
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
REVIEW article
Front. Immunol. , 17 March 2025
Sec. Mucosal Immunity
Volume 16 - 2025 | https://doi.org/10.3389/fimmu.2025.1563450
This article is part of the Research Topic Host-Microbiota Immuno-Interactions for Personalized Microbial Therapeutics View all 5 articles
 Wenwen Ding1,2†
Wenwen Ding1,2† Yiwen Cheng3†
Yiwen Cheng3† Xia Liu4†
Xia Liu4† Zhangcheng Zhu5
Zhangcheng Zhu5 Lingbin Wu6
Lingbin Wu6 Jie Gao3
Jie Gao3 Wenhui Lei7
Wenhui Lei7 Yating Li3
Yating Li3 Xin Zhou8,9,10,11
Xin Zhou8,9,10,11 Jian Wu12*
Jian Wu12* Yongtao Gao1,2*
Yongtao Gao1,2* Zongxin Ling3*†
Zongxin Ling3*† Ruilai Jiang6*
Ruilai Jiang6*Antimicrobial resistance (AMR) has become a major and escalating global health threat, undermining the effectiveness of current antibiotic and antimicrobial therapies. The rise of multidrug-resistant bacteria has led to increasingly difficult-to-treat infections, resulting in higher morbidity, mortality, and healthcare costs. Tackling this crisis requires the development of novel antimicrobial agents, optimization of current therapeutic strategies, and global initiatives in infection surveillance and control. Recent studies highlight the crucial role of the human gut microbiota in defending against AMR pathogens. A balanced microbiota protects the body through mechanisms such as colonization resistance, positioning it as a key ally in the fight against AMR. In contrast, gut dysbiosis disrupts this defense, thereby facilitating the persistence, colonization, and dissemination of resistant pathogens. This review will explore how gut microbiota influence drug-resistant bacterial infections, its involvement in various types of AMR-related infections, and the potential for novel microbiota-targeted therapies, such as fecal microbiota transplantation, prebiotics, probiotics, phage therapy. Elucidating the interactions between gut microbiota and AMR pathogens will provide critical insights for developing novel therapeutic strategies to prevent and treat AMR infections. While previous reviews have focused on the general impact of the microbiota on human health, this review will specifically look at the latest research on the interactions between the gut microbiota and the evolution and spread of AMR, highlighting potential therapeutic strategies.
Antimicrobial resistance (AMR) is a global health crisis, largely driven by the overuse and misuse of antibiotics. This has led to a significant rise in drug-resistant infections, contributing to millions of deaths each year (1, 2). Alarmingly, projections suggest that by 2050, AMR could be responsible for 10 million deaths annually (3, 4). The situation is made worse by the slow pace at which new antibiotics are developed and the growing presence of ESKAPE pathogens—Enterococcus faecium, Staphylococcus aureus, Klebsiella pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa, and Enterobacter species—which are leading contributors to AMR-related mortality (2, 5). These pathogens, although typically harmless and existing in a symbiotic relationship with the host, can become opportunistic in the event of immune system disruptions. This leads to infections that result from disturbances in the gut microbiota. Alterations in microbial composition, changes in bacterial metabolic activity, and shifts in local bacterial distribution further exacerbate gut dysbiosis, increasing the risk of infections. Furthermore, the slow pace of new antimicrobial drug development has failed to keep up with the rapid rise in AMR. This growing gap between rising resistance and limited new antibiotics highlights the urgent need for innovative strategies to manage infections and combat AMR.
As a response to this urgent challenge, researchers are increasingly turning to the human gut microbiota as a promising ally in the fight against AMR. The gut microbiota plays a vital role in supporting the immune system and protecting the body from infections (6–8). It helps regulate immune responses and maintains a delicate balance of microorganisms. However, disruptions to this balance—often caused by antibiotics or illness—can allow harmful, antibiotic-resistant pathogens to overgrow (5). Recent studies highlight how microbiota-based therapies, such as fecal microbiota transplantation (FMT) and probiotics, can help restore a healthy microbial balance and fight off drug-resistant infections. For example, Kellogg et al. find that succinate-producing microbiota drive tuft cell hyperplasia to protect against Clostridioides difficile (9). FMT has shown promise in treating recurrent C. difficile infections by outcompeting harmful bacteria and re-establishing a balanced microbiota, with approval from the U.S. FDA (10). Beyond FMT, other microbiota-based strategies, including prebiotics, synbiotics, postbiotics, and bacteriophage, are also being explored as potential weapons in the fight against AMR. These therapies work in various ways, such as fostering the growth of beneficial microbes, enhancing immune function, and directly inhibiting the growth of harmful bacteria.
This review will delve into the molecular mechanisms through which the gut microbiota influences AMR. It will examine the role of metabolites produced by gut bacteria, immune modulation, and competitive inhibition in shaping the body’s response to infections. Moreover, we will explore the diversity of the gut microbiota in different AMR bacterial infections and assess the potential of microbiota-targeting therapies as a promising approach to combatting AMR.
The human microbiota has established a mutualistic symbiosis with its host, contributing to overall health through metabolic support, immune modulation, and protection against pathogens (11, 12). A key mechanism by which the gut microbiota combats multidrug-resistant (MDR) infections is colonization resistance (CR), which prevents the colonization and overgrowth of both external pathogens and resident pathobionts (13). The gut microbiota employs a multifaceted defense strategy to protect the host, including nutritional competition, niche exclusion, contact-dependent inhibition, and the production of antimicrobial peptides and inhibitory metabolites (Figure 1). Furthermore, the microbiota contributes to mucosal barrier integrity, creates oxygen-limited environments that are inhospitable to many pathogens, and modulates immune responses to enhance immune tolerance and protection (13–15). Collectively, these mechanisms form a robust defense system that restricts the establishment and proliferation of harmful microorganisms, thereby safeguarding gut health and preventing MDR infections. A deeper understanding of how the gut microbiota limits pathogen colonization can inform the development of innovative therapeutic strategies to address the growing challenge of AMR. Harnessing the potential of the gut microbiota to enhance CR offers a promising avenue to combat MDR infections and mitigate the global threat of AMR.
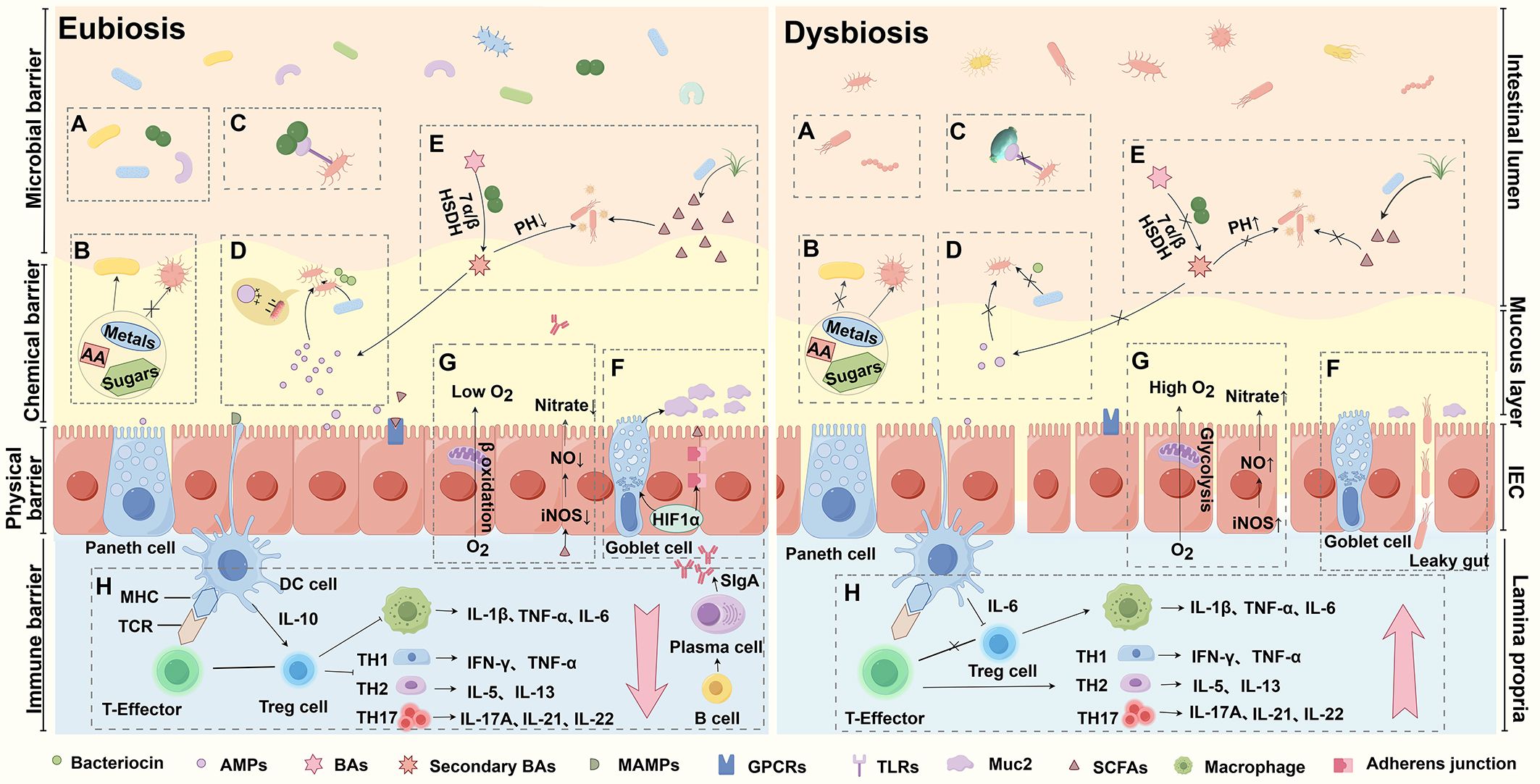
Figure 1. Mechanisms of gut microbiota in combating MDR infections. (A) Niche Exclusion: Dysbiosis reduces microbiota diversity, weakening pathogen exclusion. (B) Nutritional Competition: Healthy microbiota outcompetes pathogens for nutrients, thereby limiting the growth of pathogens by outcompeting them for resources. (C) Contact-Dependent Inhibition (CDI): Bacteria deliver toxins via cell contact, inhibiting or killing neighboring cells. (D) Antimicrobial Peptides (AMPs): Short-chain fatty acids (SCFAs) induce AMP synthesis, regulating microbiota and exerting antimicrobial effects. (E) Inhibitory Metabolites: SCFAs and secondary bile acids (BAs) inhibit pathogens or modulate immunity. (F) Mucosal Barrier: Dysbiosis reduces Muc2 production, compromising the intestinal barrier and increasing infection risk. (G) Oxygen Limitation: Inflammation, driven by iNOS and NADPH oxidase, limits oxygen availability during infection. (H) Immune Modulation: Microbiota activates Toll-like receptors (TLRs), crucial for immune signaling and defense.
The gut microbiota plays a critical role in preventing drug-resistant bacterial infections through nutritional competition, a mechanism that revolves around the depletion of essential resources required for bacterial growth. By efficiently consuming nutrients such as carbon sources, nitrogen sources, and metal ions, the gut microbiota limits their availability to potential pathogens, thereby restricting their growth and colonization. A diverse microbial community enhances this competitive edge, as highlighted by Spragge et al., who demonstrated that a rich microbial ecosystem intensifies nutrient competition, effectively outcompeting harmful pathogens (16). For example, the commensal bacterium Klebsiella michiganensis has been shown to outcompete Escherichia coli in vitro by depleting essential nutrients, limiting E. coli’s survival and growth (17). Similarly, commensal strains of E. coli, such as HS and Nissle 1917, inhibit the colonization of pathogenic E. coli O157:H7 by consuming sugars critical for the pathogen’s survival (18, 19). Genetic studies have further illuminated the role of nutrient competition in CR. For example, the gut microbiota’s uptake of dietary amino acids, such as tryptophan and arginine, plays a crucial role in preventing infections by pathogens like Citrobacter rodentium. However, a high-protein diet can enhance C. rodentium colonization in mice, underscoring the influence of nutrient availability on microbial dynamics (20). This balance is also evident in the case of C. difficile, where the gut microbiota competes for essential amino acids, restricting the pathogen’s proliferation (21). Interestingly, C. difficile attempts to counteract this by upregulating indole, a tryptophan metabolite, to promote its growth. Nevertheless, the microbiota’s efficient nutrient utilization limits the pathogen’s ability to exploit this strategy, further preventing its overgrowth (22). In addition to organic nutrients, metal ions such as iron, zinc, and manganese are critical for microbial survival and virulence. These metals are often scarce in the gut and are further sequestered by the host during inflammatory responses to limit pathogen access. Commensal bacteria, such as E. coli strain Nissle 1917, employ specialized mechanisms to acquire iron, providing protection against Salmonella infections (23). Conversely, pathogens like Vibrio cholerae and Campylobacter jejuni rely on efficient zinc uptake to survive and thrive in the gastrointestinal tract (24). By outcompeting pathogens for these vital nutrients, the gut microbiota establishes a robust defense mechanism, highlighting the critical role of nutritional competition in preventing infections and maintaining gut health. This understanding offers valuable insights into developing strategies to enhance CR and combat the growing threat of multidrug-resistant infections.
Niche exclusion is a critical strategy employed by the gut microbiota to limit pathogen colonization by occupying physical niches, depleting essential nutrients, or producing inhibitory metabolites. These competitive interactions are essential for maintaining gut homeostasis and preventing infections. Microbial species, particularly those genetically similar, often engage in direct competition. For instance, K. pneumoniae, Citrobacter, and Clostridium species inhibit the colonization of more pathogenic relatives through competitive exclusion (25–27). The “nutrient ecotope” hypothesis, introduced by Freter et al., provides a framework for understanding microbial competition in the gut. It posits that the availability of specific nutrients shapes the abundance and distribution of microbial species, with gut microbes actively consuming resources to define their niche (28). For example, Munehiro et al. isolated a symbiotic bacterial complex from healthy human fecal samples that inhibited Enterobacteriaceae, particularly Klebsiella, by regulating the availability of gluconates, a key nutrient for bacterial growth (19). Beyond nutrient competition, microbial species can coexist by acquiring novel traits. Sweeney et al. demonstrated that two closely related E. coli strains coexisted in the same niche by acquiring a high-affinity gluconate transporter, enabling them to outcompete other bacteria for the same carbon source (29). This highlights how adaptive resource uptake can shift competitive dynamics. Adhesion to mucosal surfaces is another crucial aspect of niche exclusion, as it prevents the colonization of exogenous pathogens. Kasper et al. showed that the IgA response facilitates Bacteroides fragilis in occupying a stable mucosal niche, where it inhibits harmful bacteria (30). This underscores the interplay between microbial communities and host immune responses in mediating niche occupancy. Additionally, some microbial species modify their environment to enhance colonization while excluding competitors (30, 31). For example, symbiotic bacteria limit Salmonella typhimurium colonization by occupying mucosal space and depleting nutrients like carbon sources and oxygen (32). Collectively, niche exclusion mechanisms—through resource competition, adhesion site occupation, and environmental modification—form a robust defense system that maintains microbial homeostasis and protects the host from infections. These insights highlight the intricate competitive dynamics within the gut microbiota and their role in preventing pathogen colonization.
Contact-dependent inhibition (CDI) is a sophisticated bacterial defense mechanism that enables bacteria to outcompete rivals through direct cell-to-cell interactions. This system involves the delivery of toxic effectors into neighboring cells, leading to growth inhibition or cell death. Initially discovered in E. coli strain EC93 in 2005, the CDI locus was found to contain three genes—CdiB, CdiA, and CdiI—which are sufficient to confer CDI activity to E. coli K-12 (33). Since then, CDI systems have been identified in various bacteria, including Enterobacteriaceae, Moraxellaceae, Pasteurellaceae, Neisseria meningitidis, Yersinia pestis, Dickeya dadantii, Enterobacter cloacae and Photorhabdus luminescens (34–37). The CDI system is centered on the CdiA protein, which acts as the effector by disrupting cellular processes, such as DNA degradation, in target cells (38). CdiB facilitates the transport of CdiA across the bacterial membrane, while CdiI provides immunity to the CDI+ bacterium by protecting it from its own toxins (39). Upon contact, the CdiB/CdiA two-partner secretion system delivers toxins directly into neighboring cells, inhibiting their growth or causing cell death (40). This mechanism allows bacteria to outcompete rivals in shared environments, such as during infection or biofilm formation (41). Consequently, the CDI system is increasingly viewed as a bacterial “weapon” system used to dominate microbial ecosystems (42, 43).
Another critical antibacterial mechanism is the type VI secretion system (T6SS), employed by many Gram-negative pathogens to inject toxic proteins into neighboring cells (44–47). 6SS-mediated interactions play a vital role in interbacterial competition, particularly among Bacteroides species in the gastrointestinal tract, where they compete for space and resources (48, 49). For example, B. fragilis inhibits B. polymorphicus growth in vitro through a T6SS-dependent mechanism (45) and exhibits competitive resistance against pathogenic strains (50, 51). These findings highlight the T6SS’s role in maintaining gut homeostasis and protecting against pathogenic invasion. Other secretion systems, such as Type IV and Type VII, also mediate antibacterial interactions through protein translocation and cell killing (52, 53), highlighting the importance of antagonistic interactions in the gut. Collectively, these systems demonstrate how bacteria have evolved complex strategies to shape microbial ecosystems and promote their survival. The CDI and T6SS systems represent potential targets for novel antimicrobial therapies aimed at disrupting toxin-delivery pathways, offering promising avenues to combat MDR infections and restore microbial balance in the gut (Table 1).
Antimicrobial peptides (AMPs) are essential components of the innate immune system, produced by host epithelial cells, neutrophils, and other immune cells such as mast cells and Paneth cells (54, 55). These peptides exhibit bactericidal, anti-inflammatory, and anti-endotoxin properties, serving as a first line of defense against microbial threats (56). AMPs disrupt microbial membranes and inhibit pathogen growth through diverse mechanisms. For example, nisin, produced by Lactococcus lactis, targets lipid II (a cell wall precursor) in pathogenic bacteria, forming pores in their membranes, causing cell content leakage, and disrupting membrane integrity (57). Similarly, RegIIIβ and RegIIIγ contribute to gut defense by interacting with the G protein-coupled receptor GPR43 (58, 59). AMPs also inhibit bacterial growth through specific molecular interactions. Microcin J25 targets E. coli RNA polymerase, suppressing bacterial replication (60), while colicins, produced by E. coli, degrade tRNA or disrupt proton gradients to inhibit pathogenic metabolism (61). Beyond their direct antimicrobial effects, AMPs regulate the gut microbiota, preventing the overgrowth of opportunistic pathogens and supporting immune homeostasis (62). For instance, oral administration of AMPs has been shown to alleviate intestinal inflammation induced by Enterohemorrhagic E. coli and modulate the gut microbiota (63). Bacteriocins, a subclass of AMPs produced by bacteria, selectively target closely related species to inhibit their growth or induce cell death, promoting competitive exclusion (64–66). For example, nisin is effective against Gram-positive bacteria like S. aureus and Listeria monocytogenes (67, 68), while class II bacteriocins such as Actifencin and Bacteroidetocin A selectively inhibit Lactobacillus and Bacteroides species, respectively, preserving microbiota diversity (69). Bacteriocins also enhance the resistance of beneficial bacteria, such as E. coli Nissle, against harmful microbes (70, 71). This specificity within microbial communities helps maintain balance and offers therapeutic potential, such as protecting against L. monocytogenes infections via strains like L. salivarius UCC 118 (72, 73). Therefore, AMPs play a dual role in defending against infections and shaping the gut microbial landscape. Their ability to modulate microbial abundance and diversity is critical in preventing pathogen overgrowth and maintaining gut homeostasis (74). These properties highlight the potential of AMPs as therapeutic agents to combat infections and restore microbial balance.
The gut microbiota produces a variety of inhibitory metabolites, including short-chain fatty acids (SCFAs) and secondary bile acids (BAs), which play critical roles in directly inhibiting pathogen growth and modulating host immune responses to maintain gut health (75, 76). SCFAs, primarily derived from the fermentation of dietary fibers by the gut microbiota (75), exhibit direct antimicrobial effects by disrupting bacterial membrane integrity and intracellular pH homeostasis. This mechanism inhibits pathogens such as E. coli and Salmonella under acidic conditions (77). Beyond their antimicrobial activity, SCFAs exert immunomodulatory effects, including promoting the differentiation of regulatory T cells (Tregs), which help mitigate excessive inflammation (78). For example, butyrate increases Foxp3+ Tregs in the spleen and lymph nodes of antibiotic-treated mice (79), while propionate suppresses IL-17 production in intestinal γδT cells (80). SCFAs also enhance the intestinal epithelial barrier by promoting the integrity and function of epithelial cells, thereby strengthening mucosal immunity and preventing pathogen adhesion (81, 82). Additionally, SCFAs provide energy to intestinal epithelial cells, supporting normal cellular respiration and preventing the establishment of pathogens like Citrobacter (83). A key mechanism by which SCFAs maintain gut homeostasis is through the regulation of hypoxia-inducible factor 1-alpha (HIF-1α). Butyrate stabilizes HIF-1α by inhibiting histone deacetylases (HDACs) and activating G-protein-coupled receptors such as GPR43 and GPR109A. This process upregulates tight junction proteins like occludin and claudin-1, enhancing intestinal barrier function and reducing permeability (84). HIF-1α activation also promotes the secretion of β-defensins by intestinal epithelial cells, inhibiting the overgrowth of E. coli and Salmonella while supporting beneficial bacteria like Lactobacillus and Bifidobacterium (85).
Secondary BAs, such as deoxycholic acid (DCA) and lithocholic acid (LCA), are produced by specific gut microbiota species through the metabolism of primary BAs. These metabolites exhibit antimicrobial properties, inhibiting pathogens like C. difficile (86, 87) and Salmonella while selectively promoting beneficial bacteria (88, 89). DCA exerts bactericidal effects by lowering intracellular pH and disrupting bacterial membrane integrity (90, 91), while LCA indirectly enhances gut defense by stimulating the transcription of antimicrobial peptides like LL-37 (92). Secondary BAs also modulate immune responses, with certain Bacteroidetes species promoting Treg differentiation through BA modification, further supporting immune homeostasis (93, 94). Disruptions to the gut microbiota, such as those caused by antibiotics or dietary changes, can impair the production of these protective metabolites, increasing susceptibility to infections and pathogen overgrowth (95). Therefore, maintaining a balanced gut microbiota is essential for ensuring the continuous production of inhibitory metabolites that limit pathogen growth and support immune homeostasis. These insights highlight the therapeutic potential of modulating SCFAs and secondary BAs to combat infections and promote gut health.
The gut mucosal barrier is a complex, multi-layered defense system that integrates physical, microbiological, immune, and chemical components to protect the gastrointestinal tract from pathogens. The intestinal epithelial barrier, including its mucus layer, serves as a physical shield against harmful microorganisms (96). Pathogens like E. coli can attach to epithelial cells to initiate infections, but commensal microorganisms strengthen the mucosal barrier to prevent this. Mucin 2 (Muc2), a key component of the mucus layer, is central to maintaining gut barrier integrity. Germ-free mice, which lack a normal gut microbiota, produce less Muc2 and are more susceptible to infections (97, 98). HIF-1α, a transcription factor responsive to low oxygen conditions, plays a critical role in regulating Muc2 secretion by goblet cells. This forms a protective mucus layer that prevents direct pathogen contact with epithelial cells (99). E. coli can also stimulate Muc2 production, highlighting the microbiota’s role in host defense. Muc2-deficient mice are more vulnerable to pathogen colonization and experience severe infections, such as those caused by L. monocytogenes (100). Diet significantly influences mucosal barrier integrity. Mice fed a fiber-free diet develop a thinner mucus layer, increasing susceptibility to pathogens like L. citrobacter (101, 102). Additionally, L-fucose, a sugar produced by gut bacteria, reduces the virulence of pathogens like Burkholderia citriodora, further demonstrating the microbiota’s role in modulating immune responses and pathogen defense (103). HIF-1α also regulates tight junctions between intestinal epithelial cells, enhancing barrier function. It upregulates tight junction proteins such as occludin, claudin-1, and ZO-1, preventing the penetration of pathogens and toxins (84). Disruption of these junctions by pathogens like C. difficile, which produces toxins TcdA and TcdB, increases intestinal permeability and facilitates pathogen invasion (104, 105). This underscores the critical role of the gut microbiota in maintaining epithelial barrier function. The gut microbiota further supports the mucosal barrier by producing metabolites that enhance immune function and inhibit pathogenic microbes (76). HIF-1α promotes energy metabolism in intestinal epithelial cells under hypoxic conditions, upregulating genes like GLUT1 and LDHA to reinforce barrier integrity (84). In summary, the mucosal barrier relies on a combination of mechanisms, including Muc2 secretion, tight junction regulation, and energy metabolism, to maintain gut homeostasis and protect against infections. These processes highlight the intricate interplay between the gut microbiota and host defenses in preserving intestinal health.
The gut is characterized by oxygen-limited conditions, with oxygen levels sharply decreasing in the deeper layers of the mucosal surface. This anaerobic environment is critical in shaping the composition and function of the gut microbiota. Pathogens that thrive in higher oxygen environments, such as E. coli, Salmonella, and C. difficile, struggle to establish themselves in this setting, while commensal bacteria, well-adapted to low oxygen, outcompete pathogens for space and nutrients (13, 106). Oxygen-limited conditions promote the growth of microbial species that produce SCFAs, which support mucosal immunity and strengthen the gut barrier. However, during infections, the oxygen balance can shift, creating a more favorable environment for pathogens. Infections trigger an inflammatory response, primarily through the activation of inducible nitric oxide synthase (iNOS) and NADPH oxidase. iNOS, encoded by the Nos2 gene, produces nitric oxide by oxidizing L-arginine, while NADPH oxidase generates reactive oxygen species (ROS), contributing to inflammation (15, 107). Pathogens such as S. typhimurium and E. coli can induce inflammation and dysbiosis, increasing oxygen and nitrate availability in the intestinal lumen. These molecules serve as terminal electron acceptors for pathogenic bacteria, promoting their survival and growth (108, 109). Commensal bacteria play a vital role in maintaining the gut’s hypoxic state, which limits pathogen expansion. For example, commensals that produce butyrate via β-oxidation reduce oxygen levels by stimulating aerobic respiration in intestinal cells (13), creating an oxygen-poor environment that hinders pathogen growth. However, in conditions of dysbiosis, where butyrate production is diminished, oxygen levels rise, supporting the growth of pathogens like S. typhimurium (110, 111). Inflammation and dysbiosis can further reduce butyrate-producing bacteria, exacerbating oxygen levels and promoting the growth of pathogenic anaerobes (112, 113). Therefore, oxygen-limited conditions are essential for maintaining a healthy gut microbiota and providing CR against pathogens.
The gut is a central hub of the immune system, housing approximately 70–80% of the body’s immune cells (114). It acts as a critical interface between the body and external pathogens, with the gut microbiota playing a pivotal role in maintaining immune homeostasis. This intricate interplay between the microbiota and the mucosal immune system is essential for both immune tolerance and defense mechanisms. The gut microbiota influences the development, maturation, and function of the immune system, impacting both innate and adaptive immunity (114). It educates the immune system to distinguish between harmful pathogens and harmless substances, such as food and beneficial microbes. A balanced microbiota promotes a tolerogenic immune response, preventing excessive activation that could lead to chronic inflammation or autoimmune diseases. One key mechanism by which the gut microbiota interacts with the immune system is through the activation of pattern recognition receptors (PRRs), such as Toll-like receptors (TLRs) (115). These receptors trigger the release of antimicrobial peptides, cytokines, and chemokines, initiating inflammatory responses to combat infections (116, 117). Additionally, the microbiota stimulates the production of IgA, an antibody that neutralizes pathogens, controls harmful bacterial colonization, and supports beneficial microbes (118). The absence of secretory IgT, an ancient immunoglobulin involved in mucosal immunity, disrupts the microbiota in species like rainbow trout, making them more susceptible to infections (119). This highlights the evolutionary importance of immune responses in maintaining gut microbiota balance. The gut microbiota also plays a crucial role in the differentiation of Tregs, which suppress excessive immune reactions and prevent inflammation (120). Conversely, it can stimulate the production of Th17 cells, which defend against extracellular pathogens but, when overactivated, contribute to inflammatory conditions. A balanced microbiota maintains the equilibrium between Tregs (anti-inflammatory) and Th17 cells (pro-inflammatory), supporting gut homeostasis. Beneficial microbes produce SCFAs, such as butyrate, propionate, and acetate, which possess anti-inflammatory properties and enhance Treg function (121, 122). SCFAs also contribute to immune regulation by promoting the survival of memory T cells and maintaining gut barrier integrity, preventing “leaky gut” and systemic infections (78). Overall, the gut microbiota is essential for immune homeostasis. Dysbiosis can increase susceptibility to pathogen colonization, infections, and inflammatory or autoimmune disorders.
The escalating global threat of AMR, fueled by the widespread misuse and overuse of antimicrobials, has underscored the urgent need for innovative solutions. Multidrug-resistant (MDR) pathogens, including C. difficile and the ESKAPE (ESKAPCE) group, pose significant challenges to modern healthcare systems (123). These pathogens exhibit resistance to conventional antibiotics, leading to treatment failures, recurrent infections, prolonged hospitalizations, and increased mortality rates. The scarcity of effective therapeutic options exacerbates this crisis (124). Emerging evidence highlights the pivotal role of the gut microbiota in modulating the spread and progression of AMR (124). The gut microbiota, a complex microbial ecosystem, can either facilitate or impede the colonization of drug-resistant pathogens. Understanding the intricate interactions between the gut microbiota and MDR bacteria is critical for developing novel strategies to combat resistant infections (Figure 2, Table 2).
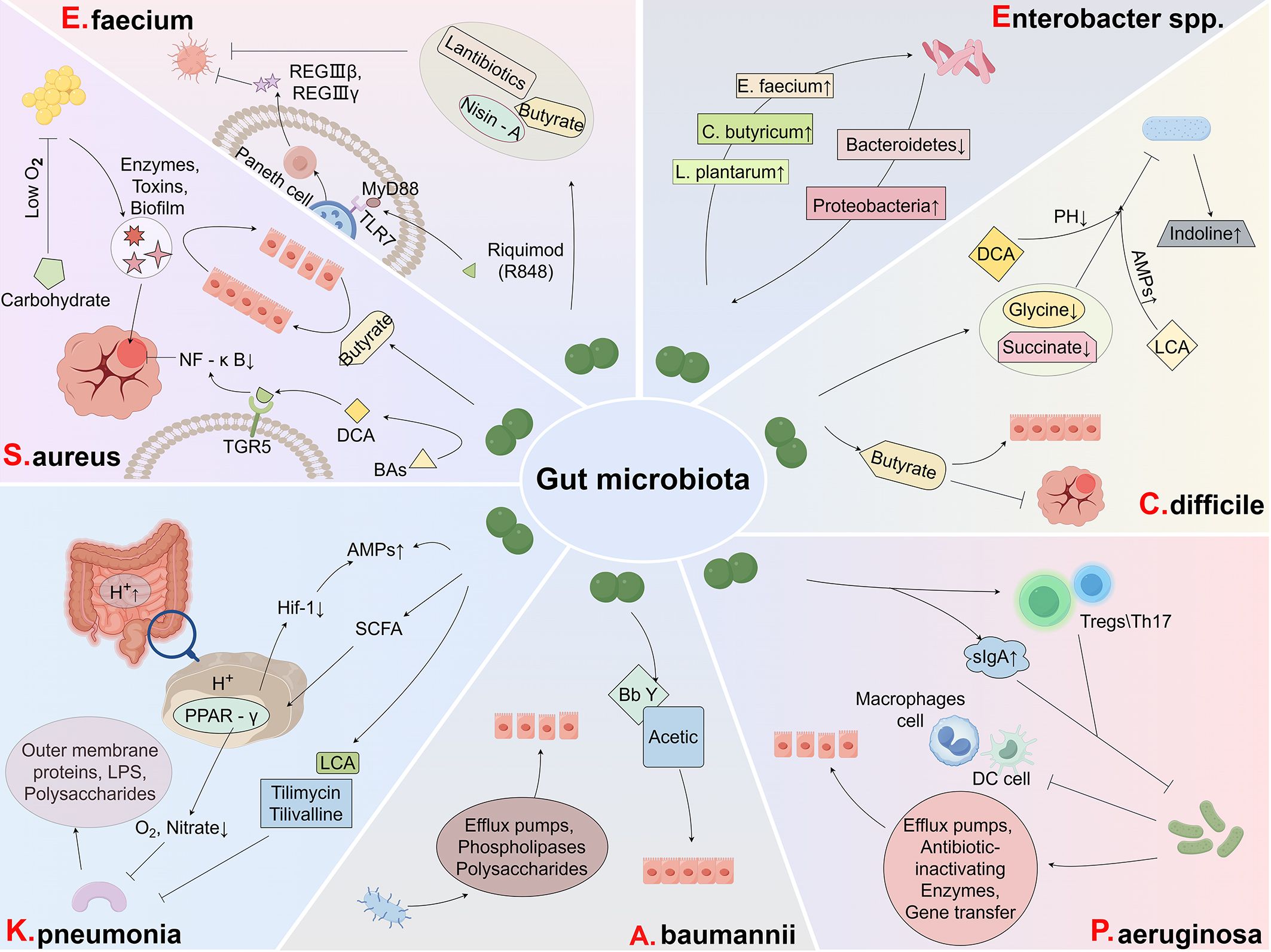
Figure 2. Roles of gut microbiota in different types of MDR infections. Enterococcus faecium: The gut microbiota resists E. faecium by secreting antibacterial substances that inhibit its growth. Staphylococcus aureus: The gut microbiota counters S. aureus by restoring the mucosal barrier and suppressing inflammation, thereby limiting bacterial colonization. Klebsiella pneumoniae: The release of beneficial metabolites and induction of intracellular acidification by the gut microbiota inhibit the growth of K. pneumoniae, curbing its spread. Acinetobacter baumannii: Restoration of the mucosal barrier by the gut microbiota prevents the invasion and colonization of A. baumannii, enhancing host defense. Pseudomonas aeruginosa: Through immune modulation, the gut microbiota strengthens the host’s immune response, effectively suppressing P. aeruginosa infections. Clostridium difficile: The gut microbiota mitigates C. difficile infections by reducing inflammation, regulating microbial metabolites, and restoring mucosal barrier integrity. Enterobacter spp.: A diverse gut microbiota inhibits Enterobacter species, contributing to the prevention of infections and maintaining microbial balance.
E. faecium, a Gram-positive facultative anaerobe, is a commensal member of the human gut microbiota, playing a role in digestion and homeostasis (149). However, under antibiotic pressure, it can transform into a pathogenic organism, particularly in immunocompromised or hospitalized individuals (150). Antibiotic-resistant strains, such as vancomycin-resistant E. faecium (VRE), pose significant clinical challenges, causing infections like urinary tract infections (UTIs), bacteremia, and endocarditis. The acquisition of resistance genes through horizontal gene transfer exacerbates treatment difficulties, leading to increased morbidity, prolonged hospital stays, and higher healthcare costs.
Studies have demonstrated that reduced expression of antimicrobial peptides RegIIIβ and RegIIIγ enhances susceptibility to VRE infections (151). Antibiotic-induced disruption of the gut microbiota diminishes RegIIIγ production, impairing VRE control. However, stimulation with the TLR7 ligand riquimod (R848) has been shown to restore RegIIIγ levels, facilitating VRE clearance in antibiotic-treated mice (152). Additionally, Kim et al. identified a four-strain Blautia producta consortium that restores resistance to VRE post-antibiotic treatment by producing a lantibiotic similar to nisin-A, which inhibits VRE growth (125). High lantibiotic gene abundance in at-risk patients correlates with reduced E. faecium levels, and lantibiotic-producing strains prevent VRE colonization in germ-free mice, highlighting their potential as probiotics. Further research has revealed that L. murinus Y74 and L. plantarum HT121 reduce VRE colonization and restore microbiota diversity in infected mice (126). Barnesiella spp. have also been effective in eliminating VRE colonization and improving survival by reshaping the gut microbiota (153). Butyrate-producing bacteria contribute to microbiota restoration and VRE suppression (127). FMT has emerged as a promising intervention, with studies reporting successful VRE decolonization in numerous patients (154–156). Moreover, Enterococcus species can exacerbate C. difficile pathogenicity by altering gut metabolism and supporting its growth through amino acids such as leucine and ornithine (157). Thus, modulating the gut microbiota and its metabolites presents a promising strategy for preventing and treating E. faecium infections.
S. aureus remains a leading cause of severe, life-threatening infections (158, 159), with methicillin-resistant S. aureus (MRSA) posing a particularly significant threat. MRSA is associated with high morbidity and mortality rates, especially among hospitalized adults (2, 160). MRSA frequently colonizes the gut, particularly in patients exposed to antibiotics or with critical illnesses, disrupting the normal gut microbiota. Its virulence factors—including enzymes, toxins, and biofilm formation—facilitate tissue invasion, induce inflammation, and impair immune responses, exacerbating infection severity and complicating treatment (161).
MRSA’s ecological adaptations and nutritional competitiveness enhance its ability to colonize the gut and initiate infections. Genetic mutations and structural changes enable MRSA to efficiently metabolize carbohydrates, synthesize its cell wall, and produce energy under low-oxygen conditions, allowing it to thrive in the gut environment. However, the presence of beneficial bacteria can counteract its growth (162). Probiotics have demonstrated potential in reducing S. aureus enterotoxin production by altering the gut environment without significantly affecting bacterial growth. For instance, S. lugdunensis produces lugdunin, which directly inhibits S. aureus growth (128, 163). In MRSA-infected mice, a decline in butyrate-producing bacteria correlates with reduced butyrate levels in the gut and bloodstream. Butyrate supplementation has been shown to restore gut mucosal integrity and enhance immune function, offering a promising therapeutic strategy (129). Additionally, Clostridium scindens converts primary bile acids (BAs) to secondary BAs, such as DCA, which may enhance the antimicrobial properties of cell membranes (164, 165). Co-culturing C. scindens with colonic cells improves cell viability and strengthens the gut barrier, mitigating damage from S. aureus infections (130). DCA has also been shown to reduce S. aureus-induced mastitis in mice by activating the TGR5 receptor, which suppresses inflammatory pathways like NF-κB and NLRP3 (131). Dysbiosis impairs DCA production and TGR5 activation, worsening MRSA infections. However, restoring the microbiota with beneficial bacteria, such as C. scindens or segmented filamentous bacteria, improves infection outcomes (166). Furthermore, targeting the microbiota through genetic modulation, such as CYP1A1 knockdown, reduces harmful metabolites like cadaverine, offering protection against MRSA-induced sepsis (167). This evidence highlights the crucial role of the gut microbiota in both preventing and treating MRSA infections, offering new therapeutic possibilities beyond antibiotics.
K. pneumoniae is a highly virulent, antibiotic-resistant Gram-negative bacterium that causes severe infections, particularly in immunocompromised individuals (168). Its pathogenicity is driven by several factors: outer membrane proteins that facilitate adhesion and immune evasion, lipopolysaccharides that induce inflammation and septic shock, and polysaccharides that prevent engulfment by immune cells. These mechanisms complicate infection management and contribute to its persistence. The K. oxytoca species complex, a component of the human microbiome, produces enterotoxins such as tilimycin and tilivalline and plays a role in antibiotic resistance (169). Studies by Osbelt et al. have shown that certain K. oxytoca strains can reduce gut colonization by MDR K. pneumoniae in antibiotic-treated and gnotobiotic mouse models. This effect is largely attributed to competition for carbohydrates, such as beta-glucosides (133), which are critical for promoting resistance. These findings suggest that K. oxytoca strains may serve as next-generation probiotics to decolonize K. pneumoniae and protect against infections (27). Further research by Shen et al. demonstrated that high concentrations of LCA inhibit K. pneumoniae growth and reduce its adhesion to Caco-2 cells (134). Sorbara et al. highlighted the role of the gut microbiota in suppressing MDR K. pneumoniae by acidifying the proximal colon, which triggers SCFA-mediated intracellular acidification (77). This process activates PPAR-γ in host epithelial cells, reducing oxygen and nitrate availability, impairing the pathogen’s respiration, and stabilizing HIF-1, which promotes antimicrobial peptide synthesis (110, 170). A recent study revealed that the gut microbiota exposed to K. pneumoniae produces sulfide via the taurine pathway, decreasing host cell respiration and preventing pathogen invasion (135). These findings highlight the crucial role of the microbiota in combating K. pneumoniae and point to potential therapeutic strategies.
A. baumannii is a formidable opportunistic pathogen, particularly in hospital settings, where its ability to acquire multidrug resistance (MDR) poses a significant threat. It is a leading cause of infections in critically ill patients, including urinary tract infections, bloodstream infections, and ventilator-associated pneumonia, contributing to increased morbidity and mortality (171). While community-acquired infections are less frequently MDR, they can still result in severe outcomes (172).
A. baumannii typically colonizes the upper respiratory tract and skin, harboring a wide array of genes that confer MDR, such as those encoding carbapenemases and broad-spectrum β-lactamases. Carbapenem-resistant A. baumannii (CRAB) is particularly problematic in intensive care units (ICUs), where it is strongly associated with ventilator-associated pneumonia (171). The CRAB genome is equipped with numerous resistance genes and virulence factors, including efflux pumps, iron acquisition systems, secretion systems, phospholipases, and polysaccharides, which enhance its survival and colonization capabilities (173). CRAB infections are linked to prolonged ICU stays, elevated healthcare costs, and increased antibiotic use (174). The limited treatment options for widespread MDR strains prompted the World Health Organization (WHO) to classify A. baumannii as a “Priority Pathogen” in 2018, emphasizing the urgent need for novel antibiotics (175). Antibiotic use increases the risk of A. baumannii colonization and infection, likely due to the disruption of commensal bacteria that compete for ecological niches (176). Modulating the gut microbiota may offer a promising strategy to restore microbial balance and limit A. baumannii infections. For example, Asahara et al. demonstrated that continuous oral administration of B. breve strain Yakult (BbY) improved survival rates and inhibited CRAB growth in infected mice. BbY also helped mitigate intestinal environmental disruptions and maintained barrier function (137). Additionally, a positive correlation between CRAB levels and acetic acid production suggests that gut microbiota metabolites play a critical role in preserving intestinal barrier integrity (177). These findings highlight the potential of probiotic interventions to reduce A. baumannii infections, improve patient outcomes, and decrease reliance on antibiotics.
P. aeruginosa is a Gram-negative opportunistic pathogen responsible for healthcare-associated infections (HAIs), including chronic obstructive pulmonary disease (COPD), cystic fibrosis, cancer, trauma, burns, sepsis, and ventilator-associated pneumonia (178, 179). It is a leading cause of HAIs globally, and the emergence of antimicrobial-resistant P. aeruginosa has led the WHO to classify it as a critical priority pathogen (180). This bacterium develops multidrug resistance (MDR) through mechanisms such as altered outer membrane permeability, efflux pump activity, production of antibiotic-inactivating enzymes, and horizontal gene transfer, making infections increasingly difficult to treat (123, 181). Carbapenem antibiotics, such as imipenem and meropenem, are commonly used against P. aeruginosa, but the rise of carbapenem-resistant P. aeruginosa (CRPA) poses a significant threat (182). CRPA infections are associated with nearly 700,000 deaths annually, with resistance rates in Europe reaching 12.9% (183), underscoring the urgent need for novel therapeutic strategies.
A key factor in P. aeruginosa infections is intestinal colonization prior to ICU admission. Gomez-Zorrilla et al. found that pre-admission intestinal colonization by P. aeruginosa increases the risk of subsequent infection by nearly 15-fold (184). Disruption of the gut microbiota, which normally provides colonization resistance through antimicrobial compounds, facilitates pathogen invasion. Gut dysbiosis can impair host immune function, disrupt the intestinal barrier, and increase susceptibility to infections. The gut microbiota plays a critical role in immune modulation, with secretory IgA (sIgA) being a key component of mucosal immunity. sIgA, produced at mucosal surfaces such as the intestines and lungs, protects against pathogens like P. aeruginosa (185, 186). However, antimicrobial treatment can reduce IgA levels in the lungs, increasing the risk of respiratory infections (187). Research has shown that certain gut bacteria, such as segmented filamentous bacteria, enhance IgA production and protect against P. aeruginosa infection (188). Asymptomatic colonization of the gut by carbapenemase-producing Enterobacterales can lead to dysbiosis, exacerbating the severity of P. aeruginosa lung infections by reducing immune cells such as alveolar macrophages and conventional dendritic cells, which are critical for fighting respiratory infections (95). Microbial ecosystem treatment (MET-2), aimed at restoring a healthy gut microbiota, has shown promise in reducing P. aeruginosa load and promoting the growth of beneficial bacteria (189). Additionally, the gut-lung axis further illustrates the influence of gut microbiota on lung immunity. Th17 cells, a subset of T-helper cells, play a vital role in protecting the lungs from P. aeruginosa infections (190). Wen et al. demonstrated that transplanting host intestinal commensal bacteria restores the balance of Tregs and Th17 cells, enhancing their metabolic functions and providing protection against P. aeruginosa pneumonia (138). Thus, restoring gut eubiosis offers promising strategies for treating P. aeruginosa infections, potentially improving patient outcomes.
C. difficile is a spore-forming, toxin-producing anaerobic bacterium and a leading cause of hospital-acquired diarrhea and pseudomembranous colitis (191). It poses a significant healthcare burden, with over 500,000 infections and approximately 30,000 deaths annually, costing over $1.5 billion each year (192). The primary risk factor is antibiotic use, which disrupts the gut microbiota, promoting C. difficile spore germination and increasing infection risk (193, 194). While most patients respond to initial antibiotic treatments, up to 25% experience recurrence, with over 60% of relapsed patients suffering further episodes (195). This recurrent nature complicates management, as antibiotics further disrupt the microbiota, increasing relapse risk.
The gut microbiota plays a central role in regulating C. difficile infections through ecological competition and nutrient availability. Non-toxigenic C. difficile strains can inhibit toxigenic strains by depleting essential metabolites like glycine (25). Similarly, Caulobacter spp. inhibit C. difficile growth by reducing luminal succinate, a key metabolite for its proliferation (141). SCFAs, particularly butyrate, particularly butyrate, are critical in protecting against C. difficile infection. Butyrate strengthens the intestinal mucosal barrier and reduces inflammation (142, 196). A deficiency in butyrate has been linked to poor outcomes in FMT for recurrent infections (143). BAs, regulated by the gut microbiota, also influence C. difficile spore germination and resistance. Farnesoid X receptor (FXR) agonists like obeticholic acid reduce infection severity, while ursodeoxycholic acid (UCA) enhances immune responses against C. difficile (144, 197). These findings highlight the potential of BA modulation in managing infections. Emerging therapies targeting the microbiota show promise. Bacteriocins from Bacillus thuringiensis and AMPs from Lactobacillus species inhibit C. difficile growth and spore germination (198, 199). Additionally, Enterococcus species, often abundant in C. difficile patients, worsen outcomes by promoting toxin production through inflammatory effects (157). These findings suggest that modulation of the microbiota, either through direct interventions such as probiotics or through indirect approaches like BAs manipulation, could offer promising new strategies for preventing and treating C. difficile infections.
The genus Enterobacter, comprising facultative anaerobic Gram-negative bacilli within the Enterobacteriaceae family (200), includes the E. cloacae complex (ECC), a significant cause of nosocomial infections, particularly in healthcare settings (123). Carbapenem-resistant Enterobacterales (CRE) are especially concerning due to their resistance to broad-spectrum antibiotics, including carbapenems. Risk factors for CRE infections include prior CRE colonization, broad-spectrum antibiotic use, ICU admission, mechanical ventilation, prolonged hospital stays, and indwelling catheters (201). CRE infections are associated with higher in-hospital mortality and longer hospital stays compared to carbapenem-susceptible strains, highlighting the urgent need for effective interventions.
The gut microbiota plays a pivotal role in the development and persistence of Enterobacter infections. Studies show that CRE-positive individuals exhibit distinct microbiota profiles, characterized by reduced diversity and increased dominance of Enterococcus, Sphingomonas, and Staphylococcus (202). CRE carriers also show elevated Proteobacteria and reduced Bacteroidetes levels compared to non-carriers (203). These shifts may facilitate CRE establishment and persistence in the gut. Broad-spectrum antibiotics, commonly used to treat infections, disrupt CR, enabling CRE expansion. Antibiotics deplete microbial metabolites that suppress CRE growth while enriching nutrients that support CRE proliferation, perpetuating dysbiosis and infection spread (147). Promisingly, certain probiotics and metabolites have shown potential in mitigating CRE colonization. Supernatants from C. butyricum, E. faecium, and L. plantarum suppress CRE growth in a dose-dependent manner, while those from B. fragilis and B. longum are less effective (204). Furthermore, SCFAs also differentially inhibit Enterobacteriaceae species, suggesting microbial metabolites play a key role in modulating CRE growth (148). These findings underscore the critical role of the gut microbiota in combating Enterobacter infections. The potential for microbiota-based therapies, such as probiotics or the restoration of microbial diversity, open an exciting avenue for future research. As our understanding of the gut’s role in resisting Enterobacter spp. and other pathogens deepens, such therapies could play a key role in reducing the burden of CRE infections and improving patient outcomes.
Advances in microbiome research, powered by omics technologies such as genomics, metagenomics, and metabolomics, have significantly enhanced our understanding of host-microbiota interactions (205). These insights highlight the potential of modulating the gut microbiota to combat MDR infections. By altering the composition and activity of the microbiota, it is possible to reduce the colonization and spread of drug-resistant pathogens, restore microbial balance, and improve gut health. Emerging strategies, including probiotics, prebiotics, synbiotics, postbiotics, FMT, and bacteriophages, are gaining traction as promising tools to address MDR infections (206–208) (Figure 3, Table 3).
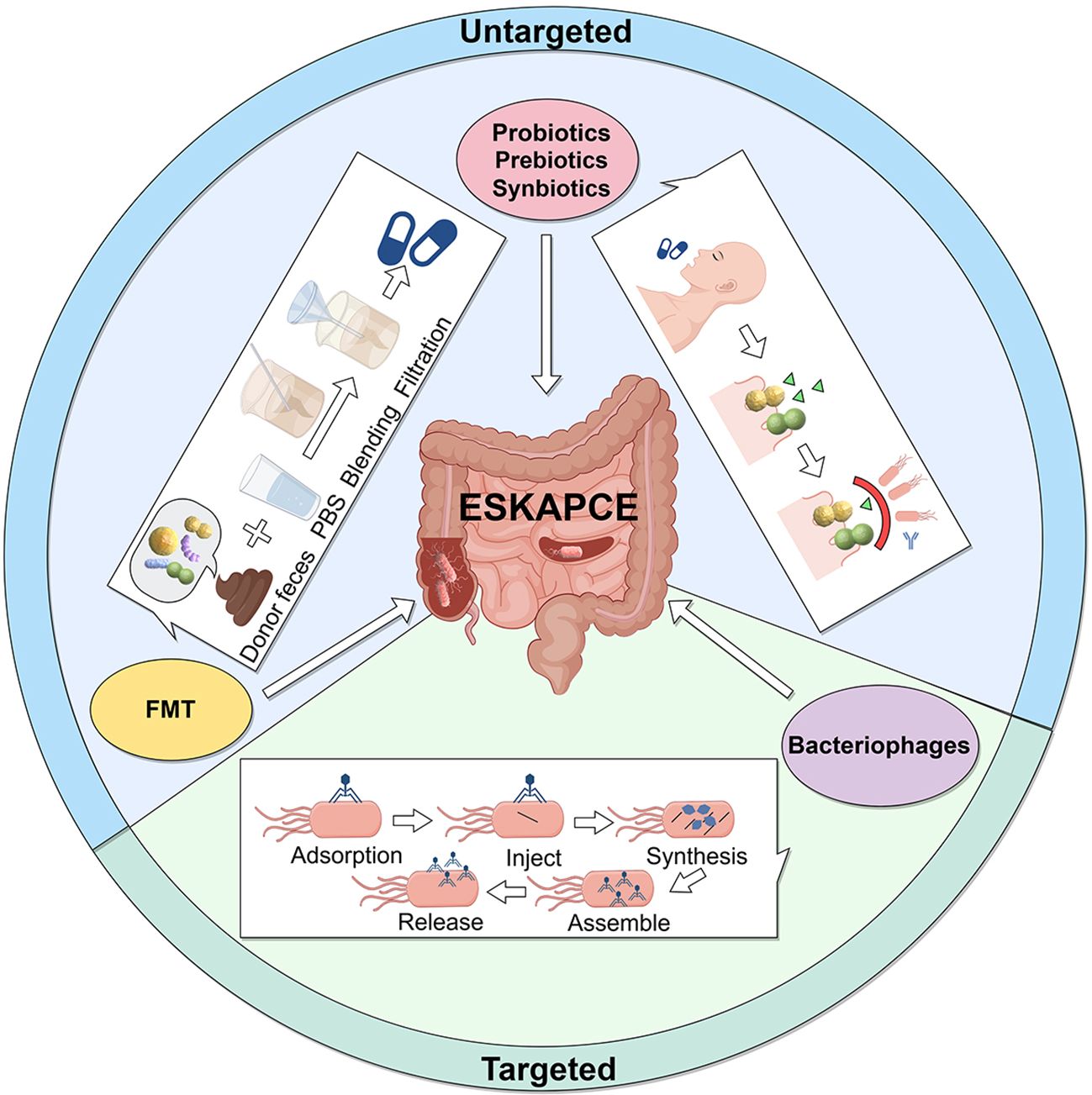
Figure 3. Gut microbiota-based therapies for MDR infections. Probiotics and Prebiotics: Administered in appropriate doses, they prevent pathogen colonization by competing for resources and binding sites, maintaining gut equilibrium, and enhancing immune responses. Fecal Microbiota Transplantation (FMT): Feces from healthy donors is processed and transplanted into patients to restore a balanced gut microbiota. Bacteriophages: Phages kill bacteria through a five-stage process: adsorption, injection, synthesis, assembly, and release, offering a targeted approach to treat bacterial infections.
Probiotics are live microorganisms that, when administered in adequate amounts, confer health benefits to the host by restoring or maintaining a healthy gut microbiota (212, 213). Common probiotic strains, such as Lactobacillus, Bifidobacterium, and Saccharomyces, exert their effects through the production of antimicrobial substances (e.g., lactic acid, hydrogen peroxide), nutrient competition, and immune modulation (214). These mechanisms reduce infection risk, making probiotics valuable in both clinical and dietary approaches to health promotion and infection prevention.
Lactobacilli species, such as L. plantarum and L. rhamnosus, exhibit antimicrobial activity against pathogens like critical role in infection prevention, with strains like MRSA (215) and VRE (156, 216). These strains enhance immune responses and promote the production of beneficial SCFAs (217, 218). For example, L. plantarum increases butyrate-producing bacteria, reduces proinflammatory cytokines, and strengthens intestinal barriers, lowering infection risks (219). Additionally, L. rhamnosus produces lactomodulin, which has bactericidal effects on resistant pathogens like MRSA and VRE (132). Certain Lactobacillus strains also alleviate antibiotic-associated diarrhea and C. difficile infections (124, 220, 221). Bifidobacterium species, particularly B. longum, improve intestinal barrier function and regulate immune responses, suppressing pathogen growth. For example, B. longum JDM301 inhibits toxigenic C. difficile growth (145). Probiotic combinations, such as S. boulardii with Lactobacillus and Bifidobacterium, effectively prevent the colonization of MDR pathogens like extended-spectrum beta-lactamase (ESBL)-producing bacteria (222). Pre-treatment with combinations like LactoLevure® (L. plantarum, L. acidophilus, S. boulardii and B. lactis) improves survival in rodent models infected with MDR P. aeruginosa (223). C. butyricum, a butyrate-producing bacterium, plays a key role in gut health by promoting beneficial bacteria and inhibiting pathogens (143). For example, C. butyricum 588 enhances antibacterial efficacy against C. difficile through immune modulation and reinforcement of gut mucosal barriers (146). Despite their potential, the effectiveness of probiotics varies depending on factors such as strain, dosage, and individual microbiota composition. Challenges in their broader application include unclear molecular mechanisms, strain-specific effects, antibiotic resistance, and stability issues. Metabiotic components may offer solutions to these limitations, advancing the field of microbiota-based therapies.
Prebiotics are non-digestible dietary compounds that promote the growth of beneficial microorganisms, enhancing gut health (224). By supporting beneficial bacteria like Bifidobacterium and Lactobacillus, prebiotics help restore microbial balance and suppress MDR pathogens (209). Common prebiotics include human milk oligosaccharides (HMOs), inulin, fructooligosaccharides (FOS), galactooligosaccharides (GOS), and dietary fibers like β-glucan, pectin, and resistant starch (55). These compounds maintain a diverse microbiota, inhibiting pathogen growth through competitive exclusion (210). For instance, GOS reduces E. coli adhesion by 70% in vitro (211, 225), while fucoidan, a marine prebiotic, promotes Bacteroides growth and eliminates P. aeruginosa in mice (139). Additionally, 1,5-anhydro-d-fructose (1,5-AF), derived from starch and glycogen, exhibits antioxidant and antibacterial properties, reduces cytokine production, and boosts F. prausnitzii growth and NAD biosynthesis genes (226).
Synbiotics combine prebiotics and probiotics to synergistically enhance the survival and colonization of beneficial microbes, improving gut health (224, 227). This approach enhances the efficacy of both components: probiotics utilize prebiotics as growth substrates, while prebiotics are more effectively utilized by targeted microbes. For instance, pre-cultivating L. plantarum with xylitol reduces C. difficile spore germination and decreases mortality in infected mice from 44% to 22% (228). Similarly, combining Lactobacillus with prebiotics like lactulose or isomalto-oligosaccharides prevents colonization by KPC-2-producing K. pneumoniae (136). In vitro, synbiotics show superior antibacterial activity against MDR pathogens like A. baumannii and E. faecalis, outperforming probiotics alone (137, 229).
Postbiotics, derived from nonviable bacteria or their metabolic by-products, include bacterial components (e.g., cell walls, enzymes, SCFAs, vitamins, peptides) and paraprobiotics (e.g., peptidoglycan, surface proteins) (213, 230). These components provide immunological benefits, with bacteriocins and AMPs disrupting bacterial membranes. For example, Nisin, produced by Lactobacillus, inhibits cell wall formation and enhances penetration when complexed with nanoparticles (231–233). SCFAs from fiber fermentation and secondary BAs inhibit C. difficile growth and boost immune defense (234, 235). Lactic acid bacteria-derived cell-free supernatants also inhibit biofilm formation, offering a potential strategy against MDR pathogens like P. aeruginosa, S. aureus, and E. coli (140).
Together, prebiotics, synbiotics, and postbiotics offer a powerful approach to managing MDR infections and restoring gut health. Prebiotics nourish beneficial microbes, synbiotics combine probiotics and prebiotics for synergistic effects, and postbiotics provide antimicrobial and anti-inflammatory benefits. These strategies collectively suppress harmful bacteria, enhance immune responses, and maintain gut microbial balance.
FMT is emerging as a promising strategy to combat MDR bacterial infections. It involves transferring gut microbiota from a healthy donor to a recipient to restore a balanced, functional microbial community. This restoration enhances CR to MDR pathogens through mechanisms like competitive inhibition, bacteriocin production, and immune modulation (206, 236). FMT has demonstrated significant success in treating recurrent C. difficile infections and severe gut dysbiosis (237, 238), with a meta-analysis reporting a 90% success rate for recurrent C. difficile infections (239). The FDA-approved FMT product Rebyota (RBX2660) has shown a 78.9% therapeutic success rate in this patient group (240). The success of FMT in C. difficile infections has spurred interest in its application to other MDR pathogens, including VRE, MRSA, and carbapenemase-producing Enterobacter spp (241, 242). FMT restores immune functions and enhances CR, improving pathogen clearance. For example, FMT has reversed lethal sepsis by restoring butyrate-producing Bacteroidetes (243) and eradicated VRE in both animal models and human patients (153, 244, 245). It has also successfully cleared NDM-1 K. pneumoniae and other CREs (246). Additionally, FMT has treated nosocomial MRSA enteritis (247) and recurrent UTIs caused by ESBL-producing K. pneumoniae (248). FMT holds considerable promise for managing MDR infections and restoring gut health, particularly in patients with recurrent infections and gut dysbiosis. By improving the gut microbiota, FMT modulates the immune system, enhances pathogen clearance, and provides systemic benefits. However, challenges remain (249, 250), including identifying optimal donor profiles (251), standardizing stool processing and administration protocols, addressing long-term safety concerns, and understanding the mechanisms behind FMT’s efficacy. Clear regulatory guidelines and ethical donor selection are also essential. Overcoming these challenges will enhance FMT’s effectiveness and safety, solidifying its role as a key therapeutic strategy in modern medicine.
Bacteriophages (phages) are viruses that specifically infect and lyse bacterial cells, offering a targeted alternative to traditional antibiotics (252, 253). Their specificity minimizes disruption to beneficial microbiota, as they often target only certain bacterial strains (254). After eradicating their bacterial hosts, phages naturally die off, reducing accumulation and toxicity risks. Unlike antibiotics, phages do not affect human cells, avoiding harm to healthy tissue. Additionally, bacterial resistance to phages develops more slowly than to antibiotics, making phage therapy a promising tool against MDR infections.
Phage therapy has evolved into various forms to combat MDR bacteria. Personalized phage therapy isolates specific phages tailored to the infecting bacterial strain, offering customized treatment for MDR infections (255). Phage cocktails, combining multiple phages targeting the same pathogen, enhance efficacy and reduce resistance risks. Phage-driven antibiotics explore how phages can complement traditional antibiotics by weakening bacterial defenses or disrupting protective biofilms (256). Clinical trials and case studies highlight the potential of phage therapy against MDR infections caused by S. aureus, P. aeruginosa, and A. baumannii. For example, a six-phage combination (including E215, E217, PAK_P1, PYO2, DEV, and PAK_P4) targeting P. aeruginosa outperformed single-phage treatments (257, 258), while a four-phage mix achieved over 98% efficacy against S. aureus and reduced the minimum inhibitory concentrations (MICs) of antibiotics like vancomycin in MRSA biofilms (259, 260). This synergy between phages and antibiotics effectively eliminates persistent infections while preserving beneficial gut microbiota (261).
Phages can also support beneficial bacteria. For example, E. faecalis V583 carries a phage that eliminates competing strains (262). Phages like ΦCD27 reduce A. baumannii bacterial load without harming non-pathogenic bacteria (194, 263). Phage P3CHA inhibits P. aeruginosa biofilm formation in mice (264, 265), though repeated exposure can lead to biofilm resistance (266). A four-phage combination disrupts C. difficile biofilms in vitro (259, 267). Recent studies highlight phages targeting K. pneumoniae, including capsular mutants, showing enhanced lethality in combination therapies (268). Corbellino et al. successfully eradicated MDR K. pneumoniae using a custom lytic phage preparation (269). These findings underscore the potential of phage therapy as an effective tool for treating infections caused by MDR bacteria. However, there are still challenges to overcome. Immune responses and bacterial resistance can limit the effectiveness of phage therapy. The immunogenicity of phage capsid proteins may lead to rapid clearance, and bacterial resistance complicates treatment outcomes. Strategies to optimize phage combinations and delivery methods are crucial, and further research is needed to establish standardized protocols and evaluate the long-term safety of phage therapy. Despite these challenges, phage therapy remains a promising approach in the fight against antibiotic-resistant infections.
The gut microbiota plays a critical yet complex role in combating MDR bacterial infections, presenting both significant challenges and untapped potential. The resistance of MDR pathogens to multiple antibiotics limits the efficacy of conventional therapies, while the intricate variability of the microbiota complicates the development of targeted treatments. The widespread overuse of antibiotics disrupts the microbiome’s balance, promoting the spread of resistance genes and increasing infection risks. Additionally, our incomplete understanding of microbiota-host immune interactions hinders the creation of effective microbiota-based therapies. Despite these challenges, leveraging the gut microbiota has emerged as a pivotal strategy in addressing MDR infections and the growing threat of AMR. A range of therapeutic approaches—including probiotics, prebiotics, synbiotics, postbiotics, FMT, bacteriophage therapy, and CRISPR-Cas-based engineered strains—show promise in modulating the microbiota to enhance host resistance. These interventions aim to restore microbial balance, strengthen protective functions, and mitigate AMR effects.
However, the complexity of the microbiota and the need for precision medicine necessitate further research into personalized microbiota-based therapies and the development of novel antibacterial agents. Future studies should focus on tailoring interventions to individual microbiome profiles, ensuring the long-term safety of microbiota modulation, and integrating microbiome-based approaches with new antibacterial compounds. Advanced technologies, such as artificial intelligence and big data analytics, can enable more precise and effective treatments for MDR infections. Ultimately, overcoming AMR requires a collaborative, multidisciplinary approach to translate innovative strategies into scalable, safe, and practical solutions. By harnessing the full potential of the gut microbiota, we can pave the way for sustainable and effective approaches to managing resistant infections, improving public health outcomes, and addressing the global antibiotic resistance crisis.
WD: Data curation, Investigation, Methodology, Writing – original draft, Writing – review & editing. YC: Funding acquisition, Investigation, Methodology, Validation, Writing – original draft, Writing – review & editing. XL: Data curation, Investigation, Methodology, Validation, Writing – original draft, Writing – review & editing. ZZ: Data curation, Investigation, Methodology, Writing – original draft. LW: Data curation, Investigation, Methodology, Writing – original draft. JG: Data curation, Investigation, Methodology, Writing – original draft. WL: Data curation, Formal Analysis, Methodology, Writing – original draft. YL: Data curation, Formal Analysis, Investigation, Methodology, Writing – original draft. XZ: Data curation, Formal Analysis, Investigation, Methodology, Writing – original draft, Conceptualization, Writing – review & editing. JW: Conceptualization, Data curation, Formal Analysis, Investigation, Methodology, Writing – original draft, Writing – review & editing. YG: Data curation, Formal Analysis, Investigation, Methodology, Writing – original draft, Writing – review & editing. ZL: Conceptualization, Funding acquisition, Methodology, Project administration, Supervision, Validation, Writing – original draft, Writing – review & editing. RJ: Conceptualization, Data curation, Funding acquisition, Investigation, Methodology, Writing – original draft, Writing – review & editing.
The author(s) declare that financial support was received for the research and/or publication of this article. This present work was funded by the grants of the National S&T Major Project of China (2023YFC2308400), the Jiangsu Provincial Research Hospital (YJXYY202204-YSB31), the Zhejiang Provincial Natural Science Foundation of China (LQ24H090005 and LQ22H030013), Shandong Provincial Laboratory Project (SYS202202), the Fundamental Research Funds for the Central Universities (2022ZFJH003), the Taishan Scholar Foundation of Shandong Province (tsqn202103119), and the Foundation of China’s State Key Laboratory for Diagnosis and Treatment of Infectious Diseases (ZZ202316 and ZZ202319).
Figures are drawn using Figdraw (https://www.figdraw.com).
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declared that they were an editorial board member of Frontiers, at the time of submission. This had no impact on the peer review process and the final decision.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
1. Naddaf M. 40 million deaths by 2050: toll of drug-resistant infections to rise by 70. Nature. (2024) 633:747–48. doi: 10.1038/d41586-024-03033-w
2. Antimicrobialresistancecollaborators. Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis. Lancet. (2022) 399:629–55. doi: 10.1016/s0140-6736(21)02724-0
3. Blackman LD, Sutherland TD, De Barro PJ, Thissen H, Locock KES. Addressing a future pandemic: how can non-biological complex drugs prepare us for antimicrobial resistance threats? Mater Horiz. (2022) 9:2076–96. doi: 10.1039/d2mh00254j
4. Gbd2021antimicrobialresistancecollaborators. Global burden of bacterial antimicrobial resistance 1990-2021: a systematic analysis with forecasts to 2050. Lancet. (2024) 404:1199–226. doi: 10.1016/s0140-6736(24)01867-1
5. Luo Q, Lu P, Chen Y, Shen P, Zheng B, Ji J, et al. ESKAPE in China: epidemiology and characteristics of antibiotic resistance. Emerg Microbes Infect. (2024) 13:2317915. doi: 10.1080/22221751.2024.2317915
6. Carvalho MJ, Sands K, Thomson K, Portal E, Mathias J, Milton R, et al. Antibiotic resistance genes in the gut microbiota of mothers and linked neonates with or without sepsis from low- and middle-income countries. Nat Microbiol. (2022) 7:1337–47. doi: 10.1038/s41564-022-01184-y
7. Zheng XQ, Wang DB, Jiang YR, Song CL. Gut microbiota and microbial metabolites for osteoporosis. Gut Microbes. (2025) 17:2437247. doi: 10.1080/19490976.2024.2437247
8. Song P, Peng Z, Guo X. Gut microbial metabolites in cancer therapy. Trends Endocrinol Metab. (2025) 36:55–69. doi: 10.1016/j.tem.2024.06.016
9. Kellogg TD, Ceglia S, Mortzfeld BM, Tanna TM, Zeamer AL, Mancini MR, et al. Succinate-producing microbiota drives tuft cell hyperplasia to protect against Clostridioides difficile. J Exp Med. (2025) 222:e20232055. doi: 10.1084/jem.20232055
10. Ghani R, Chrysostomou D, Roberts LA, Pandiaraja M, Marchesi JR, Mullish BH. Faecal (or intestinal) microbiota transplant: a tool for repairing the gut microbiome. Gut Microbes. (2024) 16:2423026. doi: 10.1080/19490976.2024.2423026
11. Pickard JM, Zeng MY, Caruso R, Núñez G. Gut microbiota: Role in pathogen colonization, immune responses, and inflammatory disease. Immunol Rev. (2017) 279:70–89. doi: 10.1111/imr.12567
12. Yang W, Cong Y. Gut microbiota-derived metabolites in the regulation of host immune responses and immune-related inflammatory diseases. Cell Mol Immunol. (2021) 18:866–77. doi: 10.1038/s41423-021-00661-4
13. Caballero-Flores G, Pickard JM, Núñez G. Microbiota-mediated colonization resistance: mechanisms and regulation. Nat Rev Microbiol. (2023) 21:347–60. doi: 10.1038/s41579-022-00833-7
14. Granato ET, Meiller-Legrand TA, Foster KR. The evolution and ecology of bacterial warfare. Curr Biol. (2019) 29:R521–r37. doi: 10.1016/j.cub.2019.04.024
15. Sorbara MT, Pamer EG. Interbacterial mechanisms of colonization resistance and the strategies pathogens use to overcome them. Mucosal Immunol. (2019) 12:1–9. doi: 10.1038/s41385-018-0053-0
16. Spragge F, Bakkeren E, Jahn MT, E BNA, Pearson CF, Wang X, et al. Microbiome diversity protects against pathogens by nutrient blocking. Science. (2023) 382:eadj3502. doi: 10.1126/science.adj3502
17. Oliveira RA, Ng KM, Correia MB, Cabral V, Shi H, Sonnenburg JL, et al. Klebsiella michiganensis transmission enhances resistance to Enterobacteriaceae gut invasion by nutrition competition. Nat Microbiol. (2020) 5:630–41. doi: 10.1038/s41564-019-0658-4
18. Maltby R, Leatham-Jensen MP, Gibson T, Cohen PS, Conway T. Nutritional basis for colonization resistance by human commensal Escherichia coli strains HS and Nissle 1917 against E. coli O157:H7 in the mouse intestine. PLoS One. (2013) 8:e53957. doi: 10.1371/journal.pone.0053957
19. Furuichi M, Kawaguchi T, Pust MM, Yasuma-Mitobe K, Plichta DR, Hasegawa N, et al. Commensal consortia decolonize Enterobacteriaceae via ecological control. Nature. (2024) 633:878–86. doi: 10.1038/s41586-024-07960-6
20. Caballero-Flores G, Pickard JM, Fukuda S, Inohara N, Núñez G. An enteric pathogen subverts colonization resistance by evading competition for amino acids in the gut. Cell Host Microbe. (2020) 28:526–33.e5. doi: 10.1016/j.chom.2020.06.018
21. Pereira FC, Wasmund K, Cobankovic I, Jehmlich N, Herbold CW, Lee KS, et al. Rational design of a microbial consortium of mucosal sugar utilizers reduces Clostridiodes difficile colonization. Nat Commun. (2020) 11:5104. doi: 10.1038/s41467-020-18928-1
22. Sinha AK, Laursen MF, Brinck JE, Rybtke ML, Hjørne AP, Procházková N, et al. Dietary fibre directs microbial tryptophan metabolism via metabolic interactions in the gut microbiota. Nat Microbiol. (2024) 9:1964–78. doi: 10.1038/s41564-024-01737-3
23. Behnsen J, Zhi H, Aron AT, Subramanian V, Santus W, Lee MH, et al. Siderophore-mediated zinc acquisition enhances enterobacterial colonization of the inflamed gut. Nat Commun. (2021) 12:7016. doi: 10.1038/s41467-021-27297-2
24. Gielda LM, Dirita VJ. Zinc competition among the intestinal microbiota. mBio. (2012) 3:e00171–12. doi: 10.1128/mBio.00171-12
25. Leslie JL, Jenior ML, Vendrov KC, Standke AK, Barron MR, O’brien TJ, et al. Protection from Lethal Clostridioides difficile Infection via Intraspecies Competition for Cogerminant. mBio. (2021) 12:e00522-21. doi: 10.1128/mBio.00522-21
26. Mullineaux-Sanders C, Carson D, Hopkins EGD, Glegola-Madejska I, Escobar-Zepeda A, Browne HP, et al. Citrobacter amalonaticus Inhibits the Growth of Citrobacter rodentium in the Gut Lumen. mBio. (2021) 12:e0241021. doi: 10.1128/mBio.02410-21
27. Osbelt L, Wende M, Almási É, Derksen E, Muthukumarasamy U, Lesker TR, et al. Klebsiella oxytoca causes colonization resistance against multidrug-resistant K. pneumoniae in the gut via cooperative carbohydrate competition. Cell Host Microbe. (2021) 29:1663–79.e7. doi: 10.1016/j.chom.2021.09.003
28. Freter R, Brickner H, Botney M, Cleven D, Aranki A. Mechanisms that control bacterial populations in continuous-flow culture models of mouse large intestinal flora. Infect Immun. (1983) 39:676–85. doi: 10.1128/iai.39.2.676-685.1983
29. Sweeney NJ, Klemm P, Mccormick BA, Moller-Nielsen E, Utley M, Schembri MA, et al. The Escherichia coli K-12 gntP gene allows E. coli F-18 to occupy a distinct nutritional niche in the streptomycin-treated mouse large intestine. Infect Immun. (1996) 64:3497–503. doi: 10.1128/iai.64.9.3497-3503.1996
30. Donaldson GP, Ladinsky MS, Yu KB, Sanders JG, Yoo BB, Chou WC, et al. Gut microbiota utilize immunoglobulin A for mucosal colonization. Science. (2018) 360:795–800. doi: 10.1126/science.aaq0926
31. Bry L, Falk PG, Midtvedt T. and Gordon, J.I. A model of host-microbial interactions in an open mammalian ecosystem. Science. (1996) 273:1380–3. doi: 10.1126/science.273.5280.1380
32. Kamada N, Kim YG, Sham HP, Vallance BA, Puente JL, Martens EC, et al. Regulated virulence controls the ability of a pathogen to compete with the gut microbiota. Science. (2012) 336:1325–9. doi: 10.1126/science.1222195
33. Aoki SK, Pamma R, Hernday AD, Bickham JE, Braaten BA, Low DA. Contact-dependent inhibition of growth in Escherichia coli. Science. (2005) 309:1245–8. doi: 10.1126/science.1115109
34. Aoki SK, Diner EJ, De Roodenbeke CT, Burgess BR, Poole SJ, Braaten BA, et al. A widespread family of polymorphic contact-dependent toxin delivery systems in bacteria. Nature. (2010) 468:439–42. doi: 10.1038/nature09490
35. Beck CM, Morse RP, Cunningham DA, Iniguez A, Low DA, Goulding CW, et al. CdiA from Enterobacter cloacae delivers a toxic ribosomal RNase into target bacteria. Structure. (2014) 22:707–18. doi: 10.1016/j.str.2014.02.012
36. Willett JL, Gucinski GC, Fatherree JP, Low DA, Hayes CS. Contact-dependent growth inhibition toxins exploit multiple independent cell-entry pathways. Proc Natl Acad Sci U S A. (2015) 112:11341–6. doi: 10.1073/pnas.1512124112
37. Webb JS, Nikolakakis KC, Willett JL, Aoki SK, Hayes CS, Low DA. Delivery of CdiA nuclease toxins into target cells during contact-dependent growth inhibition. PLoS One. (2013) 8:e57609. doi: 10.1371/journal.pone.0057609
38. Curley CL, Fedrigoni TP, Flaherty EM, Woodilla CJ, Hagan CL. Bacterial contact-Dependent inhibition protein binds near the open lateral gate in BamA prior to toxin translocation. Biochemistry. (2021) 60:2956–65. doi: 10.1021/acs.biochem.1c00337
39. Halvorsen TM, Schroeder KA, Jones AM, Hammarlöf D, Low DA, Koskiniemi S, et al. Contact-dependent growth inhibition (CDI) systems deploy a large family of polymorphic ionophoric toxins for inter-bacterial competition. PLoS Genet. (2024) 20:e1011494. doi: 10.1371/journal.pgen.1011494
40. Guerin J, Botos I, Zhang Z, Lundquist K, Gumbart JC, Buchanan SK. Structural insight into toxin secretion by contact-dependent growth inhibition transporters. Elife. (2020) 9:e58100. doi: 10.7554/eLife.58100
41. Ocasio AB, Cotter PA. CDI/CDS system-encoding genes of Burkholderia Thailandensis are located in a mobile genetic element that defines a new class of transposon. PLoS Genet. (2019) 15:e1007883. doi: 10.1371/journal.pgen.1007883
42. Hayes CS, Koskiniemi S, Ruhe ZC, Poole SJ, Low DA. Mechanisms and biological roles of contact-dependent growth inhibition systems. Cold Spring Harb Perspect Med. (2014) 4:a010025. doi: 10.1101/cshperspect.a010025
43. Boopathi S, Liu D, Jia AQ. Molecular trafficking between bacteria determines the shape of gut microbial community. Gut Microbes. (2021) 13:1959841. doi: 10.1080/19490976.2021.1959841
44. Coyne MJ, Roelofs KG, Comstock LE. Type VI secretion systems of human gut Bacteroidales segregate into three genetic architectures, two of which are contained on mobile genetic elements. BMC Genomics. (2016) 17:58. doi: 10.1186/s12864-016-2377-z
45. Russell AB, Wexler AG, Harding BN, Whitney JC, Bohn AJ, Goo YA, et al. A type VI secretion-related pathway in Bacteroidetes mediates interbacterial antagonism. Cell Host Microbe. (2014) 16:227–36. doi: 10.1016/j.chom.2014.07.007
46. Silverman JM, Brunet YR, Cascales E, Mougous JD. Structure and regulation of the type VI secretion system. Annu Rev Microbiol. (2012) 66:453–72. doi: 10.1146/annurev-micro-121809-151619
47. Motta EVS, Lariviere PJ, Jones KR, Song Y, Moran NA. Type VI secretion systems promote intraspecific competition and host interactions in a bee gut symbiont. Proc Natl Acad Sci U S A. (2024) 121:e2414882121. doi: 10.1073/pnas.2414882121
48. Chatzidaki-Livanis M, Geva-Zatorsky N, Comstock LE. Bacteroides fragilis type VI secretion systems use novel effector and immunity proteins to antagonize human gut Bacteroidales species. Proc Natl Acad Sci U S A. (2016) 113:3627–32. doi: 10.1073/pnas.1522510113
49. Coyne MJ, Comstock LE. Type VI secretion systems and the gut microbiota. Microbiol Spectr. (2019) 7:10.1128. doi: 10.1128/microbiolspec.PSIB-0009-2018
50. Hecht AL, Casterline BW, Earley ZM, Goo YA, Goodlett DR, Bubeck Wardenburg J. Strain competition restricts colonization of an enteric pathogen and prevents colitis. EMBO Rep. (2016) 17:1281–91. doi: 10.15252/embr.201642282
51. Cianciotto NP. The type II secretion system as an underappreciated and understudied mediator of interbacterial antagonism. Infect Immun. (2024) 92:e0020724. doi: 10.1128/iai.00207-24
52. Whitney JC, Peterson SB, Kim J, Pazos M, Verster AJ, Radey MC, et al. A broadly distributed toxin family mediates contact-dependent antagonism between gram-positive bacteria. Elife. (2017) 6:e26938. doi: 10.7554/eLife.26938
53. Job AM, Doran KS, Spencer BL. A group B streptococcal type VII-secreted LXG toxin mediates interbacterial competition and colonization of the murine female genital tract. mBio. (2024) 15:e0208824. doi: 10.1128/mbio.02088-24
54. Ji S, An F, Zhang T, Lou M, Guo J, Liu K, et al. Antimicrobial peptides: An alternative to traditional antibiotics. Eur J Med Chem. (2024) 265:116072. doi: 10.1016/j.ejmech.2023.116072
55. Wiertsema SP, Van Bergenhenegouwen J, Garssen J, Knippels LMJ. The interplay between the gut microbiome and the immune system in the context of infectious diseases throughout life and the role of nutrition in optimizing treatment strategies. Nutrients. (2021) 13:886. doi: 10.3390/nu13030886
56. Marissen J, Reichert L, Härtel C, Fortmann MI, Faust K, Msanga D, et al. Antimicrobial peptides (AMPs) and the microbiome in preterm infants: consequences and opportunities for future therapeutics. Int J Mol Sci. (2024) 25:6684. doi: 10.3390/ijms25126684
57. Cotter PD, Ross RP, Hill C. Bacteriocins - a viable alternative to antibiotics? Nat Rev Microbiol. (2013) 11:95–105. doi: 10.1038/nrmicro2937
58. Zhao Y, Chen F, Wu W, Sun M, Bilotta AJ, Yao S, et al. GPR43 mediates microbiota metabolite SCFA regulation of antimicrobial peptide expression in intestinal epithelial cells via activation of mTOR and STAT3. Mucosal Immunol. (2018) 11:752–62. doi: 10.1038/mi.2017.118
59. Moniri NH, Farah Q. Short-chain free-fatty acid G protein-coupled receptors in colon cancer. Biochem Pharmacol. (2021) 186:114483. doi: 10.1016/j.bcp.2021.114483
60. Nguyen HA, Dunham CM. Digging the tunnel for chemical space. Nat Chem Biol. (2017) 13:1061–62. doi: 10.1038/nchembio.2480
61. Cascales E, Buchanan SK, Duché D, Kleanthous C, Lloubès R, Postle K, et al. Colicin biology. Microbiol Mol Biol Rev. (2007) 71:158–229. doi: 10.1128/mmbr.00036-06
62. Luo Y, Song Y. Mechanism of antimicrobial peptides: antimicrobial, anti-inflammatory and antibiofilm activities. Int J Mol Sci. (2021) 22:11401. doi: 10.3390/ijms222111401
63. Zhao XQ, Wang L, Zhu CL, Xue XH, Xia XJ, Wu XL, et al. Oral administration of the antimicrobial peptide mastoparan X alleviates enterohemorrhagic Escherichia coli-induced intestinal inflammation and regulates the gut microbiota. Probiotics Antimicrob Proteins. (2024) 16:138–51. doi: 10.1007/s12602-022-10013-x
64. Heilbronner S, Krismer B, Brötz-Oesterhelt H, Peschel A. The microbiome-shaping roles of bacteriocins. Nat Rev Microbiol. (2021) 19:726–39. doi: 10.1038/s41579-021-00569-w
65. Peng Z, Wang D, He Y, Wei Z, Xie M, Xiong T. Gut distribution, impact factor, and action mechanism of bacteriocin-producing beneficial microbes as promising antimicrobial agents in gastrointestinal infection. Probiotics Antimicrob Proteins. (2024) 16:1516–27. doi: 10.1007/s12602-024-10222-6
66. Gu Q, Yan J, Lou Y, Zhang Z, Li Y, Zhu Z, et al. Bacteriocins: Curial guardians of gastrointestinal tract. Compr Rev Food Sci Food Saf. (2024) 23:e13292. doi: 10.1111/1541-4337.13292
67. Asaduzzaman SM, Sonomoto K. Lantibiotics: diverse activities and unique modes of action. J Biosci Bioeng. (2009) 107:475–87. doi: 10.1016/j.jbiosc.2009.01.003
68. Wu M, Ma Y, Dou X, Zohaib Aslam M, Liu Y, Xia X, et al. A review of potential antibacterial activities of nisin against Listeria monocytogenes: the combined use of nisin shows more advantages than single use. Food Res Int. (2023) 164:112363. doi: 10.1016/j.foodres.2022.112363
69. Elisa Heesemann Rosenkilde C, Olsen Lützhøft D, Vazquez-Uribe R, Otto Alexander Sommer M. Heterologous expression and antimicrobial potential of class II bacteriocins. Gut Microbes. (2024) 16:2369338. doi: 10.1080/19490976.2024.2369338
70. Sassone-Corsi M, Nuccio SP, Liu H, Hernandez D, Vu CT, Takahashi AA, et al. Microcins mediate competition among Enterobacteriaceae in the inflamed gut. Nature. (2016) 540:280–83. doi: 10.1038/nature20557
71. Cherrak Y, Salazar MA, Yilmaz K, Kreuzer M, Hardt WD. Commensal E. coli limits Salmonella gut invasion during inflammation by producing toxin-bound siderophores in a tonB-dependent manner. PLoS Biol. (2024) 22:e3002616. doi: 10.1371/journal.pbio.3002616
72. Corr SC, Li Y, Riedel CU, O’toole PW, Hill C, Gahan CG. Bacteriocin production as a mechanism for the antiinfective activity of Lactobacillus salivarius UCC118. Proc Natl Acad Sci U S A. (2007) 104:7617–21. doi: 10.1073/pnas.0700440104
73. Abramov VM, Kosarev IV, Machulin AV, Deryusheva EI, Priputnevich TV, Panin AN, et al. Ligilactobacillus salivarius 7247 strain: probiotic properties and anti-Salmonella effect with prebiotics. Antibiotics (Basel). (2023) 12:1535. doi: 10.3390/antibiotics12101535
74. Ismael M, Qayyum N, Gu Y, Zhezhe Y, Cui Y, Zhang Y, et al. Protective effect of plantaricin bio-LP1 bacteriocin on multidrug-resistance Escherichia Coli infection by alleviate the inflammation and modulate of gut-microbiota in BALB/c mice model. Int J Biol Macromol. (2023) 246:125700. doi: 10.1016/j.ijbiomac.2023.125700
75. Mann ER, Lam YK, Uhlig HH. Short-chain fatty acids: linking diet, the microbiome and immunity. Nat Rev Immunol. (2024) 24:577–95. doi: 10.1038/s41577-024-01014-8
76. Pan Y, Zhang H, Li M, He T, Guo S, Zhu L, et al. Novel approaches in IBD therapy: targeting the gut microbiota-bile acid axis. Gut Microbes. (2024) 16:2356284. doi: 10.1080/19490976.2024.2356284
77. Sorbara MT, Dubin K, Littmann ER, Moody TU, Fontana E, Seok R, et al. Inhibiting antibiotic-resistant Enterobacteriaceae by microbiota-mediated intracellular acidification. J Exp Med. (2019) 216:84–98. doi: 10.1084/jem.20181639
78. Bachem A, Makhlouf C, Binger KJ, De Souza DP, Tull D, Hochheiser K, et al. Microbiota-derived short-chain fatty acids promote the memory potential of antigen-activated CD8+ T cells. Immunity. (2019) 51:285–97.e5. doi: 10.1016/j.immuni.2019.06.002
79. Arpaia N, Campbell C, Fan X, Dikiy S, van der Veeken J, Deroos P, et al. Metabolites produced by commensal bacteria promote peripheral regulatory T-cell generation. Nature. (2013) 504:451–5. doi: 10.1038/nature12726
80. Dupraz L, Magniez A, Rolhion N, Richard ML, Da Costa G, Touch S, et al. Gut microbiota-derived short-chain fatty acids regulate IL-17 production by mouse and human intestinal γδ T cells. Cell Rep. (2021) 36:109332. doi: 10.1016/j.celrep.2021.109332
81. Zheng L, Kelly CJ, Battista KD, Schaefer R, Lanis JM, Alexeev EE, et al. Microbial-derived butyrate promotes epithelial barrier function through IL-10 receptor-dependent repression of claudin-2. J Immunol. (2017) 199:2976–84. doi: 10.4049/jimmunol.1700105
82. Adams L, Li X, Burchmore R, Goodwin RJA, Wall DM. Microbiome-derived metabolite effects on intestinal barrier integrity and immune cell response to infection. Microbiol (Reading). (2024) 170:001504. doi: 10.1099/mic.0.001504
83. Osbelt L, Thiemann S, Smit N, Lesker TR, Schröter M, Gálvez EJC, et al. Variations in microbiota composition of laboratory mice influence Citrobacter rodentium infection via variable short-chain fatty acid production. PLoS Pathog. (2020) 16:e1008448. doi: 10.1371/journal.ppat.1008448
84. Saeedi BJ, Liu KH, Owens JA, Hunter-Chang S, Camacho MC, Eboka RU, et al. Gut-resident lactobacilli activate hepatic nrf2 and protect against oxidative liver injury. Cell Metab. (2020) 31:956–68.e5. doi: 10.1016/j.cmet.2020.03.006
85. Zhang S, Zhao J, Xie F, He H, Johnston LJ, Dai X, et al. Dietary fiber-derived short-chain fatty acids: A potential therapeutic target to alleviate obesity-related nonalcoholic fatty liver disease. Obes Rev. (2021) 22:e13316. doi: 10.1111/obr.13316
86. Sato Y, Atarashi K, Plichta DR, Arai Y, Sasajima S, Kearney SM, et al. Novel bile acid biosynthetic pathways are enriched in the microbiome of centenarians. Nature. (2021) 599:458–64. doi: 10.1038/s41586-021-03832-5
87. Mcmillan AS, Theriot CM. Bile acids impact the microbiota, host, and C. difficile dynamics providing insight into mechanisms of efficacy of FMTs and microbiota-focused therapeutics. Gut Microbes. (2024) 16:2393766. doi: 10.1080/19490976.2024.2393766
88. Guzior DV, Quinn RA. Review: microbial transformations of human bile acids. Microbiome. (2021) 9:140. doi: 10.1186/s40168-021-01101-1
89. He Z, Ma Y, Yang S, Zhang S, Liu S, Xiao J, et al. Gut microbiota-derived ursodeoxycholic acid from neonatal dairy calves improves intestinal homeostasis and colitis to attenuate extended-spectrum β-lactamase-producing enteroaggregative Escherichia coli infection. Microbiome. (2022) 10:79. doi: 10.1186/s40168-022-01269-0
90. Sannasiddappa TH, Lund PA, Clarke SR. In vitro antibacterial activity of unconjugated and conjugated bile salts on Staphylococcus aureus. Front Microbiol. (2017) 8:1581. doi: 10.3389/fmicb.2017.01581
91. Watanabe M, Fukiya S, Yokota A. Comprehensive evaluation of the bactericidal activities of free bile acids in the large intestine of humans and rodents. J Lipid Res. (2017) 58:1143–52. doi: 10.1194/jlr.M075143
92. Termén S, Tollin M, Rodriguez E, Sveinsdóttir SH, Jóhannesson B, Cederlund A, et al. PU.1 and bacterial metabolites regulate the human gene CAMP encoding antimicrobial peptide LL-37 in colon epithelial cells. Mol Immunol. (2008) 45:3947–55. doi: 10.1016/j.molimm.2008.06.020
93. Li W, Hang S, Fang Y, Bae S, Zhang Y, Zhang M, et al. A bacterial bile acid metabolite modulates T(reg) activity through the nuclear hormone receptor NR4A1. Cell Host Microbe. (2021) 29:1366–77.e9. doi: 10.1016/j.chom.2021.07.013
94. Campbell C, Mckenney PT, Konstantinovsky D, Isaeva OI, Schizas M, Verter J, et al. Bacterial metabolism of bile acids promotes generation of peripheral regulatory T cells. Nature. (2020) 581:475–79. doi: 10.1038/s41586-020-2193-0
95. Le Guern R, Grandjean T, Stabler S, Bauduin M, Gosset P, Kipnis É, et al. Gut colonisation with multidrug-resistant Klebsiella pneumoniae worsens Pseudomonas aeruginosa lung infection. Nat Commun. (2023) 14:78. doi: 10.1038/s41467-022-35767-4
96. Renga G, Nunzi E, Stincardini C, Pariano M, Puccetti M, Pieraccini G, et al. CPX-351 exploits the gut microbiota to promote mucosal barrier function, colonization resistance, and immune homeostasis. Blood. (2024) 143:1628–45. doi: 10.1182/blood.2023021380
97. Schroeder BO, Birchenough GMH, Ståhlman M, Arike L, Johansson MEV, Hansson GC, et al. Bifidobacteria or fiber protects against diet-induced microbiota-mediated colonic mucus deterioration. Cell Host Microbe. (2018) 23:27–40.e7. doi: 10.1016/j.chom.2017.11.004
98. Di Vincenzo F, Del Gaudio A, Petito V, Lopetuso LR, Scaldaferri F. Gut microbiota, intestinal permeability, and systemic inflammation: a narrative review. Intern Emerg Med. (2024) 19:275–93. doi: 10.1007/s11739-023-03374-w
99. Pral LP, Fachi JL, Corrêa RO, Colonna M, Vinolo M. Hypoxia and HIF-1 as key regulators of gut microbiota and host interactions. Trends Immunol. (2021) 42:604–21. doi: 10.1016/j.it.2021.05.004
100. Zhang T, Sasabe J, Hullahalli K, Sit B, Waldor MK. Increased Listeria monocytogenes dissemination and altered population dynamics in muc2-deficient mice. Infect Immun. (2021) 89:e00667-20. doi: 10.1128/iai.00667-20
101. Neumann M, Steimle A, Grant ET, Wolter M, Parrish A, Willieme S, et al. Deprivation of dietary fiber in specific-pathogen-free mice promotes susceptibility to the intestinal mucosal pathogen. Citrobacter rodentium. Gut Microbes. (2021) 13:1966263. doi: 10.1080/19490976.2021.1966263
102. Wolter M, Grant ET, Boudaud M, Pudlo NA, Pereira GV, Eaton KA, et al. Diet-driven differential response of Akkermansia muciniphila modulates pathogen susceptibility. Mol Syst Biol. (2024) 20:596–625. doi: 10.1038/s44320-024-00036-7
103. Pickard JM, Maurice CF, Kinnebrew MA, Abt MC, Schenten D, Golovkina TV, et al. Rapid fucosylation of intestinal epithelium sustains host-commensal symbiosis in sickness. Nature. (2014) 514:638–41. doi: 10.1038/nature13823
104. Nusrat A, Von-Eichel-Streiber C, Turner JR, Verkade P, Madara JL, Parkos CA. Clostridium difficile toxins disrupt epithelial barrier function by altering membrane microdomain localization of tight junction proteins. Infect Immun. (2001) 69:1329–36. doi: 10.1128/iai.69.3.1329-1336.2001
105. Ghosh S, Erickson D, Chua MJ, Collins J, Jala VR. The microbial metabolite urolithin A reduces Clostridioides difficile toxin expression and toxin-induced epithelial damage. mSystems. (2024) 9:e0125523. doi: 10.1128/msystems.01255-23
106. Fachi JL, Pral LP, Assis HC, Oliveira S, Rodovalho VR, Dos Santos J, et al. Hyperbaric oxygen augments susceptibility to C. difficile infection by impairing gut microbiota ability to stimulate the HIF-1α-IL-22 axis in ILC3. Gut Microbes. (2024) 16:2297872. doi: 10.1080/19490976.2023.2297872
107. Zhao T, Liu X, Chu Z, Zhao J, Jiang D, Dong X, et al. L-arginine loading porous PEEK promotes percutaneous tissue repair through macrophage orchestration. Bioact Mater. (2024) 40:19–33. doi: 10.1016/j.bioactmat.2024.05.025
108. Winter SE, Winter MG, Xavier MN, Thiennimitr P, Poon V, Keestra AM, et al. Host-derived nitrate boosts growth of E. coli in the inflamed gut. Science. (2013) 339:708–11. doi: 10.1126/science.1232467
109. Baldelli V, Scaldaferri F, Putignani L, Del Chierico F. The role of enterobacteriaceae in gut microbiota dysbiosis in inflammatory bowel diseases. Microorganisms. (2021) 9:697. doi: 10.3390/microorganisms9040697
110. Byndloss MX, Olsan EE, Rivera-Chávez F, Tiffany CR, Cevallos SA, Lokken KL, et al. Microbiota-activated PPAR-γ signaling inhibits dysbiotic Enterobacteriaceae expansion. Science. (2017) 357:570–75. doi: 10.1126/science.aam9949
111. Rivera-Chávez F, Lopez CA, Bäumler AJ. Oxygen as a driver of gut dysbiosis. Free Radic Biol Med. (2017) 105:93–101. doi: 10.1016/j.freeradbiomed.2016.09.022
112. Litvak Y, Byndloss MX, Tsolis RM, Bäumler AJ. Dysbiotic Proteobacteria expansion: a microbial signature of epithelial dysfunction. Curr Opin Microbiol. (2017) 39:1–6. doi: 10.1016/j.mib.2017.07.003
113. Shelton CD, Byndloss MX. Gut epithelial metabolism as a key driver of intestinal dysbiosis associated with noncommunicable diseases. Infect Immun. (2020) 88:e00939-19. doi: 10.1128/iai.00939-19
114. Ziaka M, Exadaktylos A. Gut-derived immune cells and the gut-lung axis in ARDS. Crit Care. (2024) 28:220. doi: 10.1186/s13054-024-05006-x
115. Sun Y, Ho CT, Zhang X. Neuroprotection of food bioactives in neurodegenerative diseases: role of the gut microbiota and innate immune receptors. J Agric Food Chem. (2023) 71:2718–33. doi: 10.1021/acs.jafc.2c07742
116. Maynard CL, Elson CO, Hatton RD, Weaver CT. Reciprocal interactions of the intestinal microbiota and immune system. Nature. (2012) 489:231–41. doi: 10.1038/nature11551
117. Chen L, Zhang L, Hua H, Liu L, Mao Y, Wang R. Interactions between toll-like receptors signaling pathway and gut microbiota in host homeostasis. Immun Inflammation Dis. (2024) 12:e1356. doi: 10.1002/iid3.1356
118. Ost KS, O’meara TR, Stephens WZ, Chiaro T, Zhou H, Penman J, et al. Adaptive immunity induces mutualism between commensal eukaryotes. Nature. (2021) 596:114–18. doi: 10.1038/s41586-021-03722-w
119. Xu Z, Takizawa F, Casadei E, Shibasaki Y, Ding Y, Sauters TJC, et al. Specialization of mucosal immunoglobulins in pathogen control and microbiota homeostasis occurred early in vertebrate evolution. Sci Immunol. (2020) 5:eaay3254. doi: 10.1126/sciimmunol.aay3254
120. Ramanan D, Pratama A, Zhu Y, Venezia O, Sassone-Corsi M, Chowdhary K, et al. Regulatory T cells in the face of the intestinal microbiota. Nat Rev Immunol. (2023) 23:749–62. doi: 10.1038/s41577-023-00890-w
121. Zhou P, Chen C, Patil S, Dong S. Unveiling the therapeutic symphony of probiotics, prebiotics, and postbiotics in gut-immune harmony. Front Nutr. (2024) 11:1355542. doi: 10.3389/fnut.2024.1355542
122. Hays KE, Pfaffinger JM, Ryznar R. The interplay between gut microbiota, short-chain fatty acids, and implications for host health and disease. Gut Microbes. (2024) 16:2393270. doi: 10.1080/19490976.2024.2393270
123. Miller WR, Arias CA. ESKAPE pathogens: antimicrobial resistance, epidemiology, clinical impact and therapeutics. Nat Rev Microbiol. (2024) 22:598–616. doi: 10.1038/s41579-024-01054-w
124. Rahman MN, Barua N, Tin MCF, Dharmaratne P, Wong SH, Ip M. The use of probiotics and prebiotics in decolonizing pathogenic bacteria from the gut; a systematic review and meta-analysis of clinical outcomes. Gut Microbes. (2024) 16:2356279. doi: 10.1080/19490976.2024.2356279
125. Kim SG, Becattini S, Moody TU, Shliaha PV, Littmann ER, Seok R, et al. Microbiota-derived lantibiotic restores resistance against vancomycin-resistant Enterococcus. Nature. (2019) 572:665–69. doi: 10.1038/s41586-019-1501-z
126. Li X, Song L, Zhu S, Xiao Y, Huang Y, Hua Y, et al. Two strains of lactobacilli effectively decrease the colonization of VRE in a mouse model. Front Cell Infect Microbiol. (2019) 9:6. doi: 10.3389/fcimb.2019.00006
127. Mu A, Carter GP, Li L, Isles NS, Vrbanac AF, Morton JT, et al. Microbe-metabolite associations linked to the rebounding murine gut microbiome postcolonization with vancomycin-resistant Enterococcus faecium. mSystems. (2020) 5:e00452-20. doi: 10.1128/mSystems.00452-20
128. Rosenstein R, Torres Salazar BO, Sauer C, Heilbronner S, Krismer B, Peschel A. The Staphylococcus aureus-antagonizing human nasal commensal Staphylococcus lugdunensis depends on siderophore piracy. Microbiome. (2024) 12:213. doi: 10.1186/s40168-024-01913-x
129. Zhao Y, Sun H, Chen Y, Niu Q, Dong Y, Li M, et al. Butyrate protects against MRSA pneumonia via regulating gut-lung microbiota and alveolar macrophage M2 polarization. mBio. (2023) 14:e0198723. doi: 10.1128/mbio.01987-23
130. Wang H, Kim R, Wang Y, Furtado KL, Sims CE, Tamayo R, et al. In vitro co-culture of Clostridium scindens with primary human colonic epithelium protects the epithelium against Staphylococcus aureus. Front Bioeng Biotechnol. (2024) 12:1382389. doi: 10.3389/fbioe.2024.1382389
131. Zhao C, Wu K, Hao H, Zhao Y, Bao L, Qiu M, et al. Gut microbiota-mediated secondary bile acid alleviates Staphylococcus aureus-induced mastitis through the TGR5-cAMP-PKA-NF-κB/NLRP3 pathways in mice. NPJ Biofilms Microbiomes. (2023) 9:8. doi: 10.1038/s41522-023-00374-8
132. Mousa WK, Ghemrawi R, Abu-Izneid T, Ramadan A, Al-Marzooq F. Discovery of lactomodulin, a unique microbiome-derived peptide that exhibits dual anti-inflammatory and antimicrobial activity against multidrug-resistant pathogens. Int J Mol Sci. (2023) 24:6901. doi: 10.3390/ijms24086901
133. Hecht AL, Harling LC, Friedman ES, Tanes C, Lee J, Firrman J, et al. Dietary carbohydrates regulate intestinal colonization and dissemination of Klebsiella pneumoniae. J Clin Invest. (2024) 134:e174726. doi: 10.1172/jci174726
134. Shen M, Zhao H, Han M, Su L, Cui X, Li D, et al. Alcohol-induced gut microbiome dysbiosis enhances the colonization of Klebsiella pneumoniae on the mouse intestinal tract. mSystems. (2024) 9:e0005224. doi: 10.1128/msystems.00052-24
135. Stacy A, Andrade-Oliveira V, Mcculloch JA, Hild B, Oh JH, Perez-Chaparro PJ, et al. Infection trains the host for microbiota-enhanced resistance to pathogens. Cell. (2021) 184:615–27.e17. doi: 10.1016/j.cell.2020.12.011
136. Tang HJ, Chen CC, Lu YC, Huang HL, Chen HJ, Chuang YC, et al. The effect of Lactobacillus with prebiotics on KPC-2-producing Klebsiella pneumoniae. Front Microbiol. (2022) 13:1050247. doi: 10.3389/fmicb.2022.1050247
137. Asahara T, Takahashi A, Yuki N, Kaji R, Takahashi T, Nomoto K. Protective Effect of a Synbiotic against Multidrug-Resistant Acinetobacter baumannii in a Murine Infection Model. Antimicrob Agents Chemother. (2016) 60:3041–50. doi: 10.1128/aac.02928-15
138. Wen L, Shi L, Kong XL, Li KY, Li H, Jiang DX, et al. Gut Microbiota Protected Against pseudomonas aeruginosa Pneumonia via Restoring Treg/Th17 Balance and Metabolism. Front Cell Infect Microbiol. (2022) 12:856633. doi: 10.3389/fcimb.2022.856633
139. Janapatla RP, Dudek A, Chen CL, Chuang CH, Chien KY, Feng Y, et al. Marine prebiotics mediate decolonization of Pseudomonas aeruginosa from gut by inhibiting secreted virulence factor interactions with mucins and enriching Bacteroides population. J BioMed Sci. (2023) 30:9. doi: 10.1186/s12929-023-00902-w
140. Zamani H, Rahbar S, Garakoui SR, Afsah Sahebi A, Jafari H. Antibiofilm potential of Lactobacillus plantarum spp. cell free supernatant (CFS) against multidrug resistant bacterial pathogens. Pharm Biomed Res. (2017) 3:39–44. doi: 10.29252/pbr.3.2.39
141. Nagao-Kitamoto H, Leslie JL, Kitamoto S, Jin C, Thomsson KA, Gillilland MG 3rd, et al. Interleukin-22-mediated host glycosylation prevents Clostridioides difficile infection by modulating the metabolic activity of the gut microbiota. Nat Med. (2020) 26:608–17. doi: 10.1038/s41591-020-0764-0
142. Pensinger DA, Fisher AT, Dobrila HA, Van Treuren W, Gardner JO, Higginbottom SK, et al. Butyrate differentiates permissiveness to Clostridioides difficile infection and influences growth of diverse C. difficile Isolates Infect Immun. (2023) 91:e0057022. doi: 10.1128/iai.00570-22
143. Tian H, Cui J, Ye C, Zhao J, Yang B, Xu Y, et al. Depletion of butyrate-producing microbes of the Firmicutes predicts nonresponse to FMT therapy in patients with recurrent Clostridium difficile infection. Gut Microbes. (2023) 15:2236362. doi: 10.1080/19490976.2023.2236362
144. Winston JA, Rivera AJ, Cai J, Thanissery R, Montgomery SA, Patterson AD, et al. Ursodeoxycholic Acid (UDCA) Mitigates the Host Inflammatory Response during Clostridioides difficile Infection by Altering Gut Bile Acids. Infect Immun. (2020) 88:e00045-20. doi: 10.1128/iai.00045-20
145. Wei Y, Yang F, Wu Q, Gao J, Liu W, Liu C, et al. Protective effects of bifidobacterial strains against toxigenic Clostridium difficile. Front Microbiol. (2018) 9:888. doi: 10.3389/fmicb.2018.00888
146. Hagihara M, Ariyoshi T, Kuroki Y, Eguchi S, Higashi S, Mori T, et al. Clostridium butyricum enhances colonization resistance against Clostridioides difficile by metabolic and immune modulation. Sci Rep. (2021) 11:15007. doi: 10.1038/s41598-021-94572-z
147. Yip AYG, King OG, Omelchenko O, Kurkimat S, Horrocks V, Mostyn P, et al. Antibiotics promote intestinal growth of carbapenem-resistant Enterobacteriaceae by enriching nutrients and depleting microbial metabolites. Nat Commun. (2023) 14:5094. doi: 10.1038/s41467-023-40872-z
148. Chang KC, Nagarajan N, Gan YH. Short-chain fatty acids of various lengths differentially inhibit Klebsiella pneumoniae and Enterobacteriaceae species. mSphere. (2024) 9:e0078123. doi: 10.1128/msphere.00781-23
149. Krawczyk B, Wityk P, Gałęcka M, Michalik M. The many faces of enterococcus spp.-commensal, probiotic and opportunistic pathogen. Microorganisms. (2021) 9(9):1900. doi: 10.3390/microorganisms9091900
150. Xu L, Wu Y, Yang X, Pang X, Wu Y, Li X, et al. The Fe-S cluster biosynthesis in Enterococcus faecium is essential for anaerobic growth and gastrointestinal colonization. Gut Microbes. (2024) 16:2359665. doi: 10.1080/19490976.2024.2359665
151. Brandl K, Plitas G, Mihu CN, Ubeda C, Jia T, Fleisher M, et al. Vancomycin-resistant enterococci exploit antibiotic-induced innate immune deficits. Nature. (2008) 455:804–7. doi: 10.1038/nature07250
152. Abt MC, Buffie CG, Sušac B, Becattini S, Carter RA, Leiner I, et al. TLR-7 activation enhances IL-22-mediated colonization resistance against vancomycin-resistant enterococcus. Sci Transl Med. (2016) 8:327ra25. doi: 10.1126/scitranslmed.aad6663
153. Ubeda C, Bucci V, Caballero S, Djukovic A, Toussaint NC, Equinda M, et al. Intestinal microbiota containing Barnesiella species cures vancomycin-resistant Enterococcus faecium colonization. Infect Immun. (2013) 81:965–73. doi: 10.1128/iai.01197-12
154. Dubberke ER, Mullane KM, Gerding DN, Lee CH, Louie TJ, Guthertz H, et al. Clearance of Vancomycin-Resistant Enterococcus Concomitant With Administration of a Microbiota-Based Drug Targeted at Recurrent Clostridium difficile Infection. Open Forum Infect Dis. (2016) 3:ofw133. doi: 10.1093/ofid/ofw133
155. Davido B, Batista R, Fessi H, Michelon H, Escaut L, Lawrence C, et al. Fecal microbiota transplantation to eradicate vancomycin-resistant enterococci colonization in case of an outbreak. Med Mal Infect. (2019) 49:214–18. doi: 10.1016/j.medmal.2018.11.002
156. Hyun J, Lee SK, Cheon JH, Yong DE, Koh H, Kang YK, et al. Faecal microbiota transplantation reduces amounts of antibiotic resistance genes in patients with multidrug-resistant organisms. Antimicrob Resist Infect Control. (2022) 11:20. doi: 10.1186/s13756-022-01064-4
157. Smith AB, Jenior ML, Keenan O, Hart JL, Specker J, Abbas A, et al. Enterococci enhance Clostridioides difficile pathogenesis. Nature. (2022) 611:780–86. doi: 10.1038/s41586-022-05438-x
158. Dantes R, Mu Y, Belflower R, Aragon D, Dumyati G, Harrison LH, et al. National burden of invasive methicillin-resistant Staphylococcus aureus infections, United States, 2011. JAMA Intern Med. (2013) 173:1970–8. doi: 10.1001/jamainternmed.2013.10423
159. Piewngam P, Otto M. Staphylococcus aureus colonisation and strategies for decolonisation. Lancet Microbe. (2024) 5:e606–e18. doi: 10.1016/s2666-5247(24)00040-5
160. Bitrus AA, Zunita Z, Khairani-Bejo S, Othman S, Ahmad Nadzir NA. Staphylococcal cassette chromosome mec (SCCmec) and characterization of the attachment site (attB) of methicillin resistant Staphylococcus aureus (MRSA) and methicillin susceptible Staphylococcus aureus (MSSA) isolates. Microb Pathog. (2018) 123:323–29. doi: 10.1016/j.micpath.2018.07.033
161. Idrees M, Sawant S, Karodia N, Rahman A. Staphylococcus aureus biofilm: morphology, genetics, pathogenesis and treatment strategies. Int J Environ Res Public Health. (2021) 18:7602. doi: 10.3390/ijerph18147602
162. Zhou C, Pawline MB, Pironti A, Morales SM, Perault AI, Ulrich RJ, et al. Microbiota and metabolic adaptation shape Staphylococcus aureus virulence and antimicrobial resistance during intestinal colonization. bioRxiv. (2024). doi: 10.1101/2024.05.11.593044 (Preprint)
163. Zipperer A, Konnerth MC, Laux C, Berscheid A, Janek D, Weidenmaier C, et al. Human commensals producing a novel antibiotic impair pathogen colonization. Nature. (2016) 535:511–6. doi: 10.1038/nature18634
164. Kumar Y, Yadav R, Bhatia A. Can natural detergent properties of bile acids be used beneficially in tackling coronavirus disease-19? Future Virol. (2020) 15:779–82. doi: 10.2217/fvl-2020-0210
165. Ridlon JM, Harris SC, Bhowmik S, Kang DJ. and Hylemon, P.B. Consequences of bile salt biotransformations by intestinal bacteria. Gut Microbes. (2016) 7:22–39. doi: 10.1080/19490976.2015.1127483
166. Liu A, Garrett S, Hong W, Zhang J. Staphylococcus aureus infections and human intestinal microbiota. Pathogens. (2024) 13:276. doi: 10.3390/pathogens13040276
167. Ma X, Jin H, Chu X, Dai W, Tang W, Zhu J, et al. The host CYP1A1-microbiota metabolic axis promotes gut barrier disruption in methicillin-resistant Staphylococcus aureus-induced abdominal sepsis. Front Microbiol. (2022) 13:802409. doi: 10.3389/fmicb.2022.802409
168. Ali S, Alam M, Hasan GM, Hassan MI. Potential therapeutic targets of Klebsiella pneumoniae: a multi-omics review perspective. Brief Funct Genomics. (2022) 21:63–77. doi: 10.1093/bfgp/elab038
169. Osbelt L, Almási É DH, Wende M, Kienesberger S, Voltz A, Lesker TR, et al. Klebsiella oxytoca inhibits Salmonella infection through multiple microbiota-context-dependent mechanisms. Nat Microbiol. (2024) 9:1792–811. doi: 10.1038/s41564-024-01710-0
170. Hu J, Tong Y, Shen Z, Li Y, Cheng C, Au R, et al. Gegen Qinlian decoction ameliorates murine colitis by inhibiting the expansion of Enterobacteriaceae through activating PPAR-γ signaling. BioMed Pharmacother. (2022) 154:113571. doi: 10.1016/j.biopha.2022.113571
171. Yoon JG, Lim S, Hyun HJ, Seong H, Noh JY, Song JY, et al. Respiratory microbiome and clinical course of carbapenem-resistant Acinetobacter baumannii pneumonia in critically Ill patients. Med (Baltimore). (2024) 103:e38988. doi: 10.1097/md.0000000000038988
172. Ren X, Palmer LD. Acinetobacter metabolism in infection and antimicrobial resistance. Infect Immun. (2023) 91:e0043322. doi: 10.1128/iai.00433-22
173. Uppalapati SR, Sett A, Pathania R. The outer membrane proteins ompA, carO, and oprD of Acinetobacter baumannii confer a two-pronged defense in facilitating its success as a potent human pathogen. Front Microbiol. (2020) 11:589234. doi: 10.3389/fmicb.2020.589234
174. Ejaz H, Ahmad M, Younas S, Junaid K, Abosalif KOA, Abdalla AE, et al. Molecular Epidemiology of Extensively-Drug Resistant Acinetobacter baumannii Sequence Type 2 Co-Harboring bla (NDM) and bla (OXA) From Clinical Origin. Infect Drug Resist. (2021) 14:1931–39. doi: 10.2147/idr.S310478
175. Tacconelli E, Carrara E, Savoldi A, Harbarth S, Mendelson M, Monnet DL, et al. Discovery, research, and development of new antibiotics: the WHO priority list of antibiotic-resistant bacteria and tuberculosis. Lancet Infect Dis. (2018) 18:318–27. doi: 10.1016/s1473-3099(17)30753-3
176. Ren X, Clark RM, Bansah DA, Varner EN, Tiffany CR, Jaswal K, et al. Amino acid competition shapes Acinetobacter baumannii gut carriage. bioRxiv. (2024). doi: 10.1101/2024.10.19.619093
177. Lin HR, Shu HY, Lin GH. Biological roles of indole-3-acetic acid in Acinetobacter baumannii. Microbiol Res. (2018) 216:30–9. doi: 10.1016/j.micres.2018.08.004
178. Jurado-Martín I, Sainz-Mejías M, Mcclean S. Pseudomonas aeruginosa: an audacious pathogen with an adaptable arsenal of virulence factors. Int J Mol Sci. (2021) 22:3128. doi: 10.3390/ijms22063128
179. Rossi E, La Rosa R, Bartell JA, Marvig RL, Haagensen J, Sommer LM, et al. Pseudomonas aeruginosa adaptation and evolution in patients with cystic fibrosis. Nat Rev Microbiol. (2021) 19:331–42. doi: 10.1038/s41579-020-00477-5
180. Horcajada JP, Montero M, Oliver A, Sorlí L, Luque S, Gómez-Zorrilla S, et al. Epidemiology and treatment of multidrug-resistant and extensively drug-resistant Pseudomonas aeruginosa infections. Clin Microbiol Rev. (2019) 32:e00031-19. doi: 10.1128/cmr.00031-19
181. Breidenstein EB, de-la-Fuente-Núñez C, Hancock RE. Pseudomonas aeruginosa: all roads lead to resistance. Trends Microbiol. (2011) 19:419–26. doi: 10.1016/j.tim.2011.04.005
182. Wheatley R, Diaz Caballero J, Kapel N, De Winter FHR, Jangir P, Quinn A, et al. Rapid evolution and host immunity drive the rise and fall of carbapenem resistance during an acute Pseudomonas aeruginosa infection. Nat Commun. (2021) 12:2460. doi: 10.1038/s41467-021-22814-9
183. Botelho J, Grosso F, Peixe L. Antibiotic resistance in Pseudomonas aeruginosa - Mechanisms, epidemiology and evolution. Drug Resist Update. (2019) 44:100640. doi: 10.1016/j.drup.2019.07.002
184. Gómez-Zorrilla S, Camoez M, Tubau F, Cañizares R, Periche E, Dominguez MA, et al. Prospective observational study of prior rectal colonization status as a predictor for subsequent development of Pseudomonas aeruginosa clinical infections. Antimicrob Agents Chemother. (2015) 59:5213–9. doi: 10.1128/aac.04636-14
185. Kubinak JL, Petersen C, Stephens WZ, Soto R, Bake E, O’connell RM, et al. MyD88 signaling in T cells directs IgA-mediated control of the microbiota to promote health. Cell Host Microbe. (2015) 17:153–63. doi: 10.1016/j.chom.2014.12.009
186. Zhou Y, Zhang H, Verma P, Samanta D, Verma MK, Kumar K. Validation of secretory Immunoglobulin A (IgA) for early and efficient diagnosis of Pseudomonas aeruginosa lung infection. Cell Mol Biol (Noisy-le-grand). (2023) 69:35–9. doi: 10.14715/cmb/2023.69.7.6
187. Robak OH, Heimesaat MM, Kruglov AA, Prepens S, Ninnemann J, Gutbier B, et al. Antibiotic treatment-induced secondary IgA deficiency enhances susceptibility to Pseudomonas aeruginosa pneumonia. J Clin Invest. (2018) 128:3535–45. doi: 10.1172/jci97065
188. Lécuyer E, Rakotobe S, Lengliné-Garnier H, Lebreton C, Picard M, Juste C, et al. Segmented filamentous bacterium uses secondary and tertiary lymphoid tissues to induce gut IgA and specific T helper 17 cell responses. Immunity. (2014) 40:608–20. doi: 10.1016/j.immuni.2014.03.009
189. Rooney AM, Cochrane K, Fedsin S, Yao S, Anwer S, Dehmiwal S, et al. A microbial consortium alters intestinal Pseudomonadota and antimicrobial resistance genes in individuals with recurrent Clostridioides difficile infection. mBio. (2023) 14:e0348222. doi: 10.1128/mbio.03482-22
190. Lou TL, Ji T, Peng X, Ji WW, Yuan LX, Wang J, et al. Extract From Tetrastigma hemsleyanum Leaf Alleviates Pseudomonas aeruginosa Lung Infection: Network Pharmacology Analysis and Experimental Evidence. Front Pharmacol. (2021) 12:587850. doi: 10.3389/fphar.2021.587850
191. Di Bella S, Sanson G, Monticelli J, Zerbato V, Principe L, Giuffrè M, et al. Clostridioides difficile infection: history, epidemiology, risk factors, prevention, clinical manifestations, treatment, and future options. Clin Microbiol Rev. (2024) 37:e0013523. doi: 10.1128/cmr.00135-23
192. Bouwknegt M, Van Dorp S, Kuijper E. Burden of Clostridium difficile infection in the United States. N Engl J Med. (2015) 372:2368. doi: 10.1056/NEJMc1505190
193. Theriot CM, Koenigsknecht MJ, Carlson PE Jr., Hatton GE, Nelson AM, Li B, et al. Antibiotic-induced shifts in the mouse gut microbiome and metabolome increase susceptibility to Clostridium difficile infection. Nat Commun. (2014) 5:3114. doi: 10.1038/ncomms4114
194. Bratkovič T, Zahirović A, Bizjak M, Rupnik M, Štrukelj B, Berlec A. New treatment approaches for Clostridioides difficile infections: alternatives to antibiotics and fecal microbiota transplantation. Gut Microbes. (2024) 16:2337312. doi: 10.1080/19490976.2024.2337312
195. Louie T, Golan Y, Khanna S, Bobilev D, Erpelding N, Fratazzi C, et al. VE303, a defined bacterial consortium, for prevention of recurrent Clostridioides difficile infection: A randomized clinical trial. JAMA. (2023) 329:1356–66. doi: 10.1001/jama.2023.4314
196. Gurung B, Stricklin M, Wang S. Gut microbiota-gut metabolites and Clostridioides difficile infection: approaching sustainable solutions for therapy. Metabolites. (2024) 14:74. doi: 10.3390/metabo14010074
197. Jose S, Mukherjee A, Horrigan O, Setchell KDR, Zhang W, Moreno-Fernandez ME, et al. Obeticholic acid ameliorates severity of Clostridioides difficile infection in high fat diet-induced obese mice. Mucosal Immunol. (2021) 14:500–10. doi: 10.1038/s41385-020-00338-7
198. Rea MC, Dobson A, O’sullivan O, Crispie F, Fouhy F, Cotter PD, et al. Effect of broad- and narrow-spectrum antimicrobials on Clostridium difficile and microbial diversity in a model of the distal colon. Proc Natl Acad Sci U S A. (2011) 108 Suppl 1:4639–44. doi: 10.1073/pnas.1001224107
199. Lay CL, Dridi L, Bergeron MG, Ouellette M, Fliss IL. Nisin is an effective inhibitor of Clostridium difficile vegetative cells and spore germination. J Med Microbiol. (2016) 65:169–75. doi: 10.1099/jmm.0.000202
200. Davin-Regli A, Lavigne JP, Pagès JM. Enterobacter spp.: update on taxonomy, clinical aspects, and emerging antimicrobial resistance. Clin Microbiol Rev. (2019) 32:e00002-19. doi: 10.1128/cmr.00002-19
201. Palacios-Baena ZR, Giannella M, Manissero D, Rodríguez-Baño J, Viale P, Lopes S, et al. Risk factors for carbapenem-resistant Gram-negative bacterial infections: a systematic review. Clin Microbiol Infect. (2021) 27:228–35. doi: 10.1016/j.cmi.2020.10.016
202. Sindi AA, Alsayed SM, Abushoshah I, Bokhary DH, Tashkandy NR. Profile of the gut microbiome containing carbapenem-Resistant enterobacteriaceae in ICU patients. Microorganisms. (2022) 10:1309. doi: 10.3390/microorganisms10071309
203. Baek MS, Kim S, Kim WY, Kweon MN, Huh JW. Gut microbiota alterations in critically Ill patients with carbapenem-resistant Enterobacteriaceae colonization: A clinical analysis. Front Microbiol. (2023) 14:1140402. doi: 10.3389/fmicb.2023.1140402
204. Kunishima H, Ishibashi N, Wada K, Oka K, Takahashi M, Yamasaki Y, et al. The effect of gut microbiota and probiotic organisms on the properties of extended spectrum beta-lactamase producing and carbapenem resistant Enterobacteriaceae including growth, beta-lactamase activity and gene transmissibility. J Infect Chemother. (2019) 25:894–900. doi: 10.1016/j.jiac.2019.04.021
205. Van Hul M, Cani PD, Petitfils C, De Vos WM, Tilg H, El-Omar EM. What defines a healthy gut microbiome? Gut. (2024) 73:1893–908. doi: 10.1136/gutjnl-2024-333378
206. Yadegar A, Bar-Yoseph H, Monaghan TM, Pakpour S, Severino A, Kuijper EJ, et al. Fecal microbiota transplantation: current challenges and future landscapes. Clin Microbiol Rev. (2024) 37:e0006022. doi: 10.1128/cmr.00060-22
207. Bonnet M, Eckert C, Tournebize R. Decolonization of asymptomatic carriage of multi-drug resistant bacteria by bacteriophages? Front Microbiol. (2023) 14:1266416. doi: 10.3389/fmicb.2023.1266416
208. Campos-Madueno EI, Moradi M, Eddoubaji Y, Shahi F, Moradi S, Bernasconi OJ, et al. Intestinal colonization with multidrug-resistant Enterobacterales: screening, epidemiology, clinical impact, and strategies to decolonize carriers. Eur J Clin Microbiol Infect Dis. (2023) 42:229–54. doi: 10.1007/s10096-023-04548-2
209. Liu W, Li X, Zhao Z, Pi X, Meng Y, Fei D, et al. Effect of chitooligosaccharides on human gut microbiota and antiglycation. Carbohydr Polym. (2020) 242:116413. doi: 10.1016/j.carbpol.2020.116413
210. Baek GH, Kim YJ, Lee Y, Jung SC, Seo HW, Kim JS. Prebiotic potential of green banana flour: impact on gut microbiota modulation and microbial metabolic activity in a murine model. Front Nutr. (2023) 10:1249358. doi: 10.3389/fnut.2023.1249358
211. Zou Y, Wang J, Wang Y, Peng B, Liu J, Zhang B, et al. Protection of Galacto-Oligosaccharide against E. coli O157 Colonization through Enhancing Gut Barrier Function and Modulating Gut Microbiota. Foods. (2020) 9:1710. doi: 10.3390/foods9111710
212. Gao J, Zhao L, Cheng Y, Lei W, Wang Y, Liu X, et al. Probiotics for the treatment of depression and its comorbidities: A systemic review. Front Cell Infect Microbiol. (2023) 13:1167116. doi: 10.3389/fcimb.2023.1167116
213. Nataraj BH, Ali SA, Behare PV, Yadav H. Postbiotics-parabiotics: the new horizons in microbial biotherapy and functional foods. Microb Cell Fact. (2020) 19:168. doi: 10.1186/s12934-020-01426-w
214. Clemente JC, Manasson J, Scher JU. The role of the gut microbiome in systemic inflammatory disease. BMJ. (2018) 360:j5145. doi: 10.1136/bmj.j5145
215. Bhattacharya S, Dutta A, Khanra PK, Gupta N, Dutta R, Tzvetkov NT, et al. In silico exploration of 4(α-l-rhamnosyloxy)-benzyl isothiocyanate: A promising phytochemical-based drug discovery approach for combating multi-drug resistant Staphylococcus aureus. Comput Biol Med. (2024) 179:108907. doi: 10.1016/j.compbiomed.2024.108907
216. Zelante T, Iannitti RG, Cunha C, De Luca A, Giovannini G, Pieraccini G, et al. Tryptophan catabolites from microbiota engage aryl hydrocarbon receptor and balance mucosal reactivity via interleukin-22. Immunity. (2013) 39:372–85. doi: 10.1016/j.immuni.2013.08.003
217. Marco ML, Heeney D, Binda S, Cifelli CJ, Cotter PD, Foligné B, et al. Health benefits of fermented foods: microbiota and beyond. Curr Opin Biotechnol. (2017) 44:94–102. doi: 10.1016/j.copbio.2016.11.010
218. Qoms MS, Arulrajah B, Wan Ibadullah WZ, Ramli NS, Chau DM, Sarbini SR, et al. Performance of Azolla pinnata fern protein hydrolysates as an emulsifier and nutraceutical ingredient in an O/W emulsion system and their effect on human gut microbiota and mammalian cells. Food Funct. (2024) 15:6578–96. doi: 10.1039/d4fo00377b
219. Wang J, Ji H, Wang S, Liu H, Zhang W, Zhang D, et al. Probiotic Lactobacillus plantarum promotes intestinal barrier function by strengthening the epithelium and modulating gut microbiota. Front Microbiol. (2018) 9:1953. doi: 10.3389/fmicb.2018.01953
220. Ma Y, Yang JY, Peng X, Xiao KY, Xu Q, Wang C. Which probiotic has the best effect on preventing Clostridium difficile-associated diarrhea? A systematic review and network meta-analysis. J Dig Dis. (2020) 21:69–80. doi: 10.1111/1751-2980.12839
221. Alberda C, Marcushamer S, Hewer T, Journault N, Kutsogiannis D. Feasibility of a Lactobacillus casei Drink in the Intensive Care Unit for Prevention of Antibiotic Associated Diarrhea and Clostridium difficile. Nutrients. (2018) 10:539. doi: 10.3390/nu10050539
222. Wieërs G, Verbelen V, Van Den Driessche M, Melnik E, Vanheule G, Marot JC, et al. Do probiotics during in-hospital antibiotic treatment prevent colonization of gut microbiota with multi-drug-resistant bacteria? A randomized placebo-controlled trial comparing Saccharomyces to a mixture of Lactobacillus, Bifidobacterium, and Saccharomyces. Front Public Health. (2020) 8:578089. doi: 10.3389/fpubh.2020.578089
223. Machairas N, Pistiki A, Droggiti DI, Georgitsi M, Pelekanos N, Damoraki G, et al. Pre-treatment with probiotics prolongs survival after experimental infection by multidrug-resistant Pseudomonas aeruginosa in rodents: an effect on sepsis-induced immunosuppression. Int J Antimicrob Agents. (2015) 45:376–84. doi: 10.1016/j.ijantimicag.2014.11.013
224. Swanson KS, Gibson GR, Hutkins R, Reimer RA, Reid G, Verbeke K, et al. The International Scientific Association for Probiotics and Prebiotics (ISAPP) consensus statement on the definition and scope of synbiotics. Nat Rev Gastroenterol Hepatol. (2020) 17:687–701. doi: 10.1038/s41575-020-0344-2
225. Shoaf K, Mulvey GL, Armstrong GD, Hutkins RW. Prebiotic galactooligosaccharides reduce adherence of enteropathogenic Escherichia coli to tissue culture cells. Infect Immun. (2006) 74:6920–8. doi: 10.1128/iai.01030-06
226. Meng X, Kawahara K, Matsushita K, Nawa Y, Shrestha B, Kikuchi K, et al. Attenuation of LPS-induced iNOS expression by 1,5-anhydro-d-fructose. Biochem Biophys Res Commun. (2009) 387:42–6. doi: 10.1016/j.bbrc.2009.06.108
227. Gibson GR, Hutkins R, Sanders ME, Prescott SL, Reimer RA, Salminen SJ, et al. Expert consensus document: The International Scientific Association for Probiotics and Prebiotics (ISAPP) consensus statement on the definition and scope of prebiotics. Nat Rev Gastroenterol Hepatol. (2017) 14:491–502. doi: 10.1038/nrgastro.2017.75
228. Goldenberg JZ, Yap C, Lytvyn L, Lo CK, Beardsley J, Mertz D, et al. Probiotics for the prevention of Clostridium difficile-associated diarrhea in adults and children. Cochrane Database Syst Rev. (2017) 12:Cd006095. doi: 10.1002/14651858.CD006095.pub4
229. Fooks LJ, Gibson GR. In vitro investigations of the effect of probiotics and prebiotics on selected human intestinal pathogens. FEMS Microbiol Ecol. (2002) 39:67–75. doi: 10.1111/j.1574-6941.2002.tb00907.x
230. Salminen S, Collado MC, Endo A, Hill C, Lebeer S, Quigley EMM, et al. The International Scientific Association of Probiotics and Prebiotics (ISAPP) consensus statement on the definition and scope of postbiotics. Nat Rev Gastroenterol Hepatol. (2021) 18:649–67. doi: 10.1038/s41575-021-00440-6
231. Le Lay C, Fernandez B, Hammami R, Ouellette M, Fliss I. On Lactococcus lactis UL719 competitivity and nisin (Nisaplin(®)) capacity to inhibit Clostridium difficile in a model of human colon. Front Microbiol. (2015) 6:1020. doi: 10.3389/fmicb.2015.01020
232. Vukomanović M, Žunič V, Kunej Š, Jančar B, Jeverica S, Podlipec R, et al. Nano-engineering the antimicrobial spectrum of Lantibiotics: activity of nisin against gram negative bacteria. Sci Rep. (2017) 7:4324. doi: 10.1038/s41598-017-04670-0
233. Brunati C, Thomsen TT, Gaspari E, Maffioli S, Sosio M, Jabes D, et al. Expanding the potential of NAI-107 for treating serious ESKAPE pathogens: synergistic combinations against Gram-negatives and bactericidal activity against non-dividing cells. J Antimicrob Chemother. (2018) 73:414–24. doi: 10.1093/jac/dkx395
234. Gregory AL, Pensinger DA, Hryckowian AJ. A short chain fatty acid-centric view of Clostridioides difficile pathogenesis. PLoS Pathog. (2017) 17:e1009959. doi: 10.1371/journal.ppat.1009959
235. Sorg JA, Sonenshein AL. Bile salts and glycine as cogerminants for Clostridium difficile spores. J Bacteriol. (2008) 190:2505–12. doi: 10.1128/jb.01765-07
236. Van Der Waaij D, Vossen JM, Altes CK, Hartgrink C. Reconventionalization following antibiotic decontamination in man and animals. Am J Clin Nutr. (1977) 30:1887–95. doi: 10.1093/ajcn/30.11.1887
237. Green JE, Davis JA, Berk M, Hair C, Loughman A, Castle D, et al. Efficacy and safety of fecal microbiota transplantation for the treatment of diseases other than Clostridium difficile infection: a systematic review and meta-analysis. Gut Microbes. (2020) 12:1–25. doi: 10.1080/19490976.2020.1854640
238. Baunwall SMD, Andreasen SE, Hansen MM, Kelsen J, Høyer KL, Rågård N, et al. Faecal microbiota transplantation for first or second Clostridioides difficile infection (EarlyFMT): a randomised, double-blind, placebo-controlled trial. Lancet Gastroenterol Hepatol. (2022) 7:1083–91. doi: 10.1016/s2468-1253(22)00276-x
239. Orenstein R, Dubberke ER, Khanna S, Lee CH, Yoho D, Johnson S, et al. Durable reduction of Clostridioides difficile infection recurrence and microbiome restoration after treatment with RBX2660: results from an open-label phase 2 clinical trial. BMC Infect Dis. (2022) 22:245. doi: 10.1186/s12879-022-07256-y
240. Walter J, Shanahan F. Fecal microbiota-based treatment for recurrent Clostridioides difficile infection. Cell. (2023) 186:1087. doi: 10.1016/j.cell.2023.02.034
241. Benech N, Barbut F, Fitzpatrick F, Krutova M, Davies K, Druart C, et al. Update on microbiota-derived therapies for recurrent Clostridioides difficile infections. Clin Microbiol Infect. (2024) 30:462–68. doi: 10.1016/j.cmi.2023.12.007
242. Manges AR, Steiner TS, Wright AJ. Fecal microbiota transplantation for the intestinal decolonization of extensively antimicrobial-resistant opportunistic pathogens: a review. Infect Dis (Lond). (2016) 48:587–92. doi: 10.1080/23744235.2016.1177199
243. Kim SM, Defazio JR, Hyoju SK, Sangani K, Keskey R, Krezalek MA, et al. Fecal microbiota transplant rescues mice from human pathogen mediated sepsis by restoring systemic immunity. Nat Commun. (2020) 11:2354. doi: 10.1038/s41467-020-15545-w
244. Dinh A, Fessi H, Duran C, Batista R, Michelon H, Bouchand F, et al. Clearance of carbapenem-resistant Enterobacteriaceae vs vancomycin-resistant enterococci carriage after faecal microbiota transplant: a prospective comparative study. J Hosp Infect. (2018) 99:481–86. doi: 10.1016/j.jhin.2018.02.018
245. Shin J, Lee JH, Park SH, Cha B, Kwon KS, Kim H, et al. Efficacy and safety of fecal microbiota transplantation for clearance of multidrug-resistant organisms under multiple comorbidities: A prospective comparative trial. Biomedicines. (2022) 10:2404. doi: 10.3390/biomedicines10102404
246. Bilinski J, Grzesiowski P, Sorensen N, Madry K, Muszynski J, Robak K, et al. Fecal microbiota transplantation in patients with blood disorders inhibits gut colonization with antibiotic-resistant bacteria: results of a prospective, single-center study. Clin Infect Dis. (2017) 65:364–70. doi: 10.1093/cid/cix252
247. Wei Y, Gong J, Zhu W, Guo D, Gu L, Li N, et al. Fecal microbiota transplantation restores dysbiosis in patients with methicillin resistant Staphylococcus aureus enterocolitis. BMC Infect Dis. (2015) 15:265. doi: 10.1186/s12879-015-0973-1
248. Bier N, Hanson B, Jiang ZD, Dupont HL, Arias CA, Miller WR. A case of successful treatment of recurrent urinary tract infection by extended-spectrum β-lactamase producing Klebsiella pneumoniae using oral lyophilized fecal microbiota transplant. Microb Drug Resist. (2023) 29:34–8. doi: 10.1089/mdr.2022.0031
249. Defilipp Z, Bloom PP, Torres Soto M, Mansour MK, Sater MRA, Huntley MH, et al. Drug-resistant E. coli bacteremia transmitted by fecal microbiota transplant. N Engl J Med. (2019) 381:2043–50. doi: 10.1056/NEJMoa1910437
250. Pamer EG. Fecal microbiota transplantation: effectiveness, complexities, and lingering concerns. Mucosal Immunol. (2014) 7:210–4. doi: 10.1038/mi.2013.117
251. Danne C, Rolhion N, Sokol H. Recipient factors in faecal microbiota transplantation: one stool does not fit all. Nat Rev Gastroenterol Hepatol. (2021) 18:503–13. doi: 10.1038/s41575-021-00441-5
252. Anand T, Virmani N, Kumar S, Mohanty AK, Pavulraj S, Bera BC, et al. Phage therapy for treatment of virulent Klebsiella pneumoniae infection in a mouse model. J Glob Antimicrob Resist. (2020) 21:34–41. doi: 10.1016/j.jgar.2019.09.018
253. Li Z, Li W, Ma W, Ding Y, Zhang Y, Yang Q, et al. Characterization and application of a lytic phage D10 against multidrug-resistant. Salmonella Viruses. (2021) 13:1626. doi: 10.3390/v13081626
254. Weber-Dąbrowska B, Żaczek M, Łobocka M, Łusiak-Szelachowska M, Owczarek B, Orwat F, et al. Characteristics of Environmental Klebsiella pneumoniae and Klebsiella oxytoca Bacteriophages and Their Therapeutic Applications. Pharmaceutics. (2023) 15:434. doi: 10.3390/pharmaceutics15020434
255. Wang L, Tkhilaishvili T, Jiang Z, Pirlar RF, Ning Y, Millán Laleona A, et al. Phage-liposome nanoconjugates for orthopedic biofilm eradication. J Control Release. (2024) 376:949–60. doi: 10.1016/j.jconrel.2024.09.049
256. Pirnay JP, Djebara S, Steurs G, Griselain J, Cochez C, De Soir S, et al. Personalized bacteriophage therapy outcomes for 100 consecutive cases: a multicentre, multinational, retrospective observational study. Nat Microbiol. (2024) 9:1434–53. doi: 10.1038/s41564-024-01705-x
257. Forti F, Roach DR, Cafora M, Pasini ME, Horner DS, Fiscarelli EV, et al. Design of a broad-range bacteriophage cocktail that reduces Pseudomonas aeruginosa biofilms and treats acute infections in two animal models. Antimicrob Agents Chemother. (2018) 62:e02573-17. doi: 10.1128/aac.02573-17
258. Alves DR, Perez-Esteban P, Kot W, Bean JE, Arnot T, Hansen LH, et al. A novel bacteriophage cocktail reduces and disperses Pseudomonas aeruginosa biofilms under static and flow conditions. Microb Biotechnol. (2016) 9:61–74. doi: 10.1111/1751-7915.12316
259. Liu S, Hon K, Bouras GS, Psaltis AJ, Shearwin K, Wormald PJ, et al. APTC-C-SA01: A novel bacteriophage cocktail targeting Staphylococcus aureus and MRSA biofilms. Int J Mol Sci. (2022) 23:6116. doi: 10.3390/ijms23116116
260. Joo H, Wu SM, Soni I, Wang-Crocker C, Matern T, Beck JP, et al. Phage and antibiotic combinations reduce Staphylococcus aureus in static and dynamic biofilms grown on an implant material. Viruses. (2023) 15:460. doi: 10.3390/v15020460
261. Mu A, Mcdonald D, Jarmusch AK, Martino C, Brennan C, Bryant M, et al. Assessment of the microbiome during bacteriophage therapy in combination with systemic antibiotics to treat a case of staphylococcal device infection. Microbiome. (2021) 9:92. doi: 10.1186/s40168-021-01026-9
262. Duerkop BA, Clements CV, Rollins D, Rodrigues JL, Hooper LV. A composite bacteriophage alters colonization by an intestinal commensal bacterium. Proc Natl Acad Sci U S A. (2012) 109:17621–6. doi: 10.1073/pnas.1206136109
263. Badawy S, Pajunen MI, Haiko J, Baka Z, Abou-Dobara MI, El-Sayed AKA, et al. Identification and functional analysis of temperate siphoviridae bacteriophages of Acinetobacter baumannii. Viruses. (2020) 12:604. doi: 10.3390/v12060604
264. Morello E, Saussereau E, Maura D, Huerre M, Touqui L, Debarbieux L. Pulmonary bacteriophage therapy on Pseudomonas aeruginosa cystic fibrosis strains: first steps towards treatment and prevention. PLoS One. (2011) 6:e16963. doi: 10.1371/journal.pone.0016963
265. Fong SA, Drilling A, Morales S, Cornet ME, Woodworth BA, Fokkens WJ, et al. Activity of bacteriophages in removing biofilms of Pseudomonas aeruginosa isolates from chronic rhinosinusitis patients. Front Cell Infect Microbiol. (2017) 7:418. doi: 10.3389/fcimb.2017.00418
266. Martinet MG, Lohde M, Higazy D, Brandt C, Pletz MW, Middelboe M, et al. Diversification of Pseudomonas aeruginosa Biofilm Populations under Repeated Phage Exposures Decreases the Efficacy of the Treatment. Microorganisms. (2024) 12:1880. doi: 10.3390/microorganisms12091880
267. Nath A, Bhattacharjee R, Nandi A, Sinha A, Kar S, Manoharan N, et al. Phage delivered CRISPR-Cas system to combat multidrug-resistant pathogens in gut microbiome. BioMed Pharmacother. (2022) 151:113122. doi: 10.1016/j.biopha.2022.113122
268. Fang Q, Feng Y, Mcnally A, Zong Z. Characterization of phage resistance and phages capable of intestinal decolonization of carbapenem-resistant Klebsiella pneumoniae in mice. Commun Biol. (2022) 5:48. doi: 10.1038/s42003-022-03001-y
269. Corbellino M, Kieffer N, Kutateladze M, Balarjishvili N, Leshkasheli L, Askilashvili L, et al. Eradication of a multidrug-resistant, carbapenemase-producing Klebsiella pneumoniae isolate following oral and intra-rectal therapy with a custom made, lytic bacteriophage preparation. Clin Infect Dis. (2020) 70:1998–2001. doi: 10.1093/cid/ciz782
Keywords: gut microbiota, antimicrobial resistance, colonization resistance, bacteriophage, probiotics
Citation: Ding W, Cheng Y, Liu X, Zhu Z, Wu L, Gao J, Lei W, Li Y, Zhou X, Wu J, Gao Y, Ling Z and Jiang R (2025) Harnessing the human gut microbiota: an emerging frontier in combatting multidrug-resistant bacteria. Front. Immunol. 16:1563450. doi: 10.3389/fimmu.2025.1563450
Received: 20 January 2025; Accepted: 25 February 2025;
Published: 17 March 2025.
Edited by:
Nar Singh Chauhan, Maharshi Dayanand University, IndiaReviewed by:
Manu Bhambi, Zigzoo Marketing Services Pvt Ltd, IndiaCopyright © 2025 Ding, Cheng, Liu, Zhu, Wu, Gao, Lei, Li, Zhou, Wu, Gao, Ling and Jiang. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Ruilai Jiang, cnVpbGFpX2ppYW5nQHllYWgubmV0; Zongxin Ling, bGluZ3pvbmd4aW5Aemp1LmVkdS5jbg==; Yongtao Gao, eW9uZ3Rhb19nYW9AeWVhaC5uZXQ=; Jian Wu, d3VqaWFuZ2xpbnhpbmdAMTYzLmNvbQ==
†These authors have contributed equally to this work
†ORCID: Zongxin Ling, orcid.org/0000-0001-9662-099X
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.