
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Mar. Sci. , 27 March 2025
Sec. Deep-Sea Environments and Ecology
Volume 12 - 2025 | https://doi.org/10.3389/fmars.2025.1515048
DNA barcoding is a method of identifying individual organisms using short DNA fragments matched to a database of reference sequences. For metazoan plankton, a high proportion of species that reside in the deep ocean still lack reliable reference sequences for genetic markers for barcoding and systematics. We report on substantial taxonomic and barcoding efforts across major zooplankton taxonomic groups collected from surface waters to the rarely sampled abyssopelagic zone (0 – 4300 m) from the Gulf of Alaska, North Pacific Ocean. Over 1000 specimens were identified, from which the mitochondrial 16S and COI and nuclear 18S rRNA genes were sequenced. In total, 1462 sequences for 254 unique taxa were generated, adding new barcodes for 107 species, including 12 undescribed species of cnidarians, that previously lacked DNA sequences for at least one of the three genes. Additionally, we introduce the use of a new Open Nomenclature qualifier deoxyribonucleic acid abbreviation DNA (e.g., Genus DNA species, DNA Genus). This qualifier was used for specimens that could not be morphologically identified but could be assigned a low-level taxonomic identification based on the clustering of DNA barcode genes using phylogenetic trees (100% bootstrap support), where at least one of the sequences in that clade could be referred to a physical specimen (or photographs) where identification could be corroborated through morphological analyses. DNA barcodes from this work are incorporated into the MetaZooGene Atlas and Database, an open-access data and metadata portal for barcoding genes used for classifying and identifying marine organisms. As environmental sequencing (i.e., metabarcoding, metagenetics, and eDNA) becomes an increasingly common approach in marine ecosystem studies, continued population of such reference DNA sequence databases must remain a high priority.
DNA barcoding is a method of identifying individual organisms to the species level by matching short DNA sequences known as DNA barcodes (Hebert et al., 2003; Bucklin et al., 2011) to reference sequences derived from reliably identified specimens. DNA barcodes have become an exceptional tool for studying marine zooplankton communities that are fundamentally complex in terms of their biodiversity, phylogeny, and ecology. Zooplankton communities are composed of both holoplankton (organisms that are pelagic for their entire life cycle) and meroplankton (organisms that split their cycle between benthic and pelagic phases/stages) that collectively have representatives from 15 phyla and 41 functional groups (WoRMS, 2024). It is challenging for taxonomists to effectively describe the broad-spectrum of zooplankton assemblages within an ecosystem because of the presence of morphologically indistinct meroplanktonic larvae and early life stages of holoplankton, morphologically similar sibling and identical cryptic species, and fragile gelatinous organisms that are often damaged during collection. The many taxonomic challenges associated with zooplankton make it difficult to effectively describe local assemblages as well as broader communities inhabiting pelagic ecosystems. DNA barcoding and metabarcoding approaches make it easier to identify organisms to the species-level at any life stage, fostering an improved understanding of biodiversity, community structure, and biogeographical patterns that are otherwise difficult to obtain through morphological approaches.
The mitochondrial cytochrome oxidase I subunit (COI) is the most broadly applied, standard barcode gene for its ability to identify organisms to the species level across all life stages of marine metazoans (Bucklin et al., 2010c). COI has been used extensively to examine intra- and interspecific variation in almost every major taxonomic group represented within the zooplankton community (Bucklin et al., 2021). COI is a powerful marker for measuring levels of biodiversity in the pelagic ecosystem through the detection of cryptic species, a phenomenon that has emerged as being more prolific than previously thought as large-scale and global molecular analyses are conducted on species of zooplankton (Lee, 2000; Holland et al., 2004; Miyamoto et al., 2012; Cornils et al., 2017). Nonetheless, COI has some notable drawbacks when applied to all groups of zooplankton (see Bucklin et al., 2011). For example, the slower evolutionary rate of COI in anthozoan cnidarians and sponges reduces intra- and interspecific genetic distances, decreasing the barcode gap necessary for reliable species-level identifications (Meyer and Paulay, 2005; Hellberg, 2006) of pelagic stages of species in these largely benthic groups. In addition, the existence of COI pseudogenes and primer mismatch across medusozoan diversity has led to a situation where mitochondrial 16S ribosomal RNA (rRNA) has been applied far more successfully than COI to identify medusozoans to the species level (Lindsay et al., 2015). For ctenophores and larvaceans (a.k.a., appendicularians) successful amplification of COI has proven even more problematic when using “universal” COI primers. Creating reference libraries in such cases can require extensive work on primer design and experimentation (see Christianson et al., 2022 on ctenophores). While 64% of the described species of ctenophores have reference COI sequences, only 8% of larvacean species have COI references available (MetaZooGene Atlas and Database, 2024). In the absence of the application of many and varied family- and species-specific primers (e.g., COI for ctenophores by Christianson et al., 2022), alternative markers are adopted by some communities, for instance the Internal Transcribe Spacer (ITS) is being used to distinguish recently diverged lineages of larvaceans (Masunaga et al., 2022).
In addition to mitochondrial markers, the nuclear 18S rRNA gene can detect divergence across invertebrate and vertebrate taxa and is routinely analyzed in environmental sequencing studies (i.e., metabarcoding, metagenetics, and eDNA). Although 18S rRNA is not a reliable gene for obtaining species-level identification for zooplankton, and thus not a true barcode gene by definition, hypervariable regions of 18S can reliably discriminate genera, families, and higher taxonomic groups and amplify taxa that are difficult to target with COI (Bucklin et al., 2019; Blanco-Bercial, 2020; Questel et al., 2021; Matthews and Blanco-Bercial, 2023). Due to its ability to detect a broader range of taxonomic groups, 18S metabarcoding can provide a more complete characterization of a biological community than COI (Casey et al., 2021; Questel et al., 2021; Zhao et al., 2021). Thus, metabarcoding analyses that incorporate multi-gene markers can provide unbiased estimates of marine eukaryotic biodiversity.
Over the past few decades, DNA barcoding programs focused on marine zooplankton include: the global Census of Marine Zooplankton (CMarZ; Bucklin et al., 2010b), the Census of Antarctic Marine Life (CAML), Barcodes of Marine Zooplankton in China (BoMZC), Arctic Ocean Diversity (ArcOD; Bucklin et al., 2010a), Marine Barcode of Life (MarBOL; now the International Barcode of Life), the Norwegian Barcode of Life (NorBOL), and the Bureau of Ocean Energy Management (BOEM) DNA barcoding program at the Smithsonian National Museum of Natural History. These programs, and other small-scale efforts (e.g., Cheng et al., 2013; Baek et al., 2016; Questel et al., 2021) have made substantial progress in barcoding pelagic species. DNA barcodes from zooplankton collected and sequenced across the global ocean have, and continue to be, incorporated into reference DNA sequence databases (e.g., SILVA and MetaZooGene) for the taxonomic assignment of millions of unknown reads analyzed in environmental sequencing studies.
It is estimated that between 1 - 2 million eukaryotic species currently exist in the global ocean (Mora et al., 2011; Rogers et al., 2023). Given that roughly a quarter million accepted species have been named and described (WoRMS, 2024), roughly 75 – 88% of marine species remain undescribed, presenting a major challenge to understanding marine ecosystems. Much of our knowledge about the biodiversity of metazoan plankton in the global ocean comes from observations from the epipelagic zone whereas the deep ocean, defined as waters below 200 m, is the largest biome on Earth and yet remains vastly understudied (Robison, 2008; Webb et al., 2010; Levin et al., 2019). It is thus plausible that much of the undescribed marine biodiversity currently lies within the deep ocean environment.
The North Pacific Ocean is known to be inhabited by 17,264 described invertebrate species (Figure 1), of which 5,548 are crustaceans - including 911 copepod species - and 11,716 belong to non-crustacean groups (MetaZooGene Atlas and Database, 2024). Across the North Pacific Ocean, the zooplankton communities from the epipelagic zone (0 – 200 m) are well described and studied. Few studies of mid- to deep-water zooplankton communities exist, with most species’ inventories originating from sampling conducted at Ocean Station Papa in the eastern North Pacific, the Hawaii Ocean Time-series (HOT Station ALOHA), and Japanese time-series station K2 (Miller et al., 1984; Mackas et al., 1998; Yamaguchi et al., 2002; Steinberg et al., 2008) or station P in Sagami Bay (Shimode et al., 2006). Sampling below the mesopelagic has occurred at locations across the western and central North Pacific and the Bering Sea basin (Homma and Yamaguchi, 2010; Stabeno et al., 2012; Yamaguchi et al., 2015) where zooplankton communities have been described from the bathypelagic zone. However, many studies have focused on group-specific assemblages and not the entire zooplankton community, leaving a large gap in our knowledge of the true levels of pelagic diversity for the North Pacific Ocean.
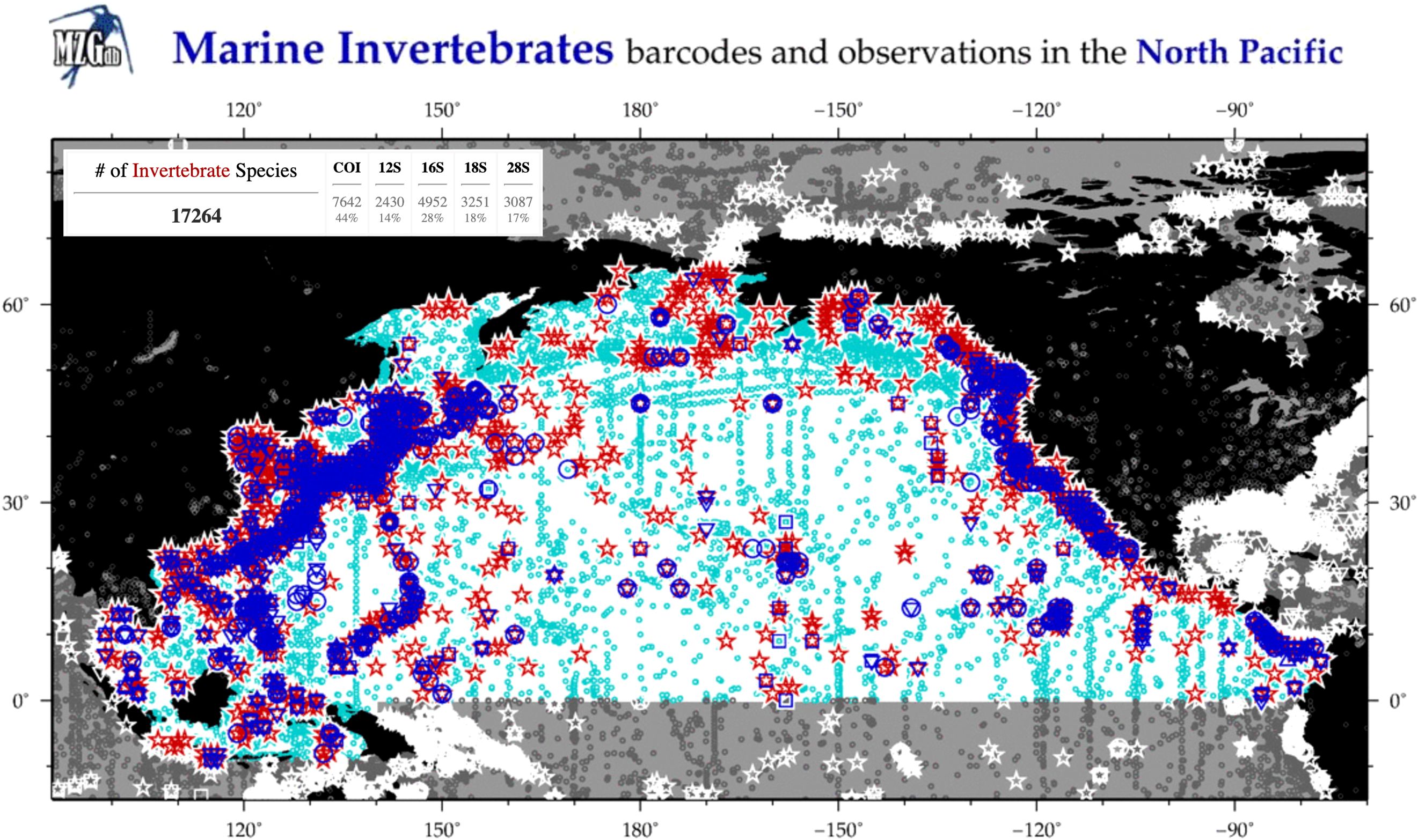
Figure 1. Map and statistics of marine invertebrate observations based on COPEPOD/OBIS (light blue circles) and DNA barcodes available in GenBank and BOLD (red stars) for the North Pacific Ocean. Data and figure from MetaZooGene Atlas and Database (https://metazoogene.org/atlas; accessed 18 October 2024).
Of the known zooplankton species documented in the North Pacific, only 44%, 28%, and 18% have DNA reads available for COI, 16S, and 18S rRNA, respectively (MetaZooGene Atlas and Database, 2024; Figure 1). Thus, the lack of studies of zooplankton communities from the global deep ocean has created a deficit in both estimates of biodiversity and DNA barcodes from currently described species, not to mention those in need of formal names and descriptions. Few studies have focused on DNA barcoding zooplankton communities from the deep ocean, with notable efforts in the Atlantic Ocean where zooplankton were barcoded from depths down to 5000 m (Wiebe et al., 2010). Our objectives were to identify and sequence the COI, 16S, and 18S rRNA genes from zooplankton collected from surface waters to the abyssopelagic zone (0 – 4300 m) in the Gulf of Alaska, North Pacific Ocean. These efforts will increase the representation of taxa for multiple genetic markers in reference DNA sequence databases routinely used in environmental sequencing studies, adding to efforts towards a global reference database for DNA barcodes of marine zooplankton (Bucklin et al., 2021).
The Gulf of Alaska is a distinct habitat within the North Pacific Ocean, bounded by the eastward flowing North Pacific Current that separates the subarctic Gulf of Alaska from the subtropical California Current System (Sutton et al., 2017). To the north and east, the Gulf of Alaska is bordered by highly productive shelf and coastal ecosystems, while offshore waters are presumed to be less productive under a High Nutrient Low Chlorophyll (HNLC) regime (Martin et al., 1989). Within these HNLC waters lies the Gulf of Alaska Seamount Province, a region characterized by hundreds of extinct submarine volcanoes (Menard and Dietz, 1951). The Seamount Province is composed of two distinct chains, the Kodiak-Bowie Seamount Chain that spans 900 km from the Aleutian Trench off eastern Kodiak Island (USA) to Queen Charlotte Island (CAN) and, to the south, the Cobb Seamount Chain that extends from the Aleutian Trench off western Kodiak Island to the Cobb hotspot on the Juan de Fuca Ridge (Menard and Dietz, 1951; Chaytor et al., 2007).
The pelagic ecosystem from surface waters to the abyssal plains was surveyed from 22 July – 2 August 2019 aboard the R/V Sikuliaq. Sampling was conducted at six oceanographic stations, including the transition zone from the Seward Line in the Northern Gulf of Alaska (https://nga.lternet.edu/) to the Quinn and Giacomini seamounts, located along the northern extent of the Kodiak-Bowie Seamount Chain (Figure 2). Surveys were conducted at the Quinn and Giacomini seamounts, sampling from surface waters to the summit (~650 m deep) and to the abyssal plain where the base of the seamounts lie between 4000 and 5000 m depth.

Figure 2. Location of the stations sampled for zooplankton in the Gulf of Alaska during the 2019 NOAA OE Seamounts cruise.
Zooplankton were collected using a variety of sampling techniques (Table 1). Two MultiNet Plankton Samplers Type Midi (0.25 m2; Hydro-Bios) each equipped with five 150-µm mesh plankton nets were fastened together and deployed in tandem (Figure 3); one MultiNet system was receiving and transmitting live feed from a 0.322 conducting wire, while the trigger depths for each net on the other MultiNet system were programmed before deployment. Securing two MultiNet systems together maximized wire time, a vital resource when conducting hours-long casts to the abyssopelagic zone. The MultiNet systems were equipped with five nets and thus required two sequential vertical casts to sample the entire water column for the following depth bins: 0 – 40 – 100 – 200 – 300 – 400 – 600 – 1200 – 2000 – 3000 – bottom (Table 1). When necessary, the top two strata were combined. Net samples from one MultiNet were preserved in 10% buffered formalin-seawater for taxonomic analysis, while net samples from the second MultiNet were split with a Folsom plankton splitter. One half of the sample was preserved in 95% ethanol while the other half was used for live sorting.
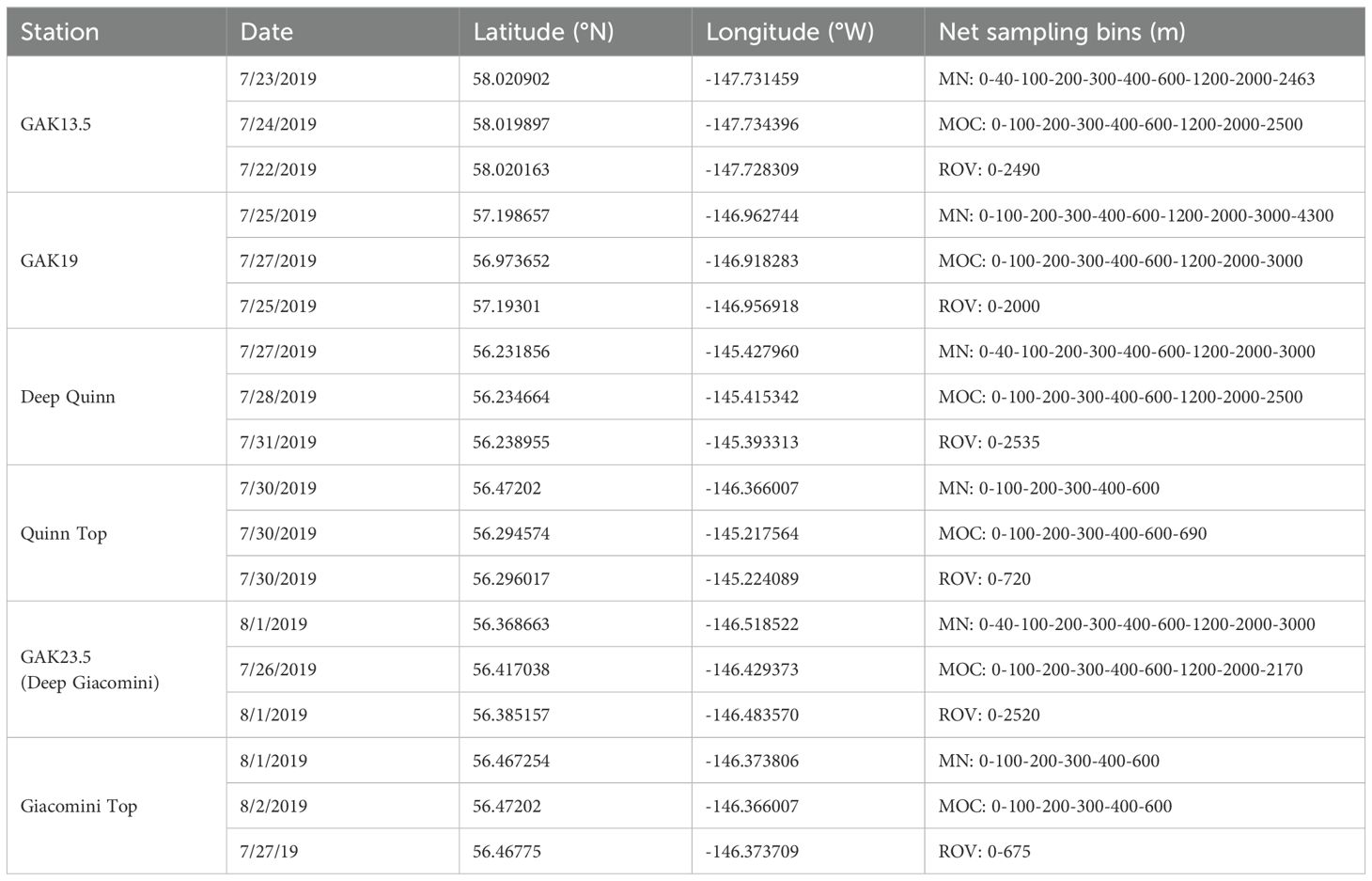
Table 1. Sampling data and net collection depth bins from stations occupied during the 2019 Gulf of Alaska Seamounts expedition. MN, MultiNet; MOC, MOCNESS; ROV, Remotely Operated Vehicle.
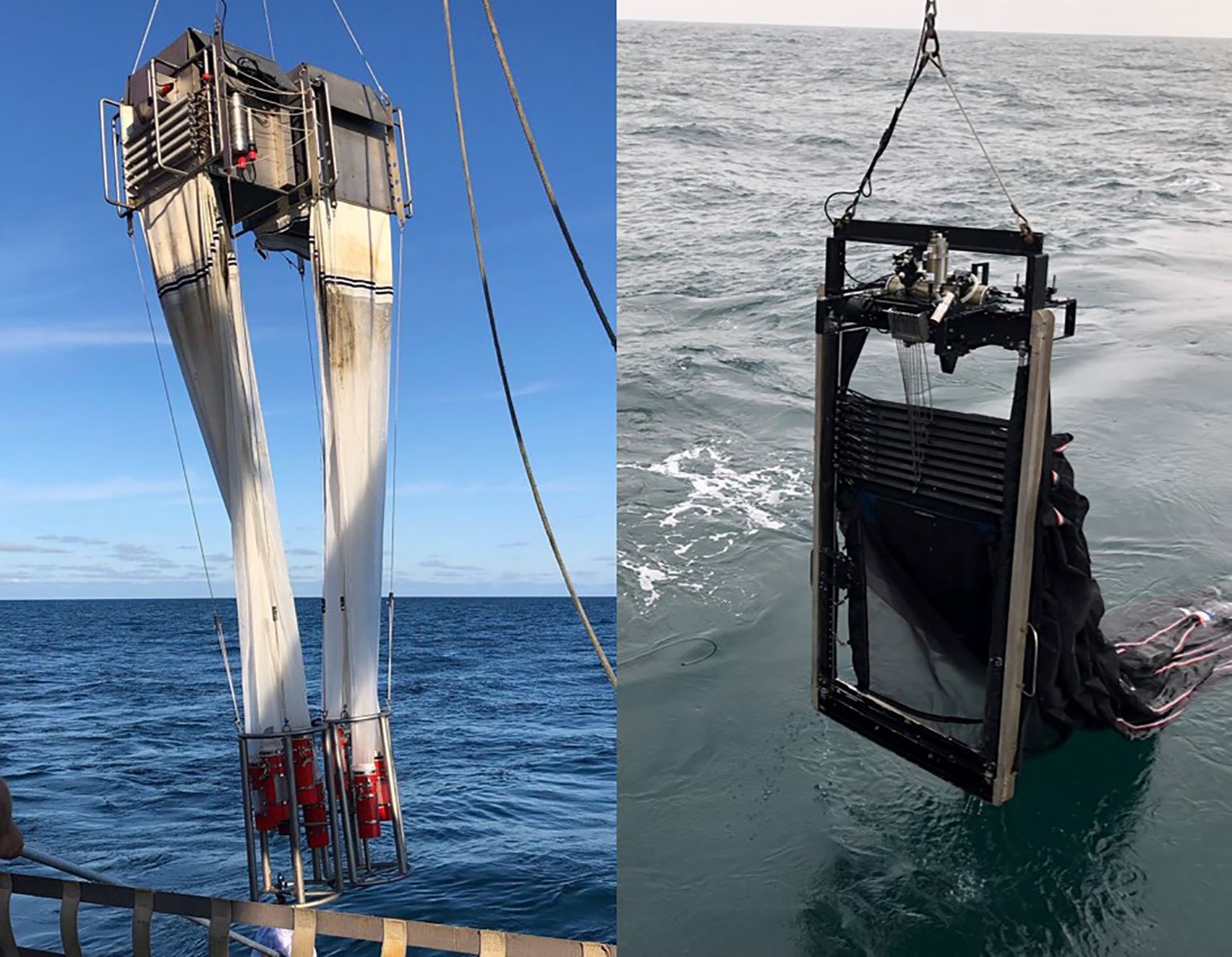

Figure 3. The tandem MultiNet (150 µm; left) and MOCNESS (505 µm; right) plankton net systems deployed from the R/V Sikuliaq during the 2019 Gulf of Alaska Seamounts expedition.
A 1 m2 Multiple Opening/Closing Net and Environmental Sensing System (MOCNESS; Wiebe et al., 1985) was deployed following the tandem MultiNet (Figure 3). The MOCNESS was equipped with 505-µm mesh nets and sampled the following depth bins: 0 –100 – 200 – 300 – 400 – 600 – 1200 – 2000 – 3000 m (Table 1). Since the MOCNESS has a larger mouth diameter than the MultiNet (1 m2 vs 0.25 m2) and thus filters a larger volume of water, samples were split twice. One half of the net sample was preserved in 10% buffered formalin-seawater for taxonomic analysis whereas the second half was split again; one quarter was preserved in 95% ethanol and the other quarter was used for live sorting. All bulk ethanol plankton net samples were stored at -20 °C and complete ethanol changes were performed ~24 hours after initial preservation.
In addition to traditional plankton net sampling, the Global Explorer ROV (Oceaneering, Houston, TX) conducted pelagic dives down to ~2550 m (Table 1). ROV operations focused on organisms larger than 1 cm and of the gelatinous zooplankton group (e.g., ctenophores, medusae, larvaceans; Bailey et al., 1995). Specimens were collected with a rotary carousel suction sampler (Youngbluth, 1984b) or static “D-samplers” (Youngbluth, 1984a). At all stations, ROV dives transpired during daytime operations (0800 – 2000) while plankton net deployments were conducted during nighttime operations (2000 – 0800).
All mesozooplankton were identified based on current taxonomic literature (e.g., Boxshall and Halsey, 2004). Organisms were identified at sea from live collections from both the plankton net samples and the ROV collections described above. Once on board, organisms from the ROV and plankton net tows were immediately transferred to a cold room maintained at 4 °C until sorting and identifications could be performed. Zooplankton were identified to the lowest taxonomic level feasible at sea using Leica MZ16, M165C, or M205C stereomicroscopes, catalogued into cryogenic vials, and preserved in 95% pre-chilled ethanol for DNA sequencing. For each catalogued specimen, a matching formalin voucher was created for taxonomic integrity. In instances when large, rare organisms from gelatinous zooplankton groups could not have a taxonomic voucher, tissue samples were cut off and preserved in ethanol while the remainder of the organism was preserved in formalin. For ROV samples, video imaging and photographs were captured in situ with an Ultra-high definition (UHD) 4K and stereo high definition (HD) camera systems prior to collection. The video clips (Apple ProRes 4:2:0 codec,.mov files with embedded timecode) serve as further voucher "material." Organisms from net collections were photographed using a 12MPix Spot Insight camera (uncompressed JPEG) mounted to a Leica stereomicroscope.
Organisms that could not be identified to the species-level at sea were more closely examined back at the University of Alaska Fairbanks using both regional and phylum- or family-specific taxonomic keys and original descriptions. Due to morphological complexity, incomplete taxonomic keys, damaged specimens, and the lack of descriptive studies on zooplankton from the deep ocean, some specimens were unable to be identified to species. For these instances we used Open Nomenclature (ON) qualifiers defined in Horton et al. (2021) and Sigovini et al. (2016). To the best of our knowledge, there is no qualifier for specimens assigned a taxonomic identification based on molecular data. Thus, we introduce the new ON qualifier deoxyribonucleic acid, abbreviation DNA (e.g., DNA Genus, Genus DNA species) for specimens assigned a genus- or species-level identification based on clustering (100% bootstrap support) of DNA barcode genes using phylogenetic trees against preexisting barcodes derived from expertly identified specimens, where at least one of the sequences in that clade could be referred to a physical specimen (or photographs) where identification could be corroborated through morphological analyses.
DNA barcoding was performed at the Smithsonian’s Laboratory of Analytical Biology (L.A.B) at the National Museum of Natural History (Washington D.C.) from the ethanol-preserved tissue samples catalogued at sea. Genomic DNA (gDNA) was extracted using a phenol-chloroform method on an AutoGenprep 965 Automated DNA Isolation System (AutoGen Inc., Holliston, MS, USA) following manufacturers’ protocols. For small taxa (e.g., copepods and larvaceans) DNA was extracted from the entire organism while, for larger organisms (e.g., Cnidarians, ctenophore, chaetognaths, and larger crustaceans), non-diagnostic features such as legs or antennae were removed for extraction where possible.
Extracted gDNA was quantified using a SpectraMax iD3 Multi-Mode Microplate reader (Molecular Devices). Mitochondrial COI and 16S rRNA and nuclear 18S rRNA were amplified using previously published primers (COI: Folmer et al., 1994; Geller et al., 2013; 16S: Cunningham and Buss, 1993; 18S rRNA: Hillis and Dixon, 1991; Borchiellini et al., 2001). All PCR reactions were carried out in 10 µL reaction volumes containing 1 µL DNA template, 3.2 µL PCR grade H2O, 5.0 µL Promega GoTaq HotStart 2X Master Mix, 0.3 µL each forward and reverse primer, 0.1 µL BSA, 0.1 µL MgCl2. PCR protocols for each amplicon were as follows: COI: initial denaturation of 95 °C 7 min; 4 cycles of 94 °C 30 sec, 50 °C 45 sec, 72 °C 1 min followed by 34 cycles of 94 °C 30 sec, 45 °C 45 sec, 72 °C 1 min; final extension of 72 °C 8 min; 16S: initial denaturation of 95 °C 2 min; 35 cycles of 95 °C 30 sec, 50 °C 30 sec, 72 °C 1 min 30 sec; final extension of 72 °C 5 min; and 18S rRNA: initial denaturation of 95 °C 2 min; 35 cycles of 95 °C 30 sec, 52 °C 30 sec, 72 °C 1 min 30 sec; final extension of 72 °C 5 min. PCR products were visualized on a 1% agarose gel. Successful PCR products were purified with ExoSAP-IT Express (37 °C 4 min and 80 °C 1 min). Cycle sequencing was carried out on purified PCR product using the Big Dye Terminator v3.1 kit (Applied Biosystems) with each primer used for PCR amplification. Due to the amplicon length of 18S, the internal primer 18SCnew (5’-CAG CCG CGG TAA TTC CAG C-3’; Miranda et al., 2016) was sequenced to bridge the gap from the 3’ end of 18SE and the 5’ end of 18SL. Cycle sequencing product was cleaned using Sephadex G-50 powder and run on an ABI PRISM 3100 Genetic Analyzer.
Data management for DNA barcodes follows the Smithsonian Institution Barcode Network guidelines (Redmond et al., 2023) that abides by the Barcode Data Standard established by the Consortium of the Barcode of Life (CBOL; Hanner, 2012). Specimen, voucher, and sequence metadata are maintained in the SI Field Information Managements System Genomic Observatories MetaDatabase (FIMS-GEOME; https://geome0db.org/) and the Laboratory Information Managements System (LIMS) databases.
For each genetic marker, sequences were assembled, trimmed, manually checked for ambiguous base calling and stop codons, and aligned using MAFFT (Katoh et al., 2002) in Geneious Prime 2023.2.1 (https://www.geneious.com). Sequence names were annotated and assigned a species name using the Biocode plugin that links sequences in Geneious Prime to the SI FIMS and LIMS databases. Sequences were run against the NCBI GenBank database using BLAST (Altschul et al., 1997) to check for accuracy of species identifications and potential contamination. Due to the lack of barcode data for species from the North Pacific deep-sea ecosystem, this QA/QC step was only informative for species with existing reads in GenBank. Therefore, neighbor-joining (NJ) trees were constructed for each genetic marker to determine clustering for all groups of zooplankton. Once identifications were checked and confirmed, sequences were submitted to the National Institute of Health National Center for Biotechnology Information (NIH NCBI) Sequence Read Archive (SRA). 18S and non-cnidarian COI reads were submitted under BioProject PRJNA971221 whereas COI and 16S reads for cnidarians were submitted under BioProject PRJNA1077907.
From this work, we generated 1462 new DNA sequences from 253 unique taxa from the Gulf of Alaska (Table 2 and Supplementary Table S1). DNA sequences were generated for COI from 781 specimens belonging to 73 families (GenBank Accession Nos. OR985931 – OR986516 & PP409229 – PP409419), 16S for 33 specimens belonging to 18 families (GenBank Accession Nos. PP356923 – PP356955), and 18S from 648 specimens belonging to 42 families (GenBank Accession Nos. PP032093 – PP032748). Sequences for all three genes encompassed organisms from 7 phyla including Arthropoda (Orders Copepoda, Amphipoda, Decapoda, Euphausiacea, Lophogastrida, and Isopoda), Cnidaria (Hydrozoa and Scyphozoa), Ctenophora (Tentaculata), Chaetognatha, Mollusca, Chordata (Tunicata), and Annelida.
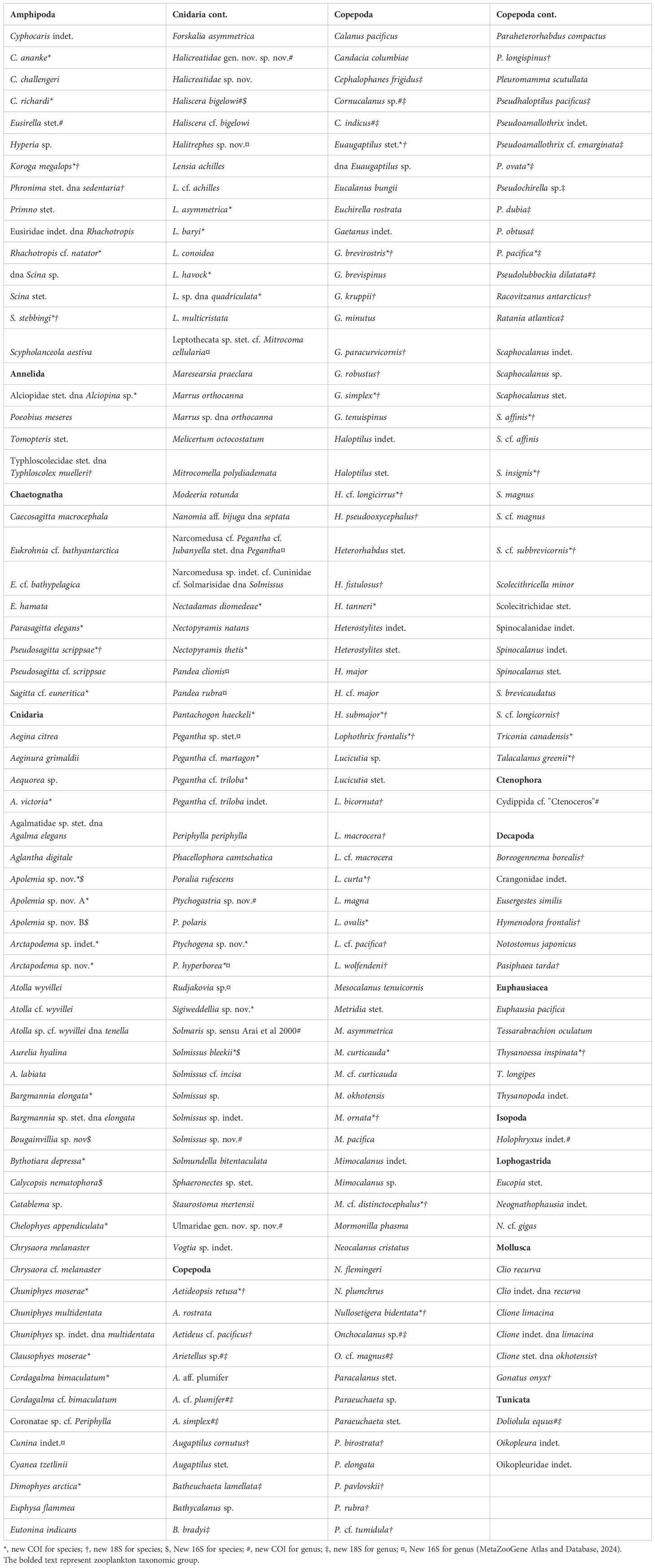
Table 2. Specimen list of zooplankton identified and sequenced for the mitochondrial COI and 16S and nuclear 18S rRNA genes from the Gulf of Alaska.
Phylogenetic relationships were analyzed for all three genes through Neighbor-Joining trees (Supplementary Figures S1 – Supplementary Figures S3). Mitochondrial COI reads clustered into major clades with >66% bootstrap support with 76% support for the order Copepoda, 50 – 100% for the phylum Cnidaria, 72% for the phylum Chaetognatha, 70 – 83% for the order Amphipoda, 66% for the order Euphausiacea, and 96 – 100% for the order Decapoda. Monophyletic relationships were resolved at 100% bootstrap support for sequences from the same species (Supplementary Figure S1). Nuclear 18S rRNA reads clustered into major clades with >89% bootstrap support (Supplementary Figure S2) with subclades forming at 97 – 100% support for the order Doliolida, 100% support for family Oikopleuridae, 70 – 100% support for the phylum Chaetognatha, 50 – 100% support for the order Pteropoda, 61 – 100% for the phylum Annelida, 60 – 100% support for the order Euphausiacea, 65 – 100% for the order Decapoda, and 51 – 100% for the order Lophogastrida. The order Amphipoda clustered – at 100% bootstrap support – into three distinct subclades with 18S reads from Koroga megalops (family Uristidae) forming the first clade, reads from Scina sp. (family Scinidae) forming the second clade, and reads from Primno sp. (family Phrosinidae) and Phronima sp. (Phronimidae) forming the third clade. All reads from the order Copepoda clustered at 100% bootstrap support with subclades clustering between 51 – 100%. (Supplementary Figure S2). Clustering patterns from COI barcodes were used in combination with morphological characteristics to help resolve taxonomic identifications for any unresolved COI and 18S reads, where possible. For Cnidarians, mitochondrial 16S reads clustered into major clades at >54% bootstrap support (Supplementary Figure S3). Reads from scyphozoans clustered at 100% bootstrap support while hydrozoans were split into two clades, represented by organisms from the subclass Hydroidolina (orders Siphonophorae and Anthoathecata) with 64% bootstrap support), then the subclass Trachylinae (orders Trachymedusae and Narcomedusae) with 96% bootstrap support.
From the zooplankton sequenced in this work, 107 species, were previously lacking barcode data from the North Pacific or the global ocean: 57 species for COI, 6 species for 16S, and 44 species for 18S (Table 2). Of the 107 species barcoded, 12 represent new species of cnidarians that are currently undergoing description. Additionally, 47 contributions were made across all markers where no sequence data were previously available for the genus level. The contributions included 11 genera of hydrozoan cnidarians: Halitrephes and Ptychogastria (Trachymedusae), Pandea (Anthoathecata), Cunina, Pegantha, Solmaris, and Solmissus (Narcomedusae), Mitrocoma and Ptychogena (Leptothecata), and Rudjakovia and Vogtia (Siphonophorae); 10 genera of copepoda: Arietellus (Arietellidae), Batheuchaeta and Pseudochirella (Aetideidae), Cephalophanes, Cornucalanus and Onchocalanus (Phaennidae), Pseudhaloptilus (Augaptilidae), Pseudoamallothrix (Scolecitrichidae), Pseudolubbockia (Lubbockiidae), and Ratania (Rataniidae); and one morphotaxon of ctenophores (Cydippida aka. "Ctenoceros"), pelagic tunicates (Doliolula), and isopods (Holophryxus).
Globally, the deep-ocean pelagic environment is severely under-sampled (Robison, 2008; Levin et al., 2019), restricting our ability to confidently measure and characterize zooplankton biodiversity below the mesopelagic zone. Current knowledge on species inventories and subsequent biodiversity trends are biased towards organisms that survive net collections relatively undamaged such as copepods (Miller and Clemons, 1984, Yamaguchi et al., 2002 & Yamaguchi et al., 2015; Homma and Yamaguchi, 2010), shrimp (Kikuchi and Omori, 1985), chaetognaths (Ozawa et al., 2007; Terazaki and Miller, 1986), and decapods (Pakhomov et al., 2019). Incorporating alternatives to plankton net sampling methodologies such as ROVs equipped with advanced collection and imaging systems used in this study, has furthered our ever-evolving knowledge of deep pelagic biodiversity. Over the past few decades, imaging-based observations have revealed gelatinous zooplankton groups such as cnidarians, ctenophores, and larvaceans, organisms with body forms adapted to withstand immense pressures at depth, comprise a substantial portion of the pelagic diversity of the deep ocean (Robison, 2008). Of the 91 cnidarian species identified in this study, of which 12 are undescribed, roughly 30% were collected at depths below 2000 m, reflecting this increase in diversity and rarity among gelatinous taxa compared to the epi- and mesopelagic zones.
The lack of systematic approaches that incorporate morphological and molecular techniques for deep-sea studies of zooplankton has created a scarcity of zooplankton collections destined for DNA barcoding work. The process of DNA barcoding requires the skills and knowledge of expert taxonomists to correctly identify specimens to the species level so that a molecular ecologist can then sequence DNA barcode genes from them (Hebert et al., 2003). The population of DNA barcodes from life on Earth into publicly available databases is giving non-expert taxonomists the ability to accurately and reliably identify an unknown organism to the species level. The work accomplished herein provided many challenges. Not only are zooplankton communities inherently complex taxonomically, but the deep-sea environment itself is challenging to sample, is very time consuming, and requires ships capable of deploying gear to thousands of meters. These challenges have contributed to the general lack of zooplankton studies in deep waters and, as a result, has created a lack of expert taxonomists and the necessary taxonomic keys needed for identifying specimens from the bathy- and abyssopelagic zones.
Despite the many challenges that come with working in the deep-sea, we were able to collect zooplankton for combined morphological identifications and DNA barcoding. From this work, we have contributed 781 COI and 33 16S barcodes and 648 18S rRNA reads for a total of 1,462 DNA sequences for zooplankton collected from the deep-water environment of the Gulf of Alaska. The DNA sequences generated in this study represent 254 unique taxa spanning 4 phyla, of which 107 species were lacking any publicly available DNA sequences prior to this work. The species list presented in Table 2 is not an exhaustive list of the >1,000 organisms collected from the region. Sequencing for the COI, 16S, and 18S genes was not successful for all specimens, or produced reads that were contaminated or of low quality. Additionally, high-quality sequences analyzed from organisms that could not be confidently identified past Family through morphological or molecular means were not submitted to GenBank. The inability for the GenBank BLAST search to further refine and aid in the genus- or species-level identification of these organisms is a direct outcome for the need to continue DNA barcoding of pelagic organisms. These results also highlight the overall importance our contributions will make in aiding future zooplankton identifications through molecular approaches.
To date, few groups of zooplankton are >50% complete for COI, 16S, or 18S reads for known species with distribution records in the North Pacific (MetaZooGene Atlas and Database, 2024; Table 3). Not surprisingly, decapods and euphausiids, the more robust yet species-poor groups of crustaceans, have COI reads for 75% and 89%, respectively, of the documented species in the region. However, copepods, the second most species-rich planktonic crustacean in the North Pacific, are only 29% complete for COI and 13% complete for 18S. Thus, continued DNA barcoding remains a high priority on both local and global scales, with further collection efforts needed from depths below 1000 m where the potential for new species discoveries is promising (Wiebe et al., 2010). Increased sampling efforts and collections destined for DNA barcoding will create more complete reference DNA sequence databases that are currently hindering taxonomic assignments of reads produced from high-throughput sequencing for environmental sequencing studies (see Bucklin et al., 2021). More complete reference DNA sequence databases will not only facilitate improved characterization of pelagic communities (see Questel et al., 2021) but will also support biodiversity, phylogeographic, and population genetic studies, as well as aid in the detection of cryptic species or morphotypes (Blanco-Bercial et al.., 2011; Blanco-Bercial et al., 2014; Questel et al., 2016; Cornils et al., 2017; Miyamoto et al., 2012; Hirai et al., 2017; Miglietta and Pruski, 2023; Moura et al., 2023, and many others).
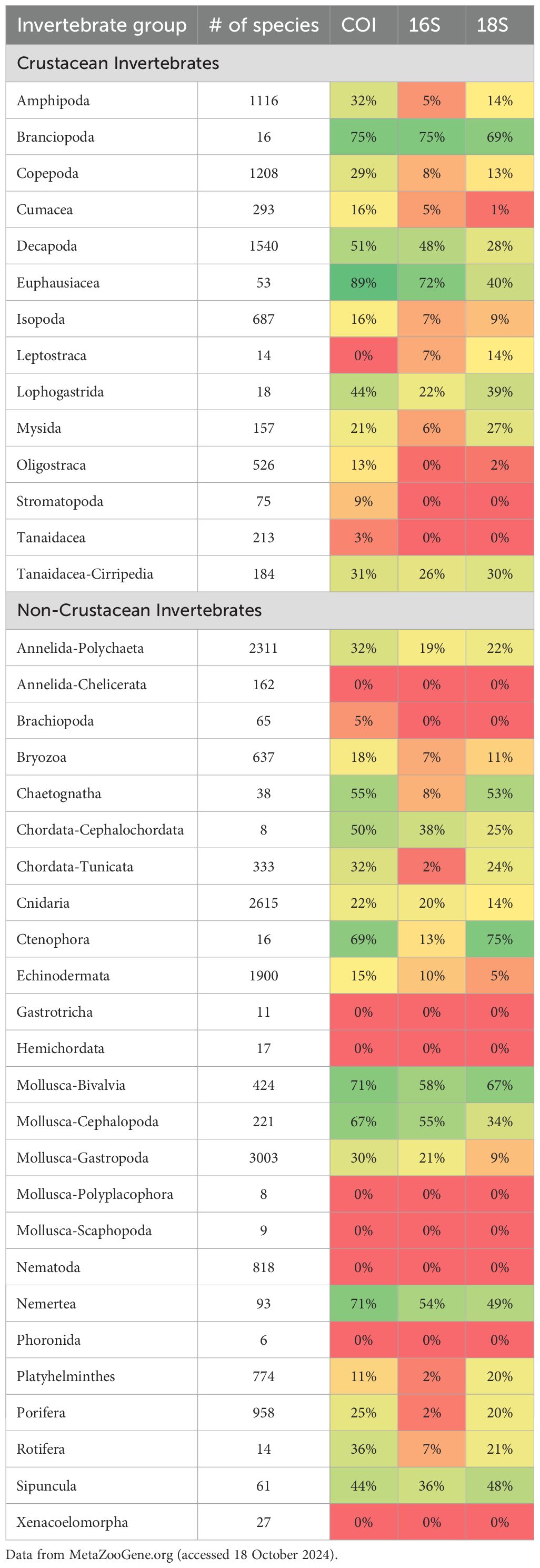
Table 3. Heatmap showing progress of DNA barcoding for invertebrate functional groups from the North Pacific Ocean. Color shading reflects completion of DNA barcoding: red = 0% and dark green = 100%.
The DNA sequences generated for this study have been deposited to the NCBI NIH Sequence Read Archive (SRA; https://www.ncbi.nlm.nih.gov/sra/) under BioProjects PRJNA971221 and PRJNA1077907. DNA sequences can be found under GenBank Accession numbers OR985931 – OR986516 & PP409229 – PP409419 (COI), PP356923 – PP356955 (16S), and PP032093 – PP032748 (18S). All metadata are stored and maintained in the Smithsonian Institution Barcode Network’s Field Information Management System – Genomic Observatories MetaData (FIMS – GEOME) and Laboratory Information Management System (LIMS) databases under BOEM project NOAA Gulf of Alaska Seamounts 2019.
The manuscript presents research on animals that do not require ethical approval for their study.
JQ: Conceptualization, Data curation, Formal analysis, Methodology, Validation, Writing – original draft, Writing – review & editing, Funding acquisition. CS: Data curation, Formal analysis, Validation, Writing – original draft, Writing – review & editing, Methodology. AC: Data curation, Formal analysis, Resources, Writing – original draft, Writing – review & editing, Funding acquisition, Investigation, Methodology, Validation. DL: Data curation, Formal analysis, Investigation, Methodology, Validation, Writing – original draft, Writing – review & editing. RH: Conceptualization, Funding acquisition, Investigation, Project administration, Writing – original draft, Writing – review & editing, Methodology.
The author(s) declare that financial support was received for the research and/or publication of this article. Support for DNA sequencing efforts reported in this publication were provided by the North Pacific Research Board (Project Number 2102). Support for field activities including sample collection and ship time on the NSF UNOLS R/V Sikuliaq was supported by National Oceanic and Atmospheric Administration (NOAA) Ocean Exploration and Research (Project Number NA15OAR0110209) and NSF NGA-LTER (OCE-1656070). Research reported in this publication was supported by an Institutional Development Award (IDeA) from the National Institute of General Medical Sciences of the National Institutes of Health (grant number P20GM103395).
All of the laboratory and computer work were conducted in and with the support of the L.A.B. facilities of the National Museum of Natural History or its partner labs. We thank the captains, crew, and marine technicians of the R/V Sikuliaq, the engineers and ROV pilots from Oceaneering, and the science personnel who helped with field activities. We graciously thank Stephanie Bush and Jeffrey Hunt at SI NMNH for their assistance with the molecular laboratory activities.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
The content is solely the responsibility of the authors and does not necessarily reflect the official views of the NIH.
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fmars.2025.1515048/full#supplementary-material
Supplementary Figure 1 | Neighbor-joining tree of 785 COI sequences. Branch labels indicate bootstrap values after 1,000 replications. The Eumetopias jubatus COI sequence (Accession No. KP992987) was included as an outgroup for rooting the tree.
Supplementary Figure 2 | Neighbor-joining tree of 33 16S sequences. Branch labels indicate bootstrap values after 1,000 replications. The Calanus glacialis 16S sequence (Accession No. FJ628357) was included as an outgroup for rooting the tree.
Supplementary Figure 3 | Neighbor-joining tree of 656 18S rRNA sequences. Branch labels indicate bootstrap values after 1,000 replications. The Eumetopias jubatus 18S sequence (Accession No. XR_003608062) was included as an outgroup for rooting the tree.
Supplementary Table 1 | List of identified specimens and the corresponding accession numbers for the COI, 16S, and 18S sequences submitted to GenBank. Taxonomic identifications were based on morphological and molecular methods. GenBank Source Organism represents the identification assigned to the specimen at the time of submission.
Altschul S. F., Madden T. L., Schäffer A. A., Zhang J., Zhang Z., Miller W., et al. (1997). Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402. doi: 10.1093/nar/25.17.3389
Baek S. Y., Jang K. H., Choi E. H., Ryu S. H., Kim S. K., Lee J. H., et al. (2016). DNA barcoding of metazoan zooplankton copepods from South Korea. PloS One 11, 1–20. doi: 10.1371/journal.pone.0157307
Bailey T. G., Youngbluth M. J., Owen G. P. (1995). Chemical composition and metabolic rates of gelatinous zooplankton from midwater and benthic boundary layer environments off Cape Hatteras, North Carolina, USA. Mar. Ecol. Prog. Ser. 122, 121–134. doi: 10.3354/meps122121
Blanco-Bercial L., Álvarez-Marqués F., Bucklin A. (2011). Comparative phylogeography and connectivity of sibling species of the marine copepod Clausocalanus (Calanoida). J Exp Mar Biol Ecol 404, 108–115. doi: 10.1016/j.jembe.2011.05.011
Blanco-Bercial L., Cornils A., Copley N., Bucklin A. (2014). DNA Barcoding of Marine Copepods: Assessment of Analytical Approaches to Species Identification. PLoS Curr. doi: 10.1371/currents.tol.cdf8b74881f87e3b01d56b43791626d2
Blanco-Bercial L. (2020). Metabarcoding analyses and seasonality of the zooplankton community at BATS. Front. Mar. Sci. 7, 1–16. doi: 10.3389/fmars.2020.00173
Borchiellini C., Manuel M., Alivon E., Boury-Esnault N., Vacelet J., Le Parco Y. (2001). Sponge paraphyly and the origin of Metazoa. J. Evol. Biol. 14, 171–179. doi: 10.1046/j.1420-9101.2001.00244.x
Boxshall G. A., Halsey S. H. (2004). An introduction to copepod diversity (London, United Kingdom: The Ray Society).
Bucklin A., Hopcroft R. R., Kosobokova K. N., Nigro L. M., Ortman B. D., Jennings R. M., et al. (2010a). DNA barcoding of Arctic Ocean holozooplankton for species identification and recognition. Deep Sea Res. 2 Top. Stud. Oceanogr 57, 40–48. doi: 10.1016/j.dsr2.2009.08.005
Bucklin A., Nishida S., Schnack-Schiel S., Wiebe P. H., Lindsay D., Machida R. J., et al. (2010b). “A census of zooplankton of the global ocean,” in Life in the world’s oceans: diversity, distribution, and abundance. Ed. McIntyre A. D. (United Kingdom: Blackwell Publishing), 361.
Bucklin A., Ortman B. D., Jennings R. M., Nigro L. M., Sweetman C. J., Copley N. J., et al. (2010c). A “Rosetta Stone“ for metazoan zooplankton: DNA barcode analysis of species diversity of the Sargasso Sea (Northwest Atlantic Ocean). Deep Sea Res. 2 Top. Stud. Oceanogr 57, 2234–2247. doi: 10.1016/j.dsr2.2010.09.025
Bucklin A., Peijnenburg K. T. C. A., Kosobokova K. N., O’Brien T. D., Blanco-B L., Cornils A., et al. (2021). Toward a global reference database of COI barcodes for marine zooplankton. Mar. Biol. 168, 1–26. doi: 10.1007/s00227-021-03887-y
Bucklin A., Steinke D., Blanco-Bercial L. (2011). DNA barcoding of marine metazoa. Ann. Rev. Mar. Sci. 3, 471–508. doi: 10.1146/annurev-marine-120308-080950
Bucklin A., Yeh H. D., Questel J. M., Richardson D. E., Reese B., Copley N. J., et al. (2019). Time-series metabarcoding analysis of zooplankton diversity of the NW Atlantic continental shelf. Zool J. Linn Soc. 76, 1162–1176. doi: 10.1093/zoolinnean/zly093
Casey J. M., Ransome E., Collins A. G., Mahardini A., Kurniasih E. M., Sembiring A., et al. (2021). DNA metabarcoding marker choice skews perception of marine eukaryotic biodiversity. Environ. DNA 00, 3, edn3.245. doi: 10.1002/EDN3.245
Chaytor J. D., Keller R. A., Duncan R. A., Dziak R. P. (2007). Seamount morphology in the Bowie and Cobb hot spot trails, Gulf of Alaska. Geochemistry Geophysics Geosystems 8, Q09016. doi: 10.1029/2007GC001712
Cheng F., Wang M., Sun S., Li C., Zhang Y. (2013). DNA barcoding of Antarctic marine zooplankton for species identification and recognition. Adv. Polar Sci. 24, 119–127. doi: 10.3724/sp.j.1085.2013.00119
Christianson L. M., Johnson S. B., Schultz D. T., Haddock S. H. D. (2022). Hidden diversity of Ctenophora revealed by new mitochondrial COI primers and sequences. Mol. Ecol. Resour. 22, 283–294. doi: 10.1111/1755-0998.13459
Cornils A., Wend-Heckmann B., Held C. (2017). Global phylogeography of Oithona similis s.l. (Crustacea, Copepoda, Oithonidae) – A cosmopolitan plankton species or a complex of cryptic lineages? Mol. Phylogenet Evol. 107, 473–485. doi: 10.1016/j.ympev.2016.12.019
Cunningham C. W., Buss L. W. (1993). Molecular evidence for multiple episodes of paedomorphosis in the family Hydractiniidae. Biochem. Systematics Ecol. 21, 57–69. doi: 10.1016/0305-1978(93)90009-G
Folmer O., Black M., Hoeh W., Lutz R., Vrijenhoek R. (1994). DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol. Mar. Biol. Biotechnol. 3, 294–299. doi: 10.1371/journal.pone.0013102
Geller J., Meyer C., Parker M., Hawk H. (2013). Redesign of PCR primers for mitochondrial cytochrome c oxidase subunit I for marine invertebrates and application in all-taxa biotic surveys. Mol. Ecol. Resour 13, 851–861. doi: 10.1111/1755-0998.12138
Hanner R., Consortium for the Barcode of Life (2012). Data Standards for BARCODE Records in INSDC (BRIs). v. 2.4 ed. Database Working Group, Consortium for the Barcode of Life. doi: 10.5479/10088/96518
Hebert P. D. N., Cywinska A., Ball S. L., deWaard J. R. (2003). Biological identifications through DNA barcodes. Biol. identifications through DNA barcodes. Pro Biol. Sci. R. Soc. 270, 313–321. doi: 10.1098/rspb.2002.2218
Hellberg M. E. (2006). No variation and low synonymous substitution rates in coral mtDNA despite high nuclear variation. BMC Evol. Biol. 6, 1–8. doi: 10.1186/1471-2148-6-24
Hillis D. M., Dixon M. T. (1991). Ribosomal DNA: molecular evolution and phylogenetic inference. Q Rev. Biol. 66, 411–453. doi: 10.2307/2831326
Hirai J., Katakura S., Kasai H., Nagai S. (2017). Cryptic zooplankton diversity revealed by a metagenetic approach to monitoring metazoan communities in the coastal waters of the Okhotsk Sea, Northeastern Hokkaido. Front. Mar. Sci. 4, 379. doi: 10.3389/fmars.2017.00379
Holland B. S., Dawson M. N., Crow G. L., Hofmann D. K. (2004). Global phylogeography of Cassiopea (Scyphozoa: Rhizostomeae): Molecular evidence for cryptic species and multiple invasions of the Hawaiian Islands. Mar. Biol. 145, 1119–1128. doi: 10.1007/s00227-004-1409-4
Homma T., Yamaguchi A. (2010). Vertical changes in abundance, biomass and community structure of copepods down to 3000m in the southern Bering Sea. Deep Sea Res. 1 Oceanogr Res. Pap 57, 965–977. doi: 10.1016/j.dsr.2010.05.002
Horton T., Marsh L., Bett B. J., Gates A. R., Jones D. O. B., Benoist N. M. A., et al. (2021). ). Recommendations for the standardisation of open taxonomic nomenclature for image-based identifications. Front. Mar. Sci. 8. doi: 10.3389/fmars.2021.620702
Katoh K., Misawa K., Kuma K.-I., Miyata T. (2002). MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 30, 3059–3066. doi: 10.1093/nar/gkf436
Kikuchi T., Omori M. (1985). Vertical distribution and migration of oceanic shrimps at two locations off the Pacific coast of Japan. Deep-Sea Res. A 32, 837–351. doi: 10.1016/0198-0149(85)90119-0
Lee C. E. (2000). Global phylogeography of a cryptic copepod species complex and reproductive isolation between genetically proximate “populations. Evolution 54, 2014–2027. doi: 10.1111/j.0014-3820.2000.tb01245.x
Levin L. A., Bett B. J., Gates A. R., Heimbach P., Howe B. M., Janssen F., et al. (2019). Global observing needs in the deep ocean. Front. Mar. Sci. 6. doi: 10.3389/fmars.2019.00241
Lindsay D. J., Grossmann M. M., Nishikawa J., Bentlage B., Collins A. G. (2015). DNA barcoding of pelagic cnidarians: current status and future prospects. Bull. Plankton Soc. Jap 62, 39–43. doi: 10.24763/bpsj.62.1_39
Mackas D. L., Goldblatt R., Lewis A. G. (1998). Interdecadal variation in developmental timing of Neocalanus plumchrus populations at Ocean Station P in the subarctic North Pacific. Can. J. Fish Aquat Sci. 55, 1878–1893. doi: 10.1139/f98-080
Martin J. H., Gordon R. M., Fitzwater S., Broenkow W. W. (1989). VERTEX: phytoplankton/iron studies in the Gulf of Alaska. Deep-Sea Res. 36, 649–680. doi: 10.1016/0198-0149(89)90144-1
Masunaga A., Mansfield M. J., Tan Y., Liu A. W., Bliznina A., Barzaghi P., et al. (2022). The cosmopolitan appendicularian Oikopleura dioica reveals hidden genetic diversity around the globe. Mar. Biol. 169, 1–17. doi: 10.1007/s00227-022-04145-5
Matthews S. A., Blanco-Bercial L. (2023). Divergent patterns of zooplankton connectivity in the epipelagic and mesopelagic zones of the eastern North Pacific. Ecol. Evol. 13, e10664. doi: 10.1002/ece3.10664
Menard H. W., Dietz R. S. (1951). Submarine geology of the Gulf of Alaska. Bull. Geol Soc. Amer 62, 1263–1285. Available at: http://pubs.geoscienceworld.org/gsa/gsabulletin/article-pdf/62/10/1263/3416444/i0016-7606-62-10-1263.pdf (Accessed October 18, 2024).
MetaZooGene (2024). Accessed through MetaZooGene. Available online at: https://www.st.nmfs.noaa.gov/copepod/collaboration/metazoogene/atlas/index-o00.html (Accessed October 18, 2024).
Meyer C. P., Paulay G. (2005). DNA barcoding: Error rates based on comprehensive sampling. PloS Biol. 3, 1–10. doi: 10.1016/0079-6611(88)90044-4
Miller C. B., Frost B. W., Batchelder H. P., Clemons M. J., Conway R. E. (1984). Life Histories of Large, Grazing Copepods in a Subarctic Ocean Gyre: Neocalanus plumchrus, Neocalanus cristatus, and Eucalanus bungii in the Northeast Pacific. Prog. Oceanogr 13, 201–243. doi: 10.1016/0079-6611(84)90009-0
Miglietta M. P., Pruski S. (2023). Cryptic species in time and space: an assessment of cryptic diversity within eight nominal species of Hydrozoa (Cnidaria). Proc. R. Soc. B Biol. Sci. 290, 20230851. doi: 10.1098/rspb.2023.0851
Miranda L. S., Hirano Y. M., Mills C. E., Falconer A., Fenwick D., Marques A. C., et al. (2016). Systematics of stalked jellyfishes (Cnidaria: Staurozoa). PeerJ 2016, 1–48. doi: 10.7717/peerj.1951
Miyamoto H., Machida R. J., Nishida S. (2012). Global phylogeography of the deep-sea pelagic chaetognath Eukrohnia hamata. Prog. Oceanogr 104, 99–109. doi: 10.1016/j.pocean.2012.06.003
Mora C., Tittensor D. P., Adl S., Simpson A. G. B., Worm B. (2011). How many species are there on earth and in the ocean? PloS Biol. 9, e1001130. doi: 10.1371/journal.pbio.1001127
Moura C. J., Magalhães B. I., Gonçalves J. M. (2023). DNA barcoding of moon jellyfish (Cnidaria, Scyphozoa, Ulmaridae, Aurelia): Two cryptic species from the Azores (NE Atlantic, Macaronesia), and evaluation of the non-indigenous species (NIS). Diversity 15, 323. doi: 10.3390/d15030323
Ozawa M., Yamaguchi A., Ikeda T., Watanabe Y., Ishizaka J. (2007). Abundance and community structure of chaetognaths from the epipelagic through abyssopelagic zones in the western North Pacific and its adjacent seas. Plankt Benthos Res. 2, 184–197. doi: 10.3800/pbr.2.184
Pakhomov E. A., Podeswa Y., Hunt B. P., Kwong L. E. (2019). Vertical distribution and active carbon transport by pelagic decapods in the North Pacific Subtropical Gyre. ICES K Mar. Sci. 76, 702–717. doi: 10.1093/icesjms/fsy134
Questel J. M., Blanco-Bercial L., Hopcroft R. R., Bucklin A. (2016). Phylogeography and connectivity of the Pseudocalanus (Copepoda: Calanoida) species complex in the eastern North Pacific and the Pacific Arctic Region. J. Plankton Res. 38, 610–623. doi: 10.1093/plankt/fbw025
Questel J. M., Hopcroft R. R., Dehart H. M., Smoot C. A., Kosobokova K. N., Bucklin A. (2021). Metabarcoding of zooplankton diversity within the Chukchi Borderland, Arctic Ocean: improved resolution from multi-gene markers and region-specific DNA databases. Mar. Biodiver 51, 1–19. doi: 10.1007/s12526-020-01136-x
Redmond N., Steier J., Trizna M., O’Mahoney M., Webb Wilcox A. (2023). SI barcode network informatics documentation release 1.0 (Washington DC: Smithsonian Institution).
Robison B. H. (2008). Conservation of deep pelagic biodiversity. Conserv. Biol. 23, 847–858. doi: 10.1111/j.1523-1739.2009.01219.x
Rogers A. D., Appiah-Madson H., Ardron J. A., Bax N. J., Bhadury P., Brandt A., et al. (2023). Accelerating ocean species discovery and laying the foundations for the future of marine biodiversity research and monitoring. Front. Mar. Sci. 10. doi: 10.3389/fmars.2023.1224471
Shimode S., Toda T., Kikuchi T. (2006). Spatio-temporal changes in diversity and community structure of planktonic copepods in Sagami Bay. Japan. Mar. Biol. 148, 581–597. doi: 10.1007/s00227-005-0093-3
Sigovini M., Keppel E., Tagliapietra D. (2016). Open Nomenclature in the biodiversity era. Methods Ecol. Evol. 7, 1217–1225. doi: 10.1111/2041-210X.12594
Stabeno P. J., Kachel N. B., Moore S. E., Napp J. M., Sigler M., Yamaguchi A., et al. (2012). Comparison of warm and cold years on the southeastern Bering Sea shelf and some implications for the ecosystem. Deep Sea Res. 2 Top. Stud. Oceanogr 65–70, 31–45. doi: 10.1016/j.dsr2.2012.02.020
Steinberg D. K., Cope J. S., Wilson S. E., Kobari T. (2008). A comparison of mesopelagic mesozooplankton community structure in the subtropical and subarctic North Pacific Ocean. Deep Sea Res. 2 Top. Stud. Oceanogr 55, 1615–1635. doi: 10.1016/j.dsr2.2008.04.025
Sutton T. T., Clark M. R., Dunn D. C., Halpin P. N., Rogers A. D., Guinotte J., et al. (2017). A global biogeographic classification of the mesopelagic zone. Deep Sea Res. Part I 126, 85–102. doi: 10.1016/0198-0149(86)90094-4
Terazaki M., Miller C. B. (1986). Life history and vertical distribution of pelagic chaetognaths at Ocean station P in the subarctic Pacific. Deep Sea Res. Part Oceanogr. Res. Pap. 33, 323–337. doi: 10.1016/0198-0149(86)90094-4
Webb T. J., vanden Berghe E., O’Dor R. (2010). Biodiversity’s big wet secret: The global distribution of marine biological records reveals chronic under-exploration of the deep pelagic ocean. PloS One 5, e10223. doi: 10.1371/journal.pone.0010223
Wiebe P. H., Bucklin A., Madin L., Angel M. V., Sutton T., Pagès F., et al. (2010). Deep-sea sampling on CMarZ cruises in the Atlantic Ocean - an Introduction. Deep Sea Res. 2 Top. Stud. Oceanogr 57, 2157–2166. doi: 10.1016/j.dsr2.2010.09.018
Wiebe P. H., Morton A. W., Bradley A. M., Backus R. H., Craddock E., Barber V., et al. (1985). New developments in the MOCNESS, an apparatus for sampling zooplankton and micronekton. Mar. Biol. 87, 313–323. doi: 10.1007/BF00397811
WoRMS (2024). Accessed through: World Register of Marine Species. Available online at: https://www.marinespecies.org/aphia.php?p=searchon2024-10-18 (Accessed October 18, 2024).
Yamaguchi A., Matsuno K., Homma T. (2015). Spatial changes in the vertical distribution of calanoid copepods down to great depths in the North Pacific. Zool Stud. 54, 1–12. doi: 10.1186/s40555-014-0091-6
Yamaguchi A., Watanabe Y., Ishida H., Harimoto T., Furusawa K., Suzuki S., et al. (2002). Community and trophic structures of pelagic copepods down to greater depths in the western subarctic Pacific (WEST-COSMIC). Deep-Sea Res. I 49, 1007–1025. doi: 10.1016/S0967-0637(02)00008-0
Youngbluth M. J. (1984a). “Manned submersibles and sophisticated instrumentation: Tools for oceanographic research,” in Proceedings of SUBTECH ’83 symposium (Society for Underwater Technology, London), 335–344. Available at: http://purl.fcla.edu/fau/fauir (Accessed October 18, 2024).
Youngbluth M. J. (1984b). “Water column ecology: in situ observations of marine zooplankton from a manned submersible,” in Divers, submersibles and marine science. Occasional papers in biology. Ed. Fleming N. C. (London: Memorial University of Newfoundland), 45–57.
Keywords: 18S, 16S, COI, DNA reference database, MetaZooGene
Citation: Questel JM, Smoot CA, Collins AG, Lindsay DJ and Hopcroft RR (2025) DNA barcoding deep-water zooplankton from the Gulf of Alaska, North Pacific Ocean. Front. Mar. Sci. 12:1515048. doi: 10.3389/fmars.2025.1515048
Received: 22 October 2024; Accepted: 06 March 2025;
Published: 27 March 2025.
Edited by:
Clara F. Rodrigues, University of Aveiro, PortugalReviewed by:
Maria Pia Miglietta, Texas A&M University at Galveston, United StatesCopyright © 2025 Questel, Smoot, Collins, Lindsay and Hopcroft. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Jennifer M. Questel, am1xdWVzdGVsQGFsYXNrYS5lZHU=
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.