
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
REVIEW article
Front. Immunol. , 24 March 2025
Sec. T Cell Biology
Volume 16 - 2025 | https://doi.org/10.3389/fimmu.2025.1563150
Dynamic changes in various forms of RNA modification are critical to the functional homeostasis of the immune system and the pathophysiology of autoimmune diseases. RNA modification-related proteins play an essential role in these processes. At present, the research methods of RNA modification in autoimmune diseases are mainly to detect the expression changes of RNA modification-related proteins in tissues or cells, but there is a lack of explorations of target RNAs and in-depth mechanisms. Considering the important role of CD4+ T cell dysfunction in the pathogenesis and progression of autoimmune diseases, the regulatory effect of abnormal RNA modification on CD4+ T cells deserves attention, which will provide a perspective for further exploring the mechanism of RNA modification in autoimmune diseases. In this Review, we discuss the abnormal RNA modification changes in patients with autoimmune diseases and highlight the effects of these abnormal changes on CD4+ T cells.
● The RNA modification related enzymes may play opposite roles in different autoimmune diseases.
● RNA modification related enzymes exhibit distinct functions across various tissues and cells in autoimmune diseases.
● RNA modification related enzymes play a crucial role in the differentiation of CD4+ T cells.
● Targeting RNA modifications may be an emerging approach for treatment of autoimmune diseases.
RNA modifications refer to the chemical labeling of bases or ribose in RNA molecules, involving a selective addition of methyl groups (1). The advancements in transcriptome technology have revealed over 170 chemical modifications in RNA molecules, with methylation being the most predominant type (2). The process of RNA methylation takes place in various kinds of coding and non-coding RNAs, including transfer RNA (tRNA), ribosomal RNA (rRNA), small nuclear RNA (snRNA) and long non-coding RNA (lncRNA). Common modifications found in RNA include N6-methyladenosine (m6A), 2’-O-methylation (Nm), and N6-2’-O-dimethyladenosine (m6Am). Additionally, there are N1-methyladenosine (m1A), 5-methylcytidine (m5C) and N7-methylguanosine (m7G) (3). The cellular functions of RNA, such as splicing, stability, and translation, are regulated by RNA methylations through their respective regulators known as “writers” (methyltransferases), “erasers” (demethylases), and “readers” (methylation recognition proteins) (4). The characteristics of the major types of RNA modification related proteins have been summarized (Supplementary Table 1). Abnormal RNA modifications are closely related to the occurrence and development of many diseases, including cancer, arteriosclerosis, and autoimmune diseases (5). Currently, many scholars focus on the diverse range of biological processes, yet the understanding about the regulatory role of RNA modifications in immune system remains limited.
CD4+ T cells are a prominent subset of T lymphocytes, characterized by their abundant presence and diverse functional states (6). They primarily regulate and activate immune responses through antigen recognition and cytokine secretion. The pivotal roles of CD4+ T cells in maintaining immune system homeostasis are valued by their associations with various diseases, including inflammatory and autoimmune diseases, such as systemic lupus erythematosus (SLE), rheumatoid arthritis (RA) and inflammatory bowel disease (IBD). The CD4+ T cells regulate the occurrence and development of autoimmune diseases through various mechanisms, such as assisting cytokine production and regulating immune response and memory (7). It is of great significance to deeply understand the function and regulatory mechanism of CD4+ T cells for further exploring the pathophysiological process of autoimmune diseases and developing new methods of diagnosis and treatment. However, the understanding of which molecular biological processes can mediate the function of CD4+ T cells is far more than that, among which the role of RNA modification on their homeostasis and differentiations cannot be ignored.
In this Review, we explore the abnormal RNA modification changes in patients with autoimmune diseases and highlight the effects of these changes on CD4+ T cells (Figure 1). In addition, we discuss the regulatory effects about RNA modification-related proteins on CD4+ T cell differentiation, and point out the key points worthy of further study in the future.

Figure 1. RNA modifications interact with autoimmune disease progression and CD4+ T cell homeostasis and differentiation. RNA modification is mediated by writers, erasers and readers. Abnormal RNA modifications are found in peripheral blood, PBMCs, focal tissues and cells in patients with autoimmune diseases or animal disease models. These changes in RNA modification may lead to abnormal CD4+ T cell homeostasis and function, which may aggravate the progression of autoimmune diseases. m6A, N6-methyladenosine; m6Am, N6-2’-O-dimethyladenosine; m1A, N1-methyladenosine; m5C, 5-methylcytidine; PBMCs, peripheral blood mononuclear cells; FLSs, fibroblast-like synovial cells; IL-17, interleukin-17; IL-22, interleukin-22; IL-10, interleukin-10; IL-4, interleukin-4; IL-5, interleukin-5; IL-9, interleukin-9; IL-13, interleukin-13; TGF-β, transforming growth factor-beta; TNF-α, tumor necrosis factor-alpha; IFN-γ, interferon gamma; Th, helper T cells; Tfh, T follicular helper cells; Tregs, regulatory T cells.
The m6A modification occurs at the 6th position of the adenine ring in RNA as the most prevalent and abundant post-transcriptional modification in eukaryotic mRNA (1). The m6A modification is predominantly localized within the 3’-untranslated region (3’-UTR) proximal to the coding sequence (CDS) and stops the codon of mRNA, highlighting the crucial role in mRNA biology (8, 9).
The m6A “writers” complex mainly comprises three core proteins: methyltransferase-like 3 (METTL3), METTL14, and Wilms’ tumor-associated protein (WTAP) (2). METTL14 and METTL3 form a heterodimer, and METTL14 is primarily responsible for recognizing and localizing subunits (10). The WTAP facilitates m6A modification by recruiting co-localized METTL3-METTL14 heterodimers within nucleolar patches. Other “writers” complex include METTL5, METTL16, RNA binding motif protein 15/15B (RBM15/15B), vir-like m6A methyltransferase associated (VIRMA), and Zinc finger CCCH-type containing 13 (ZC3H13) (11), which are crucial for the nuclear localization and stabilization.
The two “erasers” include the Fat mass and obesity-associated protein (FTO) and AlkB homologue 5 (ALKBH5), which can mediate reversible demethylation (12). The three “readers” groups include the YTH-RNA binding domain family (YTHDF), consisting of YTHDF1/2/3 and YTHDC1/2, the heterogeneous nuclear ribonucleoprotein (HNRNP) family and insulin-like growth factor-2 mRNA-binding proteins family (IGF2BPs), consisting of IGF2BP1/2/3 (13).The interactions between YTHDF3 and YTHDF1 enhance RNA translation, while the binding of YTHDF3 to YTHDF2 promotes RNA degradation (14). YTHDF1 fosters the translation of mRNA while YTHDF2 encourages the degradation of mRNA, but the mechanism remains unclear (14). IGF2BP2 stabilizes the mRNAs containing m6A and facilitates their translation (15).
The m6Am is an RNA modification located immediately after the m7G cap structure of mRNA, specifically on the first nucleotide (16).
The m6Am “writers” complex comprises phosphorylated CTD-interacting factor 1 (PCIF7) and METTL4. It’s reported that the m6A demethylase FTO could also demethylate the m6Am (17). The m6Am differs from m6A solely in the methylation on the 2’-O position of the ribose. Despite this distinction, the overall structures of m6Am and m6A are highly similar. (17). Due to the similar chemical structure of m6Am and m6A, m6A-specific antibodies often recognize m6Am as well (16). The localizations of FTO are dependent on the cell cycle stages and regulated by Casein kinase II-mediated phosphorylation. Since the higher m6A levels in cells than m6Am, the effects of FTO are attributed to m6A activity (18).
The m1A researches have revealed predominant distributions in tRNAs and rRNAs (19). The methyltransferases, known as “writers”, include TRMT6, TRMT61A/B, TRMT10C, and NML (20). The demethylases, referred to as “erasers”, include ALKBH1/3/7 and FTO, and the “readers” include YTHDF1/2/3 and YTHDC1 (21).
The TRMT6/61A complex, located in the cytoplasm, can methylate cellular tRNA m1A58 (T-loop) and mRNA with T-loop-like structure (20). The NML, localized in the nucleus, can methylate m1A on 28S rRNA (22). ALKBH1 demethylates most m1A in cyto-tRNAs with the m1A58 in tRNAs as a significant substrate (23).YTHDF1/2/3 and YTHDC1 can recognize the common modification m6A (20). However, their binding affinity to m1A is weaker than that of m6A. Researches on their function specifically as “readers” of m1A are still limited.
The m5C refers to the process by which methyltransferase uses SAM as a methyl donor and transfers methyl groups to cytosine. There are known m5C methyltransferases, including the NOL1/NOP2/sun domain (Nsun) family, DNA methyltransferase (DNMT) homologs and tRNA-specific methyltransferase (TRDMT) families (24). The majority of tRNAs are methylated by Nsun2 at the variable loop, and specifically in leucine at the wobble position (24).
To date, the role of m5C “erasers” remains a subject of controversy. Enzymes belonging to the ten-eleven translocator family (TET) have been reported to oxidize m5C in mRNA (25), resulting in the production of 5-hydroxymethylcytosine (hm5C) (25). Additionally, ALKBH1 catalyzes the formation of oxidation of 5-formylcytosine (f5C) at the mitochondrial tRNA wobble position (24). Although the f5C depositions in mitochondrial tRNAs are well-established biologically, the biological relevance in terms of hm5C deposition in mRNAs remains unclear (26). Aly/REF export factor (ALYREF) and Y-box-binding protein 1 (YBX1) appear to be m5C “reader” proteins that govern mRNA fate based on its m5C status (27).
According to locations, there are two types of m7G modification. The one type is m7G-cap at 5’-terminal and the other is internal m7G modification (28). METTL1, WD repeat domain 4 (WDR4), RNMT and RAM as “writers” can catalyze m7G modification (29). However, other enzymes still need to be identified in the future.
The known 2’-O-methylation (Nm, where N represents any nucleotide) involves the addition of a methyl group (-CH3) to the 2’-hydroxyl (-OH) of the ribose moiety. This modification is highly conserved and abundant, occurring at multiple sites in tRNA, rRNA, and snRNA (30).
The Nm “writers” FtsJ RNA 2’-O-methyltransferase 1 (FTSJ1) exhibits high expression levels in tRNAs, while fibrillarin shows elevated expression levels in rRNAs, both of which have significant promoting effects on cellular growth. The Nm “erasers” and “readers” were unknown (31). The enrichments of Nm in rRNAs confer protection against hydrolysis and enhance the stability of their conformation.
RNA modifications encompass various stages of RNA metabolism, such as RNA splicing, transport, editing, transcription, translation and degradation. Currently, primary emphasis in the field of RNA modifications lies within tumor studies, while autoimmune diseases have received comparatively less attention. Then, we summarize the role of RNA modification in autoimmune diseases and explore possible therapeutic ideas.
SLE is a complex autoimmune disease involving multiple systems, characterized by the production of numerous autoantibodies, loss of tolerance, and tissue damage (32). The involvement of m6A modification-related proteins in the regulation of SLE disease progression is multifaceted. The mRNA levels of METTL3, WTAP, ALKBH5, FTO and YTHDF2 in peripheral blood of SLE patients are significantly lower than healthy controls, while METTL14 is unchanged (33). The decreased mRNA level of ALKBH5 in peripheral blood is a risk factor of SLE, which are associated with anti-dsDNA and anti-nucleosome, rash (33). However, these changes are not exactly the same in peripheral blood mononuclear cells (PBMCs). The mRNA expression of METTL14, ALKBH5, and YTHDF2 in PBMCs of SLE patients is decreased, while others are unchanged (34). The decreased level of YTHDF2 in PBMCs is a risk factor for disease progression (34). Although the causes are complex, the reason for these differences can lie in neutrophils. Neutrophils are crucial players in SLE pathogenesis, high numbers of immature neutrophils and dysregulated neutrophil death present in the blood of SLE patients (35). The activation of neutrophils can be regulated by METTL3-mediated m6A modification (36). Although the underlying mechanism remains unclear, it is likely that the specific transcriptome locations and downstream targets play a pivotal role in determining its functionality.
RA is an autoimmune disease characterized by symmetrical inflammations in bilateral joints, leading to persistent pain and causing damage to bone and cartilage tissues (37). Researchers evaluate the levels of whole-transcriptome m6A modification in the synovium from RA patients and healthy controls, and find that the m6A is closely related to RA inflammation (37). The m6A methylation level is increased in synovial tissues and fibroblast-like synovial cells (FLSs) from RA patients (38).
In comparison to normal controls, the level of METTL3 mRNA is up-regulated in PBMCs, macrophages, synovial tissues, and FLSs from patients with RA (38–40) Elevated METTL3 mRNA level in PBMCs may indicate high disease activity (39). METTL3 promotes the tumor-like growth of RA-FLSs (38). The proliferation and invasion of METTL3-deficient FLSs are significantly attenuated (40). The NF-κB signaling pathway exerts a crucial influence on these effects through the regulation of the secretion of inflammatory mediators (39, 41). Some drugs like Artemisitene can inhibit RA progression by targeting METTL3-mediated m6A modification (42, 43).
Surprisingly, METTL14 seems to have the opposite effect to METTL3 in PBMCs. The mRNA level of METTL14 is decreased in PBMCs of RA, and the inhibitory factors of NF-κB inflammatory pathway are methylated (44). Although it is difficult to explain the mechanism, the varying stages of disease progression in patients may constitute a contributing factor. Similarly, the opposite effects of METTL3 and METTL14 are also present in the course of hepatocellular carcinoma (45). The expression of METTL14 is up-regulated in macrophages, synovial tissues, and FLSs from patients with RA (38, 46, 47). These may be related to METTL14 mRNA enhancing the migration and invasion ability of FLSs and promoting the M1 polarization of macrophage (46, 47). The mRNA level of METTL14 in peripheral blood is increased (46) or without significant change (48), which may be due to the different stages of disease progression in patients. WTAP is up-regulated in synovial tissue and FLSs from patients with RA (38). WTAP promotes FLSs pyroptosis and related inflammatory responses via NLRP3 (49). However, the peripheral blood mRNA level of METTL3, METTL14, and WTAP reveals no significant differences from RA patients and healthy individuals (48). The causes are complex, potentially due to the association with neutrophils. Neutrophil derived reactive oxygen species (ROS) and granuloprotease cause cartilage injury in RA and are associated with pathogenic post-translational modification (35). METTL3, METTL14 and WTAP may be expressed in neutrophils and play a complex role in regulating the disease progression of RA.
ALKBH5 and FTO are upregulated in FLSs and synovial tissue from RA patients (50, 51). The JARID2, one of the targeting mRNA of ALKBH5-mediated m6A modification, can negatively regulate the migration, invasion and proliferation of FLSs (50). FTO promotes ADAMTS15 mRNA stability in an m6A-IGF2BP1 dependent manner, which negatively regulates the inflammatory response, migration and invasion of FLSs (51). The upregulation of ALKBH5 in FLSs can enhance the proliferation, migration, and inflammatory response (52), while these capabilities are completely reversed in the absence of FTO (51). However, the mRNA level of ALKBH5 and FTO is significant downregulated in peripheral blood from RA patients (48). The changes in ALKBH5 and FTO, whether increasing or decreasing, exhibit a consistent trend, suggesting their potential synergistic role. The aforementioned findings indicate the variations in RNA modification-related proteins across different cells in RA, while the specific methylated genes that play a crucial role remain unknown.
The mRNA level of YTHDF2 in peripheral blood and PBMCs of RA is significantly decreased (48, 53). The expression of YTHDF2 mRNA is negatively correlated with multiple inflammatory markers (53). The NLRP3 is a target gene on ALKBH5/YTHDF2- mediated m6A modification, which has been shown to be closely related to RA (54). The level of IGF2BP3 mRNA is increased in synovial tissue and PBMCs of RA, while after treatment, IGF2BP3 mRNA level is decreased, which suggests that IGF2BP3 might be a new potential target during the treatment of RA (55). The mRNA level of IGF2BP2 is increased in synovial tissue of RA (56).
IBD, consisting of Crohn’s disease (CD) and ulcerative colitis (UC), is a chronic, relapsing inflammatory disorder (57).
The m6A-methyltransferases in IBD exhibit similar pro-inflammatory effects. METTL3 is up-regulated in the pathological tissues of IBD patients. The inhibition of Mettl3 or Mettl14 in intestinal epithelial cells (IECs) can effectively inhibit NF-κB signaling and significantly relieve inflammation (58, 59), which demonstrates the synergistic effect of Mettl3 and Mettl14. Similarly, the deficiency of Mettl3 in macrophages can ameliorate dextran sulphate sodium (DSS)-induced colitis in mice (60). Mechanistically, the depletion of Mettl3 enhances the Ythdf3-mediated PGP expression, resulting in the suppression of Th1 differentiation (60). Moreover, Wtap can promote the disease progression of DSS-induced IBD in mice (61).
The expression of ALKBH5 mRNA in intestinal mucosa of RA is contradictory and complex, due to the different stages of disease progression, the consistent conclusion could not be reached. The mRNA level of FTO is reduced in intestinal mucosa of RA and negatively correlated with Mayo score (62). FTO seems to exhibit mainly anti-inflammatory effects. Fto deficiency leads to more severe colitis in IECs under DSS induce (62).
IGF2BP1 and IGF2BP2 are down-regulated in CD or UC tissues compared to normal tissues (63). However, IGF2BP1 and IGF2BP2 appear to play an anti-inflammatory role. The Igf2bp2-knockout macrophages exhibit the enhanced M1 phenotype and promote DSS-induced colitis development (64).
The rheumatic chronic inflammatory disease known as Ankylosing spondylitis (AS) is characterized by inflammatory spondylitis, peripheral arthritis, and enthesitis (65). The mRNA expression of YTHDF2 and ALKBH5 in PBMCs of newly diagnosed AS patients is significantly decreased, and the expression of YTHDF2 mRNA in PBMCs is a risk factor for AS (66). The total m6A level and METTL14 expression are decreased in T cells of AS. Mechanistically, the expression of FOXO3a reduces with the decrease of METTL14, leading to increased autophagy and inflammation (67). The mRNA level of WTAP in PBMCs of AS patients is up-regulated. HNRNPC is down-regulated, which are potential m6A regulators of AS (65). However, the RNAs and signaling pathways targeted by these proteins remain unclear.
Primary Sjogren’s syndrome (pSS) is a chronic autoimmune disease of unknown etiology, defined by dry mouth syndrome and dry keratoconjunctivitis (68). The m6A, m1A, m5C, and m7G modification play a critical role in the diversity and complexity of the SS immune microenvironment (69). The total m6A level and METTL3 expression are increased in PBMCs of SS, and the upregulation of METTL3 is associated with disease severity (68). The elevated ALKBH5 mRNA in PBMCs of SS patients may be a risk factor (70).The mRNA levels of METTL3, ALKBH5 and YTHDF1 in T cells of SS patients are significantly up-regulated (71). The mRNA level of ALKBH5, RBMX, RBM15B and YTHDF1 is down-regulated in peripheral blood and labial salivary gland tissues of SS patients (72). METTL3 and METTL14 are elevated in salivary gland epithelial cells of SS patients, but METTL3 may play an anti-inflammatory role. The inhibition of METTL3 increased inflammatory related-gene expression, whereas inhibition of FTO presents the opposite effect (73). However, there is a lack of in vitro experiments to explore the mechanism.
YTHDF2 is significantly up-regulated in the liver of autoimmune hepatitis (AIH) patients, which is related to the degree of inflammation, and RXRα is one of the target RNA for this experiment (74). METTL3 is increased in pancreatic-β cells from patients with type 1 diabetes (T1D), the genes associated with innate immune mediators including OAS1, OAS2, OAS3, and ADAR1 exhibit hypermethylation (75).
In summary, RNA modification-related enzymes are changed in autoimmune diseases, and the role of RNA modification in autoimmune diseases deserves attention (Supplementary Table 2). Current researches have mainly concentrated on alterations in the expression of RNA modification enzymes in cells, tissues, and organs, while neglecting the upstream and downstream mechanism associated with changes in enzyme activity (33, 34, 46, 48). However, further investigations into the underlying mechanism are limited. Firstly, RNA modification-related enzymes may change differently with disease progression (46, 48). The mRNA level of METTL14 in PBMCs from SLE patients is significantly reduced, whereas it remains unchanged in the overall peripheral blood (33, 34). These two articles are from the same team, and it can be inferred from the above conclusions that the level of METTL14 mRNA in neutrophils and red blood cells is different in patients and healthy controls. In other words, the extent of RNA modification may vary among different cell types, which should be considered when developing therapeutic strategies targeting RNA modifications for disease treatment in the future. These regulatory proteins such as METTL3, METTL14 and WTAP may act simultaneously or independently, and it is necessary to explore the dominant actors causing the changes. However, the relevant researches are limited at present, which is worthy of further exploration. Additionally, existing studies have primarily focused on investigating alterations in the expression of RNA modification enzymes in cells, tissues, and organs from both patient and healthy control samples. However, the upstream and downstream mechanisms responsible for these changes in enzyme activity remain to be elucidated. Investigating the localization of RNA modifications within the transcriptome can enhance our understanding of how these proteins impact disease progression and enable exploration into upstream factors regulating these enzyme modifications. Further investigations both in vitro and in vivo are necessary to determine the localization of different RNA modifications within multiple cellular transcriptomes from patients with autoimmune diseases, to identify target RNAs affected by these modifications, and to assess their influence on disease activity as well as drug toxicity and organ damage. Lastly, current researchers focus on m6A modification that involves more than twenty regulatory proteins, only METTL3, METTL14, ALKBH5, and FTO have been elucidated thus far. The role of other RNA modification needs further investigation regarding autoimmune diseases.
The abnormality in immune cell function and composition in human diseases have been a focal point of recent researches, and significant advancements have been made. CD4+ T cells, which differentiates into helper T cells (Th), T follicular helper (Tfh) and regulatory T cells (Tregs), play an essential role in autoimmune diseases. The balance of Th1/Th2 and Th17/Treg has attracted a lot of interest over the past decades. Aspects of the imbalance between Th1/Th2 and Th17/Treg in autoimmune disease are summarized in the Reviews (76, 77). We will not discuss the above topics here. Our team is mainly committed to exploring the study of Th17 and myeloid-derived suppressor cells (MDSC) on autoimmune diseases (78). Next, we will mainly review the effect of RNA modification on CD4+ T cell function.
Recent research shows that the increased level of Th17 cells in the circulation and tissues of SLE patients is closely associated with the defects of apoptosis (79). Current researches on autoimmune disease involve the Th17 differentiation signaling pathway regulated by methylation. For example, the hypomethylation and hypermethylation of Th17-related genes serve as indicators for the presence of intestinal CD4+ T cells in patients with CD (80). SMARCA5, belonging to the SWI/SNF family, can promote the development of UC through the methylation processes (81). The recent findings demonstrate that RNF180 can exacerbate colon inflammation and disrupt the Th17/Treg cell balance with UC by modulating the ALKBH5/SMARCA5 axis (82).
The emerging T cell subsets such as Th9 and Th22 cells, link the inflammatory factors to autoimmune disease. The increased expression of IL-9 in RA is associated with heightened disease severity (83). The transcription factor PU.1 can directly target FMS like-tyrosinekinase-3 (FLT3) in macrophages and FLSs to promote the development of RA (84). In SLE patients, the expansion of CXCL13+ Tph/Tfh cells is associated with disease activity, while IL-22+ Th22 cells decrease (85). The formation of AhR and JUN in CD4+ T cells inhibits the expression of CXCL13 and induces Th22 differentiation (85). However, there is no direct evidence that Th9 and Th22 are regulated by RNA modification.
In the germinal centers (GCs), Tfh regulates B cell maturation triggered by T cells (86). Tfh cells secrete IL-21, which is to be crucial for supporting GCs. The mRNA expression of transcription factor Bcl-6 is elevated in patients with RA and IBD (87). The defective ubiquitination of CBL and CBLB leads to impaired Bcl-6 function, resulting in the dysregulation of Tfh cells in SLE patients (88). Previous studies indicate that the regulation of Bcl-6 involves histone methylation, acetylation (89) and β-hydroxybutyrylation modification (90).
In conclusion, CD4+ T cells are the central “regulators” of the immune system and play an important role in autoimmune diseases. Prior researches have demonstrated that CD4+ T cells undergo epigenetic modifications. Although these are preliminary explorations, they establish a foundational understanding of the role of RNA modifications in CD4+ T cells. Next, we delve into the role of RNA modification-associated proteins on CD4+ T cell homeostasis and differentiation.
Researchers have examined overall changes in RNA modification in patients with autoimmune diseases. The increased levels of Am, m1A, m6A, and 3 ‘OMeA in CD4+ T cells from SLE patients, while decreased levels of m3C, m5C, m1G, m5U, and t6A compared with healthy controls (91). In this study, the specific roles of various regulatory proteins remain unclear. Next, we will comprehensively examine the impact of these RNA modification-related proteins on CD4+ T cells, categorized by the three types.
Different from previous research (91), the total m6A level of CD4+ T cells is reduced in SLE patients compared with healthy controls, which is mainly caused by the decreased mRNA level of METTL3. The degree of reduced METTL3 is correlated with disease activity (92). Although these researches are difficult to explain mechanistically, it is possible that distinct subpopulations of CD4+ T cells undergo different methylation changes. In other words, the same RNA modification protein may exert diverse effects in different types of CD4+ T cells.
Mettl3 seems to exhibit important pro-inflammatory or anti-inflammatory effects in different types of CD4+ T cells. The proportion of Th2 cells in Mettl3-knockout CD4+ T cells is increased, while the Th17 is decreased (93). The Mettl3-knockout naïve T cells lose the ability to induce colitis phenotypes (93). The mechanisms are involved in the elevated levels of Janus kinase (JAK)-STAT5 signaling pathway related genes, such as SOCS1, SOCS3, and CISH. These genes belong to the SOCS family, targeted by m6A modification to prolong half-life and enhance gene stability, can negatively regular the T cell-related signaling pathway (94, 95). After inducing differentiation of different effector T cells in vitro, the expression of Mettl3 is increased dramatically in the Tregs (92). The inhibition of Mettl3 reduces Foxp3 mRNA level by decreasing its m6A modification and mRNA stability, suppressing Tregs differentiation. In addition, the vivo experiments confirmed that Mettl3 deletion increased antibody production and worsened the lupus-like phenotype in chronic graft versus host disease (cGVHD) mice (92). Importantly, the deficiency of Mettl3 in Tregs results in the elevated levels of Socs family genes and suppresses the biological activity of Tregs by inhibiting IL-2/Stat5 signaling (96). Mettl3 plays a key role in the formation of GCs by activating the endogenous pathway of Tfh programming and inhibiting Th1 lineage related genes. The deficiency of Mettl3 in CD4+ T cells impairs the differentiation of Tfh cells and the formation of GCs. Mechanistically, the Mettl3 enhances the stability of Tcf7 mRNA through catalyzed m6A methylation at 3’ UTR, improving the expression and function of TCF-1 to promote Tfh differentiation (97). In humoral immunized mice, the pharmacologically inhibiting of Mettl3 decreases the proportion of naïve T cells while increases the proportion of effector T cells. To be specially, the frequencies of Tfh cells and Treg cells in spleen are significantly reduced, which may be related to the decreased expression and stability of Foxp3 (92).
Mettl14 seems to exhibit mainly anti-inflammatory effects. The mice with the Mettl14-knockout T cells exhibit an increased proportion of Th1 and Th17, and the impaired function of Tregs. The expression of RORγt is reduced in Mettl14-knockout Tregs. In addition, the Mettl14-knockout naïve T cells lose their potential for induction into Treg cells (98, 99). Mechanistically, these changes are associated with activation of the mTOR pathway, a signaling pathway known to inhibit Tregs function (99). Moreover, in experimental transplantation mouse models (EAEs), the down-regulated expression of METTL14 facilitates the infiltration of CD4+ T cells surrounding the allograft, inducing accelerated rejection (100).
The Wtap-knockout T cells result in colitis phenotypes more severe than wild types, and the Wtap-knockout Tregs obtaine increased expression of Rorc and Il-17a, which is involved in the TCR signal activation and conduction (101). These phenotypes following Wtap deletion are reproduced on Virma or Mettl3-knockout models, suggesting that these phenotypes result from the changes in m6A signaling rather than non-m6A dependent effects of Wtap (101). At the same time, the non-m6A dependent effects of WTAP are also remarkable. For example, WTAP is upregulated in CD4+ T cells of tolerant kidney transplant recipients and positively correlated with Tregs proportion (102). However, crucial inquiries regarding the precise localization of WTAP-mediated m6A modification within the T cell transcriptome and the specific target genes influenced by m6A remain unresolved.
Among tRNA modifications, some studies have shown that the m1A58 modification of tRNA (tRNA-m1A58) can improve the initiation and elongation efficiency of protein translation (103, 104). The tRNA-m1A58 modification level is much higher than other tRNA methylation in activated T cells (105). Trmt61a and Trmt6 seem to exhibit important pro-inflammatory effects in different types of CD4+ T cells. Trmt61a and Trmt6 are up-regulated during T cells activation, which enable m1A RNA modification on specific early expression tRNAs subsets. The Trmt61-knockout mice could not be induced to develop a colitis phenotype, resulting in a predominant arrest of CD4+ T cells in the G0-G1 phase (105). However, the key questions regarding the precise localization of Trmt61a and Trmt6-mediated m1A modifications in the T cell transcriptome remain unresolved.
It has been reported that RNA m5C methylation has a potential regulatory effect on homeostasis regulation of Th17 cells (91, 106). RORγt facilitates the formation of transcription-coupled m5C modification on Th17 cell-specific cytokine mRNA by recruiting Nsun2 to the chromatin region of the target gene. Furthermore, the Nsun2-knockout T cells diminish the secretion of proinflammatory cytokines and impair the intercellular communication between Th17 cells and IL-17 receptor-expressing cells, impeding colitis progression (107). At present, the researches of m5C effect on CD4+ T cell proliferation and differentiation are limited, whether m5C “writers” independently or jointly affect T cell function are still worthy further study. It is imperative to further investigate the location of m5C modifications in the T cell transcriptomes, whether these modifications affect thymocytes development, and whether they affect IL-7 or TCR signaling in mature T cells.
Although, there are no significant changes in the mRNA levels of Fto among effector CD4+ T cells compared to the naïve CD4+ T (108), the involvement of FTO in CD4+ T cell homeostasis and differentiation should not be dismissed. Importantly, the m6A plays a crucial role in the regulation of the expression of CD40L in CD4+ T cell, and FTO directly influences the expression of CD40L (12). The primary role of CD40L is to bind with CD40 on the APCs, initiating the activation of the immune response (109). Currently, there are ongoing studies investigating the development of inhibitors targeting CD40L in SLE, RA, and other autoimmune diseases (110). In addition, the Fto-knockout IECs under the DSS-induced results in heightened severity of colitis, with a higher proportion of Th17 cells than in wild types (62). These findings suggest that FTO may indirectly regulate the function of CD4+ T cells. Similarly, there is an up-regulation of FTO in patients with aortic dissection (AD), which is inversely correlated with the abundance of activated CD4+ T cells (111). However, the mechanisms require further investigation.
It is worth noting that studies have demonstrated that FTO ubiquitously expresses in various developmental stages of human tissues, localizing to both the cytoplasm and nucleus (112, 113). The expression is highest in the hypothalamus of human brain tissue, suggesting the important role for FTO protein in metabolic control (114). Thus, the alterations mentioned above may arise from the combined influences of metabolic adjustments and RNA modification. FTO protein is initially identified as a regulator of lipid metabolism (115). The disorders of lipid metabolism have been reported in autoimmune diseases (116, 117). Fto-knockdown in IECs leads to upregulated m6A level of CerS6 (Ceramide synthase 6) and decreased C16-ceramide synthesis, accelerating pro-inflammatory macrophage and Th17 differentiation and promoting inflammation (62). In this study, FTO plays a role in regulating sphingolipid metabolism and m6A modification. On the one hand, it is needful to explore whether FTO-mediated methylation at specific loci causally impacts lipid metabolism and whether these changes are critical for T cell homeostasis. In the future, there is a need to development more precise methods to distinguish the sites between of FTO-mediated methylation and lipid metabolism. On the other hand, considering that FTO is not only involved in the regulation of m6A, but also serves as a crucial demethylase for m6Am, m1A and m5C modification (118, 119). It is crucial to distinguish the sites of these RNA modifications mediated by FTO.
ALKBH5 can decrease the m6A modification in CXCL2 and IFN-γ mRNA, leading to enhanced transcriptional stability and protein expression. Consequently, the deficiency of Alkbh5 in EAEs results in the impairment of the IL-17 signaling pathway in CD4+ T cells, and the augmented CD4+ T cell response increases the infiltration of neutrophils into the central nervous system during neuro-inflammation (108). The impacts of Alkbh5 and Mettl3 on CD4+ T cell-mediated autoimmunity models exhibit a similar pattern, yet the underlying mechanism remains incompletely understood, the possible reason could be attributed to the fine-tuning of T cell activity through a temporal-sequential ordering by these two molecules. Whether ALKBH5 and FTO participate in these processes synergistically, and whether the changes caused by ALKBH5-knockout can be reversed through overexpression of FTO, are also worth further investigation. Although ALKBH1, ALKBH 3 and ALKBH5 belong to the AlkB homologue family, the role of ALKBH1and ALKBH3 in the development, and function of CD4+ T cells remains unexplored. The self-regulation processes of CD4+ T cells are highly intricated, exploring the role of “erasers” in this series of processes will hold significant importance in bridging the gaps within related fields.
Most of the function and mechanism of RNA methyltransferase and demethylase are mediated by methylation recognition proteins. In tumor immunity, these proteins are considered potential therapeutic targets. Therefore, it is meaningful to explore the functions of “readers”. So far, no studies have directly investigated the effect of YTH family proteins on CD4+ T cell homeostasis and differentiation by knocking down YTH family genes in CD4+ T cells, and the existing studies can only provide indirect evidence. YTHDF1/2/3 proteins in naïve CD4+ T cells recognize m6A-modified SOCS family genes and promptly initiate the degradation of target genes (120). Subsequently, the expression changes of SOCS further affect the JAK/STAT signaling pathway and regulate T cell homeostasis. DCs are the main APCs responsible for T cell activation (121). Silencing Ythdf1 in DCs can impact T cell activation by diminishing the expression of co-stimulatory molecules CD40 and CD80, however, these changes don’t occur in Mettl3-knockout DC cells (122). Ythdf1 can recognize m6A-modified mRNAs and promote the mRNA translation of CD40 and CD80. In some cases, dysfunction in m6A readers may not cause changes in overall m6A modification levels, the function of “readers” should be better understood by considering “writers” and “erasers” together. Considering the crucial role of “readers” in the RNA modification system and the contributions to precise regulation of immune response, it is imperative to further investigate their underlying mechanisms.
To conclusion, despite the variety and complexity of RNA modification associated proteins, m6A-associated proteins are currently the most studied than others (123). The components of m6A modification play a crucial role in autoimmune responses and the differentiation of CD4+ T cells, which offers potential novel insights into the treatment of various diseases by regulating physiological and pathological processes associated with CD4+ T cells (Figure 2). Previous studies primarily utilize a conditional knockout mouse model targeting specific genes in CD4+ T cells to explore their impacts on cell activation, proliferation, and differentiation. However, their pro-inflammatory or anti-inflammatory effects also depend on the specific target RNAs and the type of autoimmune disease.
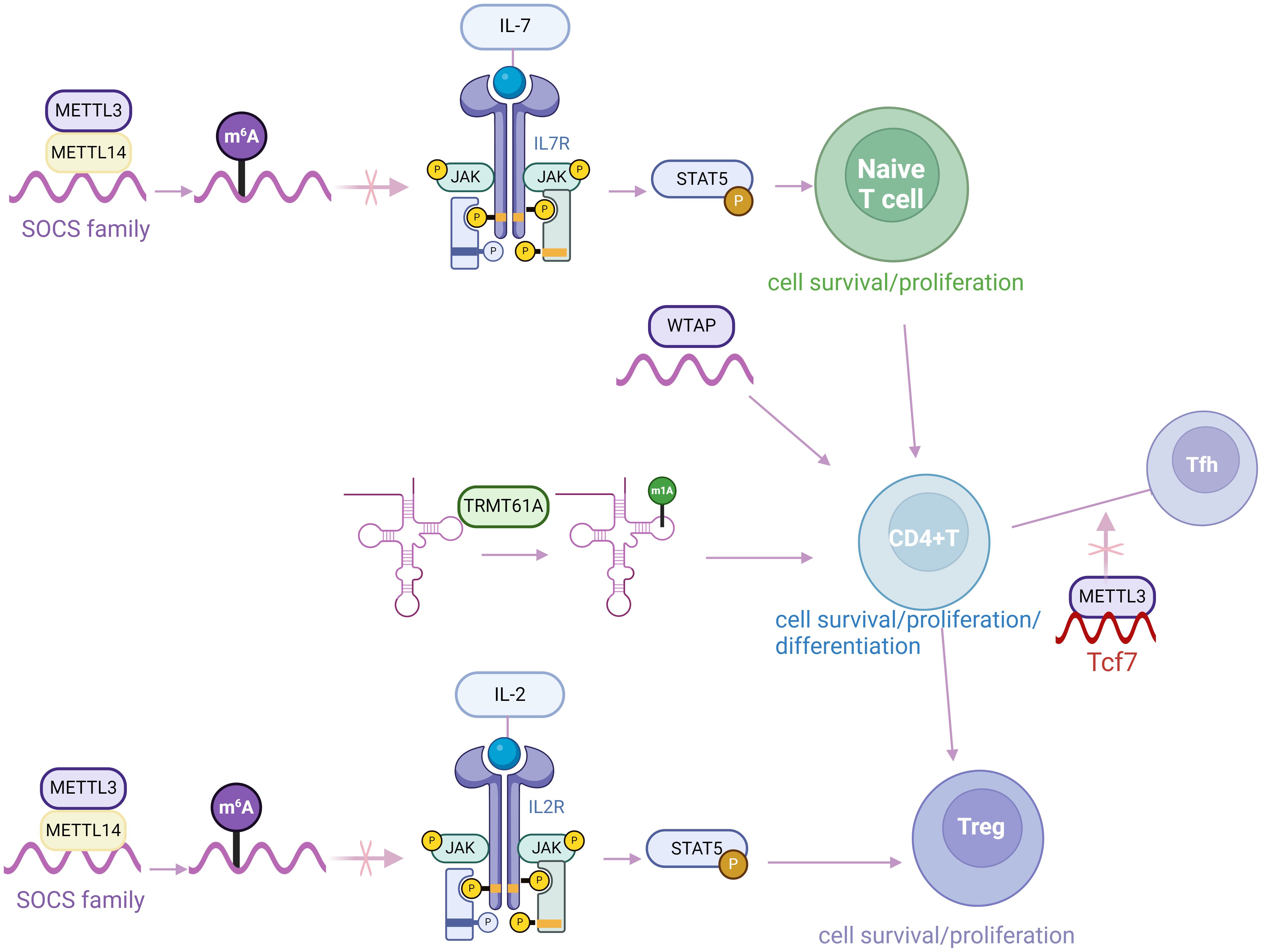
Figure 2. RNA modification plays a crucial role in regulating the development of CD4+ T cell. m6A, N6-methyladenosine; m1A, N1-methyladenosine.
Limited research has been conducted on other types of RNA modification, highlighting the need for further investigation into the role of m1A and m5C modified protein in controlling CD4+ T cell function and autoimmune disease. Nevertheless, there are modifications that can alter the biological functions in the immune system in viral infections and neoplastic diseases. The m1A methyltransferase TRMT61A can enhance the tumor-killing efficacy of CD8+ T cells through the regulation of cholesterol biosynthesis (124). Given that m1A modification shares several regulatory proteins with m6A modification, the research methodologies and findings associated with m6A can serve as a valuable reference to address the existing knowledge gaps in m1A modification within the context of autoimmune diseases. CD4+ T cells serve as the primary target for Human Immunodeficiency Virus (HIV) infection. During the latent phase of HIV infection, the m5C methyltransferase NSUN1 targets HIV TAR RNA and generates m5C methylation, which can impede HIV transcriptional and viral latency in CD4+ T cells (125). NSUN3 facilitates the infiltration of M2 macrophages while inhibiting the infiltration of M1 macrophages in head and neck squamous cell carcinoma (126). TET2 and TET3 can influence neutrophil granulation, phagocytosis, and cytokine signaling through the demethylation and destabilization of Scos3 mRNA (127). Given the complexity of RNA modifications and the diversity of immune cells, further investigation is warranted to enhance our understanding of the interactions between RNA modifications and immune cells.
Currently, research on m6Am modification in autoimmune diseases remains limited. Recent studies have highlighted the critical role of m6Am modification in inflammatory responses (128). The deletion of the m6Am methyltransferase Pcif1 mRNA impairs macrophage phagocytosis and migration via m6Am-CSF1R signaling, consequently mitigating periodontal inflammation (129). Although these studies are preliminary explorations, they offer valuable insights into the biological mechanisms by which m6Am responds to inflammation and immune cells, thereby providing a foundation for further investigation into its role in autoimmune diseases.
Despite advancements in understanding the intricate self-regulation mechanism of T cells, the significant knowledge gaps persist. Further investigations are required to determine the positions of different RNA modifications within the T cell transcriptome, identify target RNAs affected by these modifications, assess their influence on thymus cell development, and evaluate their impact on TCR signaling in mature T cells. Notably, the upstream mechanism of these enzyme activity changes should also be taken into consideration, and targeting the upstream factors of these enzymes will contribute to innovative improvements in treatment. Although these works are expected to be complex and challenging, they will hold great potential for exploring therapeutic targets.
Today, there are still many questions to be answered in order to better understand the impact of RNA modification on autoimmune diseases and T cell biology, and to apply the research to the clinic. First, investigating the extent to which RNA modifications influence disease progression in autoimmune diseases and determining whether they serve as primary regulators or auxiliary factors will be crucial for identifying potential therapeutic targets. Second, how to target RNA modification in clinical practice remains a challenge, although experiments have shown that it can work in vivo by giving nanoparticle preparations (130). Third, we cannot ignore the interaction between immune cells, and RNA modification may cause changes in immune function in all aspects by targeting one type of immune cell.
In cancer research, RNA enzymes are used as potential targets for therapy. For example, one preclinical study reported the inhibitory effect of FTO inhibitors on acute myeloid leukaemia (AML) cells by selectively inhibiting the m6A demethylase activity of FTO (131). The synthesis of selective and potent inhibitors of these enzymes is of technically feasible, it also heralds a viable new opportunity for autoimmune disease treatment. Many evidences linking dysregulation of RNA modification and autoimmune diseases suggest that developing inhibitors that target the RNA modification pathway would be fruitful. The diversity of RNA modifications and the involved molecular pathways give us hope and challenge. However, there are significant challenges in drug development, such as the selectivity of enzyme inhibitors. This problem is an inevitable challenge in the development of epigenetic modification enzyme inhibitors. For example, histone deacetylase (HDAC) is one important enzyme in the process of histone modification, which can remove the acetyl group on the histone Lysine residue, change the electric charge to make the chromosome structure more tightly, and thus inhibit the transcriptional expression of genes (132). To date, a total of five HDAC inhibitors have received market approval globally. Among these, only one is a selective inhibitor, while the remaining four are pan-inhibitors of HDAC. HDAC drugs have made a breakthrough in the clinical treatment of various subtypes of blood tumors, but the treatment of non-blood solid tumors is not effective (133). With the in-depth study of RNA modification, we have not only gained a deeper understanding of the disease mechanism and its target pathways but also uncovered its intricate relationship with the occurrence and immunity of autoimmune diseases. Many challenges remain due to the complexity of post-transcriptional modifications, epigenetic transcriptional regulation, and non-epigenetic cell signaling cascades. It is believed that with the continuous efforts of scientists, more and more efficient and low-toxic enzyme inhibitors will be applied to the treatment of autoimmune diseases and other diseases.
HYu: Writing – original draft, Writing – review & editing. ZM: Investigation, Methodology, Writing – review & editing. SS: Supervision, Visualization, Writing – review & editing. ZX: Writing – review & editing. HYi: Funding acquisition, Writing – review & editing.
The author(s) declare that no financial support was received for the research and/or publication of this article.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fimmu.2025.1563150/full#supplementary-material
1. Xu Y, Liu W, Ren L. Emerging roles and mechanism of m6A methylation in rheumatoid arthritis. BioMed Pharmacother. (2024) 170:116066. doi: 10.1016/j.biopha.2023.116066
2. Li L, Xia X, Yang T, Sun Y, Liu X, Xu W, et al. RNA methylation: A potential therapeutic target in autoimmune disease. Int Rev Immunol. (2024) 43:160–77. doi: 10.1080/08830185.2023.2280544
3. Wang X, Chen C, Sun H, Mao K, Yao J, Zhang W, et al. m(6)A mRNA modification potentiates Th17 functions to inflame autoimmunity. Sci China Life Sci. (2023) 66:2543–52. doi: 10.1007/s11427-022-2323-4
4. Li C, Xu P, Huang Y, Wang Y, Wu Y, Li H, et al. RNA methylations in depression, from pathological mechanism to therapeutic potential. Biochem Pharmacol. (2023) 215:115750. doi: 10.1016/j.bcp.2023.115750
5. Geng Q, Cao X, Fan D, Wang Q, Wang X, Zhang M, et al. Potential medicinal value of N6-methyladenosine in autoimmune diseases and tumours. Br J Pharmacol. (2023). doi: 10.1111/bph.16030
6. Cao G, Chi S, Wang X, Sun J, Zhang Y. CD4+CXCR5+PD-1+ T follicular helper cells play a pivotal role in the development of rheumatoid arthritis. Med Sci Monit. (2019) 25:3032–40. doi: 10.12659/MSM.914868
7. Behfarjam F, Sanati MH, Nasseri Moghaddam S, Ataei M, Nikfam S, Jadali Z. Role of Th1/Th2 cells and related cytokines in autoimmune hepatitis. Turk J Gastroenterol. (2017) 28:110–4. doi: 10.5152/tjg.2017.17501
8. Shan Y, Chen W, Li Y. The role of m(6)A RNA methylation in autoimmune diseases: Novel therapeutic opportunities. Genes Dis. (2024) 11:252–67. doi: 10.1016/j.gendis.2023.02.013
9. Pang J, Kuang TD, Yu XY, Novák P, Long Y, Liu M, et al. N6-methyladenosine in myeloid cells: a novel regulatory factor for inflammation-related diseases. J Physiol Biochem. (2024) 80:249–60. doi: 10.1007/s13105-023-01002-x
10. Zhang Y, Hu W, Li HB. RNA modification-mediated translational control in immune cells. RNA Biol. (2023) 20:603–13. doi: 10.1080/15476286.2023.2246256
11. Zhou H, Mao L, Xu H, Wang S, Tian J. The functional roles of m(6)A modification in T lymphocyte responses and autoimmune diseases. Cytokine Growth Factor Rev. (2022) 65:51–60. doi: 10.1016/j.cytogfr.2022.04.004
12. Van Vroonhoven ECN, Picavet LW, Scholman RC, Van Den Dungen NAM, Mokry M, Evers A, et al. N(6)-methyladenosine directly regulates CD40L expression in CD4(+) T lymphocytes. Biol (Basel). (2023) 12:1004. doi: 10.3390/biology12071004
13. Teng Y, Yi J, Chen J, Yang L. N6-methyladenosine (m6A) modification in natural immune cell-mediated inflammatory diseases. J Innate Immun. (2023) 15:804–21. doi: 10.1159/000534162
14. Shi H, Wang X, Lu Z, Zhao BS, Ma H, Hsu PJ, et al. YTHDF3 facilitates translation and decay of N(6)-methyladenosine-modified RNA. Cell Res. (2017) 27:315–28. doi: 10.1038/cr.2017.15
15. Huang H, Weng H, Sun W, Qin X, Shi H, Wu H, et al. Recognition of RNA N(6)-methyladenosine by IGF2BP proteins enhances mRNA stability and translation. Nat Cell Biol. (2018) 20:285–95. doi: 10.1038/s41556-018-0045-z
16. Benak D, Kolar F, Zhang L, Devaux Y, Hlavackova M. RNA modification m(6)Am: the role in cardiac biology. Epigenetics. (2023) 18:2218771. doi: 10.1080/15592294.2023.2218771
17. Cesaro B, Tarullo M, Fatica A. Regulation of gene expression by m6Am RNA modification. Int J Mol Sci. (2023) 24:2277. doi: 10.3390/ijms24032277
18. Annapoorna PK, Iyer H, Parnaik T, Narasimhan H, Bhattacharya A, Kumar A. FTO: an emerging molecular player in neuropsychiatric diseases. Neuroscience. (2019) 418:15–24. doi: 10.1016/j.neuroscience.2019.08.021
19. Xu L, Zhang C, Yin H, Gong S, Wu N, Ren Z, et al. RNA modifications act as regulators of cell death. RNA Biol. (2021) 18:2183–93. doi: 10.1080/15476286.2021.1925460
20. Li J, Zhang H, Wang H. N(1)-methyladenosine modification in cancer biology: Current status and future perspectives. Comput Struct Biotechnol J. (2022) 20:6578–85. doi: 10.1016/j.csbj.2022.11.045
21. Oerum S, Dégut C, Barraud P, Tisné C. m1A post-transcriptional modification in tRNAs. Biomolecules. (2017) 7:20. doi: 10.3390/biom7010020
22. Sharma S, Hartmann JD, Watzinger P, Klepper A, Peifer C, Kötter P, et al. A single N(1)-methyladenosine on the large ribosomal subunit rRNA impacts locally its structure and the translation of key metabolic enzymes. Sci Rep. (2018) 8:11904. doi: 10.1038/s41598-018-30383-z
23. Barraud P, Golinelli-Pimpaneau B, Atmanene C, Sanglier S, Van Dorsselaer A, Droogmans L, et al. Crystal structure of Thermus thermophilus tRNA m1A58 methyltransferase and biophysical characterization of its interaction with tRNA. J Mol Biol. (2008) 377:535–50. doi: 10.1016/j.jmb.2008.01.041
24. Nombela P, Miguel-López B, Blanco S. The role of m(6)A, m(5)C and Ψ RNA modifications in cancer: Novel therapeutic opportunities. Mol Cancer. (2021) 20:18. doi: 10.1186/s12943-020-01263-w
25. Fu L, Guerrero CR, Zhong N, Amato NJ, Liu Y, Liu S, et al. Tet-mediated formation of 5-hydroxymethylcytosine in RNA. J Am Chem Soc. (2014) 136:11582–5. doi: 10.1021/ja505305z
26. Guallar D, Bi X, Pardavila JA, Huang X, Saenz C, Shi X, et al. RNA-dependent chromatin targeting of TET2 for endogenous retrovirus control in pluripotent stem cells. Nat Genet. (2018) 50:443–51. doi: 10.1038/s41588-018-0060-9
27. Yang X, Yang Y, Sun BF, Chen YS, Xu JW, Lai WY, et al. 5-methylcytosine promotes mRNA export - NSUN2 as the methyltransferase and ALYREF as an m(5)C reader. Cell Res. (2017) 27:606–25. doi: 10.1038/cr.2017.55
28. Cai M, Yang C, Wang Z. N7-methylguanosine modification: from regulatory roles to therapeutic implications in cancer. Am J Cancer Res. (2023) 13:1640–55.
29. Chen Y, Lin H, Miao L, He J. Role of N7-methylguanosine (m(7)G) in cancer. Trends Cell Biol. (2022) 32:819–24. doi: 10.1016/j.tcb.2022.07.001
30. Dimitrova DG, Teysset L, Carré C. RNA 2’-O-methylation (Nm) modification in human diseases. Genes (Basel). (2019) 10:117. doi: 10.3390/genes10020117
31. Brazane M, Dimitrova DG, Pigeon J, Paolantoni C, Ye T, Marchand V, et al. The ribose methylation enzyme FTSJ1 has a conserved role in neuron morphology and learning performance. Life Sci Alliance. (2023) 6. doi: 10.26508/lsa.202201877
32. Rees F, Doherty M, Grainge MJ, Lanyon P, Zhang W. The worldwide incidence and prevalence of systemic lupus erythematosus: a systematic review of epidemiological studies. Rheumatol (Oxford). (2017) 56:1945–61. doi: 10.1093/rheumatology/kex260
33. Luo Q, Fu B, Zhang L, Guo Y, Huang Z, Li J. Decreased peripheral blood ALKBH5 correlates with markers of autoimmune response in systemic lupus erythematosus. Dis Markers. (2020) 2020:8193895. doi: 10.1155/2020/8193895
34. Luo Q, Rao J, Zhang L, Fu B, Guo Y, Huang Z, et al. The study of METTL14, ALKBH5, and YTHDF2 in peripheral blood mononuclear cells from systemic lupus erythematosus. Mol Genet Genomic Med. (2020) 8:e1298. doi: 10.1002/mgg3.1298
35. Fresneda Alarcon M, Mclaren Z, Wright HL. Neutrophils in the pathogenesis of rheumatoid arthritis and systemic lupus erythematosus: same foe different M.O. Front Immunol. (2021) 12:649693. doi: 10.3389/fimmu.2021.649693
36. Luo S, Liao C, Zhang L, Ling C, Zhang X, Xie P, et al. METTL3-mediated m6A mRNA methylation regulates neutrophil activation through targeting TLR4 signaling. Cell Rep. (2023) 42:112259. doi: 10.1016/j.celrep.2023.112259
37. Wan L, Liu J, Huang C, Zhu Z, Wang K, Sun G, et al. Comprehensive analysis and functional characteristics of differential expression of N6-methyladenosine methylation modification in the whole transcriptome of rheumatoid arthritis. Mediators Inflammation. (2022) 2022:4766992. doi: 10.1155/2022/4766992
38. Su Y, Wu Z, Liu Y, Liu X, Kang J, Jia J, et al. Increased m6A RNA methylation and METTL3 expression may contribute to the synovitis progression of rheumatoid arthritis. Exp Cell Res. (2024) 442:114237. doi: 10.1016/j.yexcr.2024.114237
39. Wang J, Yan S, Lu H, Wang S, Xu D. METTL3 attenuates LPS-induced inflammatory response in macrophages via NF-κB signaling pathway. Mediators Inflammation. (2019) 2019:3120391. doi: 10.1155/2019/3120391
40. Miao T, Qiu Y, Chen J, Li P, Li H, Zhou W, et al. METTL3 knockdown suppresses RA-FLS activation through m(6)A-YTHDC2-mediated regulation of AMIGO2. Biochim Biophys Acta Mol Basis Dis. (2024) 1870:167112. doi: 10.1016/j.bbadis.2024.167112
41. Shi W, Zheng Y, Luo S, Li X, Zhang Y, Meng X, et al. METTL3 promotes activation and inflammation of FLSs through the NF-κB signaling pathway in rheumatoid arthritis. Front Med (Lausanne). (2021) 8:607585. doi: 10.3389/fmed.2021.607585
42. Wang YT, Wu YJ, Huang YR, Xue QY, Cheng CL, Miao CG, et al. Mechanism of Huangqin Qingre Chubi Capsules in improving rheumatoid arthritis based on METTL3-SFRP4/Wnt/β-catenin pathway. Zhongguo Zhong Yao Za Zhi. (2024) 49:3081–94. doi: 10.19540/j.cnki.cjcmm
43. Chen J, Lin X, He J, Liu D, He L, Zhang M, et al. Artemisitene suppresses rheumatoid arthritis progression via modulating METTL3-mediated N6-methyladenosine modification of ICAM2 mRNA in fibroblast-like synoviocytes. Clin Transl Med. (2022) 12:e1148. doi: 10.1002/ctm2.v12.12
44. Tang J, Yu Z, Xia J, Jiang R, Chen S, Ye D, et al. METTL14-mediated m6A modification of TNFAIP3 involved in inflammation in patients with active rheumatoid arthritis. Arthritis Rheumatol. (2023) 75:2116–29. doi: 10.1002/art.v75.12
45. Ma JZ, Yang F, Zhou CC, Liu F, Yuan JH, Wang F, et al. METTL14 suppresses the metastatic potential of hepatocellular carcinoma by modulating N(6) -methyladenosine-dependent primary MicroRNA processing. Hepatology. (2017) 65:529–43. doi: 10.1002/hep.28885
46. Zhu Z, Wan L. N6−methyladenosine methyltransferase METTL14 is associated with macrophage polarization in rheumatoid arthritis. Exp Ther Med. (2024) 28:375. doi: 10.3892/etm.2024.12664
47. Li X, Xu X, Zhang Q, Ling M, Li X, Tan X. METTL14 promotes fibroblast-like synoviocytes activation via the LASP1/SRC/AKT axis in rheumatoid arthritis. Am J Physiol Cell Physiol. (2023) 324:C1089–c1100. doi: 10.1152/ajpcell.00575.2022
48. Luo Q, Gao Y, Zhang L, Rao J, Guo Y, Huang Z, et al. Decreased ALKBH5, FTO, and YTHDF2 in peripheral blood are as risk factors for rheumatoid arthritis. BioMed Res Int. (2020) 2020:5735279. doi: 10.1155/2020/5735279
49. Liu X, Xia Z, Liu L, Ren D. WTAP promotes fibroblast-like synoviocyte pyroptosis in Rheumatoid arthritis by upregulating N6-methyladenosine modification of NLRP3. J Bioenerg Biomembr. (2024) 56(5):563–71. doi: 10.1007/s10863-024-10035-w
50. Kuang Y, Li R, Wang J, Xu S, Qiu Q, Lin S, et al. ALKBH5-mediated RNA m(6) A methylation regulates the migration, invasion, and proliferation of rheumatoid fibroblast-like synoviocytes. Arthritis Rheumatol. (2024) 76:192–205. doi: 10.1002/art.42676
51. Li R, Kuang Y, Niu Y, Zhang S, Chen S, Su F, et al. FTO-mediated RNA m(6)A methylation regulates synovial aggression and inflammation in rheumatoid arthritis. Biochim Biophys Acta Mol Basis Dis. (2024) 1870:167341. doi: 10.1016/j.bbadis.2024.167341
52. Fan D, Geng Q, Wang B, Wang X, Xia Y, Yang L, et al. Hypoxia-induced ALKBH5 aggravates synovial aggression and inflammation in rheumatoid arthritis by regulating the m6A modification of CH25H. Clin Immunol. (2024) 261:109929. doi: 10.1016/j.clim.2024.109929
53. Yao F, Xu C, Gao Y, Fu B, Zhang L, Guo Y, et al. Expression and clinical significance of the m6A reader YTHDF2 in peripheral blood mononuclear cells from rheumatoid arthritis patients. J Immunotoxicol. (2022) 19:53–60. doi: 10.1080/1547691X.2022.2067916
54. Xiao J, Cai X, Wang R, Zhou W, Ye Z. ALKBH5-YTHDF2 m6A modification axis inhibits rheumatoid arthritis progression by suppressing NLRP3. Biochem Biophys Res Commun. (2023) 668:70–6. doi: 10.1016/j.bbrc.2023.05.087
55. Fan D, Liu B, Gu X, Zhang Q, Ye Q, Xi X, et al. Potential target analysis of triptolide based on transcriptome-wide m(6)A methylome in rheumatoid arthritis. Front Pharmacol. (2022) 13:843358. doi: 10.3389/fphar.2022.843358
56. Xiao J, Cai X, Wang R, Zhou W, Ye Z. Identification of synovial fibroblast-associated neuropeptide genes and m6A factors in rheumatoid arthritis using single-cell analysis and machine learning. Dis Markers. (2022) 2022:5114697. doi: 10.1155/2022/5114697
57. Rajamäki K, Taira A, Katainen R, Välimäki N, Kuosmanen A, Plaketti RM, et al. Genetic and epigenetic characteristics of inflammatory bowel disease-associated colorectal cancer. Gastroenterology. (2021) 161:592–607. doi: 10.1053/j.gastro.2021.04.042
58. Yang L, Wu G, Wu Q, Peng L, Yuan L. METTL3 overexpression aggravates LPS-induced cellular inflammation in mouse intestinal epithelial cells and DSS-induced IBD in mice. Cell Death Discovery. (2022) 8:62. doi: 10.1038/s41420-022-00849-1
59. Zhang T, Ding C, Chen H, Zhao J, Chen Z, Chen B, et al. m(6)A mRNA modification maintains colonic epithelial cell homeostasis via NF-κB-mediated antiapoptotic pathway. Sci Adv. (2022) 8:eabl5723. doi: 10.1126/sciadv.abl5723
60. Yin H, Ju Z, Zhang X, Zuo W, Yang Y, Zheng M, et al. Inhibition of METTL3 in macrophages provides protection against intestinal inflammation. Cell Mol Immunol. (2024) 21:589–603. doi: 10.1038/s41423-024-01156-8
61. Ge Y, Chen R, Ling T, Liu B, Huang J, Cheng Y, et al. Elevated WTAP promotes hyperinflammation by increasing m6A modification in inflammatory disease models. J Clin Invest. (2024) 134:e177932. doi: 10.1172/JCI177932
62. Ma Y, Zhang X, Xuan B, Li D, Yin N, Ning L, et al. Disruption of CerS6-mediated sphingolipid metabolism by FTO deficiency aggravates ulcerative colitis. Gut. (2024) 73:268–81. doi: 10.1136/gutjnl-2023-330009
63. Chen Y, Lei J, He S. m(6)A modification mediates mucosal immune microenvironment and therapeutic response in inflammatory bowel disease. Front Cell Dev Biol. (2021) 9:692160. doi: 10.3389/fcell.2021.692160
64. Wang X, Ji Y, Feng P, Liu R, Li G, Zheng J, et al. The m6A reader IGF2BP2 regulates macrophage phenotypic activation and inflammatory diseases by stabilizing TSC1 and PPARγ. Adv Sci (Weinh). (2021) 8:2100209. doi: 10.1002/advs.202100209
65. Wu F, Huang H, Sun D, Cai B, Zhou H, Quan R, et al. Identification of key genes with abnormal RNA methylation modification and selected m6A regulators in ankylosing spondylitis. Immun Inflammation Dis. (2024) 12:e1314. doi: 10.1002/iid3.v12.8
66. Luo Q, Guo Y, Xiao Q, Fu B, Zhang L, Guo Y, et al. Expression and clinical significance of the m6A RNA-binding proteins YTHDF2 in peripheral blood mononuclear cells from new-onset ankylosing spondylitis. Front Med (Lausanne). (2022) 9:922219. doi: 10.3389/fmed.2022.922219
67. Chen Y, Wu Y, Fang L, Zhao H, Xu S, Shuai Z, et al. METTL14-m6A-FOXO3a axis regulates autophagy and inflammation in ankylosing spondylitis. Clin Immunol. (2023) 257:109838. doi: 10.1016/j.clim.2023.109838
68. Ma J, Wang X, Yang X, Wang X, Tan T, Fang H, et al. Increased METTL3 expression and m(6)A RNA methylation may contribute to the development of dry eye in primary Sjögren’s syndrome. BMC Ophthalmol. (2023) 23:252. doi: 10.1186/s12886-023-02988-0
69. Liu Y, Zhu J, Ding L. Involvement of RNA methylation modification patterns mediated by m7G, m6A, m5C and m1A regulators in immune microenvironment regulation of Sjögren’s syndrome. Cell Signal. (2023) 106:110650. doi: 10.1016/j.cellsig.2023.110650
70. Xiao Q, Wu X, Deng C, Zhao L, Peng L, Zhou J, et al. The potential role of RNA N6-methyladenosine in primary Sjögren’s syndrome. Front Med (Lausanne). (2022) 9:959388. doi: 10.3389/fmed.2022.959388
71. Yin J, Fu J, Xu J, Chen C, Zhu H, Wang B, et al. Integrated analysis of m6A regulator-mediated RNA methylation modification patterns and immune characteristics in Sjögren’s syndrome. Heliyon. (2024) 10:e28645. doi: 10.1016/j.heliyon.2024.e28645
72. Cheng L, Li H, Zhan H, Liu Y, Li X, Huang Y, et al. Alterations of m6A RNA methylation regulators contribute to autophagy and immune infiltration in primary Sjögren’s syndrome. Front Immunol. (2022) 13:949206. doi: 10.3389/fimmu.2022.949206
73. Truffinet F, Arco-Hierves A, Shalabi H, Pascaud J, Mazet P, Rivière E, et al. m(6)A RNA methylation controls salivary gland epithelial cell function and has a protective role in Sjögren’s disease. Ann Rheum Dis. (2024), ard-2024-226224. doi: 10.1136/ard-2024-226224
74. Lyu Z, Huang B, Zhang J, Qian Q, Pu X, Cui N, et al. Suppression of YTHDF2 attenuates autoimmune hepatitis by expansion of myeloid-derived suppressor cells. J Autoimmun. (2023) 135:102993. doi: 10.1016/j.jaut.2023.102993
75. De Jesus DF, Zhang Z, Brown NK, Li X, Xiao L, Hu J, et al. Redox regulation of m(6)A methyltransferase METTL3 in β-cells controls the innate immune response in type 1 diabetes. Nat Cell Biol. (2024) 26:421–37. doi: 10.1038/s41556-024-01368-0
76. Knochelmann HM, Dwyer CJ, Bailey SR, Amaya SM, Elston DM, Mazza-Mccrann JM, et al. When worlds collide: Th17 and Treg cells in cancer and autoimmunity. Cell Mol Immunol. (2018) 15:458–69. doi: 10.1038/s41423-018-0004-4
77. Mills KHG. IL-17 and IL-17-producing cells in protection versus pathology. Nat Rev Immunol. (2023) 23:38–54. doi: 10.1038/s41577-022-00746-9
78. Hu C, Zhen Y, Ma Z, Zhao L, Wu H, Shu C, et al. Polyamines from myeloid-derived suppressor cells promote Th17 polarization and disease progression. Mol Ther. (2023) 31:569–84. doi: 10.1016/j.ymthe.2022.10.013
79. Ou Q, Qiao X, Li Z, Niu L, Lei F, Cheng R, et al. Apoptosis releases hydrogen sulfide to inhibit Th17 cell differentiation. Cell Metab. (2024) 36:78–89.e5. doi: 10.1016/j.cmet.2023.11.012
80. Sun Z, Braga-Neto MB, Xiong Y, Bhagwate AV, Gibbons HR, Sagstetter MR, et al. Hypomethylation and overexpression of th17-associated genes is a hallmark of intestinal CD4+ Lymphocytes in crohn’s disease. J Crohns Colitis. (2023) 17:1847–57. doi: 10.1093/ecco-jcc/jjad093
81. Xu Y, Yang J, Chen X, Deng J, Gong H, Li F, et al. MicroRNA-182-5p aggravates ulcerative colitis by inactivating the Wnt/β-catenin signaling pathway through DNMT3A-mediated SMARCA5 methylation. Genomics. (2022) 114:110360. doi: 10.1016/j.ygeno.2022.110360
82. Wang K, Liu F, Muchu B, Deng J, Peng J, Xu Y, et al. E3 ubiquitin ligase RNF180 mediates the ALKBH5/SMARCA5 axis to promote colon inflammation and Th17/Treg imbalance in ulcerative colitis mice. Arch Pharm Res. (2024) 47:645–58. doi: 10.1007/s12272-024-01507-z
83. Vyas SP, Srivastava RN, Goswami R. Calcitriol attenuates TLR2/IL-33 signaling pathway to repress Th9 cell differentiation and potentially limits the pathophysiology of rheumatoid arthritis. Mol Cell Biochem. (2021) 476:369–84. doi: 10.1007/s11010-020-03914-4
84. Tu J, Chen W, Fang Y, Han D, Chen Y, Jiang H, et al. PU.1 promotes development of rheumatoid arthritis via repressing FLT3 in macrophages and fibroblast-like synoviocytes. Ann Rheum Dis. (2023) 82:198–211. doi: 10.1136/ard-2022-222708
85. Law C, Wacleche VS, Cao Y, Pillai A, Sowerby J, Hancock B, et al. Interferon subverts an AHR-JUN axis to promote CXCL13(+) T cells in lupus. Nature. (2024) 631:857–66. doi: 10.1038/s41586-024-07627-2
86. Wei X, Niu X. T follicular helper cells in autoimmune diseases. J Autoimmun. (2023) 134:102976. doi: 10.1016/j.jaut.2022.102976
87. Yang Y, Lv X, Zhan L, Chen L, Jin H, Tang X, et al. Case report: IL-21 and bcl-6 regulate the proliferation and secretion of tfh and tfr cells in the intestinal germinal center of patients with inflammatory bowel disease. Front Pharmacol. (2020) 11:587445. doi: 10.3389/fphar.2020.587445
88. Li X, Sun W, Huang M, Gong L, Zhang X, Zhong L, et al. Deficiency of CBL and CBLB ubiquitin ligases leads to hyper T follicular helper cell responses and lupus by reducing BCL6 degradation. Immunity. (2024) 57:1603–1617.e7. doi: 10.1016/j.immuni.2024.04.023
89. Ding S, Zhang Q, Luo S, Gao L, Huang J, Lu J, et al. BCL-6 suppresses miR-142-3p/5p expression in SLE CD4(+) T cells by modulating histone methylation and acetylation of the miR-142 promoter. Cell Mol Immunol. (2020) 17:474–82. doi: 10.1038/s41423-019-0268-3
90. Guo J, Wang Y, Tang L, Tang T, Li Z, Li M, et al. The regulation of Tfh cell differentiation by β-hydroxybutyrylation modification of transcription factor Bcl6. Chromosoma. (2023) 132:257–68. doi: 10.1007/s00412-023-00799-2
91. Guo G, Wang H, Shi X, Ye L, Yan K, Chen Z, et al. Disease Activity-Associated Alteration of mRNA m(5) C Methylation in CD4(+) T Cells of Systemic Lupus Erythematosus. Front Cell Dev Biol. (2020) 8:430. doi: 10.3389/fcell.2020.00430
92. Lu S, Wei X, Zhu H, Hu Z, Zheng M, Wu J, et al. m(6)A methyltransferase METTL3 programs CD4(+) T-cell activation and effector T-cell differentiation in systemic lupus erythematosus. Mol Med. (2023) 29:46. doi: 10.1186/s10020-023-00643-4
93. Li HB, Tong J, Zhu S, Batista PJ, Duffy EE, Zhao J, et al. m(6)A mRNA methylation controls T cell homeostasis by targeting the IL-7/STAT5/SOCS pathways. Nature. (2017) 548:338–42. doi: 10.1038/nature23450
94. Palmer DC, Restifo NP. Suppressors of cytokine signaling (SOCS) in T cell differentiation, maturation, and function. Trends Immunol. (2009) 30:592–602. doi: 10.1016/j.it.2009.09.009
95. Sobah ML, Liongue C, Ward AC. SOCS proteins in immunity, inflammatory diseases, and immune-related cancer. Front Med (Lausanne). (2021) 8:727987. doi: 10.3389/fmed.2021.727987
96. Tong J, Cao G, Zhang T, Sefik E, Amezcua Vesely MC, Broughton JP, et al. m(6)A mRNA methylation sustains Treg suppressive functions. Cell Res. (2018) 28:253–6. doi: 10.1038/cr.2018.7
97. Yao Y, Yang Y, Guo W, Xu L, You M, Zhang YC, et al. METTL3-dependent m(6)A modification programs T follicular helper cell differentiation. Nat Commun. (2021) 12:1333. doi: 10.1038/s41467-021-21594-6
98. Lu TX, Zheng Z, Zhang L, Sun HL, Bissonnette M, Huang H, et al. A new model of spontaneous colitis in mice induced by deletion of an RNA m(6)A methyltransferase component METTL14 in T cells. Cell Mol Gastroenterol Hepatol. (2020) 10:747–61. doi: 10.1016/j.jcmgh.2020.07.001
99. Liu Y, Yuan Y, Zhou Z, Jiang X, He S, Wei F, et al. Mettl14 sustains FOXP3 expression to promote the differentiation and functions of induced-regulatory T cells via the mTOR signaling pathway. Immunol Lett. (2023) 258:35–44. doi: 10.1016/j.imlet.2023.04.008
100. Liu Y, Yuan Y, Zhou Z, Cui Y, Teng Y, Huang H, et al. Mettl14-mediated m6A modification enhances the function of Foxp3(+) regulatory T cells and promotes allograft acceptance. Front Immunol. (2022) 13:1022015. doi: 10.3389/fimmu.2022.1022015
101. Ito-Kureha T, Leoni C, Borland K, Cantini G, Bataclan M, Metzger RN, et al. The function of Wtap in N(6)-adenosine methylation of mRNAs controls T cell receptor signaling and survival of T cells. Nat Immunol. (2022) 23:1208–21. doi: 10.1038/s41590-022-01268-1
102. Wang Z, Qi Y, Feng Y, Xu H, Wang J, Zhang L, et al. The N6-methyladenosine writer WTAP contributes to the induction of immune tolerance post kidney transplantation by targeting regulatory T cells. Lab Invest. (2022) 102:1268–79. doi: 10.1038/s41374-022-00811-w
103. Saikia M, Fu Y, Pavon-Eternod M, He C, Pan T. Genome-wide analysis of N1-methyl-adenosine modification in human tRNAs. Rna. (2010) 16:1317–27. doi: 10.1261/rna.2057810
104. Clark WC, Evans ME, Dominissini D, Zheng G, Pan T. tRNA base methylation identification and quantification via high-throughput sequencing. Rna. (2016) 22:1771–84. doi: 10.1261/rna.056531.116
105. Liu Y, Zhou J, Li X, Zhang X, Shi J, Wang X, et al. tRNA-m(1)A modification promotes T cell expansion via efficient MYC protein synthesis. Nat Immunol. (2022) 23:1433–44. doi: 10.1038/s41590-022-01301-3
106. Wang N, Tang H, Wang X, Wang W, Feng J. Homocysteine upregulates interleukin-17A expression via NSun2-mediated RNA methylation in T lymphocytes. Biochem Biophys Res Commun. (2017) 493:94–9. doi: 10.1016/j.bbrc.2017.09.069
107. Yang WL, Qiu W, Zhang T, Xu K, Gu ZJ, Zhou Y, et al. Nsun2 coupling with RoRγt shapes the fate of Th17 cells and promotes colitis. Nat Commun. (2023) 14:863. doi: 10.1038/s41467-023-36595-w
108. Zhou J, Zhang X, Hu J, Qu R, Yu Z, Xu H, et al. m(6)A demethylase ALKBH5 controls CD4(+) T cell pathogenicity and promotes autoimmunity. Sci Adv. (2021) 7:eabg0470. doi: 10.1126/sciadv.abg0470
109. Karnell JL, Rieder SA, Ettinger R, Kolbeck R. Targeting the CD40-CD40L pathway in autoimmune diseases: Humoral immunity and beyond. Adv Drug Delivery Rev. (2019) 141:92–103. doi: 10.1016/j.addr.2018.12.005
110. Chand Dakal T, Dhabhai B, Agarwal D, Gupta R, Nagda G, Meena AR, et al. Mechanistic basis of co-stimulatory CD40-CD40L ligation mediated regulation of immune responses in cancer and autoimmune disorders. Immunobiology. (2020) 225:151899. doi: 10.1016/j.imbio.2019.151899
111. Guo R, Dai J, Xu H, Zang S, Zhang L, Ma N, et al. The diagnostic significance of integrating m6A modification and immune microenvironment features based on bioinformatic investigation in aortic dissection. Front Cardiovasc Med. (2022) 9:948002. doi: 10.3389/fcvm.2022.948002
112. Xiao Q, Lei L, Ren J, Peng M, Jing Y, Jiang X, et al. Mutant NPM1-Regulated FTO-Mediated m(6)A Demethylation Promotes Leukemic Cell Survival via PDGFRB/ERK Signaling Axis. Front Oncol. (2022) 12:817584. doi: 10.3389/fonc.2022.817584
113. Cao K, Du Y, Bao X, Han M, Su R, Pang J, et al. Glutathione-bioimprinted nanoparticles targeting of N6-methyladenosine FTO demethylase as a strategy against leukemic stem cells. Small. (2022) 18:e2106558. doi: 10.1002/smll.202106558
114. Gerken T, Girard CA, Tung YC, Webby CJ, Saudek V, Hewitson KS, et al. The obesity-associated FTO gene encodes a 2-oxoglutarate-dependent nucleic acid demethylase. Science. (2007) 318:1469–72. doi: 10.1126/science.1151710
115. Liu L, Li H, Hu D, Wang Y, Shao W, Zhong J, et al. Insights into N6-methyladenosine and programmed cell death in cancer. Mol Cancer. (2022) 21:32. doi: 10.1186/s12943-022-01508-w
116. Wójcik P, Biernacki M, Wroński A, Łuczaj W, Waeg G, Žarković N, et al. Altered lipid metabolism in blood mononuclear cells of psoriatic patients indicates differential changes in psoriasis vulgaris and psoriatic arthritis. Int J Mol Sci. (2019) 20:4249. doi: 10.3390/ijms20174249
117. Robinson G, Pineda-Torra I, Ciurtin C, Jury EC. Lipid metabolism in autoimmune rheumatic disease: implications for modern and conventional therapies. J Clin Invest. (2022) 132:e148552. doi: 10.1172/JCI148552
118. Huang Y, Zhang W, Li Q, Wang Z, Yang X. Identification of m6A/m5C/m1A-associated LncRNAs for prognostic assessment and immunotherapy in pancreatic cancer. Sci Rep. (2023) 13:3661. doi: 10.1038/s41598-023-30865-9
119. Zhou W, Wang C, Chang J, Huang Y, Xue Q, Miao C, et al. RNA methylations in cardiovascular diseases, molecular structure, biological functions and regulatory roles in cardiovascular diseases. Front Pharmacol. (2021) 12:722728. doi: 10.3389/fphar.2021.722728
120. Kumar S, Nagpal R, Kumar A, Ashraf MU, Bae YS. Immunotherapeutic potential of m6A-modifiers and microRNAs in controlling acute myeloid leukaemia. Biomedicines. (2021) 9:690. doi: 10.20944/preprints202105.0193.v1
121. Shulman Z, Stern-Ginossar N. The RNA modification N(6)-methyladenosine as a novel regulator of the immune system. Nat Immunol. (2020) 21:501–12. doi: 10.1038/s41590-020-0650-4
122. Wang H, Hu X, Huang M, Liu J, Gu Y, Ma L, et al. Mettl3-mediated mRNA m(6)A methylation promotes dendritic cell activation. Nat Commun. (2019) 10:1898. doi: 10.1038/s41467-019-09903-6
123. Cui L, Ma R, Cai J, Guo C, Chen Z, Yao L, et al. RNA modifications: importance in immune cell biology and related diseases. Signal Transduct Target Ther. (2022) 7:334. doi: 10.1038/s41392-022-01175-9
124. Miao S, Li H, Song X, Liu Y, Wang G, Kan C, et al. tRNA m1A modification regulates cholesterol biosynthesis to promote antitumor immunity of CD8+ T cells. J Exp Med. (2025) 222:e20240559. doi: 10.1084/jem.20240559
125. Kong W, Biswas A, Zhou D, Fiches G, Fujinaga K, Santoso N, et al. Nucleolar protein NOP2/NSUN1 suppresses HIV-1 transcription and promotes viral latency by competing with Tat for TAR binding and methylation. PloS Pathog. (2020) 16:e1008430. doi: 10.1371/journal.ppat.1008430
126. Jin S, Li J, Shen Y, Wu Y, Zhang Z, Ma H. RNA 5-Methylcytosine Regulator NSUN3 promotes tumor progression through regulating immune infiltration in head and neck squamous cell carcinoma. Oral Dis. (2024) 30:313–28. doi: 10.1111/odi.14357
127. Banks KM, Lan Y, Evans T. Tet Proteins Regulate Neutrophil Granulation in Zebrafish through Demethylation of socs3b mRNA. Cell Rep. (2021) 34:108632. doi: 10.1016/j.celrep.2020.108632
128. Larsson L, Giraldo-Osorno PM, Garaicoa-Pazmino C, Giannobile WV, Asa’ad F. DNA and RNA methylation in periodontal and peri-implant diseases. J Dent Res. (2025) 104:131–9. doi: 10.1177/00220345241291533
129. Song W, Liu L, Liang H, Cheng H, He W, Yin Q, et al. m(6)Am methyltransferase PCIF1 regulates periodontal inflammation. J Dent Res. (2024) 103:1130–40. doi: 10.1177/00220345241271078
130. Krienke C, Kolb L, Diken E, Streuber M, Kirchhoff S, Bukur T, et al. A noninflammatory mRNA vaccine for treatment of experimental autoimmune encephalomyelitis. Science. (2021) 371:145–53. doi: 10.1126/science.aay3638
131. Huang Y, Su R, Sheng Y, Dong L, Dong Z, Xu H, et al. Small-molecule targeting of oncogenic FTO demethylase in acute myeloid leukemia. Cancer Cell. (2019) 35:677–691.e10. doi: 10.1016/j.ccell.2019.03.006
132. Rai S, Kim WS, Ando K, Choi I, Izutsu K, Tsukamoto N, et al. Oral HDAC inhibitor tucidinostat in patients with relapsed or refractory peripheral T-cell lymphoma: phase IIb results. Haematologica. (2023) 108:811–21. doi: 10.3324/haematol.2022.280996
Keywords: post-transcriptional modification, RNA modification, autoimmune diseases, CD4+ T cells, T cell differentiation
Citation: Yu H, Ma Z, Su S, Xu Z and Yi H (2025) RNA modification: a promising code to unravel the puzzle of autoimmune diseases and CD4+ T cell differentiation. Front. Immunol. 16:1563150. doi: 10.3389/fimmu.2025.1563150
Received: 19 January 2025; Accepted: 06 March 2025;
Published: 24 March 2025.
Edited by:
Noah Isakov, Ben-Gurion University of the Negev, IsraelReviewed by:
Xianyang Liu, Chongqing Medical University, ChinaCopyright © 2025 Yu, Ma, Su, Xu and Yi. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Zheng Xu, eHV6QGpsdS5lZHUuY24=; Huanfa Yi, eWlodWFuZmFAamx1LmVkdS5jbg==
†These authors have contributed equally to this work
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.