
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Mar. Sci. , 03 April 2025
Sec. Deep-Sea Environments and Ecology
Volume 12 - 2025 | https://doi.org/10.3389/fmars.2025.1514874
Introduction: Fungal communities have only been studied in a small portion of the vast variety of habitats that exist in deepsea environments, and studies aimed at understanding fungal diversity and function are minimal.
Objective: The aim of this study is to explore both the fungal diversity in deepsea sediments and the enzymatic activities present in them, which are related to the ecological roles of the strains and their biotechnological potential.
Methods: Eighteen sediment samples from three expeditions to deepsea areas of the Eastern Tropical Pacific (ETP) of Costa Rica were analyzed. Fungi were cultured on R2A medium, followed by physical characterization and molecular analysis (ITS and whole-genome sequencing) for the taxonomic identification of the strains. Once pure cultures were established, enzymatic tests for cellobiase, chitinase, lipase, cellulase, peroxidase, and laccase activities were performed, as well as surfactant activity.
Results: Fifty-five fungal strains were isolated, and genetic analysis was conducted on 27 strains, of which 7.41% belong to the Basidiomycota group and 92.59% to Ascomycota. These strains are distributed across 14 species. Among the identified strains are Periconia LEGMi281a and Hortaea LEGMi415c. Two strains exhibited cellobiase and chitinase activity, one strain exhibited cellulase activity, and one exhibited laccase production. None of the species exhibited lipase or peroxidase activity, and no clear surfactant activity was detected. Whole-genome sequencing revealed significant size differences compared to reference genomes.
Conclusion: The enzymatic activities of the strains suggest they may play a role in the degradation of organic matter and nutrient recycling, similar to terrestrial fungal counterparts. The differences in genome sizes, with the genomes of Periconia LEGMi281a and Hortaea LEGMi415c being larger than the reference genomes, pave the way for future research into deepsea adaptations, reflected in genetic changes. Additionally, the strains were identified as having high biotechnological potential.
Fungi inhabited the oceans before they conquered terrestrial ecosystems. Considering that the deep ocean represents the largest biome on the planet, there is a scarcity of studies on the diversity and ecology of fungi in these ecosystems compared to the rest of the ocean. Furthermore, what is known about microbial ecology in deepsea sediments primarily concerns bacteria and archaea (Edgcomb et al., 2011; Nagano and Nagahama, 2012; Dekas et al., 2016; Xu et al., 2018). Therefore, detailed knowledge of deepsea fungi is required to understand their contribution to marine food webs and global biogeochemical cycles (Manohar and Raghukumar, 2013; Barone et al., 2018; Drake and Ivarsson, 2018; Grossart et al., 2019; Román et al., 2019; Hassett et al., 2020).
Fungal communities have been studied only in a small portion of the vast variety of habitats that exist in deep waters. Some of these habitats include hydrothermal vents, cold methane seeps, oxygen minimum zones, and those associated with other macronutrients (Nagahama et al., 2011; Zhang et al., 2016; Batista-García et al., 2017; Rojas-Jiménez et al., 2020). Some studies have shown that the marine sediments represents an ecosystem where micro-aerobic respiration occurs and where microbial life is present, even hundreds of meters below the seafloor (D’Hondt, 2002; D’Hondt et al., 2015; Ivarsson et al., 2016a; Nagano et al., 2016).
In general, Ascomycota and Basidiomycota are the most abundant groups in deepsea ecosystems, representing between 70-80% and 10-20%, respectively. Some of the most abundant filamentous fungal genera include Aspergillus, Cladosporium, Fusarium, and Penicillium, while among yeasts are Candida, Cryptococcus, Metschnikowia, Rhodosporidium, and Rhodotorula (Li et al., 2016, 2019; Xu et al., 2016, 2019; Zhang et al., 2016; Nagano et al., 2017; Barone et al., 2018; Wang et al., 2019; Rojas-Jiménez et al., 2020).
In the deep sea, fungi must adapt to the absence of light, low temperatures, and high hydrostatic pressure. Fungi present in deepsea sediments can survive on marine snow, which consists of organic matter derived from photosynthesis that occurs in the photic zone (Bochdansky et al., 2017). In addition to performing aerobic respiration, fungi may be capable of processes such as fermentation, sulfate reduction, methanogenesis (D’Hondt, 2002; Lenhart et al., 2012), and possibly lithoautotrophy (López-García et al., 2003; Nealson et al., 2005; Ivarsson et al., 2016b). Transcriptomic analyses also confirm fungi as active members of deepsea sediments, revealing activities related to carbon and fatty acid metabolism (Pachiadaki et al., 2016). These metabolic processes may be critical for deepsea fungi as it has been observed that with increasing depth, fungal populations exhibit more multitrophic lifestyles (Li et al., 2019).
Considering the vast area to be explored to understand fungal diversity and function in deepsea sediments, it can be said that existing research is sparse and often lacks adequate spatial and temporal resolution (Grossart and Rojas-Jimenez, 2016; Grossart et al., 2019; Morales et al., 2019). As a result, there are many underexplored geographic locations, including the Eastern Tropical Pacific (ETP).
The deep waters of the ETP are particularly important ecosystems in Costa Rica, as they represent over 90% of the national territory (Cortés, 2016, 2019). This area, along the subduction zone of the Cocos and Caribbean plates, features a high diversity of microhabitats (Lizano, 2001; Protti et al., 2012; Rojas and Alvarado, 2012; Cortés, 2016), including cold seeps of methane (Sahling et al., 2008; Levin et al., 2012, 2015), with a high diversity (Seid et al., in prep.), seamounts (Auscavitch et al, in prep.), the Coco Submarine Volcanic Range (Cortés, 2016; Alvarado-Induni, 2021). Previous studies have demonstrated high endemism in the diversity of macro and microorganisms in this region (Rusch et al., 2007; Cortés, 2008, 2019; Levin et al., 2015; Rojas-Jiménez, 2018). Additionally, the Costa Rican ETP is part of the Eastern Tropical Pacific Marine Corridor, that also includes Panamá, Colombia and Ecuador, which represents a crucial site for the conservation and marine regeneration of ETP species (Cortés, 2012). This study analyzed sediment samples from various points in the Costa Rican Exclusive Economic Zone to understand both the fungal diversity in deepsea sediments and the enzymatic activities present, which relate to the ecological role of the strains and their biotechnological potential.
Eighteen sediment samples from three deepsea expeditions in the Eastern Tropical Pacific (ETP) of Costa Rica were analyzed (Figure 1). The first expedition took place from October 24 to November 5, 2018, aboard the R/V Atlantis. It was conducted along the continental shelf edge of Costa Rica, and all samples from this expedition were collected using the manned submersible Alvin. The second expedition occurred aboard the Schmidt Ocean Institute’s R/V Falkor in January 2019, covering seamounts from the continental coast to Isla del Coco, with samples collected by the remotely-operated vehicle SuBastian. The third expedition was conducted in December 2021 aboard the R/V Sally Ride. Six samples were processed from each expedition, distributed along a bathymetric gradient of 250 m to 3474 m depth. Each sample consisted of a sediment core approximately 30 cm long. Immediately after collecting the core from the seafloor, a 2 ml aliquot of sediment from the surface of the core, specifically from the first centimeter of the sediment column, was taken. Onboard, the samples were preserved at -40°C, and upon arrival at the onshore laboratory, they were stored at -80°C. The samples were collected with permission from MINAE (SINAC-CUSBSE-PIR-032-2018; R-070-2018-OT-CONAGEBIO) and are housed at the Laboratory of Microbial Ecology and Genetics (LEGMi) of the School of Biology of the University of Costa Rica (UCR).
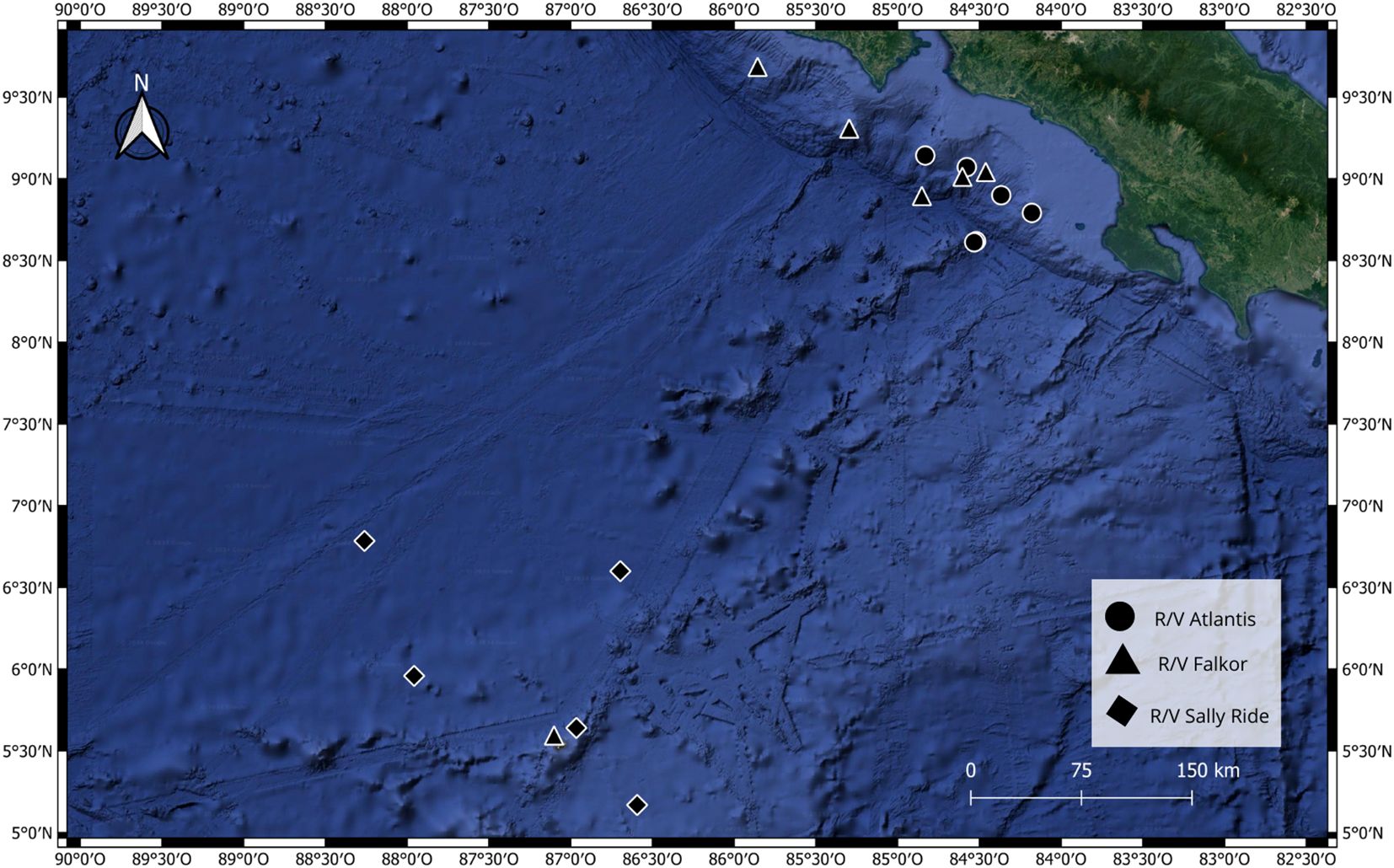
Figure 1. Sampling points of the three expeditions from which sediment samples were collected for this research.
To isolate the strains present in the sediment, the methodology of Agrawal et al. (2018) was utilized using distilled water. This was because the water collected at the sampling sites where the sediment was gathered was used in other research. For the sediment culture, 100 mg of frozen sediment was diluted in 1 ml of 1X PBS buffer to obtain a homogeneous mixture. Then, a 100 µl aliquot was placed on a Petri dish with culture medium. The medium was prepared by dissolving 4.55 g of R2A agar (Sigma-Aldrich, USA) in 250 ml of distilled water. These cultures were incubated at 28°C until they showed mycelial or yeast-like growth, for 3 weeks. The fungi from each sample were separated by morphospecies for further taxonomic and enzymatic characterization. For liquid cultures, 50 ml of MA+ culture medium (1 g/L yeast extract, 10 g/L malt extract, 1 g/L peptone, 2 g/L KH2PO4and 1 g/L MgSO4) was inoculated with 4 plugs of solid cultures (5 mm diameter), which were incubated for 7 days at 100 rpm, at room temperature (25°C).
Each sample was observed under a light microscope at 100X to identify reproductive or taxonomically significant structures. For observations, semi-permanent mounts of the species were made using polyvinyl, which were stored in the LEGMi laboratory at UCR. In addition to physical characterization, molecular analysis was performed for taxonomic identification of the species. DNA was extracted from 200 mg of tissue from the isolates using the Power Soil extraction Kit according to the manufacturer’s instructions (Qiagen, Carlsbad, CA, United States). The quality and quantity of DNA samples were assessed using the NanoDrop 2000 spectrophotometer.
PCR tests were then performed amplifying the ITS1 and ITS4 regions, with the following reaction conditions: 94°C for 2 minutes for initial denaturation, followed by 32 cycles of 94°C for 15 seconds, 53°C for 15 seconds, 72°C for 30 seconds, and a final extension at 72°C for 5 minutes (Rojas-Jimenez et al., 2017). The generated amplicons were sequenced by Macrogen Inc. Taxonomic assignment was done using BLAST with the GeneBank database from NCBI (Benson et al., 2012). Once the strains were identified, a phylogenetic tree was generated, aligning the sequences using the MAFFT v7 tool (Katoh et al., 2019). A maximum likelihood phylogenetic tree was created using FastTree version 2.1.9 (Price et al., 2010) with a GTR evolution model and a Bootstrap value of 1000. The resulting phylogenetic tree was visualized and edited in MEGA7 (Kumar et al., 2018).
Based on the sequencing results with the ITS markers and a literature review, two strains identified as Periconia sp. LEGMi281a and Hortaea sp. LEGMi415c were selected for genome sequencing. This selection was further supported by the intriguing characteristics of their closest known relatives. Periconia epilithographicola was only recently described (Coronado-Ruiz et al., 2018), and little is known about its metabolic and enzymatic activities, while Hortaea werneckii is recognized for its ability to survive the most extreme salinity ranges (Czachura et al., 2021). DNA was extracted from the selected strains using the same method mentioned earlier and sent to Novogen for sequencing using both Illumina and PacBio technologies. Raw PacBio reads were processed with Samtools v. 1.9 (Heng et al., 2018). The presence of repeats in the genome and unique or low-complexity sequences was identified using the Jellyfish v.2.3.0 tool (Marçais and Kingsford, 2011) and GenomeScope v.1.0 (Vurture et al., 2017) was used to determine the genome size, heterozygosity level, and repeat sequences level. Assemblies were carried out with Canu v.2.2 (Koren et al., 2017) using genome sizes of 46.7 Mbp and 38.2 Mbp for Periconia sp. LEGMi281a and Hortaea sp. LEGMi415c, respectively. The quality of the genomes was evaluated with Quast v. 5.2.0 (Gurevich et al., 2013), using P. macrospinosa (GenBank ID: GCA_003073855.1) and H. werneckii (GenBank ID GCA_037044905.1) as references. Additionally, the completeness of each assembly was assessed with BUSCO v.5.4.3 (Simão et al., 2015). Subsequently, the draft assemblies were optimized with Medusa v1.6-2 (Coelho et al., 2021), using different genomes associated with the species as references (Table 1). The same analyses with Quast and BUSCO were carried out with the optimized assemblies to show the improvements. Finally, a genomic alignment of the post-medusa assemblies and their respective references was performed using minimap2 v.2.17-r941 (Li, 2018) and visualized in D-Genies (Cabanettes and Klopp, 2018). The fungal genomes of this project have been deposited at GenBank under the accession number PRJNA1162099.
Six different enzymatic activities were evaluated: cellobiase, chitinase, lipase, cellulase, peroxidase, and laccase. For the chitinase, cellobiase, and lipase assays, the substrates used were 4-nitrophenyl N-acetyl-β-D-glucosaminide (Sigma), 4-nitrophenyl acetate (Sigma), and 4-nitrophenyl palmitate (Sigma), respectively. To determine enzymatic activity, 10 mM of each substrate was dissolved in 50 mM of ammonium acetate buffer pH 5, with 0.7% agar, at a temperature of 55°C. A drop of the substrate solution was placed on the mycelium, and a positive result was indicated by a color change from clear to yellow (Rojas-Jiménez and Hernández, 2015; Sandi et al., 2020). Peroxidase, laccase and cellulase activities were determined using degradation assays with Remazol Brilliant Blue (RBBR), 2,2’-azinobis-3-ethylbenzothiazoline-6-sulfonic acid (ABTS) and Carboxymethyl cellulose (CMC) respectively. These were conducted in a medium consisting of 0.94 g/L KH2PO4, 1.9 g/L K2HPO4, 1.6 g/L KCl, 1.43 g/L NaCl, 0.15 g/L NH4Cl, 0.037 g/L MgSO4, 0.1 g/L yeast extract, 10 g/L malt extract, 15 g/L agar, pH 7.0. The substrates were added to the medium as follows: 2% RBBR, 1% ABTS and 0.75% CMC according to (Rojas-Jimenez et al., 2017).
Surfactant activity was assessed using the oil displacement assay, the emulsification index E24 and the blood hemolysis test. The oil displacement assay (ODA) was implemented following the protocol of Morikawa et al. (2000) with slight modifications, as this assay is semi-quantitative, sensitive, reliable, rapid, and simple (Da Silva et al., 2021). Sterile 90 mm diameter Petri dishes were used, where 40 mL of distilled water was placed. On top of the water, 10 μL of commercial canola oil was added in a thin layer using a micropipette. Subsequently, 10 μL of liquid culture filtrate was placed in the center of the oil layer, and a ruler was used to measure the displacement. Milli-Q water and Tween 20 (v/v) were used as negative and positive controls, respectively. Three technical replicates were evaluated for each sample.
The emulsification index E24 (E24%) was determined using the Cooper and Goldenberg (1987) protocol. Conventional 10 x 100 mm test tubes were rinsed with 70% ethanol and Milli-Q water, and 2 mL of commercial canola oil and liquid culture filtrate were added to each tube. Each tube was vigorously shaken for two minutes and incubated at room temperature for 24 hours undisturbed. After this time, the E24% was calculated by dividing the height of the emulsified layer.
Finally, the blood hemolysis test (BHT) was performed to evaluate hemolytic activity. YPG plates supplemented with 5% (v/v) blood (donated by the School of Veterinary Medicine at UNA) were used; each plate was inoculated in the center with a 6 mm plug of mycelium grown in R2A growth medium and incubated at 37°C for five days. After this time, the diameter of the hemolysis halo was measured according to Thavasi et al. (2011). At least six repetitions were performed for each isolate, with an uninoculated plate serving as a negative control.
In this study, 55 fungal strains were isolated from sediment samples from the deepsea floors of the Eastern Tropical Pacific off Costa Rica. Molecular analysis was performed on 27 of these isolates, of which 7.41% belong to the Basidiomycota group and 92.59% to Ascomycota. These strains are distributed among 14 species covering 8 orders: Xylariales (the most abundant with 7 isolates), Amphisphaeriales, Eurotiales, Cladosporiales, Capnodiales, Pleosporales, Polyporales, and Serinales (Figure 2). One of the most noteworthy isolates was a melanized (black) yeast identified as Hortaea sp. LEGMi415c, which represents a new record for marine fungal diversity in Costa Rica. Similarly, the presence of Periconia sp. LEGMi281a was noted for the first time in deepsea.
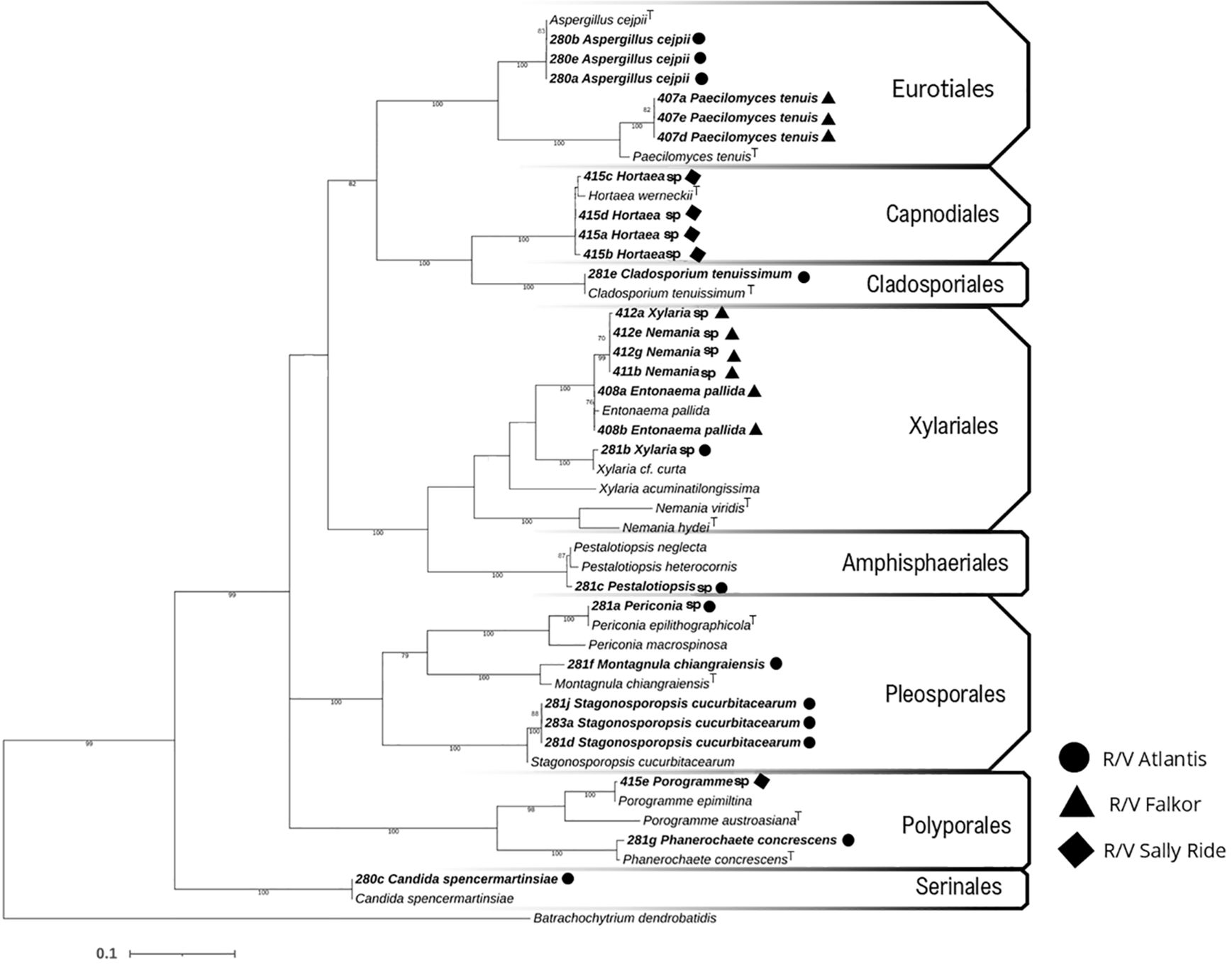
Figure 2. Phylogenetic relationship with maximum likelihood based on ITS markers, using the GTR model. Bootstrap values at nodes based on 1000 iterations, visualized in MEGA7.
Hortaea sp. LEGMi415c exhibited yeast-like and filamentous growth. Macroscopically, the colonies grow slowly (2 weeks), in a circular arrangement, with black/greenish-brown, mucous, shiny, and moist (Figure 3A). After 3 to 4 weeks, filamentous growth with brown pigments is observed at the colony’s periphery (Figure 3B). Microscopically, yeast-like cells were mostly observed as two-celled hyaline individuals, separated by a defined septum in the center of the transverse axis (Figure 3C).
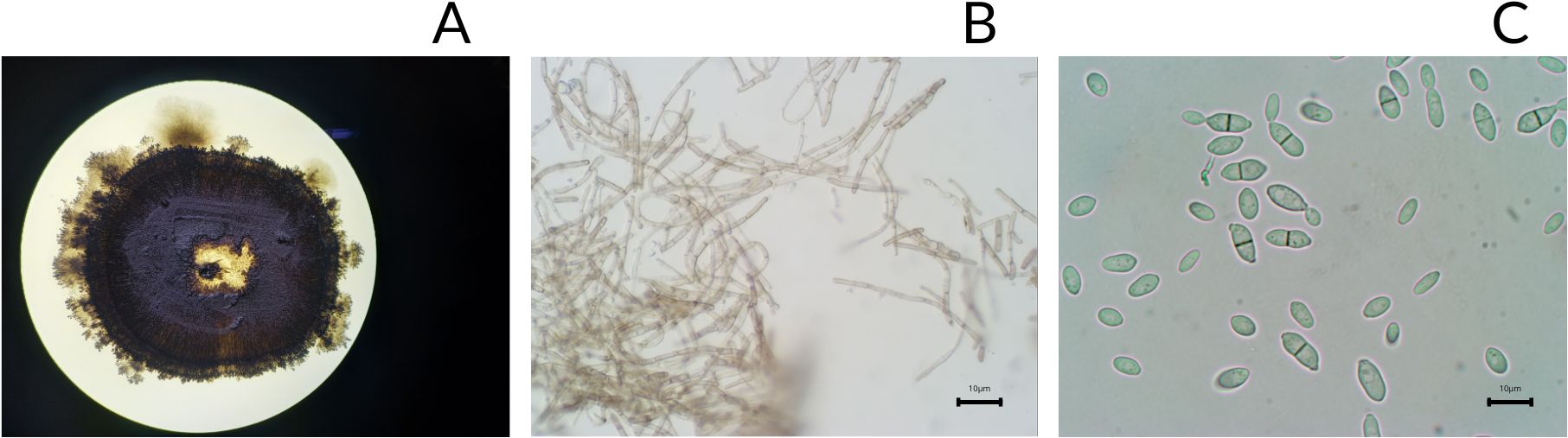
Figure 3. Photographs of Hortaea sp. LEGMi415c. Macroscopic view of the colony on a Petri dish (A), microscopic view of mycelial growth (B), microscopic view of yeast-like growth (C). The microscopy images were taken at 100X with immersion oil.
Periconia sp. LEGMi281a was characterized by growing on a plate with pink pigment mycelium. In the first week of growth, the mycelium was white, and around day 7, the first signs of pink coloration were observed (Figure 4A). Isolates grown on R2A did not show sporulation. However, two cells were observed with signs of being immature spores (Figure 4B), but the cultures did not develop further and did not reach sporulation. However, when these isolates were transferred to PDA, the specimens sporulated with spores averaging 8.5 µm in diameter (n=12) (Figure 4C, D).
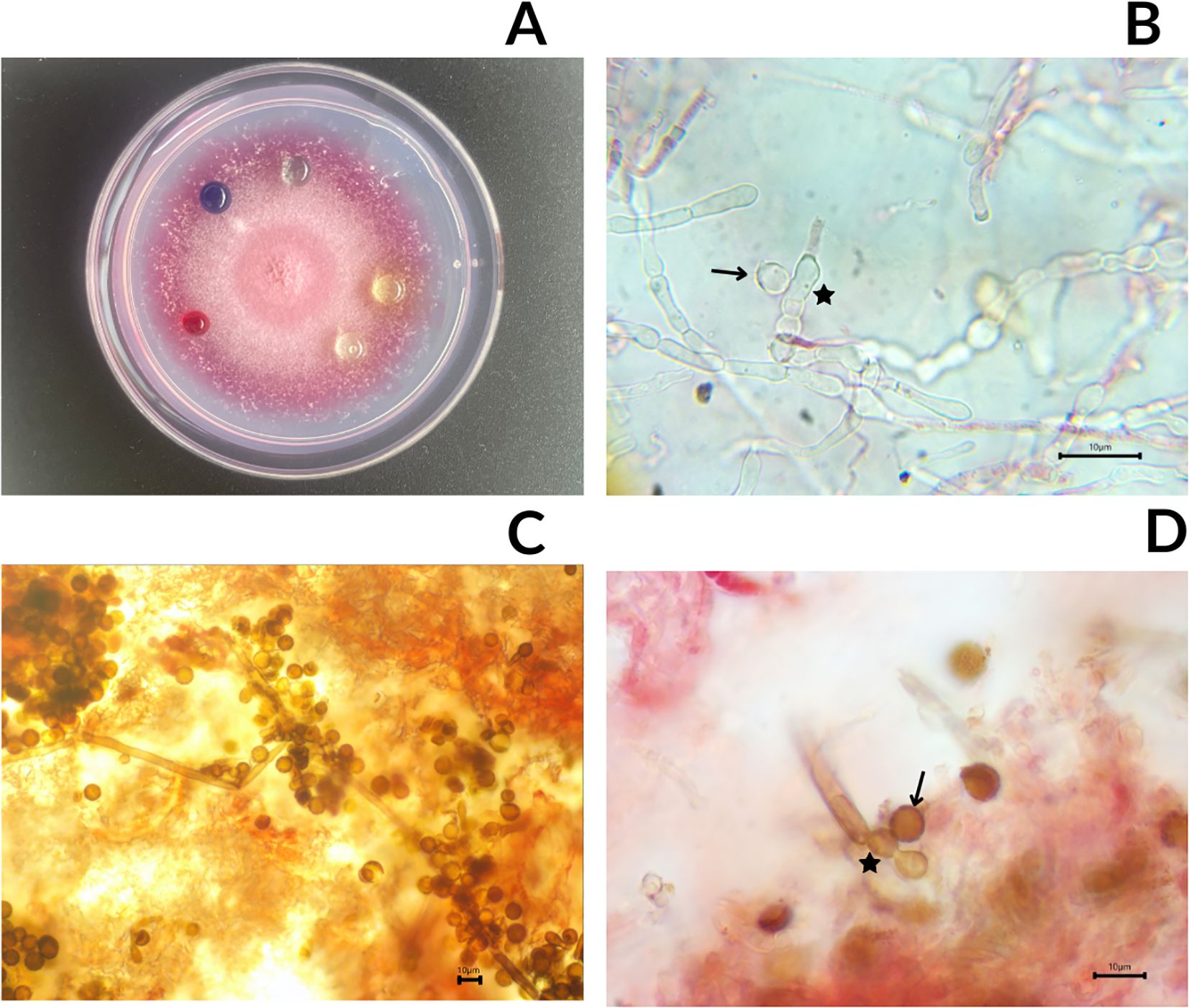
Figure 4. Photographs of Periconia sp. LEGMi281a. Macroscopic view of the colony on a Petri dish with drops from enzymatic assays (A), microscopy of immature spore marked with an arrow, in a conidiogenous cell marked with a star (B), microscopy of conidiophores (C), microscopy of mature spore marked with an arrow, in a conidiogenous cell marked with a star (D). The microscopy images were taken at 100X with immersion oil.
None of the 27 analyzed strains showed lipase or peroxidase activity. The only species with laccase activity was Hortaea sp. LEGMi415c, which exhibited a color change in the form of a halo with purple-blue tones after two weeks of growth. Regarding cellulase activity, only Periconia sp. LEGMi281a exhibited it, evident by observing colorless spots after staining the medium (with CMC) using Congo Red. Both species showed chitinase and cellobiase activity, with color changes from transparent to yellow in less than 10 minutes, indicating strong enzymatic activity.
Neither in the ODA test nor in the E24% test did Periconia sp. LEGMi281a show any activity. In the ODA tests, Hortaea sp. LEGMi415c exhibited partial displacement of the oil droplet, measuring 1.5 cm, 1.5 cm, and 2 cm in each of the replicates. In the E24% tests, two out of three replicates showed an emulsification index, with values of 31.58% and 26.32%. In the BHT tests, all three replicates of Hortaea sp. LEGMi415c were negative for hemolytic activity.
Initially, taxonomic classifications using only ITS markers identified the closest matches as Periconia epilithographicola and Hortaea werneckii. However, full genome analysis of the strains revealed discrepancies in the genomic assembly sizes compared to reference genomes (Table 2). The genomic size of Periconia sp. LEGMi281a is 68.5 Mbp, showing a difference of more than 10 Mbp compared to reference genomes, such as Periconia macrospinosa, which has a size of 55 Mbp. In the case of Hortaea sp. LEGMi415c, the size is 47.7 Mbp, which is also larger than the reference genome of Hortaea werneckii at 38.2 Mbp.
In a previous metagenomic study, 787 ASVs (Amplicon Sequence Variants) were identified in 40 sediment subsamples from deep areas of the Costa Rica ETP (Rojas-Jiménez et al., 2020). These findings anticipated a high fungal diversity and potential importance of these microorganisms to the ecosystem, which was later confirmed by the isolation of 27 strains in the present study. Similar research in the Indian Ocean reported 39 species with culture-dependent methods and 91 OTUs with molecular methods (Singh et al., 2012; Zhang et al., 2014). Molecular methods consistently reveal greater diversity than culture-dependent ones. This discrepancy arises because most microorganisms, including deepsea marine fungi, are not cultivable due to differences between culture conditions and their natural habitat (Lewis et al., 2021). While it is expected that adjusting variables such as temperature, pressure, and salinity in future studies will lead to the isolation of more strains, including new species, all isolates found in this study, regardless of identification at genus or species level are of great importance as they represent 14 new records of deep marine fungi for the country. This is a significant step in understanding the diversity of the deep ocean, particularly its mycobiota, as marine fungi account for less than 1% of known species (Raghukumar et al., 2010).
Fungi are crucial for nutrient recycling and energy flow in marine ecosystems, functioning as saprotrophs, mutualists, endobionts, and parasites (Jones et al., 2019; Grossart et al., 2019). However, their specific roles in deepsea ecosystems remain unclear, underscoring the need for isolates to study their physiology and ecological functions. Some species from this study, also found in terrestrial environments, have been reported in other marine ecosystems. For example, Aspergillus cejpii was found in hydrothermal vents (Pacheco, 2024), and Stagonosporopsis cucurbitacearum was isolated from a marine sponge (Haga et al., 2013). These strains have known saprotrophic activity in terrestrial environments (Edgcomb et al., 2011), so given their adaptations to osmotic stress and polysaccharide degradation (Gonçalves et al., 2022), it is hypothesized that these fungi primarily act as saprotrophs in sediments, recycling nutrients from marine snow and deepsea fauna waste.
The genus Hortaea, from the order Capnodiales, is commonly isolated from soil, marine products like dried fish, dust, wood, compost, and human skin (Prieto-Granada et al., 2010). While reported in marine ecosystems such as the Mediterranean water column, deepsea stations Vector and Geostar, and the Rainbow deep hydrothermal vent (Romeo et al., 2020; Cochereau et al., 2023; De Leo et al., 2019), it has not been recorded in Latin American deepsea ecosystems, making this a new report for the region.
Hortaea includes the most halotolerant fungus known, capable of growing in salinities from 0% to 32% NaCl (Butinar et al., 2005; Czachura et al., 2021). Melanin production is crucial for this halotolerance, acting as an osmoregulatory agent and maintaining cell wall integrity (Kejžar et al., 2013). The fungus alternates between yeast-like and mycelial phases (Butinar et al., 2005; Czachura et al., 2021). At low temperatures (15 ± 5°C), yeast-like growth predominates (Elsayed et al., 2016). Given that deepsea temperatures are around 2°C, Hortaea sp. LEGMi415c likely exhibits yeast-like growth in these ecosystems. Conversely, at high temperatures (30 ± 5°C), mycelial growth predominates, but yeast-like growth can occur even at elevated temperatures in the absence of NaCl (Elsayed et al., 2016). This explains the predominant yeast-like growth observed at 28°C without added NaCl. However, the transition to mycelial growth over time may be due to nutrient depletion, as mycelial growth allows for better nutrient exploration and absorption. Further research is needed to understand how temperature, salinity, and nutrient interactions affect the growth forms of Hortaea sp. LEGMi415c.
Hortaea sp. LEGMi415c exhibited laccase, chitinase, and cellobiase activities, suggesting that this strain contributes to nutrient recycling in the ecosystem. Laccases can degrade complex aromatic compounds and the ones previously reported in marine fungi can maintain their degradative capacity in marine ecosystems (Ali et al., 2020a, b; Xu et al., 1996; Bonugli-Santo et al., 2010). On the other hand, chitinase recycles nitrogen and carbon from chitin, which is found in crustacean exoskeletons and polychaetes, common in deepsea environments (Kellner et al., 2010; Choi et al., 2022). Additionally, chitinases play a role in growth modulation and spore formation and germination (El Gueddari et al., 2002; Karlsson and Stenlid, 2008). These enzymatic activities suggest that Hortaea sp. LEGMi415c is metabolically active in deepsea ecosystems, participating in nutrient recycling through both surface biomass and remains of deepsea organisms.
From a biotechnological perspective, laccases are highly versatile and have applications in the textile, pharmaceutical, materials industries, biomedicine, and bioremediation due to their ability to degrade complex compounds (Chaudhary et al., 2022). Chitinases are used in agricultural biocontrol and in the pharmaceutical industry (Sharma et al., 2023), and cellobiase is important for ethanol production (Li et al., 2020). The present study found extracellular activities of Hortaea sp. LEGMi415c, such as laccase production, despite other studies on Hortaea isolates showing no extracellular enzymatic activity with 0% NaCl (Rizk and Magdy, 2022). This suggests enzymatic plasticity related to environmental conditions. The ability to produce enzymatic extracts without adjusting salinity is advantageous industrially (Mesbah, 2022), highlighting the industrial potential of deepsea microorganisms. Although ITS markers classified the strain as Hortaea werneckii and the isolated strain shows similar behaviors, whole-genome analyses reveal significant discrepancies, with genome sizes differing by 9.5 Mbp. Further analysis is needed to determine if these differences indicate adaptations to deepsea ecosystems and if the strain represents a new species.
The genus Periconia is polymorphic and belongs to the order Pleosporales. Species of this genus are commonly isolated from plant samples, as endophytes (Su et al., 2023), and from marine and freshwater ecosystems (Morrison-Gardiner, 2002; Cai et al., 2002). This is the first report of a Periconia species in deepsea marine environments. Differences in genome sizes and GC percentages indicate that the strain isolated from deepsea sediments differs from other reference species. As there is no complete genome sequencing for P. epilithographicola at the moment of this study, it was not included in the genomic analysis, but the morphological description of the isolated strain aligns with P. epilithographicola, supporting its taxonomic classification based on ITS markers.
P. epilithographicola was initially discovered on 19th-century lithographs (Coronado-Ruiz et al., 2018). The natural history of the genus explains why Periconia sp. LEGMi281a, despite being isolated from an environment without cellulose production, is the only strain analyzed with cellulolytic capability. Cellulose is the most abundant natural polysaccharide, and marine fungi can degrade it (Le Strat et al., 2022). Degrading this polymer is advantageous since a significant portion of terrestrial cellulose ends up in the deep sea, and algal cellulose reaches deepsea areas through marine snow (Tsudome et al., 2022). Cellulase activity may provide a competitive edge by utilizing this carbon source, unavailable to other fungi without these enzymes. Additionally, the chitinase and cellobiase activities of Periconia sp. LEGMi281a suggest it, like Hortaea sp. LEGMi415c, plays a role in degrading organic remains of deepsea organisms.
Since this species was only described six years ago, there is limited research on its biotechnological potential. However, it has been reported that its presence in a community increases in the presence of humic acids (Da Silva et al., 2022), so due its cellulase, chitinase, and cellobiase activity, this species could be a candidate for bioremediation studies. Other species within the genus are already considered for biocontrol due to the presence of chitinases (Sharma et al., 2023), suggesting that Periconia sp. LEGMi281a could also be used in this area in the future. Additionally, its potential to maintain these activities under low temperatures and high hydrostatic pressure highlights its significant biotechnological potential, warranting further exploration.
This research indicates that the deepsea sediments in the ETP harbor rich fungal communities that deserve more detailed investigation. Comparisons with molecular studies show common microorganisms in these communities that were not isolated. For example, the genus Metschnikowia was the most abundant in several samples from a previous metagenomic study (Rojas-Jiménez et al., 2020) but was not present in the isolates from this study. This underscores the high potential for discovering new species.
The fungi isolated from the sediment samples suggest important ecological roles and physiological traits with biotechnological applications. These microorganisms possess enzymes with a wide range of industrial uses and may be more resilient than other deepsea fungi (Mahmoudi et al., 2020). Moreover, their ability to grow under regular environmental conditions (1 atm and 25°C) implies that their industrial production would not require high-energy systems like cooling or pressure-increasing systems.
The results raise questions about the role of these species in deepsea ecosystems. Obligate marine fungi can only grow and sporulate in marine ecosystems, while facultative fungi can thrive in other habitats due to their physiological adaptations (Kohlmeyer and Kohlmeyer, 1979). Since Periconia and Hortaea are facultative marine fungi, further research is needed on the ecological roles and enzymatic activities of Periconia sp. LEGMi281a and Hortaea sp. LEGMi415c in marine environments. Additionally, it is essential to understand the evolutionary significance of metabolic traits thought to be selected by terrestrial pressures in a deepsea context. The industrial and biomedical relevance of these microorganisms, alongside the gaps in our knowledge about their ecology and natural history in deepsea ecosystems, supports the need for conservation of these habitats.
The raw data supporting the conclusions of this article will be made available by the authors, without undue reservation.
IR-R: Conceptualization, Data curation, Investigation, Methodology, Writing – original draft. SS-G: Data curation, Investigation, Methodology, Writing – review & editing. JC: Conceptualization, Data curation, Investigation, Writing – review & editing. KR-J: Data curation, Investigation, Methodology, Writing – review & editing, Conceptualization, Project administration, Supervision.
The author(s) declare that financial support was received for the research and/or publication of this article. The financial support to make the study was given by Vicerrectoria de Investigación, Universidad de Costa Rica (Project C3027), in partnership with FAICO, Charles Darwin Foundaton and Bezos Earth Fund.
We deeply appreciate the support by Erik Cordes, Chief Scientists, the scientific team and the crew of the Atlantis (Alvin) and Falkor (SuBastian). We thank Celeste Sánchez-Noguera for collecting the Sally Ride samples.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declared that they were an editorial board member of Frontiers, at the time of submission. This had no impact on the peer review process and the final decision.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Agrawal S., Adholeya A., Barrow C. J., Deshmukh S. K. (2018). In-vitro evaluation of marine derived fungi against Cutibacterium acnes. Anaerobe 49, 5–13. doi: 10.1016/j.anaerobe.2017.10.010
Ali W. B., Ayed A. B., Turbé-Doan A., Bertrand E., Mathieu Y., Faulds C. B., et al. (2020a). Enzyme properties of a laccase obtained from the transcriptome of the marine-derived fungus Stemphylium lucomagnoense. Int. J. Mol. Sci. 21, 8402. doi: 10.3390/ijms21218402
Ali W. B., Navarro D., Kumar A., Drula E., Turbé-Doan A., Correia L. O., et al. (2020b). Characterization of the CAZy repertoire from the marine-derived fungus Stemphylium lucomagnoense in relation to saline conditions. Mar. Drugs 18, 461. doi: 10.3390/md18090461
Alvarado-Induni G. E. (2021). Costa Rica y sus volcanes (Costa Rica: Editorial UCR, Editorial UNA, Editorial TCR), 976.
Barone G., Rastelli E., Corinaldesi C., Tangherlini M., Danovaro R., Dell’Anno A. (2018). Benthic deepsea fungi in submarine canyons of the Mediterranean Sea. Prog. Oceanography 168, 57–64. doi: 10.1016/j.pocean.2018.09.011
Batista-García R. A., Sutton T., Jackson S. A., Tovar-Herrera O. E., Balcázar-López E., del-Rayo-Sánchez-Carbente M., et al. (2017). Characterization of lignocellulolytic activities from fungi isolated from the deepsea sponge stelletta normani. PloS One 12 (3), e0173750. doi: 10.1371/journal.pone.0173750
Benson D. A., Cavanaugh M., Clark K., Karsch-Mizrachi I., Lipman D. J., Ostell J., et al. (2012). GenBank. Nucleic Acids Res. 41, 36–42. doi: 10.1093/nar/gks1195
Bochdansky A. B., Clouse M. A., Herndl G. J. (2017). Eukaryotic microbes, principally fungi and labyrinthulomycetes, dominate biomass on bathypelagic marine snow. ISME J. 11, 362–373. doi: 10.1038/ismej.2016.113
Bonugli-Santos R. C., Durrant L. R., da Silva M., Sette L. D. (2010). Production of laccase, manganese peroxidase and lignin peroxidase by Brazilian marine-derived fungi. Enzyme Microbial Technol. 46, 32–37. doi: 10.1016/j.enzmictec.2009.07.014
Butinar L., Sonjak S., Zalar P., Plemenitaš A., Gunde-Cimerman N. (2005). Melanized halophilic fungi are eukaryotic members of microbial communities in hypersaline waters of solar salterns. Botanica Marina 48, 73–79. doi: 10.1515/BOT.2005.007
Cabanettes F., Klopp C. (2018). D-GENIES: dot plot large genomes in an interactive, efficient and simple way. PeerJ 6, 4958. doi: 10.7717/peerj.4958
Cai L., Tsui C. K. M., Zhang K. Q., Hyde K. D. (2002). Aquatic fungi from lake fuxian, yunnan, China. Fungal Diversity 14, 57–70.
Chaudhary S., Singh A., Varma A., Porwal S. (2022). Recent advancements in biotechnological applications of Laccase as a multifunctional enzyme. J. Pure Appl. Microbiol. 16 (3), 1479–1491. doi: 10.22207/JPAM.16.3.72
Choi H., Kim S. L., Jeong M. K., Yu O. H., Eyun S. (2022). Identification and phylogenetic analysis of chitin synthase genes from the deepsea polychaete Branchipolynoe onnuriensis genome. J. Mar. Sci. Eng. 10, 598. doi: 10.3390/jmse10050598
Cochereau B., Le Strat Y., Ji Q., Pawtowski A., Delage L., Weill A., et al. (2023). Heterologous expression and biochemical characterization of a new chloroperoxidase isolated from the deepsea hydrothermal vent black yeast Hortaea werneckii UBOCC-A-208029. Mar. Biotechnol. 25, 519–536. doi: 10.1007/s10126-023-10222-7
Coelho M. A. G., Vicente T. P., Santos S., Reis J. F., Azevedo E., Costa A. F., et al. (2021). Medusa: Multi-draft based scaffolder. Bioinformatics 37, 726–731.
Cooper D. G., Goldenberg B. G. (1987). Surface-active agents from two Bacillus species. Appl. Environ. Microbiol. 53, 224–229. doi: 10.1128/AEM.53.2.224-229.1987
Coronado-Ruiz C., Avendaño R., Escudero-Leyva E., Conejo-Barboza G., Chaverri P., Chavarría M. (2018). Two new cellulolytic fungal species isolated from a 19th-century art collection. Sci. Rep. 8. doi: 10.1038/s41598-018-24934-7
Cortés J. (2008). Historia de la investigación marina de la Isla del Coco, Costa Rica. Rev. Biol. Trop. 56, 1–18. doi: 10.15517/rbt.v56i2.26934
Cortés J. (2012). Marine biodiversity of an eastern tropical Pacific oceanic island, Isla del Coco, Costa Rica. Rev. Biol. Trop. 60, 131–185. doi: 10.15517/rbt.v60i3.28356
Cortés J. (2016). “The Pacific coastal and marine ecosystems,” in Costa rican ecosystems. Ed. Kappelle M. (The University of Chicago Press, Chicago), 97–138. doi: 10.7208/chicago/9780226121642.003.0005
Cortés J. (2019). “Isla del coco, Costa Rica, eastern tropical pacific,” in Mesophotic coral ecosystems (Switzerland: Springer, Cham), vol. 12 . Eds. Loya Y., Puglise K., Bridge &T., 465–475). doi: 10.1007/978-3-319-92735-0_26
Czachura P., Owczarek-Kościelniak M., Piątek M. (2021). Salinomyces polonicus: A moderately halophilic kin of the most extremely halotolerant fungus Hortaea werneckii. Fungal Biol. 125, 459–468. doi: 10.1016/j.funbio.2021.01.003
D’Hondt S. (2002). Metabolic activity of subsurface life in deepsea sediments. Science. 295, 2067–2070. doi: 10.1126/science.1064878
D’Hondt S., Inagaki F., Zarikian C. A., Abrams L. J., Dubois N., Engelhardt T., et al. (2015). Presence of oxygen and aerobic communities from sea floor to basement in deepsea sediments. Nat. Geosci. 8, 299–304. doi: 10.1038/ngeo2387
Da Silva A. F., Banat I. M., Giachini A. J., Robl D. (2021). Fungal biosurfactants, from nature to biotechnological product: Bioprospection, production and potential applications. Bioprocess Biosyst. Eng. 44, 2003–2034. doi: 10.1007/s00449-021-02597-5
Da Silva M. S. R. D. A., De Carvalho L. A. L., Braos L. B., de Sousa Antunes L. F., Da Silva C. S. R. D. A., Da Silva C. G. N., et al. (2022). Effect of the application of vermicompost and millicompost humic acids about the soybean microbiome under water restriction conditions. Front. Microbiol. 13. doi: 10.3389/fmicb.2022.1000222
Dekas A. E., Connon S. A., Chadwick G. L., Trembath-Reichert E., Orphan V. J. (2016). Activity and interactions of methane seep microorganisms assessed by parallel transcription and FISH-NanoSIMS analyses. ISME J. 10, 678–692. doi: 10.1038/ismej.2015.145
De Leo F., Lo Giudice A., Alaimo C., De Carlo G., Rappazzo A. C., Graziano M., et al. (2019). Occurrence of the black yeast Hortaea werneckii in the Mediterranean Sea. Extremophiles 23, 9–17. doi: 10.1007/s00792-018-1056-1
Drake H., Ivarsson M. (2018). The role of anaerobic fungi in fundamental biogeochemical cycles in the deep biosphere. Fungal Biol. Rev. 32, 20–25. doi: 10.1016/j.fbr.2017.10.001
Edgcomb V. P., Beaudoin D., Gast R., Biddle J. F., Teske A. (2011). Marine subsurface eukaryotes: The fungal majority. Environ. Microbiol. 13, 172–183. doi: 10.1111/j.1462-2920.2010.02318.x
El Gueddari N., Rauchhaus U., Moerschbacher B. M., Deising H. B. (2002). Developmentally regulated conversion of surface-exposed chitin to chitosan in cell walls of plant pathogenic fungi. New Phytol. 156, 103–112. doi: 10.1046/j.1469-8137.2002.00487.x
Elsayed A., Mowafy A. M., Soliman H. M., Gebreil A., Magdy N. I. (2016). Characterization of new strains of Hortaea werneckii isolated from salt marshes of Egypt. Egyptian J. Basic Appl. Sci. 3, 350–356. doi: 10.1016/j.ejbas.2016.09.00
Gonçalves M. F., Esteves A. C., Alves A. (2022). Marine fungi: Opportunities and challenges. Encyclopedia 2, 559–577. doi: 10.3390/encyclopedia2010037
Grossart H.-P., Rojas-Jimenez K. (2016). Aquatic fungi: Targeting the forgotten in microbial ecology. Curr. Opin. Microbiol. 31, 140–145. doi: 10.1016/j.mib.2016.03.016
Grossart H.-P., Van den Wyngaert S., Kagami M., Wurzbacher C., Cunliffe M., Rojas-Jimenez K. (2019). Fungi in aquatic ecosystems. Nat. Rev. Microbiol. 17, 339–354. doi: 10.1038/s41579-019-0175-8
Gurevich A., Saveliev V., Vyahhi N., Tesler G. (2013). QUAST: Quality assessment tool for genome assemblies. Bioinformatics 29, 1072–1075. doi: 10.1093/bioinformatics/btt086
Haga A., Tamoto H., Ishino M., Kimura E., Sugita T., Kinoshita K., et al. (2013). Pyridone alkaloids from a marine-derived fungus, Stagonosporopsis cucurbitacearum, and their activities against azole-resistant Candida albicans. J. Natural Products 76, 750–754. doi: 10.1021/np300876t
Hassett B. T., Vonnahme T. R., Peng X., Jones E. B. G., Heuzé C. (2020). Global diversity and geography of planktonic marine fungi. Botanica Marina 63, 121–139. doi: 10.1515/bot-2018-0113
Heng L., Bob H., Richard D. (2018). Samtools. (1.9) [Software]. Available online at: http://www.htslib.org (Accessed July 15, 2024).
Ivarsson M., Bengtson S., Neubeck A. (2016a). The igneous oceanic crust – Earth’s largest fungal habitat? Fungal Ecol. 20, 249–255. doi: 10.1016/j.funeco.2016.01.009
Ivarsson M., Schnürer A., Bengtson S., Neubeck A. (2016b). Anaerobic fungi: A potential source of biological H₂ in the oceanic crust. Front. Microbiol. 7. doi: 10.3389/fmicb.2016.00674
Jones E. G., Pang K.-L., Abdel-Wahab M. A., Scholz B., Hyde K. D., Boekhout T., et al. (2019). An online resource for marine fungi. Fungal Diversity 96, 347–433. doi: 10.1007/s13225-019-00426-5
Karlsson M., Stenlid J. (2008). Comparative evolutionary histories of the fungal chitinase gene family reveal non-random size expansions and contractions due to adaptive natural selection. Evolutionary Bioinf. 4, 47–60. doi: 10.4137/EBO.S604
Katoh K., Rozewicki J., Yamada K. D. (2019). MAFFT online service: Multiple sequence alignment, interactive sequence choice and visualization. Briefings Bioinf. 20, 1160–1166. doi: 10.1093/bib/bbx108
Kejžar A., Gobec S., Plemenitaš A., Lenassi M. (2013). Melanin is crucial for growth of the black yeast Hortaea werneckii in its natural hypersaline environment. Fungal Biol. 117, 368–379. doi: 10.1016/j.funbio.2013.03.006
Kellner H., Zak D. R., Vandenbol M. (2010). Fungi unearthed: Transcripts encoding lignocellulolytic and chitinolytic enzymes in forest soil. PloS One 5. doi: 10.1371/journal.pone.0010971
Koren S., Walenz B. P., Berlin K., Miller J. R., Phillippy A. M. (2017). Canu: Scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 27, 722–736. doi: 10.1101/gr.215087.116
Kumar S., Stecher G., Li M., Knyaz C., Tamura K. (2018). MEGA X: Molecular Evolutionary Genetics Analysis across computing platforms. Mol. Biol. Evol. 35, 1547–1549. doi: 10.1093/molbev/msy096
Lenhart K., Bunge M., Ratering S., Neu T. R., Schüttmann I., Greule M., et al. (2012). Evidence for methane production by saprotrophic fungi. Nat. Commun. 3, 1046. doi: 10.1038/ncomms2049
Le Strat Y., Ruiz N., Fleurence J., Pouchus Y. F., Déléris P., Dumay J. (2022). Marine fungal abilities to enzymatically degrade algal polysaccharides, proteins and lipids: A review. J. Appl. Phycology 34, 1131–1162. doi: 10.1007/s10811-022-02713-2
Levin L. A., Mendoza G. F., Grupe B. M., Gonzalez J. P., Jellison B., Rouse G., et al. (2015). Biodiversity on the rocks: Macrofauna inhabiting authigenic carbonate at Costa Rica methane seeps. PloS One 10, e0131080. doi: 10.1371/journal.pone.0131080
Levin L. A., Orphan V. J., Rouse G. W., Rathburn A. E., Ussler W., Cook G. S., et al. (2012). A hydrothermal seep on the Costa Rica margin: Middle ground in a continuum of reducing ecosystems. Proc. R. Soc. B: Biol. Sci. 279, 2580–2588. doi: 10.1098/rspb.2012.0205
Lewis W. H., Tahon G., Geesink P., Sousa D. Z., Ettema T. J. (2021). Innovations to culturing the uncultured microbial majority. Nat. Rev. Microbiol. 19, 225–240. doi: 10.1038/s41579-020-00458-8
Li H. (2018). Minimap2: Pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100. doi: 10.1093/bioinformatics/bty191
Li W., Wang M., Burgaud G., Yu H., Cai L. (2019). Fungal community composition and potential depth-related driving factors impacting distribution pattern and trophic modes from epi- to abyssopelagic zones of the Western Pacific Ocean. Microbial Ecol. 78, 820–831. doi: 10.1007/s00248-019-01374-y
Li W., Wang M. M., Wang X. G., Cheng X. L., Guo J. J., Bian X. M., et al. (2016). Fungal communities in sediments of subtropical Chinese seas as estimated by DNA metabarcoding. Sci. Rep. 6, 26528. doi: 10.1038/srep26528
Li J., Zhang Y., Li J., Sun T., Tian C. (2020). Metabolic engineering of the cellulolytic thermophilic fungus Myceliophthora thermophila to produce ethanol from cellobiase. Biotechnol. Biofuels 13, 1–15. doi: 10.1186/s13068-020-1661-y
Lizano O. (2001). Batimetría de la plataforma insular alrededor de las Isla del Coco, Costa Rica. Rev. Biología Trop. 49, 163–170.
López-García P., Philippe H., Gail F., Moreira D. (2003). Autochthonous eukaryotic diversity in hydrothermal sediment and experimental microcolonizers at the Mid-Atlantic Ridge. Proc. Natl. Acad. Sci. United States America 100, 697–702. doi: 10.1073/pnas.0235779100
Mahmoudi N., Hagen S. M., Hazen T. C., Steen A. D. (2020). Patterns in extracellular enzyme activity and microbial diversity in deepsea Mediterranean sediments. Deep Sea Res. Part I: Oceanographic Res. Papers 158, 103231. doi: 10.1016/j.dsr.2020.103231
Manohar C. S., Raghukumar C. (2013). Fungal diversity from various marine habitats deduced through culture-independent studies. FEMS Microbiol. Lett. 341, 69–78. doi: 10.1111/1574-6968.12087
Marçais G., Kingsford C. (2011). A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics 27, 764–770. doi: 10.1093/bioinformatics/btr011
Mesbah N. M. (2022). Industrial biotechnology based on enzymes from extreme environments. Front. Bioengineering Biotechnol. 10. doi: 10.3389/fbioe.2022.870083
Morales S. E., Biswas A., Herndl G. J., Baltar F. (2019). Global structuring of phylogenetic and functional diversity of pelagic fungi by depth and temperature. Front. Mar. Sci. 6. doi: 10.3389/fmars.2019.00131
Morikawa M., Hirata Y., Imanaka T. (2000). A study on the structure-function relationship of lipopeptide biosurfactants. Biochim. Biophys. Acta 1488, 211–218. doi: 10.1016/s1388-1981(00)00124-4
Morrison-Gardiner S. (2002). Dominant fungi from Australian coral reefs. Fungal Diversity 9, 105–121.
Nagahama T., Takahashi E., Nagano Y., Abdel-Wahab M. A., Miyazaki M. (2011). Molecular evidence that deep-branching fungi are major fungal components in deepsea methane cold-seep sediments. Environ. Microbiol. 13, 2359–2370. doi: 10.1111/j.1462-2920.2011.02507.x
Nagano Y., Konishi M., Nagahama T., Kubota T., Abe F., Hatada Y. (2016). Retrieval of deeply buried culturable fungi in marine subsurface sediments, Suruga-Bay, Japan. Fungal Ecol. 20, 256–259. doi: 10.1016/j.funeco.2015.12.012
Nagano Y., Miura T., Nishi S., Lima A. O., Nakayama C., Pellizari V. H., et al. (2017). Fungal diversity in deepsea sediments associated with asphalt seeps at the São Paulo Plateau. Deep Sea Res. Part II: Topical Stud. Oceanography 146, 59–67. doi: 10.1016/j.dsr2.2017.05.012
Nagano Y., Nagahama T. (2012). Fungal diversity in deepsea extreme environments. Fungal Ecol. 5, 463–471. doi: 10.1016/j.funeco.2012.01.004
Nealson K. H., Inagaki F., Takai K. (2005). Hydrogen-driven subsurface lithoautotrophic microbial ecosystems (SLiMEs): Do they exist and why should we care? Trends Microbiol. 13, 405–410. doi: 10.1016/j.tim.2005.07.010
Pacheco V. L. (2024). Evaluación del papel funcional de micromicetos aislados de ventilas hidrotermales de mar profundo (Golfo de California) en el ciclo del azufre (Master’s thesis) (Mexico: CICESE).
Pachiadaki M. G., Rédou V., Beaudoin D. J., Burgaud G., Edgcomb V. P. (2016). Fungal and prokaryotic activities in the marine subsurface biosphere at Peru margin and Canterbury Basin inferred from RNA-based analysis and microscopy. Front. Microbiol. 7. doi: 10.3389/fmicb.2016.00846
Price M. N., Dehal P. S., Arkin A. P. (2010). FastTree 2–approximately maximum-likelihood trees for large alignments. PloS One 5, e9490. doi: 10.1371/journal.pone.0009490
Prieto-Granada C. N., Lobo A. Z. C., Mihm M. C. (2010). Skin infections. In R. L. Kradin (Ed.). Diagn. Pathol. Infect. Dis. pp, 519–616. doi: 10.1016/B978-1-4160-3429-2.00019-5
Protti M., González V., Freymueller J., Doelger S. (2012). Isla del Coco, on Cocos Plate, converges with Isla de San Andrés, on the Caribbean Plate, at 78 mm/yr. Rev. Biología Trop. 60, 33–41. doi: 10.15517/rbt.v60i3.28307
Raghukumar C., Damare S., Singh P. (2010). A review on deepsea fungi: Occurrence, diversity and adaptations. Botanica Marina 53, 479–492. doi: 10.1515/bot.2010.076
Rizk S. M., Magdy M. (2022). An indigenous inland genotype of the black yeast Hortaea werneckii inhabiting the great pyramid of Giza, Egypt. Frontiers in Microbiology. 13 997495. doi: 10.3389/fmicb.2022.997495
Rojas W., Alvarado G. E. (2012). Marco geológico y tectónico de la Isla del Coco y la región marítima circunvecina. Rev. Biología Tropical 60(Supplement 3), 15–32. doi: 10.15517/rbt.v60i3.28306
Rojas-Jiménez K. (2018). Microorganismos del corredor marino Isla del Coco-Galápagos: diversidad funcional y de especies. Rev. Tecnología en Marcha 31, 157–166. doi: 10.18845/tm.v31i4.3974
Rojas-Jiménez K., Fonvielle J. A., Ma H., Grossart H. P. (2017). Transformation of humic substances by the freshwater Ascomycete. Cladosporium Limnology Oceanography 62, 1955–1962. doi: 10.1002/lno.10545
Rojas-Jiménez K., Grossart H. P., Cordes E., Cortés J. (2020). Fungal communities in sediments along a depth gradient in the Eastern Tropical Pacific. Front. Microbiol. 11. doi: 10.3389/fmicb.2020.575207
Rojas-Jiménez K., Hernández M. (2015). Isolation of fungi and bacteria associated with the guts of tropical wood-feeding Coleoptera and determination of their lignocellulolytic activities. Int. J. Microbiol. 2015, 1–11. doi: 10.1155/2015/285018
Román S., Ortiz-Álvarez R., Romano C., Casamayor E. O., Martin D. (2019). Microbial community structure and functionality in the deep sea floor: Evaluating the causes of spatial heterogeneity in a submarine canyon system (NW Mediterranean, Spain). Front. Mar. Sci. 6. doi: 10.3389/fmars.2019.00108
Romeo O., Marchetta A., Giosa D., Giuffrè L., Urzì C., De Leo F. (2020). Whole genome sequencing and comparative genome analysis of the halotolerant deep sea black yeast Hortaea werneckii. Life 10, 229. doi: 10.3390/life10100229
Rusch D. B., Halpern A. L., Sutton G., Heidelberg K. B., Williamson S., Yooseph S., et al. (2007). The sorcerer II global ocean sampling expedition: Northwest Atlantic through Eastern Tropical Pacific. PloS Biol. 5, e77. doi: 10.1371/journal.pbio.0050077
Sahling H., Masson D. G., Ranero C. R., Hühnerbach V., Weinrebe W., Klaucke I., et al. (2008). Fluid seepage at the continental margin offshore Costa Rica and southern Nicaragua. Geochemistry Geophysics Geosystems 9, 1–22. doi: 10.1029/2008GC001978
Sandi J., Mata-Araya I., Aguilar F. (2020). Diversity of lipase-producing microorganisms from tropical oilseeds Elaeis guineensis, Ricinus communis, and Jatropha curcas L. from Costa Rica. Curr. Microbiol. 77, 943–952. doi: 10.1007/s00284-020-01886-8
Sharma A., Arya S. K., Singh J., Kapoor B., Bhatti J. S., Suttee A., et al. (2023). Prospects of chitinase in sustainable farming and modern biotechnology: An update on recent progress and challenges. Biotechnol. Genet. Eng. Rev. 40, 310–340. doi: 10.1080/02648725.2023.2183593
Simão F. A., Waterhouse R. M., Ioannidis P., Kriventseva E. V., Zdobnov E. M. (2015). BUSCO: Assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212. doi: 10.1093/bioinformatics/btv351
Singh P., Raghukumar C., Meena R. M., Verma P., Shouche Y. (2012). Fungal diversity in deepsea sediments revealed by culture-dependent and culture-independent approaches. Fungal Ecol. 5, 543–553. doi: 10.1016/j.funeco.2012.01.001
Su P., Lu Z., Tian W., Chen Y., Maharachchikumbura S. S. (2023). Six additions to the genus Periconia (Dothideomycetes: Periconiaceae) from graminaceous plants in China. J. Fungi 9, 300. doi: 10.3390/jof9030300
Thavasi R., Subramanyam Nambaru V. R., Jayalakshmi S., Balasubramanian T., Banat I. M. (2011). Biosurfactant production by Pseudomonas aeruginosa from renewable resources. Indian J. Microbiol. 51, 30–36. doi: 10.1007/s12088-011-0076-7
Tsudome M., Tachioka M., Miyazaki M., Uchimura K., Tsuda M., Takaki Y., et al. (2022). An ultrasensitive nanofiber-based assay for enzymatic hydrolysis and deepsea microbial degradation of cellulose. iScience 25. doi: 10.1016/j.isci.2022.104732
Vurture M. R., Sedlazeck K. S., Fang H., Genovese C. N., Gurtowski J., Phillippy A. M., et al. (2017). GenomeScope: Fast reference-free genome profiling from short reads. Bioinformatics 33, 2202–2204. doi: 10.1093/bioinformatics/btx153
Wang Z.-P., Liu Z.-Z., Wang Y.-L., Bi W.-H., Liu L., Wang H.-Y., et al. (2019). Fungal community analysis in seawater of the Mariana Trench as estimated by Illumina HiSeq. RSC Adv. 9, 6956–6964. doi: 10.1039/C8RA10142F
Xu W., Gao Y., Gong L., Li M., Pang K.-L., Luo Z.-H. (2019). Fungal diversity in the deepsea hadal sediments of the Yap Trench by cultivation and high throughput sequencing methods based on ITS rRNA gene. Deep Sea Res. Part I: Oceanographic Res. Papers 145, 125–136. doi: 10.1016/j.dsr.2019.02.001
Xu W., Luo Z.-H., Guo S., Pang K.-L. (2016). Fungal community analysis in the deepsea sediments of the Pacific Ocean assessed by comparison of ITS, 18S, and 28S ribosomal DNA regions. Deep Sea Res. Part I: Oceanographic Res. Papers 109, 51–60. doi: 10.1016/j.dsr.2016.01.001
Xu F., Shin W., Brown S. H., Wahleithner J. A., Sundaram U. M., Solomon E. I. (1996). A study of a series of recombinant fungal laccases and bilirubin oxidase that exhibit significant differences in redox potential, substrate specificity, and stability. Biochim. Biophys. Acta (BBA) - Protein Structure Mol. Enzymology 1292, 303–311. doi: 10.1016/0167-4838(95)00210-3
Xu Z., Wang M., Wu W., Li Y., Liu Q., Han Y., et al. (2018). Vertical distribution of microbial eukaryotes from surface to the hadal zone of the Mariana Trench. Front. Microbiol. 9. doi: 10.3389/fmicb.2018.02023
Zhang X. Y., Tang G. L., Xu X. Y., Nong X. H., Qi S. H. (2014). Insights into deepsea sediment fungal communities from the East Indian Ocean using targeted environmental sequencing combined with traditional cultivation. PloS One 9, e109118. doi: 10.1371/journal.pone.0109118
Keywords: marine fungi, Hortaea, Periconia, biotechnological potential, eastern tropical Pacific
Citation: Rodríguez Ramírez I, Solano-González S, Cortés J and Rojas-Jiménez K (2025) Deepsea fungi of the eastern tropical Pacific of Costa Rica: morphological, genetic, and enzymatic characterization. Front. Mar. Sci. 12:1514874. doi: 10.3389/fmars.2025.1514874
Received: 21 October 2024; Accepted: 18 March 2025;
Published: 03 April 2025.
Edited by:
Mark Seasat Calabon, University of the Philippines Visayas, PhilippinesReviewed by:
Kandikere Ramaiah Sridhar, Mangalore University, IndiaCopyright © 2025 Rodríguez Ramírez, Solano-González, Cortés and Rojas-Jiménez. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Keilor Rojas-Jiménez, a2VpbG9yLnJvamFzQHVjci5hYy5jcg==
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.