
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
METHODS article
Front. Mar. Sci. , 20 March 2025
Sec. Marine Conservation and Sustainability
Volume 12 - 2025 | https://doi.org/10.3389/fmars.2025.1518562
The decline in eel resources, coupled with the challenges of morphological identification and the rise of illegal trade, highlights the urgent need for accurate species identification techniques. To address this, a multiplex PCR assay was developed, targeting the mitochondrial Cytochrome b gene. Species-specific primers were designed and their efficacy validated through single PCR. The multiplex PCR conditions were then optimized to enable the simultaneous amplification of five major eel species. This assay exhibited high accuracy, specificity, and sensitivity, successfully identifying all five species even when DNA concentrations were low. This multiplex PCR assay offers a rapid and cost-effective solution for eel species identification, with the potential to significantly bolster eel conservation efforts. By enabling accurate species identification, it can help combat illegal trade and support the sustainable management of eel resources.
Anguillid eels (often called freshwater eels), with their wide global distribution, hold significant ecological and economic importance (Jacoby et al., 2015). Their unique catadromous life cycle, coupled with their high economic value as a delicacy in East Asian countries like Korea, Japan, and China, has led to intense fishing pressure and concerns about their sustainability (Tsukamoto and Arai, 2001; Yuan et al., 2022). Eels are catadromous fish, meaning they migrate from freshwater to saltwater to spawn (Tsukamoto and Arai, 2001). The hatched larvae then embark on a long journey back to rivers, aided by ocean currents, where they mature into adults (Durif et al., 2022). This complex life cycle poses a significant challenge for effective eel resource management.
In recent years, eel resources have been rapidly declining due to a combination of factors, including overfishing, habitat destruction, marine pollution, climate change, and dam construction (Drouineau et al., 2018). This decline has raised international concerns, underscored by the listing of the Japanese eel (Anguilla japonica) as an endangered species on the International Union for Conservation of Nature (IUCN) Red List in 2014, and the European eel (A. anguilla) in 2020 (Jacoby and Gollock, 2014; Pike et al., 2020). These circumstances necessitate urgent and concerted efforts to effectively manage and conserve eel resources (Jacoby et al., 2015; Righton et al., 2021). The IUCN Red List, while providing crucial information on species status, does not inherently possess regulatory mechanisms. However, it serves as a critical foundation for conservation efforts, including the implementation of international agreements such as the Convention on International Trade in Endangered Species of Wild Fauna and Flora (CITES).
CITES plays a vital role in regulating international trade in endangered species, including Anguillid eels, to ensure their survival. The accurate identification of eel species is essential for CITES implementation, particularly for monitoring trade, enforcing regulations, and preventing illegal trafficking. The morphological identification of eel species, particularly challenging at the glass eel stage, has led to serious problems such as illegal fishing and smuggling, as well as origin fraud (Stein et al., 2016; Alonso and van Uhm, 2023). These issues not only pose significant obstacles to eel resource management but also disrupt the ecological balance, further increasing the risk of extinction for some species (Stein et al., 2021). This problem extends beyond glass eels to eel products in the food industry, where mislabeling and species substitution can occur (Goymer et al., 2023). Accurate species identification is crucial for combating fraud and ensuring the legality and sustainability of the eel trade. In this respect, DNA analysis can be a valuable tool for identifying the species and origin of eel products, contributing to the conservation efforts and responsible management of eel resources (Noh et al., 2018).
Therefore, the development of accurate and rapid eel species identification techniques is crucial for the sustainable use and conservation of eel resources. However, most existing eel species identification techniques have limitations. Single PCR assays can only identify one species at a time, while methods such as real-time PCR or PNA probe assays involve high costs, complex procedures, and the need for skilled personnel and expensive equipment (Sezaki et al., 2005; Itoi et al., 2005; Noh et al., 2018). While semi-multiplex PCR has been employed for eel species identification, it has been observed to produce non-specific amplification in addition to species-specific identification (Fahmi et al., 2013). These limitations hinder both the field applicability and the efficiency of eel species identification.
This study aimed to address these challenges by developing a multiplex PCR assay capable of accurately and efficiently identifying five commercially important eel species imported into Korea: Japanese eel (A. japonica), American eel (A. rostrata), Indonesian shortfin eel (A. bicolor pacifica), European eel (A. anguilla), and giant mottled eel (A. marmorata) (Noh et al., 2018). These eels are imported both live and as processed fisheries products. Live eels, primarily glass eels, are imported for aquaculture purposes, while processed fisheries products include various forms such as fresh, frozen, and processed eels. The developed assay utilizes markers that specifically bind to and amplify species-specific regions within the mitochondrial Cytochrome b (Cytb) gene (Minegishi et al., 2005).
This approach offers several advantages, including simplicity, cost-effectiveness, and easy visualization of results through gel electrophoresis (Matsunaga et al., 1999). Therefore, the findings of this study are expected to have broad applicability in various fields related to eel resource management and conservation, including on-site eel species identification, quarantine inspection of imported eels, authenticity verification of eel products, and the prevention of illegal fishing and trade (Gill, 2007; Espiñeira and Vieites, 2016), ultimately contributing to the establishment of a science-based management system for the sustainable use and conservation of eel resources, the maintenance of ecological balance, and the effective implementation of international regulations like CITES.
In this study, mitochondrial Cytb gene sequences for five eel species (A. japonica, A. rostrata, A. bicolor pacifica, A. anguilla, and A. marmorata) were obtained from the NCBI GenBank database (Supplementary Table S1). Multiple sequences per species were collected to account for intraspecific genetic variation. The obtained sequences were aligned using MegAlign Pro v17.3.0 software (DNASTAR Lasergene, Madison, WI, USA), and interspecific sequence differences were analyzed to identify species-specific regions using DnaSP v5.10.01 software (Librado and Rozas, 2009). Based on these regions, a universal reverse primer (binding to all five eel species) and species-specific forward primers were designed, with special consideration given to placing the species-specific nucleotide variations at the 3’ end of the primers to enhance specificity (Kang et al., 2015). Primer design also considered specificity, amplicon size, Tm value, and avoidance of secondary structure formation to enhance PCR efficiency and species identification accuracy (Dieffenbach et al., 1993). The designed primers were synthesized by Genotech Corporation (Seoul, Korea).
Muscle tissues were obtained from 42 eel samples, with up to 10 individuals per species, from the National Institute of Fisheries Science’s Fisheries Bio-Resources Storage (Busan, Korea) (Table 1). Each individual was an accurately identified adult specimen to ensure the reliability of the results. The collected tissue samples were incubated with 8M TNES-UREA solution and Proteinase K at 37°C for 12 hours for protein digestion and cell lysis (Chomczynski and Sacchi, 1987). Genomic DNA was then extracted using the phenol-chloroform extraction and ethanol precipitation methods (Wasko et al., 2003). The purified DNA was quantified and its purity assessed using a NanoPhotometer N60 (IMPLEN, Munich, Germany). Only high-quality DNA samples, free of PCR inhibitors, were used for further experiments. The purified DNA samples were stored at -20°C until further use.
To validate the specificity and sensitivity of the designed primers, single PCR amplifications were conducted using genomic DNA samples from each eel species. Single PCR is a technique to amplify a specific target DNA sequence using a single pair of primers. Gradient PCR was performed to determine the optimal annealing temperature for species-specific primer binding. The annealing temperature was systematically varied across a range of 50-65°C in a single PCR reaction. PCR reactions were prepared using a 2X PCR master mix (iNtRON Biotech, Seoul, Korea) according to the manufacturer’s instructions. A typical PCR master mix contains Taq DNA polymerase, dNTPs, MgCl2, and reaction buffer. Each PCR reaction contained 1 μL of template DNA, 1 μL of each primer (10 pmol), 10 μL of 2X PCR master mix, and 7 μL of distilled water, resulting in a final volume of 20 μL. The PCR amplification was performed using an VeritiTM thermal cycler (Applied Biosystems, CA, USA) under the following conditions: an initial denaturation at 94°C for 7 minutes, followed by 34 cycles of denaturation at 94°C for 30 seconds, annealing at 50-65°C (gradient) for 30 seconds, and extension at 72°C for 1 minute, with a final extension at 72°C for 7 minutes. Amplified PCR products were visualized on a 1.5% agarose gel and imaged using a GelDoc Go system (BioRad, Hercules, CA, USA). Subsequently, the PCR products were finally verified by sequencing analysis. Sequencing was performed using an ABI 3730XL DNA Analyzer (Applied Biosystems, Foster City, CA, USA), and the species were identified using NCBI BLAST (https://blast.ncbi.nlm.nih.gov/Blast.cgi).
PCR amplicon sizes for each eel species were confirmed, and multiplex PCR conditions (primer concentration and annealing temperature) were optimized stepwise to ensure efficient and specific amplification. The PCR reaction mixture contained template DNA, five species-specific primer sets, and a commercial PCR master mix. PCR products were visualized by gel electrophoresis (1.5% agarose gel) and stained with Loading star (DyneBio, Daejeon, Korea). The initial primer concentration was set at 0.3 μL (10 pmol) per primer and was adjusted based on the amplification results to achieve optimal conditions. The annealing temperature was optimized to ensure the specific binding of all five primers. To determine the sensitivity of the selected primer sets, DNA samples from each eel species were diluted to concentrations of 10 ng/μL, 1 ng/μL, 0.1 ng/μL, and 0.01 ng/μL, and then analyzed under the optimized conditions. The PCR conditions for this analysis were identical to those used in the single PCR.
Analysis of the complete 1,140 bp mitochondrial Cytb gene sequences from a total of 281 individuals representing five eel species (obtained from NCBI GenBank) revealed that each species possessed various haplotypes, with 276 nucleotide variations observed across all individuals. While there was variation within each species, distinct sequence differences were consistently observed between species, allowing for clear differentiation. The Japanese eel (A. japonica) exhibited 30 haplotypes, including seven that were unique sequence variations. The American eel (A. rostrata) had 29 haplotypes, including four that were unique sequence variations. The Indonesian shortfin eel (A. bicolor pacifica) exhibited 26 haplotypes, including 14 that were unique sequence variations. The European eel (A. anguilla) had 48 haplotypes, including two that were unique sequence variations. Finally, the giant mottled eel (A. marmorata) had 97 haplotypes, with seven unique sequence variations.
Based on the identified species-specific sequence variations, a universal reverse primer and species-specific forward primers were designed. Species-specific primers were designed based on the identified unique sequence variations, prioritizing A. anguilla, which exhibited the fewer unique variations. The primer for A. anguilla was designed to target a T nucleotide at position 329 bp (A in other species) at its 3’ end. Next, a primer for A. rostrata was designed, targeting a C nucleotide at position 510 bp (T in other species). Subsequently, the A. marmorata-specific primer was designed to target a T nucleotide at position 654 bp (C in other species). For A. japonica, the primer targeted a C nucleotide at position 115 bp (T in others), and finally, the primer for A. bicolor pacifica targeted an A nucleotide at position 789 bp (C or T in other species). A universal reverse primer was also designed to bind to a conserved region in the mitochondrial genome downstream of the Cytb gene, ensuring its compatibility with all five eel species (Table 2, Figure 1).

Figure 1. Species-specific sequence variations in the mitochondrial Cytb region of Anguilla species.
To validate the specificity and sensitivity of the designed primers, single PCR amplifications were conducted using genomic DNA samples from each eel species. Single PCR is a technique to amplify a specific target DNA sequence using a single pair of primers. Gradient PCR was performed to determine the optimal annealing temperature for species-specific primer binding. The annealing temperature was systematically varied across a range of 50-65°C in a single PCR reaction. Gradient PCR analysis revealed that an annealing temperature of 62°C resulted in species-specific binding and amplification for all selected primers. Single PCR assays using these primers on DNA samples from each species yielded PCR amplicons of the expected sizes (Japanese eel: 1,119 bp, European eel: 910 bp, American eel: 722 bp, giant mottled eel: 576 bp, Indonesian shortfin eel: 444 bp), confirming primer specificity (Figure 2). Sequencing of these amplicons confirmed their identity, with each sequence perfectly matching that of the corresponding target species.
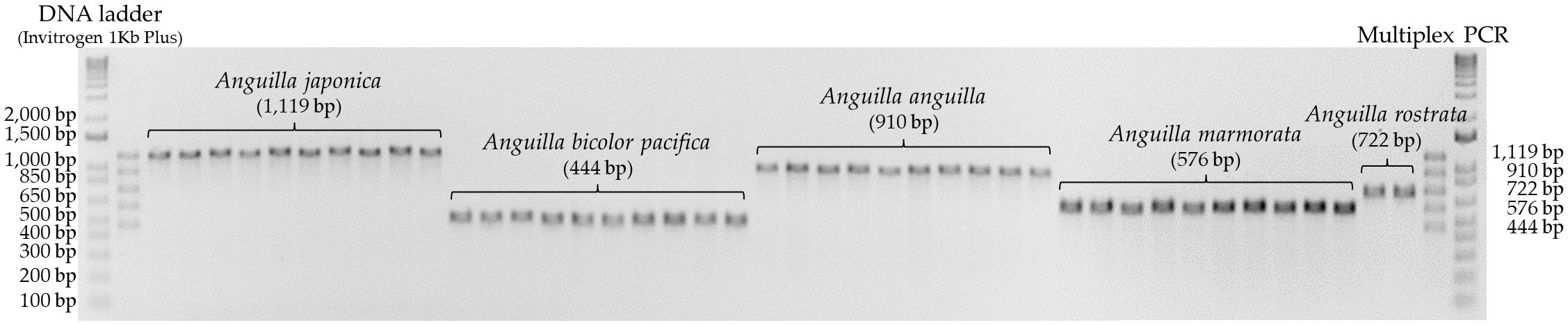
Figure 2. Results of single PCR amplification using species-specific primers for five Anguilla species.
To optimize the assay for simultaneous identification of all five eel species, optimal conditions were explored by mixing all five primers together. Based on the optimal temperature of 62°C determined in single PCR, the primer concentration was gradually increased from a minimum of 0.3 µl (10 pmol) to ensure clear interpretation of PCR amplification results. It was found that A. bicolor pacifica and A. marmorata required approximately three times higher primer concentrations than other species. In a final reaction volume of 20 µl, the optimal primer concentrations were determined to be 0.3 µl (10 pmol) for A. japonica, A. rostrata, and A. anguilla, and 1 µl (10 pmol) for A. bicolor pacifica and A. marmorata (Table 3). The optimized multiplex PCR conditions yielded clear results, allowing for the successful differentiation of all five eel species (Figure 3).
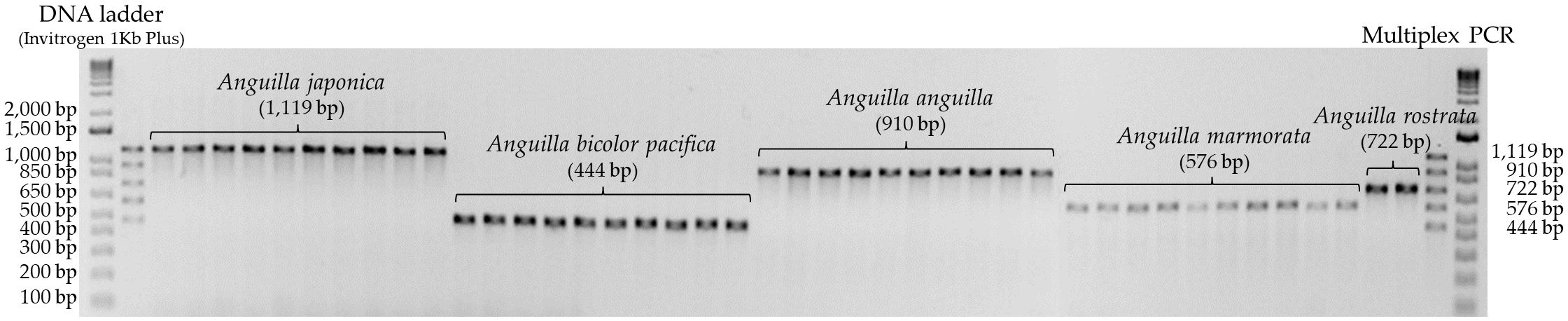
Figure 3. Results of multiplex PCR amplification using species-specific primers for five Anguilla species.
Furthermore, when DNA from all five eel species was mixed and analyzed, the species-specific PCR products were clearly separated on a 1.5% agarose gel, with no interference or cross-reactivity. The assay accurately identified all 42 eel samples (up to 10 individuals/species) based on expected amplicon patterns. Sensitivity evaluation using diluted DNA samples showed successful detection of all five species even at 0.1 ng/µL, confirming the robustness of the method (Figure 4).
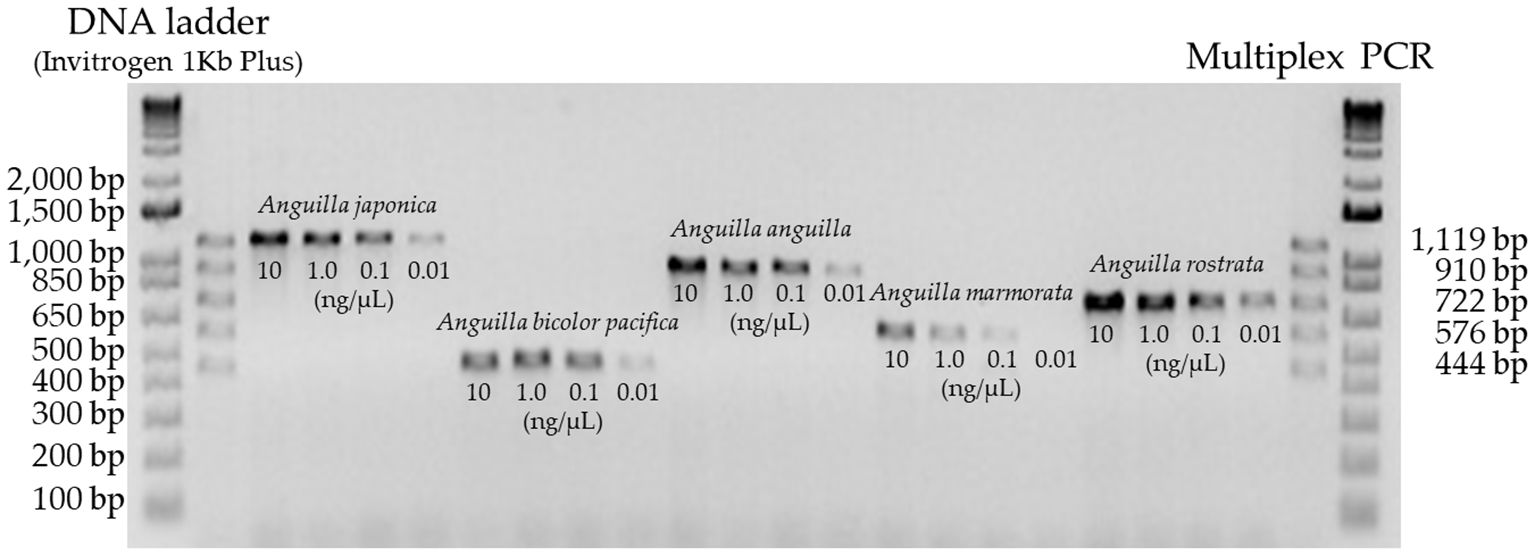
Figure 4. Sensitivity evaluation of the multiplex PCR assay for five Anguilla species using different DNA concentrations (10 ng/µL, 1.0 ng/µL, 0.1 ng/µL, and 0.01 ng/µL).
This study successfully developed a multiplex PCR assay for accurate identification of five major eel species. Through the analysis of mitochondrial Cytb gene sequences and identification of species-specific sequence variations, a set of primers was designed that enable precise identification of these five eel species within a single PCR reaction. The assay demonstrated high specificity and sensitivity, overcoming the limitations of single PCR assays and the high cost/complexity of real-time PCR or PNA probe assays. This represents a significant advancement in the field of eel species identification, particularly considering the challenges associated with morphological identification, especially at the glass eel stage.
The use of the mitochondrial Cytb gene as a genetic marker aligns with previous research demonstrating its effectiveness in species discrimination due to its high variability and maternal inheritance (Sezaki et al., 1999; Aoyama et al., 2001; Fahmi et al., 2013; Noh et al., 2018). A multiplex PCR system based on species-specific primers was established, and a comprehensive analysis of the complete Cytb gene sequences available in public database was conducted to identify species-specific sequence variation for primer design. This approach offers several distinct advantages over existing methods, including its simplicity, cost-effectiveness, and straightforward visualization of results through gel electrophoresis, rendering it highly suitable for field applications and resource-limited settings (Kim et al., 2023).
This assay can simplify and aid monitoring and enforcement efforts, particularly in combating illegal fishing and trade of endangered species (Jaser et al., 2021). Specifically, this assay can directly support the implementation of CITES by providing a reliable tool for monitoring international trade in Anguillid eels. By enabling accurate species identification, it can help to detect illegal trafficking, ensure that trade is conducted in accordance with CITES regulations, and prevent illegal fishing and smuggling of glass eels, thereby safeguarding valuable eel resources. Moreover, this assay can contribute to the establishment of science-based management system for sustainable eel resources by providing accurate species identification data for monitoring populations, assessing genetic diversity, and informing conservation strategies. Furthermore, the assay’s ability to verify the authenticity of eel products enhances consumer confidence and promotes fair trade practices, which are crucial for sustainable eel aquaculture and market regulation (Espiñeira and Vieites, 2016).
Beyond its immediate impact on eel conservation, this study highlights the power of molecular tools in addressing critical challenges in biodiversity conservation and resource management. The development of similar multiplex PCR assays for other taxa, particularly those facing threats from overexploitation or illegal trade, could significantly enhance our capacity for effective monitoring and enforcement. While this study focused on five commercially important eel species in Korea, the assay’s applicability could be expanded in future research by incorporating primers for additional species, particularly those that are threatened or play crucial ecological roles. This would enhance the assay’s utility for broader biodiversity monitoring and conservation efforts. Moreover, the integration of this assay with emerging technologies like environmental DNA (eDNA) analysis and portable sequencing platforms could further revolutionize our approach to biodiversity assessment and conservation (Cho et al., 2016; Farley et al., 2018).
Further improvements could involve simplifying DNA extraction procedures, potentially through the use of commercial kits or non-invasive sampling methods such as collecting DNA with swabs or eDNA, and refining primer design for even faster and more efficient analysis (Mason and Botella, 2020). The development of portable, field-deployable devices such as microfluidic platforms that integrate DNA extraction and multiplex PCR analysis would greatly enhance the assay’s on-site utility for rapid species identification and monitoring (Khodakov et al., 2021).
This study presents a significant advancement in eel resource management and conservation by developing a multiplex PCR assay capable of rapid and accurate identification of five major eel species. The design of species-specific primers based on the mitochondrial Cytb gene ensures high specificity and sensitivity, overcoming the limitations of existing methods. In particular, the accurate identification of species at the glass eel stage can prevent illegal fishing and smuggling, contributing to the establishment of a scientific management system for the sustainable use of eel resources.
The approach developed in this study has the potential to be applied to the conservation and management of not only eels but also other endangered or overfished species. Future expansion of the assay’s scope and technological improvements are expected to make even greater contributions to the field of biodiversity conservation and resource management.
The original contributions presented in the study are included in the article/supplementary material. Further inquiries can be directed to the corresponding author.
The animal study was approved by Institutional Animal Care and Use Committee of National Institute of Fisheries Science, Republic of Korea (2024-NIFS-IACUC-2). The study was conducted in accordance with the local legislation and institutional requirements.
EN: Conceptualization, Funding acquisition, Methodology, Project administration, Supervision, Writing – original draft, Writing – review & editing. CD: Methodology, Writing – review & editing. HP: Software, Writing – review & editing. EK: Conceptualization, Writing – review & editing. HJ: Software, Writing – review & editing. HK: Investigation, Writing – review & editing. YK: Investigation, Writing – review & editing.
The author(s) declare that financial support was received for the research and/or publication of this article. This work was supported by a grant from the National Institute of Fisheries Science (R2025018), Republic of Korea.
We thank the National Institute of Fisheries Science (NIFS), Ministry of Oceans and Fisheries, for their help in grant and research work.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fmars.2025.1518562/full#supplementary-material
Alonso A. I., van Uhm D. P. (2023). The illegal trade in European eels: outsourcing, funding, and complex symbiotic-antithetical relationships. Trends Organ. Crime 26, 293–307. doi: 10.1007/s12117-023-09490-5
Aoyama J., Nishida M., Tsukamoto K. (2001). Molecular phylogeny and evolution of the freshwater eel, genus Anguilla. Mol. Phylogenet. Evol. 20, 450–459. doi: 10.1006/mpev.2001.0959
Cho A., Morris T., Wilson C., Freeland J. (2016). Development of species-specific primers with potential for amplifying eDNA from imperilled freshwater unionid mussels. Genome 59, 1141–1149. doi: 10.1139/gen-2015-0196
Chomczynski P., Sacchi N. (1987). Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal. Biochem. 162, 156–159. doi: 10.1101/pdb.prot4455
Dieffenbach C. W., Lowe T. M., Dveksler G. S. (1993). General concepts for PCR primer design. PCR Methods Appl. 3, 30–37. doi: 10.1101/gr.3.3.s30
Drouineau H., Durif C., Castonguay M., Mateo M., Rochard E., Verreault G., et al. (2018). Freshwater eels: a symbol of the effects of global change. Fish Fish. 19, 903–930. doi: 10.1111/faf.12621
Durif C. M. F., Stockhausen H. H., Skiftesvik A. B., Cresci A., Nyqvist D., Browman H. I. (2022). A unifying hypothesis for the spawning migrations of temperate Anguillid eels. Fish Fish. 23, 358–375. doi: 10.1111/faf.12621
Espiñeira M., Vieites J. M. (2016). Genetic system for an integral traceability of European eel (Anguilla Anguilla) in aquaculture and seafood products: authentication by fast real-time PCR. Eur. Food Res. Technol. 242, 25–31. doi: 10.1007/s00217-015-2514-y
Fahmi M. R., Solihin D. D., Soewardi K., Pouyaud L., Shao Z., Berrebi P. (2013). A novel semi-multiplex PCR assay for identification of tropical eels of genus Anguilla in Indonesian waters. Fish. Sci. 79, 185–191. doi: 10.1007/s12562-012-0587-0
Farley N. J., Vasquez A. A., Kik R. IV, David S. R., Katailiha A. S., Walker X. N., et al. (2018). Primer designs for identification and environmental DNA (eDNA) detection of gars. Trans. Am. Fish. Soc 147, 687–695. doi: 10.1002/tafs.10043
Gill L. A. (2007). PCR-based methods for fish and fishery products authentication. Trends Food Sci. Technol. 18, 558–566. doi: 10.1016/j.tifs.2007.04.016
Goymer A., Steele K., Jenkins F., Burgess G., Andrews L., Baumgartner N., et al. (2023). For R-eel?! Investigating international sales of critically endangered species in freshwater eel products with DNA barcoding. Food Control 150, 109752. doi: 10.1016/j.foodcont.2023.109752
Itoi S., Nakaya M., Kaneko G., Kondo H., Sezaki K., Watabe S. (2005). Rapid identification of eels Anguilla japonica and Anguilla Anguilla by polymerase chain reaction with single nucleotide polymorphism-based specific probes. Fish. Sci. 71, 1356–1364. doi: 10.1111/j.1444-2906.2005.01102.x
Jacoby D. M. P., Casselman J. M., Crook V., DeLucia M. B., Ahn H., Kaifu K., et al. (2015). Synergistic patterns of threat and the challenges facing global Anguillid eel conservation. Glob. Ecol. Conserv. 4, 321–333. doi: 10.1016/j.gecco.2015.07.009
Jacoby D., Gollock M. (2014). Anguilla japonica. The IUCN red list of threatened species 2014. doi: 10.2305/IUCN.UK.2014-1.RLTS.T166184A1117791.en
Jaser S. K. K., Domingues R. R., Hilsdorf A. W. S. (2021). Assessing illegal fishing and trade of Atlantic billfish and swordfish by DNA-based identification. Conserv. Genet. Resour. 13, 183–190. doi: 10.1007/s12686-021-01190-y
Kang J. H., Noh E. S., Park J. Y., An C. M., Choi J. H., Kim J. K. (2015). Rapid origin determination of the Northern mauxia shrimp (Acetes chinensis) based on allele specific polymerase chain reaction of partial mitochondrial 16S rRNA Gene. Asian Australas. J. Anim. Sci. 28, 568–572. doi: 10.5713/ajas.14.0613
Khodakov D., Li J., Zhang J. X., Zhang D. Y. (2021). Highly multiplexed rapid DNA detection with single-nucleotide specificity via convective PCR in a portable device. Nat. Biomed. Eng. 5, 702–712. doi: 10.1038/s41551-021-00755-4
Kim E. M., Lee M. N., Dong C. M., Noh J. K., Noh E. S., Kim W. J., et al. (2023). Development of a multiplex Polymerase chain reaction method for rapid and accurate identification of Girella punctata and G. leonine (Teleostei: Girellidae). Fishes 8, 415. doi: 10.3390/fishes8080415
Librado P., Rozas J. (2009). DnaSP v5: A software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25, 1451–1452. doi: 10.1093/bioinformatics/btp187
Mason M. G., Botella J. R. (2020). Rapid (30-Second), Equipment-free purification of nucleic acids using easy-to-make dipsticks. Nat. Protoc. 15, 3663–3677. doi: 10.1038/s41596-020-0392-7
Matsunaga T., Chikuni K., Tanabe R., Muroya S., Shibata K., Yamada J., et al. (1999). A quick and simple method for the identification of meat species and meat products by PCR assay. Meat Sci. 51, 143–148. doi: 10.1016/S0309-1740(98)00112-0
Minegishi Y., Aoyama J., Inoue J. G., Miya M., Nishida M., Tsukamoto K. (2005). Molecular phylogeny and evolution of the freshwater eels genus Anguilla based on the whole mitochondrial genome sequences. Mol. Phylogenet. Evol. 34, 134–146. doi: 10.1016/j.ympev.2004.09.003
Noh E. S., Kang H. S., Kim E. M., Noh J. K., Park J. Y., Choi T. J., et al. (2018). Rapid differentiation of seven species of Anguilla using PNA clamping-based asymmetric PCR with fluorescence melting curve analysis. BioChip J. 12, 46–51. doi: 10.1007/s13206-017-2106-y
Pike C., Crook V., Gollock M. (2020). “Anguilla Anguilla,” in The IUCN red list of threatened species. doi: 10.2305/IUCN.UK.2020-2.RLTS.T60344A152845178.en
Righton D., Piper A., Aarestrup K., Amilhat E., Belpaire C., Casselman J., et al. (2021). Important questions to progress science and sustainable management of Anguillid eels. Fish Fish. 22, 762–788. doi: 10.1111/faf.12549
Sezaki K., Begum R. A., Wongrat P., Srivastava M. P., SriKantha S., Kikuchi K., et al. (1999). Molecular phylogeny of Asian freshwater and marine stingrays based on the DNA nucleotide and deduced amino acid Sequences of the cytochrome b gene. Fish. Sci. 65, 563–570. doi: 10.2331/fishsci.65.563
Sezaki K., Itoi S., Watabe S. (2005). A simple method to distinguish two commercially valuable eel species in Japan Anguilla japonica and A. Anguilla using polymerase chain reaction strategy with a species-specific primer. Fish. Sci. 71, 414–421. doi: 10.1111/j.1444-2906.2005.00979.x
Stein F. M., Frankowski J., Nijman V., Absil C., Kranendonk I., Dekker W. (2021). Chinese eel products in EU markets imply the effectiveness of trade regulations but expose fraudulent labelling. Mar. Pol. 132, 104651. doi: 10.1016/j.marpol.2021.104651
Stein F. S., Wong J. C. Y., Sheng V., Law C. S. W., Schröder B., Baker D. M. (2016). First genetic evidence of illegal trade in endangered european eel (Anguilla Anguilla) from Europe to Asia. Conserv. Genet. Resour. 8, 533–537. doi: 10.1007/s12686-016-0576-1
Tsukamoto K., Arai T. (2001). Facultative catadromy of the eel Anguilla japonica between freshwater and seawater habitats. Mar. Ecol. Prog. Ser. 220, 265–276. doi: 10.3354/meps220265
Wasko A. P., Martins C., Oliveira C., Foresti F. (2003). Non-destructive genetic sampling in fish. An improved method for DNA extraction from fish fins and scales. Hereditas 138, 161–165. doi: 10.1034/j.1601-5223.2003.01503.x
Keywords: anguilla, multiplex PCR, species identification, Cytochrome b, species-specific markers
Citation: Noh ES, Dong CM, Park HJ, Kim EM, Jung HS, Kong HJ and Kim YO (2025) Development of species identification techniques for anguillid eels using species-specific genetic markers. Front. Mar. Sci. 12:1518562. doi: 10.3389/fmars.2025.1518562
Received: 28 October 2024; Accepted: 03 March 2025;
Published: 20 March 2025.
Edited by:
Vitor H. Paiva, University of Coimbra, PortugalReviewed by:
Khaled Mohammed Geba, Menoufia University, EgyptCopyright © 2025 Noh, Dong, Park, Kim, Jung, Kong and Kim. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Eun Soo Noh, bGFwZXJtQGtvcmVhLmty
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.