
95% of researchers rate our articles as excellent or good
Learn more about the work of our research integrity team to safeguard the quality of each article we publish.
Find out more
ORIGINAL RESEARCH article
Front. Immunol. , 02 April 2025
Sec. Viral Immunology
Volume 16 - 2025 | https://doi.org/10.3389/fimmu.2025.1495672
 Zhi-sheng Wang1†
Zhi-sheng Wang1† Wen-jing Yu1†
Wen-jing Yu1† Xin-yi Ding1
Xin-yi Ding1 Zhan-peng Lu1
Zhan-peng Lu1 Sheng Qin1,2
Sheng Qin1,2 Xia Sun1,2
Xia Sun1,2 Xue-yang Wang1,2*
Xue-yang Wang1,2* Mu-wang Li1,2*
Mu-wang Li1,2*Autographa californica nucleopolyhedrovirus (AcMNPV) is a DNA virus with multiple host domains, and elucidating the mechanisms of its interactions with silkworms is crucial for its widespread use. Identifying key antiviral genes and analyzing their functions is an urgent task currently. Therefore, the identification and study of host genes associated with AcMNPV invasion is of great significance in solving the issue. Engulfment and cell motility (Elmo) is an identified viral infection-associated gene primarily involved in the regulation of cell motility and essential for phagocytosis and immune responses. However, its function in the silkworm response to viruses is still unclear. In this study, the sequence of BmElmo was analyzed first. It has a CED-12 functional domain that has been highly conserved among different species. Its expression peaks during the silkworm pupal stage, followed by the moth stage. Among various tissues, BmElmo expression is highest in the gonads, followed by the silk glands. BmElmo exhibits differential expression between resistant and susceptible strains. AcMNPV replication increased significantly after BmElmo knockdown in BmN cells, and decreased significantly after BmElmo overexpression. Furthermore, the expression of Janus kinase (JNK) pathway–related genes downstream of BmElmo showed altered expression that correlated positively with the expression of BmElmo. Hence, BmElmo may inhibit AcMNPV replication in the silkworm by activating the JNK pathway. The results of this study bridge the gap in understanding the role of Elmo genes in insect immunity and provides a theoretical reference for studying the interaction between insects and baculoviruses.
The silkworm Bombyx mori (Lepidoptera: Bombycidae) serves as a model insect within the Lepidoptera order and is widely utilized in biological research, which makes it a significant insect in the realms of biology and agriculture (1). Autographa californica nucleopolyhedrovirus (AcMNPV) is a baculovirus with a double-stranded DNA genome (2), which has a wide range of host domains and therefore a wide range of applications. The silkworm and AcMNPV are good models for studying the interactions between hosts and baculoviruses (3), but fewer studies have been conducted on the functional genes of host antiviral resistance, resulting in the mechanisms of interactions that remain unclear. Therefore, there is an urgent need to identify the key antiviral genes and to analyze their functions.
ELMO was first discovered in the nematode Caenorhabditis elegans as a novel phagocytosis-related gene (4). Its functional domain, CED-12, plays an important role during cell migration and phagocytosis in C. elegans, Drosophila, and mammals (5, 6). In mammalian cells, there are three Elmo isoforms (7): Elmo1, Elmo2, and Elmo3. Elmo is an evolutionarily conserved protein and has no obvious catalytic region in its protein structure (8, 9). Elmo1 can form a complex with the dedicator of cytokinesis (Dock) protein and act as a guanine nucleotide exchange factor (GEF) to activate Rac, thereby initiating the related signaling pathways downstream of Rac (10, 11), promoting cytoskeletal rearrangement and integrin-mediated phagocytosis and regulating cell migration (8, 12). In Drosophila, the CED-12 domain activates the JNK pathway to regulate ovarian nurse cell death and removal (13). In mice, Elmo1 regulates chemotaxis and the adhesion of neutrophils defects in vivo and in vitro by activating Rac, thereby affecting immunity (14, 15). However, no research has linked Elmo to viral infections, nor have we seen studies of its role in silkworms. In our previous transcriptome analysis, we found that BmElmo expression varies significantly among silkworm strains with different resistance to AcMNPV infection (16), yet the underlying mechanism remains unclear.
In this study, the function of BmElmo in AcMNPV infection was examined by knocking down and overexpressing this gene in BmN cells (a silkworm ovary cell line) and the potential downstream pathway. The findings offer valuable insights into the molecular mechanisms underlying the interaction between lepidopterans and baculoviruses.
The p50 and C108 silkworm strain was bred at the Key Laboratory of Sericulture within the School of Life Science at Jiangsu University of Science and Technology. p50 is a susceptive strain of AcMNPV with a half lethal concentration (LC50) of 4.5×104 pfu/mL, and C108 is a resistant strain of AcMNPV with a LC50 more than 6.2×108 pfu/mL. The larvae were provided with fresh mulberry leaves and kept at 26 ± 1°C, 75% ± 5% relative humidity, and a 12-h photoperiod. The temperature was reduced to 24 ± 1°C during the final two instars, while the other conditions remained unchanged.
Budded AcMNPV with enhanced green fluorescent protein (BV-eGFP) is maintained in our laboratory. The titer of BV-eGFP was calculated as described by Li et al. (17).
On the 1st day of 5th instar, each silkworm larva was injected with 2.0 μL of culture medium containing BV-eGFP at a concentration of 1.0 × 108 plaque-forming units (pfu)/μL. The control group was injected with 2.0 μL of culture medium. Various tissues, including the midgut, fat body, hemolymph, and the Malpighian tubule, were collected 36 h after BV-eGFP injection. The samples were rapidly frozen in liquid nitrogen, immediately ground into a powder, and stored at -80°C until use. BmN cells were transfected with 2 ng of DNA/RNA or treated with 5.0 μL of culture medium containing BV-eGFP (1.0× 108 pfu/μL) per well, and then collected at 24, 48, and 72 h post-transfection or BV-eGFP treatment. After centrifugation at low speed for 5 min and discarding the culture medium, phosphate-buffered saline (PBS) was added to wash the cells. Then, the cells were centrifuged at low speed for 5 min. The supernatant was discarded, and the cells were washed again. After the final wash, the cell pellet was frozen in liquid nitrogen, immediately ground into a powder, and then stored at -80°C until use.
The BmElmo nucleic acid sequence (GenBank ID: LOC101744642) and its homologous sequences in other species were retrieved from the National Center for Biotechnology Information (NCBI) website (http://www.ncbi.nlm.nih.gov/). The functional domain of BmElmo was predicted using the online SMART tool (http://smart.embl-heidelberg.de/). The homologous nucleic acid sequence of BmElmo was analyzed using the DNAMAN 8.0 software, and a neighbor-joining tree was constructed using the MEGA-X software.
BmN cells and silkworm tissues were lysed with the TransZol reagent (Transgen, Beijing, China). Then, RNA was isolated by chloroform extraction, purified by the isopropanol precipitation, and dissolved in diethyl pyrocarbonate (DEPC)-treated water. The purity of the isolated RNA was assessed based on the optical density (OD) 260/280 ratio. The RNA concentration was measured using a NanoDrop 2000 spectrophotometer (Thermo, Waltham, USA), and its integrity was verified by 1% agarose gel electrophoresis. A total of 1.0 µg of RNA was subjected to reverse transcription with the PrimeScript™ RT reagent kit (Takara, Kusatsu, Japan) according to the manufacturer’s instructions.
The relative expression of the genes of interest was detected using RT-qPCR. The specific primers were designed using the NCBI website and are presented in Table 1. Each reaction comprised 1 µL of cDNA, 1 µL of primer mix, 3 µL of double-distilled water (ddH2O), and 5 µL of 2× NovoStart® SYBR qPCR SuperMix Plus. The standard cycling protocol was 95°C for 5 min, followed by 40 cycles of denaturation at 95°C for 10 s and annealing at 60°C for 30 s, and finally a melting curve was generated at 95°C for 15 s, 60°C for 1 min, and 95°C for 15 s. To reduce experimental error, all samples underwent three repeats. The expression level of each gene was calculated using the 2−ΔΔCT method. Bombyx mori glyceraldehyde-3-phosphate dehydrogenase (BmGAPDH) was used as internal references to normalize the gene expression.
Using p50 testis cDNA as a template, the open reading frame (ORF) fragment of BmElmo was amplified using BmElmo primers (Table 1). The purified ORF was ligated into the pMD-19T vector and subsequently sent to Sangon (Shanghai, China) for sequencing. The pMD-19T-BmElmo and pIZT/V5-His-mCherry vectors were digested with KpnI and EcoRI (Takara), and the correctly sequenced ORF was ligated into the overexpression vector using T4 DNA ligase (Takara). After sequencing validation, the pIZT/V5-His-mCherry-BmElmo vector was stored at -80°C until use.
siRNA targeting the functional domain of BmElmo was designed using the BLOCK-iT™ RNAi designer website (https://rnaidesigner.thermofisher.com/), which was used to knock down the expression of BmElmo in BmN cells. siRFP served as a negative control. The primer sequences are shown in Table 2. siRNAs were synthesized using the In Vitro Transcription T7 Kit (Takara) according to the manufacturer’s instructions. Following purification, the high-quality siRNA was stored at -80°C for future use.
BmN cells were cultured at 27°C. Each 50 mL of cell culture medium comprised 44.5 mL of TC-100 (AppliChem, Gatersleben, Germany), 5 mL of fetal bovine serum (FBS; Thermo Fisher Scientific, New York, NY, USA), and 0.5 mL of dual antibiotics (penicillin and streptomycin). The culture medium of Sf9 cells was Sf-900TM (Thermo Fisher Scientific, New York, NY, USA). BmN cells were transfected with siRNA and the overexpression vector using a transfection reagent (NEOFECT, Beijing, China) according to the manufacturer’s instructions. Each 60 mm dish was transfected with 4.0 µg of plasmid or siRNA. The transfection solution for 2 mL of culture medium comprised 2 μg of DNAor RNA, 2 μL of transfection reagent (NEOFECT), and 100 μL of TC-100 solution; it was prepared 30 min before transfection. The transfection efficiency was measured at different time points. The expression level of BmElmo was detected at 24, 48, and 72 h after transfection.
All statistical analyses were performed using GraphPad Prism 8.0.1 (GraphPad Software, San Diego, CA, USA). The data are presented as the mean ± standard error of the mean of three biological replicates, and each biological replicate was derived from samples of different individuals. Student’s t-test was used to compare two groups of normally distributed data. The Kruskal–Wallis test was used to analyze the data that did not meet the normality requirement. Statistically significant differences between samples were analyzed using one-way ANOVA test. Significance is indicated by an asterisk: * P < 0.05, ** P < 0.01, and *** P < 0.001; ns means not significant.
BmElmo (GenBank ID XM_004923980.4) has a full length of 3,725 base pairs (bp) and a coding sequencing (CDS) length of 936 bp, encoding a protein of 313 amino acids. Based on functional domain analysis of BmElmo using the SMART website, there is a CED-12 domain spanning amino acids 125 to 286. Additionally, protein sequence homology alignment analysis showed that BmElmo has a sequence conservation of 86.43% (Figure 1).
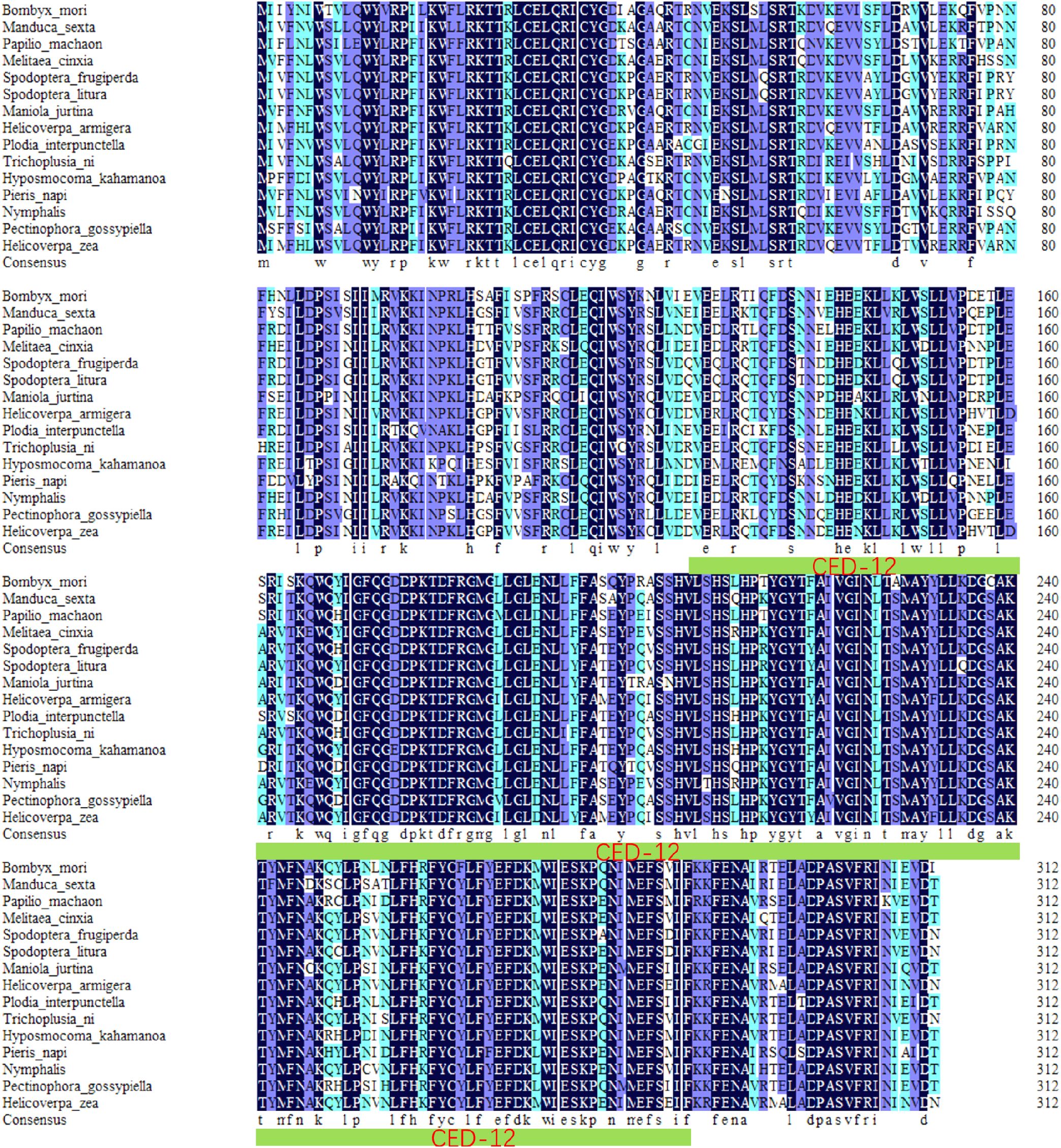

Figure 1. Analysis of BmElmo sequence conservation. A total of 15 species were compared. The green color with a thick underline indicates the functional domains of BmElmo. Conserved amino acid sequences are shown with a black background.
An evolutionary tree has been constructed by neighbor joining method in the MEGA-X software with the identified homologous Elmo protein sequences from various species, including B. mori, Drosophila melanogaster, Manduca sexta, and Melitaea cinxia (Figure 2).

Figure 2. Phylogenetic tree of ELMO among 15 species. The bootstrap values from 1000 replicates are shown on each branch. The branch length represents the evolutionary distance.
The relative expression of BmElmo across various developmental stages of the p50 strain and in different tissues of 5th instar larvae have been examined by RT qPCR. BmElmo expression was highest during the pupal stage, and there was notable expression for the 1st instar and adult stages. The egg stage presented the lowest expression (Figure 3A). Among the various tissues, BmElmo expression was highest in the testis, followed by the ovary and silk gland, while the epidermis presented the lowest expression (Figure 3B).
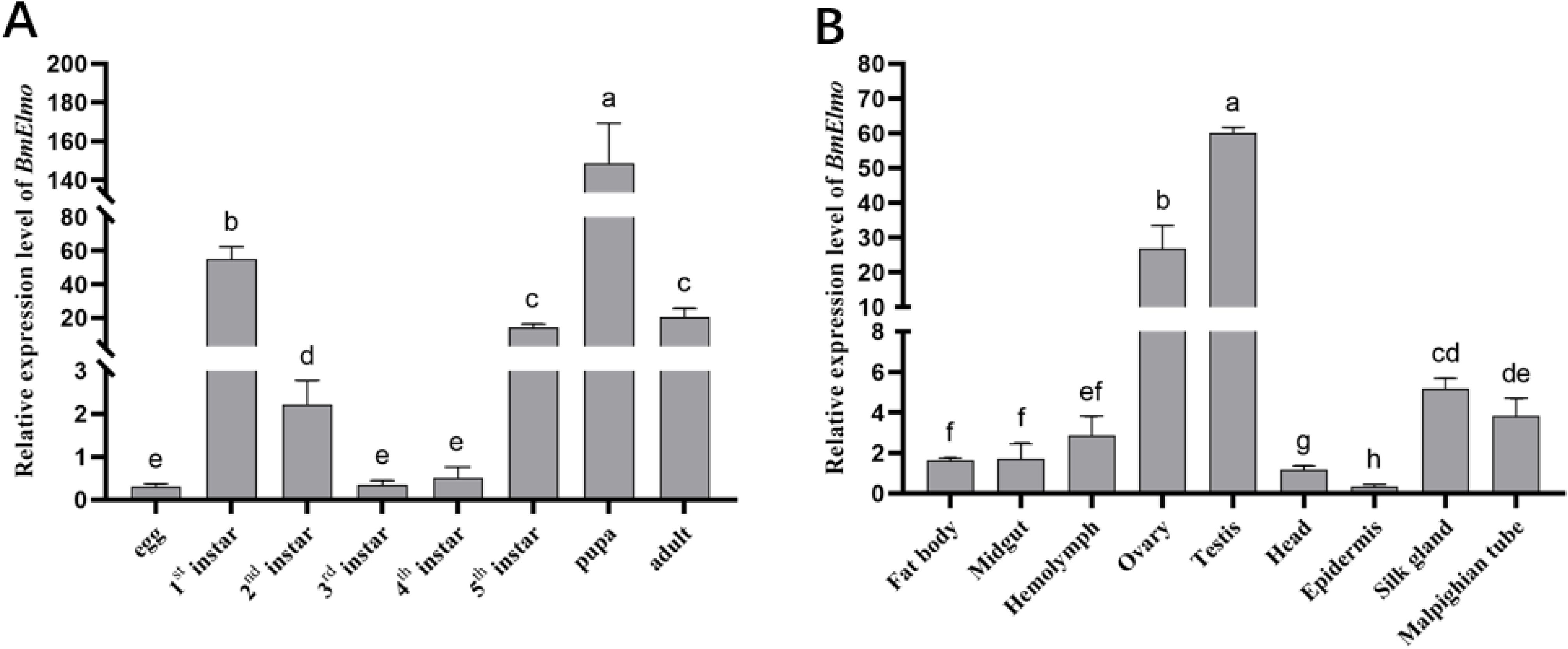
Figure 3. Analysis of BmElmo relative expression across various tissues and developmental phases based on RT-qPCR. (A) BmElmo expression at different developmental stages. (B) BmElmo expression in various tissues. BmGAPDH was used to normalize the data that were presented as the mean ± standard error of three replicates. Statistically significant differences between sample were analyzed using one-way ANOVA test. P<0,05 is indicated using different letters a, b, c, d, etc.
The relationship between BmElmo and AcMNPV has been analyzed based on BmElmo expression in four immune-related tissues (hemolymph, midgut, Malpighian tubules, and fat body) from a susceptible strain (p50) and a resistant strain (C108) 36 h after AcMNPV infection. The silkworms in the control group were injected with same volume of SF-900tm insect culture medium. BmElmo expression in the midgut, hemolymph, and Malpighian tubule of the C108 strain was significantly elevated compared with the control group, while in the p50 strain, BmElmo expression in the Malpighian tubule, hemolymph, and fat body was notably decreased relative to the control group (Figure 4).
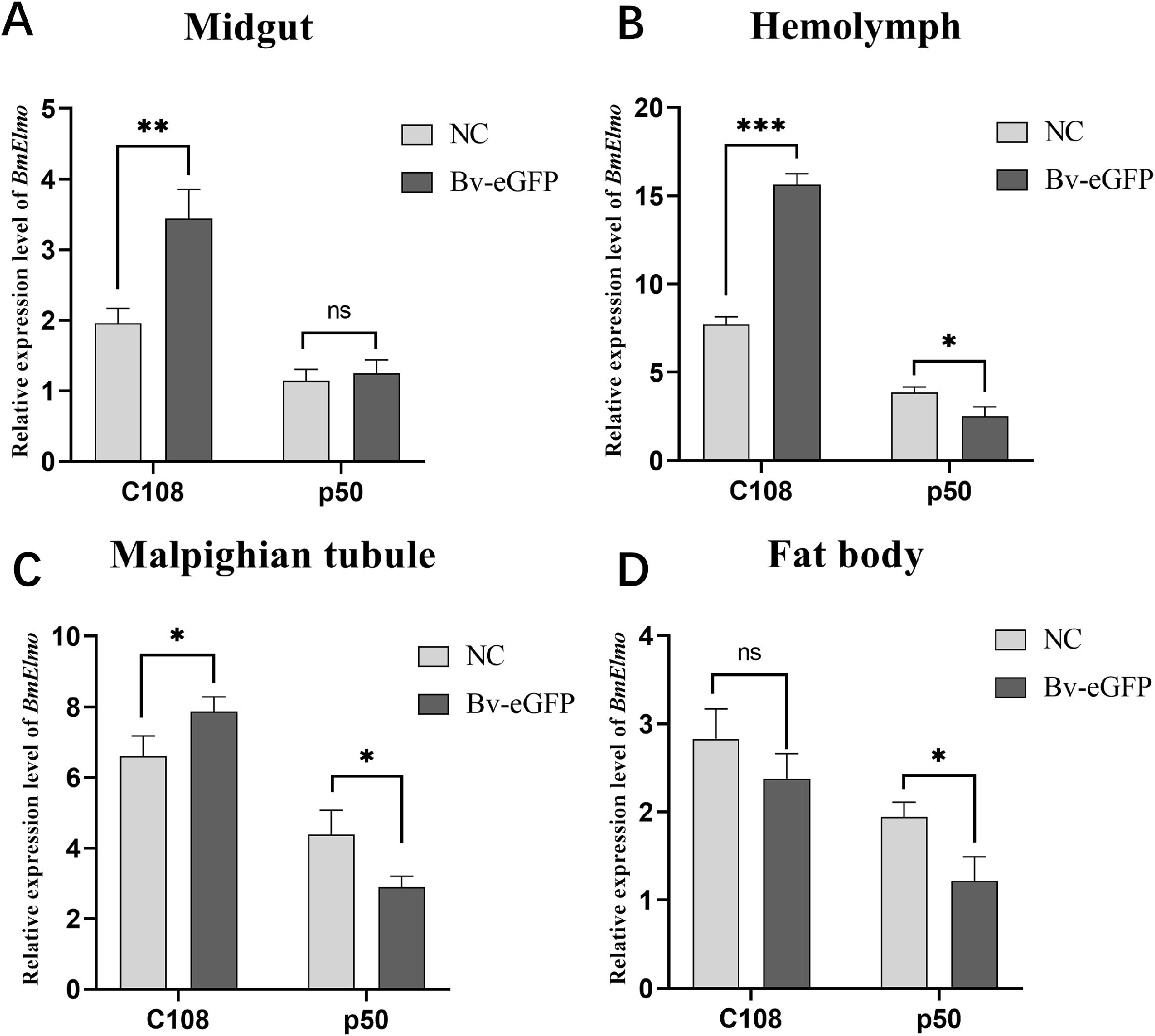
Figure 4. Analysis of BmElmo relative expression in various tissues of the susceptible (p50) and resistant (C108) strains after AcMNPV infection based on RT-qPCR. (A) Midgut, (B) Hemolymph, (C) Malpighian tubule, (D) Fat body. BmElmo expression was detected 36 h after AcMNPV infection. BmGAPDH was used to normalize the data that were presented as the mean ± standard error of three replicates. Asterisks represent significant differences. * P < 0.05, ** P < 0.01, and *** P < 0.001; ns indicates not significant.
The expression level of BmElmo has been knocked down in BmN cells by transfecting siRNA which target to its functional domain CED-12 (siBmElmo). Subsequently, these cells were infected with AcMNPV 24 h after transfection. BmElmo expression was significantly decreased in the siBmElmo group 24, 48, and 72 h after transfection (Figure 5A). AcMNPV infection was observed by fluorescence microscopy at 24, 48, and 72 h post-infection. The virus fluorescence signal rose significantly at 48 and 72 h after infection (Figure 5C). To validate the above results, the relative expression of lef3, a viral single-strand DNA-binding protein, was detected at 24, 48, and 72 h after AcMNPV infection of BmN cells. lef3 relative expression was markedly higher in the siBmElmo group compared with the control group (Figure 5B).
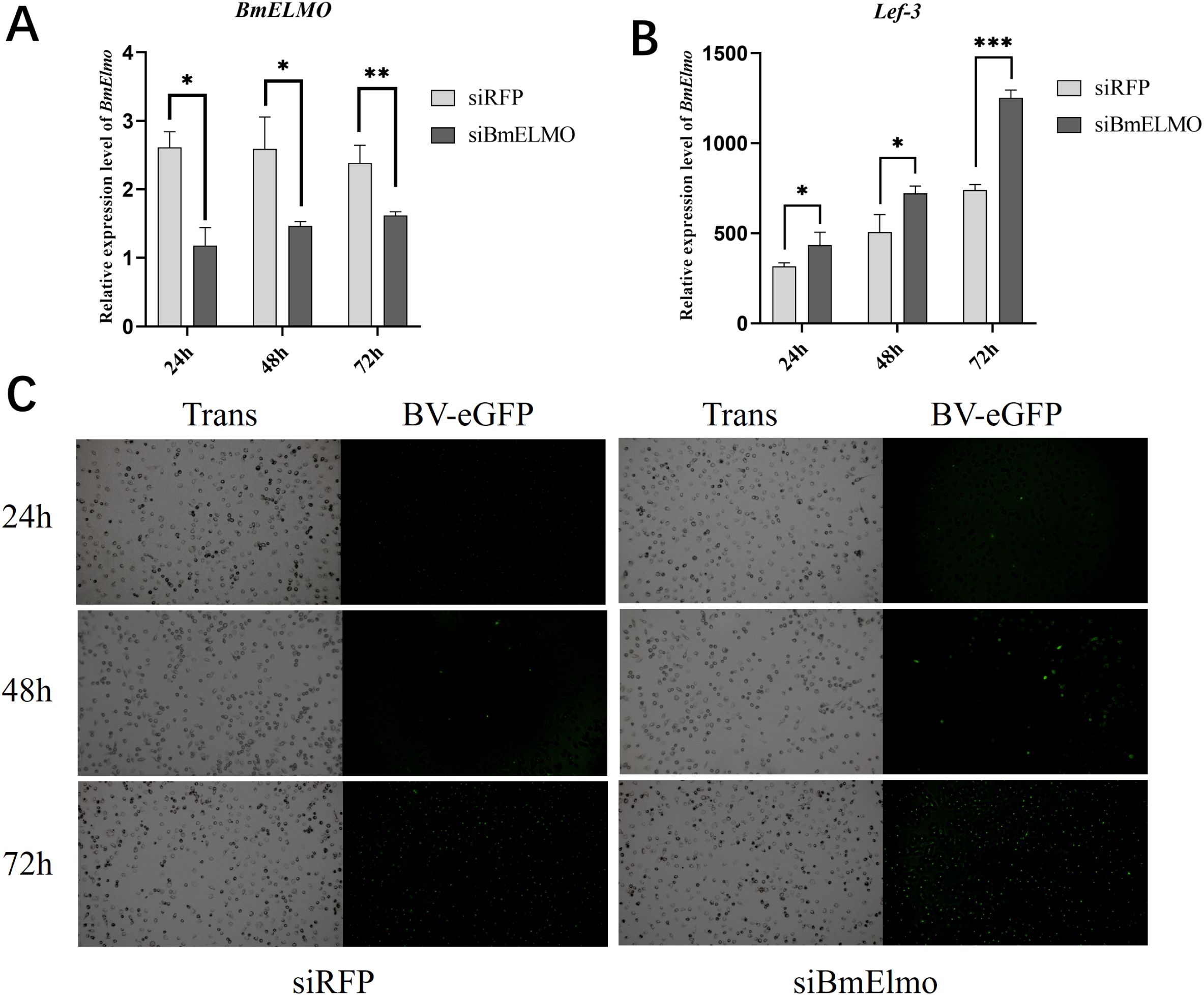
Figure 5. Analysis of AcMNPV proliferation in BmN cells at different times after BmElmo knockdown. (A) BmElmo expression at various times following siRNA transfection. (B) Expression of the viral gene lef3 24, 48, and 72 h after RNAi. (C) eGFP fluorescence signal at 24, 48, and 72 h after RNAi (the scale bar represents 100 μm). BmGAPDH was used to normalize the data that were presented as the mean ± standard error of three replicates. Asterisks represent significant differences. *P < 0.05, **P < 0.01, and ***P < 0.001.
The recombinant plasmid pIZT/V5-His-mCherry-BmElmo was constructed to further investigate BmElmo’s role in the AcMNPV response (Figure 6A) and transfected BmN cells with it to overexpress BmElmo (Figure 6B). BmElmo expression was detected by RT-qPCR following transfection with the recombinant plasmid. BmElmo expression increased significantly at different times after transfection (Figure 6C), indicating BmElmo was overexpressed successfully in BmN cells. Moreover, the green fluorescence signal of AcMNPV was lower 48 and 72 h after BmElmo overexpression compared with the control group (Figure 6E). Additionally, lef3 expression was significantly lower after BmElmo expression compared with the control group (Figure 6D).
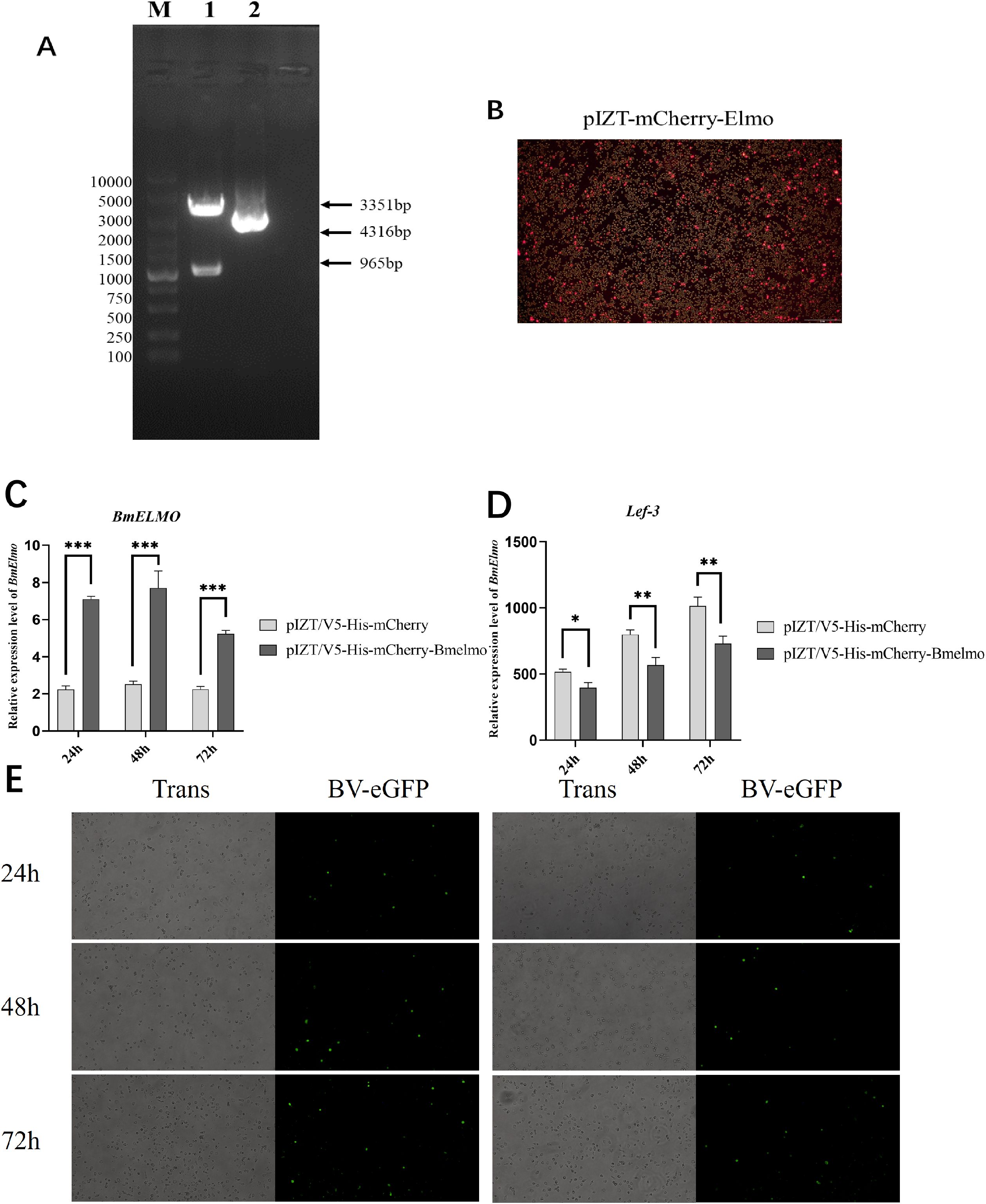
Figure 6. Analysis of AcMNPV proliferation in BmN cells at different times after BmElmo overexpression. (A) Verification of the pIZT/V5-His-mCherry-BmElmo vector by double restriction enzyme digestion. M, molecular weight of DNA. (B) Detection of the pIZT/V5-His-mCherry-BmElmo fluorescence signal in BmN cells (the scale bar represents 100 μm). (C) Expression of BmElmo 24, 48, and 72 h after BmElmo overexpression. (D) Expression of the viral gene lef3 at different times after BmElmo overexpression. (E) eGFP fluorescence signal 24, 48, and 72 h after BmElmo overexpression (the scale bar represents 100 μm). BmGAPDH was used to normalize the data that were presented as the mean ± standard error of three replicates. Asterisks represent significant differences. *P < 0.05, **P < 0.01, and ***P < 0.001.
The pathway by BmElmo in BmN cells mediates the response to AcMNPV infection was investigated. Specifically, the expression of genes that encode JNK pathway members and that participate in phagocytosis was examined 48 h after knockdown and overexpression of BmElmo. The JNK pathway–related genes BmJun, BmJnks, and BmFosS showed significantly reduced expression after BmElmo knockdown (Figures 7A–C) and significantly increased expression after BmElmo overexpression (Figures 7D–F). However, expression of the phagocytosis-related genes BmActinA1 and BmTet did not change significantly after BmElmo knockdown or overexpression of (Supplementary Figure S1).
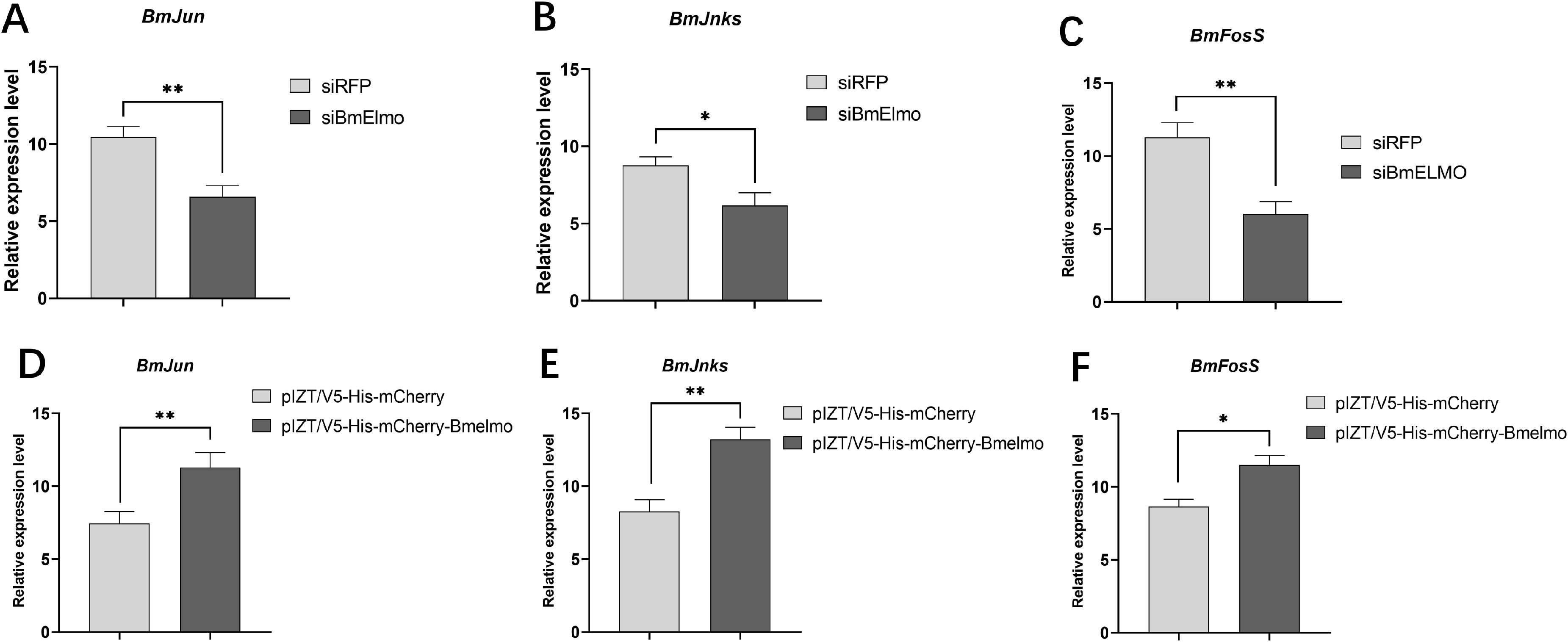
Figure 7. The relative expression of genes related to the JNK pathway in BmN cells 48 h after BmElmo knockdown or overexpression based on RT-qPCR. The relative expression levels of (A) BmJun, (B) BmJnks, and (C) BmFosS in BmN cells 48 h after BmElmo knockdown. The relative expression levels of (D) BmJun, (E) BmJnks, and (F) BmFosS in BmN cells 48 h after BmElmo overexpression. BmGAPDH was used to normalize the data that were presented as the mean ± standard error of three replicates. Asterisks represent significant differences. * P < 0.05, ** P < 0.01.
AcMNPV can infect more than 30 species of lepidopteran insects (18). Given its wide host range, it is used as a biopesticide in agricultural pest control. However, the resistance of pests to AcMNPV reduces its biocontrol ability. Numerous silkworm species can be infected with AcMNPV through subcutaneous injection. The AcMNPV-sensitive p50 strain and the AcMNPV-resistant C108 strain have been identified by our group (19). After AcMNPV injection, transcriptome analysis revealed that the p50 and C108 strains had significantly different BmElmo expression (16, 20). Based on these findings, BmElmo seems to have an important role in response to AcMNPV and confirmed in this study.
The Elmo protein sequences from different species was compared and it was found that the BmElmo amino acid sequence has been highly conserved, especially its functional domain CED-12 (Figure 1). Moreover, CED-12 has been highly phylogenetically conserved among Lepidoptera (Figure 2). CED-12 is involved in phagocytosis and apoptosis (21, 22), which provides a valuable clue for detecting the mechanism of BmElmo in response to AcMNPV infection. Silkworm larvae at 5th instar is the fastest period of weight gain that body weight increases more than 15-time compared with 1st instar (23). The very high BmElmo expression in 1st and 5th instars indicates that it plays a key role in silkworm development (Figure 3A). The tissue expression profiles showed that it is highly expressed in ovaries and testis (Figure 3B), indicating that it is involved in gonad development, which also corresponds to its high expression during the pupal stage (Figure 3A). Of note, the gonads of silkworms are in a primitive stage during the larval period and undergo several maturation steps during the pupal stage. Additionally, BmElmo expression was markedly higher compared with the control group in most of the analyzed tissues (Figure 4), indicating that BmElmo is an important gene in silkworm immunity.
The role of BmElmo in the response of BmN cells to AcMNPV infection was further investigated by knocking down and overexpressing this gene. Compared with the control, virus replication increased significantly following BmElmo knockdown (Figure 5), but decreased significantly after BmElmo overexpression (Figure 6). These findings demonstrate that BmElmo plays a critical inhibitory role in AcMNPV infection of BmN cells.
The Elmo protein participates in cellular processes such as cell migration, immunity, and phagocytosis by regulating the downstream RhoA/Rac1 (14), JNK (24) and phagocytosis pathways (6). However, prior to the present study, there had been no reports on which pathways Elmo regulates in the silkworm. In insects, the JNK pathway is closely related to infection of hosts by viruses and other microorganisms (25, 26). In silkworms, reactive oxygen species–mediated JNK phosphorylation is involved in the regulation of Bombyx mori nucleopolyhedrovirus (BmNPV) proliferation (27). Phagocytosis is a conserved evolutionary process wherein hemocytes identify, engulf, and ultimately dispose of apoptotic cells and invading microbial pathogens (28, 29). The relative expression of key genes involved in these processes was examined after BmElmo knockdown and overexpression. The relative expression of JNK pathway members, BmJun, BmJnks, and BmFosS, changed significantly after BmElmo knockdown or overexpression (Figure 7). However, the relative expression of the phagocytosis-related genes BmActinA1 and BmTet did not change when BmElmo was knocked down or overexpressed (Supplementary Figure S1). Hence, the BmElmo protein exerts its effects through the JNK pathway, but it is not clear whether the JNK pathway directly affects AcMNPV infection.
Our results indicate that the expression of BmElmo can enhance the ability of silkworms to baculoviruses infection, which might be related to the activation of JNK phosphorylation (27, 30), yet this point still needs to be further addressed.
The original contributions presented in the study are included in the article/Supplementary Material. Further inquiries can be directed to the corresponding author.
Z-SW: Conceptualization, Data curation, Investigation, Methodology, Writing – original draft, Writing – review & editing. W-JY: Project administration, Software, Supervision, Validation, Writing – review & editing. X-YD: Data curation, Formal Analysis, Methodology, Project administration, Software, Writing – review & editing. Z-PL: Data curation, Investigation, Methodology, Writing – original draft. SQ: Resources, Visualization, Writing – review & editing. XS: Project administration, Validation, Writing – review & editing. X-YW: Writing – original draft, Writing – review & editing. M-WL: Funding acquisition, Resources, Visualization, Writing – original draft, Writing – review & editing.
The author(s) declare that financial support was received for the research and/or publication of this article. This work was supported by the National Natural Science Foundation of China, (No. 32472977 and 31772523), and the Postgraduate Research & Practice Innovation Program of Jiangsu Province (KYCX23_3913).
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fimmu.2025.1495672/full#supplementary-material
1. Meng X, Zhu F, Chen K. Silkworm: A promising model organism in life science. J Insect Sci. (2017) 17:97. doi: 10.1093/jisesa/iex064
2. Zhang J, Zafar J, Kong J, Wang F, Shao X, Zhang R, et al. MicroRNA-mediated host immune genes manipulation benefits acMNPV proliferation in spodoptera frugiperda. J Agric Food Chem. (2023) 71:17175–87. doi: 10.1021/acs.jafc.3c05012
3. Pidre ML, Arrías PN, Amorós Morales LC, Romanowski V. The magic staff: A comprehensive overview of baculovirus-based technologies applied to human and animal health. Viruses. (2022) 15:80. doi: 10.3390/v15010080
4. Chung S, Gumienny TL, Hengartner MO, Driscoll M. A common set of engulfment genes mediates removal of both apoptotic and necrotic cell corpses in C. elegans. Nat Cell Biol. (2000) 2:931–7. doi: 10.1038/35046585
5. Ishimaru S, Ueda R, Hinohara Y, Ohtani M, Hanafusa H. PVR plays a critical role via JNK activation in thorax closure during Drosophila metamorphosis. EMBO J. (2004) 23:3984–94. doi: 10.1038/sj.emboj.7600417
6. Wang X, Wu YC, Fadok VA, Lee MC, Gengyo-Ando K, Cheng LC, et al. Cell corpse engulfment mediated by C. elegans phosphatidylserine receptor through CED-5 and CED-12. Sci (New York NY). (2003) 302:1563–6. doi: 10.1126/science.1087641
7. East MP, Bowzard JB, Dacks JB, Kahn RA. ELMO domains, evolutionary and functional characterization of a novel GTPase-activating protein (GAP) domain for Arf protein family GTPases. J Biol Chem. (2012) 287:39538–53. doi: 10.1074/jbc.M112.417477
8. Wu YC, Tsai MC, Cheng LC, Chou CJ, Weng NY. C. elegans CED-12 acts in the conserved crkII/DOCK180/Rac pathway to control cell migration and cell corpse engulfment. Dev Cell. (2001) 1:491–502. doi: 10.1016/S1534-5807(01)00056-9
9. E MP, B JB, D JB, K RA. ELMO domains, evolutionary and functional characterization of a novel. J Biolog Chem. 287(47):39538–53. doi: 10.1074/jbc.M112.417477
10. Ivanova AA, East MP, Slee LY, Kahn RA. Characterization of recombinant ELMOD (cell engulfment and motility domain) proteins as GTPase-activating proteins (GAPs) for ARF family GTPases. J Biol Chem. (2014) 289:11111–21. doi: 10.1074/jbc.M114.548529
11. Lu M, Ravichandran KS. Dock180-ELMO cooperation in Rac activation. Methods Enzymol. (2006) 406:388–402. doi: 10.1016/S0076-6879(06)06028-9
12. Zhou Z, Caron E, Hartwieg E, Hall A, Horvitz HR. The C. elegans PH domain protein CED-12 regulates cytoskeletal reorganization via a Rho/Rac GTPase signaling pathway. Dev Cell. (2001) 1:477–89. doi: 10.1016/S1534-5807(01)00058-2
13. Timmons AK, Mondragon AA, Meehan TL, McCall K. Control of non-apoptotic nurse cell death by engulfment genes in Drosophila. Fly. (2017) 11:104–11. doi: 10.1080/19336934.2016.1238993
14. Liang Y, Wang S, Zhang Y. Downregulation of Dock1 and Elmo1 suppresses the migration and invasion of triple? negative breast cancer epithelial cells through the RhoA/Rac1 pathway. Oncol Lett. (2018) 16:3481–8. doi: 10.3892/ol.2018.9077
15. Yu S, Geng X, Liu H, Zhang Y, Cao X, Li B, et al. ELMO1 deficiency reduces neutrophil chemotaxis in murine peritonitis. Int J Mol Sci. (2023) 24:8103. doi: 10.3390/ijms24098103
16. Ding X-y, Wang X-y, Kong Y-h, Zhao C-x, Qin S, Sun X, et al. Comparative transcriptome analysis of Bombyx mori (Lepidoptera) larval hemolymph in response to Autographa californica nucleopolyhedrovirus in differentially resistant strains. Processes. (2021) 9:1401. doi: 10.3390/pr9081401
17. Li T, Wang X, Qin S, Sun X, Wang S, Li M. The hemolymph melanization response is related to defence against the AcMNPV infection in Bombyx mori. Arch Insect Biochem Physiol. (2021) 108:e21764. doi: 10.1002/arch.v108.1
18. Vail PV, Hostetter DL, Hoffmann DF. Development of the multi-nucleocapsid nucleopolyhedroviruses (MNPVs) infectious to loopers (Lepidoptera: Noctuidae: Plusiinae) as microbial control agents. Integrated Pest Manage Rev. (1999) 4:231–57. doi: 10.1023/A:1009601212375
19. Xu J, Kusakabe T, Yamamoto K, Suetsugu Y, Mon H, Li Z, et al. A novel third chromosomal locus controls susceptibility to Autographa californica multiple nucleopolyhedrovirus in the silkworm, Bombyx mori. Appl Microbiol Biotechnol. (2014) 98:3049–58. doi: 10.1007/s00253-013-5437-1
20. Li T, Qin S, Sun X, Zhang K-x, Ding X-y, Wang X-y, et al. Transcriptome analysis reveals distinct innate immunity and ribosomal response at early stage of AcMNPV infection in haemocyte of silkworm resistant and susceptible strains. J Asia-Pac Entomol. (2022) 25:101938. doi: 10.1016/j.aspen.2022.101938
21. Park D, Tosello-Trampont AC, Elliott MR, Lu M, Haney LB, Ma Z, et al. BAI1 is an engulfment receptor for apoptotic cells upstream of the ELMO/Dock180/Rac module. Nature. (2007) 450:430–4. doi: 10.1038/nature06329
22. Gumienny TL, Brugnera E, Tosello-Trampont AC, Kinchen JM, Haney LB, Nishiwaki K, et al. CED-12/ELMO, a novel member of the CrkII/Dock180/Rac pathway, is required for phagocytosis and cell migration. Cell. (2001) 107:27–41. doi: 10.1016/S0092-8674(01)00520-7
23. Arandjelovic S, Ravichandran KS. Phagocytosis of apoptotic cells in homeostasis. Nat Immunol. (2015) 16:907–17. doi: 10.1038/ni.3253
24. Pastuhov SI, Fujiki K, Tsuge A, Asai K, Ishikawa S, Hirose K, et al. The core molecular machinery used for engulfment of apoptotic cells regulates the JNK pathway mediating axon regeneration in Caenorhabditis elegans. J Neurosci. (2016) 36:9710–21. doi: 10.1523/JNEUROSCI.0453-16.2016
25. Yu S, Luo F, Xu Y, Zhang Y, Jin LH. Drosophila innate immunity involves multiple signaling pathways and coordinated communication between different tissues. Front Immunol. (2022) 13:905370. doi: 10.3389/fimmu.2022.905370
26. Weston CR, Davis RJ. The JNK signal transduction pathway. Curr Opin Cell Biol. (2007) 19:142–9. doi: 10.1016/j.ceb.2007.02.001
27. Liu YX, Yang JY, Sun JL, Wang AC, Wang XY, Zhu LB, et al. Reactive oxygen species-mediated phosphorylation of JNK is involved in the regulation of BmFerHCH on Bombyx mori nucleopolyhedrovirus proliferation. Int J Biol Macromol. (2023) 235:123834. doi: 10.1016/j.ijbiomac.2023.123834
28. Ling E, Yu X-Q. Prophenoloxidase binds to the surface of hemocytes and is involved in hemocyte melanization in Manduca sexta. Insect Biochem Mol Biol. (2005) 35:1356–66. doi: 10.1016/j.ibmb.2005.08.007
29. Marmaras VJ, Lampropoulou M. Regulators and signalling in insect haemocyte immunity. Cell Signalling. (2009) 21:186–95. doi: 10.1016/j.cellsig.2008.08.014
Keywords: BmElmo, AcMNPV, Bombyx mori, baculovirus, JNK pathway
Citation: Wang Z-s, Yu W-j, Ding X-y, Lu Z-p, Qin S, Sun X, Wang X-y and Li M-w (2025) BmElmo is a factor for inhibiting Autographa Californica nucleopolyhedrovirus infection in silkworm, Bombyx mori. Front. Immunol. 16:1495672. doi: 10.3389/fimmu.2025.1495672
Received: 13 September 2024; Accepted: 17 March 2025;
Published: 02 April 2025.
Edited by:
Zhenhua Zheng, Chinese Academy of Sciences (CAS), ChinaReviewed by:
Zhaojun Wei, Hefei University of Technology, ChinaCopyright © 2025 Wang, Yu, Ding, Lu, Qin, Sun, Wang and Li. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Mu-wang Li, bXdsaUBqdXN0LmVkdS5jbg==; Xue-yang Wang, eHVleWFuZ3dhbmdAanVzdC5lZHUuY24=
†These authors have contributed equally to this work
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Research integrity at Frontiers

Learn more about the work of our research integrity team to safeguard the quality of each article we publish.