
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Immunol. , 03 April 2025
Sec. Cancer Immunity and Immunotherapy
Volume 16 - 2025 | https://doi.org/10.3389/fimmu.2025.1560322
This article is part of the Research Topic Cuproptosis and Tumors, Volume II: From Basic Research to Clinical Translation View all 3 articles
Background: Distant metastasis of colorectal cancer (CRC) significantly impacts patient prognosis. Cuproptosis is a new form of copper ion-dependent cell death. However, whether cuproptosis-related genes (CRGs) play a role in the metastatic potential of CRC remains unclear. This study focuses on CRGs-ACAD8 to explore its role and mechanism in metastatic CRC (mCRC).
Methods: Clinical sample data from TCGA, GEO, and Fujian Provincial Hospital patients were integrated to analyze ACAD8 expression and its association with the diagnosis and prognosis of CRC. Small interfering RNA, immunohistochemistry, colony formation, wound-healing assays and so on were used to evaluate the biological functions of ACAD8. Bioinformatics was applied to investigate its relationships with immune infiltration, chemotherapy sensitivity, and signaling pathways.
Results: ACAD8 expression was significantly reduced in mCRC and demonstrated excellent diagnostic performance. Patients with high ACAD8 expression had significantly better survival. ACAD8 was closely associated with immune cell infiltration, and enhanced chemotherapy sensitivity. Pathway enrichment analysis suggested that ACAD8 might inhibit the metastasis of CRC by regulating pathways such as response to metal ions and tight junction organization. Finally, experiments confirmed a positive correlation between copper levels and ACAD8 mRNA expression, with CuCl2 upregulating ACAD8 expression. Knockdown of ACAD8 induced cuproptosis. CuCl2 inhibited the proliferation, stemness, and migratory abilities of CRC cells, while si ACAD8 attenuated these effects. Moreover, CuCl2 enhanced the sensitivity of CRC cells to oxaliplatin and 5-fluorouracil, whereas si ACAD8 diminished this chemosensitizing effect.
Conclusion: As a novel tumor suppressor, low expression of CRGs-ACAD8 is associated with the metastasis of CRC. ACAD8 holds potential diagnostic and prognostic value and may contribute to the precise treatment of CRC by regulating immune infiltration and chemotherapy sensitivity.
Colorectal cancer (CRC) is among the most common malignancies globally, with distant metastasis, particularly to the liver and lungs, being a primary factor affecting patient outcomes (1). The metastatic potential of CRC is closely linked to several molecular signaling pathways (2). Additionally, the tumor microenvironment (TME) plays the vital role in promoting the metastasis of cancer (3). Despite considerable research advances, precise treatments for metastatic CRC (mCRC) remain challenging.
Uncovering mechanisms underlying the metastasis of CRC is key to improving treatment strategies. For mCRC patients, combined therapies such as chemotherapy, and immunotherapy have demonstrated efficacy. However, these treatments are often hindered by tumor heterogeneity and resistance (4, 5). Thus, future research should focus on elucidating metastatic mechanisms and developing novel biomarkers to enhance the survival rates of mCRC.
Cuproptosis, a new form of copper-dependent cell death, is intricately linked to intracellular copper ion concentrations (6). Excess copper ions can accumulate and induce cell death by promoting aggregation of lipoylated protein in the mitochondrial tricarboxylic acid cycle (7). Studies have shown significantly elevated copper levels in gastric cancer, where copper induces the lactylation of METTL16 at K229, leading to cuproptosis (8). Additionally, targeting ferroptosis and cuproptosis simultaneously using ferroptosis inducers and copper ionophores has shown promise in the treatment of hepatocellular carcinoma (9). However, few studies have explored cuproptosis mechanisms in CRC (10). The potential role of cuproptosis and cuproptosis-related genes (CRGs) in mCRC has garnered increasing interest as a critical anticancer strategy.
Given the novel insights that cuproptosis provides for cancer research, we hypothesize that CRGs may inhibit the metastasis of CRC. To investigate, we integrated bioinformatics and basic biological experiments to examine the clinical application and underlying mechanisms of CRGs-ACAD8 in CRC. We screened mCRC and cuproptosis-related datasets from the GEO and identified the novel CRG-ACAD8. In the context of cancer research, ACAD8 has been associated with a favorable prognosis in lung adenocarcinoma (LUAD), where it is considered a tumor-associated fibroblast-related gene associated with good survival outcomes (11). Furthermore, ACAD8 has been identified as a novel gene associated with prognosis and immunotherapy through amino acid metabolism pathways in LUAD (12). However, its functional role in CRC remains poorly characterized.
Using TCGA, we analyzed ACAD8 expression differences in mCRC and validated the findings with GEO and tumor samples from CRC patients at Fujian Provincial Hospital. ACAD8 protein levels were analyzed through the Human Protein Atlas (HPA) and verified via immunohistochemistry (IHC). The diagnostic or prognostic value of ACAD8 was evaluated by TCGA, GEO, and patient data (13). We further explored the correlation between ACAD8 expression, immune cell infiltration, immune checkpoint genes, and chemotherapy sensitivity. Pathway enrichment analysis highlighted the potential significance of ACAD8 in the metastasis of CRC. Finally, cell phenotype experiments demonstrated the ability of CRGs-ACAD8 to suppress the metastasis of CRC.
Data sets of mCRC were obtained from the GEO for analysis. Additionally, data from TCGA were cross-validated with GEO findings. CRC patient data from Fujian Provincial Hospital were utilized to validate the results of these two databases.
The human CRC cell line RKO was obtained from Haixing Bioscience Co., Ltd. (China). RKO cells were cultured in medium containing 10% fetal bovine serum (FBS) and 1% antibiotics in a humidified incubator at 37°C with 5% CO2.
RKO cells were digested with trypsin and centrifuged. The cells were resuspended in complete medium and seeded into 96-well plates. Each experimental group included five replicate wells. A total of 10μL of CCK-8 solution was added to each well, and optical density (OD) at 450 nm was measured using the microplate reader (13).
Logarithmic-phase RKO cells were digested with trypsin, centrifuged, and resuspended. RKO cells were seeded at a density of 400 or 450 cells per well in six-well plates. After visible colonies formed, cells were fixed and stained with crystal violet. Colonies were photographed and counted. Each experimental group included three replicate wells.
RKO cells were seeded into six-well plates at 99% confluence. A sterile pipette tip was used to create a straight scratch (“wound”). Debris was removed with sterile 1x PBS. Initial scratch images were captured using an inverted microscope. After 48 hours of incubation, additional images were taken, and Image J was used to quantify wound closure to evaluate the migration ability of cell.
The amplification of PCR and cycle threshold (CT) determination were performed using the Roche LightCycler 480 system. Reagents for reverse transcription and amplification were provided by Fujian HeRui Biotechnology Co., Ltd (14). The primer sequences for ACAD8 were: Forward sequence: 5’-ATGAGGAGGAGGAGGAGAGG-3’; Reverse sequence: 5’-TTTAGTTTGTCCACCTCTGC-3’.
RKO cells were seeded into six-well plates. When cells reached 65% confluence, they were transfected with siRNAs. Briefly, 200μL of diluted Lipofectamine 3000 was mixed with 200μL of diluted plasmid DNA. After mixing, the mixture was incubated at room temperature for 30 minutes to ensure complete encapsulation. A total of 400μL of the lipid-DNA complex was added to the cells and incubated for 12 hours. The forward sequence of si ACAD8 was: 5’-GGAGAGGAAGGAGAAGTTA-3’; Reverse sequence: 5’-UCAUUCUCCUUCUUCUCCAA-3’.
CRC tissue samples were fixed, dehydrated, paraffin-embedded, and sectioned. Sections were incubated at 60°C for 12 hours, followed by deparaffinization, hydration, and antigen retrieval. Anti-ACAD8 antibody (Product ID: CSB-PA892457LA01HU, CUSABIO, China) was added and incubated at 4°C for 12 hours. Secondary antibodies were added for incubation, and diaminobenzidine (DAB) was used for staining.
Protein samples were extracted from cells, subjected to electrophoresis, transferred to membranes, and blocked. Anti-ACAD8 or Anti-DLAT antibody was incubated at 4°C for 8 hours, followed by secondary antibodies (goat anti-rabbit IgG or goat anti-mouse IgG) for 2 hours. Bands were visualized using ECL substrate and an imaging system. Anti-DLAT antibody was purchased from CST (Catalog #12362).
ROC curves and area under the curve (AUC) values were analyzed and visualized using the R software packages pROC (version 1.18.0) and ggplot2 (version 3.3.6) (14). Survival data from the TCGA database were analyzed using the survival (version 3.3.1) and survminer (version 0.4.9) packages (13).
The ssGSEA algorithm from the GSVA package (version 1.46.0) was used. Spearman correlation analysis was performed to investigate the relationship between ACAD8 expression, immune cell infiltration, and immune checkpoint-related genes.
Using the GDSC, the impact of ACAD8 expression on chemotherapy response of CRC was analyzed (15). Ridge regression was applied to determine IC50 values using the pRRophetic package.
Differentially expressed genes (DEGs) associated with ACAD8 were analyzed. Functional annotation and pathway enrichment were performed using the clusterProfiler package (version 4.4.4) and visualized using ggplot2 (version 3.3.6).
A Copper Assay Kit (Abcam, ab272528, UK) was used to extract tumor tissues from 80 CRC patients. The assay followed the protocol of manufacturer, with OD measured at 359 nm.
Data analysis was performed using R software, GraphPad Prism (version 9.5.1), and SPSS (version 26). Depending on data distribution, statistical analysis included the Wilcoxon rank-sum test, Student’s t test, or chi-square test. Kaplan-Meier (KM) analysis was used to evaluate survival. The study workflow was shown in Figure 1.
Through an analysis of classic mCRC datasets from GEO (GSE81558, GSE49355, and GSE41568) and cuproptosis-related mRNA (GSE248083), we identified two CRGs associated with the metastasis of CRC: ACAD8 and HSPA8 (Figure 2A). To investigate which CRGs plays a central role in metastasis, RNA was extracted from paired primary CRC and liver metastasis samples at Fujian Provincial Hospital. qPCR revealed significantly reduced ACAD8 expression in liver metastasis compared to primary CRC (Figure 2B), while HSPA8 showed no significant difference (Figure 2C). TCGA data further confirmed that ACAD8 expression is markedly decreased in mCRC (Figures 2D, E). Thus, ACAD8 was selected for further study.
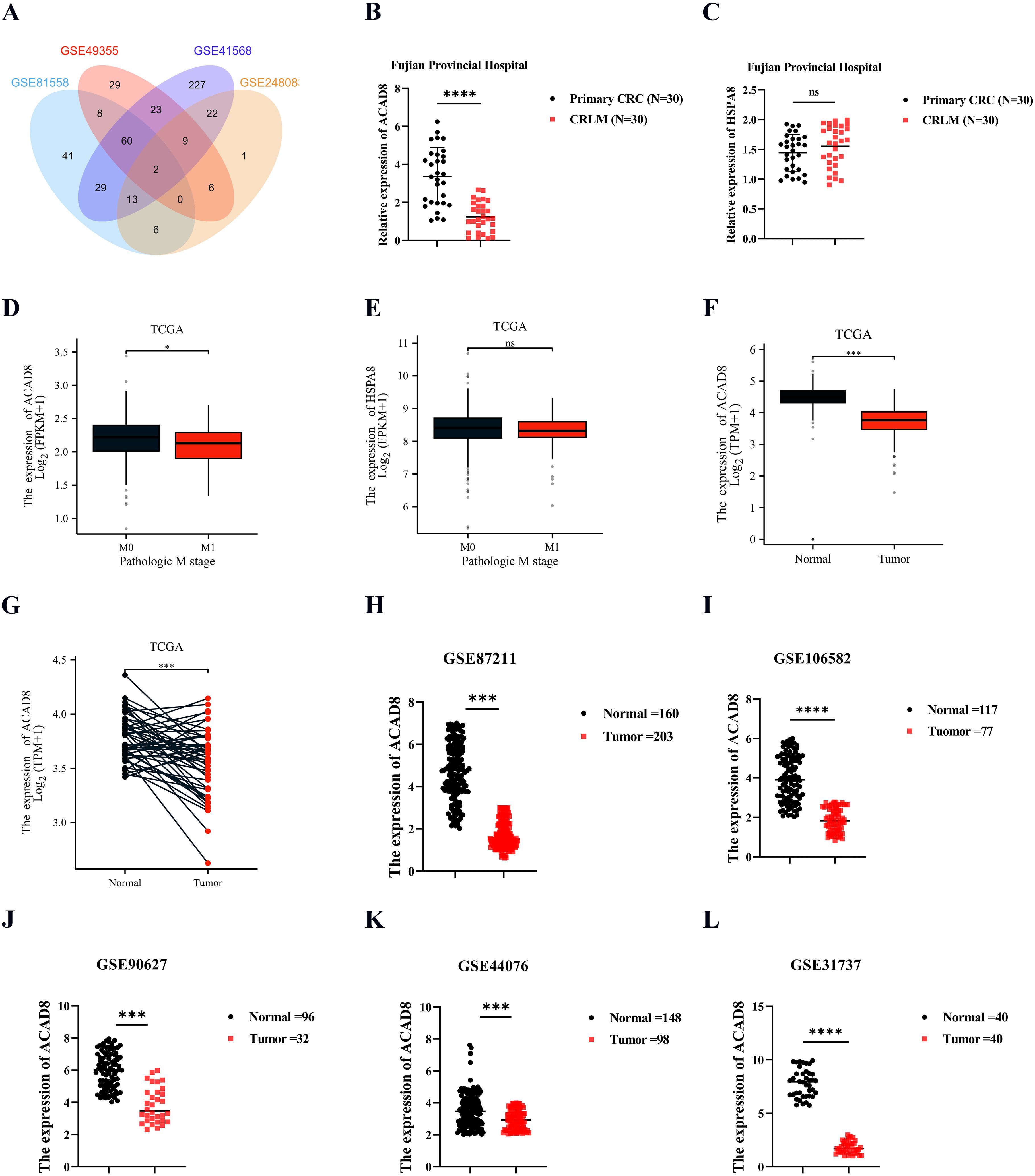
Figure 2. The RNA levels of ACAD8 in CRC. (A) ACAD8 was identified by intersecting Venn diagrams after searching the GEO dataset. (B) Analyzing the RNA levels of ACAD8 in primary CRC and matched CRLM based on data from Fujian Provincial Hospital patients. (C) Analyzing the RNA levels of HSPA8 in primary CRC and matched CRLM based on data from Fujian Provincial Hospital patients. (D) The expression difference of ACAD8 among CRC patients in different M stage. (E) The expression difference of HSPA8 among CRC patients in different M stage. (F) Analyzing the RNA levels of ACAD8 in CRC tissues and normal intestinal epithelial tissues based on TCGA. (G) Analyzing the RNA levels of ACAD8 in CRC tissues and matched normal intestinal epithelial tissues based on TCGA. Analyzing the RNA levels of ACAD8 in CRC tissues and normal intestinal epithelial tissues based on GSE87211 (H), GSE106582 (I), GSE90627 (J), GSE44076 (K), and GSE31737 (L). (P>0.05, ns. nonsignificant; P < 0.05 *; P < 0.001 ***; P < 0.0001 ****; analyses were performed using Student’ s t test or Wilcoxon rank-sum test, respectively).
To evaluate ACAD8 expression in CRC, we analyzed TCGA data comparing normal intestinal epithelium and CRC tissues, revealing significantly higher ACAD8 expression in normal tissues (Figure 2F). Paired CRC and normal epithelium samples confirmed this finding (Figure 2G). GEO datasets supported this result, demonstrating consistently higher ACAD8 expression in normal tissues than in CRC (Figures 2H–L, Supplementary Figures 1A–C). Collectively, these findings suggest that ACAD8 acts as a protective factor, with its mRNA levels significantly reduced in CRC.
After confirming ACAD8 expression in CRC, we examined whether there were also expression differences in pan-cancer. TCGA data indicated that ACAD8 was downregulated in 19 types of cancers and upregulated in 5 (Supplementary Figure S1D). Paired sample analysis across pan-cancer also indicated differential expression of ACAD8 in 14 cancer types (Supplementary Figure S1E). These results strongly suggest that ACAD8 acts as a tumor suppressor not only in CRC but also in many other cancers.
In addition to mRNA levels, we sought to determine if the protein expression of ACAD8 was also reduced in CRC. Results from the HPA database confirmed that ACAD8 protein expression was significantly lower in CRC (Figure 3A). This was further validated by IHC of patient samples from Fujian Provincial Hospital (Figure 3B). Together, these results demonstrate that both the RNA and protein levels of ACAD8 are significantly higher in normal intestinal epithelial tissues. Furthermore, the correlation between ACAD8 expression and clinical data was discussed. The low expression of ACAD8 was associated with advanced N and TNM stages (Figures 3C, D), confirming a correlation between ACAD8 and metastasis of CRC. Additionally, low expression of ACAD8 was also linked to positive surgical margins and nerve invasion (Figures 3E, F). Clinical data from Fujian Provincial Hospital confirmed the correlation between ACAD8 expression and clinical stages as observed in TCGA (Figures 3G–J).
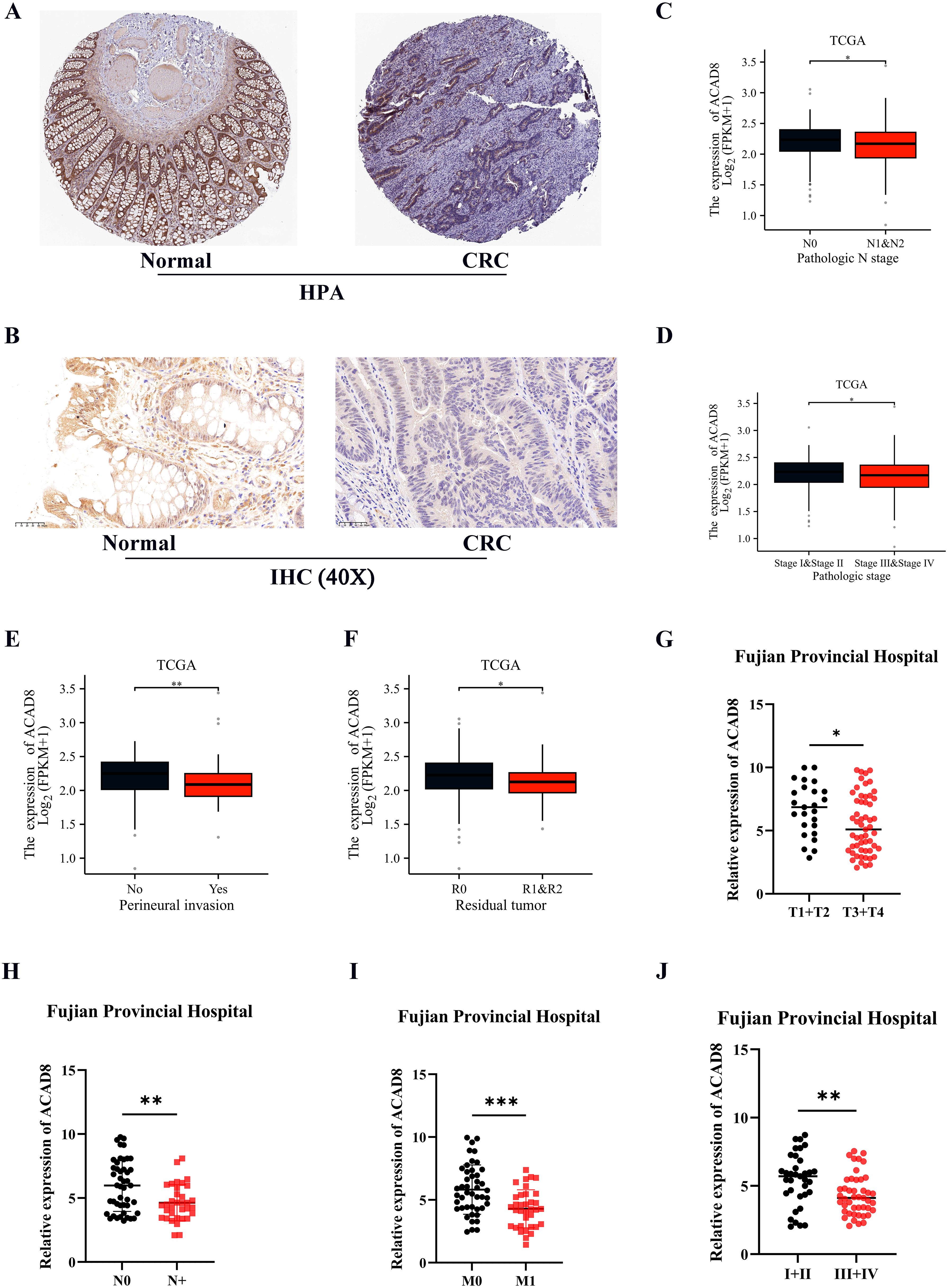
Figure 3. The protein levels of ACAD8 in CRC and association with clinical information. (A) Detected the protein expression of ACAD8 in CRC tissues and normal tissues based on the HPA database. (B) The protein expression of ACAD8 in CRC tissues and normal tissues using IHC based on patient samples. Based on TCGA, the expression difference of ACAD8 among CRC patients in different N (C), TNM stages (D), perineural invasion (E) and residual tumor (F), respectively. Based on data from Fujian Provincial Hospital patients (n=80 CRC patients), the expression difference of ACAD8 among CRC patients in different T (G), N (H), M (I), and TNM stages (J), respectively. (P>0.05, ns. nonsignificant; P < 0.05 *; P < 0.01 **; P < 0.001 ***; analyses were performed using Student’ s t test or Wilcoxon rank-sum test, respectively).
We investigated whether ACAD8 could serve as a diagnostic biomarker for CRC. ACAD8 could efficiently diagnose CRC (Figure 4A, AUC = 0.932). To cross-validate the results from a single database, we analyzed GEO datasets on normal intestinal epithelial tissues and CRC. The results supported that ACAD8 could effectively distinguish between normal tissues and CRC (Figures 4B–I, AUC values were 0.978, 0.961, 0.978, 0.905, 0.852, 0.954, 0.717, and 0.785).
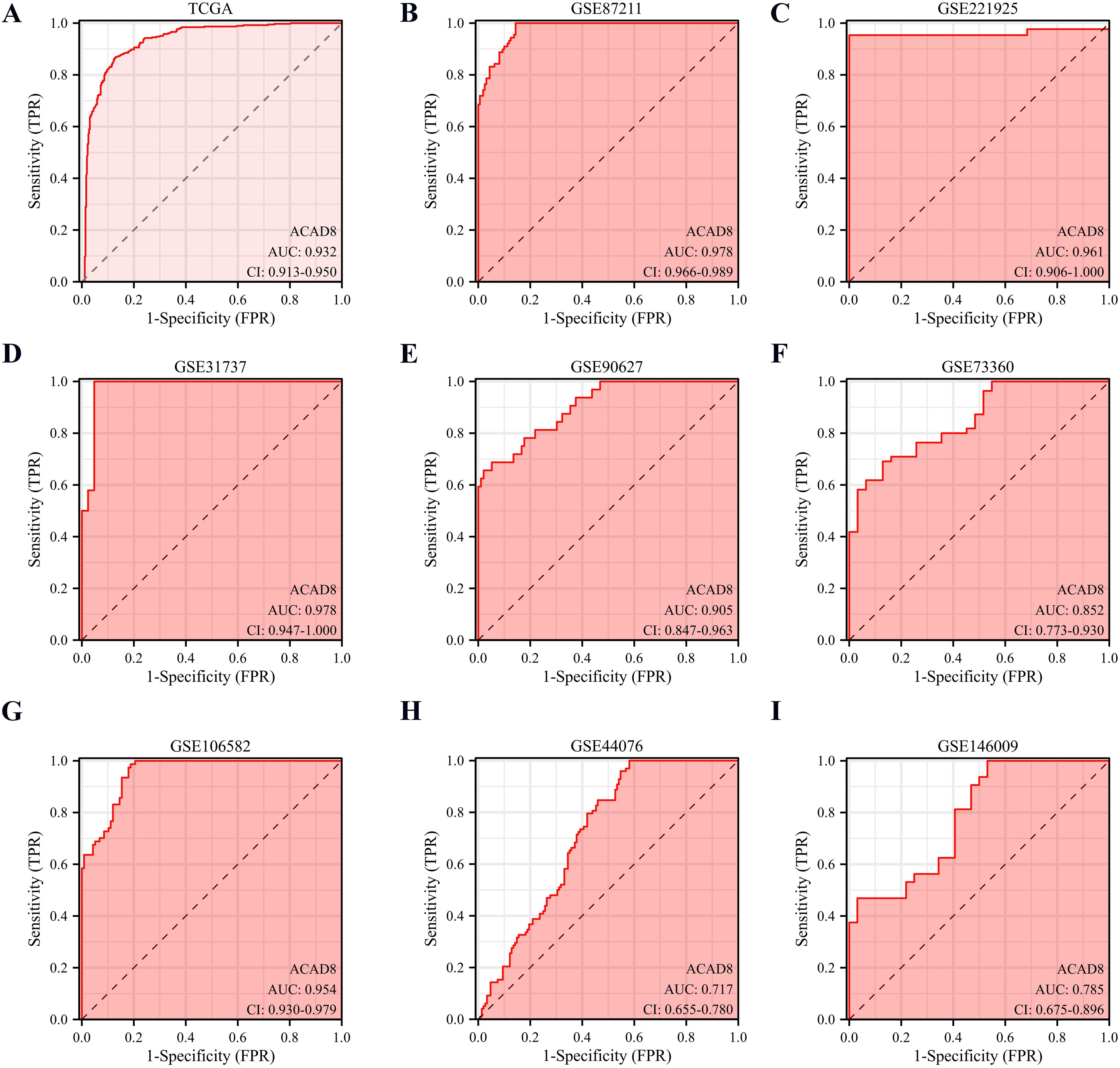
Figure 4. The diagnostic potential of ACAD8 for CRC. (A) Based on TCGA, the diagnostic capability of ACAD8 for distinguishing CRC patients from normal individuals. Based on GSE87211 (B), GSE221925 (C), GSE31737 (D), GSE90627 (E), GSE73360 (F), GSE106582 (G), GSE44076 (H) and GSE146009 (I), the diagnostic capability of ACAD8 for distinguishing CRC patients from normal individuals.
We explored whether ACAD8 could predict the prognosis of CRC. Using survival data from TCGA patients, we plotted KM curves, which showed that ACAD8 was associated with a better progression-free interval (PFI) and overall survival (OS) (Figures 5A, B). These results were further validated after excluding non-tumor-related deaths (Figure 5C). To cross-validate the findings, we analyzed the prognostic data of CRC patients from Fujian Provincial Hospital. The results confirmed that ACAD8 expression was associated with improved OS (Figure 5D). Cox regression analysis adjusted for confounding factors revealed that ACAD8 expression was significantly correlated with better PFI, disease-specific survival (DSS), and OS (Figures 5E, F, Supplementary Figure S2).
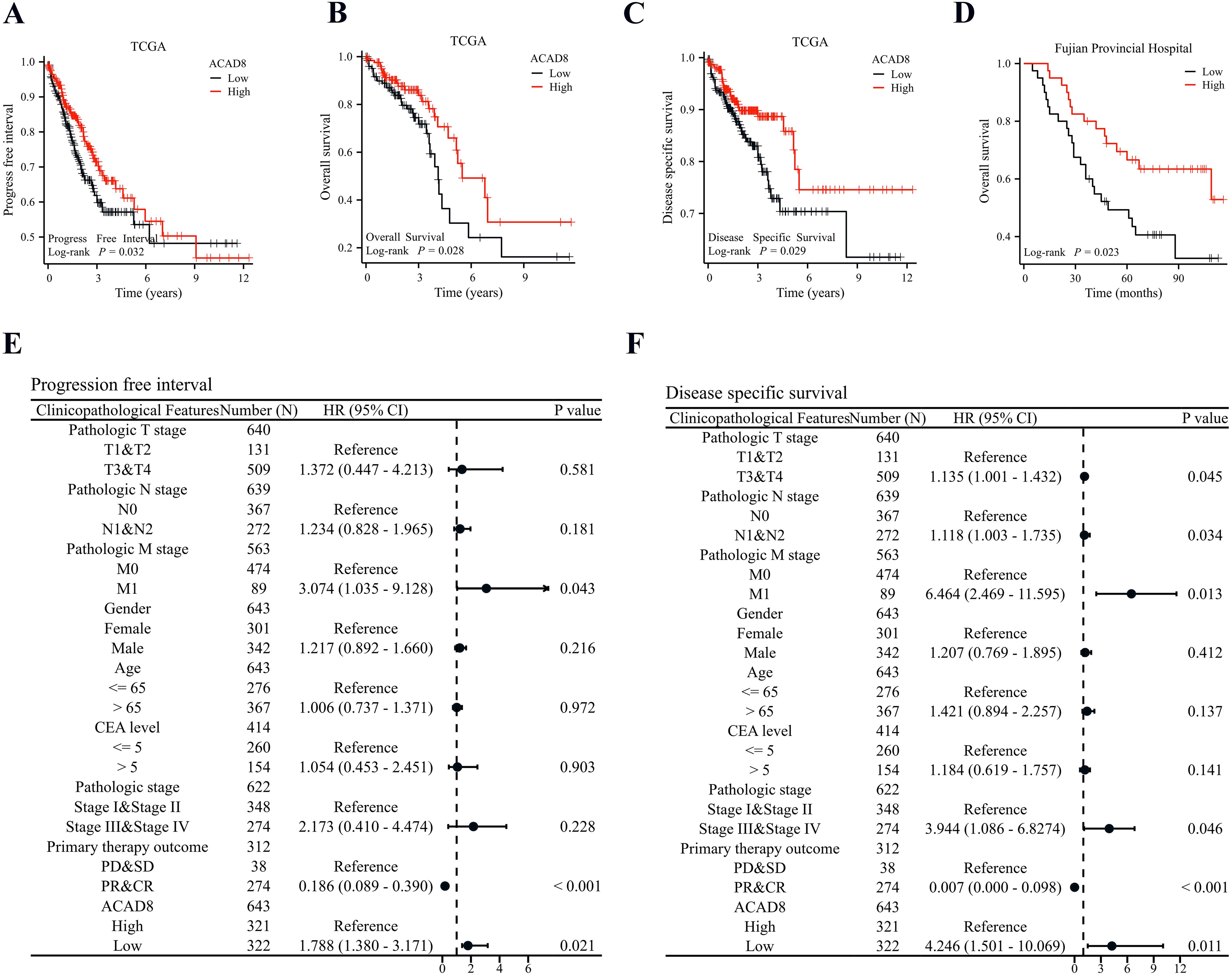
Figure 5. Low expression of ACAD8 indicates poor prognosis in CRC patients. (A) Based on TCGA, KM survival curve analysis of the impact of ACAD8 expression on PFI in CRC patients. (B) Based on TCGA, KM survival curve analysis of the impact of ACAD8 expression on OS in CRC patients. (C) Based on TCGA, KM survival curve analysis of the impact of ACAD8 expression on DSS in CRC patients. (D) Based on the data from Fujian Provincial Hospital patients, KM survival curve analysis of the impact of ACAD8 expression on OS in CRC patients. (E) A forest plot showing cox regression analysis of the impact of ACAD8 on PFI in CRC. (F) A forest plot showing cox regression analysis of the impact of ACAD8 on DSS in CRC.
The relationship between immune infiltration and tumor metastasis is a complex dynamic balance, and precise regulation of the TME is an important strategy for inhibiting tumor metastasis (16, 17). Spearman analysis revealed that ACAD8 was not correlated with 7 types of immune cells, including mast cells and NK cells (Supplementary Figures S3A–H). However, pDC cells showed a negative correlation with ACAD8 (Supplementary Figure S3I). On the other hand, B cells and DC cells showed a positive correlation with ACAD8 expression (Figures 6A–L). We further analyzed the correlation between ACAD8 expression and immune cell infiltration across pan-cancer datasets (Supplementary Figure S4).
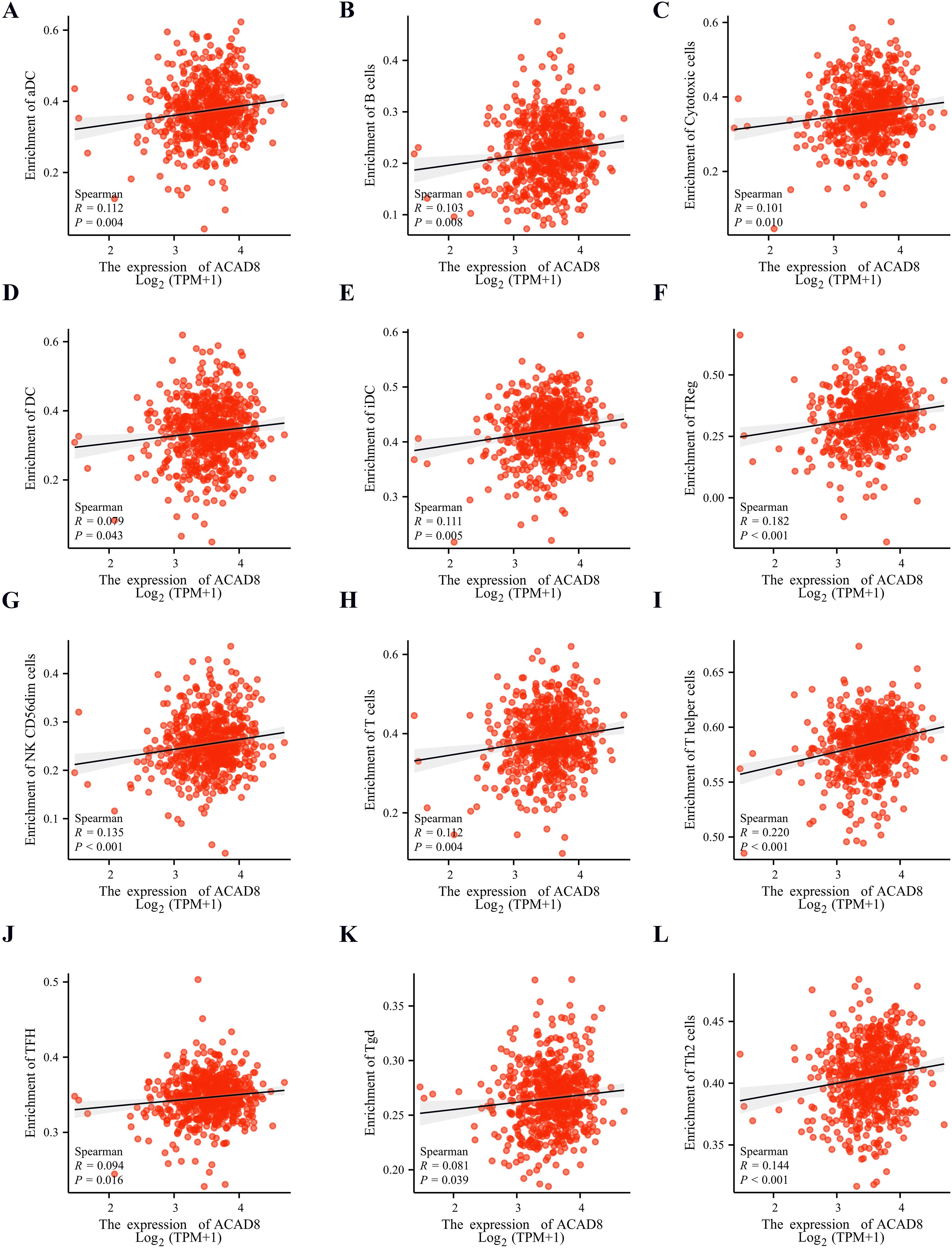
Figure 6. The relationship between expression levels of ACAD8 and immune cell infiltration. The correlation of expression levels of ACAD8 with aDC (A), B cell (B), cytotoxic cell (C), DC (D), iDC (E), Treg (F), cd56dim cell (G), T cell (H), T helper (I), TFH (J), Tgd (K), TH2 cells (L).
Immune checkpoint inhibitors (ICIs) have become an indispensable part of mCRC (18). Therefore, we explored whether ACAD8 is associated with immune checkpoint-related genes. After extracting 9 representative immune checkpoint markers from TCGA, we found that, except for SIGLEC15, all other markers showed a positive correlation with ACAD8 expression (Supplementary Figures S5A–I).
Previous studies have confirmed that copper ions can inhibit tumor chemotherapy resistance in other cancer types (19). In addition to predicting the efficacy of ICIs, we explored whether ACAD8 could enhance the sensitivity to chemotherapy drugs in CRC patients. By analyzing the GDSC database on classic chemotherapy drugs for CRC, we found that high ACAD8 levels were associated with lower IC50 values for 5-Fluorouracil, Cisplatin, Irinotecan, Gemcitabine, Rapamycin, and Paclitaxel (Figures 7A–F), suggesting that ACAD8 can increase sensitivity to chemotherapy in CRC.
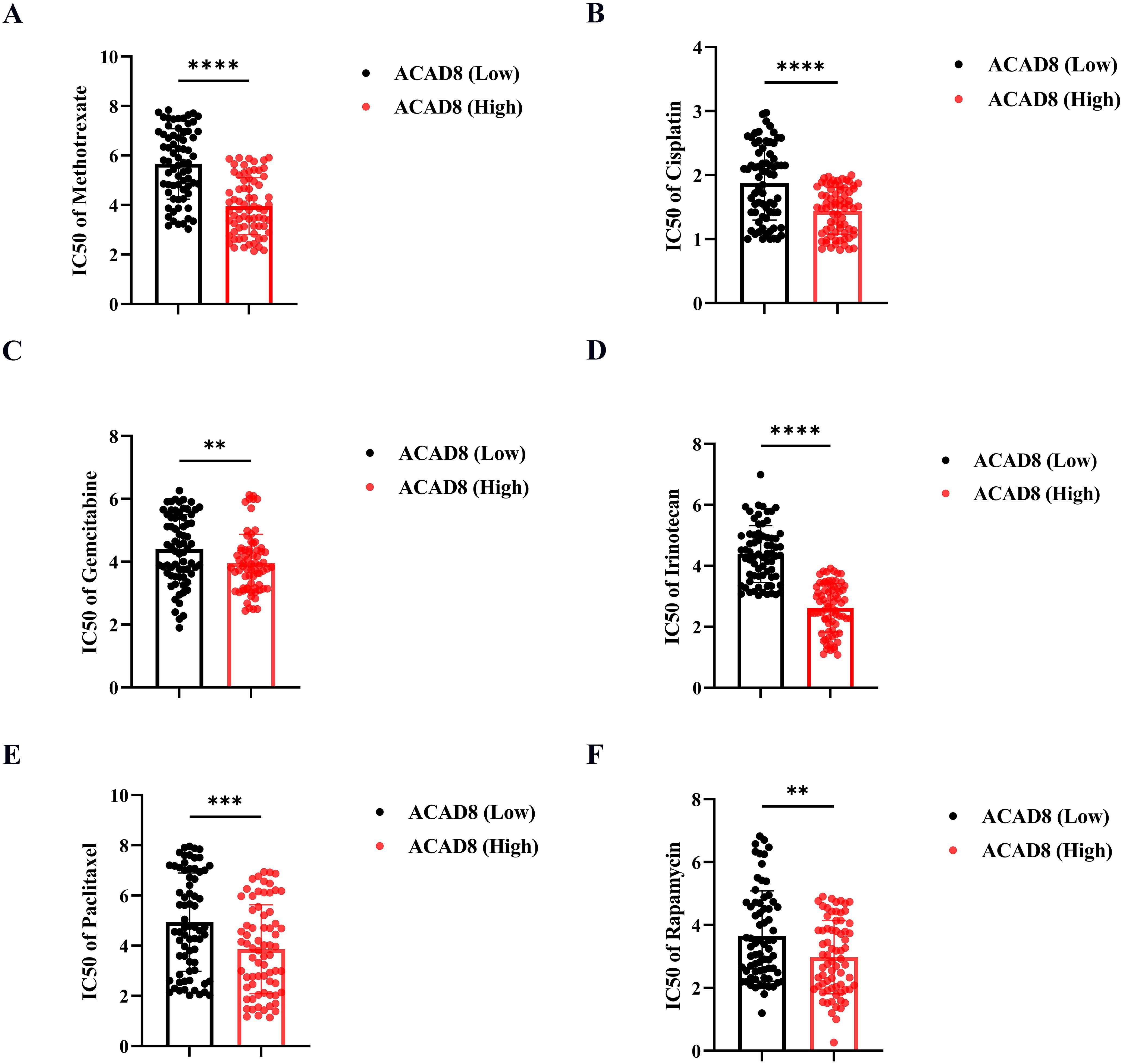
Figure 7. The correlation between ACAD8 and chemotherapy sensitivity in CRC. IC50 of response to Methotrexate (A), Cisplatin (B), Gemcitabine (C), Irinotecan (D), Paclitaxel (E), and Rapamycin (F) between the high and low expression of ACAD8 groups. (P>0.05, ns. nonsignificant; P < 0.01 **; P < 0.001 ***; P < 0.0001 ****; analyses were performed using Student’ s t test or Wilcoxon rank-sum test, respectively).
After confirming the role of ACAD8 in diagnosis, prognosis, immune infiltration, and drug response of CRC, we hypothesized potential mechanisms by which ACAD8 exerts its effects. Using TCGA data, we simulated the effects of overexpression or knockdown of ACAD8. Differential gene expression analysis revealed 761 significantly associated differentially expressed genes (DEGs) (|FC| > 1, p.adj < 0.05), with 399 genes upregulated and 365 genes downregulated (Figure 8A). Further annotation and enrichment analysis of these DEGs indicated that ACAD8 is involved in response to metal ion, cellular response to metal ion, tight junction organization and bicellular tight junction assembly (Figure 8B). KEGG analysis revealed that ACAD8 is associated with several signaling pathways, including the PI3K-Akt signaling pathway, Cell adhesion molecules, Colorectal cancer and Hippo signaling pathway (Figure 8C). These results were confirmed by GSEA analysis (Figure 8D). Additionally, TCGA data mining identified genes whose expression correlated with ACAD8. The results suggested that the expression of ACAD8 was positively correlated with FBXO40, AC134669.1, C12orf40, SMIM18, TSPYL6, IFNK, OR6B1, LIPF, AC093155.3 and TAS2R50 (Figure 8E). While ACAD8 was negatively correlated with ALB, UTF1, SRXN1, FO393400.1, FEZF2, AHSG, HBZ, NOXO1, AC004233.4, and ORM1 expression. While ACAD8 was negatively correlated with ALB, UTF1, SRXN1, FO393400.1, FEZF2, AHSG, HBZ, NOXO1, AC004233.4, and ORM1 expression (Figure 8F). These suggesting that these may be potential targets through which ACAD8 inhibits metastasis of CRC.
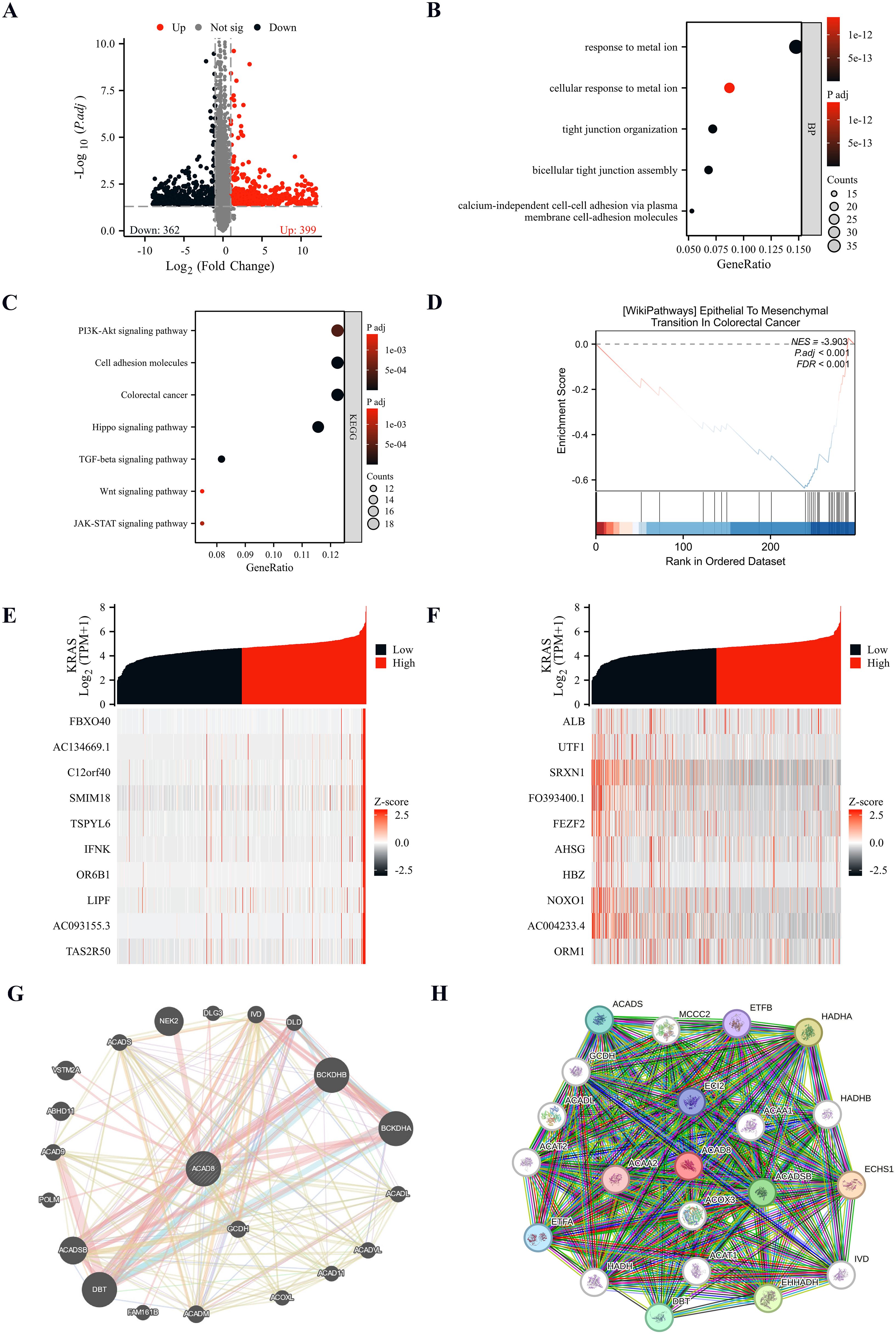
Figure 8. Pathway enrichment analysis of ACAD8 in CRC. (A) Volcano plot of ACAD8-related DEGs. (B) GO analysis of BP by ACAD8-related DEGs. (C) KEGG analysis of pathways by ACAD8-related DEGs. (D) GSEA analysis of functions affected by expression differences of ACAD8. (E, F) Analysis of co expressed genes related to ACAD8. (G) Gene interaction network diagram showing genes interacting with ACAD8 analyzed by Gene Mania. (H) Protein-protein interaction (PPI) network analysis of proteins interacting with ACAD8.
We also investigated potential interacting proteins of ACAD8. We applied GeneMANIA for a comprehensive analysis. The interaction network suggested that DBT, BCKDHA and ACADSB closely interacted with ACAD8 (Figure 8G). In addition, we further explored the possibility of interacting proteins through the STRING database, and the protein-protein interaction (PPI) network finally found that ACAA2, ACAA1, ACOX3 and other proteins interacted with ACAD8 (Figure 8H). Combined analysis of the above cooperating proteins suggests that DBT and ACADSB may be the potential interaction proteins of ACAD8 in inhibiting the metastasis and invasion of CRC.
Based on the bioinformatics findings and patient sample validations, we hypothesize that CRGs-ACAD8 may inhibit the metastasis of CRC through cuproptosis. To confirm ACAD8 as a CRG, we correlated copper levels with ACAD8 mRNA expression in CRC, finding a positive correlation (Figure 9A). We then investigated whether copper ions could upregulate ACAD8 expression. After treating CRC cells with CuCl2, we observed increased ACAD8 expression, suggesting that ACAD8 might be a key regulator in copper ion-induced cuproptosis (Figure 9B). Next, we assessed whether ACAD8 could induce cuproptosis. Given that protein lipidation is a central factor in cuproptosis, we used lipidated protein aggregation by dihydrolipoamide S-acetyltransferase (DLAT) as an indicator of cuproptosis (20). The expression level of ACAD8 was inhibited by siRNA, and the success of knocking down ACAD8 was confirmed by RT-qPCR (Figure 9C). WB showed that CuCl2 activated cuproptosis. However, inhibition of ACAD8 resulted in decreased protein aggregation of the cuproptosis marker DLAT (Figure 9D). The above experiments support that as CRGs, ACAD8 can induce cuproptosis.
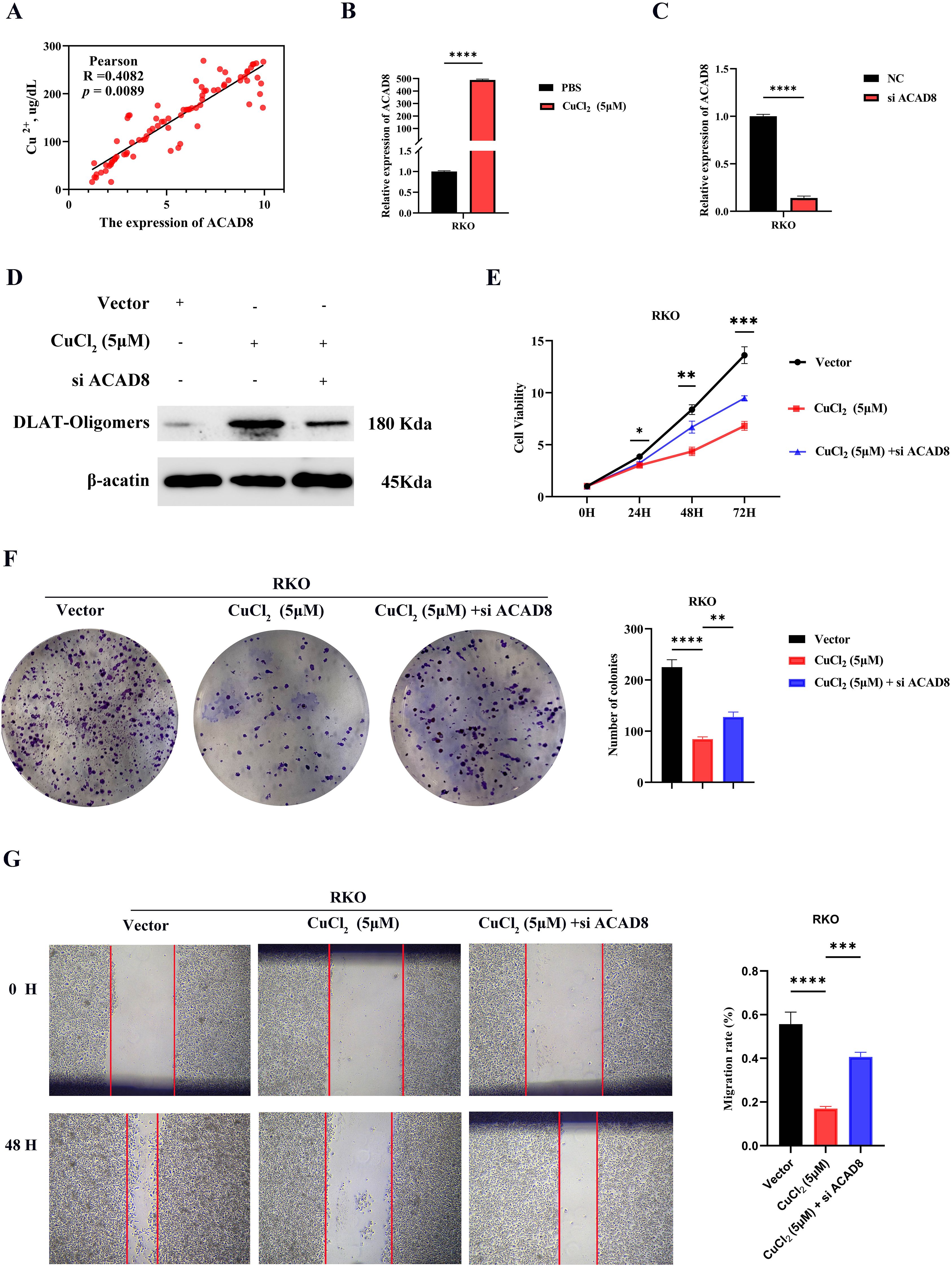
Figure 9. Knockdown of CRGs-ACAD8 promoted the metastatic ability of CRC. (A) Analysis of the correlation between Cu2+ concentration and ACAD8 expression in CRC. (B) RT-qPCR was used to observe the expression of ACAD8 in CRC cells treated with CuCl2. (C) RT-qPCR confirmed the knockdown of ACAD8. (D) WB was used to observe the changes of cuproptosis markers in CRC cells treated with CuCl2 and si ACAD8. (E) CCK8 assay detected the effect of CuCl2 and si ACAD8 on proliferation of CRC. (F) Colony formation assay detected the effect of CuCl2 and si ACAD8 on stemness capability of CRC. (G) The scratch wound assay confirmed the effect of CuCl2 and si ACAD8 on migration ability of CRC. (P>0.05, ns. nonsignificant; P < 0.05 *; P < 0.01 **; P < 0.001 ***; P < 0.0001 ****; analyses were performed using Student’s t test, Wilcoxon rank-sum test, One-way analysis of covariance or Kruskal-Wallis test, respectively).
The above experiments confirmed that as CRGs, ACAD8 can induce cuproptosis. However, whether ACAD8 can inhibit metastasis requires further experimental verification. Cell proliferation provides the cell base for metastasis, cell stemness determines the survival ability of CRC cells in heterogeneous environments, and migration ability is the core driving force for tumor metastasis (21, 22). These three factors work together to constitute the key link of tumor metastasis. Therefore, we subsequently verified the effects of cuproptosis and si ACAD8 on the metastasis ability of CRC through CCK8, colony formation, and scratch wound assay. CCK8 showed that CuCl2 inhibited the proliferation of CRC, and si ACAD8 inhibited the effect of CuCl2 (Figure 9E). The colony formation assay also found that compared with PBS, CuCl2 indeed inhibited the stemness ability of CRC, and si ACAD8 also inhibited the effect of CuCl2 (Figure 9F). In addition, the migration assay also supported the inhibition of CuCl2 and ACAD8 on the migration ability of CRC (Figure 9G). Oxaliplatin and 5-fluorouracil are classical chemotherapeutic agents for CRC. Our findings demonstrated that CuCl2 enhances the sensitivity of CRC cells to oxaliplatin and 5-fluorouracil, whereas siACAD8 attenuated the chemosensitizing effect induced by CuCl2 (Supplementary Figure S6). In conclusion, in CRC, as CRGs, ACAD8 inhibits tumor metastasis by inducing cuproptosis and suppressing cell proliferation, stemness, and migration ability.
The metastasis of CRC is a critical factor influencing patient prognosis. Comprehensive treatments, including surgery, immunotherapy, and chemotherapy, have improved the prognosis of mCRC, but tumor heterogeneity and drug resistance still limit the effectiveness of these therapies (23). Cuproptosis, a novel copper-dependent cell death mechanism, has attracted widespread attention, but its role in metastasis of CRC remains underexplored (24). By modulating the pathways associated with cuproptosis, it may be possible to inhibit the metastasis of CRC. Copper, as a target in drug delivery systems, holds promise as a new direction for future cancer therapy (25). Given that both metastatic mechanisms and CRGs are hot research topics, exploring these areas is significant for CRC.
The combination of bioinformatics and mechanistic experiments has become an indispensable component of biological research (13, 14, 26), particularly in recent years as a popular research format. Public databases offer an open and shared platform for cancer therapy research, not only accelerating data collection and analysis but also advancing the development of precision medicine (27). By deeply mining and integrating these datasets, researchers can more efficiently identify molecular mechanisms of cancer, optimize treatment strategies, and provide robust support for personalized cancer therapy in the future (28). This study, integrating both public databases and patient sample data, proposes for the first time that ACAD8, as CRG, plays a suppressive role in the distant metastasis of CRC.
We found that ACAD8 is the only gene that satisfies the criteria for involvement in both distant metastasis and cuproptosis of CRC. Previous studies have shown that regulating ACAD8 in adipocytes can induce defects in the metabolism, ultimately leading to impaired adipocyte metabolism (29). Additionally, as a gene related to lipid metabolism, ACAD8 is influenced by the inflammatory microenvironment and can manipulate specific lipid metabolism to promote skin homeostasis (30). However, there are no reports of ACAD8 in CRC, highlighting its research potential. The low expression of ACAD8 in CRC is closely associated with, advanced TNM staging, and nerve invasion, suggesting that it may act as a suppressor of metastasis in CRC. This result was confirmed by ROC analysis, prognostic analysis, and functional experiments, though the underlying mechanisms still require further exploration. Furthermore, given the advantages of ACAD8 in diagnosing CRC and predicting prognosis, we need to focus on the combined use of ACAD8 and other biomarkers (e.g., CEA) for diagnosis of CRC (31). This validation should include both early and advanced-stage CRC patients to assess the gene’s sensitivity and specificity across different disease stages.
Immune infiltration in CRC is closely related to patient prognosis. Certain immune cell types are associated with better prognosis due to their anti-tumor effects (32). In contrast, immunosuppressive cells are often associated with drug resistance, and poor prognosis (33), which aligns with our findings that ACAD8 has a negative correlation with these immune cells. Immunotherapy for CRC is continuously evolving, and the level and type of immune cell infiltration significantly impact the effectiveness of such therapies. Our analysis reveals that ACAD8 plays an important role in immune infiltration within CRC, suggesting that ACAD8 may suppress tumor metastasis by activating immune cell infiltration. Immune cell infiltration is increasingly recognized for its role in development, progression, prognosis evaluation, and immunotherapy of CRC (34). However, it remains unclear whether ACAD8 directly influences immune cell infiltration or if other mediating factors are involved, or if immune cell infiltration affects ACAD8 expression. This is a complex issue that requires further investigation. Future research will further explore the relationship between ACAD8 and immune cell types and distribution, as well as its interactions with tumor cells, to enable more personalized and precise immunotherapy strategies.
ICIs have become a crucial component in the treatment of mCRC (35). By evaluating immune cell infiltration status, and tumor immune evasion mechanisms, researchers can develop new biomarkers to identify patients who may benefit from immunotherapy. Surprisingly, the positive correlation between immune checkpoint genes and the tumor suppressor ACAD8 was found. Tumor suppressor genes help in the immune recognition of tumor cells, thereby enhancing immune surveillance (36). The positive correlation between ACAD8 and immune checkpoint gene expression could be a result of adaptive changes in the TME (37). In some cases, tumor suppressor genes may support tumor growth by inducing immune tolerance mechanisms in tumor cells. These mechanisms may include the accumulation of immunosuppressive cells, such as Tregs and MDSCs, as well as the upregulation of immune checkpoint molecules (38). The positive correlation between ACAD8 and immune checkpoint genes may also stem from complex signaling pathways within the TME (39). Overall, the positive correlation between tumor suppressor genes and immune checkpoint genes may indicate a complex adaptive regulation of the TME, which could influence immune evasion and the tumor immune response. However, this requires further mechanistic exploration to address this complex phenomenon.
Chemotherapy resistance remains a major challenge in CRC (40). Chemotherapies, such as 5-fluorouracil, cisplatin, and irinotecan, remain essential in CRC, but the onset of resistance leads to a significant reduction in treatment efficacy and affects patient survival (41). Our study reveals that copper ions can inhibit chemotherapy resistance of CRC, and that ACAD8 enhances the sensitivity to chemotherapy drugs in CRC patients, highlighting its significant clinical translational potential as a CRGs. Previous research has shown that copper ions alter the expression of various transport proteins and enzymes, modifying the response mechanisms of cell and affecting drug tolerance (19). Additionally, cuproptosis in tumor cells may enhance their sensitivity to chemotherapy drugs by regulating energy metabolism and oxidative stress responses (42). Studying the mechanisms of cuproptosis, ACAD8, and its role in chemotherapy resistance could help overcome chemotherapy resistance in tumors and improve therapeutic outcomes.
Pathway enrichment analysis aims to identify and analyze biological pathways associated with ACAD8, by comparing experimental data with known biological pathway databases, revealing how genes act together through complex signaling and metabolic pathways in cells or organisms (43). Our pathway enrichment analysis strongly indicates that ACAD8 is a CRG. These findings suggest that ACAD8 may be involved in inhibiting invasion and metastasis of CRC through various signaling pathways, particularly those related to copper ions. The most significantly enriched pathway, “response to metal ion,” has been shown to induce cell death through different mechanisms, such as triggering biocatalysis, activating immune pathways, interfering with osmotic pressure, and generating pro-oxidant effects (44). Future research should further explore how ACAD8 regulates these signaling pathways. Further protein interaction studies suggest that ACAD8 may interact with DBT and ACADSB, among others. Previous research has confirmed that DBT is a significant prognostic marker related to the Hippo pathway, and its downregulation is activated by METTL3-mediated m6A modification (45). ACADSB, a metabolic gene, regulates total cholesterol metabolism through NAT10-dependent RNA acetylation (46). Future research will need to combine immunoprecipitation and mass spectrometry to confirm the interaction sites of ACAD8 with these proteins. Furthermore, phenotypic experiments are a core part of validating bioinformatics (47). Experimental validation demonstrated that ACAD8 can induce cuproptosis and inhibit the migration of CRC, suggesting that it can suppress distant metastatic potential of CRC. These findings strongly indicate that ACAD8 plays a crucial role in cuproptosis, with its inhibitory effects mediated through this mechanism.
Although we provide insight into the potential role and mechanisms of ACAD8 in metastasis of CRC, there are still certain limitations. First, the functional experiments mainly relied on in vitro cell lines and patient samples, lacking larger-scale clinical validation. Second, while we enhanced ACAD8 expression through copper ion, whether ACAD8 can effectively induce cuproptosis in vivo and improve CRC prognosis still requires further validation in animal models. Additionally, the role of ACAD8 in other cancers and its potential therapeutic applications also require in-depth investigation. Future studies could explore the potential therapeutic applications of ACAD8 in various cancers, particularly in the context of amino acid metabolism and its impact on tumor progression and immune response.
In-depth research on cuproptosis holds promise for providing more precise and effective treatment options for mCRC. This study confirms that ACAD8, as a CRG, is a novel tumor suppressor in CRC, inhibiting metastasis by inducing cuproptosis. The potential of ACAD8 in diagnosis, prognosis, enhancing immune infiltration, and increasing chemotherapy sensitivity warrants further exploration in CRC. These findings lay the foundation for individualized therapeutic strategies targeting ACAD8 to combat CRC progression and metastasis.
The raw data supporting the conclusions of this article will be made available by the authors, without undue reservation.
Approval of the research protocol was obtained by the Institutional Reviewer Board of Fujian Provincial Hospital (Number: K2024-11-079). This study has been examined by the ethics committee and have therefore been performed in accordance with the ethical standards laid down in the Declaration of Helsinki. Written informed consent to participate in this study was provided by the participants or the participants' legal guardians/next of kin in accordance with the national legislation and the institutional requirements.
HZ: Conceptualization, Data curation, Investigation, Methodology, Writing – original draft. YC: Writing – original draft. SH: Writing – review & editing.
The author(s) declare that no financial support was received for the research and/or publication of this article.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fimmu.2025.1560322/full#supplementary-material
1. Siegel RL, Giaquinto AN, Jemal A. Cancer statistics, 2024. CA: Cancer J Clin. (2024) 74:12–49. doi: 10.3322/caac.21820
2. Jia SN, Han YB, Yang R, Yang ZC. Chemokines in colon cancer progression. Semin Cancer Biol. (2022) 86:400–7. doi: 10.1016/j.semcancer.2022.02.007
3. Wang Y, Zhong X, He X, Hu Z, Huang H, Chen J, et al. Liver metastasis from colorectal cancer: pathogenetic development, immune landscape of the tumour microenvironment and therapeutic approaches. J Exp Clin Cancer research: CR. (2023) 42:177. doi: 10.1186/s13046-023-02729-7
4. Chu X, Li X, Zhang Y, Dang G, Miao Y, Xu W, et al. Integrative single-cell analysis of human colorectal cancer reveals patient stratification with distinct immune evasion mechanisms. Nat Cancer. (2024) 5:1409–26. doi: 10.1038/s43018-024-00807-z
5. Cañellas-Socias A, Sancho E, Batlle E. Mechanisms of metastatic colorectal cancer. Nat Rev Gastroenterol Hepatol. (2024) 21:609–25. doi: 10.1038/s41575-024-00934-z
6. Xie J, Yang Y, Gao Y, He J. Cuproptosis: mechanisms and links with cancers. Mol Cancer. (2023) 22:46. doi: 10.1186/s12943-023-01732-y
7. Liu YT, Chen L, Li SJ, Wang WY, Wang YY, Yang QC, et al. Dysregulated Wnt/B-catenin signaling confers resistance to cuproptosis in cancer cells. Cell Death differentiation. (2024) 31:1452–66. doi: 10.1038/s41418-024-01341-2
8. Sun L, Zhang Y, Yang B, Sun S, Zhang P, Luo Z, et al. Lactylation of Mettl16 promotes cuproptosis via M(6)a-modification on fdx1 mrna in gastric cancer. Nat Commun. (2023) 14:6523. doi: 10.1038/s41467-023-42025-8
9. Wang W, Lu K, Jiang X, Wei Q, Zhu L, Wang X, et al. Ferroptosis inducers enhanced cuproptosis induced by copper ionophores in primary liver cancer. J Exp Clin Cancer research: CR. (2023) 42:142. doi: 10.1186/s13046-023-02720-2
10. Wang Y, Pei P, Yang K, Guo L, Li Y. Copper in colorectal cancer: from copper-related mechanisms to clinical cancer therapies. Clin Trans Med. (2024) 14:e1724. doi: 10.1002/ctm2.1724
11. Peng J, He C, Yan H, Zhou W. Prognostic value of genes related to cancer-associated fibroblasts in lung adenocarcinoma. Technol Health Care. (2023) 31:2339–54. doi: 10.3233/thc-230453
12. Xiang H, Kasajima R, Azuma K, Tagami T, Hagiwara A, Nakahara Y, et al. Multi-omics analysis-based clinical and functional significance of a novel prognostic and immunotherapeutic gene signature derived from amino acid metabolism pathways in lung adenocarcinoma. Front Immunol. (2024) 15:1361992. doi: 10.3389/fimmu.2024.1361992
13. Zhuang H, Chen Y. Integrated bioinformatics analysis and validation identify Kir2dl4 as a novel biomarker for predicting chemotherapy resistance and prognosis in colorectal cancer. Heliyon. (2024) 10:e37896. doi: 10.1016/j.heliyon.2024.e37896
14. Chen Y, Zheng Y, Liu S. Kras mutation promotes the colonization of fusobacterium nucleatum in colorectal cancer by down-regulating sertad4. J Cell Mol Med. (2024) 28:e70182. doi: 10.1111/jcmm.70182
15. Pino JC, Posso C, Joshi SK, Nestor M, Moon J, Hansen JR, et al. Mapping the proteogenomic landscape enables prediction of drug response in acute myeloid leukemia. Cell Rep Med. (2024) 5:101359. doi: 10.1016/j.xcrm.2023.101359
16. Dong D, Yu X, Xu J, Yu N, Liu Z, Sun Y. Cellular and molecular mechanisms of gastrointestinal cancer liver metastases and drug resistance. Drug resistance updates: Rev commentaries antimicrobial Anticancer chemother. (2024) 77:101125. doi: 10.1016/j.drup.2024.101125
17. Chen Y, Liu S, Tan S, Zheng Y, Chen Y, Yang C, et al. Kras mutations promote the intratumoral colonization of enterotoxigenic bacteroides fragilis in colorectal cancer through the regulation of the Mirna3655/Surf6/Irf7/Ifnβ Axis. Gut Microbes. (2024) 16:2423043. doi: 10.1080/19490976.2024.2423043
18. Lynch C, Pitroda SP, Weichselbaum RR. Radiotherapy, immunity, and immune checkpoint inhibitors. Lancet Oncol. (2024) 25:e352–e62. doi: 10.1016/s1470-2045(24)00075-5
19. Wang Y, Chen Y, Zhang J, Yang Y, Fleishman JS, Wang Y, et al. Cuproptosis: A novel therapeutic target for overcoming cancer drug resistance. Drug resistance updates: Rev commentaries antimicrobial Anticancer chemother. (2024) 72:101018. doi: 10.1016/j.drup.2023.101018
20. Tsvetkov P, Coy S, Petrova B, Dreishpoon M, Verma A, Abdusamad M, et al. Copper induces cell death by targeting lipoylated Tca cycle proteins. Sci (New York NY). (2022) 375:1254–61. doi: 10.1126/science.abf0529
21. He Y, Shi Q, Ling Y, Guo H, Fei Y, Wu R, et al. Ablim1, a novel ubiquitin E3 ligase, promotes growth and metastasis of colorectal cancer through targeting Iĸbα Ubiquitination and activating Nf-ĸb signaling. Cell Death differentiation. (2024) 31:203–16. doi: 10.1038/s41418-024-01256-y
22. Sher G, Masoodi T, Patil K, Akhtar S, Kuttikrishnan S, Ahmad A, et al. Dysregulated Foxm1 signaling in the regulation of cancer stem cells. Semin Cancer Biol. (2022) 86:107–21. doi: 10.1016/j.semcancer.2022.07.009
23. Farin HF, Mosa MH, Ndreshkjana B, Grebbin BM, Ritter B, Menche C, et al. Colorectal cancer organoid-stroma biobank allows subtype-specific assessment of individualized therapy responses. Cancer Discovery. (2023) 13:2192–211. doi: 10.1158/2159-8290.Cd-23-0050
24. Tang D, Kroemer G, Kang R. Targeting cuproplasia and cuproptosis in cancer. Nat Rev Clin Oncol. (2024) 21:370–88. doi: 10.1038/s41571-024-00876-0
25. Qiao L, Zhu G, Jiang T, Qian Y, Sun Q, Zhao G, et al. Self-destructive copper carriers induce pyroptosis and cuproptosis for efficient tumor immunotherapy against dormant and recurrent tumors. Advanced materials (Deerfield Beach Fla). (2024) 36:e2308241. doi: 10.1002/adma.202308241
26. Li A, Zhang K, Zhou J, Li M, Fan M, Gao H, et al. Bioinformatics and experimental approach identify lipocalin 2 as a diagnostic and prognostic indicator for lung adenocarcinoma. Int J Biol macromolecules. (2024) 272:132797. doi: 10.1016/j.ijbiomac.2024.132797
27. Li R, Qu H, Wang S, Chater JM, Wang X, Cui Y, et al. Cancermirnome: an interactive analysis and visualization database for mirnome profiles of human cancer. Nucleic Acids Res. (2022) 50:D1139–d46. doi: 10.1093/nar/gkab784
28. Wu S, Huang Y, Zhang M, Gong Z, Wang G, Zheng X, et al. Ascancer atlas: A comprehensive knowledgebase of alternative splicing in human cancers. Nucleic Acids Res. (2023) 51:D1196–d204. doi: 10.1093/nar/gkac955
29. Green CR, Alaeddine LM, Wessendorf-Rodriguez KA, Turner R, Elmastas M, Hover JD, et al. Impaired branched-chain amino acid (Bcaa) catabolism during adipocyte differentiation decreases glycolytic flux. J Biol Chem. (2024) 300:108004. doi: 10.1016/j.jbc.2024.108004
30. Seiringer P, Hillig C, Schäbitz A, Jargosch M, Pilz AC, Eyerich S, et al. Spatial transcriptomics reveals altered lipid metabolism and inflammation-related gene expression of sebaceous glands in psoriasis and atopic dermatitis. Front Immunol. (2024) 15:1334844. doi: 10.3389/fimmu.2024.1334844
31. Wu Q, Chen P, Shu C, Chen L, Jin Z, Huang J, et al. Survival outcomes of stage I colorectal cancer: development and validation of the aceply model using two prospective cohorts. BMC Med. (2023) 21:3. doi: 10.1186/s12916-022-02693-7
32. Zhang X, Yu D, Wu D, Gao X, Shao F, Zhao M, et al. Tissue-resident lachnospiraceae family bacteria protect against colorectal carcinogenesis by promoting tumor immune surveillance. Cell Host Microbe. (2023) 31:418–32.e8. doi: 10.1016/j.chom.2023.01.013
33. Yue T, Li J, Zhu J, Zuo S, Wang X, Liu Y, et al. Hydrogen sulfide creates a favorable immune microenvironment for colon cancer. Cancer Res. (2023) 83:595–612. doi: 10.1158/0008-5472.Can-22-1837
34. de Vries NL, van de Haar J, Veninga V, Chalabi M, Ijsselsteijn ME, van der Ploeg M, et al. [amp]]Gamma;δ T cells are effectors of immunotherapy in cancers with Hla class I defects. Nature. (2023) 613:743–50. doi: 10.1038/s41586-022-05593-1
35. Emiloju OE, Sinicrope FA. Neoadjuvant immune checkpoint inhibitor therapy for localized deficient mismatch repair colorectal cancer: A review. JAMA Oncol. (2023) 9:1708–15. doi: 10.1001/jamaoncol.2023.3323
36. Martin TD, Patel RS, Cook DR, Choi MY, Patil A, Liang AC, et al. The adaptive immune system is a major driver of selection for tumor suppressor gene inactivation. Sci (New York NY). (2021) 373:1327–35. doi: 10.1126/science.abg5784
37. Skoulidis F, Araujo HA, Do MT, Qian Y, Sun X, Cobo AG, et al. Ctla4 blockade abrogates Keap1/Stk11-related resistance to Pd-(L)1 inhibitors. Nature. (2024) 635:462–71. doi: 10.1038/s41586-024-07943-7
38. Ma Y, Qi Y, Zhou Z, Yan Y, Chang J, Zhu X, et al. Shenqi fuzheng injection modulates tumor fatty acid metabolism to downregulate mdscs infiltration, enhancing Pd-L1 antibody inhibition of intracranial growth in melanoma. Phytomed: Int J phytother phytopharmacol. (2024) 122:155171. doi: 10.1016/j.phymed.2023.155171
39. Goenka A, Khan F, Verma B, Sinha P, Dmello CC, Jogalekar MP, et al. Tumor microenvironment signaling and therapeutics in cancer progression. Cancer Commun (London England). (2023) 43:525–61. doi: 10.1002/cac2.12416
40. Johnson D, Chee CE, Wong W, Lam RCT, Tan IBH, Ma BBY. Current advances in targeted therapy for metastatic colorectal cancer - clinical translation and future directions. Cancer Treat Rev. (2024) 125:102700. doi: 10.1016/j.ctrv.2024.102700
41. Kaur E, Agrawal R, Arun R, Madhavan V, Srivastava V, Kumar D, et al. Small molecules that disrupt Rad54-Blm interaction hamper tumor proliferation in colon cancer chemoresistance models. J Clin Invest. (2024) 134:e161941. doi: 10.1172/jci161941
42. Huang Y, Liu X, Zhu J, Chen Z, Yu L, Huang X, et al. Enzyme core spherical nucleic acid that enables enhanced cuproptosis and antitumor immune response through alleviating tumor hypoxia. J Am Chem Soc. (2024) 146:13805–16. doi: 10.1021/jacs.3c14247
43. Müller J, Bayer FP, Wilhelm M, Schuh MG, Kuster B, The M. Ptmnavigator: interactive visualization of differentially regulated post-translational modifications in cellular signaling pathways. Nat Commun. (2025) 16:510. doi: 10.1038/s41467-024-55533-y
44. Hu H, Xu Q, Mo Z, Hu X, He Q, Zhang Z, et al. New anti-cancer explorations based on metal ions. J nanobiotechnol. (2022) 20:457. doi: 10.1186/s12951-022-01661-w
45. Miao D, Wang Q, Shi J, Lv Q, Tan D, Zhao C, et al. N6-methyladenosine-modified Dbt alleviates lipid accumulation and inhibits tumor progression in clear cell renal cell carcinoma through the Anxa2/Yap axis-regulated hippo pathway. Cancer Commun (London England). (2023) 43:480–502. doi: 10.1002/cac2.12413
46. Dalhat MH, Mohammed MRS, Alkhatabi HA, Rehan M, Ahmad A, Choudhry H, et al. Nat10: an Rna cytidine transferase regulates fatty acid metabolism in cancer cells. Clin Trans Med. (2022) 12:e1045. doi: 10.1002/ctm2.1045
47. Rajadnya R, Sharma N, Mahajan A, Ulhe A, Patil R, Hegde M, et al. Novel systems biology experimental pipeline reveals matairesinol’s antimetastatic potential in prostate cancer: an integrated approach of network pharmacology, bioinformatics, and experimental validation. Briefings Bioinf. (2024) 25:bbae466. doi: 10.1093/bib/bbae466
Keywords: colorectal cancer, cuproptosis, ACAD8, immunotherapy, distant metastasis
Citation: Zhuang H, Chen Y and Huang S (2025) Cuproptosis-related gene ACAD8 inhibits the metastatic ability of colorectal cancer by inducing cuproptosis. Front. Immunol. 16:1560322. doi: 10.3389/fimmu.2025.1560322
Received: 14 January 2025; Accepted: 17 March 2025;
Published: 03 April 2025.
Edited by:
Lin-Lin Bu, Wuhan University, ChinaReviewed by:
Zhaowu Ma, Yangtze University, ChinaCopyright © 2025 Zhuang, Chen and Huang. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Sifu Huang, NTkzNTA5NkBxcS5jb20=
†These authors share first authorship
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.