
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Immunol. , 19 March 2025
Sec. Immunological Tolerance and Regulation
Volume 16 - 2025 | https://doi.org/10.3389/fimmu.2025.1545605
This article is part of the Research Topic Immune Checkpoints Regulatory Mechanisms and Immunotherapy Strategies in Gastrointestinal Tumors View all 4 articles
Background: Recent years, immunotherapy has emerged as a pivotal approach in cancer treatment. However, the response of gastric cancer to immunotherapy exhibits significant heterogeneity. Therefore, the early identification of gastric cancer patients who are likely to benefit from immunotherapy and the discovery of novel therapeutic targets are of critical importance.
Materials and methods: We collected data from European Nucleotide Archive (ENA) and Gene Expression Omnibus (GEO) databases. In project PRJEB25780, we performed WGCNA analysis and Lasso regression and chose CXCR2P1 for the subsequent analysis. Then, we compared the expression difference of CXCR2P1 among different groups. Kaplan-Meier curve was used to analyze the prognostic value of CXCR2P1, which was validated by project IMvigor210 and GEO datasets. ESTIMATE and CIBERSORT algorithm were used to evaluate the reshaping effect of CXCR2P1 to immune microenvironment of tumor. Differentially expressed genes (DEG) analysis, enrichGO analysis, Gene Set Enrichment Analysis (GSEA) and co-expression analysis were used to explore the cell biological function and signaling pathway involved in CXCR2P1.
Results: WGCNA identified CXCR2P1 as a hub gene significantly associated with immune response to PD-1 inhibitors in gastric cancer. CXCR2P1 expression was elevated in responders and correlated with better prognosis. Functional analysis revealed its role in reshaping the tumor immune microenvironment by promoting immune cell infiltration, including M1 macrophages, activated CD4+ T cells, and follicular helper T cells. CXCR2P1 enhanced antigen presentation via the MHC-II complex, influenced key immune pathways, such as Toll-like receptor signaling and T-cell activation, which led to the up-regulation of expression of PD-L1. GSEA showed CXCR2P1 were correlated with microRNAs. Through DEG analysis and expression analysis, MIR215 was identified as a potential direct target of CXCR2P1.
Conclusion: High expression of CXCR2P1 is correlated with better response to PD-1 inhibitor. It reshapes the immune microenvironment by increasing immune infiltration and changing the fraction of immune cells. In tumor immune microenvironment, CXCR2P1 can promote inflammation, enhance antigen presentation and activate the PD-1/PD-L1-related signaling pathway, which might be achieved by CXCR2P1-MIR215 axis.
Gastric cancer is the fifth most common cancer in the world and causes about 770 000 deaths every year. Of note, East Asia region has the highest gastric cancer incidence (22.4/100,000 inhabitants) and mortality (14.6/100,000) (1). In recent years, immunotherapy has become a significant adjuvant treatment for gastric cancer, with programmed cell death-1 (PD-1) inhibitors emerging as an effective option, particularly for advanced or metastatic cases (2). Studies have demonstrated that PD-1 inhibitors, including nivolumab and pembrolizumab, improve patient prognosis, leading to their incorporation into the National Comprehensive Cancer Network (NCCN) guidelines (3–5). However, not all gastric cancer patients can benefit from PD-1 inhibitors.The response rate of tumors to PD-1 inhibitors varies with the microsatellite stability, the degree of immune cell infiltration, the level of PD-L1 expression and so on (6). It is reported that unselected patients who receive anti-PD-1 and anti-PD-L1 immunotherapy have a total response rate of only approximately 20% (7). Therefore, it is important to identify patients who may benefit from PD-1 inhibitors before or early in the treatment of gastric cancer.
Finding out which genes are involved in the PD-1 signaling pathway, analyzing the heterogeneity of the expression of these genes are the key to explore the interaction mechanism between tumor and immune cells, so that we can develop more targeted therapies. By analyzing genes that were differentially expressed between gastric cancer patients who responded well to PD-1 inhibitors and those who did not respond to PD-1 inhibitors, we find C-X-C Motif Chemokine Receptor 2 Pseudogene 1 (CXCR2P1), a pseudogene encoding long non-coding RNA (lncRNA) which may affect cell phenotype through microRNA according to the previous research, plays a key role in the response of gastric cancer cells to PD-1 inhibitor (8). However, based on our investigation so far, only a few researchers have mentioned CXCR2P1 in their studies. The role of CXCR2P1 in tumor development and its effect on tumor immune microenvironment still remains unknown (8–12).
In this study, we compared expression levels of CXCR2P1 in different groups from different public databases, validated the potential of CXCR2P1 as a predictor of prognosis in gastric cancer patients receiving PD-1 inhibitors. Meanwhile, we analyzed the correlation between CXCR2P1 and tumor microenvironment. Finally, the signaling pathways involved in CXCR2P1 in gastric cancer cells were analyzed by differentially expressed genes analysis, enrichment analysis and co-expression analysis. This study explored the role of CXCR2P1 in the tumor immune response in detail, which provided a new target and idea for the immunotherapy of gastric cancer.
Aspera tools were used to download fastq sequencing data of project PRJEB25780 (gastric cancer immunotherapy data set) from the European Nucleotide Archive (ENA) database (https://www.ebi.ac.uk/ena). Subreads, HISAT2 (version 2.2.1), and samtools were used to generate the counts matrix and Transcripts Per Kilobase of Exon Model Per Million Mapped Reads (tpms) from fastq sequencing data for subsequent analysis. Quantile normalized (log10) Fragments Per Kilobase of Transcript Per Million Mapped Reads (fpkm) of project PRJEB25780 were downloaded from Tumor Immune Dysfunction and Exclusion (TIDE) database. Clinical information of project PRJEB25780 were downloaded from materials of Kim’s research (13). The four gastric cancer microarray datasets (GSE193453, GSE174237, GSE84433, GSE27342) and the lung cancer immunotherapy data set (GSE135222) can be downloaded from the GEO database (https://www.ncbi.nlm.nih.gov/geo/). The expression matrix and clinical information of metastatic urothelial carcinoma immunotherapy data set (IMvigor210) were obtained through “IMvigor210CoreBiologies” R package. Gene sets of PD-1 checkpoint signaling pathway were downloaded from Kyoto Encyclopedia of Genes and Genomes (KEGG) database (https://www.kegg.jp/kegg/). All tpms matrix used for analysis were normalized by log2.
On project PRJEB25780, Weighted Correlation Network Analysis (WGCNA) was performed by “WGCNA” R package for seeking gene modules with highly synergistic changes in expression. The response to PD-1 inhibitor of gastric cancer patients were regarded as the clinical phenotypes for subsequent WGCNA. We first used the hclust function to cluster a total of 45 samples, excluded the outlier sample, and used the remaining samples for network topology analysis to determine the appropriate soft threshold. Then, gene network modules were constructed by the “blockwiseModules” function with the minimum number of 30 genes in each module. Each module was assigned a color, and genes that could not be clustered into other modules were uniformly assigned gray. The eigengenes of each module were calculated by the “moduleEigengenes” function. Finally, based on the eigengenes of each module, the correlation between the modules and the response to PD-1 was calculated. A heat map illustrating the module-phenotype correlations was generated, and the module showing the strongest correlation was chosen for further analysis.
Gene significance (GS) is defined as the absolute value of the correlation between a gene and the phenotypic trait. Module membership (MM) is defined as the correlation between the eigengene of a module and the gene expression within the module. In this study, we used the condition of GS > 0.2 and MM > 0.8 as the screening criteria for hub gene of the module we chose above.
Normalized expression matrix of hub genes screened by WGCNA was performed Least Absolute Shrinkage and Selection Operator regression (Lasso) by the “glmnet” R package. Next, cross-validation was used to select the optimal regularization parameter λ to maximize the prediction performance of the model. After obtaining the best model, the features with nonzero coefficients were retained for subsequent analysis.
Expression difference of CXCR2P1 between responders and non-responders subgroups were described in project PRJEB25780 and IMvigor210. Responder was defined as complete response (CR) or partial response (PR) and non-responder was defined as stable disease (SD) or progressive disease (PD) on clinical evaluation after treatment with PD-1 inhibitors. CXCR2P1 expression between gastric cancer tissue and normal tissue were also analyzed in four microarray datasets (GSE193453, GSE174237, GSE84433, GSE27342).
The prognostic value of CXCR2P1 was validated by overall survival (OS) from project IMvigor210 and progression free survival (PFS) from project GSE135222. OS was defined as the time from surgery to death due to any cause, PFS was defined as the time from surgery to recurrence, metastasis, or death due to disease.
Immune score was calculated based on the proportion of immune cells by ESTIMATE algorithm. The Immune score of CXCR2P1-High group and CXCR2P1-Low group were compared in patients of project PRJEB25780. Meanwhile, the fraction of immune cells of each tumor sample in project PRJEB25780 was estimated by CIBERSORT algorithm. All analysis in this section were performed using normalized tpms matrix.
Counts matrix of PRJEB25780 was used for differentially expressed genes (DEGs) analysis by the “DESeq2” R package with the filter criteria of |log2Foldchange (FC) | > 1 and false discovery rate (FDR) < 0.05. In the DEGs analysis for CXCR2P1, patients were divided into high-expression and low-expression groups based on the median expression levels. In the DEGs analysis for Response, patients were divided into Responders and Non-Responders based on their response to PD-1 inhibitor.
To explore the potential signaling pathway involved in CXCR2P1, we conducted Gene Set Enrichment Analysis (GSEA) and EnrichGO Analysis for DEGs of CXCR2P1. EnrichGO analysis was performed by R package “clusterProfiler”. Reference gene sets of GSEA included hallmark, c2wikipathways, c2KEGG and c5go. The screening conditions were |normalized enrichment score (NES)| > 1 and p value < 0.05.
DEGs of CXCR2P1 were conducted correlation analysis with the expression of CXCR2P1. DEGs with pearson correlation coefficient > 0.3 and p value < 0.05 were selected to construct the co-expression network with CXCR2P1 by the “igraph” R package. The network was then modified in Cytoscape software (version 3.10.2).
In this study, all statistical analyses were performed using R software (version 4.3.2). The comparison of the expression levels of DEG-microRNAS between different groups was performed by Mann-Whitney test, except that all comparisons between the two groups were performed by two-tailed Student t test. Survival analysis was conducted by Kaplan-Meier curve. Pearson’s analysis was used for correlation analysis between modules and response to PD-1 inhibitor or between expression of CXCR2P1 and DEGs. Spearman’s analysis was used for correlation analysis which generated scatter plots. P value of 0.05 was considered statistically significant (* means p < 0.05, ** means p < 0.01, *** means p < 0.001).
Figure 1 shows the flow chart of this study. With row data processing, we obtained the transcriptome expression matrix and clinical information of project PRJEB25780. In WGCNA analysis, we first clustered samples based on their normalized expression matrix and response to PD-1 inhibitor and exclude the sample that was apparently outlier (Figure 2A, sample code: PB-16-066). Then, a network consisting of highly cooperative gene modules was constructed by network topology analysis (Figure 2B). In the end, we got 61 effective modules in total with different colors. Figure 2C shows the correlation among the modules. Next, we conducted correlation analysis between modules and the clinical phenotype (Responder and Non-Responder) and found module lavenderblush3 had the highest correlation with the immune response of the tumor (rpearson = 0.61, p = 0.000009, shown as Figure 2D).

Figure 2. The process of screening for hub genes. (A) Clustering dendrogram of 44 samples with exclusion of PB-16-066. (B) Dendrogram of all differentially expressed genes clustered based on the measurement of dissimilarity. (C) Correlations and Clustering among modules. (D) Correlation between response to PD-1 inhibitor and modules. (E) Screening for hub genes within lavenderblush3 module. (F) Lasso coefficient path plots for the 18 hub genes. (G) Cross validation curves for lasso regression.
After loading the lavenderblush3 module, we obtained a total of 53 intra-module genes. The scatter plot shown in Figure 2E was used to represent the relationship between GS and MM of each gene in the lavenderblush3 module, whose fitting curve showed a significant correlation between GS and MM (rpearson = 0.63, p = 0.00000043). Genes with GS > 0.2 and MM > 0.8 were selected as hub genes. Finally, we obtained 18 hub genes of lavenderblush3 module (GBP4, GBP5, JAKMIP1, APOBEC3H, GBP1, GBP2, CD274, IFNG, CXCR2P1, IDO1, CXCL10, GBP1P1, CXCL9, LAP3, CXCL11, TAP1, WARS, UBE2L6).
To reduce the collinearity among hub genes, the lasso regression analysis was performed. As the penalty function (log λ) increases, the regression coefficient of each variable in the model gradually converge to 0 (Figure 2F). In the cross-validation, when the number of variables in the model was reduced to 7 (GBP4, JAKMIP1, IFNG, CXCR2P1, LAP3, CD274, APOBEC3H), the mean square error of the model was reduced to the lowest point, and the fitting effect of the equation reached the best (Figure 2G). Since the mechanism of how CXCR2P1 affects cell phenotype is still unclear, it is selected for subsequent analysis.
The prognosis of patients with gastric cancer is highly correlated with their response to the PD-1 inhibitor. Trough analyzing different data sets, expression of CXCR2P1 was found higher in responders than which in non-responders in both project PRJEB25780 (Figure 3A, p < 0.001) and project IMvigor210 (Figure 3B, p < 0.01). In four GEO datasets, expression of CXCR2P1 was higher in gastric cancer tissue than which in normal tissue (Figure 3C, p < 0.001). Through Kaplan-Meier curves, we found CXCR2P1-high group had prolonged OS in patients with metastatic urothelial carcinoma (Figure 3F, p < 0.0001) and prolonged PFS in patients with lung cancer (Figure 3E, p = 0.013) than those in CXCR2P1-low group. Based on results above, patients with higher expression of CXCR2P1 might possess better prognosis.
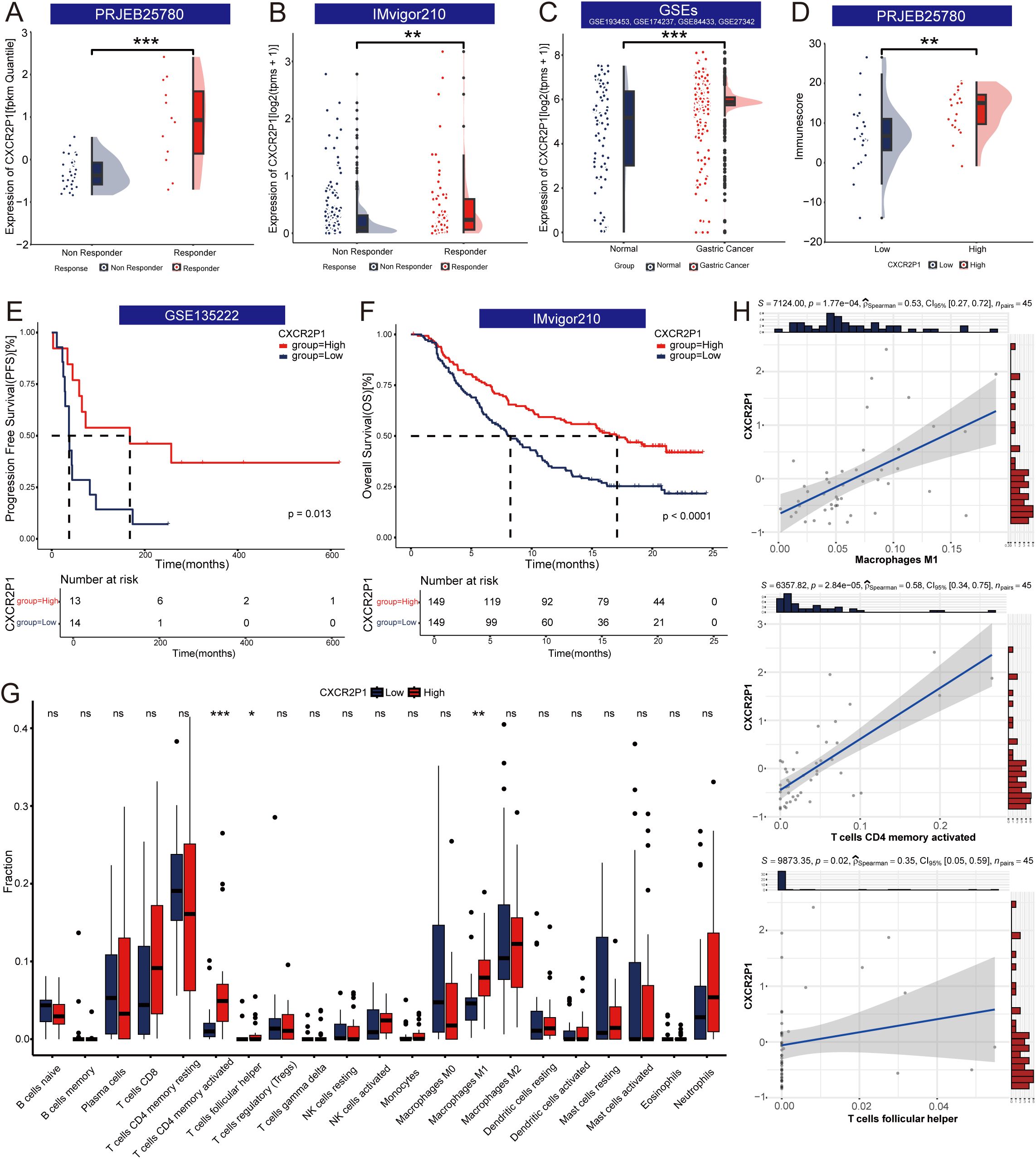
Figure 3. Exploration of the function of CXCR2P1. (A) Comparison of expression of CXCR2P1 between responders and non-responders in project PRJEB25780. (B) Comparison of expression of CXCR2P1 between responders and non-responders in project IMvigor210. (C) Comparison of expression of CXCR2P1 between gastric cancer tissue and normal gastric mucous tissue in GEO datasets. Four GEO datasets are GSE193453, GSE174237, GSE84433 and GSE27342. (D) Comparison of immunescore between CXCR2P1-High group and CXCR2P1-Low group. (E) - (F) Kaplan-Meier Curve of survival analysis of CXCR2P1. High Expression of CXCR2P1 had prolonged PFS in GSE135222 and prolonged OS in project IMvigor210. (G) Comparison of the fraction of immune cells between CXCR2P1-High group and CXCR2P1-Low group. (H) Correlation between the CXCR2P1 and Macrophages (M1, ρSpearman = 0.53, p = 0.000177), activated CD4+ memory T cells (ρSpearman = 0.58, p = 0.0000284), and follicular helper T cells (ρSpearman = 0.35, p = 0.02). PFS, progression free survival; OS, overall survival; *p < 0.05; **p < 0.01; ***p < 0.001.
In project PRJEB25780, ESTIMATE analysis showed CXCR2P1-high group had higher immune score, which revealed that high expression of CXCR2P1 increased the degree of infiltration of immune cells to tumors (Figure 3D). CIBERSORT algorithm was used to evaluate the abundance of immune cells and compared the fractions of different immune cells between CXCR2P1-high and CXCR2P1-low group (Figure 3G). Result show that the fraction of Macrophages (M1), activated CD4+ memory T cells, and follicular helper T cells was higher in CXCR2P1-high group than which in CXCR2P1-low group. Meanwhile, the linear equation of scatter plots shown as Figure 3H revealed a significant positive correlation between CXCR2P1 expression and the fraction of Macrophages (M1, ρSpearman = 0.53, p = 0.000177), activated CD4+ memory T cells (ρSpearman = 0.58, p = 0.0000284), and follicular helper T cells (ρSpearman = 0.35, p = 0.02). The expression of CXCR2P1 can influence the tumor immune microenvironment significantly.
Through differential expression analysis, 610 DEGs were found between the high- and low-expression groups of CXCR2P1 (139 up-regulated, 471 down-regulated, as shown in Figures 4A, B). To explore the cellular biological function of CXCR2P1, EnrichGO and Gene Set Enrichment Analysis (GSEA) was performed by these DEGs. Top 5 enriched biological function by EnrichGO analysis are shown as Figure 4E, and 3 of them are related with biological immune response (acute-phase response, acute inflammatory response, humoral immune response). GSEA by c2KEGG database showed that CXCR2P1 were enriched on Antigen Processing and Presentation By MHC Class II Molecules pathway (p = 0.00000103, shown as Figure 4C). In c5go database, biological processes (BP), cellular components (CC) and molecular functions (MF) of CXCR2P1 were analyzed and the top 10 enriched BP, CC, and MF are listed in Figure 4D. Results show that the BP, CC and MF of CXCR2P1 are all enriched in peptide antigen binding and the MHC-II protein complex, which indicate that CXCR2P1 might increase the infiltration of immune cells in tumor issue by enhancing the antigen presentation in tumor immune microenvironment by MHC II protein complex.
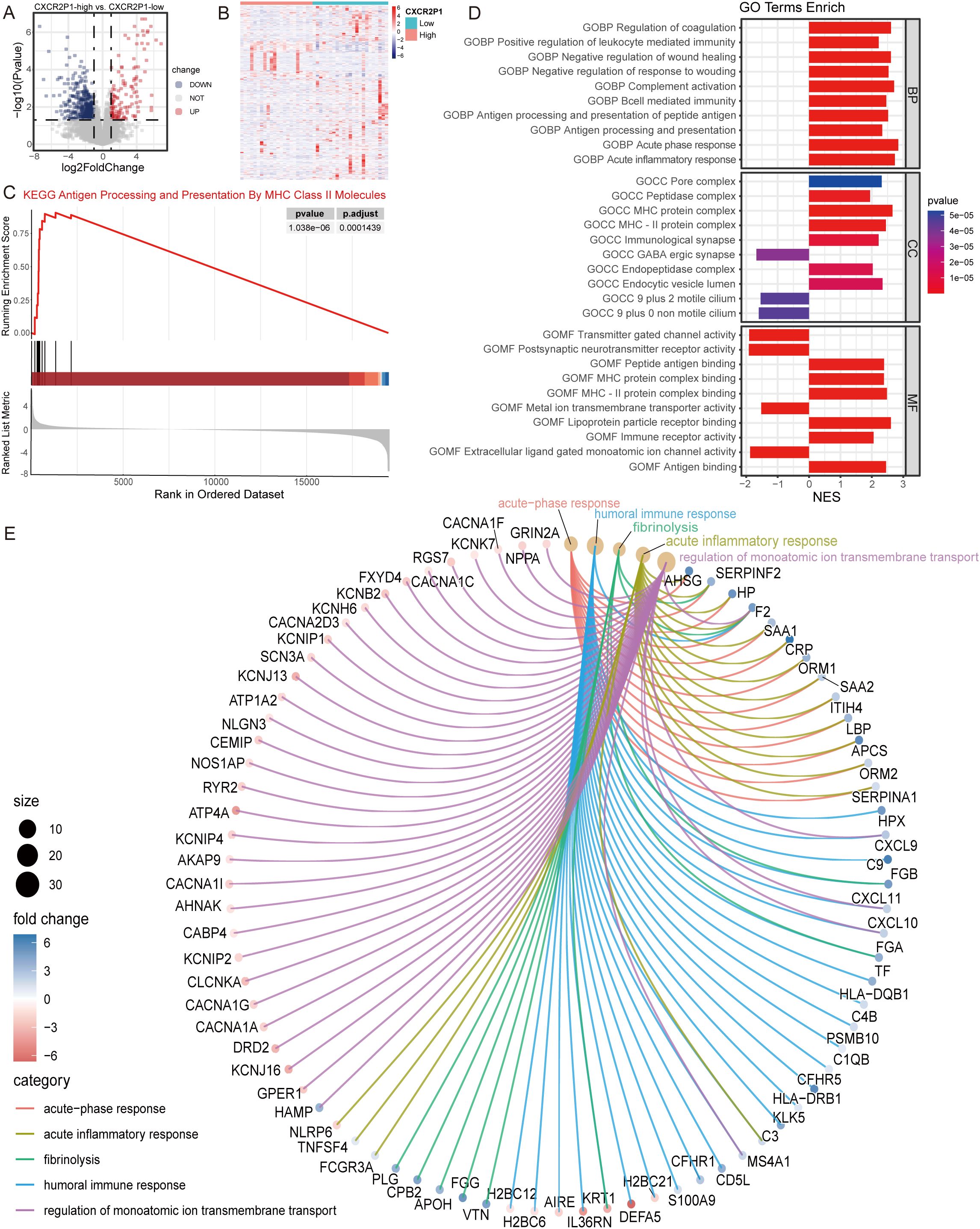
Figure 4. Analysis of cell biological function of CXCR2P1. (A) Volcano plot of DEG analysis by expression of CXCRP21. (B) Heatmap of DEG analysis by expression of CXCRP21. (C) GSEA analysis for DEGs of CXCR2P1 by c2KEGG database. (D) GSEA analysis for DEGs of CXCR2P1 by c5go database. (E) Chord diagram of top 5 enriched biological function by EnrichGO analysis for DEGs of CXCR2P1. BP, biological processes; CC, cellular components; MF, molecular functions; NES, normalized enrichment score.
To investigate the relationship between CXCR2P1 and tumor immune evasion by PD-1/PD-L1 signaling pathway, we first analyzed the correlation between the expression of CXCR2P1 and CD274. Results showed that CXCR2P1 was highly correlated with the expression of CD274 (ρSpearman = 0.60, p = 1.28 x 10-5, shown as Figure 5B) in tumor tissue. GSEA by c2wikipathway showed that CXCR2P1 was significantly enriched in Cancer Immunotherapy By PD-1 Blockade (p = 2.676 x 10-4, shown as Figure 5C). Subsequently, we obtained all 73 genes involved in PD-1/PD-L1 signaling pathway from KEGG database, and analyzed the correlation between the expression of these genes and CXCR2P1. Figure 5A shows all 20 genes which are significantly related with the expression of CXCR2P1 (p < 0.05, rpearson >0.3).
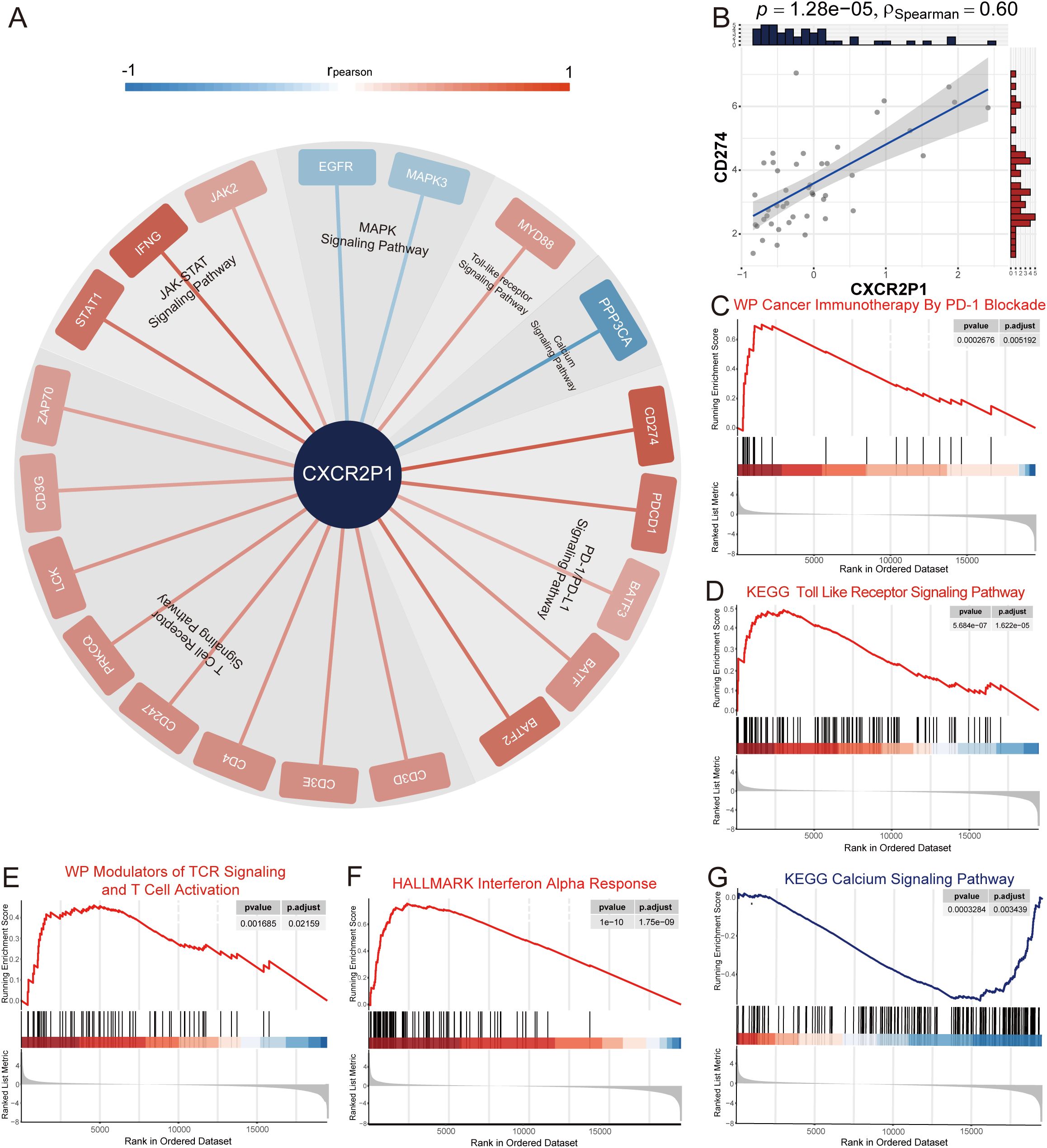
Figure 5. Analysis of signaling pathway of PD-L1. (A) Co-expression analysis between CXCR2P1 and genes involved in PD-1/PD-L1 signaling pathway. 20 genes which were significantly associated with CXCR2P1 (rpearson > 0.3, p < 0.05) were listed around the edge of pie chart and were classified according to the signaling pathway to which they belong. (B) Correlation between the expression of CXCR2P1 and CD274. (C-G) GSEA analysis for DEGs of CXCR2P1 by c2KEGG database, Hallmark database and c2wikipathway database. Results showed that CXCR2P1 can activate PD-1/PD-L1-related signaling pathways and suppress the activation of T cells.
Through GSEA analysis, we found 4 PD-1/PD-L1-related signaling pathways were enriched. Results showed that in tumor tissue, with the increasing expression of CXCR2P1, Toll Like Receptor Signaling Pathway(p = 5.684 x 10-7, shown as Figure 5D), Modulators of TCR Signaling And T Cell Activation(p = 1.685 x 10-3, shown as Figure 5E) and Interferon Alpha Signaling Pathway (p = 1.0 x 10-10, shown as Figure 5F) were significantly activated. Instead, Calcium Signaling Pathway (p = 3.284 x 10-4, shown as Figure 5G) was significantly suppressed. Our results indicated that CXCR2P1 may indirectly up-regulate the expression of PD-L1 in tumor cells by increasing immune infiltration and enhancing the activation of T cells in the tumor tissue.
GSEA by c2wikipathway database showed that CXCR2P1 was significantly enriched to Interactions Between Immune Cells And MicroRNAs In Tumor Microenvironment (2.402 x 10-3, shown as Figure 6A), which indicated that CXCR2P1 might regulate the expression of PD-L1 through microRNAs. To further clarify which microRNAs were involved in the upstream and downstream signaling pathways of CXCR2P1, we first performed DEG analysis for all microRNAs in project PRJEB25780 by the expression of CXCR2P1, and the results showed that the expression of 23 genes was down-regulated in CXCR2P1-High group (Figures 6B, C). Then another DEG analysis was performed for all microRNAs in PRJEB25780 by the response to PD-1 inhibitor, and the results showed that 2 genes were up-regulated and 12 genes were down-regulated in Responder group (Figures 6D, E). By taking the intersection of the two microRNA gene sets, we obtained a total of 5 genes, including MIR200A, MIR145, MIR215, MIR27B and MIR5707 (Figure 6F). The results showed that MIR215 expression was significantly down-regulated in both CXCR2P1-High group (p = 0.001604, Figure 6J) and Responder group (p = 0.01474, Figure 6O). MIR27B was significantly down-regulated in CXCR2P1-High group (p = 0.0253, Figure 6G), but there was no significant difference between Responder and Non-Responder groups (p = 0.09778, Figure 6L). There was no significant difference in the expression of MIR200A (Figures 6I, N), MIR145 (Figures 6H, M) and MIR5707 (Figures 6K, P) between both CXCR2P1 groups and Response groups. Our finding suggests that MIR215 might be involved in the signaling pathway of CXCR2P1 and serve as the mediator of CXCR2P1 affecting cell phenotype.

Figure 6. Correlation between CXCR2P1 and microRNA. (A) GSEA analysis for DEGs of CXCR2P1 by c2wikipathway database. (B) Volcano plot of DEG analysis for microRNAs from project PRJEB25780 by expression of CXCRP21. (C) Heatmap of DEG analysis for microRNAs from project PRJEB25780 by expression of CXCRP21. (D) Volcano plot of DEG analysis for microRNAs from project PRJEB25780 by response to PD-1 inhibitor. (E) Heatmap of DEG analysis for microRNAs from project PRJEB25780 by response to PD-1 inhibitor. (F) Venn diagram of the DEGs-microRNA by the expression of CXCR2P1 and the response to PD-1 inhibitor. The intersection contains 5 DEGs-microRNA, including MIR200A, MIR145, MIR215, MIR27B and MIR5707. (G-K) Raincloud plots showing the comparison of the expression of the 5 DEGs-microRNA between CXCR2P1-high group and CXCR2P1-low group. (L-P) Raincloud plots showing the comparison of the expression of the 5 DEGs-microRNA between Responder group and Non-Responder group. NA, not available; NS, not significant; *p < 0.05; **p < 0.01; ***p < 0.001.
In this study, we screened out a set of genes that significantly affect the response of gastric cancer to PD-1 inhibitor by WGCNA and lasso regression for the first time. Through comprehensive analysis, we explored the potential of CXCR2P1 as a prognostic predictor and its role in tumor immune response. Furthermore, we validated our findings using databases on PD-1 inhibitor response in other cancer types. This research analyzed the cell biological role of CXCR2P1 firstly, explored the effect of CXCR2P1 on the tumor immune microenvironment, and provided new insights and references for the development of targeted therapies.
The expression of CXCR2P1 might affect the stage and prognosis of tumors. CXCR2P1 was first mentioned in 2018 in the study of Lou et al., who found that CXCR2P1 encodes lncRNA through RNA transcriptomics analysis and its expression was significantly up-regulated in high-grade serous ovarian cancer tissues compared with normal ovarian tissues (12). Similar results were presented in research of Ji et al. which found CXCR2P1 was significantly up-regulated in cutaneous metastatic melanoma and high expression of CXCR2P1 was associated with prolonged overall survival (10). Sarathi et al. confirmed that CXCR2P1 is one of the key genes in determining the stage of hepatocellular carcinoma through differential analysis, which indicated low expression of CXCR2P1 will lead hepatocellular carcinoma to be more invasive and metastatic (11). Our study showed that the expression of CXCR2P1 was higher in responders and tumor tissue than those in non-responders and normal tissue. Meanwhile, in the only two databases containing both results of response to PD-1 inhibitor and survival information (lung cancer and metastatic urothelial carcinoma), CXCR2P1 showed excellent prognostic value. Patients with high expression of CXCR2P1 had prolonged OS and PFS. These findings validated that the high expression of CXCR2P1 was significantly correlated with good prognosis of tumors and the prognostic value of CXCR2P1 may be related to its effect on tumor immune response.
CXCR2P1 may help reshape the tumor immune microenvironment. As early as 2019, Choy found CXCR2P1 may have the ability to affect the response of tumor to immunotherapy through a pan-cancer machine-learning analysis (9). Our study validated that the tumors with higher expression of CXCR2P1 had higher degree of immune infiltration and immune cell abundance, and the high expression of CXCR2P1 were significantly correlated with good prognosis. The immune microenvironment and the degree of infiltration of immune cells are the key factors affecting the tumor immune response. It has been shown that tumors with pre-existing lymphocytes-infiltrating microenvironment are more likely to respond to immune-checkpoint inhibition (14). Immunotherapies are generally more effective in immunologically “hot” tumors, where abundant immune cells are already primed against tumor cells. However, “cold” tumors, which lack sufficient immune infiltration, are often less responsive to immunotherapy (15–17). Hence, tumors with high expression of CXCR2P1 may have more active tumor immune microenvironment, which laid a foundation for good immune response.
Previous studies have shown that M1 macrophages and CD4+T cells can recruit more immune cells by secreting pro-inflammatory cytokines and chemokines, presenting antigens, and inducing the development and maturation of other immune cells in the tumor immune microenvironment, thereby playing an important role in tumor’s immune surveillance (18). Our research found that the expression of CXCR2P1 was positively correlated with the number of M1 macrophages and activated CD4+ T cells in the tumor microenvironment. The reshaping of the immune microenvironment and the influence of CXCR2P1 on the response to immunotherapy may be achieved through these two types of cells.
Tumor-associated macrophages (TAMs) are crucial in shaping the tumor immune microenvironment. Based on function and phenotype, macrophages can be divided into M1 and M2 types. M1 polarization is associated with tissue inflammation, mediating tissue damage and recruiting CD8+ T cells to target tumor cells. In contrast, M2 polarization is linked to anti-inflammatory responses, which facilitate tissue repair. The M1/M2 ratio in macrophages significantly influences the tumor immune microenvironment and tumor response to immunotherapy (19, 20). TAM molecular phenotypes are shaped by chemokines, cytokines, cell metabolites, and cross-talk signaling pathways in the tumor microenvironment (21). Activation of toll-like receptor (22), interferon (23), and CD40 signaling pathways (24) promotes M1 macrophage differentiation, enhancing antigen presentation, directly phagocytosing tumor cells, and recruiting CD8+ T cells and NK cells (21). Conversely, TGF-β (25), γ-aminobutyric acid (26), and activation of myeloid checkpoint pathways (27, 28) shift macrophages toward an M2 phenotype, leading to IL-10 secretion and suppression of cytotoxic CD8+ T cells. Recent studies indicate that TAMs can also express PD-1, broadening the concept that PD-1/PD-L1 is exclusive to T cells’ immune escape mechanisms (29). This insight provides a potential explanation for CXCR2P1’s impact on PD-1 inhibitor responses via TAMs. Our study revealed a significant positive correlation between CXCR2P1 expression and acute inflammatory responses, with notable activation of MHC II antigen presentation complexes and reduced gabaergic neurotransmitter secretion in tumors with high CXCR2P1 expression. These findings suggest that CXCR2P1 fosters M1 macrophage polarization within the tumor immune microenvironment, enhancing the tumor’s responsiveness to immunotherapy.
Through cytokine secretion and direct interactions with immune and tumor cells, CD4+ T cells play a pivotal role in the tumor immune response by influencing the composition and activity of the tumor immune microenvironment (30). In the “afferent” phase of the immune response, naive CD4+ T cells are stimulated by tumor-associated antigens presented by antigen-presenting cells. This stimulation triggers their differentiation into subsets such as TH1, TH2, TH17, TH9, follicular helper T cells (TFH), and regulatory T cells (Treg) (31, 32). Differentiated CD4+ T cells then migrate to tumor tissues or adjacent lymphoid tissues to participate in tumor immune responses (33). In the tumor immune microenvironment, CD4+ T cells can stimulate and maintain the differentiation of macrophages, CD8+ T cells and NK cells by secreting cytokines such as IFN-γ, TNF and IL-2, and inhibit angiogenesis of tumor, thus playing an anti-tumor effect (34–36). At the same time, CD4+ T cells can activate antigen-presenting cells through CD40-CD40L pathway to enhance the antigen presentation of tumor-associated antigens (37, 38). Recent studies have identified cytotoxic CD4+ T cells (activated TH1 cells) that directly kill tumor cells by releasing granzyme B and perforin (39). However, not all CD4+ T cells contribute to anti-tumor immunity. Subsets such as TFH cells and Treg cells can inhibit immune responses by expressing PD-L1 and secreting TGF-β, thus fostering tumor progression (40, 41).
The balance and composition of these CD4+ T cell subsets critically influence tumor immunogenicity. In our study, tumor tissues with high CXCR2P1 expression exhibited significantly higher proportions of activated CD4+ T cells and TFH cells compared to low CXCR2P1 expression tumor, suggesting that the tumor tissues with high CXCR2P1 expression may have higher immunogenicity. In addition, an increase in the proportion of follicular helper T cells indicates increased expression of PD-L1 in the tumor, which underlies the favorable response to PD-1 inhibitors (42). It has been shown that the degree of CD4+T cell infiltration in a tumor is directly related to a favorable patient outcome regardless of immunotherapy. However, how PD-1 inhibitor affects the phenotype of CD4+T cells in the immune microenvironment and how CD4+T cells enhance the response of tumor tissues to immune responses are still unclear, and further studies are needed to clarify the cellular biological mechanisms.
At the molecular level, CXCR2P1 can activate immune-related signaling pathways and significantly increase the expression of PD-L1 in tumor tissues. Previous studies have demonstrated that increased levels of cytokines like IFN-γ (43) in the tumor microenvironment, along with the activation of the JAK-STAT (44), PI3K-Akt (45), and Toll-like receptor signaling pathways (46) can upregulate PD-L1 expression in both tumor and some immune cells, which enables tumor cells to evade immune surveillance (47). Activation of the PD-1 receptor further dampens the immune response by inhibiting key pathways in T cell receptor (TCR) signaling, such as the calcium signaling pathway (48) and MAPK signaling pathways (49), which leads to the reduction of IL-2 secretion and suppression of T cell activation.
In this study, we observed a significant positive correlation between the expression of CXCR2P1 and PD-L1 in tumor tissues. GSEA and co-expression analysis revealed that tumor tissues with high CXCR2P1 expression exhibited significant activation of the PD-1 signaling pathway, IFN signaling pathway, Toll-like receptor signaling pathway, and JAK-STAT signaling pathway. Conversely, the calcium signaling and MAPK pathways, integral to TCR signaling, were significantly suppressed. These findings suggest that CXCR2P1 plays a dual role in modulating the tumor immune microenvironment. By enhancing antigen presentation, CXCR2P1 may indirectly upregulates PD-L1 expression in tumor tissues by promoting immune cell infiltration. Elevated PD-L1 expression, coupled with enriched immune cell infiltration, enhances the tumor’s responsiveness to PD-1 inhibitors, contributing to favorable immunotherapy outcomes.
From GSEA analysis, we found CXCR2P1 was enriched into microRNAs in tumor microenvironments, which indicated that CXCR2P1 might affect the cell phenotype through microRNAs. Similar results was obtained by Zhao et al., who comprehensively analyzed the RNA transcriptome of head and neck squamous cell carcinoma and constructed a competing endogenous RNA network that significantly affected tumor immune infiltration through differential expressed genes analysis, finding that CXCR2P1 may affect the OS of patients through the lncRNA-microRNA-mRNA axis (8). Our research firstly identified MIR215 as the mediator of CXCR2P1 affecting cell phenotype.
MIR215 is an intragenic microRNA, whose host gene, isoleucyl-tRNA synthetase 2 (IARS2), is a form of alanyl-tRNA synthetase (50). Previous researches had proved that MIR215 can mediate the post-transcriptional gene silencing through receptor-interacting serine/threonine-protein kinase 1 (51), and hsa-miR-215-5p can be regulated by many lncRNAs, such as CDC6 (52), FTX (53) and UICLM (54). MIR215 is critically involved in regulating cell proliferation, apoptosis, and migration. Studies have shown that MIR215 can promote the proliferation, migration, and invasion of gastric cancer cells (55). Its expression is significantly upregulated in gastric cancer tissues compared to normal gastric mucosa, with even higher levels observed in patients with poorly differentiated tumors, advanced clinical stages (stage III or IV), or lymph node metastases (56, 57). Runt-related Transcription Factor 1 (RUNX1) has been identified as a direct target of MIR215 (58). In gastric cancer, RUNX1 inhibits malignant cell proliferation by regulating the expression of key genes such as p21 and p53, thereby suppressing cell cycle progression and promoting cellular differentiation (59). The post-transcriptional silencing of RUNX1 by MIR215 might contribute to the enhanced invasive and proliferative capacities of tumor cells. Comprehensively, our findings suggest that high CXCR2P1 expression may reduce hsa-miR-215-5p levels in tumor tissues through the lncRNA-microRNA regulatory axis, potentially improving the prognosis of gastric cancer patients. However, the precise role of MIR215 in the immune microenvironment of tumor remains unclear, further studies are required to elucidate its underlying cellular and molecular mechanisms.
Our study has several limitations. First, as the only database focusing on the response of gastric cancer to PD-1 inhibitor, project PRJEB25780 is limited by its small sample scale, which may reduce the statistical robustness of our findings. Second, all conclusions drawn in this study are based on bioinformatic analyses, lacking validation through experimental approaches. Finally, due to the inherent limitations of bioinformatics analysis, the regulatory factors involved in the upstream and downstream signaling pathways of CXCR2P1 remain unclear. Future studies should prioritize functional validation of CXCR2P1 through in vitro or in vivo models, systematically map its upstream regulators and downstream signaling cascades, and assess its clinical relevance in larger immunotherapy cohorts with matched genomic and transcriptomic data.
In gastric cancer tissues, the expression of CXCR2P1 is significantly elevated compared with normal gastric mucosa, and patients with high CXCR2P1 expression exhibit better prognosis and improved responses to PD-1 inhibitor. CXCR2P1 plays a vital role in reshaping the tumor immune microenvironment by promoting greater immune cell infiltration and significantly increasing the proportions of M1 macrophages, CD4+ T cells, and follicular helper T cells. At the cellular level, CXCR2P1 is associated with inflammatory activation and enhanced antigen presentation in tumor tissues. At the molecular level, CXCR2P1 upregulates PD-L1 expression in tumor tissues and activates key signaling pathways involved in PD-1 regulation. These effects might be mediated through the lncRNA-microRNA axis, with MIR215 identified as a potential direct target of CXCR2P1. Future studies are required to experimentally validate the upstream and downstream signaling pathways of CXCR2P1 and to further elucidate its cell biological mechanisms.
The original contributions presented in the study are included in the article/supplementary material. Further inquiries can be directed to the corresponding authors.
The studies involving humans were approved by The Ethics Committee of Peking University People’s Hospital. The studies were conducted in accordance with the local legislation and institutional requirements. Written informed consent for participation was not required from the participants or the participants’ legal guardians/next of kin in accordance with the national legislation and institutional requirements.
XW: Data curation, Writing – original draft. SH: Formal Analysis, Methodology, Writing – original draft. YY: Project administration, Resources, Writing – review & editing. ZG: Writing – review & editing.
The author(s) declare that financial support was received for the research and/or publication of this article. Horizontal Research Project of Beijing Science and Technology Planning Project: 2159000118. Horizontal Research Project of Beijing Science and Technology Planning Project: 2159000130.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declare that no Generative AI was used in the creation of this manuscript.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
1. López MJ, Carbajal J, Alfaro AL, Saravia LG, Zanabria D, Araujo JM, et al. Characteristics of gastric cancer around the world. Crit Rev Oncol Hematol. (2023) 181:103841. doi: 10.1016/j.critrevonc.2022.103841
2. Högner A, Moehler M. Immunotherapy in gastric cancer. Curr Oncol. (2022) 29:1559–74. doi: 10.3390/curroncol29030131
3. Kang YK, Boku N, Satoh T, Ryu MH, Chao Y, Kato K, et al. Nivolumab in patients with advanced gastric or gastro-oesophageal junction cancer refractory to, or intolerant of, at least two previous chemotherapy regimens (ONO-4538-12, ATTRACTION-2): a randomised, double-blind, placebo-controlled, phase 3 trial. Lancet. (2017) 390:2461–71. doi: 10.1016/S0140-6736(17)31827-5
4. Fuchs CS, Doi T, Jang RW, Muro K, Satoh T, Machado M, et al. Safety and efficacy of pembrolizumab monotherapy in patients with previously treated advanced gastric and gastroesophageal junction cancer: phase 2 clinical KEYNOTE-059 trial. JAMA Oncol. (2018) 4:e180013. doi: 10.1001/jamaoncol.2018.0013
5. Ajani JA, D’Amico TA, Bentrem DJ, Chao J, Cooke D, Corvera C, et al. Gastric cancer, version 2.2022, NCCN clinical practice guidelines in oncology. J Natl Compr Canc Netw. (2022) 20:167–92. doi: 10.6004/jnccn.2022.0008
6. Puccini A, Battaglin F, Iaia ML, Lenz HJ, Salem ME. Overcoming resistance to anti-PD1 and anti-PD-L1 treatment in gastrointestinal Malignancies. J Immunother Cancer. (2020) 8:e000404. doi: 10.1136/jitc-2019-000404
7. Long J, Lin J, Wang A, Wu L, Zheng Y, Yang X, et al. PD-1/PD-L blockade in gastrointestinal cancers: lessons learned and the road toward precision immunotherapy. J Hematol Oncol. (2017) 10:146. doi: 10.1186/s13045-017-0511-2
8. Zhao S, Peng M, Wang Z, Cao J, Zhang X, Yu R, et al. Immune infiltration-related ceRNA network revealing potential biomarkers for prognosis of head and neck squamous cell carcinoma. Dis Markers. (2022) 2022:1014347. doi: 10.1155/2022/1014347
9. Choy CT, Wong CH, Chan SL. Embedding of genes using cancer gene expression data: biological relevance and potential application on biomarker discovery. Front Genet. (2019) 9:682. doi: 10.3389/fgene.2018.00682
10. Ji C, Li Y, Yang K, Gao Y, Sha Y, Xiao D, et al. Identification of four genes associated with cutaneous metastatic melanoma. Open Med (Wars). (2020) 15:531–9. doi: 10.1515/med-2020-0190
11. Sarathi A, Palaniappan A. Novel significant stage-specific differentially expressed genes in hepatocellular carcinoma. BMC Cancer. (2019) 19:663. doi: 10.1186/s12885-019-5838-3
12. Lou Y, Jiang H, Cui Z, Wang X, Wang L, Han Y. Gene microarray analysis of lncRNA and mRNA expression profiles in patients with high-grade ovarian serous cancer. Int J Mol Med. (2018) 42:91–104. doi: 10.3892/ijmm.2018.3588
13. Kim ST, Cristescu R, Bass AJ, Kim KM, Odegaard JI, Kim K, et al. Comprehensive molecular characterization of clinical responses to PD-1 inhibition in metastatic gastric cancer. Nat Med. (2018) 24:1449–58. doi: 10.1038/s41591-018-0101-z
14. Tumeh PC, Harview CL, Yearley JH, Shintaku IP, Taylor EJ, Robert L, et al. PD-1 blockade induces responses by inhibiting adaptive immune resistance. Nature. (2014) 515:568–71. doi: 10.1038/nature13954
15. Huang P, Zhou X, Zheng M, Yu Y, Jin G, Zhang S. Regulatory T cells are associated with the tumor immune microenvironment and immunotherapy response in triple-negative breast cancer. Front Immunol. (2023) 14:1263537. doi: 10.3389/fimmu.2023.1263537
16. Lv B, Wang Y, Ma D, Cheng W, Liu J, Yong T, et al. Immunotherapy: reshape the tumor immune microenvironment. Front Immunol. (2022) 13:844142. doi: 10.3389/fimmu.2022.844142
17. Khosravi GR, Mostafavi S, Bastan S, Ebrahimi N, Gharibvand RS, Eskandari N. Immunologic tumor microenvironment modulators for turning cold tumors hot. Cancer Commun (Lond). (2024) 44:521–53. doi: 10.1002/cac2.12539
18. Fridman WH, Zitvogel L, Sautès-Fridman C, Kroemer G. The immune contexture in cancer prognosis and treatment. Nat Rev Clin Oncol. (2017) 14:717–34. doi: 10.1038/nrclinonc.2017.101
19. Locati M, Curtale G, Mantovani A. Diversity, mechanisms, and significance of macrophage plasticity. Annu Rev Pathol. (2020) 15:123–47. doi: 10.1146/annurev-pathmechdis-012418-012718
20. Sica A, Mantovani A. Macrophage plasticity and polarization: in vivo veritas. J Clin Invest. (2012) 122:787–95. doi: 10.1172/JCI59643
21. Mantovani A, Allavena P, Marchesi F, Garlanda C. Macrophages as tools and targets in cancer therapy. Nat Rev Drug Discovery. (2022) 21:799–820. doi: 10.1038/s41573-022-00520-5
22. Frega G, Wu Q, Le Naour J, Vacchelli E, Galluzzi L, Kroemer G, et al. Trial Watch: experimental TLR7/TLR8 agonists for oncological indications. Oncoimmunology. (2020) 9:1796002. doi: 10.1080/2162402X.2020.1796002
23. Biswas SK, Mantovani A. Macrophage plasticity and interaction with lymphocyte subsets: cancer as a paradigm. Nat Immunol. (2010) 11:889–96. doi: 10.1038/ni.1937
24. Beatty GL, Torigian DA, Chiorean EG, Saboury B, Brothers A, Alavi A, et al. A phase I study of an agonist CD40 monoclonal antibody (CP-870,893) in combination with gemcitabine in patients with advanced pancreatic ductal adenocarcinoma. Clin Cancer Res. (2013) 19:6286–95. doi: 10.1158/1078-0432.CCR-13-1320
25. Biglari N, Khalaj-Kondori M, Ghasemi T. Potential of hsa-miR200a-3p and hsa-miR502-3p as blood-based biomarker for Alzheimer’s disease. Mol Biol Rep. (2022) 49:11925–32. doi: 10.1007/s11033-022-07987-w
26. Zhang B, Vogelzang A, Miyajima M, Sugiura Y, Wu Y, Chamoto K, et al. B cell-derived GABA elicits IL-10+ macrophages to limit anti-tumour immunity. Nature. (2021) 599:471–6. doi: 10.1038/s41586-021-04082-1
27. Etzerodt A, Tsalkitzi K, Maniecki M, Damsky W, Delfini M, Baudoin E, et al. Specific targeting of CD163+ TAMs mobilizes inflammatory monocytes and promotes T cell-mediated tumor regression. J Exp Med. (2019) 216:2394–411. doi: 10.1084/jem.20182124
28. Barkal AA, Brewer RE, Markovic M, Kowarsky M, Barkal SA, Zaro BW, et al. CD24 signalling through macrophage Siglec-10 is a target for cancer immunotherapy. Nature. (2019) 572:392–6. doi: 10.1038/s41586-019-1456-0
29. Gordon SR, Maute RL, Dulken BW, Hutter G, George BM, McCracken MN, et al. PD-1 expression by tumour-associated macrophages inhibits phagocytosis and tumour immunity. Nature. (2017) 545:495–9. doi: 10.1038/nature22396
30. Speiser DE, Chijioke O, Schaeuble K, Münz C. CD4+ T cells in cancer. Nat Cancer. (2023) 4:317–29. doi: 10.1038/s43018-023-00521-2
31. Ruterbusch M, Pruner KB, Shehata L, Pepper M. In vivo CD4+ T cell differentiation and function: Revisiting the Th1/Th2 paradigm. Annu Rev Immunol. (2020) 38:705–25. doi: 10.1146/annurev-immunol-103019-085803
32. Kiner E, Willie E, Vijaykumar B, Chowdhary K, Schmutz H, Chandler J, et al. Gut CD4+ T cell phenotypes are a continuum molded by microbes, not by TH archetypes. Nat Immunol. (2021) 22:216–28. doi: 10.1038/s41590-020-00836-7
33. Corthay A, Skovseth DK, Lundin KU, Røsjø E, Omholt H, Hofgaard PO, et al. Primary antitumor immune response mediated by CD4+ T cells. Immunity. (2005) 22:371–83. doi: 10.1016/j.immuni.2005.02.003
34. Mumberg D, Monach PA, Wanderling S, Philip M, Toledano AY, Schreiber RD, et al. CD4(+) T cells eliminate MHC class II-negative cancer cells in vivo by indirect effects of IFN-gamma. Proc Natl Acad Sci USA. (1999) 96:8633–8. doi: 10.1073/pnas.96.15.8633
35. Huse M, Lillemeier BF, Kuhns MS, Chen DS, Davis MM. T cells use two directionally distinct pathways for cytokine secretion. Nat Immunol. (2006) 7:247–55. doi: 10.1038/ni1304
36. Müller-Hermelink N, Braumüller H, Pichler B, Wieder T, Mailhammer R, Schaak K, et al. TNFR1 signaling and IFN-gamma signaling determine whether T cells induce tumor dormancy or promote multistage carcinogenesis. Cancer Cell. (2008) 13:507–18. doi: 10.1016/j.ccr.2008.04.001
37. Ferris ST, Durai V, Wu R, Theisen DJ, Ward JP, Bern MD, et al. cDC1 prime and are licensed by CD4+ T cells to induce anti-tumour immunity. Nature. (2020) 584:624–9. doi: 10.1038/s41586-020-2611-3
38. Eickhoff S, Brewitz A, Gerner MY, Klauschen F, Komander K, Hemmi H, et al. Robust anti-viral immunity requires multiple distinct T cell-dendritic cell interactions. Cell. (2015) 162:1322–37. doi: 10.1016/j.cell.2015.08.004
39. Quezada SA, Simpson TR, Peggs KS, Merghoub T, Vider J, Fan X, et al. Tumor-reactive CD4(+) T cells develop cytotoxic activity and eradicate large established melanoma after transfer into lymphopenic hosts. J Exp Med. (2010) 207:637–50. doi: 10.1084/jem.20091918
40. Eschweiler S, Clarke J, Ramírez-Suástegui C, Panwar B, Madrigal A, Chee SJ, et al. Intratumoral follicular regulatory T cells curtail anti-PD-1 treatment efficacy. Nat Immunol. (2021) 22:1052–63. doi: 10.1038/s41590-021-00958-6
41. Zhang Y, Lazarus J, Steele NG, Yan W, Lee HJ, Nwosu ZC, et al. Regulatory T-cell depletion alters the tumor microenvironment and accelerates pancreatic carcinogenesis. Cancer Discovery. (2020) 10:422–39. doi: 10.1158/2159-8290.CD-19-0958
42. Laheurte C, Dosset M, Vernerey D, Boullerot L, Gaugler B, Gravelin E, et al. Distinct prognostic value of circulating anti-telomerase CD4+ Th1 immunity and exhausted PD-1+/TIM-3+ T cells in lung cancer. Br J Cancer. (2019) 121:405–16. doi: 10.1038/s41416-019-0531-5
43. de Kleijn S, Langereis JD, Leentjens J, Kox M, Netea MG, Koenderman L, et al. IFN-γ-stimulated neutrophils suppress lymphocyte proliferation through expression of PD-L1. PloS One. (2013) 8:e72249. doi: 10.1371/journal.pone.0072249
44. Xu YP, Lv L, Liu Y, Smith MD, Li WC, Tan XM, et al. Tumor suppressor TET2 promotes cancer immunity and immunotherapy efficacy. J Clin Invest. (2019) 129:4316–31. doi: 10.1172/JCI129317
45. Gao Y, Yang J, Cai Y, Fu S, Zhang N, Fu X, et al. IFN-γ-mediated inhibition of lung cancer correlates with PD-L1 expression and is regulated by PI3K-AKT signaling. Int J Cancer. (2018) 143:931–43. doi: 10.1002/ijc.31357
46. Boes M, Meyer-Wentrup F. TLR3 triggering regulates PD-L1 (CD274) expression in human neuroblastoma cells. Cancer Lett. (2015) 361:49–56. doi: 10.1016/j.canlet.2015.02.027
47. Sun C, Mezzadra R, Schumacher TN. Regulation and function of the PD-L1 checkpoint. Immunity. (2018) 48:434–52. doi: 10.1016/j.immuni.2018.03.014
48. Oh-hora M. Calcium signaling in the development and function of T-lineage cells. Immunol Rev. (2009) 231:210–24. doi: 10.1111/j.1600-065X.2009.00819.x
49. Jiang X, Zhou J, Giobbie-Hurder A, Wargo J, Hodi FS. The activation of MAPK in melanoma cells resistant to BRAF inhibition promotes PD-L1 expression that is reversible by MEK and PI3K inhibition. Clin Cancer Res. (2013) 19:598–609. doi: 10.1158/1078-0432.CCR-12-2731
50. Lei H, Li H, Xie H, Du C, Xia Y, Tang W. Role of MiR-215 in Hirschsprung’s disease pathogenesis by targeting SIGLEC-8. Cell Physiol Biochem. (2016) 40:1646–55. doi: 10.1159/000453214
51. Mechtler P, Singhal R, KiChina JV, Bard JE, Buck MJ, Kandel ES. MicroRNA analysis suggests an additional level of feedback regulation in the NF-κB signaling cascade. Oncotarget. (2015) 6:17097–106. doi: 10.18632/oncotarget.4005
52. Kong X, Duan Y, Sang Y, Li Y, Zhang H, Liang Y, et al. LncRNA-CDC6 promotes breast cancer progression and functions as a ceRNA to target CDC6 by sponging microRNA-215. J Cell Physiol. (2019) 234:9105–17. doi: 10.1002/jcp.27587
53. Yang Y, Zhang J, Chen X, Xu X, Cao G, Li H, et al. LncRNA FTX sponges miR-215 and inhibits phosphorylation of vimentin for promoting colorectal cancer progression. Gene Ther. (2018) 25:321–30. doi: 10.1038/s41434-018-0026-7
54. Chen DL, Lu YX, Zhang JX, Wei XL, Wang F, Zeng ZL, et al. Long non-coding RNA UICLM promotes colorectal cancer liver metastasis by acting as a ceRNA for microRNA-215 to regulate ZEB2 expression. Theranostics. (2017) 7:4836–49. doi: 10.7150/thno.20942
55. Vychytilova-Faltejskova P, Slaby O. MicroRNA-215: From biology to theranostic applications. Mol Aspects Med. (2019) 70:72–89. doi: 10.1016/j.mam.2019.03.002
56. Chen Z, Liu X, Hu Z, Wang Y, Liu M, Liu X, et al. Identification and characterization of tumor suppressor and oncogenic miRNAs in gastric cancer. Oncol Lett. (2015) 10:329–36. doi: 10.3892/ol.2015.3179
57. Wang M, Yu Y, Liu F, Ren L, Zhang Q, Zou G. Single polydiacetylene microtube waveguide platform for discriminating microRNA-215 expression levels in clinical gastric cancerous, paracancerous, and normal tissues. Talanta. (2018) 188:27–34. doi: 10.1016/j.talanta.2018.05.049
58. Li N, Zhang QY, Zou JL, Li ZW, Tian TT, Dong B, et al. miR-215 promotes Malignant progression of gastric cancer by targeting RUNX1. Oncotarget. (2016) 7:4817–28. doi: 10.18632/oncotarget.6736
Keywords: CXCR2P1, prognosis, immune microenvironment, PD-1 inhibitor, microRNAs
Citation: Wu X, Hou S, Ye Y and Gao Z (2025) CXCR2P1 enhances the response of gastric cancer to PD-1 inhibitors through increasing the immune infiltration of tumors. Front. Immunol. 16:1545605. doi: 10.3389/fimmu.2025.1545605
Received: 15 December 2024; Accepted: 03 March 2025;
Published: 19 March 2025.
Edited by:
Qing Xi, First Affiliated Hospital of Guangzhou Medical University, ChinaReviewed by:
Esther Layseca Espinosa, Autonomous University of San Luis Potosi, MexicoCopyright © 2025 Wu, Hou, Ye and Gao. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Yingjiang Ye, eWV5aW5namlhbmdAcGt1cGguZWR1LmNu; Zhidong Gao, Z2FvemRAYmptdS5lZHUuY24=
†Present address: Zhidong Gao, Peking University People’s Hospital, Beijing, China
‡These authors have contributed equally to this work and share first authorship
§These authors have contributed equally to this work
‖‖ORCID: Sen Hou, orcid.org/0000-0001-9907-1914
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.