproducts. The superiority of this system is conferred by co-utilization of xylose and acetate, the latter of which is a ubiquitous inhibitor present in plant biomass hydrolysates and serves as a redox sink for consuming surplus NADH.
Although D-xylulose, the product of XR/XDH-catalyzed conversion, can be metabolized by S. cerevisiae, it is consumed much more slowly than glucose (Senac and Hahn-Hagerdal, 1990; Yu et al., 1995). Numerous independent studies indicated that merely introducing XI or XR/XDH resulted in substantial D-xylulose accumulation in S. cerevisiae (Jin and Jeffries, 2003; Kuyper et al., 2004; Madhavan et al., 2009). These observations suggested insufficient activity of endogenous XKS. To overcome this inadequacy, genes encoding XKS from multiple sources were expressed in recombinant S. cerevisiae harboring either the XI or XR/XDH pathway (Moniruzzaman et al., 1997; Johansson et al., 2001; Toivari et al., 2001; Madhavan et al., 2009; Zhou et al., 2012). Notably, excessive XKS activity may increase metabolic burden and cause excessive ATP consumption, impairing cell growth and diminishing ethanol yield (Johansson et al., 2001; Jin et al., 2003). In this regard, fine-tuned XKS activity is essential for optimizing xylose fermentation in recombinant S. cerevisiae strains. As a result, moderate XKS activity coupled with the high activity of XR and XDH was optimal for xylose conversion (Eliasson et al., 2001; Matsushika and Sawayama, 2008).
The PPP is the entry point for integrating D-xylulose into central carbon metabolism. Therefore, the activity of PPP enzymes is crucial for efficient pentose assimilation (Hahn-Hägerdal et al., 2007). Because of the low capacity of the non-oxidative portion of the PPP (Fiaux et al., 2003), S. cerevisiae containing a xylose-assimilating pathway accumulated PPP intermediates, including 6-phosphogluconate (6-PGC), D-X5P, D-Ri5P, ribose-5-phosphate (R5P), and sedoheptulose-7-phosphate (S7P) (Senac and Hahn-Hagerdal, 1990; Kotter and Ciriacy, 1993; Matsushika et al., 2013) (Figure 1). In addition to strategies of elevating the expression level of the endogenous transketolase (TKT1) and transaldolase (TAL1), researchers have attempted to overexpress the homologs of the key enzymes involved in the PPP to improve pathway flux in xylose-fermenting yeast. Some previous studies reported that overexpression of S. stipitis TKT1 in S. cerevisiae containing the XR/XDH pathway did not affect xylose utilization, and in some cases, imposed significant metabolic burden and drastically extended the cell-doubling time on xylose (Metzger and Hollenberg, 1994; Walfridsson et al., 1995). In contrast, other studies indicated that overexpression of either the endogenous or heterologous TAL1 from S. stipitis in xylose-utilizing S. cerevisiae enhanced cell growth on xylose (Walfridsson et al., 1995; Jin et al., 2005). Specifically, S. cerevisiae transformed with XR, XDH, XKS, and TAL1 isolated from S. stipitis grew twice as fast on xylose and produced 70% more ethanol compared to the control strain overexpressing only XR/XDH/XKS (Jin et al., 2005). Moreover, two parallel studies reported the overexpression of genes encoding the entire non-oxidative PPP, including ribose-5-phosphate isomerase (RPI), ribulose-5-phosphate epimerase (RPE), TKT1, and TAL1, in the recombinant xylose-fermenting S. cerevisiae, along with deletion of the gene encoding aldose reductase (GRE3) that mediates unwanted production of xylitol (Karhumaa et al., 2005; Kuyper et al., 2005). The resulting strains demonstrated improved cell growth rate (0.22 h−1 and 0.17 h−1) and enhanced ethanol production (0.43 g g−1 and 0.29 g g−1) on xylose (Figure 1).
Arabinose is the second most abundant pentose in hemicellulose (Madhavan et al., 2012). Microorganisms metabolize arabinose to D-X5P, an intermediate of the PPP, via two distinct pathways (Figure 1) (Richard et al., 2003). Bacteria utilize arabinose isomerase (AraA), L-ribulokinase (AraB), and L-ribulose-5-phosphate 4-epimerase (AraD) to convert arabinose to D-X5P through intermediates L-ribulose and L-Ri5P (Wisselink et al., 2007). The disparate fungal alternative arabinose pathway converts arabinose to D-X5P through 5 steps via four intermediates (i.e., L-arabitol, L-xylulose, xylitol, and D-xylulose), and the corresponding reactions are catalyzed by aldose reductase (GRE3 or XR), L-arabitol 4-dehydrogenase (LAD), L-xylulose reductase (LXR), XDH, and XKS (Chiang and Knight, 1961; Witteveen et al., 1989; Richard et al., 2001, 2002). In natural arabinose-assimilating microorganisms (e.g., S. stipitis and most Candida species), L-arabitol accumulation resulted in very low ethanol titers and yields (Mcmillan and Boynton, 1994; Dien et al., 1996). Moreover, the unclear genetic background and unavailable genetic manipulation tools limit the direct application of most natural arabinose-utilizing microorganisms as production hosts in the biorenewables industry. To date, studies of arabinose utilization were mainly conducted in model microbial hosts.
Similar to xylose, arabinose does not support the growth of wild-type S. cerevisiae (Hahn-Hägerdal et al., 2007). Attempts have been made to enable arabinose utilization in S. cerevisiae by introducing the fungal arabinose pathway (Richard et al., 2003; Verho et al., 2004; Bettiga et al., 2009; Bera et al., 2010). Despite improved arabinose utilization through pathway optimization, the accumulation of L-arabitol indicated an imbalance in redox cofactors during arabinose fermentation in the engineered S. cerevisiae. The fungal arabinose pathway comprises two NADPH-dependent reductions and two NAD+-dependent oxidations. Although the expression of an NADH-dependent XR from Ambrosiozyma monospora mitigated the co-factor imbalance, excessive L-arabitol accumulation resulted in low ethanol yield (Verho et al., 2004; Bettiga et al., 2009).
Unlike its fungal counterpart, the bacterial arabinose pathway evades the cofactor imbalance and represents a viable alternative to enable arabinose utilization in S. cerevisiae. However, direct introduction of the bacterial arabinose pathway from E. coli, Bacillus subtilis or Lactobacillus plantarum into S. cerevisiae did not support robust cell growth on arabinose until extensive evolutionary engineering was applied (Sedlak and Ho, 2001; Becker and Boles, 2003; Wisselink et al., 2007, 2009; Sanchez et al., 2010; Wang et al., 2013, 2017). For example, Wisselink et al. built a recombinant S. cerevisiae strain (IMS001) containing the arabinose pathway from L. plantarum, Piromyces sp. strain E2 XI, and yeast XKS gene, coupled with overexpression of endogenous non-oxidative PPP genes, including TAL1, TKL1, RPE1, and RPI1 (Wisselink et al., 2007). However, IMS001 did not grow on arabinose, although expression of the arabinose pathway was verified by real-time PCR, albeit some genes expressed at very low levels. Only after evolutionary engineering, could the resulting strain IMS002 anaerobically grow on arabinose with a sugar consumption rate of 0.7 g h−1 g−1 drybiomass and ethanol productivity of 0.29 g h−1 g−1 dryweight. In anaerobic glucose/arabinose co-fermentation, arabinose was utilized upon glucose depletion at a rate of 0.42 g h−1 g−1 drybiomass with an ethanol yield of 0.42 g g−1. Subsequently, IMS002 was subjected to another round of evolutionary engineering to regain the xylose-assimilating ability lost during long-term selection for improved arabinose fermentation (Sanchez et al., 2010). The resulting strain converted a mixture of glucose, xylose, and arabinose to ethanol, achieving a yield of 0.43 g g-1, with a minimal byproduct accumulation (xylitol and L-arabitol) Very recently, a novel arabinose pathway from Pediococcus pentosaceus, a native arabinose-utilizing bacterium, and an arabinose transporter from Spathaspora passalidarum were isolated and introduced into S. cerevisiae (Caballero and Ramos, 2017). After one round of evolutionary engineering, arabinose was metabolized aerobically. A second round of evolutionary adaption under anaerobic conditions was carried out and the resulting strain demonstrated anaerobic growth and an ethanol production yield as high as 0.43 g g−1. To date, the arabinose-utilizing efficiency and ethanol yield have been largely improved, albeit glucose repression on arabinose utilization remains unchanged (Wisselink et al., 2007; Wang et al., 2017).
Initial efforts devoted to improving xylose assimilation were mainly focused on increasing intracellular pentose utilization. Although these efforts enhanced intracellular pentose flux into central metabolism, it was recognized that sugar uptake constrained achievable xylose assimilation (Jojima et al., 2010; Young et al., 2010, 2014). Because S. cerevisiae lacks xylose-specific transporters, the native hexose transporters, including HXT1, HXT2, HXT4, HXT5, HXT7, and GAL2, are responsible for mediating xylose transport (Hamacher et al., 2002; Sedlak and Ho, 2004; Saloheimo et al., 2007; Young et al., 2011). However, these hexose transporters possess 10-100 fold higher affinity for glucose compared to other sugars. This resultant glucose preference extends mixed-sugar fermentation process time (Van Vleet and Jeffries, 2009; Young et al., 2010; Kim et al., 2012). Therefore, mining and/or engineering xylose-specific transporters insensitive to glucose is essential for economic conversion of mixed sugars present in biomass hydrolysates.
The first non-native xylose transporters successfully expressed in S. cerevisiae were GXF1 and GXS1 from Candida intermedia (Leandro et al., 2006). Both transporters supported cell growth on xylose in the hexose transporter-null S. cerevisiae strain EBY.VW4000, which was transformed with a xylose-utilizing pathway. EBY.VW4000, deprived of 18 hexose transporters and three maltose transporters that transport hexoses, has been used as a platform for characterizing pentose transporters (Wieczorke et al., 1999). Although kinetic studies indicated that GXF1 is primarily a hexose transporter, its affinity toward xylose is 3-fold higher than those of native S. cerevisiae hexose transporters (Leandro et al., 2006; Runquist et al., 2009). Taking advantage of this property, GXF1 was expressed in xylose-fermenting S. cerevisiae strains, leading to faster xylose uptake rates (1.4–4-fold) and improved cell growth rates (1.7–4-fold) in xylose fermentation compared to the strain harboring only a xylose-utilizing pathway. Other putative xylose transporters from diverse organisms, including plants, algae, fungi, and bacteria, were expressed in S. cerevisiae to improve xylose uptake (Saloheimo et al., 2007; Hector et al., 2008; Katahira et al., 2008; Runquist et al., 2009, 2010; Du et al., 2010; Young et al., 2011). Because of potential improper localization or misfolding, many of these proteins did not enable xylose uptake in S. cerevisiae despite their high homology to known xylose transporters. For example, using GXS1 as a query, a study aimed to discover novel xylose transporters from native xylose-assimilating yeasts, S. stipitis and N. crassa (Du et al., 2010). Among a total of 18 putative xylose transporters, An25 and Xyp29 were identified as xylose-specific. Their expression on the membranes of S. cerevisiae was confirmed and xylose was accumulated intracellularly because of the scarcity of an efficient xylose-utilizing pathway in the engineered S. cerevisiae. In a subsequent study, a collection of 36 putative sugar transporters was thoroughly characterized for their substrate acceptance profiles in the EBY.VW4000 strain expressing XR and XDH from S. stipitis (Young et al., 2011). Growth-based assays demonstrated that three transporters, XUT1 and XUT3 from S. stipitis and XylHP from Debaryomyces hansenii, transported xylose efficiently and demonstrated higher xylose preference than other transporters in the hexose transporter-null strain background.
Despite the performance of these transporters in xylose media, all of these transporters were subjected to glucose repression in mixed-sugar fermentation or suffered from low xylose-uptake efficiency (Hector et al., 2008; Katahira et al., 2008; Runquist et al., 2009, 2010; Du et al., 2010; Young et al., 2011). In this regard, to overcome CCR and promote efficient biochemical production, numerous independent studies were conducted to reshape the existing transporters via rational design or evolutionary engineering (Young et al., 2012, 2014; Farwick et al., 2014; Nijland et al., 2014; Shin et al., 2015; Li et al., 2016; Reider Apel et al., 2016; Wang et al., 2016). Presumably, glucose repression can be exerted outside of the xylose-binding pocket. The very first transporter engineering work was performed by Young et al. (2012). Starting with C. intermedia GXS1 and S. stipitis XUT3, five mutants with improved xylose uptake activities were generated by employing a directed evolution method. Kinetic studies indicated that some mutated transporters obtained increased affinity to xylose (lower Km) and higher uptake rates (higher Vmax), with the best mutant amplifying the yeast cell growth rate on xylose by 1.7-fold compared to its corresponding wild-type. Relieved CCR and enhanced sugar assimilation were observed in mixed-sugar fermentation. The phenylalanine residue at position 40 of GXS1 and glutamine residue at position 538 of XUT3 were found to largely contribute to influencing the transportation characteristics.
Led by the observation that sugar transporter preference does not require a trade-off for its efficiency, multiple primary hexose transporters were engineered to be xylose-exclusive (Farwick et al., 2014; Nijland et al., 2014; Young et al., 2014; Shin et al., 2015; Li et al., 2016; Reider Apel et al., 2016; Wang et al., 2016). After evaluating 46 heterologous transporters, a conserved protein motif G-G/F-XXX-G surrounding the previously reported GXS1-F40 residue was recognized to dominate the transporters displaying high efficiency in xylose transport (Young et al., 2014). Through a series of saturation mutagenesis followed by rational mutagenesis, four xylose-exclusive transporters variants, CiGXS1F38I39M40, SsRGT2F38, SsRGT2M40, and ScHXT7I39M40M340, were obtained. The four mutants conferred robust cell growth on xylose in EBY.VW4000, albeit still inhibited by glucose, presumably via allosteric binding. Nearly concurrently, Farwick et al. built an easy yet precise xylose-specific transporter screening system by disrupting the first step of the glycolytic pathway (Farwick et al., 2014). Through growth-based screening under mixed-sugar conditions, a combinatorial strategy of evolutionary engineering and site-specific mutagenesis led to the discovery of several mutations that contributed to mitigating glucose inhibition. Specifically, simply altering threonine 219/213 and asparagine 376/370 of S. cerevisiae GAL2/HXT7 resulted in mutant transporters possessing extraordinary properties, including attenuated or completely abolished glucose affinity, enhanced xylose affinity, and decreased glucose competitive repression. This is the first study in which primary hexose transporters were engineered to be both glucose-insensitive and xylose-specific. Subsequently, various independent studies reported that hexose transporters such as S. cerevisiae HXT36 (Nijland et al., 2014), HXT11 (Shin et al., 2015), HXT7 (Reider Apel et al., 2016), C. intermedia GXS1 (Li et al., 2016), and pentose transporter [e.g., N. crassa An25 (Wang et al., 2016)] were engineered to relieve glucose inhibition and improve xylose uptake. These transporter-engineering studies suggested that direct evolution and combinatorial mutagenesis are potent protein engineering tools for expanding the collection of pentose-favoring or pentose-specific transporters.
To facilitate uptake and in situ intracellular conversion of xylose, a very intriguing study was recently conducted to construct an artificial enzyme-transporter complex in S. cerevisiae (Thomik et al., 2017). After a series of optimizations, a scaffold-based complex successfully recruited the heterologously expressed C. phytofermentans XI to the endogenous GAL2 localized on the membrane, which did not negatively affect the activities of both proteins. Because of the ameliorated substrate transportation and channeling, the xylose consumption rate and ethanol titer were remarkably increased with substantially diminished accumulation of xylitol, the undesired byproduct.
Most recent studies used EBY.VW4000 or its derivatives as the transporter-screening platform. However, this strain grows slowly because of its defects in meiosis and centromere segregation caused by large chromosome translocations (Solis-Escalante et al., 2015). In this regard, EBY.VW4000 is a good strain for transporter screening but not for industrial mixed-sugar conversion. Moreover, as described previously, xylose fermentation in engineered S. cerevisiae is accompanied by co-factor imbalance and subsequently low metabolic flux. Thus, a highly promising alternative to engineering industrial friendly model hosts to efficiently utilize mixed sugars is ameliorating innate xylose-utilizing microorganisms (Alper and Stephanopoulos, 2009). The CUG clade is a group of nonconventional yeasts that translate the CUG codon to serine rather than leucine (Santos et al., 2011). Unlike S. cerevisiae, most CUG clade yeasts can uptake and metabolize xylose much more efficiently. Specifically, S. stipitis and S. passalidarum are particularly intriguing because of their ability to convert xylose to ethanol at high yields and efficiencies (Van Vleet and Jeffries, 2009; Su et al., 2015). For instance, a newly isolated S. passalidarum strain, CMUWF1-2, converted xylose to ethanol with a yield of 0.43 g ethanol g−1 xylose and exhibited minimal glucose repression in a mixed-sugar assay (Rodrussamee et al., 2018). However, because of the dominant non-homologous end joining mechanism for repairing DNA double strand breaks and a lack of genetic manipulation tools, the exploration pace of CUG clade yeast has been limited, preventing their applications in industrial biofuel/biochemical production. Recent advances in synthetic biology have greatly expedited progress in engineering non-conventional yeasts (Lobs et al., 2017). For example, a series of genetic manipulation tools for S. stipitis were reported, including strong and constitutive promoters and terminators (Gao et al., 2017), a stable episomal expression plasmid created by successful isolation of native centromeres (Cao et al., 2017a,c), and CRISPR-enabled precise genome editing tools (Cao et al., 2017b). Through utilization of these tools, studies have demonstrated the possibility of engineering S. stipitis to an efficient platform in the near future for producing a specific group of biochemicals [i.e., shikimate pathway derivatives (Suastegui and Shao, 2016; Gao et al., 2017)] or biofuels directly from biomass hydrolysates containing hexoses and pentoses.
In addition to the CUG clade yeasts, oleaginous yeasts, such as Yarrowia lipolytica and Psedozyma hubeiensis, also exhibit native mixed-sugar consumption capabilities. Employing comprehensive metabolic and transcriptomic analysis strategies, a recent study unraveled the genes responsible for xylose and cellobiose uptake and metabolism in Y. lipolytica (Ryu et al., 2016). Simultaneous mixed-sugar utilization was achieved, which was attributed to mild glucose repression in conjunction with strong carbon catabolite activation for growth on xylose and cellobiose. In another exhaustive screening of 1,189 yeast isolates, Tanimura et al. demonstrated P. hubeiensis IPM1-10 was able to co-utilize glucose, xylose, and arabinose in the artificial hydrolysate and accumulated high amounts of lipid (Tanimura et al., 2016). These findings suggest the importance of broadening the current collection of microbial species for efficient fermentation of multiple sugars in biomass hydrolysates.
Researchers have also attempted to develop an alternative conversion scheme, which is to co-ferment oligosaccharides and pentoses (Yang et al., 2014). In contrast to S. cerevisiae, which cannot metabolize cellodextrins natively, the cellulolytic fungus N. crassa grows on cellodextrins because of its high-affinity cellodextrin transport system (Galazka et al., 2010). Thus, to bypass extracellular glucose suppression on hexose/pentose co-utilization, the integration of the newly discovered cellodextrin transport system (Tian et al., 2009; Galazka et al., 2010) in conjunction with a xylose-utilizing pathway facilitated simultaneous fermentation of cellobiose and xylose in engineered S. cerevisiae (Li et al., 2010; Ha et al., 2011). Specifically, genes encoding the cellodextrin transporter and an intracellular β-glucosidase were co-expressed in S. cerevisiae containing a functional xylose pathway. The resulting strain produced 48 g L−1 ethanol from a mixture of 10 g L−1 glucose, 80 g L−1 cellobiose, and 40 g L−1 xylose in 60 h fermentation, more productive than S. stipitis cultured under the same conditions. The major advantage of the integrated system resides in the fact that the intracellular glucose hydrolyzed from cellobiose did not repress extracellular xylose transportation, therefore enabling co-sugar uptake. Moreover, the synergistic effects of cellobiose and xylose co-utilization greatly improved ethanol yield and productivity (Ha et al., 2011). Similarly, co-fermentation of cellodextrin and galactose was successfully accomplished in S. cerevisiae, and an appreciable increase in ethanol productivity was observed during mixed-sugar fermentation (Ha et al., 2011). Taken together, this unique strategy advances the economic production of fuels and chemicals from plant biomass.
In addition to its viability on cellobiose, N. crassa also grows well on xylodextrin (e.g., xylobiose, xylotriose, and xylotetraose), owing to its innate xylodextrin transport and utilization pathway (Li et al., 2015). Reconstituting this pathway in S. cerevisiae enabled slow cell growth on xylodextrin. An ortholog of N. crassa xylodextrin transporter, ST16 from Trichoderma virens, enabled S. cerevisiae to grow more rapidly on xylodextrins as sole carbon source (Zhang et al., 2017). Moreover, ST16 was not subjected to cellobiose inhibition, making it a promising candidate for the co-fermentation of cellobiose and xylodextrins.
Similar to xylose, arabinose is exogenous to S. cerevisiae, although it can be assimilated by wild-type S. cerevisiae, facilitated mainly by the hexose transporter GAL2 (Becker and Boles, 2003; Subtil and Boles, 2011). In contrast to xylose transporters, heterologous arabinose transporters have not been extensively studied and previous studies mainly focused on introducing a functional arabinose utilization pathway into S. cerevisiae. Several studies showed that GAL2 overexpression in S. cerevisiae strains enhanced arabinose fermentation under various culture conditions (Becker and Boles, 2003; Wang et al., 2013). However, GAL2 is a non-specific, low-affinity arabinose transporter subjected to glucose repression (Kou et al., 1970; Subtil and Boles, 2011). Therefore, to improve arabinose uptake in S. cerevisiae, putative arabinose transporters from multiple species were characterized over the past several years. Verho et al. identified and characterized the first eukaryotic genes encoding arabinose-exclusive transporters, LAT1 and LAT2 from A. monospora, a native arabinose-assimilating yeast (Verho et al., 2011). Expression of LAT1 and LAT2 in the S. cerevisiae hexose transporter-null strain containing the fungal arabinose-utilizing pathway conferred cell growth on arabinose. Shortly thereafter, additional arabinose transporters were isolated, including AraT from S. stipitis, S. passalidarum, and Penicillium chrysogenum, Stp2 from Arabidopsis thaliana, LAT-1 from N. crassa and Myceliophthora thermophila, and AXT1 from Kluyveromyces marxianus and Pichia guilliermondii (Subtil and Boles, 2011; Benz et al., 2014; Knoshaug et al., 2015; Li et al., 2015; Caballero and Ramos, 2017; Bracher et al., 2018). Although these reported arabinose transporters improved arabinose uptake in S. cerevisiae, their activities in glucose/arabinose-containing cultures were either not tested or were found to be inhibited by glucose. Therefore, future targets regarding mining arabinose transporters should focus on relieving hexose repression while improving intracellular metabolic flux.
One major challenge in the efficient conversion of plant biomass to fuels/chemicals involves deconstructing complex components to release the entrained fermentable sugars. This is inherently difficult because plants have evolved recalcitrance to withstand numerous forces attempting to jeopardize their structure (Olson et al., 2012). To overcome this recalcitrance and ferment plant biomass efficiently, dedicated biomass pretreatment processes have been developed, including chemical hydrolysis, pyrolysis, and enzymatic catalysis. These additional processing steps are partially responsible for the high production cost of second-generation fuels/chemicals (Olson et al., 2012). Consolidated bioprocessing (CBP) aims to simplify and reduce the cost of biorefining by combining cellulase production, lignocellulose degradation, and sugar fermentation to desired products in one process without pretreatment or enzyme supplementation. CBP requires a microbial host that can both break down and utilize lignocellulose, as well as produce desired products at high yields/titers. Unfortunately, no natural hosts can efficiently perform both tasks. Therefore, to engineer a host for CBP, two major strategies have been developed, including: (i) engineering a fuel/chemical-producing workhorse such as E. coli or S. cerevisiae to utilize plant feedstocks and (ii) engineering a plant biomass-consuming organism (e.g., Aspergillus, Clostridium, Cellulomonas) to produce desired fuels/chemicals. This section mainly describes recent advances in engineering S. cerevisiae to utilize minimally processed plant biomass as feedstocks.
Engineering S. cerevisiae to simultaneously break down plant biomass and ferment the released carbohydrates to desired products requires the integration of lignocellulolytic capabilities. Numerous organisms exhibit lignocellulolytic activity, including various bacteria, fungi, crustaceans, and insects (Gírio et al., 2010). Various cellulases/hemicellulases enable these organisms to depolymerize plant biomass into fermentable sugars. Numerous engineering efforts to integrate lignocellulolytic activity in S. cerevisiae and other hosts have focused on heterologous expression of such enzymes either on the cell surface or through secretion to the extracellular media (Figure 3). Cellobiose can be hydrolyzed to glucose by β-glycosidase (Figure 1). Rooyen et al. expressed two β-glucosidases, BGL1 from Saccharomycopsis fibuligera and BglA from Aspergillus kawachii, on the cell surface of S. cerevisiase, enabling it to grow anaerobically on cellobiose as the sole carbon source (van Rooyen et al., 2005). Unlike glucose, cellobiose does not repress the uptake of other sugars such as xylose and galactose (Li et al., 2010). Thus, S. cerevisiae has been engineered to co-utilize cellobiose and xylose or cellobiose and galactose in the past decade. Multiple studies employed a ubiquitous strategy: expression of a surface-displayed β-glycosidase to enable cleavage of cellobiose into glucose with the resulting glucose being rapidly ingested by yeast cells, functioning to maintain a low extracellular glucose concentration to ameliorate glucose repression on ingestion of other sugars (Katahira et al., 2006; Nakamura et al., 2008; Saitoh et al., 2010). As a proof of concept, the first xylose/cellobiose co-fermenting yeast strain was built via integrating an intracellular xylose pathway and a β-glycosidase tethered to the cell surface in a single strain (Katahira et al., 2006). The resulting strain assimilated xylose in the presence of a high cellobiose concentration as rapidly as that in pure xylose media (Nakamura et al., 2008). The simultaneous utilization of xylose and cellobiose resulted in 35.9 g L−1 of ethanol from 50 g L−1 cellobiose and 50 g L−1 xylose with a yield of 0.36 g g−1 and a productivity of 0.50 g L−1 h−1, which was 34% higher than that of glucose/xylose co-fermentation. Subsequently, an improved strategy was employed to construct a more efficient cellobiose/xylose-utilizing strain starting from an industrial yeast (OC2-HUT) (Saitoh et al., 2010). Specifically, two copies of the xylose pathway and four copies of genes encoding cell-surface-displayed β-glycosidase were integrated into the genome. The resulting strain consumed 90 g L−1 cellobiose and 60 g L−1 xylose in 48 h, increasing ethanol titer, yield, and productivity to 57.4 g L−1, 0.38 g g−1, 1.2 g L−1 h−1, respectively. Because excessive extracellular cellobiose hydrolysis and non-efficient glucose transportation into the cell would result in glucose accumulation in the medium and further repress xylose uptake (Kim et al., 2012), a balanced glucose uptake rate and surface displayed β-glycosidase activity must be carefully maintained.
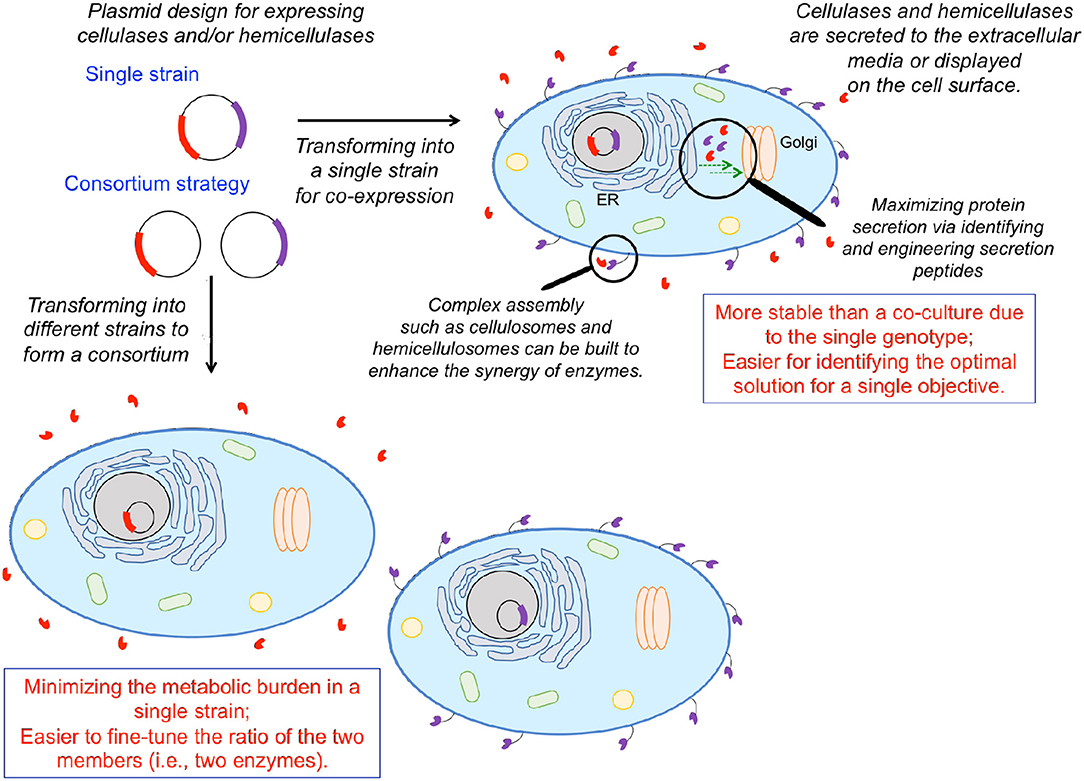
Figure 3. Solubilizing biomass to fermentable sugar substrates for biochemical production via heterologous expression of cellulases and hemicellulases. One can either engineer a single organism to express multiple cellulolytic/hemicellulolytic enzymes or employ a consortium strategy, engineering multiple organisms to each express a single enzyme. Enzymes can either be expressed on the cell surface, or secreted to the extracellular medium, depending on the application. Heterologous protein expression can be enhanced via identifying/engineering secretion tags and facilitating transport either from the endoplasmic reticulum to the Golgi, or the Golgi to the cell surface. Individual enzymes can be brought into spatial proximity via assembly of complex structures mimicking cellulosomes and hemicellulosomes to coordinate enzymatic activities.
Similarly, Jeon et al. expressed BGL1 along with EngD, an endoglucanase from Clostridium cellulovorans in S. cerevisiae, and promoted secretion of these enzymes into the extracellular media by fusing them to the α-mating factor secretion signal peptide. The resulting strain produced ethanol from β-glucan, achieving 80.3% of the theoretical yield (Jeon et al., 2009). An engineered xylose-utilizing S. cerevisiae strain co-displaying xylanase II from Trichoderma reesei and β-xylosidase from Aspergillus oryzae produced 7.1 g L−1 ethanol directly from birchwood xylan after a 62-h fermentation (Katahira et al., 2004). Sakamoto et al. co-expressed a cellulase (β-glucosidase from Aspergillus aculeatus) and hemicellulases (the xylanase II and β-xylosidase enzymes previously mentioned Katahira et al., 2004) in a xylose-utilizing S. cerevisiae strain. The resulting strain produced 8.2 g L−1 ethanol from rice straw hydrolysates after a 72-h fermentation (Sakamoto et al., 2012). Although these studies confirm that S. cerevisiae can be engineered to utilize cellulose/hemicellulose, the activities of the expressed enzymes must be coordinated to realize the full potential of CBP.
In nature, some lignocellulolytic organisms organize their cellulases into complexes known as cellulosomes (Bayer et al., 2008). The function of the cellulosome is to mediate binding to lignocellulosic biomass and coordinate enzymatic activities for efficient biomass solubilization. The discovery and characterization of cellulosomes has inspired engineering efforts to mimic these structures with the aim of synergistically increasing heterologous enzyme activities. Fan et al. engineered a S. cerevisiae strain that displayed a mini-cellulosome on its cell surface (Fan et al., 2012) (Figure 4A). This was accomplished by first secreting three different cellulases from mesophilic Clostridia along with scaffoldin I, which contains the necessary type I cohesins for cellulase anchoring using α-mating factor. Scaffoldin II was then cell surface-displayed via fusion expression with the smaller subunit (69 amino acids) of the a-agglutinin receptor, which forms two disulfide bonds with the larger subunit (725 amino acids) that is endogenously produced and then covalently attached to the yeast cell wall matrix. Scaffoldin I was also fused with a type II dockerin at the C-terminus, which subsequently bound to one of the four repeating type II cohesions carried by scaffoldin II. This enabled tethering of the engineered mini-cellulosome to the cell surface. The resulting strain directly fermented Avicel and phosphoric acid swollen cellulose (PASC) to ethanol, achieving titers of 1.41 and 1.09 g L−1 after 4-day fermentation, respectively.
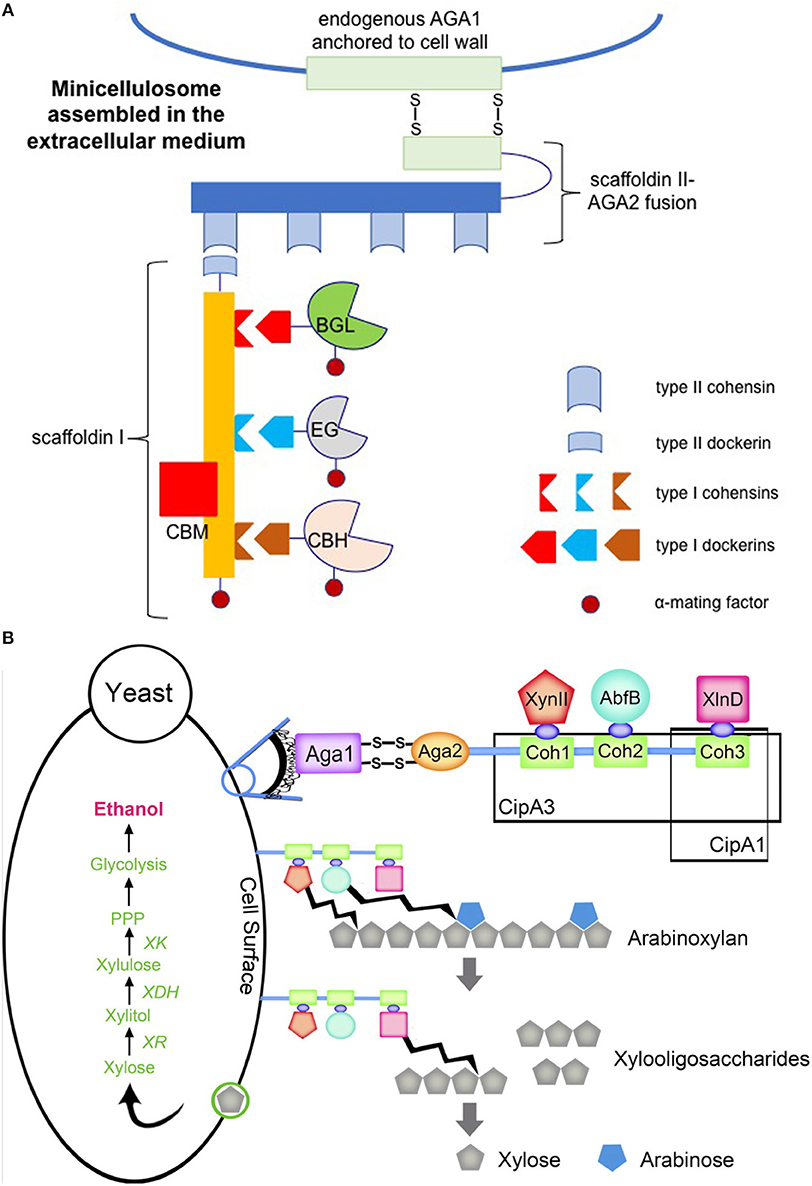
Figure 4. Engineered cellulosome (A) and hemicellulosome (B) in S. cerevisiae. (A) Three cellulases from Clostridia were heterologously expressed in S. cerevisiae, including endoglucanase (EG), cellobiohydrolase (CBH), and β-glucosidase (BGL). Each cellulase contained a type I dockerin specific to one of three type I cohesins present on scaffoldin I which was expressed by the recombinant strain. Fusion of these four components to α-mating factor enabled secretion to the extracellular medium to prevent intracellular assembly. Expression of a scaffoldin II-a-agglutinin adhesion subunit (AGA2) fusion tethered scaffoldin I-cellulase complexes to the cell wall via type II dockerin-cohesin interactions. Scaffoldin II-AGA2 anchored to the cell wall via formation of two disulfide bridges between AGA2 and endogenous a-agglutinin anchorage subunit (AGA1). The image was prepared according to the previous study carried out by Fan et al. (2012) (B) Minihemicellulosomes were expressed in S. cerevisiae carrying the xylose utilizing pathway. The miniscaffoldin CipA1 contained one type I cohesion (Coh3), whereas the miniscaffoldin CipA3 contained three type I cohesions (Coh1, Coh2, and Coh3). Each miniscaffoldin was tethered to the cell wall via AGA1 and AGA2 interaction, analogous to the cellulosome structure in (A). The endoxylanase (XynII), arabinofuranosidase (AbfB), and β-xylosidase (XlnD) were tethered to the miniscaffolidns via type 1 dockerin-cohesin interactions. The original figure published in Sun et al. (2012) was shown here with a copyright permission.
Similarly, Sun et al. constructed two separate mini-hemicellulosomes by expressing two different miniscaffoldins, one with a single cohesion module and the other with three cohesion modules, from Clostridium thermocellum in a xylose-utilizing S. cerevisiae strain (Sun et al., 2012) (Figure 4B). These miniscaffoldins were then attached to the cell surface via the a-agglutinin adhesion receptor. Heterologous expression of three hemicellulases with C-terminal dockerins mediated non-specific anchoring of the enzymes on the miniscaffoldins. With three possibilities at each cohesion, 27 different trifunctional minihemicellulosomes were engineered. In contrast, the miniscaffoldin equipped with only one cohesion module randomly adhered to one of the three hemicellulases with C-terminal dockerins. The trifunctional mini-hemicellulosome design exhibited 1.39-fold higher arabinoxylan conversion activity than the mini-hemicellulosomes bound to only one of the hemicellulases, demonstrating the synergistic benefit of placing the three hemicellulases in proximity. Fan et al. observed analogous results in their cellulosome system, noting that the trifunctional system exhibited higher activity than the sum of three unifunctional systems, resulting in a 3.6-fold increase in the ethanol titer (Fan et al., 2012).
One major culprit inhibiting the secretion of heterologous enzymes and efficient surface display is the yeast secretory pathway. Misfolding and post-translational modifications differing from the native host can jeopardize catalytic activity (Boer et al., 2000), prevent transport out of the Golgi, and/or result in degradation in the endoplasmic reticulum (ER) (Lambertz et al., 2014). Overexpression of a heterologous protein can also pose an excessive burden to a cell's protein folding and secretion machinery. Recently, Tang et al. overexpressed genes involved in vesicle trafficking from the ER to the Golgi and from the Golgi to the plasma membrane, enhancing the secretion and surface display of BGL1 and endoglucanase from C. thermocellum in S. cerevisiae (Tang et al., 2017) (Figure 3). This work also suggested that the rate-limiting step in the yeast secretory pathway is protein-specific. It was found that overexpression of genes associated with transport from the ER to the Golgi had a more pronounced effect on endoglucanase secretion, whereas overexpression of genes associated with transport from the Golgi to the cell membrane had a more pronounced effect on BGL1 secretion. This work demonstrated that modifying the processes of vesicle budding, tethering, and fusion is a useful strategy for increasing enzyme secretion and surface display. Another strategy for increasing the secretion of heterologous protein in S. cerevisiae is to fuse optimal secretion signals to the desired protein. Bae et al. developed a high-throughput screening method for selecting a translational fusion partner (TFP) for optimal protein secretion (Bae et al., 2015). By manipulating the invertase (catalyzing the hydrolysis of sucrose into fructose and glucose) fusion partner trap system, secretion of a desired protein was coupled with the growth of SUC2-disrupted S. cerevisiae on sucrose-only media. Lee et al. implemented this screening method to select TFPs for four different cellulases and then expressed each of the four cellulases with the accompanying TFP in a S. cerevisiae strain (Lee et al., 2017). These four strains were then co-fermented, producing ethanol from pre-treated rice straw at approximately 3-fold higher productivity than the control wild-type strain in the culture supplemented with 10 filter paper units commercial cellulases per gram of glucan. To implement such consortia in practical industrial applications, determining co-fermentation seed ratios (which dictates enzyme ratios) is necessary for optimizing synergistic enzymatic activity. Variance in growth rates amongst engineered strains can make this task cumbersome, presenting a technical burden to practical applications (Lee et al., 2017). Further characterization of the optimal enzyme ratios and enzyme-secreting strains are required for efficient lignocellulose solubilization.
Recently, Liu et al. proposed a novel strategy for lignocellulose saccharification. Scanning electron microscopy imaging revealed that cellulase-displaying S. cerevisiae cells adhered to PASC, a phenomenon not observed in cellulase-secreting S. cerevisiae. Additionally, the cellulase-displaying strain achieved a 9% higher ethanol yield than the cellulase-secreting strain. Adhesion of the cellulase-displaying strain mediated a “tearing” effect on PASC, increasing the rate of breakdown compared to free enzyme hydrolysis obtained from the cellulase-secreting strain (Liu et al., 2016). Interestingly, adhesion was not observed and the ethanol yields were about the same for both strains when Avicel was used as the substrate. These results indicate that the rough, high surface area properties of the amorphous PASC enables adhesion, whereas the crystalline Avicel cannot promote such a mechanism. Promoting cell adhesion to biomass is a new engineering approach to enhance saccharification rates.
Engineering microbial communities represents a new frontier of synthetic biology and has garnered increasing attention in recent years (Brenner et al., 2008). Microbial co-culture systems can perform complex tasks via labor division. Unlike monocultures, consortium enables the partitioning of multi-step metabolic pathways to each member of the consortium. Not only does this compartmentalization alleviate excessive metabolic burden on a single member, but it also provides the crucial ability to optimize segmented pathways to avoid compromise, thus minimizing byproduct accumulation (Zhang et al., 2015). In nature, communities respond more robustly to environmental fluctuations such as nutrient limitation, and this feature may be particularly useful for the fermentation of biomass hydrolysates when sugar choices and concentrations are shifting (Brenner et al., 2008). In this section, the applications of both natural and synthetic consortia for enhancing mixed-sugar consumption and improving biochemical production are discussed. Although this review mainly focuses on yeast engineering, a few interesting bacterium consortium examples are included below.
Microbial consortia are ubiquitous in nature, and synergistic interactions among community species are established after long-term evolution in specific environments (Brenner et al., 2008). Applying natural consortia in mixed-sugar utilization is beneficially associated with non-sterile fermentation at reduced operation costs (Zuroff and Curtis, 2012). For example, thermophilic bacterium consortia enriched from geothermal spring were evolved to produce hydrogen from glucose/xylose mixtures and hydrolysates of oil palm trunk with yields of 375 and 301 mL H2 g−1 sugar consumed, respectively (Hniman et al., 2011). The dominant species in the consortia were Thermoanaerobacterium sp., Thermoanaerobacter sp., Caloramater sp., and Anoxybacillus sp. In addition to the major product hydrogen, multiple byproducts were detected, including butyric acid, acetic acid, lactic acid, and butanol. Although natural consortia are durable under dynamic culture conditions because of their high self-tunability and have been applied to complex tasks such as wastewater treatment and lignocellulosic biomass deconstruction, their uncontrollability limits their wide application in producing specific biochemicals at high titers (Zuroff and Curtis, 2012). The physiology and genetic background of members of natural consortia are not well-characterized, further complicating determination of the interaction mechanisms in the microbial community.
To address this issue, significant progress has been made in the design and characterization of synthetic microbial consortia because of their relatively simple interplays (Zuroff and Curtis, 2012). The simplest design is to build a synthetic community with strains from the same species, which avoids growth incompatibility issues, including temperature, pH, nutrition requirement, and growth rate. Eiteman et al. developed an E. coli/E. coli community for simultaneous conversion of glucose and xylose to a mixture of valuable products, including formate, lactate, ethanol, and succinate (Eiteman et al., 2008). One of the two community members was engineered to be glucose-exclusive with its native XI deleted, while the other was xylose-selective with three genes involved in glucose uptake and phosphorylation disrupted. Rather than adversely competing with one another, the two E. coli strains acted in concert to co-utilize sugar mixtures, leading to a higher sugar consumption rate than the monocultures. The distinguishing hallmark of the consortium strategy is the robustness and flexibility provided by the system. Fed-batch fermentation indicated that this consortium adjusted the ratio of each strain based on fluctuations in the feed sugar concentration. For example, when xylose concentration is higher than glucose in the feed stream, the xylose-consuming strain would be dominant, and vice versa. Subsequently, the same group further extended the consortium approach to the simultaneous consumption of glucose, xylose, and arabinose along with acetate, which is a growth inhibitor typically present in lignocellulosic hydrolysates (Xia et al., 2012). Four E. coli strains engineered to be single substrate-selective were co-inoculated, and synchronous utilization of four substrates was achieved, albeit the xylose and arabinose consumption rates were decreased by 14 and 11%, respectively, compared to those in the wild-type strain.
Employing the analogous rationale to explore the synchronous conversion of glucose/xylose/arabinose mixtures, a consortium system consisting of three S. cerevisiae specialists was established recently (Verhoeven et al., 2018). Both xylose- and arabinose-exclusive specialists were constructed via disrupting hexose phosphorylation followed with adaptive evolution. Despite the rapid consumption of glucose and arabinose by the three-strain co-culture, xylose utilization was severely impaired, possibly due to (by-)product inhibition. Only after substantial anaerobic laboratory evolution was xylose utilization significantly improved, eventually leading to the simultaneous depletion of all three sugars. Moreover, the evolved three-strain consortium was more stable and robust than a pentose-fermenting yeast monoculture during long-term cultivation. Very recently, a kinetic model was constructed to provide guidelines for mixed-sugar fermentation via S. cervisiae co-cultures. This model, which incorporated pure culture models, product inhibition effects, initial substrate concentrations, and inoculum sizes, accurately predicted independent experimental results (Chen et al., 2018). Implementation of the model prediction enabled co-fermentation of 60 g L−1 cellobiose and 20 g L−1 xylose in a S. cerevisiae/S. cerevisiae co-culture.
A stable E. coli/E. coli integrated system that produced high-value compounds from glucose/xylose mixtures was constructed and optimized (Zhang et al., 2015) (Figure 5A). To overcome the lingering high leakage of pathway intermediates and non-efficient sugar utilization, the multi-step cis, cis-muconic acid (MA) synthesis pathway was split into two independent E. coli strains. One strain preferred xylose and contained the shikimate pathway, which catalyzed the conversion of xylose to the intermediate dehydroshikimate (DHS). The other strain preferred glucose and harbored the exogenous DHS-to-MA synthesis pathway together with a DHS transporter. As a result, glucose/xylose co-consumption was achieved, eliminating carbon source competition in the community. Additionally, the microbial community improved the MA titer to 4.7 g L−1 with a yield as high as 0.35 g g−1 sugar mixture. Because of the tunability and dynamic growth balance exhibited by the two constituent members, the community was more robust for handling varying mixed-sugar concentrations. Finally, the co-culture concept was extended to produce 4-hydroxybenzoic acid (4HB) using the DHS-producing strain and a new strain that converted DHS to 4HB. Through simple independent engineering of the new strain, a final titer of 2.3 g L−1 4HB was obtained, and the yield of 0.11 g g−1 was equivalent to the highest reported yield achieved in previously reported glucose fermentations (Barker and Frost, 2001).
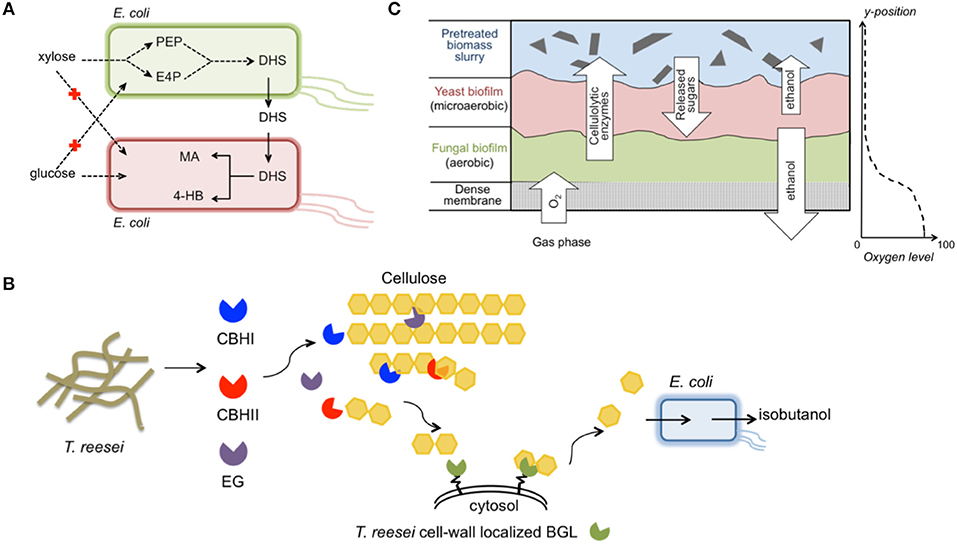
Figure 5. Exemplary engineered microbial consortia. (A) An E. coli/E. coli consortium for improved production of MA and 4HB from a glucose/xylose mixture (Zhang et al., 2015); (B) A T. reesei/E. coli consortium for direct conversion of biomass hydrolysates to isobutanol (Minty et al., 2013); (C) A T. reesei/S. cerevisiae/S. stipitis biofilm membrane reactor integrating aerobic cellulolytic enzyme production and anaerobic ethanol accumulation (Brethauer and Studer, 2014). The images were prepared according to the previous individual studies. Abbreviation of metabolites— PEP, phosphoenolpyruvate; E4P, erythrose-4-phosphate; DHS, dehydroshikimate; MA, cis, cis-muconic acid; 4-HB, 4-hydroxybenzoic acid; CBHI, cellobiohydrolase I; CBHII, cellobiohydrolase II; EG, endoglucanase; BGL, β-glucosidase.
Compared to single-species consortia, manipulating communities comprising multi-species is more complicated. This is attributed to the community instability caused by different growth rates and difficulty in building cell-cell communication across species. Solving these instability issues would be advantageous, since multi-species consortia possess unique advantages. A wider array of mixed species offer higher flexibility and capability to achieve more complex tasks. Additionally, a better understanding of multi-species consortia is beneficial for future studies to break the confines of model microorganisms and exploit non-model hosts, particularly those with desirable phenotypes but are recalcitrant to genetic modifications. Numerous studies have focused on developing microbial consortia of S. cerevisiae and a native pentose-fermenting yeast for efficient hexose/pentose co-utilization. For example, under oxygen-limited and continuous culture conditions, the co-cultured S. stipitis and respiratory-deficient S. cerevisiae system converted 35.0 g L−1 glucose and 10.4 g L−1 xylose and displayed enhanced ethanol yield at 0.42 g g−1 sugars (Laplace et al., 1993). Such a co-cultured yeast system was further optimized via adjusting the agitation rate, temperature, and continuous dilution rate (Taniguchi et al., 1997; Delgenes et al., 1998; Suriyachai et al., 2013). To identify the key determinants of the performance of this S. cerevisiae/S. stipitis co-culture system, a dynamic metabolic flux balance model was constructed (Hanly and Henson, 2013) and an 11% improvement in ethanol productivity was observed by adjusting the aeration level and inoculation concentrations predicted by the model. Slow xylose uptake and conversion were determined as the key factors restraining ethanol production by this S. cerevisiae/S. stipitis co-culture.
A unique synthetic fungal-bacterial mixed population was developed for isobutanol production directly from plant biomass (Minty et al., 2013) (Figure 5B). Isobutanol is a highly sought-after next-generation biofuel with advantageous properties as a gasoline substitute, including higher energy density, lower hygroscopicity, and lower volatility compared to ethanol (Atsumi et al., 2008). This consortium system integrated cellulase secretion, sugar saccharification, and fermentation via combining T. reesei, an active cellulase-producing host, and E. coli that was engineered to convert sugars to isobutanol. The dedicated design maintained a very low glucose concentration in the medium, analogous to ideal CBP conditions. A comprehensive ordinary differential equation model established, which included 50 parameters, revealed that T. reesei and E. coli coexisted stably despite the isobutanol toxicity effect. When using pretreated corn stover as substrate, various sugars in the hydrolysates were co-consumed and isobutanol production titer and yield reached 1.88 g L−1 and 62%, respectively. Compared to the traditional “superbug” CBP, dividing the entire isobutanol process into T. reesei and E. coli provided an optimal environment for functional expression of both the cellulase-producing pathway and sugar-conversion pathway. Moreover, the modular design could facilitate rapid adaption of the consortium system to produce a portfolio of advanced chemicals from plant biomass.
A more complicated, multi-species biofilm membrane (MBM) reactor was used to produce ethanol from undetoxified pretreated wheat straw (Brethauer and Studer, 2014) (Figure 5C). The meticulously designed MBM system maintained an optimal environment for each of the three specialists, namely T. reesei, S. cerevisiae, and S. stipitis, for cellulase production, glucose-to-ethanol conversion, and xylose-to-ethanol conversion, respectively. In MBM fermentation, a maximum of 9.8 g L−1 ethanol with a yield of 69% was obtained, which was substantially higher than the ethanol titer (4.6 g L−1) and yield (41%) obtained by co-culture of the engineered S. cerevisiae and S. stipitis using the same substrate. Despite the advantageous properties of the MBM system, limitations such as the large portion of residual cellobiose in plant biomass hydrolysates must be resolved before this system can serve as an efficient platform for economic biochemical production.
Grave concerns regarding the future availability of fossil fuels, coupled with the negative environmental ramifications of the globe's current carbon footprint, is driving the initiative toward “green” fuels and chemicals. Although a step in the right direction, first-generation biorenewable products made from food crops led to concerns regarding food security. Instead, utilization of lignocellulosic biomass as the second-generation feedstock for fuel and chemical production has numerous potential environmental and societal advantages compared to its petroleum and sugar feedstock counterparts. Since 2008, global efforts from major chemical/fuel companies (e.g., POET, Raizen, etc.) have turned dozens of commercial or semi-commercial scale cellulosic ethanol plants into operation. A typical cellulosic ethanol plant has a production capacity on the order of tens of million gallons per year, which is nowhere near the total consumption rate of petroleum. As a reference, U.S. petroleum consumption was estimated to be on average, 840 million gallons per day in 2017. Despite the price fluctuation of crude oil in the past decade (in a range of $30 to $110 per barrel), bioethanol production needs to be near $70 per barrel in order to be economically competitive, which makes the co-utilization of all the sugars present in biomass hydrolysates essential. Therefore, future efforts should focus on integrating the capacities of transporting and converting all major sugars in a CCR-free manner.
Previous studies of mixed-sugar fermentation in S. cerevisiae were mostly isolated and had limited crossovers, which have proven problematic since intracellular metabolic flux and sugar uptake can both restrict maximal production rates. Thus, future efforts should shift toward developing robust strains for industrial biomass hydrolysate conversion via applying global cellular engineering strategies. Instead of focusing solely on increasing pathway flux and sugar transportation, engineering strategies should go beyond this superficial level, aiming to expand our current understanding to unveil the native regulatory mechanisms underlying non-glucose utilization in S. cerevisiae. For example, evolutionary engineering followed by genome re-sequencing of evolved S. cerevisiae strains with beneficial phenotypes elucidated that reduced glucose phosphorylation rates enabled simultaneous glucose/xylose utilization (Lane et al., 2018; Papapetridis et al., 2018). Comparative transcriptomic analysis on the natural S. cerevisiae YB-2625 strain isolated from bagasse and the model yeast strain S288C also revealed that multiple factors contributed to the exceptional xylose-consuming capability (Cheng et al., 2018). Some factors were rather obvious, such as down-regulation of hexokinase and genes involved in glucose-repression related transcription factors. However, other factors were quite intriguing. These included up-regulation of genes encoding antioxidant enzymes, a less than obvious link to xylose uptake and utilization. Such interventions would not have been easily discovered without a global analysis approach.
Expanding the current collection of microbial factories for the highly sought-after cosugar-utilizing phenotype presents a new frontier in the biochemical industry. A high-throughput screening method of desired characteristics must first be developed to expedite the discovery of strains possessing superior phenotypes. The increasingly prevalent microbiome and metagenomic analyses will revolutionize our ability to identify new microbial species at an unprecedented rate (Shapiro et al., 2018). Newly identified nonconventional microbes will create an urgent demand for corresponding genetic manipulation tools, specifically platform technologies that can be easily transferred from one species to another. Extension of current CRISPR-Cas technology to non-conventional yeasts will enable precise genome modification, convenient pathway engineering, and wide genetic interaction analysis (Shapiro et al., 2018). Other platform technologies, such as CRISPR-mediated base-editing platforms and RNA-seq, are beneficial for expeditious mapping of superior phenotypes to genotypes.
After extensive review of the current literatures, efficient engineering of heterogeneous consortia arises as a highly promising strategy in achieving the ultimate goal of converting lignocellulosic biomass to biofuels/biochemicals in an economically competitive manner. However, several challenges must be overcome before such systems can be adapted to commercial production. Unlike their natural counterparts, synthetic communities cannot retain long-term homeostasis, particularly under industrial harsh conditions containing inhibitors and toxins (Brenner et al., 2008). Thus, combinatorial approaches, such as evolutionary engineering combined with further development of bioreactors for long-term culturing and monitoring, should be implemented to increase robustness and fine tune the efficiency of consortia (Jia et al., 2016). Additionally, computational tools should be harnessed to develop dynamic models to facilitate discovery and understanding of variables most crucial to system performance (Purnick and Weiss, 2009; Chen et al., 2018). These models can also provide rational guidelines when designing new consortia.
In addition to efficient utilization of all sugars in biomass hydrolysates, engineering efforts should also be directed toward in situ fixation of the CO2 produced through numerous decarboxylation reactions (Cheng et al., 2018). For example, various CO2 conservative pathways, such as non-oxidative glycolysis (Bogorad et al., 2013), the methanol condensation pathway (Bogorad et al., 2014), reductive PPPs (rPPPs) (Guadalupe-Medina et al., 2013; Xia et al., 2017), and the 3-hydroxypropionate pathway (Mattozzi et al., 2013), have been successfully introduced into either E. coli or S. cerevisiae. To alleviate CO2 emissions and remove the surplus NADH generated via the XR/XDH pathway, synthetic rPPPs were incorporated into xylose-utilizing S. cerevisiae strains, resulting in higher ethanol yields and lower byproduct accumulation (e.g., glycereol and xylitol) during xylose-fermentation (Li et al., 2017; Xia et al., 2017). Implementing CO2 conservative pathways, in conjunction with optimization of microbial consortia in CBP via global engineering strategies, has the potential to expedite the development of an economic, sustainable, and environmentally-friendly lignocellulosic-based biochemical industry.
MG, DP, and ZS wrote this review manuscript together. DP wrote the CBP, and cellulase secretion sections. DP and MG wrote the introduction and conclusion sections. MG wrote the remaining parts; ZS revised and polished the entire review.
ZS declares a financial relationship with Estose Biorenewables, LLC, a startup company that aims to provide technologies to build yeast platforms to produce nutraceuticals and pharmaceuticals from biorenewables.
The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
This work was partially supported by the National Science Foundation Grants (MCB 1716837, and the Graduate Research Fellowships Program GRFP). This research was also supported in part by the U.S. Department of Energy, Office of Science, Biological and Environmental Research, Biological Systems Science Division through the Ames Laboratory. The Ames Laboratory is operated for the U.S. Department of Energy by Iowa State University under Contract No. DE-AC02-07CH11358.
Aden, A., Bozell, J., Holladay, J., White, J., and Manheim, A. (2004). Top Value Added Chemicals From Biomass, eds, T. Werpy, and G. Petersen. U.S. Department of Energy, National Renewbale Energy Laboratory.
Aeling, K. A., Salmon, K. A., Laplaza, J. M., Li, L., Headman, J. R., Hutagalung, A. H., et al. (2012). Co-fermentation of xylose and cellobiose by an engineered Saccharomyces cerevisiae. J. Ind. Microbiol. Biot. 39, 1597–1604. doi: 10.1007/s10295-012-1169-y
Alper, H., and Stephanopoulos, G. (2009). Engineering for biofuels: exploiting innate microbial capacity or importing biosynthetic potential? Nat. Rev. Microbiol. 7, 715–723. doi: 10.1038/nrmicro2186
Atsumi, S., Hanai, T., and Liao, J. C. (2008). Non-fermentative pathways for synthesis of branched-chain higher alcohols as biofuels. Nature 451, 86–89. doi: 10.1038/nature06450
Attfield, P. V., and Bell, P. J. (2006). Use of population genetics to derive nonrecombinant Saccharomyces cerevisiae strains that grow using xylose as a sole carbon source. FEMS Yeast Res. 6, 862–868. doi: 10.1111/j.1567-1364.2006.00098.x
Bae, J. H., Sung, B. H., Kim, H. J., Park, S. H., Lim, K. M., Kim, M. J., et al. (2015). An efficient genome-wide fusion partner screening system for secretion of recombinant proteins in yeast. Sci. Rep. 5:12229. doi: 10.1038/srep12229
Banerjee, S., Archana, A., and Satyanarayana, T. (1994). Xylose metabolism in a thermophilic mold malbranchea-pulchella var sulfurea tmd-8. Curr. Microbiol. 29, 349–352. doi: 10.1007/BF01570228
Barker, J. L., and Frost, J. W. (2001). Microbial synthesis of p-hydroxybenzoic acid from glucose. Biotechnol. Bioeng. 76, 376–390. doi: 10.1002/bit.10160
Bayer, E. A., Lamed, R., White, B. A., and Flint, H. J. (2008). From cellulosomes to cellulosomics. Chem. Rec. 8, 364–377. doi: 10.1002/tcr.20160
Becker, J., and Boles, E. (2003). A modified Saccharomyces cerevisiae strain that consumes L-arabinose and produces ethanol. Appl. Environ. Microbiol. 69, 4144–4150. doi: 10.1128/AEM.69.7.4144-4150.2003
Bengtsson, O., Hahn-Hägerdal, B., and Gorwa-Grauslund, M. F. (2009). Xylose reductase from Pichia stipitis with altered coenzyme preference improves ethanolic xylose fermentation by recombinant Saccharomyces cerevisiae. Biotechnol. Biofuels 2:9. doi: 10.1186/1754-6834-2-9
Benz, J. P., Chau, B. H., Zheng, D., Bauer, S., Glass, N. L., and Somerville, C. R. (2014). A comparative systems analysis of polysaccharide-elicited responses in Neurospora crassa reveals carbon source-specific cellular adaptations. Mol. Microbiol. 91, 275–299. doi: 10.1111/mmi.12459
Bera, A. K., Sedlak, M., Khan, A., and Ho, N. W. (2010). Establishment of L-arabinose fermentation in glucose/xylose co-fermenting recombinant Saccharomyces cerevisiae 424A(LNH-ST) by genetic engineering. Appl. Microbiol. Biotechnol. 87, 1803–1811. doi: 10.1007/s00253-010-2609-0
Bettiga, M., Bengtsson, O., Hahn-Hägerdal, B., and Gorwa-Grauslund, M. F. (2009). Arabinose and xylose fermentation by recombinant Saccharomyces cerevisiae expressing a fungal pentose utilization pathway. Microb. Cell Fact. 8:40. doi: 10.1186/1475-2859-8-40
Betts, W. B., Dart, R. K., Ball, A. S., and Pedlar, S. L. (1991). “Biosynthesis and structure of lignocellulose,” in Biodegradation: Natural and Synthetic Material, ed W. B. Betts (London: Springer), 139–115. doi: 10.1007/978-1-4471-3470-1_7
Boer, H., Teeri, T. T., and Koivula, A. (2000). Characterization of Trichoderma reesei cellobiohydrolase Cel7A secreted from Pichia pastoris using two different promoters. Biotechnol. Bioeng. 69, 486–494. doi: 10.1002/1097-0290(20000905)69:5<486::AID-BIT3>3.0.CO;2-N
Bogorad, I. W., Chen, C. T., Theisen, M. K., Wu, T. Y., Schlenz, A. R., Lam, A. T., et al. (2014). Building carbon-carbon bonds using a biocatalytic methanol condensation cycle. Proc. Natl. Acad. Sci. U.S.A. 111, 15928–15933. doi: 10.1073/pnas.1413470111
Bogorad, I. W., Lin, T. S., and Liao, J. C. (2013). Synthetic non-oxidative glycolysis enables complete carbon conservation. Nature 502, 693–697. doi: 10.1038/nature12575
Bracher, J. M., Verhoeven, M. D., Wisselink, H. W., Crimi, B., Nijland, J. G., Driessen, A. J. M., et al. (2018). The Penicillium chrysogenum transporter PcAraT enables high-affinity, glucose-insensitive l-arabinose transport in Saccharomyces cerevisiae. Biotechnol. Biofuels 11:63. doi: 10.1186/s13068-018-1047-6
Brat, D., Boles, E., and Wiedemann, B. (2009). Functional expression of a bacterial xylose isomerase in Saccharomyces cerevisiae. Appl. Environ. Microbiol. 75, 2304–2311. doi: 10.1128/AEM.02522-08
Brenner, K., You, L., and Arnold, F. H. (2008). Engineering microbial consortia: a new frontier in synthetic biology. Trends Biotechnol. 26, 483–489. doi: 10.1016/j.tibtech.2008.05.004
Brethauer, S., and Studer, M. H. (2014). Consolidated bioprocessing of lignocellulose by a microbial consortium. Energy Environ. Sci. 7, 1446–1453. doi: 10.1039/c3ee41753k
Bruinenberg, P. M., Debot, P. H. M., Vandijken, J. P., and Scheffers, W. A. (1984). NADH-linked aldose reductase - the key to anaerobic alcoholic fermentation of xylose by yeasts. Appl. Microbiol. Biotechnol. 19, 256–260. doi: 10.1007/BF00251847
Caballero, A., and Ramos, J. L. (2017). Enhancing ethanol yields through D-xylose and L-arabinose co-fermentation after construction of a novel high efficient L-arabinose-fermenting Saccharomyces cerevisiae strain. Microbiology 163, 442–452. doi: 10.1099/mic.0.000437
Cao, M., Gao, M., Lopez-Garcia, C. L., Wu, Y., Seetharam, A. S., Severin, A. J., et al. (2017a). Centromeric DNA facilitates nonconventional yeast genetic engineering. ACS Synth. Biol. 6, 1545–1553. doi: 10.1021/acssynbio.7b00046
Cao, M., Gao, M., Ploessl, D., Song, C., and Shao, Z. (2017b). CRISPR–mediated genome editing and gene repression in Scheffersomyces stipitis. Biotechnol. J. 13:e1700598. doi: 10.1002/biot.201700598
Cao, M., Seetharam, A. S., Severin, A. J., and Shao, Z. (2017c). Rapid isolation of centromeres from Scheffersomyces stipitis. ACS Synth. Biol. 6, 2028–2034. doi: 10.1021/acssynbio.7b00166
Carriquiry, M. A., Du, X., and Timilsina, G. R. (2011). Second generation biofuels: economics and policies. Int. J. Polit. Econ. Plann. Environ. Soc. Aspect. Energy 39, 4222–4234. doi: 10.1016/j.enpol.2011.04.036
Chakravorty, M., Veiga, L. A., Horecker, B. L., and Bacila, M. (1962). Pentose metabolism in Candida. II. The diphosphopyridine nucleotide-specific polyol dehydrogenase of Candida utilis. J. Biol. Chem. 237, 1014–1020.
Chandel, A. K., Chandrasekhar, G., Radhika, K., Ravinder, R., and Ravindra, P. (2011). Bioconversion of pentose sugars into ethanol: a review and future directions. Biotechnol. Molecular Biol. Rev. 6, 8–20.
Chen, Y., Wu, Y., Zhu, B., Zhang, G., and Wei, N. (2018). Co-fermentation of cellobiose and xylose by mixed culture of recombinant Saccharomyces cerevisiae and kinetic modeling. PLoS ONE 13, e0199104. doi: 10.1371/journal.pone.0199104
Cheng, C., Tang, R. Q., Xiong, L., Hector, R. E., Bai, F. W., and Zhao, X. Q. (2018). Association of improved oxidative stress tolerance and alleviation of glucose repression with superior xylose-utilization capability by a natural isolate of Saccharomyces cerevisiae. Biotechnol. Biofuels 11:28. doi: 10.1186/s13068-018-1018-y
Chiang, C., and Knight, S. G. (1961). L-Arabinose Metabolism by cell-free extracts of Penicillium Chrysogenum. Biochim. Biophys. Acta. 46, 271–278. doi: 10.1016/0006-3002(61)90750-8
Chiang, L. C., Gong, C. S., Chen, L. F., and Tsao, G. T. (1981). D-Xylulose fermentation to ethanol by Saccharomyces cerevisiae. Appl. Environ. Microbiol. 42, 284–289.
Chu, S., and Majumdar, A. (2012). Opportunities and challenges for a sustainable energy future. Nature 488, 294–303. doi: 10.1038/nature11475
Delgenes, J. P., Escare, M. C., Laplace, J. M., Moletta, R., and Navarro, J. M. (1998). Biological production of industrial chemicals, i.e. xylitol and ethanol, from lignocelluloses by controlled mixed culture systems. Ind. Crop. Prod. 7, 101–111. doi: 10.1016/S0926-6690(97)00038-1
Dien, B. S., Kurtzman, C. P., Saha, B. C., and Bothast, R. J. (1996). Screening for L-arabinose fermenting yeasts. Appl. Biochem. Biotechnol. 57–8, 233–242. doi: 10.1007/BF02941704
Du, J., Li, S., and Zhao, H. (2010). Discovery and characterization of novel D-xylose-specific transporters from Neurospora crassa and Pichia stipitis. Mol. Biosyst. 6, 2150–2156. doi: 10.1039/c0mb00007h
Du, J., Yuan, Y., Si, T., Lian, J., and Zhao, H. (2012). Customized optimization of metabolic pathways by combinatorial transcriptional engineering. Nucleic Acids Res. 40:e142. doi: 10.1093/nar/gks549
Eiteman, M. A., Lee, S. A., and Altman, E. (2008). A co-fermentation strategy to consume sugar mixtures effectively. J. Biol. Eng. 2:3. doi: 10.1186/1754-1611-2-3
Eliasson, A., Hofmeyr, J. S., Pedler, S., and Hahn-Hägerdal, B. (2001). The xylose reductase/xylitol dehydrogenase/xylulokinase ratio affects product formation in recombinant xylose-utilising Saccharomyces cerevisiae. Enzyme Microb. Technol. 29, 288–297. doi: 10.1016/S0141-0229(01)00386-6
Fan, L. H., Zhang, Z. J., Yu, X. Y., Xue, Y. X., and Tan, T. W. (2012). Self-surface assembly of cellulosomes with two miniscaffoldins on Saccharomyces cerevisiae for cellulosic ethanol production. Proc. Natl. Acad. Sci. U.S.A. 109, 13260–13265. doi: 10.1073/pnas.1209856109
Farwick, A., Bruder, S., Schadeweg, V., Oreb, M., and Boles, E. (2014). Engineering of yeast hexose transporters to transport D-xylose without inhibition by D-glucose. Proc. Natl. Acad. Sci. U.S.A. 111, 5159–5164. doi: 10.1073/pnas.1323464111
Fengel, D., and Wegener, G. (1983). Wood: Chemistry, Ultrastructure, Reactions. Berlin: Walter de Gruyter. doi: 10.1515/9783110839654
Fiaux, J., Cakar, Z. P., Sonderegger, M., Wüthrich, K., Szyperski, T., and Sauer, U. (2003). Metabolic-flux profiling of the yeasts Saccharomyces cerevisiae and Pichia stipitis. Eukaryot. Cell 2, 170–180. doi: 10.1128/EC.2.1.170-180.2003
Galazka, J. M., Tian, C., Beeson, W. T., Martinez, B., Glass, N. L., and Cate, J. H. (2010). Cellodextrin transport in yeast for improved biofuel production. Science 330, 84–86. doi: 10.1126/science.1192838
Gao, M. R., Cao, M. F., Suástegui, M., Walker, J., Quiroz, N. R., Wu, Y., et al. (2017). Innovating a nonconventional yeast platform for producing shikimate as the building block of high-value aromatics. ACS Synth. Biol. 6, 29–38. doi: 10.1021/acssynbio.6b00132
Gírio, F. M., Fonseca, C., Carvalheiro, F., Duarte, L. C., Marques, S., and Bogel-Łukasik, R. (2010). Hemicelluloses for fuel ethanol: A review. Bioresour. Technol. 101, 4775–4800. doi: 10.1016/j.biortech.2010.01.088
Graves, T., Narendranath, N. V., Dawson, K., and Power, R. (2006). Effect of pH and lactic or acetic acid on ethanol productivity by Saccharomyces cerevisiae in corn mash. J. Ind. Microbiol. Biotechnol. 33, 469–474. doi: 10.1007/s10295-006-0091-6
Guadalupe-Medina, V., Wisselink, H. W., Luttik, M. A., de Hulster, E., Daran, J. M., Pronk, J. T., et al. (2013). Carbon dioxide fixation by Calvin-Cycle enzymes improves ethanol yield in yeast. Biotechnol. Biofuels 6:125. doi: 10.1186/1754-6834-6-125
Ha, S. J., Galazka, J. M., Kim, S. R., Choi, J. H., Yang, X., Seo, J. H., et al. (2011). Engineered Saccharomyces cerevisiae capable of simultaneous cellobiose and xylose fermentation. Proc. Natl. Acad. Sci. U.S.A. 108, 504–509. doi: 10.1073/pnas.1010456108
Ha, S. J., Wei, Q., Kim, S. R., Galazka, J. M., Cate, J. H., and Jin, Y. S. (2011). Cofermentation of cellobiose and galactose by an engineered Saccharomyces cerevisiae strain. Appl. Environ. Microbiol. 77, 5822–5825. doi: 10.1128/AEM.05228-11
Hahn-Hägerdal, B., Karhumaa, K., Jeppsson, M., and Gorwa-Grauslund, M. F. (2007). Metabolic engineering for pentose utilization in Saccharomyces cerevisiae. Biofuels 108, 147–177. doi: 10.1007/10_2007_062
Hamacher, T., Becker, J., Gárdonyi, M., Hahn-Hägerdal, B., and Boles, E. (2002). Characterization of the xylose-transporting properties of yeast hexose transporters and their influence on xylose utilization. Microbiology 148, 2783–2788. doi: 10.1099/00221287-148-9-2783
Hanly, T. J., and Henson, M. A. (2013). Dynamic metabolic modeling of a microaerobic yeast co-culture: predicting and optimizing ethanol production from glucose/xylose mixtures. Biotechnol. Biofuels 6:44. doi: 10.1186/1754-6834-6-44
Harhangi, H. R., Akhmanova, A. S., Emmens, R., van der Drift, C., de Laat, W. T., van Dijken, J. P., et al. (2003). Xylose metabolism in the anaerobic fungus Piromyces sp. strain E2 follows the bacterial pathway. Arch Microbiol. 180, 134–141. doi: 10.1007/s00203-003-0565-0
Hector, R. E., Dien, B. S., Cotta, M. A., and Mertens, J. A. (2013). Growth and fermentation of D-xylose by Saccharomyces cerevisiae expressing a novel D-xylose isomerase originating from the bacterium Prevotella ruminicola TC2-24. Biot. Biofuels 6:84. doi: 10.1186/1754-6834-6-84
Hector, R. E., Qureshi, N., Hughes, S. R., and Cotta, M. A. (2008). Expression of a heterologous xylose transporter in a Saccharomyces cerevisiae strain engineered to utilize xylose improves aerobic xylose consumption. Appl. Microbiol. Biotechnol. 80, 675–684. doi: 10.1007/s00253-008-1583-2
Heux, S., Cachon, R., and Dequin, S. (2006). Cofactor engineering in Saccharomyces cerevisiae: Expression of a H2O-forming NADH oxidase and impact on redox metabolism. Metab. Eng. 8, 303–314. doi: 10.1016/j.ymben.2005.12.003
Hniman, A., O.-,Thong, S., and Prasertsan, P. (2011). Developing a thermophilic hydrogen-producing microbial consortia from geothermal spring for efficient utilization of xylose and glucose mixed substrates and oil palm trunk hydrolysate. Int. J. Hydrogen Energy 36, 8785–8793. doi: 10.1016/j.ijhydene.2010.09.067
Hou, J., Suo, F., Wang, C., Li, X., Shen, Y., and Bao, X. (2014). Fine-tuning of NADH oxidase decreases byproduct accumulation in respiration deficient xylose metabolic Saccharomyces cerevisiae. BMC Biotechnol. 14:13. doi: 10.1186/1472-6750-14-13
Jeon, E., Hyeon, Je., Eun, L. S., Park, B. S., Kim, S. W., Lee, J., et al. (2009). Cellulosic alcoholic fermentation using recombinant Saccharomyces cerevisiae engineered for the production of Clostridium cellulovorans endoglucanase and Saccharomycopsis fibuligera beta-glucosidase. Fems. Microbiol. Lett. 301, 130–136. doi: 10.1111/j.1574-6968.2009.01808.x
Jeppsson, M., Bengtsson, O., Franke, K., Lee, H., ahn-Hägerdal, B., and Gorwa-Grauslund, M. F. (2006). The expression of a Pichia stipitis xylose reductase mutant with higher K-M for NADPH increases ethanol production from xylose in recombinant Saccharomyces cerevisiae. Biotechnol. Bioeng. 93, 665–673. doi: 10.1002/bit.20737
Jia, X., Liu, C., Song, H., Ding, M., Du, J., Ma, Q., et al. (2016). Design, analysis and application of synthetic microbial consortia. Synth. Syst. Biotechnol. 1, 109–117. doi: 10.1016/j.synbio.2016.02.001
Jin, Y. S., Alper, H., Yang, Y. T., and Stephanopoulos, G. (2005). Improvement of xylose uptake and ethanol production in recombinant Saccharomyces cerevisiae through an inverse metabolic engineering approach. Appl. Environ. Microbiol. 71, 8249–8256. doi: 10.1128/AEM.71.12.8249-8256.2005
Jin, Y. S., and Jeffries, T. W. (2003). Changing flux of xylose metabolites by altering expression of xylose reductase and xylitol dehydrogenase in recombinant Saccharomyces cerevisiae. Appl. Biochem. Biotechnol. 105–108, 277–286. doi: 10.1385/ABAB:106:1-3:277
Jin, Y. S., Lee, T. H., Choi, Y. D., Ryu, Y. W., and Seo, J. H. (2000). Conversion of xylose to ethanol by recombinant Saccharomyces cerevisiae containing genes for xylose reductase and xylitol dehydrogenase from Pichia stipitis. J. Microbiol. Biotechn. 10, 564–567.
Jin, Y. S., Ni, H., Laplaza, J. M., and Jeffries, T. W. (2003). Optimal growth and ethanol production from xylose by recombinant Saccharomyces cerevisiae require moderate D-xylulokinase activity. Appl. Environ. Microbiol. 69, 495–503. doi: 10.1128/AEM.69.1.495-503.2003
Johansson, B., Christensson, C., Hobley, T., and ahn-Hägerdal, B. (2001). Xylulokinase overexpression in two strains of Saccharomyces cerevisiae also expressing xylose reductase and xylitol dehydrogenase and its effect on fermentation of xylose and lignocellulosic hydrolysate. Appl. Environ. Microbiol. 67, 4249–4255. doi: 10.1128/AEM.67.9.4249-4255.2001
Jojima, T., Omumasaba, C. A., Inui, M., and Yukawa, H. (2010). Sugar transporters in efficient utilization of mixed sugar substrates: current knowledge and outlook. Appl. Microbiol. Biotechnol. 85, 471–480. doi: 10.1007/s00253-009-2292-1
Kang, M. H., Ni, H., and Jeffries, T. W. (2003). Molecular characterization of a gene for aldose reductase (CbXYL1) from Candida boidinii and its expression in Saccharomyces cerevisiae. Appl. Biochem. Biotechnol. 105–108, 265–276. doi: 10.1385/ABAB:106:1-3:265
Karhumaa, K., Garcia Sanchez, R., Hahn-Hägerdal, B., and Gorwa-Grauslund, M. F. (2007). Comparison of the xylose reductase-xylitol dehydrogenase and the xylose isomerase pathways for xylose fermentation by recombinant Saccharomyces cerevisiae. Microb. Cell Fact. 6:5. doi: 10.1186/1475-2859-6-5
Karhumaa, K., Hahn-Hägerdal, B., and Gorwa-Grauslund, M. F. (2005). Investigation of limiting metabolic steps in the utilization of xylose by recombinant Saccharomyces cerevisiae using metabolic engineering. Yeast 22, 359–368. doi: 10.1002/yea.1216
Katahira, S., Fujita, Y., Mizuike, A., Fukuda, H., and Kondo, A. (2004). Construction of a xylan-fermenting yeast strain through codisplay of xylanolytic enzymes on the surface of xylose-utilizing Saccharomyces cerevisiae cells. Appl. Environ. Microbiol. 70, 5407–5414. doi: 10.1128/AEM.70.9.5407-5414.2004
Katahira, S., Ito, M., Takema, H., Fujita, Y., Tanino, T., Tanaka, T., et al. (2008). Improvement of ethanol productivity during xylose and glucose co-fermentation by xylose-assimilating S. cerevisiae via expression of glucose transporter Sut1, Enzyme Microb. Technol. 43, 115–119. doi: 10.1016/j.enzmictec.2008.03.001
Katahira, S., Mizuike, A., Fukuda, H., and Kondo, A. (2006). Ethanol fermentation from lignocellulosic hydrolysate by a recombinant xylose- and cellooligosaccharide-assimilating yeast strain. Appl. Microbiol. Biotechnol. 72, 1136–1143. doi: 10.1007/s00253-006-0402-x
Khoury, G. A., Fazelinia, H., Chin, J. W., Pantazes, R. J., Cirino, P. C., and Maranas, C. D. (2009). Computational design of Candida boidinii xylose reductase for altered cofactor specificity. Protein Sci. 18, 2125–2138. doi: 10.1002/pro.227
Kim, S. R., Ha, S. J., Wei, N., Oh, E. J., and Jin, Y. S. (2012). Simultaneous co-fermentation of mixed sugars: a promising strategy for producing cellulosic ethanol. Trends Biotechnol. 30, 274–282. doi: 10.1016/j.tibtech.2012.01.005
Knoshaug, E. P., Vidgren, V., Magalhaes, F., Jarvis, E. E., Franden, M. A., Zhang, M., et al. (2015). Novel transporters from Kluyveromyces marxianus and Pichia guilliermondii expressed in Saccharomyces cerevisiae enable growth on L-arabinose and D-xylose. Yeast 32, 615–628. doi: 10.1002/yea.3084
Kotter, P., Amore, R., Hollenberg, C. P., and Ciriacy, M. (1990). Isolation and characterization of the Pichia stipitis xylitol dehydrogenase gene. XYL2, and construction of a xylose-utilizing Saccharomyces cerevisiae transformant. Curr. Genet. 18, 493–500. doi: 10.1007/BF00327019
Kotter, P., and Ciriacy, M. (1993). Xylose fermentation by Saccharomyces cerevisiae. Appl. Microbiol. Biotechnol. 38, 776–783. doi: 10.1007/BF00167144
Kou, S. C., Christensen, M. S., and Cirillo, V. P. (1970). Galactose transport in Saccharomyces cerevisiae. II. Characteristics of galactose uptake and exchange in galactokinaseless cells. J. Bacteriol. 103, 671–678.
Kristo, P., Saarelainen, R., Fagerstrom, R., Aho, S., and Korhola, M. (1996). Protein purification, and cloning and characterization of the cDNA and gene for xylose isomerase of barley. Eur. J. Biochem. 237, 240–246. doi: 10.1111/j.1432-1033.1996.0240n.x
Kuyper, M., Harhangi, H. R., Stave, A. K., Winkler, A. A., Jetten, M. S., de Laat, W. T., et al. (2003). High-level functional expression of a fungal xylose isomerase: the key to efficient ethanolic fermentation of xylose by Saccharomyces cerevisiae?, FEMS Yeast Res. 4, 69–78. doi: 10.1016/S1567-1356(03)00141-7
Kuyper, M., Hartog, M. M., Toirkens, M. J., Almering, M. J., Winkler, A. A., van Dijken, J. P., et al. (2005). Metabolic engineering of a xylose-isomerase-expressing Saccharomyces cerevisiae strain for rapid anaerobic xylose fermentation. FEMS Yeast Res. 5, 399–409. doi: 10.1016/j.femsyr.2004.09.010
Kuyper, M., Winkler, A. A., van Dijken, J. P., and Pronk, J. T. (2004). Minimal metabolic engineering of Saccharomyces cerevisiae for efficient anaerobic xylose fermentation: a proof of principle. FEMS Yeast Res. 4, 655–664. doi: 10.1016/j.femsyr.2004.01.003
Lambertz, C., Garvey, M., Klinger, J., Heesel, D., Klose, H., Fischer, R., et al. (2014). Challenges and advances in the heterologous expression of cellulolytic enzymes: a review. Biotechnol. Biofuels 7:135. doi: 10.1186/s13068-014-0135-5
Lane, S., Xu, H. Q., Oh, E. J., Kim, H., Lesmana, A., Jeong, D., et al. (2018). Glucose repression can be alleviated by reducing glucose phosphorylation rate in Saccharomyces cerevisiae. Sci. Rep. 8:2613. doi: 10.1038/s41598-018-20804-4
Laplace, J. M., Delgenes, J. P., Moletta, R., and Navarro, J. M. (1993). Cofermentation of glucose and xylose to ethanol by a respiratory-deficient mutant of Saccharomyces cerevisiae co-cultivated with a xylose-fermenting yeast. J. Ferment. Bioeng. 75, 207–212. doi: 10.1016/0922-338X(93)90117-Q
Latimer, L. N., Lee, M. E., Medina-Cleghorn, D., Kohnz, R. A., Nomura, D. K., and Dueber, J. E. (2014). Employing a combinatorial expression approach to characterize xylose utilization in Saccharomyces cerevisiae. Metab. Eng. 25, 20–29. doi: 10.1016/j.ymben.2014.06.002
Leandro, M. J., Goncalves, P., and Spencer-Martins, I. (2006). Two glucose/xylose transporter genes from the yeast Candida intermedia: first molecular characterization of a yeast xylose-H+ symporter. Biochem. J. 395, 543–549. doi: 10.1042/BJ20051465
Lee, C. R., Sung, B. H., Lim, K. M., Kim, M. J., Sohn, M. J., Bae, J. H., et al. (2017). Co-fermentation using recombinant Saccharomyces cerevisiae yeast strains hyper-secreting different cellulases for the production of cellulosic bioethanol. Sci. Rep. 7:4428. doi: 10.1038/s41598-017-04815-1
Li, H. B., Schmitz, O., and Alper, H. S. (2016). Enabling glucose/xylose co-transport in yeast through the directed evolution of a sugar transporter. Appl. Microbiol. Biotechnol. 100, 10215–10223. doi: 10.1007/s00253-016-7879-8
Li, J., Xu, J., Cai, P., Wang, B., Ma, Y., Benz, J. P., et al. (2015). Functional analysis of two L-arabinose transporters from filamentous fungi reveals promising characteristics for improved pentose utilization in Saccharomyces cerevisiae. Appl. Environ. Microbiol. 81, 4062–4070. doi: 10.1128/AEM.00165-15
Li, S., Du, J., Sun, J., Galazka, J. M., Glass, N. L., Cate, J. H., et al. (2010). Overcoming glucose repression in mixed sugar fermentation by co-expressing a cellobiose transporter and a beta-glucosidase in Saccharomyces cerevisiae. Mol. Biosyst. 6, 2129–2132. doi: 10.1039/c0mb00063a
Li, X., Yu, V. Y., Lin, Y., Chomvong, K., Estrela, R., Park, A., et al. (2015). Expanding xylose metabolism in yeast for plant cell wall conversion to biofuels. Elife 4:05896. doi: 10.7554/eLife.05896
Li, Y. J., Wang, M. M., Chen, Y. W., Wang, M., Fan, L. H., Tan, T. W. (2017) Engineered yeast with a CO2-fixation pathway to improve the bio-ethanol production from xylose-mixed sugars. Sci. Rep. 7:43875., et al.
Liu, Z., Ho, S. H., Sasaki, K., den Haan, R., Inokuma, K., Ogino, C., et al. (2016). Engineering of a novel cellulose-adherent cellulolytic Saccharomyces cerevisiae for cellulosic biofuel production. Sci. Rep. 6:24550. doi: 10.1038/srep24550
Lobs, A. K., Schwartz, C., and Wheeldon, I. (2017). Genome and metabolic engineering in non-conventional yeasts: current advances and applications. Synth. Syst. Biotechnol. 2, 198–207. doi: 10.1016/j.synbio.2017.08.002
Madhavan, A., Srivastava, A., Kondo, A., and Bisaria, V. S. (2012). Bioconversion of lignocellulose-derived sugars to ethanol by engineered Saccharomyces cerevisiae. Crit. Rev. Biotechnol. 32, 22–48. doi: 10.3109/07388551.2010.539551
Madhavan, A., Tamalampudi, S., Srivastava, A., Fukuda, H., Bisaria, V. S., and Kondo, A. (2009). Alcoholic fermentation of xylose and mixed sugars using recombinant Saccharomyces cerevisiae engineered for xylose utilization. Appl. Microbiol. Biotechnol. 82, 1037–1047. doi: 10.1007/s00253-008-1818-2
Maehara, T., Takabatake, K., and Kaneko, S. (2013). Expression of Arabidopsis thaliana xylose isomerase gene and its effect on ethanol production in Flammulina velutipes. Fungal Biol. 117, 776–782. doi: 10.1016/j.funbio.2013.09.005
Matsushika, A., Nagashima, A., Goshima, T., and Hoshino, T. (2013). Fermentation of xylose causes inefficient metabolic state due to carbon/energy starvation and reduced glycolytic flux in recombinant industrial Saccharomyces cerevisiae. PLoS One 8:e69005. doi: 10.1371/journal.pone.0069005
Matsushika, A., and Sawayama, S. (2008). Efficient bioethanol production from xylose by recombinant Saccharomyces cerevisiae requires high activity of xylose reductase and moderate xylulokinase activity. J. Biosci. Bioeng. 106, 306–309. doi: 10.1263/jbb.106.306
Mattozzi, M., Ziesack, M., Voges, M. J., Silver, P. A., and Way, J. C. (2013). Expression of the sub-pathways of the Chloroflexus aurantiacus 3-hydroxypropionate carbon fixation bicycle in E. coli: toward horizontal transfer of autotrophic growth. Metab. Eng. 16, 130–139. doi: 10.1016/j.ymben.2013.01.005
Mcmillan, J. D., and Boynton, B. L. (1994). Arabinose utilization by xylose-fermenting yeasts and fungi. Appl. Biochem. Biotechnol. 45–6, 569–584. doi: 10.1007/BF02941831
Metzger, M. H., and Hollenberg, C. P. (1994). Isolation and characterization of the Pichia stipitis transketolase gene and expression in a xylose-utilising Saccharomyces cerevisiae transformant. Appl. Microbiol. Biotechnol. 42, 319–325. doi: 10.1007/s002530050257
Metzger, M. H., and Hollenberg, C. P. (1995). Amino acid substitutions in the yeast Pichia stipitis xylitol dehydrogenase coenzyme-binding domain affect the coenzyme specificity. Eur. J. Biochem. 228, 50–54. doi: 10.1111/j.1432-1033.1995.tb20227.x
Minty, J. J., Singer, M. E., Scholz, S. A., Bae, C. H., Ahn, J. H., Foster, C. E., et al. (2013). Design and characterization of synthetic fungal-bacterial consortia for direct production of isobutanol from cellulosic biomass. Proc. Natl. Acad. Sci. U. S. A. 110, 14592–14597. doi: 10.1073/pnas.1218447110
Moniruzzaman, M., Dien, B. S., Skory, C. D., Chen, Z. D., Hespell, R. B., Ho, N. W. Y., et al. (1997). Fermentation of corn fibre sugars by an engineered xylose utilizing Saccharomyces yeast strain. World J. Microbiol. Biotechnol. 13, 341–346. doi: 10.1023/A:1018547427078
Nakamura, N., Yamada, R., Katahira, S., Tanaka, T., Fukuda, H., and Kondo, A. (2008). Effective xylose/cellobiose co-fermentation and ethanol production by xylose-assimilating S. cerevisiae via expression of beta-glucosidase on its cell surface. Enzyme Microb. Technol. 43, 233–236. doi: 10.1016/j.enzmictec.2008.04.003
Nijland, J. G., Shin, H. Y., de Jong, R. M., de Waal, P. P., Klaassen, P., and Driessen, A. J. (2014). Engineering of an endogenous hexose transporter into a specific D-xylose transporter facilitates glucose-xylose co-consumption in Saccharomyces cerevisiae. Biotechnol. Biofuels 7:168. doi: 10.1186/s13068-014-0168-9
Olson, D. G., McBride, J. E., Shaw, A. J., and Lynd, L. R. (2012). Recent progress in consolidated bioprocessing. Curr. Opin. Biotechnol. 23, 396–405. doi: 10.1016/j.copbio.2011.11.026
Palmqvist, E., Grage, H., Meinander, N. Q., and Hahn-Hagerdal, B. (1999). Main and interaction effects of acetic acid, furfural, and p-hydroxybenzoic acid on growth and ethanol productivity of yeasts. Biotechnol. Bioeng. 63, 46–55. doi: 10.1002/(SICI)1097-0290(19990405)63:1<46::AID-BIT5>3.0.CO;2-J
Papapetridis, I., Verhoeven, M. D., Wiersma, S. J., Goudriaan, M., van Maris, A. J. A., and Pronk, J. T. (2018). Laboratory evolution for forced glucose-xylose co-consumption enables identification of mutations that improve mixed-sugar fermentation by xylose-fermenting Saccharomyces cerevisiae. FEMS Yeast Res. 18:foy056. doi: 10.1093/femsyr/foy056
Peng, B. Y., Huang, S. C., Liu, T. T., and Geng, A. L. (2015). Bacterial xylose isomerases from the mammal gut Bacteroidetes cluster function in Saccharomyces cerevisiae for effective xylose fermentation. Microb. Cell Fact. 14:70. doi: 10.1186/s12934-015-0253-1
Peters, M. S., Timmerhaus, K. D., and West, R. E. (2003). Plant Design and Economics for Chemical Engineers, 5th ed. New York, NY: McGraw-Hill.
Petschacher, B., and Nidetzky, B. (2008). Altering the coenzyme preference of xylose reductase to favor utilization of NADH enhances ethanol yield from xylose in a metabolically engineered strain of Saccharomyces cerevisiae. Microb. Cell Fact. 7:9. doi: 10.1186/1475-2859-7-9
Purnick, P. E., and Weiss, R. (2009). The second wave of synthetic biology: from modules to systems. Nat. Rev. Mol. Cell. Biol. 10, 410–422. doi: 10.1038/nrm2698
Rawat, U., Phadtare, S., Deshpande, V., and Rao, M. (1996). A novel xylose isomerase from Neurospora crassa. Biotechnol. Lett. 18, 1267–1270. doi: 10.1007/BF00129952
Reider Apel, A., Ouellet, M., Szmidt-Middleton, H., Keasling, J. D., and Mukhopadhyay, A. (2016). Evolved hexose transporter enhances xylose uptake and glucose/xylose co-utilization in Saccharomyces cerevisiae. Sci. Rep. 6:19512. doi: 10.1038/srep19512
Richard, P., Londesborough, J., Putkonen, M., Kalkkinen, N., and Penttila, M. (2001). Cloning and expression of a fungal L-arabinitol 4-dehydrogenase gene. J. Biol. Chem. 276, 40631–40637. doi: 10.1074/jbc.M104022200
Richard, P., Putkonen, M., Vaananen, R., Londesborough, J., and Penttila, M. (2002). The missing link in the fungal L-arabinose catabolic pathway, identification of the L-xylulose reductase gene. Biochemistry 41, 6432–6437. doi: 10.1021/bi025529i
Richard, P., Verho, R., Putkonen, M., Londesborough, J., and Penttila, M. (2003). Production of ethanol from L-arabinose by Saccharomyces cerevisiae containing a fungal L-arabinose pathway. FEMS Yeast Res. 3, 185–189. doi: 10.1016/S1567-1356(02)00184-8
Rodriguez-Pena, J. M., Cid, V. J., Arroyo, J., and Nombela, C. (1998). The YGR194c (XKS1) gene encodes the xylulokinase from the budding yeast Saccharomyces cerevisiae. FEMS Microbiol. Lett. 162, 155–160. doi: 10.1016/S0378-1097(98)00118-9
Rodrussamee, N., Sattayawat, P., and Yamada, M. (2018). Highly efficient conversion of xylose to ethanol without glucose repression by newly isolated thermotolerant Spathaspora passalidarum CMUWF1-2. BMC Microbiol. 18:73. doi: 10.1186/s12866-018-1218-4
Runquist, D., Fonseca, C., Radstrom, P., Spencer-Martins, I., and Hahn-Hagerdal, B. (2009). Expression of the Gxf1 transporter from Candida intermedia improves fermentation performance in recombinant xylose-utilizing Saccharomyces cerevisiae. Appl. Microbiol. Biotechnol. 82, 123–130. doi: 10.1007/s00253-008-1773-y
Runquist, D., Hahn-Hagerdal, B., and Radstrom, P. (2010). Comparison of heterologous xylose transporters in recombinant Saccharomyces cerevisiae. Biotechnol. Biofuels 3:5. doi: 10.1186/1754-6834-3-5
Ryu, S., Hipp, J., and Trinh, C. T. (2016). Activating and elucidating metabolism of complex sugars in Yarrowia lipolytica. Appl. Environ. Microbiol. 82, 1334–1345. doi: 10.1128/AEM.03582-15
Saha, B. C. (2003). Hemicellulose bioconversion. J. Ind. Microbiol. Biotechnol. 30, 279–291. doi: 10.1007/s10295-003-0049-x
Saitoh, S., Hasunuma, T., Tanaka, T., and Kondo, A. (2010). Co-fermentation of cellobiose and xylose using beta-glucosidase displaying diploid industrial yeast strain OC-2, Appl. Microbiol. Biotechnol. 87, 1975–1982. doi: 10.1007/s00253-010-2714-0
Sakamoto, T., Hasunuma, T., Hori, Y., Yamada, R., and Kondo, A. (2012). Direct ethanol production from hemicellulosic materials of rice straw by use of an engineered yeast strain codisplaying three types of hemicellulolytic enzymes on the surface of xylose-utilizing Saccharomyces cerevisiae cells. J. Biotechnol. 158, 203–210. doi: 10.1016/j.jbiotec.2011.06.025
Saloheimo, A., Rauta, J., Stasyk, O. V., Sibirny, A. A., Penttila, M., and Ruohonen, L. (2007). Xylose transport studies with xylose-utilizing Saccharomyces cerevisiae strains expressing heterologous and homologous permeases. Appl. Microbiol. Biotechnol. 74, 1041–1052. doi: 10.1007/s00253-006-0747-1
Sanchez, R. G., Karhumaa, K., Fonseca, C., Nogue, V. S., Almeida, J. R. M., Larsson, C. U., et al. (2010). Improved xylose and arabinose utilization by an industrial recombinant Saccharomyces cerevisiae strain using evolutionary engineering. Biotechnol. Biofuels 3:13. doi: 10.1186/1754-6834-3-13
Santos, M. A., Gomes, A. C., Santos, M. C., Carreto, L. C., and Moura, G. R. (2011). The genetic code of the fungal CTG clade. C. R. Biol. 334, 607–611. doi: 10.1016/j.crvi.2011.05.008
Schellenberg, G. D., Sarthy, A., Larson, A. E., Backer, M. P., Crabb, J. W., Lidstrom, M., et al. (1984). Xylose isomerase from Escherichia coli. Characterization of the protein and the structural gene. J. Biol. Chem. 259, 6826–6832.
Sedlak, M., and Ho, N. W. (2004). Characterization of the effectiveness of hexose transporters for transporting xylose during glucose and xylose co-fermentation by a recombinant Saccharomyces yeast. Yeast 21, 671–684. doi: 10.1002/yea.1060
Sedlak, M., and Ho, N. W. Y. (2001). Expression of E-coli araBAD operon encoding enzymes for metabolizing L-arabinose in Saccharomyces cerevisiae. Enzyme Microb. Technol. 28, 16–24. doi: 10.1016/S0141-0229(00)00282-9
Senac, T., and Hahn-Hagerdal, B. (1990). Intermediary metabolite concentrations in xylulose- and glucose-fermenting Saccharomyces cerevisiae cells. Appl. Environ. Microbiol. 56, 120–126.
Shapiro, R. S., Chavez, A., and Collins, J. J. (2018). CRISPR-based genomic tools for the manipulation of genetically intractable microorganisms. Nat. Rev. Microbiol. 16, 333–339. doi: 10.1038/s41579-018-0002-7
Shin, H. Y., Nijland, J. G., de Waal, P. P., de Jong, R. M., Klaassen, P., and Driessen, A. J. (2015). An engineered cryptic Hxt11 sugar transporter facilitates glucose-xylose co-consumption in Saccharomyces cerevisiae. Biotechnol. Biofuels 8:176. doi: 10.1186/s13068-015-0360-6
Solis-Escalante, D., van den Broek, M., Kuijpers, N. G., Pronk, J. T., Boles, E., Daran, J. M., et al. (2015). The genome sequence of the popular hexose-transport-deficient Saccharomyces cerevisiae strain EBY.VW4000 reveals LoxP/Cre-induced translocations and gene loss. FEMS Yeast Res. 15:fou004. doi: 10.1093/femsyr/fou004
Su, Y. K., Willis, L. B., and Jeffries, T. W. (2015). Effects of aeration on growth, ethanol and polyol accumulation by Spathaspora passalidarum NRRL Y-27907 and Scheffersomyces stipitis NRRL Y-7124, Biotechnol. Bioeng. 112, 457–469. doi: 10.1002/bit.25445
Suastegui, M., and Shao, Z. (2016). Yeast factories for the production of aromatic compounds: from building blocks to plant secondary metabolites. J. Ind. Microbiol. Biotechnol. 43, 1611–1624. doi: 10.1007/s10295-016-1824-9
Subtil, T., and Boles, E. (2011). Improving L-arabinose utilization of pentose fermenting Saccharomyces cerevisiae cells by heterologous expression of L-arabinose transporting sugar transporters. Biotechnol. Biofuels 4:38. doi: 10.1186/1754-6834-4-38
Sun, J., Wen, F., Si, T., Xu, J. H., and Zhao, H. (2012). Direct conversion of xylan to ethanol by recombinant Saccharomyces cerevisiae strains displaying an engineered minihemicellulosome. Appl. Environ. Microbiol. 78, 3837–3845. doi: 10.1128/AEM.07679-11
Suriyachai, N., Weerasaia, K., Laosiripojana, N., Champreda, V., and Unrean, P. (2013). Optimized simultaneous saccharification and co-fermentation of rice straw for ethanol production by Saccharomyces cerevisiae and Scheffersomyces stipitis co-culture using design of experiments. Bioresour. Technol. 142, 171–178. doi: 10.1016/j.biortech.2013.05.003
Tang, H., Song, M., He, Y., Wang, J., Wang, S., Shen, Y., et al. (2017). Engineering vesicle trafficking improves the extracellular activity and surface display efficiency of cellulases in Saccharomyces cerevisiae. Biotechnol. Biofuels 10:53. doi: 10.1186/s13068-017-0738-8
Taniguchi, M., Tohma, T., Itaya, T., and Fujii, M. (1997). Ethanol production from a mixture of glucose and xylose by co-culture of Pichia stipitis and a respiratory-deficient mutant of Saccharomyces cerevisiae. J. Ferment. Bioeng. 83, 364–370. doi: 10.1016/S0922-338X(97)80143-2
Tanimura, A., Takashima, M., Sugita, T., Endoh, R., Ohkuma, M., Kishino, S., et al. (2016). Lipid production through simultaneous utilization of glucose, xylose, and l-arabinose by Pseudozyma hubeiensis: a comparative screening study. AMB Expr. 6:58. doi: 10.1186/s13568-016-0236-6
Tantirungkij, M., Nakashima, N., Seki, T., and Yoshida, T. (1993). Construction of xylose-assimilating Saccharomyces cerevisiae. J. Ferment. Bioeng. 75, 83–88. doi: 10.1016/0922-338X(93)90214-S
Thomik, T., Wittig, I., Choe, J. Y., Boles, E., and Oreb, M. (2017). An artificial transport metabolon facilitates improved substrate utilization in yeast. Nat. Chem. Biol. 13, 1158–1163. doi: 10.1038/nchembio.245
Tian, C., Beeson, W. T., Iavarone, A. T., Sun, J., Marletta, M. A., Cate, J. H., et al. (2009). Systems analysis of plant cell wall degradation by the model filamentous fungus Neurospora crassa. Proc. Natl. Acad. Sci. U.S.A. 106, 22157–22162. doi: 10.1073/pnas.0906810106
Toivari, M. H., Aristidou, A., Ruohonen, L., and Penttila, M. (2001). Conversion of xylose to ethanol by recombinant Saccharomyces cerevisiae: importance of xylulokinase (XKS1) and oxygen availability. Metab. Eng. 3, 236–249. doi: 10.1006/mben.2000.0191
Toivari, M. H., Salusjarvi, L., Ruohonen, L., and Penttila, M. (2004). Endogenous xylose pathway in Saccharomyces cerevisiae. Appl. Environ. Microbiol. 70, 3681–3686. doi: 10.1128/AEM.70.6.3681-3686.2004
van Rooyen, R., Hahn-Hagerdal, B., La Grange, D. C., and van Zyl, W. H. (2005). Construction of cellobiose-growing and fermenting Saccharomyces cerevisiae strains. J. Biotechnol. 120, 284–295. doi: 10.1016/j.jbiotec.2005.06.013
Van Vleet, J. H., and Jeffries, T. W. (2009). Yeast metabolic engineering for hemicellulosic ethanol production. Curr. Opin. Biotechnol. 20, 300–306. doi: 10.1016/j.copbio.2009.06.001
Vemuri, G. N., Eiteman, M. A., McEwen, J. E., Olsson, L., and Nielsen, J. (2007). Increasing NADH oxidation reduces overflow metabolism in Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. U.S.A. 104, 2402–2407. doi: 10.1073/pnas.0607469104
Verho, R., Londesborough, J., Penttila, M., and Richard, P. (2003). Engineering redox cofactor regeneration for improved pentose fermentation in Saccharomyces cerevisiae. Appl. Environ. Microbiol. 69, 5892–5897. doi: 10.1128/AEM.69.10.5892-5897.2003
Verho, R., Penttila, M., and Richard, P. (2011). Cloning of two genes (LAT1,2) encoding specific L-arabinose transporters of the L-arabinose fermenting yeast Ambrosiozyma monospora. Appl. Biochem. Biotechnol. 164, 604–611. doi: 10.1007/s12010-011-9161-y
Verho, R., Putkonen, M., Londesborough, J., Penttila, M., and Richard, P. (2004). A novel NADH-linked l-xylulose reductase in the L-arabinose catabolic pathway of yeast. J. Biol. Chem. 279, 14746–14751. doi: 10.1074/jbc.M312533200
Verhoeven, M. D., de Valk, S. C., Daran, J. G., van Maris, A. J. A., and Pronk, J. T. (2018). Fermentation of glucose-xylose-arabinose mixtures by a synthetic consortium of single-sugar-fermenting Saccharomyces cerevisiae strains. FEMS Yeast Res. 18:oy075. doi: 10.1093/femsyr/foy075
Walfridsson, M., Hallborn, J., Penttila, M., Keranen, S., and Hahn-Hagerdal, B. (1995). Xylose-metabolizing Saccharomyces cerevisiae strains overexpressing the TKL1 and TAL1 genes encoding the pentose phosphate pathway enzymes transketolase and transaldolase. Appl. Environ. Microbiol. 61, 4184–4190.
Wang, C. Q., Shen, Y., Zhang, Y. Y., Suo, F., Hou, J., and Bao, X. M. (2013). Improvement of L-arabinose fermentation by modifying the metabolic pathway and transport in Saccharomyces cerevisiae. Biomed Res. Int. 2013:461204. doi: 10.1155/2013/461204
Wang, C. Q., Zhao, J. Z., Qiu, C. X., Wang, S. H., Shen, Y., Du, B. H., et al. (2017). Coutilization of D-glucose. D-xylose, and L-arabinose in Saccharomyces cerevisiae by coexpressing the metabolic pathways and evolutionary engineering. Biomed Res. Int. 2017:5318232. doi: 10.1155/2017/5318232
Wang, M., Yu, C., and Zhao, H. (2016). Directed evolution of xylose specific transporters to facilitate glucose-xylose co-utilization. Biotechnol. Bioeng. 113, 484–491. doi: 10.1002/bit.25724
Watanabe, S., Abu Saleh, A., Pack, S. P., Annaluru, N., Kodaki, T., and Makino, K. (2007). Ethanol production from xylose by recombinant Saccharomyces cerevisiae expressing protein-engineered NADH-preferring xylose reductase from Pichia stipitis. Microbiol-Sgm 153, 3044–3054. doi: 10.1099/mic.0.2007/007856-0
Watanabe, S., Kodaki, T., and Makino, K. (2005). Complete reversal of coenzyme specificity of xylitol dehydrogenase and increase of thermostability by the introduction of structural zinc. J. Biol. Chem. 280, 10340–10349. doi: 10.1074/jbc.M409443200
Watanabe, S., Saleh, A. A., Pack, S. P., Annaluru, N., Kodaki, T., and Makino, K. (2007). Ethanol production from xylose by recombinant Saccharomyces cerevisiae expressing protein engineered NADP+-dependent xylitol dehydrogenase. J. Biotechnol. 130, 316–319. doi: 10.1016/j.jbiotec.2007.04.019
Wei, N., Quarterman, J., Kim, S. R., Cate, J. H., and Jin, Y. S. (2013). Enhanced biofuel production through coupled acetic acid and xylose consumption by engineered yeast. Nat. Commun. 4:2580. doi: 10.1038/ncomms3580
Wieczorke, R., Krampe, S., Weierstall, T., Freidel, K., Hollenberg, C. P., and Boles, E. (1999). Concurrent knock-out of at least 20 transporter genes is required to block uptake of hexoses in Saccharomyces cerevisiae. FEBS Lett. 464, 123–128. doi: 10.1016/S0014-5793(99)01698-1
Wilhelm, M., and Hollenberg, C. P. (1984). Selective cloning of Bacillus subtilis xylose isomerase and xylulokinase in Escherichia coli genes by IS5-mediated expression. EMBO J. 3, 2555–2560. doi: 10.1002/j.1460-2075.1984.tb02173.x
Wisselink, H. W., Toirkens, M. J., del Rosario Franco Berriel, M., Winkler, A. A., van Dijken, J. P., Pronk, J. T., et al. (2007). Engineering of Saccharomyces cerevisiae for efficient anaerobic alcoholic fermentation of L-arabinose. Appl. Environ. Microbiol. 73, 4881–4891. doi: 10.1128/AEM.00177-07
Wisselink, H. W., Toirkens, M. J., Wu, Q., Pronk, J. T., and van Maris, A. J. A. (2009). Novel evolutionary engineering approach for accelerated utilization of glucose, xylose, and arabinose mixtures by engineered Saccharomyces cerevisiae strains. Appl. Environ. Microbiol. 75, 907–914. doi: 10.1128/AEM.02268-08
Witteveen, C. F. B., Busink, R., Vandevondervoort, P., Dijkema, C., Swart, K., and Visser, J. (1989). L-Arabinose and D-xylose catabolism in Aspergillus niger. J. Gen. Microbiol. 135, 2163–2171.
Wooley, R., Ruth, M., Sheehan, J., and Ibsen, K. (1999). Lignocellulosic Biomass to Ethanol Process Design and Economics Utilizing Co-current Dilute Acid Prehydrolysis and Enzymatic Hydrolysis Current and Futuristic Scenarios. Golden, CO: Biotechnology Center for Fuels and Chemicals.
Xia, P. F., Zhang, G. C., Walker, B., Seo, S. O., Kwak, S., Liu, J. J., et al. (2017). Recycling carbon dioxide during xylose fermentation by engineered Saccharomyces cerevisiae. ACS Synth. Biol. 6, 276–283. doi: 10.1021/acssynbio.6b00167
Xia, T., Eiteman, M. A., and Altman, E. (2012). Simultaneous utilization of glucose, xylose and arabinose in the presence of acetate by a consortium of Escherichia coli strains. Microb. Cell Fact. 11:77. doi: 10.1186/1475-2859-11-77
Xue, X. D., and Ho, N. W. Y. (1990). Xylulokinase activity in various yeasts including Saccharomyces cerevisiae containing the cloned cylulokinase gene. Appl. Biochem. Biotechnol. 24–5, 193–199.
Yang, B., Dai, Z. Y., Ding, S. Y., and Wyman, C. E. (2014). Enzymatic hydrolysis of cellulosic biomass. Adv. Biochem. Eng. Biot. 2, 421–449.
Young, E., Lee, S. M., and Alper, H. (2010). Optimizing pentose utilization in yeast: the need for novel tools and approaches. Biotechnol. Biofuels 3:24. doi: 10.1186/1754-6834-3-24
Young, E., Poucher, A., Comer, A., Bailey, A., and Alper, H. (2011). Functional survey for heterologous sugar transport proteins, using Saccharomyces cerevisiae as a host. Appl. Environ. Microbiol. 77, 3311–3319. doi: 10.1128/AEM.02651-10
Young, E. M., Comer, A. D., Huang, H., and Alper, H. S. (2012). A molecular transporter engineering approach to improving xylose catabolism in Saccharomyces cerevisiae. Metab. Eng. 14, 401–411. doi: 10.1016/j.ymben.2012.03.004
Young, E. M., Tong, A., Bui, H., Spofford, C., and Alper, H. S. (2014). Rewiring yeast sugar transporter preference through modifying a conserved protein motif. Proc. Natl. Acad. Sci. U.S.A. 111, 131–136. doi: 10.1073/pnas.1311970111
Yu, S., Jeppsson, H., and HahnHagerdal, B. (1995). Xylulose fermentation by Saccharomyces cerevisiae and xylose-fermenting yeast strains. Appl. Microbiol. Biotechnol. 44, 314–320. doi: 10.1007/BF00169922
Zaldivar, J., Nielsen, J., and Olsson, L. (2001). Fuel ethanol production from lignocellulose: a challenge for metabolic engineering and process integration. Appl. Microbiol. Biotechnol. 56, 17–34. doi: 10.1007/s002530100624
Zhang, C., Acosta-Sampson, L., Yu, V. Y., and Cate, J. H. (2017). Screening and directed evolution of transporters to improve xylodextrin utilization in the yeast Saccharomyces cerevisiae. PLoS ONE 12:e0184730. doi: 10.1371/journal.pone.0184730
Zhang, G. C., Liu, J. J., and Ding, W. T. (2012). Decreased xylitol formation during xylose fermentation in Saccharomyces cerevisiae due to overexpression of water-forming NADH oxidase. Appl. Environ. Microbiol. 78, 1081–1086. doi: 10.1128/AEM.06635-11
Zhang, H., Pereira, B., Li, Z., and Stephanopoulos, G. (2015). Engineering Escherichia coli coculture systems for the production of biochemical products. Proc. Natl. Acad. Sci. U.S.A. 112, 8266–8271. doi: 10.1073/pnas.1506781112
Zhou, H., Cheng, J. S., Wang, B. L., Fink, G. R., and Stephanopoulos, G. (2012). Xylose isomerase overexpression along with engineering of the pentose phosphate pathway and evolutionary engineering enable rapid xylose utilization and ethanol production by Saccharomyces cerevisiae. Metab. Eng. 14, 611–622. doi: 10.1016/j.ymben.2012.07.011
Keywords: mixed-sugar utilization, carbon catabolite repression, non-glucose transporters, consolidated bioprocessing, microbial consortium, cellulase, hemicellulase
Citation: Gao M, Ploessl D and Shao Z (2019) Enhancing the Co-utilization of Biomass-Derived Mixed Sugars by Yeasts. Front. Microbiol. 9:3264. doi: 10.3389/fmicb.2018.03264
Received: 25 June 2018; Accepted: 14 December 2018;
Published: 22 January 2019.
Edited by:
Ryan S. Senger, Virginia Tech, United StatesReviewed by:
Soo Rin Kim, Kyungpook National University, South KoreaCopyright © 2019 Gao, Ploessl and Shao. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Zengyi Shao, enlzaGFvQGlhc3RhdGUuZWR1
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.